text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Evaluate for pretrain model
```
import sys
sys.path.append("..")
sys.path.append("../../")
import paddle
import paddlenlp
from paddlenlp.transformers import ErnieForMaskedLM, ErnieTokenizer
```
load model paramerters
```
MODEL_NAME = "ernie-1.0"
model = ErnieForMaskedLM.from_pretrained(MODEL_NAME, num_classes=2)
tokenizer = ErnieTokenizer.from_pretrained(MODEL_NAME)
```
## Prepare for Interpretations
```
from trustai.interpretation.token_level import IntGradInterpreter
import numpy as np
from assets.utils import convert_example, load_data
from paddlenlp.data import Stack, Tuple, Pad
def masked_one_hot(input_ids, mask_id):
res = []
for x in input_ids:
if x == mask_id:
res.append(1)
else:
res.append(0)
return res
# preprocess data functions
def preprocess_fn(data):
examples = []
data_trans = []
for key in data:
data_trans.append(data[key])
for text in data_trans:
input_ids, segment_ids = convert_example(text, tokenizer, max_seq_length=128, is_test=True)
masked = masked_one_hot(input_ids, tokenizer.convert_tokens_to_ids('[MASK]'))
examples.append((input_ids, segment_ids, masked))
batchify_fn = lambda samples, fn=Tuple(
Pad(axis=0, pad_val=tokenizer.pad_token_id), # input id
Pad(axis=0, pad_val=tokenizer.pad_token_id), # segment id
Pad(axis=0, pad_val=tokenizer.pad_token_id), # masked_one_hot
): fn(samples)
input_ids, segment_ids, masked = batchify_fn(examples)
return paddle.to_tensor(input_ids, stop_gradient=False), paddle.to_tensor(segment_ids, stop_gradient=False), paddle.to_tensor(masked, stop_gradient=False)
# download data
!wget --no-check-certificate -c https://trustai.bj.bcebos.com/data_samples/pretrain_predict -P ../assets/
!wget --no-check-certificate -c https://trustai.bj.bcebos.com/data_samples/pretrain_golden -P ../assets/
# predict data for predict
data = load_data("../assets/pretrain_predict")
print("data:\n", list(data.values())[:2])
# golden data for evluate
goldens = load_data("../assets/pretrain_golden")
print("goldens:\n", list(goldens.values())[:2])
```
## IG Interpreter
```
from trustai.interpretation.token_level.common import get_word_offset
from trustai.interpretation.token_level.data_processor import VisualizationTextRecord, visualize_text
contexts = []
batch_words = []
for example in data.values():
contexts.append("[CLS]" + " " + example['context'] + " " + "[SEP]")
batch_words.append(["[CLS]"] + example['sent_token'] + ["[SEP]"])
word_offset_maps = []
subword_offset_maps = []
for i in range(len(contexts)):
word_offset_maps.append(get_word_offset(contexts[i], batch_words[i]))
subword_offset_maps.append(tokenizer.get_offset_mapping(contexts[i]))
from trustai.interpretation.token_level.common import ig_predict_fn_on_paddlenlp_pretrain
ig = IntGradInterpreter(model, device="gpu", predict_fn=ig_predict_fn_on_paddlenlp_pretrain)
result = ig(preprocess_fn(data), steps=100)
align_res = ig.alignment(result, contexts, batch_words, word_offset_maps, subword_offset_maps, special_tokens=["[CLS]", '[SEP]', '[MASK]'])
def prepare_eval_data(data, results, paddle_model):
res = {}
for data_id, inter_res in zip(data, results):
eval_data = {}
eval_data['id'] = data_id
eval_data['pred_label'] = inter_res.pred_label
eval_data['pred_proba'] = inter_res.pred_proba
eval_data['rationale'] = [inter_res.rationale]
eval_data['non_rationale'] = [inter_res.non_rationale]
eval_data['rationale_tokens'] = [inter_res.rationale_tokens]
eval_data['non_rationale_tokens'] = [inter_res.non_rationale_tokens]
rationale_context = "".join(inter_res.rationale_tokens)
non_rationale_context = "".join(inter_res.non_rationale_tokens)
res[data_id] = eval_data
return res
predicts = prepare_eval_data(data, align_res, model)
print(list(predicts.values())[0])
```
evaluate for interpretation result
```
from trustai.evaluation import Evaluator
evaluator = Evaluator()
result = evaluator.cal_map(goldens, predicts)
print("map score:",result)
result = evaluator.cal_f1(goldens, predicts)
print("plausibility f1:", result)
result = evaluator.calc_iou_f1(goldens, predicts)
print("plausibility iou f1:",result)
```
## Attention Interpreter
```
from trustai.interpretation.token_level.common import attention_predict_fn_on_paddlenlp_pretrain
from trustai.interpretation.token_level import AttentionInterpreter
att = AttentionInterpreter(model, device="gpu", predict_fn=attention_predict_fn_on_paddlenlp_pretrain)
result = att(preprocess_fn(data))
align_res = att.alignment(result, contexts, batch_words, word_offset_maps, subword_offset_maps, special_tokens=["[CLS]", '[SEP]', '[MASK]'])
predicts = prepare_eval_data(data, align_res, model)
result = evaluator.cal_map(goldens, predicts)
print("map score:",result)
result = evaluator.cal_f1(goldens, predicts)
print("plausibility f1:", result)
result = evaluator.calc_iou_f1(goldens, predicts)
print("plausibility iou f1:", result)
```
| github_jupyter |
author: leezeeyee
date: 2021/4/15
link: [github](https://github.com/easilylazy/pattern-recognition)
initial: can't using solve_qp in qpsolevers: Q should be positive
因为把svm转化成QP问题时构造的矩阵不是正定的,但增加一个近似0的偏置数据即可解决
```
from numpy import array, dot
from qpsolvers import solve_qp
M = array([[1., 2., 0.], [-8., 3., 2.], [0., 1., 1.]])
P = dot(M.T, M) # this is a positive definite matrix
q = dot(array([3., 2., 3.]), M).reshape((3,))
G = array([[1., 2., 1.], [2., 0., 1.], [-1., 2., -1.]])
h = array([3., 2., -2.]).reshape((3,))
A = array([1., 1., 1.])
b = array([1.])
x = solve_qp(P, q, G, h, A, b)
print("QP solution: x = {}".format(x))
import numpy as np
import matplotlib.pyplot as plt
import qpsolvers
```
## QP solver

$$
\begin{equation}
\begin{array}{ll}
\underset{x}{\operatorname{minimize}} & \frac{1}{2} x^{T} P x+q^{T} x \\
\text { subject to } & G x \leq h \\
& A x=b \\
& l b \leq x \leq u b
\end{array}
\end{equation}
$$
```
def addSingleBias(X):
N=X.shape[0]
X=np.reshape(X,[N,1])
bias_vector=np.ones([1,1])
return np.row_stack((bias_vector,X))
def addBias(X):
'''
为维数为N,d的向量X添加偏置向量
Args:
X(numpy.ndarray):shape为[N,d]
Returns:
numpy.ndarray:shape为[N,d+1],且第一列列向量全为1
'''
try:
N=X.shape[0]
d=X.shape[1]
if d == 1:
return addSingleBias(X)
bias_vector=np.ones([N,1])
return np.column_stack((bias_vector,X))
except:
return addSingleBias(X)
```
## loadData
```
#中国沿海城市的坐标集
X1 = np.array([
[119.1,34.4],
[121.3,29.5],
[120.2,36.1],
[119.4,39.6],
[113.5,22.3],
[118.1,24.5],
[121.4,34.5],
[114.1,22.6],
[110.1,20.3],
[123.3,38.6],
[117.2,39.1]])
X1_add = np.array([
[114,30],
[116,24],
[117,32]])
#日本沿海城市的坐标集
X2 = np.array([
[130.3,32.3],
[130.6,33.6],
[139.4,35.3],
[135.1,34.4],
[136.6,35.1],
[130.4,33.6],
[132.3,34.2],
[130.5,33.4],
[129.9,32.8],
[135.3,34.4],
[139.5,35.4]])
X2_add = np.array([
[141,38],
[138,36],
[135,35]])
island=np.asarray([123,25])
D=np.r_[X1,X2]
Y=np.r_[np.ones([len(X1),1]),-np.ones([len(X2),1])]
D_add=np.r_[X1,X1_add,X2,X2_add]
Y=np.r_[np.ones([len(X1),1]),-np.ones([len(X2),1])]
Y_add=np.r_[np.ones([len(X1)+len(X1_add),1]),-np.ones([len(X2)+len(X2_add),1])]
```
## basic
原始数据,皆为临海城市
```
d=2
I=np.eye(d)
smallBias=1e-6
Q= np.block([[smallBias,smallBias*np.ones([1,d])],
[smallBias*np.ones([d,1]),I]])
p=np.zeros(d+1)
A=-np.multiply(addBias(D),Y)
C=-np.ones(len(A))
x = solve_qp(Q, p, A, C)#G, h, A, b)
# solve_qp(solver= CVXOPT, OSQP and quadprog))
print("QP solution: x = {}".format(x))
x
```
### support vector
```
np.dot(x,addBias(D).transpose())
support_vector_site=np.where(np.abs(np.abs(np.dot(x,addBias(D).transpose()))-1)<=1e-6)[0]
for site in support_vector_site:
print(D[site])
```
### result
```
print(np.dot(x,addBias(island)))
if np.dot(x.transpose(),addBias(island))>0:
print('China')
else:
print('Japan')
```
### display
```
class Point():
def __init__(self,x,y):
self.x=x
self.y=y
W1=x.reshape(3)
# print(W1)
k=-W1[1]/W1[2]
b=-W1[0]/W1[2]
scale=1000
plt.figure(figsize=(7,5))
P1=Point(scale,k*scale+b)
P2=Point(-scale,-k*scale+b)
plt.scatter(D[:len(X1),0],D[:len(X1),1],marker = 'o', s=20, cmap = plt.cm.Spectral)
plt.scatter(D[len(X1):,0],D[len(X1):,1],marker = 'o', s=20, cmap = plt.cm.Spectral)
plt.scatter(D[support_vector_site][:,0],D[support_vector_site][:,1],marker = '+', s=50, cmap = plt.cm.Spectral)
plt.scatter(island[0],island[1],marker = '*', s=100, cmap = plt.cm.Spectral)
plt.plot([P1.x,P2.x],[P1.y,P2.y],'-')
plt.legend(['Classify plane','China','Japan','support vector','island'])
plt.grid(True)
plt.xlim((100, 150))
plt.ylim((0, 60))
plt.show()
```
## pro
增加非沿海城市数据
```
d=2
I=np.eye(d)
smallBias=1e-6
Q= np.block([[smallBias,smallBias*np.ones([1,d])],
[smallBias*np.ones([d,1]),I]])
p=np.zeros(d+1)
A=-np.multiply(addBias(D_add),Y_add)
C=-np.ones(len(A))
x = solve_qp(Q, p, A, C)#G, h, A, b)
# solve_qp(solver= CVXOPT, OSQP and quadprog))
print("QP solution: x = {}".format(x))
```
### support vector
```
support_vector_site=np.where(np.abs(np.abs(np.dot(x,addBias(D).transpose()))-1)<=1e-6)[0]
for site in support_vector_site:
print(D[site])
```
### result
```
print(np.dot(x,addBias(island)))
if np.dot(x.transpose(),addBias(island))>0:
print('China')
else:
print('Japan')
```
### display
```
class Point():
def __init__(self,x,y):
self.x=x
self.y=y
W1=x.reshape(3)
# print(W1)
k=-W1[1]/W1[2]
b=-W1[0]/W1[2]
scale=1000
plt.figure(figsize=(7,5))
P1=Point(scale,k*scale+b)
P2=Point(-scale,-k*scale+b)
plt.scatter(D[:len(X1),0],D[:len(X1),1],marker = 'o', s=20, cmap = plt.cm.Spectral)
plt.scatter(D[len(X1):,0],D[len(X1):,1],marker = 'o', s=20, cmap = plt.cm.Spectral)
plt.scatter(D[support_vector_site][:,0],D[support_vector_site][:,1],marker = '+', s=50, cmap = plt.cm.Spectral)
plt.scatter(island[0],island[1],marker = '*', s=100, cmap = plt.cm.Spectral)
plt.plot([P1.x,P2.x],[P1.y,P2.y],'-')
plt.xlim((100, 150))
plt.ylim((0, 60))
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/wisrovi/pyimagesearch-buy/blob/main/pokedex_find_screen_part_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

# Building a Pokedex in Python: Finding the Game Boy Screen (Step 4 of 6)
### by [PyImageSearch.com](http://www.pyimagesearch.com)
## Welcome to **[PyImageSearch Plus](http://pyimg.co/plus)** Jupyter Notebooks!
This notebook is associated with the [Building a Pokedex in Python: Finding the Game Boy Screen (Step 4 of 6)](https://www.pyimagesearch.com/2014/04/21/building-pokedex-python-finding-game-boy-screen-step-4-6/) blog post published on 2014-04-21.
Only the code for the blog post is here. Most codeblocks have a 1:1 relationship with what you find in the blog post with two exceptions: (1) Python classes are not separate files as they are typically organized with PyImageSearch projects, and (2) Command Line Argument parsing is replaced with an `args` dictionary that you can manipulate as needed.
We recommend that you execute (press ▶️) the code block-by-block, as-is, before adjusting parameters and `args` inputs. Once you've verified that the code is working, you are welcome to hack with it and learn from manipulating inputs, settings, and parameters. For more information on using Jupyter and Colab, please refer to these resources:
* [Jupyter Notebook User Interface](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface)
* [Overview of Google Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
As a reminder, these PyImageSearch Plus Jupyter Notebooks are not for sharing; please refer to the **Copyright** directly below and **Code License Agreement** in the last cell of this notebook.
Happy hacking!
*Adrian*
<hr>
***Copyright:*** *The contents of this Jupyter Notebook, unless otherwise indicated, are Copyright 2020 Adrian Rosebrock, PyimageSearch.com. All rights reserved. Content like this is made possible by the time invested by the authors. If you received this Jupyter Notebook and did not purchase it, please consider making future content possible by joining PyImageSearch Plus at http://pyimg.co/plus/ today.*
### Download the code zip file
```
!wget https://www.pyimagesearch.com/wp-content/uploads/2014/03/pokedex-find-screen-part-1.zip
!unzip -qq pokedex-find-screen-part-1.zip
%cd pokedex-find-screen-part-1
```
## Blog Post Code
### Import Packages
```
# import the necessary packages
from matplotlib import pyplot as plt
import numpy as np
import argparse
import imutils
import cv2
```
### Function to display images in Jupyter Notebooks and Google Colab
```
def plt_imshow(title, image):
# convert the image frame BGR to RGB color space and display it
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.imshow(image)
plt.title(title)
plt.grid(False)
plt.show()
```
### Building a Pokedex in Python: Finding the Game Boy Screen
```
# construct the argument parser and parse the arguments
#ap = argparse.ArgumentParser()
#ap.add_argument("-q", "--query", required = True,
# help = "Path to the query image")
#args = vars(ap.parse_args())
# since we are using Jupyter Notebooks we can replace our argument
# parsing code with *hard coded* arguments and values
args = {
"query": "queries/query_marowak.jpg"
}
# load the query image, compute the ratio of the old height
# to the new height, clone it, and resize it
image = cv2.imread(args["query"])
ratio = image.shape[0] / 300.0
orig = image.copy()
image = imutils.resize(image, height = 300)
# convert the image to grayscale, blur it, and find edges
# in the image
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
gray = cv2.bilateralFilter(gray, 11, 17, 17)
edged = cv2.Canny(gray, 30, 200)
# find contours in the edged image, keep only the largest
# ones, and initialize our screen contour
cnts = cv2.findContours(edged.copy(), cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
cnts = sorted(cnts, key = cv2.contourArea, reverse = True)[:10]
screenCnt = None
# loop over our contours
for c in cnts:
# approximate the contour
peri = cv2.arcLength(c, True)
approx = cv2.approxPolyDP(c, 0.01 * peri, True)
# if our approximated contour has four points, then
# we can assume that we have found our screen
if len(approx) == 4:
screenCnt = approx
break
# draw a rectangle around the screen
orig = image.copy()
cv2.drawContours(image, [screenCnt], -1, (0, 255, 0), 3)
plt_imshow("Game Boy Screen", image)
# create a mask for the screen
mask = np.zeros(image.shape[:2], dtype = "uint8")
cv2.drawContours(mask, [screenCnt], -1, 255, -1)
plt_imshow("Masked", cv2.bitwise_and(orig, orig, mask = mask))
```
For a detailed walkthrough of the concepts and code, be sure to refer to the full tutorial, [*Building a Pokedex in Python: Finding the Game Boy Screen (Step 4 of 6)*](https://www.pyimagesearch.com/2014/04/21/building-pokedex-python-finding-game-boy-screen-step-4-6/) published on 2014-04-21.
# Code License Agreement
```
Copyright (c) 2020 PyImageSearch.com
SIMPLE VERSION
Feel free to use this code for your own projects, whether they are
purely educational, for fun, or for profit. THE EXCEPTION BEING if
you are developing a course, book, or other educational product.
Under *NO CIRCUMSTANCE* may you use this code for your own paid
educational or self-promotional ventures without written consent
from Adrian Rosebrock and PyImageSearch.com.
LONGER, FORMAL VERSION
Permission is hereby granted, free of charge, to any person obtaining
a copy of this software and associated documentation files
(the "Software"), to deal in the Software without restriction,
including without limitation the rights to use, copy, modify, merge,
publish, distribute, sublicense, and/or sell copies of the Software,
and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be
included in all copies or substantial portions of the Software.
Notwithstanding the foregoing, you may not use, copy, modify, merge,
publish, distribute, sublicense, create a derivative work, and/or
sell copies of the Software in any work that is designed, intended,
or marketed for pedagogical or instructional purposes related to
programming, coding, application development, or information
technology. Permission for such use, copying, modification, and
merger, publication, distribution, sub-licensing, creation of
derivative works, or sale is expressly withheld.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND,
EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES
OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND
NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS
BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN
ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
```
| github_jupyter |
# Advanced Automatic Differentiation in JAX
[](https://colab.research.google.com/github/google/jax/blob/main/docs/jax-101/04-advanced-autodiff.ipynb)
*Authors: Vlatimir Mikulik & Matteo Hessel*
Computing gradients is a critical part of modern machine learning methods. This section considers a few advanced topics in the areas of automatic differentiation as it relates to modern machine learning.
While understanding how automatic differentiation works under the hood isn't crucial for using JAX in most contexts, we encourage the reader to check out this quite accessible [video](https://www.youtube.com/watch?v=wG_nF1awSSY) to get a deeper sense of what's going on.
[The Autodiff Cookbook](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html) is a more advanced and more detailed explanation of how these ideas are implemented in the JAX backend. It's not necessary to understand this to do most things in JAX. However, some features (like defining [custom derivatives](https://jax.readthedocs.io/en/latest/notebooks/Custom_derivative_rules_for_Python_code.html)) depend on understanding this, so it's worth knowing this explanation exists if you ever need to use them.
## Higher-order derivatives
JAX's autodiff makes it easy to compute higher-order derivatives, because the functions that compute derivatives are themselves differentiable. Thus, higher-order derivatives are as easy as stacking transformations.
We illustrate this in the single-variable case:
The derivative of $f(x) = x^3 + 2x^2 - 3x + 1$ can be computed as:
```
import jax
f = lambda x: x**3 + 2*x**2 - 3*x + 1
dfdx = jax.grad(f)
```
The higher-order derivatives of $f$ are:
$$
\begin{array}{l}
f'(x) = 3x^2 + 4x -3\\
f''(x) = 6x + 4\\
f'''(x) = 6\\
f^{iv}(x) = 0
\end{array}
$$
Computing any of these in JAX is as easy as chaining the `grad` function:
```
d2fdx = jax.grad(dfdx)
d3fdx = jax.grad(d2fdx)
d4fdx = jax.grad(d3fdx)
```
Evaluating the above in $x=1$ would give us:
$$
\begin{array}{l}
f'(1) = 4\\
f''(1) = 10\\
f'''(1) = 6\\
f^{iv}(1) = 0
\end{array}
$$
Using JAX:
```
print(dfdx(1.))
print(d2fdx(1.))
print(d3fdx(1.))
print(d4fdx(1.))
```
In the multivariable case, higher-order derivatives are more complicated. The second-order derivative of a function is represented by its [Hessian matrix](https://en.wikipedia.org/wiki/Hessian_matrix), defined according to
$$(\mathbf{H}f)_{i,j} = \frac{\partial^2 f}{\partial_i\partial_j}.$$
The Hessian of a real-valued function of several variables, $f: \mathbb R^n\to\mathbb R$, can be identified with the Jacobian of its gradient. JAX provides two transformations for computing the Jacobian of a function, `jax.jacfwd` and `jax.jacrev`, corresponding to forward- and reverse-mode autodiff. They give the same answer, but one can be more efficient than the other in different circumstances – see the [video about autodiff](https://www.youtube.com/watch?v=wG_nF1awSSY) linked above for an explanation.
```
def hessian(f):
return jax.jacfwd(jax.grad(f))
```
Let's double check this is correct on the dot-product $f: \mathbf{x} \mapsto \mathbf{x} ^\top \mathbf{x}$.
if $i=j$, $\frac{\partial^2 f}{\partial_i\partial_j}(\mathbf{x}) = 2$. Otherwise, $\frac{\partial^2 f}{\partial_i\partial_j}(\mathbf{x}) = 0$.
```
import jax.numpy as jnp
def f(x):
return jnp.dot(x, x)
hessian(f)(jnp.array([1., 2., 3.]))
```
Often, however, we aren't interested in computing the full Hessian itself, and doing so can be very inefficient. [The Autodiff Cookbook](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html) explains some tricks, like the Hessian-vector product, that allow to use it without materialising the whole matrix.
If you plan to work with higher-order derivatives in JAX, we strongly recommend reading the Autodiff Cookbook.
## Higher order optimization
Some meta-learning techniques, such as Model-Agnostic Meta-Learning ([MAML](https://arxiv.org/abs/1703.03400)), require differentiating through gradient updates. In other frameworks this can be quite cumbersome, but in JAX it's much easier:
```python
def meta_loss_fn(params, data):
"""Computes the loss after one step of SGD."""
grads = jax.grad(loss_fn)(params, data)
return loss_fn(params - lr * grads, data)
meta_grads = jax.grad(meta_loss_fn)(params, data)
```
## Stopping gradients
Auto-diff enables automatic computation of the gradient of a function with respect to its inputs. Sometimes, however, we might want some additional control: for instance, we might want to avoid back-propagating gradients through some subset of the computational graph.
Consider for instance the TD(0) ([temporal difference](https://en.wikipedia.org/wiki/Temporal_difference_learning)) reinforcement learning update. This is used to learn to estimate the *value* of a state in an environment from experience of interacting with the environment. Let's assume the value estimate $v_{\theta}(s_{t-1}$) in a state $s_{t-1}$ is parameterised by a linear function.
```
# Value function and initial parameters
value_fn = lambda theta, state: jnp.dot(theta, state)
theta = jnp.array([0.1, -0.1, 0.])
```
Consider a transition from a state $s_{t-1}$ to a state $s_t$ during which we observed the reward $r_t$
```
# An example transition.
s_tm1 = jnp.array([1., 2., -1.])
r_t = jnp.array(1.)
s_t = jnp.array([2., 1., 0.])
```
The TD(0) update to the network parameters is:
$$
\Delta \theta = (r_t + v_{\theta}(s_t) - v_{\theta}(s_{t-1})) \nabla v_{\theta}(s_{t-1})
$$
This update is not the gradient of any loss function.
However, it can be **written** as the gradient of the pseudo loss function
$$
L(\theta) = [r_t + v_{\theta}(s_t) - v_{\theta}(s_{t-1})]^2
$$
if the dependency of the target $r_t + v_{\theta}(s_t)$ on the parameter $\theta$ is ignored.
How can we implement this in JAX? If we write the pseudo loss naively we get:
```
def td_loss(theta, s_tm1, r_t, s_t):
v_tm1 = value_fn(theta, s_tm1)
target = r_t + value_fn(theta, s_t)
return (target - v_tm1) ** 2
td_update = jax.grad(td_loss)
delta_theta = td_update(theta, s_tm1, r_t, s_t)
delta_theta
```
But `td_update` will **not** compute a TD(0) update, because the gradient computation will include the dependency of `target` on $\theta$.
We can use `jax.lax.stop_gradient` to force JAX to ignore the dependency of the target on $\theta$:
```
def td_loss(theta, s_tm1, r_t, s_t):
v_tm1 = value_fn(theta, s_tm1)
target = r_t + value_fn(theta, s_t)
return (jax.lax.stop_gradient(target) - v_tm1) ** 2
td_update = jax.grad(td_loss)
delta_theta = td_update(theta, s_tm1, r_t, s_t)
delta_theta
```
This will treat `target` as if it did **not** depend on the parameters $\theta$ and compute the correct update to the parameters.
The `jax.lax.stop_gradient` may also be useful in other settings, for instance if you want the gradient from some loss to only affect a subset of the parameters of the neural network (because, for instance, the other parameters are trained using a different loss).
## Straight-through estimator using `stop_gradient`
The straight-through estimator is a trick for defining a 'gradient' of a function that is otherwise non-differentiable. Given a non-differentiable function $f : \mathbb{R}^n \to \mathbb{R}^n$ that is used as part of a larger function that we wish to find a gradient of, we simply pretend during the backward pass that $f$ is the identity function. This can be implemented neatly using `jax.lax.stop_gradient`:
```
def f(x):
return jnp.round(x) # non-differentiable
def straight_through_f(x):
# Create an exactly-zero expression with Sterbenz lemma that has
# an exactly-one gradient.
zero = x - jax.lax.stop_gradient(x)
return zero + jax.lax.stop_gradient(f(x))
print("f(x): ", f(3.2))
print("straight_through_f(x):", straight_through_f(3.2))
print("grad(f)(x):", jax.grad(f)(3.2))
print("grad(straight_through_f)(x):", jax.grad(straight_through_f)(3.2))
```
## Per-example gradients
While most ML systems compute gradients and updates from batches of data, for reasons of computational efficiency and/or variance reduction, it is sometimes necessary to have access to the gradient/update associated with each specific sample in the batch.
For instance, this is needed to prioritise data based on gradient magnitude, or to apply clipping / normalisations on a sample by sample basis.
In many frameworks (PyTorch, TF, Theano) it is often not trivial to compute per-example gradients, because the library directly accumulates the gradient over the batch. Naive workarounds, such as computing a separate loss per example and then aggregating the resulting gradients are typically very inefficient.
In JAX we can define the code to compute the gradient per-sample in an easy but efficient way.
Just combine the `jit`, `vmap` and `grad` transformations together:
```
perex_grads = jax.jit(jax.vmap(jax.grad(td_loss), in_axes=(None, 0, 0, 0)))
# Test it:
batched_s_tm1 = jnp.stack([s_tm1, s_tm1])
batched_r_t = jnp.stack([r_t, r_t])
batched_s_t = jnp.stack([s_t, s_t])
perex_grads(theta, batched_s_tm1, batched_r_t, batched_s_t)
```
Let's walk through this one transformation at a time.
First, we apply `jax.grad` to `td_loss` to obtain a function that computes the gradient of the loss w.r.t. the parameters on single (unbatched) inputs:
```
dtdloss_dtheta = jax.grad(td_loss)
dtdloss_dtheta(theta, s_tm1, r_t, s_t)
```
This function computes one row of the array above.
Then, we vectorise this function using `jax.vmap`. This adds a batch dimension to all inputs and outputs. Now, given a batch of inputs, we produce a batch of outputs -- each output in the batch corresponds to the gradient for the corresponding member of the input batch.
```
almost_perex_grads = jax.vmap(dtdloss_dtheta)
batched_theta = jnp.stack([theta, theta])
almost_perex_grads(batched_theta, batched_s_tm1, batched_r_t, batched_s_t)
```
This isn't quite what we want, because we have to manually feed this function a batch of `theta`s, whereas we actually want to use a single `theta`. We fix this by adding `in_axes` to the `jax.vmap`, specifying theta as `None`, and the other args as `0`. This makes the resulting function add an extra axis only to the other arguments, leaving `theta` unbatched, as we want:
```
inefficient_perex_grads = jax.vmap(dtdloss_dtheta, in_axes=(None, 0, 0, 0))
inefficient_perex_grads(theta, batched_s_tm1, batched_r_t, batched_s_t)
```
Almost there! This does what we want, but is slower than it has to be. Now, we wrap the whole thing in a `jax.jit` to get the compiled, efficient version of the same function:
```
perex_grads = jax.jit(inefficient_perex_grads)
perex_grads(theta, batched_s_tm1, batched_r_t, batched_s_t)
%timeit inefficient_perex_grads(theta, batched_s_tm1, batched_r_t, batched_s_t).block_until_ready()
%timeit perex_grads(theta, batched_s_tm1, batched_r_t, batched_s_t).block_until_ready()
```
| github_jupyter |
# 循环
- 循环是一种控制语句块重复执行的结构
- while 适用于广度遍历
- for 开发中经常使用
## while 循环
- 当一个条件保持真的时候while循环重复执行语句
- while 循环一定要有结束条件,否则很容易进入死循环
- while 循环的语法是:
while loop-contunuation-conndition:
Statement
```
i = 0
while i<5:
print('hahaha')
i += 1 #i=i+1
sum = 0
i = 0
while i <10:
sum = sum + i
i = i + 1
print(i)
```
## 示例:
sum = 0
i = 1
while i <10:
sum = sum + i
i = i + 1
## 错误示例:
sum = 0
i = 1
while i <10:
sum = sum + i
i = i + 1
- 一旦进入死循环可按 Ctrl + c 停止
## EP:


# 验证码
- 随机产生四个字母的验证码,如果正确,输出验证码正确。如果错误,产生新的验证码,用户重新输入。
- 验证码只能输入三次,如果三次都错,返回“别爬了,我们小网站没什么好爬的”
- 密码登录,如果三次错误,账号被锁定
```
import random
q=random.randint(1000,4444)
print(q)
a=eval(input('请输入验证码'))
while 1:
if a==q:
print('正确')
break
if a!=q:
print('错误请重新输入')
q=random.randint(1000,4444)
print(q)
a=eval(input('请输入验证码'))
import random
i=0
q=random.randint(1111,6666)
print(q)
d=eval(input('请输入验证码'))
while i<3:
if q==d:
print('正确')
break
if q!=d:
print('错误请再输入一次')
q=random.randint(1111,6666)
print(q)
if i==3:
print('回去')
a=eval(input('请输入验证码'))
i+=1
if i==3:
print('回去')
i=0
while i<3:
index1=random.randint(97,122)
index2=random.randint(97,122)
index3=random.randint(97,122)
index4=random.randint(97,122)
n1=chr(index1)
n2=chr(index2)
n3=chr(index3)
n4=chr(index4)
N=n1+n2+n2+n4
print(N)
me=input('>>')
if me == N:
print('正确')
else:
print('错误')
i+=1
if i==3:
print('回去')
a=[chr(random.randint(97,122)) for i in range(4)]
N="".join(a)
i=0
while i<3:
index1=random.randint(65,122)
index2=random.randint(65,122)
index3=random.randint(65,122)
index4=random.randint(65,122)
n1=chr(index1)
n2=chr(index2)
n3=chr(index3)
n4=chr(index4)
N=n1+n2+n2+n4
print(N)
me=input('请输入验证码')
if me == N:
print('正确')
break
else:
print('错误')
i+=1
if i==3:
print('回去')
n=random.randint(65,122)
N=""
i=0
while 1:
if 91<=n<=96:
n=random.randint(65,122)
else:
N+=chr(n)
i+=1
n=random.randint(65,122)
if i==4:
break
print(N)
```
## 尝试死循环
## 实例研究:猜数字
- 你将要编写一个能够随机生成一个0到10之间的且包括两者的数字程序,这个程序
- 提示用户连续地输入数字直到正确,且提示用户输入的数字是过高还是过低
## 使用哨兵值来控制循环
- 哨兵值来表明输入的结束
- 
## 警告

```
i=1
while i!=0:
i-=0.1
```
## for 循环
- Python的for 循环通过一个序列中的每个值来进行迭代
- range(a,b,k), a,b,k 必须为整数
- a: start
- b: end
- k: step
- 注意for 是循环一切可迭代对象,而不是只能使用range
```
for i in range(0,10,2):
print(i)
for i in range(1,10,2):
print(i)
for i in range(10,1,-1):
print(i)
for i in range(11,1,-2):
print(i)
a='jnsdv'
for i in a:
print(i)
a={1,2,3}
a.__iter__
for i in a:
print(i)
```
# 在Python里面一切皆对象
## EP:
- 
```
sum_=0
i=0
while i < 1001:
sum_ = sum_ + i
i += 1
print(sum)
sum_=0
for i in range(1,10001):
sum_+=i
if sum_>10000:
break
print(sum_)
```
## 嵌套循环
- 一个循环可以嵌套另一个循环
- 每次循环外层时,内层循环都会被刷新重新完成循环
- 也就是说,大循环执行一次,小循环会全部执行一次
- 注意:
> - 多层循环非常耗时
- 最多使用3层循环
## EP:
- 使用多层循环完成9X9乘法表
- 显示50以内所有的素数
```
for e in range(1,10):
for y in range(1,e+1):
jiujiu=e*y
print(y,'x',e,'=',jiujiu,end=' ')
print()
import random
for n in range(50):
for h in range(2,n-1):
if (n/h)!=(n/h):
print(n)
```
## 关键字 break 和 continue
- break 跳出循环,终止循环
- continue 跳出此次循环,继续执行
```
for i in range(10):
if i==5:
break
print(i)
for i in range(10):
if i==5:
continue
print(i)
for i in range(10):
if i==5:
pass
else:
print(i)
for i in range(10):
for j in range(10):
print(i,j)
```
## 注意


# Homework
- 1

```
zs=0
fs=0
while 1:
number=eval(input('请输入:'))
if number==0:
break
if number>0:
zs=zs+number
print(zs)
if number<0:
fs=fs+number
print(fs)
```
- 2

```
sum_four=0
now=10000
for i in range(1,15):
if i==10:
ten=now*(1+0.05)**i
print(ten)
if i>10:
i+=1
four=now*(1+0.05)**i
sum_four=sum_four+four
print(sum_four)
```
- 3

- 4

```
for i in range(100,1000):
if i%5==0 and i%6==0:
print(i,end=' ')
```
- 5

```
n=0
n+=1
while n**3<12000:
break
```
- 6

```
d_money=eval(input('Loan Amount:'))
time_money=eval(input('Number of Year: '))
result='Interest Rate Monthly Payment Total Payment'
print(result)
n=5
mon=188.71
toal=11322.74
for i in range(24):
n+=0.125
print()
print(n,'%')
for j in range(1):
mon+=0.57
print(' ',round(mon,2))
for k in range(1):
toal+=34.39
print(' ',round(toal,2))
```
- 7

```
i = 0
sum_left = 0
while i < 50000:
i = i + 1
sum_left=sum_left+(1/i)
print(sum_left)
j = 50001
sum_right = 0
while j>1:
j-=1
sum_right=sum_right+(1/j)
print(sum_right)
sum_left=0
sum_right=0
for i in range(1,50001):
sum_left=sum_left+(1/i)
print(sum_left)#从左到右
for j in range(50000,0,-1):
sum_right=sum_right+(1/j)#从右到左
print(sum_right)
```
- 8

```
for i in range(1,98,2):
for j in range(3,100,2):
sum_=i/j
print(sum_)
```
- 9

```
for i in range(10000,100001,10000):
mi=(-1)**(i+1)
for j in range(1,2*i-1,2):
pin_=4*(mi/j)
print(pin_)
```
- 10

```
for i in range(1,10000):
for n in range(1,i):
if i%n==0:
print(n,i/n)
```
- 11

```
n=0
for i in range(1,8):
for j in range(1,8):
if i==j:
continue
elif (i//10==j%10) and (i%10==j//10):
continue
print(i,'',j)
n=n+(i+j)*(1/(i+j))
print('总共有',n)
```
- 12

```
number1=eval(input('Enter ten number: '))
number2=eval(input(''))
number3=eval(input(''))
number4=eval(input(''))
number5=eval(input(''))
number6=eval(input(''))
number7=eval(input(''))
number8=eval(input(''))
number9=eval(input(''))
number10=eval(input(''))
mean=(number1+number2+number3+number4+number5+number6+number7+number8+number9+number10)/10
agv=(((number1-mean)**2+(number2-mean)**2+(number3-mean)**2+(number4-mean)**2+
(number5-mean)**2+(number6-mean)**2+(number7-mean)**2+(number8-mean)**2+
(number9-mean)**2+(number10-mean)**2)/(10-1))**0.5
print(mean)
print(agv)
```
| github_jupyter |
```
#3.5 The Image Classification Dataset
import torch
import torchvision
from torch.utils import data
from torchvision import transforms
from d2l import torch as d2l
d2l.use_svg_display()
#3.5.1 Reading the Dataset
trans = transforms.ToTensor() #將圖像從PIL類型轉換為32-bit浮點數的tensor(將像素值/255,因此全部的值都介在0到1之間)
mnist_train = torchvision.datasets.FashionMNIST(root="../data", train=True, transform=trans,download=True)
mnist_test = torchvision.datasets.FashionMNIST(root="../data", train=False, transform=trans, download=True)
len(mnist_train), len(mnist_test)
#Fashion-MNIST 由10個類別的圖像組成,每個類別由"訓練數據集"中的6000張圖像和"測試數據集"中的1000張圖像表示
mnist_train[0][0].shape
#每個input img的(height,width)==(28,28)
def get_fashion_mnist_labels(labels): #使用labels(index)對應label
"""Return text labels for the Fashion-MNIST dataset."""
text_labels = [
't-shirt', 'trouser', 'pullover', 'dress', 'coat', 'sandal', 'shirt',
'sneaker', 'bag', 'ankle boot']
return [text_labels[int(i)] for i in labels]
def show_images(imgs, num_rows, num_cols, titles=None, scale=1.5):
"""Plot a list of images."""
figsize = (num_cols * scale, num_rows * scale)
_, axes = d2l.plt.subplots(num_rows, num_cols, figsize=figsize)
axes = axes.flatten()
for i, (ax, img) in enumerate(zip(axes, imgs)):
if torch.is_tensor(img):
# Tensor Image
ax.imshow(img.numpy())
else:
# PIL Image
ax.imshow(img)
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
if titles:
ax.set_title(titles[i])
return axes
X, y = next(iter(data.DataLoader(mnist_train, batch_size=18)))
show_images(X.reshape(18, 28, 28), 3, 6, titles=get_fashion_mnist_labels(y));
#3.5.2 Reading a Minibatch
batch_size = 256
def get_dataloader_workers():
"""Use 4 processes to read the data."""
return 4
train_iter = data.DataLoader(mnist_train, batch_size, shuffle=True, num_workers=get_dataloader_workers())
timer = d2l.Timer()
for X, y in train_iter:
continue
f'讀取training data的時間: {timer.stop():.2f} sec'
#3.5.3 Putting All Things Together
def load_data_fashion_mnist(batch_size, resize=None): #獲取和讀取 Fashion-MNIST 數據集
"""Download the Fashion-MNIST dataset and then load it into memory."""
trans = [transforms.ToTensor()]
if resize:
trans.insert(0, transforms.Resize(resize))
trans = transforms.Compose(trans)
mnist_train = torchvision.datasets.FashionMNIST(root="../data", train=True, transform=trans, download=True)
mnist_test = torchvision.datasets.FashionMNIST(root="../data", train=False, transform=trans, download=True)
return (data.DataLoader(mnist_train, batch_size, shuffle=True, num_workers=get_dataloader_workers()),
data.DataLoader(mnist_test, batch_size, shuffle=False, num_workers=get_dataloader_workers()))
train_iter, test_iter = load_data_fashion_mnist(32, resize=64)
for X, y in train_iter:
print(X.shape, X.dtype, y.shape, y.dtype)
break
```
| github_jupyter |
```
import csv
import matplotlib.pyplot as plt
import numpy as np
# decompose accuracies
# pred = [3, 4, 5] gt = [] = wrong
# pred = [], gt = [1] = wrong
# pred = [], gt = [] =
my_data
# Load the data into a np array
my_data = np.genfromtxt('results_train.csv', delimiter=' ', dtype=float)
#plots = csv.reader(csvfile, delimiter=' ')
x = range(len(my_data) -1) # all the training accuracies
plt.plot(x,my_data[:-1,1], label='Agg accuracy')
plt.plot(x,my_data[:-1,2], label='Sel accuracy')
plt.plot(x,my_data[:-1,3], label='Cond accuracy')
plt.plot(x,my_data[:-1,4], label='Group accuracy')
plt.plot(x,my_data[:-1,5], label='Having accuracy')
plt.plot(x, my_data[:-1, 6], label='Orderby accuracy')
plt.plot(x, my_data[:-1, 7], label='Limit accuracy')
plt.plot(x, my_data[:-1,0], label = 'Total accuracy')
plt.xlabel('Iteration')
plt.ylabel('Accuracy')
plt.title('Training Accuracy')
plt.legend(loc=9, bbox_to_anchor=(1.3, 0.5))
#pylab.legend(loc=9, bbox_to_anchor=(0.5, -0.1))
plt.show()
# Load the data into a np array
dev_data = np.genfromtxt('results_dev.csv', delimiter=' ', dtype=float)
plt.plot(x,dev_data[:-2,1], label='Agg accuracy')
plt.plot(x,dev_data[:-2,2], label='Sel accuracy')
plt.plot(x,dev_data[:-2,3], label='Cond accuracy')
plt.plot(x,dev_data[:-2,4], label='Group accuracy')
plt.plot(x,dev_data[:-2,5], label='Having accuracy')
plt.plot(x, dev_data[:-2, 6], label="Orderby accuracy")
plt.plot(x, dev_data[:-2,0], label='Overall accuracy')
plt.plot(x, dev_data[:-2, 7], label="Limit accuracy")
plt.xlabel('Iteration')
plt.ylabel('Accuracy')
plt.title('Dev Accuracy')
plt.legend(loc=9, bbox_to_anchor=(1.3, 0.5))
#pylab.legend(loc=9, bbox_to_anchor=(0.5, -0.1))
plt.show()
best_train_acc = np.nanmax(my_data, axis=0)
best_dev_acc = np.nanmax(dev_data,axis=0) # Overall_acc, Agg_acc, sel_acc, Cond_acc, Group_acc,
#Having_Acc, OrderBy_acc, Limit_acc
print(best_train_acc)
print(best_dev_acc)
```
## APPEND DATA TO BEST_RESULTS_FILE
```
import csv
import pandas
# with open('best_results_train.csv', 'wb') as csvfile:
# spamwriter = csv.writer(csvfile, delimiter=' ', quotechar='|', quoting=csv.QUOTE_MINIMAL)
# spamwriter.writerow(['total', 'agg', 'sel', 'cond', 'group', 'epoch_num', 'toy', 'changes_made'])
# with open('best_results_dev.csv', 'wb') as csvfile:
# new_writer = csv.writer(csvfile, delimiter=' ', quotechar='|', quoting=csv.QUOTE_MINIMAL)
# new_writer.writerow(['total', 'agg', 'sel', 'cond', 'group', 'epoch_num', 'toy', 'changes_made'])
# Train_Data
to_write = np.append(np.nanmax(my_data, axis=0), [300, True])
# Get Data
to_app = {}
names = ['total', 'agg', 'sel', 'cond', 'group', 'epoch_num', 'toy', 'changes_made']
for i, item in enumerate(to_write):
to_app[names[i]] = item
to_app['toy'] = to_app['toy'] == 1
to_app['changes_made'] = 'moving agg accuracy back to before'
# Convert to DataFrame - one row
to_add = pandas.DataFrame(columns=names)
to_add.loc[0] = to_app
# Append to CSV
with open('best_results_train.csv', 'a') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=' ', quotechar='|', quoting=csv.QUOTE_MINIMAL)
spamwriter.writerow(to_add.loc[0])
# Dev_data
to_write = np.append(np.nanmax(dev_data, axis=0), [300, True])
# Get Data
to_app = {}
names = ['total', 'agg', 'sel', 'cond', 'group', 'epoch_num', 'toy', 'changes_made']
for i, item in enumerate(to_write):
to_app[names[i]] = item
to_app['toy'] = to_app['toy'] == 1
to_app['changes_made'] = 'moving agg accuracy back to before'
# Convert to DataFrame - one row
to_add = pandas.DataFrame(columns=names)
to_add.loc[0] = to_app
# Append to CSV
with open('best_results_dev.csv', 'a') as csvfile:
spamwriter = csv.writer(csvfile, delimiter=' ', quotechar='|', quoting=csv.QUOTE_MINIMAL)
spamwriter.writerow(to_add.loc[0])
```
## See Total History of Changes
```
train_hist = np.genfromtxt('best_results_train.csv', delimiter=' ',usecols=np.arange(0,5), dtype=float)
dev_hist = np.genfromtxt('best_results_dev.csv', delimiter=' ', usecols = np.arange(0, 5), dtype=float)
x = range(len(train_hist) - 1)
plt.plot(x,train_hist[1:,1],'--o', label='Agg accuracy' )
plt.plot(x,train_hist[1:,2], '--o', label='Sel accuracy')
plt.plot(x,train_hist[1:,3], '--o',label='Cond accuracy')
plt.plot(x,train_hist[1:,4], '--o',label='Group accuracy')
plt.plot(x, train_hist[1:,0], '--o',label = 'Total accuracy')
plt.xlabel('Attempt')
plt.ylabel('Accuracy')
plt.title('Training Accuracy')
plt.legend(loc=9, bbox_to_anchor=(1.3, 0.5))
#pylab.legend(loc=9, bbox_to_anchor=(0.5, -0.1))
plt.show()
dev_hist[1:,2] # SELECTION ACCURACY
dev_hist[1:,1] # AGGREGATION ACCURACY
```
| github_jupyter |
# TME 10 : Compression d'images avec le codage de Huffman
> Consignes: le fichier **TME10_Sujet.ipynb** est à déposer sur le site Moodle de l'UE https://moodle-sciences.upmc.fr/moodle-2019/course/view.php?id=4248. Si vous êtes en binôme, renommez-le en **TME10_nom1_nom2.ipynb**.
N'oubliez pas de sauvegarder fréquemment votre notebook !! ...
Pour chaque questions, écrivez des commentaires brefs et concis lorsque demandé dans les cases **Réponse**.
```
# Chargement des modules et fonctions utiles.
from PIL import Image
import matplotlib.pyplot as plt
import numpy as np
def affichage_14(affichages,titres=None, axis=True):
# effectue entre 1 et 4 affichages avec leurs titres, pour des images ou courbes
# paramètres :
# - liste des affichages (entre 1 et 4)
# - liste des titres (entre 1 et 4, autant que de affichages) Optionnelle
if not type(affichages) == type([]):
affichages = [affichages]
if titres is None:
titres = ['',]*len(affichages)
if not type(titres) == type([]):
titres = [titres]
nb_affichages = len(affichages)
if nb_affichages >4 or nb_affichages < 1 :
raise ValueError('affichage_14 nécéssite 1 à 4 entrées en paramètre')
if nb_affichages != len(titres):
raise ValueError('affichage_14 nécéssite autant de titres que d\'affichages')
courbes = False
for i in range(0,nb_affichages):
s = plt.subplot(101+10*nb_affichages+i)
s.set_title(titres[i])
if len(affichages[i].shape)>=2 and affichages[i].shape[0] > 1 and affichages[i].shape[1] > 1:
# on affiche une image
s.imshow(affichages[i], cmap="gray",interpolation='nearest', aspect='equal')
else :
# il s'agit d'une seule ligne, à afficher comme une courbe
plt.plot(affichages[i])
courbes=True
# ne pas afficher les axes si demandé
if not axis:
plt.axis('off')
agrandissement_h = nb_affichages
agrandissement_v = nb_affichages*2 if courbes else nb_affichages
params = plt.gcf()
plSize = params.get_size_inches()
params.set_size_inches( (plSize[0]*agrandissement_v, plSize[1]*agrandissement_h) )
plt.show()
def litbit(s,b):
""" bytearray*int->str
lit la valeur du bit b dans la table d'octets s
et retourne '1' ou '0'
"""
byte , bit = b//8 , 7-b%8
if s[byte] & (1<<bit):
return '1'
else:
return '0'
def ecritbit(s,b,val):
""" bytearray*int*str -> NoneType
len(str) == 1
écrit la valeur du bit b dans la table d'octets s
"""
byte , bit = b // 8 , 7 - b % 8
if val == '1':
s[byte] = s[byte] | (1<<bit)
else:
s[byte] = s[byte] & ~(1<<bit)
```
## Calcul d'entropie sur des images
Considérons un ensemble d'images en nuances de gris, notre source, et considérons la valeur de leurs pixels comme une variable aléatoire X ($X\in[0,255]$ la plus-part du temps).
On va dans un premier temps calculer l'entropie de X pour différentes sources, i.e différents groupes d'images.
### Question 1 :
1. Charger plusieurs images dans des tableaux ```np.array()``` dont une image binaire (par exemple ``rectangle.png``), puis affichez-les (on pourra grouper les images par groupe de 4 maximum et utiliser la fonction ```affichage_14()```).
2. Calculer l'entropie pour chaque image (une image est vue comme la réalisation d'une variable aléatoire, et donc une suite de symboles). Indication: utiliser la fonction ``np.histogram(img,256)`` du module Numpy.
3. Quelle différence y a-t-il entre l'entropie de l'image binaire et celle des images non binaires ? Pourquoi ?
```
### Reponse 1.1
rectangle = np.array(Image.open("images/rectangle.png"))
couloir = np.array(Image.open("images/couloir.png"))
bureau = np.array(Image.open("images/bureau.png"))
affichage_14([rectangle, couloir, bureau], ["rectangle", "couloir", "bureau"])
### Réponse 1.2
for img, titre in zip([rectangle, couloir, bureau], ["rectangle", "couloir", "bureau"]):
# histograme
N = img.shape[0]*img.shape[1]
hist, _ = np.histogram(img, 256)
hist = hist/N
plt.figure(figsize=(20, 10))
plt.subplot(121)
plt.title(titre)
plt.imshow(img, cmap='gray')
# entropy
plt.subplot(122)
plt.bar(np.arange(256), hist)
hist[hist == 0] = 1 # log(1) = 0
entropy = -hist.dot(np.log2(hist))
plt.title("Entropy = " + str(entropy))
plt.show()
```
### Réponse 1.3
- On remarque que l'image binaire a une entropie relativement petite comparé aux images réelles.
- cela est du au fait qu'on distingue exactement 2 couleurs dans les images binaires et qui ont des probabilités assez large et donc plus certain, comparé au images réelle qui elles couvrent un spectre plus important de couleurs avec des probabilités plus ou moins égales, et donc plus uncertain et necessiterait plus d'information pour conclure la valeur d'un pixel.
## Encodage des pixels avec le code de Huffman
On suppose ici connaître la dimension des images (donc leur nombre de pixels et leur nombre de lignes), et on souhaite encoder ces images en séquences binaires les plus compactes possibles.
On va pour cela utiliser le codage de Huffman.
### Question 2
1. Écrire la fonction ``huffman()`` vue en TD.
```
# type Noeud = tuple[int,float,list[Noeud]]
import pdb
import heapq
class Noeud(object):
def __init__(self, val=-1, freq=0, left=None, right=None):
self.val = val
self.freq = freq
self.right = right
self.left = left
def __repr__(self):
if self.val != -1:# feuille
# pdb.set_trace()
return f'{self.freq}({self.val})'
else: # Feuille ...
return f'{self.left} {self.right}'
def __lt__(self, other):
#print(self.freq, other.freq ,self.freq < other.freq)
return self.freq < other.freq
def huffman(hisn):
""" Array[float] -> Noeud
construit l'arbre de huffman correspondant a
l'histogramme normalise hisn et retourne la racine.
"""
N = len(hisn)
L = [Noeud(i, hisn[i]) for i in range(N)]
heapq.heapify(L) # pour reduire de la complexité
#print(L)
while len(L) > 1:
n1 = heapq.heappop(L)
n2 = heapq.heappop(L)
n = Noeud(freq=n1.freq+n2.freq, left=n1, right=n2)
#print("left n =", n.left, "right n =", n.right)
heapq.heappush(L, n)
return L[0]
```
2. Testez la sur l'exemple du TD en imprimant l'arbre.
```
histo_td = np.zeros(256)
for v,f in [(0,12046),(36,3325),(73,2390),(109,8716),(146,19544),(182,18748),(219,594),(255,178)]:
histo_td[v] = f
###
# histo_td = np.zeros(8)
# for v,f in [(0,12046),(1,3325),(2,2390),(3,8716),(4,19544),(5,18748),(6,594),(7,178)]:
# histo_td[v] = f
print(huffman(histo_td))
```
### Question 3
1. Écrire la fonction ```hufftable()``` vu en TD.
```
def hufftable(racine):
""" Noeud -> dict[int,str] """
def lirecode(nd: Noeud, code: str):
if nd.val == -1:
lirecode(nd.left, code+'0')
lirecode(nd.right, code+'1')
else:
table[nd.val] = code
del nd
table = dict()
lirecode(racine, '')
return table
```
2. Tester la sur l'arbre calculé précédemment et afficher la table
```
table = hufftable(huffman(histo_td))
table
```
### Question 4
1. Écrire la fonction ``encode()`` vue en TD.
```
def encode(img,table):
""" Array[int] * dict[int,str] -> tuple[bytearray,int]
Encode l'image img selon la table de Huffman table
retourne un tableau d'octets et la véritable longueur
en bit des données encodés
"""
def ecritbit(s, b, val):
"""bytearray*int*str -> None"""
byte, bit = b // 8, 7-(b%8)
if val == '1':
s[byte] = s[byte] | (1<<bit)
else:
s[byte] = s[byte] & ~(1<<bit)
nbits = 0
out = bytearray([0]*img.size)
img_r = img.reshape(img.size)
for val in img_r:
for s in table[val]:
ecritbit(out, nbits, s)
nbits += 1
return out, nbits
```
2. Tester la sur la portion d'image utilisée en TD. Afficher la taille en bit avant et après encodage.
```
img_td = np.array([
[109, 182, 109, 109, 219, 109, 146, 182],
[146, 219, 182, 182, 255, 219, 219, 182],
[219, 219, 219, 219, 219, 219, 219, 182],
[36, 182, 73, 73, 219, 73, 109, 182],
[36, 146, 73, 73, 182, 73, 109, 182],
[36, 109, 73, 73, 109, 73, 146, 182],
[73, 36, 36, 36, 182, 146, 182, 182],
[182, 146, 146, 146, 182, 182, 182, 182]
],dtype=int)
out, nbits = encode(img_td, table)
print("taille avant :", 8*img_td.size, ",taille aprés :", nbits)
```
3. Écrire une fonction ``print_encode()`` qui imprime le tampon encodé sous la forme d'une suite de caractères '0' et
'1'. Tester la sur l'exemple précédent.
```
def print_encoded( encoded, l):
""" bytearray*int->str"""
return ''.join(format(byte, '08b') for byte in encoded)[:l]
print_encoded(out, nbits)
```
### Question 5
1. Écrire le code de la fonction ```decode()``` vu en TD.
```
def decode(encoded,shape,table):
""" bytearray*tuple[int,int]*dict[int,str]->Array[int]
decode une image de huffman
"""
def litbit(s, b):
byte, bit = b//8, 7-(b%8)
return '1' if s[byte] & (1<<bit) else '0'
out = np.zeros(shape[0]*shape[1])
bits = 0
table2 = {v: k for k, v in table.items()}
for i in range(out.size):
code = ''
while not code in table2:
code = code + litbit(encoded, bits)
bits += 1
out[i] = table2[code]
return out.reshape(shape)
```
2. Tester la sur la portion d''image précédemment encodée et vérifier qu'elle est bien égale à la portion d'image originale.
```
out2 = decode(out, img_td.shape, table)
out2 == img_td
```
### Question 5: expérimentations
1. Vérifier, pour les trois images, que la compression huffman fonctionne correctement
2. On définit le taux de compression d'une image par : *1 - (taille_bit_code / taille_bit_image) %* Calculez les taux de compression des trois images. Commentez.
3. Quel serait le taux de compression d'une image avec pixels aléatoires de loi uniforme [0,255] ? Quel est le lien avec la valeur de l'entropie des pixels ?
```
# Réponse questions 5.1 et 5.2
for img, titre in zip([rectangle, couloir, bureau], ["rectangle", "couloir", "bureau"]):
# histograme
N = img.shape[0]*img.shape[1]
hist, _ = np.histogram(img, 256)
hist = hist/N
racine = huffman(hist)
table = hufftable(racine)
out, nbits = encode(img, table)
out2 = decode(out, img.shape, table)
np.testing.assert_array_equal(img, out2) # test de fonctionnement
taux = 1 - (nbits/(8*img.size))
print("Image ", titre, "Taux de compression", taux)
plt.figure(figsize=(10, 10))
plt.title(titre)
plt.imshow(img, cmap='gray')
plt.show()
```
**Réponse question 5.2**
### On remarque que le taux de compression est superieur pour l'image binaire comparé au images réelles
**Réponse question 5.3**
### le taux de compression d'une image avec pixel aléatoire suivant une loi uniforme [0,255] serait = 0 (arbre binaire complet d'hauteur log255 = 8)
| github_jupyter |
# Trax : Ungraded Lecture Notebook
In this notebook you'll get to know about the Trax framework and learn about some of its basic building blocks.
## Background
### Why Trax and not TensorFlow or PyTorch?
TensorFlow and PyTorch are both extensive frameworks that can do almost anything in deep learning. They offer a lot of flexibility, but that often means verbosity of syntax and extra time to code.
Trax is much more concise. It runs on a TensorFlow backend but allows you to train models with 1 line commands. Trax also runs end to end, allowing you to get data, model and train all with a single terse statements. This means you can focus on learning, instead of spending hours on the idiosyncrasies of big framework implementation.
### Why not Keras then?
Keras is now part of Tensorflow itself from 2.0 onwards. Also, trax is good for implementing new state of the art algorithms like Transformers, Reformers, BERT because it is actively maintained by Google Brain Team for advanced deep learning tasks. It runs smoothly on CPUs,GPUs and TPUs as well with comparatively lesser modifications in code.
### How to Code in Trax
Building models in Trax relies on 2 key concepts:- **layers** and **combinators**.
Trax layers are simple objects that process data and perform computations. They can be chained together into composite layers using Trax combinators, allowing you to build layers and models of any complexity.
### Trax, JAX, TensorFlow and Tensor2Tensor
You already know that Trax uses Tensorflow as a backend, but it also uses the JAX library to speed up computation too. You can view JAX as an enhanced and optimized version of numpy.
**Watch out for assignments which import `import trax.fastmath.numpy as np`. If you see this line, remember that when calling `np` you are really calling Trax’s version of numpy that is compatible with JAX.**
As a result of this, where you used to encounter the type `numpy.ndarray` now you will find the type `jax.interpreters.xla.DeviceArray`.
Tensor2Tensor is another name you might have heard. It started as an end to end solution much like how Trax is designed, but it grew unwieldy and complicated. So you can view Trax as the new improved version that operates much faster and simpler.
### Resources
- Trax source code can be found on Github: [Trax](https://github.com/google/trax)
- JAX library: [JAX](https://jax.readthedocs.io/en/latest/index.html)
## Installing Trax
Trax has dependencies on JAX and some libraries like JAX which are yet to be supported in [Windows](https://github.com/google/jax/blob/1bc5896ee4eab5d7bb4ec6f161d8b2abb30557be/README.md#installation) but work well in Ubuntu and MacOS. We would suggest that if you are working on Windows, try to install Trax on WSL2.
Official maintained documentation - [trax-ml](https://trax-ml.readthedocs.io/en/latest/) not to be confused with this [TraX](https://trax.readthedocs.io/en/latest/index.html)
```
#!pip install trax==1.3.1 Use this version for this notebook
```
## Imports
```
import numpy as np # regular ol' numpy
from trax import layers as tl # core building block
from trax import shapes # data signatures: dimensionality and type
from trax import fastmath # uses jax, offers numpy on steroids
# Trax version 1.3.1 or better
!pip list | grep trax
```
## Layers
Layers are the core building blocks in Trax or as mentioned in the lectures, they are the base classes.
They take inputs, compute functions/custom calculations and return outputs.
You can also inspect layer properties. Let me show you some examples.
### Relu Layer
First I'll show you how to build a relu activation function as a layer. A layer like this is one of the simplest types. Notice there is no object initialization so it works just like a math function.
**Note: Activation functions are also layers in Trax, which might look odd if you have been using other frameworks for a longer time.**
```
# Layers
# Create a relu trax layer
relu = tl.Relu()
# Inspect properties
print("-- Properties --")
print("name :", relu.name)
print("expected inputs :", relu.n_in)
print("promised outputs :", relu.n_out, "\n")
# Inputs
x = np.array([-2, -1, 0, 1, 2])
print("-- Inputs --")
print("x :", x, "\n")
# Outputs
y = relu(x)
print("-- Outputs --")
print("y :", y)
```
### Concatenate Layer
Now I'll show you how to build a layer that takes 2 inputs. Notice the change in the expected inputs property from 1 to 2.
```
# Create a concatenate trax layer
concat = tl.Concatenate()
print("-- Properties --")
print("name :", concat.name)
print("expected inputs :", concat.n_in)
print("promised outputs :", concat.n_out, "\n")
# Inputs
x1 = np.array([-10, -20, -30])
x2 = x1 / -10
print("-- Inputs --")
print("x1 :", x1)
print("x2 :", x2, "\n")
# Outputs
y = concat([x1, x2])
print("-- Outputs --")
print("y :", y)
```
## Layers are Configurable
You can change the default settings of layers. For example, you can change the expected inputs for a concatenate layer from 2 to 3 using the optional parameter `n_items`.
```
# Configure a concatenate layer
concat_3 = tl.Concatenate(n_items=3) # configure the layer's expected inputs
print("-- Properties --")
print("name :", concat_3.name)
print("expected inputs :", concat_3.n_in)
print("promised outputs :", concat_3.n_out, "\n")
# Inputs
x1 = np.array([-10, -20, -30])
x2 = x1 / -10
x3 = x2 * 0.99
print("-- Inputs --")
print("x1 :", x1)
print("x2 :", x2)
print("x3 :", x3, "\n")
# Outputs
y = concat_3([x1, x2, x3])
print("-- Outputs --")
print("y :", y)
```
**Note: At any point,if you want to refer the function help/ look up the [documentation](https://trax-ml.readthedocs.io/en/latest/) or use help function.**
```
#help(tl.Concatenate) #Uncomment this to see the function docstring with explaination
```
## Layers can have Weights
Some layer types include mutable weights and biases that are used in computation and training. Layers of this type require initialization before use.
For example the `LayerNorm` layer calculates normalized data, that is also scaled by weights and biases. During initialization you pass the data shape and data type of the inputs, so the layer can initialize compatible arrays of weights and biases.
```
# Uncomment any of them to see information regarding the function
# help(tl.LayerNorm)
# help(shapes.signature)
# Layer initialization
norm = tl.LayerNorm()
# You first must know what the input data will look like
x = np.array([0, 1, 2, 3], dtype="float")
# Use the input data signature to get shape and type for initializing weights and biases
norm.init(shapes.signature(x)) # We need to convert the input datatype from usual tuple to trax ShapeDtype
print("Normal shape:",x.shape, "Data Type:",type(x.shape))
print("Shapes Trax:",shapes.signature(x),"Data Type:",type(shapes.signature(x)))
# Inspect properties
print("-- Properties --")
print("name :", norm.name)
print("expected inputs :", norm.n_in)
print("promised outputs :", norm.n_out)
# Weights and biases
print("weights :", norm.weights[0])
print("biases :", norm.weights[1], "\n")
# Inputs
print("-- Inputs --")
print("x :", x)
# Outputs
y = norm(x)
print("-- Outputs --")
print("y :", y)
```
## Custom Layers
This is where things start getting more interesting!
You can create your own custom layers too and define custom functions for computations by using `tl.Fn`. Let me show you how.
```
help(tl.Fn)
# Define a custom layer
# In this example you will create a layer to calculate the input times 2
def TimesTwo():
layer_name = "TimesTwo" #don't forget to give your custom layer a name to identify
# Custom function for the custom layer
def func(x):
return x * 2
return tl.Fn(layer_name, func)
# Test it
times_two = TimesTwo()
# Inspect properties
print("-- Properties --")
print("name :", times_two.name)
print("expected inputs :", times_two.n_in)
print("promised outputs :", times_two.n_out, "\n")
# Inputs
x = np.array([1, 2, 3])
print("-- Inputs --")
print("x :", x, "\n")
# Outputs
y = times_two(x)
print("-- Outputs --")
print("y :", y)
```
## Combinators
You can combine layers to build more complex layers. Trax provides a set of objects named combinator layers to make this happen. Combinators are themselves layers, so behavior commutes.
### Serial Combinator
This is the most common and easiest to use. For example could build a simple neural network by combining layers into a single layer using the `Serial` combinator. This new layer then acts just like a single layer, so you can inspect intputs, outputs and weights. Or even combine it into another layer! Combinators can then be used as trainable models. _Try adding more layers_
**Note:As you must have guessed, if there is serial combinator, there must be a parallel combinator as well. Do try to explore about combinators and other layers from the trax documentation and look at the repo to understand how these layers are written.**
```
# help(tl.Serial)
# help(tl.Parallel)
# Serial combinator
serial = tl.Serial(
tl.LayerNorm(), # normalize input
tl.Relu(), # convert negative values to zero
times_two, # the custom layer you created above, multiplies the input recieved from above by 2
### START CODE HERE
# tl.Dense(n_units=2), # try adding more layers. eg uncomment these lines
# tl.Dense(n_units=1), # Binary classification, maybe? uncomment at your own peril
# tl.LogSoftmax() # Yes, LogSoftmax is also a layer
### END CODE HERE
)
# Initialization
x = np.array([-2, -1, 0, 1, 2]) #input
serial.init(shapes.signature(x)) #initialising serial instance
print("-- Serial Model --")
print(serial,"\n")
print("-- Properties --")
print("name :", serial.name)
print("sublayers :", serial.sublayers)
print("expected inputs :", serial.n_in)
print("promised outputs :", serial.n_out)
print("weights & biases:", serial.weights, "\n")
# Inputs
print("-- Inputs --")
print("x :", x, "\n")
# Outputs
y = serial(x)
print("-- Outputs --")
print("y :", y)
```
## JAX
Just remember to lookout for which numpy you are using, the regular ol' numpy or Trax's JAX compatible numpy. Both tend to use the alias np so watch those import blocks.
**Note:There are certain things which are still not possible in fastmath.numpy which can be done in numpy so you will see in assignments we will switch between them to get our work done.**
```
# Numpy vs fastmath.numpy have different data types
# Regular ol' numpy
x_numpy = np.array([1, 2, 3])
print("good old numpy : ", type(x_numpy), "\n")
# Fastmath and jax numpy
x_jax = fastmath.numpy.array([1, 2, 3])
print("jax trax numpy : ", type(x_jax))
```
## Summary
Trax is a concise framework, built on TensorFlow, for end to end machine learning. The key building blocks are layers and combinators. This notebook is just a taste, but sets you up with some key inuitions to take forward into the rest of the course and assignments where you will build end to end models.
| github_jupyter |
<a href="https://colab.research.google.com/github/gzc/spark/blob/main/PySpark_Regression_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Running Pyspark in Colab**
To run spark in Colab, we need to first install all the dependencies in Colab environment i.e. Apache Spark 2.3.2 with hadoop 2.7, Java 8 and Findspark to locate the spark in the system. The tools installation can be carried out inside the Jupyter Notebook of the Colab. One important note is that if you are new in Spark, it is better to avoid Spark 2.4.0 version since some people have already complained about its compatibility issue with python.
Follow the steps to install the dependencies:
```
!apt-get install openjdk-8-jdk-headless -qq > /dev/null
!wget -q https://www-us.apache.org/dist/spark/spark-2.4.7/spark-2.4.7-bin-hadoop2.7.tgz
!tar xf /content/spark-2.4.7-bin-hadoop2.7.tgz
!pip install -q findspark
```
Now that you installed Spark and Java in Colab, it is time to set the environment path which enables you to run Pyspark in your Colab environment. Set the location of Java and Spark by running the following code:
```
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.4.7-bin-hadoop2.7"
```
Run a local spark session to test your installation:
```
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
```
Congrats! Your Colab is ready to run Pyspark. Let's build a simple Linear Regression model.
# Linear Regression Model
Linear Regression model is one the oldest and widely used machine learning approach which assumes a relationship between dependent and independent variables. For example, a modeler might want to predict the forecast of the rain based on the humidity ratio. Linear Regression consists of the best fitting line through the scattered points on the graph and the best fitting line is known as the regression line.
The goal of this exercise to predict the housing prices by the given features. Let's predict the prices of the Boston Housing dataset by considering MEDV as the output variable and all the other variables as input.
Download the dataset from [here](https://github.com/gzc/spark/blob/main/BostonHousing.csv) and keep it somewhere on your computer. Load the dataset into your Colab directory from your local system:
```
from google.colab import files
files.upload()
```
Check the dataset is uploaded correctly in the system by the following command
```
!ls
```
Now that we have uploaded the dataset, we can start analyzing.
For our linear regression model we need to import two modules from Pyspark i.e. Vector Assembler and Linear Regression. Vector Assembler is a transformer that assembles all the features into one vector from multiple columns that contain type double. We could have used StringIndexer if any of our columns contains string values to convert it into numeric values. Luckily, the BostonHousing dataset only contains double values, so we don't need to worry about StringIndexer for now.
```
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.regression import LinearRegression
dataset = spark.read.csv('BostonHousing.csv',inferSchema=True, header =True)
```
Notice that we used InferSchema inside read.csv mofule. InferSchema enables us to infer automatically different data types for each column.
Let us print look into the dataset to see the data types of each column:
```
dataset.printSchema()
```
Next step is to convert all the features from different columns into a single column and let's call this new vector column as 'Attributes' in the outputCol.
```
#Input all the features in one vector column
assembler = VectorAssembler(inputCols=['crim', 'zn', 'indus', 'chas', 'nox', 'rm', 'age', 'dis', 'rad', 'tax', 'ptratio', 'b', 'lstat'], outputCol = 'Attributes')
output = assembler.transform(dataset)
#Input vs Output
finalized_data = output.select("Attributes","medv")
finalized_data.show()
```
Here, 'Attributes' are in the input features from all the columns and 'medv' is the target column.
Next, we should split the training and testing data according to our dataset (0.8 and 0.2 in this case).
```
#Split training and testing data
train_data,test_data = finalized_data.randomSplit([0.8,0.2])
regressor = LinearRegression(featuresCol = 'Attributes', labelCol = 'medv')
#Learn to fit the model from training set
regressor = regressor.fit(train_data)
#To predict the prices on testing set
pred = regressor.evaluate(test_data)
#Predict the model
pred.predictions.show()
```
We can also print the coefficient and intercept of the regression model by using the following command:
```
#coefficient of the regression model
coeff = regressor.coefficients
#X and Y intercept
intr = regressor.intercept
print ("The coefficient of the model is : %a" %coeff)
print ("The Intercept of the model is : %f" %intr)
```
# Basic Statistical Analysis
Once we are done with the basic linear regression operation, we can go a bit further and analyze our model statistically by importing RegressionEvaluator module from Pyspark.
```
from pyspark.ml.evaluation import RegressionEvaluator
eval = RegressionEvaluator(labelCol="medv", predictionCol="prediction", metricName="rmse")
# Root Mean Square Error
rmse = eval.evaluate(pred.predictions)
print("RMSE: %.3f" % rmse)
# Mean Square Error
mse = eval.evaluate(pred.predictions, {eval.metricName: "mse"})
print("MSE: %.3f" % mse)
# Mean Absolute Error
mae = eval.evaluate(pred.predictions, {eval.metricName: "mae"})
print("MAE: %.3f" % mae)
# r2 - coefficient of determination
r2 = eval.evaluate(pred.predictions, {eval.metricName: "r2"})
print("r2: %.3f" %r2)
```
| github_jupyter |
## Questionário 42 (Q42)
Orientações:
- Registre suas respostas no questionário de mesmo nome no SIGAA.
- O tempo de registro das respostas no questionário será de 10 minutos. Portanto, resolva primeiro as questões e depois registre-as.
- Haverá apenas 1 (uma) tentativa de resposta.
- Submeta seu arquivo-fonte (utilizado para resolver as questões) em formato _.ipynb_ pelo SIGAA anexando-o à Tarefa denominada "Envio de arquivo" correspondente ao questionário.
*Nota:* o arquivo-fonte será utilizado apenas como prova de execução da tarefa. Nenhuma avaliação será feita quanto ao estilo de programação.
<hr>
**Questão 1.** No _dataset_ [enem2019.xlsx](https://github.com/gcpeixoto/ICD/blob/main/database/enem2019.xlsx), estão disponíveis as notas médias por estado obtidas nas provas do ENEM 2019. Supondo que _x_ é a diferença entre a amplitude da quantidade de inscritos na região Sudeste e a amplitude da quantidade de inscritos na região Norte, e que _y_ é o desvio médio para a série da quantidade total de inscritos de ensino médio público apenas para os estados do sul, assinale a alternativa que corretamente expressa os valores de _x_ e _y_, nesta sequência.
**Obs.:** considere apenas a parte inteira do desvio médio.
A. 149465 e 5690
B. 169265 e 6593
C. 149465 e 0
D. 173921 e 2
## GABARITO
$n =$ **_Quantidade de inscritos da região_**
$\overline{X}$ = **_Média do total de inscritos de Ensino Público_**
$X_i$ = $Estado_i$
Alterei o **n** e o desvio da média; alterei o **n** que deveria ser para evitar que os alunos apenas usem _.mad()_ e realizem de fato o desvio médio
Alternativa **C**
<hr>
```
import pandas as pd
import numpy as np
df = pd.read_excel("../database/enem2019.xlsx")
df = df.set_index("Estado")
df.head()
norte0 = ["AC","AP","AM","PA","RO","RR","TO"]
alvos = ["Quantidade de Inscritos"]
nortedf = df.loc[norte0,alvos]
sudeste0 = ["ES","MG","RJ","SP"]
sudestedf = df.loc[sudeste0,alvos]
R_norte = nortedf.values.max() - nortedf.values.min()
R_sudeste = sudestedf.values.max() - sudestedf.values.min()
R_dif = R_sudeste - R_norte
alvos = ["Quantidade de Inscritos","Total de inscritos de Ensino Médio Público","Total de inscritos de Ensino Médio Privado"]
sul0 = ["PR","RS","SC"]
suldf = df.loc[sul0,alvos]
n = sum(suldf["Quantidade de Inscritos"])
media = suldf["Total de inscritos de Ensino Médio Público"].mean()
DM = sum((suldf["Total de inscritos de Ensino Médio Público"] - media)/n)
f'Resultado esperado: {R_dif} // {DM}'
```
<hr>
**Questão 2.** Calcule o percentual _p_ de inscritos para o ENEM 2019 provenientes do ensino privado de todos os Estados em relação ao total de inscritos no exame, bem como o valor do quociente _v/V_, onde _v_ é a variância para a série do total de inscritos provenientes do ensino público e _V_ a variância para a série do total de inscritos provenientes do ensino privado. Assinale a alternativa correta para _p_ e _v/V_.
A. 11.4% e 34.48
B. 15% e 33.45
C. 12.5% e 36.78
D. 13.54% e 34.6
<hr>
## GABARITO
Alternativa **A**
```
#RESPOSTA CERTA A)
df4 = df.filter(["Quantidade de Inscritos", "Total de inscritos de Ensino Médio Público", "Total de inscritos de Ensino Médio Privado"])
p = (df4["Total de inscritos de Ensino Médio Privado"].sum() / df4["Quantidade de Inscritos"].sum())*100
v = df4["Total de inscritos de Ensino Médio Público"].var() / df4["Total de inscritos de Ensino Médio Privado"].var()
print("{:.2f}% é a porcentagem do total de inscritos no ENEM do Ensino Médio Privado" .format(p))
print("{:.2f} é o quociente da variância do Total de inscritos de Ensino Médio Público sob o Total de inscritos de Ensino Médio Privado" .format(v))
```
<hr>
**Questão 3.** Defina a nota média $N(x)$ de cada região brasileira $x$ como a média das notas $N_i$ de cada uma das $Q$ grandes áreas de conhecimento que constam da prova do ENEM 2019, isto é,
$$N(x) = \frac{ \sum_{i=1}^Q N_i(x)}{Q},$$
e assinale a alternativa cujas regiões detém o primeiro e o segundo maiores valores de desvio padrão.
A. Nordeste e Sudeste
B. Sudeste e Nordeste
C. Norte e Sul
D. Sul e Centro-Oeste
<hr>
## GABARITO
Na verdade é apenas o desvio padrão da nota de cada região.
Alternativa **B**
```
df2 = df.iloc[:,4:]
nordeste0 = ["AL","BA","CE","MA","PB","PE","PI","RN","SE"]
centro_oeste0 = ["GO","MT","MS","DF"]
norte1 = df2.loc[norte0]
sudeste1 = df2.loc[sudeste0]
nordeste1 = df2.loc[nordeste0]
centro_oeste1 = df2.loc[centro_oeste0]
sul1 = df2.loc[sul0]
df3 = pd.DataFrame({"Nordeste": nordeste1.mean(), "Sudeste":sudeste1.mean(), "Centro-Oeste":centro_oeste1.mean(),
"Norte": norte1.mean(), "Sul":sul1.mean()},index=nordeste1.mean().index)
df3.head()
df4 = pd.concat([df3.mean(),df3.std()], axis=1)
df4.columns=['Média', 'Desvio padrão']
df4.sort_values(by='Desvio padrão',ascending=False)
```
| github_jupyter |
**Exercise set 4**
==============
>The goal of this exercise is to perform **least-squares regression** and to see how we can estimate errors in the parameters we find.
**Exercise 4.1**
In this exercise we will use least-squares regression to investigate a physical phenomenon: the decay of beer froth with time. The file [erdinger.txt](Data/erdinger.txt) (located at 'Data/erdinger.txt') contains [measured heights](https://doi.org/10.1088/0143-0807/23/1/304) for beer froth as a function of time, along with the errors in the measured heights.
**(a)** Use least-squares regression to create a linear model that predicts the beer froth height as a function of time. Plot your linear model together with the raw data.
```
# Your code here
```
**Your answer to question 4.1(a):** *Double click here*
**(b)** Obtain the [coefficient of determination](https://en.wikipedia.org/wiki/Coefficient_of_determination), $R^2$, for your model.
```
# Your code here
```
**Your answer to question 4.1(b):** *Double click here*
**(c)** It is reasonable to assume that the change in the volume of froth is proportional
to the volume present at a given time. One can show that this leads
to exponential decay,
\begin{equation}
h(t) = h(0) \exp \left(-\frac{t}{\tau} \right),
\label{eq:hard}
\tag{1}\end{equation}
where $h(t)$ is the height of the froth as a function of time $t$, and $\tau$ is a parameter.
In the following, consider $h(0)$ as an unknown parameter to be determined. Show
how you can transform the equation above to a linear equation of the form,
\begin{equation}
y = a + b x,
\tag{2}\end{equation}
and give the relation(s) between the variables $h, h(0), t, \tau$ and
$a, b, x, y$.
**Your answer to question 4.1(c):** *Double click here*
**(d)** Using the transformation found above, create a new linear model, and estimate $h(0)$ and $\tau$.
Further, calculate the coefficient of determination for this case, and compare the two
linear models you have found so far.
```
# Your code here
```
**Your answer to question 4.1(d):** *Double click here*
**(e)** From the analytical model Eq. (1), $h(0)$ is a known constant, equal to the height of
the froth at time zero. Thus, we can reformulate our model and fit it to just obtain
one parameter, $b$. Essentially, we are defining $y^\prime = y - a$ and using the model,
\begin{equation}
y^\prime = y - a = b x,
\tag{3}\end{equation}
that is, we have a linear model *without* the constant term.
Show that the least-squares solution for $b$ when fitting $y^\prime = bx$ is given by,
\begin{equation}
b = \frac{
\sum_{i=1}^n y_i^\prime x_i
}{\sum_{i=1}^n x_i^2
},
\label{eq:bexpr}
\tag{4}\end{equation}
where $n$ is the number of measurements and $x_i$ and $y_i^\prime$ are the
measured values.
**Your answer to question 4.1(e):** *Double click here*
**(f)** Do the fitting a final time, but use Eq. (4)
to obtain the parameter $b$.
Calculate the coefficient of determination and compare the three linear models you have found.
```
# Your code here
```
**Your answer to question 4.1(f):** *Double click here*
**Exercise 4.2**
In this exercise, we will consider a linear model where we have one variable:
\begin{equation}
y = a + bx,
\end{equation}
and we have determined $a$ and $b$ using the least-squares equations. We further have
$n$ data points $(x_1, y_1), (x_2, y_2), \ldots, (x_n, y_n)$ where the $x_i$'s do not have
any uncertainty, while the uncertainty in the $y_i$'s are all equal to $\sigma_y$.
> Our goal here is to find expressions for estimating
> the errors in the parameters $a$ and $b$,
> given the error in our measurements of $y$.
**Background information: Propagation of errors**
To be able to estimate the errors in $a$ and $b$, we will use [propagation of errors](https://en.wikipedia.org/wiki/Propagation_of_uncertainty).
For simplicity, consider a function, $f$, of two variables $u$ and $v$: $f = f(u, v)$.
By doing a Taylor expansion about the average values, $\bar{u}$ and $\bar{v}$, we can
show that the uncertainty (or "error") in the function $f$, $\sigma_f$, due to the uncertainties in $u$
and $v$ ($\sigma_u$ and $\sigma_v$, respectively) is given by:
\begin{equation}
\sigma_f^2 = \left(\frac{\partial f}{\partial u} \right)^2 \sigma_u^2 +
\left(\frac{\partial f}{\partial v} \right)^2 \sigma_v^2 +
2 \frac{\partial f}{\partial u} \frac{\partial f}{\partial v} \sigma_{uv} + \text{higher-order terms},
\end{equation}
where $\sigma_{uv}$ is the *covariance* between $u$ and $v$. Typically, the errors are "small"
and this motivates us to neglect the higher-order terms. Further, we will assume that the
variables $u$ and $v$ are *not* correlated: $\sigma_{uv} = 0$. We then arrive at the
(perhaps well-known) approximate propagation-of-errors-expression for the uncertainty in $f$:
\begin{equation}
\sigma_f^2 \approx \left(\frac{\partial f}{\partial u} \right)^2 \sigma_u^2 +
\left(\frac{\partial f}{\partial v} \right)^2 \sigma_v^2 .
\end{equation}
This can be generalized to $k$ variables, say $f=f(z_1, z_2, \ldots, z_k)$. The approximate
expression for the uncertainty in $f$, $\sigma_f$, due to the uncertainties
in the $z_i$'s, $\sigma_{z_{i}}$, is then:
\begin{equation}
\sigma_f^2 \approx \sum_{i=1}^{k} \left(\frac{\partial f}{\partial z_{i}} \right)^2 \sigma_{z_{i}}^2 .
\label{eq:errorp}
\tag{5}\end{equation}
We will use this expression to estimate the uncertainties in $a$ and $b$.
**Deriving expressions for the uncertainties in $a$ and $b$**
**(a)** Show that the error in the $b$ parameter, $\sigma_b$,
is given by the following expression:
\begin{equation}
\sigma_b^2 = \frac{\sigma_y^2}{\sum_{i=1}^n \left(x_i - \bar{x}\right)^2},
\end{equation}
where $\bar{x} = \frac{1}{n} \sum_{i=1}^{n} x_i$ is the average of $x$.
***Hint:*** Use the least-squares expression for $b$:
\begin{equation}
b = \frac{
\sum_{i=1}^n (x_i - \bar{x}) (y_i - \bar{y})
}{
\sum_{i=1}^n (x_i - \bar{x})^2
},
\end{equation}
together with the propagation-of-errors expression (Eq. (5)), and consider $b$ as a
function of the $y_i$'s: $b = f(y_1, y_2, \ldots, y_n)$. You might find it helpful to determine
$\frac{\partial b}{\partial y_j}$
as an intermediate step in your derivation.
**Your answer to question 4.2(a):** *Double click here*
**(b)** Show that the error in the $a$ parameter, $\sigma_a$, is given by the following expression:
\begin{equation}
\sigma_a^2 = \frac{\sigma_y^2}{n} \times
\frac{
\sum_{i=1}^{n} x_i^2
}{
\sum_{i=1}^{n} (x_i - \bar{x})^2
} .
\end{equation}
***Hint:*** Use the least-squares expression for $a$:
\begin{equation}
a = \bar{y} - b \bar{x},
\end{equation}
together with the propagation-of-errors expression (Eq. (5)), and consider $a$ as a
function of the $y_i$'s *and* $b$: $a = f(y_1, y_2, \ldots, y_n,b)$. You might find it
helpful to determine
$\frac{\partial a}{\partial y_j}$ and $\frac{\partial a}{\partial b}$ as intermediate steps
in your derivation.
**Your answer to question 4.2(b):** *Double click here*
| github_jupyter |
The purpose of this notebook is to read the full enzyme database into a notebook, and to begin to explore promiscuous enzymes, and how to pull data on linked reactions
```
# imports
from Bio.KEGG import REST
from Bio.KEGG import Enzyme
from Bio.KEGG import Compound
import gzip
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import re
# function to read data directly from the text file instead
def enzyme_records_to_df(file_path):
"""
Input should be a filepath string pointing to a gzipped text file of KEGG enzyme records.
Function parses all records using Biopython.Bio.KEGG.Enzyme parser, and returns a pandas dataframe.
"""
enzyme_fields = [method for method in dir(Enzyme.Record()) if not method.startswith('_')]
data_matrix = []
with gzip.open(file_path, 'rt') as file:
for record in Enzyme.parse(file):
data_matrix.append([getattr(record, field) for field in enzyme_fields])
enzyme_df = pd.DataFrame(data_matrix, columns=enzyme_fields)
return enzyme_df
# read in enzyme data
enzyme_df = enzyme_records_to_df('../metamoles/tests/test_kegg_enzyme_records.txt.gz')
enzyme_df.columns == ['classname', 'cofactor', 'comment', 'dblinks', 'disease', 'effector',
'entry', 'genes', 'inhibitor', 'name', 'pathway', 'product', 'reaction',
'structures', 'substrate', 'sysname']
enzyme_df['entry'].tolist() == ['1.1.1.1', '1.1.1.2', '1.1.1.3']
enzyme_df.head()
# apply a boolean mask to the dataframe to select only rows in which the number of reaction entries is > 1
promiscuous_df = enzyme_df[[True if len(rxn) > 1 else False for rxn in enzyme_df['reaction']]]
# how many promiscuous enzymes are there?
promiscuous_df.shape
# histogram of total reactions per enzyme EC category
reactions = [len(rxn) for rxn in promiscuous_df['reaction']]
hist = sns.distplot(reactions, bins=20, kde=False, norm_hist=False)
# example product field
prod_example = enzyme_df.iloc[0]['product']
prod_example
# test pull CPD record from KEGG
# note: this doesn't appear to work if CPD is in caps
cpd_entry_71 = (REST.kegg_get('cpd:c00071').read())
cpd_entry_71
# fiddle with regex to get CPD number
prod_example.append('[cpd:c00071]')
regex = '\[CPD:(C\d{5})]'
print(re.findall(regex, str(prod_example), re.IGNORECASE))
prod_example
# function that extracts list of KEGG compound IDs from product or substrate field of enzyme entry
# regex reference: https://regex101.com/
# changed this to leave out the brackets
def extract_KEGG_compound_IDs(field):
"""
This function uses regular expressions to extract the KEGG compound IDs from a product or substrate field
in an enzyme record (or other KEGG record)
"""
cpd_list = []
regex = 'CPD:(C\d+)' # matches '[CPD: ' characters exactly, captures 'C' + any number of digits (\d+), befor another literal ']' character
for entry in field:
ids = re.findall(regex, str(entry), re.IGNORECASE)
for i in ids:
cpd_list.append(i)
return cpd_list
enzyme_df.head()
l = extract_KEGG_compound_IDs(enzyme_df['product'])
print(l)
# # test out KEGG compound extraction function
# # this works but takes a very long time
# test_df = promiscuous_df[:5]
# test_df['compound_ids'] = test_df['product'].apply(extract_KEGG_compound_IDs)
# test_df
# try extracting entry, classname, and name for each product compound id, and creating new dataframe
def explode_dataframe(dataframe, explosion_function, explosion_target_field, fields_to_include):
"""
This function applies the input explosion_function to the target field in each row of a dataframe.
Each item in the output of the explosion_function is an anchor for a new row in the new dataframe.
All of the supplied fields_to_include are added to the explosion item, and appended to the new dataframe row.
dataframe,
explosion_function,
explosion_target_field,
fields_to_include
"""
new_rows = []
for _, row in dataframe.iterrows():
explosion_list = explosion_function(row[explosion_target_field])
for item in explosion_list:
row_data = [row[field] for field in fields_to_include]
row_data.append(item)
new_rows.append(row_data)
fields_to_include.append(explosion_target_field)
new_df = pd.DataFrame(new_rows, columns=fields_to_include)
return new_df
# test explode_dataframe function
exploded_df = explode_dataframe(enzyme_df, extract_KEGG_compound_IDs, 'product', ['entry'])
exploded_df.shape
print(exploded_df['product'].tolist())
# run explode dataframe function on full promiscuous enzyme dataframe
expanded_prom_enzymes_df = explode_dataframe(promiscuous_df, extract_KEGG_compound_IDs, 'product', ['entry'])
# looks like a total of 1878 product compound/enzyme pairs
expanded_prom_enzymes_df.head()
# no empty cells - this is good
expanded_prom_enzymes_df.isnull().sum()
```
## Starting work here
```
expanded_prom_enzymes_df.shape
#cofactor removal
cofactor_df = pd.read_csv("../datasets/cofactor_list.csv")
cofactor_list = extract_KEGG_compound_IDs(cofactor_df['CPD'])
len(cofactor_list)
# remove rows that contain cofactors
bool_mask = [False if cpd in cofactor_list else True for cpd in expanded_prom_enzymes_df['product']]
clean_enzyme_df = expanded_prom_enzymes_df[bool_mask]
clean_enzyme_df.shape
# one duplicate enzyme - compound entry
deduplicated = clean_enzyme_df.drop_duplicates()
deduplicated.shape
clean_enzyme_df.head()
# check PubChem id number lengths & for empty cells
# looks like there are some empty fields, as well as some longer id numbers
# we should look into these more, but for now I will charge ahead
def check_length_dist(dataframe, field):
id_lengths = [len(data) for data in dataframe[field]]
values, counts = np.unique(id_lengths, return_counts=True)
for i in range(len(values)):
print("{}: {}".format(values[i], counts[i]))
print("\ntotal numbers: {}\ncdataframe shape: {}".format(counts.sum(), dataframe.shape))
check_length_dist(clean_enzyme_df, 'entry')
# function to create two dataframes - one of positive data (enzyme - compound pairs that exist in the dataset)
# and one of negative data (all enzyme - compound pairs that do not exist in the dataset)
def neg_data_matchmaker(dataframe, enzyme_field, compound_field):
"""neg_data_matchmaker creates two dataframes. One dataframe is positive data that contains all the
enzyme-compound pairs that exist in the dataset. The second data frame is negative data made from matching
all enzyme-compound pairs that do not exist in the dataset.
Inputs
dataframe: pandas dataframe
enzyme_field: string identifier denoting which column contains enzyme ids
compound_field: string identifier denoting which column contains compound ids
Outputs
positive_df: pandas dataframe with fields 'enzyme', 'product', 'reacts'
negative_df: pandas dataframe with fields 'enzyme', 'product', 'reacts'
"""
unique_enzymes = set(dataframe[enzyme_field].unique())
# set of all unique enzymes in provided dataframes
unique_cpds = set(dataframe[compound_field].unique())
# set of all unique compounds in provided dataframe
positive_data = []
negative_data = []
# initialize empty lists
for enzyme in unique_enzymes:
# iterate through unique enzyme set
working_prods = set(dataframe[dataframe[enzyme_field] == enzyme][compound_field].unique())
# unique set of all products reported to reaction with this enzyme in provided dataset
non_working_prods = (unique_cpds - working_prods)
# set math of all remaining products in the dataset minus those reported to react
reactions = [{'reacts':1.0, 'enzyme':enzyme, 'product':product} for product in working_prods]
# create new entry for each positive reaction
non_reactions = [{'reacts':0.0, 'enzyme':enzyme, 'product':product} for product in non_working_prods]
# create new entry for each negative reaction
positive_data.extend(reactions)
# add positive reactions to master list
negative_data.extend(non_reactions)
# add negative reactions to master list
positive_df = pd.DataFrame(positive_data)
negative_df = pd.DataFrame(negative_data)
return positive_df, negative_df
pos_df, neg_df = neg_data_matchmaker(exploded_df, 'entry', 'product')
pos_df.shape
neg_df.shape
print(neg_df['product'].tolist())
unique_cpds.remove('C00022')
len(unique_cpds)
# function to read compound data from text file directly into dataframe
def compound_records_to_df(file_path):
"""
Input should be a filepath string pointing to a gzipped text file of KEGG enzyme records.
Function parses all records using Biopython.Bio.KEGG.Compound parser, and returns a pandas dataframe.
"""
compound_fields = [method for method in dir(Compound.Record()) if not method.startswith('_')]
data_matrix = []
with gzip.open(file_path, 'rt') as file:
for record in Compound.parse(file):
data_matrix.append([getattr(record, field) for field in compound_fields])
compound_df = pd.DataFrame(data_matrix, columns=compound_fields)
return compound_df
!ls ../metamoles/tests/
# read in compound data
compound_df = compound_records_to_df('../metamoles/tests/test_kegg_compound_records.txt.gz')
# looks like there are 18505 entries
compound_df.shape
compound_df.columns
print(compound_df['entry'].tolist())
# function to extract PubChem compound ID:
def extract_PubChem_id(field):
"""
This function uses regular expressions to extract the PubChem compound IDs from a field in a record
"""
regex = "'PubChem', \[\'(\d+)\'\]\)" # matches "'PubChem', ['" characters exactly, then captures any number of digits (\d+), before another literal "']" character match
ids = re.findall(regex, str(field), re.IGNORECASE)
if len(ids) > 0:
pubchem_id = ids[0]
else:
pubchem_id = ''
return pubchem_id
# extract PubChem compound IDs to a new field in compound dataframe
PubChemID_list = []
for _, row in compound_df.iterrows():
pubchem_id = extract_PubChem_id(row['dblinks'])
PubChemID_list.append(pubchem_id)
compound_df['PubChem'] = PubChemID_list
print(PubChemID_list)
# check PubChem id number lengths & for empty cells
# looks like there are some empty fields, as well as some longer id numbers
# we should look into these more, but for now I will charge ahead
id_lengths = [len(field) for field in compound_df['PubChem']]
values, counts = np.unique(id_lengths, return_counts=True)
for i in range(len(values)):
print("{}: {}".format(values[i], counts[i]))
print("\ntotal numbers: {}\ncompound_df shape: {}".format(counts.sum(), compound_df.shape))
# final step: join compound dataframe onto enzyme dataframe
joint_enzyme_compound_df = expanded_prom_enzymes_df.merge(compound_df, left_on='product', right_on='entry')
joint_enzyme_compound_df.head()
# clean up columns, etc
joint_enzyme_compound_df.drop(['entry_y', 'enzyme', 'pathway', 'structures'], axis=1, inplace=True)
# rename columns
new_col_names = ['ec_number', 'enzyme_name', 'product_cpd', 'dblinks', 'product_formula', 'product_mass', 'product_name', 'pubchem_id']
new_col_dict = dict(zip(joint_enzyme_compound_df.columns, new_col_names))
joint_enzyme_compound_df.rename(columns=new_col_dict, inplace=True)
joint_enzyme_compound_df.sort_values(by=['ec_number']).head()
# check PubChem id number lengths & empty fields
id_lengths = [len(field) for field in joint_enzyme_compound_df['pubchem_id']]
values, counts = np.unique(id_lengths, return_counts=True)
for i in range(len(values)):
print("{}: {}".format(values[i], counts[i]))
print("\ntotal numbers: {}\ncompound_df shape: {}".format(counts.sum(), joint_enzyme_compound_df.shape))
# examine 0 fields
# looks like these entries don't have a dblink for PubChem
# may be able to deal with these later, but for now we'll just have to move on
bool_mask = [True if length == 0 else False for length in id_lengths]
empty_pubchems = joint_enzyme_compound_df[bool_mask]
empty_pubchems
# write to csv file
joint_enzyme_compound_df.to_csv('pubchem_ids_promiscuous_enzyme_products.csv')
```
| github_jupyter |
[Index](Index.ipynb) - [Back](Widget Basics.ipynb) - [Next](Output Widget.ipynb)
# Widget List
```
import ipywidgets as widgets
```
## Numeric widgets
There are many widgets distributed with ipywidgets that are designed to display numeric values. Widgets exist for displaying integers and floats, both bounded and unbounded. The integer widgets share a similar naming scheme to their floating point counterparts. By replacing `Float` with `Int` in the widget name, you can find the Integer equivalent.
### IntSlider
- The slider is displayed with a specified, initial `value`. Lower and upper bounds are defined by `min` and `max`, and the value can be incremented according to the `step` parameter.
- The slider's label is defined by `description` parameter
- The slider's `orientation` is either 'horizontal' (default) or 'vertical'
- `readout` displays the current value of the slider next to it. The options are **True** (default) or **False**
- `readout_format` specifies the format function used to represent slider value. The default is '.2f'
```
widgets.IntSlider(
value=7,
min=0,
max=10,
step=1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
```
### FloatSlider
```
widgets.FloatSlider(
value=7.5,
min=0,
max=10.0,
step=0.1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
```
An example of sliders **displayed vertically**.
```
widgets.FloatSlider(
value=7.5,
min=0,
max=10.0,
step=0.1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='vertical',
readout=True,
readout_format='.1f',
)
```
### FloatLogSlider
The `FloatLogSlider` has a log scale, which makes it easy to have a slider that covers a wide range of positive magnitudes. The `min` and `max` refer to the minimum and maximum exponents of the `base`, and the `value` refers to the actual value of the slider.
```
widgets.FloatLogSlider(
value=10,
base=10,
min=-10, # max exponent of base
max=10, # min exponent of base
step=0.2, # exponent step
description='Log Slider'
)
```
### IntRangeSlider
```
widgets.IntRangeSlider(
value=[5, 7],
min=0,
max=10,
step=1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d',
)
```
### FloatRangeSlider
```
widgets.FloatRangeSlider(
value=[5, 7.5],
min=0,
max=10.0,
step=0.1,
description='Test:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
```
### IntProgress
```
widgets.IntProgress(
value=7,
min=0,
max=10,
description='Loading:',
bar_style='', # 'success', 'info', 'warning', 'danger' or ''
style={'bar_color': 'maroon'},
orientation='horizontal'
)
```
### FloatProgress
```
widgets.FloatProgress(
value=7.5,
min=0,
max=10.0,
description='Loading:',
bar_style='info',
style={'bar_color': '#ffff00'},
orientation='horizontal'
)
```
The numerical text boxes that impose some limit on the data (range, integer-only) impose that restriction when the user presses enter.
### BoundedIntText
```
widgets.BoundedIntText(
value=7,
min=0,
max=10,
step=1,
description='Text:',
disabled=False
)
```
### BoundedFloatText
```
widgets.BoundedFloatText(
value=7.5,
min=0,
max=10.0,
step=0.1,
description='Text:',
disabled=False
)
```
### IntText
```
widgets.IntText(
value=7,
description='Any:',
disabled=False
)
```
### FloatText
```
widgets.FloatText(
value=7.5,
description='Any:',
disabled=False
)
```
## Boolean widgets
There are three widgets that are designed to display a boolean value.
### ToggleButton
```
widgets.ToggleButton(
value=False,
description='Click me',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Description',
icon='check' # (FontAwesome names without the `fa-` prefix)
)
```
### Checkbox
- `value` specifies the value of the checkbox
- `indent` parameter places an indented checkbox, aligned with other controls. Options are **True** (default) or **False**
```
widgets.Checkbox(
value=False,
description='Check me',
disabled=False,
indent=False
)
```
### Valid
The valid widget provides a read-only indicator.
```
widgets.Valid(
value=False,
description='Valid!',
)
```
## Selection widgets
There are several widgets that can be used to display single selection lists, and two that can be used to select multiple values. All inherit from the same base class. You can specify the **enumeration of selectable options by passing a list** (options are either (label, value) pairs, or simply values for which the labels are derived by calling `str`).
<div class="alert alert-info">
Changes in *ipywidgets 8*:
Selection widgets no longer accept a dictionary of options. Pass a list of key-value pairs instead.
</div>
### Dropdown
```
widgets.Dropdown(
options=['1', '2', '3'],
value='2',
description='Number:',
disabled=False,
)
```
The following is also valid, displaying the words `'One', 'Two', 'Three'` as the dropdown choices but returning the values `1, 2, 3`.
```
widgets.Dropdown(
options=[('One', 1), ('Two', 2), ('Three', 3)],
value=2,
description='Number:',
)
```
### RadioButtons
```
widgets.RadioButtons(
options=['pepperoni', 'pineapple', 'anchovies'],
# value='pineapple', # Defaults to 'pineapple'
# layout={'width': 'max-content'}, # If the items' names are long
description='Pizza topping:',
disabled=False
)
```
#### With dynamic layout and very long labels
```
widgets.Box(
[
widgets.Label(value='Pizza topping with a very long label:'),
widgets.RadioButtons(
options=[
'pepperoni',
'pineapple',
'anchovies',
'and the long name that will fit fine and the long name that will fit fine and the long name that will fit fine '
],
layout={'width': 'max-content'}
)
]
)
```
### Select
```
widgets.Select(
options=['Linux', 'Windows', 'macOS'],
value='macOS',
# rows=10,
description='OS:',
disabled=False
)
```
### SelectionSlider
```
widgets.SelectionSlider(
options=['scrambled', 'sunny side up', 'poached', 'over easy'],
value='sunny side up',
description='I like my eggs ...',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True
)
```
### SelectionRangeSlider
The value, index, and label keys are 2-tuples of the min and max values selected. The options must be nonempty.
```
import datetime
dates = [datetime.date(2015, i, 1) for i in range(1, 13)]
options = [(i.strftime('%b'), i) for i in dates]
widgets.SelectionRangeSlider(
options=options,
index=(0, 11),
description='Months (2015)',
disabled=False
)
```
### ToggleButtons
```
widgets.ToggleButtons(
options=['Slow', 'Regular', 'Fast'],
description='Speed:',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltips=['Description of slow', 'Description of regular', 'Description of fast'],
# icons=['check'] * 3
)
```
### SelectMultiple
Multiple values can be selected with <kbd>shift</kbd> and/or <kbd>ctrl</kbd> (or <kbd>command</kbd>) pressed and mouse clicks or arrow keys.
```
widgets.SelectMultiple(
options=['Apples', 'Oranges', 'Pears'],
value=['Oranges'],
#rows=10,
description='Fruits',
disabled=False
)
```
## String widgets
There are several widgets that can be used to display a string value. The `Text`, `Textarea`, and `Combobox` widgets accept input. The `HTML` and `HTMLMath` widgets display a string as HTML (`HTMLMath` also renders math). The `Label` widget can be used to construct a custom control label.
### Text
```
widgets.Text(
value='Hello World',
placeholder='Type something',
description='String:',
disabled=False
)
```
### Textarea
```
widgets.Textarea(
value='Hello World',
placeholder='Type something',
description='String:',
disabled=False
)
```
### Combobox
```
widgets.Combobox(
# value='John',
placeholder='Choose Someone',
options=['Paul', 'John', 'George', 'Ringo'],
description='Combobox:',
ensure_option=True,
disabled=False
)
```
### Password
The `Password` widget hides user input on the screen. **This widget is not a secure way to collect sensitive information because:**
+ The contents of the `Password` widget are transmitted unencrypted.
+ If the widget state is saved in the notebook the contents of the `Password` widget is stored as plain text.
```
widgets.Password(
value='password',
placeholder='Enter password',
description='Password:',
disabled=False
)
```
### Label
The `Label` widget is useful if you need to build a custom description next to a control using similar styling to the built-in control descriptions.
```
widgets.HBox([widgets.Label(value="The $m$ in $E=mc^2$:"), widgets.FloatSlider()])
```
### HTML
```
widgets.HTML(
value="Hello <b>World</b>",
placeholder='Some HTML',
description='Some HTML',
)
```
### HTML Math
```
widgets.HTMLMath(
value=r"Some math and <i>HTML</i>: \(x^2\) and $$\frac{x+1}{x-1}$$",
placeholder='Some HTML',
description='Some HTML',
)
```
## Image
```
file = open("images/WidgetArch.png", "rb")
image = file.read()
widgets.Image(
value=image,
format='png',
width=300,
height=400,
)
```
## Button
```
button = widgets.Button(
description='Click me',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Click me',
icon='check' # (FontAwesome names without the `fa-` prefix)
)
button
```
The `icon` attribute can be used to define an icon; see the [fontawesome](https://fontawesome.com/icons) page for available icons.
A callback function `foo` can be registered using `button.on_click(foo)`. The function `foo` will be called when the button is clicked with the button instance as its single argument.
## Output
The `Output` widget can capture and display stdout, stderr and [rich output generated by IPython](http://ipython.readthedocs.io/en/stable/api/generated/IPython.display.html#module-IPython.display). For detailed documentation, see the [output widget examples](https://ipywidgets.readthedocs.io/en/latest/examples/Output Widget.html).
## Play (Animation) widget
The `Play` widget is useful to perform animations by iterating on a sequence of integers with a certain speed. The value of the slider below is linked to the player.
```
play = widgets.Play(
value=50,
min=0,
max=100,
step=1,
interval=500,
description="Press play",
disabled=False
)
slider = widgets.IntSlider()
widgets.jslink((play, 'value'), (slider, 'value'))
widgets.HBox([play, slider])
```
## Date picker
For a list of browsers that support the date picker widget, see the [MDN article for the HTML date input field](https://developer.mozilla.org/en-US/docs/Web/HTML/Element/input/date#Browser_compatibility).
```
widgets.DatePicker(
description='Pick a Date',
disabled=False
)
```
## Time picker
For a list of browsers that support the time picker widget, see the [MDN article for the HTML time input field](https://developer.mozilla.org/en-US/docs/Web/HTML/Element/input/time#Browser_compatibility).
```
widgets.TimePicker(
description='Pick a Time',
disabled=False
)
```
## Datetime picker
For a list of browsers that support the datetime picker widget, see the [MDN article for the HTML datetime-local input field](https://developer.mozilla.org/en-US/docs/Web/HTML/Element/input/datetime-local#Browser_compatibility). For the browsers that do not support the datetime-local input, we try to fall back on displaying separate date and time inputs.
### Time zones
There are two points worth to note with regards to timezones for datetimes:
- The browser always picks datetimes using *its* timezone.
- The kernel always gets the datetimes in the default system timezone of the kernel (see https://docs.python.org/3/library/datetime.html#datetime.datetime.astimezone with `None` as the argument).
This means that if the kernel and browser have different timezones, the default string serialization of the timezones might differ, but they will still represent the same point in time.
```
widgets.DatetimePicker(
description='Pick a Time',
disabled=False
)
```
## Color picker
```
widgets.ColorPicker(
concise=False,
description='Pick a color',
value='blue',
disabled=False
)
```
## File Upload
The `FileUpload` allows to upload any type of file(s) into memory in the kernel.
```
widgets.FileUpload(
accept='', # Accepted file extension e.g. '.txt', '.pdf', 'image/*', 'image/*,.pdf'
multiple=False # True to accept multiple files upload else False
)
```
The upload widget exposes a `value` attribute that contains the files uploaded. The value attribute is a tuple with a dictionary for each uploaded file. For instance:
```python
uploader = widgets.FileUpload()
display(uploader)
# upload something...
# once a file is uploaded, use the `.value` attribute to retrieve the content:
uploader.value
#=> (
#=> {
#=> 'name': 'example.txt',
#=> 'type': 'text/plain',
#=> 'size': 36,
#=> 'last_modified': datetime.datetime(2020, 1, 9, 15, 58, 43, 321000, tzinfo=datetime.timezone.utc),
#=> 'content': <memory at 0x10c1b37c8>
#=> },
#=> )
```
Entries in the dictionary can be accessed either as items, as one would any dictionary, or as attributes:
```
uploaded_file = uploader.value[0]
uploaded_file["size"]
#=> 36
uploaded_file.size
#=> 36
```
The contents of the file uploaded are in the value of the `content` key. They are a [memory view](https://docs.python.org/3/library/stdtypes.html#memory-views):
```python
uploaded_file.content
#=> <memory at 0x10c1b37c8>
```
You can extract the content to bytes:
```python
uploaded_file.content.tobytes()
#=> b'This is the content of example.txt.\n'
```
If the file is a text file, you can get the contents as a string by [decoding it](https://docs.python.org/3/library/codecs.html):
```python
import codecs
codecs.decode(uploaded_file.content, encoding="utf-8")
#=> 'This is the content of example.txt.\n'
```
You can save the uploaded file to the filesystem from the kernel:
```python
with open("./saved-output.txt", "wb") as fp:
fp.write(uploaded_file.content)
```
To convert the uploaded file into a Pandas dataframe, you can use a [BytesIO object](https://docs.python.org/3/library/io.html#binary-i-o):
```python
import io
import pandas as pd
pd.read_csv(io.BytesIO(uploaded_file.content))
```
If the uploaded file is an image, you can visualize it with an [image](#Image) widget:
```python
widgets.Image(value=uploaded_file.content.tobytes())
```
<div class="alert alert-info">
Changes in *ipywidgets 8*:
The `FileUpload` changed significantly in ipywidgets 8:
- The `.value` traitlet is now a list of dictionaries, rather than a dictionary mapping the uploaded name to the content. To retrieve the original form, use `{f["name"]: f.content.tobytes() for f in uploader.value}`.
- The `.data` traitlet has been removed. To retrieve it, use `[f.content.tobytes() for f in uploader.value]`.
- The `.metadata` traitlet has been removed. To retrieve it, use `[{k: v for k, v in f.items() if k != "content"} for f in w.value]`.
</div>
<div class="alert alert-warning">
Warning: When using the `FileUpload` Widget, uploaded file content might be saved in the notebook if widget state is saved.
</div>
## Controller
The `Controller` allows a game controller to be used as an input device.
```
widgets.Controller(
index=0,
)
```
## Container/Layout widgets
These widgets are used to hold other widgets, called children. Each has a `children` property that may be set either when the widget is created or later.
### Box
```
items = [widgets.Label(str(i)) for i in range(4)]
widgets.Box(items)
```
### HBox
```
items = [widgets.Label(str(i)) for i in range(4)]
widgets.HBox(items)
```
### VBox
```
items = [widgets.Label(str(i)) for i in range(4)]
left_box = widgets.VBox([items[0], items[1]])
right_box = widgets.VBox([items[2], items[3]])
widgets.HBox([left_box, right_box])
```
### GridBox
This box uses the HTML Grid specification to lay out its children in two dimensional grid. The example below lays out the 8 items inside in 3 columns and as many rows as needed to accommodate the items.
```
items = [widgets.Label(str(i)) for i in range(8)]
widgets.GridBox(items, layout=widgets.Layout(grid_template_columns="repeat(3, 100px)"))
```
### Accordion
```
accordion = widgets.Accordion(children=[widgets.IntSlider(), widgets.Text()], titles=('Slider', 'Text'))
accordion
```
### Tabs
In this example the children are set after the tab is created. Titles for the tabs are set in the same way they are for `Accordion`.
```
tab_contents = ['P0', 'P1', 'P2', 'P3', 'P4']
children = [widgets.Text(description=name) for name in tab_contents]
tab = widgets.Tab()
tab.children = children
tab.titles = [str(i) for i in range(len(children))]
tab
```
### Stacked
The `Stacked` widget can have multiple children widgets as for `Tab` and `Accordion`, but only shows one at a time depending on the value of ``selected_index``:
```
button = widgets.Button(description='Click here')
slider = widgets.IntSlider()
stacked = widgets.Stacked([button, slider])
stacked # will show only the button
```
This can be used in combination with another selection-based widget to show different widgets depending on the selection:
```
dropdown = widgets.Dropdown(options=['button', 'slider'])
widgets.jslink((dropdown, 'index'), (stacked, 'selected_index'))
widgets.VBox([dropdown, stacked])
```
### Accordion, Tab, and Stacked use `selected_index`, not value
Unlike the rest of the widgets discussed earlier, the container widgets `Accordion` and `Tab` update their `selected_index` attribute when the user changes which accordion or tab is selected. That means that you can both see what the user is doing *and* programmatically set what the user sees by setting the value of `selected_index`.
Setting `selected_index = None` closes all of the accordions or deselects all tabs.
In the cells below try displaying or setting the `selected_index` of the `tab` and/or `accordion`.
```
tab.selected_index = 3
accordion.selected_index = None
```
### Nesting tabs and accordions
Tabs and accordions can be nested as deeply as you want. If you have a few minutes, try nesting a few accordions or putting an accordion inside a tab or a tab inside an accordion.
The example below makes a couple of tabs with an accordion children in one of them
```
tab_nest = widgets.Tab()
tab_nest.children = [accordion, accordion]
tab_nest.titles = ('An accordion', 'Copy of the accordion')
tab_nest
```
[Index](Index.ipynb) - [Back](Widget Basics.ipynb) - [Next](Output Widget.ipynb)
| github_jupyter |
# Basic Examples with Different Protocols
## Prerequisites
* A kubernetes cluster with kubectl configured
* curl
* grpcurl
* pygmentize
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](seldon_core_setup.ipynb) to setup Seldon Core with an ingress - either Ambassador or Istio.
Then port-forward to that ingress on localhost:8003 in a separate terminal either with:
* Ambassador: `kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080`
* Istio: `kubectl port-forward $(kubectl get pods -l istio=ingressgateway -n istio-system -o jsonpath='{.items[0].metadata.name}') -n istio-system 8003:80`
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
import json
```
## Seldon Protocol REST Model
```
!pygmentize resources/model_seldon_rest.yaml
!kubectl apply -f resources/model_seldon_rest.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=rest-seldon -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep rest-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/rest-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
!kubectl delete -f resources/model_seldon_rest.yaml
```
## Seldon Protocol GRPC Model
```
!pygmentize resources/model_seldon_grpc.yaml
!kubectl apply -f resources/model_seldon_grpc.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=grpc-seldon \
-o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep grpc-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!cd ../executor/proto && grpcurl -d '{"data":{"ndarray":[[1.0,2.0,5.0]]}}' \
-rpc-header seldon:grpc-seldon -rpc-header namespace:seldon \
-plaintext \
-proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict
d=json.loads("".join(X))
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
!kubectl delete -f resources/model_seldon_grpc.yaml
```
## Tensorflow Protocol REST Model
```
!pygmentize resources/model_tfserving_rest.yaml
!kubectl apply -f resources/model_tfserving_rest.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=rest-tfserving \
-o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep rest-tfserving -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"instances": [1.0, 2.0, 5.0]}' \
-X POST http://localhost:8003/seldon/seldon/rest-tfserving/v1/models/halfplustwo/:predict \
-H "Content-Type: application/json"
d=json.loads("".join(X))
print(d)
assert(d["predictions"][0] == 2.5)
!kubectl delete -f resources/model_tfserving_rest.yaml
```
## Tensorflow Protocol GRPC Model
```
!pygmentize resources/model_tfserving_grpc.yaml
!kubectl apply -f resources/model_tfserving_grpc.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=grpc-tfserving \
-o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep grpc-tfserving -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!cd ../executor/proto && grpcurl \
-d '{"model_spec":{"name":"halfplustwo"},"inputs":{"x":{"dtype": 1, "tensor_shape": {"dim":[{"size": 3}]}, "floatVal" : [1.0, 2.0, 3.0]}}}' \
-rpc-header seldon:grpc-tfserving -rpc-header namespace:seldon \
-plaintext -proto ./prediction_service.proto \
0.0.0.0:8003 tensorflow.serving.PredictionService/Predict
d=json.loads("".join(X))
print(d)
assert(d["outputs"]["x"]["floatVal"][0] == 2.5)
!kubectl delete -f resources/model_tfserving_grpc.yaml
```
| github_jupyter |
# Soft Computing
## Vežba 5 - Klasifikacija zvuka
### Zvuk
Zvuk je mehanički talas frekvencija od 16Hz do 20kHZ, tj. u rasponu u kojem ga čuje ljudsko uvo. Zvuk frekvencije:
* niže od 16Hz - infrazvuk
* više od 20kHz - ultrazvuk
* više od 1GHz - hiperzvuk.
Zvuk nastaje više ili manje periodičnim oscilovanjem izvora zvuka koji u neposrednoj okolini menja pritisak medijuma, poremećaj pritiska prenosi se na susedne čestice medijuma i tako se širi u obliku:
* uglavnom longitudinalnih talasa u gasovima i tečnostima i
* longitudinalnih i transferzalnih talasa u čvrstim telima.
Talasi su vibracije koje prenose energiju sa mesta na mesto bez prenošenja materije.

Zvuk su kompresije i razređivanja u vazduhu koje će uvo pokupiti. Zvuk je kretanje vazduha. Često se izražava talasnim oblikom koji pokazuje šta se dešava sa česticama vazduha koje se tokom vremena kreću napred-nazad. Vertikalna osa pokazuje kako se vazduh kreće unazad ili unapred u odnosu na nultu poziciju. Horizontalna osa pokazuje vreme.
### Uzorkovanje
U obradi signala, uzorkovanje je redukcija kontinualnog signala u niz diskrenih vrednosti. Frekvencija ili brzina uzorkovanja je broj uzoraka uzetih tokom određenog vremenskog perioda. Visoka frekvencija uzorkovanja rezultira sa manjim gubitkom informacija, ali većim troškovima izračunavanja. Niske frekvencije uzorkovanja imaju veći gubitak informacija, ali su brze i jeftine za izračunavanje.

Frekvencija ili brzina uzorkovanja (eng. *sampling rate*) i bitna dubina (eng. *bit depth*) su dva najvažnija elementa kod diskretizacije zvučnog signala. Frekvencija uzorkovanja određuje koliko često će uzimati uzorke, a bitna dubina određuje kako detaljno će uzimati uzorke, kao što je prikazano na slici ispod:

Obično, CD ima 44.1kHz frekvenciju uzorkovanja sa 16-bitnom dubinom. To znači da se uzorci uzimaju 44100 puta u sekundi i da bilo koji uzorak može uzeti vrednosti iz raspona 65536 vrednosti, što će odgovarati njegovoj amplitudi.
Pored frekvencije uzorkovanja i bitne dubine, još se najčešće spominje i broj kanala (eng. *channels*). Najčešće vrednosti za broj kanala su 1 (mono) i 2 (stereo).
### Amplituda
Amplituda zvučnog talasa je mera njegove promene tokom određenog perioda. Druga uobičajena definicija amplitude je funkcija veličine razlike između ekstremnih vrednosti varijable.
### Furijeova transformacija
Furijeova transformacija razlaže funkciju vremena (signal) u frekvencije koje ga čine.

Na isti način kao što se muzički akord može predstaviti glasnoćom i frekvencijama njegovih sastavnih nota, Furijeova transformacija funkcije prikazuje amplitudu svake frekvencije prisutne u osnovnoj funkciji (signalu).

### Periodogram
U obradi signala, periodogram je procena spektralne gustine signala.

Periodogram iznad pokazuje spektar snage dve sinusoidne funkcije od ~ 30Hz i ~ 50Hz. Izlaz Furijeove transformacije se može zamisliti kao periodogram.
### Spektralna gustina
Spektar jačine (eng. *power spectrum*) talasnog oblika je način za opisivanje distribucije snage u diskrentim frekvencijskim komponentama koje čine taj signal. Statistički prosek signala, meren njegovim frekvencijskim sadržajem, naziva se njegovim spektrom. Spektralna gustina (eng. *spectral density*) digitalnog signala opisuje frekvencijski sadržaj signala.

### Mel-skala
Mel-skala (eng. *Mel-scale*) je skala visine tonova za koje slušaoci procenjuju da su jednaki u udaljenosti jedan od drugog. Referentna tačka između mel-skale i normalnog merenja frekvencije je proizvoljno definisana dodeljivanjem perceptivnog tona od 1000 mela na 1000Hz.

