text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Title
The title of the notebook should be coherent with file name. Namely, file name should be:