Formula za konverziju f Hz u m melova je:

### Cepstrum
Cepstrum je rezultat uzimanja Furijeove transformacije logartima procenjenog spektra snage signala.
### Zadatak - Klasifikacija urbanih zvukova
Kreiranje modela mašinskog učenja za klasifikaciju, opis ili generisanje zvuka obično se odnosi na modelovanje gde su ulazni podaci zvučni uzorci.
Dat je deo [UrbanSound8K](https://urbansounddataset.weebly.com/urbansound8k.html) skupa podataka koji sadrži označene zvučne uzorke (dužine <= 4 sekunde) urbanih zvukova iz 10 klasa:
* klima uređaj (*air conditioner*)
* auto sirena (*car horn*)
* dečija igra (*children playing*)
* lavež pasa (*dog bark*)
* bušenje (*drilling*)
* motor u praznom hodu (*engine idling*)
* pucanj (*gun shot*)
* sirena (*siren*)
* ulična muzika (*street music*)
* pneumatska bušilica (*jackhammer*, ili na [francuskom](https://www.youtube.com/watch?v=JqnPlH7Aol4)).
Primer za svaku klasu je dat u **samples/** folderu.
Skup podataka se nalazi u **data/** folderu tako da se:
* zvučni fajlovi nalaze u **data/audio/** folderu
* metapodaci nalaze u **data/metadata.csv** (opis svih kolona iz metapodataka dat je na gorenavedenom linku).
Potrebno je kreirati klasifikator koji će klasifikovati zvučne odlomke u odgovarajuću klasu sa što većom tačnošću.
### LibROSA
Za rad sa zvukom ćemo koristiti Python-ov paket za zvučnu i muzičku analizu - **LibROSA**:
* [Dokumentacija](https://librosa.github.io/librosa/)
* [Tutorial](https://librosa.github.io/librosa/tutorial.html)
* [Naučni rad](http://conference.scipy.org/proceedings/scipy2015/pdfs/brian_mcfee.pdf)
```
import numpy as np
import pandas as pd
import IPython.display as ipd
import librosa
import librosa.display
import matplotlib.pyplot as plt
%matplotlib inline
```
##### Metapodaci
```
df = pd.read_csv("data/metadata.csv")
```
Prikaz zaglavlja i prvih 5 redova metapodataka:
```
df.head()
```
Ukupan broj redova u skupu podataka, informacije o kolonama (tip podataka, da li sadrži nedostajuće vrednosti) i ukupna zauzetost RAM memorije od strane metapodataka:
```
df.info()
```
Distribucija klasa u skupu podataka može se utvrditi brojanjem pojavljivanja svake klase u **class** koloni:
```
df["class"].value_counts()
```
##### Primeri za svaku klasu
Zvučni uzorci se obično predstavljaju kao vremenske serije (eng. *time series*), gde y-osa predstavlja amplitudu talasnog oblika. Amplituda se obično meri kao funkcija promene pritiska oko mikrofona ili prijemnog uređaja koji je prvobitno pokupio zvuk.
Ako nema metapodataka povezanih sa zvučnim uzorcima, ovi signali vremenske serije (eng. *time series signals*) su često jedini ulazni podaci za treniranje modela.
Slede primeri svake klase iz našeg skupa podataka. Primer su propraćeni brzinom uzorkovanja, opsegom amplitude i grafičkim prikazom vremenske serije signala.
```
def display_sample(file_path):
plt.figure(figsize=(12, 4))
data, sample_rate = librosa.load(file_path)
print("Sample rate: ", sample_rate)
print("min-max range: ", np.min(data), 'to', np.max(data))
_ = librosa.display.waveplot(data, sr=sample_rate)
ipd.Audio("samples/air_con.wav")
display_sample("samples/air_con.wav")
ipd.Audio("samples/car_horn.wav")
display_sample("samples/car_horn.wav")
ipd.Audio("samples/child_play.wav")
display_sample("samples/child_play.wav")
ipd.Audio("samples/dog_bark.wav")
display_sample("samples/dog_bark.wav")
ipd.Audio("samples/drilling.wav")
display_sample("samples/drilling.wav")
ipd.Audio("samples/eng_idle.wav")
display_sample("samples/eng_idle.wav")
ipd.Audio("samples/gun.wav")
display_sample("samples/gun.wav")
ipd.Audio("samples/jackhammer.wav")
display_sample("samples/jackhammer.wav")
ipd.Audio("samples/siren.wav")
display_sample("samples/siren.wav")
ipd.Audio("samples/street_music.wav")
display_sample("samples/street_music.wav")
```
Pregledom primera iznad, jasno je da sam talasni oblik ne mora nužno da daje jasne podatke o identifikaciji klase. Talasni oblici za motor u praznom hodu, sirenu i pneumatsku bušilicu izgledaju prilično slično.
#### Izdvajanje osobina
Jedna od najboljih tehnika za izdvajanje osobina iz talasnih oblika (i signala uopšte) jeste tehnika iz 1980-te godine - **Mel Frequency Cepstral Coefficients** *(MFCCs)*, koju su osmislili [*Davis* i *Mermelstein*](https://users.cs.northwestern.edu/~pardo/courses/eecs352/papers/Davis1980-MFCC.pdf).
Koraci kod MFCCs su:
1. Furijeova transformacija signala
2. Mapiranje snage spektra dobijenog u koraku 1 na Mel-skalu
3. Logaritmovanje snage svake frekvencije na Mel-skali
4. Diskretna kosinusna transformacija liste iz koraka 3, kao da je signal
5. MFCCs su amplitude rezultujućeg spektra.
```
librosa_audio, librosa_sample_rate = librosa.load("samples/air_con.wav")
mfccs = librosa.feature.mfcc(y=librosa_audio, sr=librosa_sample_rate, n_mfcc=40)
print(mfccs.shape)
print(mfccs)
```
Vizuelizovaćemo dobijeni rezultat putem spektograma (eng. *spectogram*). Spektogram je vizuelni prikaz spektra frekvencija signala koji varira sa vremenom. Dobar način ilustrovanja spektograma jeste posmatrati ga kao složen prikaz periodograma preko nekog vremenskog intervala digitalnog signala.
```
librosa.display.specshow(mfccs, sr=librosa_sample_rate, x_axis="time")
def extract_features(file_name):
try:
audio, sample_rate = librosa.load(file_name, res_type="kaiser_fast")
mfccs = librosa.feature.mfcc(y=audio, sr=sample_rate, n_mfcc=40)
mfccscaled = np.mean(mfccs.T, axis=0)
except Exception as e:
print("Error encountered while parsing file: ", file_name)
return None
return mfccscaled
folder_path = "data/audio/"
features = []
for index, row in df.iterrows():
file_name = folder_path + row["slice_file_name"]
class_label = row["class"]
data = extract_features(file_name)
features.append([file_name, data, class_label])
features_df = pd.DataFrame(features, columns=["file", "feature", "class_label"])
features_df.head()
```
##### Klasifikacija
Prvi korak jeste da imena klasa konvertujemo u numeričke vrednosti. Za konverziju ćemo koristiti [Label Encoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html#sklearn.preprocessing.LabelEncoder).
Nako konverzije, vršimo podelu skupa podataka na trening i validacioni.
```
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
X = np.array(features_df.feature.tolist())
y = np.array(features_df.class_label.tolist())
le = LabelEncoder()
yy = le.fit_transform(y)
x_train, x_test, y_train, y_test = train_test_split(X, yy, test_size=0.2, shuffle=True, random_state=42)
```
#### SVM
Kao prvi klasifikator, koristićemo SVM sa linearnim kernelom.
```
from sklearn.svm import SVC
from sklearn import metrics
model = SVC(kernel="linear")
model.fit(x_train, y_train)
y_pred_train = model.predict(x_train)
print("Train Accuracy:", metrics.accuracy_score(y_train, y_pred_train))
y_pred_test = model.predict(x_test)
print("Test Accuracy:", metrics.accuracy_score(y_test, y_pred_test))
```
Bolju interpretaciju rezultata na testnom skupu možemo dobiti generisanjem [Classification report-a](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html#sklearn.metrics.classification_report).
```
print(metrics.classification_report(y_test, y_pred_test))
```
Ukoliko želimo da vidimo stvarni naziv svake klase, potrebno je da prosledimo i mapiranja koja je generisao Label Encoder.
```
print(metrics.classification_report(y_test, y_pred_test, target_names=le.classes_))
```
##### Neuronska mreža
Kao drugi klasifikator, koristićemo *feed-forward* neuronsku mrežu.
```
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.optimizers import Adam
from keras.utils import to_categorical
```
Prvi korak koji moramo da uradimo jeste konverzija labela (izlaza) u format pogodan za neuronsku mrežu (tj. klasa 1 će se konvertovati u [0, 1, 0, 0, 0, 0, 0, 0, 0, 0] itd.).
```
num_classes = 10
y_train = to_categorical(y_train, num_classes=num_classes)
y_test = to_categorical(y_test, num_classes=num_classes)
model = Sequential()
model.add(Dense(256, input_shape=(40, )))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer='adam')
```
Vizuelizacija našeg modela sa brojem parametara koje je potrebno "istrenirati".
```
model.summary()
```
Kao pomoć prilikom treniranja možemo koristiti neki od [callback-ova iz Keras biblioteke](https://keras.io/callbacks/). U ovom slučaju, koristićemo **ModelCheckpoint** koji čuva model nakon svake epohe. Mi ćemo ga modifikovati tako da čuva samo poslednji najbolji model.
```
from keras.callbacks import ModelCheckpoint
from datetime import datetime
num_epochs = 100
batch_size = 32
checkpointer = ModelCheckpoint(filepath="models/best_weights.hdf5", verbose=1, save_best_only=True)
start = datetime.now()
model.fit(x_train, y_train, batch_size=batch_size, epochs=num_epochs, validation_data=(x_test, y_test),
callbacks=[checkpointer], verbose=1)
duration = datetime.now() - start
print("Training completed in: ", duration)
train_score = model.evaluate(x_train, y_train, verbose=1)
print("Training Accuracy: ", train_score[1])
test_score = model.evaluate(x_test, y_test, verbose=1)
print("Test Accuracy: ", test_score[1])
```
Model neuronske mreže je ostvario manju trening tačnost, ali je ostvario veću tačnost na testnom skupu podataka od SVM modela. Možemo zaključiti da je model neuronske mreže bolje "naučio" da generalizuje na novim (nevidljivim) podacima.
| github_jupyter |
<img src="../../img/logo_white_bkg_small.png" align="left" />
# Tuning your Classifier
This worksheet covers concepts covered in the second part of day 3 - Tuning Classifiers. It should take no more than 20-30 minutes to complete. Please raise your hand if you get stuck.
## Import the Libraries
For this exercise, we will be using:
* Pandas (http://pandas.pydata.org/pandas-docs/stable/)
* Numpy (https://docs.scipy.org/doc/numpy/reference/)
* Matplotlib (http://matplotlib.org/api/pyplot_api.html)
* Scikit-learn (http://scikit-learn.org/stable/documentation.html)
* YellowBrick (http://www.scikit-yb.org/en/latest/)
* Seaborn (https://seaborn.pydata.org)
* Lime (https://github.com/marcotcr/lime)
```
# Load Libraries - Make sure to run this cell!
import pandas as pd
import numpy as np
import time
from collections import Counter
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from scipy.stats import uniform as sp_rand
from yellowbrick.classifier import ClassificationReport
from yellowbrick.classifier import ConfusionMatrix
import matplotlib.pyplot as plt
import matplotlib
import lime
%matplotlib inline
```
## Prepare the Data
For this exercise, we are going to focus on building a pipeline and then tuning the resultant model, so we're going to use our data from the last worksheet.
```
df_final = pd.read_csv('../../Data/dga_features_final_df.csv')
target = df_final['isDGA']
feature_matrix = df_final.drop(['isDGA'], axis=1)
feature_matrix.sample(5)
```
### Split the data into training and testing sets.
We're going to need a training and testing dataset, so you know the drill, split the data..
```
# Simple Cross-Validation: Split the data set into training and test data
feature_matrix_train, feature_matrix_test, target_train, target_test = train_test_split(feature_matrix,
target,
test_size=0.25)
```
## Build a Model
For this exercise, we're going to create a K-NN Classifier for the DGA data and tune it, but first, create a classifier with the default options and calculate the accuracy score for it. (http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html)
The default parameters are shown below.
```python
KNeighborsClassifier(algorithm='auto', leaf_size=30, metric='minkowski',
metric_params=None, n_jobs=1, n_neighbors=5, p=2,
weights='uniform')
```
```
# Your code here ...
#Store the predictions
```
## Improving Performance
Out of the box, the model achieves approximately 85% accuracy. Better than chance but let's see if we can do better.
**Note: This notebook is written without using fixed random seeds, so you might get slightly different results.**
### Scaling the Features
K-NN is a distance-based classifier and hence it is necessary to scale the features prior to training the model. For this exercise however, let's create a simple pipeline with two steps:
1. StandardScaler
2. Train the classifier
Once you've done that, calculate the accuracy and see if it has improved.
```
#Your code here...
```
Scaling the features did result in a small improvement: .85 accuracy to .88. But let's see if we can't do even better.
### Using RandomSearchCV and GridSearchCV to tune Hyperparameters
Now that we've scaled the features and built a simple pipeline, let's try to tune the hyperparameters to see if we can improve the model performance. Scikit-learn provides two methods for accomplishing this task: `RandomizedSearchCV` and `GridSearchCV`.
* `GridSearchCV`: GridSearch iterates through all possible combinations of tuning parameters to find the optimal combination. (http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html)
* `RandomizedSearchCV`: RandomizedSearch interates through random combinations of paremeters to find the optimal combination. While RandomizedSearch does not try every possible combination, is considerably faster than GridSearch and has been shown to get very close to the optimal combination in considerably less time. (http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html)
You can see in the results below, that the model was able to achieve **91.9%** accuracy with RandomSearch!
```
[INFO] randomized search took 0.85 seconds
[INFO] grid search accuracy: 91.93%
[INFO] randomized search best parameters: {'clf__weights': 'uniform', 'clf__p': 1, 'clf__n_neighbors': 27, 'clf__metric': 'euclidean', 'clf__leaf_size': 25, 'clf__algorithm': 'kd_tree'}
```
Both `RandomizedSearchCV` and `GridSearchCV` require you to provide a grid of parameters. You will need to refer to the documentation for the classifier you are using to get a list of paramenters for that particular model. Also since we will be using the pipeline, you have to format the parameters correctly. The name of the variable must be preceeded by the name of the step in your pipeline and two underscores. For example. If the classifier in the pipeline is called `clf`, and you have a tuning parameter called `metric`, the parameter grid would be as follows:
```python
params = {
"clf__n_neighbors": np.arange(1, 50, 2),
"clf__metric": ["euclidean", "cityblock"]
}
```
### Your Task
Using either GridSearchCV or RandomizedSearchCV, improve the performance of your model.
```
#Your code here...
```
## Model Comparison
Your final task is to:
1. Using RandomForest, create a classifier for the DGA dataset
2. Use either GridSearchCV or RandomizedSearchCV to find the optimal parameters for this model.
How does this model compare with the first K-NN classifier for this data?
```
#Your code here...
```
| github_jupyter |
# Preparing the Datasets for Image Classification using the Apache MXNet Vision Datasets Functions
<img align="left" width="130" src="https://raw.githubusercontent.com/PacktPublishing/Amazon-SageMaker-Cookbook/master/Extra/cover-small-padded.png"/>
This notebook contains the code to help readers work through one of the recipes of the book [Machine Learning with Amazon SageMaker Cookbook: 80 proven recipes for data scientists and developers to perform ML experiments and deployments](https://www.amazon.com/Machine-Learning-Amazon-SageMaker-Cookbook/dp/1800567030)
### How to do it...
```
%%bash
mkdir -p tmp/train/0 tmp/train/1 tmp/train/2 tmp/train/3 tmp/train/4
mkdir -p tmp/train/5 tmp/train/6 tmp/train/7 tmp/train/8 tmp/train/9
mkdir -p tmp/validation/0 tmp/validation/1 tmp/validation/2 tmp/validation/3 tmp/validation/4
mkdir -p tmp/validation/5 tmp/validation/6 tmp/validation/7 tmp/validation/8 tmp/validation/9
%%bash
mkdir -p tmp/train_lst
mkdir -p tmp/validation_lst
mkdir -p tmp/test
%%bash
ls -1F tmp/train
%%bash
ls -1F tmp/validation
import mxnet as mx
mx.random.seed(21)
def transform_fxn(data, label):
data = data.astype('float32')
data = data / 255
return data, label
ds = mx.gluon.data.vision.datasets.MNIST(
train=True,
transform=transform_fxn
)
training_and_validation_dataset = ds
ds = mx.gluon.data.vision.datasets.MNIST(
train=False,
transform=transform_fxn
)
test_dataset = ds
print(len(training_and_validation_dataset))
print(len(test_dataset))
def get_training_row_indexes(row_count,
percent=0.5,
ratio=0.8):
training_index_start = 0
end = int(row_count * ratio * percent)
training_index_end = end
print("Range Index Start:",
training_index_start)
print("Range Index End:",
training_index_end)
output = list(range(training_index_start,
training_index_end))
print("Output Length:", len(output))
print("Last Index:", output[-1])
return output
def get_validation_row_indexes(row_count,
percent=0.5,
ratio=0.8):
start = int(row_count * ratio)
validation_index_start = start
count = int((1 - ratio) * row_count * percent) + 1
element_count = count
validation_index_end = validation_index_start + element_count
print("Range Index Start:",
validation_index_start)
print("Element Count:",
element_count)
print("Range Index End:",
validation_index_end)
output = list(range(validation_index_start,
validation_index_end))
print("Output Length:", len(output))
print("Last Index:", output[-1])
return output
def get_test_row_indexes(row_count,
percent=0.5):
test_index_start = 0
test_index_end = int(row_count * percent)
print("Range Index Start:",
test_index_start)
print("Range Index End:",
test_index_end)
output = list(range(test_index_start,
test_index_end))
print("Output Length:", len(output))
print("Last Index:", output[-1])
return output
get_training_row_indexes(row_count=60000,
percent=0.5)
get_validation_row_indexes(row_count=60000,
percent=0.5)
get_test_row_indexes(row_count=10000,
percent=0.1)
import string
import random
def generate_random_string():
return ''.join(
random.sample(
string.ascii_uppercase,12)
)
generate_random_string()
import matplotlib
import matplotlib.pyplot
def save_image(image_data, filename):
matplotlib.pyplot.imsave(
f"tmp/{filename}",
image_data[:,:,0].asnumpy())
def generate_image_files_and_lst_dict(
dataset,
indexes,
tag
):
list_of_lst_dicts = []
for index in indexes:
image_label_pair = dataset[index]
image_data = image_label_pair[0]
label = image_label_pair[1]
random_string = generate_random_string()
if tag == "test":
rp = f"{random_string}.png"
relative_path = rp
filename = f"{tag}/{relative_path}"
else:
rp = f"{label}/{random_string}.png"
relative_path = rp
filename = f"{tag}/{relative_path}"
save_image(
image_data,
filename=filename
)
lst_dict = {
'relative_path': relative_path,
'class': label
}
list_of_lst_dicts.append(lst_dict)
return list_of_lst_dicts
train_dataset_length = len(
training_and_validation_dataset
)
train_indexes = get_training_row_indexes(
row_count=train_dataset_length,
percent=0.01)
t = generate_image_files_and_lst_dict(
dataset=training_and_validation_dataset,
indexes=train_indexes,
tag = "train"
)
train_lst_dict = t
train_lst_dict
train_dataset_length = len(
training_and_validation_dataset
)
validation_indexes = get_validation_row_indexes(
row_count=train_dataset_length,
percent=0.01)
v = generate_image_files_and_lst_dict(
dataset=training_and_validation_dataset,
indexes=validation_indexes,
tag = "validation"
)
validation_lst_dict = v
validation_lst_dict
test_dataset_length = len(test_dataset)
test_indexes = get_test_row_indexes(
row_count=test_dataset_length,
percent=0.01)
test_lst_dict = generate_image_files_and_lst_dict(
dataset=test_dataset,
indexes=test_indexes,
tag = "test"
)
test_lst_dict
def save_lsts_to_file(values, filename):
with open(filename, 'w') as output:
for index, row in enumerate(
values,
start=1
):
relative_path = row['relative_path']
cls = row['class']
tmp = f"{index}\t{cls}\t{relative_path}\n"
output.write(tmp)
save_lsts_to_file(
train_lst_dict,
filename="tmp/train_lst/train.lst"
)
save_lsts_to_file(
validation_lst_dict,
filename="tmp/validation_lst/validation.lst"
)
%%bash
head tmp/train_lst/train.lst
s3_bucket = "<insert S3 bucket name here>"
prefix = "image-experiments"
!aws s3 cp tmp/. s3://{s3_bucket}/{prefix}/ --recursive
%store s3_bucket
%store prefix
```
| github_jupyter |
```
# Model train loss = 0.44 (approx)
# Importing the required libraries
import cv2
import numpy as np
import pandas as pd
import os
import glob
import sys
import time
import torch
sys.path.insert(0, "/kaggle/input/blazeface-pytorch")
from blazeface import BlazeFace
from tensorflow.keras.applications.inception_v3 import InceptionV3
from keras.applications.inception_v3 import preprocess_input
from tensorflow.keras.layers import Input
from tensorflow.keras.models import load_model
# Initializing the paths
input_path = '/kaggle/input/deepfake-detection-challenge/'
deepfake_model_path = '/kaggle/input/deepfake-model044/deepfake_predictor_0.44.h5'
inception_weigths_path = '/kaggle/input/inception-pretrained/inception_v3.h5'
test_dir = glob.glob(input_path + 'test_videos/*.mp4')
class VideoReader:
"""Helper class for reading one or more frames from a video file."""
def __init__(self, verbose=True, insets=(0, 0)):
"""Creates a new VideoReader.
Arguments:
verbose: whether to print warnings and error messages
insets: amount to inset the image by, as a percentage of
(width, height). This lets you "zoom in" to an image
to remove unimportant content around the borders.
Useful for face detection, which may not work if the
faces are too small.
"""
self.verbose = verbose
self.insets = insets
def read_frames(self, path, num_frames, jitter=0, seed=None):
"""Reads frames from 90th frame continuously.
Arguments:
path: the video file
num_frames: how many frames to read, -1 means the entire video
(warning: this will take up a lot of memory!)
jitter: if not 0, adds small random offsets to the frame indices;
this is useful so we don't always land on even or odd frames
seed: random seed for jittering; if you set this to a fixed value,
you probably want to set it only on the first video
"""
assert num_frames > 0
capture = cv2.VideoCapture(path)
# frame_count = int(capture.get(cv2.CAP_PROP_FRAME_COUNT))
# if frame_count <= 0: return None
frame_idxs = np.linspace(90, 90 + num_frames - 1, num_frames, endpoint=True, dtype=np.int)
# frame_idxs = np.linspace(0, frame_count - 1, num_frames, endpoint=True, dtype=np.int)
if jitter > 0:
np.random.seed(seed)
jitter_offsets = np.random.randint(-jitter, jitter, len(frame_idxs))
frame_idxs = np.clip(frame_idxs + jitter_offsets, 0, frame_count - 1)
result = self._read_frames_at_indices(path, capture, frame_idxs)
capture.release()
return result
def read_frames_at_indices(self, path, frame_idxs):
"""Reads frames from a video and puts them into a NumPy array.
Arguments:
path: the video file
frame_idxs: a list of frame indices. Important: should be
sorted from low-to-high! If an index appears multiple
times, the frame is still read only once.
Returns:
- a NumPy array of shape (num_frames, height, width, 3)
- a list of the frame indices that were read
Reading stops if loading a frame fails, in which case the first
dimension returned may actually be less than num_frames.
Returns None if an exception is thrown for any reason, or if no
frames were read.
"""
assert len(frame_idxs) > 0
capture = cv2.VideoCapture(path)
result = self._read_frames_at_indices(path, capture, frame_idxs)
capture.release()
return result
def _read_frames_at_indices(self, path, capture, frame_idxs):
try:
frames = []
idxs_read = []
for frame_idx in range(frame_idxs[0], frame_idxs[-1] + 1):
# Get the next frame, but don't decode if we're not using it.
ret = capture.grab()
if not ret:
if self.verbose:
print("Error grabbing frame %d from movie %s" % (frame_idx, path))
break
# Need to look at this frame?
current = len(idxs_read)
if frame_idx == frame_idxs[current]:
ret, frame = capture.retrieve()
if not ret or frame is None:
if self.verbose:
print("Error retrieving frame %d from movie %s" % (frame_idx, path))
break
frame = self._postprocess_frame(frame)
frames.append(frame)
idxs_read.append(frame_idx)
if len(frames) > 0:
return np.stack(frames), idxs_read
if self.verbose:
print("No frames read from movie %s" % path)
return None
except:
if self.verbose:
print("Exception while reading movie %s" % path)
return None
def _postprocess_frame(self, frame):
frame = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
if self.insets[0] > 0:
W = frame.shape[1]
p = int(W * self.insets[0])
frame = frame[:, p:-p, :]
if self.insets[1] > 0:
H = frame.shape[1]
q = int(H * self.insets[1])
frame = frame[q:-q, :, :]
return frame
class FaceExtractor:
"""Wrapper for face extraction workflow."""
def __init__(self, video_read_fn, facedetn):
"""Creates a new FaceExtractor.
Arguments:
video_read_fn: a function that takes in a path to a video file
and returns a tuple consisting of a NumPy array with shape
(num_frames, H, W, 3) and a list of frame indices, or None
in case of an error
facedet: the face detector object
"""
self.video_read_fn = video_read_fn
self.facedet = facedet
def process_videos(self, input_dir, filenames, video_idxs):
"""For the specified selection of videos, grabs one or more frames
from each video, runs the face detector, and tries to find the faces
in each frame.
The frames are split into tiles, and the tiles from the different videos
are concatenated into a single batch. This means the face detector gets
a batch of size len(video_idxs) * num_frames * num_tiles (usually 3).
Arguments:
input_dir: base folder where the video files are stored
filenames: list of all video files in the input_dir
video_idxs: one or more indices from the filenames list; these
are the videos we'll actually process
Returns a list of dictionaries, one for each frame read from each video.
This dictionary contains:
- video_idx: the video this frame was taken from
- frame_idx: the index of the frame in the video
- frame_w, frame_h: original dimensions of the frame
- faces: a list containing zero or more NumPy arrays with a face crop
- scores: a list array with the confidence score for each face crop
If reading a video failed for some reason, it will not appear in the
output array. Note that there's no guarantee a given video will actually
have num_frames results (as soon as a reading problem is encountered for
a video, we continue with the next video).
"""
target_size = self.facedet.input_size
videos_read = []
frames_read = []
frames = []
tiles = []
resize_info = []
for video_idx in video_idxs:
# Read the full-size frames from this video.
filename = filenames[video_idx]
video_path = os.path.join(input_dir, filename)
result = self.video_read_fn(video_path)
# Error? Then skip this video.
if result is None: continue
videos_read.append(video_idx)
# Keep track of the original frames (need them later).
my_frames, my_idxs = result
frames.append(my_frames)
frames_read.append(my_idxs)
# Split the frames into several tiles. Resize the tiles to 128x128.
my_tiles, my_resize_info = self._tile_frames(my_frames, target_size)
tiles.append(my_tiles)
resize_info.append(my_resize_info)
# Put all the tiles for all the frames from all the videos into
# a single batch.
batch = np.concatenate(tiles)
# Run the face detector. The result is a list of PyTorch tensors,
# one for each image in the batch.
all_detections = self.facedet.predict_on_batch(batch, apply_nms=False)
result = []
offs = 0
for v in range(len(tiles)):
# Not all videos may have the same number of tiles, so find which
# detections go with which video.
num_tiles = tiles[v].shape[0]
detections = all_detections[offs:offs + num_tiles]
offs += num_tiles
# Convert the detections from 128x128 back to the original frame size.
detections = self._resize_detections(detections, target_size, resize_info[v])
# Because we have several tiles for each frame, combine the predictions
# from these tiles. The result is a list of PyTorch tensors, but now one
# for each frame (rather than each tile).
num_frames = frames[v].shape[0]
frame_size = (frames[v].shape[2], frames[v].shape[1])
detections = self._untile_detections(num_frames, frame_size, detections)
# The same face may have been detected in multiple tiles, so filter out
# overlapping detections. This is done separately for each frame.
detections = self.facedet.nms(detections)
for i in range(len(detections)):
# Crop the faces out of the original frame.
faces = self._add_margin_to_detections(detections[i], frame_size, 0.2)
faces = self._crop_faces(frames[v][i], faces)
# Add additional information about the frame and detections.
scores = list(detections[i][:, 16].cpu().numpy())
frame_dict = { "video_idx": videos_read[v],
"frame_idx": frames_read[v][i],
"frame_w": frame_size[0],
"frame_h": frame_size[1],
"faces": faces,
"scores": scores }
result.append(frame_dict)
# TODO: could also add:
# - face rectangle in original frame coordinates
# - the keypoints (in crop coordinates)
return result
def process_video(self, video_path):
"""Convenience method for doing face extraction on a single video."""
input_dir = os.path.dirname(video_path)
filenames = [ os.path.basename(video_path) ]
return self.process_videos(input_dir, filenames, [0])
def _tile_frames(self, frames, target_size):
"""Splits each frame into several smaller, partially overlapping tiles
and resizes each tile to target_size.
After a bunch of experimentation, I found that for a 1920x1080 video,
BlazeFace works better on three 1080x1080 windows. These overlap by 420
pixels. (Two windows also work but it's best to have a clean center crop
in there as well.)
I also tried 6 windows of size 720x720 (horizontally: 720|360, 360|720;
vertically: 720|1200, 480|720|480, 1200|720) but that gives many false
positives when a window has no face in it.
For a video in portrait orientation (1080x1920), we only take a single
crop of the top-most 1080 pixels. If we split up the video vertically,
then we might get false positives again.
(NOTE: Not all videos are necessarily 1080p but the code can handle this.)
Arguments:
frames: NumPy array of shape (num_frames, height, width, 3)
target_size: (width, height)
Returns:
- a new (num_frames * N, target_size[1], target_size[0], 3) array
where N is the number of tiles used.
- a list [scale_w, scale_h, offset_x, offset_y] that describes how
to map the resized and cropped tiles back to the original image
coordinates. This is needed for scaling up the face detections
from the smaller image to the original image, so we can take the
face crops in the original coordinate space.
"""
num_frames, H, W, _ = frames.shape
# Settings for 6 overlapping windows:
# split_size = 720
# x_step = 480
# y_step = 360
# num_v = 2
# num_h = 3
# Settings for 2 overlapping windows:
# split_size = min(H, W)
# x_step = W - split_size
# y_step = H - split_size
# num_v = 1
# num_h = 2 if W > H else 1
split_size = min(H, W)
x_step = (W - split_size) // 2
y_step = (H - split_size) // 2
num_v = 1
num_h = 3 if W > H else 1
splits = np.zeros((num_frames * num_v * num_h, target_size[1], target_size[0], 3), dtype=np.uint8)
i = 0
for f in range(num_frames):
y = 0
for v in range(num_v):
x = 0
for h in range(num_h):
crop = frames[f, y:y+split_size, x:x+split_size, :]
splits[i] = cv2.resize(crop, target_size, interpolation=cv2.INTER_AREA)
x += x_step
i += 1
y += y_step
resize_info = [split_size / target_size[0], split_size / target_size[1], 0, 0]
return splits, resize_info
def _resize_detections(self, detections, target_size, resize_info):
"""Converts a list of face detections back to the original
coordinate system.
Arguments:
detections: a list containing PyTorch tensors of shape (num_faces, 17)
target_size: (width, height)
resize_info: [scale_w, scale_h, offset_x, offset_y]
"""
projected = []
target_w, target_h = target_size
scale_w, scale_h, offset_x, offset_y = resize_info
for i in range(len(detections)):
detection = detections[i].clone()
# ymin, xmin, ymax, xmax
for k in range(2):
detection[:, k*2 ] = (detection[:, k*2 ] * target_h - offset_y) * scale_h
detection[:, k*2 + 1] = (detection[:, k*2 + 1] * target_w - offset_x) * scale_w
# keypoints are x,y
for k in range(2, 8):
detection[:, k*2 ] = (detection[:, k*2 ] * target_w - offset_x) * scale_w
detection[:, k*2 + 1] = (detection[:, k*2 + 1] * target_h - offset_y) * scale_h
projected.append(detection)
return projected
def _untile_detections(self, num_frames, frame_size, detections):
"""With N tiles per frame, there also are N times as many detections.
This function groups together the detections for a given frame; it is
the complement to tile_frames().
"""
combined_detections = []
W, H = frame_size
split_size = min(H, W)
x_step = (W - split_size) // 2
y_step = (H - split_size) // 2
num_v = 1
num_h = 3 if W > H else 1
i = 0
for f in range(num_frames):
detections_for_frame = []
y = 0
for v in range(num_v):
x = 0
for h in range(num_h):
# Adjust the coordinates based on the split positions.
detection = detections[i].clone()
if detection.shape[0] > 0:
for k in range(2):
detection[:, k*2 ] += y
detection[:, k*2 + 1] += x
for k in range(2, 8):
detection[:, k*2 ] += x
detection[:, k*2 + 1] += y
detections_for_frame.append(detection)
x += x_step
i += 1
y += y_step
combined_detections.append(torch.cat(detections_for_frame))
return combined_detections
def _add_margin_to_detections(self, detections, frame_size, margin=0.2):
"""Expands the face bounding box.
NOTE: The face detections often do not include the forehead, which
is why we use twice the margin for ymin.
Arguments:
detections: a PyTorch tensor of shape (num_detections, 17)
frame_size: maximum (width, height)
margin: a percentage of the bounding box's height
Returns a PyTorch tensor of shape (num_detections, 17).
"""
offset = torch.round(margin * (detections[:, 2] - detections[:, 0]))
detections = detections.clone()
detections[:, 0] = torch.clamp(detections[:, 0] - offset*2, min=0) # ymin
detections[:, 1] = torch.clamp(detections[:, 1] - offset, min=0) # xmin
detections[:, 2] = torch.clamp(detections[:, 2] + offset, max=frame_size[1]) # ymax
detections[:, 3] = torch.clamp(detections[:, 3] + offset, max=frame_size[0]) # xmax
return detections
def _crop_faces(self, frame, detections):
"""Copies the face region(s) from the given frame into a set
of new NumPy arrays.
Arguments:
frame: a NumPy array of shape (H, W, 3)
detections: a PyTorch tensor of shape (num_detections, 17)
Returns a list of NumPy arrays, one for each face crop. If there
are no faces detected for this frame, returns an empty list.
"""
faces = []
for i in range(len(detections)):
ymin, xmin, ymax, xmax = detections[i, :4].cpu().numpy().astype(np.int)
face = frame[ymin:ymax, xmin:xmax, :]
faces.append(face)
return faces
def keep_only_best_face(self, crops):
"""For each frame, only keeps the face with the highest confidence.
This gets rid of false positives, but obviously is problematic for
videos with two people!
This is an optional postprocessing step. Modifies the original
data structure.
"""
for i in range(len(crops)):
frame_data = crops[i]
if len(frame_data["faces"]) > 0:
frame_data["faces"] = frame_data["faces"][:1]
frame_data["scores"] = frame_data["scores"][:1]
def isotropically_resize_image(img, size, resample=cv2.INTER_AREA):
h, w = img.shape[:2]
if w > h:
h = h * size // w
w = size
else:
w = w * size // h
h = size
resized = cv2.resize(img, (w, h), interpolation=resample)
return resized
def make_square_image(img):
h, w = img.shape[:2]
size = max(h, w)
t = 0
b = size - h
l = 0
r = size - w
return cv2.copyMakeBorder(img, t, b, l, r, cv2.BORDER_CONSTANT, value=0)
frames_per_video = 20 # 20 frames starting from 90th frame will be captured
input_size = 229 # 229x229x3 (square-fitted)
# Taking the base model as Inception V3 and initializing its weight with imagenet
input_tensor = Input(shape = (229, 229, 3))
cnn_model = InceptionV3(input_tensor = input_tensor, weights = None, include_top = False, pooling = 'avg')
cnn_model.load_weights(inception_weigths_path)
cnn_model.summary()
# Loading trained RNN model
rnn = load_model(deepfake_model_path)
rnn.summary()
# Loading the BlazeFace model weights
gpu = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
facedet = BlazeFace().to(gpu)
facedet.load_weights("/kaggle/input/blazeface-pytorch/blazeface.pth")
facedet.load_anchors("/kaggle/input/blazeface-pytorch/anchors.npy")
_ = facedet.train(False)
def preprocessing(video_path):
try:
# Find the faces for frames_per_video captured frames in the video
faces = face_extractor.process_video(video_path)
# In case only one person in the video, the other detections are false positives
# The face with highest confidence is taken
face_extractor.keep_only_best_face(faces)
# TODO: def sort_by_histogram(self, crops) for videos with 2 people
features = []
for frame_data in faces:
for face in frame_data["faces"]:
face = make_square_image(isotropically_resize_image(face, input_size))
# BGR to RGB
features.append(np.asarray(face)[:,:,::-1])
x = frames_per_video - np.array(features).shape[0]
# No face found
if(x == frames_per_video):
return None, False
if(x != 0):
# Pre-padding for efficient training (Done when features found is less than frames_per_video)
features = np.concatenate((np.zeros((x, 229, 229, 3)), np.array(features)))
# Preprocessing specific to Keras implementation of Inception-V3
features = preprocess_input(np.array(features))
return features, True
except:
return None, False
def predict_on_video(video_path):
features, success = preprocessing(video_path)
if(success == False):
return 0.5
return(rnn.predict(np.expand_dims(cnn_model.predict(np.array(features)), axis = 0))[0][1])
def predict_on_video_set(video_paths, output_list):
output = []
for v in video_paths:
output.append([v.split('/')[-1], predict_on_video(v)])
return output
video_reader = VideoReader()
video_read_fn = lambda x: video_reader.read_frames(x, num_frames = frames_per_video)
face_extractor = FaceExtractor(video_read_fn, facedet)
t1 = time.time()
output_list = predict_on_video_set(test_dir, [])
print('Time taken (Preprocessing + Prediction): ', time.time() - t1)
df = pd.DataFrame(output_list, columns = ['filename', 'label'])
df.head()
# Submission file
df.to_csv('submission.csv', index = False)
```
| github_jupyter |
<figure>
<IMG SRC="https://mamba-python.nl/images/logo_basis.png" WIDTH=125 ALIGN="right">
</figure>
# 1. Read and write text files
This notebook is a brief introduction to reading and writing text files.
Note: For these exercises we assume you are working on a windows computer. Some things will be different for mac or linux.
### Inhoudsopgave<a id="top"></a>
1. [Reading](#1)
2. [File path](#2)
3. [Writing](#3)
4. [Modifying](#4)
5. [Higher level reading/writing (NOT YET AVAILABLE)](#5)
6. [Answers](#6)
## [1. Reading a text file ](#top)<a id="1"></a>
If you want to read the data from a text file with Python you follow these steps:
1. Open the file with Python
2. Read data from the file
3. Close the file
In the example below we read the file [file1.txt](./file1.txt). We use the built-in function `open` to open the file.
```
#1 open the file
f = open('file1.txt')
```
We read the text from the file as a string. We save the string in the variable `data`.
```
#2 read data from the file
data = f.read()
data
```
You can see that the string contains the characters `\n` where there is an enter in the file. `\n` is the code for a line ending in a string. When you print the variable `data` the `\n` is replaced by an enter.
```
print(data)
```
When you are done reading the file you have to close it. Otherwise the file remains open in Python. Windows treats open files as locked so you cannot edit them or open them again.
```
#3 close the file
f.close()
```
Now that you've closed the file you canot read it anymore.
```
print(f.read())
```
#### Exercise 1 <a name="opdr1"></a>
Read the text from the file [file2.txt](./file2.txt). Print the data in the file.
<a href="#antw1">Answer Exercise 1</a>
### with statement
Opening and closing files can be annoying. If you forget to close the file unexpected things may happen. Therefore the `with` statement is very usefull. If you use a `with` statement to open a file, the file will be closed automatically once the code within the with-statement is executed. Therefore you don't have to explicitely close the file. For example:
```
with open('file1.txt') as f:
data = f.read()
print(data)
#check to see if it is still open
f.read()
```
#### Exercise 2 <a name="opdr2"></a>
Open and read the text in `file2.txt` using a with statement. Store the text in a variable.
<a href="#antw2">Answer Exercise 2</a>
## [2. file path](#top)<a id="2"></a>
In [chapter 1](#1) we've read 2 files using only the filename: `file1.txt` and `file2.txt`. Python knows which file to read because the files are in the same directory as this jupyter notebook (`01_read_write_text_files.ipynb`). If you want to read files from other directories you have roughly 2 options:
1. specify the full file path
2. specify the relative path to the file. This is the path relative to the directory of the jupyter notebook you are working in.
#### terms & definitions
In this chapter we use a few terms that can be confusing. Here is a short summary of the terms and their meaning:
- **file extension** the file extension contains information about the type of file and the program you can use to open the file e.g. '.txt', '.xlsx'
- **filename** the name of the file including the extension e.g. 'file1.txt', 'calculation.csv'.
- **directory** the folder in which a file is stored. Commonly used directory names in Windows are 'Documents' or 'Downloads'.
- **path** the unique location of a file or directory on your local file system, e.g.
- directory path: 'C:\Users\onno_\Documents\Exercises_Python_Basic'
- file path: 'C:\Users\onno_\Downloads\pycharm-community-2020.1.1.exe'
- **current working directory (cwd)** the directory that you are currently working in. In this case it is the directory that contains this jupyter notebook ('01_read_write_text_files.ipynb').
#### 1. full path
You can read a file by specifying the full path using the code below. Note, the code below probably gives a `FileNotFoundError` because your course-material is in a different directory.
```
with open(r'C:\Users\noel\02_git_repos\course-material\Exercise_notebooks\on_topic\07_read_write_text_files\file1.txt') as f:
data = f.read()
print(data)
```
Note that we use the character `r` before the string `'C:\Users\noel\...'`. If we omit this character we get an error because the backslash `\` is a special character in a string. For example `'\n'` represents an enter. With the `r` in front of the string we indicate that the string should be interpreted as a raw string without any special characters. It basically converts all backslashes to double backslashes `\\`. A double backslash is intepreted as a backslash in Python.
```
# using 'r' to create a raw string (with double backslashes)
s1 = r'C:\Users\noel\02_git_repos\course-material\Exercise_notebooks\on_topic\07_read_write_text_files\file1.txt'
s1
```
When you try to create a string with backslashes and without the `'r'` you can run into errors. In the example below we get an error because the `\U` is a special character that you use to create a unicode string. We get this error because the characters after `\U` cannot be interpreted as unicode (because it is not unicode).
```
# trying to create a string with single backslashes gives an error
s2 = 'C:\Users\noel\02_git_repos\course-material\Exercise_notebooks\on_topic\07_read_write_text_files\file1.txt'
s2
```
#### Exercise 3 <a name="opdr3"></a>
Find a text file on your computer. Read the content of the file using the full file path. Print the content of the file.
<a href="#antw3">Answer exercise 3</a>
### 2. relative path
You can also specify the path of the file relative to the current working directory. In the example below we open the file [`holland_seawater.dat`](../02_matplotlib/holland_seawater.dat) which is located in the `02_matplotlib` directory. You can see in the diagram below that the file and this Jupyter Notebook share the same parent directory named `On_topic`.
<br>
<figure>
<IMG SRC="directory_structure.PNG" WIDTH=500>
</figure>
When you specify a relative path you use `..\` to go to the parent directory of the current working directory.
```
with open(r'..\02_matplotlib\holland_seawater.dat') as f:
data = f.read()
print(data)
```
#### Exercise 4<a name="opdr4"></a>
Read the data from the file `xypoints.dat` in the directory `Exercise_notebooks\Basic\basic3_arrays`.
Tip: You can use `'..\..\'` to go to the parent of the parent directory.
<a href="#antw4">Answer exercise 4</a>
#### why use relative paths?
Using relative paths can help when you work together on a project. In a project the scripts and text files are usually in different directories within the same project directory. This makes it possible to use relative paths to read the files in your script. Now your colleague can use a copy of the project directory and run all the scripts seamlessly without any FileNotFoundErrors.
When you would've used absolute paths FileNotFoundErrors will occur when others run the scripts. Their project directory path is different from yours. With relative paths your scripts will work for every user regardless of where others have put the project directory on their local file system.
## [3. Writing text files](#top)<a id="3"></a>
Writing a text file has steps similar to reading a file:
1. Open the file with Python
2. Write data to the file
3. Close the file
In the example below we write the string `I ate a clock yesterday, it was very time-consuming.` to the file `my_diary.txt`.
```
#1 open the file
f = open('my_diary.txt', 'w')
#2 write data to the file
f.write('I ate a clock yesterday, it was very time-consuming.')
#3 close the file
f.close()
#or using the with statement
with open('my_diary.txt', 'w') as f:
f.write('I ate a clock yesterday, it was very time-consuming.')
```
Note that the second argument of the `open` function is the `mode`. You can choose from the following modes:
- `'r'`: read from the file
- `'w'`: write to the file
- `'a'`: append to the file
The default mode is `'r'`, that's why we did not specify a mode when we were reading files. Now that we want to write a file we set the mode to `'w'`.
When you have run the cells above you see that the file 'my_diary.txt' appeared in the current working directory. You can open the file to check if it contains the correct text.
#### Exercise 5 <a name="opdr5"></a>
Create a text file in the `personal_application` directory (`Exercise_notebooks\personal_application`) named `readme.md`. Write the following text to the file `this folder contains my personal application. In the Jupyter Notebook 'eigen_toepassing' you can find more information.`.
<a href="#antw5">Answer exercise 5</a>
## [4. Modifying](#top)<a id="4"></a>
Fairly often we want to modify an existing file rather than creating a new file. For this we can use a combination of reading and writing. In the example below we modify the file `file2.txt` by replacing the word `Iraq` with `Malawi`. We save the new file under a new name `file2_mod.txt`. Note that we did not overwrite the original file. This is common practice to avoid loosing data. If something goes wrong we will still have the original file.
```
# reading the whole file as a string and write the modified string to the file
with open('file2.txt', 'r') as f:
txt = f.read()
new_txt = txt.replace('Iraq', 'Malawi')
with open('file2_mod.txt', 'w') as f:
f.write(new_txt)
```
#### Exercise 6 <a name="opdr6"></a>
Use a combination of reading an writing to add the string `Then I went back for seconds.` to the file `my_diary.txt`. This is the file that we've created in [chapter 3](#3). Decide for yourself which method you want to use.
<a href="#antw6">Answer exercise 6</a>
#### Append
In the example above we read the whole file as and write the modified string to a new file. Below is another way to make the same modifications to the file. Now we read the file line by line. After a line is read and modified it is written to a file. Note that we use `'a'` in the `open()` function to indicate that we want to append to this file rather than overwrite. This is necesarry because we write to the file line by line.
```
# use the append, reading and writing the file line by line
with open('file2.txt', 'r') as f_in:
with open('file2_mod.txt', 'a') as f_out:
for line in f_in:
if 'Iraq' in line:
f_out.writelines(line.replace('Iraq', 'Malawi'))
else:
f_out.writelines(line)
```
#### Exercise 7 <a name="opdr7"></a>
With the append mode you can add text at the end of an existing text file. Repeat Exercise 6 but now use the append mode to add the string to the file.
In the example above we
```
```
<a href="#antw7">Answer exercise 7</a>
### [5. Higher level reading/writing (NOT YET AVAILABLE)](#top)<a id="5"></a>
## [6. Answers](#top)<a id="6"></a>
#### <a href="#opdr1">Answer exercise 1</a> <a name="antw1"></a>
Don't forget to close the file after you're done reading!
```
f = open('file2.txt')
print(f.read())
f.close()
```
#### <a href="#opdr2">Answer Exercise 2</a> <a name="antw2"></a>
```
with open('file2.txt') as f:
data = f.read()
```
#### <a href="#opdr3">Answer exercise 3</a> <a name="antw3"></a>
Replace `<your_file_path>` in the code below with your full file path.
```
with open(r'<your_file_path>') as f:
data = f.read()
print(data)
```
#### <a href="#opdr4">Answer exercise 4</a> <a name="antw4"></a>
```
with open(r'..\..\Basic\basic3_arrays\xypoints.dat') as f:
data = f.read()
print(data)
```
#### <a href="#opdr5">Answer exercise 5</a> <a name="antw5"></a>
In the answer below we use the relative directory to write the file `readme.md`. You can also use the absolute path of course.
Note: because the string has many characters we use the `\` at the end of the string to indicate that the string continues on the following line.
```
with open(r'..\..\personal_application\readme.md', 'w') as f:
f.write("this folder contains my personal application. In the Jupyter Notebook"\
"'eigen_toepassing' you can find more information.")
```
#### <a href="#opdr6">Answer exercise 6</a> <a name="antw6"></a>
```
with open(r'my_diary.txt', 'r') as f:
txt = f.read()
new_txt = txt + ' Then I went back for seconds.'
with open(r'my_diary_mod.txt', 'w') as f:
f.write(new_txt)
```
#### <a href="#opdr7">Answer exercise 7</a> <a name="antw7"></a>
Note that when you use the append method there is no way of renaming the file. If we rerun this cell the text will be added once more.
```
with open(r'my_diary.txt', 'a') as f:
f.write(' Then I went back for seconds.')
```
#### <a href="#opdr8">Answer exercise 8</a> <a name="antw8"></a>
| github_jupyter |
# Logistic Regression using Gradient Descent
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.preprocessing import PolynomialFeatures
```
## Import Data Set
```
train_data = pd.read_csv('micro_data_train.csv')
train_data.head()
train_data.info()
train_data.describe()
test_data = pd.read_csv('micro_data_test.csv')
test_data.head()
train_data_0 = train_data[train_data.accepted == 0]
train_data_1 = train_data[train_data.accepted == 1]
plt.scatter(train_data_1.iloc[:, 0], train_data_1.iloc[:, 1], marker='+', color='black')
plt.scatter(train_data_0.iloc[:, 0], train_data_0.iloc[:, 1], marker='o', color='y')
plt.xlabel('Micro Test 1')
plt.ylabel('Micro Test 2')
plt.legend(labels=['Accepted', 'Rejected'])
plt.show()
```
## Feature Separation and Normalization
```
X_train = train_data.iloc[:, :-1].values
y_train = train_data.iloc[:, -1].values
m = y_train.shape[0]
X_test = test_data.iloc[:, :].values
poly = PolynomialFeatures(6)
X_train_mapped = poly.fit_transform(X_train[:, 0:2])
X_test_mapped = poly.fit_transform(X_test)
```
## Gradient Descent
```
def sigmoid(input_var):
sigmoid_result = 1 / (1 + np.exp(-input_var))
return sigmoid_result
def safe_ln(x, minval=0.0000000001):
return np.log(x.clip(min=minval))
def compute_cost_reg(theta, X, y):
z = np.dot(X, theta)
hx = sigmoid(z)
neg_0_cost = (-1 * (1 - y)) * safe_ln((1 - hx))
pos_1_cost = (-1 * y) * np.log(hx)
cost_normal = (pos_1_cost + neg_0_cost) / m
cost_reg = (lambda_value / (2 * m)) * np.sum(np.power(theta[1:], 2))
cost_normal_sum = np.sum(cost_normal)
cost = cost_normal_sum + cost_reg
return cost.flatten()
def gradient_reg(theta, X, y):
z = np.dot(X, theta)
hx = sigmoid(z)
error_value = hx - y
error_value_final = np.matmul(error_value.T, X)
delta = error_value_final / m
delta_reg = (lambda_value / m) * theta.reshape(-1, 1).T
delta_reg[0] = 0
delta_total = np.multiply((delta + delta_reg), (alpha / m))
grad = delta_total.flatten()
return grad.flatten()
def custom_optimizer(theta, X, y, iterations):
iteration_array = np.array([itr for itr in range(iterations)])
cost_history = []
for i in range(iterations):
theta = theta - gradient_reg(theta, X, y)
cost_history.append(compute_cost_reg(theta, X, y))
return [theta, np.column_stack((iteration_array, np.asarray(cost_history)))]
def predict(theta, X, threshold=0.5):
p = sigmoid(X.dot(theta.T)) >= threshold
return p.astype('int')
```
## Tests with few lambda values
```
alpha = m
lambda_value = 1
theta_value = np.zeros(X_train_mapped.shape[1])
cost = compute_cost_reg(theta_value, X_train_mapped, y_train)
grad = gradient_reg(theta_value, X_train_mapped, y_train)
print("For lambda = 1 and theta = zeros")
print("Cost: " + str(cost))
print("Grad (First 5): " + str(grad[:5]))
lambda_value = 10
theta_value = np.ones(X_train_mapped.shape[1])
cost = compute_cost_reg(theta_value, X_train_mapped, y_train)
grad = gradient_reg(theta_value, X_train_mapped, y_train)
print("For lambda = 10 and theta = ones")
print("Cost: " + str(cost))
print("Grad (First 5): " + str(grad[:5]))
```
## Use Custom Optimizer
```
lambda_value = 1
theta_value = np.zeros(X_train_mapped.shape[1])
result_theta, cost_history = custom_optimizer(theta_value, X_train_mapped, y_train, 400)
print(result_theta)
accuracy = 100 * sum(predict(result_theta, X_train_mapped) == y_train.ravel()) / y_train.size
print(accuracy)
```
## Cost Function
```
plt.plot(cost_history)
plt.xlabel("Iteration")
plt.ylabel("Cost")
plt.show()
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# ONNX Runtime: Tutorial for Nuphar execution provider
**Accelerating model inference via compiler, using Docker Images for ONNX Runtime with Nuphar**
This example shows how to accelerate model inference using Nuphar, an execution provider that leverages just-in-time compilation to generate optimized executables.
For more background about Nuphar, please check [Nuphar-ExecutionProvider.md](https://github.com/microsoft/onnxruntime/blob/master/docs/execution_providers/Nuphar-ExecutionProvider.md) and its [build instructions](https://www.onnxruntime.ai/docs/how-to/build.html#nuphar).
#### Tutorial Roadmap:
1. Prerequistes
2. Create and run inference on a simple ONNX model, and understand how ***compilation*** works in Nuphar.
3. Create and run inference on a model using ***LSTM***, run symbolic shape inference, edit LSTM ops to Scan, and check Nuphar speedup.
4. ***Quantize*** the LSTM model and check speedup in Nuphar (CPU with AVX2 support is required).
5. Working on real models from onnx model zoo: ***BERT squad***, ***GPT-2*** and ***Bidirectional Attention Flow ([BiDAF](https://arxiv.org/pdf/1611.01603))***.
6. ***Ahead-Of-Time (AOT) compilation*** to save just-in-time compilation cost on model load.
7. Performance tuning for single thread inference.
## 1. Prerequistes
Please make sure you have installed following Python packages. Besides, C++ compiler/linker is required for ahead-of-time compilation. Please make sure you have g++ if running on Linux, or Visual Studio 2017 on Windows.
For simplicity, you may use [Nuphar docker image](https://github.com/microsoft/onnxruntime/blob/master/dockerfiles/README.md) from Microsoft Container Registry.
```
import cpufeature
import hashlib
import numpy as np
import onnx
from onnx import helper, numpy_helper
import os
from timeit import default_timer as timer
import shutil
import subprocess
import sys
import tarfile
import urllib.request
def is_windows():
return sys.platform.startswith('win')
if is_windows():
assert shutil.which('cl.exe'), 'Please make sure MSVC compiler and liner are in PATH.'
else:
assert shutil.which('g++'), 'Please make sure g++ is installed.'
def print_speedup(name, delta_baseline, delta):
print("{} speed-up {:.2f}%".format(name, 100*(delta_baseline/delta - 1)))
print(" Baseline: {:.3f} s, Current: {:.3f} s".format(delta_baseline, delta))
def create_cache_dir(cache_dir):
# remove any stale cache files
if os.path.exists(cache_dir):
shutil.rmtree(cache_dir)
os.makedirs(cache_dir, exist_ok=True)
def md5(file_name):
hash_md5 = hashlib.md5()
with open(file_name, "rb") as f:
for chunk in iter(lambda: f.read(4096), b""):
hash_md5.update(chunk)
return hash_md5.hexdigest()
```
And Nuphar package in onnxruntime is required too. Please make sure you are using Nuphar enabled build.
```
import onnxruntime
from onnxruntime.nuphar.model_editor import convert_to_scan_model
from onnxruntime.nuphar.model_quantizer import convert_matmul_model
from onnxruntime.nuphar.rnn_benchmark import generate_model, perf_test
from onnxruntime.tools.symbolic_shape_infer import SymbolicShapeInference
```
## 2. Create and run inference on a simple ONNX model
Let's start with a simple model: Y = ((X + X) * X + X) * X + X
```
model = onnx.ModelProto()
opset = model.opset_import.add()
opset.domain == 'onnx'
opset.version = 7 # ONNX opset 7 is required for LSTM op later
model.ir_version = onnx.IR_VERSION
graph = model.graph
X = 'input'
Y = 'output'
# declare graph input/ouput with shape [seq, batch, 1024]
dim = 1024
model.graph.input.add().CopyFrom(helper.make_tensor_value_info(X, onnx.TensorProto.FLOAT, ['seq', 'batch', dim]))
model.graph.output.add().CopyFrom(helper.make_tensor_value_info(Y, onnx.TensorProto.FLOAT, ['seq', 'batch', dim]))
# create nodes: Y = ((X + X) * X + X) * X + X
num_nodes = 5
for i in range(num_nodes):
n = helper.make_node('Mul' if i % 2 else 'Add',
[X, X if i == 0 else 'out_'+str(i-1)],
['out_'+str(i) if i < num_nodes - 1 else Y],
'node'+str(i))
model.graph.node.add().CopyFrom(n)
# save the model
simple_model_name = 'simple.onnx'
onnx.save(model, simple_model_name)
```
We will use nuphar execution provider to run the inference for the model that we created above, and use settings string to check the generated code.
Because of the redirection of output, we dump the lowered code from a subprocess to a log file:
```
code_to_run = '''
import onnxruntime
s = 'codegen_dump_lower:verbose'
providers = [('NupharExecutionProvider', {'nuphar_settings': s}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession('simple.onnx', providers=providers)
'''
log_file = 'simple_lower.log'
with open(log_file, "w") as f:
subprocess.run([sys.executable, '-c', code_to_run], stdout=f, stderr=f)
```
The lowered log is similar to C source code, but the whole file is lengthy to show here. Let's just check the last few lines that are most important:
```
with open(log_file) as f:
log_lines = f.readlines()
log_lines[-10:]
```
The compiled code showed that the nodes of Add/Mul were fused into a single function, and vectorization was applied in the loop. The fusion was automatically done by the compiler in the Nuphar execution provider, and did not require any manual model editing.
Next, let's run inference on the model and compare the accuracy and performance with numpy:
```
seq = 128
batch = 16
input_data = np.random.rand(seq, batch, dim).astype(np.float32)
sess = onnxruntime.InferenceSession(simple_model_name)
simple_feed = {X:input_data}
simple_output = sess.run([], simple_feed)
np_output = ((((input_data + input_data) * input_data) + input_data) * input_data) + input_data
assert np.allclose(simple_output[0], np_output)
simple_repeats = 100
start_ort = timer()
for i in range(simple_repeats):
sess.run([], simple_feed)
end_ort = timer()
start_np = timer()
for i in range(simple_repeats):
np_output = ((((input_data + input_data) * input_data) + input_data) * input_data) + input_data
end_np = timer()
print_speedup('Fusion', end_np - start_np, end_ort - start_ort)
```
## 3. Create and run inference on a model using LSTM
Now, let's take one step further to work on a 4-layer LSTM model, created from onnxruntime.nuphar.rnn_benchmark module.
```
lstm_model = 'LSTMx4.onnx'
input_dim = 256
hidden_dim = 1024
generate_model('lstm', input_dim, hidden_dim, bidirectional=False, layers=4, model_name=lstm_model)
```
**IMPORTANT**: Nuphar generates code before knowing shapes of input data, unlike other execution providers that do runtime shape inference. Thus, shape inference information is critical for compiler optimizations in Nuphar. To do that, we run symbolic shape inference on the model. Symbolic shape inference is based on the ONNX shape inference, and enhanced by sympy to better handle Shape/ConstantOfShape/etc. ops using symbolic computation.
**IMPORTANT**: When running multi-threaded inference, Nuphar currently uses TVM's parallel schedule with has its own thread pool that's compatible with OpenMP and MKLML. The TVM thread pool has not been integrated with ONNX runtime thread pool, so intra_op_num_threads won't control it. Please make sure the build is with OpenMP or MKLML, and use OMP_NUM_THREADS to control thread pool.
```
onnx.save(SymbolicShapeInference.infer_shapes(onnx.load(lstm_model)), lstm_model)
```
Now, let's check baseline performance on the generated model, using CPU execution provider.
```
sess_baseline = onnxruntime.InferenceSession(lstm_model, providers=['CPUExecutionProvider'])
seq = 128
input_data = np.random.rand(seq, 1, input_dim).astype(np.float32)
lstm_feed = {sess_baseline.get_inputs()[0].name:input_data}
lstm_output = sess_baseline.run([], lstm_feed)
```
To run RNN models in Nuphar execution provider efficiently, LSTM/GRU/RNN ops need to be converted to Scan ops. This is because Scan is more flexible, and supports quantized RNNs.
```
lstm_scan_model = 'Scan_LSTMx4.onnx'
convert_to_scan_model(lstm_model, lstm_scan_model)
```
After conversion, let's compare performance and accuracy with baseline:
```
sess_nuphar = onnxruntime.InferenceSession(lstm_scan_model)
output_nuphar = sess_nuphar.run([], lstm_feed)
assert np.allclose(lstm_output[0], output_nuphar[0])
lstm_repeats = 10
start_lstm_baseline = timer()
for i in range(lstm_repeats):
sess_baseline.run([], lstm_feed)
end_lstm_baseline = timer()
start_nuphar = timer()
for i in range(lstm_repeats):
sess_nuphar.run([], lstm_feed)
end_nuphar = timer()
print_speedup('Nuphar Scan', end_lstm_baseline - start_lstm_baseline, end_nuphar - start_nuphar)
```
## 4. Quantize the LSTM model
Let's get more speed-ups from Nuphar by quantizing the floating point GEMM/GEMV in LSTM model to int8 GEMM/GEMV.
**NOTE:** For inference speed of quantizated model, a CPU with AVX2 instructions is preferred.
```
cpufeature.CPUFeature['AVX2'] or 'No AVX2, quantization model might be slow'
```
We can use onnxruntime.nuphar.model_quantizer to quantize floating point GEMM/GEMVs. Assuming GEMM/GEMV takes form of input * weights, weights are statically quantized per-column, and inputs are dynamically quantized per-row.
```
lstm_quantized_model = 'Scan_LSTMx4_int8.onnx'
convert_matmul_model(lstm_scan_model, lstm_quantized_model)
```
Now run the quantized model, and check accuracy. Please note that quantization may cause accuracy loss, so we relax the comparison threshold a bit.
```
sess_quantized = onnxruntime.InferenceSession(lstm_quantized_model)
output_quantized = sess_quantized.run([], lstm_feed)
assert np.allclose(lstm_output[0], output_quantized[0], rtol=1e-3, atol=1e-3)
```
Now check quantized model performance:
```
start_quantized = timer()
for i in range(lstm_repeats):
sess_quantized.run([], lstm_feed)
end_quantized = timer()
print_speedup('Quantization', end_nuphar - start_nuphar, end_quantized - start_quantized)
```
To check RNN quantization performance, please use rnn_benchmark.perf_test.
```
rnn_type = 'lstm' # could be 'lstm', 'gru' or 'rnn'
num_threads = cpufeature.CPUFeature['num_physical_cores'] # no hyper thread
input_dim = 80 # size of input dimension
hidden_dim = 512 # size of hidden dimension in cell
bidirectional = True # specify RNN being bidirectional
layers = 6 # number of stacked RNN layers
seq_len = 40 # length of sequence
batch_size = 1 # size of batch
original_ms, scan_ms, int8_ms = perf_test(rnn_type, num_threads, input_dim, hidden_dim, bidirectional, layers, seq_len, batch_size)
print_speedup('Nuphar Quantization speed up', original_ms / 1000, int8_ms / 1000)
```
## 5. Working on real models
### 5.1 BERT Squad
BERT (Bidirectional Encoder Representations from Transformers) applies Transformers to language modelling. With Nuphar, we may fuse and compile the model to accelerate inference on CPU.
#### Download model and test data
```
# download BERT squad model
cwd = os.getcwd()
bert_model_url = 'https://onnxzoo.blob.core.windows.net/models/opset_10/bert_squad/download_sample_10.tar.gz'
bert_model_local = os.path.join(cwd, 'download_sample_10.tar.gz')
if not os.path.exists(bert_model_local):
urllib.request.urlretrieve(bert_model_url, bert_model_local)
with tarfile.open(bert_model_local, 'r') as f:
f.extractall(cwd)
```
#### Run symbolic shape inference
Note that this model has computations like `min(100000, seq_len)` which could be simplified to `seq_len` if we know `seq_len` is not going to be too big. We can do this by setting int_max. Besides, auto_merge is used to make sure the all nodes in the entire model could have shape inferenced by merging symbolic dims when broadcasting.
```
bert_model_dir = os.path.join(cwd, 'download_sample_10')
bert_model = os.path.join(bert_model_dir, 'bertsquad10.onnx')
bert_model_with_shape_inference = os.path.join(bert_model_dir, 'bertsquad10_shaped.onnx')
# run symbolic shape inference
onnx.save(SymbolicShapeInference.infer_shapes(onnx.load(bert_model), auto_merge=True, int_max=100000), bert_model_with_shape_inference)
```
#### Run inference on original model, using CPU execution provider, with maximum optimization
```
sess_options = onnxruntime.SessionOptions()
sess_options.graph_optimization_level = onnxruntime.GraphOptimizationLevel.ORT_ENABLE_ALL
sess_baseline = onnxruntime.InferenceSession(bert_model, sess_options=sess_options, providers=['CPUExecutionProvider'])
# load test data
test_data_dir = os.path.join(bert_model_dir, 'test_data_set_1')
tps = [onnx.load_tensor(os.path.join(test_data_dir, 'input_{}.pb'.format(i))) for i in range(len(sess_baseline.get_inputs()))]
bert_feed = {tp.name:numpy_helper.to_array(tp) for tp in tps}
bert_output_baseline = sess_baseline.run([], bert_feed)
bert_repeats = 20
start_bert_baseline = timer()
for i in range(bert_repeats):
sess_baseline.run([], bert_feed)
end_bert_baseline = timer()
```
#### Run inference on the model with symbolic shape inference, using Nuphar execution provider
First let's check accuracy:
```
sess = onnxruntime.InferenceSession(bert_model_with_shape_inference)
output = sess.run([], bert_feed)
assert all([np.allclose(o, ob, atol=1e-4) for o, ob in zip(output, bert_output_baseline)])
```
Then check speed:
```
start_nuphar = timer()
for i in range(bert_repeats):
sess.run([], bert_feed)
end_nuphar = timer()
print_speedup('Nuphar BERT squad', end_bert_baseline - start_bert_baseline, end_nuphar - start_nuphar)
```
### 5.2 GPT-2 with fixed batch size
GPT-2 is a language model using Generative Pre-Trained Transformer for text generation. With Nuphar, we may fuse and compile the model to accelerate inference on CPU.
#### Download model and test data
```
# download GPT-2 model
cwd = os.getcwd()
gpt2_model_url = 'https://onnxzoo.blob.core.windows.net/models/opset_10/GPT2/GPT-2.tar.gz'
gpt2_model_local = os.path.join(cwd, 'GPT-2.tar.gz')
if not os.path.exists(gpt2_model_local):
urllib.request.urlretrieve(gpt2_model_url, gpt2_model_local)
with tarfile.open(gpt2_model_local, 'r') as f:
f.extractall(cwd)
```
#### Change batch dimension to fixed value, and run symbolic shape inference
The GPT-2 model from model zoo has a symbolic batch dimension. By replacing it with a fixed value, compiler would be able to generate better code.
```
gpt2_model_dir = os.path.join(cwd, 'GPT2')
gpt2_model = os.path.join(gpt2_model_dir, 'model.onnx')
# edit batch dimension from symbolic to int value for better codegen
mp = onnx.load(gpt2_model)
mp.graph.input[0].type.tensor_type.shape.dim[0].dim_value = 1
onnx.save(mp, gpt2_model)
gpt2_model_with_shape_inference = os.path.join(gpt2_model_dir, 'model_shaped.onnx')
# run symbolic shape inference
onnx.save(SymbolicShapeInference.infer_shapes(onnx.load(gpt2_model), auto_merge=True), gpt2_model_with_shape_inference)
```
#### Run inference and compare accuracy/performance to CPU provider
```
sess_options = onnxruntime.SessionOptions()
sess_options.graph_optimization_level = onnxruntime.GraphOptimizationLevel.ORT_ENABLE_ALL
sess_baseline = onnxruntime.InferenceSession(gpt2_model, sess_options=sess_options, providers=['CPUExecutionProvider'])
# load test data, note the tensor proto name in data does not match model, so override it in feed
input_name = [i.name for i in sess_baseline.get_inputs()][0] # This model only has one input
test_data_dir = os.path.join(gpt2_model_dir, 'test_data_set_0')
tp = onnx.load_tensor(os.path.join(test_data_dir, 'input_0.pb'))
gpt2_feed = {input_name:numpy_helper.to_array(tp).reshape(1,-1)} # the test data missed batch dimension
gpt2_output_baseline = sess_baseline.run([], gpt2_feed)
gpt2_repeats = 100
start_gpt2_baseline = timer()
for i in range(gpt2_repeats):
sess_baseline.run([], gpt2_feed)
end_gpt2_baseline = timer()
sess = onnxruntime.InferenceSession(gpt2_model_with_shape_inference)
output = sess.run([], gpt2_feed)
assert all([np.allclose(o, ob, atol=1e-4) for o, ob in zip(output, gpt2_output_baseline)])
start_nuphar = timer()
for i in range(gpt2_repeats):
output = sess.run([], gpt2_feed)
end_nuphar = timer()
print_speedup('Nuphar GPT-2', end_gpt2_baseline - start_gpt2_baseline, end_nuphar - start_nuphar)
```
### 5.3 BiDAF with quantization
BiDAF is a machine comprehension model that uses LSTMs. The inputs to this model are paragraphs of contexts and queries, and the outputs are start/end indices of words in the contexts that answers the queries.
First let's download the model:
```
# download BiDAF model
cwd = os.getcwd()
bidaf_url = 'https://onnxzoo.blob.core.windows.net/models/opset_9/bidaf/bidaf.tar.gz'
bidaf_local = os.path.join(cwd, 'bidaf.tar.gz')
if not os.path.exists(bidaf_local):
urllib.request.urlretrieve(bidaf_url, bidaf_local)
with tarfile.open(bidaf_local, 'r') as f:
f.extractall(cwd)
```
Now let's check the performance of the CPU provider:
```
bidaf_dir = os.path.join(cwd, 'bidaf')
bidaf = os.path.join(bidaf_dir, 'bidaf.onnx')
sess_baseline = onnxruntime.InferenceSession(bidaf, providers=['CPUExecutionProvider'])
# load test data
test_data_dir = os.path.join(cwd, 'bidaf', 'test_data_set_3')
tps = [onnx.load_tensor(os.path.join(test_data_dir, 'input_{}.pb'.format(i))) for i in range(len(sess_baseline.get_inputs()))]
bidaf_feed = {tp.name:numpy_helper.to_array(tp) for tp in tps}
bidaf_output_baseline = sess_baseline.run([], bidaf_feed)
```
The context in this test data:
```
' '.join(list(bidaf_feed['context_word'].reshape(-1)))
```
The query:
```
' '.join(list(bidaf_feed['query_word'].reshape(-1)))
```
And the answer:
```
' '.join(list(bidaf_feed['context_word'][bidaf_output_baseline[0][0]:bidaf_output_baseline[1][0]+1].reshape(-1)))
```
Now put all steps together:
```
# editing
bidaf_converted = 'bidaf_mod.onnx'
onnx.save(SymbolicShapeInference.infer_shapes(onnx.load(bidaf)), bidaf_converted)
convert_to_scan_model(bidaf_converted, bidaf_converted)
# When quantizing, there's an only_for_scan option to quantize only the GEMV inside Scan ops.
# This is useful when the input dims of LSTM being much bigger than hidden dims.
# BiDAF has several LSTMs with input dim being 800/1400/etc, while hidden dim is 100.
# So unlike the LSTMx4 model above, we use only_for_scan here
convert_matmul_model(bidaf_converted, bidaf_converted, only_for_scan=True)
# inference and verify accuracy
sess = onnxruntime.InferenceSession(bidaf_converted)
output = sess.run([], bidaf_feed)
assert all([np.allclose(o, ob) for o, ob in zip(output, bidaf_output_baseline)])
```
Check performance after all these steps:
```
bidaf_repeats = 100
start_bidaf_baseline = timer()
for i in range(bidaf_repeats):
sess_baseline.run([], bidaf_feed)
end_bidaf_baseline = timer()
start_nuphar = timer()
for i in range(bidaf_repeats):
sess.run([], bidaf_feed)
end_nuphar = timer()
print_speedup('Nuphar quantized BiDAF', end_bidaf_baseline - start_bidaf_baseline, end_nuphar - start_nuphar)
```
The benefit of quantization in BiDAF is not as great as in the LSTM sample above, because BiDAF has relatively small hidden dimensions, which limited the gain from optimization inside Scan ops. However, this model still benefits from fusion/vectorization/etc.
## 6. Ahead-Of-Time (AOT) compilation
Nuphar runs Just-in-time (JIT) compilation when loading models. The compilation may lead to slow cold start. We can use create_shared script to build dll from JIT code and accelerate model loading.
```
start_jit = timer()
sess = onnxruntime.InferenceSession(bidaf_converted)
end_jit = timer()
'JIT took {:.3f} seconds'.format(end_jit - start_jit)
# use settings to enable JIT cache
bidaf_cache_dir = os.path.join(bidaf_dir, 'cache')
create_cache_dir(bidaf_cache_dir)
settings = 'nuphar_cache_path:{}'.format(bidaf_cache_dir)
providers = [('NupharExecutionProvider', {'nuphar_settings': settings}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(bidaf_converted, providers=providers)
```
Now object files of JIT code is stored in cache_dir, let's link them into dll:
```
bidaf_cache_versioned_dir = os.path.join(bidaf_cache_dir, os.listdir(bidaf_cache_dir)[0])
# use onnxruntime.nuphar.create_shared module to create dll
subprocess.run([sys.executable, '-m', 'onnxruntime.nuphar.create_shared', '--input_dir', bidaf_cache_versioned_dir], check=True)
os.listdir(bidaf_cache_versioned_dir)
```
Check the model loading speed-up with AOT dll:
```
start_aot = timer()
settings = 'nuphar_cache_path:{}'.format(bidaf_cache_dir)
providers = [('NupharExecutionProvider', {'nuphar_settings': settings}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(bidaf_converted, providers=providers)
end_aot = timer()
print_speedup('AOT', end_jit - start_jit, end_aot - start_aot)
```
Moreover, Nuphar AOT also supports:
* Generate JIT cache with AVX/AVX2/AVX-512 and build a AOT dll including support for all these CPUs, which makes deployment easier when targeting different CPUs in one package.
* Bake model checksum into AOT dll to validate model with given AOT dll.
```
# create object files for different CPUs
cache_dir = os.path.join(os.getcwd(), 'lstm_cache')
model_name = lstm_quantized_model
model_checksum = md5(model_name)
repeats = lstm_repeats
feed = lstm_feed
time_baseline = end_lstm_baseline - start_lstm_baseline
multi_isa_so = 'avx_avx2_avx512.so'
create_cache_dir(cache_dir)
settings = 'nuphar_cache_path:{}'.format(cache_dir)
for isa in ['avx512', 'avx2', 'avx']:
settings_with_isa = settings + ', nuphar_codegen_target:' + isa
providers = [('NupharExecutionProvider', {'nuphar_settings': settings_with_isa}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(model_name, providers=providers)
cache_versioned_dir = os.path.join(cache_dir, os.listdir(cache_dir)[0])
# link object files to AOT dll
subprocess.run([sys.executable, '-m', 'onnxruntime.nuphar.create_shared', '--input_dir', cache_versioned_dir, '--input_model', model_name, '--output_name', multi_isa_so], check=True)
# now load the model with AOT dll
# NOTE: when nuphar_codegen_target is not set, it defaults to current CPU ISA
settings = 'nuphar_cache_path:{}, nuphar_cache_so_name:{}, nuphar_cache_model_checksum:{}, nuphar_cache_force_no_jit:on'.format(cache_dir, multi_isa_so, model_checksum)
providers = [('NupharExecutionProvider', {'nuphar_settings': settings}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(model_name, providers=providers)
# force to a different ISA which is a subset of current CPU
# NOTE: if an incompatible ISA is used, exception on invalid instructions would be thrown
for valid_isa in ['avx2', 'avx']:
settings_with_isa = 'nuphar_cache_path:{}, nuphar_cache_so_name:{}, nuphar_cache_model_checksum:{}, nuphar_codegen_target:{}, nuphar_cache_force_no_jit:on'.format(cache_dir, multi_isa_so, model_checksum, valid_isa)
providers = [('NupharExecutionProvider', {'nuphar_settings': settings_with_isa}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(model_name, providers=providers)
start_nuphar = timer()
for i in range(repeats):
sess.run([], feed)
end_nuphar = timer()
print_speedup('{} in {}'.format(model_name, valid_isa), time_baseline, end_nuphar - start_nuphar)
```
## 7. Performance tuning for single thread inference.
By default, Nuphar enables parallel schedule for lower inference latency with multiple threads, when building with MKLML or OpenMP. For some models, user may want to run single-thread inference for better throughput with multiple concurrent inference threads, and turning off parallel schedule may make inference a bit faster in single thread.
```
# set OMP_NUM_THREADS to 1 for single thread inference
# this would mak
os.environ['OMP_NUM_THREADS'] = '1'
sess = onnxruntime.InferenceSession(bidaf_converted)
start_baseline = timer()
for i in range(bidaf_repeats):
sess_baseline.run([], bidaf_feed)
end_baseline = timer()
# use NUPHAR_PARALLEL_MIN_WORKLOADS=0 to turn off parallel schedule, using settings string
# it can be set from environment variable too: os.environ['NUPHAR_PARALLEL_MIN_WORKLOADS'] = '0'
settings = 'nuphar_parallel_min_workloads:0'
providers = [('NupharExecutionProvider', {'nuphar_settings': settings}), 'CPUExecutionProvider']
sess = onnxruntime.InferenceSession(bidaf_converted, providers=providers)
start = timer()
for i in range(bidaf_repeats):
sess_baseline.run([], bidaf_feed)
end = timer()
print_speedup('Single thread perf w/o parallel schedule', end_baseline - start_baseline, end - start)
del os.environ['OMP_NUM_THREADS']
```
| github_jupyter |
# Creating a Rock Paper Scissor Game
## 1. Load Required Libraries
```
import keras
import os
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Conv2D, MaxPool2D, Dense, Flatten , Dropout
from keras.models import Sequential
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint
from keras.models import load_model
import cv2
import copy
import time
```
## 2. Prepare data
```
train_dir = "rps/rps"
test_dir = "rps-test-set/rps-test-set"
train_rock = os.listdir(train_dir +"/rock")
train_paper = os.listdir(train_dir +"/paper")
train_scissors = os.listdir(train_dir +"/scissors")
test_rock = os.listdir(train_dir +"/rock")
test_scissors = os.listdir(train_dir +"/scissors")
test_paper = os.listdir(train_dir +"/paper")
print("Number of images in the train-set:", len(train_rock) + len(train_paper) + len(train_scissors))
print("Number of images in the test-set:", len(test_rock) + len(test_paper) + len(test_scissors))
print("\nNumber of rocks in the train-set:", len(train_rock))
print("Number of papers in the train-set:", len(train_paper))
print("Number of scissors in the train-set:", len(train_scissors))
print("\nNumber of rocks in the test-set:", len(test_rock))
print("Number of papers in the test-set:", len(test_paper))
print("Number of scissors in the test-set:", len(test_scissors))
```
### Image Augmentation using Keras
```
train_datagen= ImageDataGenerator(rescale= 1/255.0,
width_shift_range =0.2 ,
height_shift_range=0.2 ,
zoom_range=0.2 ,
rotation_range=40 ,
shear_range=0.2,
horizontal_flip= True)
test_datagen = ImageDataGenerator(rescale= 1/255.0)
train_generator = train_datagen.flow_from_directory(train_dir, target_size=(75,75), class_mode="categorical", batch_size=128)
test_generator = test_datagen.flow_from_directory(test_dir, target_size=(75,75), class_mode="categorical", batch_size=128)
```
## 3. Build Convolution Neural Net Model
```
model = Sequential()
model.add(Conv2D(64, (3,3), activation="relu", input_shape=(75,75,3), padding="same"))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(64, (3,3), activation="relu", padding="same"))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(128, (3,3), activation="relu", padding="same"))
model.add(MaxPool2D((2,2)))
model.add(Conv2D(128, (3,3), activation="relu", padding="same"))
model.add(MaxPool2D((2,2)))
model.add(Flatten())
model.add(Dense(512, activation="relu"))
model.add(Dropout(0.2))
model.add(Dense(512, activation= "relu"))
model.add(Dropout(0.2))
model.add(Dense(3, activation="softmax"))
model.summary()
## Create list of keras callbacks
my_callbacks_es = EarlyStopping(monitor='val_accuracy', patience=5, verbose=1)
my_callbacks_rlr = ReduceLROnPlateau(monitor='val_accuracy', pateince=2, factor=0.5, min_lr=0.00001, verbose=1)
my_callbacks_mc = ModelCheckpoint("model.h5",monitor='val_accuracy', save_best_only=True, verbose=1, mode='max')
my_callbacks = [my_callbacks_es, my_callbacks_rlr, my_callbacks_mc]
## compile model
model.compile(loss= "categorical_crossentropy", optimizer='adam', metrics=['accuracy'])
## train model
history= model.fit(train_generator, validation_data=test_generator,
epochs = 25, verbose=1, steps_per_epoch= 20, validation_steps=3,
callbacks= my_callbacks)
## load our trained model
from keras.models import load_model
model = load_model("model.h5")
```
## 4. Using OpenCV to test model with real time video from webcam.
```
## start webcam
cap = cv2.VideoCapture(0)
start = True
while True:
ret, frame = cap.read()
img = copy.deepcopy(frame)
cv2.rectangle(img, (20,50), (320, 350), (255,0,0), 3)
roi = frame[40:340, 10:310]
input_img = cv2.resize(roi, (75,75))
input_img = np.expand_dims(input_img, axis=0)
list1 = ['PAPER', "ROCK", "SCISSORS"]
array= model.predict(input_img)
id= np.argmax(array)
text = list1[id]
cv2.putText(img, "Put your hand in the box", (20,30), cv2.FONT_HERSHEY_SIMPLEX, 0.5 , (0,0,255), 2)
cv2.putText(img, text, (20,380), cv2.FONT_HERSHEY_SIMPLEX, 0.5 , (0,255,255), 2)
cv2.imshow("frame", img)
key = cv2.waitKey(1) & 0xFF
if key== ord('q'):
break
cap.release()
cv2.destroyAllWindows()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import wfdb
import ast
from sklearn.preprocessing import StandardScaler, MultiLabelBinarizer
import os
from sklearn.utils import shuffle
import math
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler,normalize, MinMaxScaler
import os
import wandb
from sklearn.metrics import roc_auc_score, classification_report, accuracy_score
import warnings
### Preprocessing
# Using the super classes, multi label classification, excluding samples with no labels and considering atleast one label
path = 'ptb/'
Y = pd.read_csv(path+ 'ptbxl_database.csv', index_col = 'ecg_id')
data = np.array([wfdb.rdsamp(path+f)[0] for f in Y.filename_lr])
Y.scp_codes = Y.scp_codes.apply(lambda x: ast.literal_eval(x))
agg_df = pd.read_csv(path+ 'scp_statements.csv', index_col = 0)
agg_df = agg_df[agg_df.diagnostic == 1]
def agg(y_dic):
temp =[]
for key in y_dic.keys():
if key in agg_df.index:
c = agg_df.loc[key].diagnostic_class
if str(c) != 'nan':
temp.append(c)
return list(set(temp))
Y['diagnostic_superclass'] = Y.scp_codes.apply(agg)
Y['superdiagnostic_len'] = Y['diagnostic_superclass'].apply(lambda x: len(x))
#########
counts = pd.Series(np.concatenate(Y.diagnostic_superclass.values)).value_counts()
Y['diagnostic_superclass'] = Y['diagnostic_superclass'].apply(lambda x: list(set(x).intersection(set(counts.index.values))))
X_data = data[Y['superdiagnostic_len'] >= 1]
Y_data = Y[Y['superdiagnostic_len'] >= 1]
mlb = MultiLabelBinarizer()
mlb.fit(Y_data['diagnostic_superclass'])
y = mlb.transform(Y_data['diagnostic_superclass'].values)
########
## Stratify split
X_train = X_data[Y_data.strat_fold < 9]
y_train = y[Y_data.strat_fold < 9]
X_val = X_data[Y_data.strat_fold == 9]
y_val = y[Y_data.strat_fold == 9]
X_test = X_data[Y_data.strat_fold == 10]
y_test = y[Y_data.strat_fold == 10]
del X_data, Y_data, y
#########
# Standardizing
def apply_scaler(X, scaler):
X_tmp = []
for x in X:
x_shape = x.shape
X_tmp.append(scaler.transform(x.flatten()[:,np.newaxis]).reshape(x_shape))
X_tmp = np.array(X_tmp)
return X_tmp
scaler = StandardScaler()
scaler.fit(np.vstack(X_train).flatten()[:,np.newaxis].astype(float))
X_train_scale = apply_scaler(X_train, scaler)
X_test_scale = apply_scaler(X_test, scaler)
X_val_scale = apply_scaler(X_val, scaler)
del X_train, X_test, X_val
## Shuffling
X_train_scale, y_train = shuffle(X_train_scale, y_train, random_state = 42)
class DataGen(tf.keras.utils.Sequence):
def __init__(self, X, y,batch_size = 16):
self.batch_size = batch_size
self.X = X
self.y = y
def __len__(self):
return math.ceil(len(self.X) / self.batch_size)
def __getitem__(self,idx):
X_full = self.X[idx * self.batch_size:(idx + 1) *self.batch_size]
batch_y = self.y[idx * self.batch_size:(idx + 1) *self.batch_size]
return np.transpose(X_full[..., np.newaxis], (0, 2, 1, 3)) ,batch_y
## Params
batch_size = 32
train_gen = DataGen(X_train_scale, y_train, batch_size = batch_size)
test_gen = DataGen(X_test_scale, y_test, batch_size = batch_size)
test = train_gen[0][0].shape
print(test)
import tensorflow.keras.backend as K
class attention(tf.keras.layers.Layer):
def __init__(self, return_sequences = False, dim = 32, **kwargs):
self.return_sequences = return_sequences
self.dim = dim
super(attention,self).__init__(**kwargs)
def build(self, input_shape):
self.W=self.add_weight(name="att_weight", shape=(input_shape[-1], self.dim),
initializer="normal")
self.b=self.add_weight(name="att_bias", shape=(input_shape[1], self.dim),
initializer="zeros")
self.V = self.add_weight(name = "Vatt", shape = (self.dim, 1), initializer = "normal")
super(attention,self).build(input_shape)
def call(self, x):
e = K.tanh(K.dot(x,self.W)+self.b)
e = K.dot(e, self.V)
a = K.softmax(e, axis=1)
output = x*a
if self.return_sequences :
return output, a
return K.sum(output, axis=1), a
def get_config(self):
base_config = super().get_config()
config = {"return sequences" : tf.keras.initializers.serialize(self.return_sequences), "att dim" : tf.keras.initializers.serialize(self.dim)}
return dict(list(base_config.items()) + list(config.items()))
## Resnet blocks
def relu_bn(inputs: tf.Tensor) -> tf.Tensor:
dp = Dropout(0.5)(inputs)
relu = ReLU()(dp)
bn = BatchNormalization()(relu)
return bn
def residual_block(x: tf.Tensor, downsample: bool, filters: int, kernel_size: int = 12) -> tf.Tensor:
y = Conv1D(kernel_size=kernel_size,
strides= (1 if not downsample else 2),
filters=filters,
padding="same")(x)
y = relu_bn(y)
y = Conv1D(kernel_size=kernel_size,
strides=1,
filters=filters,
padding="same")(y)
if downsample:
x = Conv1D(kernel_size=1,
strides=2,
filters=filters,
padding="same")(x)
out = Add()([x, y])
out = relu_bn(out)
return out
## Params
sig_len = 1000
beat_size = 50
from tensorflow.keras.layers import Conv1D, Input, Attention, LSTM, Activation, Dense, Average,ReLU, BatchNormalization,Add, Reshape, Bidirectional, Concatenate
num_channel = 12
num_filters = 32
num_blocks_list = [2, 2, 2]
inputs = Input(shape = (num_channel, sig_len, 1), batch_size = None)
#### Beat Level
x = K.reshape(inputs, (-1, beat_size,1 ))
x = Conv1D(32 ,12 ,padding = 'same')(x)
x = Activation('relu')(x)
for i in range(len(num_blocks_list)):
num_blocks = num_blocks_list[i]
for j in range(num_blocks):
x = residual_block(x, downsample=(j==0 and i!=0), filters=num_filters)
num_filters *= 2
x, _ = attention(name = "beat_att")(x)
##### Rhythm level
x = K.reshape(x,(-1, int(sig_len/beat_size) , 64))
x = Bidirectional(LSTM(32, return_sequences = True))(x)
x, _ = attention(name = "rhythm_att")(x)
#### Channel level
x = K.reshape(x, (-1, num_channel, 64))
x, _ = attention(name = "channel_att")(x)
outputs = Dense(5, activation = 'sigmoid')(x)
model = tf.keras.models.Model(inputs = inputs, outputs = outputs)
model.compile(optimizer = tf.keras.optimizers.Adam(learning_rate = 0.001), loss = tf.keras.losses.BinaryCrossentropy(), metrics = ['accuracy',tf.keras.metrics.AUC(multi_label = True)])
model.summary()
if not os.path.exists('3_level_att_saves'):
os.mkdir('3_level_att_saves')
wandb.init(project = '3_level_att', name = 'run_1')
### Accuracy metric
def metrics(y_true, y_scores):
y_pred = y_scores >= 0.5
acc = np.zeros(y_pred.shape[-1])
for i in range(y_pred.shape[-1]):
acc[i] = accuracy_score(y_true[:,i], y_pred[:,i])
return acc, np.mean(acc)
## Callback for logging and metrics
class model_checkpoint(tf.keras.callbacks.Callback):
def __init__(self, filepath, gen, monitor='loss', options=None, **kwargs):
super().__init__()
self.filepath = filepath
self.monitor = monitor
self.test_data = gen
def on_epoch_end(self, epoch, logs = {}) :
test_len = len(self.test_data)
score = []
gt =[]
for i in range(test_len):
X,y = self.test_data[i][0], self.test_data[i][1]
temp_score = self.model.predict(X)
score.append(temp_score)
gt.append(y)
score = np.concatenate(score, axis = 0)
gt = np.concatenate(gt, axis = 0)
roc_auc = roc_auc_score(gt, score, average = 'macro')
_, accuracy = metrics(gt, score)
temp_path = f"{epoch+1}_roc_{roc_auc:.4f}.h5"
path = os.path.join(self.filepath, temp_path)
if epoch > 5 :
self.model.save_weights(path)
wandb.log({'train_loss' : logs['loss'], 'epoch' : epoch})
wandb.log({'train_keras_auroc' : logs.get(self.monitor), 'epoch' : epoch})
wandb.log({'test_loss' : logs['val_loss'], 'epoch' : epoch})
wandb.log({'test_keras_auroc' : logs['val_auc'], 'epoch' : epoch})
wandb.log({'test_roc_score' : roc_auc, 'epoch' : epoch})
wandb.log({'test_accuracy_score' : accuracy, 'epoch' : epoch})
logs['val_roc_auc'] = roc_auc
logs['val_accuracy_score'] = accuracy
def set_model(self, model):
self.model = model
metric = 'auc'
checkpoint_filepath = '3_level_att_saves'
checkpoint = model_checkpoint(checkpoint_filepath, monitor = metric, gen = test_gen )
reduce_lr = tf.keras.callbacks.ReduceLROnPlateau(
factor=0.1,
patience=10,
min_lr=0.001 * 0.001)
callbacks = [checkpoint, reduce_lr]
history = model.fit(train_gen, epochs = 60, validation_data = test_gen, workers = 5 )
path_weights = r'49_roc_0.9216.h5'
model.load_weights(path_weights)
test_gen = DataGen(X_test_scale, y_test, batch_size = len(y_test))
pred = model.predict(test_gen[0][0])
roc_auc_score(y_test, pred, average='macro')
### Accuracy metric
def metrics(y_true, y_scores):
y_pred = y_scores >= 0.5
acc = np.zeros(y_pred.shape[-1])
for i in range(y_pred.shape[-1]):
acc[i] = accuracy_score(y_true[:,i], y_pred[:,i])
return acc, np.mean(acc)
acc, mean_acc = metrics(y_test, pred)
print(f'class wise accuracy: {acc}')
print(f'accuracy: {mean_acc}')
### Class wise AUC
roc_score = roc_auc_score(y_test, pred, average='macro')
print(f'roc_score : {roc_score}')
def AUC(y_true: np.ndarray, y_pred: np.ndarray, verbose=False) -> float:
"""Computes the macro-average AUC score.
Args:
y_true (np.ndarray): list of labels
y_pred (np.ndarray): list of predicted probabilities
Returns:
float: macro-average AUC score.
"""
aucs = []
assert len(y_true.shape) == 2 and len(y_pred.shape) == 2, 'Predictions and labels must be 2D.'
for col in range(y_true.shape[1]):
try:
aucs.append(roc_auc_score(y_true[:, col], y_pred[:, col]))
except ValueError as e:
if verbose:
print(
f'Value error encountered for label {col}, likely due to using mixup or '
f'lack of full label presence. Setting AUC to accuracy. '
f'Original error was: {str(e)}.'
)
aucs.append((y_pred == y_true).sum() / len(y_pred))
return np.array(aucs)
class_auc = AUC(y_test, pred)
print(f'class wise AUC : {class_auc}')
pred_values = pred >= 0.5
report = classification_report(y_test, pred_values, target_names = mlb.classes_)
print(report)
def multi_threshold_precision_recall(y_true: np.ndarray, y_pred: np.ndarray, thresholds: np.ndarray) :
# Expand analysis to number of thresholds
y_pred_bin = np.repeat(y_pred[None, :, :], len(thresholds), axis=0) >= thresholds[:, None, None]
y_true_bin = np.repeat(y_true[None, :, :], len(thresholds), axis=0)
# Compute true positives
TP = np.sum(np.logical_and(y_true, y_pred_bin), axis=2)
# Compute macro-average precision handling all warnings
with np.errstate(divide='ignore', invalid='ignore'):
den = np.sum(y_pred_bin, axis=2)
precision = TP / den
precision[den == 0] = np.nan
with warnings.catch_warnings(): # for nan slices
warnings.simplefilter("ignore", category=RuntimeWarning)
av_precision = np.nanmean(precision, axis=1)
# Compute macro-average recall
recall = TP / np.sum(y_true_bin, axis=2)
av_recall = np.mean(recall, axis=1)
return av_precision, av_recall
def metric_summary(y_true: np.ndarray, y_pred: np.ndarray, num_thresholds: int = 10) :
thresholds = np.arange(0.00, 1.01, 1. / (num_thresholds - 1), float)
average_precisions, average_recalls = multi_threshold_precision_recall(
y_true, y_pred, thresholds
)
f_scores = 2 * (average_precisions * average_recalls) / (average_precisions + average_recalls)
auc = np.array(AUC(y_true, y_pred, verbose=True)).mean()
return (
f_scores[np.nanargmax(f_scores)],
auc,
f_scores,
average_precisions,
average_recalls,
thresholds
)
metric_summary(y_test, pred)
```
| github_jupyter |
# XGBoost Cross Validation
The Python wrap around XGBoots implements a scikit-learn interface and this interface, more or less, support the scikit-learn cross validation system. More, XGBoost have is own cross validation system and the Python wrap support it. In other words, we have two cross validation systems. They are partialy supported and the functionalities supported for XGBoost are not the same for LightGBM. Currently, it's a puzzle.
The example presented covers both cases. The first, step_GradientBoostingCV, call the XGBoost cross validation. The second, step_GridSearchCV, call the scikit-learn cross validation.
The data preparation is the same as for the nbex_xgb_model.ipynb example. We take only two images to speed up the process.
The 'Tune' class manages everything.
The step_GradientBoostingCV method call the XGBoost cv() function.
The step_GridSearchCV method call the scikit-learn GridSearchCV() function.
Take note that this is in development and that changes can be significant.
```
%matplotlib inline
from __future__ import print_function
import os
import os.path as osp
import numpy as np
import pysptools.ml as ml
import pysptools.skl as skl
from sklearn.model_selection import train_test_split
home_path = os.environ['HOME']
source_path = osp.join(home_path, 'dev-data/CZ_hsdb')
result_path = None
def print_step_header(step_id, title):
print('================================================================')
print('{}: {}'.format(step_id, title))
print('================================================================')
print()
# img1
img1_scaled, img1_cmap = ml.get_scaled_img_and_class_map(source_path, result_path, 'img1',
[['Snow',{'rec':(41,79,49,100)}]],
skl.HyperGaussianNB, None,
display=False)
# img2
img2_scaled, img2_cmap = ml.get_scaled_img_and_class_map(source_path, result_path, 'img2',
[['Snow',{'rec':(83,50,100,79)},{'rec':(107,151,111,164)}]],
skl.HyperLogisticRegression, {'class_weight':{0:1.0,1:5}},
display=False)
def step_GradientBoostingCV(tune, update, cv_params, verbose):
print_step_header('Step', 'GradientBoosting cross validation')
tune.print_params('input')
tune.step_GradientBoostingCV(update, cv_params, verbose)
def step_GridSearchCV(tune, params, title, verbose):
print_step_header('Step', 'scikit-learn cross-validation')
tune.print_params('input')
tune.step_GridSearchCV(params, title, verbose)
tune.print_params('output')
```
X_train and y_train sets are built
The class Tune is created with the HyperXGBClassifier estimator. It's ready for cross validation, we can call Tune methods repeatedly with differents cv hypothesis.
```
verbose = False
n_shrink = 3
snow_fname = ['img1','img2']
nosnow_fname = ['imga1','imgb1','imgb6','imga7']
all_fname = snow_fname + nosnow_fname
snow_img = [img1_scaled,img2_scaled]
nosnow_img = ml.batch_load(source_path, nosnow_fname, n_shrink)
snow_cmap = [img1_cmap,img2_cmap]
M = snow_img[0]
bkg_cmap = np.zeros((M.shape[0],M.shape[1]))
X,y = skl.shape_to_XY(snow_img+nosnow_img,
snow_cmap+[bkg_cmap,bkg_cmap,bkg_cmap,bkg_cmap])
seed = 5
train_size = 0.25
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=train_size,
random_state=seed)
start_param = {'max_depth':10,
'min_child_weight':1,
'gamma':0,
'subsample':0.8,
'colsample_bytree':0.5,
'scale_pos_weight':1.5}
# Tune can be call with HyperXGBClassifier or HyperLGBMClassifier,
# but hyperparameters and cv parameters are differents
t = ml.Tune(ml.HyperXGBClassifier, start_param, X_train, y_train)
```
We set an hypothesis and call the Gradient Boosting cross validation
```
# Step 1: Fix learning rate and number of estimators for tuning tree-based parameters
step_GradientBoostingCV(t, {'learning_rate':0.2,'n_estimators':500,'silent':1},
{'verbose_eval':False},
True)
# After reading the cross validation results we manually set n_estimator
t.p_update({'n_estimators':9})
t.print_params('output')
```
Same but this time we call the scikit-learn cross validation
```
# Step 2: Tune max_depth and min_child_weight
step_GridSearchCV(t, {'max_depth':[24,25, 26], 'min_child_weight':[1]}, 'Step 2', True)
```
Finally, the result
```
print(t.get_p_current())
```
| github_jupyter |
```
import gc
import gzip
import json
import math
import numpy as np
import pandas as pd
import pickle
import time
```
## Split Wikipedia files to more sub-files
Part 00 lines: 1378917
Part 01 lines: 1380085
Part 02 lines: 1384443
Part 03 lines: 1160277
```
config = dict()
config["part00"]=[int(1378917/40),"../data/DAWT/wiki_annotations_json_en_part_00","../data/DAWT/part00/split"]
config["part01"]=[int(1380085/40),"../data/DAWT/wiki_annotations_json_en_part_01","../data/DAWT/part01/split"]
config["part02"]=[int(1384443/40),"../data/DAWT/wiki_annotations_json_en_part_02","../data/DAWT/part02/split"]
config["part03"]=[int(1160277/40),"../data/DAWT/wiki_annotations_json_en_part_03","../data/DAWT/part03/split"]
```
Run the following cell for every DAWT part folder (e.g. part00, part01, part02, and part03).
```
for part, par in config.items():
print("------"+part+"------")
path = par[1]
split_length = par[0]
num_of_splits = 40
current_split = 1
with open(path) as fin:
fout = par[2]
lines = '' # []
start_time = time.time()
# loop over the input_file and append the lines in a list
for i, line in enumerate(fin):
lines += line
# lines.append(line)
if (i + 1) % split_length == 0 and current_split < num_of_splits:
# pickle the list after split length steps
with open(fout + "_" + str(int(i / (split_length - 1))), 'w') as f2:
f2.write(lines)
# pickle.dump(lines, f2)
print("Split at " + str(i + 1) + " lines")
lines = ''
current_split += 1
elapsed_time = time.time() - start_time
print(elapsed_time)
with open(fout + "_" + str(int(i / (split_length - 1))), 'w') as f2:
f2.write(lines)
print("Split at " + str(i + 1) + " lines")
elapsed_time = time.time() - start_time
print(elapsed_time)
```
#### Replace some of the generated files with preprocessed ones
```
import wget
# part 00
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart00&files=split_40",
out="../data/DAWT/part00/")
# part 01
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart01&files=split_1",
out="../data/DAWT/part01/")
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart01&files=split_40",
out="../data/DAWT/part01/")
# part 02
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart02&files=split_1",
out="../data/DAWT/part02/")
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart02&files=split_40",
out="../data/DAWT/part02/")
# part 03
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart03&files=split_1",
out="../data/DAWT/part03/")
wget.download("https://surfdrive.surf.nl/files/index.php/s/hiNeHPTuFZ3HtEp/download?path=%2Fkgsqa_for_unseen_domains%2FDAWT%2Fpart00&files=split_40",
out="../data/DAWT/part00/")
```
| github_jupyter |
# Note
This notebook can be run on google colab for improved performance. The code changes necessary for running on this system are commented over the code.
## Data preprocessing
```
# ! pip install sentence_transformers==0.4.0
import pandas as pd
import sys
from sklearn.model_selection import train_test_split
from sentence_transformers import SentencesDataset, SentenceTransformer, InputExample, losses
from sentence_transformers.evaluation import LabelAccuracyEvaluator
from torch import nn, Tensor
from typing import Iterable, Dict
from torch.utils.data import DataLoader
import math
# from google.colab import drive
# drive.mount('/content/drive')
def country_labeled_sentences(excel_map):
result = {}
sent_num = 0
for country, dataframe in excel_map.items():
new_sents_col = dataframe["Sentence"].dropna()
new_labels_col= dataframe["Primary Instrument"].dropna()
sentences = list(new_sents_col.apply(lambda x: x.replace("\n", "").strip()))
label_col = new_labels_col.apply(lambda x: x.replace("(PES)", "").replace("(Bond)", "").strip())
labels = [[string.strip() for string in label.split(", ")][0] for label in label_col]
result[country] = {}
for sent, label in zip(sentences, labels):
if sent_num not in result[country]:
result[country][sent_num] = {"text": sent, "labels": [label]}
else:
result[country][sent_num]["text"] = sent
result[country][sent_num]["labels"] = [label]
sent_num += 1
return result
def merge_labels(all_labels, labels_to_merge):
return [f"{labels_to_merge[0]} & {labels_to_merge[1]}" if label in labels_to_merge else label for label in all_labels]
# Reading data from excel
data_excel = pd.read_excel("../../input/WRI_Policy_Tags.xlsx", engine="openpyxl", sheet_name=None)
# data_excel = pd.read_excel("/content/drive/MyDrive/WRI-LatinAmerica-Talent/Cristina_Policy_Files/WRI_Policy_Tags.xlsx", engine="openpyxl", sheet_name=None)
# Formatting the data
all_labeled_sentences = country_labeled_sentences(data_excel)
labeled_sents = dict()
for sents in all_labeled_sentences.values():
labeled_sents.update(sents)
# Fitlering out General Incentive and Unknown sentences
filtered_sents_maps = [sent for sent in labeled_sents.values() if sent['labels'][0] not in ["General incentive", "Unknown"]]
all_sents = [sent['text'] for sent in filtered_sents_maps]
all_labels = [sent['labels'][0] for sent in filtered_sents_maps]
all_labels = merge_labels(all_labels, ["Credit", "Guarantee"])
label_names = list(set(all_labels))
label_names
```
## Fine-tuning the embedding model on the labeled data
### Something we can try out:
https://www.sbert.net/examples/training/data_augmentation/README.html#extend-to-your-own-datasets
### Links:
https://github.com/UKPLab/sentence-transformers/issues/350
https://omoindrot.github.io/triplet-loss
### Possible tasks for fine-tuning:
1) Given a pair of sentence embeddings, do they belong to the same category (binary)?
2) Given a sentence and a category embedding, does the sentence belong to the category (binary)?
3) Given a sentence embedding, use a classifier to predict its category (multiclass) [https://github.com/UKPLab/sentence-transformers/blob/master/examples/training/nli/training_nli.py](https://github.com/UKPLab/sentence-transformers/blob/master/examples/training/nli/training_nli.py)
4) Use a triplet loss approach such that sentences (texts) that have the same labels will become close in vector space, while sentences with a different label will be further away [https://github.com/UKPLab/sentencetransformers/blob/master/examples/training/other/training_batch_hard_trec_continue_training.py](https://github.com/UKPLab/sentencetransformers/blob/master/examples/training/other/training_batch_hard_trec_continue_training.py)
#### In this notebook **task number 3** is used to fine-tune the model.
```
# Train test split stratified
X_train, X_test, y_train, y_test = train_test_split(all_sents, all_labels, test_size=0.15, stratify=all_labels, random_state=42)
# Define model to fine-tune
model = SentenceTransformer('stsb-xlm-r-multilingual')
# model = SentenceTransformer('xlm-r-100langs-bert-base-nli-stsb-mean-tokens')
class SoftmaxClassifier(nn.Module):
"""
This loss adds a softmax classifier on top of the output of the transformer network.
It takes a sentence embedding and learns a mapping between it and the corresponding category.
:param model: SentenceTransformer model
:param sentence_embedding_dimension: Dimension of your sentence embeddings
:param num_labels: Number of different labels
"""
def __init__(self,
model: SentenceTransformer,
sentence_embedding_dimension: int,
num_labels: int):
super(SoftmaxClassifier, self).__init__()
self.model = model
self.num_labels = num_labels
self.classifier = nn.Linear(sentence_embedding_dimension, num_labels)
def forward(self, sentence_features: Iterable[Dict[str, Tensor]], labels: Tensor):
# Get batch sentence embeddings
features = self.model(sentence_features[0])['sentence_embedding']
# Get batch loss
output = self.classifier(features)
loss_fct = nn.CrossEntropyLoss()
if labels is not None:
loss = loss_fct(output, labels.view(-1))
return loss
else:
return features, output
# Load data samples into batches
train_batch_size = 16
label2int = dict(zip(label_names, range(len(label_names))))
train_samples = []
for sent, label in zip(X_train, y_train):
label_id = label2int[label]
train_samples.append(InputExample(texts=[sent], label=label_id))
train_dataset = SentencesDataset(train_samples, model=model)
train_dataloader = DataLoader(train_dataset, shuffle=True, batch_size=train_batch_size)
# Define the way the loss is computed
classifier = SoftmaxClassifier(model=model, sentence_embedding_dimension=model.get_sentence_embedding_dimension(), num_labels=len(label2int))
# Configure the dev set evaluator - still need to test whether this works
dev_samples = []
for sent, label in zip(X_test, y_test):
label_id = label2int[label]
dev_samples.append(InputExample(texts=[sent], label=label_id))
dev_dataset = SentencesDataset(dev_samples, model=model)
dev_dataloader = DataLoader(dev_dataset, shuffle=True, batch_size=train_batch_size)
dev_evaluator = LabelAccuracyEvaluator(dataloader=dev_dataloader, softmax_model=classifier, name='lae-dev')
# Configure the training
num_epochs = 1
warmup_steps = math.ceil(len(train_dataset) * num_epochs / train_batch_size * 0.1) # 10% of train data for warm-up
model_save_path = "../../output/FineTuning"
# model_save_path = "/content/drive/MyDrive/WRI-LatinAmerica-Talent/Modeling/FineTuning"
# Train the model
model.fit(train_objectives=[(train_dataloader, classifier)],
evaluator=dev_evaluator,
epochs=num_epochs,
evaluation_steps=1000,
warmup_steps=warmup_steps,
output_path=model_save_path
)
# Load the saved model and obtain random sentence embedding
load_model = SentenceTransformer(model_save_path)
load_model.encode(all_sents[0])
```
| github_jupyter |
<a href="http://cocl.us/pytorch_link_top">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
</a>
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
<h1>Loss Function with a High Condition Number with and Without Momentum</h1>
<h2>Table of Contents</h2>
<p>In this lab, we will generate data that will produce a Loss Function with a High Condition Number. You will create two models; one with the momentum term and one without the momentum term.</p>
<ul>
<li><a href="#Makeup_Data">Make Some Data </a></li>
<li><a href="#Model_Cost">Create two Models, Two Optimizers and a Cost Function</a></li>
<li><a href="#BGD">Train the Model: Batch Gradient Descent</a></li>
</ul>
<p>Estimated Time Needed: <strong>30 min</strong></p>
<hr>
<h2>Preparation</h2>
We'll need the following libraries:
```
# Import the libraries we need for this lab
import torch
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
from torch.utils.data import Dataset, DataLoader
from torch import nn, optim
torch.manual_seed(1)
```
The class <code>plot_error_surfaces</code> is just to help you visualize the data space and the parameter space during training and has nothing to do with Pytorch.
```
# Define the class for plot out the surface
class plot_error_surfaces(object):
# Constructor
def __init__(self, w_range, b_range, X, Y, n_samples=30, go=True):
W = np.linspace(-w_range, w_range, n_samples)
B = np.linspace(-b_range, b_range, n_samples)
w, b = np.meshgrid(W, B)
Z = np.zeros((n_samples, n_samples))
count1 = 0
self.y = Y.numpy()
self.x = X.numpy()
for w1, b1 in zip(w, b):
count2 = 0
for w2, b2 in zip(w1, b1):
Z[count1, count2] = np.mean((self.y - w2 * self.x + b2) ** 2)
count2 += 1
count1 += 1
self.Z = Z
self.w = w
self.b = b
self.LOSS_list = {}
# Setter
def set_para_loss(self, model, name, loss):
if (not (name in self.LOSS_list)):
self.LOSS_list[name] = []
w = list(model.parameters())[0].item()
b = list(model.parameters())[1].item()
self.LOSS_list[name].append({"loss": loss, "w": w, "b": b})
# Plot the diagram
def plot_ps(self, iteration=0):
plt.contour(self.w, self.b, self.Z)
count = 1
if (len(self.LOSS_list) > 0):
for key, value in self.LOSS_list.items():
w = [v for d in value for (k, v) in d.items() if "w" == k]
b = [v for d in value for (k, v) in d.items() if "b" == k]
plt.scatter(w, b, cmap='viridis', marker='x', label=key)
plt.title('Loss Surface Contour not to scale, Iteration: ' + str(iteration))
plt.legend()
plt.xlabel('w')
plt.ylabel('b')
plt.show()
```
<!--Empty Space for separating topics-->
<h2 id="Makeup_Data">Make Some Data</h2>
Generate values from -2 to 2 that create a line with a slope of 0.1 and a bias of 10000. This is the line that you need to estimate. Add some noise to the data:
```
# Define a class to create the dataset
class Data(Dataset):
# Constructor
def __init__(self):
self.x = torch.arange(-2, 2, 0.1).view(-1, 1)
self.f = 1 * self.x + 10000
self.y = self.f + 0.1 * torch.randn(self.x.size())
self.len = self.x.shape[0]
# Getter
def __getitem__(self, index):
return self.x[index], self.y[index]
# Get Length
def __len__(self):
return self.len
```
Create a dataset object:
```
# Create a dataset object
dataset = Data()
```
Plot the data
```
# Plot the data
plt.plot(dataset.x.numpy(), dataset.y.numpy(), 'rx', label='y')
plt.plot(dataset.x.numpy(), dataset.f.numpy(), label='f')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
```
<!--Empty Space for separating topics-->
<h2 id="Model_Cost">Create the Model and Total Loss Function (Cost)</h2>
Create a linear regression class
```
# Define linear regression class
class linear_regression(nn.Module):
# Constructor
def __init__(self, input_size, output_size):
super(linear_regression, self).__init__()
self.linear = nn.Linear(input_size, output_size)
# Prediction
def forward(self, x):
yhat = self.linear(x)
return yhat
```
We will use PyTorch's build-in function to create a criterion function; this calculates the total loss or cost
```
# Use the build-in function to create a criterion function
criterion = nn.MSELoss()
```
Create a linear regression object, and an SGD optimizer object with no momentum.
```
# Create a linear regression object and the optimizer without momentum
model = linear_regression(1, 1)
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
Create a linear regression object, and an SGD optimiser object with momentum .
```
# Create a linear regression object and the optimizer with momentum
model_momentum = linear_regression(1, 1)
optimizer_momentum = optim.SGD(model_momentum.parameters(), lr=0.01, momentum=0.2)
```
Create a dataloader object:
```
# Create a data loader
trainloader = DataLoader(dataset=dataset, batch_size=1, shuffle=True)
```
PyTorch randomly initializes your model parameters. If we use those parameters, the result will not be very insightful as convergence will be extremely fast. In order to prevent that, we will initialize the parameters such that it will take longer to converge.
```
# Set parameters
model.state_dict()['linear.weight'][0] = -5000
model.state_dict()['linear.bias'][0] = -100000
model_momentum.state_dict()['linear.weight'][0] = -5000
model_momentum.state_dict()['linear.bias'][0] = -100000
```
Create a plotting object, not part of PyTorch, only used to help visualize
```
# Plot the surface
get_surface = plot_error_surfaces(5000, 100000, dataset.x, dataset.y, 100, go=False)
get_surface.plot_ps()
```
<!--Empty Space for separating topics-->
<h2 id="BGD">Train the Model via Stochastic Gradient Descent</h2>
Run 1 epochs of stochastic gradient descent and view parameter space.
```
# Train the model
def train_model(epochs=1):
for epoch in range(epochs):
for i, (x, y) in enumerate(trainloader):
#no momentum
yhat = model(x)
loss = criterion(yhat, y)
#momentum
yhat_m = model_momentum(x)
loss_m = criterion(yhat_m, y)
#apply optimization to momentum term and term without momentum
#for plotting
#get_surface.get_stuff(model, loss.tolist())
#get_surface.get_stuff1(model_momentum, loss_m.tolist())
get_surface.set_para_loss(model=model_momentum, name="momentum" ,loss=loss_m.tolist())
get_surface.set_para_loss(model=model, name="no momentum" , loss=loss.tolist())
optimizer.zero_grad()
optimizer_momentum.zero_grad()
loss.backward()
loss_m.backward()
optimizer.step()
optimizer_momentum.step()
get_surface.plot_ps(iteration=i)
train_model()
```
The plot above shows the different parameter values for each model in different iterations of SGD. The values are overlaid over the cost or total loss surface. The contour lines somewhat miss scaled but it is evident that in the vertical direction they are much closer together implying a larger gradient in that direction. The model trained with momentum shows somewhat more displacement in the hozontal direction.
The plot below shows the log of the cost or total loss, we see that the term with momentum converges to a minimum faster and to an overall smaller value. We use the log to make the difference more evident.
```
# Plot the loss
loss = [v for d in get_surface.LOSS_list["no momentum"] for (k, v) in d.items() if "loss" == k]
loss_m = [v for d in get_surface.LOSS_list["momentum"] for (k, v) in d.items() if "loss" == k]
plt.plot(np.log(loss), 'r', label='no momentum' )
plt.plot(np.log(loss_m), 'b', label='momentum' )
plt.title('Cost or Total Loss' )
plt.xlabel('Iterations ')
plt.ylabel('Cost')
plt.legend()
plt.show()
```
<!--Empty Space for separating topics-->
<a href="http://cocl.us/pytorch_link_bottom">
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
</a>
<h2>About the Authors:</h2>
<a href="https://www.linkedin.com/in/joseph-s-50398b136/">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/">Michelle Carey</a>, <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a">Mavis Zhou</a>
<hr>
Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
| github_jupyter |
```
#code to make pairwise rmsd heatmaps for the centroids nicely
import mdtraj as md
import glob
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.pyplot import cm
import matplotlib as mplt
from mpl_toolkits.axes_grid1 import make_axes_locatable
#getting a combined trajectory of all the apo centroids
methods = ['TICA', 'TICA_CBA', 'PCA', 'PCA_CBA', 'GROMOS', 'GROMOS_CBA']
traj = md.load('/home/jegan/final_centroids/XTAL/XTAL_0.pdb')
for method in methods:
for cen in glob.glob('/home/jegan/final_centroids/'+method+'/*.pdb'):
for num in range(10):
if str(num) in cen.split('/')[5]:
centroid = md.load(cen)
traj = centroid.join(traj)
traj = traj.superpose(traj,0)
#traj.save_pdb('/home/jegan/centroid_analysis/pairwise_rmsd/centroids/all_APO_cens.pdb')
#getting a pairwise rmsd matrix based on the rmsd of the binding atom indices
binding_atoms = [1024, 1027, 306, 1029, 1031, 1033, 2851, 1035, 1036, 1037, 1038, 1039, 1034, 1040, 1042, 1043, 1044, 1045, 1041, 2854, 1049, 2076,
311, 1060, 1061, 1062, 1072, 1074, 1076, 1087, 2112, 2116, 2120, 2122, 2123, 2126, 2135, 323, 2138, 324, 325, 1133, 1134, 1135, 1136,
1137, 2170, 330, 3206, 3207, 3208, 3210, 3323, 334, 3219, 336, 2207, 2209, 2210, 2211, 2213, 2214, 2215, 2217, 2219, 2220, 2221, 3244,
1715, 1716, 1743, 1745, 1747, 1749, 2897, 3312, 3313, 3314, 3315, 3316, 3317, 3318, 3319, 3320, 3321, 3322, 2837, 328, 329, 2899, 339,
342, 343, 350, 2911, 354, 359, 361, 362, 363, 364, 365, 366, 367, 379, 385, 386, 395, 398, 2449, 2451, 2452, 2453, 2454, 2455, 2456,
2460, 2465, 2466, 2467, 2468, 2469, 2470, 2471, 2472, 2473, 2474, 2475, 2477, 2478, 941, 2480, 2481, 946, 2834, 944, 2486, 2487, 2488,
2835, 2492, 2836, 2494, 963, 2838, 2506, 2507, 2508, 2839, 2510, 972, 978, 979, 980, 2841, 2842, 2844, 997, 2845, 1003, 2846, 1008,
1007, 1010, 1009, 2847, 1012, 1013, 1011, 1016, 1017, 1018, 1022]
print(traj.n_frames)
distances = np.empty((traj.n_frames, traj.n_frames))
for i in range(traj.n_frames):
distances[i] = md.rmsd(traj, traj, i,binding_atoms) * 10
print('Max pairwise rmsd: %f angstroms' % np.max(distances))
#putting the numpy array data into a pairwise heatmap
import matplotlib as mplt
font = {'family' : 'sans-serif',
'sans-serif': 'Arial'}
mplt.rc('font', **font)
#plotting the heatmap based of the numpy array calculated above
fig, ax = plt.subplots(figsize = (3.5,3.5)) #figsize (x,y)
im = ax.imshow(distances,cmap = cm.viridis)
#setting axis labels and vertical lines to indicate which clustering method
ax.set_xticks([0,9,18,27,37,49,60])
indx = [1,11,21,31,41,51]
for ind in indx:
ax.axvline(ind, c = 'black', linestyle = '--', linewidth = 1)
ax.set_yticks([0,7,16,26,36,46,55])
for ind in indx:
ax.axhline(ind, c = 'black', linestyle = '--', linewidth = 1)
x_labels = ['XTAL','TICA','TICA CBA','PCA','PCA CBA','GROMOS','GROMOS CBA']
y_labels = ['XTAL','TICA','TICA\nCBA','PCA','PCA\nCBA','GROMOS','GROMOS\nCBA']
ax.set_xticklabels(x_labels, rotation = 40, fontsize = 8, ha = 'right', family = 'Arial')
ax.set_yticklabels(y_labels, fontsize = 8, family = 'Arial')
ax.tick_params(axis=u'both', which=u'both',length=0)
#color bar parameters
divider = make_axes_locatable(ax)
cax = divider.new_vertical(size="5%", pad=0.05, pack_start=False)
fig.add_axes(cax)
cbar = fig.colorbar(im, cax=cax, orientation="horizontal")
cbar.ax.tick_params(labelsize=8)
cax.xaxis.set_label_position('top')
cax.xaxis.set_ticks_position('top')
cbar.set_label('RMSD ('+r'$\AA$'+')', labelpad = 5, family = 'Arial', fontsize = 8)
#saving out
plt.tight_layout()
#plt.savefig('/home/jegan/centroid_analysis/pairwise_rmsd/figs/pair_rmsd_1.png', dpi = 300)
```
| github_jupyter |
# ** DIscBIO: a user-friendly pipeline for biomarker discovery in single-cell transcriptomics**
The pipeline consists of four successive steps: data pre-processing, cellular clustering and pseudo-temporal ordering, determining differential expressed genes and identifying biomarkers.

# CTC Notebook [PART 1]
# Data Pre-processing and k-means clustering
## Required Packages
```
library(DIscBIO)
```
## Loading dataset
The "CTCdataset" dataset consisting of single migratory circulating tumor cells (CTCs) collected from patients with breast cancer. Data are available in the GEO database with accession numbers GSE51827, GSE55807, GSE67939, GSE75367, GSE109761, GSE111065 and GSE86978. The dataset should be formatted in a data frame where columns refer to samples and rows refer to genes. We provide here the possibility to load the dataset either as ".csv" or ".rda" extensions.
```
FileName<-"CTCdataset" # Name of the dataset
#CSV=TRUE # If the dataset has ".csv", the user shoud set CSV to TRUE
CSV=FALSE # If the dataset has ".rda", the user shoud set CSV to FALSE
if (CSV==TRUE){
DataSet <- read.csv(file = paste0(FileName,".csv"), sep = ",",header=T)
rownames(DataSet)<-DataSet[,1]
DataSet<-DataSet[,-1]
} else{
load(paste0(FileName,".rda"))
DataSet<-get(FileName)
}
cat(paste0("The ", FileName," contains:","\n","Genes: ",length(DataSet[,1]),"\n","cells: ",length(DataSet[1,]),"\n"))
sc<- DISCBIO(DataSet) # The DISCBIO class is the central object storing all information generated throughout the pipeline
```
## 1. Data Pre-processing
Prior to applying data analysis methods, it is standard to pre-process the raw read counts resulted from the sequencing. The preprocessing approach depends on the existence or absence of ERCC spike-ins. In both cases, it includes normalization of read counts and gene filtering.
#### Normalization of read counts
To account for RNA composition and sequencing depth among samples (single-cells), the normalization method “median of ratios” is used. This method takes the ratio of the gene instantaneous median to the total counts for all genes in that cell (column median). The gene instantaneous median is the product of multiplying the median of the total counts across all cells (row median) with the read of the target gene in each cell. This normalization method makes it possible to compare the normalized counts for each gene equally between samples.
#### Gene filtering
The key idea in filtering genes is to appoint the genes that manifest abundant variation across samples. Filtering genes is a critical step due to its dramatic impact on the downstream analysis. In case the raw data includes ERCC spike-ins, genes will be filtered based on variability in comparison to a noise level estimated from the ERCC spike-ins according to an algorithm developed by Brennecke et al (Brennecke et al., 2013). This algorithm utilizes the dependence of technical noise on the average read count and fits a model to the ERCC spike-ins. Further gene filtering can be implemented based on gene expression.
In case the raw data does not include ERCC spike-ins, genes will be only filtered based on minimum expression in certain number of cells.

### 1.2. Filtering and normalizing the raw data that does not include ERCCs
To normalize and filter the raw data that does not include ERCCs can be done by applying the function Normalizedata() and giving the parameters minexpr and minnumber some values. This function will discard cells with less than mintotal transcripts. Genes that are not expressed at minexpr transcripts in at least minnumber cells are discarded. Furthermore, it will normalize the count reads using the normalization method “median of ratios”.
To Finalize the preprocessing the function FinalPreprocessing() should be implemented by setting the parameter "GeneFlitering" to ExpF.
```
# Estimating a value for the "mintotal" parameters
# As a common practice, mintotal is set to 1000
S1<-summary(colSums(DataSet,na.rm=TRUE)) # It gives an idea about the number of reads across cells
print(S1)
# Estimating a value for the "minexpr" parameter
S2<-summary(rowMeans(DataSet,na.rm=TRUE)) # It gives an idea about the overall expression of the genes
print(S2)
minexpr= S2[3] # S2[3] is referring to the median whereas S2[4] is referring to the mean
# Estimating a value for the "minnumber" parameters
minnumber= round(length(DataSet[1,])/10) # To be expressed in at 10% of the cells.
print(minnumber)
sc<-Normalizedata(sc, mintotal=1000, minexpr=minexpr, minnumber=minnumber, maxexpr=Inf, downsample=FALSE, dsn=1, rseed=17000)
sc<-FinalPreprocessing(sc,GeneFlitering="ExpF",export = TRUE) # The GeneFiltering should be set to "ExpF"
```
## 2. Cellular Clustering and Pseudo Time ordering
Cellular clustering is performed according to the gene expression profiles to detect cellular sub-population with unique properties. After clustering, pseudo-temporal ordering is generated to indicate the cellular differentiation degree.

## 2.1. K-means Clustering
Rare cell type Identification algorithm (RaceID) was used to cluster the pre-processed data using k-means on a similarity distance matrix, which was based on Pearson correlation and the similarity matrix was computed as “1 – Pearson correlation”. The approach of the proposed clustering, i.e., applying k-means on a similarity distance matrix using the Euclidean metric, improves cluster separation. RaceID estimates the number of clusters by finding the minimal clusters' number at the saturation level of gap statistics, which standardizes the within-cluster dispersion.
The Clustexp() functions has several parameters:
- object: the outcome of running the DISCBIO() function.
- clustnr Maximum number of clusters for the derivation of the cluster number by the saturation of mean within-cluster dispersion. Default is 20.
- bootnr A numeric value of booststrapping runs for \code{clusterboot}. Default is 50.
- metric Is the method to transform the input data to a distance object.
- Metric has to be one of the following: ["spearman","pearson","kendall","euclidean","maximum","manhattan","canberra","binary","minkowski"].
- do.gap A logical vector that allows generating the number of clusters based on the gap statistics. Default is TRUE.
- SE.method The SE.method determines the first local maximum of the gap statistics.
- The SE.method has to be one of the following:["firstSEmax","Tibs2001SEmax","globalSEmax","firstmax","globalmax"]. Default is "Tibs2001SEmax"
- SE.factor A numeric value of the fraction of the standard deviation by which the local maximum is required to differ from the neighboring points it is compared to. Default is 0.25.
- B.gap Number of bootstrap runs for the calculation of the gap statistics. Default is 50
- cln Number of clusters to be used. Default is \code{NULL} and the cluster number is inferred by the saturation criterion.
- rseed Integer number. Random seed to enforce reproducible clustering results. Default is 17000.
- quiet if `TRUE`, intermediate output is suppressed

#### 2.1.1. Defining the Cells in the clusters generated by k-means clustering
```
load("SC.RData") # Loading the "SC" object that has include the data of the k-means clustering
load("Ndata.RData") # Loading the "Ndata" object and stored in the @ndata will be used to plot the expression of genes
load("expdata.RData") # Loading the "expdata" object and stored in the @expdata will be used to plot the expression of genes
sc<-SC # Storing the data of SC in the sc
sc@ndata<-Ndata
sc@expdata<-expdata
########## Removing the unneeded objects
rm(Ndata)
rm(expdata)
rm(DataSet)
rm(SC)
#sc<- Clustexp(sc,cln=4,quiet=T,clustnr=6,rseed=17000)
plotGap(sc) ### Plotting gap statistics
```
#### Cluster plotting using tSNE maps
Here you visualize the K-means clusters using t-distributed stochastic neighbor embedding (tSNE), which is a non-linear dimensionality reduction method that places neighbor cells close to each other.
```
############ Plotting the clusters
plottSNE(sc)
```
#### Cellular pseudo-time ordering based on k-means clusters
```
sc<-pseudoTimeOrdering(sc,quiet = TRUE, export = FALSE)
plotOrderTsne(sc)
```
#### Plotting the gene expression of a particular gene in a tSNE map
```
g='ENSG00000104413' #### Plotting the log expression of ESRP1
plotExptSNE(sc,g)
g='ENSG00000251562' #### Plotting the log expression of MALAT1
plotExptSNE(sc,g)
```
| github_jupyter |
ERROR: type should be string, got "https://yangtavares.com/2017/07/31/creating-a-simple-overlay-for-pynq-z1-board-from-vivado-hlx/\n\n```\n#This notebook also uses the `(some) LaTeX environments for Jupyter`\n#https://github.com/ProfFan/latex_envs wich is part of the\n#jupyter_contrib_nbextensions package\n\nfrom myhdl import *\nfrom myhdlpeek import Peeker\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\n%matplotlib inline\n\nfrom sympy import *\ninit_printing()\n\nimport itertools\n\nfrom IPython.display import clear_output\n\n\n#https://github.com/jrjohansson/version_information\n%load_ext version_information\n%version_information myhdl, myhdlpeek, numpy, pandas, matplotlib, sympy, itertools, IPython\n#helper functions to read in the .v and .vhd generated files into python\ndef VerilogTextReader(loc, printresult=True):\n with open(f'{loc}.v', 'r') as vText:\n VerilogText=vText.read()\n if printresult:\n print(f'***Verilog modual from {loc}.v***\\n\\n', VerilogText)\n return VerilogText\n\ndef VHDLTextReader(loc, printresult=True):\n with open(f'{loc}.vhd', 'r') as vText:\n VerilogText=vText.read()\n if printresult:\n print(f'***VHDL modual from {loc}.vhd***\\n\\n', VerilogText)\n return VerilogText\n\ndef ConstraintXDCTextReader(loc, printresult=True):\n with open(f'{loc}.xdc', 'r') as xdcText:\n ConstraintText=xdcText.read()\n if printresult:\n print(f'***Constraint file from {loc}.xdc***\\n\\n', ConstraintText)\n return ConstraintText\n```\n\n# IP ClockDivider\n\n## Non SoC Test\n\n```\n@block\ndef ClockDivider(Divisor, clkOut, clk,rst):\n \"\"\"\n Simple Clock Divider based on the Digilint Clock Divider\n https://learn.digilentinc.com/Documents/262\n \n Input:\n Divisor(32 bit): the clock frequncy divide by value\n clk(bool): The input clock\n rst(bool): clockDivider Reset\n \n Ouput:\n clkOut(bool): the divided clock ouput\n count(32bit): the value of the internal counter\n \"\"\"\n \n count_i=Signal(modbv(0)[32:])\n @always(clk.posedge, rst.posedge)\n def counter():\n if rst:\n count_i.next=0\n elif count_i==(Divisor-1):\n count_i.next=0\n else:\n count_i.next=count_i+1\n \n clkOut_i=Signal(bool(0))\n @always(clk.posedge, rst.posedge)\n def clockTick():\n if rst:\n clkOut_i.next=0\n elif count_i==(Divisor-1):\n clkOut_i.next=not clkOut_i\n else:\n clkOut_i.next=clkOut_i\n \n \n \n @always_comb\n def OuputBuffer():\n clkOut.next=clkOut_i\n \n return instances()\nRefClkFreq=125e6\nTargetClkFreq=40\nDivsionFactor=int(RefClkFreq/TargetClkFreq)\nDivsionFactor\nPeeker.clear()\nclk=Signal(bool(0)); Peeker(clk, 'clk')\nDivisor=Signal(intbv(DivsionFactor)[32:]); Peeker(Divisor, 'Divisor')\nclkOut=Signal(bool(0)); Peeker(clkOut, 'clkOut')\nrst=Signal(bool(0)); Peeker(rst, 'rst')\n\nDUT=ClockDivider(Divisor, clkOut, clk,rst)\nDUT.convert()\nVerilogTextReader('ClockDivider');\nConstraintXDCTextReader('ClockAXI');\n```\n\n## AXI SoC\n\nThe IP Project is `myClockDividerIP_v1_0`\nAfter adding the Verilog Clock Divider Module under sources there are two addintal modules that where created with the IP that are the AXI Lite Slave IP Connection Header `myClockDividerIP_v1_0.v` and the AXI Slave BUS controler `myClockDividerIP_v1_0_S00_AXI_inst.v`\n\n" | github_jupyter |
# Tutorial 1 - Introduction to PyRossGeo
- [Skip to **Tutorial 2: Handling PyRossGeo output**](../tutorial2-handling-PyRossGeo-output/tutorial2-handling-PyRossGeo-output.ipynb)
- [Skip to **Tutorial 3: Handling the simulation output**](../tutorial3-interventions/tutorial3-interventions.ipynb)
- [Go to the PyRossGeo documentation](https://github.com/lukastk/PyRossGeo/blob/master/docs/Documentation.md)
<b>Note: The various model parameters used in this tutorial were chosen for illustrative purposes, and are not based on figures from medical literature. Therefore the results of the simulations in the tutorial are not indicative of reality.</b>
This tutorial assumes knowledge of the PyRossGeo model. Please take a look at the [model description](https://github.com/lukastk/PyRossGeo/blob/master/docs/model.pdf) if you have not yet done so.
In this tutorial we will introduce the core functionalities of PyRossGeo. We will first simulate a single node with SIR dynamics, and then we will proceed to simulate a network of nodes with SEAIR dynamics. If you are unfamiliar with SIR dynamics, or of its numerous variations, please see [this](https://github.com/rajeshrinet/pyross/blob/master/docs/models.pdf).
```
%%capture
# Compile and import local pyrossgeo module
import os, sys
owd = os.getcwd()
os.chdir('../../')
sys.path.insert(0,'../../')
!python setup.py build_ext --inplace
os.chdir(owd)
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import pyrossgeo
import pandas as pd
import json
```
## 1.1 Age-structured SIR on a single node
The following configuration files are used in a PyRossGeo simulation:
- [model.json](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#modeljson)
- [node_parameters.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.tex.md#node_parameterscsv)
- [cnode_parameters.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#cnode_parameterscsv)
- [contact_matrices.json](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#contact_matricesjson)
- [node_cmatrices.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#node_cmatricescsv)
- [cnode_cmatrices.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#cnode_cmatricescsv)
- [node_populations.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#node_populationscsv)
- [commuter_networks.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#commuter_networkscsv)
We will go through the structure of these files briefly in this tutorial, but for more detailed descriptions please click the links above. The configuration files used for section 1.1 of this tutorial can be found [here](https://github.com/lukastk/PyRossGeo/tree/master/examples/tutorial1-introduction-to-pyrossgeo/SIR_single_node).
### 1.1.1 Defining the model ([model.json](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#modeljson))
`model.json` defines the model to use for the local epidemiological dynamics at each node. Below you can see the `model.json` for SIR.
```
with open('./SIR_single_node/model.json', 'r') as f:
print(f.read())
```
The list of all epidemiological classes are defined using in the list `"classes" : ["S", "I", "R"]`. The dynamics of each individual class is then defined using the
```json
"I" : {
"linear" : [ ["I", "-gamma"] ],
"infection" : [ ["I", "beta"] ]
},
```
construction, where each linear and each non-linear infection term are given. For example, we can read off the dynamics of the $I$ class as
$$
\dot{I}^\mu = \beta \sum_\nu C_{\mu \nu} \frac{I^\nu}{N^\nu} - \gamma I^\mu
$$
where the $\mu$ corresponds the age bracket. Indices giving the home and locations $(i,j)$ have been omitted. Note that the models are automatically given an age-structure. The age-contact structure of the simulation is defined in section 1.1.3.
The whole system of equations are:
$$
\begin{aligned}
\dot{S}^\mu & = - \beta \sum_\nu C_{\mu \nu} \frac{I^\nu}{N^\nu} S^\mu \\
\dot{I}^\mu & = \beta \sum_\nu C_{\mu \nu} \frac{I^\nu}{N^\nu} - \gamma I^\mu \\
\dot{R}^\mu & = \gamma I^\mu
\end{aligned}
$$
### 1.1.2 Setting the model parameters ([node_parameters.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.tex.md#node_parameterscsv) and [cnode_parameters.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#cnode_parameterscsv))
Next, we must define the model parameters `beta` and `gamma`. Below you can see the `node_parameters.csv` for this example.
```
pd.read_csv('./SIR_single_node/node_parameters.csv')
```
Here we have set $\beta_{ij}^\alpha = 0.01$ and $\gamma_{ij}^\alpha = 0.03$ for all nodes $(\alpha, i, j)$. We have also set the area to $A_i = 1km^2$ for all locations $i$. The area is used in PyRossGeo to scale the infectivity of the SIR dynamics with population density.
Recall that a *node* in PyRossGeo is not a geographical location, but rather the 3-tuple $(\alpha, i, j)$, corresponding to age-group, residence and location. In other words, PyRossGeo not only keeps track of how many people populate each location $j$, but also where they live, and what age they are.
The parameters of the commuterverse is set in a similar way, using `cnode_parameters.csv`.
```
pd.read_csv('./SIR_single_node/cnode_parameters.csv')
```
Since we are only modelling a single node, `cnode_parameters.csv` serves no purpose in the simulation. Regardless, the file is still required for the initialisation of the simulation.
### 1.1.3 Contact structure ([contact_matrices.json](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#contact_matricesjson), [node_cmatrices.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#node_cmatricescsv) and [cnode_cmatrices.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#cnode_cmatricescsv))
The contact matrices that are to be used in the simulation are defined in the file `contact_matrices.json`:
```
with open('./SIR_single_node/contact_matrices.json', 'r') as f:
print(f.read())
```
Here we see that we have a single contact matrix `C_main`, with 4 age groups. Let's interpret $\alpha=0,1,2,3,4$ as *child*, *young adult*, *adult* and *elderly* respectively. We can then read off from `C_main` that:
- $C^\text{home}_{\mu \mu}=10$. All age-groups have 10 encounters with members from their own group each day
- $C^\text{home}_{12}=C^\text{home}_{21}=5$. Children and young adults have 5 mutual interactions every day
- $C^\text{home}_{32}=C^\text{home}_{23}=3$. Young adults and adults have 3 mutual interactions every day
- $C^\text{home}_{43}=C^\text{home}_{34}=6$. Adults and the elderly have 6 mutual interactions every day
Which contact matrix to use for which node and commuter nodes is set using `node_cmatrices.csv` and `cnode_cmatrices.csv`.
```
pd.read_csv('./SIR_single_node/node_cmatrices.csv')
pd.read_csv('./SIR_single_node/cnode_cmatrices.csv')
```
Note the `NaN` under the $S$ and $R$ classes. This is because contact matrices are only applicable for infection classes (which is $I$ in this case).
### 1.1.4 Populating the simulation ([node_populations.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#node_populationscsv))
The population of each node is set using `node_populations.csv`.
```
pd.read_csv('./SIR_single_node/node_populations.csv')
```
Here we have assigned various numbers of susceptible children, young adults, adults and elderly to our single node. We have also seeded the node with a single infected child.
### 1.1.5 The commuter network ([commuter_networks.csv](https://github.com/lukastk/PyRossGeo/blob/master/docs/Configuration%20files.md#commuter_networkscsv))
As we are only considering a single node in this example, the commuter network file is empty. The file is still required for the initialisation of the simulation.
```
pd.read_csv('./SIR_single_node/commuter_networks.csv')
```
### 1.1.6 Initializing and running the simulation
To run the simulation we first create an instance of the `pyrossgeo.Simulation` class, and then initialize it by passing the path of the folder containing the configuration files.
**General note of caution:** <i>At the moment PyRossGeo does not have any checks in place to detect formatting errors in the configuration files. Any mistakes in the configuration files may lead to uncaught exceptions or unexpected simulation behaviour. This will be rectified in the future.</i>
```
sim = pyrossgeo.Simulation()
X_state = sim.initialize(sim_config_path='SIR_single_node')
```
`X_state` is a one-dimensional vector containing the initial conditions of the simulation, as specified by the configuration files.
An alternative way to initialize is by specifically passing the path for each file.
```
sim = pyrossgeo.Simulation()
X_state = sim.initialize(model_dat = 'SIR_single_node/model.json',
commuter_networks_dat = 'SIR_single_node/commuter_networks.csv',
node_parameters_dat = 'SIR_single_node/node_parameters.csv',
cnode_parameters_dat = 'SIR_single_node/cnode_parameters.csv',
contact_matrices_dat = 'SIR_single_node/contact_matrices.json',
node_cmatrices_dat = 'SIR_single_node/node_cmatrices.csv',
cnode_cmatrices_dat = 'SIR_single_node/cnode_cmatrices.csv',
node_populations_dat = 'SIR_single_node/node_populations.csv')
```
Calling `pyrossgeo.Simulation.initialize` without any arguments is equivalent to setting `sim_config_path='.'`.
To run the simulation, we simply define a starting time, an end time, a time-step, and call `pyrossgeo.Simulation.simulate`.
```
t_start = 0
t_end = 200*24*60 # Time is measured in units of minutes
dt = 1 # The time-step is 10 minutes
sim_data = sim.simulate(X_state, t_start, t_end, dt, steps_per_save=1)
```
By specifying `steps_per_save=1` the state of the simulation is saved every simulation step and returned in `sim_data`. More on the structure of the `sim_data` in the [subsequent tutorial](../tutorial2-handling-PyRossGeo-output/tutorial2-handling-PyRossGeo-output.ipynb).
### 1.1.7 Plotting the results
We can now plot the result of the simulation using the data returned in `sim_data`.
```
ts, node_data, cnode_data, location_data, community_data, network_data = pyrossgeo.utils.extract_simulation_data(sim_data)
ts_days = ts / (24*60)
plt.figure( figsize=(8,3) )
S = np.sum(network_data[:,:,0], axis=1) # Sum over all age-groups
I = np.sum(network_data[:,:,1], axis=1)
R = np.sum(network_data[:,:,2], axis=1)
N = S + I + R
plt.plot(ts_days, S, label="S")
plt.plot(ts_days, I, label="I")
plt.plot(ts_days, R, label="R")
plt.legend(loc='upper right', fontsize=12)
plt.xlabel('Days')
plt.show()
```
Plotting the individual age groups:
```
plt.figure( figsize=(14,5) )
S0 = network_data[:,0,0]
I0 = network_data[:,0,1]
R0 = network_data[:,0,2]
S1 = network_data[:,1,0]
I1 = network_data[:,1,1]
R1 = network_data[:,1,2]
S2 = network_data[:,2,0]
I2 = network_data[:,2,1]
R2 = network_data[:,2,2]
S3 = network_data[:,3,0]
I3 = network_data[:,3,1]
R3 = network_data[:,3,2]
plt.plot(ts_days, S0, label="S0")
plt.plot(ts_days, I0, label="I0")
plt.plot(ts_days, R0, label="R0")
plt.plot(ts_days, S1, label="S1")
plt.plot(ts_days, I1, label="I1")
plt.plot(ts_days, R1, label="R1")
plt.plot(ts_days, S2, label="S2")
plt.plot(ts_days, I2, label="I2")
plt.plot(ts_days, R2, label="R2")
plt.plot(ts_days, S3, label="S3")
plt.plot(ts_days, I3, label="I3")
plt.plot(ts_days, R3, label="R3")
plt.legend(loc='upper right', fontsize=12)
plt.xlabel('Days')
plt.show()
```
## 1.2 Age-structured SEAIR on three nodes
Now that we have covered the simulation of single-node SIR, we can go on to more interesting things.
### 1.2.1 Configuration files
The SEAIR model is defined as
$$
\begin{aligned}
\dot{S}^\mu & = - \lambda^\mu(t) S^\mu \\
\dot{E}^\mu & = \lambda^\mu(t) S^\mu - \gamma_E E^\mu \\
\dot{A}^\mu & = \gamma_E E^\mu - \gamma_A A^\mu \\
\dot{I}^\mu & = \gamma_A A^\mu - \gamma_I I^\mu \\
\dot{R}^\mu & = \gamma_I I^\mu
\end{aligned}
$$
where
$$
\lambda^\mu(t) = \sum_\nu \left( \beta_I C^I_{\mu\nu} \frac{I^\nu}{N^\nu} + \beta_A C^A_{\mu\nu} \frac{A^\nu}{N^\nu} \right)
$$
```
with open('./SEAIR_network/model.json', 'r') as f:
print(f.read())
```
The model parameters for each node
```
pd.read_csv('./SEAIR_network/node_parameters.csv')
```
The model parameters for each commuter node. We set the area of commuter nodes to be $0.1 km^2$ to reflect the fact that commutes tend to pack people more closely.
```
pd.read_csv('./SEAIR_network/cnode_parameters.csv')
```
We define three different contact matrices, corresponding to people who are currently at their home nodes, are away, or are commuting.
```
with open('./SEAIR_network/contact_matrices.json', 'r') as f:
print(f.read())
```
Set contact matrices for each node
```
pd.read_csv('./SEAIR_network/node_cmatrices.csv')
```
Note the keyword `HOME` in the `Location` column, which signifies that the value in the `Home` column should be copied. So the second row matches all nodes $(\alpha,i,j)$ where $i=j$.
Set contact matrices for each commuter node
```
pd.read_csv('./SEAIR_network/cnode_cmatrices.csv')
```
Node populations
```
pd.read_csv('./SEAIR_network/node_populations.csv')
```
#### 1.2.1.1 Defining the commuter network
We will be using the commtuer network below:
```
pd.read_csv('./SEAIR_network/commuter_networks.csv')
```
Each row of the file defines a specific commute that occurs every day.
**First row:**
- It defines a commute that takes 210 people from node $(\alpha=0,i=0,j=0)$ to the node $(\alpha=0,i=0,j=1)$. In other words, it moves 210 people who live on location 0, and are currently at location 0, to location 1.
- The amount of people that are commuting is specified by the `# to move` column. Alternatively, we can also specify percentages of populations to commute using the `% to move`. It's important to note that both columns cannot be used at the same time. The column that is not used must be set to `-1`.
- The departure window is between `t1=7` and `t2=8`, meaning that people will start leaving at `t1`, and by `t2` all 210 people will have left.$^*$ The transport is modeled using a Gaussian pulse function (see the [model description](model.pdf) for more details). Time is given in units of hours, and should be in the range of a single day `[0, 24]`.
- People leaving the $(0,0,0)$ node will first enter the commuting node $(0,0,0 \to 1)$. The latter should be read as *the commuting node of residents of location 0, of age group 0, who are travelling from location 0 to 1*.
- The arrival window is between `ct1=8` and `ct2=9`. At `ct1` people will start to be moved from the commuting node to the destination node $(0,0,1)$, by $ct2$ everyone will have arrived.
- Using `Allow S`, `Allow E`... we can block certain classes from going on the commute. We have set `Allow I` to 0, in order to model the fact that symptomatic infecteds are likely to stay home and recuperate.
**Second row:**
- This is the return commute for the people who went to node $(0,0,1)$ from $(0,0,0)$ in the morning. We see that people will start leaving at 6PM, and arrive at home at 7PM.
- As opposed to the morning commute, we specify that 100% of the people at node $(0,0,1)$ should return to $(0,0,0)$.
- Note that we allow members of all classes to return home. This is important, as we would expect people who catch the virus away from home would return.
The rest of the rows defines commutes for other age classes and locations.
<i>$^*$ In practice, it is possible that fewer than the specified amount of people will travel. This is because of the fact that we are disallowing infected classes to move across the network (for example, if all residents of 0 are currently infected, then no people should be moving). See the [model description](https://github.com/lukastk/PyRossGeo/blob/master/docs/model.pdf) for more details on this.</i>
Note that we always specify `# to move` in the forward commute, and `% to move` in the return to commute. In most use-cases this is how commuting networks should be constructed. The reason why we specify percentages rather than absolute numbers in the return commute is because we might not know at run-time how many people actually went on the morning commute, due to the fact that we are barring certain classes from commuting.
### 1.2.2 Simulating the network
Before we start the simulation, we will create an adapted time-stepping schedule for the simulation using `pyrossgeo.utils.get_dt_schedule`.
```
ts, dts = pyrossgeo.utils.get_dt_schedule([
(0, 2*60), # Use a 2h time-step between 0-7
(5*60, 1), # Use a 10min time-step between 5-11
(11*60, 2*60), # Use a 2h time-step between 11-16
(16*60, 1), # Use a 10min time-step between 16-22
(22*60, 2*60) # Use a 2h time-step between 22-24
], end_time=24*60)
```
We pass a list of tuples `[(t0, dt0), (t1, dt1), ..., (tn, dtn)]`, and the function returns a tuple `(ts, dts)`, containing the corresponding times and the time-steps. Between `t0` and `t1`, the time-steps will be `dt0`. Between `t1` and `t2`, the time-steps will be `dt1`, and so on.
As transport generally requires a larger time-step than the epidemics, we define the time-stepping schedule with a 10 minute time-step during the commuting hours, and a two hour time-step anytime else.
We are now ready to run the simulation.
```
t_start = 0
t_end = 24*60*300 # Run for 200 days
sim = pyrossgeo.Simulation()
X_state = sim.initialize(sim_config_path='SEAIR_network')
sim_data = sim.simulate(X_state, t_start, t_end, dts, steps_per_save=1)
```
Various relevant data can be extracted from the simulation output `sim_data` using the function `pyrossgeo.utils.extract_simulation_data`.
```
ts, node_data, cnode_data, location_data, community_data, network_data = pyrossgeo.utils.extract_simulation_data(sim_data)
```
We can plot the epidemic for the network as a whole
```
ts_days = ts / (24*60)
plt.figure( figsize=(8,3) )
S = np.sum(network_data[:,:,0], axis=1) # Sum over all age-groups
E = np.sum(network_data[:,:,1], axis=1)
A = np.sum(network_data[:,:,2], axis=1)
I = np.sum(network_data[:,:,3], axis=1)
R = np.sum(network_data[:,:,4], axis=1)
plt.plot(ts_days, S, label="S")
plt.plot(ts_days, E, label="E")
plt.plot(ts_days, A, label="A")
plt.plot(ts_days, I, label="I")
plt.plot(ts_days, R, label="R")
plt.legend(loc='upper right', fontsize=12)
plt.xlabel('Days')
plt.show()
```
We can plot for a single location.
```
ts_days = ts / (24*60)
loc = 0
plt.figure( figsize=(8,3) )
S = np.sum(location_data[:,:,0,loc], axis=1)
E = np.sum(location_data[:,:,1,loc], axis=1)
A = np.sum(location_data[:,:,2,loc], axis=1)
I = np.sum(location_data[:,:,3,loc], axis=1)
R = np.sum(location_data[:,:,4,loc], axis=1)
plt.plot(ts_days, S, label="S")
plt.plot(ts_days, E, label="E")
plt.plot(ts_days, A, label="A")
plt.plot(ts_days, I, label="I")
plt.plot(ts_days, R, label="R")
#plt.plot(ts_days, S+E+A+I+R, label="N")
plt.legend(loc='upper right', fontsize=12)
plt.xlabel('Days')
plt.title("Location %s" % loc)
plt.show()
```
The thickness in the curves stem from the oscillations due to the daily commute. Zooming in, we get:
```
ts_days = ts / (24*60)
loc = 0
plt.figure( figsize=(8,3) )
S = np.sum(location_data[:,:,0,loc], axis=1)
E = np.sum(location_data[:,:,1,loc], axis=1)
A = np.sum(location_data[:,:,2,loc], axis=1)
I = np.sum(location_data[:,:,3,loc], axis=1)
R = np.sum(location_data[:,:,4,loc], axis=1)
for i in range(int(np.max(ts_days))):
plt.axvline(i, color='black')
plt.plot(ts_days, S+E+A+I+R, label="N")
plt.legend(loc='upper right', fontsize=12)
plt.xlabel('Days')
plt.title("Location %s" % loc)
plt.xlim(0, 5)
plt.show()
```
This concludes the first part of the PyRossGeo tutorial.
- [Continue to **Tutorial 2: Handling PyRossGeo output**](../tutorial2-handling-PyRossGeo-output/tutorial2-handling-PyRossGeo-output.ipynb)
- [Skip to **Tutorial 3: Handling the simulation output**](../tutorial3-interventions/tutorial3-interventions.ipynb)
- [Go to the PyRossGeo documentation](https://github.com/lukastk/PyRossGeo/blob/master/docs/Documentation.md)
| github_jupyter |
```
"""Process Leif Anderson's debris thickness data (https://zenodo.org/record/4317470#.X-TlbOlKhTa)"""
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import debrisglobal.globaldebris_input as debris_prms
df_fp = ('/Users/drounce/Documents/DebrisGlaciers_WG/Melt_Intercomparison/hd_obs/datasets/Leif_comparisons/' +
'Alps_debris_thicknesses/')
# df_fn = '11.01450_grosseraletsch_anderson2019-welev.csv'
# df_fn = '11.01509_oberaar_anderson2019-welev.csv'
# df_fn = '11.01827_oberaletsch_anderson2019-welev.csv'
# df_fn = '11.02749_cheilon_anderson2019-welev.csv'
# df_fn = '11.02771_piece_anderson2019-welev.csv'
# df_fn = '11.02796_brenay_anderson2019-welev.csv'
df_fn = '11.03005_miage_anderson2019-welev.csv'
df = pd.read_csv(df_fp + df_fn)
df_all = None
for nrow in range(df.shape[0]):
df_pt = df.loc[[nrow],:]
# Personal communication with Leif Anderson (12/11/2020) assume 5 measurements (3 mean, then 1 for min and max)
for ncount in range(5):
if ncount < 3:
df_pt['hd_m'] = df_pt['mean'] / 100
elif ncount == 3:
df_pt['hd_m'] = df_pt['min'] / 100
elif ncount == 4:
df_pt['hd_m'] = df_pt['max'] / 100
if df_all is None:
df_all = df_pt
else:
df_all = pd.concat([df_all, df_pt], axis=0)
df_all.reset_index(inplace=True, drop=True)
df_all.to_csv(df_fp + '../../' + df_fn.replace('-welev.csv', '-hd.csv'), index=False)
df_fp + '../../' + df_fn.replace('-welev.csv', '-hd.csv')
df_all
# Load point data
df_fp = '/Users/drounce/Documents/DebrisGlaciers_WG/Melt_Intercomparison/hd_obs/datasets/hd_pt_data/'
# df_fn = '11.02749_cheilon_anderson2019-hd_pt.csv'
df_fn = '15.03733_gibson-hd_pt.csv'
df_pt = pd.read_csv(df_fp + df_fn)
# drop null values
df_pt.loc[df_pt['hd_mod_m'] > debris_prms.hd_max, 'hd_mod_m'] = np.nan
df_pt = df_pt.dropna(subset=['hd_obs_m', 'hd_mod_m'])
df_pt.reset_index(inplace=True, drop=True)
glac_str = df_fn.split('_')[0]
# Load bin data
bin_fp = '/Users/drounce/Documents/DebrisGlaciers_WG/Melt_Intercomparison/hd_obs/hd_processed/'
# bin_fn = '11.02749_cheilon_anderson2019-hd-processed.csv'
bin_fn = '15.03733_gibson-hd-processed.csv'
df_bin = pd.read_csv(bin_fp + bin_fn)
# Uncertainty dataframe and dictionary for bounds
hd_uncertainty_fullfn = debris_prms.output_fp + 'hd_uncertainty_bnds-1std.csv'
hd_uncertainty_df = pd.read_csv(hd_uncertainty_fullfn)
hd_uncertainty_dict_low = dict(zip([int(np.round(x*100)) for x in hd_uncertainty_df['hd_m']],
list(hd_uncertainty_df['hd_bndlow_both'].values)))
hd_uncertainty_dict_low[0] = 0
hd_uncertainty_dict_low[1] = 0
hd_uncertainty_dict_high = dict(zip([int(np.round(x*100)) for x in hd_uncertainty_df['hd_m']],
list(hd_uncertainty_df['hd_bndhigh_both'].values)))
hd_uncertainty_dict_high[0] = hd_uncertainty_df.loc[0,'hd_bndhigh_both']
hd_uncertainty_dict_high[1] = hd_uncertainty_df.loc[0,'hd_bndhigh_both']
rmse_pts = (np.sum((df_pt['hd_obs_m'] - df_pt['hd_mod_m'])**2) / df_pt.shape[0])**0.5
print('rmse:', np.round(rmse_pts,2), 'points:', df_pt.shape[0])
# Add uncertainty
df_pt['hd_mod_m_low'] = [hd_uncertainty_dict_low[x]
for x in list(np.round(np.array(df_pt['hd_mod_m'])*100,0).astype(int))]
df_pt['hd_mod_m_high'] = [hd_uncertainty_dict_high[x]
for x in list(np.round(np.array(df_pt['hd_mod_m'])*100,0).astype(int))]
df_pt['hd_mod_m_diflow'] = np.abs(df_pt['hd_mod_m'] - df_pt['hd_mod_m_low'])
df_pt['hd_mod_m_difhigh'] = np.abs(df_pt['hd_mod_m'] - df_pt['hd_mod_m_high'])
df_pt['hd_mod_m_std'] = (df_pt['hd_mod_m_diflow'] + df_pt['hd_mod_m_difhigh']) / 2
hd_min, hd_max = 0.004, 5
n_obs_min = 5
hd_compare_fp = debris_prms.main_directory + '/../hd_obs/figures/hd_obs_compare/'
fig, ax = plt.subplots(1, 1, squeeze=False, gridspec_kw = {'wspace':0, 'hspace':0})
# Points
ax[0,0].scatter(df_pt['hd_obs_m'].values,
df_pt['hd_mod_m'].values,
color='grey', marker='o', linewidth=0.5, facecolor='none',
# s=s_plot, zorder=zorder,
# label=label_str, clip_on=True
)
ax[0,0].errorbar(df_pt['hd_obs_m'],
df_pt['hd_mod_m'],
yerr=df_pt['hd_mod_m_std'],
capsize=1, capthick=lw_err, elinewidth=lw_err, linewidth=0, color='grey', alpha=1, zorder=2)
# # Bins
# for ndata in df_bin.index.values:
# label_str = None
# marker = 's'
# # Size thresholds
# s_sizes = [20, 40, 80]
# lws = [0.5, 1, 1]
# lws_err = [0.1, 0.5, 0.5]
# colors = ['grey', '#31a354', '#3182bd']
# zorders = [3,4,5]
# obs_count = df_bin.loc[ndata,'obs_count']
# if obs_count >= n_obs_min and obs_count < 25:
# s_plot = s_sizes[0]
# lw = lws[0]
# lw_err = lws_err[0]
# color = colors[0]
# zorder = zorders[0]
# elif obs_count >= 25 and obs_count < 100:
# s_plot = s_sizes[1]
# lw = lws[1]
# lw_err = lws_err[1]
# color = colors[1]
# zorder = zorders[1]
# elif obs_count >= 100:
# s_plot = s_sizes[2]
# lw = lws[2]
# lw_err = lws_err[2]
# color = colors[2]
# zorder = zorders[2]
# else:
# print('NO COLOR')
# ax[0,0].scatter(df_bin.loc[ndata,'hd_obs_med'],
# df_bin.loc[ndata,'hd_ts_med_m'],
# color=color, marker=marker, linewidth=lw, facecolor='none', s=s_plot, zorder=zorder,
# label=label_str, clip_on=True)
# ax[0,0].errorbar(df_bin.loc[ndata,'hd_obs_med'],
# df_bin.loc[ndata,'hd_ts_med_m'],
# xerr=1.483*df_bin.loc[ndata,'hd_obs_mad'],
# yerr=1.483*df_bin.loc[ndata,'hd_ts_mad_m'],
# capsize=1, capthick=lw_err, elinewidth=lw_err, linewidth=0, color=color, alpha=1, zorder=2)
# Log scale
ax[0,0].set_xscale('log')
ax[0,0].set_yscale('log')
# Labels
ax[0,0].set_xlabel('Observed $h_{d}$ (m)', size=12)
ax[0,0].set_ylabel('Modeled $h_{d}$ (m)', size=12)
ax[0,0].set_xlim(hd_min,hd_max)
ax[0,0].set_ylim(hd_min,hd_max)
ax[0,0].plot([hd_min, hd_max], [hd_min, hd_max], color='k', linewidth=0.5, zorder=1)
fig.set_size_inches(3.45,3.45)
fig_fullfn = hd_compare_fp + glac_str + '-hd_compare.png'
fig.savefig(fig_fullfn, bbox_inches='tight', dpi=300)
# POINT DATASETS - process such that Leif's data has 5 measurements per site in agreement with his measurements
df_fp = '/Users/drounce/Documents/DebrisGlaciers_WG/Melt_Intercomparison/hd_obs/datasets/hd_pt_data/'
# df_fn = '11.01450_grosseraletsch_anderson2019-hd_pt-old.csv'
# df_fn = '11.01509_oberaar_anderson2019-hd_pt-old.csv'
# df_fn = '11.01827_oberaletsch_anderson2019-hd_pt-old.csv'
# df_fn = '11.02749_cheilon_anderson2019-hd_pt-old.csv'
# df_fn = '11.02771_piece_anderson2019-hd_pt-old.csv'
df_fn = '11.02796_brenay_anderson2019-hd_pt-old.csv'
# # df_fn = '11.03005_miage_anderson2019-hd_pt-old.csv'
df = pd.read_csv(df_fp + df_fn)
df_all = None
for nrow in range(df.shape[0]):
df_pt = df.loc[[nrow],:]
# Personal communication with Leif Anderson (12/11/2020) assume 5 measurements (3 mean, then 1 for min and max)
for ncount in range(5):
if ncount < 3:
df_pt['hd_m'] = df_pt['mean'] / 100
elif ncount == 3:
df_pt['hd_m'] = df_pt['min'] / 100
elif ncount == 4:
df_pt['hd_m'] = df_pt['max'] / 100
if df_all is None:
df_all = df_pt
else:
df_all = pd.concat([df_all, df_pt], axis=0)
df_all.reset_index(inplace=True, drop=True)
df_all.to_csv(df_fp + df_fn.replace('-old.csv', '.csv'), index=False)
```
| github_jupyter |
## Example for FloPy methods note
Import the `modflow` and `utils` subpackages of FloPy and give them the aliases `fpm` and `fpu`, respectively
```
import os
import sys
import numpy as np
import flopy
import flopy.modflow as fpm
import flopy.utils as fpu
import matplotlib as mpl
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
```
Create a MODFLOW model object. Here, the MODFLOW model object is stored in a Python variable called {\tt model}, but this can be an arbitrary name. This object name is important as it will be used as a reference to the model in the remainder of the FloPy script. In addition, a {\tt modelname} is specified when the MODFLOW model object is created. This {\tt modelname} is used for all the files that are created by FloPy for this model.
```
exe = 'mf2005'
ws = os.path.join('temp')
model = fpm.Modflow(modelname='gwexample', exe_name=exe, model_ws=ws)
```
The discretization of the model is specified with the discretization file (DIS) of MODFLOW. The aquifer is divided into 201 cells of length 10 m and width 1 m. The first input of the discretization package is the name of the model object. All other input arguments are self explanatory.
```
fpm.ModflowDis(model, nlay=1, nrow=1, ncol=201,
delr=10, delc=1, top=50, botm=0)
```
Active cells and the like are defined with the Basic package (BAS), which is required for every MODFLOW model. It contains the {\tt ibound} array, which is used to specify which cells are active (value is positive), inactive (value is 0), or fixed head (value is negative). The {\tt numpy} package (aliased as {\tt np}) can be used to quickly initialize the {\tt ibound} array with values of 1, and then set the {\tt ibound} value for the first and last columns to -1. The {\tt numpy} package (and Python, in general) uses zero-based indexing and supports negative indexing so that row 1 and column 1, and row 1 and column 201, can be referenced as [0, 0], and [0, -1], respectively. Although this simulation is for steady flow, starting heads still need to be specified. They are used as the head for fixed-head cells (where {\tt ibound} is negative), and as a starting point to compute the saturated thickness for cases of unconfined flow.
```
ibound = np.ones((1, 201))
ibound[0, 0] = ibound[0, -1] = -1
fpm.ModflowBas(model, ibound=ibound, strt=20)
```
The hydraulic properties of the aquifer are specified with the Layer Properties Flow (LPF) package (alternatively, the Block Centered Flow (BCF) package may be used). Only the hydraulic conductivity of the aquifer and the layer type ({\tt laytyp}) need to be specified. The latter is set to 1, which means that MODFLOW will calculate the saturated thickness differently depending on whether or not the head is above the top of the aquifer.
```
fpm.ModflowLpf(model, hk=10, laytyp=1)
```
Aquifer recharge is simulated with the Recharge package (RCH) and the extraction of water at the two ditches is simulated with the Well package (WEL). The latter requires specification of the layer, row, column, and injection rate of the well for each stress period. The layers, rows, columns, and the stress period are numbered (consistent with Python's zero-based numbering convention) starting at 0. The required data are stored in a Python dictionary ({\tt lrcQ} in the code below), which is used in FloPy to store data that can vary by stress period. The {\tt lrcQ} dictionary specifies that two wells (one in cell 1, 1, 51 and one in cell 1, 1, 151), each with a rate of -1 m$^3$/m/d, will be active for the first stress period. Because this is a steady-state model, there is only one stress period and therefore only one entry in the dictionary.
```
fpm.ModflowRch(model, rech=0.001)
lrcQ = {0: [[0, 0, 50, -1], [0, 0, 150, -1]]}
fpm.ModflowWel(model, stress_period_data=lrcQ)
```
The Preconditioned Conjugate-Gradient (PCG) solver, using the default settings, is specified to solve the model.
```
fpm.ModflowPcg(model)
```
The frequency and type of output that MODFLOW writes to an output file is specified with the Output Control (OC) package. In this case, the budget is printed and heads are saved (the default), so no arguments are needed.
```
fpm.ModflowOc(model)
```
Finally the MODFLOW input files are written (eight files for this model) and the model is run. This requires, of course, that MODFLOW is installed on your computer and FloPy can find the executable in your path.
```
model.write_input()
model.run_model()
```
After MODFLOW has responded with the positive {\tt Normal termination of simulation}, the calculated heads can be read from the binary output file. First a file object is created. As the modelname used for all MODFLOW files was specified as {\tt gwexample} in step 1, the file with the heads is called {\tt gwexample.hds}. FloPy includes functions to read data from the file object, including heads for specified layers or time steps, or head time series at individual cells. For this simple mode, all computed heads are read.
```
fpth = os.path.join(ws, 'gwexample.hds')
hfile = fpu.HeadFile(fpth)
h = hfile.get_data(totim=1.0)
```
The heads are now stored in the Python variable {\tt h}. FloPy includes powerful plotting functions to plot the grid, boundary conditions, head, etc. This functionality is demonstrated later. For this simple one-dimensional example, a plot is created with the matplotlib package
```
import matplotlib.pyplot as plt
ax = plt.subplot(111)
x = model.dis.sr.xcentergrid[0]
ax.plot(x,h[0,0,:])
ax.set_xlim(0,x.max())
ax.set_xlabel("x(m)")
ax.set_ylabel("head(m)")
plt.show()
```
| github_jupyter |
# Solution b.
Create a inference script. Let's call it `inference.py`.
Let's also create the `input_fn`, `predict_fn`, `output_fn` and `model_fn` functions.
Copy the cells below and paste in [the main notebook](../deployment_hosting.ipynb).
```
%%writefile inference.py
import os
import pickle
import xgboost
import sagemaker_xgboost_container.encoder as xgb_encoders
# Same as in the training script
def model_fn(model_dir):
"""Load a model. For XGBoost Framework, a default function to load a model is not provided.
Users should provide customized model_fn() in script.
Args:
model_dir: a directory where model is saved.
Returns:
A XGBoost model.
XGBoost model format type.
"""
model_files = (file for file in os.listdir(model_dir) if os.path.isfile(os.path.join(model_dir, file)))
model_file = next(model_files)
try:
booster = pickle.load(open(os.path.join(model_dir, model_file), 'rb'))
format = 'pkl_format'
except Exception as exp_pkl:
try:
booster = xgboost.Booster()
booster.load_model(os.path.join(model_dir, model_file))
format = 'xgb_format'
except Exception as exp_xgb:
raise ModelLoadInferenceError("Unable to load model: {} {}".format(str(exp_pkl), str(exp_xgb)))
booster.set_param('nthread', 1)
return booster
def input_fn(request_body, request_content_type):
"""
The SageMaker XGBoost model server receives the request data body and the content type,
and invokes the `input_fn`.
The input_fn that just validates request_content_type and prints
"""
print("Hello from the PRE-processing function!!!")
if request_content_type == "text/csv":
return xgb_encoders.csv_to_dmatrix(request_body)
else:
raise ValueError(
"Content type {} is not supported.".format(request_content_type)
)
def predict_fn(input_object, model):
"""
SageMaker XGBoost model server invokes `predict_fn` on the return value of `input_fn`.
"""
return model.predict(input_object)[0]
def output_fn(prediction, response_content_type):
"""
After invoking predict_fn, the model server invokes `output_fn`.
An output_fn that just adds a column to the output and validates response_content_type
"""
print("Hello from the POST-processing function!!!")
appended_output = "hello from post-processing function!!!"
predictions = [prediction, appended_output]
if response_content_type == "text/csv":
return ','.join(str(x) for x in predictions)
else:
raise ValueError("Content type {} is not supported.".format(response_content_type))
```
Deploy the new model with the inference script:
- find the S3 bucket where the artifact is stored (you can create a tarball and upload it to S3 or use another model that was previously created in SageMaker)
#### Finding a previously trained model:
Go to the Experiments tab in Studio again:

Choose another trained model, such as the one trained with Framework mode (right-click and choose `Open in trial details`):

Click on `Artifacts` and look at the `Output artifacts`:

Copy and paste your `SageMaker.ModelArtifact` of the S3 URI where the model is saved:
In this example:
```
s3_artifact="s3://sagemaker-studio-us-east-2-<AWS_ACCOUNT_ID>/xgboost-churn/output/demo-xgboost-customer-churn-2021-04-13-18-51-56-144/output/model.tar.gz"
```
```
s3_artifact="s3://<YOUR-BUCKET>/PATH/TO/model.tar.gz"
%store -r docker_image_name
```
**Deploy it:**
```
from sagemaker.xgboost.model import XGBoostModel
xgb_inference_model = XGBoostModel(
entry_point="inference.py",
model_data=s3_artifact,
role=role,
image_uri=docker_image_name,
framework_version="0.90-2",
py_version="py3"
)
data_capture_prefix = '{}/datacapture'.format(prefix)
endpoint_name = "model-xgboost-customer-churn-" + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("EndpointName = {}".format(endpoint_name))
predictor = xgb_inference_model.deploy( initial_instance_count=1,
instance_type='ml.m4.xlarge',
endpoint_name=endpoint_name,
data_capture_config=DataCaptureConfig(
enable_capture=True,
sampling_percentage=100,
destination_s3_uri='s3://{}/{}'.format(bucket, data_capture_prefix)
)
)
## Updating an existing endpoint
# model_name = xgb_inference_model.name
# from sagemaker.predictor import Predictor
# predictor = Predictor(endpoint_name=endpoint_name)
# predictor.update_endpoint(instance_type='ml.m4.xlarge',
# initial_instance_count=1,
# model_name=model_name,
# data_capture_config=DataCaptureConfig(
# enable_capture=True,
# sampling_percentage=100,
# destination_s3_uri='s3://{}/{}'.format(bucket, data_capture_prefix)
# )
# )
```
**Send some requests:**
```
runtime_client = boto3.client("sagemaker-runtime")
with open('/root/amazon-sagemaker-workshop/4-Deployment/RealTime/config/test_sample.csv', 'r') as f:
for row in f:
payload = row.rstrip('\n')
print(f"Sending: {payload}")
response = runtime_client.invoke_endpoint(EndpointName=endpoint_name,
ContentType='text/csv',
Accept='text/csv',
Body=payload)
print(f"\nReceived: {response['Body'].read()}")
break
```
Go to CloudWatch logs and check the inference logic:
[Link to CloudWatch Logs](https://us-east-2.console.aws.amazon.com/cloudwatch/home?region=us-east-2#logsV2:log-groups$3FlogGroupNameFilter$3D$252Faws$252Fsagemaker$252FEndpoints$252F)
| github_jupyter |
```
import sys
from matplotlib import pyplot as plt
from sklift.metrics import uplift_at_k
import seaborn as sns
import numpy as np
import pandas as pd
# install uplift library scikit-uplift and other libraries
#!{sys.executable} -m pip install scikit-uplift dill catboost
%config InlineBackend.figure_format = 'svg'
%matplotlib inline
```
## Load Dataset
We are going to use a Hillstrom dataset from the MineThatData [hosted](https://blog.minethatdata.com/2008/03/minethatdata-e-mail-analytics-and-data.html) in march 2008 by the president of this company Kevin Hillstrom.
MineThatData is a consulting company that helps CEO understand the complex relationship between Customers, Advertising, Products, Brands, and Channels.
## Data description
Dataset can be loaded from sklift.datasets module using fetch_hillstrom function.
Read more about dataset in the [api docs](https://www.uplift-modeling.com/en/latest/api/datasets/fetch_hillstrom.html).
This dataset contains 64,000 customers who last purchased within twelve months. The customers were involved in an e-mail test
**Major columns**
- **visit** (binary): target. 1/0 indicator, 1 = Customer visited website in the following two weeks.
- **conversion** (binary): target. 1/0 indicator, 1 = Customer purchased merchandise in the following two weeks.
- **spend** (float): target. Actual dollars spent in the following two weeks.
- **segment** (str): treatment. The e-mail campaign the customer received
Read more in the [docs](https://www.uplift-modeling.com/en/latest/api/datasets/fetch_hillstrom.html#hillstrom)
```
from sklift.datasets import fetch_hillstrom
# returns sklearn Bunch object
# with data, target, treatment keys
# data features (pd.DataFrame), target (pd.Series), treatment (pd.Series) values
dataset = fetch_hillstrom()
print(f"Dataset type: {type(dataset)}\n")
print(f"Dataset features shape: {dataset.data.shape}")
print(f"Dataset target shape: {dataset.target.shape}")
print(f"Dataset treatment shape: {dataset.treatment.shape}")
```
## EDA
```
dataset.data.head().append(dataset.data.tail())
#info about types and null cells in dataset
dataset.data.info()
```
There is no missing data in the cells!
# Categorical data
```
cat_features = ['channel', 'zip_code', 'history_segment', 'newbie']
dataset.data.channel.unique()
dataset.data.zip_code.unique()
dataset.data.history_segment.unique()
```
**Zip code**
```
dataset.data.zip_code.value_counts().plot(kind = 'bar', grid=True)
```
**Channel**
```
dataset.data.channel.value_counts().plot(kind = 'bar', grid=True)
```
**History segment**
```
dataset.data.history_segment.value_counts().plot(kind = 'bar', grid=True)
#As option I propose to apply following function for transformation data in column "historic_segment"
def historic_segment_transform(dataset):
for payment in dataset.data['history_segment'].unique():
if payment =='1) $0 - $100':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 50
elif payment =='2) $100 - $200':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 150
elif payment =='3) $200 - $350':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 275
elif payment =='4) $350 - $500':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 425
elif payment =='5) $500 - $750':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 575
elif payment =='5) $750 - $1000':
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 825
else:
dataset.data.loc[dataset.data['history_segment'] == payment, 'history_segment'] = 1000
return dataset.data.history_segment
dataset.data.history_segment.value_counts()
dataset.data.history_segment.value_counts().plot(kind = 'bar', grid=True)
#It's better to proceed from categorical to numeric data
# For example, we could replace 0-100 on average value 50
dataset.data.womens.value_counts().plot(kind = 'bar', grid=True)
dataset.data.mens.value_counts().plot(kind = 'bar', grid=True)
dataset.data.womens.value_counts()
dataset.data.groupby('womens').size()/dataset.data['womens'].count()*100
#55% - womens purchases
#44% - mens purchases
dataset.data.mens.value_counts()
dataset.data.groupby('mens').size()/dataset.data['mens'].count()*100
#55% - mens purchases
#44% - womens purchases
plt.figure(figsize = (14,8))
sns.set(font_scale=0.75)
sns.heatmap(dataset.data.corr().round(3), annot=True, square = True, linewidths=.75, cmap='RdPu', fmt = '.2f',annot_kws = {"size": 10} )
plt.title('Correlation matrix')
plt.show()
# womens and mens are in inverse correlation. I propose to make 1 column "gender" and merge.
# As we can see, there is high correlation between 'history_segment' and 'history'. Could we merge it also and transform columns to numeric data?
```
# Numeric data
```
dataset.data.loc[:, 'recency'].hist(figsize=(8, 4), bins=12, grid=True)
dataset.data.history.value_counts()
dataset.data.loc[:, 'history'].hist(figsize=(8, 4), bins=20, grid=True);
```
# Target data
```
#dataset_segment
dataset.treatment.head()
dataset.treatment.unique()
dataset.treatment.value_counts().plot(kind = 'bar', grid = 'True')
#dataset_visit
dataset.target.head()
dataset.target.value_counts().plot(kind = 'bar')
#Target is disbalanced
import pandas as pd
pd.crosstab(dataset.treatment, dataset.target, normalize='index')
dataset.target.unique()
crosstab = pd.crosstab(dataset.treatment, dataset.target, normalize='index')
sns.heatmap(crosstab, annot=True, fmt=".2f", linewidths=1, square = True, cmap = 'RdPu')
plt.xlabel('Target')
plt.title("Treatment & Target")
#Let's consider two cases:
#1) Womens E-mail - No E-mail
#2) Mens E-mail - No E-mail
```
## Womens E-mail - No E-mail
```
# make treatment binary
treat_dict_womens = {
'Womens E-Mail': 1,
'No E-Mail': 0,
'Mens E-Mail': 0
}
dataset.treatment_womens = dataset.treatment.map(treat_dict_womens)
dataset.treatment_womens.value_counts().plot(kind = 'bar', grid = 'True')
stratify_cols = pd.concat([dataset.treatment_womens, dataset.target], axis=1)
stratify_cols.head(5)
from sklearn.model_selection import train_test_split
stratify_cols = pd.concat([dataset.treatment_womens, dataset.target], axis=1)
X_train, X_val, trmnt_train, trmnt_val, y_train, y_val = train_test_split(
dataset.data,
dataset.treatment_womens,
dataset.target,
stratify=stratify_cols,
test_size=0.3,
random_state=42
)
print(f"Train shape: {X_train.shape}")
print(f"Validation shape: {X_val.shape}")
from sklift.models import ClassTransformation
from catboost import CatBoostClassifier
estimator = CatBoostClassifier(verbose=100,
cat_features=['womens', 'mens','channel', 'zip_code', 'history_segment', 'newbie'],
random_state=42,
thread_count=1
)
ct_model = ClassTransformation(estimator=estimator)
ct_model.fit(
X=X_train,
y=y_train,
treatment=trmnt_train
)
#in progress with multiclassclassifier for Catboost.
from sklift.metrics import uplift_at_k
uplift_predictions = ct_model.predict(X_val)
# k = 10%
k = 0.1
# strategy='overall' sort by uplift treatment and control together
uplift_overall = uplift_at_k(y_val, uplift_predictions, trmnt_val, strategy='overall', k=k)
# strategy='by_group' sort by uplift treatment and control separately
uplift_bygroup = uplift_at_k(y_val, uplift_predictions, trmnt_val, strategy='by_group', k=k)
print(f"uplift@{k * 100:.0f}%: {uplift_overall:.4f} (sort groups by uplift together)")
print(f"uplift@{k * 100:.0f}%: {uplift_bygroup:.4f} (sort groups by uplift separately)")
```
## Mens E-mail - No E-mail
```
treat_dict_mens = {
'Mens E-Mail': 1,
'No E-Mail': 0,
'Womens E-Mail': 0
}
dataset.treatment_mens = dataset.treatment.map(treat_dict_mens)
dataset.treatment_mens = dataset.treatment.map(treat_dict_mens)
dataset.treatment_mens.value_counts().plot(kind = 'bar', grid = 'True')
stratify_cols = pd.concat([dataset.treatment_mens, dataset.target], axis=1)
from sklearn.model_selection import train_test_split
stratify_cols = pd.concat([dataset.treatment_mens, dataset.target], axis=1)
X_train, X_val, trmnt_train, trmnt_val, y_train, y_val = train_test_split(
dataset.data,
dataset.treatment_mens,
dataset.target,
stratify=stratify_cols,
test_size=0.3,
random_state=42
)
print(f"Train shape: {X_train.shape}")
print(f"Validation shape: {X_val.shape}")
from sklift.models import ClassTransformation
from catboost import CatBoostClassifier
estimator = CatBoostClassifier(verbose=100,
cat_features=['womens', 'mens','channel', 'zip_code', 'history_segment', 'newbie'],
random_state=42,
thread_count=1
)
ct_model_mens = ClassTransformation(estimator=estimator)
ct_model_mens.fit(
X=X_train,
y=y_train,
treatment=trmnt_train
)
uplift_predictions_mens = ct_model_mens.predict(X_val)
# k = 10%
k = 0.1
# strategy='overall' sort by uplift treatment and control together
uplift_overall_mens = uplift_at_k(y_val, uplift_predictions_mens, trmnt_val, strategy='overall', k=k)
# strategy='by_group' sort by uplift treatment and control separately
uplift_bygroup_mens = uplift_at_k(y_val, uplift_predictions_mens, trmnt_val, strategy='by_group', k=k)
print(f"uplift@{k * 100:.0f}%: {uplift_overall_mens:.4f} (sort groups by uplift_mens together)")
print(f"uplift@{k * 100:.0f}%: {uplift_bygroup_mens:.4f} (sort groups by uplift_mens separately)")
```
| github_jupyter |
```
import numpy as np
folder_name = "fake"
num_row = 10
num_col = 10
O_prob = 0.4
D_prob = 0.4
cap = 2000
length = 0.5
rhoj = 200
num_lane = 2
spd_lb = 20
spd_ub = 50
assign_horizon = 5
demand = 15
link_type = 'CTM'
node_type = 'FWJ'
connector_type = 'PQ'
O_type = 'DMOND'
D_type = 'DMDND'
file_attr = "MNM_input_"
node_file = file_attr + "node"
link_file = file_attr + "link"
snap_file = "Snap_graph"
od_file = file_attr + "od"
demand_file = file_attr + 'demand'
def add_node_str(node_num, type_str):
return str(node_num) + ' ' + type_str + '\n'
def add_snap_str(link_num, from_node, to_node):
return str(link_num) + ' ' + str(from_node) + ' ' + str(to_node) + '\n';
def add_link_str(link_num, link_type):
# CTM 0.8 45 2200 200 1
# PQ 1 99999 99999 99999 1
if link_type != 'PQ':
spd = np.round(np.random.random_sample() * (spd_ub - spd_lb) + np.float(spd_lb))
return str(link_num) + ' ' + str(link_type) + ' ' + str(length) + ' ' + str(spd) + ' ' + str(cap) + ' ' + str(rhoj) + ' ' + str(num_lane) + '\n'
else:
return str(link_num) + ' PQ 1 99999 99999 99999 1\n'
def add_OD(od_id, node_id):
return str(od_id) + ' ' + str(node_id) + '\n'
link_str_list = []
node_str_list = []
snap_str_list = []
O_dict = dict()
D_dict = dict()
O_str_list = []
D_str_list = []
node_counter = 1
link_counter = 1
O_counter = 1
D_counter = 1
for i in xrange(num_row):
for j in xrange(num_col):
node_str_list.append(add_node_str(node_counter, node_type))
if (j != num_col - 1):
node1 = num_col * i + j + 1
node2 = node1 + 1
link_str_list.append(add_link_str(link_counter, link_type))
snap_str_list.append(add_snap_str(link_counter, node1, node2))
link_counter += 1
link_str_list.append(add_link_str(link_counter, link_type))
snap_str_list.append(add_snap_str(link_counter, node2, node1))
link_counter += 1
if(i != num_row - 1):
node1 = num_col * i + j + 1
node2 = node1 + num_col
link_str_list.append(add_link_str(link_counter, link_type))
snap_str_list.append(add_snap_str(link_counter, node1, node2))
link_counter += 1
link_str_list.append(add_link_str(link_counter, link_type))
snap_str_list.append(add_snap_str(link_counter, node2, node1))
link_counter += 1
node_counter += 1
for i in xrange(num_row):
for j in xrange(num_col):
node1 = num_col * i + j + 1
if(np.random.random_sample() < O_prob or (i == 0 and j ==0)):
node_str_list.append(add_node_str(node_counter, O_type))
O_dict[O_counter] = node1
O_str_list.append(add_OD(O_counter, node_counter))
link_str_list.append(add_link_str(link_counter, connector_type))
snap_str_list.append(add_snap_str(link_counter, node_counter, node1))
link_counter += 1
node_counter += 1
O_counter += 1
if(np.random.random_sample() < D_prob or (i == num_row -1 and j == num_col - 1)):
node_str_list.append(add_node_str(node_counter, D_type))
D_dict[D_counter] = node1
D_str_list.append(add_OD(D_counter, node_counter))
link_str_list.append(add_link_str(link_counter, connector_type))
snap_str_list.append(add_snap_str(link_counter, node1, node_counter))
link_counter += 1
node_counter += 1
D_counter += 1
print link_str_list
print node_str_list
print snap_str_list
print O_str_list
print D_str_list
print O_dict
def add_demand_str(O, D):
l = [str(O), str(D)]
d = [str(demand) for i in range(assign_horizon)]
l = l + d
return ' '.join(l) + '\n'
# Origin_ID Destination_ID <demand by interval>
demand_str_list = []
for O, O_node in O_dict.iteritems():
for D, D_node in D_dict.iteritems():
if O_node != D_node:
demand_str_list.append(add_demand_str(O, D))
f = file(link_file, 'w')
f.write("#ID Type LEN(mile) FFS(mile/h) Cap(v/hour) RHOJ(v/miles) Lane\n")
for line in link_str_list:
f.write(line)
f.close()
f = file(node_file, 'w')
f.write('#ID Type\n')
for line in node_str_list:
f.write(line)
f.close()
f = file(snap_file, 'w')
f.write('# EdgeID FromNodeId ToNodeId\n')
for line in snap_str_list:
f.write(line)
f.close()
f = file(od_file, 'w')
f.write('#Origin_ID <-> node_ID\n')
for line in O_str_list:
f.write(line)
f.write('#Dest_ID <-> node_ID\n')
for line in D_str_list:
f.write(line)
f.close()
f = file(demand_file, 'w')
f.write('#Origin_ID Destination_ID <demand by interval>\n')
for line in demand_str_list:
f.write(line)
f.close()
print 'maximum_interval:', assign_horizon
print 'num_of_link:', len(link_str_list)
print 'num_of_node:', len(node_str_list)
print 'num_of_O:', len(O_dict)
print 'num_of_D:', len(D_dict)
print 'num_of_od:', len(demand_str_list)
```
| github_jupyter |
<div style="text-align:center">
<h1> Expressions </h1>
<h2> CS3100 Fall 2019 </h2>
</div>
## Recap
<h4> Last Time: </h4>
* Why functional programming matters?
<h4> Today: </h4>
* Expressions, Values, Definitions.
## Expressions
Every kind of expression has:
* **Syntax**
* **Semantics:**
+ Type-checking rules (static semantics): produce a type or fail with an error message
+ Evaluation rules (dynamic semantics): produce a value
* (or exception or infinite loop)
* **Used only on expressions that type-check** (static vs dynamic languages)
## Values
A *value* is an expression that does not need further evaluation.
<center>
<img src="images/val-expr.svg" width="300">
</center>
## Values in OCaml
```
42
"Hello"
3.1415
```
* Observe that the values have
+ static semantics: types `int`, `string`, `float`.
+ dynamic semantics: the value itself.
## Type Inference and annotation
* OCaml compiler **infers** types
+ Compilation fails with type error if it can't
+ Hard part of language design: guaranteeing compiler can infer types when program is correctly written
* You can manually annotate types anywhere – Replace `e` with `(e : t)`
+ Useful for resolving type errors
```
(42.4 : float)
```
## More values
OCaml also support other values. See [manual](https://caml.inria.fr/pub/docs/manual-ocaml/values.html).
```
()
(1,"hello", true, 3.4)
[1;2;3]
[|1;2;3|]
```
## Static vs Dynamic distinction
Static typing helps catch lots errors at compile time.
Which of these is static error?
```
23 = 45.0
23 = 45
```
## If expression
```ocaml
if e1 then e2 else e3
```
* **Static Semantics:** If `e1` has type `bool`, and `e2` has type `t2` and `e3` has type `t2` then `if e1 then e2 else e3` has type `t2`.
* **Dynamic Semantics:** If `e1` evaluates to true, then evaluate `e2`, else evaluate `e3`
```
if 32 = 31 then "Hello" else "World"
if true then 13 else 13.4
```
## More Formally
<script type="text/x-mathjax-config">
MathJax.Hub.Config({ TeX: { extensions: ["color.js"] }});
</script>
$
\newcommand{\inferrule}[2]{\displaystyle{\frac{#1}{#2}}}
\newcommand{\ite}[3]{\text{if }{#1}\text{ then }{#2}\text{ else }{#3}}
\newcommand{\t}[1]{\color{green}{#1}}
\newcommand{\true}{\color{purple}{true}}
\newcommand{\false}{\color{purple}{false}}
\newcommand{\letin}[3]{\text{let }{{#1} = {#2}}\text{ in }{#3}}
$
**Static Semantics of if expression**
\\[
\inferrule{e1:\t{bool} \quad e2:\t{t} \quad e3:\t{t}}{\ite{e1}{e2}{e3} : \t{t}}
\\]
(omits some details which we will cover in later lectures)
#### to be read as
\\[
\inferrule{Premise_1 \quad Premise_2 \quad \ldots \quad Premise_N}{Conclusion}
\\]
Such rules are known as inference rules.
## Dynamic semantics of if expression
For the case when the predicate evaluates to `true`:
\\[
\inferrule{e1 \rightarrow \true \quad e2 \rightarrow v}{\ite{e1}{e2}{e3} \rightarrow v}
\\]
For the case when the predicate evaluates to `false`:
\\[
\inferrule{e1 \rightarrow \false \quad e3 \rightarrow v}{\ite{e1}{e2}{e3} \rightarrow v}
\\]
Read $\rightarrow$ as *evaluates to*.
## Let expression
```ocaml
let x = e1 in e2
```
* `x` is an identifier
* `e1` is the binding expression
* `e2` is the body expression
* `let x = e1 in e2` is itself an expression
```
let x = 5 in x + 5
let x = 5 in
let y = 10 in
x + y
let x = 5 in
let x = 10 in
x
```
## Scopes & shadowing
```ocaml
let x = 5 in
let x = 10 in
x
```
is parsed as
```ocaml
let x = 5 in
(let x = 10 in
x)
```
* Importantly, `x` is not mutated; there are two `x`s in different **scopes**.
* Inner definitions **shadow** the outer definitions.
```
let x = 5 in
let y =
let x = 10 in
x
in
x+y
```
## let at the top-level
```ocaml
let x = e
```
is implicitly, "**in** the rest of the program text"
```
let a = "Hello"
let b = "World"
let c = a ^ b
```
## Definitions
* The top-level `let x = e` are known as **definitions**.
* Definitions give name to a value.
* Definitions are not expressions, or vice versa.
* But definitions syntactically contain expressions.
<center>
<img src="images/val-expr-defn.svg">
</center>
## Let expression
```ocaml
let x = e1 in e2
```
**Static semantics**
\\[
\inferrule{x : \t{t1} \quad e1 : \t{t1} \quad e2 : \t{t2}}{\letin{x}{e1}{e2} : \t{t2}}
\\]
(again omits some details)
**Dynamic semantics**
\\[
\inferrule{e1 \rightarrow v1 \quad \text{substituting } v1 \text{ for } x \text{ in } e2 \rightarrow v2}
{\letin{x}{e1}{e2} \rightarrow v2}
\\]
## Exercise
In OCaml, we cannot use `+` for floating point addition, and instead have to use `+.`. Why do you think this is the case?
```
5.4 +. 6.0
```
## Exercise
Write down the static semantics for `+` and `+.`.
<div style="text-align:center">
<h1> <i> Fin. </i> </h1>
</div>
| github_jupyter |
# Integrate and fire model
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
from bokeh.plotting import figure
from bokeh.io import output_notebook, show
from numba import jit
output_notebook()
# Define a dictionary to store all parameters
param_dict = {'C': 0.2,
'gL': 10/1000,
'EL': -60,
'sigma': 0.12,
'Vreset': -61,
'Vthresh': -55,
'tau_ref': 2,
'EI': -75,
'EE': -5,
'V': -56,
'dt': 0.1,
'gE': 2.0/1000,
'gI': 1/1000,
'nsteps': 1000,
'smax': 7,
'tau_I': 80,
'wI': 0.0011575}
def update_V(spikes, s, V, params):
'''Update V according to the integrate and fire model'''
smax = params['smax']
Vreset = params['Vreset']
Vthresh = params['Vthresh']
tau_ref = params['tau_ref']
C = params['C']
gL = params['gL']
EL = params['EL']
gE = params['gE']
EE = params['EE']
gI = params['gI']
sigma = params['sigma']
tau_I = params['tau_I']
dt = params['dt']
EI = params['EI']
t = len(spikes)
if np.sum(np.nonzero(spikes)) == 0:
lastfire = -np.inf
else:
lastfire = np.max(np.nonzero(spikes))
if (t - lastfire) * dt < tau_ref:
return 0, Vreset, s - s / tau_I * dt
elif V >= Vthresh:
return 1, Vreset, s + (smax - s) / smax
else:
noise = np.random.normal()
Vdot = (gL * (EL - V) + gE * (EE - V) + gI * (EI - V) + sigma * noise) / C
return 0, V + dt * Vdot, s - s / tau_I * dt
```
## Simulate a spike train with no inhibitory input
```
nsteps = 10000
spikes = []
Vlst = []
slst = []
V = param_dict['EL']
s = 0
for i in range(nsteps):
state, V, s = update_V(spikes, s, V, param_dict)
spikes.append(state)
Vlst.append(V)
slst.append(s)
np.mean(slst)
plt.plot(Vlst)
np.sum(spikes)
p = figure(width=500, height=500)
p.line(np.arange(len(slst)), slst)
show(p)
plt.plot(np.arange(nsteps) * 0.1 / 1000, np.array(Vlst) + np.array(spikes) * 20,
label='Voltage');
plt.plot(np.arange(nsteps) * 0.1 / 1000, np.array(slst) * 3 - 40,
label='Output')
#plt.ylim(-65, -40)
plt.xlabel('Time (s)')
plt.ylabel('voltage (mV)');
plt.legend();
```
## Plot the IO function
```
gI_list = np.arange(8) / 1000
param_for_S2
rate_lst = []
output_lst = []
nsteps = 10000
param_for_S2 = param_dict.copy()
param_for_S2['gE'] = 2/1000
#Vlst = []
for gI in gI_list:
print('Doing gI=', gI)
param_for_S2['gI'] = gI
spikes = []
s = 0
V = param_for_S2['EL']
slst = []
for i in range(nsteps):
state, V, s = update_V(spikes, s, V, param_for_S2)
#Vlst.append(V)
spikes.append(state)
slst.append(s)
output_lst.append(np.mean(slst))
rate_lst.append(np.sum(spikes))
plt.plot(Vlst)
plt.plot(gI_list * 1000, rate_lst)
plt.xlabel('gI (nS)')
plt.ylabel('Average firing rate (Hz)')
plt.plot(gI_list * 1000, output_lst)
plt.xlabel('gI (nS)')
plt.ylabel('Synaptic Output')
plt.plot(rate_lst, output_lst)
plt.xlabel('gI (nS)')
plt.ylabel('Synaptic Output')
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import xgboost as xgb
import matplotlib.pylab as plt
%matplotlib inline
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.datasets.samples_generator import make_classification
# X为样本特征,y为样本类别输出, 共10000个样本,每个样本20个特征,输出有2个类别,没有冗余特征,每个类别一个簇
X, y = make_classification(n_samples=10000, n_features=20, n_redundant=0,
n_clusters_per_class=1, n_classes=2, flip_y=0.1)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
print (X_train.shape)
print (y_train.shape)
print (X_test.shape)
print (y_test.shape)
```
<h1>XGBoost 使用原生API</h1>
```
dtrain = xgb.DMatrix(X_train,y_train)
dtest = xgb.DMatrix(X_test,y_test)
param = {'max_depth':5, 'eta':0.5, 'verbosity':1, 'objective':'binary:logistic'}
raw_model = xgb.train(param, dtrain, num_boost_round=20)
from sklearn.metrics import accuracy_score
pred_train_raw = raw_model.predict(dtrain)
for i in range(len(pred_train_raw)):
if pred_train_raw[i] > 0.5:
pred_train_raw[i]=1
else:
pred_train_raw[i]=0
print (accuracy_score(dtrain.get_label(), pred_train_raw))
pred_test_raw = raw_model.predict(dtest)
for i in range(len(pred_test_raw)):
if pred_test_raw[i] > 0.5:
pred_test_raw[i]=1
else:
pred_test_raw[i]=0
print (accuracy_score(dtest.get_label(), pred_test_raw))
```
<h1>XGBoost 使用sklearn wrapper,仍然使用原始API的参数</h1>
```
sklearn_model_raw = xgb.XGBClassifier(**param)
sklearn_model_raw.fit(X_train, y_train, early_stopping_rounds=10, eval_metric="error",
eval_set=[(X_test, y_test)])
```
<h1>XGBoost 使用sklearn wrapper,使用sklearn风格的参数(推荐)</h1>
```
sklearn_model_new = xgb.XGBClassifier(max_depth=5,learning_rate= 0.5, verbosity=1, objective='binary:logistic',random_state=1)
sklearn_model_new.fit(X_train, y_train, early_stopping_rounds=10, eval_metric="error",
eval_set=[(X_test, y_test)])
```
<h1>使用sklearn网格搜索调参</h1>
一般固定步长,先调好框架参数n_estimators,再调弱学习器参数max_depth,min_child_weight,gamma等,接着调正则化相关参数subsample,colsample_byXXX, reg_alpha以及reg_lambda,最后固定前面调好的参数,来调步长learning_rate
```
gsCv = GridSearchCV(sklearn_model_new,
{'max_depth': [4,5,6],
'n_estimators': [5,10,20]})
gsCv.fit(X_train,y_train)
print(gsCv.best_score_)
print(gsCv.best_params_)
sklearn_model_new2 = xgb.XGBClassifier(max_depth=4,n_estimators=10,verbosity=1, objective='binary:logistic',random_state=1)
gsCv2 = GridSearchCV(sklearn_model_new2,
{'learning_rate ': [0.3,0.5,0.7]})
gsCv2.fit(X_train,y_train)
print(gsCv2.best_score_)
print(gsCv2.best_params_)
sklearn_model_new2 = xgb.XGBClassifier(max_depth=4,learning_rate= 0.3, verbosity=1, objective='binary:logistic',n_estimators=10)
sklearn_model_new2.fit(X_train, y_train, early_stopping_rounds=10, eval_metric="error",
eval_set=[(X_test, y_test)])
pred_test_new = sklearn_model_new2.predict(X_test)
print (accuracy_score(dtest.get_label(), pred_test_new))
```
| github_jupyter |
# Support Vector Classification with Robust Scaler & Power Transformer
This Code template is for the Classification task using Support Vector Classifier(SVC) based on the Support Vector Machine algorithm with PowerTransformer as Feature Transformation Technique and rescaling technique as Robust Scaler in a pipeline.
### Required Packages
```
!pip install imblearn
import warnings
import seaborn as se
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import RandomOverSampler
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import LabelEncoder, PowerTransformer, RobustScaler
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
Support vector machines (SVMs) are a set of supervised learning methods used for classification, regression and outliers detection.
A Support Vector Machine is a discriminative classifier formally defined by a separating hyperplane. In other terms, for a given known/labelled data points, the SVM outputs an appropriate hyperplane that classifies the inputted new cases based on the hyperplane. In 2-Dimensional space, this hyperplane is a line separating a plane into two segments where each class or group occupied on either side.
Here we have used SVC, the svc implementation is based on libsvm.<br/>
Model Tuning Parameters
C -> Regularization parameter. The strength of the regularization is inversely proportional to C. Must be strictly positive.
kernel -> Specifies the kernel type to be used in the algorithm. It must be one of ‘linear’, ‘poly’, ‘rbf’, ‘sigmoid’, ‘precomputed’ or a callable. If none is given, ‘rbf’ will be used. If a callable is given it is used to pre-compute the kernel matrix from data matrices; that matrix should be an array of shape (n_samples, n_samples).
gamma -> Gamma is a hyperparameter that we have to set before the training model. Gamma decides how much curvature we want in a decision boundary.
degree -> Degree of the polynomial kernel function (‘poly’). Ignored by all other kernels.Using degree 1 is similar to using a linear kernel. Also, increasing degree parameter leads to higher training times.
#### Rescaling technique
Standardization of a dataset is a common requirement for many machine learning estimators. Typically this is done by removing the mean and scaling to unit variance. However, outliers can often influence the sample mean / variance in a negative way. In such cases, the median and the interquartile range often give better results.
The Robust Scaler removes the median and scales the data according to the quantile range (defaults to IQR: Interquartile Range). The IQR is the range between the 1st quartile (25th quantile) and the 3rd quartile (75th quantile).
#### Feature Transformation
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired. The optimal parameter for stabilizing variance and minimizing skewness is estimated through maximum likelihood.
<br/><br/>
For more information on Power transformers [click here](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PowerTransformer.html?highlight=power%20transformer#sklearn.preprocessing.PowerTransformer)
```
model = make_pipeline(RobustScaler(),PowerTransformer(),SVC(random_state=123))
model.fit(x_train,y_train)
```
### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
where:
Precision:- Accuracy of positive predictions.
Recall:- Fraction of positives that were correctly identified.
f1-score:- percent of positive predictions were correct
support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Arpit Somani , Github: [Profile](https://github.com/arpitsomani8)
| github_jupyter |
# COVID19-related literature SQL database
In this notebook, we create a relational database dump of a set of COVID19-related publication datasets. These include:
* CORD19: https://pages.semanticscholar.org/coronavirus-research
* Dimensions: https://docs.google.com/spreadsheets/d/1-kTZJZ1GAhJ2m4GAIhw1ZdlgO46JpvX0ZQa232VWRmw/edit#gid=2034285255
```
# magics, warnings and imports
%load_ext autoreload
%autoreload 2
import warnings; warnings.simplefilter('ignore')
import os, random, codecs, json, math, re
import pandas as pd
import numpy as np
from tqdm.notebook import tqdm
import pymysql
from sqlalchemy import create_engine
from sqlalchemy import Integer,String,Boolean,DateTime
seed = 99
random.seed(seed)
np.random.seed(seed)
import nltk, sklearn
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="white")
sns.set_context("notebook", font_scale=1.2, rc={"lines.linewidth": 2.5})
```
#### Load datasets
```
# point here to the versions of the datasets you want to use
dimensions_filename = "datasets_input/Dimensions_01_07_2020.csv"
cord19_folder = "datasets_input/CORD19_01_07_2020"
df_dimensions = pd.read_csv(dimensions_filename, dtype=str, sep=";")
df_cord = pd.read_csv(os.path.join(cord19_folder,"metadata.csv"), dtype=str)
df_cord.shape
df_dimensions.shape
```
### Prepare dataframes for ingestion
#### Clean-up data frames
##### Dimensions
```
df_dimensions.head()
df_dimensions.columns
df_dimensions.drop(columns=['Date added', 'Publisher', 'Authors', 'Corresponding Authors',
'Authors Affiliations', 'Research Organizations - standardized',
'GRID IDs', 'City of Research organization',
'Country of Research organization', 'Funder',
'UIDs of supporting grants', 'Times cited', 'Altmetric',
'Source Linkout'], inplace=True)
df_dimensions.columns
df_dimensions.rename(columns={'Publication ID':'dimensions_id', 'DOI':'doi', 'PMID':'pmid', 'PMCID':'pmcid', 'Title':'title', 'Abstract':'abstract',
'Source title':'journal', 'Source UID':'source_uid', 'MeSH terms':'mesh_terms', 'Publication Date':'publication_date',
'PubYear':'publication_year', 'Volume':'volume', 'Issue':'issue', 'Pagination':'pages', 'Open Access':'open_access',
'Publication Type':'publication_type', 'Dimensions URL':'dimensions_url'}, inplace=True)
df_dimensions[pd.isnull(df_dimensions.abstract)].shape
def get_year(date):
if isinstance(date,str) and len(date)>3 and date[:4].isdigit():
return date[:4]
return ""
month_to_number = {"Jan":"1","Feb":"2","Mar":"3","Apr":"4","May":"5","Jun":"6","Jul":"7","Aug":"8","Sep":"9","Oct":"10","Nov":"11","Dec":"12"}
def get_month(date):
if isinstance(date,str) and len(date)>6:
if "-" in date and date.split("-")[1].isdigit():
return str(int(date.split("-")[1]))
else:
try:
return month_to_number[date.split()[1]]
except:
return ""
return ""
def sanitize_string(s):
return " ".join(s.split())
df_dimensions["publication_year"] = df_dimensions["publication_year"].apply(get_year)
df_dimensions["publication_month"] = df_dimensions["publication_date"].apply(get_month)
df_dimensions["arxiv_id"] = ""
df_dimensions.drop(columns="publication_date", inplace=True)
df_dimensions = df_dimensions.fillna('')
```
Retrieve arXiv ID for Dimensions papers
```
# Sync option, quite slow
"""
import requests, re
for index, row in df_dimensions[df_dimensions.journal=="arXiv"].iterrows():
r = requests.get(row["dimensions_url"])
if r.status_code == 200:
arxiv_url = re.search("https://arxiv.org/pdf/arXiv:[0-9]+.[0-9]+",r.text).group()
arxiv_id = arxiv_url.split(":")[-1]
row["arxiv_id"] = arxiv_id
"""
# from https://towardsdatascience.com/fast-and-async-in-python-accelerate-your-requests-using-asyncio-62dafca83c33
# aSync option, fast
import aiohttp
import requests
import asyncio
from aiohttp import ClientSession
def extract_fields_from_response(response):
"""Extract arxiv id from API's response"""
search_res = re.search("arXiv:[0-9]+.[0-9]+",response)
if search_res:
arxiv_url = search_res.group()
arxiv_id = arxiv_url.split(":")[-1]
return arxiv_id
return ""
async def get_details_async(session, payload):
response = await session.request(method='GET', url=payload[1]["dimensions_url"])
if response.status != 200:
print(payload[0], response.status)
pass
return await response.text()
async def run_program(session, payload):
"""Wrapper for running program in an asynchronous manner"""
response = await get_details_async(session, payload)
arxiv_id = extract_fields_from_response(response)
return (arxiv_id,payload[1]["dimensions_url"])
payloads = df_dimensions[df_dimensions.journal=="arXiv"]
connector = aiohttp.TCPConnector(limit=5)
async with aiohttp.ClientSession(connector=connector) as session:
results = await asyncio.gather(*[run_program(session, payload) for payload in payloads.iterrows()])
df_tmp = pd.DataFrame.from_dict({"arxiv_id":[x[0] for x in results],"dimensions_url":[x[1] for x in results]})
df_tmp.shape
df_dimensions = df_dimensions.merge(df_tmp, how="left", left_on="dimensions_url", right_on="dimensions_url")
df_dimensions.drop(columns="arxiv_id_x", inplace=True)
df_dimensions.rename(columns={"arxiv_id_y":"arxiv_id"}, inplace=True)
df_dimensions = df_dimensions.fillna('')
df_dimensions.shape
df_dimensions[df_dimensions.journal=="arXiv"].shape
df_dimensions.head()
df_dimensions[df_dimensions.doi==""].shape
```
##### CORD19
```
df_cord.head()
df_cord["license"].value_counts()
df_cord["source_x"].value_counts()
# how many full text files
df_cord[(pd.notnull(df_cord.pdf_json_files))|(pd.notnull(df_cord.pmc_json_files))].shape
df_cord[(pd.notnull(df_cord.pdf_json_files))|(pd.notnull(df_cord.pmc_json_files))].shape[0]/df_cord.shape[0]
# check Medline
df_cord[(df_cord.source_x=="Medline")&((pd.notnull(df_cord.pdf_json_files))|(pd.notnull(df_cord.pmc_json_files)))].shape[0]/df_cord[(df_cord.source_x=="Medline")].shape[0]
df_cord[(df_cord.source_x=="Medline")].shape[0]
# check NOT Medline
df_cord[(df_cord.source_x!="Medline")&((pd.notnull(df_cord.pdf_json_files))|(pd.notnull(df_cord.pmc_json_files)))].shape[0]/df_cord[(df_cord.source_x!="Medline")].shape[0]
df_cord[(df_cord.source_x!="Medline")].shape[0]
# NEW columns (for now, we drop)
df_cord.drop(columns=["cord_uid","url","pdf_json_files","pmc_json_files","s2_id"],inplace=True)
df_cord.columns
df_cord.drop(columns='authors', inplace=True)
df_cord = df_cord.fillna('')
df_cord.rename(columns={'source_x':'source', 'pubmed_id': 'pmid',
'mag_id': 'ms_academic_id', 'who_covidence_id': 'who_covidence'}, inplace=True)
df_cord["publication_year"] = df_cord["publish_time"].apply(get_year)
df_cord["publication_month"] = df_cord["publish_time"].apply(get_month)
df_cord.drop(columns='publish_time', inplace=True)
df_cord['pages'] = ""
df_cord['volume'] = ""
df_cord['issue'] = ""
df_cord["dimensions_id"] = ""
df_cord.head()
df_cord[(df_cord.doi=="") & ((df_cord.sha!="") | (df_cord.pmid!="") | (df_cord.pmcid!="") | df_cord.arxiv_id!="")].shape
df_cord[(df_cord.doi=="") & (df_cord.pmid=="") & (df_cord.pmcid=="")].shape
df_cord[((df_cord.doi=="") & (df_cord.pmid=="") & (df_cord.pmcid==""))&(df_cord.arxiv_id!="")].shape
df_dimensions.shape
df_cord.shape
df_dimensions[((df_dimensions.doi=="") & (df_dimensions.pmid=="") & (df_dimensions.pmcid==""))&(df_dimensions.journal=="arXiv")].shape
df_dimensions[df_dimensions.journal=="arXiv"].shape
df_cord[df_cord.arxiv_id!=""].shape
df_cord[df_cord.title=="CORD-19: The Covid-19 Open Research Dataset"]
df_dimensions[df_dimensions.title=="CORD-19: The Covid-19 Open Research Dataset"]
# license
df_sub = df_cord[df_cord.license.isin(df_cord.license.value_counts()[:30].index.tolist())]
b = sns.countplot(y="license", data=df_sub, order=df_sub['license'].value_counts().index)
#b.axes.set_title("Title",fontsize=50)
b.set_xlabel("Count",fontsize=15)
b.set_ylabel("License",fontsize=15)
b.tick_params(labelsize=15)
```
### Prepare tables
```
# the main table: pub
pub_table_columns = ['title','abstract','publication_year','publication_month','journal','volume','issue','pages','doi','pmid','pmcid','dimensions_id','arxiv_id']
df_pub = df_dimensions[pub_table_columns].append(df_cord[pub_table_columns], ignore_index=True)
df_pub["title"] = df_pub["title"].apply(sanitize_string)
df_pub["abstract"] = df_pub["abstract"].apply(sanitize_string)
df_pub["doi"] = df_pub["doi"].apply(str.lower)
df_pub["pmid"] = df_pub["pmid"].apply(str.lower)
df_pub["pmcid"] = df_pub["pmcid"].apply(str.lower)
df_pub["dimensions_id"] = df_pub["dimensions_id"].apply(str.lower)
df_pub["arxiv_id"] = df_pub["arxiv_id"].apply(str.lower)
df_pub.shape
df_pub[(df_pub.doi=="") & (df_pub.pmid=="") & (df_pub.pmcid=="") & (df_pub.dimensions_id=="") & (df_pub.arxiv_id=="")].shape
# check to have at least one valid identifier per publication
# we drop publications which do not: hopefully, they will be equipped with an identifier in future releases
df_pub = df_pub[~((df_pub.doi=="") & (df_pub.pmid=="") & (df_pub.pmcid=="") & (df_pub.dimensions_id=="") & (df_pub.arxiv_id==""))]
df_pub[df_pub.doi=="0.1126/science.abb7331"]
# drop duplicates, first on dois then pmids then pmcids. We need this to keep empty values!
df_tmp = df_pub[df_pub.doi==""]
df_pub1 = df_pub[df_pub.doi!=""].groupby('doi').first()
df_pub1.reset_index(inplace=True)
df_tmp2 = df_tmp[df_tmp.pmid==""]
df_pub2 = df_tmp[df_tmp.pmid!=""].groupby('pmid').first()
df_pub2.reset_index(inplace=True)
df_tmp3 = df_tmp2[df_tmp2.pmcid==""]
df_pub3 = df_tmp2[df_tmp2.pmcid!=""].groupby('pmcid').first()
df_pub3.reset_index(inplace=True)
df_tmp4 = df_tmp3[df_tmp3.arxiv_id==""]
df_pub4 = df_tmp3[df_tmp3.arxiv_id!=""].groupby('arxiv_id').first()
df_pub4.reset_index(inplace=True)
df_pub5 = df_tmp4[df_tmp4.dimensions_id!=""].groupby('dimensions_id').first()
df_pub5.reset_index(inplace=True)
df_pub1[df_pub1.doi=="0.1126/science.abb7331"]
df_pub = pd.concat([df_pub1,df_pub2,df_pub3,df_pub4,df_pub5])
# add PK and reset index
df_pub.reset_index(drop=True,inplace=True)
df_pub["pub_id"] = df_pub.index.values
df_pub.shape
df_pub[df_pub.arxiv_id=="2003.08720"]
# create other tables via joins
df_datasource = pd.DataFrame.from_dict({"source":["CORD19","Dimensions"],"url":["https://pages.semanticscholar.org/coronavirus-research","https://docs.google.com/spreadsheets/d/1-kTZJZ1GAhJ2m4GAIhw1ZdlgO46JpvX0ZQa232VWRmw/edit#gid=2034285255"]})
df_cord_metadata = df_cord[['source','license','ms_academic_id','who_covidence','doi','pmid','pmcid','sha','arxiv_id']]
df_dimensions_metadata = df_dimensions[['dimensions_id', 'doi', 'pmid', 'pmcid','arxiv_id', 'source_uid', 'mesh_terms',
'open_access', 'publication_type', 'dimensions_url']]
df_cord_metadata["doi"] = df_cord_metadata["doi"].apply(str.lower)
df_cord_metadata["pmid"] = df_cord_metadata["pmid"].apply(str.lower)
df_cord_metadata["pmcid"] = df_cord_metadata["pmcid"].apply(str.lower)
df_dimensions_metadata["doi"] = df_dimensions_metadata["doi"].apply(str.lower)
df_dimensions_metadata["pmid"] = df_dimensions_metadata["pmid"].apply(str.lower)
df_dimensions_metadata["pmcid"] = df_dimensions_metadata["pmcid"].apply(str.lower)
df_datasource.head()
# CORD19 metadata
df_cord_metadata.shape
df_pub[df_pub.doi=="0.1126/science.abb7331"]
#We need this to keep empty values!
df_tmp = df_cord_metadata[df_cord_metadata.doi==""]
df_cord_metadata1 = pd.merge(df_cord_metadata[df_cord_metadata.doi!=""], df_pub[['pub_id','doi']], how='inner', left_on=['doi'], right_on=['doi'])
df_tmp2 = df_tmp[df_tmp.pmid==""]
df_cord_metadata2 = pd.merge(df_tmp[df_tmp.pmid!=""], df_pub[['pub_id','pmid']], how='inner', left_on=['pmid'], right_on=['pmid'])
df_tmp3 = df_tmp2[df_tmp2.pmcid==""]
df_cord_metadata3 = pd.merge(df_tmp2[df_tmp2.pmcid!=""], df_pub[['pub_id','pmcid']], how='inner', left_on=['pmcid'], right_on=['pmcid'])
df_cord_metadata4 = pd.merge(df_tmp3[df_tmp3.arxiv_id!=""], df_pub[['pub_id','arxiv_id']], how='inner', left_on=['arxiv_id'], right_on=['arxiv_id'])
df_cord_metadata1 = df_cord_metadata1.groupby("doi").first()
df_cord_metadata1.reset_index(inplace=True)
df_cord_metadata2 = df_cord_metadata2.groupby("pmid").first()
df_cord_metadata2.reset_index(inplace=True)
df_cord_metadata3 = df_cord_metadata3.groupby("pmcid").first()
df_cord_metadata3.reset_index(inplace=True)
df_cord_metadata4 = df_cord_metadata4.groupby("arxiv_id").first()
df_cord_metadata4.reset_index(inplace=True)
df_cord_metadata = pd.concat([df_cord_metadata1,df_cord_metadata2,df_cord_metadata3,df_cord_metadata4])
df_cord_metadata.shape
# read full texts in
folders = ['document_parses/pdf_json']
shas = list()
full_texts = list()
for folder in folders:
for root, dirs, files in os.walk(os.path.join(cord19_folder,folder)):
for file in tqdm(files):
if ".json" in file: # read
data = json.loads(codecs.open(os.path.join(root,file)).read())
sha = data["paper_id"]
full_text = " ".join(sanitize_string(section["text"]) for section in data["body_text"])
shas.append(sha)
full_texts.append(full_text)
df_cord_fulltext = pd.DataFrame.from_dict({"sha":shas,"full_text":full_texts})
df_cord_fulltext.shape
df_cord_metadata = pd.merge(df_cord_metadata, df_cord_fulltext, how='left', left_on=['sha'], right_on=['sha'])
df_cord_metadata = df_cord_metadata.fillna('')
df_cord_metadata.rename(columns={"id":"pub_id"},inplace=True)
df_cord_metadata.head()
df_cord_metadata[df_cord_metadata.arxiv_id!=""].head()
# Dimensions metadata
df_tmp = df_dimensions_metadata[df_dimensions_metadata.dimensions_id==""]
df_dimensions_metadata1 = pd.merge(df_dimensions_metadata[df_dimensions_metadata.dimensions_id!=""], df_pub[['pub_id','dimensions_id']], how='inner', left_on=['dimensions_id'], right_on=['dimensions_id'])
df_dimensions_metadata1 = df_dimensions_metadata1.groupby("dimensions_id").first()
df_dimensions_metadata1.reset_index(inplace=True)
df_dimensions_metadata = pd.concat([df_dimensions_metadata1])
df_dimensions_metadata.shape
df_dimensions_metadata.rename(columns={"id":"pub_id"},inplace=True)
# Create datasource tables
cord_source_id = df_datasource[df_datasource.source=="CORD19"].index.values[0]
dimensions_source_id = df_datasource[df_datasource.source=="Dimensions"].index.values[0]
df_cord_metadata["source_id"] = cord_source_id
df_dimensions_metadata["source_id"] = dimensions_source_id
df_pub_to_datasource = df_cord_metadata[["pub_id","source_id"]]
df_pub_to_datasource = df_pub_to_datasource.append(df_dimensions_metadata[["pub_id","source_id"]],ignore_index=True)
df_pub_to_datasource.drop_duplicates(inplace=True)
df_pub_to_datasource.rename(columns={"source_id":"datasource_id"},inplace=True)
df_pub_to_datasource.shape
df_pub_to_datasource[df_pub_to_datasource.pub_id==22787]
# remove unnecessary columns
df_cord_metadata.drop(columns=['doi','pmid','pmcid','arxiv_id','source_id'],inplace=True)
df_dimensions_metadata.drop(columns=['doi','pmid','pmcid','arxiv_id','source_id'],inplace=True)
# reset all indexes which will become PKs
df_cord_metadata.reset_index(drop=True,inplace=True)
df_dimensions_metadata.reset_index(drop=True,inplace=True)
df_datasource.reset_index(drop=True,inplace=True)
df_cord_metadata["cord19_metadata_id"] = df_cord_metadata.index.values
df_dimensions_metadata["dimensions_metadata_id"] = df_dimensions_metadata.index.values
df_datasource["datasource_metadata_id"] = df_datasource.index.values
# make numeric where needed
df_pub["publication_year"] = pd.to_numeric(df_pub["publication_year"])
df_pub["publication_month"] = pd.to_numeric(df_pub["publication_month"])
df_pub["pmid"] = pd.to_numeric(df_pub["pmid"])
# add timestamp
df_pub["timestamp"] = pd.Timestamp.now()
# clean-up text (optional)
replaces = [""]
def clean_up(txt):
for r in replaces:
txt = txt.replace(r,"")
return txt.encode('utf8', 'ignore').decode('utf8')
df_pub["abstract"] = [clean_up(a) for a in df_pub["abstract"].values]
df_pub.head()
# reorder the columns to match the SQL schema
df_datasource.columns
df_pub = df_pub[['pub_id', 'title', 'abstract', 'publication_year', 'publication_month', 'journal',
'volume', 'issue', 'pages', 'doi', 'pmid', 'pmcid', 'dimensions_id', 'arxiv_id',
'timestamp']]
df_dimensions_metadata = df_dimensions_metadata[['dimensions_metadata_id', 'dimensions_id', 'source_uid', 'open_access',
'publication_type', 'dimensions_url', 'mesh_terms', 'pub_id']]
df_cord_metadata = df_cord_metadata[[ 'cord19_metadata_id', 'source', 'license', 'ms_academic_id',
'who_covidence', 'sha', 'full_text', 'pub_id']]
df_datasource = df_datasource[['datasource_metadata_id', 'source', 'url']]
df_pub.doi.value_counts()
df_pub.arxiv_id.value_counts()
df_pub[df_pub.doi == "10.1016/s0140-6736(20)30607-3"].doi.to_string()
df_cord_metadata.columns
```
### Dump to CSV
```
### Export the df_pub dataframe for further use
df_pub.to_csv("datasets_output/df_pub.csv", compression="gzip", index=False)
# export TSV for ingestion
df_pub.to_csv("datasets_output/sql_tables/pub.csv",index=False,sep="\t",header=False)
df_cord_metadata.to_csv("datasets_output/sql_tables/cord19_metadata.csv",index=False,sep="\t",header=False)
df_dimensions_metadata.to_csv("datasets_output/sql_tables/dimensions_metadata.csv",index=False,sep="\t",header=False)
df_datasource.to_csv("datasets_output/sql_tables/datasource.csv",index=False,sep="\t",header=False)
df_pub_to_datasource.to_csv("datasets_output/sql_tables/pub_datasource.csv",index=False,sep="\t",header=False)
```
### Dump to MySQL
Use this if you want to create a MySQL db.
```
dtype_dict = {'pub_id':Integer, 'title':String, 'abstract':String, 'publication_year':Integer, 'publication_month':Integer, 'journal':String,
'volume':String, 'issue':String, 'pages':String, 'doi':String, 'pmid':Integer, 'pmcid':String, 'timestamp':DateTime}
# get API key
import configparser
config = configparser.ConfigParser()
config.read("credentials/conf.ini")
mysql_username = config["MYSQL"]["username"]
mysql_password = config["MYSQL"]["password"]
mysql_database = config["MYSQL"]["database"]
sqlEngine = create_engine('mysql+pymysql://%s:%s@127.0.0.1/%s'%(mysql_username,mysql_password,mysql_database), pool_recycle=3600)
dbConnection = sqlEngine.connect()
# main table
table_name = "pub"
try:
frame = df_pub.to_sql(table_name, dbConnection, if_exists='append', index=False, index_label="pub_id", dtype=dtype_dict);
except ValueError as vx:
print(vx)
except Exception as ex:
print(ex)
else:
print("Table %s created successfully."%table_name);
finally:
dbConnection.close()
sqlEngine = create_engine('mysql+pymysql://%s:%s@127.0.0.1/%s'%(mysql_username,mysql_password,mysql_database), pool_recycle=3600)
dbConnection = sqlEngine.connect()
# other tables
try:
frame = df_cord_metadata.to_sql("cord19_metadata", dbConnection, if_exists='append', index=True, index_label="cord19_metadata_id")
frame = df_who_metadata.to_sql("who_metadata", dbConnection, if_exists='append', index=True, index_label="who_metadata_id")
frame = df_dimensions_metadata.to_sql("dimensions_metadata", dbConnection, if_exists='append', index=True, index_label="dimensions_metadata_id")
frame = df_datasource.to_sql("datasource", dbConnection, if_exists='append', index=True, index_label="datasource_id")
except ValueError as vx:
print(vx)
except Exception as ex:
print(ex)
else:
print("Tables created successfully.");
finally:
dbConnection.close()
sqlEngine = create_engine('mysql+pymysql://%s:%s@127.0.0.1/%s'%(mysql_username,mysql_password,mysql_database), pool_recycle=3600)
dbConnection = sqlEngine.connect()
# last table
try:
frame = df_pub_to_datasource.to_sql("pub_datasource", dbConnection, if_exists='append', index=False, index_label=["pub_id","datasource_id"])
except ValueError as vx:
print(vx)
except Exception as ex:
print(ex)
else:
print("Table created successfully.");
finally:
dbConnection.close()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from tqdm import tqdm_notebook as tqdm
# dataset_final.zip is the final dataset generated
# please download it from the link provided in readme before running this cell
from zipfile import ZipFile
path = '/Dataset/dataset_final.zip'
with ZipFile(path, 'r') as zip:
zip.printdir()
data = pd.read_csv(path)
# This block should be used to visualise the dataset as it is a highly compressed zip file
print(data.shape)
data.head()
#data.rename(columns=lambda x: x[0:3], inplace=True)
#data.drop(columns=['Unn'],inplace=True)
#data.rename(columns={'A n':'title','AIM':'abstract'},inplace=True)
data['title'].replace('', np.nan, inplace=True)
data['abstract'].replace('', np.nan, inplace=True)
data.dropna(axis=0,how='any',inplace=True)
import re
data = data.applymap(lambda x: re.sub(' +',' ',x))
p = data.title.str.count(' ')
from collections import Counter
cntr = Counter(p)
tot = 0
for i in range(7):
tot+=cntr[i]
print(i,tot)
for i in range(16,18):
tot+=cntr[i]
print(i,tot)
col = []
for i in tqdm(range(data.shape[0])):
tw = data.iloc[i]['title'].count(' ')
aw = data.iloc[i]['abstract'].count(' ')
if tw<7 or tw>15 or aw<80 or aw>250:
col.append(i)
len(col),data.shape[0]-len(col)
data.drop(col,inplace=True)
data.drop_duplicates(inplace=True)
path = '/Dataset/dataset_final.zip'
data.to_csv(path,index=False,compression='zip')
title,abstract = list(data['title']),list(data['abstract'])
def remove_newline(arr):
for i in tqdm(range(len(arr))):
arr[i] = arr[i].replace('\n',' ')
return arr
title = remove_newline(title)
abstract = remove_newline(abstract)
from sklearn.model_selection import train_test_split
titleTrain, titleValid, abstractTrain, abstractValid = train_test_split(title, abstract, test_size = 0.2, random_state = 42)
titleValid, titleTest,abstractValid, abstractTest = train_test_split(titleValid, abstractValid, test_size = 0.5, random_state = 42)
print(len(title),len(abstract))
print(len(titleTrain),len(abstractTrain))
print(len(titleValid),len(abstractValid))
print(len(titleTest),len(abstractTest))
from bz2 import open
def store_to_file(arr,fl):
arr = '\n'.join(arr)
with open(fl,'w') as f:
f.write(arr.encode())
path = '/Dataset/'
store_to_file(titleTrain[:600000],path+'title.train.bz2')
store_to_file(titleTest[:100000],path+'title.test.bz2')
store_to_file(titleValid[:100000],path+'title.valid.bz2')
store_to_file(abstractTrain[:600000],path+'abstract.train.bz2')
store_to_file(abstractTest[:100000],path+'abstract.test.bz2')
store_to_file(abstractValid[:100000],path+'abstract.valid.bz2')
```
| github_jupyter |
### This jupyter notebooks provides the code for classifying signals using the Discrete Wavelet Transform.
### To get some more background information, please have a look at the accompanying blog-post:
### http://ataspinar.com/2018/12/21/a-guide-for-using-the-wavelet-transform-in-machine-learning/
```
import os
import time
import numpy as np
import pandas as pd
import scipy.io as sio
from IPython.display import display
import matplotlib.pyplot as plt
import pywt
import scipy.stats
import datetime as dt
from collections import defaultdict, Counter
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
from sklearn.neural_network import MLPClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import AdaBoostClassifier
from sklearn.gaussian_process import GaussianProcessClassifier
dict_classifiers = {
"Gradient Boosting Classifier": GradientBoostingClassifier(),
"Random Forest": RandomForestClassifier(),
"Logistic Regression": LogisticRegression(),
"Nearest Neighbors": KNeighborsClassifier(),
"Decision Tree": DecisionTreeClassifier(),
"Linear SVM": SVC(),
"Neural Net": MLPClassifier(alpha = 1),
"Naive Bayes": GaussianNB(),
"AdaBoost": AdaBoostClassifier(),
"Gaussian Process": GaussianProcessClassifier()
}
def batch_classify(X_train, Y_train, X_test, Y_test, no_classifiers = 5, verbose = True):
"""
This method, takes as input the X, Y matrices of the Train and Test set.
And fits them on all of the Classifiers specified in the dict_classifier.
Usually, the SVM, Random Forest and Gradient Boosting Classifier take quiet some time to train.
So it is best to train them on a smaller dataset first and
decide whether you want to comment them out or not based on the test accuracy score.
"""
dict_models = {}
for classifier_name, classifier in list(dict_classifiers.items())[:no_classifiers]:
t_start = time.clock()
classifier.fit(X_train, Y_train)
t_end = time.clock()
t_diff = t_end - t_start
train_score = classifier.score(X_train, Y_train)
test_score = classifier.score(X_test, Y_test)
dict_models[classifier_name] = {'model': classifier, 'train_score': train_score, 'test_score': test_score, 'train_time': t_diff}
if verbose:
print("trained {c} in {f:.2f} s".format(c=classifier_name, f=t_diff))
return dict_models
def get_train_test(df, y_col, x_cols, ratio):
"""
This method transforms a dataframe into a train and test set, for this you need to specify:
1. the ratio train : test (usually 0.7)
2. the column with the Y_values
"""
mask = np.random.rand(len(df)) < ratio
df_train = df[mask]
df_test = df[~mask]
Y_train = df_train[y_col].values
Y_test = df_test[y_col].values
X_train = df_train[x_cols].values
X_test = df_test[x_cols].values
return df_train, df_test, X_train, Y_train, X_test, Y_test
def display_dict_models(dict_models, sort_by='test_score'):
cls = [key for key in dict_models.keys()]
test_s = [dict_models[key]['test_score'] for key in cls]
training_s = [dict_models[key]['train_score'] for key in cls]
training_t = [dict_models[key]['train_time'] for key in cls]
df_ = pd.DataFrame(data=np.zeros(shape=(len(cls),4)), columns = ['classifier', 'train_score', 'test_score', 'train_time'])
for ii in range(0,len(cls)):
df_.loc[ii, 'classifier'] = cls[ii]
df_.loc[ii, 'train_score'] = training_s[ii]
df_.loc[ii, 'test_score'] = test_s[ii]
df_.loc[ii, 'train_time'] = training_t[ii]
display(df_.sort_values(by=sort_by, ascending=False))
def calculate_entropy(list_values):
counter_values = Counter(list_values).most_common()
probabilities = [elem[1]/len(list_values) for elem in counter_values]
entropy=scipy.stats.entropy(probabilities)
return entropy
def calculate_statistics(list_values):
n5 = np.nanpercentile(list_values, 5)
n25 = np.nanpercentile(list_values, 25)
n75 = np.nanpercentile(list_values, 75)
n95 = np.nanpercentile(list_values, 95)
median = np.nanpercentile(list_values, 50)
mean = np.nanmean(list_values)
std = np.nanstd(list_values)
var = np.nanvar(list_values)
rms = np.nanmean(np.sqrt(list_values**2))
return [n5, n25, n75, n95, median, mean, std, var, rms]
def calculate_crossings(list_values):
zero_crossing_indices = np.nonzero(np.diff(np.array(list_values) > 0))[0]
no_zero_crossings = len(zero_crossing_indices)
mean_crossing_indices = np.nonzero(np.diff(np.array(list_values) > np.nanmean(list_values)))[0]
no_mean_crossings = len(mean_crossing_indices)
return [no_zero_crossings, no_mean_crossings]
def get_features(list_values):
entropy = calculate_entropy(list_values)
crossings = calculate_crossings(list_values)
statistics = calculate_statistics(list_values)
return [entropy] + crossings + statistics
def get_uci_har_features(dataset, labels, waveletname):
uci_har_features = []
for signal_no in range(0, len(dataset)):
features = []
for signal_comp in range(0,dataset.shape[2]):
signal = dataset[signal_no, :, signal_comp]
list_coeff = pywt.wavedec(signal, waveletname)
for coeff in list_coeff:
features += get_features(coeff)
uci_har_features.append(features)
X = np.array(uci_har_features)
Y = np.array(labels)
return X, Y
```
# 1. Loading the UCI HAR dataset
download dataset from https://archive.ics.uci.edu/ml/datasets/Human+Activity+Recognition+Using+Smartphones
```
activities_description = {
1: 'walking',
2: 'walking upstairs',
3: 'walking downstairs',
4: 'sitting',
5: 'standing',
6: 'laying'
}
def read_signals(filename):
with open(filename, 'r') as fp:
data = fp.read().splitlines()
data = map(lambda x: x.rstrip().lstrip().split(), data)
data = [list(map(float, line)) for line in data]
return data
def read_labels(filename):
with open(filename, 'r') as fp:
activities = fp.read().splitlines()
activities = list(map(int, activities))
return activities
def randomize(dataset, labels):
permutation = np.random.permutation(labels.shape[0])
shuffled_dataset = dataset[permutation, :, :]
shuffled_labels = labels[permutation]
return shuffled_dataset, shuffled_labels
INPUT_FOLDER_TRAIN = './data/UCI_HAR/train/InertialSignals/'
INPUT_FOLDER_TEST = './data/UCI_HAR/test/InertialSignals/'
INPUT_FILES_TRAIN = ['body_acc_x_train.txt', 'body_acc_y_train.txt', 'body_acc_z_train.txt',
'body_gyro_x_train.txt', 'body_gyro_y_train.txt', 'body_gyro_z_train.txt',
'total_acc_x_train.txt', 'total_acc_y_train.txt', 'total_acc_z_train.txt']
INPUT_FILES_TEST = ['body_acc_x_test.txt', 'body_acc_y_test.txt', 'body_acc_z_test.txt',
'body_gyro_x_test.txt', 'body_gyro_y_test.txt', 'body_gyro_z_test.txt',
'total_acc_x_test.txt', 'total_acc_y_test.txt', 'total_acc_z_test.txt']
LABELFILE_TRAIN = './data/UCI_HAR/train/y_train.txt'
LABELFILE_TEST = './data/UCI_HAR/test/y_test.txt'
train_signals, test_signals = [], []
for input_file in INPUT_FILES_TRAIN:
signal = read_signals(INPUT_FOLDER_TRAIN + input_file)
train_signals.append(signal)
train_signals = np.transpose(np.array(train_signals), (1, 2, 0))
for input_file in INPUT_FILES_TEST:
signal = read_signals(INPUT_FOLDER_TEST + input_file)
test_signals.append(signal)
test_signals = np.transpose(np.array(test_signals), (1, 2, 0))
train_labels = read_labels(LABELFILE_TRAIN)
test_labels = read_labels(LABELFILE_TEST)
[no_signals_train, no_steps_train, no_components_train] = np.shape(train_signals)
[no_signals_test, no_steps_test, no_components_test] = np.shape(train_signals)
no_labels = len(np.unique(train_labels[:]))
print("The train dataset contains {} signals, each one of length {} and {} components ".format(no_signals_train, no_steps_train, no_components_train))
print("The test dataset contains {} signals, each one of length {} and {} components ".format(no_signals_test, no_steps_test, no_components_test))
print("The train dataset contains {} labels, with the following distribution:\n {}".format(np.shape(train_labels)[0], Counter(train_labels[:])))
print("The test dataset contains {} labels, with the following distribution:\n {}".format(np.shape(test_labels)[0], Counter(test_labels[:])))
uci_har_signals_train, uci_har_labels_train = randomize(train_signals, np.array(train_labels))
uci_har_signals_test, uci_har_labels_test = randomize(test_signals, np.array(test_labels))
```
# 2. Generating features for the UCI-HAR features
```
waveletname = 'rbio3.1'
X_train, Y_train = get_uci_har_features(uci_har_signals_train, uci_har_labels_train, waveletname)
X_test, Y_test = get_uci_har_features(uci_har_signals_test, uci_har_labels_test, waveletname)
```
# 3. Classifying the train and test sets
```
models = batch_classify(X_train, Y_train, X_test, Y_test)
display_dict_models(models)
```
| github_jupyter |
# About this kernel
+ resnext50_32x4d
+ ArcFace
+ Mish() activation
+ Ranger (RAdam + Lookahead) optimizer
+ margin = 0.8
## Imports
```
import sys
sys.path.append('../input/shopee-competition-utils')
sys.path.insert(0,'../input/pytorch-image-models')
import numpy as np
import pandas as pd
import torch
from torch import nn
from torch.nn import Parameter
from torch.nn import functional as F
from torch.utils.data import Dataset, DataLoader
import albumentations
from albumentations.pytorch.transforms import ToTensorV2
from custom_scheduler import ShopeeScheduler
from custom_activation import replace_activations, Mish
from custom_optimizer import Ranger
import math
import cv2
import timm
import os
import random
import gc
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import GroupKFold
from sklearn.neighbors import NearestNeighbors
from tqdm.notebook import tqdm
```
## Config
```
class CFG:
DATA_DIR = '../input/shopee-product-matching/train_images'
TRAIN_CSV = '../input/shopee-product-matching/train.csv'
# data augmentation
IMG_SIZE = 512
MEAN = [0.485, 0.456, 0.406]
STD = [0.229, 0.224, 0.225]
SEED = 2021
# data split
N_SPLITS = 5
TEST_FOLD = 0
VALID_FOLD = 1
EPOCHS = 8
BATCH_SIZE = 8
NUM_WORKERS = 4
DEVICE = 'cuda:3'
CLASSES = 6609
SCALE = 30
MARGIN = 0.8
MODEL_NAME = 'resnext50_32x4d'
MODEL_PATH = f'{MODEL_NAME}_arc_face_epoch_{EPOCHS}_bs_{BATCH_SIZE}_margin_{MARGIN}.pt'
FC_DIM = 512
SCHEDULER_PARAMS = {
"lr_start": 1e-5,
"lr_max": 1e-5 * 32,
"lr_min": 1e-6,
"lr_ramp_ep": 5,
"lr_sus_ep": 0,
"lr_decay": 0.8,
}
```
## Augmentations
```
def get_train_transforms():
return albumentations.Compose(
[
albumentations.Resize(CFG.IMG_SIZE,CFG.IMG_SIZE,always_apply=True),
albumentations.HorizontalFlip(p=0.5),
albumentations.VerticalFlip(p=0.5),
albumentations.Rotate(limit=120, p=0.8),
albumentations.RandomBrightness(limit=(0.09, 0.6), p=0.5),
albumentations.Normalize(mean=CFG.MEAN, std=CFG.STD),
ToTensorV2(p=1.0),
]
)
def get_valid_transforms():
return albumentations.Compose(
[
albumentations.Resize(CFG.IMG_SIZE,CFG.IMG_SIZE,always_apply=True),
albumentations.Normalize(mean=CFG.MEAN, std=CFG.STD),
ToTensorV2(p=1.0)
]
)
def get_test_transforms():
return albumentations.Compose(
[
albumentations.Resize(CFG.IMG_SIZE,CFG.IMG_SIZE,always_apply=True),
albumentations.Normalize(mean=CFG.MEAN, std=CFG.STD),
ToTensorV2(p=1.0)
]
)
```
## Reproducibility
```
def seed_everything(seed):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = True # set True to be faster
seed_everything(CFG.SEED)
```
## Dataset
```
class ShopeeDataset(torch.utils.data.Dataset):
"""for training
"""
def __init__(self,df, transform = None):
self.df = df
self.root_dir = CFG.DATA_DIR
self.transform = transform
def __len__(self):
return len(self.df)
def __getitem__(self,idx):
row = self.df.iloc[idx]
img_path = os.path.join(self.root_dir,row.image)
image = cv2.imread(img_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
label = row.label_group
if self.transform:
augmented = self.transform(image=image)
image = augmented['image']
return {
'image' : image,
'label' : torch.tensor(label).long()
}
class ShopeeImageDataset(torch.utils.data.Dataset):
"""for validating and test
"""
def __init__(self,df, transform = None):
self.df = df
self.root_dir = CFG.DATA_DIR
self.transform = transform
def __len__(self):
return len(self.df)
def __getitem__(self,idx):
row = self.df.iloc[idx]
img_path = os.path.join(self.root_dir,row.image)
image = cv2.imread(img_path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
label = row.label_group
if self.transform:
augmented = self.transform(image=image)
image = augmented['image']
return image,torch.tensor(1)
```
## ArcMarginProduct
```
class ArcMarginProduct(nn.Module):
r"""Implement of large margin arc distance: :
Args:
in_features: size of each input sample
out_features: size of each output sample
s: norm of input feature
m: margin
cos(theta + m)
"""
def __init__(self, in_features, out_features, s=30.0, m=0.50, easy_margin=False, ls_eps=0.0):
super(ArcMarginProduct, self).__init__()
self.in_features = in_features
self.out_features = out_features
self.s = s
self.m = m
self.ls_eps = ls_eps # label smoothing
self.weight = Parameter(torch.FloatTensor(out_features, in_features))
nn.init.xavier_uniform_(self.weight)
self.easy_margin = easy_margin
self.cos_m = math.cos(m)
self.sin_m = math.sin(m)
self.th = math.cos(math.pi - m)
self.mm = math.sin(math.pi - m) * m
def forward(self, input, label):
# --------------------------- cos(theta) & phi(theta) ---------------------------
cosine = F.linear(F.normalize(input), F.normalize(self.weight))
sine = torch.sqrt(1.0 - torch.pow(cosine, 2))
phi = cosine * self.cos_m - sine * self.sin_m
if self.easy_margin:
phi = torch.where(cosine > 0, phi, cosine)
else:
phi = torch.where(cosine > self.th, phi, cosine - self.mm)
# --------------------------- convert label to one-hot ---------------------------
# one_hot = torch.zeros(cosine.size(), requires_grad=True, device='cuda')
one_hot = torch.zeros(cosine.size(), device=CFG.DEVICE)
one_hot.scatter_(1, label.view(-1, 1).long(), 1)
if self.ls_eps > 0:
one_hot = (1 - self.ls_eps) * one_hot + self.ls_eps / self.out_features
# -------------torch.where(out_i = {x_i if condition_i else y_i) -------------
output = (one_hot * phi) + ((1.0 - one_hot) * cosine)
output *= self.s
return output, nn.CrossEntropyLoss()(output,label)
class ShopeeModel(nn.Module):
def __init__(
self,
n_classes = CFG.CLASSES,
model_name = CFG.MODEL_NAME,
fc_dim = CFG.FC_DIM,
margin = CFG.MARGIN,
scale = CFG.SCALE,
use_fc = True,
pretrained = True):
super(ShopeeModel,self).__init__()
print('Building Model Backbone for {} model'.format(model_name))
self.backbone = timm.create_model(model_name, pretrained=pretrained)
if 'efficientnet' in model_name:
final_in_features = self.backbone.classifier.in_features
self.backbone.classifier = nn.Identity()
self.backbone.global_pool = nn.Identity()
elif 'resnet' in model_name:
final_in_features = self.backbone.fc.in_features
self.backbone.fc = nn.Identity()
self.backbone.global_pool = nn.Identity()
elif 'resnext' in model_name:
final_in_features = self.backbone.fc.in_features
self.backbone.fc = nn.Identity()
self.backbone.global_pool = nn.Identity()
elif 'nfnet' in model_name:
final_in_features = self.backbone.head.fc.in_features
self.backbone.head.fc = nn.Identity()
self.backbone.head.global_pool = nn.Identity()
self.pooling = nn.AdaptiveAvgPool2d(1)
self.use_fc = use_fc
if use_fc:
self.dropout = nn.Dropout(p=0.0)
self.fc = nn.Linear(final_in_features, fc_dim)
self.bn = nn.BatchNorm1d(fc_dim)
self._init_params()
final_in_features = fc_dim
self.final = ArcMarginProduct(final_in_features,
n_classes,
s=scale,
m=margin)
def _init_params(self):
nn.init.xavier_normal_(self.fc.weight)
nn.init.constant_(self.fc.bias, 0)
nn.init.constant_(self.bn.weight, 1)
nn.init.constant_(self.bn.bias, 0)
def forward(self, image, label):
feature = self.extract_feat(image)
logits = self.final(feature,label)
return logits
def extract_feat(self, x):
batch_size = x.shape[0]
x = self.backbone(x)
x = self.pooling(x).view(batch_size, -1)
if self.use_fc:
x = self.dropout(x)
x = self.fc(x)
x = self.bn(x)
return x
```
## Engine
```
def train_fn(model, data_loader, optimizer, scheduler, i):
model.train()
fin_loss = 0.0
tk = tqdm(data_loader, desc = "Epoch" + " [TRAIN] " + str(i+1))
for t,data in enumerate(tk):
for k,v in data.items():
data[k] = v.to(CFG.DEVICE)
optimizer.zero_grad()
_, loss = model(**data)
loss.backward()
optimizer.step()
fin_loss += loss.item()
tk.set_postfix({'loss' : '%.6f' %float(fin_loss/(t+1)), 'LR' : optimizer.param_groups[0]['lr']})
scheduler.step()
return fin_loss / len(data_loader)
def eval_fn(model, data_loader, i):
model.eval()
fin_loss = 0.0
tk = tqdm(data_loader, desc = "Epoch" + " [VALID] " + str(i+1))
with torch.no_grad():
for t,data in enumerate(tk):
for k,v in data.items():
data[k] = v.to(CFG.DEVICE)
_, loss = model(**data)
fin_loss += loss.item()
tk.set_postfix({'loss' : '%.6f' %float(fin_loss/(t+1))})
return fin_loss / len(data_loader)
def read_dataset():
df = pd.read_csv(CFG.TRAIN_CSV)
df['matches'] = df.label_group.map(df.groupby('label_group').posting_id.agg('unique').to_dict())
df['matches'] = df['matches'].apply(lambda x: ' '.join(x))
gkf = GroupKFold(n_splits=CFG.N_SPLITS)
df['fold'] = -1
for i, (train_idx, valid_idx) in enumerate(gkf.split(X=df, groups=df['label_group'])):
df.loc[valid_idx, 'fold'] = i
labelencoder= LabelEncoder()
df['label_group'] = labelencoder.fit_transform(df['label_group'])
train_df = df[df['fold']!=CFG.TEST_FOLD].reset_index(drop=True)
train_df = train_df[train_df['fold']!=CFG.VALID_FOLD].reset_index(drop=True)
valid_df = df[df['fold']==CFG.VALID_FOLD].reset_index(drop=True)
test_df = df[df['fold']==CFG.TEST_FOLD].reset_index(drop=True)
train_df['label_group'] = labelencoder.fit_transform(train_df['label_group'])
return train_df, valid_df, test_df
def precision_score(y_true, y_pred):
y_true = y_true.apply(lambda x: set(x.split()))
y_pred = y_pred.apply(lambda x: set(x.split()))
intersection = np.array([len(x[0] & x[1]) for x in zip(y_true, y_pred)])
len_y_pred = y_pred.apply(lambda x: len(x)).values
precision = intersection / len_y_pred
return precision
def recall_score(y_true, y_pred):
y_true = y_true.apply(lambda x: set(x.split()))
y_pred = y_pred.apply(lambda x: set(x.split()))
intersection = np.array([len(x[0] & x[1]) for x in zip(y_true, y_pred)])
len_y_true = y_true.apply(lambda x: len(x)).values
recall = intersection / len_y_true
return recall
def f1_score(y_true, y_pred):
y_true = y_true.apply(lambda x: set(x.split()))
y_pred = y_pred.apply(lambda x: set(x.split()))
intersection = np.array([len(x[0] & x[1]) for x in zip(y_true, y_pred)])
len_y_pred = y_pred.apply(lambda x: len(x)).values
len_y_true = y_true.apply(lambda x: len(x)).values
f1 = 2 * intersection / (len_y_pred + len_y_true)
return f1
def get_valid_embeddings(df, model):
model.eval()
image_dataset = ShopeeImageDataset(df,transform=get_valid_transforms())
image_loader = torch.utils.data.DataLoader(
image_dataset,
batch_size=CFG.BATCH_SIZE,
pin_memory=True,
num_workers = CFG.NUM_WORKERS,
drop_last=False
)
embeds = []
with torch.no_grad():
for img,label in tqdm(image_loader):
img = img.to(CFG.DEVICE)
label = label.to(CFG.DEVICE)
feat,_ = model(img,label)
image_embeddings = feat.detach().cpu().numpy()
embeds.append(image_embeddings)
del model
image_embeddings = np.concatenate(embeds)
print(f'Our image embeddings shape is {image_embeddings.shape}')
del embeds
gc.collect()
return image_embeddings
def get_valid_neighbors(df, embeddings, KNN = 50, threshold = 0.36):
model = NearestNeighbors(n_neighbors = KNN, metric = 'cosine')
model.fit(embeddings)
distances, indices = model.kneighbors(embeddings)
predictions = []
for k in range(embeddings.shape[0]):
idx = np.where(distances[k,] < threshold)[0]
ids = indices[k,idx]
posting_ids = ' '.join(df['posting_id'].iloc[ids].values)
predictions.append(posting_ids)
df['pred_matches'] = predictions
df['f1'] = f1_score(df['matches'], df['pred_matches'])
df['recall'] = recall_score(df['matches'], df['pred_matches'])
df['precision'] = precision_score(df['matches'], df['pred_matches'])
del model, distances, indices
gc.collect()
return df, predictions
```
# Training
```
def run_training():
train_df, valid_df, test_df = read_dataset()
train_dataset = ShopeeDataset(train_df, transform = get_train_transforms())
train_dataloader = torch.utils.data.DataLoader(
train_dataset,
batch_size = CFG.BATCH_SIZE,
pin_memory = True,
num_workers = CFG.NUM_WORKERS,
shuffle = True,
drop_last = True
)
print(train_df['label_group'].nunique())
model = ShopeeModel()
model = replace_activations(model, torch.nn.SiLU, Mish())
model.to(CFG.DEVICE)
optimizer = Ranger(model.parameters(), lr = CFG.SCHEDULER_PARAMS['lr_start'])
#optimizer = torch.optim.Adam(model.parameters(), lr = config.SCHEDULER_PARAMS['lr_start'])
scheduler = ShopeeScheduler(optimizer,**CFG.SCHEDULER_PARAMS)
best_valid_f1 = 0.
for i in range(CFG.EPOCHS):
avg_loss_train = train_fn(model, train_dataloader, optimizer, scheduler, i)
valid_embeddings = get_valid_embeddings(valid_df, model)
valid_df, valid_predictions = get_valid_neighbors(valid_df, valid_embeddings)
valid_f1 = valid_df.f1.mean()
valid_recall = valid_df.recall.mean()
valid_precision = valid_df.precision.mean()
print(f'Valid f1 score = {valid_f1}, recall = {valid_recall}, precision = {valid_precision}')
if valid_f1 > best_valid_f1:
best_valid_f1 = valid_f1
print('Valid f1 score improved, model saved')
torch.save(model.state_dict(),CFG.MODEL_PATH)
run_training()
def get_test_embeddings(test_df):
model = ShopeeModel()
model.eval()
model = replace_activations(model, torch.nn.SiLU, Mish())
model.load_state_dict(torch.load(CFG.MODEL_PATH))
model = model.to(CFG.DEVICE)
image_dataset = ShopeeImageDataset(test_df,transform=get_test_transforms())
image_loader = torch.utils.data.DataLoader(
image_dataset,
batch_size=CFG.BATCH_SIZE,
pin_memory=True,
num_workers = CFG.NUM_WORKERS,
drop_last=False
)
embeds = []
with torch.no_grad():
for img,label in tqdm(image_loader):
img = img.cuda()
label = label.cuda()
feat,_ = model(img,label)
image_embeddings = feat.detach().cpu().numpy()
embeds.append(image_embeddings)
del model
image_embeddings = np.concatenate(embeds)
print(f'Our image embeddings shape is {image_embeddings.shape}')
del embeds
gc.collect()
return image_embeddings
```
## Best threshold Search
```
train_df, valid_df, test_df = read_dataset()
print("Searching best threshold...")
search_space = np.arange(10, 50, 1)
model = ShopeeModel()
model.eval()
model = replace_activations(model, torch.nn.SiLU, Mish())
model.load_state_dict(torch.load(CFG.MODEL_PATH))
model = model.to(CFG.DEVICE)
valid_embeddings = get_valid_embeddings(valid_df, model)
best_f1_valid = 0.
best_threshold = 0.
for i in search_space:
threshold = i / 100
valid_df, valid_predictions = get_valid_neighbors(valid_df, valid_embeddings, threshold=threshold)
valid_f1 = valid_df.f1.mean()
valid_recall = valid_df.recall.mean()
valid_precision = valid_df.precision.mean()
print(f"threshold = {threshold} -> f1 score = {valid_f1}, recall = {valid_recall}, precision = {valid_precision}")
if (valid_f1 > best_f1_valid):
best_f1_valid = valid_f1
best_threshold = threshold
print("Best threshold =", best_threshold)
print("Best f1 score =", best_f1_valid)
BEST_THRESHOLD = best_threshold
print("Searching best knn...")
search_space = np.arange(40, 80, 2)
best_f1_valid = 0.
best_knn = 0
for knn in search_space:
valid_df, valid_predictions = get_valid_neighbors(valid_df, valid_embeddings, KNN=knn, threshold=BEST_THRESHOLD)
valid_f1 = valid_df.f1.mean()
valid_recall = valid_df.recall.mean()
valid_precision = valid_df.precision.mean()
print(f"knn = {knn} -> f1 score = {valid_f1}, recall = {valid_recall}, precision = {valid_precision}")
if (valid_f1 > best_f1_valid):
best_f1_valid = valid_f1
best_knn = knn
print("Best knn =", best_knn)
print("Best f1 score =", best_f1_valid)
BEST_KNN = best_knn
test_embeddings = get_valid_embeddings(test_df,model)
test_df, test_predictions = get_valid_neighbors(test_df, test_embeddings, KNN = BEST_KNN, threshold = BEST_THRESHOLD)
test_f1 = test_df.f1.mean()
test_recall = test_df.recall.mean()
test_precision = test_df.precision.mean()
print(f'Test f1 score = {test_f1}, recall = {test_recall}, precision = {test_precision}')
```
| github_jupyter |
```
import numpy as np
from scipy import signal
from scipy.linalg import eig
from matplotlib import pyplot as plt
# problem 1
A = [[-0.5,-0.25],[1.0,0.0]]
print(np.linalg.eig(A)[0])
B = [[0.75],[0.0]]
C = [0.0,1.0]
D = [0.0]
system = signal.StateSpace(A,B,C,D)
t = np.linspace(0,60,200)
x = np.zeros(len(t))
x[10:40] = 1.0 # first step input
x[100:130] = 1.0 # first step input
tr,y,ur = signal.lsim(system,x,t)
plt.clf()
plt.plot(x, color='red')
plt.plot(y, color='black')
plt.show()
result = np.stack((t, x, y), axis=1)
np.savetxt("test1.csv", result, delimiter=",")
# problem 2
K1 = 1
K2 = 1.2
A = [[-1., -K1, 0., 0.],
[ 0., -5, -5*K2, -5*K2],
[-0.4*K1, -0.4*K1, -0.5, 0.0],
[0.0, 0.0, -4*K2, -2-4*K2]]
print(np.linalg.eig(A)[0])
B = [[K1, 0.0],
[0.0, 5*K2],
[0.4*K1, 0.0],
[0.0, 4*K2]]
C = [[1.,1.,0.,0.],
[0.0,0.0,1.0,1.0]]
D = [[0.0,0.0],[0.0,0.0]]
system = signal.StateSpace(A,B,C,D)
t = np.linspace(0,60,100)
x = np.zeros((len(t),2))
x[10:40,0] = 1.0 # first step input
x[30:70,1] = 1.0 # first step input
tr,y,ur = signal.lsim(system,x,t)
plt.clf()
plt.figure(figsize=(10,8))
plt.plot(x[:,0], c='lightgreen', label='x0')
plt.plot(x[:,1], c='darkgreen', label='x1')
plt.plot(y[:,0], c='darkgrey', label='y0')
plt.plot(y[:,1], c='black', label='y1')
plt.legend()
plt.show()
result = np.column_stack((t.reshape(-1,1), x, y))
np.savetxt("test2.csv", result, delimiter=",")
# problem 3
A = [[-0.25,-0.25],[1.0,0.0]]
print(np.linalg.eig(A)[0])
B = [[0.75],[0.0]]
C = [0.0,1.0]
D = [0.0]
system = signal.StateSpace(A,B,C,D)
t = np.linspace(0,150,1200)
x = np.zeros(len(t))
x[0:30] = 0.5 # first step input
x[90:150] = 1.0 # second step input
x[200:260] = .50
x[300:370] = -1.0
x[400:740] = 1.0
tr,y,u = signal.lsim(system,x,t)
plt.clf()
plt.plot(x, color='green')
plt.plot(y, color='black')
plt.show()
result = np.stack((t, x, y), axis=1)
np.savetxt("test3.csv", result, delimiter=",")
# problem 4
A = [[-0.2,-0.25],[1.0,0.0]]
print(np.linalg.eig(A)[0])
B = [[0.75],[0.0]]
C = [0.0,.1]
D = [0.0]
system = signal.StateSpace(A,B,C,D)
t = np.linspace(0,50,200)
x = np.zeros(len(t))
x[0:30] = 1.0 # first step input
tr,y,u = signal.lsim(system,x,t)
plt.clf()
plt.plot(x, color='green')
plt.plot(y, color='black')
plt.show()
result = np.stack((t, x, y), axis=1)
np.savetxt("test4.csv", result, delimiter=",")
# problem 5
A = [[-0.9,0.0],[0.0,-0.5]]
print(np.linalg.eig(A)[0])
B = [[1.],[1.]]
C = [1.0,1.0]
D = [0.0]
system = signal.StateSpace(A,B,C,D)
t = np.linspace(0,50,200)
x = np.random.random(len(t))
tr,y,u = signal.lsim(system, x,t)
plt.clf()
plt.plot(x, color='green')
plt.plot(y, color='black')
plt.show()
result = np.stack((t, x, y), axis=1)
np.savetxt("test5.csv", result, delimiter=",")
```
| github_jupyter |
# Getting Started with ctapipe
This hands-on was presented at the Paris CTA Consoritum meeting (K. Kosack)
## Part 1: load and loop over data
```
from ctapipe.io import event_source
from ctapipe import utils
from matplotlib import pyplot as plt
%matplotlib inline
path = utils.get_dataset_path("gamma_test_large.simtel.gz")
for event in event_source(path, max_events=4):
print(event.count, event.r0.event_id, event.mc.energy)
event
event.r0
for event in event_source(path, max_events=4):
print(event.count, event.r0.tels_with_data)
event.r0.tel[2]
r0tel = event.r0.tel[2]
r0tel.waveform
r0tel.waveform.shape
```
note that this is ($N_{channels}$, $N_{pixels}$, $N_{samples}$)
```
plt.pcolormesh(r0tel.waveform[0])
plt.plot(r0tel.waveform[0,10])
from ipywidgets import interact
@interact
def view_waveform(chan=0, pix_id=200):
plt.plot(r0tel.waveform[chan, pix_id])
```
try making this compare 2 waveforms
## Part 2: Explore the instrument description
This is all well and good, but we don't really know what camera or telescope this is... how do we get instrumental description info?
Currently this is returned *inside* the event (it will soon change to be separate in next version or so)
```
subarray = event.inst.subarray # soon EventSource will give you event, subarray separate
subarray
subarray.peek()
subarray.to_table()
subarray.tel[2]
subarray.tel[2].camera
subarray.tel[2].optics
tel = subarray.tel[2]
tel.camera
tel.optics
tel.camera.pix_x
tel.camera.to_table()
tel.optics.mirror_area
from ctapipe.visualization import CameraDisplay
disp = CameraDisplay(tel.camera)
disp = CameraDisplay(tel.camera)
disp.image = r0tel.waveform[0,:,10] # display channel 0, sample 0 (try others like 10)
```
** aside: ** show demo using a CameraDisplay in interactive mode in ipython rather than notebook
## Part 3: Apply some calibration and trace integration
```
from ctapipe.calib import CameraCalibrator
calib = CameraCalibrator(subarray=subarray)
for event in event_source(path, max_events=4):
calib(event) # fills in r1, dl0, and dl1
print(event.dl1.tel.keys())
event.dl1.tel[2]
dl1tel = event.dl1.tel[2]
dl1tel.image.shape # note this will be gain-selected in next version, so will be just 1D array of 1855
dl1tel.pulse_time
CameraDisplay(tel.camera, image=dl1tel.image)
CameraDisplay(tel.camera, image=dl1tel.pulse_time)
```
Now for Hillas Parameters
```
from ctapipe.image import hillas_parameters, tailcuts_clean
image = dl1tel.image
mask = tailcuts_clean(tel.camera, image, picture_thresh=10, boundary_thresh=5)
mask
CameraDisplay(tel.camera, image=mask)
cleaned = image.copy()
cleaned[~mask] = 0
disp = CameraDisplay(tel.camera, image=cleaned)
disp.cmap = plt.cm.coolwarm
disp.add_colorbar()
plt.xlim(-1.0,0)
plt.ylim(0,1.0)
params = hillas_parameters(tel.camera, cleaned)
print(params)
disp = CameraDisplay(tel.camera, image=cleaned)
disp.cmap = plt.cm.coolwarm
disp.add_colorbar()
plt.xlim(-1.0,0)
plt.ylim(0,1.0)
disp.overlay_moments(params, color='white', lw=2)
```
## Part 4: Let's put it all together:
- loop over events, selecting only telescopes of the same type (e.g. LST:LSTCam)
- for each event, apply calibration/trace integration
- calculate Hillas parameters
- write out all hillas paremeters to a file that can be loaded with Pandas
first let's select only those telescopes with LST:LSTCam
```
subarray.telescope_types
subarray.get_tel_ids_for_type("LST_LST_LSTCam")
```
Now let's write out program
```
data = utils.get_dataset_path("gamma_test_large.simtel.gz")
source = event_source(data, allowed_tels=[1,2,3,4], max_events=10) # remove the max_events limit to get more stats
for event in source:
calib(event)
for tel_id, tel_data in event.dl1.tel.items():
tel = event.inst.subarray.tel[tel_id]
mask = tailcuts_clean(tel.camera, tel_data.image)
params = hillas_parameters(tel.camera[mask], tel_data.image[mask])
from ctapipe.io import HDF5TableWriter
with HDF5TableWriter(filename='hillas.h5', group_name='dl1', overwrite=True) as writer:
for event in event_source(data, allowed_tels=[1,2,3,4], max_events=10):
calib(event)
for tel_id, tel_data in event.dl1.tel.items():
tel = event.inst.subarray.tel[tel_id]
mask = tailcuts_clean(tel.camera, tel_data.image)
params = hillas_parameters(tel.camera[mask], tel_data.image[mask])
writer.write("hillas", params)
```
### We can now load in the file we created and plot it
```
!ls *.h5
import pandas as pd
hillas = pd.read_hdf("hillas.h5", key='/dl1/hillas')
hillas
_ = hillas.hist(figsize=(8,8))
```
If you do this yourself, loop over more events to get better statistics
| github_jupyter |
# DTI-derived metrics (FA, MD, AD, RD) within FS-segmented ROIs
BMED360-2021 `05-dmr-dti-feature-extraction-roi-wise.ipynb`
(assuming the `02-dmri-find-affine-fs-brainmask2dwi.ipynb` and `03-dmri-reconstruction-dti.ipynb` and `04-dmri-do-affine-reg-anat2dwi.ipynb` notebooks have been executed)
<a href="https://colab.research.google.com/github/computational-medicine/BMED360-2021/blob/main/Lab3-diffusion-MRI/05-dmr-dti-feature-extraction-roi-wise.ipynb">
<img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/>
</a>
### Learning objectives
#### Check [FreeSurferColorLUT.txt](https://surfer.nmr.mgh.harvard.edu/fswiki/FsTutorial/AnatomicalROI/FreeSurferColorLUT) for the ROI-label for the different regions (left and right hemisphere)
Relevant ROIs:
```
# Below is the color table for a lobar white matter parcelation
# obtained from running
# mri_annotation2label --subject subject --hemi lh --lobesStrict lobes
# mri_annotation2label --subject subject --hemi rh --lobesStrict lobes
# mri_aparc2aseg --s subject --labelwm --hypo-as-wm --rip-unknown \
# --volmask --o wmparc.lobes.mgz --ctxseg aparc+aseg.mgz \
# --annot lobes --base-offset 200 [--base-offset must be last arg]
3201 wm-lh-frontal-lobe 235 35 95 0
3203 wm-lh-cingulate-lobe 35 75 35 0
3204 wm-lh-occiptal-lobe 135 155 195 0
3205 wm-lh-temporal-lobe 115 35 35 0
3206 wm-lh-parietal-lobe 35 195 35 0
3207 wm-lh-insula-lobe 20 220 160 0
4201 wm-rh-frontal-lobe 235 35 95 0
4203 wm-rh-cingulate-lobe 35 75 35 0
4204 wm-rh-occiptal-lobe 135 155 195 0
4205 wm-rh-temporal-lobe 115 35 35 0
4206 wm-rh-parietal-lobe 35 195 35 0
4207 wm-rh-insula-lobe 20 220 160 0
```
### For using Colab
**--> (some of) the following libraries must be `pip installed` (i.e. uncommet these among the following pip commands):**
```
#!pip install gdown
#!pip install nilearn
#!pip install dipy
```
**Download a data file from Google Drive using gdown** (https://github.com/wkentaro/gdown)
```
import gdown
import shutil
import sys
import os
from os.path import expanduser, join, basename, split
import glob
import shutil
import platform
```
Check your platform for running this notebook
```
if platform.system() == 'Darwin':
print(f'OK, you are running on MacOS ({platform.version()})')
if platform.system() == 'Linux':
print(f'OK, you are running on Linux ({platform.version()})')
if platform.system() == 'Windows':
print(f'OK, but consider to install WSL for Windows10 since you are running on {platform.system()}')
print('Check https://docs.microsoft.com/en-us/windows/wsl/install-win10')
cwd = os.getcwd()
working_dir = join(cwd, 'data')
bids_dir = '%s/bids_bg_bmed360' % (working_dir)
dmri_res = '%s/dmri_results' % (working_dir)
# Download zip-file if ./data does not exist (as when running in Colab)
if os.path.isdir(working_dir) == False:
## Download data.zip for Google Drive
# https://drive.google.com/file/d/1pX6Mx_9P8fojDXmbTz-th5FKFYZDVFuO/view?usp=sharing
file_id = '1pX6Mx_9P8fojDXmbTz-th5FKFYZDVFuO'
url = 'https://drive.google.com/uc?id=%s' % file_id
output = './data.zip'
gdown.download(url, output, quiet=False)
## Unzip the assets file into `./data`
shutil.unpack_archive(output, '.')
## Delete the `data.zip` file
os.remove(output)
else:
print(f'./data exists already!')
# Download zip-file if ./data/dmri_results does not exist (as when running in Colab)
if os.path.isdir(dmri_res) == False:
## Download dmri_results.zip for Google Drive
# https://drive.google.com/file/d/1pX6Mx_9P8fojDXmbTz-th5FKFYZDVFuO/view?usp=sharing
file_id = '1wu5pzAcE2hyZymq-IzuzKYGK_lMYbnJy'
url = 'https://drive.google.com/uc?id=%s' % file_id
output = 'dmri_results.zip'
gdown.download(url, output, quiet=False)
## Unzip the assets file into `./data/dmri_results`
shutil.unpack_archive(output, './data/')
## Delete the `dmri_results.zip` file
os.remove(output)
else:
print(f'./data/dmri_results exists already!')
```
## Import libraries
```
import os
import pathlib
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import nibabel as nib
from os.path import expanduser, join, basename, split
import time
from dipy.io.image import save_nifti
def get_dti_features_from_roi(inpdir, sub, ses, roi_name, roi_num, outdir):
r"""
Extraction of DTI-derived featurevalues, i.e. FA, MD, AD, RD from a given antomical region, named 'roi_name'
and numbered by Freesurfer as 'roi_num' with respect to the coregistered wmparc segmented image for
given subject ('sub') and session ('ses' = 1)
Parameters
----------
inpdir : input directory e.g. inpdir = 'data/dmri_results' where the moving files
from Freesurfer segmentation mapped to T1w native space are stored.
sub : subject id, e.g. 102 for sub-102
ses : session number, e.g. 1 for ses-1
roi_name : name of a (white matter) region in wmparc, e.g. 'wm-lh-insula'
roi_num : number of that region according to the Freesurfer LUT, e.g. 3035
outdir : output directory e.g. outdir = inpdir, where resulting ... are stored,
Returns
-------
meas_name : list of DTI-derived features, for now ['fa', 'md', 'ad', 'rd']
meas_num : number of voxels in the named 3D ROI region (same accross all features)
meas_val : the feature values of each feature i in ['fa', 'md', 'ad', 'rd']
Notes
-----
"""
# Preparations
# Load WMPARC:
wmparc_fn = join(inpdir, 'sub_%d_tp%d_wmparc_in_native_space_aff_to_dwi_S0_mean.nii.gz' % (sub, ses))
wmparc_img= nib.load(wmparc_fn)
wmparc_data = wmparc_img.get_fdata()
wmparc_affine = wmparc_img.affine
#a = wmparc_data.flatten()
#unique, counts = np.unique(a, return_counts=True)
#dict(zip(unique, counts))
roi = np.where(wmparc_data == roi_num)
roi_flatten = np.where(wmparc_data.flatten() == roi_num)
# Metrics from DTI in native DWI space
dti_metrics = ['fa', 'md', 'ad', 'rd']
meas_name = []
meas_num = []
meas_val = []
for i, meas in enumerate(dti_metrics):
fname = join(inpdir, 'sub_%d_tp%d_dwi_brainmask_tensor_%s.nii.gz' % (sub, ses, meas))
meas_img= nib.load(fname)
meas_data = meas_img.get_fdata()
meas_affine = meas_img.affine
meas_data_flatten = meas_data.flatten()
meas_name.append(meas)
meas_num.append(len(roi_flatten[0]))
meas_val.append(meas_data_flatten[roi_flatten])
#plt.hist(meas_val[i],bins=50)
#plt.title('sub-%d | %s | %s | n=%d' % (sub, roi_name, meas_name[i], len(roi_flatten[0])))
#plt.show()
#fig = plt.figure(figsize = [14, 12])
#for i, meas in enumerate(dti_metrics):
# plt.subplot(2,2,i+1)
# plt.hist(meas_val[i],bins=50, color='black')
# plt.title('sub-%d | %s | %s | n=%d | median=%.5f' %
# (sub, roi_name, meas_name[i].upper(), len(roi_flatten[0]), np.median(meas_val[i])))
#plt.show()
#fig.savefig('%s/tmp/FIGURES/sub_%d_wmparc_%s_dti_features.png' % (home, sub, roi_name),
# transparent=False, dpi=300, bbox_inches="tight")
return dti_metrics, meas_name, meas_num, meas_val
# get_dti_features_from_roi?
inp_dir = dmri_res
outp_dir = inp_dir
```
### Test the function
```
roi_name = 'wm-lh-insula'
roi_num = 3035
sub = 102
ses = 1
%%time
dti_metrics, meas_name, meas_num, meas_val = \
get_dti_features_from_roi(inp_dir, sub, ses, roi_name, roi_num, outp_dir)
fig = plt.figure(figsize = [14, 12])
for i, meas in enumerate(dti_metrics):
plt.subplot(2,2,i+1)
plt.hist(meas_val[i],bins=50, color='black')
plt.title('sub-%d | %s | %s | n=%d | median=%.5f' %
(sub, roi_name, meas_name[i].upper(), len(meas_val[i]), np.median(meas_val[i])))
plt.show()
#fig.savefig('./assets/sub_%d_wmparc_%s_dti_features.png' % (sub, roi_name),
# transparent=False, dpi=300, bbox_inches="tight")
```
## Run for all subjects
```
# Relevant WM regions:
rois = [3035, 4035, 10, 49, 13, 52]
wm_dict = {3035: 'wm-lh-insula',
4035: 'wm-rh-insula',
10: 'Left-Thalamus-Proper',
49: 'Right-Thalamus-Proper',
13: 'Left-Pallidum',
52: 'Right-Pallidum'}
wm_dict[rois[0]], wm_dict[rois[-1]]
```
```python
# Mechanism to save dictionaries and lists as a binary .npy file
import numpy as np
# Save
dictionary = {'hello':'world'}
np.save('%s/tmp/my_file.npy' % (home), dictionary)
# Load
read_dictionary = np.load('%s/tmp/my_file.npy' % (home),allow_pickle='TRUE').item()
print(read_dictionary['hello']) # displays "world"
```
-->
world
```
%%time
ses=1
for j in range(len(rois)):
roi_num = rois[j]
roi_name = wm_dict[rois[j]]
dic = {}
dipy_dic = []
for sub in [102, 103, 111, 123]:
print(f'Computing sub:{sub} ses:{ses} roi:{roi_name}')
dti_metrics, meas_name, meas_num, meas_val = \
get_dti_features_from_roi(inp_dir, sub, ses, roi_name, roi_num, outp_dir)
dic = {'subj': sub, 'roi_num': roi_num, 'roi_name': roi_name, \
'meas_name': meas_name, 'meas_num': meas_num, 'meas_val': meas_val}
dipy_dic.append(dic)
# np.save('%s/%s_%d_dipy_dic.npy' % (outp_dir, roi_name, roi_num), dipy_dic)
np.save('%s/%s_%d_dipy_dic.npy' % (outp_dir, roi_name, roi_num), dipy_dic)
# Load
for j in range(len(rois)):
roi_num = rois[j]
roi_name = wm_dict[rois[j]]
read_dipy_dic = np.load('%s/%s_%d_dipy_dic.npy' % (outp_dir, roi_name, roi_num), allow_pickle='TRUE')
print('ROI #%d: %s %d' % (j, wm_dict[rois[j]], rois[j]))
print('- Subject', read_dipy_dic[0]['subj']) # displays 101
print('-', read_dipy_dic[0]['meas_name'][0]) # displays fa
print('-', read_dipy_dic[0]['meas_num'][0]) # displays number of voxels in region
print('-', read_dipy_dic[0]['meas_val'][0]) # displays FA vector of length number of voxels in region
print('-', read_dipy_dic[0]['meas_val'][0].mean().round(4)) # displays mean value of FA vector in region
print('.....')
print('- Subject', read_dipy_dic[-1]['subj']) # displays 130
print('-', read_dipy_dic[-1]['meas_name'][0]) # displays fa
print('-', read_dipy_dic[-1]['meas_num'][0]) # displays number of voxels in region
print('-', read_dipy_dic[-1]['meas_val'][0]) # displays FA vector of length number of voxels in region
print('-', read_dipy_dic[-1]['meas_val'][0].mean().round(4)) # displays mean value of FA vector in region
```
| github_jupyter |
# Mask R-CNN - Inspect Ballon Trained Model
Code and visualizations to test, debug, and evaluate the Mask R-CNN model.
```
import os
import sys
import random
import math
import re
import time
import numpy as np
import tensorflow as tf
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
# Root directory of the project
ROOT_DIR = os.path.abspath("../../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
from mrcnn import visualize
from mrcnn.visualize import display_images
import mrcnn.model as modellib
from mrcnn.model import log
from samples.balloon import balloon
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Path to Ballon trained weights
# You can download this file from the Releases page
# https://github.com/matterport/Mask_RCNN/releases
BALLON_WEIGHTS_PATH = "/path/to/mask_rcnn_balloon.h5" # TODO: update this path
```
## Configurations
```
config = balloon.BalloonConfig()
BALLOON_DIR = os.path.join(ROOT_DIR, "datasets/balloon")
# Override the training configurations with a few
# changes for inferencing.
class InferenceConfig(config.__class__):
# Run detection on one image at a time
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
```
## Notebook Preferences
```
# Device to load the neural network on.
# Useful if you're training a model on the same
# machine, in which case use CPU and leave the
# GPU for training.
DEVICE = "/cpu:0" # /cpu:0 or /gpu:0
# Inspect the model in training or inference modes
# values: 'inference' or 'training'
# TODO: code for 'training' test mode not ready yet
TEST_MODE = "inference"
def get_ax(rows=1, cols=1, size=16):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Adjust the size attribute to control how big to render images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
```
## Load Validation Dataset
```
# Load validation dataset
dataset = balloon.BalloonDataset()
dataset.load_balloon(BALLOON_DIR, "val")
# Must call before using the dataset
dataset.prepare()
print("Images: {}\nClasses: {}".format(len(dataset.image_ids), dataset.class_names))
```
## Load Model
```
# Create model in inference mode
with tf.device(DEVICE):
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR,
config=config)
# Set path to balloon weights file
# Download file from the Releases page and set its path
# https://github.com/matterport/Mask_RCNN/releases
# weights_path = "/path/to/mask_rcnn_balloon.h5"
# Or, load the last model you trained
weights_path = model.find_last()[1]
# Load weights
print("Loading weights ", weights_path)
model.load_weights(weights_path, by_name=True)
```
## Run Detection
```
image_id = random.choice(dataset.image_ids)
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset, config, image_id, use_mini_mask=False)
info = dataset.image_info[image_id]
print("image ID: {}.{} ({}) {}".format(info["source"], info["id"], image_id,
dataset.image_reference(image_id)))
# Run object detection
results = model.detect([image], verbose=1)
# Display results
ax = get_ax(1)
r = results[0]
visualize.display_instances(image, r['rois'], r['masks'], r['class_ids'],
dataset.class_names, r['scores'], ax=ax,
title="Predictions")
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
```
## Color Splash
This is for illustration. You can call `balloon.py` with the `splash` option to get better images without the black padding.
```
splash = balloon.color_splash(image, r['masks'])
display_images([splash], cols=1)
```
## Step by Step Prediction
## Stage 1: Region Proposal Network
The Region Proposal Network (RPN) runs a lightweight binary classifier on a lot of boxes (anchors) over the image and returns object/no-object scores. Anchors with high *objectness* score (positive anchors) are passed to the stage two to be classified.
Often, even positive anchors don't cover objects fully. So the RPN also regresses a refinement (a delta in location and size) to be applied to the anchors to shift it and resize it a bit to the correct boundaries of the object.
### 1.a RPN Targets
The RPN targets are the training values for the RPN. To generate the targets, we start with a grid of anchors that cover the full image at different scales, and then we compute the IoU of the anchors with ground truth object. Positive anchors are those that have an IoU >= 0.7 with any ground truth object, and negative anchors are those that don't cover any object by more than 0.3 IoU. Anchors in between (i.e. cover an object by IoU >= 0.3 but < 0.7) are considered neutral and excluded from training.
To train the RPN regressor, we also compute the shift and resizing needed to make the anchor cover the ground truth object completely.
```
# Generate RPN trainig targets
# target_rpn_match is 1 for positive anchors, -1 for negative anchors
# and 0 for neutral anchors.
target_rpn_match, target_rpn_bbox = modellib.build_rpn_targets(
image.shape, model.anchors, gt_class_id, gt_bbox, model.config)
log("target_rpn_match", target_rpn_match)
log("target_rpn_bbox", target_rpn_bbox)
positive_anchor_ix = np.where(target_rpn_match[:] == 1)[0]
negative_anchor_ix = np.where(target_rpn_match[:] == -1)[0]
neutral_anchor_ix = np.where(target_rpn_match[:] == 0)[0]
positive_anchors = model.anchors[positive_anchor_ix]
negative_anchors = model.anchors[negative_anchor_ix]
neutral_anchors = model.anchors[neutral_anchor_ix]
log("positive_anchors", positive_anchors)
log("negative_anchors", negative_anchors)
log("neutral anchors", neutral_anchors)
# Apply refinement deltas to positive anchors
refined_anchors = utils.apply_box_deltas(
positive_anchors,
target_rpn_bbox[:positive_anchors.shape[0]] * model.config.RPN_BBOX_STD_DEV)
log("refined_anchors", refined_anchors, )
# Display positive anchors before refinement (dotted) and
# after refinement (solid).
visualize.draw_boxes(image, boxes=positive_anchors, refined_boxes=refined_anchors, ax=get_ax())
```
### 1.b RPN Predictions
Here we run the RPN graph and display its predictions.
```
# Run RPN sub-graph
pillar = model.keras_model.get_layer("ROI").output # node to start searching from
# TF 1.4 introduces a new version of NMS. Search for both names to support TF 1.3 and 1.4
nms_node = model.ancestor(pillar, "ROI/rpn_non_max_suppression:0")
if nms_node is None:
nms_node = model.ancestor(pillar, "ROI/rpn_non_max_suppression/NonMaxSuppressionV2:0")
rpn = model.run_graph([image], [
("rpn_class", model.keras_model.get_layer("rpn_class").output),
("pre_nms_anchors", model.ancestor(pillar, "ROI/pre_nms_anchors:0")),
("refined_anchors", model.ancestor(pillar, "ROI/refined_anchors:0")),
("refined_anchors_clipped", model.ancestor(pillar, "ROI/refined_anchors_clipped:0")),
("post_nms_anchor_ix", nms_node),
("proposals", model.keras_model.get_layer("ROI").output),
])
# Show top anchors by score (before refinement)
limit = 100
sorted_anchor_ids = np.argsort(rpn['rpn_class'][:,:,1].flatten())[::-1]
visualize.draw_boxes(image, boxes=model.anchors[sorted_anchor_ids[:limit]], ax=get_ax())
# Show top anchors with refinement. Then with clipping to image boundaries
limit = 50
ax = get_ax(1, 2)
visualize.draw_boxes(image, boxes=rpn["pre_nms_anchors"][0, :limit],
refined_boxes=rpn["refined_anchors"][0, :limit], ax=ax[0])
visualize.draw_boxes(image, refined_boxes=rpn["refined_anchors_clipped"][0, :limit], ax=ax[1])
# Show refined anchors after non-max suppression
limit = 50
ixs = rpn["post_nms_anchor_ix"][:limit]
visualize.draw_boxes(image, refined_boxes=rpn["refined_anchors_clipped"][0, ixs], ax=get_ax())
# Show final proposals
# These are the same as the previous step (refined anchors
# after NMS) but with coordinates normalized to [0, 1] range.
limit = 50
# Convert back to image coordinates for display
h, w = config.IMAGE_SHAPE[:2]
proposals = rpn['proposals'][0, :limit] * np.array([h, w, h, w])
visualize.draw_boxes(image, refined_boxes=proposals, ax=get_ax())
```
## Stage 2: Proposal Classification
This stage takes the region proposals from the RPN and classifies them.
### 2.a Proposal Classification
Run the classifier heads on proposals to generate class propbabilities and bounding box regressions.
```
# Get input and output to classifier and mask heads.
mrcnn = model.run_graph([image], [
("proposals", model.keras_model.get_layer("ROI").output),
("probs", model.keras_model.get_layer("mrcnn_class").output),
("deltas", model.keras_model.get_layer("mrcnn_bbox").output),
("masks", model.keras_model.get_layer("mrcnn_mask").output),
("detections", model.keras_model.get_layer("mrcnn_detection").output),
])
# Get detection class IDs. Trim zero padding.
det_class_ids = mrcnn['detections'][0, :, 4].astype(np.int32)
det_count = np.where(det_class_ids == 0)[0][0]
det_class_ids = det_class_ids[:det_count]
detections = mrcnn['detections'][0, :det_count]
print("{} detections: {}".format(
det_count, np.array(dataset.class_names)[det_class_ids]))
captions = ["{} {:.3f}".format(dataset.class_names[int(c)], s) if c > 0 else ""
for c, s in zip(detections[:, 4], detections[:, 5])]
visualize.draw_boxes(
image,
refined_boxes=utils.denorm_boxes(detections[:, :4], image.shape[:2]),
visibilities=[2] * len(detections),
captions=captions, title="Detections",
ax=get_ax())
```
### 2.c Step by Step Detection
Here we dive deeper into the process of processing the detections.
```
# Proposals are in normalized coordinates. Scale them
# to image coordinates.
h, w = config.IMAGE_SHAPE[:2]
proposals = np.around(mrcnn["proposals"][0] * np.array([h, w, h, w])).astype(np.int32)
# Class ID, score, and mask per proposal
roi_class_ids = np.argmax(mrcnn["probs"][0], axis=1)
roi_scores = mrcnn["probs"][0, np.arange(roi_class_ids.shape[0]), roi_class_ids]
roi_class_names = np.array(dataset.class_names)[roi_class_ids]
roi_positive_ixs = np.where(roi_class_ids > 0)[0]
# How many ROIs vs empty rows?
print("{} Valid proposals out of {}".format(np.sum(np.any(proposals, axis=1)), proposals.shape[0]))
print("{} Positive ROIs".format(len(roi_positive_ixs)))
# Class counts
print(list(zip(*np.unique(roi_class_names, return_counts=True))))
# Display a random sample of proposals.
# Proposals classified as background are dotted, and
# the rest show their class and confidence score.
limit = 200
ixs = np.random.randint(0, proposals.shape[0], limit)
captions = ["{} {:.3f}".format(dataset.class_names[c], s) if c > 0 else ""
for c, s in zip(roi_class_ids[ixs], roi_scores[ixs])]
visualize.draw_boxes(image, boxes=proposals[ixs],
visibilities=np.where(roi_class_ids[ixs] > 0, 2, 1),
captions=captions, title="ROIs Before Refinement",
ax=get_ax())
```
#### Apply Bounding Box Refinement
```
# Class-specific bounding box shifts.
roi_bbox_specific = mrcnn["deltas"][0, np.arange(proposals.shape[0]), roi_class_ids]
log("roi_bbox_specific", roi_bbox_specific)
# Apply bounding box transformations
# Shape: [N, (y1, x1, y2, x2)]
refined_proposals = utils.apply_box_deltas(
proposals, roi_bbox_specific * config.BBOX_STD_DEV).astype(np.int32)
log("refined_proposals", refined_proposals)
# Show positive proposals
# ids = np.arange(roi_boxes.shape[0]) # Display all
limit = 5
ids = np.random.randint(0, len(roi_positive_ixs), limit) # Display random sample
captions = ["{} {:.3f}".format(dataset.class_names[c], s) if c > 0 else ""
for c, s in zip(roi_class_ids[roi_positive_ixs][ids], roi_scores[roi_positive_ixs][ids])]
visualize.draw_boxes(image, boxes=proposals[roi_positive_ixs][ids],
refined_boxes=refined_proposals[roi_positive_ixs][ids],
visibilities=np.where(roi_class_ids[roi_positive_ixs][ids] > 0, 1, 0),
captions=captions, title="ROIs After Refinement",
ax=get_ax())
```
#### Filter Low Confidence Detections
```
# Remove boxes classified as background
keep = np.where(roi_class_ids > 0)[0]
print("Keep {} detections:\n{}".format(keep.shape[0], keep))
# Remove low confidence detections
keep = np.intersect1d(keep, np.where(roi_scores >= config.DETECTION_MIN_CONFIDENCE)[0])
print("Remove boxes below {} confidence. Keep {}:\n{}".format(
config.DETECTION_MIN_CONFIDENCE, keep.shape[0], keep))
```
#### Per-Class Non-Max Suppression
```
# Apply per-class non-max suppression
pre_nms_boxes = refined_proposals[keep]
pre_nms_scores = roi_scores[keep]
pre_nms_class_ids = roi_class_ids[keep]
nms_keep = []
for class_id in np.unique(pre_nms_class_ids):
# Pick detections of this class
ixs = np.where(pre_nms_class_ids == class_id)[0]
# Apply NMS
class_keep = utils.non_max_suppression(pre_nms_boxes[ixs],
pre_nms_scores[ixs],
config.DETECTION_NMS_THRESHOLD)
# Map indicies
class_keep = keep[ixs[class_keep]]
nms_keep = np.union1d(nms_keep, class_keep)
print("{:22}: {} -> {}".format(dataset.class_names[class_id][:20],
keep[ixs], class_keep))
keep = np.intersect1d(keep, nms_keep).astype(np.int32)
print("\nKept after per-class NMS: {}\n{}".format(keep.shape[0], keep))
# Show final detections
ixs = np.arange(len(keep)) # Display all
# ixs = np.random.randint(0, len(keep), 10) # Display random sample
captions = ["{} {:.3f}".format(dataset.class_names[c], s) if c > 0 else ""
for c, s in zip(roi_class_ids[keep][ixs], roi_scores[keep][ixs])]
visualize.draw_boxes(
image, boxes=proposals[keep][ixs],
refined_boxes=refined_proposals[keep][ixs],
visibilities=np.where(roi_class_ids[keep][ixs] > 0, 1, 0),
captions=captions, title="Detections after NMS",
ax=get_ax())
```
## Stage 3: Generating Masks
This stage takes the detections (refined bounding boxes and class IDs) from the previous layer and runs the mask head to generate segmentation masks for every instance.
### 3.a Mask Targets
These are the training targets for the mask branch
```
display_images(np.transpose(gt_mask, [2, 0, 1]), cmap="Blues")
```
### 3.b Predicted Masks
```
# Get predictions of mask head
mrcnn = model.run_graph([image], [
("detections", model.keras_model.get_layer("mrcnn_detection").output),
("masks", model.keras_model.get_layer("mrcnn_mask").output),
])
# Get detection class IDs. Trim zero padding.
det_class_ids = mrcnn['detections'][0, :, 4].astype(np.int32)
det_count = np.where(det_class_ids == 0)[0][0]
det_class_ids = det_class_ids[:det_count]
print("{} detections: {}".format(
det_count, np.array(dataset.class_names)[det_class_ids]))
# Masks
det_boxes = utils.denorm_boxes(mrcnn["detections"][0, :, :4], image.shape[:2])
det_mask_specific = np.array([mrcnn["masks"][0, i, :, :, c]
for i, c in enumerate(det_class_ids)])
det_masks = np.array([utils.unmold_mask(m, det_boxes[i], image.shape)
for i, m in enumerate(det_mask_specific)])
log("det_mask_specific", det_mask_specific)
log("det_masks", det_masks)
display_images(det_mask_specific[:4] * 255, cmap="Blues", interpolation="none")
display_images(det_masks[:4] * 255, cmap="Blues", interpolation="none")
```
## Visualize Activations
In some cases it helps to look at the output from different layers and visualize them to catch issues and odd patterns.
```
# Get activations of a few sample layers
activations = model.run_graph([image], [
("input_image", model.keras_model.get_layer("input_image").output),
("res2c_out", model.keras_model.get_layer("res2c_out").output),
("res3c_out", model.keras_model.get_layer("res3c_out").output),
("res4w_out", model.keras_model.get_layer("res4w_out").output), # for resnet100
("rpn_bbox", model.keras_model.get_layer("rpn_bbox").output),
("roi", model.keras_model.get_layer("ROI").output),
])
# Input image (normalized)
_ = plt.imshow(modellib.unmold_image(activations["input_image"][0],config))
# Backbone feature map
display_images(np.transpose(activations["res2c_out"][0,:,:,:4], [2, 0, 1]), cols=4)
```
| github_jupyter |
# Gathering Earthquake Info with Libcomcat
### EQ of the Week, 15 March, 2022
Libcomcat is a python interface for searching the ANSS earthquake catalog. Using this package you can:
- search for earthquakes over a region
- search for earthquake info based on a known ID
- pull origin, moment tensor, DYFI, etc results for an event
- download event related data (ex: Finite-Fault)
### Acquiring Libcomcat:
The easiest way to download is through conda. It is recommended that you create a seperate conda environment.
```
conda install libcomcat
```
More info about libcomcat and how to download here: https://github.com/usgs/libcomcat#readme
```
# importing necessary packages
import libcomcat
from datetime import datetime
import pandas as pd
import numpy as np
# for visualization
#import matplotlib.pyplot as plt
import pygmt # <---- unsure how to use pygmt? Check out Utpal's notebook in the EWQ Github repository
```
### Searching the ComCat catalog for earthquakes
You can search the ANSS earthquake catalog for known events by setting a search region. The region can either be set as a bounding box, or by selecting a point and defining a radius. In this example, I am searching for all events over the year 2021 along the Alaskan Peninsula.
```
from libcomcat.search import search, get_event_by_id
from libcomcat.dataframes import (get_detail_data_frame, get_dyfi_data_frame,
get_history_data_frame, get_magnitude_data_frame,
get_pager_data_frame, get_phase_dataframe,
get_summary_data_frame)
# search params
search_region = [-162,-150, 52,58] # lonmin, lonmax, latmin, latmax
search_start_time = datetime(2021, 1, 1, 1, 1) # start of search timeframe
search_end_time = datetime(2021, 12, 31, 23, 59) # end of search timeframe
minmagnitude=3 # min magnitude (set to limit getting tons of small events)
maxmagnitude=10 # max magnitude (set here to be unreasonably high)
earthquakes = search(starttime=search_start_time, endtime=search_end_time,
minlatitude=search_region[2], maxlatitude=search_region[3],
minlongitude=search_region[0], maxlongitude=search_region[1],
minmagnitude=minmagnitude,maxmagnitude=maxmagnitude)
print('Querey found ',len(earthquakes),' earthquakes!')
print(earthquakes[0])
# convert the output to a more usable format:
catalog_df = pd.DataFrame( {'ID':[eq.id for eq in earthquakes],
'time':[eq.time for eq in earthquakes],
'longitude':[eq.longitude for eq in earthquakes],
'latitude':[eq.latitude for eq in earthquakes],
'depth':[eq.depth for eq in earthquakes],
'magnitude':[eq.magnitude for eq in earthquakes]})
catalog_df.head()
```
### Plot the results
```
fig = pygmt.Figure()
pygmt.config(MAP_FRAME_TYPE="plain")
fig.basemap(projection="M8c",region=search_region,frame=["a4f1",'WSne']) # set blank map over search region
fig.coast(shorelines=["1/0.5p","2/0.5p"],land="#e6e6e6", water="#e6f2ff") # add coast, land, sea colors
pygmt.makecpt(cmap="magma", series=(minmagnitude, maxmagnitude, 1)) # make color scale
fig.plot(x = catalog_df['longitude'],y=catalog_df['latitude'],
pen='0.1p',size=0.01*(1.75**catalog_df['magnitude']),
style='cc',color=catalog_df['magnitude'],cmap=True,transparency=30) # plot catalog as mag-scaled circles
fig.colorbar(frame=["x+lmagnitude"])
fig.show()
```
### Look at a single (larger) event:
Here I am grabbing the largest event from this catalog. It was the M8.2 Chignik earthquake that occured last summer.
```
idx = np.argmax(catalog_df['magnitude']) # get the largest magnitude earthquake in this catalog
event_id =catalog_df['ID'].iloc[idx]
event = get_event_by_id(event_id)
event
USGS_products, event = get_history_data_frame(event) # this can take ~10+ seconds for bigger events with more products
USGS_products['Product'].value_counts() # show what products are available
```
### Who felt this earthquake?
Here, I will grab the did you feel it reports and plot them on a map. The DYFI data is accessed through the "get_dyfi_data_frame" function
```
dyfi_df = get_dyfi_data_frame(event)
dyfi_df.head()
```
### Plot DYFI results
Here I set a region near the earthquake and expected affected communities. You can instead set your view region to be the as big as the furthest responses. Beware, though, sometimes extremely far-field responses dminish map quality.
```
dyfi_region = [-172, -140,50,63] # I set a region near the earthquake.
fig = pygmt.Figure()
pygmt.config(MAP_FRAME_TYPE="plain")
fig.basemap(projection="M8c",region=dyfi_region,frame=["a4f1",'WSne'])
fig.coast(shorelines=["1/0.1p","2/0.1p"],land="#e6e6e6", water="#e6f2ff")
pygmt.makecpt(cmap="jet", series=(1, 9, 1))
fig.plot(x = dyfi_df['lon'],y = dyfi_df['lat'],color=dyfi_df['intensity'],
style='d.2c',transparency=40,cmap=True)
fig.plot(x = event.longitude, y = event.latitude, pen='0.1p,red',style='a.5c',color='white')
fig.colorbar(frame=["x+lintensity"])
fig.show()
```
### Search for and download relevant data
```
products = event.getProducts('finite-fault', version='preferred') # get preferred finite-fault info
products[0].contents
filename = './'+event_id+'_finite_fault.param' # Downloading to this folder
products[0].getContent('basic_inversion.param',filename)
```
| github_jupyter |
# Character level language model - Dinosaurus land
Welcome to Dinosaurus Island! 65 million years ago, dinosaurs existed, and in this assignment they are back. You are in charge of a special task. Leading biology researchers are creating new breeds of dinosaurs and bringing them to life on earth, and your job is to give names to these dinosaurs. If a dinosaur does not like its name, it might go beserk, so choose wisely!
<table>
<td>
<img src="images/dino.jpg" style="width:250;height:300px;">
</td>
</table>
Luckily you have learned some deep learning and you will use it to save the day. Your assistant has collected a list of all the dinosaur names they could find, and compiled them into this [dataset](dinos.txt). (Feel free to take a look by clicking the previous link.) To create new dinosaur names, you will build a character level language model to generate new names. Your algorithm will learn the different name patterns, and randomly generate new names. Hopefully this algorithm will keep you and your team safe from the dinosaurs' wrath!
By completing this assignment you will learn:
- How to store text data for processing using an RNN
- How to synthesize data, by sampling predictions at each time step and passing it to the next RNN-cell unit
- How to build a character-level text generation recurrent neural network
- Why clipping the gradients is important
We will begin by loading in some functions that we have provided for you in `rnn_utils`. Specifically, you have access to functions such as `rnn_forward` and `rnn_backward` which are equivalent to those you've implemented in the previous assignment.
```
import numpy as np
from utils import *
import random
```
## 1 - Problem Statement
### 1.1 - Dataset and Preprocessing
Run the following cell to read the dataset of dinosaur names, create a list of unique characters (such as a-z), and compute the dataset and vocabulary size.
```
data = open('dinos.txt', 'r').read()
data= data.lower()
chars = list(set(data))
data_size, vocab_size = len(data), len(chars)
print('There are %d total characters and %d unique characters in your data.' % (data_size, vocab_size))
```
The characters are a-z (26 characters) plus the "\n" (or newline character), which in this assignment plays a role similar to the `<EOS>` (or "End of sentence") token we had discussed in lecture, only here it indicates the end of the dinosaur name rather than the end of a sentence. In the cell below, we create a python dictionary (i.e., a hash table) to map each character to an index from 0-26. We also create a second python dictionary that maps each index back to the corresponding character character. This will help you figure out what index corresponds to what character in the probability distribution output of the softmax layer. Below, `char_to_ix` and `ix_to_char` are the python dictionaries.
```
char_to_ix = { ch:i for i,ch in enumerate(sorted(chars)) }
ix_to_char = { i:ch for i,ch in enumerate(sorted(chars)) }
print(ix_to_char)
```
### 1.2 - Overview of the model
Your model will have the following structure:
- Initialize parameters
- Run the optimization loop
- Forward propagation to compute the loss function
- Backward propagation to compute the gradients with respect to the loss function
- Clip the gradients to avoid exploding gradients
- Using the gradients, update your parameter with the gradient descent update rule.
- Return the learned parameters
<img src="images/rnn.png" style="width:450;height:300px;">
<caption><center> **Figure 1**: Recurrent Neural Network, similar to what you had built in the previous notebook "Building a RNN - Step by Step". </center></caption>
At each time-step, the RNN tries to predict what is the next character given the previous characters. The dataset $X = (x^{\langle 1 \rangle}, x^{\langle 2 \rangle}, ..., x^{\langle T_x \rangle})$ is a list of characters in the training set, while $Y = (y^{\langle 1 \rangle}, y^{\langle 2 \rangle}, ..., y^{\langle T_x \rangle})$ is such that at every time-step $t$, we have $y^{\langle t \rangle} = x^{\langle t+1 \rangle}$.
## 2 - Building blocks of the model
In this part, you will build two important blocks of the overall model:
- Gradient clipping: to avoid exploding gradients
- Sampling: a technique used to generate characters
You will then apply these two functions to build the model.
### 2.1 - Clipping the gradients in the optimization loop
In this section you will implement the `clip` function that you will call inside of your optimization loop. Recall that your overall loop structure usually consists of a forward pass, a cost computation, a backward pass, and a parameter update. Before updating the parameters, you will perform gradient clipping when needed to make sure that your gradients are not "exploding," meaning taking on overly large values.
In the exercise below, you will implement a function `clip` that takes in a dictionary of gradients and returns a clipped version of gradients if needed. There are different ways to clip gradients; we will use a simple element-wise clipping procedure, in which every element of the gradient vector is clipped to lie between some range [-N, N]. More generally, you will provide a `maxValue` (say 10). In this example, if any component of the gradient vector is greater than 10, it would be set to 10; and if any component of the gradient vector is less than -10, it would be set to -10. If it is between -10 and 10, it is left alone.
<img src="images/clip.png" style="width:400;height:150px;">
<caption><center> **Figure 2**: Visualization of gradient descent with and without gradient clipping, in a case where the network is running into slight "exploding gradient" problems. </center></caption>
**Exercise**: Implement the function below to return the clipped gradients of your dictionary `gradients`. Your function takes in a maximum threshold and returns the clipped versions of your gradients. You can check out this [hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.clip.html) for examples of how to clip in numpy. You will need to use the argument `out = ...`.
```
### GRADED FUNCTION: clip
def clip(gradients, maxValue):
'''
Clips the gradients' values between minimum and maximum.
Arguments:
gradients -- a dictionary containing the gradients "dWaa", "dWax", "dWya", "db", "dby"
maxValue -- everything above this number is set to this number, and everything less than -maxValue is set to -maxValue
Returns:
gradients -- a dictionary with the clipped gradients.
'''
dWaa, dWax, dWya, db, dby = gradients['dWaa'], gradients['dWax'], gradients['dWya'], gradients['db'], gradients['dby']
### START CODE HERE ###
# clip to mitigate exploding gradients, loop over [dWax, dWaa, dWya, db, dby]. (≈2 lines)
for gradient in [dWax, dWaa, dWya, db, dby]:
np.clip(gradient, -maxValue, maxValue, out=gradient)
### END CODE HERE ###
gradients = {"dWaa": dWaa, "dWax": dWax, "dWya": dWya, "db": db, "dby": dby}
return gradients
np.random.seed(3)
dWax = np.random.randn(5,3)*10
dWaa = np.random.randn(5,5)*10
dWya = np.random.randn(2,5)*10
db = np.random.randn(5,1)*10
dby = np.random.randn(2,1)*10
gradients = {"dWax": dWax, "dWaa": dWaa, "dWya": dWya, "db": db, "dby": dby}
gradients = clip(gradients, 10)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("gradients[\"dWax\"][3][1] =", gradients["dWax"][3][1])
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
```
** Expected output:**
<table>
<tr>
<td>
**gradients["dWaa"][1][2] **
</td>
<td>
10.0
</td>
</tr>
<tr>
<td>
**gradients["dWax"][3][1]**
</td>
<td>
-10.0
</td>
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td>
0.29713815361
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td>
[ 10.]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>
[ 8.45833407]
</td>
</tr>
</table>
### 2.2 - Sampling
Now assume that your model is trained. You would like to generate new text (characters). The process of generation is explained in the picture below:
<img src="images/dinos3.png" style="width:500;height:300px;">
<caption><center> **Figure 3**: In this picture, we assume the model is already trained. We pass in $x^{\langle 1\rangle} = \vec{0}$ at the first time step, and have the network then sample one character at a time. </center></caption>
**Exercise**: Implement the `sample` function below to sample characters. You need to carry out 4 steps:
- **Step 1**: Pass the network the first "dummy" input $x^{\langle 1 \rangle} = \vec{0}$ (the vector of zeros). This is the default input before we've generated any characters. We also set $a^{\langle 0 \rangle} = \vec{0}$
- **Step 2**: Run one step of forward propagation to get $a^{\langle 1 \rangle}$ and $\hat{y}^{\langle 1 \rangle}$. Here are the equations:
$$ a^{\langle t+1 \rangle} = \tanh(W_{ax} x^{\langle t \rangle } + W_{aa} a^{\langle t \rangle } + b)\tag{1}$$
$$ z^{\langle t + 1 \rangle } = W_{ya} a^{\langle t + 1 \rangle } + b_y \tag{2}$$
$$ \hat{y}^{\langle t+1 \rangle } = softmax(z^{\langle t + 1 \rangle })\tag{3}$$
Note that $\hat{y}^{\langle t+1 \rangle }$ is a (softmax) probability vector (its entries are between 0 and 1 and sum to 1). $\hat{y}^{\langle t+1 \rangle}_i$ represents the probability that the character indexed by "i" is the next character. We have provided a `softmax()` function that you can use.
- **Step 3**: Carry out sampling: Pick the next character's index according to the probability distribution specified by $\hat{y}^{\langle t+1 \rangle }$. This means that if $\hat{y}^{\langle t+1 \rangle }_i = 0.16$, you will pick the index "i" with 16% probability. To implement it, you can use [`np.random.choice`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.random.choice.html).
Here is an example of how to use `np.random.choice()`:
```python
np.random.seed(0)
p = np.array([0.1, 0.0, 0.7, 0.2])
index = np.random.choice([0, 1, 2, 3], p = p.ravel())
```
This means that you will pick the `index` according to the distribution:
$P(index = 0) = 0.1, P(index = 1) = 0.0, P(index = 2) = 0.7, P(index = 3) = 0.2$.
- **Step 4**: The last step to implement in `sample()` is to overwrite the variable `x`, which currently stores $x^{\langle t \rangle }$, with the value of $x^{\langle t + 1 \rangle }$. You will represent $x^{\langle t + 1 \rangle }$ by creating a one-hot vector corresponding to the character you've chosen as your prediction. You will then forward propagate $x^{\langle t + 1 \rangle }$ in Step 1 and keep repeating the process until you get a "\n" character, indicating you've reached the end of the dinosaur name.
```
# GRADED FUNCTION: sample
def sample(parameters, char_to_ix, seed):
"""
Sample a sequence of characters according to a sequence of probability distributions output of the RNN
Arguments:
parameters -- python dictionary containing the parameters Waa, Wax, Wya, by, and b.
char_to_ix -- python dictionary mapping each character to an index.
seed -- used for grading purposes. Do not worry about it.
Returns:
indices -- a list of length n containing the indices of the sampled characters.
"""
# Retrieve parameters and relevant shapes from "parameters" dictionary
Waa, Wax, Wya, by, b = parameters['Waa'], parameters['Wax'], parameters['Wya'], parameters['by'], parameters['b']
vocab_size = by.shape[0]
n_a = Waa.shape[1]
### START CODE HERE ###
# Step 1: Create the one-hot vector x for the first character (initializing the sequence generation). (≈1 line)
x = np.zeros((vocab_size, 1))
# Step 1': Initialize a_prev as zeros (≈1 line)
a_prev = np.zeros((n_a,1))
# Create an empty list of indices, this is the list which will contain the list of indices of the characters to generate (≈1 line)
indices = []
# Idx is a flag to detect a newline character, we initialize it to -1
idx = -1
# Loop over time-steps t. At each time-step, sample a character from a probability distribution and append
# its index to "indices". We'll stop if we reach 50 characters (which should be very unlikely with a well
# trained model), which helps debugging and prevents entering an infinite loop.
counter = 0
newline_character = char_to_ix['\n']
while (idx != newline_character and counter != 50):
# Step 2: Forward propagate x using the equations (1), (2) and (3)
a = np.tanh(np.dot(Wax, x) + np.dot(Waa, a_prev) + b)
z = np.dot(Wya, a) + by
y = softmax(z)
# for grading purposes
np.random.seed(counter+seed)
# Step 3: Sample the index of a character within the vocabulary from the probability distribution y
idx = np.random.choice(np.arange(vocab_size), p=y.ravel())
# Append the index to "indices"
indices.append(idx)
# Step 4: Overwrite the input character as the one corresponding to the sampled index.
x = np.zeros((vocab_size, 1))
x[idx,0] = 1
# Update "a_prev" to be "a"
a_prev = a
# for grading purposes
seed += 1
counter +=1
### END CODE HERE ###
if (counter == 50):
indices.append(char_to_ix['\n'])
return indices
np.random.seed(2)
_, n_a = 20, 100
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
indices = sample(parameters, char_to_ix, 0)
print("Sampling:")
print("list of sampled indices:", indices)
print("list of sampled characters:", [ix_to_char[i] for i in indices])
```
** Expected output:**
<table>
<tr>
<td>
**list of sampled indices:**
</td>
<td>
[12, 17, 24, 14, 13, 9, 10, 22, 24, 6, 13, 11, 12, 6, 21, 15, 21, 14, 3, 2, 1, 21, 18, 24, <br>
7, 25, 6, 25, 18, 10, 16, 2, 3, 8, 15, 12, 11, 7, 1, 12, 10, 2, 7, 7, 11, 5, 6, 12, 25, 0, 0]
</td>
</tr><tr>
<td>
**list of sampled characters:**
</td>
<td>
['l', 'q', 'x', 'n', 'm', 'i', 'j', 'v', 'x', 'f', 'm', 'k', 'l', 'f', 'u', 'o', <br>
'u', 'n', 'c', 'b', 'a', 'u', 'r', 'x', 'g', 'y', 'f', 'y', 'r', 'j', 'p', 'b', 'c', 'h', 'o', <br>
'l', 'k', 'g', 'a', 'l', 'j', 'b', 'g', 'g', 'k', 'e', 'f', 'l', 'y', '\n', '\n']
</td>
</tr>
</table>
## 3 - Building the language model
It is time to build the character-level language model for text generation.
### 3.1 - Gradient descent
In this section you will implement a function performing one step of stochastic gradient descent (with clipped gradients). You will go through the training examples one at a time, so the optimization algorithm will be stochastic gradient descent. As a reminder, here are the steps of a common optimization loop for an RNN:
- Forward propagate through the RNN to compute the loss
- Backward propagate through time to compute the gradients of the loss with respect to the parameters
- Clip the gradients if necessary
- Update your parameters using gradient descent
**Exercise**: Implement this optimization process (one step of stochastic gradient descent).
We provide you with the following functions:
```python
def rnn_forward(X, Y, a_prev, parameters):
""" Performs the forward propagation through the RNN and computes the cross-entropy loss.
It returns the loss' value as well as a "cache" storing values to be used in the backpropagation."""
....
return loss, cache
def rnn_backward(X, Y, parameters, cache):
""" Performs the backward propagation through time to compute the gradients of the loss with respect
to the parameters. It returns also all the hidden states."""
...
return gradients, a
def update_parameters(parameters, gradients, learning_rate):
""" Updates parameters using the Gradient Descent Update Rule."""
...
return parameters
```
```
# GRADED FUNCTION: optimize
def optimize(X, Y, a_prev, parameters, learning_rate = 0.01):
"""
Execute one step of the optimization to train the model.
Arguments:
X -- list of integers, where each integer is a number that maps to a character in the vocabulary.
Y -- list of integers, exactly the same as X but shifted one index to the left.
a_prev -- previous hidden state.
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
b -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
learning_rate -- learning rate for the model.
Returns:
loss -- value of the loss function (cross-entropy)
gradients -- python dictionary containing:
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dWya -- Gradients of hidden-to-output weights, of shape (n_y, n_a)
db -- Gradients of bias vector, of shape (n_a, 1)
dby -- Gradients of output bias vector, of shape (n_y, 1)
a[len(X)-1] -- the last hidden state, of shape (n_a, 1)
"""
### START CODE HERE ###
# Forward propagate through time (≈1 line)
loss, cache = rnn_forward(X, Y, a_prev, parameters)
# Backpropagate through time (≈1 line)
gradients, a = rnn_backward(X, Y, parameters, cache)
# Clip your gradients between -5 (min) and 5 (max) (≈1 line)
gradients = clip(gradients, 5)
# Update parameters (≈1 line)
parameters = update_parameters(parameters, gradients, learning_rate)
### END CODE HERE ###
return loss, gradients, a[len(X)-1]
np.random.seed(1)
vocab_size, n_a = 27, 100
a_prev = np.random.randn(n_a, 1)
Wax, Waa, Wya = np.random.randn(n_a, vocab_size), np.random.randn(n_a, n_a), np.random.randn(vocab_size, n_a)
b, by = np.random.randn(n_a, 1), np.random.randn(vocab_size, 1)
parameters = {"Wax": Wax, "Waa": Waa, "Wya": Wya, "b": b, "by": by}
X = [12,3,5,11,22,3]
Y = [4,14,11,22,25, 26]
loss, gradients, a_last = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
print("Loss =", loss)
print("gradients[\"dWaa\"][1][2] =", gradients["dWaa"][1][2])
print("np.argmax(gradients[\"dWax\"]) =", np.argmax(gradients["dWax"]))
print("gradients[\"dWya\"][1][2] =", gradients["dWya"][1][2])
print("gradients[\"db\"][4] =", gradients["db"][4])
print("gradients[\"dby\"][1] =", gradients["dby"][1])
print("a_last[4] =", a_last[4])
```
** Expected output:**
<table>
<tr>
<td>
**Loss **
</td>
<td>
126.503975722
</td>
</tr>
<tr>
<td>
**gradients["dWaa"][1][2]**
</td>
<td>
0.194709315347
</td>
<tr>
<td>
**np.argmax(gradients["dWax"])**
</td>
<td> 93
</td>
</tr>
<tr>
<td>
**gradients["dWya"][1][2]**
</td>
<td> -0.007773876032
</td>
</tr>
<tr>
<td>
**gradients["db"][4]**
</td>
<td> [-0.06809825]
</td>
</tr>
<tr>
<td>
**gradients["dby"][1]**
</td>
<td>[ 0.01538192]
</td>
</tr>
<tr>
<td>
**a_last[4]**
</td>
<td> [-1.]
</td>
</tr>
</table>
### 3.2 - Training the model
Given the dataset of dinosaur names, we use each line of the dataset (one name) as one training example. Every 100 steps of stochastic gradient descent, you will sample 10 randomly chosen names to see how the algorithm is doing. Remember to shuffle the dataset, so that stochastic gradient descent visits the examples in random order.
**Exercise**: Follow the instructions and implement `model()`. When `examples[index]` contains one dinosaur name (string), to create an example (X, Y), you can use this:
```python
index = j % len(examples)
X = [None] + [char_to_ix[ch] for ch in examples[index]]
Y = X[1:] + [char_to_ix["\n"]]
```
Note that we use: `index= j % len(examples)`, where `j = 1....num_iterations`, to make sure that `examples[index]` is always a valid statement (`index` is smaller than `len(examples)`).
The first entry of `X` being `None` will be interpreted by `rnn_forward()` as setting $x^{\langle 0 \rangle} = \vec{0}$. Further, this ensures that `Y` is equal to `X` but shifted one step to the left, and with an additional "\n" appended to signify the end of the dinosaur name.
```
# GRADED FUNCTION: model
def model(data, ix_to_char, char_to_ix, num_iterations = 35000, n_a = 50, dino_names = 7, vocab_size = 27):
"""
Trains the model and generates dinosaur names.
Arguments:
data -- text corpus
ix_to_char -- dictionary that maps the index to a character
char_to_ix -- dictionary that maps a character to an index
num_iterations -- number of iterations to train the model for
n_a -- number of units of the RNN cell
dino_names -- number of dinosaur names you want to sample at each iteration.
vocab_size -- number of unique characters found in the text, size of the vocabulary
Returns:
parameters -- learned parameters
"""
# Retrieve n_x and n_y from vocab_size
n_x, n_y = vocab_size, vocab_size
# Initialize parameters
parameters = initialize_parameters(n_a, n_x, n_y)
# Initialize loss (this is required because we want to smooth our loss, don't worry about it)
loss = get_initial_loss(vocab_size, dino_names)
# Build list of all dinosaur names (training examples).
with open("dinos.txt") as f:
examples = f.readlines()
examples = [x.lower().strip() for x in examples]
# Shuffle list of all dinosaur names
np.random.seed(0)
np.random.shuffle(examples)
# Initialize the hidden state of your LSTM
a_prev = np.zeros((n_a, 1))
# Optimization loop
for j in range(num_iterations):
### START CODE HERE ###
# Use the hint above to define one training example (X,Y) (≈ 2 lines)
index = j % len(examples)
X = [None] + [char_to_ix[ch] for ch in examples[index]] #YD: Add [None] so X and Y are shifted 1 step
Y = X[1:] + [char_to_ix['\n']]
# Perform one optimization step: Forward-prop -> Backward-prop -> Clip -> Update parameters
# Choose a learning rate of 0.01
curr_loss, gradients, a_prev = optimize(X, Y, a_prev, parameters, learning_rate = 0.01)
### END CODE HERE ###
# Use a latency trick to keep the loss smooth. It happens here to accelerate the training.
loss = smooth(loss, curr_loss)
# Every 2000 Iteration, generate "n" characters thanks to sample() to check if the model is learning properly
if j % 2000 == 0:
print('Iteration: %d, Loss: %f' % (j, loss) + '\n')
# The number of dinosaur names to print
seed = 0
for name in range(dino_names):
# Sample indices and print them
sampled_indices = sample(parameters, char_to_ix, seed)
print_sample(sampled_indices, ix_to_char)
seed += 1 # To get the same result for grading purposed, increment the seed by one.
print('\n')
return parameters
```
Run the following cell, you should observe your model outputting random-looking characters at the first iteration. After a few thousand iterations, your model should learn to generate reasonable-looking names.
```
parameters = model(data, ix_to_char, char_to_ix)
```
## Conclusion
You can see that your algorithm has started to generate plausible dinosaur names towards the end of the training. At first, it was generating random characters, but towards the end you could see dinosaur names with cool endings. Feel free to run the algorithm even longer and play with hyperparameters to see if you can get even better results. Our implemetation generated some really cool names like `maconucon`, `marloralus` and `macingsersaurus`. Your model hopefully also learned that dinosaur names tend to end in `saurus`, `don`, `aura`, `tor`, etc.
If your model generates some non-cool names, don't blame the model entirely--not all actual dinosaur names sound cool. (For example, `dromaeosauroides` is an actual dinosaur name and is in the training set.) But this model should give you a set of candidates from which you can pick the coolest!
This assignment had used a relatively small dataset, so that you could train an RNN quickly on a CPU. Training a model of the english language requires a much bigger dataset, and usually needs much more computation, and could run for many hours on GPUs. We ran our dinosaur name for quite some time, and so far our favoriate name is the great, undefeatable, and fierce: Mangosaurus!
<img src="images/mangosaurus.jpeg" style="width:250;height:300px;">
## 4 - Writing like Shakespeare
The rest of this notebook is optional and is not graded, but we hope you'll do it anyway since it's quite fun and informative.
A similar (but more complicated) task is to generate Shakespeare poems. Instead of learning from a dataset of Dinosaur names you can use a collection of Shakespearian poems. Using LSTM cells, you can learn longer term dependencies that span many characters in the text--e.g., where a character appearing somewhere a sequence can influence what should be a different character much much later in ths sequence. These long term dependencies were less important with dinosaur names, since the names were quite short.
<img src="images/shakespeare.jpg" style="width:500;height:400px;">
<caption><center> Let's become poets! </center></caption>
We have implemented a Shakespeare poem generator with Keras. Run the following cell to load the required packages and models. This may take a few minutes.
```
from __future__ import print_function
from keras.callbacks import LambdaCallback
from keras.models import Model, load_model, Sequential
from keras.layers import Dense, Activation, Dropout, Input, Masking
from keras.layers import LSTM
from keras.utils.data_utils import get_file
from keras.preprocessing.sequence import pad_sequences
from shakespeare_utils import *
import sys
import io
```
To save you some time, we have already trained a model for ~1000 epochs on a collection of Shakespearian poems called [*"The Sonnets"*](shakespeare.txt).
Let's train the model for one more epoch. When it finishes training for an epoch---this will also take a few minutes---you can run `generate_output`, which will prompt asking you for an input (`<`40 characters). The poem will start with your sentence, and our RNN-Shakespeare will complete the rest of the poem for you! For example, try "Forsooth this maketh no sense " (don't enter the quotation marks). Depending on whether you include the space at the end, your results might also differ--try it both ways, and try other inputs as well.
```
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y, batch_size=128, epochs=1, callbacks=[print_callback])
# Run this cell to try with different inputs without having to re-train the model
generate_output()
```
The RNN-Shakespeare model is very similar to the one you have built for dinosaur names. The only major differences are:
- LSTMs instead of the basic RNN to capture longer-range dependencies
- The model is a deeper, stacked LSTM model (2 layer)
- Using Keras instead of python to simplify the code
If you want to learn more, you can also check out the Keras Team's text generation implementation on GitHub: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py.
Congratulations on finishing this notebook!
**References**:
- This exercise took inspiration from Andrej Karpathy's implementation: https://gist.github.com/karpathy/d4dee566867f8291f086. To learn more about text generation, also check out Karpathy's [blog post](http://karpathy.github.io/2015/05/21/rnn-effectiveness/).
- For the Shakespearian poem generator, our implementation was based on the implementation of an LSTM text generator by the Keras team: https://github.com/keras-team/keras/blob/master/examples/lstm_text_generation.py
| github_jupyter |
# 机器学习工程师纳米学位
## 强化学习
## 项目 4: 训练智能出租车学会驾驶
欢迎来到机器学习工程师纳米学位的第四个项目!在这个notebook文件中,模板代码已经提供给你,有助于你对*智能出租车*的分析和实现学习算法。你无须改动已包含的代码,除非另有要求。 你需要回答notebook文件中给出的与项目或可视化相关的问题。每一个你要回答的问题前都会冠以**'问题 X'**。仔细阅读每个问题,并在后面**'回答'**文本框内给出完整的回答。你提交的项目会根据你对于每个问题的回答以及提交的`agent.py`的实现来进行评分。
>**提示:** Code 和 Markdown 单元格可通过 **Shift + Enter** 快捷键来执行。此外,Markdown可以通过双击进入编辑模式。
-----
## 开始
在这个项目中,你将构建一个优化的Q-Learning驾驶代理程序,它会操纵*智能出租车* 通过它的周边环境到达目的地。因为人们期望*智能出租车*要将乘客从一个地方载到另一个地方,驾驶代理程序会以两个非常重要的指标来评价:**安全性**和**可靠性**。驾驶代理程序在红灯亮时仍然让*智能出租车*行驶往目的地或者勉强避开事故会被认为是**不安全**的。类似的,驾驶代理程序频繁地不能适时地到达目的地会被认为**不可靠**。最大化驾驶代理程序的**安全性**和**可靠性**保证了*智能出租车*会在交通行业获得长期的地位。
**安全性**和**可靠性**用字母等级来评估,如下:
| 等级 | 安全性 | 可靠性 |
|:-----: |:------: |:-----------: |
| A+ | 代理程序没有任何妨害交通的行为,<br/>并且总是能选择正确的行动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的100%。 |
| A | 代理程序有很少的轻微妨害交通的行为,<br/>如绿灯时未能移动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的90%。 |
| B | 代理程序频繁地有轻微妨害交通行为,<br/>如绿灯时未能移动。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的80%。 |
| C | 代理程序有至少一次重大的妨害交通行为,<br/>如闯红灯。| 代理程序在合理时间内到达目的地的次数<br />占行驶次数的70%。 |
| D | 代理程序造成了至少一次轻微事故,<br/>如绿灯时在对面有车辆情况下左转。 | 代理程序在合理时间内到达目的地的次数<br />占行驶次数的60%。 |
| F | 代理程序造成了至少一次重大事故,<br/>如有交叉车流时闯红灯。 | 代理程序在合理时间内到达目的地的次数<br />未能达到行驶次数的60%。 |
为了协助评估这些重要的指标,你会需要加载可视化模块的代码,会在之后的项目中用到。运行下面的代码格来导入这个代码,你的分析中会需要它。
```
# 检查你的Python版本
from sys import version_info
if version_info.major != 2 and version_info.minor != 7:
raise Exception('请使用Python 2.7来完成此项目')
# Import the visualization code
import visuals as vs
# Pretty display for notebooks
%matplotlib inline
```
### 了解世界
在开始实现你的驾驶代理程序前,首先需要了解*智能出租车*和驾驶代理程序运行的这个世界(环境)。构建自我学习的代理程序重要的组成部分之一就是了解代理程序的特征,包括代理程序如何运作。原样直接运行`agent.py`代理程序的代码,不需要做任何额外的修改。让结果模拟运行一段时间,以观察各个不同的工作模块。注意在可视化模拟程序(如果启用了),**白色车辆**就是*智能出租车*。
### 问题 1
用几句话,描述在运行默认的`agent.py`代理程序中,你在模拟程序里观察到了什么。一些你可以考虑的情况:
- *在模拟过程中,智能出租车究竟移动了吗?*
- *驾驶代理程序获得了什么样的奖励?*
- *交通灯的颜色改变是如何影响奖励的?*
**提示:** 从顶层的`/smartcab/`目录(这个notebook所在的地方),运行命令
```bash
'python smartcab/agent.py'
```
**回答:**
- 在模拟过程中,智能小车没有移动
- 代理程序分别获得了正奖励和负奖励
- 交通灯为红色时,获得正奖励,为绿色时,获得负奖励
### 理解代码
除了要了解世界之外,还需要理解掌管世界、模拟程序等等如何运作的代码本身。如果一点也不去探索一下*“隐藏”*的器件,就试着去创建一个驾驶代理程序会很难。在顶层的`/smartcab/`的目录下,有两个文件夹:`/logs/` (之后会用到)和`/smartcab/`。打开`/smartcab/`文件夹,探索每个下面的Python文件,然后回答下面的问题。
### 问题 2
- *在*`agent.py`* Python文件里,选择 3 个可以设定的 flag,并描述他们如何改变模拟程序的。*
- *在*`environment.py`* Python文件里,当代理程序执行一个行动时,调用哪个Environment类的函数?*
- *在*`simulator.py`* Python 文件里,*`'render_text()'`*函数和*`'render()'`*函数之间的区别是什么?*
- *在*`planner.py`* Python文件里,*`'next_waypoint()`* 函数会先考虑南北方向还是东西方向?*
**回答:**
-
- update_delay 用于设定更新速率
- display 用于设定是否启用窗口显示
- log_metrics 用于输出训练日志
- 当代理程序执行一个行动时,会调用act这个函数
- render_text这个函数用于无窗口的,而render用于用窗口的
- 会优先考虑东西方向
-----
## 实现一个基本的驾驶代理程序
创建一个优化Q-Learning的驾驶代理程序的第一步,是让代理程序确实地执行有效的行动。在这个情况下,一个有效的行动是`None`(不做任何行动)、`'Left'`(左转)、`'Right'`(右转)或者`'Forward'`(前进)。作为你的第一个实现,到`'choose_action()'`代理程序函数,使驾驶代理程序随机选择其中的一个动作。注意你会访问到几个类的成员变量,它们有助于你编写这个功能,比如`'self.learning'`和`'self.valid_actions'`。实现后,运行几次代理程序文件和模拟程序来确认你的驾驶代理程序每步都执行随机的动作。
### 基本代理程序模拟结果
要从最初的模拟程序获得结果,你需要调整下面的标志:
- `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
- `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
- `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件。
- `'n_test'` - 将此标志设定为`'10'`则执行10次测试试验。
可选的,你还可以通过将`'display'`标志设定为`False`来禁用可视化模拟(可以使得试验跑得更快)。调试时,设定的标志会返回到他们的默认设定。重要的是要理解每个标志以及它们如何影响到模拟。
你成功完成了最初的模拟后(有20个训练试验和10个测试试验),运行下面的代码单元格来使结果可视化。注意运行同样的模拟时,日志文件会被覆写,所以留意被载入的日志文件!在 projects/smartcab 下运行 agent.py 文件。
```
# Load the 'sim_no-learning' log file from the initial simulation results
vs.plot_trials('sim_no-learning.csv')
```
### 问题 3
利用上面的从你初始模拟中得到的可视化结果,给出关于驾驶代理程序的分析和若干观察。确保对于可视化结果上的每个面板你至少给出一条观察结果。你可以考虑的一些情况:
- *驾驶代理程序多频繁地做出不良决策?有多少不良决策造成了事故?*
- *假定代理程序是随机驾驶,那么可靠率是否合理?*
- *代理程序对于它的行动会获得什么样的奖励?奖励是否表明了它收到严重的惩罚?*
- *随着试验数增加,结果输出是否有重大变化?*
- *这个智能出租车对于乘客来说,会被人为是安全的且/或可靠的吗?为什么或者为什么不?*
**答案:**
- 非常频繁的做出不良决策,大概有40%造成了事故
- 假定代理程序是随机驾驶的,那么可靠率比较合理
- 代理程序对于它的行动会得到一些负面奖励,说明了收到了严重的惩罚
- 随着实验的增加,结果并没有太大改变,因为是随机的
- 这个智能车对于乘客来说,不会被认为是安全的。因为所有动作都是随机的,没有安全性可言。
-----
## 通知驾驶代理程序
创建一个优化Q-Learning的驾驶代理程序的第二步,是定义一系列代理程序会在环境中发生的状态。根据输入、感知数据和驾驶代理程序可用的变量,可以为代理程序定义一系列状态,使它最终可以*学习*在一个状态下它需要执行哪个动作。对于每个状态的`'如果这个处于这个状态就那个行动'`的状况称为**策略**,就是最终驾驶代理程序要学习的。没有定义状态,驾驶代理程序就不会明白哪个动作是最优的——或者甚至不会明白它要关注哪个环境变量和条件!
### 识别状态
查看`'build_state()'`代理程序函数,它显示驾驶代理函数可以从环境中获得下列数据:
- `'waypoint'`,*智能出租车*去向目的地应该行驶的方向,它是*智能出租车*车头方向的相对值。
- `'inputs'`,*智能出租车*的感知器数据。它包括
- `'light'`,交通灯颜色。
- `'left'`,*智能出租车*左侧车辆的目的方向。如果没有车辆,则返回`None`。
- `'right'`,*智能出租车*右侧车辆的目的方向。如果没有车辆,则返回`None`。
- `'oncoming'`,*智能出租车*交叉方向车辆的目的方向。如果没有车辆,则返回`None`。
- `'deadline'`,*智能出租车*在时间之内到达目的地还所需的剩余动作数目。
### 问题 4
*代理程序的哪些可用特征与学习**安全性**和**效率**相关性最高?你为什么认为这些特征适合在环境中对**智能出租车**建模?如果你没有选择某些特征,放弃他们的原因是什么?*
**回答:**
waypoint,oncoming和light这两个特征对学习安全性和效率相关性最高。因为交通灯颜色和对面车辆的方向对于智能车的安全性很重要,而智能车应该去的目的地方向则能很好的引导智能车行驶到指定位置。left,righ这两个状态是智能车的左右车辆的目的方向,加上这两个可以更好的掌握安全性。而剩余动作数目虽然对于效率有作用,但是剩余数目的类别太多,会使状态空间非常大,造成智能车的训练可能不足,而且好多的状态并不是频繁出现,所以不选择deadline。
### 定义状态空间
当定义一系列代理程序会处于的状态,必需考虑状态空间的*大小*。就是说,如果你期望驾驶代理程序针对每个状态都学习一个**策略**,你会需要对于每一个代理状态都有一个最优的动作。如果所有可能状态的数量非常大,最后会变成这样的状况,驾驶代理程序对于某些状态学不到如何行动,会导致未学习过的决策。例如,考虑用下面的特征定义*智能出租车*的状态的情况:
`('is_raining', 'is_foggy', 'is_red_light', 'turn_left', 'no_traffic', 'previous_turn_left', 'time_of_day')`.
发生如`(False, True, True, True, False, False, '3AM')`的状态的频次如何?没有近乎无限数量的训练,很怀疑代理程序会学到一个合适的动作!
### 问题 5
*如果用你在**问题4**中选择的特征来定义一个状态,状态空间的大小是多少?假定你了解环境以及它是如何模拟的,你觉得经过合理数量的训练之后,代理驾驶能学到一个较好的策略吗?(遇见绝大部分状态都能作出正确决策。)*
**提示:** 考虑特征*组合*来计算状态的总数!
**回答:**
按照问题四定义状态,状态空间是3X2X4X4X4 = 384个。我认为通过三百次以上数量的训练,代理驾驶才能能学到一个比较好的策略
### 更新驾驶代理程序的状态
要完成你的第二个实现,去到`'build_state()'`代理程序函数。根据你在**问题4**给出的判断,你现在要将`'state'`变量设定为包含所有Q-Learning所需特征的元组。确认你的驾驶代理程序通过运行代理程序文件和模拟会更新它的状态,注意状态是否显示了。如果用了可视化模拟,确认更新的状态和在模拟程序里看到的一致。
**注意:** 观察时记住重置模拟程序的标志到默认设定!
-----
## 实现Q-Learning驾驶代理程序
创建一个优化Q-Learning的驾驶代理程序的第三步,是开始实现Q-Learning自身的功能。Q-Learning的概念相当直接:每个访问的状态,为所有可用的状态-行动配对在Q-table里创建一条记录。然后,当代理程序遇到一个状态并执行了一个动作,基于获得的奖励和设定的相互的更新规则,来更新关联的状态-动作配对的Q-value。当然,Q-Learning还带来其他的收益,如此我们可以让代理程序根据每个可能的状态-动作配对的Q-values,来为每个状态选择*最佳*动作。在这个项目里,你会实现一个*衰减* $\epsilon$ *-贪心* 的Q-learning算法,不含折扣因子。遵从每个代理程序函数的**TODO**下的实现指导。
注意代理程序的属性`self.Q`是一个字典:这就是Q-table的构成。每个状态是`self.Q`字典的键,每个值是另一个字典,包含了*action*和*Q-value*。这里是个样例:
```
{ 'state-1': {
'action-1' : Qvalue-1,
'action-2' : Qvalue-2,
...
},
'state-2': {
'action-1' : Qvalue-1,
...
},
...
}
```
此外,注意你要求利用一个*衰减*$\epsilon$*(探索)因子*。因此,随着试验的增加,$\epsilon$会向0减小。这是因为,代理程序会从它的行为中学习,然后根据习得的行为行动。而且当$\epsilon$达到特定阈值后(默认阈值为0.01),代理程序被以它所学到的东西来作检测。作为初始的Q-Learning实现,你将实现一个线性衰减$\epsilon$的函数。
### Q-Learning模拟结果
要从最初的Q-learning程序获得结果,你需要调整下面的标志和设置:
- `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
- `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
- `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件,Q-table存为`.txt`文件。
- `'n_test'` - 将此标志设定为`'10'`则执行10次测试试验。
- `'learning'` - 将此标志设定为`'True'`来告诉驾驶代理使用你的Q-Learning实现。
此外,使用下面的$\epsilon$衰减函数:
$$ \epsilon_{t+1} = \epsilon_{t} - 0.05, \hspace{10px}\textrm{for trial number } t$$
如果你在实施时遇到困难,尝试把`'verbose'`标志设为`True`来调试。调试时,在这里设定的标志会返回到它们的默认设定。重要的是你要理解每个标志做什么并且解释它们怎么影响模拟!
当你成功完成初始的Q-Learning模拟程序后,运行下面代码单元格来使结果可视化。注意当相同的模拟运行时,log文件会被覆写,所以要留意载入的log文件!
```
# Load the 'sim_default-learning' file from the default Q-Learning simulation
vs.plot_trials('sim_default-learning.csv')
```
### 问题 6
利用上面的从你默认的Q-Learning模拟中得到的可视化结果,像在**问题3**那样,给出关于驾驶代理程序的分析和若干观察。注意模拟程序应该也产生了Q-table存在一个文本文件中,可以帮到你观察代理程序的算法。你可以考虑的一些情况:
- *有没有观察到基本驾驶代理程序和默认的Q-Learning代理程序的相似之处?*
- *在测试之前驾驶代理大约需要做多少训练试验?在给定的$\epsilon$ 容忍度下,这个数字是否合理?*
- *你实现的$\epsilon$(探索因子)衰减函数是否准确地在参数面板中显示?*
- *随着试验数增加,不良动作的数目是否减少?平均奖励是否增加?*
- *与初始的驾驶代理程序相比,安全性和可靠性评分怎样?*
**回答:**
- 在训练初期,默认的q学习代理和基本的很相似,因为处于高探索率,q学习代理程序也会进行随机决策
- 大约进行了20次训练实验,在给定的容忍度下,这个数字不够合理
- 我实现的探索因子衰减函数准确的在参数面板中显示了
- 随着实验数增加,不良动作的数目逐渐减少,平均奖励也在增加
- 与初始驾驶代理程序相比,安全性和可靠性都没有明显增加,原因可能是训练数量不够。分别是F和F
-----
## 改进Q-Learning驾驶代理程序
创建一个优化Q-Learning的驾驶代理程序的第三步,是执行优化!现在Q-Learning算法已经实现并且驾驶代理程序已经成功学习了,需要调整设定、调节参数让驾驶代理程序学习**安全性**和**效率**。通常这一步需要很多试验和错误,因为某些设定必定会造成更糟糕的学习。要记住的一件事是学习的行为本身和需要的时间:理论上,我们可以允许代理程序用非常非常长的时间来学习;然而,Q-Learning另一个目的是*将没有习得行为的试验试验变为有习得行为的行动*。例如,训练中总让代理程序执行随机动作(如果$\epsilon = 1$并且永不衰减)当然可以使它*学习*,但是不会让它*行动*。当改进你的Q-Learning实现时,要考虑做一个特定的调整的意义,以及它是否逻辑上是否合理。
### 改进Q-Learning的模拟结果
要从最初的Q-learning程序获得结果,你需要调整下面的标志和设置:
- `'enforce_deadline'` - 将此标志设定为`True`来强制驾驶代理程序捕获它是否在合理时间内到达目的地。
- `'update_delay'` - 将此标志设定为较小数值(比如`0.01`)来减少每次试验中每步之间的时间。
- `'log_metrics'` - 将此标志设定为`True`将模拟结果记录为在`/logs/`目录下的`.csv`文件,Q-table存为`.txt`文件。
- `'learning'` - 将此标志设定为`'True'`来告诉驾驶代理使用你的Q-Learning实现。
- `'optimized'` - 将此标志设定为`'True'`来告诉驾驶代理你在执行一个优化版本的Q-Learning实现。
优化Q-Learning代理程序可以调整的额外的标志:
- `'n_test'` - 将此标志设定为某个正数(之前是10)来执行那么多次测试试验。
- `'alpha'` - 将此标志设定为0 - 1之间的实数来调整Q-Learning算法的学习率。
- `'epsilon'` - 将此标志设定为0 - 1之间的实数来调整Q-Learning算法的起始探索因子。
- `'tolerance'` - 将此标志设定为某个较小的大于0的值(默认是0.05)来设定测试的epsilon阈值。
此外,使用一个你选择的$\epsilon$ (探索因子)衰减函数。注意无论你用哪个函数,**一定要以合理的速率衰减**到`'tolerance'`。Q-Learning代理程序到此才可以开始测试。某个衰减函数的例子($t$是试验的数目):
$$ \epsilon = a^t, \textrm{for } 0 < a < 1 \hspace{50px}\epsilon = \frac{1}{t^2}\hspace{50px}\epsilon = e^{-at}, \textrm{for } 0 < a < 1 \hspace{50px} \epsilon = \cos(at), \textrm{for } 0 < a < 1$$
如果你想的话,你也可以使用$\alpha$ (学习率) 的衰减函数,当然这通常比较少见。如果你这么做了,确保它满足不等式$0 \leq \alpha \leq 1$。
如果你在实施时遇到困难,尝试把`'verbose'`标志设为`True`来调试。调试时,在这里设定的标志会返回到它们的默认设定。重要的是你要理解每个标志做什么并且解释它们怎么影响模拟!
当你成功完成初始的Q-Learning模拟程序后,运行下面代码单元格来使结果可视化,请注意为了达到项目要求你需要在安全性和可靠性上获得至少都为A的评分。注意当相同的模拟运行时,log文件会被覆写,所以要留意载入的log文件!
```
# Load the 'sim_improved-learni ng' file from the improved Q-Learning simulation
vs.plot_trials('sim_improved-learning.csv')
```
### 问题7
利用上面的从你改进的Q-Learning模拟中得到的可视化结果,像在**问题6**那样,给出关于改进的驾驶代理程序的最终分析和观察。你需要回答的问题:
- *使用了什么epsilon(探索因子)的衰减函数?*
- *在测试之前驾驶代理大约需要做多少训练试验?*
- *你用了什么epsilon-tolerance和alpha(学习率)值?为什么?*
- *与之前的默认Q-Learning学习器相比,这个Q-Learning学习器有多少改进? *
- *你会说Q-Learning学习器的结果表明了你的驾驶代理程序成功地学习了一个合适的策略吗?*
- *你对*智能出租车*的安全性和可靠性评分满意吗?*
**回答:**
- 我使用了$$ \epsilon = \epsilon - 0.002 $$ 这个衰减函数
- 在测试之前,代理驾驶大约需要做500次训练实验
- epsilon为1,tolerance为0.002,alpha我也采用了衰减方式,初始值是0.95,每次衰减0.0015,线性衰减。
- 因为epsilon会根据衰减函数逐渐衰减,所以调整tolerance的值会影响训练次数,太少的训练不足以覆盖所有状态的Q值,而且学习力度不够,所以我选择0.002,大概训练500次,这样智能小车才能学到合适的决策。
- alpha的取值在0-1之间,alpha越大则新的尝试对于Q值更重要,而alpha越小则说明以前的经验对于Q值最为重要,因为之前的审阅说alpha也应该是逐渐衰减的,我认为有道理,在前期需要探索的时候,新的尝试对于Q值作用更大,而在后面,智能车已经学到了很多,所以经验会比较重要。
- 与之前的相比,这学习器有很大的改进,两个分数分别达到了双A+
- 我任务这个学习器的结果表明了这个驾驶代理程序成功的学习了一个合适的策略
- 我对智能出租车的安全性和可靠性评分满意
### 定义一个最优策略
有时,对于重要的问题*“我要让我的代理程序学习什么?”*的答案,只是理论性的,无法具体描述。然而这里,你可以具体定义代理程序要学什么,就是美国通行权交通法案。这些法律是已知信息,你可以基于这些法律,为*智能出租车*进一步定义每一个状态所做的最优动作。在那种情况下,我们称这一系列最优状态-动作配对为**最优策略**。因此,不像那些理论性的回答,不仅通过收到的奖励(惩罚),而且纯观察,代理程序是否在“错误”地行动能很清晰地得知。如果代理程序闯了红灯,我们既看见它获得了一个负面奖励,也知道这是一个错误的行为。这可以用来帮你验证驾驶代理程序习得的**策略**是否正确,或只是个**次优策略**。
### 问题 8
给出几个关于最优策略是什么样子的例子(用你已定义的状态)。之后,查看`'sim_improved-learning.txt'`文本文件,看你的改进的Q-Learning算法的结果。_每个从模拟中纪录的状态,对于给定的状态,**策略**(得分最高的动作)是否正确?是否对于有些状态,有策略不同于预期的最优策略?_给出一个状态和记录的状态-动作的奖励,解释为什么正是个正确的策略。
**回答:**
- 最有策略就是在当前状态下,做出最正确的决策,比如红灯的时候不能前进和转弯,只能停止等待,而处于绿灯的情况下,停止不前则不是明智的选择,会造成交通拥堵,所以要根据不同的状态选择最正确的决策,对于此而言,最正确的就是不妨碍交通规则,并且快速到达目的地。
- 在已定义的状态中,('forward', 'green', 'forward', 'left', None),则应该选择前进,因为目的地是前方,绿灯,右边车辆需要左转弯,所以最好的选择就是左转弯或者前进,但是我们的智能车需要安全的快速的驶向目的地,所以最好的选择是前进,在Q值中最高的分别为前进(1.80)左转(1.14)和预期相符。
- 在已定义状态中,('right', 'red', 'forward', 'right', None),则应该选择停止等待,因为是红灯,不应该前进和转弯,在Q值中,只有None是正值2.17,其中前进的惩罚为-34.64。
- 在已定义状态中,('left', 'green', None, 'forward', 'left'),则应选择左转,因为是绿灯,并且目的地方向在左边,而前方车辆和左右方车辆都没有干扰,所以应该选择左转在保证安全的前提下以最快的速度驶向目的地。在Q值中最高分为左转(1.42),和预期相符
-----
### 选做:未来奖励 - 折扣因子 `'gamma'`
也许你会好奇,作为Q-Learning算法的一部分,之前要求你在实现中**不要**使用折扣引子`'gamma'`。在算法中包含未来奖励能有助于在未来状态回溯到当前状态时的反向正面奖励。本质上,如果给予驾驶代理程序执行若干动作到达不同状态的选择,包含未来奖励会是代理程序偏向可以得到更多奖励的状态。一个例子是驶向目的的驾驶代理程序:所有行动和奖励都相等,那么理论上如果到达目的地会有额外奖励,驶向目的会获得更好的奖励。然而,即使在这个项目里,驾驶代理程序也要在规定的时间里到达目的地,包含未来奖励不会有益于代理程序。实际上,如果代理程序给予多次试验学习,它甚至会给Q-value带来负面影响!
### 可选问题 9
*在项目中有两个特点使得未来奖励在这个Q-Learning算法无效。一个特点是关于*智能出租车*本身,另一个是关于环境。你能指出它们是什么以及为什么未来奖励不会在这个项目中起效?*
**回答:**
- 我认为一个原因是因为起点和终点不固定,智能小车并不知道重点距离当前位置还有多远,只知道一个大概方向,所以未来奖励很难传播到起点
- 第二个是有规定的时间限制,这样未来奖励很难会让小车做出正确选择,因为距离终点的距离不定,时间限制也不一样,所以未来奖励不会对小车产生很大的益处
> **注意**:当你写完了所有的代码,并且回答了所有的问题。你就可以把你的 iPython Notebook 导出成 HTML 文件。你可以在菜单栏,这样导出**File -> Download as -> HTML (.html)**把这个 HTML 和这个 iPython notebook 一起做为你的作业提交。
| github_jupyter |
# Putting It All Together: A Realistic Example
In this section we're going to work through a realistic example of a deep learning workflow. We'll be working with a smallish dataset featuring different kinds of flowers from Kaggle. We're going to apply data augmentation to synthetically expand the size of our dataset. And we'll attempt transfer learning using networks pretrained on the ImageNet dataset, which includes some flower species already.
```
# All of this should look familiar from previous notebooks:
import matplotlib.pyplot as plt
import numpy as np
import os
from PIL import Image, ImageOps
from tensorflow.keras.applications.mobilenet_v2 import MobileNetV2, preprocess_input
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D, Dropout
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.utils import to_categorical
# Our function to load an image from a path and fix it up, but
# modified slightly to accomodate MobileNetV2.
def load_maintain_aspect_ratio(input_image_path, target_size):
image = Image.open(input_image_path)
width, height = image.size
w_pad = 0
h_pad = 0
bonus_h_pad = 0
bonus_w_pad = 0
if width > height:
pix_diff = (width - height)
h_pad = pix_diff // 2
bonus_h_pad = pix_diff % 2 # If the difference was odd, add one pixel on one side.
elif height > width:
pix_diff = (height - width)
w_pad = pix_diff // 2
bonus_w_pad = pix_diff % 2 # If the difference was odd, add one pixel on one side.
# else: image is already square. Both pads stay 0
image = ImageOps.expand(image, (w_pad, h_pad, w_pad+bonus_w_pad, h_pad+bonus_h_pad))
image = image.resize(target_size)
# Get the image data as a numpy array.
image_data = np.array(image.getdata()).reshape(image.size[0], image.size[1], 3)
# The preprocess function from MobileNetV2
# It expects a numpy array with RGB values between 0-255
return preprocess_input(image_data)
# Our recurring plot function.
def plot_training_history(history, model):
figure = plt.figure()
plt.subplot(1, 2, 1)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.tight_layout()
plt.subplot(1, 2, 2)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.tight_layout()
figure.tight_layout()
plt.show()
# Some constants:
# 224x224 is MobileNetV2's default, so lets stick with it
image_size = 224
batch_size = 32
validation_split = 0.2
# Start small, for the sake of learning speed.
num_epochs = 5
# The dataset is too large to reasonably redistribute as part of this repository, so you will
# have to download it separately from: https://www.kaggle.com/alxmamaev/flowers-recognition/
# The download as a flowers folder, this variable should point to the
# loccation of that folder. Inside that folder there should be 5 folders
# each named for the type of flower.
flower_dataset_directory = 'flowers_dataset/flowers/'
# The image classes
classes = {
'daisy': 0,
'dandelion': 1,
'rose': 2,
'sunflower': 3,
'tulip': 4
}
# Process all of the images into an array
images = []
labels = []
for subdir in classes.keys():
current_location = os.path.join(flower_dataset_directory, subdir)
print(f'Processing {subdir}')
sub_dir_count = 0
for file in os.listdir(current_location):
try:
image = load_maintain_aspect_ratio(os.path.join(current_location, file), (image_size, image_size))
images.append(image)
labels.append(classes[subdir])
sub_dir_count += 1
except:
print(f'Failed to load image: {subdir}/{file}. Ignored it.')
print(f'Found {sub_dir_count} images of type {subdir}')
# Just double check.
assert len(images) == len(labels)
# This is a little bit crude, but we'll just randomly select each image/label pair
# to be in the validation set based on our validation split. We could take greater
# care here to ensure that the right amount are represented from each class
# but it will probably be okay...
x_train = []
y_train = []
x_validation = []
y_validation = []
for image, label in zip(images, labels):
if np.random.random() > validation_split:
x_train.append(image)
y_train.append(label)
else:
x_validation.append(image)
y_validation.append(label)
# Properly format the images into a np array
x_train = np.array(x_train)
x_validation = np.array(x_validation)
# Make the labels one-hot encoded:
y_train = to_categorical(y_train, len(classes))
y_validation = to_categorical(y_validation, len(classes))
print(f'Loaded {len(images)}')
print(f'Training size: {len(x_train)}, validation size: {len(x_validation)}')
# Lots of possible augmentation to the training data
# hopefully allowing us to avoid overfitting
train_generator = ImageDataGenerator(
rotation_range=360,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=10,
zoom_range=0.5,
fill_mode='constant',
cval=0.0
)
# Don't transform the validation images.
validation_generator = ImageDataGenerator()
# Fit them both, best practice
train_generator.fit(x_train)
validation_generator.fit(x_validation)
# Lets do some sanity checking:
print(x_train.shape)
print(x_validation.shape)
print(y_train.shape)
print(y_validation.shape)
# View a couple, validations are never augmented
for _ in range(3):
plt.imshow(next(validation_generator.flow(x_validation))[0])
plt.show()
# But training data is
for _ in range(3):
plt.imshow(next(train_generator.flow(x_train))[0])
plt.show()
# Loading our pretrained mobilenet
base_model = MobileNetV2(weights='imagenet', include_top=False, input_shape=(image_size, image_size, 3))
# Make a very simple new classifier
old_top = base_model.output
old_top = GlobalAveragePooling2D()(old_top)
new_top = Dense(len(classes), activation='softmax')(old_top)
model = Model(inputs=base_model.input, outputs=new_top)
# We have a small amount of data, but the data is pretty similar
# to imagenet, which does train on many flower images, so we can
# expect the existing weights to be pretty good. Freeze all.
for layer in base_model.layers:
layer.trainable = False
# Go for it!
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit_generator(train_generator.flow(x_train, y_train, batch_size=batch_size),
steps_per_epoch=len(x_train) // batch_size,
epochs=num_epochs,
validation_data=validation_generator.flow(x_validation, y_validation),
validation_steps=len(x_validation) // batch_size
)
plot_training_history(history, model)
# Things are going great! Lets unfreeze some of our model's layers and see if "forgetting"
# some of the stuff our network learned about dogs, buildings, and waterbottles can
# improve our results further...
# This number was chosen specifically for MobileNetV2, it is the
# start of the 15th block.
for layer in model.layers[134:]:
layer.trainable = True
# Recompile to ensure the layers get set to trainable
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit_generator(train_generator.flow(x_train, y_train, batch_size=batch_size),
steps_per_epoch=len(x_train) // batch_size,
epochs=num_epochs,
validation_data=validation_generator.flow(x_validation, y_validation),
validation_steps=len(x_validation) // batch_size
)
plot_training_history(history, model)
```
Right on, things still look okay. The dip in validation accuracy is definitely concerning though. After this maybe we could get an even bigger dataset and buy some cloud compute time to train the model for longer... But if we continue to see the validation accuracy decline (overfitting) then we NEED to try something to set it right. More data, more augmentation of the data would both be good ideas. We could also try an alternate model or adjust our classification layers — but that should probably be **in addition** to more data and more data.
```
# I ran this one overnight, just for fun, and to see if we continued to overfit
history = model.fit_generator(train_generator.flow(x_train, y_train, batch_size=batch_size),
steps_per_epoch=len(x_train) // batch_size,
epochs=30,
validation_data=validation_generator.flow(x_validation, y_validation),
validation_steps=len(x_validation) // batch_size
)
plot_training_history(history, model)
```
I love this result — we do see validation continue to diverge from training, implying we are overfitting even with the augmentation tactics. But we also see that validation is really unstable. Sometimes we're getting lucky, but some of the adjustments are hurting us. We probably need more data to improve much more.
| github_jupyter |
y-tunnus strings can be converted to the following formats via the `output_format` parameter:
* `compact`: only number strings without any seperators or whitespace, like "20774740"
* `standard`: y-tunnus strings with proper whitespace in the proper places, like "2077474-0"
Invalid parsing is handled with the `errors` parameter:
* `coerce` (default): invalid parsing will be set to NaN
* `ignore`: invalid parsing will return the input
* `raise`: invalid parsing will raise an exception
The following sections demonstrate the functionality of `clean_fi_ytunnus()` and `validate_fi_ytunnus()`.
### An example dataset containing y-tunnus strings
```
import pandas as pd
import numpy as np
df = pd.DataFrame(
{
"ytunnus": [
"20774740",
"2077474-1",
"51824753556",
"51 824 753 556",
"hello",
np.nan,
"NULL"
],
"address": [
"123 Pine Ave.",
"main st",
"1234 west main heights 57033",
"apt 1 789 s maple rd manhattan",
"robie house, 789 north main street",
"(staples center) 1111 S Figueroa St, Los Angeles",
"hello",
]
}
)
df
```
## 1. Default `clean_fi_ytunnus`
By default, `clean_fi_ytunnus` will clean ytunnus strings and output them in the standard format with proper separators.
```
from dataprep.clean import clean_fi_ytunnus
clean_fi_ytunnus(df, column = "ytunnus")
```
## 2. Output formats
This section demonstrates the output parameter.
### `standard` (default)
```
clean_fi_ytunnus(df, column = "ytunnus", output_format="standard")
```
### `compact`
```
clean_fi_ytunnus(df, column = "ytunnus", output_format="compact")
```
## 3. `inplace` parameter
This deletes the given column from the returned DataFrame.
A new column containing cleaned y-tunnus strings is added with a title in the format `"{original title}_clean"`.
```
clean_fi_ytunnus(df, column="ytunnus", inplace=True)
```
## 4. `errors` parameter
### `coerce` (default)
```
clean_fi_ytunnus(df, "ytunnus", errors="coerce")
```
### `ignore`
```
clean_fi_ytunnus(df, "ytunnus", errors="ignore")
```
## 4. `validate_fi_ytunnus()`
`validate_fi_ytunnus()` returns `True` when the input is a valid y-tunnus. Otherwise it returns `False`.
The input of `validate_fi_ytunnus()` can be a string, a Pandas DataSeries, a Dask DataSeries, a Pandas DataFrame and a dask DataFrame.
When the input is a string, a Pandas DataSeries or a Dask DataSeries, user doesn't need to specify a column name to be validated.
When the input is a Pandas DataFrame or a dask DataFrame, user can both specify or not specify a column name to be validated. If user specify the column name, `validate_fi_ytunnus()` only returns the validation result for the specified column. If user doesn't specify the column name, `validate_fi_ytunnus()` returns the validation result for the whole DataFrame.
```
from dataprep.clean import validate_fi_ytunnus
print(validate_fi_ytunnus("20774740"))
print(validate_fi_ytunnus("2077474-1"))
print(validate_fi_ytunnus("51824753556"))
print(validate_fi_ytunnus("51 824 753 556"))
print(validate_fi_ytunnus("hello"))
print(validate_fi_ytunnus(np.nan))
print(validate_fi_ytunnus("NULL"))
```
### Series
```
validate_fi_ytunnus(df["ytunnus"])
```
### DataFrame + Specify Column
```
validate_fi_ytunnus(df, column="ytunnus")
```
### Only DataFrame
```
validate_fi_ytunnus(df)
```
| github_jupyter |
# Rekall Tutorial: Empty Parking Space Detection
In this tutorial, you'll learn how to use Rekall's programming model to detect empty parking spaces in a fixed-angle camera feed of a parking lot -- using nothing more than the outputs of an off-the-shelf object detector!
You should complete this tutorial after the Cyclist Detection tutorial.
Again, let's start by importing Rekall and a few important classes:
```
%load_ext autoreload
%autoreload 2
from rekall import Interval, IntervalSet, IntervalSetMapping, Bounds3D
from rekall.predicates import *
```
We'll again provide some helpers to handle data loading and visualization. For more details about what's going on with these helpers, check out the data loading and visualization tutorial.
```
from empty_parking_space_tutorial_helpers import *
```
And now let's load up the pre-computed bounding box detections:
```
bboxes = get_maskrcnn_bboxes()
```
And visualize them:
```
visualize_helper([bboxes])
```
If you click on the second video, you can see that there are sometimes empty parking spaces in this parking lot. Our goal is to detect these by creating an `IntervalSetMapping` object that contains all the empty parking spaces in these videos.
# Task: detect all empty parking spaces
Your goal is to write a Rekall program to detect all empty parking spaces (visualized in the second timeline above).
We're starting with a `IntervalSetMapping` object, `bounding_boxes`, that contains detections from Mask R-CNN. The Intervals contain 3D bounds, and the payloads contain the class and the class score:
```
bboxes[0].get_intervals()[0]
```
The bounding boxes are sampled every thirty seconds (hence why the Interval above has time bounds of 0 to 30), and so are the ground truth annotations.
The goal is to write a query that detects all the **empty parking spaces** in our videos.
This task is inspired by [this Medium blog post](https://medium.com/@ageitgey/snagging-parking-spaces-with-mask-r-cnn-and-python-955f2231c400):
* They use an off-the-shelf object detector to detect cars (like what we have in `bboxes`)
* They take a timestamp where all the parking spots are full, and use car detections to get parking spots
* Then empty parking spots are just parking spots where there are no cars
We'll be following these steps in this tutorial!
# Step 1: Detect Parking Spaces
Before we detect empty parking spaces, we first need to detect parking spaces! Luckily, every parking space is filled at the beginning of the first video. We can use the **car detections** at the beginning of this video to construct an `IntervalSetMapping` object corresponding to parking spaces in both videos.
Let's begin by looking at the video ID's of our videos:
```
bboxes.keys()
```
Videos are sorted by ID, so we know that the video on the left is video 0.
Let's start by getting all the car detections at the beginning of the video. We can reference the `IntervalSet` corresponding to video `0` as follows:
```Python
bboxes[0]
```
And filter down to the car detections at the beginning of video `0` like this:
```Python
parking_spot_candidates = bboxes[0].filter(
lambda intrvl: intrvl['t1'] == 0.0 and intrvl['payload']['class'] == 'car'
)
```
Try it yourself below!
```
# Construct parking spot candidates!
parking_spot_candidates = bboxes[0].filter(
lambda intrvl: intrvl['t1'] == 0.0 and intrvl['payload']['class'] == 'car'
)
```
`parking_spot_candidates` contains all the car detections at time `0` of video `0`. Next, we want to create `parking_spots`, an `IntervalSetMapping` object that represents all the parking spots (empty or not) in both of our videos.
We'll need to create a new `IntervalSetMapping` object called `parking_spots` that contains:
* Two keys (`0` and `2`), each of which points to an `IntervalSet` that contains parking spots
* Each `IntervalSet` should have `Intervals` for each parking spot in the parking lot
* Since our object detections are sampled once every 30 seconds, we can sample the parking spot objects once every thirty seconds as well
Go ahead and give this a try now! This is very similar to the checkpoint exercise in the cyclist detection tutorial. Solution is below.
```
# Construct parking spots!
video_lengths = {
key: bboxes[key].get_intervals()[-1]['t2']
for key in bboxes.keys()
}
parking_spots = IntervalSetMapping({
key: IntervalSet([
Interval(Bounds3D(
t1 = t,
t2 = t + 30, # Make the interval last 30 seconds
x1 = parking_spot['x1'],
x2 = parking_spot['x2'],
y1 = parking_spot['y1'],
y2 = parking_spot['y2']
))
for parking_spot in parking_spot_candidates.get_intervals() # For each parking spot
for t in range(0, int(video_lengths[key]), 30)
])
for key in bboxes.keys()
})
```
Exercise solution:
```Python
video_lengths = {
key: bboxes[key].get_intervals()[-1]['t2']
for key in bboxes.keys()
}
parking_spots = IntervalSetMapping({
key: IntervalSet([
Interval(Bounds3D(
t1 = t,
t2 = t + 30, # Make the interval last 30 seconds
x1 = parking_spot['x1'],
x2 = parking_spot['x2'],
y1 = parking_spot['y1'],
y2 = parking_spot['y2']
))
for parking_spot in parking_spot_candidates.get_intervals() # For each parking spot
for t in range(0, int(video_lengths[key]), 30)
])
for key in bboxes.keys()
})
```
Let's take a moment to visualize `parking_spots` and make sure it looks right -- you should have bounding boxes over each parking spot for the duration of the video.
```
visualize_helper([parking_spots])
```
# Step 2: Detect Empty Parking Spaces using `minus`
Now we have `parking_spots`, which represents all the parking spots in our video feeds, and we want to detect the empty parking spots.
This is a great use of Rekall's [`minus`](https://rekallpy.readthedocs.io/en/latest/index.html#rekall.IntervalSet.minus) function:
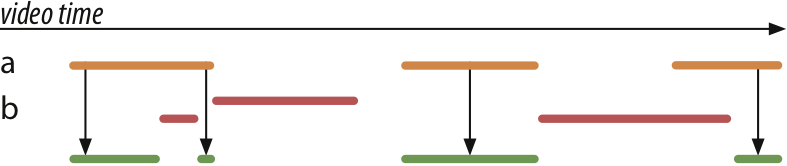
We can use the `minus` function to find all the instances where a parking spot does not have sufficient overlap with a car:
```Python
cars = bboxes.filter(
lambda intrvl: intrvl['payload']['class'] == 'car'
)
empty_parking_spots = parking_spots.minus(
cars,
predicate = and_pred(
Bounds3D.T(overlaps()),
Bounds3D.X(overlaps()),
Bounds3D.Y(overlaps()),
iou_at_least(0.25)
),
window=0.0,
progress_bar=True
)
```
This code takes any parking spot detection that overlaps with a detected car Interval with [IOU](https://www.pyimagesearch.com/2016/11/07/intersection-over-union-iou-for-object-detection/) at least 0.25, and removes it from the set. The remaining Intervals are empty parking spots.
Try it yourself below (it'll take about ten seconds)!
```
# Detect empty parking spots
cars = bboxes.filter(
lambda intrvl: intrvl['payload']['class'] == 'car'
)
empty_parking_spots = parking_spots.minus(
cars,
predicate = and_pred(
Bounds3D.T(overlaps()),
Bounds3D.X(overlaps()),
Bounds3D.Y(overlaps()),
iou_at_least(0.25)
),
window=0.0,
progress_bar=True
)
```
And let's visualize the results:
```
visualize_helper([empty_parking_spots])
```
# Step 3: Remove False Positives with `coalesce` and `filter_size`
So this is pretty good! But we have some false positives due to errors in the object detector; sometimes the cars are not detected. This results in parking spots that appear for 30 seconds at a time before disappearing. Let's remove some of these false positives by removing any parking spots that appear for less than a few minutes!
This is a great chance to use Rekall's [`coalesce`](https://rekallpy.readthedocs.io/en/latest/index.html#rekall.IntervalSet.coalesce) function:
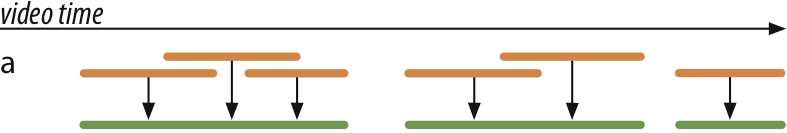
We'll use the `coalesce` function to merge all parking spots that are adjacent in time and overlapping in space:
```Python
empty_parking_spots_merged = empty_parking_spots.coalesce(
('t1', 't2'),
bounds_merge_op = Bounds3D.span,
predicate = iou_at_least(0.5)
)
```
Let's break down what's happening here:
* `coalesce` recursively merges all overlapping or touching Intervals along some axis (in this case, time).
* We merge the bounds by taking the span of the existing Intervals and the new Interval.
* We add in a predicate that we should only merge in new intervals if the IOU is at least 0.5.
* At the end, we have a Interval that covers the entire contiguous spacetime volume of an empty parking spot.
Then we can filter the resulting set by length to remove any parking spots that appear for less than four minutes:
```Python
empty_parking_spots_filtered = empty_parking_spots_merged.filter_size(
min_size = 60 * 4
)
```
Try it yourself below!
```
empty_parking_spots_merged = empty_parking_spots.coalesce(
('t1', 't2'),
bounds_merge_op = Bounds3D.span,
predicate = iou_at_least(0.5)
)
empty_parking_spots_filtered = empty_parking_spots_merged.filter_size(
min_size = 60 * 4
)
visualize_helper([empty_parking_spots_filtered])
```
This turns out to do pretty well! We have completely removed all the false detections from the first video (hence why it doesn't appear anymore), and we've removed many of the false detections from the second video.
# Step 4: Continue Debugging On Your Own
Now it's your turn! Can you modify the above code to get rid of even more false positives? Here's a few hints:
* We've fixed some issues with objects not being detected using the `coalesce` and `filter_size` functions, but what other issues might the object detector have?
* In particular, are there cases where a car could be mis-classified as something else?
* Try visualizing object classes like people, cars, and trucks to see what could be going on...
* This task was discussed in the [Rekall tech report](http://www.danfu.org/projects/rekall-tech-report/) (with pseudocode), so take a look at that for our solution!
```
# Play around on your own!
```
# Congratulations!
You've now written a pretty complex Rekall query to detect empty parking spots. If you haven't already, check out the [Rekall tech report](http://www.danfu.org/projects/rekall-tech-report/) to read about some of the other cool things we've been able to do with Rekall.
Next, check out the data loading and visualization tutorial to get more familiar with the nitty-gritty of how to visualize data with Rekall and Vgrid.
| github_jupyter |
Copyright 2018 The Dopamine Authors.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
https://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# Dopamine: How to create and train a custom agent
This colab demonstrates how to create a variant of a provided agent (Example 1) and how to create a new agent from
scratch (Example 2).
Run all the cells below in order.
```
# @title Install necessary packages.
!pip install --upgrade --no-cache-dir dopamine-rl
!pip install cmake
!pip install atari_py
!pip install gin-config
# @title Necessary imports and globals.
import numpy as np
import os
from dopamine.agents.dqn import dqn_agent
from dopamine.discrete_domains import run_experiment
from dopamine.colab import utils as colab_utils
from absl import flags
import gin.tf
BASE_PATH = '/tmp/colab_dope_run' # @param
GAME = 'Asterix' # @param
# @title Load baseline data
!gsutil -q -m cp -R gs://download-dopamine-rl/preprocessed-benchmarks/* /content/
experimental_data = colab_utils.load_baselines('/content')
```
## Example 1: Train a modified version of DQN
Asterix is one of the standard agents provided with Dopamine.
The purpose of this example is to demonstrate how one can modify an existing agent. The modification
we are doing here (choosing actions randomly) is for illustrative purposes: it will clearly perform very
poorly.
```
# @title Create an agent based on DQN, but choosing actions randomly.
LOG_PATH = os.path.join(BASE_PATH, 'random_dqn', GAME)
class MyRandomDQNAgent(dqn_agent.DQNAgent):
def __init__(self, sess, num_actions):
"""This maintains all the DQN default argument values."""
super(MyRandomDQNAgent, self).__init__(sess, num_actions)
def step(self, reward, observation):
"""Calls the step function of the parent class, but returns a random action.
"""
_ = super(MyRandomDQNAgent, self).step(reward, observation)
return np.random.randint(self.num_actions)
def create_random_dqn_agent(sess, environment, summary_writer=None):
"""The Runner class will expect a function of this type to create an agent."""
return MyRandomDQNAgent(sess, num_actions=environment.action_space.n)
random_dqn_config = """
import dopamine.discrete_domains.atari_lib
import dopamine.discrete_domains.run_experiment
atari_lib.create_atari_environment.game_name = '{}'
atari_lib.create_atari_environment.sticky_actions = True
run_experiment.Runner.num_iterations = 200
run_experiment.Runner.training_steps = 10
run_experiment.Runner.max_steps_per_episode = 100
""".format(GAME)
gin.parse_config(random_dqn_config, skip_unknown=False)
# Create the runner class with this agent. We use very small numbers of steps
# to terminate quickly, as this is mostly meant for demonstrating how one can
# use the framework.
random_dqn_runner = run_experiment.TrainRunner(LOG_PATH, create_random_dqn_agent)
# @title Train MyRandomDQNAgent.
print('Will train agent, please be patient, may be a while...')
random_dqn_runner.run_experiment()
print('Done training!')
# @title Load the training logs.
random_dqn_data = colab_utils.read_experiment(
LOG_PATH, verbose=True, summary_keys=['train_episode_returns'])
random_dqn_data['agent'] = 'MyRandomDQN'
random_dqn_data['run_number'] = 1
experimental_data[GAME] = experimental_data[GAME].merge(random_dqn_data,
how='outer')
# @title Plot training results.
import seaborn as sns
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(16,8))
sns.tsplot(data=experimental_data[GAME], time='iteration', unit='run_number',
condition='agent', value='train_episode_returns', ax=ax)
plt.title(GAME)
plt.show()
```
## Example 2: Train an agent built from scratch.
The purpose of this example is to demonstrate how one can create an agent from scratch. The agent
created here is meant to demonstrate the bare minimum functionality that is expected from agents. It is
selecting actions in a very suboptimal way, so it will clearly do poorly.
```
# @title Create a completely new agent from scratch.
LOG_PATH = os.path.join(BASE_PATH, 'sticky_agent', GAME)
class StickyAgent(object):
"""This agent randomly selects an action and sticks to it. It will change
actions with probability switch_prob."""
def __init__(self, sess, num_actions, switch_prob=0.1):
self._sess = sess
self._num_actions = num_actions
self._switch_prob = switch_prob
self._last_action = np.random.randint(num_actions)
self.eval_mode = False
def _choose_action(self):
if np.random.random() <= self._switch_prob:
self._last_action = np.random.randint(self._num_actions)
return self._last_action
def bundle_and_checkpoint(self, unused_checkpoint_dir, unused_iteration):
pass
def unbundle(self, unused_checkpoint_dir, unused_checkpoint_version,
unused_data):
pass
def begin_episode(self, unused_observation):
return self._choose_action()
def end_episode(self, unused_reward):
pass
def step(self, reward, observation):
return self._choose_action()
def create_sticky_agent(sess, environment, summary_writer=None):
"""The Runner class will expect a function of this type to create an agent."""
return StickyAgent(sess, num_actions=environment.action_space.n,
switch_prob=0.2)
sticky_config = """
import dopamine.discrete_domains.atari_lib
import dopamine.discrete_domains.run_experiment
atari_lib.create_atari_environment.game_name = '{}'
atari_lib.create_atari_environment.sticky_actions = True
run_experiment.Runner.num_iterations = 200
run_experiment.Runner.training_steps = 10
run_experiment.Runner.max_steps_per_episode = 100
""".format(GAME)
gin.parse_config(sticky_config, skip_unknown=False)
# Create the runner class with this agent. We use very small numbers of steps
# to terminate quickly, as this is mostly meant for demonstrating how one can
# use the framework.
sticky_runner = run_experiment.TrainRunner(LOG_PATH, create_sticky_agent)
# @title Train StickyAgent.
print('Will train sticky agent, please be patient, may be a while...')
sticky_runner.run_experiment()
print('Done training!')
# @title Load the training logs.
sticky_data = colab_utils.read_experiment(
LOG_PATH, verbose=True, summary_keys=['train_episode_returns'])
sticky_data['agent'] = 'StickyAgent'
sticky_data['run_number'] = 1
experimental_data[GAME] = experimental_data[GAME].merge(sticky_data,
how='outer')
# @title Plot training results.
import seaborn as sns
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(16,8))
sns.tsplot(data=experimental_data[GAME], time='iteration', unit='run_number',
condition='agent', value='train_episode_returns', ax=ax)
plt.title(GAME)
plt.show()
```
| github_jupyter |
# Introduction to Python
Guillaume Lemaitre
## Basic usage
### An interprated language
Python is an interpreted language. Each line of code is evaluated.
```
print('Hello world')
x = 20
```
The previous cell call the funcion `print` which will return the parameter which pass to it. This function is directly evaluated by the Python interpreter without the need of an extra step.
```
print(x)
```
### An untyped language
There is no need to specify the type of variables in Python
```
# This is an example of C++ declaration.
# Note that it will fail during the execution of the cell.
# We are programming in Python!!!
int a = 10;
```
Python will infer the appropriate type.
```
x = 2
type(x)
```
We can first try to check which built-in types Python offers.
```
x = 2
type(x)
x = 2.0
type(x)
x = 'two'
type(x)
x = True
type(x)
x = False
type(x)
```
`True` and `False` are booleans. In addition, these types can be obtained when making some comparison.
```
x = (3 > 4)
type(x)
x
```
### Python as a calculator
Python provides some built-in operators as in other languages.
```
2 * 3
2 / 3
2 + 3
2 - 3
```
As previously mentioned, Python will infer the most appropriate data type when doing the operation.
```
type(2 * 3)
type(2 * 3.0)
type(2 / 3)
```
Other useful operators are availble and differ from other languages.
```
3 % 2
3 // 2
3 ** 2
```
### Python to make some logic operations
Python provides some common logic oprators `and`, `or`, `not`. & / | / ~
Let's look at the Carnaugh table of the `and` operator.
```
True and True
True and False
False and True
False and False
```
### Exercise:
* Check the Carnaugh table for the `or` operator and spot the difference.
`not` will inverse the boolean value.
```
not True
```
Be aware that `not` with other type then boolean.
An empty string `''`, `0`, `False`, will be interpretated as `False` when doing some boolean operation. We will see later what are lists, but an empty list `[]` will also be interpretated as `False`.
```
bool(0)
bool('')
bool([])
```
In the same manner, non-zero numbers, non-empty list or string will be interpreted as `True` in logical operations.
```
bool(1)
bool(50)
bool('xxx')
bool([1, 2, 3])
```
## The standard library
### The example of the `math` module
Up to now, we saw that Python allows to make some simple operation. What if you want to make some advance operations, e.g. compute a cosine.
```
cos(2 * pi)
```
These functionalities are organised into differnt **modules** from which you have to first import them before to use them.
```
import math
math.cos(2 * math.pi)
```
The main question is how to we find out which module to use and which function to use. The answer is the Python documentation:
* The Python Language Reference: http://docs.python.org/3/reference/index.html
* The Python Standard Library: http://docs.python.org/3/library/
Never try to reinvente the wheel by coding your own sorting algorithm (apart of of didactic reason). Most of what you need are already efficiently implemented. If you don't know where to search in the Python documentation, Google it, Bing it, Yahoo it (this will not work).
In matlab, you are used to have the function in the main namespace. You can have something similar in Python.
```
from math import cos, pi
cos(2 * pi)
```
Python allows to use `alias` during import to avoid name collision.
```
import math
import numpy
```
Both package provide an implementation of `cos`
```
math.cos(1)
numpy.cos(1)
```
However, the NumPy implementation support transforming several values at one.
```
math.cos([1, 2])
numpy.cos([1, 2])
```
One issue with name collision would have happen if we would have import the `cos` function directly from each package or module.
### Exercise:
* import `cos` directly from `numpy` and `math` and check which function will be used if you call `cos`. You might want to use `type(cos)` to guess which function will be used. Deduce how the importing mechanism works.
What if you need to find the documentation and that Google is broken or you simply don't have internet. You can use the `help` function.
```
import math
help(math)
```
This command will just give you the same documentation than the one you have on internet. The only issue is that it could be less readable. If you are using `ipython` or `jupyter notebook`, you can use the `?` or `??` magic functions.
```
math.log?
math.log??
```
### Exercise:
* Write a small code to compute the volume of a sphere of radius 2.5. Round the results after the second digits.
### Other modules which are in the standard library
There is more than the `math` module. You can interact with the system, make regular expression, etc: `os`, `sys`, `math`, `shutil`, `re`, etc.
Refer to https://docs.python.org/3/library/ for a full list of the available tools.
## Containers: strings, lists, tuples, set (let's skip it), dictionary
### Strings
We already introduce the string but we give an example again.
```
s = 'Hello world!'
s
type(s)
```
A string can be seen as a table of characters. Therefore, we can actually get an element from the string. Let's take the first element.
```
s[0]
```
As in some other languages, the indexing start at 0 in Python. Unlike other language, you can easily iterate backward using negative indexing.
```
s[-1]
```
#### `slice` function
If you come from Matlab you already are aware of the slicing function, e.g. `start:end:step`. Let see the full story how does it works in Python.
The idea of slicing is to take a part of the data, with a regular structure. This structure is defined by: (i) the start of the slice (the starting index), (ii) the end of the slice (the ending index), and (iii) the step to take to go from the start to the end. In Python, the function used is called `slice`.
```
type(slice)
help(slice)
s
```
So I can select a sub-string using this slice.
```
my_slice = slice(2, 7, 3)
s[my_slice]
```
What if I don't want to mention the `step`. Then, you can they that the step should be `None`.
```
my_slice = slice(2, 7, None)
s[my_slice]
```
Similar thing for the `start` or `end`.
```
s[slice(None, 7, None)]
my_slice = slice(7)
s[my_slice]
```
However, this syntax is a bit long and we can use the well-known `[start:end:step]` instead.
```
s[2:7:2]
```
Similarly, we can use `None`.
```
s[None:7:None]
```
Since `None` mean nothing, we can even remove it.
```
s[:7:]
```
And if the last `:` are followed by nothing, we can even skip them.
```
s[:7]
```
Now, you know why the slice has this syntax.
**Be aware**: Be aware that the `stop` index is not including within your data which sliced.
```
s[2:]
```
The third character (index 2) is discarded. Why so? Because:
```
start = 0
end = 2
print((end - start) == len(s[start:end]))
```
#### String manipulation
We already saw that we can easily print anything using the `print` function.
```
print(10)
```
This `print` function can even take care about converting into the string format some variables or values.
```
print("str", 10, 2.0)
```
Sometimes, we are interested to add the value of a variable in a string. There is several way to do that. Let's start with the old fashion way.
```
s = "val1 = %.2f, val2 = %d" % (3.1415, 1.5)
s
import math
s = "the number %s is equal to %s"
print(s % ("pi", math.pi))
print(s % ("e", math.exp(1.)))
```
But more recently, there is the `format` function to do such thing.
```
s = "Pi is equal to {:.2f} while e is equal to {}".format(
math.pi, math.e
)
print(s)
```
And in the future, you will use the format string.
```
s = f'Pi is equal to {math.pi} while e is equal to {math.e}'
print(s)
```
A previously mentioned, string is a container. Thus, it has some specific functions associated with it.
```
print("str1" + "str2" + "str2")
print("str1" * 3)
```
In addition, a string has is own methods. You can access them using the auto-completion using Tab after writing the name of the variable and a dot.
```
s = 'hello world'
s.ljust?
```
But we will comeback on this later on.
#### Exercise
* Write the following code with the shortest way that you think is the best:
`'Hello DSSP! Hello DSSP! Hello DSSP! Hello DSSP! Hello DSSP! GO GO GO!'`
```
str_1 = 'Hello DSSP! ' * 5
str_2 = 'GO ' * 3
print(repr(str_2))
print(str_1 + str_2[:-1] + '!')
```
### Lists
Lists are similar to strings. However, they can contain whatever types. The squared brackets are used to identified lists.
```
l = [1, 2, 3, 4]
l
type(l)
l = [1, '2', 3.0]
l
print(f'The element {l[0]} is of type {type(l[0])}')
print(f'The element {l[1]} is of type {type(l[1])}')
print(f'The element {l[2]} is of type {type(l[2])}')
l = [1, 2, 3, 4, 5]
```
We can use the same syntax to index and slice the lists.
```
l[0]
l[-1]
l[2:5:2]
```
### Exercise:
* A list is also a container. Therefore, we would expect the same behavior for `+` and `*` operators. Check the behavior of both operators.
#### Append, insert, modify, and delete elements
In addition, a list also have some specific methods. Let's use the auto-completion
```
l = []
len(l)
l.append("A")
l
len(l)
```
`append` is adding an element at the end of the list.
```
l.append("x")
l
l[-1]
```
`insert` will let you choose where to insert the element.
```
l.insert(1, 'c')
l
```
We did not try to modify an element from string before. We can check what would happen.
```
s
s[0] = "H"
s2 = s.capitalize()
s
```
So we call the `string` an immutable container since it cannot be changed.
What happens with a list?
```
l
l[1] = 2
l
```
A list is therefore mutable. We can change any element in the list. So we can also remove an element from it.
```
l.remove(2)
l2 = [1, 2, 3, 4, 5, 2]
l2.remove(2)
l2
l2.remove?
l
```
Or directly using an index.
```
del l[-1]
l
```
### Tuples
```
l = [1, 2, 3]
```
Tuple can be seen as an immutable list. The syntax used is `(values1, ...)`.
```
t = (1, 2, 3)
t
type(t)
```
### Exercise:
* Try to assign the value `0` to the first element of the tuple `t`.
However, tuples are not only used as such. They are mainly used for unpacking variable. For instance they are usually returned by function when there is several values.
We can easily unpack tuple with the associated number of variables.
```
x, y, z = (1, 2, 3)
x
y
z
out = (1, 2, 3)
out
x, y, z = out
x
y
z
```
### Dictionary
Dictionaries are used to map a key to a value. The syntax used is `{key1: value1, ...}`.
```
d = {
'param1': 1.0,
'param2': 2.0,
'param3': 3.0
}
d
type(d)
```
To access a value associated to a key, you index using the key:
```
d['param1']
```
Dictionaries are mutable. Thus, you can change the value associated to a key.
```
d['param1'] = 4.0
d
```
You can add a new key-value relationship in a dictionary.
```
d['param4'] = 5.0
d
```
And you can as well remove relationship.
```
del d['param4']
d
```
You can also know if a key is inside the dictionary.
```
'param2' in d
```
You can also know about the key and values with the following methods:
```
d.keys()
d.values()
d.items()
```
It can allows to iterate:
```
keys = list(d.keys())
d[keys[0]]
d[keys[1]]
items = list(d.items())
items[0]
key, value = items[0]
print(f"key: {key} -> value: {value}")
```
### Built-in functions
Now that we introduced the `list` and `string`, we can check the so called built-in functions: https://docs.python.org/3/library/functions.html
These functions are a set of functions which are commonly used. For instance, we already presented the `slice` functions. From this list, we will present three functions: `in`, `range`, `enumerate`, and `sorted`. You can check the other functions later on.
#### `sorted` function
The sorted function will allow us to introduce the difference between inplace and copy operation. Let's take the following list:
```
l = [1, 5, 3, 4, 2]
from copy import copy
l_sorted = copy(l)
l_sorted.sort()
l = l_sorted
l
```
We can call the function `sorted` to sort the list.
```
sorted?
l_sorted = sorted(l)
l_sorted
```
We can observe that a sorted list is returned by the function. We can also check that the original list is actually unchanged:
```
l
```
It means that the `sorted` function made a copy of `l`, sorted it, and return us the result. The operation was not made inplace. However, we saw that a list is mutable. Therefore, it should be possible to make the operation inplace without making a copy. We can check the method of the list and we will see a method `sort`.
```
l.sort()
l
```
We see that this `sort` method did not return anything and that the list was changed inplace.
Thus, if the container is mutable, calling a method will try to do the operation inplace while calling the function will make a copy.
#### `range` function
It is sometimes handy to be able to generate number with regular interval (e.g. start:end:step).
```
range?
list(range(5, 10, 2))
```
#### `enumerate` function
The `enumerate` function allows to get the index associated with the element extracted from a container. Let see what we mean:
```
list(enumerate([5, 7, 9]))
enum = list(enumerate([5, 7, 9]))
indice, value = enum[0]
print(f"indice: {indice} -> value: {value}")
```
#### `in` function
The function `in` allows to know if a value is in the container.
```
l = [1, 2, 3, 4, 5]
5 in l
6 in l
s = 'Hello world'
'h' in s
'H' in s
s.find('e')
```
## Conditions and loop
### `if`, `elif`, and `else` condtions
Python delimates code block using indentation.
```
x = (1, 2, 3)
a = 3
b = 3
if a < b:
print('a is smaller than b')
print('xxxx')
elif a > b:
print('a is bigger than b')
else:
print('a is equal to b')
```
Be aware that if you do not indent properly your code, then you will get some nasty errors.
```
if True:
print('whatever')
print('wrong indentation')
```
### `for` loop
In Python you can get the element from a container.
```
for elt in [5, 7, 9]:
print(f'value: {elt}')
```
And if you wish to get the corresponding indices, you can always use `enumerate`.
```
for idx, elt in enumerate([5, 7, 9]):
print(f'idx: {idx} => value: {elt}')
```
You can have nested loop.
```
for word in ["calcul", "scientifique", "en", "python"]:
for letter in word:
if letter in ['c', 'e', 'i']:
continue
print(letter)
```
#### Exercise
* Count the number of occurences of each character in the string `'HelLo WorLd!!'`. Return a dictionary associating a letter to its number of occurences.
* Given the following encoding, encode the string `s`.
* Once the string encoded, decode it by inversing the dictionary.
```
code = {'e':'a', 'l':'m', 'o':'e', 'a': 'e'}
```
### `while` loop
If your loop should stop at a condition rather than using a number of iterations, you can use the `while` loop.
```
i = 0
while i < 5:
print(i)
i = i + 1
print("OK")
```
#### Exercise
* Code the Wallis formula to compute $\pi$:
$$
\pi = 2 \prod_{i=1}^{\infty} \frac{4 i^2}{4 i^2 - 1}
$$
## Fonctions
We already used functions above. So we will give a formal introduction. Fonction in Python are using the keyword `def` and define a list of parameters.
```
def func(x, y):
print(f'x={x}; y={y}')
x = func(1, 2)
print(x)
```
These parameters can be positional or use a default values.
```
def func(x, y, z=0):
print(f'x={x}; y={y}; z={z};')
func(1, 2)
func(1, 2, z=3)
func(1)
```
Functions can return one or more values. The output is a tuple if there is several values.
```
def square(x):
return x ** 2
x = square(2)
x
def square(x, y):
return x ** 2, y ** 2
square(2, 3)
x_2, y_2 = square(2, 3)
x_2
y_2
```
How does the documentation is working in Python.
```
help(square)
```
We can easily define what should be our inputs and outputs such that people can use our function documentation.
```
def square(x, y):
"""Square a pair of numbers.
Parameters
----------
x : real
First number.
y : real
Second number.
Returns
-------
squared_numbers : tuple of real
The squared x and y.
"""
return x ** 2, y ** 2
help(square)
```
## Classes
### Recognize classes
Here, we are only interesting to know about recognizing classes and use them. We will probably not have to program any.
A typical scikit-learn example:
```
from sklearn.datasets import load_iris
data, target = load_iris(return_X_y=True)
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(max_iter=1000).fit(data, target)
model.coef_
```
* `LogisticRegression` is a class: leading capital letter is a Python convention.
* `model` is an instance of the `LogisticRegression` class.
* `model` will have some methods (simply function belonging to the class) and attributes (simply variable belonging to the class).
* `fit` is a class method and `coef_` is an class attribute.
However, you already manipulated classes with Python built-in types.
```
mylist = [1, 2, 3, 4]
mylist.
```
### Program your own class
This introduction is taken from the scipy lecture notes:
https://scipy-lectures.org/intro/language/oop.html
Python supports object-oriented programming (OOP). The goals of OOP are:
* to organize the code, and
* to re-use code in similar contexts.
Here is a small example: we create a Student class, which is an object gathering several custom functions (methods) and variables (attributes), we will be able to use:
```
class Student:
def __init__(self, name):
self.name = name
def set_age(self, age):
self.age = age
def set_major(self, major):
self.major = major
anna = Student('anna')
anna.set_age(21)
anna.set_major('physics')
anna.
```
In the previous example, the Student class has `__init__`, `set_age` and `set_major` methods. Its attributes are `name`, `age` and `major`. We can call these methods and attributes with the following notation: `classinstance.method` or `classinstance.attribute`. The `__init__` constructor is a special method we call with: `MyClass(init parameters if any)`.
Now, suppose we want to create a new class MasterStudent with the same methods and attributes as the previous one, but with an additional `internship` attribute. We won’t copy the previous class, but **inherit** from it:
```
class MasterStudent(Student):
internship = 'mandatory, from March to June'
james = MasterStudent('james')
james.internship
james.set_age(23)
james.age
```
The MasterStudent class inherited from the Student attributes and methods.
Thanks to classes and object-oriented programming, we can organize code with different classes corresponding to different objects we encounter (an Experiment class, an Image class, a Flow class, etc.), with their own methods and attributes. Then we can use inheritance to consider variations around a base class and **re-use** code. Ex : from a Flow base class, we can create derived StokesFlow, TurbulentFlow, PotentialFlow, etc.
| github_jupyter |
# How can I upload a file?
### Overview
Here we introduce file upload via API. Specifically we will:
1. create a new project
2. check that there are no files
3. upload some files
4. set any metadata we like
5. search for the files via metadata
### Prerequisites
1. You need your _authentication token_ and the API needs to know about it. See <a href="Setup_API_environment.ipynb">**Setup_API_environment.ipynb**</a> for details.
3. You downloaded/cloned the whole repo so the files we will try to upload exist
## Imports
We import the _Api_ class from the official sevenbridges-python bindings below.
```
import sevenbridges as sbg
```
## Initialize the object
The `Api` object needs to know your **auth\_token** and the correct path. Here we assume you are using the credentials file in your home directory. For other options see <a href="Setup_API_environment.ipynb">Setup_API_environment.ipynb</a>
```
# [USER INPUT] specify credentials file profile {cgc, sbg, default}
prof = 'default'
config_file = sbg.Config(profile=prof)
api = sbg.Api(config=config_file)
```
## Create a shiny, new project
To avoid any copy-errors with the app, we will make a new project. If this project name already exists, the code below will raise an interupt and fail. Be _creative_ with your project names, it's something you will look back on and laugh.
#### PROTIPS
This next cell is more extensively detailed in this [recipe](projects_makeNew.ipynb)
```
# [USER INPUT] Set project name and billing group index here:
# Note that you can have multiple apps or projects with the same name. It is best practice to reference entities by ID.
new_project_name = 'Shiny and Newer'
index_billing = 0
# Check if this project already exists. LIST all projects and check for name match
my_project = api.projects.query(name=new_project_name)
if my_project: # exploit fact that empty list is False
print('A project named {} exists, please choose a unique name'
.format(new_project_name))
raise KeyboardInterrupt
else:
# Create a new project
# What are my funding sources?
billing_groups = api.billing_groups.query()
print((billing_groups[index_billing].name + \
' will be charged for computation and storage (if applicable)'))
# Set up the information for your new project
new_project = {
'billing_group': billing_groups[index_billing].id,
'description': """A project created by the API recipe (apps_installFromJSON).
This also supports **markdown**
_Pretty cool_, right?
""",
'name': new_project_name
}
my_project = api.projects.create(
name=new_project['name'], billing_group=new_project['billing_group'],
description=new_project['description']
)
# (re)list all projects, and get your new project
my_project = [p for p in api.projects.query(limit=100).all()
if p.name == new_project_name][0]
print('Your new project {} has been created.'.format(
my_project.name))
# Print description if it exists
if hasattr(my_project, 'description'):
print('Project description: \n {}'.format(my_project.description))
```
## Sanity-check: do I have any files?
Since you have just created the project, there will be **no** _Files_, _Apps_, or _Tasks_ in it. But just to be sure, let's query the apps in our project.
#### PROTIPS
This next cell is more extensively detailed in this [recipe](files_listAll.ipynb)
```
my_files = api.files.query(project = my_project)
print('In project {}, you have {} files.'.format(
my_project.name, my_files.total))
```
## Upload some toy files
Here we are using some of the recipes from the [ok, API](https://github.com/sbg/okAPI) repository. This **synchronous** upload will not return any information. Next, we set the _same metadata_ to all of the files (except one). What is really excellent about this **flexible metadata** is that it is searchable, you can use it to build tasks later. Furthermore, you can set **tags** via API. These will be visable on the GUI and via API.
#### Notes:
* The search by metadata function does **not** work with booleans or integers right now. This is a **known** bug so you **know** we are on it! However, I'm confident you will be able to do something clever like change True to 'True' or 1 to 'one' if you really need it
* Alternatively, and **orders of magnitude** more slowly, you could get the metadata of each file individually and search it (including booleans and integers) in Python. An example of that is [here](files_listByMetadata.ipynb)
```
# [USER INPUT] file names to upload:
file_list = ['files_listAll.ipynb',
'files_copyFromMyProject.ipynb',
'files_copyFromPublicReference.ipynb',
'files_detailOne.ipynb',
'files_upload_and_setMetadata.ipynb']
for f in file_list:
api.files.upload(project=my_project, path=f)
# List all files in the project
my_files = api.files.query(project=my_project)
print('In project {}, you have {} files.\n'.format(
my_project.name, my_files.total))
for f in my_files:
print(f.name)
# Set file metadata
base_md = {
'toy_example': False,
'extension': 'ipynb',
'revision_number': 7,
'Hello':'Nope!'
}
# We could go through each individual file and set metadata
# for f in my_files:
# f.metadata = base_md
# f.save()
# But note that this means one API request for each f.save()
# But it is much more efficient if it is done in bulk, we can update up to 100 files with one request
for f in my_files:
f.metadata = base_md
api.files.bulk_update(my_files)
# change one file's metadata to look for it later
f = my_files[2]
f.metadata['Hello'] = "is it me you're looking for?"
f.save()
# Also set a tag on that file
f.tags = ['example']
f.save()
# List files based on metadata
my_matched_files = api.files.query(
project=my_project,
metadata = {'Hello' : "is it me you're looking for?"}
)
print('In project {}, you have {} matching files.\n'.format(
my_project.name, my_matched_files.total))
for f in my_matched_files:
print("""File named ({}) matched search criteria.
File metadata is {}.
File tags are {}'"""
.format(f.name, f.metadata, f.tags))
```
## (optional) Upload real-sized files
Toy files are great, but are not going to rock the genomic world. What about _hundreds of Gb_? The API uploader deals with that pretty well too. However, the method above doesn't give any indications of progress, which is rather unsettling. So let's use **asynchronous uploads** with a **progress bar**.
_iPython_ is **not very reliable with printing to screen**, so I would recommend using this in _Python_. (in fact, the code below **does not work**). It is _especially unreliable_ in a for loop as the progress bar from the prior upload can interfere with the current one. So here we are only showing a _single file_ (which you need bring from your own files).
#### Note
By setting
``` python
wait=False
```
we are using _ayschronous_ uploading. This also means we need to **.start()** each upload when we are ready. This is different than the prior cell where we used _synchronous_ uploads which started automatically.
```
# [USER INPUT] file to upload:
file_name = 'heavy.sites.vcf' # TODO: Replace with your own large, local file
from sevenbridges.transfer.utils import simple_progress_bar
upload = api.files.upload(
path = file_name, project = my_project, wait=False
)
upload.status
upload.add_progress_callback(simple_progress_bar)
upload.status
upload.start()
```
## Additional Information
Detailed documentation of this particular REST architectural style request is available in this [section](http://docs.sevenbridges.com/docs/upload-files)
| github_jupyter |
# Starbucks Capstone Challenge
# Part 3 - Supervised Machine Learning to predict offer completed rate
In this notebooks contains
- Feature Preprosessing & transforming
- Supervised Machine Learning Model with regression
- Model evaluation
- Conclusion
## Problem Understanding
In this section, I use the clean profile with cluster as defined by part 2 to predict the offer completed rate. I try to aim how well we can predict customer response for each offer - bogo, discount, and informational. Using the model, I try to find another information about their demographics and relationship with the feature.
## Project Metrics :
A Supervised Machine learning using regression algorithm is used in this section.
The regression metrics are :
1. Mean Squared Error (MSE):
2. Coefficient of Determination (R^2)
## 1. Import Library
```
import pandas as pd
import numpy as np
import math
import json
import seaborn as sns
from tqdm import tqdm
import matplotlib.pyplot as plt
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
%load_ext autoreload
%autoreload 2
import helpers as h
```
## 2. Load Data
The clean profile data as a result of part 2 is saved in `data/main_cluster.csv`
```
main_cluster = pd.read_csv('data/main_cluster.csv')
main_cluster = main_cluster.set_index(main_cluster.columns[0])
#rename cluster name to m_1, m_2, etc
main_cluster['cluster'] = main_cluster['cluster'].apply(lambda x: 'm_'+ str(x))
main_cluster.shape
main_cluster.head()
# check for null data
main_cluster.isnull().sum().sum()
```
## 3. Features Preprocessing and Selection
Most of features have been preprocessed in part 1 and 2. So There is very few preprocessing required.
The targets for predictions are `rate_completed_received_bogo`, `rate_completed_received_discount`, `rate_viewed_informational`
The features that are related to the targets will be removed from the feature to avoid data leakage, so the remaining features are 46 features.
```
features = ['age',
'avg_spending',
'gender_F',
'gender_M',
'gender_O',
'income',
'informational_received',
'invalid',
'member_days_since',
'member_year_2013',
'member_year_2014',
'member_year_2015',
'member_year_2016',
'member_year_2017',
'member_year_2018',
'offer received_bogo_10_10_5',
'offer received_bogo_10_10_7',
'offer received_bogo_5_5_5',
'offer received_bogo_5_5_7',
'offer received_discount_10_2_10',
'offer received_discount_10_2_7',
'offer received_discount_20_5_10',
'offer received_discount_7_3_7',
'offer received_informational_0_0_3',
'offer received_informational_0_0_4',
'offer viewed_bogo_10_10_5',
'offer viewed_bogo_10_10_7',
'offer viewed_bogo_5_5_5',
'offer viewed_bogo_5_5_7',
'offer viewed_discount_10_2_10',
'offer viewed_discount_10_2_7',
'offer viewed_discount_20_5_10',
'offer viewed_discount_7_3_7',
'rate_offer_viewed_bogo_10_10_5',
'rate_offer_viewed_bogo_10_10_7',
'rate_offer_viewed_bogo_5_5_5',
'rate_offer_viewed_bogo_5_5_7',
'rate_offer_viewed_discount_10_2_10',
'rate_offer_viewed_discount_10_2_7',
'rate_offer_viewed_discount_20_5_10',
'rate_offer_viewed_discount_7_3_7',
'rate_viewed_bogo',
'rate_viewed_discount',
'sum_spending',
'transaction_count', 'cluster']
X = main_cluster[features]
target_cols = ['rate_completed_received_bogo',
'rate_completed_received_discount','rate_viewed_informational']
y = main_cluster[target_cols]
X.shape
```
## 4. Feature Transformations
### 4.1 One-hot encoding on cluster column
```
X_one_hot = pd.get_dummies(X)
def one_hot(df):
return pd.get_dummies(df)
X_one_hot.columns
```
### 4.2 Feature Scaling
```
"""
StandardScaler Procedure:
"""
from sklearn.preprocessing import StandardScaler, RobustScaler
def scaling(features):
scale = StandardScaler()
scale.fit(features)
features_std = pd.DataFrame(scale.transform(features), index= features.index, columns=features.columns)
return features_std
X_std = scaling(X_one_hot)
```
Feature scaling will be included in Model Pipeline
### 4.3 Split Data : Train & Test
```
def get_train_test(features, target):
""" Split : Train - Test """
from sklearn.model_selection import train_test_split as tts
X_train, X_test, y_train, y_test = tts(features, target, test_size=0.33, random_state=42)
return X_train, X_test, y_train, y_test
# as feature scaling will be included in model pipeline
# the data will be split after performing one hot encoding.
X_train, X_test, y_train, y_test = get_train_test(X_one_hot, y)
```
# 5. Supervised Machine Learning Model
## 5.1 Base Model
```
from sklearn.model_selection import cross_val_score, cross_validate
from sklearn.model_selection import KFold
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import FunctionTransformer
from sklearn.metrics import mean_squared_error
from sklearn.multioutput import MultiOutputRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.preprocessing import StandardScaler, RobustScaler, MinMaxScaler
from sklearn.linear_model import LinearRegression
seed=42
np.random.seed(seed)
estimators = [('standardize', StandardScaler()),
('reg', MultiOutputRegressor(LinearRegression()))]
pipeline = Pipeline(estimators)
kfold = KFold(n_splits=10, random_state=seed)
results = cross_validate(pipeline, X_train, y_train, cv=kfold, scoring=['r2','neg_mean_squared_error'])
r2_arr = results['test_r2']
mse_arr = results['test_neg_mean_squared_error']*-1
print(f"R^2: {r2_arr.mean():.4f} with stdev {r2_arr.std():.4f}")
print(f"MSE: {mse_arr.mean():.4f} with stdev {mse_arr.std():.4f}")
```
## 5.2 Refinement
#### 5.2.1 Spot Check Alogrithm
Finding the best alogrithm.
The code is a modified version to accomodate MultiOuputRegressor.
References : [here](https://machinelearningmastery.com/rescaling-data-for-machine-learning-in-python-with-scikit-learn/)
```
# get model list
models = h.get_models_multioutput()
# evaluate models
results = h.evaluate_models(X_train, y_train, models, metric='neg_mean_squared_error')
# summarize results
h.summarize_results(results)
```
The Random Forest Regressor and Bagging regressor are the best models that have the lowest MSE (highest negative MSE). However Random Forest has lower IQR (Interquantile Range). Therefore Random Forest Regressor is choosen as the main model for this data.
#### 5.2.2 The Selected Model - Random Forest Regressor
```
from sklearn.ensemble import RandomForestRegressor
seed=42
np.random.seed(seed)
estimators = [('standardize', StandardScaler()),
('reg', MultiOutputRegressor(RandomForestRegressor(random_state=seed)))]
pipeline = Pipeline(estimators)
kfold = KFold(n_splits=10, random_state=seed)
results = cross_validate(pipeline, X_train, y_train, cv=kfold, scoring=['r2','neg_mean_squared_error'])
r2_arr = results['test_r2']
mse_arr = results['test_neg_mean_squared_error']*-1
print(f"R^2: {r2_arr.mean():.4f} with stdev {r2_arr.std():.4f}")
print(f"MSE: {mse_arr.mean():.4f} with stdev {mse_arr.std():.4f}")
```
#### 5.2.3 Remove Scaler for easier to interprate the model as Random Forest is not sensitive to scaling process.
```
from sklearn.ensemble import RandomForestRegressor
seed=42
np.random.seed(seed)
rf = MultiOutputRegressor(RandomForestRegressor(random_state=seed))
kfold = KFold(n_splits=10, random_state=seed)
results = cross_validate(rf, X_train, y_train, cv=kfold, scoring=['r2','neg_mean_squared_error'])
r2_arr = results['test_r2']
mse_arr = results['test_neg_mean_squared_error']*-1
print(f"R^2: {r2_arr.mean():.4f} with stdev {r2_arr.std():.4f}")
print(f"MSE: {mse_arr.mean():.4f} with stdev {mse_arr.std():.4f}")
```
The The model performance without scaler is equivalent.
#### 5.2.4 Parameters tuning
```
""" Model Tuning """
# Import
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
# Initialize the classifier
seed=42
np.random.seed(seed)
reg = MultiOutputRegressor(RandomForestRegressor(random_state=seed))
#kfold = KFold(n_splits=5, random_state=seed)
#Create the parameters list to tune
parameters = {'estimator__n_estimators':[50, 75,100,200,300, 500],
'estimator__max_depth' : [2,3,5,8,10, None],
'estimator__min_samples_split' : [2,5,7,9,12],
'estimator__min_samples_leaf' : [1,3,5,7,9]
}
# Perform grid search
grid_obj = RandomizedSearchCV(reg, param_distributions=parameters,n_iter = 50,scoring='neg_mean_squared_error', verbose=10, n_jobs=-1)
# Fit the grid search object to the training data
grid_fit = grid_obj.fit(X_train, y_train)
# Get the estimator
best_reg = grid_fit.best_estimator_
print('Tuned Model')
print('R^2 score : ', best_reg.score(X_test, y_test))
print('MSE : ', mean_squared_error(y_test, best_reg.predict(X_test)))
best_reg
h.save(best_reg, 'sav/best_reg.sav')
```
## 6. Evaluation
### 6.1 Model Performance Summary
| Metric | Linear Regression | Base RF | Tuned RF |
| :------------: | :---------------: | :-------------: |:-------------: |
| R2 | 0.6099 | 0.7574 | 0.7848 |
| MSE | 0.0783 | 0.0484 | 0.0435 |
The tuned model is signifantly better than Linear Regression and better than the base random forest regressor.
### 6.2 Feature Importances
#### 6.2.1 BOGO Offer Completed Rate
```
"""
Feature Permutation Importance using eli5
https://eli5.readthedocs.io/en/latest/
"""
import eli5
from eli5.sklearn import PermutationImportance
perm = PermutationImportance(best_reg.estimators_[0], random_state=1).fit(X_test, y_test.iloc[:,0])
eli5.show_weights(perm, feature_names = X_test.columns.tolist())
"""
Partial Dependence Plot with PDPbox
https://pdpbox.readthedocs.io/en/latest/
"""
from pdpbox import pdp, get_dataset, info_plots
def plot_pdp(model, X, feature, title):
"""
INPUT :
model : fittet model sklearn object
X : features df
feature : feature name to plot, str
title : title for the plot
RETURN : None, just to plot the pdp given the feature
"""
feature_names = X.columns.tolist()
# Create the data that we will plot
pdp_goals = pdp.pdp_isolate(model=model, dataset=X, model_features=feature_names, feature=feature)
# plot it
f, ax = pdp.pdp_plot(pdp_goals, feature)
ax['pdp_ax'].set_xlabel(feature,size=15)
ax['pdp_ax'].set_title(title)
# save it
#f.savefig("graph/best"+feature+".png", bbox_inches='tight')
return ax['pdp_ax']
features = ['cluster_m_7', 'cluster_m_1', 'cluster_m_3']
bogo_model = best_reg.estimators_[0]
for feature in features :
plot_pdp(bogo_model, X_test, feature,'BOGO')
```
Cluster 1,3, and 7 in MAIN Profile are the people who are very not responsive to BOGO offer. So if a customer is in those clusters, he will have lower offer completed rate for BOGO.
```
"""sum_spending"""
plot_pdp(bogo_model, X_test, 'sum_spending','BOGO')
"""avg_spending"""
ax = plot_pdp(bogo_model, X_test, 'avg_spending','BOGO')
ax.set_xlim([0,100]);
```
Based on this graph, we see that increasing number of total spending & avg_spending increases the chances of higher offer completed rate but have little impact when it is more than ~200 dollars and ~ 12 dollars. This is the case when customers become regulars.
#### 6.3.2 DISCOUNT
```
""" Feature Permutation Importance """
import eli5
from eli5.sklearn import PermutationImportance
perm = PermutationImportance(best_reg.estimators_[1], random_state=1).fit(X_test, y_test.iloc[:,0])
eli5.show_weights(perm, feature_names = X_test.columns.tolist())
features = ['cluster_m_7', 'invalid']
discount_model = best_reg.estimators_[1]
for feature in features :
plot_pdp(discount_model, X_test, feature,'DISCOUNT')
```
Increasing Both features, `cluster_m_7` and `invalid` decreases the lower offered rate of discount. Cluster m_7 is not a very responsive for both both Bogo and Discount offer. While increasing number of invalid transactions means they are regulars and make purchasing even there is no offer.
```
"""sum_spending"""
plot_pdp(discount_model, X_test, 'sum_spending','DISCOUNT')
"""avg_spending"""
ax = plot_pdp(discount_model, X_test, 'avg_spending','DISCOUNT')
ax.set_xlim([0,100]);
```
Based on above graph, we see that increasing number of total spending & avg_spending increases the chances of higher offer completed rate. However for sum_spending, it will have little impact when it is more than ~180 dollars. And for `avg_spending` it will decrease the prediction impact when more than ~18 dollars.
#### 6.3.3 INFORMATIONAL
```
""" Feature Permutation Importance """
import eli5
from eli5.sklearn import PermutationImportance
perm = PermutationImportance(best_reg.estimators_[2], random_state=1).fit(X_test, y_test.iloc[:,0])
eli5.show_weights(perm, feature_names = X_test.columns.tolist())
info_model = best_reg.estimators_[2]
"""sum_spending"""
plot_pdp(info_model, X_test, 'offer received_informational_0_0_4','INFORMATIONAL')
```
Increasing offer informational sent to all customers, decrease the viewed rate. This is the case when we need to be selective enough when sending informational offer to improved the viewed rate and engagement.
```
"""sum_spending"""
plot_pdp(info_model, X_test, 'sum_spending','INFORMATIONAL')
ax.set_xlim([0,100]);
"""avg_spending"""
ax = plot_pdp(info_model, X_test, 'avg_spending','INFORMATIONAL')
ax.set_xlim([0,100]);
```
The customers who have total spending (`sum_spending`) less than ~ 20 dollars are likely to have lower viewed rate. But increases when higher than ~ 20 dollars, until 100 dollars, beyond that it has little impact.
It is similar with `avg_spending`, lower viewed rates is likely when the `avg_spending` is less than ~7 dollars, and higher when more than ~7 up to ~12 dollars, beyond that it has little impact or may decreases the viewed rate.
## 7. Conclusion
### Model Performance
| Metric | Linear Regression | Base RF | Tuned RF |
| :------------: | :---------------: | :-------------: |:-------------: |
| R2 | 0.6099 | 0.7574 | 0.7848 |
| MSE | 0.0783 | 0.0484 | 0.0435 |
### Summary
Increasing number of total spending & avg_spending increases the chances of higher offer completed rate for bogo and discount but have little impact when it achieved certain threshold. The threshold may indicated when the customers became regulars, they purchase even without offer anyway.
Like in part 2, Cluster 7 in MAIN profile is the cluster who are not very responsive to both offer, bogo and discount, and become a good parameter to predict a lower offer completed rate.
Given more informational offer did not guarantee the higher viewed rate, instead it caused lower viewed rate. So, we may need to be selective when sending informational offer in order to improve their engagement. Part 2 provided the targeted cluster for informational offer.
### Reflection
An unsupervised machine learning can really help us to identify and understand more about customers. Using this knowledge, we can make a better prediction about offer completed rate, how likely they are to response to the offer.
While developing model, I find it difficulty to choose the suitable algorithm in supervise machine learning. By using an available pipeline to spot check the algorithm, I can quickly screen the better model. Then start improving the model from there.
Using the feature importance in the improved model, I can have a better understanding about customers behaviour, adding a valuable information about their demographic.
### Improvement
The suggestion for improvement is to develop a pipeline model supervised machine learning to predict their cluster thru classifier then to predict their offer completed rate thru regression. A model can utilize the available neural networks technology. Advanced feature engineering can also be applied for improvement.
### References
https://machinelearningmastery.com/rescaling-data-for-machine-learning-in-python-with-scikit-learn/
| github_jupyter |
## Install
`pip install -U tabular_ml_toolkit`
*Here we are using XGBClassifier, on [Kaggle TPS Challenge (Nov 2021) data](https://www.kaggle.com/c/tabular-playground-series-nov-2021/data)*
```
from tabular_ml_toolkit.tmlt import *
from xgboost import XGBClassifier
import numpy as np
import gc
import pandas as pd
from sklearn.metrics import roc_auc_score, accuracy_score, log_loss
# Dataset file names and Paths
DIRECTORY_PATH = "/home/pankaj/kaggle_datasets/tpc_dec_2021/"
TRAIN_FILE = "train.csv"
TEST_FILE = "test.csv"
SAMPLE_SUB_FILE = "sample_submission.csv"
OUTPUT_PATH = "kaggle_tps_dec_output/"
# create tmlt
tmlt = TMLT().prepare_data(
train_file_path= DIRECTORY_PATH + TRAIN_FILE,
test_file_path= DIRECTORY_PATH + TEST_FILE,
#make sure to use right index and target columns
idx_col="Id",
target="Cover_Type",
random_state=42,
problem_type="multi_class_classification"
)
print(type(tmlt.dfl.X))
print(tmlt.dfl.X.shape)
print(type(tmlt.dfl.y))
print(tmlt.dfl.y.shape)
print(type(tmlt.dfl.X_test))
print(tmlt.dfl.X_test.shape)
print(dict(pd.Series(tmlt.dfl.y).value_counts()))
```
#### PreProcess X, y and X_test
<!-- and apply SMOTEENN combine technique (oversample+undersample) to resample imbalance classses -->
```
X_np, y_np, X_test_np = tmlt.pp_fit_transform(tmlt.dfl.X, tmlt.dfl.y, tmlt.dfl.X_test)
print(X_np.shape)
print(type(X_np))
print(y_np.shape)
print(type(y_np))
print(X_test_np.shape)
print(type(X_test_np))
print(dict(pd.Series(y_np).value_counts()))
gc.collect()
```
### For Simple TabNet Models Training
```
from pytorch_tabnet.tab_model import TabNetClassifier
tabnet_params = {
'max_epochs': 30,
'patience': 5,
'batch_size': 4096*6*tmlt.IDEAL_CPU_CORES,
'virtual_batch_size' : 512*6*tmlt.IDEAL_CPU_CORES
}
#choose model
tabnet_model = TabNetClassifier(optimizer_params=dict(lr=0.1), verbose=1)
```
#### Let's Use K-Fold Training with best params
```
%%time
# k-fold training
tabnet_model_metrics_score, tabnet_model_test_preds = tmlt.do_kfold_training(X_np, y_np, X_test=X_test_np,
n_splits=5, model=tabnet_model,
kfold_metric=accuracy_score,
eval_metric = 'accuracy',
tabnet_params=tabnet_params)
gc.collect()
```
2021-12-20 17:08:30,867 INFO Training Finished!
2021-12-20 17:08:30,868 INFO Predicting Val Probablities!
2021-12-20 17:08:32,559 INFO Predicting Val Score!
2021-12-20 17:08:34,262 INFO fold: 5 accuracy_score : 0.9588027059864701
2021-12-20 17:08:34,263 INFO Predicting Test Scores!
2021-12-20 17:08:36,422 INFO Mean Metrics Results from all Folds are: {'accuracy_score': 0.9581629704082029}
```
# predict on test dataset
if tabnet_model_test_preds is not None:
print(tabnet_model_test_preds.shape)
```
#### Create Kaggle Predictions
```
test_preds = tabnet_model_test_preds
print(type(test_preds))
test_preds_round = np.around(test_preds).astype(int)
test_preds_round[:1000]
print(f"{dict(pd.Series(test_preds_round).value_counts())}")
# target encoding changes 1 to 7 classes to 0 to 6
test_preds_round = test_preds_round + 1
print(type(test_preds_round))
print(f"{dict(pd.Series(test_preds_round).value_counts())}")
submission_file_name = 'tue_dec_21_1957_submission.csv'
sub = pd.read_csv(DIRECTORY_PATH + SAMPLE_SUB_FILE)
sub['Cover_Type'] = test_preds_round
sub.to_csv(submission_file_name, index=False)
print(f"{submission_file_name} saved!")
```
| github_jupyter |
```
PATH = "d:\\git-nlp\\ner-uk\\"
# Read tokens and positions of tokens from a file
def read_tokens(filename):
tokens = []
pos = 0
with open(filename, "r", encoding='utf-8') as f:
text = f.read().split("\n")
for line in text:
if len(line) == 0:
pos += 1
else:
tokens.append(("<S>", pos, pos))
for token in line.split(" "):
tokens.append((token, pos, pos + len(token)))
pos += len(token) + 1
tokens.append(("</S>", pos, pos))
return tokens
# Read annotations and positions of annotations from a file
def read_annotations(filename):
anno = []
with open(filename, "r", encoding='utf-8') as f:
for line in f.readlines():
annotations = line.split()
#print(annotations)
anno.append((annotations[1], int(annotations[2]), int(annotations[3])))
return anno
# Using positions of tokens and annotations, extract a list of token labels
def cyr_to_lat(label:str):
res = label.replace('ОРГ', 'ORG').replace('ЛОК', 'LOC').replace('ПЕРС', 'PERS').replace('РІЗН', 'MISC')
return res
def extract_labels(anno, tokens):
labels = []
ann_id = 0
for token in tokens:
if ann_id < len(anno):
label, beg, end = anno[ann_id]
label = cyr_to_lat(label)
if token[0] in ["<S>", "</S>"]:
labels.append("--")
elif token[1] < beg:
labels.append("--")
else:
if token[1] == beg:
labels.append("B-" + label)
else:
labels.append("I-" + label)
if token[2] == end:
ann_id += 1
else:
labels.append("--")
return labels
tokens = read_tokens(PATH + "data/A_alumni.krok.edu.ua_Prokopenko_Vidrodzhennia_velotreku(5).tok.txt")
anno = read_annotations(PATH + "data/A_alumni.krok.edu.ua_Prokopenko_Vidrodzhennia_velotreku(5).tok.ann")
labels = extract_labels(anno, tokens)
for i, j in zip(tokens, labels):
print(i[0], j)
# Extract list of files for training and testing
dev_test = {"dev": [], "test": []}
category = ""
with open(PATH + "doc/dev-test-split.txt", "r") as f:
for line in f.readlines():
line = line.strip()
if line in ["DEV", "TEST"]:
category = line.lower()
elif len(line) == 0:
continue
else:
dev_test[category].append(line)
print(len(dev_test["dev"]), len(dev_test["test"]))
# Get train and test data and labels
train_tokens, test_tokens, train_labels, test_labels = [], [], [], []
for filename in dev_test["dev"]:
try:
tokens = read_tokens(PATH + "data/" + filename + ".txt")
train_tokens += [token[0] for token in tokens]
train_labels += extract_labels(read_annotations(PATH + "data/" + filename + ".ann"), tokens)
except:
pass
for filename in dev_test["test"]:
try:
tokens = read_tokens(PATH + "data/" + filename + ".txt")
test_tokens += [token[0] for token in tokens]
test_labels += extract_labels(read_annotations(PATH + "data/" + filename + ".ann"), tokens)
except:
pass
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.naive_bayes import MultinomialNB, BernoulliNB
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.decomposition import TruncatedSVD
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.preprocessing import StandardScaler, Normalizer
```
## https://sklearn-crfsuite.readthedocs.io/en/latest/tutorial.html
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
from itertools import chain
import nltk
import sklearn
import scipy.stats
from sklearn.metrics import make_scorer
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RandomizedSearchCV
import numpy as np
import sklearn_crfsuite
from sklearn_crfsuite import scorers
from sklearn_crfsuite import metrics
from bpemb import BPEmb
bpemb_uk = BPEmb(lang="uk", dim=100)
def calc_emb(text):
res = np.zeros(bpemb_uk.vectors.shape[1], dtype=np.float32)
# tokens = word_tokenize(text)
# for t in tokens:
embs = bpemb_uk.embed(text)
for e in embs:
res += e
n = len(embs)
if n:
res /= n
return res/2
def word2features(tokens, labels, i):
word = tokens[i]
# print(word)
features = {
'bias': 1.0,
'word.lower()': word.lower(),
'word[-3:]': word[-3:],
'word[-2:]': word[-2:],
'word[:-3]': word[:-3],
'word[:-2]': word[:-2],
'word.isupper()': word.isupper(),
'word.istitle()': word.istitle(),
'word.isdigit()': word.isdigit(),
}
#emb = calc_emb(word)
#emb_features = {f'e{k}':v for k, v in enumerate(emb)}
#features.update(emb_features)
if i > 0 and tokens[i-1]!='<S>':
word1 = tokens[i-1]
features.update({
'-1:word.lower()': word1.lower(),
'-1:word.istitle()': word1.istitle(),
'-1:word.isupper()': word1.isupper(),
'-1:label': labels[i-1],
'-1:qstart': word1=='«',
})
else:
features['BOS'] = True
features['-1:label']='--'
if i < len(tokens)-1 and tokens[i+1]!='</S>':
word1 = tokens[i+1]
features.update({
'+1:word.lower()': word1.lower(),
'+1:word.istitle()': word1.istitle(),
'+1:word.isupper()': word1.isupper(),
'+1:qend': word1=='»',
})
else:
features['EOS'] = True
return features
def tokens2features(tokens, labels):
return [word2features(tokens, labels, i) for i in range(len(tokens))]
def sent2labels(sent):
return [label for token, postag, label in sent]
def sent2tokens(sent):
return [token for token, postag, label in sent]
# emb = calc_emb('слово')
# print(emb)
# emb_features = {f'e{k}':v for k, v in enumerate(emb)}
tokens2features(train_tokens, train_labels)[:3]
X_train = [tokens2features(train_tokens, train_labels)]
y_train = [train_labels]
X_test = [tokens2features(test_tokens, test_labels)]
y_test = [test_labels]
crf = sklearn_crfsuite.CRF(
algorithm='lbfgs',
c1=0.1,
c2=0.1,
max_iterations=100,
all_possible_transitions=True,
verbose=1
)
crf.fit(X_train, y_train)
labels = list(crf.classes_)
labels.remove('--')
labels
y_pred = crf.predict(X_test)
metrics.flat_f1_score(y_test, y_pred,
average='weighted', labels=labels)
# group B and I results
sorted_labels = sorted(
labels,
key=lambda name: (name[1:], name[0])
)
print(metrics.flat_classification_report(
y_test, y_pred, labels=sorted_labels, digits=3
))
```
| github_jupyter |
# PCA Clustering For Iris Dataset
```
#Importing libraries from SKLEARN
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.decomposition import PCA
iris = datasets.load_iris()
X = iris.data
y = iris.target
target_names = iris.target_names
pca = PCA(n_components=3)
X_r = pca.fit(X).transform(X)
```
# PCA Cluster plot
```
plt.scatter(X_r[y == 0, 0], X_r[y == 0, 1], s =80, c = 'orange', label = 'Iris-setosa')
plt.scatter(X_r[y == 1, 1], X_r[y == 1, 0], s =80, c = 'yellow', label = 'Iris-versicolour')
plt.scatter(X_r[y == 2, 0], X_r[y == 2, 1], s =80, c = 'green', label = 'Iris-virginica')
plt.title('PCA plot for Iris Dataset')
plt.legend()
from sklearn.model_selection import train_test_split
from sklearn import neighbors, datasets, preprocessing
X = iris.data
y = iris.target
Xtrain, Xtest, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
scaler = preprocessing.StandardScaler().fit(Xtrain)
Xtrain = scaler.transform(Xtrain)
Xtest = scaler.transform(Xtest)
# Using knn model for accuracy
knn = neighbors.KNeighborsClassifier(n_neighbors=5)
knn.fit(Xtrain, y_train)
y_pred = knn.predict(Xtest)
y_pred
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
print('Accuracy Score:', accuracy_score(y_test, y_pred))
print('Confusion matrix \n', confusion_matrix(y_test, y_pred))
print('Classification \n', classification_report(y_test, y_pred))
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
Xtrain, Xtest, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
```
# Logistic Regression Accuracy
```
#Logistic Regression
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
accuracy = accuracy_score(y_test,y_pred)
print("Logistic Regression :")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For LR
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
```
# K Nearest Neighbors Accuracy
```
#K Nearest Neighbors
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier(n_neighbors=5, p=2, metric='minkowski')
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
accuracy = accuracy_score(y_test,y_pred)
print("K Nearest Neighbors :")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For KNN
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
Xtrain, Xtest, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=0)
```
# Support Vector Machine Accuracy
```
#Support Vector Machine
from sklearn.svm import SVC
classifier = SVC(kernel='linear',random_state=123)
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
accuracy = accuracy_score(y_test,y_pred)
print("Support Vector Machine:")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For SVM
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
```
# Gaussian Naive Bayes Accuracy
```
#Gaussian Naive Bayes
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
accuracy = accuracy_score(y_test,y_pred)
print("Gaussian Naive Bayes :")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For GNB
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
```
# Decision Tree Classifier Accuracy
```
#Decision Tree Classifier
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier as DT
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
classifier = DT(criterion='entropy',max_depth=3, random_state=0)
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
print("Decision Tree Classifier :")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For DTC
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
```
# Random Forest Classifier Accuracy
```
#Random Forest Classifier
from sklearn.ensemble import RandomForestClassifier as RF
classifier = RF(n_estimators=10, criterion='entropy', random_state=0)
classifier.fit(Xtrain,y_train)
y_pred = classifier.predict(Xtest)
cm = confusion_matrix(y_test,y_pred)
print("Random Forest Classifier :")
print("Accuracy = ", accuracy)
print(cm)
```
# Cohen Kappa Accuracy For RFC
```
from sklearn.metrics import cohen_kappa_score
cluster = cohen_kappa_score(y_test, y_pred)
cluster
```
| github_jupyter |
# 1-D Parabolic Equations
Recall that the governing equation (in 1D) is:
$$\frac{\partial u}{\partial t}=\kappa\left(\frac{\partial^2 u}{\partial x^2}\right)$$
To solve the equation we need both boundary condition values at _u(x0,t)_ and _u(x1,t)_ and also initial conditions at _u(x,0)_.
We will need a class to store the 1D grid, set the initial conditions and update the boundary conditions. Fortunately the grid generation is simpler than the 2D case.
To set up the initial condtions we will pass a function _f(x)_ to the function, this will be called inside a loop passing the _x_ orfdinate of evergy grid point. A similar idea will be used for Dirichlet boundary condtions where a function _f(t)_ will be passed as a parameter. Here is the code for the initial conditions:
def initial_conditions(self,fun):
for i in range(self.N_i):
self.u[0,i] = fun(self.x[i])
Finally we need to know the time step that can be used. As we saw in the lectures this is characterised by the CFL condition
$$\Delta t \le \nu \frac{\Delta x^2}{2\kappa}$$
The choice of ν depends on the solver being used so we will pass it as a parameter:
def set_Delta_t(self,Coutrant_number):
self.Delta_t = Coutrant_number * self.Delta_x()**2/(2*self.kappa)
```
import numpy as np
import matplotlib.pyplot as plt
import time
def one(t):
'''simple function of x and t which allways returns 1.0'''
return 1.0
class Grid:
'''Class defining the grid for a 1D parabolic solver. Solving
$$\frac{\partial u}{\partial t}=\kappa\left(\frac{\partial^2 u}{\partial x^2}\right)$$
The grid object contains the number of grid points (Ni), left
and right ordinates (x0 and x1), flags for wether the boundaries
are Dirichlet or Neumann boundary conditions and functions for
plotting the solution both at a particular time level and in the
form of an x-t plot. The grid also contains the solution vector (u)
which is stored as an array with differnt time levels, together with
the number of time steps N_t and the (uniform) time step Delta_t.
Written by Prof David Ingram, School of Engineering
(c) 2021 The University of Edinburgh
Licensed under CC-BY-NC.
'''
DIRICHLET_BC = 0
NEUMANN_BC = 1
BC_NAME = ['left', 'right']
def __init__(self,ni):
# set up information about the grid
self.x0 = 0.0 # left
self.x1 = 1.0 # right
self.N_i = ni # grid points in i direction
self.N_t = 0 # how many time steps have we performed
self.Delta_t = 0.0 # no dt value yet
self.kappa = 1.0 # Thermal Diffusivity
# initialse x,y and u arrays
self.u = np.zeros((1,ni))
self.x = np.zeros(ni)
self.t = np.zeros(1)
# boundary conditions (left and right )
self.BC = [ self.DIRICHLET_BC, self.DIRICHLET_BC ]
self.BC_fun = [ one, one ]
def set_x0(self,x0):
self.x0 = x0
def set_x1(self,x1):
self.x1 = x1
def generate(self,Quiet=True):
'''generate a uniformly spaced grid covering the domain from the
x0 to the x1. We are going to do this using linspace from
numpy to create a list of x ordinates.'''
self.x = np.linspace(self.x0, self.x1, self.N_i)
if not Quiet:
print(self)
def initial_conditions(self,fun):
'''set the initial conditions by calling the specified function
fun(x) at every ordinate'''
for i in range(self.N_i):
self.u[0,i] = fun(self.x[i])
def Delta_x(self):
# calculate delta x
return self.x[1]-self.x[0]
def set_Neumann_bc(self,side):
try:
self.BC[self.BC_NAME.index(side)] = self.NEUMANN_BC
except:
print('error {} must be one of {}'.format(side,self.BC_NAME))
def set_Dirichlet_bc(self,side,fun):
'''set the Dirichlet boundary condition. As well as the side
this must be passed a function fun(t) which returns the value
associated with the Dirichlet boundary at the time t.'''
try:
self.BC[self.BC_NAME.index(side)] = self.DIRICHLET_BC
except:
print('error {} must be one of {}'.format(side,self.BC_NAME))
self.BC_fun[self.BC_NAME.index(side)] = fun
def set_Delta_t(self,Coutrant_number):
# set the time step using the Courant-Friedirchs-Lewey condition
self.Delta_t = Coutrant_number * self.Delta_x()**2/(2*self.kappa)
def apply_BC(self,t):
'''Apply the left and right boundary conditions to the values in
the latest value of u at the current time level'''
# left boundary
if self.BC[0]==self.NEUMANN_BC:
self.u[-1,0] = self.u[-1,2]
else:
self.u[-1,0] = self.BC_fun[0](t)
# right
if self.BC[1]==self.NEUMANN_BC:
self.u[-1,-1] = self.u[-1,-3]
else:
self.u[-1,-1] = self.BC_fun[1](t)
def integrate(self,t_stop,solver):
'''call the integrator defined by solver to integrate
the solution from t=0 to t=t_stop.'''
time = 0.0
o_time = t_stop/10
print('Integratung to {:.3g} seconds Δt={:.3g} :'.format(t_stop,self.Delta_t),end='')
# save the initial conditions as the first saved result
self.u = np.vstack([self.u, self.u])
self.t = np.vstack([self.t, 0.0])
# time loop
while True:
# short time step at the end
dt = min(self.Delta_t,t_stop-time)
# update the boundary conditions and call the solver
self.apply_BC(time)
# update the time and save the time and results
time += dt
self.u[-1] = solver(self)
self.t[-1] = time
# progress monitor
if (time >= o_time) & (time<t_stop):
print('#',end='')
o_time += t_stop/10
self.t = np.vstack([self.t, time])
self.u = np.vstack([self.u, self.u[-1]])
self.N_t += 1
# are we done
if time >= t_stop:
print('. ')
break
def plot(self,title):
'''plot the solution at the current time level. If there are
more than 30 points in the x-direction then a line is used.'''
if self.N_i<30:
plt.plot(self.x,self.u[self.N_t],'ob')
else:
plt.plot(self.x,self.u[self.N_t],'-b')
plt.title(title+', t={:.2f}'.format(self.N_t*self.Delta_t))
plt.show()
def report_BC(self):
'''compile a string listing the boundary conditions on each side.
We build up a string of two {side name}: {BC type} pairs and
return it'''
# initialise the string
string = ''
# loop over the sides
for side in range(2):
# add the side name
string = string + self.BC_NAME[side]
# and the bounday condition type
if self.BC[side] == self.DIRICHLET_BC:
string = string + ': Dirichlet, '
elif self.BC[side] == self.NEUMANN_BC:
string = string + ': Neumann, '
return string[:-2] +'.' # lose the last comma and space.
def __str__(self):
# describe the object when asked to print it
describe = 'Parabolic problem, 𝜅={:.3g}\n'.format(self.kappa)
describe += 'Uniform {} node grid from {} to {}.\n'.format(self.N_i, self.x0, self.x1)
describe += 'Boundaries conditions are - ' + self.report_BC()
if self.N_t==0:
describe += '\nInitial conditions at t=0.0, Δt={:.3g} seconds'.format(self.Delta_t)
else:
describe += '\n{} time steps from t=0.0 to t={:.3g}, Δt={:.3g} seconds'.format(self.N_t+1,self.N_t*self.Delta_t,self.Delta_t)
return describe
```
### A test problem
We have a 1m long domain from _x_=0.0 to _x_=1.0 with 𝜅=1.0. The initial conditions are
$$u(x,0)=\sin \frac{3\pi x}{2}$$
and we have a Dirichlet boundary condition with _u_(0,_t_)=0.0 at _x_=0.0 and a Neumann boundary condition _u'_(1,_t_)=0.0 at _x_=1.0.
This problem has an analytical solution
$$u(x,t)=\exp\left(-\frac{9\kappa\pi^2 t}{4}\right)\sin\frac{3\pi x}{2}.$$
Let's set it up
```
def u_bc(x):
return 0.0
def u_init(x):
return np.sin(3*np.pi*x/2)
test = Grid(20)
test.generate()
test.set_Delta_t(0.95)
test.set_Neumann_bc('right')
test.set_Dirichlet_bc('left',u_bc)
test.initial_conditions(u_init)
test.plot('Problem 9.20')
print(test)
```
## The FTCS Scheme
Replacing the time derivities with the first order, forward, finite difference approximation and the second derivative with the 2nd order central finite difference apporximation and introducing the index _n_ to represent the time level we obtain:
$$\frac{u_{i}^{n+1}-u_{i}^n}{\Delta t}=\kappa\frac{u^n_{i-1}-2u^n_i+u^n_{i+1}}{\Delta x^2}.$$
This can be re-arranged to obtain the explicit time-marching method:
$$u_i^{n+1}=u^n_i+\frac{\kappa\Delta t}{\Delta x^2}\left(u^n_{i-1}-2u^n_i+u^n_{i+1}\right).$$
This is the well known Forward-Time, Centred-Space (FTCS) scheme. If is formally 1<sup>st</sup> order in time and 2<sup>nd</sup> order in space.
```
def FTCS(grid):
'''perform one time step using the FTCS scheme on the parabolic
problem defined by grid. We are using whole array opperations
for speed.'''
Rx = grid.kappa*grid.Delta_t/grid.Delta_x()**2
u = grid.u[-1].copy()
u[1:-1]=u[1:-1]+Rx*(u[0:-2]-2*u[1:-1]+u[2:])
return u
test.integrate(0.1, FTCS)
test.plot('Example 9.20')
print(test)
# do annother 0.1 seconds
test.integrate(0.1, FTCS)
test.plot('Example 9.20')
print(test)
# more grid points
test1 = Grid(201)
test1.generate()
test1.set_Delta_t(0.95)
test1.set_Neumann_bc('right')
test1.set_Dirichlet_bc('left',u_bc)
test1.initial_conditions(u_init)
test1.integrate(0.1, FTCS)
test1.plot('Example 9.20')
print(test1)
```
### X-t plot
Because we have stored every time step, we can plot a contour plot of the solution with time on one axis and space on the other. To do this we must:
1. create a <code>meshgrid</code> using _x_ and _t_
2. plot the contour plot
3. add a colour bar
4. add titles and captions
```
X,T = np.meshgrid(test1.x,test1.t)
fig, ax1 = plt.subplots()
cmap = plt.get_cmap('jet')
cf = ax1.contourf(X,T,test1.u,cmap=cmap, levels = 21)
fig.colorbar(cf, ax=ax1)
ax1.set_title('Example 9.20: x-t plot')
ax1.set_xlabel('x')
ax1.set_ylabel('t')
plt.show()
```
### Analysis
The _x_-_t_ plot shows the expected behaviour with the amplitude of the oscilation decaying as time increases.
We can compare the solution with the analytical one as well
```
plt.plot(test1.x,test1.u[-1],'r-',label='Numerical')
u_exact = np.exp(-9*test1.kappa*np.pi**2*test1.t[-1]/4)*np.sin(1.5*np.pi*test1.x)
plt.plot(test1.x,u_exact,'b-',label='Analytical')
plt.title('Example 9.20 $\kappa=1.0$, $\Delta x={:.3g}$ $t=0.1$ seconds'.format(test.Delta_x()))
plt.legend()
plt.show()
```
## Mesh refinement study
In this case we are going to use the range of _u(x)_ as the integrating quantity.
$$\text{Range}(u)=\max_i u - \min_i u$$
We will run the solution to _t_=0.1 seconds and use 𝜅=1.0 As with the Laplace Solvers I'm going to use a sequence of grids with
$$N_i = 20\times2^n + 1 \ n=1 \ldots 8.$$
The finest grid will have 5121 grid points and the coarsest 81. The finest grid will take about 5 mins to run. We are also going to plot the solution on every grid (together with the analytical solution).
```
import datetime # just seconds may not be enough
# we need some lists u and dx values
U_val = []
dx_val = []
run_time = []
n_pts =[]
for grid_index in range(8,1,-1):
ni = 20 * 2**grid_index + 1
n_pts.append(ni)
# set up the problem
test = Grid(ni)
test.generate()
test.set_Delta_t(0.95)
test.set_Neumann_bc('right')
test.set_Dirichlet_bc('left',u_bc)
test.initial_conditions(u_init)
print(test)
# run the solver
start = time.process_time()
test.integrate(0.1, FTCS)
stop = time.process_time()
elapsed = datetime.timedelta(seconds=stop-start)
print("The solver took ",elapsed)
# save dx and the range into the list for evalutation
dx_val.append(test.Delta_x())
U_val.append(np.max(test.u[-1])-np.min(test.u[-1]))
run_time.append(stop-start)
print('Range(u) is ',U_val[-1],'\n')
# plot a graph of the solution and the analytical solution on the same grid.
plt.plot(test.x,test.u[-1],'r-',label='Numerical')
u_exact = np.exp(-9*test.kappa*np.pi**2*test.t[-1]/4)*np.sin(1.5*np.pi*test.x)
plt.plot(test.x,u_exact,'b-',label='Analytical')
plt.title('Example 9.20 $\kappa=1.0$, $\Delta x={:.3g}$ $t=0.1$ seconds'.format(test.Delta_x()))
plt.legend()
plt.show()
# now run the mesh refinement analysis
from refinement_analysis import refinement_analysis
# lets to the refinement analysis
analysis = refinement_analysis(dx_val,U_val)
analysis.report('range(u)')
analysis.plot(True,'range(u)')
```
### Analysis
The method has an apparent accuracy of first order. We expect this since it's 2nd order in space and 1st order in time. We've taken a lot of time steps so accuracy is at most 1st order. On the finest grids (_Ni_>320) the solution is grid converged (and it's prety close on the other grids). So what about the execution time
```
# plot the runtime
plt.plot(n_pts,run_time,'o')
plt.title('Runtime')
plt.xlabel('N')
plt.yscale('log',base=10)
plt.ylabel('runtime (s)')
plt.show()
```
| github_jupyter |
# Tutorial: The Basic Tools of Private Deep Learning
Welcome to PySyft's introductory tutorial for privacy preserving, decentralized deep learning. This series of notebooks is a step-by-step guide for you to get to know the new tools and techniques required for doing deep learning on secret/private data/models without centralizing them under one authority.
**Scope:** Note that we'll not just be talking about how to decentralized / encrypt data, but we'll be addressing how PySyft can be used to help decentralize the entire ecosystem around data, even including the Databases where data is stored and queried, and the neural models which are used to extract information from data. As new extensions to PySyft are created, these notebooks will be extended with new tutorials to explain the new functionality.
Authors:
- Andrew Trask - Twitter: [@iamtrask](https://twitter.com/iamtrask)
## Outline:
- Part 1: The Basic Tools of Private Deep Learning
## Why Take This Tutorial?
**1) A Competitive Career Advantage** - For the past 20 years, the digital revolution has made data more and more accessible in ever larger quantities as analog processes have become digitized. However, with new regulation such as [GDPR](https://eugdpr.org/), enterprises are under pressure to have less freedom with how they use - and more importantly how they analyze - personal information. **Bottom Line:** Data Scientists aren't going to have access to as much data with "old school" tools, but by learning the tools of Private Deep Learning, YOU can be ahead of this curve and have a competitive advantage in your career.
**2) Entrepreneurial Opportunities** - There are a whole host of problems in society that Deep Learning can solve, but many of the most important haven't been explored because it would require access to incredibly sensitive information about people (consider using Deep Learning to help people with mental or relationship issues!). Thus, learning Private Deep Learning unlocks a whole host of new startup opportunities for you which were not previously available to others without these toolsets.
**3) Social Good** - Deep Learning can be used to solve a wide variety of problems in the real world, but Deep Learning on *personal information* is Deep Learning about people, *for people*. Learning how to do Deep Learning on data you don't own represents more than a career or entrepreneurial opportunity, it is the opportunity to help solve some of the most personal and important problems in people's lives - and to do it at scale.
## How do I get extra credit?
- Star PySyft on GitHub! - [https://github.com/OpenMined/PySyft](https://github.com/OpenMined/PySyft)
- Make a Youtube video teaching this notebook!
... ok ... let's do this!
# Part -1: Prerequisites
- Know PyTorch - if not then take the http://fast.ai course and come back
- Read the PySyft Framework Paper https://arxiv.org/pdf/1811.04017.pdf! This will give you a thorough background on how PySyft is constructed which will help things make more sense.
# Part 0: Setup
To begin, you'll need to make sure you have the right things installed. To do so, head on over to PySyft's readme and follow the setup instructions. TLDR for most folks is.
- Install Python 3.6 or higher
- Install PyTorch 1.4
- Clone PySyft (git clone https://github.com/OpenMined/PySyft.git)
- cd PySyft
- pip install -r pip-dep/requirements.txt
- pip install -r pip-dep/requirements_udacity.txt
- python setup.py install udacity
- python setup.py test
If any part of this doesn't work for you (or any of the tests fail) - first check the [README](https://github.com/OpenMined/PySyft.git) for installation help and then open a GitHub Issue or ping the #beginner channel in our slack! [slack.openmined.org](http://slack.openmined.org/)
```
# Run this cell to see if things work
import sys
import torch
from torch.nn import Parameter
import torch.nn as nn
import torch.nn.functional as F
import syft as sy
hook = sy.TorchHook(torch)
torch.tensor([1,2,3,4,5])
```
If this cell executed, then you're off to the races! Let's do this!
# Part 1: The Basic Tools of Private, Decentralized Data Science
So - the first question you may be wondering is - How in the world do we train a model on data we don't have access to?
Well, the answer is surprisingly simple. If you're used to working in PyTorch, then you're used to working with torch.Tensor objects like these!
```
x = torch.tensor([1,2,3,4,5])
y = x + x
print(y)
```
Obviously, using these super fancy (and powerful!) tensors is important, but also requires you to have the data on your local machine. This is where our journey begins.
# Section 1.1 - Sending Tensors to Bob's Machine
Whereas normally we would perform data science / deep learning on the machine which holds the data, now we want to perform this kind of computation on some **other** machine. More specifically, we can no longer simply assume that the data is on our local machine.
Thus, instead of using Torch tensors, we're now going to work with **pointers** to tensors. Let me show you what I mean. First, let's create a "pretend" machine owned by a "pretend" person - we'll call him Bob.
```
bob = sy.VirtualWorker(hook, id="bob")
```
Let's say Bob's machine is on another planet - perhaps on Mars! But, at the moment the machine is empty. Let's create some data so that we can send it to Bob and learn about pointers!
```
x = torch.tensor([1,2,3,4,5])
y = torch.tensor([1,1,1,1,1])
```
And now - let's send our tensors to Bob!!
```
x_ptr = x.send(bob)
y_ptr = y.send(bob)
x_ptr
```
BOOM! Now Bob has two tensors! Don't believe me? Have a look for yourself!
```
bob._objects
z = x_ptr + x_ptr
z
bob._objects
```
Now notice something. When we called `x.send(bob)` it returned a new object that we called `x_ptr`. This is our first *pointer* to a tensor. Pointers to tensors do NOT actually hold data themselves. Instead, they simply contain metadata about a tensor (with data) stored on another machine. The purpose of these tensors is to give us an intuitive API to tell the other machine to compute functions using this tensor. Let's take a look at the metadata that pointers contain.
```
x_ptr
```
Check out that metadata!
There are two main attributes specific to pointers:
- `x_ptr.location : bob`, the location, a reference to the location that the pointer is pointing to
- `x_ptr.id_at_location : <random integer>`, the id where the tensor is stored at location
They are printed in the format `<id_at_location>@<location>`
There are also other more generic attributes:
- `x_ptr.id : <random integer>`, the id of our pointer tensor, it was allocated randomly
- `x_ptr.owner : "me"`, the worker which owns the pointer tensor, here it's the local worker, named "me"
```
x_ptr.location
bob
bob == x_ptr.location
x_ptr.id_at_location
x_ptr.owner
```
You may wonder why the local worker which owns the pointer is also a VirtualWorker, although we didn't create it.
Fun fact, just like we had a VirtualWorker object for Bob, we (by default) always have one for us as well. This worker is automatically created when we called `hook = sy.TorchHook()` and so you don't usually have to create it yourself.
```
me = sy.local_worker
me
me == x_ptr.owner
```
And finally, just like we can call .send() on a tensor, we can call .get() on a pointer to a tensor to get it back!!!
```
x_ptr
x_ptr.get()
y_ptr
y_ptr.get()
z.get()
bob._objects
```
And as you can see... Bob no longer has the tensors anymore!!! They've moved back to our machine!
# Section 1.2 - Using Tensor Pointers
So, sending and receiving tensors from Bob is great, but this is hardly Deep Learning! We want to be able to perform tensor _operations_ on remote tensors. Fortunately, tensor pointers make this quite easy! You can just use pointers like you would normal tensors!
```
x = torch.tensor([1,2,3,4,5]).send(bob)
y = torch.tensor([1,1,1,1,1]).send(bob)
z = x + y
z
```
And voilà!
Behind the scenes, something very powerful happened. Instead of x and y computing an addition locally, a command was serialized and sent to Bob, who performed the computation, created a tensor z, and then returned the pointer to z back to us!
If we call .get() on the pointer, we will then receive the result back to our machine!
```
z.get()
```
### Torch Functions
This API has been extended to all of Torch's operations!!!
```
x
y
z = torch.add(x,y)
z
z.get()
```
### Variables (including backpropagation!)
```
x = torch.tensor([1,2,3,4,5.], requires_grad=True).send(bob)
y = torch.tensor([1,1,1,1,1.], requires_grad=True).send(bob)
z = (x + y).sum()
z.backward()
x = x.get()
x
x.grad
```
So as you can see, the API is really quite flexible and capable of performing nearly any operation you would normally perform in Torch on *remote data*. This lays the groundwork for our more advanced privacy preserving protocols such as Federated Learning, Secure Multi-Party Computation, and Differential Privacy !
# Congratulations!!! - Time to Join the Community!
Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
### Star PySyft on GitHub
The easiest way to help our community is just by starring the GitHub repos! This helps raise awareness of the cool tools we're building.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
### Join a Code Project!
The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
- [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
[OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
(ADBLUCO)=
# 3.2 Algoritmos de descenso y búsqueda de línea en *Unconstrained Convex Optimization* (UCO)
```{admonition} Notas para contenedor de docker:
Comando de docker para ejecución de la nota de forma local:
nota: cambiar `<ruta a mi directorio>` por la ruta de directorio que se desea mapear a `/datos` dentro del contenedor de docker y `<versión imagen de docker>` por la versión más actualizada que se presenta en la documentación.
`docker run --rm -v <ruta a mi directorio>:/datos --name jupyterlab_optimizacion -p 8888:8888 -d palmoreck/jupyterlab_optimizacion:<versión imagen de docker>`
password para jupyterlab: `qwerty`
Detener el contenedor de docker:
`docker stop jupyterlab_optimizacion`
Documentación de la imagen de docker `palmoreck/jupyterlab_optimizacion:<versión imagen de docker>` en [liga](https://github.com/palmoreck/dockerfiles/tree/master/jupyterlab/optimizacion).
```
---
Nota generada a partir de [liga](https://drive.google.com/file/d/16-_PvWNaO0Zc9x04-SRsxCRdn5fxebf2/view).
```{admonition} Al final de esta nota el y la lectora:
:class: tip
* Comprenderá el uso de la información de primer y segundo orden para resolver problemas *small scale* de minimización de funciones convexas mediante los algoritmos general de descenso y de búsqueda de línea por *backtracking*.
* Aprenderá la importancia y relación con ecuaciones no lineales al resolver los problemas que involucran aproximar mínimos locales de funciones.
```
En esta nota consideramos problemas de optimización *small scale*. Aunque el término *small scale* es ambiguo pues depende la máquina en la que se realice el cómputo e involucra el número de variables o parámetros y cantidad de almacenamiento para datos, tomamos como *small scale* aquel problema de optimización **sin restricciones** en el que se tiene un número de variables del orden menor o igual a $10^3$.
## Ejemplos de problemas de optimización *small scale*
En optimización la búsqueda del (o los) **óptimo(s)** involucran el cálculo de información de primer o segundo orden, ver {ref}`Definición de función, continuidad y derivada <FCD>`, de la función $f_o$ de acuerdo a lo revisado en los {ref}`resultados útiles<RESUT>`. Tal información para problemas *small scale* es calculada utilizando todos los datos en un enfoque por *batch* o lote.
### Ejemplo
$$\displaystyle \min_{x \in \mathbb{R}^2} x_1^4+2x_1^2x_2+x_2^2$$
```
import numpy as np
import sympy
from sympy.tensor.array import derive_by_array
import matplotlib.pyplot as plt
from scipy.optimize import fsolve
from scipy.optimize import fmin
import pandas as pd
import cvxpy as cp
from pytest import approx
np.set_printoptions(precision=3, suppress=True)
```
Los candidatos a ser mínimos los encontramos al calcular el gradiente de $f_o$ que podemos calcular con *SymPy*
```
x1, x2 = sympy.symbols("x1, x2")
fo_sympy = x1**4+2*x1**2*x2+x2**2
sympy.pprint(fo_sympy)
gf = derive_by_array(fo_sympy, (x1, x2))
sympy.pprint(gf)
```
y plantear:
$$
\nabla f_o(x) =
\left [
\begin{array}{c}
4x_1^3+4x_1x_2\\
2x_1^2+2x_2
\end{array}
\right ]=0
$$
la cual es una ecuación de dos variables y dos incógnitas **no lineal**. Resolviendo para $x_2$ se obtiene la relación: $x_2 = -x_1^2$. Entonces todos los puntos con coordenadas $x = (x_1, x_2)$ que satisfacen tal relación cumplen $\nabla f_o(x) = 0$. ¿Todos serán mínimos locales?
**Gráfica de la superficie $f_o$**
```
def fo_numpy(x):
return x[0]**4 + 2*x[0]**2*x[1] + x[1]**2
x1_plot,x2_plot = np.meshgrid(np.linspace(-2,2,100), np.linspace(-2,2,100))
z_plot = x1_plot**4 + 2*x1_plot**2*x2_plot + x2_plot**2
x1_np = 0
x2_np = 0
z_np = fo_numpy([x1_np, x2_np])
point = (x1_np, x2_np, z_np)
print(point)
# Create the figure
fig = plt.figure()
# Add an axes
ax = fig.gca(projection='3d')
ax.plot_surface(x1_plot, x2_plot, z_plot, alpha=0.2)
ax.scatter(point[0], point[1], point[2], color='green')
plt.title("$f_o(x) = x_1^4+2x_1^2x_2+x_2^2$")
plt.show()
```
**Gráfica de las curvas de nivel de $f_o$**
```
x1_plot,x2_plot = np.meshgrid(np.linspace(-2,2,100), np.linspace(-4, 1,100))
z_plot = x1_plot**4 + 2*x1_plot**2*x2_plot + x2_plot**2
plt.contour(x1_plot,x2_plot,z_plot)
plt.scatter(point[0], point[1], color="green")
plt.title("Curvas de nivel de $f_o$")
plt.show()
```
Resolvamos con [scipy.optimize.fsolve](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.fsolve.html#scipy.optimize.fsolve) la ecuación no lineal $\nabla f_o(x) = 0$
```
def eqn(x):
x1,x2=x
return [4*x1**3+4*x1*x2, 2*x1**2+2*x2]
```
```{margin}
Elegimos diferentes puntos iniciales.
```
```
root1 = fsolve(eqn, (1, 1))
root2 = fsolve(eqn, (-1, 1))
root3 = fsolve(eqn, (2, 0))
dic_roots = {"root1": root1,
"root2": root2,
"root3": root3}
```
```{margin}
Obsérvese que los puntos `root1`, `root2`, `root3` satisfacen la relación $x_2 = -x_1^2$.
```
```
print(pd.DataFrame(dic_roots))
```
Al evaluar el gradiente en cada punto obtenemos cero (o cercano a cero):
```
gf_eval = lambda x: np.array([partial_derivative.subs({"x1": x[0],
"x2": x[1]}) for partial_derivative in gf],
dtype=float)
dic = {"root1": gf_eval(root1),
"root2": gf_eval(root2),
"root3": gf_eval(root3)}
```
```{margin}
Los puntos `root1`, `root2` y `root3` resuelven la ecuación no lineal $\nabla f(x) = 0$ .
```
```
print(pd.DataFrame(dic).round(3))
```
**¿Cómo podemos identificar si son mínimos? ...** usamos la Hessiana de $f_o$
```
Hf = derive_by_array(gf, (x1, x2))
sympy.pprint(Hf)
```
y revisamos eigenvalores de la Hessiana evaluada en los puntos `root1`, `root2`, `root3`
```
Hf_eval = lambda x: np.array([second_partial_derivative.subs({"x1": x[0],
"x2": x[1]}) for second_partial_derivative in Hf],
dtype=float)
Hf_root1 = Hf_eval(root1)
Hf_root2 = Hf_eval(root2)
Hf_root3 = Hf_eval(root3)
```
```{margin}
La Hessiana en `root1` es semidefinida positiva por lo que no podemos concluir que sea mínimo local de $f_o$.
```
```
print(np.linalg.eigvals(Hf_root1))
```
```{margin}
La Hessiana en `root2` es semidefinida positiva por lo que no podemos concluir que sea mínimo local de $f_o$.
```
```
print(np.linalg.eigvals(Hf_root2))
```
```{margin}
La Hessiana en `root3` es semidefinida positiva por lo que no podemos concluir que sea mínimo local de $f_o$.
```
```
print(np.linalg.eigvals(Hf_root3))
```
Tenemos un criterio para $2$ dimensiones:
```{admonition} Comentario
Sea $f \in \mathcal{C}^2(\text{intdom}f)$, $\det(\nabla^2 f(x))$ determinante de la Hessiana y $x \in \mathbb{R}^2$ un punto crítico o estacionario de $f$:
* Si $\frac{\partial^2f(x)}{\partial x_1^2} < 0$ y $\det(\nabla^2 f(x)) >0$ entonces $f$ tiene un **máximo local** en $x$.
* Si $\frac{\partial^2f(x)}{\partial x_1^2} > 0$ y $\det(\nabla^2 f(x)) >0$ entonces $f$ tiene un **mínimo local** en $x$.
* Si $\det(\nabla^2 f(x)) < 0$ entonces $f$ tiene un **punto silla o [*saddle point*](https://en.wikipedia.org/wiki/Saddle_point)** en $x$.
* Si $\det(\nabla^2 f(x)) = 0$ no podemos concluir si $x$ es extremo.
```
```{admonition} Observaciones
:class: tip
* Al determinante de la Hessiana de $f$ se le nombra **Hessiano** de $f$.
* Lo anterior es un caso particular de los resultados descritos en {ref}`sobre puntos críticos <SPCRITICOS>`.
```
En el ejemplo el Hessiano es:
```
sympy.pprint(sympy.Matrix(Hf).det())
```
```{margin}
Los tres puntos `root1`, `root2` y `root3` satisfacen $x_2 = -x_1^2$.
```
El cual se anula justo en los puntos que cumplen: $x_2 = -x_1^2$
$8x_1^2 + 8x_2 = 8 x_1^2 + 8(-x_1^2) = 0$
por lo que no podemos concluir...
Usemos una función de *SciPy* [scipy.optimize.fmin](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.fmin.html)
```{margin}
Elegimos un punto inicial.
```
```
res_fmin = fmin(fo_numpy, [1,1])
```
```{margin}
El punto `res_fmin` satisface $\nabla f_o(x) = 0$ y la relación $x_2 = -x_1^2$.
```
```
print(res_fmin)
print(gf_eval(res_fmin))
Hf_fmin = Hf_eval(res_fmin)
```
```{margin}
La Hessiana en `res_fmin` es semidefinida positiva por lo que no podemos concluir que sea mínimo local de $f_o$.
```
```
print(np.linalg.eigvals(Hf_fmin))
```
Grafiquemos los puntos que cumplen $x_2=-x_1^2$
```
x1_plot,x2_plot = np.meshgrid(np.linspace(-2,2,100), np.linspace(-4,2,100))
z_plot = x1_plot**4 + 2*x1_plot**2*x2_plot + x2_plot**2
point1 = (root1[0], root1[1], fo_numpy(root1))
point2 = (root2[0], root2[1], fo_numpy(root2))
point3 = (root3[0], root3[1], fo_numpy(root3))
#another point:
point4 = (-2, -4, 0)
x1_plot2 = np.linspace(-2,2,100)
# Create the figure
fig = plt.figure()
# Add an axes
ax = fig.gca(projection='3d')
ax.plot_surface(x1_plot, x2_plot, z_plot, alpha=0.2)
ax.scatter(point[0], point[1], point[2], color='green')
ax.scatter(point1[0], point1[1], point1[2], color='green')
ax.scatter(point2[0], point2[1], point2[2], color='green')
ax.scatter(point3[0], point3[1], point3[2], color='green')
ax.scatter(point4[0], point4[1], point4[2], color='green')
ax.plot(x1_plot2, -x1_plot2**2, color="red")
plt.title("$f_o(x) = x_1^4+2x_1^2x_2+x_2^2$")
plt.show()
```
Evaluemos en un rango más amplio la función objetivo $f_o$ y realicemos una gráfica
```
x1_plot,x2_plot = np.meshgrid(np.linspace(-100,100,100), np.linspace(-100,100,100))
z_plot = x1_plot**4 + 2*x1_plot**2*x2_plot + x2_plot**2
# Create the figure
fig = plt.figure()
# Add an axes
ax = fig.gca(projection='3d')
ax.plot_surface(x1_plot, x2_plot, z_plot, alpha=0.2)
plt.title("$f_o(x) = x_1^4+2x_1^2x_2+x_2^2$")
plt.show()
np.sum(z_plot < 0)
```
**El mínimo valor de $f_o$ es $0$ por lo que tenemos un conjunto de mínimos dados por la curva $x_2=-x_1^2$.**
### ¿Por qué fue un poco complicado determinar el conjunto de mínimos de $f_o$?
**$f_o(x)=x_1^4+2x_1^2x_2+x_2^2$ no es una función convexa en su $\text{dom}f_o$**, $\mathbb{R}^2$, esto es, no satisface la desigualdad:
$$f_o(\theta x + (1-\theta) y) \leq \theta f_o(x) + (1-\theta)f_o(y)$$
$\forall x,y$ en su dominio y $\forall \theta \in [0,1]$.
```
pointx = np.array([-.5, -1.5])
pointy = np.array([.5, -1.5])
theta = 1/2
point_convex_combination = theta*pointx + (1-theta)*pointy
print(fo_numpy(point_convex_combination))
```
```{margin}
Se tiene $f(\theta x + (1-\theta)y) > \theta f(x) + (1-\theta)f(y)$ con la elección $\theta=0.5$, $x=[-0.5, -1.5]^T$, $y=[0.5, -1.5]^T$ lo que indica que $f$ no es convexa sino cóncava para esta elección de puntos $x,y$.
```
```
print(theta*fo_numpy(pointx) + (1-theta)*fo_numpy(pointy))
```
```{admonition} Observación
:class: tip
Recordar que si $f_o$ es una función convexa, el gradiente de $f_o$ nos ayuda a determinar si un punto es un mínimo local de forma necesaria y suficiente.
```
```{admonition} Ejercicio
:class: tip
Realizar un análisis similar al anterior para la función $f_o(x) = x_2^4+2x_2^2x_1^2+x_1^2$.
```
### Ejemplo importante
$$\displaystyle \min_{x \in \mathbb{R}^2} \frac{1}{2}x^TPx+q^Tx+r$$
donde: $P=\left [\begin{array}{cc} 5 & 4\\ 4 & 5 \end{array} \right ]$, $q=\left [\begin{array}{c} -1\\ 1 \end{array} \right]$, $r=3$.
Haciendo las multiplicaciones de matriz-vector y productos punto
```
P = sympy.Matrix([[5, 4],
[4, 5]])
x = sympy.Matrix(sympy.symbols("x1, x2"))
q = sympy.Matrix([-1,1])
r = 3
fo_sympy = (1/2*x.T*P*x + q.T*x)[0] + r
sympy.pprint(fo_sympy.expand())
```
se **reescribe** el problema de optimización como:
$$\displaystyle \min_{x \in \mathbb{R}^2} \frac{5}{2}x_1^2 + \frac{5}{2}x_2^2+4x_1x_2 -x_1 + x_2+3$$
La función objetivo es una **función estrictamente convexa** en $\mathbb{R}^2$ (de hecho **fuertemente convexa**) pues:
```{margin}
Los eigenvalores de $P$ son $1$ y $9$ de multiplicidad simple cada uno.
```
```
sympy.pprint(P.eigenvals())
```
y la Hessiana de $f_o$ es:
```
sympy.pprint(derive_by_array(derive_by_array(fo_sympy, (x1,x2)),
(x1,x2))
)
```
El gradiente de $f_o$ es:
```
sympy.pprint(fo_sympy.diff(x))
```
El mínimo debe satisfacer la ecuación **lineal** con dos variables y dos ecuaciones
$$\nabla f_o(x) = Px +q =\left [ \begin{array}{cc}
5 & 4\\
4 & 5
\end{array}
\right ]
\left [ \begin{array}{c}
x_1\\
x_2
\end{array}
\right ]
+ \left [ \begin{array}{c}
-1\\
1
\end{array}
\right ]=
\left [ \begin{array}{cc}
5x_1+4x_2-1\\
4x_1+5x_2+1
\end{array}
\right ]
=0
$$
```{admonition} Observación
:class: tip
En algunos casos especiales es posible resolver la ecuación no lineal $\nabla f_o(x) = 0$ para $x$ de forma analítica o cerrada. Este es el caso de este ejemplo cuya solución está dada por $x^* = -P^{-1}q$.
```
```
P = np.array([[5,4],[4,5]])
q = np.array([-1,1])
print(np.linalg.solve(P,-q))
```
El problema anterior también lo podemos resolver con [cvxpy](https://github.com/cvxgrp/cvxpy) pues es un **problema convexo**.
```{margin}
Definición de variables y función objetivo: $\frac{1}{2}x^TPx+q^Tx+r$
```
```
n = 2 #number of variables
x = cp.Variable(n) #optimization variable
fo_cvxpy = (1/2)*cp.quad_form(x, P) + q.T @ x + r#objective function
opt_objective = cp.Minimize(fo_cvxpy) #optimization objective
prob = cp.Problem(opt_objective) #optimization problem
print(prob.solve())
print("status:", prob.status)
print("optimal value", prob.value)
print("optimal var", x.value)
```
````{admonition} Observaciones
:class: tip
* El paquete cvxpy requiere que se especifique el problema de optimización a resolver siguiendo reglas establecidas en [Disciplined Convex Programming](https://dcp.stanford.edu/). En el ejemplo anterior resulta en un error si se ejecutan las líneas siguientes:
```python
x1 = cp.Variable()
x2 = cp.Variable()
fo = 2.5*x1**2 + 4*x1*x2 - x1 + 2.5*x2**2 + x2 + 3
obj = cp.Minimize(fo)
prob = cp.Problem(obj)
prob.solve()
```
La última línea produce
```
Problem does not follow DCP rules. Specifically: The objective is not DCP.
```
* En la liga de [ejemplos](https://www.cvxpy.org/examples/index.html) hay muchos problemas típicos en optimización convexa y en [Atomic Functions](https://www.cvxpy.org/tutorial/functions/index.html) ejemplos de funciones atómicas que pueden aplicarse a expresiones de *CVXPY*.
````
```{admonition} Ejercicio
:class: tip
Utilizando las herramientas (teóricas y prácticas) del ejemplo anterior, resolver el problema de optimización:
$$\min_{x \in \mathbb{R}^2} ||Ax-b||_2^2$$
con $A=\left [ \begin{array}{ccc} 1 & 6 & 2.5\\ 1 & 2 & 8 \\ 1 & 10 & -1\\ 1 & -9 & 3\\ 1 & -1 & 2 \end{array} \right ]$, $b=\left [ \begin{array}{c} -1 \\ 0 \\ 2 \\ 3.5 \\ -1.7 \end{array} \right ]$.
```
## Métodos de descenso para funciones convexas
Los ejemplos anteriores mostraron la importancia de la información de primer y segundo orden de la función objetivo $f_o$ y las ecuaciones no lineales para resolver el problema de optimización. El primer ejemplo utilizó las funciones [scipy.optimize.fsolve](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.fsolve.html#scipy.optimize.fsolve) y [scipy.optimize.fmin](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.fmin.html) para este propósito. Tales funciones utilizan métodos iterativos para resolver ecuaciones no lineales y aproximar un mínimo local respectivamente en un esquema que para el caso de minimización satisface:
$$f_o(x^{(0)}) > f_o(x^{(1)}) > f_o(x^{(2)}) > \cdots > f_o(x^{(k)}) > \cdots$$
con $x^{(0)}$ punto inicial.
En lo siguiente se asume que $f_o$ cumple $f_o \in \mathcal{C}^2(\text{dom}f_o)$ y es convexa en un conjunto convexo y cerrado que contiene a $x^*$. Ver {ref}`conjunto abierto, cerrado, cerradura e interior <CACCI>` para definición de conjunto cerrado.
### Ejemplo de función objetivo convexa
Encontrar el mínimo del siguiente problema con un **método iterativo**.
$$\displaystyle \min_{x \in \mathbb{R}^4} (x_1-2)^2+(2-x_2)^2+x_3^2+x_4^4$$
```
x1, x2, x3, x4 = sympy.symbols("x1, x2, x3, x4")
fo_sympy = (x1-2)**2 + (2-x2)**2 + x3**2 + x4**4
gf = derive_by_array(fo_sympy, (x1, x2, x3, x4))
sympy.pprint(gf)
Hf = derive_by_array(gf, (x1, x2, x3, x4))
sympy.pprint(Hf)
```
Como $f_o$ es una función convexa (de hecho **estrictamente convexa**) en su dominio $\mathbb{R}^4$, se tiene que su óptimo se obtiene igualando y resolviendo la **ecuación no lineal** $\nabla f_o(x) = 0$ :
$$\nabla f_o(x) =
\left[ \begin{array}{c}
2x_1-4 \\
2x_2-4\\
2x_3\\
4x_4^3
\end{array}
\right]
= 0
$$
El óptimo $x^* \in \mathbb{R}^4$ está dado por:
$$x^*=
\left[ \begin{array}{c}
2\\
2\\
0\\
0
\end{array}
\right]
$$
**¿Cómo encontramos numéricamente el óptimo con un método iterativo?**
## Métodos de descenso
Los métodos que utilizan esquemas iterativos para calcular una **secuencia de minimización** de puntos $x^{(0)}, x^{(1)}, \dots \in \text{dom}f_o$ con la característica $f_o(x^{(k)}) \rightarrow p^*$ si $k \rightarrow \infty$ se conocen con el nombre de **métodos de descenso**.
```{admonition} Definición
Un método de descenso es aquel que genera la secuencia de minimización $x^{(0)}, x^{(1)}, \dots \in \text{dom}f_o$ la cual cumple con la desigualdad: $f_o(x^{(k+1)}) < f_o(x^{(k)})$ excepto para $x^{(k)}$ óptimo y $f_o(x^{(k)}) \rightarrow p^*$ si $k \rightarrow \infty$.
```
```{admonition} Observación
:class: tip
La definición de una dirección de descenso anterior aplica para funciones en general, no es necesario que $f_o$ sea convexa.
```
```{margin}
Recuérdese que si $f_o$ es fuertemente convexa en el conjunto $\mathcal{S}$ entonces $\nabla^2 f_o (x) \in \mathbb{S}^n_{++}$ y $\text{cond}(\nabla ^2 f_o(x))$ está acotado por arriba por una constante para $x \in \mathcal{S}$.
```
```{admonition} Comentario
Asumiendo que la función $f_o$ es convexa, típicamente se asume lo siguiente para tener métodos iterativos confiables y exactos:
* Los puntos iniciales $x^{(0)}$ están en $\text{dom}f_o$.
* Que el conjunto $f_o(x^{(0)})$-subnivel sea cerrado pues así se garantiza que la secuencia de minimización está en el conjunto $f_o(x^{(0)})$-subnivel para todas las iteraciones.
* $f_o$ fuertemente convexa en el conjunto $f_o(x^{(0)})$-subnivel para tener propiedades dadas en los {ref}`resultados que son posibles probar para funciones fuertemente convexas <RESFFUERTCON>`.
```
### Condición para que un paso o dirección de búsqueda sea de descenso
La idea de los métodos de optimización es calcular direcciones $\Delta x$ de búsqueda que sean de descenso, esto es, que al movernos de un punto a otro en tal dirección, el valor de $f_o$ decrece. Existen muchas direcciones de descenso (de hecho infinitas) una que se muestra en el dibujo siguiente es la dirección de descenso de Newton $\Delta x_{nt}$:
<img src="https://dl.dropboxusercontent.com/s/25bmebx645howjw/direccion_de_descenso_de_Newton_1d.png?dl=0" heigth="600" width="600">
En el dibujo $f = f_o$ y $\hat{f}$ es un **modelo cuadrático**. Del punto $(x,f(x))$ nos debemos mover al punto $(x+\Delta x_{nt}, f(x + \Delta x_{nt}))$ para llegar al óptimo. En tal dirección $f$ decrece: $f(x+\Delta x_{nt}) < f(x)$ y obsérvese que $\Delta x_{nt}$ es mínimo de $\hat{f}$.
```{margin}
Ver {ref}`teorema de Taylor para una función de varias variables <TEOTAYLORNVARIABLES>`
```
```{admonition} Comentario
El modelo cuadrático del dibujo anterior está dado por la aproximación de segundo orden a la función $f_o$ por el teorema de Taylor con centro en $x$:
$$m(x + v) = \hat{f}_o(x + v) = f_o(x) + \nabla f_o(x)^T v + \frac{1}{2} v^T \nabla^2f_o(x)v$$
con único mínimo si $\nabla ^2 f_o(x) \in \mathbb{S}^n_{++}$ dado por $v^* = \Delta x_{nt}$ y $\Delta x_{nt}$ dirección de Newton cuya expresión está más adelante. En cada iteración se construye un modelo cuadrático:
$$m(x^{(k)} + v) = \hat{f}_o(x^{(k)} + v) = f_o(x^{(k)}) + \nabla f_o(x^{(k)})^T v + \frac{1}{2} v^T \nabla^2f_o(x^{(k)})v$$
```
Geométricamente las direcciones de descenso forman un **ángulo agudo** con $-\nabla f_o(x)$:
<img src="https://dl.dropboxusercontent.com/s/eednhn6lj1rag1j/zone_for_descent_directions.png?dl=0" heigth="350" width="350">
En el dibujo $f = f_o$.
```{admonition} Observación
:class: tip
Aunque se tienen una cantidad infinita de direcciones de descenso, las direcciones de descenso que típicamente son elegidas no son cercanas a ser ortogonales con el gradiente de $f_o$.
```
Tenemos una condición para garantizar que una dirección sea de descenso:
```{admonition} Definición
Si el paso o dirección de búsqueda satisface: $\nabla f_o(x)^T\Delta x < 0$ se le nombra **dirección de descenso**.
```
```{margin}
Recuérdese que el teorema de Taylor nos ayuda a aproximar a una función de forma **local**.
```
```{admonition} Comentarios
* Recuérdese que $\nabla f_o(x)^T \Delta x$ es una **derivada direccional** de $f_o$ en $x$ en la dirección $\Delta x$, ver {ref}`ejemplo función restringida a una línea <EJRestriccionALinea>`.
* La definición anterior se justifica pues recuérdese que por la aproximación del teorema de Taylor a primer orden se tiene:
$$f_o(x + \Delta x) \approx f_o(x) + \nabla f_o(x) ^T \Delta x$$
y si $\Delta x$ es dirección de descenso entonces: $f_o(x) + \nabla f_o(x) ^T \Delta x < f_o(x)$.
```
```{admonition} Observación
:class: tip
Obsérvese que si $x^*$ es mínimo local entonces $\nabla f_o(x^*) = 0$ (condición necesaria de primer orden) por lo que no existen direcciones de descenso.
```
### Ejemplos de direcciones de descenso
```{sidebar} La dirección del gradiente...
Una forma de obtener la dirección de Newton es encontrando el mínimo del modelo cuadrático referido anteriormente asumiendo $\nabla ^2 f_o(x) \in \mathbb{S}^n_{++}$. La dirección del gradiente se obtiene al resolver el problema de optimización con restricciones siguiente:
$$\min_{v \in \mathbb{R}^n} \nabla f_o(x)^T v$$
$$\text{sujeto a:} ||v|| = 1$$
para la norma $2$. Se utiliza una restricción del tipo normalización pues la función objetivo involucra un producto punto (derivada direccional) que es dependiente o proporcional a la longitud de $v$.
```
* $\Delta x = - \nabla f_o \left (x^{(k)} \right )$ que da lugar al **método de descenso en gradiente** para $x^{(k)}$ no óptimo.
* $\Delta x = - \nabla^2 f_o \left (x^{(k)} \right )^{-1} \nabla f_o\left(x^{(k)} \right)$ que da lugar al **método de descenso por Newton** con $\nabla^2 f_o \left (x^{(k)} \right ) \in \mathbb{S}^n_{++}$ y $x^{(k)}$ no óptimo.
* $\Delta x = - H_k ^{-1} \nabla f_o\left(x^{(k)}\right)$ con $H_k$ aproximación a la Hessiana de $f_o$ con $\nabla^2 f_o \left (x^{(k)} \right ) \in \mathbb{S}^n_{++}$ y $x^{(k)}$ no óptimo.
```{admonition} Observaciones
:class: tip
* La definición de una dirección de descenso aplica para funciones en general, no es necesario que $f_o$ sea convexa.
* Para funciones en general, la dirección de Newton es de descenso si la Hessiana es definida positiva y análogamente para las direcciones en las que se utilicen aproximaciones a la Hessiana. Esto asegura que el **modelo cuadrático** tenga un único mínimo y que $f_o$ decrezca su valor en tal dirección.
* Comúnmente los métodos que utilizan aproximaciones a la Hessiana se conocen con el nombre de **métodos Cuasi-Newton**, ver [Quasi-Newton_method](https://en.wikipedia.org/wiki/Quasi-Newton_method).
```
### Continuando con el ejemplo anterior
Encontrar el mínimo del siguiente problema con un **método iterativo**.
$$\displaystyle \min_{x \in \mathbb{R}^4} (x_1-2)^2+(2-x_2)^2+x_3^2+x_4^4$$
**Opción descenso en gradiente**: usando la dirección del gradiente de $f_o$ se tiene:
$$x^{(k+1)} = x^{(k)} - \nabla f_o(x^{(k)})$$
Tomando $x^{(0)} = \left [ \begin{array}{c} 5 \\ 5 \\1 \\ 0.1 \\ \end{array} \right ]$ como punto inicial:
```
x_0 = np.array([5,5,1,0.1])
```
```{margin}
Definición de $f_o$
```
```
f_o_np = lambda x: (x[0]-2)**2 + (2-x[1])**2 + x[2]**2 + x[3]**2
```
```{margin}
Evaluando $f_o$ en $x^{(0)}$.
```
```
print(f_o_np(x_0))
gf_eval = lambda x: np.array([partial_derivative.subs({"x1": x[0],
"x2": x[1],
"x3": x[2],
"x4": x[3]}) for partial_derivative in gf],
dtype=float)
```
```{margin}
Evaluando el gradiente en $x^{(0)}$: $\nabla f_o(x^{(0)})$.
```
```
print(gf_eval(x_0))
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(0)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla f_o \left (x^{(0)} \right )$.
```
```
print(gf_eval(x_0).dot(-gf_eval(x_0)))
```
**Primera iteración**
```{margin}
Esquema iterativo: $x_1 = x_0 + \Delta x = x_0 - \nabla f_o(x^{(0)})$.
```
```
x_1 = x_0 - gf_eval(x_0)
print(x_1)
```
```{margin}
Evaluando $f_o$ en $x^{(1)}$ se observa que $f_o$ decreció muy poco, de hecho $f_o(x^{(0)}) \approx f_o(x^{(1)})$.
```
```
print(f_o_np(x_0))
print(f_o_np(x_1))
```
Obsérvese que la aproximación a primer orden por Taylor no es correcta: $f_o(x_0 + \Delta x) \neq f_o(x_0) + \nabla f_o(x_0)^T \Delta x$.
```
print(f_o_np(x_0) + gf_eval(x_0).dot(-gf_eval(x_0)))
```
**Segunda iteración**
```{margin}
Evaluando el gradiente en $x^{(1)}$: $\nabla f_o(x^{(1)})$.
```
```
print(gf_eval(x_1))
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(1)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla f_o \left (x^{(1)} \right )$.
```
```
print(gf_eval(x_1).dot(-gf_eval(x_1)))
```
```{margin}
Esquema iterativo: $x_2 = x_1 + \Delta x = x_1 - \nabla f_o(x^{(1)})$.
```
```
x_2 = x_1 - gf_eval(x_1)
print(x_2)
```
```{margin}
Evaluando $f_o$ en $x^{(1)}$ se observa que $f_o$ decreció muy poco, de hecho $f_o(x^{(2)}) \approx f_o(x^{(1)})$.
```
```
print(f_o_np(x_1))
print(f_o_np(x_2))
```
Obsérvese que la aproximación a primer orden por Taylor no es correcta: $f_o(x_1 + \Delta x) \neq f_o(x_1) + \nabla f_o(x_1)^T \Delta x$.
```
print(f_o_np(x_1) + gf_eval(x_1).dot(-gf_eval(x_1)))
```
**Tercera iteración**
```{margin}
Esquema iterativo: $x_3 = x_2 + \Delta x = x_2 - \nabla f_o(x^{(2)})$.
```
```
x_3 = x_2 - gf_eval(x_2)
```
```{margin}
Obsérvese que $x_3 \approx x_1$.
```
```
print(x_1)
print(x_3)
```
**Cuarta iteración**
```{margin}
Esquema iterativo: $x_4 = x_3 + \Delta x = x_3 - \nabla f_o(x^{(3)})$.
```
```
x_4 = x_3 - gf_eval(x_3)
```
```{margin}
Obsérvese que $x_4 \approx x_2$.
```
```
print(x_2)
print(x_4)
```
y así nos quedaremos ciclando por muchas iteraciones...
```{admonition} Comentario
El método de descenso en gradiente para el ejemplo anterior no convergerá al óptimo $x^* = \left [ \begin{array}{c} 2 \\ 2 \\0 \\ 0 \\ \end{array} \right ]$
```
```{admonition} Ejercicio
:class: tip
¿Converge el método de descenso en gradiente al óptimo para un punto inicial $x^{(0)} = \left [ \begin{array}{c} 2.5 \\ 2.5 \\0.001 \\ 0.001 \\ \end{array} \right ]$?
```
**Opción descenso por dirección de Newton:** usando la dirección de descenso de Newton de $f_o$ se tiene:
$$x^{(k+1)} = x^{(k)} - \nabla^2 f_o \left (x^{(k)} \right )^{-1} \nabla f_o\left(x^{(k)} \right)$$
Con
$$
\nabla ^2 f_o(x) =
\left [
\begin{array}{cccc}
2 & 0 & 0 & 0 \\
0 & 2 & 0 & 0 \\
0 & 0 & 2 & 0 \\
0 & 0 & 0 & 12x_4^2
\end{array}
\right ]
$$
Tomando $x^{(0)} = \left [ \begin{array}{c} 5 \\ 5 \\1 \\ 0.1 \\ \end{array} \right ]$ como punto inicial y **no calculando la inversa** de la Hessiana pues en su lugar resolvemos el sistema de ecuaciones lineales $\nabla ^2 f_o \left ( x^{(k)} \right ) \Delta x = - \nabla f_o \left(x^{(k)}\right )$ resulta
```
x_0 = np.array([5,5,1,0.1])
Hf_eval = lambda x: np.array([second_partial_derivative.subs({"x1": x[0],
"x2": x[1],
"x3": x[2],
"x4": x[3]}) for second_partial_derivative in Hf],
dtype=float)
```
```{margin}
Evaluando la Hessiana en $x^{(0)}$: $\nabla^2f_o(x^{(0)})$.
```
```
Hf_sympy_eval = sympy.Matrix(Hf_eval(x_0))
sympy.pprint(Hf_sympy_eval)
```
```{margin}
Evaluando el gradiente en $x^{(0)}$: $\nabla f_o(x^{(0)})$.
```
```
gf_sympy_eval = gf_eval(x_0)
sympy.pprint(gf_sympy_eval)
```
El sistema de **ecuaciones lineales** a resolver es:
$$
\left [
\begin{array}{cccc}
2 & 0 & 0 & 0 \\
0 & 2 & 0 & 0 \\
0 & 0 & 2 & 0 \\
0 & 0 & 0 & 0.12
\end{array}
\right ]
\Delta x =
-\left [
\begin{array}{c}
6 \\
6 \\
2 \\
0.004
\end{array}
\right ]
$$
Resolviendo con *NumPy* el sistema de ecuaciones lineales:
```{margin}
Aquí convertimos de un objeto de *SymPy* a un *array* de *NumPy* pues si bien podríamos resolver el sistema con *SymPy* es menos costoso utilizar *arrays* de *NumPy*.
```
```
Hf_np_eval = np.array(Hf_sympy_eval, dtype=float)
_, n = Hf_np_eval.shape
```
```{margin}
Aquí convertimos de un objeto de *SymPy* a un *array* de *NumPy* pues si bien podríamos resolver el sistema con *SymPy* es menos costoso utilizar *arrays* de *NumPy*.
```
```
gf_np_eval = np.array(gf_sympy_eval, dtype = float)
gf_np_eval
```
```{margin}
Resolvemos $\nabla ^2 f_o \left ( x^{(0)} \right ) \Delta x = - \nabla f_o \left(x^{(0)}\right )$ para obtener $\Delta x$ dirección de Newton.
```
```
dir_Newton = np.linalg.solve(Hf_np_eval, -gf_np_eval)
print(dir_Newton)
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(0)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla ^2f_o(x^{(0)})^{-1} \nabla f_o(x^{(0)})$.
```
```
print(gf_np_eval.dot(dir_Newton))
```
**Primera iteración**
```{margin}
Esquema iterativo: $x_1 = x_0 + \Delta x = x_0 - \nabla ^2f_o(x^{(0)})^{-1} \nabla f_o(x^{(0)})$
```
```
x_1 = x_0 + dir_Newton
print(x_1)
```
Recuérdese que **siempre** es útil monitorear el número de condición de la matriz del sistema de ecuaciones lineales que en este caso es la Hessiana de $f_o$ en $x^{(0)}$ para confiabilidad de nuestros cálculos al resolver el sistema de ecuaciones lineales asociado, ver {ref}`Número de condición de una matriz <NCM>`:
```
print(np.linalg.cond(Hf_np_eval))
```
```{margin}
Evaluando $f_o$ en $x^{(1)}$ se observa que $f_o$ sí decrece $f_o(x^{(1)}) < f_o(x^{(0)})$.
```
```
print(f_o_np(x_0))
print(f_o_np(x_1))
```
Obsérvese que la aproximación a segundo orden por Taylor es cercana: $f_o(x_0 + \Delta x) \approx f_o(x_0) + \nabla f_o(x_0)^T \Delta x + \frac{1}{2}\Delta x ^T\nabla^2 f_o(x_0) \Delta x$.
```
print(f_o_np(x_0) + gf_np_eval.dot(dir_Newton) + 1/2*dir_Newton.dot(Hf_np_eval@dir_Newton))
```
**Segunda iteración**
```{margin}
Evaluando la Hessiana en $x^{(1)}$: $\nabla^2f_o(x^{(1)})$, evaluando el gradiente en $x^{(1)}$: $\nabla f_o(x^{(1)})$ y resolviendo el sistema de ecuaciones $\nabla ^2 f_o \left ( x^{(1)} \right ) \Delta x = - \nabla f_o \left(x^{(1)}\right )$.
```
```{margin}
Esquema iterativo: $x_2 = x_1 + \Delta x = x_1 - \nabla ^2f_o(x^{(1)})^{-1} \nabla f_o(x^{(1)})$
```
```
Hf_sympy_eval = sympy.Matrix(Hf_eval(x_1))
gf_sympy_eval = gf_eval(x_1)
Hf_np_eval = np.array(Hf_sympy_eval, dtype=float)
gf_np_eval = np.array(gf_sympy_eval, dtype = float)
dir_Newton = np.linalg.solve(Hf_np_eval, -gf_np_eval)
x_2 = x_1 + dir_Newton
print(x_2)
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(1)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla ^2f_o(x^{(1)})^{-1} \nabla f_o(x^{(1)})$.
```
```
print(gf_np_eval.dot(dir_Newton))
```
```{margin}
Evaluando $f_o$ en $x^{(2)}$ se observa que $f_o$ sí decrece $f_o(x^{(2)}) < f_o(x^{(1)})$.
```
```
print(f_o_np(x_1))
print(f_o_np(x_2))
```
Obsérvese que la aproximación a segundo orden por Taylor es cercana: $f_o(x_1 + \Delta x) \approx f_o(x_1) + \nabla f_o(x_1)^T \Delta x + \frac{1}{2}\Delta x ^T\nabla^2 f_o(x_1) \Delta x$.
```
print(f_o_np(x_1) + gf_np_eval.dot(dir_Newton) + 1/2*dir_Newton.dot(Hf_np_eval@dir_Newton))
```
```{margin}
Número de condición de la Hessiana de $f_o$ en $x^{(1)}$.
```
```
print(np.linalg.cond(Hf_np_eval))
```
**Tercera iteración**
```{margin}
Evaluando la Hessiana en $x^{(2)}$: $\nabla^2f_o(x^{(2)})$, evaluando el gradiente en $x^{(2)}$: $\nabla f_o(x^{(2)})$ y resolviendo el sistema de ecuaciones $\nabla ^2 f_o \left ( x^{(2)} \right ) \Delta x = - \nabla f_o \left(x^{(2)}\right )$.
```
```{margin}
Esquema iterativo: $x_3 = x_2 + \Delta x = x_2 - \nabla ^2f_o(x^{(2)})^{-1} \nabla f_o(x^{(2)})$
```
```
Hf_sympy_eval = sympy.Matrix(Hf_eval(x_2))
gf_sympy_eval = gf_eval(x_2)
Hf_np_eval = np.array(Hf_sympy_eval, dtype=float)
gf_np_eval = np.array(gf_sympy_eval, dtype = float)
dir_Newton = np.linalg.solve(Hf_np_eval, -gf_np_eval)
x_3 = x_2 + dir_Newton
print(x_3)
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(2)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla ^2f_o(x^{(2)})^{-1} \nabla f_o(x^{(2)})$.
```
```
print(gf_np_eval.dot(dir_Newton))
```
```{margin}
Evaluando $f_o$ en $x^{(3)}$ se observa que $f_o$ sí decrece $f_o(x^{(3)}) < f_o(x^{(2)})$.
```
```
print(f_o_np(x_2))
print(f_o_np(x_3))
```
Obsérvese que la aproximación a segundo orden por Taylor es cercana: $f_o(x_2 + \Delta x) \approx f_o(x_2) + \nabla f_o(x_2)^T \Delta x + \frac{1}{2}\Delta x ^T\nabla^2 f_o(x_2) \Delta x$.
```
print(f_o_np(x_2) + gf_np_eval.dot(dir_Newton) + 1/2*dir_Newton.dot(Hf_np_eval@dir_Newton))
```
```{margin}
Número de condición de la Hessiana de $f_o$ en $x^{(2)}$.
```
```
print(np.linalg.cond(Hf_np_eval))
```
**Cuarta iteración**
```{margin}
Evaluando la Hessiana en $x^{(3)}$: $\nabla^2f_o(x^{(3)})$, evaluando el gradiente en $x^{(3)}$: $\nabla f_o(x^{(3)})$ y resolviendo el sistema de ecuaciones $\nabla ^2 f_o \left ( x^{(3)} \right ) \Delta x = - \nabla f_o \left(x^{(3)}\right )$.
```
```{margin}
Esquema iterativo: $x_4 = x_3 + \Delta x = x_3 - \nabla ^2f_o(x^{(3)})^{-1} \nabla f_o(x^{(3)})$
```
```
Hf_sympy_eval = sympy.Matrix(Hf_eval(x_3))
gf_sympy_eval = gf_eval(x_3)
Hf_np_eval = np.array(Hf_sympy_eval, dtype=float)
gf_np_eval = np.array(gf_sympy_eval, dtype = float)
dir_Newton = np.linalg.solve(Hf_np_eval, -gf_np_eval)
x_4 = x_3 + dir_Newton
print(x_4)
```
```{margin}
Verificando que es dirección de descenso: $\nabla f_o \left (x^{(3)} \right )^T \Delta x < 0$ con $\Delta x = -\nabla ^2f_o(x^{(3)})^{-1} \nabla f_o(x^{(3)})$.
```
```{margin}
Obsérvese que el gradiente y la dirección de Newton son cercanos a ser ortogonales.
```
```
print(gf_np_eval.dot(dir_Newton))
```
```{margin}
Evaluando $f_o$ en $x^{(4)}$ se observa que $f_o$ sí decrece $f_o(x^{(4)}) < f_o(x^{(3)})$.
```
```
print(f_o_np(x_3))
print(f_o_np(x_4))
```
Obsérvese que la aproximación a segundo orden por Taylor es cercana: $f_o(x_3 + \Delta x) \approx f_o(x_3) + \nabla f_o(x_3)^T \Delta x + \frac{1}{2}\Delta x ^T\nabla^2 f_o(x_3) \Delta x$.
```
print(f_o_np(x_3) + gf_np_eval.dot(dir_Newton) + 1/2*dir_Newton.dot(Hf_np_eval@dir_Newton))
```
```{margin}
Obsérvese cómo va aumentando el número de condición de la Hessiana conforme nos aproximamos a la solución, en este paso se ha calculado la Hessiana de $f_o$ en $x^{(3)}$.
```
```
print(np.linalg.cond(Hf_np_eval))
print(x_4 == approx(np.array([2,2,0,0.0]), abs=1e-1, rel=1e-1))
```
```{admonition} Comentario
El método por dirección de Newton sí convergerá al óptimo $x^* = \left [ \begin{array}{c} 2 \\ 2 \\0 \\ 0 \\ \end{array} \right ]$ pero la convergencia será lenta.
```
Si hubiéramos elegido como punto inicial $x^{(0)} = \left [ \begin{array}{c} 5 \\ 5 \\1 \\ 0 \\ \end{array} \right ]$
```
x_0 = np.array([5, 5, 1, 0])
```
```{margin}
Evaluando la Hessiana en $x^{(0)}$: $\nabla^2f_o(x^{(0)})$.
```
```
Hf_sympy_eval = sympy.Matrix(Hf_eval(x_0))
sympy.pprint(Hf_sympy_eval)
```
```{margin}
Evaluando el gradiente en $x^{(0)}$: $\nabla f_o(x^{(0)})$.
```
```
gf_sympy_eval = sympy.Matrix(gf_eval(x_0))
sympy.pprint(gf_sympy_eval)
```
El sistema de **ecuaciones lineales** a resolver es:
$$
\left [
\begin{array}{cccc}
2 & 0 & 0 & 0 \\
0 & 2 & 0 & 0 \\
0 & 0 & 2 & 0 \\
0 & 0 & 0 & 0
\end{array}
\right ]
\Delta x =
-\left [
\begin{array}{c}
6 \\
6 \\
2 \\
0
\end{array}
\right ]
$$
**Obsérvese que la última ecuación es redundante por lo que una solución al sistema anterior es considerar $x_4=0$ y resolver:**
$$
\left [
\begin{array}{ccc}
2 & 0 & 0 \\
0 & 2 & 0 \\
0 & 0 & 2 \\
\end{array}
\right ]
\Delta x =
-\left [
\begin{array}{c}
6 \\
6 \\
2 \\
\end{array}
\right ]
$$
```
x = sympy.Matrix([x1, x2, x3, x4])
```
```{margin}
*SymPy* nos permite obtener soluciones a sistemas de ecuaciones lineales que tienen un renglón y columna de ceros.
```
```
sympy.pprint(sympy.solve(Hf_sympy_eval*x - (-gf_sympy_eval), x))
```
```{margin}
Esquema iterativo: $x_1 = x_0 + \Delta x = x_0 - \nabla ^2f_o(x^{(0)})^{-1} \nabla f_o(x^{(0)})$
```
```
x_1 = x_0 + np.array([-3, -3, -1, 0])
```
```{margin}
$x_1$ es el óptimo del problema.
```
```
print(x_1)
```
```{margin}
El número de condición es $\infty$, con el `print` de *SymPy* se ve así.
```
```
sympy.pprint(Hf_sympy_eval.condition_number())
print(x_1 == approx(np.array([2,2,0,0.0]), abs=1e-4, rel=1e-4))
```
```{admonition} Comentarios
De acuerdo al ejemplo anterior:
* Utilizar información de primer o segundo orden nos ayuda a encontrar óptimo(s) de funciones.
* El método de descenso en gradiente no converge para el punto inicial elegido. En el caso del método de Newton sí hay convergencia pero es lenta si el punto inicial tiene en la última entrada un número cercano a $0$.
* La ventaja que tiene utilizar la dirección del gradiente vs la dirección de Newton es que el gradiente involucra menos almacenamiento en memoria que el almacenamiento de la Hessiana: $\mathcal{O}(n)$ vs $\mathcal{O}(n^2)$.
```
### Resolviendo el problema con [*CVXPY*](https://github.com/cvxgrp/cvxpy)
```
x1 = cp.Variable()
x2 = cp.Variable()
x3 = cp.Variable()
x4 = cp.Variable()
fo_cvxpy = (x1 -2)**2 + (2-x2)**2 + x3**2 + x4**2
obj = cp.Minimize(fo_cvxpy)
prob = cp.Problem(obj)
print(prob.solve())
print("status:", prob.status)
print("optimal value", prob.value)
print("optimal var", x1.value, x2.value, x3.value, x4.value)
```
## Tamaño o longitud de paso
En el ejemplo anterior en el que se aproximó al mínimo del siguiente problema con el método de Newton
$$\displaystyle \min_{x \in \mathbb{R}^4} (x_1-2)^2+(2-x_2)^2+x_3^2+x_4^4$$
se concluyó que tal método converge de forma lenta y el método de descenso en gradiente no converge para el punto inicial elegido en ambos métodos. La pequeña reducción que se obtenía en $f_o$ por cada iteración fue la razón de tal situación en el caso del descenso en gradiente. Una metodología que resuelve la no convergencia del método de descenso en gradiente utiliza el siguiente esquema iterativo:
$$x^{(k+1)} = x^{(k)} + t^{(k)}\Delta x$$
con $t^{(k)}>0$.
```{admonition} Comentario
Para el caso del método de Newton, la convergencia cuadrática depende además de elegir tamaños de pasos adecuados que:
* las iteraciones vayan aproximándose a $x^*$,
* la función objetivo sea dos veces diferenciable y su Hessiana sea definida positiva en un conjunto abierto que contenga a $x^*$ y sea *Lipschitz* continua en tal conjunto, ver [Lipschitz_continuity ](https://en.wikipedia.org/wiki/Lipschitz_continuity), que ayuda a acotar la diferencia entre $f_o$ y el modelo cuadrático $m$.
```
### Continuando con el ejemplo anterior
Usando el método de descenso en gradiente de $f_o$ y el esquema iterativo:
$$x^{(k+1)} = x^{(k)} - t^{(k)} \nabla f_o(x^{(k)})$$
con $t^{(0)} = 0.5$ con punto inicial $x^{(0)} = \left [ \begin{array}{c} 5 \\ 5 \\1 \\ 0.1 \\ \end{array} \right ]$ se tiene:
```
x_0 = np.array([5,5,1,0.1])
t_0=0.5
x_1 = x_0 - t_0*gf_eval(x_0)
print(x_1)
print(x_1 == approx(np.array([2,2,0,0.0]), abs=1e-1, rel=1e-1))
```
por lo que llegamos a aproximar al óptimo en una iteración.
```{admonition} Observación
:class: tip
Para problemas *small scale* el método de Newton o Cuasi-Newton son muy superiores al método de descenso en gradiente. Sin embargo para problemas *large scale* el método de descenso en gradiente es utilizado ampliamente en aplicaciones de *machine learning*.
```
```{admonition} Definición
Al escalar $t^{(k)}$ se le nombra **tamaño o longitud de paso** y siempre es positivo salvo en el caso en que $x^{(k)}$ sea óptimo.
```
El valor $t^{(k)}$ se calcula con metodologías como búsqueda de línea o regiones de confianza, ver [line search](https://en.wikipedia.org/wiki/Line_search), [trust region](https://en.wikipedia.org/wiki/Trust_region) y en esta nota se revisa la búsqueda de línea con *backtracking*.
### ¿Por qué funciona lo anterior?
La condición para que una dirección produzca descenso en $f_o$, $f_o(x^{(k+1)}) < f_o(x^{(k)})$, no es la única para que la secuencia de minimización converja a un mínimo de $f_o$. El siguiente ejemplo muestra el comportamiento del ejemplo trabajado antes en el plano.
Considérese $f_o(x) = x^2$, $x^{(0)} = 2$ y los pasos dados por la secuencia: $(-1)^k(1+2^{-k})$. Entonces:
```
f_o = lambda x:x**2
n = 10
def minimization_sequence():
for k in range(n):
yield (-1)**k*(1+2**(-k))
t = np.linspace(-2.3, 2.3, 100)
plt.plot(t, f_o(t))
[plt.scatter(s, f_o(s)) for s in minimization_sequence()]
plt.title("Secuencia de minimización que no converge al mínimo de $f_o$")
plt.annotate('$x^{(0)}$',(2, f_o(2)),fontsize=12)
plt.annotate('$x^{(1)}$',(-1.5, f_o(-1.5)),fontsize=12)
plt.annotate('$x^{(2)}$',(1.25, f_o(1.25)),fontsize=12)
plt.show()
```
para esta secuencia se cumple $f_o(x^{(k+1)}) < f_o(x^{(k)})$ por lo que es una secuencia de minimización pero no converge tal secuencia pues oscila en los valores $\approx -1, 1$.
El problema en este caso es que en cada iteración es muy pequeña la reducción que se obtiene en $f_o$ relativo a la longitud de los pasos. Esto puede arreglarse al requerir que se cumpla una reducción en $f_o$ al moverse de $f_o(x^{(k)})$ hacia $f_o(x^{(k+1)})$ de al menos una cantidad (reducción suficiente).
```{admonition} Comentario
Otra dificultad relacionada con la reducción de $f_o$ y la longitud de los pasos es la situación en la que los pasos sean muy pequeños relativos a la reducción de $f_o$. En la implementación por *backtracking* que se revisa a continuación sólo se soluciona la reducción suficiente de $f_o$.
```
(MBUSLINBACK)=
## Método de búsqueda de línea por *backtracking*
```{margin}
Ver [Backtracking_line_search](https://en.wikipedia.org/wiki/Backtracking_line_search).
```
En esta sección consideramos $f = f_o$.
La idea del método de búsqueda de línea por *backtracking* es moverse de un punto a otro siguiendo una dirección de descenso con un tamaño de paso completo y si tal tamaño no satisface un criterio entonces realizar un *backtrack* de forma sistemática a lo largo de tal dirección hasta satisfacer el criterio resultando en el tamaño de paso correspondiente.
### Descenso suficiente
Para entender el método de búsqueda de línea por *backtracking* supóngase que $f$ tiene una forma siguiente:
<img src="https://dl.dropboxusercontent.com/s/0woqoj8foo5eco9/level_set_of_func.png?dl=0" heigth="300" width="300">
```{margin}
Un rayo es el conjunto definido por $\{x + \theta v : \theta \geq 0 , v \neq 0, v \in \mathbb{R}^n\}$ para $x \in \mathbb{R}^n$.
<img src="https://dl.dropboxusercontent.com/s/l3z9j49ldzknmif/ej_rayo.png?dl=0" heigth="200" width="200">
```
Y considérese una función $g: \mathbb{R} \rightarrow \mathbb{R}$ igual a $f$ pero restringida al **rayo** $x + t\Delta x$, esto es: $g(t) = f(x+t \Delta x)$ con $t>0$, $\Delta x$ dirección de descenso. Lo anterior se visualiza como sigue:
<img src="https://dl.dropboxusercontent.com/s/18udjmzmmd7drrz/line_search_backtracking_1.png?dl=0" heigth="300" width="300">
```{admonition} Observación
:class: tip
Obsérvese en el dibujo que $\nabla f(x) \neq 0$.
```
```{margin}
Ver {ref}`ejemplo<EJRestriccionALinea>` de la nota {ref}`Definición de función, continuidad y derivada <FCD>` para expresión de la derivada $g'(t)$.
```
Y como $f$ es continua y diferenciable, $g$ también lo es y $g(0)=f(x)$, $g'(t) = \nabla f(x+t\Delta x)^T \Delta x$. Si graficamos $g$ se tendría:
<img src="https://dl.dropboxusercontent.com/s/sgj7eqr2qysi8hs/line_search_backtracking_2.png?dl=0" heigth="300" width="300">
En la búsqueda de línea se construyen dos rectas. Una recta es $g(0) + \alpha g'(0)(t-0)$ con $\alpha \in (0,\frac{1}{2})$. La otra recta es $g(0)+g'(0)(t-0)$. Ambas rectas tienen pendiente negativa. Esto se visualiza como sigue:
<img src="https://dl.dropboxusercontent.com/s/11y008lq0fd6jl6/line_search_backtracking_3.png?dl=0" heigth="500" width="500">
Se busca $t^{(k)}$ tal que $f$ **decrezca suficientemente**. Lo anterior se establece con la desigualdad:
$$f(x+t \Delta x) < f(x) + \alpha t \nabla f(x)^T \Delta x$$
que se nombra **condición de Armijo**:
<img src="https://dl.dropboxusercontent.com/s/o4f341x1y5sqxt8/line_search_backtracking_4.png?dl=0" heigth="500" width="500">
Obsérvese en el dibujo anterior que la región en la que se elegirá $t^{(k)}$ está a la izquierda de la línea punteada vertical de color verde.
Y visualmente en $R^3$ se tiene:
<img src="https://dl.dropboxusercontent.com/s/t3yn7kkpd4il8hx/line_search_backtracking_5.png?dl=0" heigth="300" width="300">
```{admonition} Comentario
La desigualdad de descenso suficiente establece que la reducción debe ser proporcional al tamaño de paso y la derivada direccional $\nabla f(x)^T \Delta x$.
```
### Algoritmo: búsqueda de línea por *backtracking*
El método depende de dos constantes $\alpha$ y $\beta$ con $\alpha \in (0,\frac{1}{2})$ y $\beta \in (0,1)$.
> **Dados** $\Delta x$ dirección de descenso para $f$ en $x \in \text{dom}f$, $\alpha \in (0,\frac{1}{2})$, $\beta \in (0,1)$.
>
> **Asignar** t=1.
>
> **Mientras** $f(x+t\Delta x) > f(x) + \alpha t \nabla f(x) ^T\Delta x$.
>>
>> 1. **Reducir** $t: t= \beta t$.
```{admonition} Comentarios
* El *backtracking* permite tomar tamaños de paso completos, esto es, $t = 1$.
* El valor $\alpha$ típicamente se elige entre $.01$ y $.3$ que indica que se acepta un decrecimiento en el valor de $f$ entre el $1 \%$ y el $30 \%$. La constante $\beta$ comúnmente se elige entre $.1$ (que modifica fuertemente $t$) y $.8$ (que realiza una modificación menos drástica de $t$).
* Aunque la condición de descenso suficiente por sí sola no garantiza que un algoritmo que la utilice realice progreso razonable (pues también hay que añadir una segunda condición para no tomar valores de $t$ muy pequeños) en la implementación del *backtracking* puede incluirse que se revise que $t$ no sea muy pequeño (por ejemplo menor a $10^{-6}$) relativo a $x$, $\Delta x$ y $\nabla f(x)$.
```
## Función en Python para el método de *backtracking*
```python
def line_search_by_backtracking(f,dir_desc,x,
der_direct, alpha=.15, beta=.5):
"""
Line search that sufficiently decreases f restricted to a ray in the direction dir_desc.
Args:
alpha (float): parameter in line search with backtracking, tipically .15
beta (float): parameter in line search with backtracking, tipically .5
f (lambda expression): definition of function f.
dir_desc (array): descent direction.
x (array): numpy array that holds values where line search will be performed.
der_direct (float): directional derivative of f.
Returns:
t (float): positive number for stepsize along dir_desc that sufficiently decreases f.
"""
t = 1
if alpha > 1/2:
print("alpha must be less than or equal to 1/2")
t = -1
if beta > 1:
print("beta must be less than 1")
t = -1;
if t != -1:
eval1 = f(x+t*dir_desc)
eval2 = f(x) + alpha*t*der_direct
while eval1 > eval2:
t = beta*t
eval1 = f(x+t*dir_desc)
eval2 = f(x)+alpha*t*der_direct
return t
```
```{admonition} Ejercicio
:class: tip
Realizar $5$ iteraciones para minimizar la función $log(e^{x_1^2} + e^{x_2^2})$ del método de descenso en gradiente utilizando búsqueda de línea tomando como punto inicial $(10,1)$ y calcular el error relativo de cada iteración. Para calcular $x^*$ utilizar [cvxpy](https://github.com/cvxgrp/cvxpy).
```
## Algoritmo: método general de descenso para problemas UCO
```{margin}
Los pasos de un algoritmo representan una guía para la implementación, no implica que se tengan que implementar uno a continuación del otro como se describe. Si una implementación respeta la lógica y al mismo algoritmo, entonces pueden seguirse los pasos de una forma distinta.
```
> **Dado** un **punto inicial** $x$ en $\text{dom}f_o$
>
> **Repetir** el siguiente bloque para $k=0,1,2,...$
>>
>> 1. Determinar una dirección de descenso $\Delta x$.
>> 2. Búsqueda de línea. Elegir un tamaño de paso $t > 0$.
>> 3. Hacer la actualización: $x = x + t\Delta x$.
>
> **hasta** convergencia (satisfacer criterio de paro).
```{admonition} Comentarios.
* El algoritmo termina si $f_o(x^{(k)})-p^* \leq \epsilon$ con $\epsilon >0$, esto es, $x^{(k)}$ es $\epsilon$-subóptimo.
* El criterio de paro típicamente es de la forma $||\nabla f_o(x)|| \leq tol$ donde: $tol$ es una cantidad pequeña y positiva (comúnmente menor o igual a $10^{-8}$). También se involucra el número máximo de iteraciones en el criterio de paro.
* El paso $2$ busca reducir $f_o$ lo **suficiente** o minimizarla aproximadamente a lo largo de un rayo. La **búsqueda de línea por [*backtracking*](https://en.wikipedia.org/wiki/Backtracking_line_search)** permite esto.
```
**Preguntas de comprehensión.**
1)¿Qué es un método de descenso?
2)Describe los elementos del esquema iterativo de un método de descenso.
3)¿Cómo se obtiene la dirección de Newton en un método de descenso?
4)¿Cómo se obtiene la dirección del gradiente en un método de descenso?
5)¿Qué es un método Cuasi-Newton?
6)¿Cuál es la condición matemática y geométrica que verifica que un paso de búsqueda sea de descenso?
7)¿Qué problemática está resolviendo el método de búsqueda de línea con *backtracking* en cuanto la reducción que se obtiene en $f_o$ relativo a la longitud de los pasos?
**Referencias:**
1. S. P. Boyd, L. Vandenberghe, Convex Optimization, Cambridge University Press, 2009.
2. J. Dennis, R. B. Schnabel, Numerical Methods for Unconstrained Optimization and Nonlinear Equations, SIAM, 1996.
| github_jupyter |
This notebook introduces the notion of computing the general linear model using linear algebra. First we load the necessarily libraries.
```
import numpy,pandas
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats
import statsmodels.api as sm
import statsmodels
from statsmodels.formula.api import ols,glsar
from statsmodels.tsa.arima_process import arma_generate_sample
from scipy.linalg import toeplitz
from IPython.display import display, HTML
%matplotlib inline
```
### A simple example
We start with a simple example of an independent samples t-test. First, let's make a function that will generate some data. We will assume that there are two conditions with specified means and standard deviations
```
def make_ttest_data(n_obs=[50,50],mean_obs=[10,10.1],sd_obs=[2,2]):
"""
function to generate independent-sample data with two conditions
"""
n_obs=[50,50] # number of observations in each condition
n_obs_total=numpy.sum(n_obs)
mean_obs=[10,11]
sd_obs=[1,1]
condition=numpy.zeros(n_obs_total)
condition[:n_obs[0]]=0
condition[n_obs[0]:n_obs_total]=1
data=numpy.zeros(n_obs_total)
data[:n_obs[0]]=mean_obs[0]
data[n_obs[0]:n_obs_total]=mean_obs[1]
# doublecheck our work
assert numpy.sum(data==mean_obs[0])==n_obs[0]
assert numpy.sum(data==mean_obs[1])==n_obs[1]
noise=numpy.zeros(n_obs_total)
noise[:n_obs[0]]=numpy.random.randn(n_obs[0])*sd_obs[0]
noise[n_obs[0]:n_obs_total]=numpy.random.randn(n_obs[1])*sd_obs[1]
df=pandas.DataFrame({'data':data+noise,'condition':condition})
return df
```
Make some data and plot the distributions for the two conditions
```
data=make_ttest_data()
Y=data.data.values
X=data.condition.values
f = plt.figure()
sns.distplot(Y[X==0], hist=False, label="condition 1")
sns.distplot(Y[X==1], hist=False, label="condition 2")
```
Now we want to perform a t-test to ask whether the means of the two conditions are different. Let's try to compute it on our own, using linear algebra. Remember that the formula for the GLM is:
$Y = X * B + e$
where Y is an N X 1 matrix containing the data that we generated, and X is an N X c "design matrix" that describes the conditions (in this case, a single vector indicating condition 1 or 2). Using the normal equations, we can estimate B using:
$\hat{B} = (X'X)^{-1}X'Y$
Before we dive into these computations, we need to go over how to do linear algebra in Python. The following borrows liberally from https://www.ibm.com/developerworks/community/blogs/jfp/entry/Elementary_Matrix_Operations_In_Python?lang=en
#### Making arrays/matrices in Python
```
# to make an array, we give a list to numpy.array
y = numpy.array([1,3,2])
print(y)
print('y shape:',y.shape)
# we can add a dimension with the None operator
z=y[:,None]
print(z)
print('z shape:',z.shape)
# one option to create a matrix is to give a vector and reshape to a matrix
print('A')
A = numpy.array([1,1,2,3,5,8,13,21,34]).reshape(3,3)
print(A)
# another alternative is to pass a list of lists
print('B')
B = numpy.array([[1,1,2],[3,5,8],[13,21,34]])
print(B)
# to transpose a matrix, use the .T operator
print('B.T')
print(B.T)
```
There are some useful functions to generate specific types of matrices
```
# create a matrix full of zeros
# note that the shape is passed as a tuple if you want multiple dimensions
a=numpy.zeros((2,4))
print('a')
print(a)
#create a matrix full of ones
b=numpy.ones((2,4))
print('b')
print(b)
# create a matrix full of any other number:
c=b*12
print('c')
print(c)
# create a range of numbers:
d=numpy.arange(10)
print('d')
print(d)
e=numpy.arange(3,5,0.33)
print('e')
print(e)
```
Now let's look at some basic arithmetic operations
```
print('a+5')
print(a+5)
print('c/2')
print(c/2)
print('a+b+c')
print(a+b+c)
print('a*b*c')
print(a*b*c)
print('b/c')
print(b/c)
```
#### Matrix multiplication
Matrix multiplication is performed on numpy arrays using the .dot() operator.
```
x=numpy.array([[1,2],[3,4]])
y=numpy.array([[1,0],[0,2]])
print('x')
print(x)
print('y')
print(y)
print('scalar product of x and y: x*y')
print(x*y)
print('matrix product of x and y: x.dot(y)')
print(x.dot(y))
print('or use numpy.matrix')
print(numpy.matrix(x)*numpy.matrix(y))
```
__Exercise__: We know that the variance of a matrix X is computed as $mean((X-mean(X))*(X-mean(X))')$. Fill in the appropriate code in the function below so that it returns a value that equals the value obtained from the numpy.var() command.
```
def variance(Y):
# insert code here to estimate variance using matrix multiplication
return var
# use allclose rather than == to deal with numerical errors
assert numpy.allclose(numpy.var(Y),variance(Y))
```
__Exercise__: Write a function to compute the correlation coefficient using matrix algebra. The equation to compute the correlation using matrix algebra is:
$r = \frac{X\cdot Y}{\sqrt{(X\cdot X)*(Y\cdot Y)}}$
assuming that X and Y have zero mean, so you need to remove the mean before computing this.
```
def corrcoef(x,y):
assert len(x)==len(y)
# add code here to compute correlation
return r
print('My result:',corrcoef(X,Y))
print('Numpy result:',numpy.corrcoef(X,Y))
assert numpy.allclose(numpy.corrcoef(X,Y)[0,1],corrcoef(X,Y))
```
#### Matrix inversion
We also need to know how to compute the inverse of a matrix, which we do using numpy.linalg.inv().
__Exercise:__ In the cell below, create a matrix containing the following numbers:
[1,0]
[0,2]
and print out the original matrix along with the inverted matrix.
```
# Exercise code here
```
Now that we know how to perform the necessary matrix operations, let's do our t-test on the data generated above. We first have to fix a problem: we need both X and Y to be matrices for our computation to work, but right now they are 1-dimensional vectors rather than two-dimensional matrices. We can fix this using numpy - let's go ahead and create a function to compute the ordinary least squares estimates, that includes code to reformat the data into matrices. We also include an option to add an intercept (i.e. a column of ones) to the model if it doesn't already exist.
```
def ols_estimate(X,Y,add_intercept=True,verbose=False,
ddof=1,use_two_sided=True):
"""
function to estimate parameters for a general linear model
"""
# first we need to set up the matrices in the proper shape
# Y should be N X 1
# X should be X X c
if verbose:
print('original Y shape:',Y.shape)
Y=Y.reshape((len(Y),1))
if verbose:
print('new Y shape:',Y.shape)
if verbose:
print('original X shape:',X.shape)
if len(X.shape)==1:
X=X.reshape((len(X),1))
Xnames=['X%d'%int(i+1) for i in range(X.shape[1])]
if verbose:
print('new X shape:',X.shape)
# add an intercept to the model if specified
if add_intercept:
X=sm.add_constant(X)
Xnames=Xnames.append('Intercept')
# make sure that the design matrix is full rank
assert numpy.linalg.matrix_rank(X)==X.shape[1]
# estimate the parameters using the normal equations
b_hat=numpy.linalg.inv(X.T.dot(X)).dot(X.T.dot(Y))
if verbose:
print('b_hat=',b_hat)
# compute residuals and their variance
resid=Y-X.dot(b_hat)
sigma2=resid.T.dot(resid)/(X.shape[0] - X.shape[1]) # variance of the residuals
# now compute the t statistic and p values for for each variable in X
t=numpy.zeros(X.shape[1])
p=numpy.zeros(X.shape[1])
for i in range(X.shape[1]):
c=numpy.zeros(X.shape[1])
c[i]=1
t[i]=c.dot(b_hat)/numpy.sqrt(c.dot(numpy.linalg.inv(X.T.dot(X))).dot(c.T)*sigma2)
if t[i]<0:
p[i]=scipy.stats.distributions.t.cdf(t[i],len(Y)-1)
else:
p[i]=1-scipy.stats.distributions.t.cdf(t[i],len(Y)-1)
if use_two_sided:
p[i]=p[i]*2
if verbose:
print('t=',t)
df=pandas.DataFrame({'bhat':b_hat.ravel(),'t':t.ravel(),'p':p.ravel()},index=Xnames)
return df
```
Now let's estimate the model parameters using our function.
```
e=ols_estimate(X,Y)
display(e)
```
Let's compute the same test using a couple of other canned procedures. First, we use the t-test procedure within the scipy package.
```
t,p=scipy.stats.ttest_ind(Y[X==1],Y[X==0])
print('t/p computed by scipy:',t,p)
assert numpy.allclose(t,e.t.values[1])
```
We can also compute it via the general linear model, using the ordinary least squares (OLS) method from statsmodels.
```
X=sm.add_constant(X)
ols_result=sm.OLS(Y,X).fit()
print(ols_result.summary())
# make sure our result is close to the one from statsmodels
for i in range(len(e.t.values)):
assert numpy.allclose(e.t.values[i],ols_result.tvalues[i])
```
__Exercise:__ Confirm that the dot product between the residuals from OLS and the X variable is zero.
```
residual=Y - X.dot(e.bhat.values)
## insert code here
```
### Multiple regression
Let's now look at how we can fit a more complex model using the GLM. Let's make some data based on two regressors plus noise. We will make one of the regressors smooth across observations, for reasons that will become clearer later.
```
def mkar1noise(tslen,coef,noisesd):
"""
function to return AR(1) autocorrelated noise
"""
varcorrect = numpy.sqrt(1-coef**2)
noise=numpy.random.randn(tslen)*noisesd
for i in range(1,tslen):
noise[i]=noise[i]*varcorrect+noise[i-1]*coef
return noise
def make_regression_data(nobs=100,regsmooth=[1],
regsmoothcoef=0.8,
beta=[0.1,0.5,10],noisesd=1.,
noisecorr=0):
"""
function to generate regression data
with option to add autocorrelated noise
beta reflects two conditions plus intercept
"""
regs=numpy.random.randn(nobs,len(beta)-1)
regvarcorrect = numpy.sqrt(1-regsmoothcoef**2)
for r in regsmooth:
for i in range(1,nobs):
regs[i,r]=regs[i,r]*regvarcorrect+regs[i-1,r]*regsmoothcoef
regs=numpy.hstack((regs,numpy.ones((regs.shape[0],1))))
data=regs.dot(numpy.array(beta))
if noisecorr==0:
noise=numpy.random.randn(len(data))*noisesd
else:
noise=mkar1noise(len(data),noisecorr,noisesd)
return data+noise,regs
Y,X=make_regression_data()
#X=X-numpy.mean(X,0)
plt.imshow(X,interpolation='nearest')
plt.axis('auto')
plt.ylim([0,100])
plt.figure()
plt.scatter(X[:,0],Y)
plt.xlabel('first X regressor - X[:,0]')
plt.ylabel('Y')
plt.figure()
plt.scatter(X[:,1],Y)
plt.xlabel('first X regressor - X[:,1]')
plt.ylabel('Y')
e=ols_estimate(X,Y)
display(e)
```
Let's run the same analysis using a canned function from the statsmodels package to compare the results. Note that statsmodels automatically adds an intercept, so we don't pass that column from the design matrix.
```
ols_result=sm.OLS(Y,X).fit()
print(ols_result.summary())
for i in range(len(e.t.values)):
assert numpy.allclose(e.t.values[i],ols_result.tvalues[i])
```
### Beyond ordinary least squares
In the foregoing, we used ordinary least squares estimation, which is the best linear unbiased estimator in the case of uncorrelated and homoscedastic (equal variance) errors (according to the Gauss-Markov theorem). However, there are common situations where these assumptions fail, in which case we need to use more sophisticated models. The case that is most relevant to fMRI is when there are correlated errors, which we will explore below.
First, let's simulate performance using OLS when the assumptions are upheld - the Type I error rate should be about 0.05.
```
nruns=1000
pval=numpy.zeros((nruns,3))
bhat=numpy.zeros((nruns,3))
for i in range(nruns):
Y,X=make_regression_data(beta=[0,0,0])
e=ols_estimate(X,Y)
pval[i,:]=e.p.values
bhat[i,:]=e.bhat.values
df=pandas.DataFrame({'Type 1 error':[numpy.mean(pval[:,i]<0.05) for i in range(3)],
'Variance of bhat':[numpy.std(bhat[:,i]) for i in range(3)]},
index=['X1','X2','intercept'])
display(df)
```
Now let's introduce some correlated noise, using the function created above which smooths the noise across observations using a first-order autoregressive (AR(1)) model. We do this for a range of levels of autocorrelation; because we have set the true beta values to zero, and the resulting proportion of significant results tells us the Type 1 error. We also assess the variance of the estimates.
```
nruns=1000
ncvals=numpy.arange(0.0,0.9,0.1)
pval=numpy.zeros((nruns,3,len(ncvals)))
bhat=numpy.zeros((nruns,3,len(ncvals)))
for nc in range(len(ncvals)):
for i in range(nruns):
Y,X=make_regression_data(beta=[0,0,0],noisecorr=ncvals[nc])
e=ols_estimate(X,Y,add_intercept=False)
pval[i,:,nc]=e.p.values
bhat[i,:,nc]=e.bhat.values
pval_exc=pval<0.05
meanpval=numpy.mean(pval_exc,0)
f=plt.figure(figsize=(8,5))
plt.subplot(1,2,1)
plt.plot(ncvals,meanpval.T)
plt.plot([0,1],[0.05,0.05],'--')
plt.xlabel('autocorrelation')
plt.ylabel('Type I error (% of significant tests)')
plt.legend(['X1','X2','Intercept'])
plt.ylim([0,1])
plt.subplot(1,2,2)
bhvar=numpy.std(bhat,0)
plt.plot(ncvals,bhvar.T)
plt.xlabel('autocorrelation')
plt.ylabel('std of parameter estimates')
plt.legend(['X1','X2','Intercept'])
plt.ylim([0,1])
```
__Exercise__: What do you see? Why do the effects of correlation in the data differ between regressors?
### Generalized least squares
In cases where the data do not adhere to the assumptions of OLS, we can use generalized least squares to obtain BLUE estimates. This requires that we have a model of the autocorrelation structure. Let's use a Toeplitz matrix, allows us to create an AR(1) covariance matrix.
The Toeplitz matrix has this form (in this case for a dataset with 4 observations):
```
print(toeplitz(range(4)))
```
The AR1 covariance has this form:
$V = \sigma^2 \begin{bmatrix}1 & \rho & \rho^2 & \rho^3\\
\rho & 1 & \rho & \rho^2\\
\rho^2 & \rho & 1 & \rho \\
\rho^3 & \rho^2 & \rho & 1 \\\end{bmatrix}$
where $\rho$ is the first-order autocorrelation and $\sigma^2$ is the variance. Note that we still assume that the variances are homogenous across datapoints. Thus, to generate such a matrix we simply exponentiate $\rho$ by the Toeplitz matrix (which is acheived using the $**$ operator) in Python.
```
rho=0.3
print(rho**toeplitz(range(4)))
```
Now let's build a version of our estimator that uses GLS rather than OLS. We do this using an interative approach. We first run OLS to estimate the model and obtain the residuals, and then we estimate the autocorrelation structure from the residuals. Then we estimate the model using GLS with the autocorrelation structure estimated above. The GLS estimator is:
$\hat{B} = (X'V^{-1}X)^{-1}X'V^{-1}Y$
where $V$ is the covariance matrix (which in OLS we assumed was simply $\sigma^2I$). This is akin to "whitening" the data by removing the covariance structure.
```
def gls_estimate(X,Y,add_intercept=True,verbose=False,
ddof=1,use_two_sided=True):
"""
estimate generalized least squares
using a Toeplitz matrix to generate AR(1) covariance
"""
# first we need to set up the matrices in the proper shape
# Y should be N X 1
# X should be X X c
if verbose:
print('original Y shape:',Y.shape)
Y=Y.reshape((len(Y),1))
if verbose:
print('new Y shape:',Y.shape)
if verbose:
print('original X shape:',X.shape)
if len(X.shape)==1:
X=X.reshape((len(X),1))
Xnames=['X%d'%int(i+1) for i in range(X.shape[1])]
if verbose:
print('new X shape:',X.shape)
# add an intercept to the model if specified
if add_intercept:
X=sm.add_constant(X)
# make sure that the design matrix is full rank
assert numpy.linalg.matrix_rank(X)==X.shape[1]
# first fit OLS to get residuals for AC estimation
e=ols_estimate(X,Y)
resid=Y - X.dot(e.bhat.values[:,numpy.newaxis])
ar1_coef=statsmodels.tsa.stattools.acf(resid)[1] # get the first-order autocorrelation estimate
# compute the inverse covariance matrix
order=toeplitz(range(len(Y)))
sigma=ar1_coef**order
Vinv=numpy.linalg.inv(sigma)
# re-estimate the model using GLS
b_hat=numpy.linalg.inv(X.T.dot(Vinv).dot(X)).dot(X.T.dot(Vinv).dot(Y))
if verbose:
print('b_hat=',b_hat)
resid=Y-X.dot(b_hat)
sigma2=resid.T.dot(resid)/(X.shape[0] - X.shape[1]) # variance of the residuals
# now compute the t statistic and p values for for each variable in X
t=numpy.zeros(X.shape[1])
p=numpy.zeros(X.shape[1])
for i in range(X.shape[1]):
c=numpy.zeros(X.shape[1])
c[i]=1
t[i]=c.dot(b_hat)/numpy.sqrt(c.dot(numpy.linalg.inv(X.T.dot(Vinv).dot(X))).dot(c.T)*sigma2)
if t[i]<0:
p[i]=scipy.stats.distributions.t.cdf(t[i],len(Y)-1)
else:
p[i]=1-scipy.stats.distributions.t.cdf(t[i],len(Y)-1)
if use_two_sided:
p[i]=p[i]*2
if verbose:
print('t=',t)
df=pandas.DataFrame({'bhat':b_hat.ravel(),'t':t.ravel(),'p':p.ravel()},index=Xnames)
return df
order=toeplitz(range(len(Y)))
sigma=0.5**order
Y,X=make_regression_data(beta=[1,0.1,10],noisecorr=0.5)
e=gls_estimate(X,Y)
display(e)
gls_result=sm.GLS(Y,X,sigma=sigma).fit()
gls_result.summary()
```
What do you see in this comparison?
Now let's simulate datasets under the null and estimate the model, across different levels of autocorrelation, as we did above. Because the estimation is a bit more complex this will take a couple of minutes.
```
nruns=1000
ncvals=numpy.arange(0.0,0.9,0.1)
pval=numpy.zeros((nruns,2,len(ncvals)))
bhat=numpy.zeros((nruns,2,len(ncvals)))
for nc in range(len(ncvals)):
for i in range(nruns):
Y,X=make_regression_data(beta=[0,0,0],noisecorr=ncvals[nc])
e=gls_estimate(X,Y)
pval[i,:,nc]=e.p.values[:2]
bhat[i,:,nc]=e.bhat.values[:2]
pval_exc=pval<0.05
meanpval=numpy.mean(pval_exc,0)
f=plt.figure(figsize=(12,5))
f=plt.subplot(1,2,1)
plt.plot(ncvals,meanpval.T)
plt.plot([0,1],[0.05,0.05],'--')
plt.xlabel('autocorrelation')
plt.ylabel('% of significant tests')
plt.legend(['X1','X2','Intercept'])
plt.ylim([0,1])
bhvar=numpy.std(bhat,0)
f=plt.subplot(1,2,2)
plt.plot(ncvals,bhvar.T)
plt.xlabel('autocorrelation')
plt.ylabel('std of parameter estimates')
plt.legend(['X1','X2','Intercept'])
plt.ylim([0,1])
```
| github_jupyter |
```
from youtube_transcript_api import *
from iso639 import languages
import math
def get_transcript_list(youtube_id):
try:
return YouTubeTranscriptApi.list_transcripts(youtube_id)
except Exception:
raise Exception("Check The Link")
def get_langs(transcript_list,manual=True):
langs = {}
if manual:
langs = transcript_list._manually_created_transcripts.copy()
else:
langs = transcript_list._generated_transcripts.copy()
for lang_code in list(langs.keys()):
try:
langs[lang_code] = languages.get(alpha2=lang_code).name
except:
pass
return {value:key for key, value in langs.items()}
def get_transcript(transcript_list,lang='en'):
return transcript_list.find_transcript([lang]).fetch()
def translate_transcript(transcript_list,to_lang):
transcript = transcript_list.find_transcript(['en'])
return transcript.translate(to_lang).fetch()
def get_time_stamps(transcript,word):
time_stamps = []
for sentence in transcript:
if word in sentence["text"].lower().split():
time = sentence["start"]
duration = int(sentence["duration"])
time_formated = ''
if time < 60:
secs = int(time)
time_formated = f'00:00:{secs} - 00:00:{secs+duration}'
elif 60 <= time < 3600:
mins = int(time/60)
secs = int((time % 60) * 60)
secs_lenght = len(str(secs))
secs = secs if secs_lenght < 2 else int(secs / pow(10,secs_lenght-2))
time_formated = f'00:{mins}:{secs} - 00:{mins}:{secs+duration}'
else:
hours = int(time/3600)
mins = int( (time % 3600) * 3600) / 60
print(time)
mins_lenght = len(str(mins))
mins = mins if mins_lenght < 2 else int(mins / pow(10,mins_lenght-2))
secs = int((time % 3600) * 3600)
secs_lenght = len(str(secs))
secs = secs if secs_lenght < 2 else int(secs / pow(10,secs_lenght-2))
time_formated = f'{hours}:{mins}:{secs} - {hours}:{mins}:{secs+duration}'
time_stamps.append(time_formated)
return None if len(time_stamps) == 0 else time_stamps
def get_translation_langs(transcript_list):
translation_langs = {}
for lang in transcript_list._translation_languages:
try:
translation_langs[languages.get(alpha2=lang["language_code"]).name] = lang["language_code"]
except:
pass
return translation_langs
def get_youtubeId(link):
'''
Examples of Youtube links:
1- https://youtu.be/zPF4coJ7pvU
2- https://www.youtube.com/watch?v=zPF4coJ7pvU
3- https://youtu.be/zPF4coJ7pvU?t=40
4- https://www.youtube.com/embed/zPF4coJ7pvU
5- https://www.youtube-nocookie.com/embed/zPF4coJ7pvU
6- https://www.youtube-nocookie.com/embed/zPF4coJ7pvU?start=40
7- https://www.youtube.com/embed/zPF4coJ7pvU?controls=0&start=40
'''
id_part = link.split("/")[-1]
id = id_part
if 'watch' in id_part:
id = id_part.split("v=")[-1]
elif '?' in id_part:
id = id_part.split('?')[0]
return id
def get_time_stamps_dict(transcript, words):
time_stamps_dict = {}
for word in words:
time_stamp = get_time_stamps(transcript,word)
if time_stamp == None:
time_stamps_dict[word] = "Not Found"
else:
time_stamps_dict[word] = time_stamp
return time_stamps_dict
if __name__ == '__main__':
link = input("Enter YouTube Link That You Want To Search: ")
id = get_youtubeId(link)
transcript_list = get_transcript_list(id)
manul_dict = get_langs(transcript_list)
auto_dict = get_langs(transcript_list, manual=False)
translation_dict = get_translation_langs(transcript_list)
manul_names = list(manul_dict.keys())
auto_names = list(auto_dict.keys())
translation_names = list(translation_dict.keys())
auto_names = list(filter(lambda elm: elm not in manul_names, auto_names))
transcript = None
print("Recommended: " + str(manul_names))
print("Not Recommended: " + str(auto_names))
choosen_lang = input("Choose Language To Search In (Type translate if you want another language, Not Recommended): ")
if choosen_lang.lower() == "translate":
try:
choosen_lang = input("Choose Language To Translate To: ")
transcript = translate_transcript(transcript_list, translation_dict[choosen_lang.capitalize()])
except:
transcript = print("Language Not Found Defaulting To English")
else:
found_in_manul = choosen_lang.capitalize() in manul_names
found_in_auto = choosen_lang.capitalize() in auto_names
lang = manul_dict[choosen_lang.capitalize()] if found_in_manul else auto_dict[choosen_lang.capitalize()]
if found_in_manul or found_in_auto:
transcript = get_transcript(transcript_list , lang=lang)
flag = True
while flag:
word = input("Choose Words To Search For, Put One Space Between Each Word: ").lower().split()
time_stamps = get_time_stamps_dict(transcript,word)
print("You May Not Find The Word In These Intervals But It Colud Be Close To Them")
for (word, time_stamp) in time_stamps.items():
print(f"Intervals For {word}: ")
print(time_stamp)
ans = input("Do You Want To Search For Another Words?(y/n): ")
flag = True if ans.lower() == 'y' else False
```
| github_jupyter |
<a href="https://colab.research.google.com/github/wikistat/AI-Frameworks/blob/master/IntroductionDeepReinforcementLearning/Policy_Gradient.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# [IA Frameworks](https://github.com/wikistat/AI-Frameworks) - Introduction to Deep Reinforcement Learning
<center>
<a href="http://www.insa-toulouse.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/logo-insa.jpg" style="float:left; max-width: 120px; display: inline" alt="INSA"/></a>
<a href="http://wikistat.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/wikistat.jpg" width=400, style="max-width: 150px; display: inline" alt="Wikistat"/></a>
<a href="http://www.math.univ-toulouse.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/logo_imt.jpg" width=400, style="float:right; display: inline" alt="IMT"/> </a>
</center>
# Part 2 : Policy Gradient Algorithm
The objectives of this notebook are the following :
* Implement Hard-Coded And Neural network policy to solve the *CartPole* Game
* Implement Policy gradient algorithm to solve the *CartPole* Game
# Files & Data (Google Colab)
If you're running this notebook on Google colab, you do not have access to the `solutions` folder you get by cloning the repository locally.
The following lines will allow you to build the folders and the files you need for this TP.
**WARNING 1** Do not run this line localy. <br>
**WARNING 2** The magic command `%load` does not work work on google colab, you will have to copy-paste the solution on the notebook.
```
! mkdir solution
! wget -P solution https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/solutions/pg_simple_policy.py
! wget -P solution https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/solutions/pg_neural_network_policy.py
! wget -P solution https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/solutions/pg_learn_given_policy.py
! wget -P solution https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/solutions/PG_class.py
! wget -P solution https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/solutions/discounted_loss.py
! wget -P . https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/discounted_rewards.py
! wget -P . https://github.com/wikistat/AI-Frameworks/raw/master/IntroductionDeepReinforcementLearning/keras_model.py
```
# Import librairies
```
import numpy as np
import random
import math
from tqdm import tqdm
# To plot figures and animations
import matplotlib
import matplotlib.animation as animation
import matplotlib.pyplot as plt
from IPython.display import HTML
#Tensorflow/Keras utils
import tensorflow as tf
import tensorflow.keras.models as km
import tensorflow.keras.layers as kl
import tensorflow.keras.initializers as ki
import tensorflow.keras.optimizers as ko
import tensorflow.keras.backend as K
import tensorflow.keras.losses as klo
import tensorflow.keras.metrics as kme
# Gym Library
import gym
```
The following functions enable to build a video from a list of images. <br>
They will be used to build video of the game you will played.
```
def update_scene(num, frames, patch):
patch.set_data(frames[num])
return patch,
def plot_animation(frames, repeat=False, interval=40):
plt.close() # or else nbagg sometimes plots in the previous cell
fig = plt.figure()
patch = plt.imshow(frames[0])
plt.axis('off')
return animation.FuncAnimation(fig, update_scene, fargs=(frames, patch), frames=len(frames), repeat=repeat, interval=interval)
```
# AI Gym Librairie
<a href="https://gym.openai.com/" ><img src="https://gym.openai.com/assets/dist/home/header/home-icon-54c30e2345.svg" style="float:left; max-width: 120px; display: inline" alt="INSA"/></a>
In this notebook we will be using [OpenAI gym](https://gym.openai.com/), a great toolkit for developing and comparing Reinforcement Learning algorithms. It provides many environments for your learning *agents* to interact with.
# A simple environment: the Cart-Pole
## Description
A pole is attached by an un-actuated joint to a cart, which moves along a frictionless track. The pendulum starts upright, and the goal is to prevent it from falling over by increasing and reducing the cart's velocity.
### Observation
Num | Observation | Min | Max
---|---|---|---
0 | Cart Position | -2.4 | 2.4
1 | Cart Velocity | -Inf | Inf
2 | Pole Angle | ~ -41.8° | ~ 41.8°
3 | Pole Velocity At Tip | -Inf | Inf
### Actions
Num | Action
--- | ---
0 | Push cart to the left
1 | Push cart to the righ&t
Note: The amount the velocity is reduced or increased is not fixed as it depends on the angle the pole is pointing. This is because the center of gravity of the pole increases the amount of energy needed to move the cart underneath it
### Reward
Reward is 1 for every step taken, including the termination step
### Starting State
All observations are assigned a uniform random value between ±0.05
### Episode Termination
1. Pole Angle is more than ±12°
2. Cart Position is more than ±2.4 (center of the cart reaches the edge of the display)
3. Episode length is greater than 200
### Solved Requirements
Considered solved when the average reward is greater than or equal to 195.0 over 100 consecutive trials.
The description above if part of the official description of this environemtn. Read full description [here](https://github.com/openai/gym/wiki/CartPole-v0).
The following command will load the `CartPole` environment.
```
env = gym.make("CartPole-v0")
env.reset()
img = env.render(mode = "rgb_array")
env.close()
print("Environemnt is a %dx%dx%d images" %img.shape)
plt.imshow(img)
plt.axis("off")
```
If you have forgotten how the `CartPole` environment works, open the `Deep_Q_Learning_CartPole.ipynb` notebook to run explanation's cell.
The functions below are generic functions that enable to:
* `run_one_epsiode`: run a complete episode of a game according to a policy function as an input.
* `play_games` : play `n_games`parameter and print mean reward of this `n_games`.
```
def run_one_episode(policy, return_frames=False):
frames = []
observation = env.reset()
reward_episod = 0
done = False
while not(done):
action = policy(observation)
observation, reward, done, _ = env.step(action)
reward_episod += reward
if return_frames:
img = env.render(mode = "rgb_array")
env.close()
frames.append(img)
return reward_episod, frames
def play_games(policy, n_games=100):
all_reward_sum = []
n_game = 0
while n_game < n_games:
reward_episod, _ = run_one_episode(policy)
if n_game %10 == 0:
print("Game played : %d. Mean and Standart deviation's reward for the last 10 episode: %.1f - %.1f" %(n_game, np.mean(all_reward_sum[-10:]), np.std(all_reward_sum[-10:])) )
all_reward_sum.append(reward_episod)
n_game += 1
print("Over %d episodes, mean reward: %.1f, std : %.1f" %(n_games, np.mean(all_reward_sum), np.std(all_reward_sum)))
```
# Hard coded policy
How can we make the poll remain upright? We will need to define a _policy_ for that.
This is the strategy that the agent will use to select an action at each step. It can use all the past actions and observations to decide what to do.
Let's first implement **Hard Coded policies**, *i.e.* simple rules that defines which action to takes according to the parameters.
## Random policy
Let's start with a completly random policy and see how much time the poll will remain upright over 100 episodes.
```
def policy_random(state):
return env.action_space.sample()
play_games(policy = policy_random)
```
#### Visualize a complete game
Let's run one pisode with his random policy and save all images representing the environment at each step.
```
reward_episod, frames = run_one_episode(policy = policy_random, return_frames=True)
HTML(plot_animation(frames).to_html5_video())
```
### Simple strategy
Let's hard code a simple strategy: if the pole is tilting to the left, then push the cart to the left, and _vice versa_. Let's see if that works.
**Exercise** implement this policy and play 100 games with this policy. What are the means and std deviation of the reward sum over the 100 games?
```
# %load solutions/pg_simple_policy.py
play_games(policy = simple_policy)
```
**Q** What can you say about this strategy?
**Exercise** Vizualize a complete game:
```
reward_episod, frames = run_one_episode(policy = simple_policy, return_frames=True)
HTML(plot_animation(frames).to_html5_video())
```
# Neural Network Policies
Let's create a neural network to build a better policy.
This network will take the observations as inputs, and output the probability of the action to take for each observation. <br>
In the case of the Cart-Pole environment, there are just two possible actions (left or right), so we only need one output neuron: it will output the probability `p` of the action 1 (right), and of course the probability of action 0 (right) will be `1 - p`.
Let's first see how this neural network policy work without training it and then let's try to learn the simple policy define above.
## The architecture
Because this problem is simple, we can define a very simple architecture for our neural network.<bR>
Here it's simple MLP with 1 hidden layer and four neurons.
```
# Specify the network architecture
n_inputs = 4 # == env.observation_space.shape[0]
n_hidden = 9 # it's a simple task, we don't need more than this
n_outputs = 1 # only outputs the probability of accelerating right
# Build the neural network
policy_network=km.Sequential()
policy_network.add(kl.Dense(n_hidden, input_shape = (n_inputs,), activation = "relu"))
policy_network.add(kl.Dense(n_outputs, activation = "sigmoid"))
policy_network.summary()
```
Note that the model is not compiled so far, no loss function is defined.
## Policy from a neural network
We can now easily predict the probability of both actions given the observation.
**Exercise** Define a function to choose the action to take from an observation and the neural network.
```
# %load solutions/pg_neural_network_policy.py
```
***NB*** : In this particular environment, the past actions and observations can safely be ignored, since each observation contains the environment's full state. If there were some hidden state then you may need to consider past actions and observations in order to try to infer the hidden state of the environment.
For example, if the environment only revealed the position of the cart but not its velocity, you would have to consider not only the current observation but also the previous observation in order to estimate the current velocity. <br> Another example: if the observations are noisy: you may want to use the past few observations to estimate the most likely current state. Our problem is thus as simple as can be: the current observation is noise-free and contains the environment's full state.
## Random neural network policy.
Let's see how this neural network policy perform.
```
play_games(policy = lambda obs : neural_network_policy(obs, model=policy_network), n_games=10)
```
Let's randomly initialize this policy neural network and use it to play one game:
```
reward_episod, frames = run_one_episode(policy = lambda obs : neural_network_policy(obs, model=policy_network), return_frames=True)
HTML(plot_animation(frames).to_html5_video())
```
The neural network is working. But it's still acting randomly because we do not train the neural network. Let's try to make it learn better policy.
## Learn a given policy
In this part we will train the neural network in order that it learns the simple strategy we hard coded before : if the pole is tilting to the left, then push the cart to the left, and _vice versa_. <br>
The class defined below enables to train the neural network in order to learn this simple policy.
The **pseudo code** is quite simple here:
while *n_episode_max* is not reached:
* Play and episode and return the observation and target
* Train the network from these observation and target.
**Exercicse**: Complete the code below: <br>
* Choose a loss to compile the model within the `init_model`method.
* Write the `play_one_episode`method that will:
* Play an episode until it's end with the hard coded policy
* Return observation an target, i.e, the action to take according to each observation, of this episode.
You can do this exercise on this notebook or with the `PG_learn_a_policy.py`and the `PG_learn_a_policy_solution.py` files.
```
class PG:
def __init__(self):
# Environment
self.env = gym.make("CartPole-v0")
self.dim_input = self.env.observation_space.shape[0]
# Model
self.model = self.init_model()
self.n_episode_max = 1000
def init_model(self):
# Build the neural network
policy_network = km.Sequential()
policy_network.add(kl.Dense(9, input_shape=(self.dim_input,), activation="relu"))
policy_network.add(kl.Dense(1, activation="sigmoid"))
policy_network.compile(loss=, optimizer=ko.Adam(), metrics=['accuracy'])
return policy_network
def play_one_episode(self):
# Todo
return train_data
def train(self):
for iteration in tqdm(range(self.n_episode_max)):
train_data = self.play_one_episode()
n_step = len(train_data)
target = np.array([x[1] for x in train_data]).reshape((n_step, 1))
observations = np.array([x[0] for x in train_data])
self.model.train_on_batch(observations, target)
pg = PG()
pg.train()
# %load solutions/pg_learn_given_policy.py
play_games(policy = lambda obs : neural_network_policy(obs, model=pg.model), n_games=10)
reward_episod, frames = run_one_episode(policy = lambda obs : neural_network_policy(obs, model=pg.model), return_frames=True)
HTML(plot_animation(frames).to_html5_video())
```
Looks like it learned the policy correctly! <br>
Let's now reach our final target : The neural network has to find a better policy by its own.
# Policy Gradients
The idea behind *Policy Gradients* its quite simple : The _Policy Gradients_ algorithm tackles this problem by first playing multiple games, then making the actions in good games slightly more likely, while actions in bad games are made slightly less likely. First we play, then we go back and think about what we did.
### Algorithm
* Run an episode untill it's done and save at each iteration the observation, action and reward.
* When an episode it's done. Compute the discounted rewards for all the episode, and save it.
* If you have done *batch_size=50* episodes train your model on this batch.
* Stop if you have reach *num episodes* or *goal* target.
### Parameters
| Variable | Value | Description |
|---|---|---|
|Gamma | 0.99 | The discounted rate apply for the discounted reward |
|batch_size | 50 | Number of episode to run before training model on a batch of episode |
| Num episodes | 10.000 | Maximum number of episode to run before stopping the training |
| goal | 190 | Number of step to achieve on one episode to stop the training. |
Those parameters are fixed for this TP, they are common value for this kind of problem based on experiences. They are not definitive nor results or any research.
## Discounted rewards
To train this neural network we will then used the observation of the experiences as an inputs and the actions taken as an output.
But how do we provide to the neural network the information the choosen actions was good or bad?
The problem is that most actions have delayed effects, so when you win or lose points in a game, it is not clear which actions contributed to this result: was it just the last action? Or the last 10? Or just one action 50 steps earlier? <br>
This is called the _credit assignment problem_.
To tackle this problem, a common strategy is to evaluate an action based on the sum of all the rewards that come after it, usually applying a discount rate r at each step. It's call the **discounted rewards**
$$ R_t = \sum_{i=0}^{\infty}\gamma^i r_{t+i}$$
This rate will the be applied to the loss function of the neural network :
* A high discounted reward will lead to higher gradient which will increase the importance of this action
* A low discounted reward will lead to lower gradient which will decrease the importance of this action
**Exercise** : Implement the discount_rewards function.
```
def discount_rewards(r, gamma=0.99):
"""Takes 1d float array of rewards and computes discounted reward
e.g. f([1, 1, 1], 0.99) -> [2.9701, 1.99, 1]
"""
TODO
return discounted_rewards
# %load discounted_rewards.py
assert np.all(discount_rewards([1,1,1], gamma=0.99) == [2.9701, 1.99, 1])
assert np.all(discount_rewards([3,2,1],gamma=0.99) == [5.960100000000001, 2.99, 1.0])
```
## Architecture & Loss Function
As before we will define a very simple architecture to our neural network : A MLP with only one hidden layer and 8 neurons.
We want to write a custom loss for this game : a *weighted binary cross-entropy* where the weights are the discounted rewards of the predicted action at the given state.
To write this loss, a `discountedLoss` class that inherit the keras.losses.Loss class features is defined below.
**Exercise**: Write the loss function within the `call`method using keras backend (*i.e*, using `K.log` and `K.mean`function that works like numpy functions.
The method takes the *prediction*, the *true_value* and the *discounted rewards* as an input.
```
class discountedLoss(klo.Loss):
"""
Args:
pos_weight: Scalar to affect the positive labels of the loss function.
weight: Scalar to affect the entirety of the loss function.
from_logits: Whether to compute loss from logits or the probability.
reduction: Type of tf.keras.losses.Reduction to apply to loss.
name: Name of the loss function.
"""
def __init__(self,
reduction=klo.Reduction.AUTO,
name='discountedLoss'):
super().__init__(reduction=reduction, name=name)
def call(self, y_true, y_pred, disc_r):
#TODO
return loss
#%load solutions/discounted_loss.py
```
A `kerasModel`class below is defined below. This class inherit the `keras.models.Model` class.
The **metric**, and **optimizer** of this class are defined using native keras function that you are use to use to define and compile a keras model.
The **loss** takes the discountedLoss we defined above.
```
class kerasModel(km.Model):
def __init__(self):
super(kerasModel, self).__init__()
self.layersList = []
self.layersList.append(kl.Dense(9, activation="relu",
input_shape=(4,),
use_bias=False,
kernel_initializer=ki.VarianceScaling(),
name="dense_1"))
self.layersList.append(kl.Dense(1,
activation="sigmoid",
kernel_initializer=ki.VarianceScaling(),
use_bias=False,
name="out"))
self.loss = discountedLoss()
self.optimizer = ko.Adam(lr=1e-2)
self.train_loss = kme.Mean(name='train_loss')
self.validation_loss = kme.Mean(name='val_loss')
self.metric = kme.Accuracy(name="accuracy")
@tf.function()
def predict(x):
"""
This is where we run
through our whole dataset and return it, when training and testing.
"""
for l in self.layersList:
x = l(x)
return x
self.predict = predict
@tf.function()
def train_step(x, labels, disc_r):
"""
This is a TensorFlow function, run once for each epoch for the
whole input. We move forward first, then calculate gradients with
Gradient Tape to move backwards.
"""
with tf.GradientTape() as tape:
predictions = self.predict(x)
loss = self.loss.call(
y_true=labels,
y_pred = predictions,
disc_r = disc_r)
gradients = tape.gradient(loss, self.trainable_variables)
self.optimizer.apply_gradients(zip(gradients, self.trainable_variables))
self.train_loss(loss)
return loss
self.train_step = train_step
```
## PG class
The `PG` class contains the implementation of the **Policy Gradient** algorithm. The code is incomplete and you will have to fill it!
**GENERAL INSTRUCTION**:
* Read the init of the `PG` class.
* Various variable are set with their definition, make sure you understand all of its.
* The *game environment*, the *experiences list* and the *keras model* are initialised.
* Read the `train` method. It contains the main code corresponding to the **pseudo code** below. YOU DO NOT HAVE TO MODIFY IT! But make sure you understand it.
* The `train` method use methods that are not implemented.
* You will have to complete the code of 3 functions. (read instruction of each exercise below)
* After the cell of the `PG` class code below there is a **test cells**. <br>
This cell should be executed after all the methods have been completed. This cell will check that the function you implemented take input and output in the desired format. <br> DO NOT MODIFY this cell. It will work if you're code is good <br> **Warning** The test cell does not guarantee that your code is correct. It just test than input and output are in the good format.
#### Pseudo code
We will consider that the game is **completed** if the mean score over 10 games is above 190.
While you didn't reach the expected *goal* reward or the max *num_episodes* allow to be played:
* Run `one_episode` and save all experiences in the `experiences` list (**Exercise 1 & 2**):
* Every `batch_size` episodes played:
* train model over a batch of experiences (**Exercise 3**)
**Exercise 1**: Implement `choose_action`<br>
This method chooses an action according to this rules:<br>
* let $p$ be the probability of the output of the model that we play the action $right(=1)$,
* Then choose action to play right with probability $p$ else play left.
* Hence, the more the model will be good about it's prediction, the less exploration we will perform.
**Exercise 2**: Implement `run_one_episode` <br>
This method:<br>
* play an complete episode until it's done. At each step of the episode, it :
* chooses an action
* save all the actions, state and reward
* once the episode is done and all rewards are know, it compute all the discounted rewards.
* fill the `experiences's` list with all experience of the episode = '[state, action, discounted_reward]'
**Exercise 3**: Implement `run_one_batch_train`<br>
This method:<br>
* call the on `train_step` method of the `model`with the argument in the `experiences` list.
* Empty the `experiences` list
* return the loss of this batch step.
You can do these exercises on this notebook or with the `PG.py`and the `PG_solution.py` files.
```
import tensorflow as tf
tf.config.experimental_run_functions_eagerly(True)
tf.keras.backend.set_floatx('float64')
class PG:
def __init__(self, gamma=.99, batch_size=50, num_episodes=10000, goal=190, n_test=10, print_every=100):
# Environment
self.env = gym.make("CartPole-v0")
self.dim_input = self.env.observation_space.shape[0]
# Parameters
self.gamma = gamma # -> Discounted reward
self.batch_size = batch_size # -> Size of episode before training on a batch
# Stop factor
self.num_episodes = num_episodes # Max number of iterations
self.goal = goal # Stop if our network achieve this goal over *n_test*
self.n_test = n_test
self.print_every = print_every # ?Numbe rof episode before trying if our model perform well.
# Init Model to be trained
self.model = kerasModel()
# Placeholders for our observations, outputs and rewards
self.experiences = []
self.losses = []
def choose_action(self, state):
# TODO
return action
def run_one_episode(self):
# TODO
return score
def run_one_batch_train(self):
# TODO
return loss
def score_model(self, model, num_tests, dimen, ):
scores = []
for num_test in range(num_tests):
observation = self.env.reset()
reward_sum = 0
while True:
state = np.reshape(observation, [1, dimen])
predict = model.predict(state)[0]
action = 1 if predict > 0.5 else 0
observation, reward, done, _ = self.env.step(action)
reward_sum += reward
if done:
break
scores.append(reward_sum)
return np.mean(scores)
def train(self):
metadata = []
i_batch = 0
# Number of episode and total score
num_episode = 0
train_score_sum = 0
while num_episode < self.num_episodes:
train_score = self.run_one_episode()
train_score_sum += train_score
num_episode += 1
if num_episode % self.batch_size == 0:
i_batch += 1
loss = self.run_one_batch_train()
self.losses.append(loss)
metadata.append([i_batch, self.score_model(self.model, self.n_test, self.dim_input)])
# Print results periodically
if num_episode % self.print_every == 0:
test_score = self.score_model(self.model, self.n_test, self.dim_input)
print(
"Average reward for training episode {}: {:0.2f} Mean test score over {:d} episode: {:0.2f} Loss: {:0.6f} ".format(
num_episode, train_score_sum / self.print_every, self.n_test,
test_score,
self.losses[-1]))
reward_sum = 0
if test_score >= self.goal:
print("Solved in {} episodes!".format(num_episode))
break
return metadata
pg = PG()
score = pg.run_one_episode()
assert type(score) is float
for state, action, dreward in pg.experiences:
assert np.all(state.shape==(1,4))
assert type(action)==int
assert type(dreward)==np.float64
loss = pg.run_one_batch_train()
assert type(loss) == float
assert len(pg.experiences)==0
# %load solutions/PG_class.py
```
### Training
Let's train the model !
```
pg = PG(goal=200)
metadata = pg.train()
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sb
sb.set_style("whitegrid")
fig = plt.figure(figsize=(20,6))
ax = fig.add_subplot(1,1,1)
ax.plot(list(range(len(metadata))),[x[1] for x in metadata])
ax.set_yticks(np.arange(0,210,10))
ax.set_xticks(np.arange(0,100,25))
ax.set_title("Score/Lenght of episode over Iteration", fontsize=20)
ax.set_xlabel("Number of iteration", fontsize=14)
plt.yticks(fontsize=12)
plt.xticks(fontsize=12)
ax.set_ylabel("Score/Length of episode", fontsize=16)
plt.savefig("pg_normalized.png", bbox_to_anchor="tigh", dpi=200)
```
**Exercise**
* Use the model to play 100 games and check how it performs compare to previous policy tested
* Register a video of a game and display it
```
play_games(policy = lambda obs : neural_network_policy(obs, model=pg.model), n_games=10)
reward_episod, frames = run_one_episode(policy = lambda obs : neural_network_policy(obs, model=pg.model), return_frames=True)
HTML(plot_animation(frames).to_html5_video())
```
| github_jupyter |
Simulate some data via Splatter to test BBKNN. Let's go for three batches of three cell types, and port the counts and cell type/batch metadata back to python.
```
%load_ext rpy2.ipython
%%R -o counts -o meta
suppressMessages(library(splatter))
params <- newSplatParams()
params <- setParam(params, "nGenes", 5000)
params <- setParam(params, "batchCells", c(500,500,500))
params <- setParam(params, "batch.facLoc", 0.5)
params <- setParam(params, "batch.facScale", 0.5)
params <- setParam(params, "group.prob", c(1/3,1/3,1/3))
sim <- splatSimulate(params, method="groups", verbose=FALSE)
counts = data.frame(counts(sim))
meta = data.frame(colData(sim))
import anndata
import scanpy.api as sc
import bbknn
sc.settings.set_figure_params(dpi=80) # low dpi (dots per inch) yields small inline figures
```
Do some basic data processing (scale to 10,000, log-transform), get a PCA.
```
adata = anndata.AnnData(X=counts.values.T, obs=meta)
sc.pp.normalize_per_cell(adata,counts_per_cell_after=10000)
sc.pp.log1p(adata)
sc.tl.pca(adata,svd_solver='arpack')
```
Standard neighbour inference procedure without any batch correction leads to a pretty heavy technical effect corrupting the visualisation.
```
sc.pp.neighbors(adata, n_neighbors=9)
sc.tl.umap(adata)
sc.pl.umap(adata,color=['Batch','Group'])
```
Applying BBKNN leads to the cell types being reconnected across the batches.
```
bbknn.bbknn(adata, batch_key='Batch', metric='euclidean')
sc.tl.umap(adata)
sc.pl.umap(adata,color=['Batch','Group'])
```
Let's inspect the influence of the trimming parameter. To do this, let's trim down the simulated data to a smaller setup - two batches, one has two cell types, the third has three. This is what we're left with on the original UMAP space.
```
to_del = (adata.obs['Batch'] == 'Batch3') | ((adata.obs['Batch'] == 'Batch2') & (adata.obs['Group'] == 'Group3'))
adata2 = adata[[not i for i in to_del]]
sc.pl.umap(adata2,color=['Batch','Group'])
```
Running BBKNN (after re-running PCA, as we have a subset of the original cells here) without trimming the graph leads to the population present in only one of the samples being erroneously integrated with an unrelated population.
```
sc.tl.pca(adata2,svd_solver='arpack')
bbknn.bbknn(adata2, batch_key='Batch', metric='euclidean')
sc.tl.umap(adata2)
sc.pl.umap(adata2,color=['Batch','Group'])
```
Trimming the graph leads to the population's independence being correctly restored, while not splitting up any of the correctly merged cell types.
```
bbknn.bbknn(adata2, batch_key='Batch', trim=10, metric='euclidean')
sc.tl.umap(adata2)
sc.pl.umap(adata2,color=['Batch','Group'])
```
| github_jupyter |
```
import pandas as pd
df = pd.read_csv('~/data/ynacc_proc/articles/articles_fixed_4.csv')
urls = df[df['text'].duplicated()]['url'].tolist()
# manually copied & pasted text from websites
texts = ["""
A gorilla was caught on camera dancing up a storm inside a British zoo.
Read: Ruh Roh! Owners Return Home to Discover Mischievous Dog Found Their Calligraphy Ink
Lope, a three-year-old, silverback gorilla was caught by Helen Fairhead dancing inside the Twycross Zoo in Leicestershire, England.
Video of the dancing primate was posted online on Monday and has been seen over one million times.
Read: Goat Escapes Its Home, Wanders Into a Starbucks
The zoo’s director of life science, Dr. Chartlotte Macdonald told BBC the gorilla's dancing is evidence of him playing, which according to her, is an important aspect of a young ape's behavior.
Watch: Mother Gorilla Dies At Zoo Shortly After Giving Birth Via Emergency C-Section""","""The woman who recorded a video of a panther whizzing right by her called the encounter “amazing.”
Read: Diver Straddles Shark to Untangle It From Rope
Tina Dorschel was taking an early morning stroll through Corkscrew Swamp Sanctuary in Naples, Florida, when, out of nowhere a panther, came right in her direction.
She told IE: “It was extremely surprising.”
Though nervous, she kept right on recording the big cat earlier this week.
"I was hoping it wasn't angry," she said.
Read: Panda Poses For Selfie With Caretaker
She posted the video on Facebook and it has racked up more than four million views.
On on early morning nature walk we saw a gator, a snake, frogs, pretty birds, and had this unexpected encounter. (Warning...curse word at end!)(For licensing or usage, please contact licensing@viralhog.com)
Posted by Tina Dorschel on Tuesday, March 29, 2016
The panther may have been more frightened than she was.
On the sanctuary’s Facebook page, staff wrote: “Panthers are shy creatures and this kind of encounter was a lucky and extremely rare experience.”
Watch: Abandoned Sloth Clings to Stuffed Animal for Emotional Support""","""Prince’s body is scheduled to undergo an autopsy on Friday to confirm what caused his death.
Read: World is Bathed in Purple for Prince As His Sister Says: 'Thank You For Loving Him'
He passed away just days after his private plane made an emergency landing in Moline, Illinois last Friday -- supposedly because he was suffering from a terrible case of the flu.
But reports now say the medical crisis was actually a drug overdose. According to the reports, Prince may have been brought back from the brink of death by a “save shot.”
The shots, also known as Narcan, are used to counteract the effects of opiates.
According to TMZ, doctors advised Prince to stay in the hospital for 24 hours, but he left after three.
Prince was found unresponsive Thursday morning in his elevator.
Reports suggest that Prince may have overdosed on the painkiller Percocet. The drug contains acetaminophen and oxycodone, an opioid, and is known to be highly addictive.
According to TMZ, Prince was taking the painkillers for a hip problem. The singer had hip surgery in 2010.
Read: Singer Prince Found Dead Inside Elevator At His Home, Age 57
Musician Sheila E, once Prince's fiancée, has flown to Minnesota to investigate the death and settle some of mysterious circumstances.
As she boarded a flight at LAX, she told TMZ that she is determined to find out what happened to her former lover and close friend.
Watch: Remembering the Most Iconic Moments of Prince's Lengthy Career""","""
Here are 5 stocks added to the Zacks Rank #1 (Strong Buy) List today:
American Superconductor Corporation (AMSC - Free Report)
BlackBerry Ltd
Blueprint Medicines Corp (BPMC - Free Report)
Darden Restaurants, Inc (DRI - Free Report)
eGain Corp (EGAN - Free Report)
View the entire Zacks Rank #1 List.
Zacks Restaurant Recommendations: In addition to dining at these special places, you can feast on their stock shares. A Zacks Special Report spotlights 5 recent IPOs to watch plus 2 stocks that offer immediate promise in a booming sector. Download it free »""",
"""Two tandem skydivers died in California on Saturday after the parachute failed to open.
One of the victims, whose name has not been released, was a teenager whose family was watching from below as he and the instructor hit the ground at about 10 a.m.
It was the teen's first-ever jump.
Read: Daredevil With No Parachute Survives 25,000 Foot Leap From Plane Into Net
Both victims died on impact in a Lodi vineyard, according to reports.
Tragically, the San Joaquine County Sheriff's Office said it appeared that the parachute deployed after impact.
Read: 16 Dead After Hot Air Balloon Hits High Voltage Power Lines
Authorities on Sunday were still trying to reach one of the men's families for notification, sheriff's Sgt. Brandon Riley. The instructor was in his mid-20s, Riley said.
Sheriff’s deputies first responded to a 911 call from a citizen reporting that one of his field workers called him and said a skydiver hit the ground without an open parachute.
The FAA has been contacted and will be taking over the investigation into the circumstances surrounding the accident.
Watch: World War II Vet Celebrates 90th Birthday by Skydiving, Days After His Brother Dies
"""]
for [url, text] in zip(urls, texts):
df.loc[df['url'] == url, ['text']] = text
df[df['text'].duplicated()]['url'].tolist()
df[df['url'].duplicated()]
df[df['text'].str.contains('Accept')]
df.to_csv('~/data/ynacc_proc/articles/articles_fixed_5.csv')
```
| github_jupyter |
# DefinedAEpTandZ0 media example
```
%load_ext autoreload
%autoreload 2
import skrf as rf
import skrf.mathFunctions as mf
import numpy as np
from numpy import real, log, log10, sum, absolute, pi, sqrt
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator
from scipy.optimize import minimize
rf.stylely()
```
## Measurement of two CPWG lines with different lenghts
The measurement where performed the 21th March 2017 on a Anritsu MS46524B 20GHz Vector Network Analyser. The setup is a linear frequency sweep from 1MHz to 10GHz with 10'000 points. Output power is 0dBm, IF bandwidth is 1kHz and neither averaging nor smoothing are used.
CPWGxxx is a L long, W wide, with a G wide gap to top ground, T thick copper coplanar waveguide on ground on a H height substrate with top and bottom ground plane. A closely spaced via wall is placed on both side of the line and the top and bottom ground planes are connected by many vias.
| Name | L (mm) | W (mm) | G (mm) | H (mm) | T (um) | Substrate |
| :--- | ---: | ---: | ---: | ---: | ---: | :--- |
| MSL100 | 100 | 1.70 | 0.50 | 1.55 | 50 | FR-4 |
| MSL200 | 200 | 1.70 | 0.50 | 1.55 | 50 | FR-4 |
The milling of the artwork is performed mechanically with a lateral wall of 45°.
The relative permittivity of the dielectric was assumed to be approximatively 4.5 for design purpose.

```
# Load raw measurements
TL100 = rf.Network('CPWG100.s2p')
TL200 = rf.Network('CPWG200.s2p')
TL100_dc = TL100.extrapolate_to_dc(kind='linear')
TL200_dc = TL200.extrapolate_to_dc(kind='linear')
plt.figure()
plt.suptitle('Raw measurement')
TL100.plot_s_db()
TL200.plot_s_db()
plt.figure()
t0 = -2
t1 = 4
plt.suptitle('Time domain reflexion step response (DC extrapolation)')
ax = plt.subplot(1, 1, 1)
TL100_dc.s11.plot_z_time_step(pad=2000, window='hamming', z0=50, label='TL100', ax=ax, color='0.0')
TL200_dc.s11.plot_z_time_step(pad=2000, window='hamming', z0=50, label='TL200', ax=ax, color='0.2')
ax.set_xlim(t0, t1)
ax.xaxis.set_minor_locator(AutoMinorLocator(10))
ax.yaxis.set_minor_locator(AutoMinorLocator(5))
ax.patch.set_facecolor('1.0')
ax.grid(True, color='0.8', which='minor')
ax.grid(True, color='0.4', which='major')
plt.show()
```
Impedance from the line and from the connector section may be estimated on the step response.
The line section is not flat, there is some variation in the impedance which may be induced by manufacturing tolerances and dielectric inhomogeneity.
Note that the delay on the reflexion plot are twice the effective section delays because the wave travel back and forth on the line.
Connector discontinuity is about 50 ps long. TL100 line plateau (flat impedance part) is about 450 ps long.
```
Z_conn = 53.2 # ohm, connector impedance
Z_line = 51.4 # ohm, line plateau impedance
d_conn = 0.05e-9 # s, connector discontinuity delay
d_line = 0.45e-9 # s, line plateau delay, without connectors
```
## Dielectric effective relative permittivity extraction by multiline method
```
#Make the missing reflect measurement
#Reflect only affects sign of the corrected
reflect = TL100.copy()
reflect.s[:,0,0] = 1
reflect.s[:,1,1] = 1
reflect.s[:,1,0] = 0
reflect.s[:,0,1] = 0
# Perform NISTMultilineTRL algorithm
cal = rf.NISTMultilineTRL([TL100, reflect, TL200], [1], [100e-3, 200e-3], er_est=3.0, refl_offset=[0])
plt.figure()
plt.title('Corrected lines')
cal.apply_cal(TL100).plot_s_db()
cal.apply_cal(TL200).plot_s_db()
plt.show()
```
Calibration results shows a very low residual noise floor. The error model is well fitted.
```
from skrf.media import DefinedAEpTandZ0
freq = TL100.frequency
f = TL100.frequency.f
f_ghz = TL100.frequency.f/1e9
L = 0.1
A = 0.0
f_A = 1e9
ep_r0 = 2.0
tanD0 = 0.001
f_ep = 1e9
x0 = [ep_r0, tanD0]
ep_r_mea = cal.er_eff.real
A_mea = 20/log(10)*cal.gamma.real
def model(x, freq, ep_r_mea, A_mea, f_ep):
ep_r, tanD = x[0], x[1]
m = DefinedAEpTandZ0(frequency=freq, ep_r=ep_r, tanD=tanD, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
ep_r_mod = m.ep_r_f.real
A_mod = m.alpha * log(10)/20
return sum((ep_r_mod - ep_r_mea)**2) + 0.001*sum((20/log(10)*A_mod - A_mea)**2)
res = minimize(model, x0, args=(TL100.frequency, ep_r_mea, A_mea, f_ep),
bounds=[(2, 4), (0.001, 0.013)])
ep_r, tanD = res.x[0], res.x[1]
print('epr={:.3f}, tand={:.4f} at {:.1f} GHz.'.format(ep_r, tanD, f_ep * 1e-9))
m = DefinedAEpTandZ0(frequency=freq, ep_r=ep_r, tanD=tanD, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
plt.figure()
plt.suptitle('Effective relative permittivity and attenuation')
plt.subplot(2,1,1)
plt.ylabel('$\epsilon_{r,eff}$')
plt.plot(f_ghz, ep_r_mea, label='measured')
plt.plot(f_ghz, m.ep_r_f.real, label='model')
plt.legend()
plt.subplot(2,1,2)
plt.xlabel('Frequency [GHz]')
plt.ylabel('A (dB/m)')
plt.plot(f_ghz, A_mea, label='measured')
plt.plot(f_ghz, 20/log(10)*m.alpha, label='model')
plt.legend()
plt.show()
```
Relative permittivity $\epsilon_{e,eff}$ and attenuation $A$ shows a reasonnable agreement.
A better agreement could be achieved by implementing the Kirschning and Jansen miscrostripline dispersion model or using a linear correction.
## Connectors effects estimation
```
# note: a half line is embedded in connector network
coefs = cal.coefs
r = mf.sqrt_phase_unwrap(coefs['forward reflection tracking'])
s1 = np.array([[coefs['forward directivity'],r],
[r, coefs['forward source match']]]).transpose()
conn = TL100.copy()
conn.name = 'Connector'
conn.s = s1
# delay estimation,
phi_conn = (np.angle(conn.s[:500,1,0]))
z = np.polyfit(f[:500], phi_conn, 1)
p = np.poly1d(z)
delay = -z[0]/(2*np.pi)
print('Connector + half thru delay: {:.0f} ps'.format(delay * 1e12))
print('TDR readed half thru delay: {:.0f} ps'.format(d_line/2 * 1e12))
d_conn_p = delay - d_line/2
print('Connector delay: {:.0f} ps'.format(d_conn_p * 1e12))
# connector model with guessed loss
half = m.line(d_line/2, 's', z0=Z_line)
mc = DefinedAEpTandZ0(m.frequency, ep_r=1, tanD=0.025, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
left = mc.line(d_conn_p, 's', z0=Z_conn)
right = left.flipped()
check = mc.thru() ** left ** half ** mc.thru()
plt.figure()
plt.suptitle('Connector + half thru comparison')
plt.subplot(2,1,1)
conn.plot_s_deg(1, 0, label='measured')
check.plot_s_deg(1, 0, label='model')
plt.ylabel('phase (rad)')
plt.legend()
plt.subplot(2,1,2)
conn.plot_s_db(1, 0, label='Measured')
check.plot_s_db(1, 0, label='Model')
plt.xlabel('Frequency (GHz)')
plt.ylabel('Insertion Loss (dB)')
plt.legend()
plt.show()
```
Connector + thru plots shows a reasonable agreement between calibration results and model. There is a phase jump in the calibration results.
## Final check
```
DUT = m.line(d_line, 's', Z_line)
DUT.name = 'model'
Check = m.thru() ** left ** DUT ** right ** m.thru()
Check.name = 'model with connectors'
plt.figure()
TL100.plot_s_db()
Check.plot_s_db(1,0, color='k')
Check.plot_s_db(0,0, color='k')
plt.show()
Check_dc = Check.extrapolate_to_dc(kind='linear')
plt.figure()
plt.suptitle('Time domain step-response')
ax = plt.subplot(1,1,1)
TL100_dc.s11.plot_z_time_step(pad=2000, window='hamming', label='Measured', ax=ax, color='k')
Check_dc.s11.plot_z_time_step(pad=2000, window='hamming', label='Model', ax=ax, color='b')
t0 = -2
t1 = 4
ax.set_xlim(t0, t1)
ax.xaxis.set_minor_locator(AutoMinorLocator(10))
ax.yaxis.set_minor_locator(AutoMinorLocator(5))
ax.patch.set_facecolor('1.0')
ax.grid(True, color='0.8', which='minor')
ax.grid(True, color='0.5', which='major')
```
The plots shows a reasonnable agreement between model and measurement up to 4 GHz.
Further works may include implementing CPWG medium or modeling the line by more sections to account the impedance variation vs. position.
| github_jupyter |
# Getting an Overview of Regular 3D Data
In this notebook, we're going to talk a little bit about how you might get an overview of regularized 3D data, specifically using matplotlib.
In a subsequent notebook we'll address the next few steps, specifically how you might use tools like ipyvolume and yt.
To start with, let's generate some fake data! (Now, I say 'fake,' but that's a bit pejorative, isn't it? Data is data! Ours is just synthetic.)
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
import scipy.special
```
We'll use the scipy [spherical harmonics](https://docs.scipy.org/doc/scipy/reference/generated/scipy.special.sph_harm.html) function to make some data, but first we need a reference coordinate system. We'll start with $x, y, z$ and then transform them into spherical coordinates.
**Note**: we'll be using the convention that $\theta \in [0, \pi]$ and $\phi \in[0,2\pi)$, which is reverse from what SciPy expects. So if you compare to the docstring for sph_harm, keep that in mind. Feel free to switch the definitions if you like!
```
N = 64
x = np.mgrid[-1.0:1.0:N*1j][:,None,None]
y = np.mgrid[-1.0:1.0:N*1j][None,:,None]
z = np.mgrid[-1.0:1.0:N*1j][None,None,:]
r = np.sqrt(x*x + y*y + z*z)
theta = np.arctan2(np.sqrt(x*x + y*y), z)
phi = np.arctan2(y, x)
np.abs(x - r * np.sin(theta)*np.cos(phi)).max()
np.abs(y - r * np.sin(theta)*np.sin(phi)).max()
np.abs(z - r * np.cos(theta)).max()
data = {}
for n in [1, 4]:
for m in range(n + 1):
data[f"sph_n{n}_m{m}"] = np.absolute(scipy.special.sph_harm(m, n, phi, theta))
```
Now we have some data! And, we can use matplotlib to visualize it in *reduced* form. Let's try this out:
```
plt.imshow(data["sph_n4_m4"][:,:,N//4], norm=LogNorm())
plt.colorbar()
phi.min(), phi.max()
plt.imshow(data["sph_n1_m0"].max(axis=0), norm=LogNorm())
plt.colorbar()
```
This is getting a bit cumbersome, though! Let's try using the [`ipywidgets`](https://ipywidgets.readthedocs.org) library to speed this up just a bit.
We're going to use the `ipywidgets.interact` decorator around our function to add some inputs. This is a pretty powerful decorator, as it sets up new widgets based on the info that you feed it, and then re-executes the function every time those inputs change.
```
import ipywidgets
@ipywidgets.interact(dataset = list(sorted(data.keys())), slice_position = (0, N, 1))
def make_plots(dataset, slice_position):
plt.imshow(data[dataset][slice_position,:,:], norm=LogNorm())
plt.colorbar()
```
We still have some artifacts here we want to get rid of; let's see if we can restrict our colorbar a bit.
```
print(min(_.min() for _ in data.values()), max(_.max() for _ in data.values()))
```
Typically in these cases, the more interesting values are the ones at the top -- the bottom are usually falling off rather quickly to zero. So let's set our maximum, and then drop 5 orders of magnitude for the minimum. I'm changing the colorbar's "extend" value to reflect this.
```
@ipywidgets.interact(dataset = list(sorted(data.keys())), slice_position = (0, N, 1))
def make_plots(dataset, slice_position):
plt.imshow(data[dataset][slice_position,:,:], norm=LogNorm(vmin=1e-5, vmax=1.0))
plt.colorbar(extend = 'min')
```
We're going to do one more thing for getting an overview, and then we'll see if we can do some other, cooler things with it using plotly.
We're going to change our `slice_position` to be in units of actual coordinates, instead of integers, and we'll add on a multiplot so we can see all three at once.
```
@ipywidgets.interact(dataset = list(sorted(data.keys())), x = (-1.0, 1.0, 2.0/N), y = (-1.0, 1.0, 2.0/N), z = (-1.0, 1.0, 2.0/N))
def make_plots(dataset, x, y, z):
xi, yi, zi = (int(_*N + 1.0) for _ in (x, y, z))
fig, axes = plt.subplots(nrows=2, ncols=2, dpi = 200)
datax = data[dataset][xi,:,:]
datay = data[dataset][:,yi,:]
dataz = data[dataset][:,:,zi]
vmax = max(_.max() for _ in (datax, datay, dataz))
vmin = max( min(_.min() for _ in (datax, datay, dataz)), vmax / 1e5)
imx = axes[0][0].imshow(datax, norm=LogNorm(vmin=vmin, vmax=vmax), extent = [-1.0, 1.0, -1.0, 1.0])
imy = axes[0][1].imshow(datay, norm=LogNorm(vmin=vmin, vmax=vmax), extent = [-1.0, 1.0, -1.0, 1.0])
imz = axes[1][0].imshow(dataz, norm=LogNorm(vmin=vmin, vmax=vmax), extent = [-1.0, 1.0, -1.0, 1.0])
fig.delaxes(axes[1][1])
fig.colorbar(imx, ax=axes, extend = 'min', fraction = 0.1)
import plotly.graph_objects as go
plt.hist(data["sph_n4_m3"].flatten())
iso_data=go.Isosurface(
x=(x * np.ones((N,N,N))).flatten(),
y=(y * np.ones((N,N,N))).flatten(),
z=(z * np.ones((N,N,N))).flatten(),
value=data["sph_n4_m3"].flatten(),
isomin=0,
isomax=data["sph_n4_m3"].max(),
surface_count=5, # number of isosurfaces, 2 by default: only min and max
colorbar_nticks=5, # colorbar ticks correspond to isosurface values
caps=dict(x_show=False, y_show=False))
fig = go.Figure(data = iso_data)
fig
```
One thing I've run into with plotly while making this notebook has been that in many cases, the 3D plots strain a bit under large data sizes. This is to be expected, and is completely understandable! One of the really nice things about regular mesh data like this is that you can usually cut it down quite effectively with slices. Unfortunately, what I have found -- and I may have done something completely wrong! -- is that plotly some times appears to almost work, and then doesn't quite make it when I throw too much data at it. I've found that it seems to work best in the neighborhood of $64^3$ zones, maybe a bit more.
## Other Summary Techniques
There are, of course, other ways you can take a look at a set of values! Given a regular mesh, it's straightforward with numpy to apply any of the reduction operations along one of the axes. For instance, you might take the min, the max, the sum, the mean and so forth. If we do this with our spherical harmonics data:
```
plt.imshow(data["sph_n4_m3"].sum(axis=0), extent=[-1.0, 1.0, -1.0, 1.0])
```
One thing you might keep in mind, when doing things like sums, is that if your cells aren't equally spaced along an axis, your sum will not necessarily be what you expect! You may want to integrate instead, where you multiple by a path length.
| github_jupyter |
```
%run notebook_setup
```
*If you have not already read it, you may want to start with the first tutorial: [Getting started with The Joker](1-Getting-started.ipynb).*
# Continue generating samples with standard MCMC
When many prior samples are used with *The Joker*, and the sampler returns one sample, or the samples returned are within the same mode of the posterior, the posterior *pdf* is likely unimodal. In these cases, we can use standard MCMC methods to generate posterior samples, which will typically be much more efficient than *The Joker* itself. In this example, we will use `pymc3` to "continue" sampling for data that are very constraining.
First, some imports we will need later:
```
import astropy.coordinates as coord
import astropy.table as at
from astropy.time import Time
import astropy.units as u
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
import corner
import pymc3 as pm
import pymc3_ext as pmx
import exoplanet as xo
import arviz as az
import thejoker as tj
# set up a random number generator to ensure reproducibility
rnd = np.random.default_rng(seed=42)
```
Here we will again load some pre-generated data meant to represent well-sampled, precise radial velocity observations of a single luminous source with a single companion (we again downsample the data set here just for demonstration):
```
data_tbl = at.QTable.read('data.ecsv')
sub_tbl = data_tbl[rnd.choice(len(data_tbl), size=18, replace=False)] # downsample data
data = tj.RVData.guess_from_table(sub_tbl, t_ref=data_tbl.meta['t_ref'])
_ = data.plot()
```
We will use the default prior, but feel free to play around with these values:
```
prior = tj.JokerPrior.default(
P_min=2*u.day, P_max=1e3*u.day,
sigma_K0=30*u.km/u.s,
sigma_v=100*u.km/u.s)
```
The data above look fairly constraining: it would be hard to draw many distinct orbital solutions through the RV data plotted above. In cases like this, we will often only get back 1 or a few samples from *The Joker* even if we use a huge number of prior samples. Since we are only going to use the samples from *The Joker* to initialize standard MCMC, we will only use a moderate number of prior samples:
```
prior_samples = prior.sample(size=250_000,
random_state=rnd)
joker = tj.TheJoker(prior, random_state=rnd)
joker_samples = joker.rejection_sample(data, prior_samples,
max_posterior_samples=256)
joker_samples
joker_samples.tbl
_ = tj.plot_rv_curves(joker_samples, data=data)
```
The sample that was returned by *The Joker* does look like it is a reasonable fit to the RV data, but to fully explore the posterior *pdf* we will use standard MCMC through `pymc3`. Here we will use the NUTS sampler, but you could also experiment with other backends (e.g., Metropolis-Hastings, or even `emcee` by [following this blog post](https://dfm.io/posts/emcee-pymc3/)):
```
with prior.model:
mcmc_init = joker.setup_mcmc(data, joker_samples)
trace = pmx.sample(tune=500, draws=500,
start=mcmc_init,
cores=1, chains=2)
```
If you get warnings from running the sampler above, they usually indicate that we should run the sampler for many more steps to tune the sampler and for our main run, but let's ignore that for now. With the MCMC traces in hand, we can summarize the properties of the chains using `pymc3.summary`:
```
az.summary(trace, var_names=prior.par_names)
```
To convert the trace into a `JokerSamples` instance, we can use the `TheJoker.trace_to_samples()` method. Note here that the sign of `K` is arbitrary, so to compare to the true value, we also call `wrap_K()` to store only the absolute value of `K` (which also increases `omega` by π, to stay consistent):
```
mcmc_samples = joker.trace_to_samples(trace, data)
mcmc_samples.wrap_K()
mcmc_samples
```
We can now compare the samples we got from MCMC to the true orbital parameters used to generate this data:
```
import pickle
with open('true-orbit.pkl', 'rb') as f:
truth = pickle.load(f)
# make sure the angles are wrapped the same way
if np.median(mcmc_samples['omega']) < 0:
truth['omega'] = coord.Angle(truth['omega']).wrap_at(np.pi*u.radian)
if np.median(mcmc_samples['M0']) < 0:
truth['M0'] = coord.Angle(truth['M0']).wrap_at(np.pi*u.radian)
df = mcmc_samples.tbl.to_pandas()
truths = []
colnames = []
for name in df.columns:
if name in truth:
colnames.append(name)
truths.append(truth[name].value)
_ = corner.corner(df[colnames], truths=truths)
```
Overall, it looks like we do recover the input parameters!
| github_jupyter |
<h1>Accidents Example</h1>
```
from GreyNsights.analyst import Pointer, Command, Analyst
from GreyNsights.client import DataWorker, DataSource
from GreyNsights.frameworks import framework
import numpy as np
```
This notebook demonstrates how to use GreyNSights on a remote dataset hosted by some dataowner. The primary aim of this example is to show pandas could be used as it is across a wide range of queries to analyze and explore a remote datasource. For running this example first run datasource.py , this begins the datasource server which hosts the dataset and executes the requests made from this notebook.
```
#Pandas version of GreyNsights that performs queries remotely
frameworks = framework()
pandas = frameworks.pandas
```
The analyst identity doesn't actually have any underlying functionality for now , but it is a placeholder for the future such as providing an actual identity in terms of certificate.
```
identity = Analyst("Alice", port=65441, host="127.0.0.1")
```
This connects to the remote dataowner
```
worker = DataWorker(port=65441, host="127.0.0.1")
dataset = DataSource(identity, worker, "Sample Data")
```
Get the config of data owner to understand the limitations set on the private dataset for querying
```
a = dataset.get_config()
print(a)
a = a.approve().init_pointer()
```
Create a dataframe from the dataset (Its already a dataframe but to demonstrate GreyNSights pandas remote execution)
```
df = pandas.DataFrame(a)
```
Variables and functions can be sent remotely for execution using function send(). The send() returns a pointer to the variable that now lives remotely.
```
p = 3
p = dataset.send(p)
# last 5 rows
print(df.tail(p))
print(df)
```
The below operation performs operation on the pointer which ensures the operation is executed remotely by datasource. The original results are returned only when the get function is called.The exact same functionalities as Pandas dataframes can be performed.
```
print(df["TMC"])
print(df["TMC"].sum())
print(df["TMC"].sum().get())
print("TMC sum: ", df["TMC"].sum().get())
print("TMC std: ", df["TMC"].std().get())
print("Severity mean: ", df["Severity"].mean().get())
```
The number of rows should be queried as a differentially private count. This reflects dimension of dataset but not the number of rows.
```
df.shape
print("COLUMNS: ", df.columns)
df.columns = [
"ID",
"Source",
"TMC",
"Severity",
"Start_Time",
"End_Time",
"Start_Lat",
"Start_Lng",
"End_Lat",
"End_Lng",
"Distance_mi",
"Description",
"Number",
"Street",
"Side",
"City",
"County",
"State",
"Zipcode",
"Country",
"Timezone",
"Airport_Code",
"Weather_Timestamp",
"Temperature_F",
"Wind_Chill_F",
"Humidity_%",
"Pressure_in",
"Visibility_mi",
"Wind_Direction",
"Wind_Speed_mph",
"Precipitation_in",
"Weather_Condition",
"Amenity",
"Bump",
"Crossing",
"Give_Way",
"Junction",
"No_Exit",
"Railway",
"Roundabout",
"Station",
"Stop",
"Traffic_Calming",
"Traffic_Signal",
"Turning_Loop",
"Sunrise_Sunset",
"Civil_Twilight",
"Nautical_Twilight",
"Astronomical_Twilight",
]
```
<h3>Transforming original dataset into a subset of columns</h3>
```
df = df[
[
"ID",
"Source",
"TMC",
"Severity",
"Start_Time",
"End_Time",
"Start_Lat",
"Start_Lng",
"End_Lat",
"End_Lng",
]
]
```
<h3>A wide range of data transformations applied on pointers</h3>
```
df["Somecol"] = (df["TMC"] + df["Severity"] / 10) / 2
(df["TMC"] + df["Severity"])
df["Somecol"] = df["TMC"] + df["Severity"]
(df["TMC"] + df["Severity"] / 10) / 2
df["TMC"] > 2
(df["Severity"] > 8) | (df["TMC"] > 200)
df[df["TMC"] > 200]
df[(df["Severity"] > 8) | (df["TMC"] > 200)]
And_df = df[(df["TMC"] > 200)]
# Multiple conditions: OR
Or_df = df[(df["Severity"] > 8) | (df["TMC"] > 200)]
And_df["TMC"].mean().get()
Or_df["TMC"].mean().get()
```
<h3>Sending a function across and passing pointers as arguments</h3>
```
def somefunc(x):
return x + 2
somefunc_pt = dataset.send(somefunc)
df["Somecol"] = df["TMC"].apply(somefunc_pt)
print(df["Somecol"])
df["Somecol"].mean().get()
```
| github_jupyter |
# NumPy under MinPy, with GPU
This part of tutorial is also available in step-by-step notebook version on [github](https://github.com/dmlc/minpy/blob/master/examples/tutorials/numpy_under_minpy.ipynb). Please try it out!
## Basic NDArray Operation
MinPy has the same syntax as NumPy, which is the language of choice for numerical computing, and in particular deep learning. The popular [Stanford course cs231n](http://cs231n.stanford.edu/syllabus.html)
uses NumPy as its main coursework. To use NumPy under MinPy, you only need to replace `import numpy as np` with `import minpy.numpy as np` at the header of your NumPy program. if you are not familiar with NumPy, you may want to look up [NumPy Quickstart Tutorial](https://docs.scipy.org/doc/numpy-dev/user/quickstart.html) for more details.
Using NumPy under MinPy has two simple but important reasons, one for productivity and another for performance: 1) Auto-differentiation, and 2) GPU/CPU co-execution. We will discuss them in this tutorial.
But first, let us review some of the most common usages of NumPy.
### Array Creation
An array can be created in multiple ways. For example, we can create an array from a regular Python list or tuple by using the `array` function
```
import minpy.numpy as np
a = np.array([1,2,3]) # create a 1-dimensional array with a python list
b = np.array([[1,2,3], [2,3,4]]) # create a 2-dimensional array with a nested python list
```
Here are some useful ways to create arrays with initial placeholder content.
```
a = np.zeros((2,3)) # create a 2-dimensional array full of zeros with shape (2,3)
b = np.ones((2,3)) # create a same shape array full of ones
c = np.full((2,3), 7) # create a same shape array with all elements set to 7
d = np.empty((2,3)) # create a same shape whose initial content is random and depends on the state of the memory
```
### Basic Operations
Arithmetic operators on arrays apply *elementwise*, with a new array holding result.
```
a = np.ones((2,3))
b = np.ones((2,3))
c = a + b # elementwise plus
d = - c # elementwise minus
print(d)
e = np.sin(c**2).T # elementwise pow and sin, and then transpose
print(e)
f = np.maximum(a, c) # elementwise max
print(f)
```
### Indexing and Slicing
The slice operator `[]` applies on axis 0.
```
a = np.arange(6)
a = np.reshape(a, (3,2))
print(a[:])
# assign -1 to the 2nd row
a[1:2] = -1
print(a)
```
We can also slice a particular axis with the method `slice_axis`
```
# slice out the 2nd column
d = np.slice_axis(a, axis=1, begin=1, end=2)
print(d)
```
## AutoGrad Feature
If you work in a policy mode called `NumpyOnlyPolicy` (refer [here](https://minpy.readthedocs.io/en/latest/feature/policy.html) for more details), MinPy is almost compatible with the most of NumPy usages. But what makes MinPy awesome is that it give you the power of autograd, saving you from writing the most tedious and error prone part of deep net implementation:
```
from minpy.core import grad
# define a function: f(x) = 5*x^2 + 3*x - 2
def foo(x):
return 5*(x**2) + 3*x - 2
# f(4) = 90
print(foo(4))
# get the derivative function by `grad`: f'(x) = 10*x + 3
d_foo = grad(foo)
# f'(4) = 43.0
print(d_foo(4))
```
More details about this part can be found in [Autograd Tutorial](http://minpy.readthedocs.io/en/latest/tutorial/autograd_tutorial.html).
## GPU Support
But we do not stop here, we want MinPy not only friendly to use, but also fast. To this end, MinPy leverages GPU's parallel computing ability. The code below shows our GPU support and a set of API to make you freely to change the runnning context (i.e. to run on CPU or GPU). You can refer to [Select Context for MXNet](http://minpy.readthedocs.io/en/latest/feature/context.html) for more details.
```
import minpy.numpy as np
import minpy.numpy.random as random
from minpy.context import cpu, gpu
import time
n = 100
with cpu():
x_cpu = random.rand(1024, 1024) - 0.5
y_cpu = random.rand(1024, 1024) - 0.5
# dry run
for i in xrange(10):
z_cpu = np.dot(x_cpu, y_cpu)
z_cpu.asnumpy()
# real run
t0 = time.time()
for i in xrange(n):
z_cpu = np.dot(x_cpu, y_cpu)
z_cpu.asnumpy()
t1 = time.time()
with gpu(0):
x_gpu0 = random.rand(1024, 1024) - 0.5
y_gpu0 = random.rand(1024, 1024) - 0.5
# dry run
for i in xrange(10):
z_gpu0 = np.dot(x_gpu0, y_gpu0)
z_gpu0.asnumpy()
# real run
t2 = time.time()
for i in xrange(n):
z_gpu0 = np.dot(x_gpu0, y_gpu0)
z_gpu0.asnumpy()
t3 = time.time()
print("run on cpu: %.6f s/iter" % ((t1 - t0) / n))
print("run on gpu: %.6f s/iter" % ((t3 - t2) / n))
```
The `asnumpy()` call is somewhat mysterious, implying `z_cpu` is not NumPy's `ndarray` type. Indeed this is true. For fast execution, MXNet maintains its own datastrcutre `NDArray`. This calls re-synced `z_cpu` into NumPy array.
As you can see, there is a gap between the speeds of matrix multiplication in CPU and GPU. That's why we set default policy mode as `PreferMXNetPolicy`, which means MinPy will dispatch the operator to MXNet as much as possible for you, and achieve transparent fallback while there is no MXNet implementation. MXNet operations run on GPU, whereas the fallbacks run on CPU.
See [Transparent Fallback](https://minpy.readthedocs.io/en/latest/tutorial/transparent_fallback.html) for more details.
## Something You Need to Know
With [Transparent Fallback](http://minpy.readthedocs.io/en/latest/tutorial/transparent_fallback.html), we hope to transparently upgrade the running speed without your changing a line of code. This can be done by expanding the MXNet GPU operators.
However, there are some important [pitfalls](http://minpy.readthedocs.io/en/latest/feature/limitation.html) you should know when you try to use MinPy, we strongly suggest that you should read it next.
| github_jupyter |
### Dependencies
```
import os
import cv2
import math
import random
import shutil
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import albumentations as albu
import matplotlib.pyplot as plt
from tensorflow import set_random_seed
from sklearn.model_selection import train_test_split
from keras import optimizers
from keras import backend as K
from keras.utils import Sequence
from keras.losses import binary_crossentropy
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint, Callback
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(seed)
seed = 0
seed_everything(seed)
warnings.filterwarnings("ignore")
!pip install segmentation-models
import segmentation_models as sm
```
### Load data
```
train = pd.read_csv('../input/understanding_cloud_organization/train.csv')
submission = pd.read_csv('../input/understanding_cloud_organization/sample_submission.csv')
print('Number of train samples:', train.shape[0])
print('Number of test samples:', submission.shape[0])
# Preprocecss data
train['image'] = train['Image_Label'].apply(lambda x: x.split('_')[0])
train['label'] = train['Image_Label'].apply(lambda x: x.split('_')[1])
submission['image'] = submission['Image_Label'].apply(lambda x: x.split('_')[0])
test = pd.DataFrame(submission['image'].unique(), columns=['image'])
train_pivoted = pd.pivot_table(train, index=['image'], values=['EncodedPixels'], columns=['label'], aggfunc=np.min).reset_index()
train_pivoted.columns = ['image', 'Fish', 'Flower', 'Gravel', 'Sugar']
display(train_pivoted.head())
```
### Split train and validation sets
```
X_train, X_val = train_test_split(train_pivoted, test_size=0.2, random_state=seed)
X_train['set'] = 'train'
X_val['set'] = 'validation'
test['set'] = 'test'
print('Number of train samples:', X_train.shape[0])
print('Number of validation samples:', X_val.shape[0])
```
# Model parameters
```
model_path = '../working/uNet_ResNet34_baseline.h5'
BACKBONE = 'resnet34'
BATCH_SIZE = 32
EPOCHS = 15
LEARNING_RATE = 1e-4
HEIGHT = 320
WIDTH = 480
CHANNELS = 3
N_CLASSES = train['label'].nunique()
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
LR_WARMUP_EPOCHS = 5
WARMUP_BATCHES = (LR_WARMUP_EPOCHS * len(X_train)) // BATCH_SIZE
```
### Auxiliary functions
```
def np_resize(img, input_shape):
height, width = input_shape
return cv2.resize(img, (width, height))
def mask2rle(img):
pixels= img.T.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
def build_rles(masks, reshape=None):
width, height, depth = masks.shape
rles = []
for i in range(depth):
mask = masks[:, :, i]
if reshape:
mask = mask.astype(np.float32)
mask = np_resize(mask, reshape).astype(np.int64)
rle = mask2rle(mask)
rles.append(rle)
return rles
def build_masks(rles, input_shape, reshape=None):
depth = len(rles)
if reshape is None:
masks = np.zeros((*input_shape, depth))
else:
masks = np.zeros((*reshape, depth))
for i, rle in enumerate(rles):
if type(rle) is str:
if reshape is None:
masks[:, :, i] = rle2mask(rle, input_shape)
else:
mask = rle2mask(rle, input_shape)
reshaped_mask = np_resize(mask, reshape)
masks[:, :, i] = reshaped_mask
return masks
def rle2mask(rle, input_shape):
width, height = input_shape[:2]
mask = np.zeros( width*height ).astype(np.uint8)
array = np.asarray([int(x) for x in rle.split()])
starts = array[0::2]
lengths = array[1::2]
current_position = 0
for index, start in enumerate(starts):
mask[int(start):int(start+lengths[index])] = 1
current_position += lengths[index]
return mask.reshape(height, width).T
def dice_coefficient(img1, img2):
img1 = np.asarray(img1).astype(np.bool)
img2 = np.asarray(img2).astype(np.bool)
intersection = np.logical_and(img1, img2)
return 2. * intersection.sum() / (img1.sum() + img2.sum())
def dice_coef(y_true, y_pred, smooth=1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2. * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
def post_process(probability, threshold=0.5, min_size=10000):
mask = cv2.threshold(probability, threshold, 1, cv2.THRESH_BINARY)[1]
num_component, component = cv2.connectedComponents(mask.astype(np.uint8))
predictions = np.zeros(probability.shape, np.float32)
for c in range(1, num_component):
p = (component == c)
if p.sum() > min_size:
predictions[p] = 1
return predictions
def preprocess_image(image_id, base_path, save_path, HEIGHT=HEIGHT, WIDTH=WIDTH):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (WIDTH, HEIGHT))
cv2.imwrite(save_path + image_id, image)
def preprocess_data(df, HEIGHT=HEIGHT, WIDTH=WIDTH):
df = df.reset_index()
for i in range(df.shape[0]):
item = df.iloc[i]
image_id = item['image']
item_set = item['set']
if item_set == 'train':
preprocess_image(image_id, train_base_path, train_images_dest_path)
if item_set == 'validation':
preprocess_image(image_id, train_base_path, validation_images_dest_path)
if item_set == 'test':
preprocess_image(image_id, test_base_path, test_images_dest_path)
def get_metrics(model, df, generator, set_name='Complete set'):
generator.shuffle = False
generator.augment = None
generator.batch_size = 1
column_names = ['Fish', 'Flower', 'Gravel', 'Sugar', set_name]
index_name = ['Dice Coeff']
dice = []
dice_post = []
for sample in range(len(df)):
x, y = generator.__getitem__(sample)
preds = model.predict(x)[0]
sample_dice = []
sample_dice_post = []
for class_index in range(N_CLASSES):
label_mask = y[..., class_index]
pred_mask = preds[..., class_index]
class_dice = dice_coefficient(pred_mask, label_mask)
pred_mask_post = post_process(pred_mask)
class_dice_post = dice_coefficient(pred_mask_post, label_mask)
if math.isnan(class_dice_post):
class_dice_post = 0.0
sample_dice.append(class_dice)
sample_dice_post.append(class_dice_post)
dice.append(sample_dice)
dice_post.append(sample_dice_post)
dice_class = np.mean(dice, axis=0)
dice = np.mean(dice_class, axis=0)
metrics = np.append(dice_class, dice)
metrics = pd.DataFrame(metrics.reshape(1, metrics.shape[0]), columns=column_names, index=index_name)
dice_class_post = np.mean(dice_post, axis=0)
dice_post = np.mean(dice_class_post, axis=0)
metrics_post = np.append(dice_class_post, dice_post)
metrics_post = pd.DataFrame(metrics_post.reshape(1, metrics_post.shape[0]), columns=column_names, index=index_name)
return metrics, metrics_post
def plot_metrics(history):
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, sharex='col', figsize=(20, 14))
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['dice_coef'], label='Train Dice coefficient')
ax2.plot(history['val_dice_coef'], label='Validation Dice coefficient')
ax2.legend(loc='best')
ax2.set_title('Dice coefficient')
ax3.plot(history['score'], label='Train F-Score')
ax3.plot(history['val_score'], label='Validation F-Score')
ax3.legend(loc='best')
ax3.set_title('F-Score')
plt.xlabel('Epochs')
sns.despine()
plt.show()
class WarmUpLearningRateScheduler(Callback):
def __init__(self, warmup_batches, init_lr, verbose=0):
"""
Constructor for warmup learning rate scheduler
:param warmup_batches {int}: Number of batch for warmup.
:param init_lr {float}: Learning rate after warmup.
:param verbose {int}: 0: quiet, 1: update messages. (default: {0})
"""
super(WarmUpLearningRateScheduler, self).__init__()
self.warmup_batches = warmup_batches
self.init_lr = init_lr
self.verbose = verbose
self.batch_count = 0
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.batch_count = self.batch_count + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
if self.batch_count <= self.warmup_batches:
lr = self.batch_count * self.init_lr / self.warmup_batches
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: WarmUpLearningRateScheduler setting learning rate to %s.' % (self.batch_count + 1, lr))
class RAdam(optimizers.Optimizer):
"""RAdam optimizer.
# Arguments
lr: float >= 0. Learning rate.
beta_1: float, 0 < beta < 1. Generally close to 1.
beta_2: float, 0 < beta < 1. Generally close to 1.
epsilon: float >= 0. Fuzz factor. If `None`, defaults to `K.epsilon()`.
decay: float >= 0. Learning rate decay over each update.
weight_decay: float >= 0. Weight decay for each param.
amsgrad: boolean. Whether to apply the AMSGrad variant of this
algorithm from the paper "On the Convergence of Adam and
Beyond".
# References
- [Adam - A Method for Stochastic Optimization](https://arxiv.org/abs/1412.6980v8)
- [On the Convergence of Adam and Beyond](https://openreview.net/forum?id=ryQu7f-RZ)
- [On The Variance Of The Adaptive Learning Rate And Beyond](https://arxiv.org/pdf/1908.03265v1.pdf)
"""
def __init__(self, lr=0.001, beta_1=0.9, beta_2=0.999,
epsilon=None, decay=0., weight_decay=0., amsgrad=False, **kwargs):
super(RAdam, self).__init__(**kwargs)
with K.name_scope(self.__class__.__name__):
self.iterations = K.variable(0, dtype='int64', name='iterations')
self.lr = K.variable(lr, name='lr')
self.beta_1 = K.variable(beta_1, name='beta_1')
self.beta_2 = K.variable(beta_2, name='beta_2')
self.decay = K.variable(decay, name='decay')
self.weight_decay = K.variable(weight_decay, name='weight_decay')
if epsilon is None:
epsilon = K.epsilon()
self.epsilon = epsilon
self.initial_decay = decay
self.initial_weight_decay = weight_decay
self.amsgrad = amsgrad
def get_updates(self, loss, params):
grads = self.get_gradients(loss, params)
self.updates = [K.update_add(self.iterations, 1)]
lr = self.lr
if self.initial_decay > 0:
lr = lr * (1. / (1. + self.decay * K.cast(self.iterations, K.dtype(self.decay))))
t = K.cast(self.iterations, K.floatx()) + 1
ms = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='m_' + str(i)) for (i, p) in enumerate(params)]
vs = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='v_' + str(i)) for (i, p) in enumerate(params)]
if self.amsgrad:
vhats = [K.zeros(K.int_shape(p), dtype=K.dtype(p), name='vhat_' + str(i)) for (i, p) in enumerate(params)]
else:
vhats = [K.zeros(1, name='vhat_' + str(i)) for i in range(len(params))]
self.weights = [self.iterations] + ms + vs + vhats
beta_1_t = K.pow(self.beta_1, t)
beta_2_t = K.pow(self.beta_2, t)
sma_inf = 2.0 / (1.0 - self.beta_2) - 1.0
sma_t = sma_inf - 2.0 * t * beta_2_t / (1.0 - beta_2_t)
for p, g, m, v, vhat in zip(params, grads, ms, vs, vhats):
m_t = (self.beta_1 * m) + (1. - self.beta_1) * g
v_t = (self.beta_2 * v) + (1. - self.beta_2) * K.square(g)
m_corr_t = m_t / (1.0 - beta_1_t)
if self.amsgrad:
vhat_t = K.maximum(vhat, v_t)
v_corr_t = K.sqrt(vhat_t / (1.0 - beta_2_t) + self.epsilon)
self.updates.append(K.update(vhat, vhat_t))
else:
v_corr_t = K.sqrt(v_t / (1.0 - beta_2_t) + self.epsilon)
r_t = K.sqrt((sma_t - 4.0) / (sma_inf - 4.0) *
(sma_t - 2.0) / (sma_inf - 2.0) *
sma_inf / sma_t)
p_t = K.switch(sma_t > 5, r_t * m_corr_t / v_corr_t, m_corr_t)
if self.initial_weight_decay > 0:
p_t += self.weight_decay * p
p_t = p - lr * p_t
self.updates.append(K.update(m, m_t))
self.updates.append(K.update(v, v_t))
new_p = p_t
# Apply constraints.
if getattr(p, 'constraint', None) is not None:
new_p = p.constraint(new_p)
self.updates.append(K.update(p, new_p))
return self.updates
def get_config(self):
config = {
'lr': float(K.get_value(self.lr)),
'beta_1': float(K.get_value(self.beta_1)),
'beta_2': float(K.get_value(self.beta_2)),
'decay': float(K.get_value(self.decay)),
'weight_decay': float(K.get_value(self.weight_decay)),
'epsilon': self.epsilon,
'amsgrad': self.amsgrad,
}
base_config = super(RAdam, self).get_config()
return dict(list(base_config.items()) + list(config.items()))
```
# Pre-process data
```
train_base_path = '../input/understanding_cloud_organization/train_images/'
test_base_path = '../input/understanding_cloud_organization/test_images/'
train_images_dest_path = 'base_dir/train_images/'
validation_images_dest_path = 'base_dir/validation_images/'
test_images_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_images_dest_path):
shutil.rmtree(train_images_dest_path)
if os.path.exists(validation_images_dest_path):
shutil.rmtree(validation_images_dest_path)
if os.path.exists(test_images_dest_path):
shutil.rmtree(test_images_dest_path)
# Creating train, validation and test directories
os.makedirs(train_images_dest_path)
os.makedirs(validation_images_dest_path)
os.makedirs(test_images_dest_path)
n_cpu = mp.cpu_count()
train_n_cnt = X_train.shape[0]//n_cpu
val_n_cnt = X_val.shape[0]//n_cpu
test_n_cnt = test.shape[0]//n_cpu
# Pre-procecss train set
pool = mp.Pool(n_cpu)
dfs = [X_train.iloc[train_n_cnt*i:train_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_train.iloc[train_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss validation set
pool = mp.Pool(n_cpu)
dfs = [X_val.iloc[val_n_cnt*i:val_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_val.iloc[val_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss test set
pool = mp.Pool(n_cpu)
dfs = [test.iloc[test_n_cnt*i:test_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = test.iloc[test_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
preprocessing = sm.backbones.get_preprocessing(BACKBONE)
augmentation = albu.Compose([albu.HorizontalFlip(p=0.5),
albu.VerticalFlip(p=0.5)
])
```
### Data generator
```
class DataGenerator(Sequence):
def __init__(self, df, target_df=None, mode='fit', base_path=train_images_dest_path,
batch_size=BATCH_SIZE, n_channels=CHANNELS, reshape=(HEIGHT, WIDTH),
n_classes=N_CLASSES, random_state=seed, shuffle=True, preprocessing=None, augmentation=None):
self.batch_size = batch_size
self.df = df
self.mode = mode
self.base_path = base_path
self.target_df = target_df
self.reshape = reshape
self.n_channels = n_channels
self.n_classes = n_classes
self.shuffle = shuffle
self.augmentation = augmentation
self.preprocessing = preprocessing
self.list_IDs = self.df.index
self.random_state = random_state
self.mask_shape = (1400, 2100)
if self.random_state is not None:
np.random.seed(self.random_state)
self.on_epoch_end()
def __len__(self):
return len(self.list_IDs) // self.batch_size
def __getitem__(self, index):
indexes = self.indexes[index*self.batch_size:(index+1)*self.batch_size]
list_IDs_batch = [self.list_IDs[k] for k in indexes]
X = self.__generate_X(list_IDs_batch)
if self.mode == 'fit':
Y = self.__generate_y(list_IDs_batch)
if self.augmentation:
X, Y = self.__augment_batch(X, Y)
return X, Y
elif self.mode == 'predict':
return X
def on_epoch_end(self):
self.indexes = np.arange(len(self.list_IDs))
if self.shuffle == True:
np.random.shuffle(self.indexes)
def __generate_X(self, list_IDs_batch):
X = np.empty((self.batch_size, *self.reshape, self.n_channels))
for i, ID in enumerate(list_IDs_batch):
im_name = self.df['image'].loc[ID]
img_path = self.base_path + im_name
img = cv2.imread(img_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
if self.preprocessing:
img = self.preprocessing(img)
# img = img.astype(np.float32) / 255.
X[i,] = img
return X
def __generate_y(self, list_IDs_batch):
Y = np.empty((self.batch_size, *self.reshape, self.n_classes), dtype=int)
for i, ID in enumerate(list_IDs_batch):
im_name = self.df['image'].loc[ID]
image_df = self.target_df[self.target_df['image'] == im_name]
rles = image_df['EncodedPixels'].values
masks = build_masks(rles, input_shape=self.mask_shape, reshape=self.reshape)
Y[i, ] = masks
return Y
def __augment_batch(self, img_batch, masks_batch):
for i in range(img_batch.shape[0]):
img_batch[i, ], masks_batch[i, ] = self.__random_transform(img_batch[i, ], masks_batch[i, ])
return img_batch, masks_batch
def __random_transform(self, img, masks):
composed = self.augmentation(image=img, mask=masks)
aug_img = composed['image']
aug_masks = composed['mask']
return aug_img, aug_masks
train_generator = DataGenerator(
base_path=train_images_dest_path,
df=X_train,
target_df=train,
batch_size=BATCH_SIZE,
reshape=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
augmentation=augmentation,
random_state=seed)
valid_generator = DataGenerator(
base_path=validation_images_dest_path,
df=X_val,
target_df=train,
batch_size=BATCH_SIZE,
reshape=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
random_state=seed)
```
# Model
```
model = sm.Unet(encoder_name=BACKBONE,
encoder_weights='imagenet',
classes=N_CLASSES,
activation='sigmoid',
input_shape=(HEIGHT, WIDTH, CHANNELS))
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True, save_weights_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
warmup_lr = WarmUpLearningRateScheduler(WARMUP_BATCHES, LEARNING_RATE)
loss = sm.losses.bce_dice_loss
metric_list = [dice_coef, sm.metrics.f1_score]
callback_list = [checkpoint, es, rlrop, warmup_lr]
optimizer = RAdam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss=loss, metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = len(X_train)//BATCH_SIZE
STEP_SIZE_VALID = len(X_val)//BATCH_SIZE
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
callbacks=callback_list,
epochs=EPOCHS,
verbose=2).history
```
## Model loss graph
```
plot_metrics(history)
```
# Model evaluation
## Without post processing
```
# Train metrics
train_metrics, train_metrics_post = get_metrics(model, X_train, train_generator, 'Train')
display(train_metrics)
# Validation metrics
validation_metrics, validation_metrics_post = get_metrics(model, X_val, valid_generator, 'Validation')
display(validation_metrics)
```
## With post processing
```
display(train_metrics_post)
display(validation_metrics_post)
```
# Apply model to test set
```
test_df = []
for i in range(0, test.shape[0], 500):
batch_idx = list(range(i, min(test.shape[0], i + 500)))
batch_set = test[batch_idx[0]: batch_idx[-1]+1]
test_generator = DataGenerator(
base_path=test_images_dest_path,
df=batch_set,
target_df=submission,
batch_size=1,
reshape=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
random_state=seed,
mode='predict',
shuffle=False)
preds = model.predict_generator(test_generator)
for index, b in enumerate(batch_idx):
filename = test['image'].iloc[b]
image_df = submission[submission['image'] == filename].copy()
pred_masks = preds[index, ].round().astype(int)
pred_rles = build_rles(pred_masks, reshape=(350, 525))
image_df['EncodedPixels'] = pred_rles
### Post procecssing
pred_masks_post = preds[index, ].astype('float32')
for k in range(pred_masks_post.shape[-1]):
pred_mask = pred_masks_post[...,k]
pred_mask = post_process(pred_mask)
pred_masks_post[...,k] = pred_mask
pred_rles_post = build_rles(pred_masks_post, reshape=(350, 525))
image_df['EncodedPixels_post'] = pred_rles_post
###
test_df.append(image_df)
sub_df = pd.concat(test_df)
```
### Regular submission
```
submission_df = sub_df[['Image_Label' ,'EncodedPixels']]
submission_df.to_csv('submission.csv', index=False)
display(submission_df.head())
```
### Submission with post processing
```
submission_df_post = sub_df[['Image_Label' ,'EncodedPixels_post']]
submission_df_post.columns = ['Image_Label' ,'EncodedPixels']
submission_df_post.to_csv('submission_post.csv', index=False)
display(submission_df_post.head())
# Cleaning created directories
if os.path.exists(train_images_dest_path):
shutil.rmtree(train_images_dest_path)
if os.path.exists(validation_images_dest_path):
shutil.rmtree(validation_images_dest_path)
if os.path.exists(test_images_dest_path):
shutil.rmtree(test_images_dest_path)
```
| github_jupyter |
# mParticle Real Time Data + Personalize
In this module you are going to be adding the ability to maintain a real-time dataset that represents the latest user behavior for users of the Retail Demo Store. You will then connect that dataset to the Personalize dataset groups that you built in the first part of the workshop. This will enable your Personalize models to be kept up to date with the latest events your users are performing.
This workshop will use the mParticle web sdk npm library (https://github.com/mParticle/mparticle-web-sdk) to collect real-time data from the Retail Demo Store, and then feed that event data into the mParticle platform, where it can be routed directly to a Personalize Tracker, and then used to maintain the latest behavioral data for your personalization user-item interaction data.
*Recommended Time: 45 Minutes*
## Prerequisites
In order to complete this workshop, you will need to complete the 1.1-Personalize workbook in this directory. You will also need a mParticle workspace. If you are doing this workshop as part of a live workshop event, ask your moderator how to set up a mParticle workspace.
If you are running this workshop on your own, you can email or reach out to [dl-mparticle-workshops@mparticle.com](mailto:dl-mparticle-workshops@mparticle.com) to request for the creation of a mParticle account. We do not recommend using your production mParticle workspace for this workshop.
## mParticle Platform Overview
mParticle is a customer data platform (CDP) that helps you collect, clean, and control your customer data. mParticle provides several types of Sources which you can use to collect your data, and which you can choose from based on the needs of your app or site. For websites, you can use a javascript library to collect data. If you have a mobile app, you can embed one of mParticle’s Mobile SDKs, and if you’d like to create messages directly on a server (if you have, for example a dedicated .NET server that processes payments), mParticle has several server-based libraries that you can embed directly into your backend code. With mParticle, you can also use cloud-sources to import data about your app or site from other tools like Zendesk or Salesforce, to enrich the data sent through mParticle. By using mParticle to decouple data collection from data use, you can create a centralized data supply chain based on organized and modular data.
<img src="images/mparticle/mparticle_overview.png" height="1280" width="720">
## Setup
If you have already entered your mParticle API key and secret into your Cloud Formation deployment, you can skip to the next section.
mParticle uses *connections* as a way to organize data inputs into the platform. Configuring a input will allow you to collect real-time event data from the Retail Demo Store user interface, and pass that information to mParticle. You need to be signed into your mParticle workspace to begin this process. Once you are signed in to the mParticle console (https://app.mparticle.com), click on your workspace, and then ‘Setup’ in the left hand navigation bar of the screen. Then, click ‘Inputs’.
<img src="images/mparticle/mparticle-step-1.png" height="1280" width="720">
Select the ‘Web’ type within Platforms.
<img src="images/mparticle/mparticle-step-2.png" height="1280" width="720">
And click ‘Web’ then click Issue Keys.
<img src="images/mparticle/mparticle-step-3.png" height="1280" width="720">
mParticle will generate a pair of key and secret which you will use as part of the cloud formation template setup earlier.
<img src="images/mparticle/mparticle-step-4.png" height="1280" width="720">
Now that you are here, set the write key for your new source in the environment variable below.
You will need this in a few minutes, when you enable mParticle events collection in the Retail Demo Store.
Make sure you run the cell after you paste the key. This will allow us to set the mParticle API key in the web UI deployment, and pass the keys securely to other back-end services via SSM.
```
# THIS IS ONLY REQUIRED IF YOU DID NOT SET THE mPARTICLE API KEYS AND ORG ID IN YOUR ORIGINAL DEPLOYMENT
# IF YOU ARE RUNNING THIS IN A GUIDED WORKSHOP, YOU WILL NEED TO SET THESE VALUES BEFORE CONTINUING
mparticle_api_key = "us1-57db4f7a516ef34aa93df13e4ec11836"
mparticle_secret_key = "lC3b_BT3O9TEHuox2sRAWxesJx9uW1RNovvl6x8YrKSSxEIx9owdEEWsU9hT-P7c"
import boto3
import json
ssm = boto3.client('ssm')
iam = boto3.client('iam')
sts = boto3.client('sts')
aws_account_id = sts.get_caller_identity().get('Account')
region_name = boto3.Session().region_name
if mparticle_api_key:
response = ssm.put_parameter(
Name='/retaildemostore/webui/mparticle_api_key',
Value='{}'.format(mparticle_api_key),
Type='String',
Overwrite=True
)
if mparticle_secret_key:
response = ssm.put_parameter(
Name='/retaildemostore/webui/mparticle_secret_key',
Value='{}'.format(mparticle_secret_key),
Type='String',
Overwrite=True
)
print("mParticle API Key:")
print(mparticle_api_key)
print("mParticle Secret Key:")
print(mparticle_secret_key)
print("AWS Account ID:")
print(aws_account_id)
print("AWS Region:")
print(region_name)
```
You now have an environment variable that will enable the mParticle data collection library in the code in `AnalyticsHandler.js`. All we need to do now, is force a re-deploy of the Retail Demo Store.
To do that, go back to your AWS Console tab or window, and select Code Pipeline from the Services search. Then, find the pipeline name that contains `WebUIPipeline` and click that link.
Then, select the ‘Release Change’ button, and confirm the release once the popup shows up. You will see a confirmation that the pipeline is re-deploying the web ui for the Retail Demo Store.
This process should complete in a few minutes. Once this is done, you will see the bottom tile confirm that your deployment has completed.
Now that you have a working source, let's try to see if the Events from the Retail Demo Store are flowing into mParticle.
## Sending Real-Time Events via the Retail Demo Store
Navigate to your Retail Demo Store Web App and refresh the screen to reload the user interface. This will load the libraries you just deployed, and will allow your instance of the Retail Demo Store to send events to mParticle.
mParticle provides a variety of ways to collect real time events, and a full discussion of how this works is beyond the scope of this document, however the Retail Demo Store represents a fairly typical deployment for most web applications, in that it uses the mParticle Web SDK library, loaded via NPM, to inject their code into the web application.
To verify if mParticle JS is fully instantiated within the Retail Demo Store, just open developer console of your web browser and type
```javascript window.mParticle.Identity.getCurrentUser().getMPID()```
You should get the following response:
<img src="images/mparticle/mparticle-verification-step.png" height="1280" width="720">
Then, open another tab to the mParticle console, and select Live Stream under Data Master:
<img src="images/mparticle/mparticle-live-stream.png" height="1280" width="720">
You should see events collected by the mParticle SDK being streamed real-time within your mParticle instance. Feel free to view the events and see the actualy information hold per each event.
You can also try logging in or creating an account within the AWS Retail Demo Store Web application. Feel free to check the difference between a logged in user vs a user who is a guest within the retail demo store web app.
## Configure the mParticle Personalize Destination
mParticle uses Outputs to route real-time event data to a data consumer application. In this case, you will be using Amazon Kinesis as the destination. This destination will take real-time events from mParticle, pass them through an AWS Lambda function, and then into the user-item interactions dataset in your Retail Demo Store.
For this use case you will need to bring together mParticle and three AWS resources:
1.) An Amazon Kinesis stream to receive real-time events from mParticle
2.) An Amazon Personalize campaign to create product recommendations
3.) A Lambda function to act as a broker to transform data from Kinesis into a format accepted by Amazon Personalize (and ingested via a Personalize Event Tracker)
When you deployed the Retail Demo Store, a Cloud Formation template deployed a Kinesis stream and Lambda function for this workshop, as well as the necessary IAM account and policy for mParticle to write to your Kinesis stream. Let's connect these to your mParticle environment.
### Connect mParticle to Kinesis
mParticle offers an "event" output for streaming event data to Kinesis in real time. This can be set up and controlled from the mParticle dashboard without writing code. You can read an overview of event outputs in the mParticle docs (https://docs.mparticle.com/guides/getting-started/connect-an-event-output/).
Amazon Kinesis is an AWS service for processing streaming data. mParticle will forward commerce event data to Kinesis, where it will be picked up by the Lambda function you will set up in a moment.
Click ‘Directory’ in the left hand navigation bar of the screen, and then search ‘Amazon Kinesis’.
<img src="images/mparticle/mparticle-step-5.png" height="1280" width="720">
#### Create configuration
First, you will need to create an overall configuration for Kinesis. This holds all the settings that will remain the same for every input you connect.
<img src="images/mparticle/mparticle-step-6.png" height="1280" width="720">
To obtain an Access Key ID and Secret Access Key, please run the following code below and enter the generated Access Key ID and Secret in mParticle.
```
# Create keys for Kinesis
# The Uid is a unique ID and we need it to find the role made by CloudFormation
with open('/opt/ml/metadata/resource-metadata.json') as f:
data = json.load(f)
sagemaker = boto3.client('sagemaker')
sagemakerResponce = sagemaker.list_tags(ResourceArn=data["ResourceArn"])
for tag in sagemakerResponce["Tags"]:
if tag['Key'] == 'Uid':
Uid = tag['Value']
break
print('Uid:', Uid)
# policy JSON
# replace the region and account id
# arn:aws:kinesis:us-east-1:683819462896:stream/finalbuildtest-us-east-1-mParticlePersonalizeEventsKinesisStream
kinesisarn = "arn:aws:kinesis:"+region_name+":"+aws_account_id+":stream/"+Uid+"-mParticlePersonalizeEventsKinesisStream"
print('kinesisarn:', kinesisarn)
customPolicy = {
"Version": "2012-10-17",
"Statement": [
{
"Effect": "Allow",
"Action": [
"kinesis:PutRecord"
],
"Resource": [
kinesisarn
]
}
]
}
# create the policy
policy = iam.create_policy(
PolicyName='KinesismParticlePolicy',
PolicyDocument=json.dumps(customPolicy)
)
policy_arn = policy['Policy']['Arn']
print(policy_arn)
user_name = 'mParticleRetailDemoStoreKinesis'
#create user
created_user = iam.create_user(
UserName=user_name
)
print(created_user)
response = iam.attach_user_policy(
UserName=user_name,
PolicyArn=policy_arn
)
print(response)
#create programmatic access_key for mParticle
response = iam.create_access_key(
UserName=user_name
)
access_key_id = response['AccessKey']['AccessKeyId']
print(response)
# The AWS region you are running in is:
print(f'AWS Region: {region_name}')
```
#### Connect all sources
Next, you will connect the Retail Demo Store Web UI as an input: Web to Kinesis. To do that, click Connections then Connect. Select JS Web Platform as the Input
<img src="images/mparticle/mparticle-step-7.png" height="1280" width="720">
Click + Connect Output
<img src="images/mparticle/mparticle-step-8.png" height="1280" width="720">
Select Kinesis and the configuration You've just recently created earlier
<img src="images/mparticle/mparticle-step-9.png" height="1280" width="720">
The settings you need to provide here are the Amazon Region in which you deployed the Retail Demo Store and the name of the Kinesis stream in your environment. The region will depend on which region you are using in your AWS account or workshop account. The name of the Kinesis stream will be `mParticlePersonalizeEventsKinesisStream`. This was deployed for you when you deployed the workshop environment.
<img src="images/mparticle/mparticle-step-10.png" height="1280" width="720">
After setting up the Kinesis service region, make sure you untick all the checkboxes but leave Send eCommerce Events only ticked or selected. Click Save to save your settings. We only need to send eCommerece Events to Kinesis as these are the only events relevant for AWS Personalize.
<img src="images/mparticle/mparticle-kinesis-connection-config2.png" height="1280" width="720">
## Configure Lambda Parameters and Review Code
Before the destination can send events to your Amazon Personalize events tracker, you will need to tell the destination lambda where to send the events. It looks for an environment variable called 'personalize_tracking_id'.
Let's set that. Run the following cell to look up the relevant Amazon Personalize tracker from the Personalize workbook.
We can then set the appropriate value in the destination Lambda.
Within the mParticle Platform, navigate to Directory and within the search for Custom Feed.
<img src="images/mparticle/CustomFeed.png" height="1280" width="720">
Click Setup and it should generate you a pair of key and secret. The key and secret generated here will be the keys you'll used for your lambda environment configuration.
<img src="images/mparticle/CustomFeedDetails.png" height="1280" width="720">
```
# Set the Custom Feed Server to Server API Key and Secret from mParticle
mparticle_s2s_api_key = "us1-18cdb48aae1fb2459b805df5122f60a3"
mparticle_s2s_secret_key = "17VD4QWXLezi7_Qc5-RWF5kyxM4BgKN9Y_s5Krd4qspmCdtltU7fH-gNrFwGlBUN"
# Let's look up the appropriate tracking string
response = ssm.get_parameter(
Name='retaildemostore-personalize-event-tracker-id'
)
tracking_id = response['Parameter']['Value']
# Get the Campaign ARN
response = ssm.get_parameter(
Name='retaildemostore-product-recommendation-campaign-arn'
)
product_recommendation_arn = response['Parameter']['Value']
# set the Parameters via SSM
if mparticle_s2s_api_key:
response = ssm.put_parameter(
Name='/retaildemostore/webui/mparticle_s2s_api_key',
Value='{}'.format(mparticle_s2s_api_key),
Type='String',
Overwrite=True
)
if mparticle_s2s_secret_key:
response = ssm.put_parameter(
Name='/retaildemostore/webui/mparticle_s2s_secret_key',
Value='{}'.format(mparticle_s2s_secret_key),
Type='String',
Overwrite=True
)
#Print
print("mParticle S2S API Key:")
print(mparticle_s2s_api_key)
print("mParticle S2S Secret Key:")
print(mparticle_s2s_secret_key)
print("AWS Personalize Tracking ID:")
print(tracking_id)
print("AWS Product Recommendation ARN:")
print(product_recommendation_arn)
```
Go to your AWS console tab or window, and select Lambda from the Services menu.
Find the mParticlePersonalizeLambda, and click on it in the list.
<img src="images/mparticle/mparticle-find-lambda-function.png" height="1280" width="720">
Feel free to look at the Lambda code. Make sure the Kinesis component is added as a trigger in the Lambda. If the Kinesis component is not added, make sure to add it.
<img src="images/mparticle/addKinesisStreamLambda.png" height="1280" width="720">
If the Kinesis component is already added, verify if it is set to Enabled. If its not set to Enabled, you would need to enable the Kinesis configuration in your Lambda function. Most likely the Kinesis component will be in a disabled state when it was initially created via the cloud formation template. Click the Enable Button.
<img src="images/mparticle/enableKinesisStreamLambda.png" height="1280" width="720">
Take some time to look at the code that this Lambda uses to send events to Personalize. You can use this code in your own deployment, however you may need to change the event parameters sent to Amazon Personalize depending on the dataset you set up.
```javascript
/ Copyright Amazon.com, Inc. or its affiliates. All Rights Reserved.
// SPDX-License-Identifier: MIT-0
const AWS = require('aws-sdk');
const SSM = new AWS.SSM();
const mParticle = require('mparticle');
const reportActions = ["purchase", "view_detail", "add_to_cart", "checkout","add_to_wishlist"];
const personalizeEvents = new AWS.PersonalizeEvents();
const personalizeRuntime = new AWS.PersonalizeRuntime();
const axios = require('axios');
exports.handler = async function (event, context) {
// Load all of our variables from SSM
try {
let params = {
Names: ['/retaildemostore/services_load_balancers/products',
'/retaildemostore/webui/mparticle_s2s_api_key',
'/retaildemostore/webui/mparticle_s2s_secret_key',
'retaildemostore-personalize-event-tracker-id',
'retaildemostore-product-recommendation-campaign-arn'],
WithDecryption: false
};
let responseFromSSM = await SSM.getParameters(params).promise();
for(const param of responseFromSSM.Parameters) {
if( param.Name === '/retaildemostore/services_load_balancers/products') {
var productsServiceURL = param.Value;
} else if (param.Name === '/retaildemostore/webui/mparticle_s2s_api_key') {
var mpApiKey = param.Value;
} else if (param.Name === '/retaildemostore/webui/mparticle_s2s_secret_key') {
var mpApiSecret = param.Value;
} else if (param.Name === 'retaildemostore-personalize-event-tracker-id') {
var personalizeTrackerID = param.Value;
} else if (param.Name === 'retaildemostore-product-recommendation-campaign-arn') {
var personalizeCampaignARN = param.Value;
}
}
// Init mParticle libraries for the function invocation
var mpApiInstance = new mParticle.EventsApi(new mParticle.Configuration(mpApiKey, mpApiSecret));
} catch (e) {
console.log("Error getting SSM parameter for loadbalancer.");
console.log(e);
throw e;
}
for (const record of event.Records) {
const payloadString = Buffer.from(record.kinesis.data, 'base64').toString('ascii');
const payload = JSON.parse(payloadString);
const events = payload.events;
var amazonPersonalizeUserId;
console.log(`EVENTS: ${JSON.stringify(events)}`);
// First, get the mParticle user ID from the events payload. In this example, mParticle will send all the events
// for a particular user in a batch to this lambda.
// retreive the mParticle user id which is available for anonymous and known customer profiles
var anonymousID = events[0].data.custom_attributes.mpid.toString();
// if the customer profile is known then replace the amazon Personalize User id with the actual
// personalize Id captured from the user's profile
if(payload.user_attributes && payload.user_attributes.amazonPersonalizeId)
amazonPersonalizeUserId = payload.user_attributes.amazonPersonalizeId;
else
amazonPersonalizeUserId = anonymousID;
// Verify in mParticle's payload if there is a customer id set within the customer profile
// this will be used for identity resolution later on within mParticle.
var customerId = null;
if(payload.user_identities){
for (const identityRecord of payload.user_identities)
{
if(identityRecord.identity_type==="customer_id")
customerId = identityRecord.identity;
}
}
var params = {
sessionId: payload.message_id,
userId: amazonPersonalizeUserId,
trackingId: personalizeTrackerID,
eventList: []
};
// Check for variant and assign one if not already assigned
/*var variantAssigned;
var variant;
if(payload.user_attributes && payload.user_attributes.ml_variant) {
variantAssigned = Boolean(payload.user_attributes.ml_variant);
variant = variantAssigned ? payload.user_attributes.ml_variant : Math.random() > 0.5 ? "A" : "B";
}*/
for (const e of events) {
if (e.event_type === "commerce_event" && reportActions.indexOf(e.data.product_action.action) >= 0) {
const timestamp = Math.floor(e.data.timestamp_unixtime_ms / 1000);
const action = e.data.product_action.action;
const event_id = e.data.event_id;
let params = {
sessionId: payload.message_id,
userId: amazonPersonalizeUserId,
trackingId: personalizeTrackerID,
eventList: []
};
// Build the list of events for the user session...
for (const product of e.data.product_action.products) {
const purchasedItem = { itemId: product.id };
params.eventList.push({
properties: purchasedItem,
sentAt: timestamp,
eventId: event_id,
eventType: action
});
}
}
}
console.log(JSON.stringify(params));
// Send the events to Amazon Personalize for training purposes
try {
await personalizeEvents.putEvents(params).promise();
} catch (e) {
console.log(`ERROR - Could not put events - ${e}`);
}
// Get Recommendations from Personalize for the user ID we got up top
let recommendationsParams = {
// Select campaign based on variant
campaignArn: personalizeCampaignARN,
numResults: '5',
userId: amazonPersonalizeUserId
};
try {
var recommendations = await personalizeRuntime.getRecommendations(recommendationsParams).promise();
console.log(`RECOMMENDATIONS - ${JSON.stringify(recommendations)}`);
} catch (e) {
console.log(`ERROR - Could not get recommendations - ${e}`);
}
// Reverse Lookup the product ids to actual product name using the product service url
let itemList = [];
var productNameList = [];
for (let item of recommendations.itemList) {
itemList.push(item.itemId);
var productRequestURL = `${productsServiceURL}/products/id/${item.itemId}`;
var productInfo = await axios.get(productRequestURL);
productNameList.push(productInfo.data.name);
}
//build the mParticle object and send it to mParticle
let batch = new mParticle.Batch(mParticle.Batch.Environment.development);
// if the customer profile is anonymous, we'll use the mParticle ID to tie this recommendation back to the anonymous user
// else we will use the customer Id which was provided earlier
if(customerId == null) {
batch.mpid = anonymousID;
} else {
batch.user_identities = new mParticle.UserIdentities();
batch.user_identities.customerid = customerId; // identify the user via the customer id
}
batch.user_attributes = {};
batch.user_attributes.product_recs = itemList;
batch.user_attributes.product_recs_name=productNameList;
// Create an Event Object using an event type of Other
let event = new mParticle.AppEvent(mParticle.AppEvent.CustomEventType.other, 'AWS Product Personalization Recs Update');
event.custom_attributes = {product_recs: itemList.join()};
batch.addEvent(event);
var body = [batch]; // {[Batch]} Up to 100 Batch objects
console.log(event);
console.log(batch);
let mp_callback = function(error, data, response) {
if (error) {
console.error(error);
} else {
console.log('API called successfully.');
}
};
// Send to Event to mParticle
mpApiInstance.bulkUploadEvents(body, mp_callback);
}
};
```
## Validate that Real-Time Events are Flowing to AWS Kinesis
To validate if events being captured from mParticle are being sent to Kinesis, you would need to go back to the mParticle UI/Platform and Click Data Master then Livestream.
Under Message Direction, select Both In and Out
<img src="images/mparticle/mparticle-verification-step-kinesis.png" height="1280" width="720">
Go back to the Retail Demo Store Web App, and do a eCommerce event. This can be done by viewing a product, click add to Cart, Checkout or Purchase.
You should see the following entries within Livestream which will contain Amazon Kinesis with an outward arrow.
<img src="images/mparticle/mparticle-verification-step-kinesis2.png" height="1280" width="720">
If you haven't seen any outbound events generated to Amazon Kinesis, you might need to wait for a while before the settings are applied properly.
## Save AWS Personalize Recommended Products back to mParticle
Aside from just sending events from the AWS Retail Demo Store to mParticle, the Lambda function above also sends the commerce events to AWS Personalize. This allows AWS Personalize to receive specific commerce events made by a anonymous and known user and from there allow AWS Personalize to do its magic by providing product recommendation information back to mParticle. Once AWS Personalize has finished computing the relevant products that is associated to the recent events the customer has made, the said product recommendation information will be sent back to mParticle using the mParticle NodeJS SDK. The said code snippet below will set the product_recommendation information as a user attribute (product_recs) within the user's profile.
```javascript
// if Events are more than 10 splice the events
if(params.eventList.length > 10)
{
var lastTenRecords = params.eventList.length / 2;
params.eventList = params.eventList.slice(lastTenRecords);
}
if (params.eventList.length > 0) {
// Reverse Lookup the product ids to actual product name using the product service url
let itemList = [];
var productNameList = [];
for (let item of recommendations.itemList) {
itemList.push(item.itemId);
var productRequestURL = '{productsServiceURL}/products/id/${item.itemId}';
var productInfo = await axios.get(productRequestURL);
productNameList.push(productInfo.data.name);
}
//build the mParticle object and send it to mParticle
let batch = new mParticle.Batch(mParticle.Batch.Environment.development);
// if the customer profile is anonymous, we'll use the mParticle ID to tie this recommendation back to the anonymous user
// else we will use the customer Id which was provided earlier
if(customerId == null){
batch.mpid = anonymousID;
}
else{
batch.user_identities = new mParticle.UserIdentities();
batch.user_identities.customerid = customerId; // identify the user via the customer id
}
batch.user_attributes = {};
batch.user_attributes.product_recs = itemList;
batch.user_attributes.product_recs_name=productNameList;
// Create an Event Object using an event type of Other
let event = new mParticle.AppEvent(mParticle.AppEvent.CustomEventType.other, 'AWS Product Personalization Recs Update');
event.custom_attributes = {product_recs: itemList.join()};
batch.addEvent(event);
var body = [batch]; // {[Batch]} Up to 100 Batch objects
console.log(event);
console.log(batch);
let mp_callback = async function(error, data, response) {
if (error) {
console.error(error);
} else {
console.log('API called successfully.');
}
};
// Send to Event to mParticle
await mpApiInstance.bulkUploadEvents(body, mp_callback);
}
};
```
<img src="images/mparticle/mparticle-product_recs1.png" height="1280" width="720">
<img src="images/mparticle/mparticle-product_recs2.png" height="1280" width="720">
```
# When you are done with this module, you can delete the user and policies you created earlier
# detach the policy from the user we created
result = iam.detach_user_policy(
PolicyArn=policy_arn,
UserName=user_name )
result = iam.delete_access_key(
AccessKeyId = access_key_id,
UserName = user_name )
# delete the kinesis policy
policy = iam.delete_policy(
PolicyArn=policy_arn )
# delete the user we created
created_user = iam.delete_user(
UserName=user_name
)
```
| github_jupyter |
# Part 3: Reverse Mode Automatic Differentiation with PyTorch
```
# Execute this code block to install dependencies when running on colab
try:
import torch
except:
from os.path import exists
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
cuda_output = !ldconfig -p|grep cudart.so|sed -e 's/.*\.\([0-9]*\)\.\([0-9]*\)$/cu\1\2/'
accelerator = cuda_output[0] if exists('/dev/nvidia0') else 'cpu'
!pip install -q http://download.pytorch.org/whl/{accelerator}/torch-1.0.0-{platform}-linux_x86_64.whl torchvision
```
PyTorch implements Dynamic Reverse Mode Automatic Differentiation, much like we did in the previous exercise. There is one really major difference in what PyTorch provides over our simple example: it works directly with matrices (`Tensor`s) rather than with scalars (although obviously a matrix can represent a scalar).
In this tutorial, we'll explore PyTorch's AD implementation. Note that we're using the API of PyTorch 0.4 or later which simplifies use of AD (previous versions required wrapping all `Tensor`s that you wanted to compute gradients of in `Variable` objects; PyTorch 0.4 removes the need to do this and allows `Tensor`s themselves to track gradients).
We'll start with the simple example we tried earlier in the code block below:
__Task:__ Run the following code and verify the solution is correct
```
import torch
# set up the problem
x = torch.tensor(0.5, requires_grad=True)
y = torch.tensor(4.2, requires_grad=True)
z = x * y + torch.sin(x)
print("z = " + str(z.item()))
z.backward() # this goes through the computation graph and accumulates the gradients in the cached .grad attributes
print("dz/dx = " + str(x.grad.item()))
print("dz/dy = " + str(y.grad.item()))
```
As with our own AD implementation, PyTorch lets us differentiate through an algorithm.
__Task__: Use the block below to compute the gradient $\partial z/\partial x$ of the following pseudocode algorithm and store the result in the `dzdx` variable:
x = 0.5
z = 1
i = 0
while i<2:
z = (z + i) * x * x
i = i + 1
```
dzdx = None
# YOUR CODE HERE
# raise NotImplementedError()
z=1
for i in range(0,2):
x = torch.tensor(0.5, requires_grad=True)
z=(z+i)*x*x
z.backward(retain_graph=True)
dzdx=x.grad.item()
print(dzdx)
dzdx = None
# YOUR CODE HERE
# raise NotImplementedError()
z = torch.tensor(1.0)
for i in range(0,2):
x = torch.tensor(0.5, requires_grad=True)
z=(z+i)*x*x
z.backward()
dzdx=x.grad.item()
print(dzdx)
import math
print((math.pow(0.5,2)+1)*2*0.5)
assert dzdx == ((math.pow(0.5,2)+1)*2*0.5)
```
## PyTorch limitations: in-place operations and aliasing
PyTorch will throw an error at runtime if you try to differentiate through an in-place operation on a tensor.
__Task__: Run the following code to see this in action.
```
x = torch.tensor(1.0, requires_grad=True)
y = x.tanh()
y.add_(3) # inplace addition
y.backward()
```
Aliasing is also something that can't be differentiated through and will result in a slightly more cryptic error.
__Task__: Run the following code to see this in action. If you don't understand what this code does add some `print` statements to show the values of `x` and `y` at various points.
```
x = torch.tensor([1, 2, 3, 4], requires_grad=True, dtype=torch.float)
print(x)
y = x[:1]
print(y)
y.add_(3)
print(y)
y.backward()
```
## Dealing with multiple outputs
PyTorch can deal with the case where there are multiple output variables if we can formulate the expression in terms of tensor operations. Consider the example from the presentation for example:
$$\begin{cases}
z = 2x + \sin x\\
v = 4x + \cos x
\end{cases}$$
We could formulate this as:
$$
\begin{bmatrix}z \\ v\end{bmatrix} = \begin{bmatrix}2 \\ 4\end{bmatrix} \odot \bar{x} + \begin{bmatrix}1 \\ 0\end{bmatrix} \odot \sin\bar x + \begin{bmatrix}0 \\ 1\end{bmatrix} \odot \cos\bar x
$$
where
$$
\bar x = \begin{bmatrix}x \\ x\end{bmatrix}
$$
and $\odot$ represents the Hadamard or element-wise product. This is demonstrated using PyTorch in the following code block.
__Task:__ run the code below.
```
x = torch.tensor([[1.0],[1.0]], requires_grad=True)
zv = ( torch.tensor([[2.0],[4.0]]) * x +
torch.tensor([[1.0], [0.0]]) * torch.sin(x) +
torch.tensor([[0.0], [1.0]]) * torch.cos(x) )
zv.backward(torch.tensor([[1.0],[1.0]])) # Note as we have "multiple outputs" we must pass in a tensor of weights of the correct size
print(x.grad)
```
__Task:__ Use the following box to write down the derivative of the expression for $\begin{bmatrix}z \\ v\end{bmatrix}$ and verify the gradients $\partial z/\partial x$ and $\partial v/\partial x$ are correct for $x=1$.
```
import math
print(2+math.cos(1.0),4-math.sin(1.0))
```
**Answer: The derivatives of the expression for $\begin{bmatrix}z \\ v\end{bmatrix}$ is
$\begin{cases}
\partial z/\partial x = 2 + \cos x\\
\partial v/\partial x = 4 - \sin x
\end{cases}$, and the gradients $\partial z/\partial x$ and $\partial v/\partial x$ for $x=1$ is $\begin{bmatrix}2.5403 \\ 3.1585\end{bmatrix}$**
## Gradient descent & gradients with respect to a vector
Let's put everything together and using automatically computed gradients to find the minima of a function by taking steps down the gradient from an initial position. Rather than explicitly creating each input variable as a scalar as in the previous examples, we'll use a vector instead (so our gradients will be with respect to each element of the vector).
__Task:__ work through the following example to see how taking gradients with respect to a vector works & how simple gradient descent optimisation can be implemented.
```
# This is our starting vector
initial=[[2.0], [1.0], [10.0]]
# This is the function we will optimise (feel free to work out the analytic minima!)
def function(x):
return x[0]**2 + x[1]**2 + x[2]**2
x = torch.tensor(initial, requires_grad=True, dtype=torch.float)
for i in range(0,100):
# manually dispose of the gradient (in reality it would be better to detach and zero it to reuse memory)
x.grad=None
# evaluate the function
J = function(x)
# auto-compute the gradients at the previously evaluated point x
J.backward()
# compute the update
z = x - x.grad*0.1
x.data = z
if i%10 == 0:
print((x, function(x).item()))
```
__Task__: Answer the following question in the box below: Why must the update in the code above be assigned to a different variable (the `z`) before being assigned to the `data` value of `x`?
- **Answer:**
**Because the function falue changes after the gradients descent, we need to get the changing gradient for this function when the function value changes. Hence, we update the data value of x**
## Differentiating through random operations
We'll end with an example that will be important later in the course: differentiating with respect to the parameters of a random number generator.
Assume that as some part of a differentiable program that we write we wish to incorporate a random element where we sample values, $z$ from a Normal distribution: $z \sim \mathcal{N}(\mu,\sigma^2)$. We want to learn the parameters of the generator $\mu$ and $\sigma^2$, but how can we do this? In a differentiable program setting we want to differentiate with respect to these parameters, but at first glance it isn't at all obvious what this means as the generator _just_ produces numbers which themselves don't have gradients.
The answer is often called the _reparameterisation trick_: If we note that sampling a Normal distribution is equivalent to drawing numbers from a Uniform distribution and scaling and shifting them: $z \sim \mathcal{N}(\mu,\sigma^2) \equiv z \sim \mu + \sigma\mathcal{U}(0,1)\equiv z = \mu + \sigma \zeta\, \rm{where}\, \zeta\sim\mathcal{U}(0,1)$. With this reparameterisation the gradients with respect to the parameters are obvious.
The following code block demonstrates this in practice; each of the gradients can be interpreted as how much an infintesimal change in $\mu$ or $\sigma$ would change $z$ if we could repeat the sampling operation again with the same value of `torch.rand(1)` being produced:
```
mu = torch.tensor(1.0, requires_grad=True)
sigma = torch.tensor(1.0, requires_grad=True)
for i in range(10):
mu.grad = None
sigma.grad = None
z = mu + sigma * torch.randn(1)
z.backward()
print("z:", z.item(), "\tdzdmu:", mu.grad.item(), "\tdzdsigma:", sigma.grad.item())
```
| github_jupyter |
```
import sys
sys.path.append("../../src")
sys.path.append("../")
import os
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
import datetime
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import pickle
import pdb
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
%matplotlib inline
from analyze import get_names, read_results, delete_results
from training import load_model, TrainModel
from os import listdir
import pandas as pd
# from IPython.display import display
```
### Print available cases
```
directory = listdir('../data/')
case_list = ['lock']
for name in directory:
if '.' not in name and 'lorenz' in name:
casename = '_'.join(name.split('_')[2:])
if casename not in case_list:
case_list.append(casename)
for case in case_list: print(case)
cases = ['initv_intloss']
path = '../data/'
my_params = ['loss_weight_integral', 'sindy_pert', 'svd_dim', 'model']
primary_params = ['case', 'coefficient_initialization', 'exact_features', 'fix_coefs', 'input_dim', 'latent_dim',
'loss_weight_integral', 'loss_weight_rec', 'loss_weight_sindy_regularization', 'loss_weight_sindy_x',
'loss_weight_sindy_z', 'loss_weight_x0', 'model', 'n_ics', 'widths_ratios', 'svd_dim']
secondary_params = ['activation', 'actual_coefficients', 'coefficient_threshold', 'dt', 'fixed_coefficient_mask', 'library_dim',
'max_epochs', 'model_order', 'noise', 'option', 'patience', 'poly_order', 'print_frequency',
'save_checkpoints', 'save_freq', 'scale', 'sindy_pert']
tertiary_params = ['batch_size', 'data_path', 'include_sine', 'learning_rate', 'learning_rate_sched', 'print_progress']
```
### Get names for a given case
```
name_list = get_names(cases, path)
for idx, name in enumerate(name_list): print(idx, name)
end_time = 100
end_time_plot = 100
display_params = my_params #primary_params + secondary_params + tertiary_params
t0_frac = 0.2
query_remove = False
non_existing_files, remove_files = read_results(name_list[:],
path,
end_time=end_time,
display_params=display_params,
t0_frac=t0_frac,
end_time_plot=end_time_plot,
query_remove=query_remove)
print(non_existing_files)
print(remove_files)
# delete_results(non_existing_files+remove_files, '../data/')
```
| github_jupyter |
# Getting Started with pandas
```
import pandas as pd
from pandas import Series, DataFrame
import numpy as np
np.random.seed(12345)
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
PREVIOUS_MAX_ROWS = pd.options.display.max_rows
pd.options.display.max_rows = 20
np.set_printoptions(precision=4, suppress=True)
```
## Introduction to pandas Data Structures
### Series
```
obj = pd.Series([4, 7, -5, 3])
obj
obj.values
obj.index # like range(4)
obj2 = pd.Series([4, 7, -5, 3], index=['d', 'b', 'a', 'c'])
obj2
# obj2.index
obj2['a']
obj2['d'] = 6
obj2.loc[['c', 'a', 'd']]
obj2[obj2 > 0]
obj2 * 2
np.exp(obj2)
'b' in obj2
'e' in obj2
sdata = {'Ohio': 35000, 'Texas': 71000, 'Oregon': 16000, 'Utah': 5000}
obj3 = pd.Series(sdata)
obj3
states = ['California', 'Ohio', 'Oregon', 'Texas']
obj4 = pd.Series(sdata, index=states)
obj4
pd.isnull(obj4)
pd.notnull(obj4)
obj4.isnull()
obj3
obj4
obj3 + obj4
obj4.name = 'population'
obj4.index.name = 'state'
obj4
obj
obj.index = ['Bob', 'Steve', 'Jeff', 'Ryan']
obj
```
### DataFrame
```
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
frame = pd.DataFrame(data)
frame
frame.head()
pd.DataFrame(data, columns=['year', 'state', 'pop'])
frame2 = pd.DataFrame(data, columns=['year', 'state', 'pop', 'debt'],
index=['one', 'two', 'three', 'four',
'five', 'six'])
frame2
# frame2.columns
frame2['state']
frame2.year
frame2.loc['three']
frame2['debt'] = 16.5
frame2
frame2['debt'] = np.arange(6.)
frame2
val = pd.Series([-1.2, -1.5, -1.7], index=['two', 'four', 'five'])
frame2['debt'] = val
frame2
frame2['eastern'] = frame2.state == 'Ohio'
frame2
del frame2['eastern']
frame2.columns
pop = {'Nevada': {2001: 2.4, 2002: 2.9},
'Ohio': {2000: 1.5, 2001: 1.7, 2002: 3.6}}
frame3 = pd.DataFrame(pop)
frame3
frame3.T
pd.DataFrame(pop, index=[2001, 2002, 2003])
pdata = {'Ohio': frame3['Ohio'][:-1],
'Nevada': frame3['Nevada'][:2]}
pd.DataFrame(pdata)
frame3.index.name = 'year'; frame3.columns.name = 'state'
frame3
frame3.values
frame2.values
```
### Index Objects
```
obj = pd.Series(range(3), index=['a', 'b', 'c'])
index = obj.index
index
index[1:]
```
index[1] = 'd' # TypeError
```
labels = pd.Index(np.arange(3))
labels
obj2 = pd.Series([1.5, -2.5, 0], index=labels)
obj2
obj2.index is labels
frame3
frame3.columns
'Ohio' in frame3.columns
2003 in frame3.index
dup_labels = pd.Index(['foo', 'foo', 'bar', 'bar'])
dup_labels
```
## Essential Functionality
### Reindexing
```
obj = pd.Series([4.5, 7.2, -5.3, 3.6], index=['d', 'b', 'a', 'c'])
obj
obj2 = obj.reindex(['a', 'b', 'c', 'd', 'e'])
obj2
obj3 = pd.Series(['blue', 'purple', 'yellow'], index=[0, 2, 4])
obj3
obj3.reindex(range(6), method='ffill')
frame = pd.DataFrame(np.arange(9).reshape((3, 3)),
index=['a', 'c', 'd'],
columns=['Ohio', 'Texas', 'California'])
frame
frame2 = frame.reindex(['a', 'b', 'c', 'd'])
frame2
states = ['Texas', 'Utah', 'California']
frame.reindex(columns=states)
frame.loc[['a', 'b', 'c', 'd'], states]
```
### Dropping Entries from an Axis
```
obj = pd.Series(np.arange(5.), index=['a', 'b', 'c', 'd', 'e'])
obj
new_obj = obj.drop('c')
new_obj
obj.drop(['d', 'c'])
data = pd.DataFrame(np.arange(16).reshape((4, 4)),
index=['Ohio', 'Colorado', 'Utah', 'New York'],
columns=['one', 'two', 'three', 'four'])
data
data.drop(['Colorado', 'Ohio'])
data.drop('two', axis=1)
data.drop(['two', 'four'], axis='columns')
obj.drop('c', inplace=True)
obj
```
### Indexing, Selection, and Filtering
```
obj = pd.Series(np.arange(4.), index=['a', 'b', 'c', 'd'])
obj
obj['b']
obj[1]
obj[2:4]
obj[['b', 'a', 'd']]
obj[[1, 3]]
obj[obj < 2]
obj['b':'c']
obj['b':'c'] = 5
obj
data = pd.DataFrame(np.arange(16).reshape((4, 4)),
index=['Ohio', 'Colorado', 'Utah', 'New York'],
columns=['one', 'two', 'three', 'four'])
data
data['two']
data[['three', 'one']]
data[:2]
data[data['three'] > 5]
data < 5
data[data < 5] = 0
data
```
#### Selection with loc and iloc
```
data.loc['Colorado', ['two', 'three']]
data.iloc[2, [3, 0, 1]]
data.iloc[2]
data.iloc[[1, 2], [3, 0, 1]]
data.loc[:'Utah', 'two']
data.iloc[:, :3][data.three > 5]
```
### Integer Indexes
ser = pd.Series(np.arange(3.))
ser
ser[-1]
```
ser = pd.Series(np.arange(3.))
ser
ser2 = pd.Series(np.arange(3.), index=['a', 'b', 'c'])
ser2[-1]
ser[:1]
ser.loc[:1]
ser.iloc[:1]
```
### Arithmetic and Data Alignment
```
s1 = pd.Series([7.3, -2.5, 3.4, 1.5], index=['a', 'c', 'd', 'e'])
s2 = pd.Series([-2.1, 3.6, -1.5, 4, 3.1],
index=['a', 'c', 'e', 'f', 'g'])
s1
s2
s1 + s2
df1 = pd.DataFrame(np.arange(9.).reshape((3, 3)), columns=list('bcd'),
index=['Ohio', 'Texas', 'Colorado'])
df2 = pd.DataFrame(np.arange(12.).reshape((4, 3)), columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
df1
df2
df1 + df2
df1 = pd.DataFrame({'A': [1, 2]})
df2 = pd.DataFrame({'B': [3, 4]})
df1
df2
df1 - df2
```
#### Arithmetic methods with fill values
```
df1 = pd.DataFrame(np.arange(12.).reshape((3, 4)),
columns=list('abcd'))
df2 = pd.DataFrame(np.arange(20.).reshape((4, 5)),
columns=list('abcde'))
df2.loc[1, 'b'] = np.nan
df1
df2
df1 + df2
df1.add(df2, fill_value=0)
1 / df1
df1.rdiv(1)
df1.reindex(columns=df2.columns, fill_value=0)
```
#### Operations between DataFrame and Series
```
arr = np.arange(12.).reshape((3, 4))
arr
arr[0]
arr - arr[0]
frame = pd.DataFrame(np.arange(12.).reshape((4, 3)),
columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
series = frame.iloc[0]
frame
series
frame - series
series2 = pd.Series(range(3), index=['b', 'e', 'f'])
frame + series2
series3 = frame['d']
frame
series3
frame.sub(series3, axis='index')
```
### Function Application and Mapping
```
frame = pd.DataFrame(np.random.randn(4, 3), columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
frame
np.abs(frame)
f = lambda x: x.max() - x.min()
frame.apply(f)
frame.apply(f, axis='columns')
def f(x):
return pd.Series([x.min(), x.max()], index=['min', 'max'])
frame.apply(f)
format = lambda x: '%.2f' % x
frame.applymap(format)
frame['e'].map(format)
```
### Sorting and Ranking
```
obj = pd.Series(range(4), index=['d', 'a', 'b', 'c'])
obj.sort_index()
frame = pd.DataFrame(np.arange(8).reshape((2, 4)),
index=['three', 'one'],
columns=['d', 'a', 'b', 'c'])
frame.sort_index()
frame.sort_index(axis=1)
frame.sort_index(axis=1, ascending=False)
obj = pd.Series([4, 7, -3, 2])
obj.sort_values()
obj = pd.Series([4, np.nan, 7, np.nan, -3, 2])
obj.sort_values()
frame = pd.DataFrame({'b': [4, 7, -3, 2], 'a': [0, 1, 0, 1]})
frame
frame.sort_values(by='b')
frame.sort_values(by=['a', 'b'])
obj = pd.Series([7, -5, 7, 4, 2, 0, 4])
obj.rank()
obj.rank(method='first')
# Assign tie values the maximum rank in the group
obj.rank(ascending=False, method='max')
frame = pd.DataFrame({'b': [4.3, 7, -3, 2], 'a': [0, 1, 0, 1],
'c': [-2, 5, 8, -2.5]})
frame
frame.rank(axis='columns')
```
### Axis Indexes with Duplicate Labels
```
obj = pd.Series(range(5), index=['a', 'a', 'b', 'b', 'c'])
obj
obj.index.is_unique
obj['a']
obj['c']
df = pd.DataFrame(np.random.randn(4, 3), index=['a', 'a', 'b', 'b'])
df
df.loc['b']
```
## Summarizing and Computing Descriptive Statistics
```
df = pd.DataFrame([[1.4, np.nan], [7.1, -4.5],
[np.nan, np.nan], [0.75, -1.3]],
index=['a', 'b', 'c', 'd'],
columns=['one', 'two'])
df
df.sum()
df.sum(axis='columns')
df.mean(axis='columns', skipna=False)
df.idxmax()
df.cumsum()
df.describe()
obj = pd.Series(['a', 'a', 'b', 'c'] * 4)
obj.describe()
```
### Correlation and Covariance
conda install pandas-datareader
```
price = pd.read_pickle('examples/yahoo_price.pkl')
volume = pd.read_pickle('examples/yahoo_volume.pkl')
```
import pandas_datareader.data as web
all_data = {ticker: web.get_data_yahoo(ticker)
for ticker in ['AAPL', 'IBM', 'MSFT', 'GOOG']}
price = pd.DataFrame({ticker: data['Adj Close']
for ticker, data in all_data.items()})
volume = pd.DataFrame({ticker: data['Volume']
for ticker, data in all_data.items()})
```
returns = price.pct_change()
returns.tail()
returns['MSFT'].corr(returns['IBM'])
returns['MSFT'].cov(returns['IBM'])
returns.MSFT.corr(returns.IBM)
returns.corr()
returns.cov()
returns.corrwith(returns.IBM)
returns.corrwith(volume)
```
### Unique Values, Value Counts, and Membership
```
obj = pd.Series(['c', 'a', 'd', 'a', 'a', 'b', 'b', 'c', 'c'])
uniques = obj.unique()
uniques
obj.value_counts()
pd.value_counts(obj.values, sort=False)
obj
mask = obj.isin(['b', 'c'])
mask
obj[mask]
to_match = pd.Series(['c', 'a', 'b', 'b', 'c', 'a'])
unique_vals = pd.Series(['c', 'b', 'a'])
pd.Index(unique_vals).get_indexer(to_match)
data = pd.DataFrame({'Qu1': [1, 3, 4, 3, 4],
'Qu2': [2, 3, 1, 2, 3],
'Qu3': [1, 5, 2, 4, 4]})
data
result = data.apply(pd.value_counts).fillna(0)
result
```
## Conclusion
```
pd.options.display.max_rows = PREVIOUS_MAX_ROWS
```
| github_jupyter |
## Programming Exercise 7: K-means Clustering and Principal Component Analysis
#### Author - Rishabh Jain
```
import warnings,os
warnings.simplefilter('ignore')
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
%matplotlib inline
from PIL import Image,ImageFilter
from scipy.io import loadmat
```
### 1 K-means Clustering
#### Problem Statement
In this part of the exercise, we will implement the K-means algorithm and use it for image compression by reducing the number of colors that occur in an image to only those that are most common in that image.
#### 1.1 Implementing K-means
The K-means algorithm is a method to autmatically cluster similar data together. We have a training set $\{x^{(1)}...x^{(m)}\}$ where $x^{(i)}\epsilon R^{n}$. The intution behind K-means is an iterative procedure that starts by guessing the initial centroids, and then refines the guess by repeatedly assigning examples to their closest centroids and then recomputing the centroids based on assignments.
```
# Initialize Centroids
centroids=kMeansInitCentroids(X,K)
for i in range(iter):
# Cluster assignment step: Assign each data point to the closest centroid. idx(i) corrsponds to c(i),
# the index of the centroid assigned to example i
idx=findClosestCentroids(X,centroids)
# Move centroid step: Compute means based on centroid assignments
centroids=computeMeans(X,idx,K);
```
The inner loop of the algoritm carries two steps:
1. Assigning each training example $x^{(i)}$ to its closest centroid.
2. Recomputing the mean of each centroid using the points assigned to it.
The K-means algorithm will always converge to some final set of means for the centroids.
##### 1.1.1 Find closest centroids
In the "Cluster assignment" phase of the K-means algorithm, **the algorithm assigns every training example $x^{(i)}$ to its closest centroid, given the current position of centroid.**
$$c^{(i)}:=j\text{ that minimizes }||x^{(i)}-\mu||^{2}$$
where $c^{(i)}$ is the index of the centroid that is closes to $x^{(i)}$, $\mu^{(j)}$ is the position of $j^{th}$ centroid.
```
def findClosestCentroids(X,centroids):
'''Computes and returns the closest centroid label for the given samples'''
m,n=X.shape
# Number of clusters(K)
K=centroids.shape[0]
c=np.zeros(m)
for i in range(m):
x=X[i,:]
norm=np.zeros(K)
for k in range(K):
norm[k]=np.dot((x-centroids[k,:]),(x-centroids[k,:]).T)
c[i]=np.argmin(norm)
return c
```
##### 1.1.2 Computing centroid means
Here, we will calculate the centroid new location based on the examples assigned to that centroid. The cordinates of that centroid is mean of the cordinates of all the samples assigned to it.
$$\mu^{(j)}=\frac{\sum_{i\epsilon c_k}x^{(i)}}{|C_k|}$$
where $C_k$ is the set of examples assigned to that centroid.
```
def computeCentroids(X,c,K):
'''Computes and returns the new centroids by computing the means of samples assigned to that centroid'''
m,n=X.shape
centroids=np.zeros(shape=(K,n))
for k in range(K):
x=X[c==k,:]
centroids[k,:]=x.mean(axis=0)
return centroids
```
#### 1.2 K-means on example dataset
#### 1.3 Random Initialization
A good strategy for initializing the centroids is to select a random sample from the training set. Example used above first select the random K centroids, finds and compute the closest centroid for each sample in training set.
```
def runKMeans(X,K,maxIters,plotProgress=False):
m,n=X.shape
# Randomly selecting K centroids from X
indices=np.random.randint(0,m,K)
centroids=X[indices,:]
history={}
for i in range(1,maxIters+1):
print(f'ITERATION : {i}',end='\r')
c=findClosestCentroids(X,centroids)
history[i]=centroids
centroids=computeCentroids(X,c,K)
if plotProgress==True and X.shape[1]==2:
fig,ax=plt.subplots(figsize=(10,8))
# Plotting points belonging to same cluster
for k in range(K):
x=X[c==k,:]
sns.scatterplot(x[:,0],x[:,1],ax=ax,legend=False)
# Plotting centroids history
for i in range(1,maxIters):
for k in range(K):
temp=np.array([history[i][k,:],history[i+1][k,:]])
sns.lineplot(temp[:,0],temp[:,1],color='black')
sns.scatterplot(temp[:,0],temp[:,1],color='black',marker='x',s=50)
if i+1==maxIters:
sns.scatterplot([temp[1,0]],[temp[1,1]],color='red',marker='x',s=100)
return c,centroids
mat=loadmat('ex7data2.mat')
X=mat['X']
sns.scatterplot(X[:,0],X[:,1]);
K=3
maxIter=10
labels,centroids=runKMeans(X,K,maxIter,plotProgress=True)
```
**Note : Red Cross markers are the final centroids of that cluster.**
#### 1.4 Image compression with K-means
In this example, we will apply K-means for image compression. In a straightforward 24-bit color representation of an image, each pixel is represented as 8-bit unsigned integers (ranging from 0 to 255) that specify the red, green and blue intensity values. This encoding is often referred to as RGB encoding. **Our image contains thousands of colors, and we will reduce the number of colors to 16 colors.**
Concretely, we will treat every pixel in the original image as a part of our dataset and use the K-means algorithm to find the 16 colors that best group (cluster) the pixels in 3-dimensional space (RGB).
##### 1.4.1 K-means on pixels
On loading the image, we get a 3-D matrix whose first two indices identify a pixel position and whose last index represent a red,green and blue value. Here will reshape the 3-D matrix into mX3 matrix of pixel colors and call our K-means function on it.
**EXAMPLE 1:**
```
K=16
maxIter=5
img=Image.open('images/bird.png')
mat=np.asarray(img)
print(f'IMAGE DIMENSION : {mat.shape}')
X=mat.reshape((-1,3))
print(f'X SHAPE : {X.shape}')
labels,centroids=runKMeans(X,K,maxIter)
print(f'\n\nTOP {K} COLORS :\n\n',centroids)
```
After finding the top $K=16$ colors to represent the image, we will now assign each pixel position to its closest centroids. This allows us to represent the original image using centroid assignment of each pixel. We have significantly reduced the number of bits that are required to describe the image. The original image required 24 bits for each of 128 x 128 pixels, resulting in total of 128 x 128 x 4 = 363,216 bits. The new representation requires some overhead storage in form of a dictionary of 16 colors, each of which require 24 bits, but the image itself then only requires 4 bits per pixel location. The final number of bits used is therefore 16 x 24 + 128 x 128 X 4 = 65,920 bits, which corresponds to compressing the original image by about a factor of 6.
```
xCompressed=centroids[labels.astype(int),:]
xCompressed=np.uint8(xCompressed.reshape(mat.shape))
fig,ax=plt.subplots(1,2)
ax[0].imshow(img)
ax[0].set_title('Original')
ax[0].axis('off')
ax[1].imshow(Image.fromarray(xCompressed))
ax[1].set_title('Compressed')
ax[1].axis('off');
```
**EXAMPLE 2:**
```
K=6
maxIter=10
img=Image.open('images/parrot.png')
mat=np.asarray(img)
print(f'IMAGE DIMENSION : {mat.shape}')
X=mat.reshape((-1,3))
print(f'X SHAPE : {X.shape}')
labels,centroids=runKMeans(X,K,maxIter)
print(f'\n\nTOP {K} COLORS :\n\n',centroids)
xCompressed=centroids[labels.astype(int),:]
xCompressed=np.uint8(xCompressed.reshape(mat.shape))
fig,ax=plt.subplots(1,2)
ax[0].imshow(img)
ax[0].set_title('Original')
ax[0].axis('off')
ax[1].imshow(Image.fromarray(xCompressed))
ax[1].set_title('Compressed')
ax[1].axis('off');
```
### 2 Principal Component Analysis
<img src="images/PCA1.png">
As we can see in figure (A), three axes: L(Length), W(Width) and H(Height) are used to represent the ellipse in 3-D world. So each data point on ellipse can be written as a function of three variables :
$$Data(i)=f(L(i),W(i),H(i))$$
But, this is not the best way to represent ellipse. We can do the following improvements:
1. Find the geometric mean of the ellipse, and set it as the coordinate origin.
2. Find the direction along which the ellipse has the longest radius (large variance) and call it as 'Principal Component 1'.
3. Find the another direction perpendicular to the first one and along which the ellipse has the second longest radius and call it 'Principal Component 2'
4. Re-plot the ellipse under the new coordinate system defined by principal component C1 and C2.
In the new coordinate system, data points on ellipse can re-written as a function of two variables:
$$Data(j)=g(C1(j),C2(j))$$
After the coordinate system transformation, we get:
- Fewer variables (lower dimension of variables) in function g as compared to function f.
- No information lost.
>- f$\approx$g
>- The relative geometric positions of all data points remains unchanged.
**That's exactly what PCA do. The term 'Principal Component' denotes new variables we choose to describe our data in lower dimension. All PCA must satisfy two conditions :**
1. They must be perpendicular (or mathematically orthogonal) to each other.
>- This means that the principal components are NOT linearly correlated with each other.
>- And that's why PCA can reduce the number of variables without losing much information because the variables in raw data are not independent and correlated variables cause redundancy.
2. These principal components are ordered by the variance in the data points along them. So our data must have the largest variance along the axes of compoenent 1.
>- This means the higher order a component have, the more important it is.
>- **Sometimes we sacrifice minor components to further reduce the number of variables. For example: If the first two components contributed 90% of the variance in the data, we might want to focus on them and discard the rest of the components.**
**Useful Resources**
- [Linear Transformation](https://www.youtube.com/watch?v=kYB8IZa5AuE&list=PL0-GT3co4r2y2YErbmuJw2L5tW4Ew2O5B&index=4)
- [Matrix multiplication as composition](https://www.youtube.com/watch?v=XkY2DOUCWMU&list=PL0-GT3co4r2y2YErbmuJw2L5tW4Ew2O5B&index=5)
- [Eigenvectors and eigenvalues](https://www.youtube.com/watch?v=PFDu9oVAE-g&list=PL0-GT3co4r2y2YErbmuJw2L5tW4Ew2O5B&index=14)
#### 2.1 Example Dataset
We will start with a 2D dataset which has one direction of large variation and one of smaller variation.
```
mat=loadmat('./ex7data1.mat')
X=mat['X']
fig=sns.scatterplot(X[:,0],X[:,1]);
fig.set(title='Original 2D Dataset');
```
#### 2.2 Implementing PCA
##### PCA can be implemented in two ways.
1. **EVD (Eigen Value Decomposition)**
>1. First, the values in Data matrix (X of m x n shape) are centered and scaled by subtracting the mean of each features and dividing the resultant by the standard deviation of each feature.
$$X=\frac{X-\bar{X}}{\sigma(X)}$$
>2. Covariance matrix (C of n x n shape) is calculated for the normalized matrix.
```
C=np.cov(X.T)
```
>3. Finally, we calculate the eigen decomposition of the covariance matrix (C). This results in a list of eigenvalues and eigenvectors. The eigenvectors represent the directions or principal components for the reduced subspace of the data matrix whereas the eigenvalues represents the magnitude for the directions.
```
eigenValues,eigenVectors=np.linalg.eig(C)
```
>4. The eigenvectors can be sorted by the eigenvalues in descending order to provide a ranking of the components or axes of the new subspace for data matrix (X).
```
idx=eigenValues.argsort()[::-1]
eigenValues=eigenValues[idx]
eigenVectors=eigenVectors[:idx]
```
>5. A total of m or less components must be selected to comprise the chosen subspace. Ideally, we would select k eigenvectors, called principal components (m x k shape), that have the k largest eigenvalues.
```
principalComponents=eigenVectors[:,:k]
```
>6. Once chosen, data matrix (of shape m X n) can be projected into the subspace (of shape m x k) via matrix multiplication.
```
Z=X.dot(principalComponents)
```
2. **SVD (Singular Value Decomposition)**
>1. First, the values in Data matrix (X of m x n shape) are centered and scaled by subtracting the mean of each features and dividing the resultant by the standard deviation of each feature.
$$X=\frac{X-\bar{X}}{\sigma(X)}$$
>2. Finally, we calculate the singular value decomposition of the normalized matrix (X of shape m x n). This returns the U, Sigma and $V^T$ elements. This U is an m x m matrix, Sigma is a list of singular values and $V^T$ is the transpose of an n x n matrix.
```
U,S,VT=np.linalg.svd(X)
V=VT.T
```
>3. The singular values are sorted in descending order to provide a ranking of the components or axes of the new subspace for data matrix (X).
```
idx=S.argsort()[::-1]
S=S[idx]
V=V[:idx]
```
>4. A total of m or less components must be selected to comprise the chosen subspace. Ideally, we would select k singular vectors, called principal components (m x k shape), that have the k largest eigenvalues.
```
principalComponents=V[:,:k]
```
>5. Once chosen, data matrix (of shape m x n) can be projected into the subspace (of shape m x k) via matrix multiplication.
```
Z=X.dot(principalComponents)
```
Note:
- Eigen vectors from EVD and Singular vectors from SVD are both same.
- SVD is computationally more efficient as $XX^T$ calculation is not required in SVD whereas it is required in EVD.
```
def featureNormalize(X):
'''Returns the normalized matrix'''
mean=np.mean(X,axis=0)
std=np.std(X,axis=0)
xNormalized=(X-mean)/std
return xNormalized
```
**PCA using EVD**
```
# Normalizing
X=featureNormalize(X)
# Computing covariance
C=np.cov(X.T)
# Computing eigenvectors
eigenValues,eigenVectors=np.linalg.eig(C)
# Sorting eigenvectors based on eigenvalues
idx=eigenValues.argsort()[::-1]
eigenValues=eigenValues[idx]
eigenVectors=eigenVectors[:,idx]
print("EIGENVALUES : ",eigenValues,sep='\n',end='\n\n')
print("EIGENVECTORS : ",eigenVectors,sep='\n')
fig=sns.scatterplot(X[:,0],X[:,1]);
points=list(zip([0,0],1.5*eigenValues[0]*eigenVectors[:,0]))
sns.lineplot(*points,label='PC1',color='red')
points=list(zip([0,0],1.5*eigenValues[1]*eigenVectors[:,1]))
sns.lineplot(*points,label='PC2',color='purple');
fig.set(title='Normalized Dataset with 2 Principal Components');
```
#### 2.3 Dimensionality Reduction with PCA
After computing the principal components, we can use them to reduce the feature dimension of our dataset by projecting each example onto a lower dimensional space $x^{(i)}\rightarrow z^{(i)}$. In this part of the exercise, **we will use the computed eigenvectors and project the example dataset into a 1-dimensional space.**
In practice, if we were using a learning algorithm such as linear regression or prehaps Nerual network, we could now use the projected data instead of the original data as there are less dimensions in the input.
##### 2.3.1 Projecting the data onto the principal components
```
# Selecting top k principal components from eigenvectors matrix
k=1
principalComponents=eigenVectors[:,:k]
# Projecting the centered data matrix using only the top K eigenvectors
Z=X.dot(principalComponents)
print(Z.shape)
fig=sns.scatterplot(Z.reshape(-1),0);
fig.set(title='Data projected in 1D (PC1)');
```
##### 2.3.2 Reconstructing an approximation of the data
After projecting the data onto the lower dimensional space, we can approximately recover the data by projecting them back onto the original high dimensional space.
```
xRec=Z.dot(principalComponents.T)
print(xRec.shape)
xRec[:5]
```
##### 2.3.3 Visualizing the projections
Here we will plot the projection and the approximate reconstruction to show how the projection affects the data.
```
fig,ax=plt.subplots(1,1,figsize=(10,8))
sns.scatterplot(X[:,0],X[:,1],label='Original');
sns.scatterplot(xRec[:,0],xRec[:,1],label='Projection');
for i in range(X.shape[0]):
points=list(zip(X[i,:],xRec[i,:]))
sns.lineplot(*points,color='gray')
ax.lines[i].set_linestyle('--')
ax.set(title='Original Data and reconstructed projection');
```
The reconstructed projection effectively only retains the information in the direction given by PC1.
#### 2.4 Face Image Dataset
In this part of the exercise, we will run PCA on face images to see how it can be used in practice for dimension reduction. The dataset contains 5000 grayscale face images of 32 X 32 pixel size.
```
mat=loadmat('./ex7faces.mat')
print(*mat.keys(),sep='\n')
# Loading data from mat to Dataframe
X=mat['X']
m,n=X.shape
data=pd.DataFrame()
for i in range(n):
data[f'x{i+1}']=X[:,i]
print('TRAINING DATASET SHAPE : {0} X {1}'.format(*data.shape))
data.sample(5)
```
**Visualizing data**
```
def displayData(X,title=None):
m,n=X.shape
width=int(np.sqrt(n))
height=int(n/width)
rows=int(np.floor(np.sqrt(m)))
cols=int(np.ceil(m/rows))
totalWidth=cols+cols*width
displayArray=np.zeros((1,totalWidth))
rowPadding=np.ones((1,totalWidth))*np.min(X)
colPadding=np.ones((height,1))*np.min(X)
index=0
for i in range(rows):
row=colPadding*0
for j in range(cols):
if index<m:
x=X[index].reshape((width,height)).T
index=index+1
else:
x=np.zeros((width,height)).T
row=np.column_stack((row,x))
if j<cols-1:
row=np.column_stack((row,colPadding))
displayArray=np.row_stack((displayArray,row))
if i<rows-1:
displayArray=np.row_stack((displayArray,rowPadding))
displayArray=np.row_stack((displayArray,rowPadding*0))
plt.imshow(displayArray,cmap='gray')
plt.title(title)
plt.axis('off')
randomIndices=np.random.randint(0,X.shape[0],49)
displayData(X[randomIndices,:],'Face Dataset');
```
**Before running PCA, let's take a look at mean of all human faces.**
```
plt.imshow(X.mean(axis=0).reshape((32,32)).T,cmap='gray')
plt.axis('off');
```
##### 2.4.1 PCA on Faces
To run PCA on faces, we will first normalize the dataset. After running PCA, we will obtain the principal components of the dataset. Notice that each principal component in VT is of length n=1024. It
```
# Normalizing
X=featureNormalize(X)
# SVD
U,S,VT=np.linalg.svd(X)
V=VT.T
# Sorting singular vector based on singular values
idx=S.argsort()[::-1]
S=S[idx]
V=V[:,idx]
print(f'U : {U.shape}')
print(f'S : {S.shape}')
print(f'V : {V.shape}')
fig,ax=plt.subplots(nrows=1,ncols=2,figsize=(10,4))
chart1=sns.distplot(S,kde=False,ax=ax[0]);
chart1.set(xlabel='Singular Values',ylabel='Occurences',title='Histogram')
s=S/np.sum(S)
chart2=sns.lineplot(list(range(len(s))),np.cumsum(s),drawstyle='steps',ax=ax[1])
chart2.set(xlabel='Singular Values',ylabel='Cumulative Sum',title='Cumulative Distribution');
```
From the above graph, we can conclude few things:
1. There are a lot of singular values that seems to matter for this dataset.
2. In order to get 95% variance of the data, we will end up selecting almost 800 out of 1024 singular values or principal components which are majority of them.
3. This data is very noisy and has a lot of variability which can only be explained by a dataset with relatively larger dimension.
Let's take a look at the first 36 principal components that describe the largest variations.
```
k=36
principalComponents=V[:,:k]
displayData(principalComponents.T,f'First {k} Principal Components');
```
##### 2.4.2 Dimensionality Reduction
Now that we have computed the principal components for the face dataset, we can use it to reduce the dimension of the face dataset. This allows us to use our learning algorithm with a smaller input size (e.g. 100 dimensions) instead of the original 1024 dimensions. This can help speed our learning algorithm.
```
# Selecting top k principal components from singular vector matrix
k=100
principalComponents=V[:,:k]
# Projecting the centered data matrix using only the top K singular vectors
Z=X.dot(principalComponents)
Z.shape
```
Reconstructing data matrix with original dimension from the lower dimension.
```
xRec=Z.dot(principalComponents.T)
print(xRec.shape)
displayData(X[:36],'Original Face Images');
displayData(xRec[:36],f'Reconstructed Face Images from \nTop {k} Principal Components')
```
From the reconstrution, we can observe that the genral structure and appearence of the face are kept while the fine details are lost. This is a remarkable reduction (more than 10x) in the dataset size that can help us speed our learning algorithm significantly. For example, if we were training a neural network to perform person recognition, we can use the dimension reduced input of only a 100 dimensions instead of the original dataset.
<img src="images/meme.jpg" width="400">
| github_jupyter |
# Mandatory Assignment 1
This is the first of two mandatory assignments which must be completed during the course. First some practical information:
* When is the assignment due?: **23:59, Sunday, August 19, 2018.**
* How do you grade the assignment?: You will **peergrade** each other as primary grading.
* Can i work with my group?: **yes**
The assigment consist of one to tree problems from each of the exercise sets you have solved so far (excluding Set 1). We've tried to select problems which are self contained, but it might be nessecary to solve some of the previous exercises in each set to fully answer the problems in this assignment.
## Problems from Exercise Set 2:
> **Ex. 2.2**: Make two lists. The first should be numbered. The second should be unnumbered and contain at least one sublevel.
```
# [Answer to Ex. 2.2 here] (convert to markdown cell)
```
## Problems from Exercise set 3:
> **Ex. 3.1.3:** Let `l1 = ['r ', 'Is', '>', ' < ', 'g ', '?']`. Create from `l1` the sentence `"Is r > g?"` using your knowledge about string formatting. Store this new string in a variable called `answer_31`. Make sure there is only one space in between worlds.
>
>> _Hint:_ You should be able to combine the above informations to solve this exercise.
```
# [Answer to Ex. 3.1.3 here]
l1 = ['r ', 'Is', '>', ' < ', 'g ', '?']
# answer_31 =
# YOUR CODE HERE
raise NotImplementedError()
assert answer_31 == "Is r > g?"
```
> **Ex. 3.1.4**: Create an empty dictionary `words` using the `dict()`function. Then add each of the words in `['animal', 'coffee', 'python', 'unit', 'knowledge', 'tread', 'arise']` as a key, with the value being a boolean indicator for whether the word begins with a vowel. The results should look like `{'bacon': False, 'asynchronous': True ...}`. Store the result in a new variable called `answer_32`.
>
>> _Hint:_ You might want co first construct a function that asseses whether a given word begins with a vowel or not.
```
# [Answer to Ex. 3.1.4 here]
W = ['animal', 'coffee', 'python', 'unit', 'knowledge', 'tread', 'arise']
# answer_32 =
# YOUR CODE HERE
raise NotImplementedError()
assert answer_32 == {i: i[0] in 'aeiou' for i in W}
assert sorted(answer_32) == sorted(W)
```
> **Ex. 3.3.2:** use the `requests` module (get it with `pip install requests`) and `construct_link()` which you defined in the previous question (ex 3.3.1) to request birth data from the "FOD" table. Get all available years (variable "Tid"), but only female births (BARNKON=P) . Unpack the json payload and store the result. Wrap the whole thing in a function which takes an url as input and returns the corresponding output.
>
> Store the birth data in a new variable called `answer_33`.
>
>> _Hint:_ The `requests.response` object has a `.json()` method.
>
>> _Note:_ you wrote `construct_link()` in 3.3.1, if you didn't heres the link you need to get: `https://api.statbank.dk/v1/data/FOLK1A/JSONSTAT?lang=en&Tid=*`
```
# [Answer to Ex. 3.3.2 here]
# answer_33 =
# YOUR CODE HERE
raise NotImplementedError()
assert sorted(answer_33['dataset'].keys()) == ['dimension', 'label', 'source', 'updated', 'value']
assert 'BARNKON' in answer_33['dataset']['dimension'].keys()
```
## Problems from exercise set 4
```
import numpy as np
import pandas as pd
```
> **Ex. 4.1.1:** Use Pandas' CSV reader to fetch daily data weather from 1864 for various stations - available [here](https://www1.ncdc.noaa.gov/pub/data/ghcn/daily/by_year/). Store the dataframe in a variable called `answer_41`.
>
>> *Hint 1*: for compressed files you may need to specify the keyword `compression`.
>
>> *Hint 2*: keyword `header` can be specified as the CSV has no column names.
>
>> *Hint 3*: Specify the path, as the URL linking directly to the 1864 file.
```
# [Answer to Ex. 4.1.1 here]
# answer_41 =
# YOUR CODE HERE
raise NotImplementedError()
assert answer_41.shape == (27349, 8)
assert list(answer_41.columns) == list(range(8))
```
> **Ex. 4.1.2:** Structure your weather DataFrame by using only the relevant columns (station identifier, data, observation type, observation value), rename them. Make sure observations are correctly formated (how many decimals should we add? one?).
>
> Store the resulting dataframe in a new variable called `answer_42`.
>
>> *Hint:* rename can be done with `df.columns=COLS` where `COLS` is a list of column names.
```
# [Answer to Ex. 4.1.2 here]
# answer_42 =
# YOUR CODE HERE
raise NotImplementedError()
assert answer_42.shape == (27349, 4)
assert 144.8 in [answer_42[i].max() for i in answer_42]
assert -666.0 in [answer_42[i].min() for i in answer_42]
assert 18640101 in [answer_42[i].min() for i in answer_42]
```
> **Ex. 4.1.3:** Select data for the station `ITE00100550` and only observations for maximal temperature. Make a copy of the DataFrame. Explain in a one or two sentences how copying works.
>
> Store the subsetted dataframe in a new variable called `answer_43`.
>
>> *Hint 1*: the `&` operator works elementwise on boolean series (like `and` in core python).
>
>> *Hint 2*: copying of the dataframe is done with the `copy` method for DataFrames.
```
# [Answer to Ex. 4.1.3 here]
# answer_43 =
# YOUR CODE HERE
raise NotImplementedError()
assert 'ITE00100550' in [answer_43[i].min() for i in answer_43]
assert 'ITE00100550' in [answer_43[i].max() for i in answer_43]
assert 'TMAX' in [answer_43[i].min() for i in answer_43]
assert 'TMAX' in [answer_43[i].max() for i in answer_43]
```
> **Ex. 4.1.4:** Make a new column in `answer_44` called `TMAX_F` where you have converted the temperature variables to Fahrenheit. Make sure not to overwrite `answer_43`.
>
> Store the resulting dataframe in a variable called `answer_44`.
>
>> *Hint*: Conversion is $F = 32 + 1.8*C$ where $F$ is Fahrenheit and $C$ is Celsius.
```
# [Answer to Ex. 4.1.4 here]
answer_44 = answer_43.copy()
# answer_44 =
# YOUR CODE HERE
raise NotImplementedError()
assert set(answer_44.columns) - set(answer_43.columns) == {'TMAX_F'}
```
## Problems from exercise set 5
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
%matplotlib inline
iris = sns.load_dataset('iris')
titanic = sns.load_dataset('titanic')
```
> **Ex. 5.1.1:**: Show the first five rows of the titanic dataset. What information is in the dataset? Use a barplot to show the probability of survival for men and women within each passenger class. Can you make a boxplot showing the same information (why/why not?). _Bonus:_ show a boxplot for the fare-prices within each passenger class.
>
> Spend five minutes discussing what you can learn about the survival-selection aboard titanic from the figure(s).
>
> > _Hint:_ https://seaborn.pydata.org/generated/seaborn.barplot.html, specifically the `hue` option.
```
# [Answer to Ex. 5.1.1 here]
# YOUR CODE HERE
raise NotImplementedError()
```
> **Ex. 5.1.2:** Using the iris flower dataset, draw a scatterplot of sepal length and petal length. Include a second order polynomial fitted to the data. Add a title to the plot and rename the axis labels.
> _Discuss:_ Is this a meaningful way to display the data? What could we do differently?
>
> For a better understanding of the dataset this image might be useful:
> <img src="iris_pic.png" alt="Drawing" style="width: 200px;"/>
>
>> _Hint:_ use the `.regplot` method from seaborn.
```
# [Answer to Ex. 5.1.2 here]
# YOUR CODE HERE
raise NotImplementedError()
```
> **Ex. 5.1.3:** Combine the two of the figures you created above into a two-panel figure similar to the one shown here:
> <img src="Example.png" alt="Drawing" style="width: 600px;"/>
>
> Save the figure as a png file on your computer.
>> _Hint:_ See [this question](https://stackoverflow.com/questions/41384040/subplot-for-seaborn-boxplot) on stackoverflow for inspiration.
```
# [Answer to Ex. 5.1.3 here]
# YOUR CODE HERE
raise NotImplementedError()
```
> **Ex. 5.1.4:** Use [pairplot with hue](https://seaborn.pydata.org/generated/seaborn.pairplot.html) to create a figure that clearly shows how the different species vary across measurements. Change the color palette and remove the shading from the density plots. _Bonus:_ Try to explain how the `diag_kws` argument works (_hint:_ [read here](https://stackoverflow.com/questions/1769403/understanding-kwargs-in-python))
```
# [Answer to Ex. 5.1.4 here]
# YOUR CODE HERE
raise NotImplementedError()
```
## Problems from exercise set 6
> _Note:_ In the exercises we asked you to download weather data from the NOAA website. For this assignment the data are loaded in the following code cell into two pandas dataframes.
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
weather_1864 = pd.read_csv('weather_data_1864.csv')
```
> **Ex. 6.1.4:** Extract the country code from the station name into a separate column.
>
> Create a new column in `weather_1864` called `answer_61` and store the country codes here.
>
>> _Hint:_ The station column contains a GHCND ID, given to each weather station by NOAA. The format of these ID's is a 2-3 letter country code, followed by a integer identifying the specific station. A simple approach is to assume a fixed length of the country ID. A more complex way would be to use the [`re`](https://docs.python.org/2/library/re.html) module.
```
# [Answer to Ex. 6.1.4]
# weather_1864['answer_61'] =
# YOUR CODE HERE
raise NotImplementedError()
assert sorted(weather_1864['answer_61'].str[:2].unique()) == sorted(['SZ', 'CA', 'EZ', 'GM', 'AU', 'IT', 'BE', 'UK', 'EI', 'AG', 'AS'])
```
> **Ex. 6.1.5:** Make a function that downloads and formats the weather data according to previous exercises in Exercise Section 4.1, 6.1. You should use data for ALL stations but still only select maximal temperature. _Bonus:_ To validate that your function works plot the temperature curve for each country in the same window. Use `plt.legend()` to add a legend.
>
> Name your function `prepareWeatherData`.
```
# [Answer to Ex. 6.1.5]
def prepareWeatherData(year):
# Your code here
return
# YOUR CODE HERE
raise NotImplementedError()
assert prepareWeatherData('1864').shape == (5686, 6)
```
## Problems from exercise set 7
> _Note:_ Once again if you haven't managed to download the data from NOAA, you can refer to the github repo to get csv-files containing the required data.
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
# Increases the plot size a little
mpl.rcParams['figure.figsize'] = 11, 6
```
> **Ex. 7.1.1:** Plot the monthly max,min, mean, first and third quartiles for maximum temperature for our station with the ID _'ITE00100550'_ in 1864.
> *Hint*: the method `describe` computes all these measures.
```
# [Answer to Ex. 7.1.1]
# YOUR CODE HERE
raise NotImplementedError()
```
> **Ex. 7.1.2:** Get the processed data from years 1864-1867 as a list of DataFrames. Convert the list into a single DataFrame by concatenating vertically.
>
> Name the concatenated data `answer_72`
```
# [Answer to Ex. 7.1.2]
# YOUR CODE HERE
raise NotImplementedError()
assert answer_72.shape == (30003, 7)
```
> **Ex. 7.1.3:** Parse the station location data which you can find at https://www1.ncdc.noaa.gov/pub/data/ghcn/daily/ghcnd-stations.txt. Merge station locations onto the weather data spanning 1864-1867.
>
> Store the merged data in a new variable called `answer_73`.
>
> _Hint:_ The location data have the folllowing format,
```
------------------------------
Variable Columns Type
------------------------------
ID 1-11 Character
LATITUDE 13-20 Real
LONGITUDE 22-30 Real
ELEVATION 32-37 Real
STATE 39-40 Character
NAME 42-71 Character
GSN FLAG 73-75 Character
HCN/CRN FLAG 77-79 Character
WMO ID 81-85 Character
------------------------------
```
> *Hint*: The station information has fixed width format - does there exist a pandas reader for that?
```
# [Answer to Ex. 7.1.3]
# YOUR CODE HERE
raise NotImplementedError()
assert answer_73.shape == (5686, 15) or answer_73.shape == (30003, 15)
```
## Problems from exercise set 8
> **Ex. 8.1.2.:** Use the `request` module to collect the first page of job postings.
>
> Store the response.json() object in a new variable called `answer_81`.
>
```
# [Answer to Ex. 8.1.2]
# YOUR CODE HERE
raise NotImplementedError()
assert sorted(answer_81.keys()) == sorted(['Expression', 'Facets', 'JobPositionPostings', 'TotalResultCount'])
```
> **Ex. 8.1.3.:** Store the 'TotalResultCount' value for later use. Also create a dataframe from the 'JobPositionPostings' field in the json. Name this dataframe `answer_82`.
```
# [Answer to Ex. 8.1.3]
# answer_82 =
# YOUR CODE HERE
raise NotImplementedError()
assert answer_82.shape == (20,44)
```
| github_jupyter |
## Figure 16
The classification performance metrics for
BMC (blue), TPC (green), morphology (red), and HB (purple)
as applied to the CFHTLenS data in the VVDS field
with various magnitude cuts.
The top panel shows the number of sources in the training set
at corresponding magnitude cuts.
We show only one of the four combination methods, BMC,
which has the best overall performance.
```
%matplotlib inline
from __future__ import division, print_function, unicode_literals
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.rc('legend', fontsize=10)
truth_train = np.loadtxt('../../data/truth_train.dat')
truth_test = np.loadtxt('../../data/truth_test.dat')
# load base classifiers
mag_cut_str = ['99', '24_0', '23_5', '23_0', '22_5', '22_0', '21_5', '21_0', '20_5', '20_0']
tpc = [np.loadtxt('../../data/vvds_{0}_tpc_test.mlz'.format(i), unpack=True, usecols=(2,)) for i in mag_cut_str]
som = [np.loadtxt('../../data/vvds_{0}_som_test.mlz'.format(i), unpack=True, usecols=(2,)) for i in mag_cut_str]
hbc = [np.loadtxt('../../data/vvds_{0}_median.hbc'.format(i), unpack=True, usecols=(2,)) for i in mag_cut_str]
hbc = [i[-len(truth_test):] for i in hbc]
bmc = [np.loadtxt('../../data/vvds_{0}.bmc'.format(i), unpack=True, usecols=(0,)) for i in mag_cut_str]
def calc_completeness_purity(truth, classif, mag, p_cut=0.5, bins=np.arange(16, 26, 0.5)):
'''
'''
# true galaxies classified as stars
gs_bin, _ = np.histogram(mag[(classif > p_cut) & (truth == 0)], bins=bins)
# true galaxies classified as galaxies
gg_bin, _ = np.histogram(mag[(classif < p_cut) & (truth == 0)], bins=bins)
# true stars classified as galaxies
sg_bin, _ = np.histogram(mag[(classif < p_cut) & (truth == 1)], bins=bins)
# true stars classified as stars
ss_bin, _ = np.histogram(mag[(classif > p_cut) & (truth == 1)], bins=bins)
# galaxy completeness
g_comp_bin = gg_bin / (gg_bin + gs_bin)
g_comp_bin[~np.isfinite(g_comp_bin)] = 1
# galaxy purity
g_pur_bin = gg_bin / (gg_bin + sg_bin)
g_pur_bin[~np.isfinite(g_pur_bin)] = 1
# star completeness
s_comp_bin = ss_bin / (ss_bin + sg_bin)
s_comp_bin[~np.isfinite(s_comp_bin)] = 1
# star purity
s_pur_bin = ss_bin / (ss_bin + gs_bin)
s_pur_bin[~np.isfinite(s_pur_bin)] = 1
return g_comp_bin, g_pur_bin, s_comp_bin, s_pur_bin
def find_purity_at(truth_test, clf, step=0.001, gc=None, sc=None):
if bool(gc) == bool(sc):
raise Exception('Specify only one of gp or sp parameter.')
pbin = np.arange(0, 1, step)
pure_all = np.zeros(len(pbin))
comp_all = np.zeros(len(pbin))
for i, p in enumerate(pbin):
# true galaxies classified as stars
gs = ((clf >= p) & (truth_test == 0)).sum()
# true galaxies classified as galaxies
gg = ((clf < p) & (truth_test == 0)).sum()
# true stars classified as galaxies
sg = ((clf < p) & (truth_test == 1)).sum()
# true stars classified as stars
ss = ((clf >= p) & (truth_test == 1)).sum()
if gc is not None:
if gg == 0 and gg + sg == 0:
pure_all[i] = 1
else:
pure_all[i] = gg / (gg + sg)
if gg == 0 and gg + gs == 0:
comp_all[i] = 1
else:
comp_all[i] = gg / (gg + gs)
if sc is not None:
if ss == 0 and ss + sg == 0:
comp_all[i] = 1
else:
comp_all[i] = ss / (ss + sg)
if ss == 0 and ss + gs == 0:
pure_all[i] = 1
else:
pure_all[i] = ss / (ss + gs)
if gc is not None:
ibin = np.argmin(np.abs(comp_all - gc))
return pbin[ibin], pure_all[ibin]
if sc is not None:
ibin = np.argmin(np.abs(comp_all - sc))
return pbin[ibin], pure_all[ibin]
from sklearn.metrics import roc_auc_score
tpc_auc = [roc_auc_score(truth_test, i) for i in tpc]
som_auc = [roc_auc_score(truth_test, i) for i in som]
hbc_auc = [roc_auc_score(truth_test, i) for i in hbc]
bmc_auc = [roc_auc_score(truth_test, i) for i in bmc]
from sklearn.metrics import mean_squared_error
tpc_mse = [mean_squared_error(truth_test, i) for i in tpc]
som_mse = [mean_squared_error(truth_test, i) for i in som]
hbc_mse = [mean_squared_error(truth_test, i[-len(truth_test):]) for i in hbc]
bmc_mse = [mean_squared_error(truth_test, i) for i in bmc]
def find_gal_pur(truth, clf, gc=0.9964, step=0.001):
result = []
for k in clf:
_, j = find_purity_at(truth, k, gc=gc, step=step)
result += [j]
return result
tpc_gpur = find_gal_pur(truth_test, tpc)
bmc_gpur= find_gal_pur(truth_test, bmc)
hbc_gpur= find_gal_pur(truth_test, hbc)
som_gpur= find_gal_pur(truth_test, som)
def find_star_pur(truth, clf, sc=0.7145, step=0.001):
result = []
for k in clf:
_, j = find_purity_at(truth, k, sc=sc, step=step)
result += [j]
return result
tpc_spur = find_star_pur(truth_test, tpc)
bmc_spur = find_star_pur(truth_test, bmc)
hbc_spur = find_star_pur(truth_test, hbc)
som_spur = find_star_pur(truth_test, som)
p = sns.color_palette()
sns.set_style("ticks")
markersize = 4
fig = plt.figure(figsize=(6, 10))
ax1 = plt.subplot2grid((5, 3), (0, 0), colspan=3)
ax2 = plt.subplot2grid((5, 3), (1, 0), colspan=3)
ax3 = plt.subplot2grid((5, 3), (2, 0), colspan=3)
ax4 = plt.subplot2grid((5, 3), (3, 0), colspan=3)
ax5 = plt.subplot2grid((5, 3), (4, 0), colspan=3)
plt.setp(ax1.get_xticklabels(), visible=False)
plt.setp(ax2.get_xticklabels(), visible=False)
plt.setp(ax3.get_xticklabels(), visible=False)
plt.setp(ax4.get_xticklabels(), visible=False)
plt.setp(ax5.get_xticklabels(), rotation=45)
ax1.plot([10456, 9968, 9268, 8506, 7686, 6361, 4949, 3619, 2565, 1755], ls='-', marker='o', markersize=markersize)
ax1.set_ylabel('training size')
ax1.legend(loc='upper right')
ax1.set_yticks([0, 2000, 4000, 6000, 8000, 10000])
ax1.set_yticklabels([r'$0$', r'$2 \times 10^3$', r'$4 \times 10^3$', r'$6 \times 10^3$', r'$8 \times 10^3$',
r'$1 \times 10^4$'])
for ticks in ax1.get_yaxis().majorTicks[1:]:
ticks.set_pad(0)
ax2.plot(bmc_auc, label='BMC',
color=p[0], ls='-', marker='o', markersize=markersize)
ax2.plot(tpc_auc, label='TPC',
color=p[1], ls='-', marker='o', markersize=markersize)
ax2.plot(hbc_auc, label='HB',
color=p[3], ls='-', marker='o', markersize=markersize)
ax2.set_ylim(0.93, 1.0)
ax2.set_yticks([0.94, 0.96, 0.98, 1.0])
ax2.legend(loc='upper left', ncol=3)
ax2.set_ylabel('AUC')
ax2.set_ylabel('AUC')
ax3.plot(bmc_mse, label='BMC',
color=p[0], ls='-', marker='o', markersize=markersize)
ax3.plot(tpc_mse, label='TPC',
color=p[1], ls='-', marker='o', markersize=markersize)
ax3.plot([0.0397] * len(tpc_mse),
color=p[2], label='Morphology', ls='-', marker='o', markersize=markersize)
ax3.plot(hbc_mse, label='HB',
color=p[3], ls='-', marker='o', markersize=markersize)
ax3.set_ylim(0.02, 0.15)
ax3.set_yticks([0.04, 0.08, 0.12])
ax3.set_ylabel('MSE')
ax3.legend(loc='upper left', fancybox=True, framealpha=0.5, ncol=2)
ax4.plot(bmc_gpur, label='BMC',
color=p[0], ls='-', marker='o', markersize=markersize)
ax4.plot(tpc_gpur, label='TPC',
color=p[1], ls='-', marker='o', markersize=markersize)
ax4.plot([0.9597] * len(tpc_gpur),
color=p[2], label='Morphology', marker='o', markersize=markersize)
ax4.plot(hbc_gpur, label='HB',
color=p[3], ls='-', marker='o', markersize=markersize)
ax4.set_ylim(0.92, 0.98)
ax4.set_yticks([0.93, 0.95, 0.97])
ax4.set_ylabel(r'$p_g\left(c_g=0.9964\right)$', fontsize=12)
ax5.plot(bmc_spur, label='BMC',
color=p[0], ls='-', marker='o', markersize=markersize)
ax5.plot(tpc_spur, label='TPC',
color=p[1], ls='-', marker='o', markersize=markersize)
ax5.plot([0.9666] * len(tpc_spur),
color=p[2], label='Morphology', ls='-', marker='o', markersize=markersize)
ax5.plot(hbc_spur, label='HB',
color=p[3], ls='-', marker='o', markersize=markersize)
ax5.set_ylim(0.65, 1.04)
ax5.set_yticks([0.7, 0.8, 0.9, 1.0])
ax5.set_ylabel(r'$p_s\left(c_s=0.7145\right)$', fontsize=12)
ax5.set_xticklabels(['', '< 24.0', '< 23.5', '< 23.0', '< 22.5',
'< 22.0', '< 21.5', '< 21.0', '< 20.5', '< 20.0'])
ax5.set_xlabel(r'$i$ magnitude cut')
plt.savefig('../../figures/perform_mag_cut.pdf')
plt.show()
```
| github_jupyter |
# California Weather Analysis
This is a report on the historical analysis of weather patterns in an area that approximately overlaps the Central area of California.
The data we will use here comes from [NOAA](https://www.ncdc.noaa.gov/). Specifically, it was downloaded from This [FTP site](ftp://ftp.ncdc.noaa.gov/pub/data/ghcn/daily/).
We focused on six measurements:
* **TMIN, TMAX:** the daily minimum and maximum temperature.
* **TOBS:** The average temperature for each day.
* **PRCP:** Daily Percipitation (in mm)
* **SNOW:** Daily snowfall (in mm)
* **SNWD:** The depth of accumulated snow.
## 1. Span of Weather Stations
By plotting the latitude,longitude of the weather stations in the dataset, it was found that all the weather stations belong to the coastal side of central California.
<p><img style='height:300px' src="myfigs/location_map.png" /></p>
### 1.1 Sanity-check: comparison with outside sources
<p>As a quick sanity check, I picked one of the weather stations from my dataset at Oakley, california and got it's weather statistics from the <a href='http://www.usclimatedata.com/climate/oakley/california/united-states/usca2070'>US climate data</a> website.The graph below shows the daily minimum and maximum temperatures for each month, as well as the total precipitation for each month.</p>
<p> </p>
<p><img style='height:300px' src="myfigs/sanity_check_oakley.png" /></p>
<p> </p>
<p>We see that the min and max daily temperature approximately agrees with the pattern we got from our data(shown in the below figure). Same is the case with the precipitation data, the mean precipitation is approximately similar to the data obtained from US climate data website. A point to be noted here is that the statistics obtained from US climate data is only pertaining to a single weather station while the mean precipitation shown in our graph is an average over the entire Central California region.</p>
<p> </p>
<p><img alt="TMIN,TMAX.png" src="myfigs/sanity_check_oakley2.png" style="height:300px;" /></p>
<p> <img alt="PRCP.png" src="myfigs/sanity_check_oakley3.png" style="height:300px;" /></p>
## 2. PCA analysis
For each of the six measurement, we compute the percentate of the variance explained as a function of the number of eigen-vectors used.
### Percentage of variance explained.

We see that the top 5 eigen-vectors explain 38% of variance for TMIN, 59% for TOBS and 47% for TMAX.
We conclude that of the three, TOBS is best explained by the top 5 eigenvectors. This is especially true for the first eigen-vector which, by itself, explains 48% of the variance. TMAX and TMIN also have a pretty good approximation using the first five eigen vectors since they contribute to a significant share of the total variance. On the whole it can be inferred that the temperature statistics follow a pattern with majority of the variance along a few principal axes.
Based on this initial promise, we will dig deeper into the PCA analysis of TOBS. However we can expect that the TOBS data is going to be noisy since the temperatures on consecutive days of the year do not follow a strictly increasing or decreasing sequence over a considerably big window.
<p></p>

The top 5 eigenvectors explain only 16% of the variance for PRCP. On the other hand the top 5 eigenvectors explain 72% of the variance for SNWD and 68% for SNOW. This means that these top 5 eigenvectors capture most of the variation in the snow signals. However this is not statistically significant since the majority of the weather stations are located in parts of California where it never really snows. As we can see from the below graphs the mean snow depth is close to zero for major parts of the year apart from minor spikes. Also the top three eigen vectors remain zero almost the whole year except for a few sporadic spikes. This means that any reconstruction using the top few eigen vectors will will get 70% of the data correct simply owing to the fact that the snow is zero always.
<p> <img src="myfigs/snwd-stats.png" style="height:300px;" /></p>
### 2.1 Analysis of TOBS
We choose to analyze the eigen-decomposition for TOBS because the first 3 eigen-vectors explain more than 55% of the variance.
First, we graph the mean and the top 3 eigen-vectors.
We observe that the average temperatures conform to the seasonal pattern with the temperature being maximum between mid june and end of october which is the Summer season. Likewise the minimum temperature is observed between December and March which is the Winter season.
<p> <img src="myfigs/tobs-stats.png" style="height:300px;" /></p>
Next we interpret the eigen-functions. The first eigen-vector has a shape very similar to the mean function. The interpretation of this shape is that eig1 represents the deviation of temperature above/below the mean, but without changing the distribution over time.
**eig2 and eig3** are similar in the following way. They peak during a certain period of the year. In other words, they correspond to the deviation in temperature distribition between different months.
They can be interpreted as follows:
* **eig2:** less temperature in june - october than the rest of the year.
* **eig3:** more temperature in march - july, than the rest of the year.
#### 2.1.1 Examples of reconstructions
In all of the below graphs we find that the reconstruction of the target vector gets more and more accurate with additional eigen vectors. As stated in the earlier section,the average daily temperatures is not a smoothly increasing/decreasing function and the reconstruction from the eigen vectors minimizes the noise in the function.
#### Coeff1
Coeff1: small values

Coeff1: large values

Large values of coeff1 correspond to more than average temperature and low values correspond to less than average temperature.
#### Coeff2
Coeff2: small values

Coeff2: large values

Large values of coeff2 correspond to low summer temperatures between june and october. Small values for coeff2 correspond to high summer temperatures.
#### Coeff3
Coeff3: small values

Coeff3: large values

Large values of coeff3 correspond to a high temperatures during the march-july period of the year and small values of coeff3 correspond to low temperatures during the march-july timeperiod.
#### 2.1.2 Cumulative Distribution of residuals and coefficients
The below graph shows a plot of the cumulative distribution function of the first three residuals. As expected the residual2 is better than residual1 since the first two eigen vectors combined capture more variance than the first eigen vector alone. In other words the residual error from reconstruction using only the first eigen vector grows faster than the residual error from reconstruction using the first two eigen vectors combined. However we can see that there is not much difference in the cumulative residual errors of res_2 and res_3. This behaviour is as expected,conforming with what we saw in the percentage variance explained plot of TOBS, where the increase in the percentage of variance explained between 2 and 3 eigen vectors is very small.<br/>

The below graph shows a plot of the cumulative distribution function of the first three coefficients of the eigen vectors. Since the first eigen vector gives the direction of maximum variation it is natural that many data points are dominated by a large coefficient1 compared to the other coefficients. As we can see there is not much difference between the coefficients 2 and 3, for the same reason as explained above.

#### 2.1.3 Visualizing data distribution for TOBS
The below map shows a distribution of the different weather stations that have collected the TOBS measurement in the central California region. The size of the circles in the map is proportional to the number of datapoints contributed by a particular weather station. The fill color of the circles is a heatmap denoting the average value of coefficient1 which is the deviation of temperature from the mean. A red weather station indicates a high average temperature while a blue weather station indicates a relatively lower average temperature.

## 3. Analysis of Precipitation
There is an average rainfall of 13.31 mm/day in the given region. As shown in the below graph, most of the rain occurs during the period of November to February. The CDF plot shows that it rains for about 20% of the days in our region. The first eigen vector represents the deviation in rainfall from mean. The second and third eigen vectors represent seasonal rain.
<p> <img src="myfigs/prcp-stats.png" style="height:300px;float:left;" /><img src="myfigs/cdf-rain.png" style="height:258px;" /></p>
Since the weather stations are all close to one another there is good chance that precipitation in one station guarantees precipitation in a nearby station. To accept/reject our hypothesis we begin with plotting a correlation matrix of Log probabilities where each of the values represent the probability of a coincidental rain in two weather stations.
<p> <img src="myfigs/7-correlation.png" style="height:400px;" /></p>
We can see from the above graph that the first 30 weather stations are correlated. To find more correlations we use PCA of this correlation matrix and cluster the weather stations based on the first few principal components. As shown in the below graph the top 10 eigen vectors of the correlation matrix explain about 90% of the of the square magnitude of the matrix.
<p> <img src="myfigs/7-correlation-decomp.png" style="height:300px;" /></p>
For the purpose of clustering, we consider only the first four principal components. We sort the weather stations in the correlation matrix according to increasing order of the magnitude of dimensions of the first eigen vector. The resultant correlation matrix visualized as a heatmap shows clusters of weather stations that are correlated based on the first eigen vector. We repeat this process for the 2nd,3rd and 4th eigen vectors. Below are the new heatmaps obtained after sorting. From the upper left heatmap we can see that the first 40 stations are correlated and especially the first 20 are highly correlated.
<p> <img src="myfigs/7-heatmap.png" style="height:700px;" /></p>
Plotting the first few correlated weather stations on a geo map, we get the below plot. From our analysis, it is evident that the weather stations that are nearby have a good chance of experiencing rain on the same day of the year.
<p> <img src="myfigs/7-heatmap-geo.png" style="height:300px;" /></p>
An alernate method of visualizing the data is to plot the weather stations along with the coefficients of the first principal components. The below geo plot indicates the principal components by triangles with the size of the triangle representing the magnitude of the coefficient and the opacity of the triangle representing the sign of the coefficient (filled triangles for negative and un-filled triangles for positive coefficients). As we can see there are a few close weather stations with similar triangle structures,that is, similar principal components. Both the visualizations(the above map and the below map) conform that the weather stations near the region of Concord and Pleasant Hill are correlated and experience precipitaion on the same days of the year.
<p> <img src="myfigs/5.5-map.png" style="height:300px;" /></p>
## 4. Temporal Vs Spatial Analysis of Precipitation
In the previous section we see the variation of Coeff1, which corresponds to the total amount of rain, with respect to location. We now estimate the relative importance of location-to-location variation relative to year-by-year variation.
These are measured using the fraction by which the variance is reduced when we subtract from each station/year entry the average-per-year or the average-per-station respectively. Here are the results:
coeff_1<br/>
total RMS = 194.754604183<br/>
RMS removing mean-by-station= 173.891026199 Fraction explained= 10.71%<br/>
RMS removing mean-by-year = 120.264234979 Fraction explained= 38.24%<br/>
coeff_2 <br/>
total RMS = 180.793723228<br/>
RMS removing mean-by-station= 172.563345122 Fraction explained= 4.55%<br/>
RMS removing mean-by-year = 80.9796786501 Fraction explained= 55.20%<br/>
coeff_3<br/>
total RMS = 171.693528795<br/>
RMS removing mean-by-station= 167.550306474 Fraction explained= 2.41%<br/>
RMS removing mean-by-year = 70.5968252719 Fraction explained= 58.88%<br/>
We see that the variation by year explains more than the variation by station. However this effect is weaker consider coeff_1, which has to do with the total rainfall, vs. coeff_2,3 which, as we saw above have to do with the timining of rainfall. We see that for coeff_2,3 the stations explain 2-5% of the variance while the year explaines 55-60%.
| github_jupyter |
# Numpy
---
```
import numpy as np
```
## Converting shape of an array
```
help(np.reshape)
```
Create 1-dimensional array
```
arr = np.arange(0, 12)
arr
```
Reshape to new 2 dimensional array
```
arr.reshape(3, 4)
```
*reshapes one array to another with given dimension in row, column format*
Reshape to new 3 dimensional array
```
arr.reshape(3, 2, 2)
```
*reshapes one array to another with given dimension in depth, row, column format*
*size must be same after reshaping, cannot reshape to arbitrary size array*
```
arr.size
arr.reshape(3, 4).size
arr.reshape(3, 2, 2).size
```
## Indexing
Select by index number, index starts with zero "$0$"
```
arr[0]
arr[2]
arr[-1]
arr[12]
```
2 dimensional array
```
arr2 = arr.reshape(4, 3)
arr2
arr2[0]
arr2[-1]
```
*Numpy has another syntax for accessing multidimensional array more elegantly*
`arr[row, column]`
```
arr2[0, 1]
```
*select item from 1st row and 2nd column.*
> *Which is same as `arr[0][1]` but above syntax should be preferrable*
```
arr2[2, 2]
```
3 - dimensional array
```
arr3 = arr.reshape(2, 2, 3)
arr3
arr3[1, 0, 1]
```
*select from 2nd depth, 1st row and 2nd column*
> `arr[depth, row, column]`
## Slicing
```
arr2
```
Get all items from 3rd row
```
arr2[2, :]
arr2[2]
```
*empty colon $:$ is redundant here*
Get all items from 2nd column
```
arr2[:, 1]
# get all items from 3rd row except first
arr[2, 1:]
# get all items from 3rd row except last
arr[2, :-1]
arr
# get all items from 2nd column
arr[:, 1]
# get all items from 2nd column except first
arr[1:, 0]
# get all items from 2nd column except last
arr[:-1, 1]
# get 2nd and 3rd column items from 2nd and 3rd row
arr[1:3, 1:3]
```
### Assigning values to array
```
arr
arr[0, 0] = 20
arr
arr[1, 1:]
arr[1, 1:] = [55, 66, 77]
arr
arr[1:3, 1:3] = [[555, 666], [999, 100]]
arr
```
**slices are references**
```
ab = arr[1, 2:]
ab
ab[0] = 667
arr
```
> *`667` is reflected on `arr`, because `ab` is just a reference to slice of `arr`*
### Three dimensional array
```
arr3 = np.arange(24).reshape(2, 3, 4)
arr3
```
> *`depth, rows, columns`*
**Indexing**
`arr3[depth, row, column]`
```
arr3[1]
arr3[1, 0]
arr3[1, 0, 3]
```
**slicing**
```
arr3[1, 1:, :-1]
arr3[:, 1:, :-1]
```
### Fancy Indexing
```
arr
# select 2nd and 3rd row
arr[[1, 2]]
# select 1st, 3rd and 4th item of 2nd row
arr[1, [0, 2, 3]]
# select 1st and 4th item of 2nd and 3rd column respectively
arr[[0, -1], [0, -1]]
arr[::2, [0, -1]]
arr[1, 2]
```
### Boolean operation and Masking
```
abc = np.arange(30)
abc
abc < 10
```
> *gives us boolean array, with True on each element which satifies the condition*
*Only select that satisfy condition*
```
abc[abc < 10]
abc[(abc < 10) | (abc > 20)]
```
use __&__ for "_and_", __|__ for "_or_". Each condition must be enclosed in brackets*
*each condition should be enclosed in brackets, to remove ambiguity created by operator precedence*
```
abc[abc % 2 == 0]
```
## Numpy Methods
```
abc
# sum of all elements
abc.sum()
# product
abc.prod()
abc[(abc > 1) & (abc < 10)].prod()
# index of minimum value
abc.argmin()
# index of maximum value
abc.argmax()
```
**argmin/argmax in multidimensional array**
```
arr
# index of minimum value as if given array is flat
arr.argmin()
arr.argmax()
np.unravel_index(arr.argmin(), arr.shape)
np.unravel_index(arr.argmax(), arr.shape)
help(np.unravel_index)
np.unravel_index(6, arr.shape)
np.unravel_index(6, (4, 2))
```
> return tuple is index within multidimensional array, ie. 0th row and 1st column
*sum across rows or across columns*
```
arr
# across rows
arr.sum(axis=0)
# across columns
arr.sum(axis=1)
```
*can be used with other methods as well*
```
arr.mean(axis=1)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# `GiRaFFE_NRPy_staggered`: Source Terms
## Author: Patrick Nelson
**Notebook Status:** <font color='green'><b>Validated</b></font>
**Validation Notes:** This code is a port from the old `GiRaFFE`
### NRPy+ Source Code for this module:
* [GiRaFFE_NRPy/GiRaFFE_NRPy_staggered_Source_Terms.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_staggered_Source_Terms.py)
## Introduction:
This notebook presents an alternate algorithm for computing the source terms in a staggered prescription. It is a direct port of the old `GiRaFFE` implementation.
Our goal here will to be to add the following terms to the appropriate right hand side:
* $\partial_t A_i -= \partial_i (\alpha \Phi - \beta^j A_j)$
* $\partial_t [\sqrt{\gamma} \Phi] + \partial_j (\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi]) = - \xi \alpha [\sqrt{\gamma} \Phi]$
(Note, the *whole* of the $\partial_t [\sqrt{\gamma} \Phi]$ evolution equation is of interest here.)
This will be done using basic second-order finite-difference derivatives; however, there is a significant wrinkle here: Not all of these quantities are sampled at cell centers! In particular, at code index `i,j,k`, the following quantities are what is actually available:
* Centers: $B^x(i,j,k)$, $B^y(i,j,k)$, $B^z(i,j,k)$
* Faces: $B^x(i+1/2,j,k)$, $B^y(i,j+1/2,k)$, $B^z(i,j,k+1/2)$
* Edges: $A_x(i,j+1/2,k+1/2)$, $A_y(i+1/2,j,k+1/2)$, $A_z(i+1/2,j+1/2,k)$
* Vertices: $[\sqrt{\gamma} \Phi](i+1/2,j+1/2,k+1/2)$
(Unless otherwise specified above, the quantity is available at cell centers.)
To overcome this, we will make heavy use of interpolation; it is sufficient for our purposes to merely average over the nearest points.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#prelim): Preliminaries
1. [Step 2](#code): Write the C code
1. [Step 2.a](#define_constants): Function definitions and useful constants
1. [Step 2.b](#operands): Compute derivative operands
1. [Step 2.b.i](#interp): Read in quantities needed for interpolation
1. [Step 2.b.ii](#raise_a): Compute $\alpha \sqrt{\gamma} A^j$
1. [Step 2.b.iii](#a_gauge_term): Compute $\alpha \Phi - \beta^j A_j$
1. [Step 2.c](#finite_difference): Take the finite difference derivatives
1. [Step 2.c.i](#a_rhs): Compute the $A_i$ gauge term
1. [Step 2.c.ii](#phi_rhs): Compute the $[\psi^6 \Phi]$ gauge term
1. [Step 2.d](#interp_func): The interpolation function
1. [Step 3](#code_validation): Code Validation
1. [Step 4](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='prelim'></a>
# Step 1: Preliminaries \[Back to [top](#toc)\]
$$\label{prelim}$$
This first block of code just sets up a subdirectory within `GiRaFFE_standalone_Ccodes/` to which we will write the C code.
```
# Step 0: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
Ccodesdir = "GiRaFFE_standalone_Ccodes/RHSs"
cmd.mkdir(os.path.join(Ccodesdir))
```
<a id='code'></a>
# Step 2: Write the C code \[Back to [top](#toc)\]
$$\label{code}$$
<a id='define_constants'></a>
## Step 2.a: Function definitions and useful constants \[Back to [top](#toc)\]
$$\label{define_constants}$$
We will first declare the crucial function `avg()`, our interpolator; its inputs are a grid function as will as the limits over which we will loop. [It is defined below](#interp_func).
We will also `#define` some constants. We will frequently need to loop *near* a given gridpoint by up to two gridpoints in any direction, so these constants will allow us to define these limits more intuitively. The rest of these provide similar heuristics to access memory within the construct `in_vars` and `INTERP_VARS` that we will use to pass data without a large mess of function inputs.
```
%%writefile $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
static inline REAL avg(const REAL f[PLUS2+1][PLUS2+1][PLUS2+1],const int imin,const int imax, const int jmin,const int jmax, const int kmin,const int kmax);
#define MINUS2 0
#define MINUS1 1
#define PLUS0 2
#define PLUS1 3
#define PLUS2 4
// The "I" suffix denotes interpolation. In other words, these
// definitions are used for interpolation ONLY. The order here
// matters as well!
static const int SHIFTXI=0,SHIFTYI=1,SHIFTZI=2,GUPXXI=3,GUPXYI=4,GUPXZI=5,GUPYYI=6,GUPYZI=7,GUPZZI=8,
PSII=9,LAPM1I=10,A_XI=11,A_YI=12,A_ZI=13,LAPSE_PSI2I=14,LAPSE_OVER_PSI6I=15;
#define MAXNUMINTERP 16
// GiRaFFE_NRPy does not store the inverse metric. So, the actual inputs to the function will be
// just the metric, which we can invert in an early step. To keep things consistent, we'll label the
// components with the same labels as the inverse:
static const int GXXI=3,GXYI=4,GXZI=5,GYYI=6,GYZI=7,GZZI=8;
```
Our function includes the C parameters `params`, an array of pointers to our input variables `in_vars`, the input `psi6phi`, eight arrays for temporary variables, and the output arrays for the potential.
Finally, we also declare variables from the paramstruct in the normal way with the `#include` directive. (If you're reading this code and can't figure out where a variable is set, check here!)
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
static void Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs(const paramstruct *params,REAL **in_vars,const REAL *psi6phi,
/* TEMPS: */
REAL *shiftx_iphjphkph,REAL *shifty_iphjphkph,REAL *shiftz_iphjphkph,
REAL *alpha_iphjphkph,REAL *alpha_Phi_minus_betaj_A_j_iphjphkph,REAL *alpha_sqrtg_Ax_interp,
REAL *alpha_sqrtg_Ay_interp,REAL *alpha_sqrtg_Az_interp,
/* END TEMPS, 8 total! */
REAL *psi6phi_rhs,REAL *Ax_rhs,REAL *Ay_rhs,REAL *Az_rhs) {
#include "../set_Cparameters.h"
```
<a id='operands'></a>
## Step 2.b: Compute derivative operands \[Back to [top](#toc)\]
$$\label{operands}$$
We will first compute $ \partial_t [\psi^6\Phi] = -\partial_j ( \alpha \sqrt{\gamma} A^j - \beta^j \psi^6\Phi)$, taken from Eq. 13 of [this](https://arxiv.org/pdf/1110.4633.pdf) paper (the Lorentz gauge evolution). Since all input variables to this are defined at all gridpoints, we can safely start out by looping over the entire grid *except* for the outermost ghostzone in each direction. We will define the index in our data storage of the point $(i,j,k)$ for the iteration and declare storage for the values that we will use to do the interpolation.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
/* Compute
* \partial_t psi6phi = -\partial_j ( \alpha \sqrt{\gamma} A^j - \beta^j psi6phi)
* (Eq 13 of http://arxiv.org/pdf/1110.4633.pdf), using Lorenz gauge.
* Note that the RHS consists of a shift advection term on psi6phi and
* a term depending on the vector potential.
* psi6phi is defined at (i+1/2,j+1/2,k+1/2), but instead of reconstructing
* to compute the RHS of \partial_t psi6phi, we instead use standard
* interpolations.
*/
// The stencil here is {-1,1},{-1,1},{-1,1} for x,y,z directions, respectively.
// Note that ALL input variables are defined at ALL gridpoints, so no
// worries about ghostzones.
#pragma omp parallel for
for(int k=1;k<Nxx_plus_2NGHOSTS2-1;k++) for(int j=1;j<Nxx_plus_2NGHOSTS1-1;j++) for(int i=1;i<Nxx_plus_2NGHOSTS0-1;i++) {
const int index=IDX3S(i,j,k);
REAL INTERP_VARS[MAXNUMINTERP][PLUS2+1][PLUS2+1][PLUS2+1];
```
<a id='interp'></a>
### Step 2.b.i: Read in quantities needed for interpolation \[Back to [top](#toc)\]
$$\label{interp}$$
We will code this one term at a time, starting with $\partial_j (\alpha \sqrt{\gamma} A^j)$. To do so, we will need to find this quantity at the cell vertex $(i+1/2,j+1/2,k+1/2)$. Since $A_x$ exists at $(i,j+1/2,k+1/2)$ it makes sense to do this by finding $(\alpha \sqrt{\gamma} A^x)$ at $(i,j+1/2,k+1/2)$ and $(i+1,j+1/2,k+1/2)$ and using those values to perform the finite differencing. (And so on, for the other components of $A_i$.)
Note also that $\alpha \sqrt{\gamma} A^j = \alpha \sqrt{\gamma} \gamma^{ij} A_i$; so, we will need to interpolate the inverse metric $\gamma^{ij}$, the square root of the metric determinant $\sqrt{\gamma}$, and the lapse function $\alpha$ to the cell edges.
So, we must prepare for all these interpolations. We will read in the values of the above specified metric gridfunctions in a small cube two gridpoints in every direction and write them to `INTERP_VARS`.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// First compute \partial_j \alpha \sqrt{\gamma} A^j (RHS of \partial_i psi6phi)
// FIXME: Would be much cheaper & easier to unstagger A_i, raise, then interpolate A^i.
// However, we keep it this way to be completely compatible with the original
// Illinois GRMHD thorn, called mhd_evolve.
//
//Step 1) j=x: Need to raise A_i, but to do that, we must have all variables at the same gridpoints:
// The goal is to compute \partial_j (\alpha \sqrt{\gamma} A^j) at (i+1/2,j+1/2,k+1/2)
// We do this by first interpolating (RHS1x) = (\alpha \sqrt{\gamma} A^x) at
// (i,j+1/2,k+1/2)and (i+1,j+1/2,k+1/2), then taking \partial_x (RHS1x) =
// [ RHS1x(i+1,j+1/2,k+1/2) - RHS1x(i,j+1/2,k+1/2) ]/dX.
// First bring gup's, psi, and alpha to (i,j+1/2,k+1/2):
int num_vars_to_interp;
int vars_to_interpolate[MAXNUMINTERP] = {GUPXXI,GUPXYI,GUPXZI,GUPYYI,GUPYZI,GUPZZI,LAPM1I,PSII,SHIFTXI,SHIFTYI,SHIFTZI};
num_vars_to_interp = 11;
// We may set interp_limits to be more general than we need.
int interp_limits[6] = {-1,1,-1,1,-1,1}; SET_INDEX_ARRAYS_NRPY_3DBLOCK(interp_limits);
// Major change here to invert the metric on the spot!
// Read in variable at interp. stencil points from main memory, store in INTERP_VARS.
for(int kk=PLUS0;kk<=PLUS1;kk++) for(int jj=PLUS0;jj<=PLUS1;jj++) for(int ii=PLUS0;ii<=PLUS1;ii++) {
// First, we will read in each component of the metric, then find the determinant.
// We write the twelfth root to psi_bssn. Then, we invert the metric and store these values.
const REAL gammaDD00 = in_vars[GXXI][index_arr_3DB[kk][jj][ii]];
const REAL gammaDD01 = in_vars[GXYI][index_arr_3DB[kk][jj][ii]];
const REAL gammaDD02 = in_vars[GXZI][index_arr_3DB[kk][jj][ii]];
const REAL gammaDD11 = in_vars[GYYI][index_arr_3DB[kk][jj][ii]];
const REAL gammaDD12 = in_vars[GYZI][index_arr_3DB[kk][jj][ii]];
const REAL gammaDD22 = in_vars[GZZI][index_arr_3DB[kk][jj][ii]];
// Generated by NRPy+:
/*
* NRPy+ Finite Difference Code Generation, Step 2 of 1: Evaluate SymPy expressions and write to main memory:
*/
const double FDPart3_0 = cbrt(gammaDD00*gammaDD11*gammaDD22 - gammaDD00*((gammaDD12)*(gammaDD12)) - ((gammaDD01)*(gammaDD01))*gammaDD22 + 2*gammaDD01*gammaDD02*gammaDD12 - ((gammaDD02)*(gammaDD02))*gammaDD11);
const double FDPart3_1 = (1.0/(FDPart3_0));
const REAL gamma_bssnDD00 = FDPart3_1*gammaDD00;
const REAL gamma_bssnDD01 = FDPart3_1*gammaDD01;
const REAL gamma_bssnDD02 = FDPart3_1*gammaDD02;
const REAL gamma_bssnDD11 = FDPart3_1*gammaDD11;
const REAL gamma_bssnDD12 = FDPart3_1*gammaDD12;
const REAL gamma_bssnDD22 = FDPart3_1*gammaDD22;
const double tmp_5 = gamma_bssnDD00*gamma_bssnDD11*gamma_bssnDD22 - gamma_bssnDD00*((gamma_bssnDD12)*(gamma_bssnDD12)) - ((gamma_bssnDD01)*(gamma_bssnDD01))*gamma_bssnDD22 + 2*gamma_bssnDD01*gamma_bssnDD02*gamma_bssnDD12 - ((gamma_bssnDD02)*(gamma_bssnDD02))*gamma_bssnDD11;
const double tmp_6 = (1.0/(tmp_5));
INTERP_VARS[GUPXXI][kk][jj][ii] = tmp_6*(gamma_bssnDD11*gamma_bssnDD22 - ((gamma_bssnDD12)*(gamma_bssnDD12)));
INTERP_VARS[GUPXYI][kk][jj][ii] = tmp_6*(-gamma_bssnDD01*gamma_bssnDD22 + gamma_bssnDD02*gamma_bssnDD12);
INTERP_VARS[GUPXZI][kk][jj][ii] = tmp_6*(gamma_bssnDD01*gamma_bssnDD12 - gamma_bssnDD02*gamma_bssnDD11);
INTERP_VARS[GUPYYI][kk][jj][ii] = tmp_6*(gamma_bssnDD00*gamma_bssnDD22 - ((gamma_bssnDD02)*(gamma_bssnDD02)));
INTERP_VARS[GUPYZI][kk][jj][ii] = tmp_6*(-gamma_bssnDD00*gamma_bssnDD12 + gamma_bssnDD01*gamma_bssnDD02);
INTERP_VARS[GUPZZI][kk][jj][ii] = tmp_6*(gamma_bssnDD00*gamma_bssnDD11 - ((gamma_bssnDD01)*(gamma_bssnDD01)));
INTERP_VARS[PSII][kk][jj][ii] = pow(FDPart3_0,1.0/4.0);
// Now, we read in the lapse function.
int whichvar=vars_to_interpolate[6];
INTERP_VARS[whichvar][kk][jj][ii] = in_vars[whichvar][index_arr_3DB[kk][jj][ii]]-1.0; // Input alpha, expect alpha-1
// Finally, we read in the shift vector into the array.
for(int ww=8;ww<num_vars_to_interp;ww++) {
int whichvar=vars_to_interpolate[ww];
INTERP_VARS[whichvar][kk][jj][ii] = in_vars[whichvar][index_arr_3DB[kk][jj][ii]];
}
}
```
While we are at it, we will also interpolate the lapse function $\alpha$ to the cell vertices.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// Next set \alpha at (i+1/2,j+1/2,k+1/2). Will come in handy when computing damping term later.
alpha_iphjphkph[index] = avg(INTERP_VARS[LAPM1I] , PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1)+1.0;
```
Here, we set up the interpolation stencils for the vector potential. These are defined by the maximum extents of the interpolations that we do below, where the bounds are explained more depth.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
//A_x needs a stencil s.t. interp_limits={0,1,-1,1,-1,1}:
for(int kk=MINUS1;kk<=PLUS1;kk++) for(int jj=MINUS1;jj<=PLUS1;jj++) for(int ii=PLUS0;ii<=PLUS1;ii++) {
INTERP_VARS[A_XI][kk][jj][ii] = in_vars[A_XI][index_arr_3DB[kk][jj][ii]]; }
//A_y needs a stencil s.t. interp_limits={-1,1,0,1,-1,1}:
for(int kk=MINUS1;kk<=PLUS1;kk++) for(int jj=PLUS0;jj<=PLUS1;jj++) for(int ii=MINUS1;ii<=PLUS1;ii++) {
INTERP_VARS[A_YI][kk][jj][ii] = in_vars[A_YI][index_arr_3DB[kk][jj][ii]]; }
//A_z needs a stencil s.t. interp_limits={-1,1,-1,1,0,1}:
for(int kk=PLUS0;kk<=PLUS1;kk++) for(int jj=MINUS1;jj<=PLUS1;jj++) for(int ii=MINUS1;ii<=PLUS1;ii++) {
INTERP_VARS[A_ZI][kk][jj][ii] = in_vars[A_ZI][index_arr_3DB[kk][jj][ii]]; }
```
<a id='raise_a'></a>
### Step 2.b.ii: Compute $\alpha \sqrt{\gamma} A^j$ \[Back to [top](#toc)\]
$$\label{raise_a}$$
Since we are starting with $A_x$, we care about the edge $(i,j+1/2,k+1/2)$ right now. As was stated above, our interpolation is an average over the nearest points, which will be $(i,j,k)$, $(i,j+1,k)$, $(i,j,k+1)$, and $(i,j+1,k+1)$ for the metric gridfunctions, which are already at cell centers.
We will also need $A_i$ at $(i,j+1/2,k+1/2)$. Because $A_x$ is already there, nothing needs to be done with it. We have $A_y(i+1/2,j,k+1/2)$, so we must average `i-1,j,k`, `i,j,k`, `i-1,j+1,k`, and `i,j+1,k`. We have $A_z(i+1/2,j+1/2,k)$, so we must average `i-1,j,k`, `i,j,k`, `i-1,j,k+1`, and `i,j,k+1`.
Then, we are free to calculate $\alpha \sqrt{\gamma} A^x = \alpha \sqrt{\gamma} (\gamma^{xx} A_x + \gamma^{xy} A_y + \gamma^{xz} A_z)$ at every gridpoint.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// FIRST DO A^X TERM (interpolate to (i,j+1/2,k+1/2) )
// \alpha \sqrt{\gamma} A^x = \alpha psi^6 A^x (RHS of \partial_i psi6phi)
// Note that gupij is \tilde{\gamma}^{ij}, so we need to multiply by \psi^{-4}.
const REAL gupxx_jphkph = avg(INTERP_VARS[GUPXXI], PLUS0,PLUS0, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL gupxy_jphkph = avg(INTERP_VARS[GUPXYI], PLUS0,PLUS0, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL gupxz_jphkph = avg(INTERP_VARS[GUPXZI], PLUS0,PLUS0, PLUS0,PLUS1, PLUS0,PLUS1);
for(int kk=PLUS0;kk<=PLUS1;kk++) for(int jj=PLUS0;jj<=PLUS1;jj++) for(int ii=PLUS0;ii<=PLUS1;ii++) {
const REAL Psi2 = INTERP_VARS[PSII][kk][jj][ii]*INTERP_VARS[PSII][kk][jj][ii];
const REAL alpha = INTERP_VARS[LAPM1I][kk][jj][ii]+1.0;
INTERP_VARS[LAPSE_PSI2I][kk][jj][ii]=alpha*Psi2;
INTERP_VARS[LAPSE_OVER_PSI6I][kk][jj][ii]=alpha/(Psi2*Psi2*Psi2);
}
const REAL lapse_Psi2_jphkph = avg(INTERP_VARS[LAPSE_PSI2I], PLUS0,PLUS0, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL A_x_jphkph = avg(INTERP_VARS[A_XI], PLUS0,PLUS0, PLUS0,PLUS0, PLUS0,PLUS0); // @ (i,j+1/2,k+1/2)
const REAL A_y_jphkph = avg(INTERP_VARS[A_YI],MINUS1,PLUS0, PLUS0,PLUS1, PLUS0,PLUS0); // @ (i+1/2,j,k+1/2)
const REAL A_z_jphkph = avg(INTERP_VARS[A_ZI],MINUS1,PLUS0, PLUS0,PLUS0, PLUS0,PLUS1); // @ (i+1/2,j+1/2,k)
alpha_sqrtg_Ax_interp[index] = lapse_Psi2_jphkph*
( gupxx_jphkph*A_x_jphkph + gupxy_jphkph*A_y_jphkph + gupxz_jphkph*A_z_jphkph );
```
As above, except we are now calculating $\alpha \sqrt{\gamma} A^y = \alpha \sqrt{\gamma} (\gamma^{yx} A_x + \gamma^{yy} A_y + \gamma^{yz} A_z)$ at every gridpoint, making the appropriate cyclic permutations.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// DO A^Y TERM (interpolate to (i+1/2,j,k+1/2) )
// \alpha \sqrt{\gamma} A^y = \alpha psi^6 A^y (RHS of \partial_i psi6phi)
// Note that gupij is \tilde{\gamma}^{ij}, so we need to multiply by \psi^{-4}.
const REAL gupxy_iphkph = avg(INTERP_VARS[GUPXYI], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1);
const REAL gupyy_iphkph = avg(INTERP_VARS[GUPYYI], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1);
const REAL gupyz_iphkph = avg(INTERP_VARS[GUPYZI], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1);
const REAL lapse_Psi2_iphkph = avg(INTERP_VARS[LAPSE_PSI2I], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1);
//REAL lapse_iphkph = avg(INTERP_VARS[LAPM1I], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1)+1.0;
//REAL psi_iphkph = avg(INTERP_VARS[PSII ], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS1);
const REAL A_x_iphkph = avg(INTERP_VARS[A_XI], PLUS0,PLUS1,MINUS1,PLUS0, PLUS0,PLUS0); // @ (i,j+1/2,k+1/2)
const REAL A_y_iphkph = avg(INTERP_VARS[A_YI], PLUS0,PLUS0, PLUS0,PLUS0, PLUS0,PLUS0); // @ (i+1/2,j,k+1/2)
const REAL A_z_iphkph = avg(INTERP_VARS[A_ZI], PLUS0,PLUS0,MINUS1,PLUS0, PLUS0,PLUS1); // @ (i+1/2,j+1/2,k)
alpha_sqrtg_Ay_interp[index] = lapse_Psi2_iphkph*
( gupxy_iphkph*A_x_iphkph + gupyy_iphkph*A_y_iphkph + gupyz_iphkph*A_z_iphkph );
```
As above, except we are now calculating $\alpha \sqrt{\gamma} A^z = \alpha \sqrt{\gamma} (\gamma^{zx} A_x + \gamma^{zy} A_y + \gamma^{zz} A_z)$ at every gridpoint, making the appropriate cyclic permutations.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// DO A^Z TERM (interpolate to (i+1/2,j+1/2,k) )
// \alpha \sqrt{\gamma} A^z = \alpha psi^6 A^z (RHS of \partial_i psi6phi)
// Note that gupij is \tilde{\gamma}^{ij}, so we need to multiply by \psi^{-4}.
const REAL gupxz_iphjph = avg(INTERP_VARS[GUPXZI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0);
const REAL gupyz_iphjph = avg(INTERP_VARS[GUPYZI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0);
const REAL gupzz_iphjph = avg(INTERP_VARS[GUPZZI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0);
//REAL lapse_iphjph = avg(INTERP_VARS[LAPM1I], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0)+1.0;
//REAL psi_iphjph = avg(INTERP_VARS[PSII ], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0);
const REAL lapse_Psi2_iphjph = avg(INTERP_VARS[LAPSE_PSI2I], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS0);
const REAL A_x_iphjph = avg(INTERP_VARS[A_XI], PLUS0,PLUS1, PLUS0,PLUS0,MINUS1,PLUS0); // @ (i,j+1/2,k+1/2)
const REAL A_y_iphjph = avg(INTERP_VARS[A_YI], PLUS0,PLUS0, PLUS0,PLUS1,MINUS1,PLUS0); // @ (i+1/2,j,k+1/2)
const REAL A_z_iphjph = avg(INTERP_VARS[A_ZI], PLUS0,PLUS0, PLUS0,PLUS0, PLUS0,PLUS0); // @ (i+1/2,j+1/2,k)
alpha_sqrtg_Az_interp[index] = lapse_Psi2_iphjph*
( gupxz_iphjph*A_x_iphjph + gupyz_iphjph*A_y_iphjph + gupzz_iphjph*A_z_iphjph );
```
<a id='a_gauge_term'></a>
### Step 2.b.iii: Compute $\alpha \Phi - \beta^j A_j$ \[Back to [top](#toc)\]
$$\label{a_gauge_term}$$
Now, we will calculate $\alpha \Phi - \beta^j A_j$ at every gridpoint for the $A_k$ evolution equation. This follows fairly similarly to what we did above, except we must interpolate all the inputs (lapse $\alpha$, shift $\beta^i$, vector potential $A_i$) to the cell vertices $(i+1/2,j+1/2,k+1/2)$. While $\psi^6\Phi$ is already sampled at cell vertices, the equation specifically calls for only $\Phi$; fortunately, we have precomputed $\alpha/\psi^6$ at cell centers.
Thus, we ultimately will be interpolating
* the shift and $\alpha/\psi^6$ from the cell centers to the cell vertices, using indices `i,j,k`, `i+1,j,k`, `i,j+1,k`, `i,j,k+1`, `i,j+1,k+1`, `i+1,j,k+1`, `i+1,j+1,k`, and `i+1,j+1,k+1`
* $A_x$ from the cell edge $(i,j+1/2,k+1/2)$ using `i,j,k` and `i+1,j,k`
* $A_y$ from the cell edge $(i+1/2,j,k+1/2)$ using `i,j,k` and `i,j+1,k`
* $A_z$ from the cell edge $(i+1/2,j+1/2,k)$ using `i,j,k` and `i,j,k+1`
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// Next set \alpha \Phi - \beta^j A_j at (i+1/2,j+1/2,k+1/2):
// We add a "L" suffix to shifti_iphjphkph to denote "LOCAL", as we set
// shifti_iphjphkph[] gridfunction below.
const REAL shiftx_iphjphkphL = avg(INTERP_VARS[SHIFTXI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL shifty_iphjphkphL = avg(INTERP_VARS[SHIFTYI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL shiftz_iphjphkphL = avg(INTERP_VARS[SHIFTZI], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1);
const REAL lapse_over_Psi6_iphjphkphL = avg(INTERP_VARS[LAPSE_OVER_PSI6I], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1);
//REAL psi_iphjphkph = avg(INTERP_VARS[PSII ], PLUS0,PLUS1, PLUS0,PLUS1, PLUS0,PLUS1);
//REAL psi2_iphjphkph= psi_iphjphkph*psi_iphjphkph;
//REAL psi6_iphjphkph= psi2_iphjphkph*psi2_iphjphkph*psi2_iphjphkph;
const REAL A_x_iphjphkph = avg(INTERP_VARS[A_XI], PLUS0,PLUS1, PLUS0,PLUS0, PLUS0,PLUS0); // @ (i,j+1/2,k+1/2)
const REAL A_y_iphjphkph = avg(INTERP_VARS[A_YI], PLUS0,PLUS0, PLUS0,PLUS1, PLUS0,PLUS0); // @ (i+1/2,j,k+1/2)
const REAL A_z_iphjphkph = avg(INTERP_VARS[A_ZI], PLUS0,PLUS0, PLUS0,PLUS0, PLUS0,PLUS1); // @ (i+1/2,j+1/2,k)
alpha_Phi_minus_betaj_A_j_iphjphkph[index] = psi6phi[index]*lapse_over_Psi6_iphjphkphL
- (shiftx_iphjphkphL*A_x_iphjphkph + shifty_iphjphkphL*A_y_iphjphkph + shiftz_iphjphkphL*A_z_iphjphkph);
```
We will also store the shift function at $(i+1/2,j+1/2,k+1/2)$ (which we have already interpolated) for $\partial_j \beta^j [\psi^6 \Phi]$, which is part of the $\psi^6 \Phi$ evolution equation.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// Finally, save shifti_iphjphkph, for \partial_j \beta^j psi6phi
shiftx_iphjphkph[index]=shiftx_iphjphkphL;
shifty_iphjphkph[index]=shifty_iphjphkphL;
shiftz_iphjphkph[index]=shiftz_iphjphkphL;
}
```
<a id='finite_difference'></a>
## Step 2.c: Take the finite difference derivatives \[Back to [top](#toc)\]
$$\label{finite_difference}$$
We restate the equations we want to solve:
* $\partial_t A_i -= \partial_i (\alpha \Phi - \beta^j A_j)$
* $\partial_t [\sqrt{\gamma} \Phi] + \partial_j (\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi]) = - \xi \alpha [\sqrt{\gamma} \Phi]$
So far, we have calculated only the operands of the differentiation operator, i.e. the parts within the parentheses as they are written above, and repeated here:
* $\alpha \Phi - \beta^j A_j$
* $\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi]$
So, we must now take the derivatives using finite-difference methods. We will thus loop over the grid interior, and define the index to which we will be writing our results.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// This loop requires two additional ghostzones in every direction. Hence the following loop definition:
#pragma omp parallel for
for(int k=NGHOSTS;k<Nxx_plus_2NGHOSTS2-NGHOSTS;k++) for(int j=NGHOSTS;j<Nxx_plus_2NGHOSTS1-NGHOSTS;j++) for(int i=NGHOSTS;i<Nxx_plus_2NGHOSTS0-NGHOSTS;i++) {
const int index = IDX3S(i,j,k);
```
<a id='a_rhs'></a>
### Step 2.c.i: Compute the $A_i$ gauge term \[Back to [top](#toc)\]
$$\label{a_rhs}$$
We start with the right hand side of $\partial_t A_i$, adding on the gauge term $-\partial_i (\alpha \Phi - \beta^j A_j)$. Recall that as we calculated the operand, we interpolated it to the cell vertices $(i+1/2,j+1/2,k+1/2)$. Thus, since $A_x$ is sampled at $(i,j+1/2,k+1/2)$, the relevant finite-difference template here is
$$-(Q_{i,j,k} - Q_{i-1,j,k})/dx = (Q_{i-1,j,k} - Q_{i,j,k})/dx$$
for a dummy quantity $Q$, and so on for the $y$ and $z$ directions.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// \partial_t A_i = [reconstructed stuff] + [gauge stuff],
// where [gauge stuff] = -\partial_i (\alpha \Phi - \beta^j A_j)
const REAL alpha_Phi_minus_betaj_A_j_iphjphkphL = alpha_Phi_minus_betaj_A_j_iphjphkph[index];
// - partial_i -> - (A_{i} - A_{i-1})/dX = (A_{i-1} - A_{i})/dX, for Ax
Ax_rhs[index] += invdx0*(alpha_Phi_minus_betaj_A_j_iphjphkph[IDX3S(i-1,j,k)] - alpha_Phi_minus_betaj_A_j_iphjphkphL);
Ay_rhs[index] += invdx1*(alpha_Phi_minus_betaj_A_j_iphjphkph[IDX3S(i,j-1,k)] - alpha_Phi_minus_betaj_A_j_iphjphkphL);
Az_rhs[index] += invdx2*(alpha_Phi_minus_betaj_A_j_iphjphkph[IDX3S(i,j,k-1)] - alpha_Phi_minus_betaj_A_j_iphjphkphL);
```
<a id='phi_rhs'></a>
### Step 2.c.ii: Compute the $[\psi^6 \Phi]$ gauge term \[Back to [top](#toc)\]
$$\label{phi_rhs}$$
The operand of the gradient operator in the $\partial_t [\psi^6 \Phi]$ equation is the sum of two terms; we will first compute the shift advection term $\partial_j (\beta^j \psi^6 \Phi)$.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// \partial_t psi6phi = [shift advection term] + \partial_j (\alpha \sqrt{\gamma} A^j)
// Here we compute [shift advection term] = \partial_j (\beta^j psi6phi)
// Cache misses are likely more expensive than branch mispredictions here,
// which is why we use if() statements and array lookups inside the if()'s.
REAL psi6phi_rhsL=0.0;
const REAL psi6phiL=psi6phi[index];
const REAL shiftx_iphjphkphL=shiftx_iphjphkph[index];
const REAL shifty_iphjphkphL=shifty_iphjphkph[index];
const REAL shiftz_iphjphkphL=shiftz_iphjphkph[index];
// \partial_x (\beta^x psi6phi) :
if(shiftx_iphjphkphL < 0.0) {
psi6phi_rhsL+=0.5*invdx0*(+ shiftx_iphjphkph[IDX3S(i-2,j,k)]*psi6phi[IDX3S(i-2,j,k)]
-4.0*shiftx_iphjphkph[IDX3S(i-1,j,k)]*psi6phi[IDX3S(i-1,j,k)]
+3.0*shiftx_iphjphkphL* psi6phiL);
} else {
psi6phi_rhsL+=0.5*invdx0*(- shiftx_iphjphkph[IDX3S(i+2,j,k)]*psi6phi[IDX3S(i+2,j,k)]
+4.0*shiftx_iphjphkph[IDX3S(i+1,j,k)]*psi6phi[IDX3S(i+1,j,k)]
-3.0*shiftx_iphjphkphL* psi6phiL);
}
// \partial_y (\beta^y psi6phi) :
if(shifty_iphjphkphL < 0.0) {
psi6phi_rhsL+=0.5*invdx1*(+ shifty_iphjphkph[IDX3S(i,j-2,k)]*psi6phi[IDX3S(i,j-2,k)]
-4.0*shifty_iphjphkph[IDX3S(i,j-1,k)]*psi6phi[IDX3S(i,j-1,k)]
+3.0*shifty_iphjphkphL* psi6phiL);
} else {
psi6phi_rhsL+=0.5*invdx1*(- shifty_iphjphkph[IDX3S(i,j+2,k)]*psi6phi[IDX3S(i,j+2,k)]
+4.0*shifty_iphjphkph[IDX3S(i,j+1,k)]*psi6phi[IDX3S(i,j+1,k)]
-3.0*shifty_iphjphkphL* psi6phiL);
}
// \partial_z (\beta^z psi6phi) :
if(shiftz_iphjphkphL < 0.0) {
psi6phi_rhsL+=0.5*invdx2*(+ shiftz_iphjphkph[IDX3S(i,j,k-2)]*psi6phi[IDX3S(i,j,k-2)]
-4.0*shiftz_iphjphkph[IDX3S(i,j,k-1)]*psi6phi[IDX3S(i,j,k-1)]
+3.0*shiftz_iphjphkphL* psi6phiL);
} else {
psi6phi_rhsL+=0.5*invdx2*(- shiftz_iphjphkph[IDX3S(i,j,k+2)]*psi6phi[IDX3S(i,j,k+2)]
+4.0*shiftz_iphjphkph[IDX3S(i,j,k+1)]*psi6phi[IDX3S(i,j,k+1)]
-3.0*shiftz_iphjphkphL* psi6phiL);
}
```
The operand of the gradient operator in the $\partial_t [\psi^6 \Phi]$ equation is the sum of two terms; we will add the term $-\partial_j (\alpha \sqrt{\gamma} A^j)$. Recall that we computed $\alpha \sqrt{\gamma} A^x$ at $(i,j+1/2,k+1/2)$; thus, the appropriate finite-difference template here is
$$-(Q_{i+1,j,k} - Q_{i,j,k})/dx = (Q_{i,j,k} - Q_{i+1,j,k})/dx$$
for a dummy quantity $Q$, and so on for the $y$ and $z$ directions.
Finally, we will add the damping factor to $\partial_t [\psi^6 \Phi]$, which is given as $\xi \alpha \psi^6 \Phi$, where $\xi$ is some scalar damping parameter. Since we interpolated the lapse $\alpha$ to cell vertices earlier, everything is exactly where we need it!
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
// Next we add \partial_j (\alpha \sqrt{\gamma} A^j) to \partial_t psi6phi:
psi6phi_rhsL+=invdx0*(alpha_sqrtg_Ax_interp[index] - alpha_sqrtg_Ax_interp[IDX3S(i+1,j,k)])
+ invdx1*(alpha_sqrtg_Ay_interp[index] - alpha_sqrtg_Ay_interp[IDX3S(i,j+1,k)])
+ invdx2*(alpha_sqrtg_Az_interp[index] - alpha_sqrtg_Az_interp[IDX3S(i,j,k+1)]);
// *GENERALIZED* LORENZ GAUGE:
// Finally, add damping factor to \partial_t psi6phi
//subtract lambda * alpha psi^6 Phi
psi6phi_rhsL+=-xi_damping*alpha_iphjphkph[index]*psi6phiL;
psi6phi_rhs[index] = psi6phi_rhsL;
}
}
```
<a id='interp_func'></a>
## Step 2.d: The interpolation function \[Back to [top](#toc)\]
$$\label{interp_func}$$
Here, we give the function definition for our interpolator, `avg`. It takes the arithmetic mean of the points of the gridfunction `f` specified by `imin`, `imax`, `jmin`, `jmax`, `kmin`, and `kmax`.
```
%%writefile -a $Ccodesdir/Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h
static inline REAL avg(const REAL f[PLUS2+1][PLUS2+1][PLUS2+1],const int imin,const int imax, const int jmin,const int jmax, const int kmin,const int kmax) {
REAL retval=0.0,num_in_sum=0.0;
for(int kk=kmin;kk<=kmax;kk++) for(int jj=jmin;jj<=jmax;jj++) for(int ii=imin;ii<=imax;ii++) {
retval+=f[kk][jj][ii]; num_in_sum++;
}
return retval/num_in_sum;
}
```
<a id='code_validation'></a>
# Step 3: Code Validation \[Back to [top](#toc)\]
$$\label{code_validation}$$
To validate the code in this tutorial we check for agreement between the files
1. that were written in this tutorial and
1. those that are generated by the python module
```
# Define the directory that we wish to validate against:
valdir = "GiRaFFE_NRPy/GiRaFFE_Ccode_library/RHSs/"
import GiRaFFE_NRPy.GiRaFFE_NRPy_staggered_Source_Terms as source
source.GiRaFFE_NRPy_Source_Terms(valdir)
import difflib
import sys
print("Printing difference between original C code and this code...")
# Open the files to compare
files = ["Lorenz_psi6phi_rhs__add_gauge_terms_to_A_i_rhs.h"]
for file in files:
print("Checking file " + file)
with open(os.path.join(valdir,file)) as file1, open(os.path.join(Ccodesdir,file)) as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
num_diffs = 0
for line in difflib.unified_diff(file1_lines, file2_lines, fromfile=os.path.join(valdir+file), tofile=os.path.join(Ccodesdir+file)):
sys.stdout.writelines(line)
num_diffs = num_diffs + 1
if num_diffs == 0:
print("No difference. TEST PASSED!")
else:
print("ERROR: Disagreement found with .py file. See differences above.")
sys.exit(1)
```
<a id='latex_pdf_output'></a>
# Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-GiRaFFE_NRPy_staggered-Source_Terms.pdf](Tutorial-GiRaFFE_NRPy_staggered-Source_Terms.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-GiRaFFE_NRPy_staggered-Source_Terms",location_of_template_file=os.path.join(".."))
```
| github_jupyter |
```
import nltk
import matplotlib.pyplot as plt
import collections as c
import operator
nltk.download('book')
from nltk.book import *
texts()
sents()
text3
sent6
len(sent1)
len(text1)
len(set(text1))
type(text1)
list(set(text1))[:10]
dist = c.OrderedDict(FreqDist(text1))
sorted_dist = sorted(dist.items(), key = operator.itemgetter(1), reverse = True)
vocab1 = dist.keys()
vocab1
dist['late']
freqwords = [w for w in vocab1 if len(w)>5 and dist[w] > 100]
len(freqwords)
freqwords
input1 = "Madhuri Dixit (born 15 May 1967), also known by her married name Madhuri Dixit Nene,[1] is an Indian actress, producer, and television personality. One of the most popular and highest-paid Hindi film actresses in the late 1980s, 1990s and early 2000s,[2][3] she has been praised by critics for her acting and dancing skills.[4] She is the recipient of numerous accolades, including six Filmfare Awards. In 2008, the Government of India awarded her with the Padma Shri, the fourth highest civilian honour of the country.Dixit made her acting debut with a leading role in the 1984 drama Abodh. This was followed by a series of commercial failures, which led to a setback in her career. The 1988 blockbuster Tezaab marked a turning point for Dixit, establishing her as a leading actress of Bollywood. She achieved further success with starring roles in several top-grossing productions, including the crime dramas Ram Lakhan (1989) and Parinda (1989), the action thrillers Tridev (1989), Thanedaar (1990) and Khalnayak (1993), the action comedy Kishen Kanhaiya (1990), and the romantic dramas Prem Pratigyaa (1989), Saajan (1991) and Raja (1995).Her portrayal of a supercilious girl in the romantic drama Dil (1990), an incorrigible woman in the social drama Beta (1992), a playful conservative in the romantic comedy Hum Aapke Hain Koun..! (1994) "
input1
#normalization of words - convert into same case. split the words
words1 = input1.lower().split(" ")
words1 = [w for w in words1 if len(w) > 3]
list(words1)
len(set(words1))
#stemming of words - remove the prefix/suffix and convert them into same root words
porter = nltk.PorterStemmer()
stemmed_words1 = [porter.stem(t) for t in words1]
len(set(stemmed_words1))
#lemmatization - make the stemmed words meaningful
lemma = nltk.stem.WordNetLemmatizer()
lemma_words1 = [lemma.lemmatize(t) for t in words1]
len(set(lemma_words1))
[w for w in set(words1) if w not in set(stemmed_words1)]
#better word tokenization with nltk than a simple split
words2 = nltk.word_tokenize(input1)
words2
print(len(words1), len(words2))
#sentence splitting using NLTK
sentences = nltk.sent_tokenize(input1)
sentences
```
| github_jupyter |
# Correlation between Detected Breeding Sites and Larval Survey
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from plotly import tools
from plotly.graph_objs import *
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
import plotly.graph_objs as go
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os, graphviz
from sklearn import *
from copy import deepcopy
from scipy.stats.stats import pearsonr, spearmanr
from shapely.geometry import Polygon
from collections import Counter
loo = model_selection.LeaveOneOut()
month = ['Jan','Feb','Mar','Apr','May','Jun','Jul','Aug','Sep','Oct','Nov','Dec']
categories = np.array(['bin','bowl','bucket','cup','jar','pottedplant','tire','vase']).reshape(-1,1)
import visualizer
import data_loader
df_loader = data_loader.df_loader()
df_survey = df_loader.load_survey()
df_filtered = df_loader.load_filterd('bi')
df_area = df_loader.load_area()
df_detect = df_loader.load_detect()
df_population = df_loader.load_population()
df_dengue_cases = df_loader.load_cases()
df_survey.head()
print('Total:', len(df_survey))
df_filtered.head()
len(df_filtered)
df_detect.head()
print('Total:', len(df_detect))
df_area.head()
len(df_area)
df_population.head()
len(df_population)
```
## Visualize average breteau index value for each month
```
key, val = [], []
for m in range(12):
key.append(month[m])
val.append(round(df_filtered[df_filtered.index.month == m+1]['bi'].mean(),0))
trace_bar = go.Bar(
x = key,
y = val,
text = val,
textposition = 'auto',
marker=dict(
color='rgb(158,202,225)',
line=dict(
color='rgb(8,48,107)',
width=1.5),
),
opacity=0.8
)
layout = go.Layout(
title='Average BI for each Month',
height=550,
width=750,
yaxis= dict(title='avg bi'),
xaxis= dict(title='month')
)
fig = go.Figure(data=[trace_bar], layout=layout)
iplot(fig)
```
## Visualize number of detected breeding sites (containers) for each month
```
key, val = [], []
for m in range(12):
key.append(month[m])
val.append(round(df_detect[df_detect.index.month == m+1]['total'].mean(),0))
trace_bar = go.Bar(
x = key,
y = val,
text = val,
textposition = 'auto',
marker=dict(
color='rgb(158,202,225)',
line=dict(
color='rgb(8,48,107)',
width=1.5),
),
opacity=0.8
)
layout = go.Layout(
title='Number of detected breeding sites for each Month',
height=550,
width=750,
yaxis= dict(title='Number of containers'),
xaxis= dict(title='month')
)
fig = go.Figure(data=[trace_bar], layout=layout)
iplot(fig)
# trace = go.Scatter(
# x = df_area['area'], y = df_population['area'], mode = 'markers',
# marker = dict(size = 12, opacity = 0.3)
# )
# iplot(go.Figure(data=[trace]))
```
# Correlation
Perform correlation between number of detected containers and breteau index
## Filter outliers in each class
```
# df_category_stat = []
# for column in np.squeeze(categories):
# mean_det, std_det = df_detect[column].mean(), df_detect[column].std()
# df_category_stat.append([column, mean_det, std_det])
# df_category_stat = pd.DataFrame.from_records(df_category_stat)
# df_category_stat.columns = ['class','mean','std']
# df_category_stat = df_category_stat.set_index('class')
# df_category_stat
```
## Regression Models
```
def plot_regression(regr, name):
regr.fit(X, y)
try: coef = regr.coef_.reshape(-1,1)
except: pass
try: coef = regr.feature_importances_.reshape(-1,1)
except: pass
# df_coef = pd.DataFrame.from_records(np.concatenate((features_name, coef), axis=1))
# df_coef.columns = ['class', 'coef']
# df_coef['coef'] = df_coef['coef'].astype(float).round(4)
y_pred, y_true = [], []
for train_index, test_index in loo.split(X):
X_train, X_test = X[train_index], X[test_index]
Y_train, Y_test = y[train_index], y[test_index]
_=regr.fit(X_train, Y_train)
pred = regr.predict(X_test)
y_pred.append(np.squeeze(pred))
y_true.append(np.squeeze(Y_test))
y_pred = np.array(y_pred)
y_true = np.array(y_true)
print('R-squared:', metrics.r2_score(y_true, y_pred))
print('Person:', pearsonr(y_true, y_pred))
print(spearmanr(y_true, y_pred),'\n')
trace_1 = go.Scatter(
x = y_true, y = y_pred, mode = 'markers', name='Scatter',
marker = dict(size = 12, opacity = 0.5)
)
xs = np.array(y_true)
ys = np.array(y_pred)
regr = linear_model.LinearRegression()
regr.fit(xs.reshape(-1, 1), ys.reshape(-1, 1))
ys_pred = regr.predict(xs.reshape(-1, 1))
trace_2 = go.Scatter(
x = xs, y = np.squeeze(ys_pred), name='Regression',
mode = 'lines', line = dict(width = 4)
)
# coef = np.squeeze(coef)
# coef = 100.0 * (coef / coef.max())
# sorted_idx = np.argsort(coef)[::-1]
# pos = np.arange(sorted_idx.shape[0]) + .5
# trace_3 = go.Bar(
# x = np.squeeze(features_name[sorted_idx]),
# y = coef[sorted_idx],
# name='Variable Importance',
# marker=dict(
# color='rgb(158,202,225)',
# line=dict(
# color='rgb(8,48,107)',
# width=1.5),
# ),
# opacity=0.8
# )
fig = tools.make_subplots(
rows=1, cols=1,
# subplot_titles=(
# 'Correlation Subplot',
# 'Variable Importance Subplot',
# ),
vertical_spacing = 0.15
)
fig.append_trace(trace_1, 1, 1)
fig.append_trace(trace_2, 1, 1)
# fig.append_trace(trace_3, 1, 1)
fig['layout'].update(width=650,title=name, font=dict(size=16))
fig['layout']['xaxis1'].update(title='Breteau index')
# fig['layout']['xaxis1'].update(title='Breeding site')
fig['layout']['yaxis1'].update(title='Predicted')
# fig['layout']['yaxis1'].update(title='Relative Importance')
iplot(fig)
# return df_coef
def plot_one_regression(regr, name):
regr.fit(X, y)
y_pred, y_true = [], []
for train_index, test_index in loo.split(X):
X_train, X_test = X[train_index], X[test_index]
Y_train, Y_test = y[train_index], y[test_index]
_=regr.fit(X_train, Y_train)
pred = regr.predict(X_test)
y_pred.append(np.squeeze(pred))
y_true.append(np.squeeze(Y_test))
y_pred = np.array(y_pred)
y_true = np.array(y_true)
r2 = round(metrics.r2_score(y_true, y_pred),4)
pearson = round(pearsonr(y_true, y_pred)[0],4)
spearman = round(spearmanr(y_true, y_pred)[0],4)
print('R-squared:', metrics.r2_score(y_true, y_pred))
print('Person:', pearsonr(y_true, y_pred))
print(spearmanr(y_true, y_pred),'\n')
trace_1 = go.Scatter(
x = y_true, y = y_pred, mode = 'markers', name='Scatter',
marker = dict(size = 12, opacity = 0.5)
)
xs = np.array(y_true)
ys = np.array(y_pred)
regr = linear_model.LinearRegression()
regr.fit(xs.reshape(-1, 1), ys.reshape(-1, 1))
ys_pred = regr.predict(xs.reshape(-1, 1))
trace_2 = go.Scatter(
x = xs, y = np.squeeze(ys_pred), name='Regression',
mode = 'lines', line = dict(width = 4)
)
fig = tools.make_subplots(
rows=1, cols=1
)
fig.append_trace(trace_1, 1, 1)
fig.append_trace(trace_2, 1, 1)
name += 'R-squared: ' + str(r2) + \
', Pearson: ' + str(pearson) + \
', Spearman: ' + str(spearman)
fig['layout'].update(width=650,title=name, font=dict(size=16))
fig['layout']['xaxis1'].update(title='Breteau index')
fig['layout']['yaxis1'].update(title='Predicted')
iplot(fig)
```
## Selecte Features Used
```
# features_name = np.concatenate((categories,[['month']]), axis=0)
# features_name = np.concatenate((categories,[['month'], ['area'], ['popluation']]), axis=0)
# features_name = np.array([['bucket'], ['jar'], ['pottedplant'], ['month'], ['area'], ['popluation']])
# features_name = np.array([['bucket'], ['jar'], ['pottedplant']])
# features_name = np.array([['bucket'], ['jar'], ['pottedplant'], ['month']])
features_name = np.array([['bin'], ['bowl'], ['bucket'], ['jar'], ['pottedplant'], ['tire']])
# features_name = deepcopy(categories)
features_name
features_name.shape
df_detect['brd_total'] = df_detect['total']-df_detect['cup']-df_detect['bowl']
df_detect.head()
x_train, y_train = [], []
xs, ys = [], []
column = 'total'
mean_det, std_det = df_detect[column].mean(), df_detect[column].std()
subdist_list = df_survey['subdist'].unique()
for subdist in subdist_list:
detect = round(df_detect.loc[df_detect['subdist'] == subdist][column].mean(),2)
area = round(df_area.loc[df_area['subdist'] == subdist]['area'].mean(),2)
population = round(df_population.loc[df_population['subdist'] == subdist]['population'].mean(),2)
n_villages = round(df_population.loc[df_population['subdist'] == subdist]['n_villages'].mean(),2)
survey = round(df_filtered.loc[(df_filtered['subdist'] == subdist)
# & (df_filtered.index.month.isin([6,7,8,9,10,11]))
]['bi'].mean(), 2)
if np.isnan(detect) or np.isnan(survey): continue
if detect > mean_det+1*std_det or detect < mean_det-1*std_det: continue
xs.append(survey)
ys.append((detect)/(area))
x = df_detect.loc[df_detect['subdist'] == subdist].copy()
# x = x[['bin','bowl','bucket','cup','jar','pottedplant','tire','vase']].copy()
# x = x[['bin','bowl','bucket', 'jar','pottedplant','tire']].copy()
x = x[['bucket','jar','pottedplant']].copy()
month = df_detect.loc[df_detect['subdist'] == subdist].index.month[0]
y_dengue_season = 1 if month in [6,7,8,9,10] else 0
n_dengue_season = 0 if month in [6,7,8,9,10] else 1
features = list(np.squeeze(x.values)/area) + [month]
# features = list(np.squeeze(x.values))
# features = np.array(detect/area)
x_train.append(np.array(features))
y_train.append(survey)
X = np.array(x_train)
y = np.array(y_train)
print('X_train.shape:', X.shape)
print('\nR-squared:', metrics.r2_score(xs, ys))
print('Person:', pearsonr(xs, ys))
print(spearmanr(xs, ys),'\n')
trace = go.Scatter(
x = xs, y = ys, mode = 'markers', name='Subdistrict',
marker = dict(size = 15, opacity = 0.4)
)
xs = np.array(xs)
ys = np.array(ys)
regr = linear_model.LinearRegression()
regr.fit(xs.reshape(-1, 1), ys.reshape(-1, 1))
ys_pred = regr.predict(xs.reshape(-1, 1))
trace_2 = go.Scatter(
x = xs, y = np.squeeze(ys_pred), mode = 'lines', line = dict(width = 4), name='Regression'
)
layout = dict(
title = 'Entire year (53 data points): Linear Regression<br>' \
'Pearson: 0.350, Spearman: 0.358',
width=650,
xaxis = dict(title = 'Breteau index'),
yaxis = dict(title = 'Total # of detected containers<br>divided by area'),
font=dict(size=16)
)
iplot(go.Figure(data=[trace,trace_2], layout=layout))
regr.fit(ys.reshape(-1, 1), xs.reshape(-1, 1))
pred = np.squeeze(regr.predict(ys.reshape(-1, 1)))
print('\nR-squared:', metrics.r2_score(xs, pred))
print('Person:', pearsonr(xs, pred))
print(spearmanr(xs, pred),'\n')
# parameter_grid_gb = {
# 'max_depth': [3, 4, 5, 6, 7, 8],
# 'max_features': [2, 3, 4],
# 'subsample': [0.6, 0.8, 1],
# 'learning_rate':[0.03, 0.05, 0.1]
# }
# parameter_grid_tree = {
# 'max_depth': [3, 4, 5, 6, 7, 8],
# 'max_features': [2, 3, 4],
# }
# parameter_grid_svr = {
# 'kernel': ['linear','poly','rbf'],
# 'degree': [1,2,3,4,5,6]
# }
# parameter_grid_ada = {
# 'base_estimator': [svr, dt],
# 'n_estimators': [5, 10, 15, 20, 25],
# 'loss': ['linear', 'square', 'exponential'],
# 'learning_rate':[0.1]
# }
# grid_search = model_selection.GridSearchCV(
# estimator=ensemble.GradientBoostingRegressor(),
# param_grid=parameter_grid_gb,
# cv=loo,
# n_jobs=8
# )
# _=grid_search.fit(X, y)
# grid_search.best_score_
# grid_search.best_params_
# grid_search = model_selection.GridSearchCV(
# estimator=ensemble.RandomForestRegressor(),
# param_grid=parameter_grid_tree,
# cv=loo,
# n_jobs=1
# )
# _=grid_search.fit(X, y)
# grid_search.best_score_
# grid_search.best_params_
# grid_search = model_selection.GridSearchCV(estimator=svm.SVR(),
# param_grid=parameter_grid_svr,
# cv=10,
# n_jobs=8)
# _=grid_search.fit(X, y)
# grid_search.best_score_
# grid_search.best_params_
X = X.reshape(-1,1)
X[0]
X.shape
svr = svm.SVR(kernel='poly', degree=3)
rf = ensemble.RandomForestRegressor(max_depth=3, max_features=3)
dt = tree.DecisionTreeRegressor(max_depth=3, max_features=3)
gb = ensemble.GradientBoostingRegressor(learning_rate=0.00/9, max_depth=3, max_features=3, subsample=1)
linear = linear_model.LinearRegression()
bayes = linear_model.BayesianRidge()
knn = neighbors.KNeighborsRegressor()
bag_rf = ensemble.BaggingRegressor(rf)
bag_svr = ensemble.BaggingRegressor(svr)
ada = ensemble.AdaBoostRegressor()
ada_svr = ensemble.AdaBoostRegressor(svm.NuSVR(kernel='poly', degree=3, tol=12.3, gamma=0.28), learning_rate=0.001, loss='linear')
ada_dt = ensemble.AdaBoostRegressor(dt, learning_rate=0.03, loss='linear')
ada_rf = ensemble.AdaBoostRegressor(rf, learning_rate=0.03, loss='linear')
regrs = [
# [linear, 'Linear Regression'],
# [svm.NuSVR(kernel='poly', degree=3, tol=12.3, gamma=0.3), 'NuSVR'],
# [svm.SVR(kernel='poly', degree=3, tol=1), 'SVR'],
# [bayes, 'Bayesian Ridge'],
# [rf, 'Random Forest'],
# [dt, 'Decision Tree'],
# [gb, 'Gradient Boosting'],
# [ada_svr, 'Ada SVR'],
# [ada_dt, 'Ada DT'],
# [ada_rf, 'Ada RF'],
# [knn, 'KNeighbors'],
# [bag_rf, 'Bagging RF'],
# [bag_svr, 'Bagging SVR']
]
regrs = [
# [linear, 'Linear Regression'],
[svm.NuSVR(kernel='poly', degree=3, tol=12.3, gamma=0.28), 'NuSVR'],
# [svm.SVR(kernel='poly', degree=3, tol=1), 'SVR'],
# [bayes, 'Bayesian Ridge'],
# [rf, 'Random Forest'],
# [dt, 'Decision Tree'],
# [gb, 'Gradient Boosting'],
# [ada_svr, 'Ada SVR'],
]
df_selection = []
for k in range(1):
df_compare = []
for regr, name in regrs:
y_pred, y_true = [], []
for train_index, test_index in loo.split(X):
X_train, X_test = X[train_index], X[test_index]
Y_train, Y_test = y[train_index], y[test_index]
_=regr.fit(X_train, Y_train)
pred = regr.predict(X_test)
y_true.append(np.squeeze(Y_test))
y_pred.append(np.squeeze(pred))
y_true = np.array(y_true)
y_pred = np.array(y_pred)
df_compare.append([
name+'-'+str(k+1),
metrics.r2_score(y_true, y_pred),
pearsonr(y_true, y_pred)[0],
spearmanr(y_true, y_pred)[0]
])
df_compare = pd.DataFrame.from_records(df_compare)
df_compare.columns = ['Model','R-squared','Pearson','Spearman']
df_compare = df_compare.set_index('Model')
df_compare = df_compare.round(4)
df_selection.append(df_compare)
df_selection = pd.concat(df_selection, axis=0)
tmp = pd.DataFrame([[df_selection['R-squared'].mean(),
df_selection['Pearson'].mean(),
df_selection['Spearman'].mean()]])
tmp.columns = ['R-squared','Pearson','Spearman']
tmp.index = ['Average']
df_selection = df_selection.append(tmp)
df_selection
k = 0
plot_one_regression(
regrs[k][0],
'Entire year ('+str(X.shape[0])+' data points): '+regrs[k][1]+'<br>'
)
visualizer.plot_importance(regrs[2][0], regrs[2][1], X, y, loo, features_name)
```
| github_jupyter |
```
import re
import lxml
import nltk
import pandas as pd
import pymongo
from pymongo import MongoClient
from bs4 import BeautifulSoup
from nltk.tokenize import word_tokenize
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import CountVectorizer
import seaborn as sns
import csv
from collections import Counter
import spacy
nlp = spacy.load("en_core_web_sm")
```
## Returns Data Frame for given Flair
```
def flairToDataFrame(flair, flairList,combinedList,posts):
start = -1
end = -1
for i in range(len(flairList)):
if (flair==flairList[i]):
if start == -1:
start = i
else:
if start != -1:
end = i
break
return posts["combined"][start:end]
```
## Returns Top 10 words from Corpus
```
def topWords(corpus):
vec = CountVectorizer(ngram_range=(1, 1)).fit(corpus)
bag_of_words = vec.transform(corpus)
sum_words = bag_of_words.sum(axis=0)
words_freq = [(word, sum_words[0, idx])
for word, idx in vec.vocabulary_.items()]
words_freq =sorted(words_freq, key = lambda x: x[1], reverse=True)
print(words_freq[:10])
return words_freq[:10]
```
## Performs Name-Entity Analysis for Corpus
```
def ner(text): #returns entity for specific words in text
doc=nlp(text)
return [X.label_ for X in doc.ents]
def classifyNamedEntity(data): #returns count for each entity in corpus
ent=data.apply(lambda x : ner((x)))
ent=[x for sub in ent for x in sub]
counter=Counter(ent)
count=counter.most_common()
print(count)
return count
```
## Fetch Cleaned Data and Separating Data by Flair
```
posts = pd.read_csv('../data/cleansedData300.csv')
posts["combined"] = posts.title.astype("str")+" "+posts.textBody.astype("str")+" "+posts.comments.astype("str")
combinedList = posts["combined"].values.tolist()
flairList = posts["flair"].values.tolist()
flairs= ["Scheduled","Politics","Photography","Policy/Economy","AskIndia","Sports",
"Non-Political","Science/Technology","Food","Business/Finance","Coronavirus"]
scheduledCleaned = flairToDataFrame(flairs[0], flairList,combinedList,posts)
politicsCleaned = flairToDataFrame(flairs[1], flairList,combinedList,posts)
photographyCleaned= flairToDataFrame(flairs[2], flairList,combinedList,posts)
policyEconomyCleaned = flairToDataFrame(flairs[3], flairList,combinedList,posts)
askIndiaCleaned = flairToDataFrame(flairs[4], flairList,combinedList,posts)
sportsCleaned = flairToDataFrame(flairs[5], flairList,combinedList,posts)
nonPoliticalCleaned = flairToDataFrame(flairs[6], flairList,combinedList,posts)
scienceTechCleaned = flairToDataFrame(flairs[7], flairList,combinedList,posts)
foodCleaned = flairToDataFrame(flairs[8], flairList,combinedList,posts)
buisnessFinanceCleaned = flairToDataFrame(flairs[9], flairList,combinedList,posts)
coronavirusCleaned = flairToDataFrame(flairs[10], flairList,combinedList,posts)
```
## Observing Clean Data
```
posts.head(50)
```
## Plotting Top 10 words in each Flair Data
```
scheduledTopWords=topWords(scheduledCleaned)[:10]
x,y=map(list,zip(*scheduledTopWords))
sns.barplot(x=y,y=x)
politicsTopWords=topWords(politicsCleaned)[:10]
x,y=map(list,zip(*politicsTopWords))
sns.barplot(x=y,y=x)
photographyTopWords=topWords(photographyCleaned)[:10]
x,y=map(list,zip(*photographyTopWords))
sns.barplot(x=y,y=x)
policyEconomyTopWords=topWords(policyEconomyCleaned)[:10]
x,y=map(list,zip(*policyEconomyTopWords))
sns.barplot(x=y,y=x)
askIndiaTopWords=topWords(askIndiaCleaned)[:10]
x,y=map(list,zip(*askIndiaTopWords))
sns.barplot(x=y,y=x)
sportsTopWords=topWords(sportsCleaned)[:10]
x,y=map(list,zip(*sportsTopWords))
sns.barplot(x=y,y=x)
nonPoliticalTopWords=topWords(nonPoliticalCleaned)[:10]
x,y=map(list,zip(*nonPoliticalTopWords))
sns.barplot(x=y,y=x)
scienceTechTopWords=topWords(scienceTechCleaned)[:10]
x,y=map(list,zip(*scienceTechTopWords))
sns.barplot(x=y,y=x)
foodTopWords=topWords(foodCleaned)[:10]
x,y=map(list,zip(*foodTopWords))
sns.barplot(x=y,y=x)
buisnessFinanceTopWords=topWords(buisnessFinanceCleaned)[:10]
x,y=map(list,zip(*buisnessFinanceTopWords))
sns.barplot(x=y,y=x)
coronavirusTopWords=topWords(coronavirusCleaned)[:10]
x,y=map(list,zip(*coronavirusTopWords))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity((scheduledCleaned))))
print(x,y)
sns.barplot(x=y,y=x)
```
## Top Entities in Each Flair Data
```
x,y=map(list,zip(*classifyNamedEntity(politicsCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(photographyCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(policyEconomyCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(askIndiaCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(sportsCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(nonPoliticalCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(scienceTechCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(foodCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(buisnessFinanceCleaned)))
sns.barplot(x=y,y=x)
x,y=map(list,zip(*classifyNamedEntity(coronavirusCleaned)))
sns.barplot(x=y,y=x)
```
| github_jupyter |
### Machine Learning for Engineers: [Computer Vision Introduction](https://www.apmonitor.com/pds/index.php/Main/ComputerVisionIntro)
- [Computer Vision Introduction](https://www.apmonitor.com/pds/index.php/Main/ComputerVisionIntro)
- Source Blocks: 13
- Description: Computer vision is how computers automate tasks that mimic human response to visual information. Computers gain high-level understanding and take actions from digital images or videos.
- [Course Overview](https://apmonitor.com/pds)
- [Course Schedule](https://apmonitor.com/pds/index.php/Main/CourseSchedule)
<img width=400px align=left src='https://apmonitor.com/pds/uploads/Main/computer_vision_opencv.png'>
```
pip install opencv-python
import urllib.request
# download image
url = 'http://apmonitor.com/pds/uploads/Main/students_walking.jpg'
urllib.request.urlretrieve(url, 'students_walking.jpg')
import cv2 as cv
im = cv.imread('students_walking.jpg')
h,w,c = im.shape
im1 = cv.resize(im,(300,200))
im2 = cv.resize(im,None,fx=0.5,fy=0.5)
width = 300 # new width
h,w,c = im.shape # get image size
scale = width/w # scaling factor
height = int(h * scale)
dim = (width, height)
im3 = cv.resize(im,dim)
import matplotlib.pyplot as plt
im2 = plt.imread('students_walking.jpg')
plt.imshow(im2)
plt.imshow(im[:,:,[2,1,0]])
name='image'
cv.imshow(name,im)
cv.namedWindow(name, cv.WINDOW_AUTOSIZE)
cv.waitKey(0) # waits for key press
cv.destroyAllWindows()
import cv2 as cv
import matplotlib.pyplot as plt
im = cv.imread('students_walking.jpg')
im2 = plt.imread('students_walking.jpg')
plt.subplot(1,2,1)
plt.imshow(im)
plt.title('BGR (OpenCV)')
plt.subplot(1,2,2)
plt.imshow(im2)
plt.title('RGB (Matplotlib)')
cv.imwrite('students.png',im)
import cv2 as cv
import time
# Get camera Object
camera = cv.VideoCapture(0)
w = int(camera.get(cv.CAP_PROP_FRAME_WIDTH))
h = int(camera.get(cv.CAP_PROP_FRAME_HEIGHT))
# Write video.avi
out = cv.VideoWriter('video.avi', \
cv.VideoWriter_fourcc(*'XVID'), \
25, (w,h))
# Create Window to video frames
WindowName = 'View'
cv.namedWindow(WindowName, cv.WINDOW_AUTOSIZE)
# Save and view 5 second video
start = time.time()
while time.time()-start<=5.0:
ret0, frame = camera.read()
cv.imshow(WindowName, frame)
out.write(frame)
if cv.waitKey(1) & 0xFF == ord('q'):
break
cv.imwrite('frame.jpg', frame)
# Release camera and video file
camera.release(); out.release()
cv.destroyAllWindows()
```
| github_jupyter |
# Pivot_Longer : One function to cover transformations from wide to long form.
```
import janitor
import pandas as pd
import numpy as np
```
Unpivoting(reshaping data from wide to long form) in Pandas is executed either through [pd.melt](https://pandas.pydata.org/docs/reference/api/pandas.melt.html), [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html), or [pd.DataFrame.stack](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.stack.html). However, there are scenarios where a few more steps are required to massage the data into the long form that we desire. Take the dataframe below, copied from [Stack Overflow](https://stackoverflow.com/questions/64061588/pandas-melt-multiple-columns-to-tabulate-a-dataset#64062002):
```
df = pd.DataFrame(
{
"id": [1, 2, 3],
"M_start_date_1": [201709, 201709, 201709],
"M_end_date_1": [201905, 201905, 201905],
"M_start_date_2": [202004, 202004, 202004],
"M_end_date_2": [202005, 202005, 202005],
"F_start_date_1": [201803, 201803, 201803],
"F_end_date_1": [201904, 201904, 201904],
"F_start_date_2": [201912, 201912, 201912],
"F_end_date_2": [202007, 202007, 202007],
}
)
df
```
Below is a [beautiful solution](https://stackoverflow.com/a/64062027/7175713), from Stack Overflow :
```
df1 = df.set_index('id')
df1.columns = df1.columns.str.split('_', expand=True)
df1 = (df1.stack(level=[0,2,3])
.sort_index(level=[0,1], ascending=[True, False])
.reset_index(level=[2,3], drop=True)
.sort_index(axis=1, ascending=False)
.rename_axis(['id','cod'])
.reset_index())
df1
```
We propose an alternative, based on [pandas melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) and [concat](https://pandas.pydata.org/docs/reference/api/pandas.concat.html), that abstracts the reshaping mechanism, allows the user to focus on the task, can be applied to other scenarios, and is chainable :
```
result = (df.pivot_longer(index="id",
names_to=("cod", ".value", 'dates'),
names_pattern="(M|F)_(start|end)_(.+)",
sort_by_appearance=True)
.drop(columns='dates')
)
result
df1.equals(result)
```
[pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html#janitor.pivot_longer) is not a new idea; it is a combination of ideas from R's [tidyr](https://tidyr.tidyverse.org/reference/pivot_longer.html) and [data.table](https://rdatatable.gitlab.io/data.table/) and is built on the powerful pandas' [melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) and [concat](https://pandas.pydata.org/docs/reference/api/pandas.concat.html) functions.
[pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html#janitor.pivot_longer) can melt dataframes easily; It is just a wrapper around pandas' [melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html).
[Source Data](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html#reshaping-by-melt)
```
index = pd.MultiIndex.from_tuples([('person', 'A'), ('person', 'B')])
df = pd.DataFrame({'first': ['John', 'Mary'],
'last': ['Doe', 'Bo'],
'height': [5.5, 6.0],
'weight': [130, 150]},
index=index)
df
df.pivot_longer(index=['first','last'])
```
If you want the data unpivoted in order of appearance, you can set `sort_by_appearance` to `True``:
```
df.pivot_longer(
index=['first','last'],
sort_by_appearance = True
)
```
If you wish to reuse the original index, you can set `ignore_index` to `False``; note that the index labels will be repeated as necessary:
```
df.pivot_longer(
index=['first','last'],
ignore_index = False
)
```
You can also unpivot MultiIndex columns, the same way you would with pandas' [melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html#pandas.melt):
[Source Data](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html#pandas.melt)
```
df = pd.DataFrame({'A': {0: 'a', 1: 'b', 2: 'c'},
'B': {0: 1, 1: 3, 2: 5},
'C': {0: 2, 1: 4, 2: 6}})
df.columns = [list('ABC'), list('DEF')]
df
df.pivot_longer(
index = [("A", "D")],
values_to = "num"
)
df.pivot_longer(
index = [("A", "D")],
column_names = [("B", "E")]
)
```
And just like [melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html#pandas.melt), you can unpivot on a specific level, with `column_level`:
```
df.pivot_longer(
index = "A",
column_names = "B",
column_level = 0
)
```
Note that when unpivoting MultiIndex columns, you need to pass a list of tuples to the `index` or `column_names` parameters.
Also, if `names_sep` or `names_pattern` is not None, then unpivoting on MultiIndex columns is not supported.
You can dynamically select columns, using regular expressions with the `janitor.patterns` function (inspired by R's data.table's [patterns](https://rdatatable.gitlab.io/data.table/reference/patterns.html) function, and is really just a wrapper around `re.compile`), especially if it is a lot of column names, and you are *lazy* like me 😄
```
url = 'https://raw.githubusercontent.com/tidyverse/tidyr/main/data-raw/billboard.csv'
df = pd.read_csv(url)
df
# unpivot all columns that start with 'wk'
df.pivot_longer(column_names = janitor.patterns("^(wk)"),
names_to='week')
```
You can also use [pyjanitor's](https://pyjanitor-devs.github.io/pyjanitor/) [select_columns](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.select_columns.html#janitor.select_columns) syntax:
```
df.pivot_longer(column_names = "wk*",
names_to = 'week')
```
[pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html#janitor.pivot_longer) can also unpivot paired columns. In this regard, it is like pandas' [wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html), but with more flexibility and power. Let's look at an example from pandas' [wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) docs :
```
df = pd.DataFrame({
'famid': [1, 1, 1, 2, 2, 2, 3, 3, 3],
'birth': [1, 2, 3, 1, 2, 3, 1, 2, 3],
'ht1': [2.8, 2.9, 2.2, 2, 1.8, 1.9, 2.2, 2.3, 2.1],
'ht2': [3.4, 3.8, 2.9, 3.2, 2.8, 2.4, 3.3, 3.4, 2.9]
})
df
```
In the data above, the `height`(ht) is paired with `age`(numbers). [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) can handle this easily:
```
pd.wide_to_long(df, stubnames='ht', i=['famid', 'birth'], j='age')
```
Now let's see how [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) handles this:
```
df.pivot_longer(index=['famid','birth'],
names_to=('.value', 'age'),
names_pattern=r"(ht)(\d)")
```
The first observable difference is that [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) is method chainable, while [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) is not. Now, let's learn more about the `.value` variable.
When `.value` is used in `names_to`, a pairing is created between `names_to` and `names_pattern``. For the example above, we get this pairing :
{".value": ("ht"), "age": (\d)}
This tells the [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) function to keep values associated with `.value`(`ht`) as the column name, while values not associated with `.value`, in this case, the numbers, will be collated under a new column `age``. Internally, pandas `str.extract` is used to get the capturing groups before reshaping. This level of abstraction, we believe, allows the user to focus on the task, and get things done faster.
Note that if you want the data returned in order of appearance you can set `sort_by_appearance` to `True`:
```
df.pivot_longer(
index = ['famid','birth'],
names_to = ('.value', 'age'),
names_pattern = r"(ht)(\d)",
sort_by_appearance = True,
)
```
Note that you are likely to get more speed when `sort_by_appearance` is `False``.
Note also that the values in the `age` column are of `object` dtype. You can change the dtype, using pandas' [astype](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.astype.html) method.
We've seen already that [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) handles this already and very well, so why bother? Let's look at another scenario where [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) would need a few more steps. [Source Data](https://community.rstudio.com/t/pivot-longer-on-multiple-column-sets-pairs/43958):
```
df = pd.DataFrame(
{
"off_loc": ["A", "B", "C", "D", "E", "F"],
"pt_loc": ["G", "H", "I", "J", "K", "L"],
"pt_lat": [
100.07548220000001,
75.191326,
122.65134479999999,
124.13553329999999,
124.13553329999999,
124.01028909999998,
],
"off_lat": [
121.271083,
75.93845266,
135.043791,
134.51128400000002,
134.484374,
137.962195,
],
"pt_long": [
4.472089953,
-144.387785,
-40.45611048,
-46.07156181,
-46.07156181,
-46.01594293,
],
"off_long": [
-7.188632000000001,
-143.2288569,
21.242563,
40.937416999999996,
40.78472,
22.905889000000002,
],
}
)
df
```
We can unpivot with [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) by first reorganising the columns :
```
df1 = df.copy()
df1.columns = ["_".join(col.split("_")[::-1])
for col in df1.columns]
df1
```
Now, we can unpivot :
```
pd.wide_to_long(
df1.reset_index(),
stubnames=["loc", "lat", "long"],
sep="_",
i="index",
j="set",
suffix=".+",
)
```
We can get the same transformed dataframe, with less lines, using [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html):
```
df.pivot_longer(
names_to = ["set", ".value"],
names_pattern = "(.+)_(.+)"
)
# Another way to see the pairings,
# to see what is linked to `.value`,
# names_to = ["set", ".value"]
# names_pattern = "(.+)_(.+)"
# column _names = off_loc
# off_lat
# off_long
```
Again, the key here is the `.value` symbol. Pairing `names_to` with `names_pattern` and its results from [pd.str.extract](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.str.extract.html), we get :
set--> (.+) --> [off, pt] and
.value--> (.+) --> [loc, lat, long]
All values associated with `.value`(loc, lat, long) remain as column names, while values not associated with `.value`(off, pt) are lumped into a new column `set``.
Notice that we did not have to reset the index - [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) takes care of that internally; [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) allows you to focus on what you want, so you can get it and move on.
Note that the unpivoting could also have been executed with `names_sep`:
```
df.pivot_longer(
names_to = ["set", ".value"],
names_sep = "_",
ignore_index = False,
sort_by_appearance = True
)
```
Let's look at another example, from [Stack Overflow](https://stackoverflow.com/questions/45123924/convert-pandas-dataframe-from-wide-to-long/45124130) :
```
df = pd.DataFrame([{'a_1': 2, 'ab_1': 3,
'ac_1': 4, 'a_2': 5,
'ab_2': 6, 'ac_2': 7}])
df
```
The data above requires extracting `a`, `ab` and `ac` from `1` and `2`. This is another example of a paired column. We could solve this using [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html); infact there is a very good solution from [Stack Overflow](https://stackoverflow.com/a/45124775/7175713)
```
df1 = df.copy()
df1['id'] = df1.index
pd.wide_to_long(df1, ['a','ab','ac'],i='id',j='num',sep='_')
```
Or you could simply pass the buck to [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html):
```
df.pivot_longer(
names_to = ('.value', 'num'),
names_sep = '_'
)
```
In the solution above, we used the `names_sep` argument, as it is more convenient. A few more examples to get you familiar with the `.value` symbol.
[Source Data](https://stackoverflow.com/questions/55403008/pandas-partial-melt-or-group-melt)
```
df = pd.DataFrame([[1,1,2,3,4,5,6],
[2,7,8,9,10,11,12]],
columns=['id', 'ax','ay','az','bx','by','bz'])
df
df.pivot_longer(
index = 'id',
names_to = ('name', '.value'),
names_pattern = '(.)(.)'
)
```
For the code above `.value` is paired with `x`, `y`, `z`(which become the new column names), while `a`, `b` are unpivoted into the `name` column.
In the dataframe below, we need to unpivot the data, keeping only the suffix `hi`, and pulling out the number between `A` and `g`. [Source Data](https://stackoverflow.com/questions/35929985/melt-a-data-table-with-a-column-pattern)
```
df = pd.DataFrame([{'id': 1, 'A1g_hi': 2,
'A2g_hi': 3, 'A3g_hi': 4,
'A4g_hi': 5}])
df
df.pivot_longer(
index = 'id',
names_to = ['time','.value'],
names_pattern = "A(\d)g_(hi)")
```
Let's see an example where we have multiple values in a paired column, and we wish to split them into separate columns. [Source Data](https://stackoverflow.com/questions/64107566/how-to-pivot-longer-and-populate-with-fields-from-column-names-at-the-same-tim?noredirect=1#comment113369419_64107566) :
```
df = pd.DataFrame(
{
"Sony | TV | Model | value": {0: "A222", 1: "A234", 2: "A4345"},
"Sony | TV | Quantity | value": {0: 5, 1: 5, 2: 4},
"Sony | TV | Max-quant | value": {0: 10, 1: 9, 2: 9},
"Panasonic | TV | Model | value": {0: "T232", 1: "S3424", 2: "X3421"},
"Panasonic | TV | Quantity | value": {0: 1, 1: 5, 2: 1},
"Panasonic | TV | Max-quant | value": {0: 10, 1: 12, 2: 11},
"Sanyo | Radio | Model | value": {0: "S111", 1: "S1s1", 2: "S1s2"},
"Sanyo | Radio | Quantity | value": {0: 4, 1: 2, 2: 4},
"Sanyo | Radio | Max-quant | value": {0: 9, 1: 9, 2: 10},
}
)
df
```
The goal is to reshape the data into long format, with separate columns for `Manufacturer`(Sony,...), `Device`(TV, Radio), `Model`(S3424, ...), `maximum quantity` and `quantity``.
Below is the [accepted solution](https://stackoverflow.com/a/64107688/7175713) on Stack Overflow :
```
df1 = df.copy()
# Create a multiIndex column header
df1.columns = pd.MultiIndex.from_arrays(
zip(*df1.columns.str.split("\s?\|\s?"))
)
# Reshape the dataframe using
# `set_index`, `droplevel`, and `stack`
(df1.stack([0, 1])
.droplevel(1, axis=1)
.set_index("Model", append=True)
.rename_axis([None, "Manufacturer", "Device", "Model"])
.sort_index(level=[1, 2, 3])
.reset_index()
.drop("level_0", axis=1)
)
```
Or, we could use [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html), along with `.value` in `names_to` and a regular expression in `names_pattern` :
```
df.pivot_longer(
names_to = ("Manufacturer", "Device", ".value"),
names_pattern = r"(.+)\|(.+)\|(.+)\|.*",
)
```
The cleanup (removal of whitespace in the column names) is left as an exercise for the reader.
What if we are interested in unpivoting only a part of the entire dataframe? [Source Data](https://stackoverflow.com/questions/63044119/converting-wide-format-data-into-long-format-with-multiple-indices-and-grouped-d)
```
df = pd.DataFrame({'time': [1, 2, 3],
'factor': ['a','a','b'],
'variable1': [0,0,0],
'variable2': [0,0,1],
'variable3': [0,2,0],
'variable4': [2,0,1],
'variable5': [1,0,1],
'variable6': [0,1,1],
'O1V1': [0,0.2,-0.3],
'O1V2': [0,0.4,-0.9],
'O1V3': [0.5,0.2,-0.6],
'O1V4': [0.5,0.2,-0.6],
'O1V5': [0,0.2,-0.3],
'O1V6': [0,0.4,-0.9],
'O1V7': [0.5,0.2,-0.6],
'O1V8': [0.5,0.2,-0.6],
'O2V1': [0,0.5,0.3],
'O2V2': [0,0.2,0.9],
'O2V3': [0.6,0.1,-0.3],
'O2V4': [0.5,0.2,-0.6],
'O2V5': [0,0.5,0.3],
'O2V6': [0,0.2,0.9],
'O2V7': [0.6,0.1,-0.3],
'O2V8': [0.5,0.2,-0.6],
'O3V1': [0,0.7,0.4],
'O3V2': [0.9,0.2,-0.3],
'O3V3': [0.5,0.2,-0.7],
'O3V4': [0.5,0.2,-0.6],
'O3V5': [0,0.7,0.4],
'O3V6': [0.9,0.2,-0.3],
'O3V7': [0.5,0.2,-0.7],
'O3V8': [0.5,0.2,-0.6]})
df
```
What is the task? This is copied verbatim from the source:
<blockquote>Each row of the data frame represents a time period. There are multiple 'subjects' being monitored, namely O1, O2, and O3. Each subject has 8 variables being measured. I need to convert this data into long format where each row contains the information for one subject at a given time period, but with only the first 4 subject variables, as well as the extra information about this time period in columns 2-4, but not columns 5-8.</blockquote>
Below is the accepted solution, using [wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html):
```
df1 = df.rename(columns={x: x[2:]+x[1:2] for x in df.columns[df.columns.str.startswith('O')]})
df1 = pd.wide_to_long(df1, i=['time', 'factor']+[f'variable{i}' for i in range(1,7)],
j='id', stubnames=[f'V{i}' for i in range(1,9)], suffix='.*')
df1 = (df1.reset_index()
.drop(columns=[f'V{i}' for i in range(5,9)]
+[f'variable{i}' for i in range(3,7)]))
df1
```
We can abstract the details and focus on the task with [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html):
```
df.pivot_longer(
index = slice("time", "variable2"),
column_names = janitor.patterns(".+V[1-4]$"),
names_to = ("id", ".value"),
names_pattern = ".(.)(.+)$",
sort_by_appearance = True
)
```
One more example on the `.value` symbol for paired columns [Source Data](https://stackoverflow.com/questions/59477686/python-pandas-melt-single-column-into-two-seperate) :
```
df = pd.DataFrame({'id': [1, 2],
'A_value': [50, 33],
'D_value': [60, 45]})
df
df.pivot_longer(
index = 'id',
names_to = ('value_type', '.value'),
names_sep = '_'
)
```
There are scenarios where we need to unpivot the data, and group values within the column names under new columns. The values in the columns will not become new column names, so we do not need the `.value` symbol. Let's see an example below: [Source Data](https://stackoverflow.com/questions/59550804/melt-column-by-substring-of-the-columns-name-in-pandas-python)
```
df = pd.DataFrame({'subject': [1, 2],
'A_target_word_gd': [1, 11],
'A_target_word_fd': [2, 12],
'B_target_word_gd': [3, 13],
'B_target_word_fd': [4, 14],
'subject_type': ['mild', 'moderate']})
df
```
In the dataframe above, `A` and `B` represent conditions, while the suffixes `gd` and `fd` represent value types. We are not interested in the words in the middle (`_target_word`). We could solve it this way (this is the chosen solution, copied from [Stack Overflow](https://stackoverflow.com/a/59550967/7175713)) :
```
new_df =(pd.melt(df,
id_vars=['subject_type','subject'],
var_name='abc')
.sort_values(by=['subject', 'subject_type'])
)
new_df['cond']=(new_df['abc']
.apply(lambda x: (x.split('_'))[0])
)
new_df['value_type']=(new_df
.pop('abc')
.apply(lambda x: (x.split('_'))[-1])
)
new_df
```
Or, we could just pass the buck to [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html):
```
df.pivot_longer(
index = ["subject", "subject_type"],
names_to = ("cond", "value_type"),
names_pattern = "([A-Z]).*(gd|fd)",
)
```
In the code above, we pass in the new names of the columns to `names_to`('cond', 'value_type'), and pass the groups to be extracted as a regular expression to `names_pattern`.
Here's another example where [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) abstracts the process and makes reshaping easy.
In the dataframe below, we would like to unpivot the data and separate the column names into individual columns(`vault` should be in an `event` column, `2012` should be in a `year` column and `f` should be in a `gender` column). [Source Data](https://dcl-wrangle.stanford.edu/pivot-advanced.html)
```
df = pd.DataFrame(
{
"country": ["United States", "Russia", "China"],
"vault_2012_f": [
48.132,
46.366,
44.266,
],
"vault_2012_m": [46.632, 46.866, 48.316],
"vault_2016_f": [
46.866,
45.733,
44.332,
],
"vault_2016_m": [45.865, 46.033, 45.0],
"floor_2012_f": [45.366, 41.599, 40.833],
"floor_2012_m": [45.266, 45.308, 45.133],
"floor_2016_f": [45.999, 42.032, 42.066],
"floor_2016_m": [43.757, 44.766, 43.799],
}
)
df
```
We could achieve this with a combination of [pd.melt](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.melt.html) and pandas string methods (or janitor's [deconcatenate_columns](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.deconcatenate_column.html#janitor.deconcatenate_column) method); or we could, again, pass the buck to [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html):
```
df.pivot_longer(
index = "country",
names_to = ["event", "year", "gender"],
names_sep = "_",
values_to = "score",
)
```
Again, if you want the data returned in order of appearance, you can turn on the `sort_by_appearance` parameter:
```
df.pivot_longer(
index = "country",
names_to = ["event", "year", "gender"],
names_sep = "_",
values_to = "score",
sort_by_appearance = True
)
```
One more feature that [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) offers is to pass a list of regular expressions to `names_pattern`. This comes in handy when one single regex cannot encapsulate similar columns for reshaping to long form. This idea is inspired by the [melt](https://rdatatable.gitlab.io/data.table/reference/melt.data.table.html) function in R's [data.table](https://rdatatable.gitlab.io/data.table/). A couple of examples should make this clear.
[Source Data](https://stackoverflow.com/questions/61138600/tidy-dataset-with-pivot-longer-multiple-columns-into-two-columns)
```
df = pd.DataFrame(
[{'title': 'Avatar',
'actor_1': 'CCH_Pound…',
'actor_2': 'Joel_Davi…',
'actor_3': 'Wes_Studi',
'actor_1_FB_likes': 1000,
'actor_2_FB_likes': 936,
'actor_3_FB_likes': 855},
{'title': 'Pirates_of_the_Car…',
'actor_1': 'Johnny_De…',
'actor_2': 'Orlando_B…',
'actor_3': 'Jack_Daven…',
'actor_1_FB_likes': 40000,
'actor_2_FB_likes': 5000,
'actor_3_FB_likes': 1000},
{'title': 'The_Dark_Knight_Ri…',
'actor_1': 'Tom_Hardy',
'actor_2': 'Christian…',
'actor_3': 'Joseph_Gor…',
'actor_1_FB_likes': 27000,
'actor_2_FB_likes': 23000,
'actor_3_FB_likes': 23000},
{'title': 'John_Carter',
'actor_1': 'Daryl_Sab…',
'actor_2': 'Samantha_…',
'actor_3': 'Polly_Walk…',
'actor_1_FB_likes': 640,
'actor_2_FB_likes': 632,
'actor_3_FB_likes': 530},
{'title': 'Spider-Man_3',
'actor_1': 'J.K._Simm…',
'actor_2': 'James_Fra…',
'actor_3': 'Kirsten_Du…',
'actor_1_FB_likes': 24000,
'actor_2_FB_likes': 11000,
'actor_3_FB_likes': 4000},
{'title': 'Tangled',
'actor_1': 'Brad_Garr…',
'actor_2': 'Donna_Mur…',
'actor_3': 'M.C._Gainey',
'actor_1_FB_likes': 799,
'actor_2_FB_likes': 553,
'actor_3_FB_likes': 284}]
)
df
```
Above, we have a dataframe of movie titles, actors, and their facebook likes. It would be great if we could transform this into a long form, with just the title, the actor names, and the number of likes. Let's look at a possible solution :
First, we reshape the columns, so that the numbers appear at the end.
```
df1 = df.copy()
pat = r"(?P<actor>.+)_(?P<num>\d)_(?P<likes>.+)"
repl = lambda m: f"""{m.group('actor')}_{m.group('likes')}_{m.group('num')}"""
df1.columns = df1.columns.str.replace(pat, repl, regex=True)
df1
```
Now, we can reshape, using [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) :
```
pd.wide_to_long(df1,
stubnames = ['actor', 'actor_FB_likes'],
i = 'title',
j = 'group',
sep = '_')
```
We could attempt to solve it with [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html), using the `.value` symbol :
```
df1.pivot_longer(
index = 'title',
names_to = (".value", "group"),
names_pattern = "(.+)_(\d)$"
)
```
What if we could just get our data in long form without the massaging? We know our data has a pattern to it --> it either ends in a number or *likes*. Can't we take advantage of that? Yes, we can (I know, I know; it sounds like a campaign slogan 🤪)
```
df.pivot_longer(
index = 'title',
names_to = ("actor", "num_likes"),
names_pattern = ('\d$', 'likes$'),
)
```
A pairing of `names_to` and `names_pattern` results in:
{"actor": '\d$', "num_likes": 'likes$'}
The first regex looks for columns that end with a number, while the other looks for columns that end with *likes*. [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) will then look for columns that end with a number and lump all the values in those columns under the `actor` column, and also look for columns that end with *like* and combine all the values in those columns into a new column -> `num_likes`. Underneath the hood, [numpy select](https://numpy.org/doc/stable/reference/generated/numpy.select.html) and [pd.Series.str.contains](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.str.contains.html) are used to pull apart the columns into the new columns.
Again, it is about the goal; we are not interested in the numbers (1,2,3), we only need the names of the actors, and their facebook likes. [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) aims to give as much flexibility as possible, in addition to ease of use, to allow the end user focus on the task.
Let's take a look at another example. [Source Data](https://stackoverflow.com/questions/60439749/pair-wise-melt-in-pandas-dataframe) :
```
df = pd.DataFrame({'id': [0, 1],
'Name': ['ABC', 'XYZ'],
'code': [1, 2],
'code1': [4, np.nan],
'code2': ['8', 5],
'type': ['S', 'R'],
'type1': ['E', np.nan],
'type2': ['T', 'U']})
df
```
We cannot directly use [pd.wide_to_long](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.wide_to_long.html) here without some massaging, as there is no definite suffix(the first `code` does not have a suffix), neither can we use `.value` here, again because there is no suffix. However, we can see a pattern where some columns start with `code`, and others start with `type`. Let's see how [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) solves this, using a sequence of regular expressions in the `names_pattern` argument :
```
df.pivot_longer(
index = ["id", "Name"],
names_to = ("code_all", "type_all"),
names_pattern = ("^code", "^type")
)
```
The key here is passing the right regular expression, and ensuring the names in `names_to` is paired with the right regex in `names_pattern`; as such, every column that starts with `code` will be included in the new `code_all` column; the same happens to the `type_all` column. Easy and flexible, right?
Let's explore another example, from [Stack Overflow](https://stackoverflow.com/questions/12466493/reshaping-multiple-sets-of-measurement-columns-wide-format-into-single-columns) :
```
df = pd.DataFrame(
[
{
"ID": 1,
"DateRange1Start": "1/1/90",
"DateRange1End": "3/1/90",
"Value1": 4.4,
"DateRange2Start": "4/5/91",
"DateRange2End": "6/7/91",
"Value2": 6.2,
"DateRange3Start": "5/5/95",
"DateRange3End": "6/6/96",
"Value3": 3.3,
}
])
df
```
In the dataframe above, we need to reshape the data to have a start date, end date and value. For the `DateRange` columns, the numbers are embedded within the string, while for `value` it is appended at the end. One possible solution is to reshape the columns so that the numbers are at the end :
```
df1 = df.copy()
pat = r"(?P<head>.+)(?P<num>\d)(?P<tail>.+)"
repl = lambda m: f"""{m.group('head')}{m.group('tail')}{m.group('num')}"""
df1.columns = df1.columns.str.replace(pat, repl, regex=True)
df1
```
Now, we can unpivot:
```
pd.wide_to_long(df1,
stubnames = ['DateRangeStart',
'DateRangeEnd',
'Value'],
i = 'ID',
j = 'num')
```
Using the `.value` symbol in pivot_longer:
```
df1.pivot_longer(
index = 'ID',
names_to = [".value",'num'],
names_pattern = "(.+)(\d)$"
)
```
Or, we could allow pivot_longer worry about the massaging; simply pass to `names_pattern` a list of regular expressions that match what we are after :
```
df.pivot_longer(
index = 'ID',
names_to = ("DateRangeStart", "DateRangeEnd", "Value"),
names_pattern = ("Start$", "End$", "^Value")
)
```
The code above looks for columns that end with *Start*(`Start$`), aggregates all the values in those columns into `DateRangeStart` column, looks for columns that end with *End*(`End$`), aggregates all the values within those columns into `DateRangeEnd` column, and finally looks for columns that start with *Value*(`^Value`), and aggregates the values in those columns into the `Value` column. Just know the patterns, and pair them accordingly. Again, the goal is a focus on the task, to make it simple for the end user.
Let's look at another example [Source Data](https://stackoverflow.com/questions/64316129/how-to-efficiently-melt-multiple-columns-using-the-module-melt-in-pandas/64316306#64316306) :
```
df = pd.DataFrame({'Activity': ['P1', 'P2'],
'General': ['AA', 'BB'],
'm1': ['A1', 'B1'],
't1': ['TA1', 'TB1'],
'm2': ['A2', 'B2'],
't2': ['TA2', 'TB2'],
'm3': ['A3', 'B3'],
't3': ['TA3', 'TB3']})
df
```
This is a [solution](https://stackoverflow.com/a/64316306/7175713) provided by yours truly :
```
(pd.wide_to_long(df,
i = ["Activity", "General"],
stubnames = ["t", "m"],
j = "number")
.set_axis(["Task", "M"],
axis = "columns")
.droplevel(-1)
.reset_index()
)
```
Or, we could use [pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html), abstract the details, and focus on the task :
```
df.pivot_longer(
index = ['Activity','General'],
names_pattern = ['^m','^t'],
names_to = ['M','Task']
)
```
Alright, one last example :
[Source Data](https://stackoverflow.com/questions/64159054/how-do-you-pivot-longer-columns-in-groups)
```
df = pd.DataFrame({'Name': ['John', 'Chris', 'Alex'],
'activity1': ['Birthday', 'Sleep Over', 'Track Race'],
'number_activity_1': [1, 2, 4],
'attendees1': [14, 18, 100],
'activity2': ['Sleep Over', 'Painting', 'Birthday'],
'number_activity_2': [4, 5, 1],
'attendees2': [10, 8, 5]})
df
```
The task here is to unpivot the data, and group the data under three new columns ("activity", "number_activity", and "attendees").
We can see that there is a pattern to the data; let's create a list of regular expressions that match the patterns and pass to `names_pattern``:
```
df.pivot_longer(
index = 'Name',
names_to = ('activity','number_activity','attendees'),
names_pattern = ("^activity","^number_activity","^attendees")
)
```
Alright, let's look at one final example:
[Source Data](https://stackoverflow.com/questions/60387077/reshaping-and-melting-dataframe-whilst-picking-up-certain-regex)
```
df = pd.DataFrame({'Location': ['Madrid', 'Madrid', 'Rome', 'Rome'],
'Account': ['ABC', 'XYX', 'ABC', 'XYX'],
'Y2019:MTD:January:Expense': [4354, 769867, 434654, 632556456],
'Y2019:MTD:January:Income': [56456, 32556456, 5214, 46724423],
'Y2019:MTD:February:Expense': [235423, 6785423, 235423, 46588]})
df
df.pivot_longer(index = ['Location','Account'],
names_to=("year", "month", ".value"),
names_pattern=r"Y(.+):MTD:(.{3}).+(Income|Expense)",
sort_by_appearance=True)
```
[pivot_longer](https://pyjanitor-devs.github.io/pyjanitor/reference/janitor.functions/janitor.pivot_longer.html) does not solve all problems; no function does. Its aim is to make it easy to unpivot dataframes from wide to long form, while offering a lot of flexibility and power.
| github_jupyter |
# Case Study: Hydrologic models of soil physical processes from the NWM and TOPMODEL
Input files for the HBV, SWMM, and TOPMODEL case studies are available [here](https://drive.google.com/drive/folders/1p96I1m88nDhiwVyOoEX0ywuL5g0oBNuk?usp=sharing). To run, copy these files into the input folder within the directory of the case study.
The Sleepers River Research Watershed (SRRW) in Vermont is an active hydrologic research site since 1959 and was the setting where Dunne and Black (1970) determined the controls of saturation-excess overland flow (SOF) on streamflow generation.
This jupyter notebook has code cells to:
1. Load data for use in R and Python scripts
2. Create a historical time series and magnitude percentile plots for simulated data
3. Generate a priori and a posteriori parameter distribution plots via Approximate Bayesian Computation
4. Compute Sobol first-order, delta, and OLS sensitivity indices
5. Produce portrait, scatter, and spider plots
To complete and visualize a comprehensive sensitivity analysis on the SWMM simulations, we use packages from both R and Python. This command allows us to run R scripts in the Python jupyter notebook.
```
%load_ext rpy2.ipython
```
Now we load the model simulation data, observation/truth data, parameter sets, time stamps, and objective function values into Python as data frames using the pandas library.
```
import pandas as pd
sim = pd.read_csv("input/simulation_ts.csv", index_col = 0)
pars = pd.read_csv("input/params.csv", header = 0)
OF = pd.read_csv("input/OF_values.csv")
obs = pd.read_csv("input/observation_ts.csv")
```
### 2. Approximate Bayesian Computation
One method integrated into the workflow was Approximate Bayesian Computation (ABC), which represents the combination of model parameter values that maximize the probability of representing the observed data. ABC applies [Bayesian theory](https://en.wikipedia.org/wiki/Bayes%27_theorem) to parameter spaces to estimate posterior distributions of model parameters. ABC is advantageous because it bypasses calculating the likelihood function by using the model simulations compared to the observed data ([Engeland and Gottschalk 2002](https://www.hydrol-earth-syst-sci.net/6/883/2002/), [Kavetski et al. 2006](https://agupubs.onlinelibrary.wiley.com/doi/10.1029/2005WR004368), [Sunnåker et al. 2013](https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1002803), [Vrugt and Sadegh 2013](https://agupubs.onlinelibrary.wiley.com/doi/full/10.1002/wrcr.20354)). The steps to compute ABC are: 1) Calculate the observed data’s statistics (e.g. mean, standard deviation) and choose model specific objective functions (e.g. NSE). 2) Assume a uniform sampling interval for the parameter space. Draw a total of n parameters from prior and simulate the model for each of the parameter points, this results in n sequences of simulated data. 3) Calculate objective functions for each sequence of simulated data. 4) Determine the distance between the observed and simulated transition frequencies for all parameter points. Remove parameter points beyond a user specified tolerance interval (e.g. NSE ≥ 0.0) to approximate samples from the posterior distribution. 5) Estimate the posterior distribution with the parameter points within the tolerance interval ([Sunnåker et al. 2013](https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1002803), [Vrugt and Sadegh 2013](https://agupubs.onlinelibrary.wiley.com/doi/full/10.1002/wrcr.20354)).
For the python script below, specify the number of model runs, tolerance of the objective functions, number of histogram bins and the figure colors. Here we used pre-defined objective functions, but the code can be modified to calculate a variety of objective functions. The plots produced are histograms of the various parameters illustrating the difference between original modeled output and the ABC constrained parameter sets.
```
# %load approx_bayes_calc_of_defined.py
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
'''
Approximate Baysian Calculation requires:
1) observation dataset (df_obs)
2) parameter sets (df_parms)
3) model output (df_model)
4) objective functions (df_OFs)
5) tolerance
6) number of model runs
'''
def approx_bayes_calc_OF(parms,OFs,simulations):
keep_nse = []
for i in np.arange(simulations):
# User can redefine tolerance and OF here
if OFs.iloc[i,0] >= tolerance_nse:
keep_nse.append(parms.iloc[i])
return keep_nse
def make_histograms(df_parms,bayes_approx,bins,alpha,cc1,cc2,parameters,metric):
plt.figure(figsize=(15,12))
gridsize = math.ceil((np.sqrt((df_parms.iloc[0,:]).size))) # graph columns
for col in np.arange(1,((df_parms.iloc[0,:]).size)+1):
plt.subplot(gridsize-1,gridsize,col)
ax = df_parms.iloc[:,col-1].plot.hist(bins=bins,alpha=alpha,color=cc1,linewidth=4)
ax = bayes_approx.iloc[:,col-1].plot.hist(bins=bins,alpha=alpha,color=cc2,linewidth=4)
ax.set_xlabel(str(parameters[col-1]))
plt.legend(['Output','ABC'],fancybox=True)
plt.tight_layout()
plt.savefig('output/plots/ABC/'+metric+'_histogram.png',dpi=1000)
def make_cdfs_pdfs(df_parms,bayes_approx,bins,alpha,cc1,cc2,parameters,metric):
plt.figure(figsize=(15,12))
gridsize = math.ceil((np.sqrt((df_parms.iloc[0,:]).size)))
for col in np.arange(1,((df_parms.iloc[0,:]).size)+1):
plt.subplot(gridsize-1,gridsize,col)
ax = df_parms.iloc[:,col-1].plot.hist(cumulative=True, density=1,bins=bins,alpha=alpha,color=cc1,linewidth=4)
ax = bayes_approx.iloc[:,col-1].plot.hist(cumulative=True, density=1,bins=bins,alpha=alpha,color=cc2,linewidth=4)
ax.set_xlabel(str(parameters[col-1]))
plt.legend(['Output','ABC'],fancybox=True)
plt.tight_layout()
plt.savefig('output/plots/ABC/'+metric+'_cdf.png',dpi=1000)
plt.figure(figsize=(15,12))
gridsize = math.ceil((np.sqrt((df_parms.iloc[0,:]).size)))
for col in np.arange(1,((df_parms.iloc[0,:]).size)+1):
plt.subplot(gridsize-1,gridsize,col)
ax = df_parms.iloc[:,col-1].plot.kde(alpha=alpha,color=cc1,linewidth=4)
ax = bayes_approx.iloc[:,col-1].plot.kde(alpha=alpha,color=cc2,linewidth=4)
ax.set_xlabel(str(parameters[col-1]))
plt.legend(['Output','ABC'],fancybox=True)
plt.tight_layout()
plt.savefig('output/plots/ABC/'+metric+'_pdf.png',dpi=1000)
def runABC(df_parms,df_OFs,runs,bins,color1,color2):
# models with objective functions within tolerance thresholds
results_nse = np.array(approx_bayes_calc_OF(df_parms,df_OFs,runs))
# saves models with objective functions within tolerance thresholds
bayes_approx_nse = pd.DataFrame(results_nse,columns=None)
bayes_approx_nse.to_csv('output/bayes_parameters_NSE.csv',index=False)
parameters = list(df_parms.columns.values)
# print ABC results and make figures
print('precent of models with NSE >= to',str(tolerance_nse),'are:',str(len(results_nse)/runs),'%')
make_histograms(df_parms,bayes_approx_nse,bins,0.5,color1,color2,parameters,'NSE')
make_cdfs_pdfs(df_parms,bayes_approx_nse,bins,0.5,color1,color2,parameters,'NSE')
# Specify tolerance for objective functions (OF)
tolerance_nse = 0.0 # OF >= tolerance (NSE)
runs = 5000 # specify number of model runs
bins = 100 # specify number of histogram bins
color1 = 'b' # color of original model output
color2 = 'k' # color of 1st ABC applied to OF (NSE)
# Runs function that evaluates models outputs with approximate Bayesian computation
runABC(pars, OF, runs, bins, color1, color2)
```
### 3. Sensitivity Analysis
Two sensitivity analyses are incorporated into the workflow: a moment-independent sensitivity analysis and an ordinary least squares regression.
The Delta index ([Borgonovo, 2007](https://doi.org/10.1016/j.ress.2006.04.015); [Borgonovo et al., 2012](https://www.sciencedirect.com/science/article/pii/S1364815211001617?via%3Dihub); [Plischke et al., 2013](https://doi.org/10.1016/j.ejor.2012.11.047)) is a moment-independent global sensitivity analysis. While less robust than indices returned by a variance-based sensitivity analysis, a moment-independent sensitivity analysis was a popular technique due to its computational efficiency and insensitivity to dependent parameters ([Pannell, 1997](https://doi.org/10.1016/S0169-5150(96)01217-0)). The Delta sensitivity analysis searches for parameters with the greatest impact on the probability density function of model output. Delta indices capture non-linear and non-monotonic parameter-output dynamics. Lastly, the ordinary least squares (OLS) regression yields an R2 coefficient, which quantified the linear effects of model input parameters on model output variance. OLS regressions have long been employed throughout model sensitivity analyses and assume an explicit interaction between model output and any given parameter ([Kleijnen, 1995](https://doi.org/10.1002/sdr.4260110403); [Pannell, 1997](https://doi.org/10.1016/S0169-5150(96)01217-0); [Zobitz et al., 2006](https://doi.org/10.1016/j.agrformet.2006.01.003)).
To visualize objective function sensitivity to model input parameters, the following code produces radial convergence plots, scatter plots, portrait plots, and spider plots based on the outputs from the sensitivity analyses.
The Sobol and delta sensitivity indices are calculated using a modified version of the Python sensitivity analysis library ([SALib](https://salib.readthedocs.io/en/latest/index.html)) and the OLS regression is calculated using [StatsModels](https://www.statsmodels.org/dev/generated/statsmodels.regression.linear_model.OLS.html) library in Python.
```
# %load SensIndices_RCPlots.py
#!/usr/bin/env python3
# import python libraries
import pandas as pd
import os
# back out a directory to load python functions from "Scripts" folder
org_dir_name = os.path.dirname(os.path.realpath('SensIndices_RCPlots.py'))
parent_dir_name = os.path.dirname(os.path.dirname(os.path.realpath('SensIndices_RCPlots.py')))
os.chdir(parent_dir_name + "/Scripts")
# load python functions from ‘Scripts’ folder
import delta
import ols
# move back into case study 0 folder
os.chdir(org_dir_name)
# load in model parameters and OF values
pars = pd.read_csv("input/params.csv", index_col = 0)
OF = pd.read_csv("input/OF_values.csv")
# Define the model inputs
problem = {
'num_vars': 10,
'names': ['qs0', 'lnTe', 'm', 'Sr0', 'Srmax', 'td', 'vch', 'vr', 'k0', 'CD']
}
# save the parameter names
param_names = problem['names']
# calculate delta indices and sobol first-order indices
results_delta = []
results_delta = delta.objective_function_delta(problem, pars, OF)
# calculate R^2 from OLS regression
results_R2 = []
results_R2 = ols.objective_function_OLS(OF, pars, param_names)
```
From the sensitivity analysis results (calculated and exported from python), we can create portrait plots, scatter plots, and spider plots for various objective functions and parameter values.
First, the data is loaded and formatted into a usable format and then exported to a .csv file. Then the script creates additional plots to help visualize and convey parameter sensitivity.
```
%%R
# %load Portrait_Scatter_Spider.R
# This script loads in data from Sobol, Delta, and OLS sensitivity analyses calculated in
# Python script for Case Study 0: Fall Creek, NY.
library(dplyr)
# load in parameter sets, objective functions, observation, simulation, and time steps
pars <- read.csv("input/params.csv", header = TRUE) %>%
dplyr::select(-1)
# "model_runs" rows, "num_pars" columns
OF <- read.csv("input/OF_values.csv", header = TRUE)
# "model_runs" rows, "num_OF" columns
# save names of objective functions and parameters
OF_names <- colnames(OF)
param_names <- colnames(pars)
# set variables of number of model runs, time steps, and number of parameters
model_runs <- nrow(pars)
num_pars <- ncol(pars)
num_OF <- ncol(OF)
# load in results from delta, sobol, and ols sensitivity analyses (calculated in python script)
source("../Scripts/python_to_r_results.R")
results_delta <- python_to_r_results(data_type = "delta", param_names, OF_names)
results_ols <- python_to_r_results(data_type = "ols", param_names, OF_names)
# save as csv files
lapply(results_delta, function(x) write.table(data.frame(x), 'output/formatted_delta.csv', append = T, sep = ',' ))
lapply(results_ols, function(x) write.table(data.frame(x), 'output/formatted_ols.csv', append = T, sep = ',' ))
# scatter plots of objective functions versus parameter values
source("../Scripts/scatterplots.R")
for (i in 1:num_OF) {
# subset by objective function, i
objective_fun <- OF[, i]
# create scatterplots of all parameters versus objective function, i
par_OF_scatter(params = pars, objective_fun, OF_name = colnames(OF)[i])
}
# portrait plots of objective functions versus parameter values
source("../Scripts/portrait_plots.R")
portrait_plot(results_delta, "delta")
portrait_plot(results_ols, "ols")
# spiders plots of objective functions versus parameter values
source("../Scripts/spider_plots.R")
spiderplot(results_delta)
spiderplot(results_ols)
```
| github_jupyter |
## Latent Distribution Two-Graph Testing
```
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(8888)
from graspy.inference import LatentDistributionTest
from graspy.embed import AdjacencySpectralEmbed
from graspy.simulations import sbm, rdpg
from graspy.utils import symmetrize
from graspy.plot import heatmap, pairplot
%matplotlib inline
```
### Generate a stochastic block model graph
We generate a stochastic block model graph (SBM), which is shown below.
```
n_components = 4 # the number of embedding dimensions for ASE
P = np.array([[0.9, 0.11, 0.13, 0.2],
[0, 0.7, 0.1, 0.1],
[0, 0, 0.8, 0.1],
[0, 0, 0, 0.85]])
P = symmetrize(P)
csize = [50] * 4
A = sbm(csize, P)
X = AdjacencySpectralEmbed(n_components=n_components).fit_transform(A)
heatmap(A, title='4-block SBM adjacency matrix')
pairplot(X, title='4-block adjacency spectral embedding')
```
### Latent distribution test where null is true
Now, we want to know whether the above two graphs were generated from the same latent position. We know that they were, so the test should predict that the differences between SBM 1 and 2 (up to a rotation) are no greater than those differences observed by chance.
In other words, we are testing
\begin{align*}
H_0:&X_1 = X_2\\
H_\alpha:& X_1 \neq X_2
\end{align*}
and want to see that the p-value for the unmatched test is high (fail to reject the null)
We generate a second SBM in the same way, and run an unmatched test on it, generating a distance between the two graphs as well as a null distribution of distances between permutations of the graph. We can see this below.
```
A1 = sbm(csize, P)
heatmap(A1, title='4-block SBM adjacency matrix A1')
X1 = AdjacencySpectralEmbed(n_components=n_components).fit_transform(A1)
pairplot(X1, title='4-block adjacency spectral embedding A1')
```
### Plot of Null Distribution
We plot the null distribution shown in blue and the test statistic shown red vertical line. We see that the test static is small, resulting in p-value of 0.94. Thus, we cannot reject the null hypothesis that the two graphs come from the same generating distributions.
```
ldt = LatentDistributionTest()
p = ldt.fit(A, A1)
fig, ax = plt.subplots(figsize=(10, 6))
ax.hist(ldt.null_distribution_, 50)
ax.axvline(ldt.sample_T_statistic_, color='r')
ax.set_title("P-value = {}".format(p), fontsize=20)
plt.show();
```
### Latent distribution test where null is false
We generate a seconds SBM with different block probabilities, and run a latent distribution test comaring the previous graph with the new one.
```
P2 = np.array([[0.8, 0.2, 0.2, 0.5],
[0, 0.9, 0.3, 0.2],
[0, 0, 0.5, 0.2],
[0, 0, 0, 0.5]])
P2 = symmetrize(P2)
A2 = sbm(csize, P2)
heatmap(A2, title='4-block SBM adjacency matrix A2')
X2 = AdjacencySpectralEmbed(n_components=n_components).fit_transform(A2)
pairplot(X2, title='4-block adjacency spectral embedding A2')
```
### Plot of Null Distribution
We plot the null distribution shown in blue and the test statistic shown red vertical line. We see that the test static is small, resulting in p-value of 0. Thus, we reject the null hypothesis that the two graphs come from the same generating distributions.
```
ldt = LatentDistributionTest()
p = ldt.fit(A, A2)
fig, ax = plt.subplots(figsize=(10, 6))
ax.hist(ldt.null_distribution_, 50)
ax.axvline(ldt.sample_T_statistic_, color='r')
ax.set_title("P-value = {}".format(p), fontsize=20)
plt.show();
```
| github_jupyter |
# Acquire the Data
```
import numpy as np
import pandas as pd
import requests, zipfile, io
```
## Get the Files
```
# Get the url for ml-100k
zip_file_url = "http://files.grouplens.org/datasets/movielens/ml-100k.zip"
# Get the files and extract them
print("Downloading movielens data...")
r = requests.get(zip_file_url, stream=True)
z = zipfile.ZipFile(io.BytesIO(r.content))
z.extractall()
print("Done.")
```
## Users, Items & Ratings
**Users** from the README
`
u.user -- Demographic information about the users; this is a tab
separated list of
user id | age | gender | occupation | zip code
The user ids are the ones used in the u.data data set.
`
```
users_cols = ['user_id', 'age', 'sex', 'occupation', 'zip_code']
users = pd.read_csv('ml-100k/u.user', sep='|', names=users_cols, encoding='latin-1')
users.shape
users.head()
```
**Items** from the README
`
u.item -- Information about the items (movies); this is a tab separated
list of
movie id | movie title | release date | video release date |
IMDb URL | unknown | Action | Adventure | Animation |
Children's | Comedy | Crime | Documentary | Drama | Fantasy |
Film-Noir | Horror | Musical | Mystery | Romance | Sci-Fi |
Thriller | War | Western |
The last 19 fields are the genres, a 1 indicates the movie
is of that genre, a 0 indicates it is not; movies can be in
several genres at once.
The movie ids are the ones used in the u.data data set.
`
```
genre_cols = [
"genre_unknown", "Action", "Adventure", "Animation", "Children", "Comedy",
"Crime", "Documentary", "Drama", "Fantasy", "FilmNoir", "Horror",
"Musical", "Mystery", "Romance", "SciFi", "Thriller", "War", "Western"
]
items_cols = ['movie_id', 'title', 'release_date', "video_release_date", "imdb_url"] + genre_cols
items_raw = pd.read_csv('ml-100k/u.item', sep='|', names=items_cols, encoding='latin-1')
items_raw.shape
items_raw.head()
```
**Ratings** from the README
`
u.data -- The full u data set, 100000 ratings by 943 users on 1682 items.
Each user has rated at least 20 movies. Users and items are
numbered consecutively from 1. The data is randomly
ordered. This is a tab separated list of
user id | item id | rating | timestamp.
The time stamps are unix seconds since 1/1/1970 UTC
`
```
ratings_cols = ['user_id', 'movie_id', 'rating', 'unix_timestamp']
ratings = pd.read_csv('ml-100k/u.data', sep='\t', names=ratings_cols, encoding='latin-1')
ratings.shape
ratings.head()
```
## Save the Data
```
from utils import create_directory
### Create Feature Directory
create_directory("/data")
# Save the data
users.to_csv("data/users.csv", index=None)
items_raw.to_csv("data/items_raw.csv", index=None)
ratings.to_csv("data/ratings.csv", index=False)
```
| github_jupyter |
```
# Installing gutenberg is tricky, so it is not included in the requirements.txt
# For it to work we need a berkelydb version <= 6 for licensing reasons. On OSX
# using brew you can do:
# brew install berkeley-db@4
!pip install gutenberg
# If this doesn't work, this notebook should still run, just not fetching data
# from gutenberg.
try:
GUTENBERG = True
from gutenberg.acquire import load_etext
from gutenberg.query import get_etexts, get_metadata
from gutenberg.acquire import get_metadata_cache
from gutenberg.acquire.text import UnknownDownloadUriException
from gutenberg.cleanup import strip_headers
from gutenberg._domain_model.exceptions import CacheAlreadyExistsException
except ImportError:
GUTENBERG = False
print('Gutenberg is not installed. See instructions at https://pypi.python.org/pypi/Gutenberg')
from keras.models import Input, Model
from keras.layers import Dense, Dropout
from keras.layers import LSTM
from keras.layers.wrappers import TimeDistributed
import keras.callbacks
import keras.backend as K
import scipy.misc
import json
import os, sys
import re
import PIL
from PIL import ImageDraw
from keras.optimizers import RMSprop
import random
import numpy as np
import tensorflow as tf
from keras.utils import get_file
from IPython.display import clear_output, Image, display, HTML
try:
from io import BytesIO
except ImportError:
from StringIO import StringIO as BytesIO
if GUTENBERG:
cache = get_metadata_cache()
try:
cache.populate()
except CacheAlreadyExistsException:
pass
if GUTENBERG:
for text_id in get_etexts('author', 'Shakespeare, William'):
print(text_id, list(get_metadata('title', text_id))[0])
if GUTENBERG:
shakespeare = strip_headers(load_etext(100))
else:
path = get_file('shakespeare', 'https://storage.googleapis.com/deep-learning-cookbook/100-0.txt')
shakespeare = open(path).read()
training_text = shakespeare.split('\nTHE END', 1)[-1]
len(training_text)
chars = list(sorted(set(training_text)))
char_to_idx = {ch: idx for idx, ch in enumerate(chars)}
len(chars)
def char_rnn_model(num_chars, num_layers, num_nodes=512, dropout=0.1):
input = Input(shape=(None, num_chars), name='input')
prev = input
for i in range(num_layers):
lstm = LSTM(num_nodes, return_sequences=True, name='lstm_layer_%d' % (i + 1))(prev)
if dropout:
prev = Dropout(dropout)(lstm)
else:
prev = lstm
dense = TimeDistributed(Dense(num_chars, name='dense', activation='softmax'))(prev)
model = Model(inputs=[input], outputs=[dense])
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
return model
model = char_rnn_model(len(chars), num_layers=2, num_nodes=640, dropout=0)
model.summary()
CHUNK_SIZE = 160
def data_generator(all_text, char_to_idx, batch_size, chunk_size):
X = np.zeros((batch_size, chunk_size, len(char_to_idx)))
y = np.zeros((batch_size, chunk_size, len(char_to_idx)))
while True:
for row in range(batch_size):
idx = random.randrange(len(all_text) - chunk_size - 1)
chunk = np.zeros((chunk_size + 1, len(char_to_idx)))
for i in range(chunk_size + 1):
chunk[i, char_to_idx[all_text[idx + i]]] = 1
X[row, :, :] = chunk[:chunk_size]
y[row, :, :] = chunk[1:]
yield X, y
next(data_generator(training_text, char_to_idx, 4, chunk_size=CHUNK_SIZE))
early = keras.callbacks.EarlyStopping(monitor='loss',
min_delta=0.03,
patience=3,
verbose=0, mode='auto')
BATCH_SIZE = 256
model.fit_generator(
data_generator(training_text, char_to_idx, batch_size=BATCH_SIZE, chunk_size=CHUNK_SIZE),
epochs=40,
callbacks=[early,],
steps_per_epoch=2 * len(training_text) / (BATCH_SIZE * CHUNK_SIZE),
verbose=2
)
with open('zoo/06/shakespeare.json', 'w') as fout:
json.dump({
'chars': ''.join(chars),
'char_to_idx': char_to_idx,
'chunk_size': CHUNK_SIZE,
}, fout)
model.save('zoo/06/shakespeare.h5')
def generate_output(model, training_text, start_index=None, diversity=None, amount=400):
if start_index is None:
start_index = random.randint(0, len(training_text) - CHUNK_SIZE - 1)
generated = training_text[start_index: start_index + CHUNK_SIZE]
yield generated + '#'
for i in range(amount):
x = np.zeros((1, len(generated), len(chars)))
for t, char in enumerate(generated):
x[0, t, char_to_idx[char]] = 1.
preds = model.predict(x, verbose=0)[0]
if diversity is None:
next_index = np.argmax(preds[len(generated) - 1])
else:
preds = np.asarray(preds[len(generated) - 1]).astype('float64')
preds = np.log(preds) / diversity
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
next_index = np.argmax(probas)
next_char = chars[next_index]
yield next_char
generated += next_char
return generated
for ch in generate_output(model, training_text):
sys.stdout.write(ch)
print()
def find_python(rootdir):
matches = []
for root, dirnames, filenames in os.walk(rootdir):
for fn in filenames:
if fn.endswith('.py'):
matches.append(os.path.join(root, fn))
return matches
# + find_python(os.path.join(sys.executable.rsplit('/', 2)[0], 'lib'))
srcs = find_python(random.__file__.rsplit('/', 1)[0])
len(srcs)
def replacer(value):
value = ''.join(ch for ch in value if ord(ch) < 127)
if not ' ' in value:
return value
if sum(1 for ch in value if ch.isalpha()) > 6:
return 'MSG'
return value
def replace_literals(st):
res = []
start_text = start_quote = i = 0
quote = ''
while i < len(st):
if quote:
if st[i: i + len(quote)] == quote:
quote = ''
start_text = i
res.append(replacer(st[start_quote: i]))
elif st[i] in '"\'':
quote = st[i]
if i < len(st) - 2 and st[i + 1] == st[i + 2] == quote:
quote = 3 * quote
start_quote = i + len(quote)
res.append(st[start_text: start_quote])
if st[i] == '\n' and len(quote) == 1:
start_text = i
res.append(quote)
quote = ''
if st[i] == '\\':
i += 1
i += 1
return ''.join(res) + st[start_text:]
#replace_literals('print("hel\\"lo")') + replace_literals("print('hel\\'lo world')")
replace_literals('this = "wrong\n')
COMMENT_RE = re.compile('#.*')
python_code = []
for fn in srcs:
try:
with open(fn, 'r') as fin:
src = fin.read()
except UnicodeDecodeError:
print('Could not read %s' % fn)
src = replace_literals(src)
src = COMMENT_RE.sub('', src)
python_code.append(src)
python_code = '\n\n\n'.join(python_code)
len(python_code)
py_chars = list(sorted(set(python_code)))
py_char_to_idx = {ch: idx for idx, ch in enumerate(py_chars)}
len(py_chars)
py_model = char_rnn_model(len(py_chars), num_layers=2, num_nodes=640, dropout=0)
py_model.summary()
early = keras.callbacks.EarlyStopping(monitor='loss',
min_delta=0.03,
patience=3,
verbose=0, mode='auto')
BATCH_SIZE = 256
py_model.fit_generator(
data_generator(python_code, py_char_to_idx, batch_size=BATCH_SIZE, chunk_size=160),
epochs=40,
callbacks=[early,],
steps_per_epoch=2 * len(python_code) / (BATCH_SIZE * 160),
verbose=2
)
def generate_code(model, start_with='\ndef ', end_with='\n\n', diversity=1.0):
generated = start_with
yield generated
for i in range(2000):
x = np.zeros((1, len(generated), len(py_chars)))
for t, char in enumerate(generated):
x[0, t, py_char_to_idx[char]] = 1.
preds = model.predict(x, verbose=0)[0]
preds = np.asarray(preds[len(generated) - 1]).astype('float64')
preds = np.log(preds) / diversity
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
next_index = np.argmax(probas)
next_char = py_chars[next_index]
yield next_char
generated += next_char
if generated.endswith(end_with):
break
for i in range(20):
for ch in generate_code(py_model):
sys.stdout.write(ch)
st += ch
print()
BATCH_SIZE = 512
flat_model = char_rnn_model(len(py_chars), num_layers=1, num_nodes=512, dropout=0)
early = keras.callbacks.EarlyStopping(monitor='loss',
min_delta=0.03,
patience=3,
verbose=0, mode='auto')
flat_model.fit_generator(
data_generator(python_code, py_char_to_idx, batch_size=BATCH_SIZE, chunk_size=160),
epochs=40,
callbacks=[early,],
steps_per_epoch=2 * len(python_code) / (BATCH_SIZE * 160),
verbose=2
)
example_code = 'if a == 2:\n b=1\nelse:\n b=2\n'
#example_code = 'a=(2 * 3)\nb=(4 * 6 + 7)\nreturn C'
def activations(model, code):
x = np.zeros((1, len(code), len(py_char_to_idx)))
for t, char in enumerate(code):
x[0, t, py_char_to_idx[char]] = 1.
output = model.get_layer('lstm_layer_1').output
f = K.function([model.input], [output])
return f([x])[0][0]
act = activations(flat_model, example_code)
act.shape
def interesting_neurons(act):
res = []
for n in np.argmax(act, axis=1):
if not n in res:
res.append(n)
return res
neurons = interesting_neurons(act)
len(neurons)
def visualize_neurons(neurons, code, act, cell_size=12):
img = np.full((len(neurons) + 1, len(code), 3), 128)
scores = (act[:, neurons].T + 1) / 2
img[1:, :, 0] = 255 * (1 - scores)
img[1:, :, 1] = 255 * scores
f = BytesIO()
img = scipy.misc.imresize(img, float(cell_size), interp='nearest')
pil_img = PIL.Image.fromarray(img)
draw = ImageDraw.Draw(pil_img)
for idx, ch in enumerate(code):
draw.text((idx * cell_size + 2, 0), ch)
pil_img.save(f, 'png')
return Image(data=f.getvalue())
img = visualize_neurons(neurons, example_code, act)
display(img)
def image_for_code(code):
act = activations(flat_model, code)
neurons = interesting_neurons(act)
return visualize_neurons(neurons, code, act)
display(image_for_code('if (a == 2) and ((b == 1) or (c==2)):'))
code = 'if (a == 2) and ((b == 1) or (c==2)):'
mask = ' ________ ____________________ '
act = activations(flat_model, code)
positive = [idx for idx, ch in enumerate(mask) if ch == '_']
negative = [idx for idx, ch in enumerate(mask) if ch != '_']
neurons = np.argsort(act[positive].sum(axis=0) - act[negative].sum(axis=0))[-5:]
img = visualize_neurons(neurons, code, act)
display(img)
neurons
act[negative, 108].sum()
x0 = 0
x1 = 0
for idx, ch in enumerate(mask):
if ch == '_':
x0 += act[idx, 108]
else:
x1 += act[idx, 108]
x0, x1
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import pickle
import itertools
import xgboost as xgb
from sklearn.svm import LinearSVC
from sklearn.naive_bayes import MultinomialNB
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import SelectFromModel
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer, TfidfVectorizer
from sklearn.metrics import confusion_matrix, f1_score, accuracy_score, precision_score, recall_score
from IPython.display import display
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('data/t2e/text_train.csv')
df = df.append(pd.read_csv('data/t2e/text_test.csv'))
display(df.head())
col = ['label', 'transcription']
df = df[col]
df.columns
df.columns = ['label', 'transcription']
category_to_id = {'ang': 0,
'hap': 1,
'sad': 2,
'fea': 3,
'sur': 4,
'neu': 5}
id_to_category = {0: 'ang', 1: 'hap', 2: 'sad', 3: 'fea', 4: 'sur', 5: 'neu'}
fig = plt.figure(figsize=(8,6))
df.groupby('label').transcription.count().plot.bar(ylim=0)
plt.show()
tfidf = TfidfVectorizer(sublinear_tf=True, min_df=5, norm='l2', encoding='latin-1', ngram_range=(1, 2), stop_words='english')
features = tfidf.fit_transform(df.transcription).toarray()
labels = df.label
print(features.shape)
x_train, x_test, y_train, y_test = train_test_split(features, labels, test_size=0.20)
# count_vect = CountVectorizer()
# x_train_counts = count_vect.fit_transform(x_train)
# tfidf_transformer = TfidfTransformer()
# x_train_tfidf = tfidf_transformer.fit_transform(x_train_counts)
emotion_dict = {'ang': 0,
'hap': 1,
'sad': 2,
'fea': 3,
'sur': 4,
'neu': 5}
emo_keys = list(['ang', 'hap', 'sad', 'fea', 'sur', 'neu'])
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
# plt.figure(figsize=(8,8))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
def one_hot_encoder(true_labels, num_records, num_classes):
temp = np.array(true_labels[:num_records])
true_labels = np.zeros((num_records, num_classes))
true_labels[np.arange(num_records), temp] = 1
return true_labels
def display_results(y_test, pred_probs, cm=True):
pred = np.argmax(pred_probs, axis=-1)
one_hot_true = one_hot_encoder(y_test, len(pred), len(emotion_dict))
print('Test Set Accuracy = {0:.3f}'.format(accuracy_score(y_test, pred)))
print('Test Set F-score = {0:.3f}'.format(f1_score(y_test, pred, average='macro')))
print('Test Set Precision = {0:.3f}'.format(precision_score(y_test, pred, average='macro')))
print('Test Set Recall = {0:.3f}'.format(recall_score(y_test, pred, average='macro')))
if cm:
plot_confusion_matrix(confusion_matrix(y_test, pred), classes=emo_keys)
rf_classifier = RandomForestClassifier(n_estimators=600, min_samples_split=25)
rf_classifier.fit(x_train, y_train)
# Predict
pred_probs = rf_classifier.predict_proba(x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/text_rf_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
xgb_classifier = xgb.XGBClassifier(max_depth=7, learning_rate=0.008, objective='multi:softprob',
n_estimators=600, sub_sample=0.8, num_class=len(emotion_dict),
booster='gbtree', n_jobs=4)
xgb_classifier.fit(x_train, y_train)
# Predict
pred_probs = xgb_classifier.predict_proba(x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/text_xgb_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
svc_classifier = LinearSVC()
svc_classifier.fit(x_train, y_train)
# Predict
pred = svc_classifier.predict(x_test)
# Results
one_hot_true = one_hot_encoder(y_test, len(pred), len(emotion_dict))
print('Test Set Accuracy = {0:.3f}'.format(accuracy_score(y_test, pred)))
print('Test Set F-score = {0:.3f}'.format(f1_score(y_test, pred, average='macro')))
print('Test Set Precision = {0:.3f}'.format(precision_score(y_test, pred, average='macro')))
print('Test Set Recall = {0:.3f}'.format(recall_score(y_test, pred, average='macro')))
plot_confusion_matrix(confusion_matrix(y_test, pred), classes=emotion_dict.keys())
(y_test, pred_probs)
with open('pred_probas/text_svc_classifier_model.pkl', 'wb') as f:
pickle.dump(svc_classifier, f)
mnb_classifier = MultinomialNB()
mnb_classifier.fit(x_train, y_train)
# Predict
pred_probs = mnb_classifier.predict_proba(x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/text_mnb_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
mlp_classifier = MLPClassifier(hidden_layer_sizes=(500, ), activation='relu', solver='adam', alpha=0.0001,
batch_size='auto', learning_rate='adaptive', learning_rate_init=0.01,
power_t=0.5, max_iter=1000, shuffle=True, random_state=None, tol=0.0001,
verbose=False, warm_start=True, momentum=0.8, nesterovs_momentum=True,
early_stopping=False, validation_fraction=0.1, beta_1=0.9, beta_2=0.999,
epsilon=1e-08)
mlp_classifier.fit(x_train, y_train)
# Predict
pred_probs = mlp_classifier.predict_proba(x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/text_mlp_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
lr_classifier = LogisticRegression(solver='lbfgs', multi_class='multinomial', max_iter=1000)
lr_classifier.fit(x_train, y_train)
# Predict
pred_probs = lr_classifier.predict_proba(x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/text_lr_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
ax = xgb.plot_importance(xgb_classifier, max_num_features=10, height=0.5, show_values=False)
contribution_scores = xgb_classifier.feature_importances_
contribution_scores
with open('pred_probas/text_lstm_classifier.pkl', 'rb') as f:
lstm_pred_probs = pickle.load(f)
display_results(y_test, lstm_pred_probs)
# Load predicted probabilities
with open('pred_probas/text_rf_classifier.pkl', 'rb') as f:
rf_pred_probs = pickle.load(f)
with open('pred_probas/text_xgb_classifier.pkl', 'rb') as f:
xgb_pred_probs = pickle.load(f)
with open('pred_probas/text_svc_classifier_model.pkl', 'rb') as f:
svc_preds = pickle.load(f)
with open('pred_probas/text_mnb_classifier.pkl', 'rb') as f:
mnb_pred_probs = pickle.load(f)
with open('pred_probas/text_mlp_classifier.pkl', 'rb') as f:
mlp_pred_probs = pickle.load(f)
with open('pred_probas/text_lr_classifier.pkl', 'rb') as f:
lr_pred_probs = pickle.load(f)
# Average of the predicted probabilites
ensemble_pred_probs = (xgb_pred_probs +
mlp_pred_probs +
rf_pred_probs +
mnb_pred_probs +
lr_pred_probs)/5.0
# Show metrics
display_results(y_test, ensemble_pred_probs)
top_n = [500, 1000, 1500, 2000] # number of features
for n in top_n:
threshold = np.argsort(contribution_scores)[::-1][:n][-1]
print('Stats for top {} features:'.format(n))
# Select features using threshold
selection = SelectFromModel(xgb_classifier, threshold=contribution_scores[threshold], prefit=True)
select_x_train = selection.transform(x_train)
select_x_test = selection.transform(x_test)
# Train
select_xgb_classifier = xgb.XGBClassifier(max_depth=7, learning_rate=0.008, objective='multi:softprob',
n_estimators=600, sub_sample = 0.8, num_class = len(emotion_dict),
booster='gbtree', n_jobs=4)
select_xgb_classifier.fit(select_x_train, y_train)
# Predict
pred_probs = select_xgb_classifier.predict_proba(select_x_test)
# Results
display_results(y_test, pred_probs, cm = False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Norod/hebrew-gpt_neo/blob/main/Demo_of_GRADIO2_Hebrew_GPT_Neo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Using Gradio to wrap a text to text interface around GPT-2
Check out the library on [github](https://github.com/gradio-app/gradio-UI) and see the [getting started](https://gradio.app/getting_started.html) page for more demos.
### Installs and Imports
```
!pip install -q gradio
!pip install -q transformers
!pip install -q Tokenizers
import gradio as gr
import numpy as np
import torch
from transformers import AutoTokenizer, AutoModelForCausalLM
```
### Loading the model and creating the generate function
Note: You can also change to `hebrew-gpt_neo-tiny`, `hebrew-gpt_neo-small` or `hebrew-gpt_neo-xl`
---
> Indented block
```
#model_name = "Norod78/hebrew-gpt_neo-tiny"
model_name = "Norod78/hebrew-gpt_neo-small"
#model_name = "Norod78/hebrew-gpt_neo-xl"
tokenizer = AutoTokenizer.from_pretrained(model_name)
model = AutoModelForCausalLM.from_pretrained(model_name, pad_token_id=tokenizer.eos_token_id)
seed = 1000
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
n_gpu = 0 if torch.cuda.is_available()==False else torch.cuda.device_count()
#print(f"device: {device}, n_gpu: {n_gpu}")
np.random.seed(seed)
torch.manual_seed(seed)
if n_gpu > 0:
torch.cuda.manual_seed_all(seed)
model.to(device)
def generate_text(inp):
encoded_prompt = tokenizer.encode(
inp, add_special_tokens=False, return_tensors="pt")
encoded_prompt = encoded_prompt.to(device)
if encoded_prompt.size()[-1] == 0:
input_ids = None
else:
input_ids = encoded_prompt
#print("input_ids = " + str(input_ids))
beam_output = model.generate(input_ids, max_length=50, num_beams=5, no_repeat_ngram_size=2, early_stopping=True)
output = tokenizer.decode(beam_output[0], skip_special_tokens=True, clean_up_tokenization_spaces=True)
return ".".join(output.split(".")[:-1]) + "."
generate_text('שלום, זאת הדגמה של')
```
###Creating the interface and launching!
```
output_text = gr.outputs.Textbox()
gr.Interface(generate_text,"textbox", output_text, title=model_name,
description="Go ahead and input a sentence and see what it completes \
it with! Takes around 20s to run.").launch(debug=True)
```
#### The model is now live on the gradio.app link shown above. Go ahead and open that in a new tab!
Please contact us [here](mailto:team@gradio.app) if you have any questions, or [open an issue](https://github.com/gradio-app/gradio-UI/issues/new/choose) at our github repo.
| github_jupyter |
# Epicurious Data Prep
## This notebook contains EDA for the Epicurious data
### It also contains functions to prepare the data for word vectorization
```
import json
import csv
import re
import pandas as pd
import numpy as np
import nltk
from nltk.corpus import stopwords
from nltk.stem.wordnet import WordNetLemmatizer
import string
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.metrics.pairwise import cosine_similarity, pairwise_distances
from sklearn.feature_extraction.text import TfidfVectorizer
```
## Using some stopwords from https://github.com/AlludedCrabb/sound-tasty
```
stopwords_list
```
# Define functions to use
```
def cuisine_namer(text):
if text == 'Central American/Caribbean':
return 'Caribbean'
elif text == 'Jewish':
return 'Kosher'
elif text == 'Eastern European/Russian':
return 'Eastern European'
elif text in ['Spanish/Portuguese', 'Greek']:
return 'Mediterranean'
elif text == 'Central/South American':
return 'Latin American'
elif text == 'Sushi':
return 'Japanese'
elif text == 'Southern Italian':
return 'Italian'
elif text in ['Southern', 'Tex-Mex']:
return 'American'
elif text in ['Southeast Asian', 'Korean']:
return 'Asian'
else:
return text
filename = "../raw_data/recipes-en-201706/epicurious-recipes_m2.json"
with open(filename, 'r') as f:
datastore = json.load(f)
def load_data(filepath, test_size=0.1, random_state=10):
""" This function uses a filepath, test_size, and random_state
to load the Epicurious JSON into a dataframe and then split into
train/test sets."""
with open(filepath, 'r') as f:
datastore = json.load(f)
datastore_df = pd.DataFrame(datastore)
X_train, X_test = train_test_split(datastore_df,
test_size=test_size,
random_state=random_state)
return X_train, X_test
def prep_data(X):
""" This function takes a dataframe X, drops columns that will not be used,
expands the hierarchical column into the dataframe, renames the columns
to be more human-readable, and drops one column created during dataframe
expansion"""
X.drop(['pubDate', 'author', 'type', 'aggregateRating', 'reviewsCount',
'willMakeAgainPct', 'dateCrawled'],
axis=1,
inplace=True)
concat = pd.concat([X.drop(['tag'], axis=1),
X['tag'].apply(pd.Series)],
axis=1)
concat.drop([0, 'photosBadgeAltText', 'photosBadgeFileName', 'photosBadgeID',
'photosBadgeRelatedUri'],
axis=1,
inplace=True)
cols = ['id', 'description', 'title', 'url', 'photo_data', 'ingredients',
'steps', 'category', 'name', 'remove']
concat.columns = cols
concat.drop('remove', axis=1, inplace=True)
cuisine_only = concat[concat['category'] == 'cuisine']
cuisine_only.dropna(axis=0, inplace=True)
cuisine_only['imputed_label'] = cuisine_only['name'].apply(cuisine_namer)
return cuisine_only
def fit_transform_tfidf_matrix(X_df, stopwords_list):
tfidf = sklearn.feature_extraction.text.TfidfVectorizer(stop_words=stopwords_list,
min_df=2,
token_pattern=r'(?u)\b[a-zA-Z]{2,}\b',
preprocessor=lemmatizer.lemmatize,
)
temp = X_df['ingredients'].apply(' '.join).str.lower()
tfidf.fit(temp)
response = tfidf.transform(temp)
print(response.shape)
word_matrix = pd.DataFrame(response.toarray(),
columns=tfidf.get_feature_names(),
index=X_df.index)
return tfidf, word_matrix
def transform_tfidf(tfidf, recipe):
response = tfidf.transform(recipe['ingredients'])
transformed_recipe = pd.DataFrame(response.toarray(),
columns=tfidf.get_feature_names(),
index=recipe.index)
return transformed_recipe
def transform_from_test_tfidf(tfidf, df, idx):
recipe = [' '.join(df.iloc[idx]['ingredients'])]
response = tfidf.transform(recipe)
transformed_recipe = pd.DataFrame(response.toarray(),
columns=tfidf.get_feature_names())
return transformed_recipe
def filter_out_cuisine(ingred_word_matrix, X_df, cuisine_name, tfidf):
combo = pd.concat([ingred_word_matrix, X_df['imputed_label']], axis=1)
filtered_ingred_word_matrix = combo[combo['imputed_label'] != cuisine_name].drop('imputed_label',
axis=1)
return filtered_ingred_word_matrix
def find_closest_recipes(filtered_ingred_word_matrix, recipe_tfidf, X_df):
search_vec = np.array(recipe_tfidf).reshape(1,-1)
res_cos_sim = cosine_similarity(filtered_ingred_word_matrix, search_vec)
top_five = np.argsort(res_cos_sim.flatten())[-5:][::-1]
proximity = res_cos_sim[top_five]
recipe_ids = [filtered_ingred_word_matrix.iloc[idx].name for idx in top_five]
suggest_df = X_df.loc[recipe_ids]
return suggest_df, proximity
```
# Create the dataframe
```
X_train, X_test = load_data(filename)
prepped = prep_data(X_train)
prepped.head()
print(prepped['ingredients'].apply(' '.join))
```
# Create the ingredients TFIDF matrix that will be the database
```
ingred_tfidf, ingred_word_matrix = fit_transform_tfidf_matrix(prepped, stopwords_list)
ingred_word_matrix
```
# X_test is using the test split from train_test_split to return test recipes
```
X_test
test_prepped = prep_data(X_test)
test_prepped.head()
sample_recipe = test_prepped.iloc[300]
sample_recipe
sample_recipe['ingredients']
remove_cuisine = sample_recipe['imputed_label']
remove_cuisine
sample_words = transform_from_test_tfidf(ingred_tfidf, test_prepped, 300)
sample_words
```
# Make a sub DataFrame without the target's cuisine style
```
filtered_ingred_matrix = filter_out_cuisine(ingred_word_matrix,
prepped,
remove_cuisine,
ingred_tfidf)
filtered_ingred_matrix
```
# Calculate Cosine Similarity between provided recipe and database
```
filtered_ingred_matrix.iloc[10201].name
res_cos_sim, proximity = find_closest_recipes(filtered_ingred_matrix,
sample_words,
prepped)
res_cos_sim
proximity
sample_recipe['ingredients'], [res_cos_sim['ingredients'].iloc[idx] for idx in range(0,5)]
avo_pesto_query = [27135, 19328, 2031, 11527, 26330]
prepped.loc[avo_pesto_query]
prepped.loc[27135]['ingredients']
```
---
```
brint = 74
brian_tries = test_prepped.iloc[brint]
brian_tries
brian_sample_words = transform_from_test_tfidf(ingred_tfidf,
test_prepped,
brint)
brian_filtered_ingred_matrix = filter_out_cuisine(ingred_word_matrix,
prepped,
remove_cuisine,
ingred_tfidf)
br_res_cos_sim, br_proximity = find_closest_recipes(brian_filtered_ingred_matrix,
brian_sample_words,
prepped)
br_res_cos_sim
br_proximity
br_res_cos_sim['ingredients'].iloc[4]
```
---
```
eint = 200
erin_tries = test_prepped.iloc[eint]
erin_tries
erin_sample_words = transform_from_test_tfidf(ingred_tfidf,
test_prepped,
eint)
erin_filtered_ingred_matrix = filter_out_cuisine(ingred_word_matrix,
prepped,
remove_cuisine,
ingred_tfidf)
er_res_cos_sim, er_proximity = find_closest_recipes(erin_filtered_ingred_matrix,
erin_sample_words,
prepped)
er_res_cos_sim
er_proximity
```
---
```
cint = 3
c_tries = test_prepped.iloc[cint]
c_tries
c_sample_words = transform_from_test_tfidf(ingred_tfidf,
test_prepped,
cint)
c_filtered_ingred_matrix = filter_out_cuisine(ingred_word_matrix,
prepped,
remove_cuisine,
ingred_tfidf)
c_res_cos_sim, c_proximity = find_closest_recipes(c_filtered_ingred_matrix,
c_sample_words,
prepped)
c_res_cos_sim
```
| github_jupyter |
# Optimizing Multiple Objectives using Vertex Vizier
## Overview
This tutorial demonstrates [Vertex Vizier](https://cloud.google.com/vertex-ai/docs/vizier/overview) multi-objective optimization. Multi-objective optimization is concerned with mathematical optimization problems involving more than one objective function to be optimized simultaneously
## Objective
The goal is to __`minimize`__ the objective metric:
```
y1 = r*sin(theta)
```
and simultaneously __`maximize`__ the objective metric:
```
y2 = r*cos(theta)
```
that you will evaluate over the parameter space:
- __`r`__ in [0,1],
- __`theta`__ in [0, pi/2]
## Introduction
In this notebook, you will use [Vertex Vizier](https://cloud.google.com/vertex-ai/docs/vizier/overview) multi-objective optimization. Multi-objective optimization is concerned with mathematical optimization problems involving more than one objective function to be optimized simultaneously.
Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/gapic-vizier-multi-objective-optimization.ipynb).
**Make sure to enable the Vertex AI API**
#### Install Vertex AI library
Download and install Vertex AI library.
```
# Setup your dependencies
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
# Upgrade the specified package to the newest available version
! pip install {USER_FLAG} --upgrade google-cloud-aiplatform
import os
if not os.getenv("IS_TESTING"):
# Restart the kernel after pip installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
### Import libraries and define constants
```
# Import necessary libraries
import datetime
import json
from google.cloud import aiplatform_v1beta1
```
## Tutorial
This section defines some parameters and util methods to call Vertex Vizier APIs. Please fill in the following information to get started.
```
# Fill in your project ID and region
REGION = "[region]" # @param {type:"string"}
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# These will be automatically filled in.
STUDY_DISPLAY_NAME = "{}_study_{}".format(
PROJECT_ID.replace("-", ""), datetime.datetime.now().strftime("%Y%m%d_%H%M%S")
) # @param {type: 'string'}
ENDPOINT = REGION + "-aiplatform.googleapis.com"
PARENT = "projects/{}/locations/{}".format(PROJECT_ID, REGION)
print("ENDPOINT: {}".format(ENDPOINT))
print("REGION: {}".format(REGION))
print("PARENT: {}".format(PARENT))
# If you don't know your project ID, you might be able to get your project ID
# using gcloud command by executing the second cell below.
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
### Create the study configuration
The following is a sample study configuration, built as a hierarchical python dictionary. It is already filled out. Run the cell to configure the study.
```
# Parameter Configuration
param_r = {"parameter_id": "r", "double_value_spec": {"min_value": 0, "max_value": 1}}
param_theta = {
"parameter_id": "theta",
"double_value_spec": {"min_value": 0, "max_value": 1.57},
}
# Objective Metrics
metric_y1 = # TODO -- Your code goes here
# Objective Metrics
metric_y2 = # TODO -- Your code goes here
# Put it all together in a study configuration
study = {
"display_name": STUDY_DISPLAY_NAME,
"study_spec": {
"algorithm": "RANDOM_SEARCH",
"parameters": [
param_r,
param_theta,
],
"metrics": [metric_y1, metric_y2],
},
}
print(json.dumps(study, indent=2, sort_keys=True))
```
### Create the study
Next, create the study, which you will subsequently run to optimize the two objectives.
```
# Create the study using study configuration and send request through VizierServiceClient
vizier_client = # TODO -- Your code goes here(
client_options=dict(api_endpoint=ENDPOINT)
)
study = vizier_client.create_study(parent=PARENT, study=study)
STUDY_ID = study.name
print("STUDY_ID: {}".format(STUDY_ID))
```
### Metric evaluation functions
Next, define some functions to evaluate the two objective metrics.
```
import math
# r * sin(theta)
def Metric1Evaluation(r, theta):
"""Evaluate the first metric on the trial."""
return r * math.sin(theta)
# r * cos(theta)
def Metric2Evaluation(r, theta):
"""Evaluate the second metric on the trial."""
return r * math.cos(theta)
def CreateMetrics(trial_id, r, theta):
print(("=========== Start Trial: [{}] =============").format(trial_id))
# Evaluate both objective metrics for this trial
y1 = # TODO -- Your code goes here(r, theta)
y2 = # TODO -- Your code goes here(r, theta)
print(
"[r = {}, theta = {}] => y1 = r*sin(theta) = {}, y2 = r*cos(theta) = {}".format(
r, theta, y1, y2
)
)
metric1 = {"metric_id": "y1", "value": y1}
metric2 = {"metric_id": "y2", "value": y2}
# Return the results for this trial
return [metric1, metric2]
```
### Set configuration parameters for running trials
__`client_id`__: The identifier of the client that is requesting the suggestion. If multiple SuggestTrialsRequests have the same `client_id`, the service will return the identical suggested trial if the trial is `PENDING`, and provide a new trial if the last suggested trial was completed.
__`suggestion_count_per_request`__: The number of suggestions (trials) requested in a single request.
__`max_trial_id_to_stop`__: The number of trials to explore before stopping. It is set to 4 to shorten the time to run the code, so don't expect convergence. For convergence, it would likely need to be about 20 (a good rule of thumb is to multiply the total dimensionality by 10).
```
client_id = "client1" # @param {type: 'string'}
suggestion_count_per_request = 5 # @param {type: 'integer'}
max_trial_id_to_stop = 4 # @param {type: 'integer'}
print("client_id: {}".format(client_id))
print("suggestion_count_per_request: {}".format(suggestion_count_per_request))
print("max_trial_id_to_stop: {}".format(max_trial_id_to_stop))
```
### Run Vertex Vizier trials
Run the trials.
```
trial_id = 0
while int(trial_id) < max_trial_id_to_stop:
suggest_response = vizier_client.suggest_trials(
{
"parent": STUDY_ID,
"suggestion_count": suggestion_count_per_request,
"client_id": client_id,
}
)
for suggested_trial in suggest_response.result().trials:
trial_id = suggested_trial.name.split("/")[-1]
trial = vizier_client.get_trial({"name": suggested_trial.name})
if trial.state in ["COMPLETED", "INFEASIBLE"]:
continue
for param in trial.parameters:
if param.parameter_id == "r":
r = param.value
elif param.parameter_id == "theta":
theta = param.value
print("Trial : r is {}, theta is {}.".format(r, theta))
# Store your measurement and send the request
# TODO -- Your code goes here(
{
"trial_name": suggested_trial.name,
"measurement": {
"metrics": # TODO -- Your code goes here(suggested_trial.name, r, theta)
},
}
)
response = vizier_client.complete_trial(
{"name": suggested_trial.name, "trial_infeasible": False}
)
```
### List the optimal solutions
list_optimal_trials returns the pareto-optimal Trials for multi-objective Study or the optimal Trials for single-objective Study. In the case, we define mutliple-objective in previeous steps, pareto-optimal trials will be returned.
```
# List all the pareto-optimal trails
optimal_trials = # TODO -- Your code goes here({"parent": STUDY_ID})
print("optimal_trials: {}".format(optimal_trials))
```
## Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial. You can also manually delete resources that you created by running the following code.
```
vizier_client.delete_study({"name": STUDY_ID})
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_08_3_keras_hyperparameters.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 8: Kaggle Data Sets**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 8 Material
* Part 8.1: Introduction to Kaggle [[Video]](https://www.youtube.com/watch?v=v4lJBhdCuCU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_1_kaggle_intro.ipynb)
* Part 8.2: Building Ensembles with Scikit-Learn and Keras [[Video]](https://www.youtube.com/watch?v=LQ-9ZRBLasw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_2_keras_ensembles.ipynb)
* **Part 8.3: How Should you Architect Your Keras Neural Network: Hyperparameters** [[Video]](https://www.youtube.com/watch?v=1q9klwSoUQw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_3_keras_hyperparameters.ipynb)
* Part 8.4: Bayesian Hyperparameter Optimization for Keras [[Video]](https://www.youtube.com/watch?v=sXdxyUCCm8s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_4_bayesian_hyperparameter_opt.ipynb)
* Part 8.5: Current Semester's Kaggle [[Video]](https://www.youtube.com/watch?v=PHQt0aUasRg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_5_kaggle_project.ipynb)
# Google CoLab Instructions
The following code ensures that Google CoLab is running the correct version of TensorFlow.
```
# Startup CoLab
try:
%tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
```
# Part 8.3: How Should you Architect Your Keras Neural Network: Hyperparameters
* [Guide to choosing Hyperparameters for your Neural Networks](https://towardsdatascience.com/guide-to-choosing-hyperparameters-for-your-neural-networks-38244e87dafe)
### Number of Hidden Layers and Neuron Counts
* [Keras Layers](https://keras.io/layers/core/)
Layer types and when to use them:
* **Activation** - Layer that simply adds an activation function, the activation function can also be specified as part of a Dense (or other) layer type.
* **ActivityRegularization** Used to add L1/L2 regularization outside of a layer. L1 and L2 can also be specified as part of a Dense (or other) layer type.
* **Dense** - The original neural network layer type. Every neuron is connected to the next layer. The input vector is one-dimensional and placing certain inputs next to each other does not have an effect.
* **Dropout** - Dropout consists in randomly setting a fraction rate of input units to 0 at each update during training time, which helps prevent overfitting. Dropout only occurs during training.
* **Flatten** - Flattens the input to 1D. Does not affect the batch size.
* **Input** - A Keras tensor is a tensor object from the underlying backend (Theano, TensorFlow or CNTK), which we augment with certain attributes that allow us to build a Keras model just by knowing the inputs and outputs of the model.
* **Lambda** - Wraps arbitrary expression as a Layer object.
* **Masking** - Masks a sequence by using a mask value to skip timesteps.
* **Permute** - Permutes the dimensions of the input according to a given pattern. Useful for e.g. connecting RNNs and convnets together.
* **RepeatVector** - Repeats the input n times.
* **Reshape** - Similar to Numpy reshapes.
* **SpatialDropout1D** - This version performs the same function as Dropout, however it drops entire 1D feature maps instead of individual elements.
* **SpatialDropout2D** - This version performs the same function as Dropout, however it drops entire 2D feature maps instead of individual elements
* **SpatialDropout3D** - This version performs the same function as Dropout, however it drops entire 3D feature maps instead of individual elements.
### Activation Functions
* [Keras Activation Functions](https://keras.io/activations/)
* [Activation Function Cheat Sheets](https://ml-cheatsheet.readthedocs.io/en/latest/activation_functions.html)
As follows:
* **softmax** - Used for multi-class classification. Ensures all output neurons behave as probabilities and sum to 1.0.
* **elu** - Exponential linear unit. Exponential Linear Unit or its widely known name ELU is a function that tend to converge cost to zero faster and produce more accurate results. Can produce negative outputs.
* **selu** - Scaled Exponential Linear Unit (SELU), essentially **elu** multiplied by a scaling constant.
* **softplus** - Softplus activation function. $log(exp(x) + 1)$ [Introduced](https://papers.nips.cc/paper/1920-incorporating-second-order-functional-knowledge-for-better-option-pricing.pdf) in 2001.
* **softsign** Softsign activation function. $x / (abs(x) + 1)$ Similar to tanh, but not widely used.
* **relu** - Very popular neural network activation function. Used for hidden layers, cannot output negative values. No trainable parameters.
* **tanh** Classic neural network activation function, though often replaced by relu family on modern networks.
* **sigmoid** - Classic neural network activation. Often used on output layer of a binary classifier.
* **hard_sigmoid** - Less computationally expensive variant of sigmoid.
* **exponential** - Exponential (base e) activation function.
* **linear** - Pass through activation function. Usually used on the output layer of a regression neural network.
### Advanced Activation Functions
* [Keras Advanced Activation Functions](https://keras.io/layers/advanced-activations/)
The advanced activation functions contain parameters that are trained during neural network fitting. As follows:
* **LeakyReLU** - Leaky version of a Rectified Linear Unit. It allows a small gradient when the unit is not active, controlled by alpha hyperparameter.
* **PReLU** - Parametric Rectified Linear Unit, learns the alpha hyperparameter.
### Regularization: L1, L2, Dropout
* [Keras Regularization](https://keras.io/regularizers/)
* [Keras Dropout](https://keras.io/layers/core/)
### Batch Normalization
* [Keras Batch Normalization](https://keras.io/layers/normalization/)
* Ioffe, S., & Szegedy, C. (2015). [Batch normalization: Accelerating deep network training by reducing internal covariate shift](https://arxiv.org/abs/1502.03167). *arXiv preprint arXiv:1502.03167*.
Normalize the activations of the previous layer at each batch, i.e. applies a transformation that maintains the mean activation close to 0 and the activation standard deviation close to 1. Can allow learning rate to be larger.
### Training Parameters
* [Keras Optimizers](https://keras.io/optimizers/)
* **Batch Size** - Usually small, such as 32 or so.
* **Learning Rate** - Usually small, 1e-3 or so.
```
import pandas as pd
from scipy.stats import zscore
# Read the data set
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/jh-simple-dataset.csv",
na_values=['NA','?'])
# Generate dummies for job
df = pd.concat([df,pd.get_dummies(df['job'],prefix="job")],axis=1)
df.drop('job', axis=1, inplace=True)
# Generate dummies for area
df = pd.concat([df,pd.get_dummies(df['area'],prefix="area")],axis=1)
df.drop('area', axis=1, inplace=True)
# Missing values for income
med = df['income'].median()
df['income'] = df['income'].fillna(med)
# Standardize ranges
df['income'] = zscore(df['income'])
df['aspect'] = zscore(df['aspect'])
df['save_rate'] = zscore(df['save_rate'])
df['age'] = zscore(df['age'])
df['subscriptions'] = zscore(df['subscriptions'])
# Convert to numpy - Classification
x_columns = df.columns.drop('product').drop('id')
x = df[x_columns].values
dummies = pd.get_dummies(df['product']) # Classification
products = dummies.columns
y = dummies.values
import pandas as pd
import os
import numpy as np
import time
import tensorflow.keras.initializers
import statistics
import tensorflow.keras
from sklearn import metrics
from sklearn.model_selection import StratifiedKFold
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Dropout
from tensorflow.keras import regularizers
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.model_selection import StratifiedShuffleSplit
from tensorflow.keras.layers import LeakyReLU,PReLU
from tensorflow.keras.optimizers import Adam
def evaluate_network(dropout,lr,neuronPct,neuronShrink):
SPLITS = 2
# Bootstrap
boot = StratifiedShuffleSplit(n_splits=SPLITS, test_size=0.1)
# Track progress
mean_benchmark = []
epochs_needed = []
num = 0
neuronCount = int(neuronPct * 5000)
# Loop through samples
for train, test in boot.split(x,df['product']):
start_time = time.time()
num+=1
# Split train and test
x_train = x[train]
y_train = y[train]
x_test = x[test]
y_test = y[test]
# Construct neural network
# kernel_initializer = tensorflow.keras.initializers.he_uniform(seed=None)
model = Sequential()
layer = 0
while neuronCount>25 and layer<10:
#print(neuronCount)
if layer==0:
model.add(Dense(neuronCount,
input_dim=x.shape[1],
activation=PReLU()))
else:
model.add(Dense(neuronCount, activation=PReLU()))
model.add(Dropout(dropout))
neuronCount = neuronCount * neuronShrink
model.add(Dense(y.shape[1],activation='softmax')) # Output
model.compile(loss='categorical_crossentropy', optimizer=Adam(lr=lr))
monitor = EarlyStopping(monitor='val_loss', min_delta=1e-3,
patience=100, verbose=0, mode='auto', restore_best_weights=True)
# Train on the bootstrap sample
model.fit(x_train,y_train,validation_data=(x_test,y_test),
callbacks=[monitor],verbose=0,epochs=1000)
epochs = monitor.stopped_epoch
epochs_needed.append(epochs)
# Predict on the out of boot (validation)
pred = model.predict(x_test)
# Measure this bootstrap's log loss
y_compare = np.argmax(y_test,axis=1) # For log loss calculation
score = metrics.log_loss(y_compare, pred)
mean_benchmark.append(score)
m1 = statistics.mean(mean_benchmark)
m2 = statistics.mean(epochs_needed)
mdev = statistics.pstdev(mean_benchmark)
# Record this iteration
time_took = time.time() - start_time
tensorflow.keras.backend.clear_session()
return (-m1)
print(evaluate_network(
dropout=0.2,
lr=1e-3,
neuronPct=0.2,
neuronShrink=0.2))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.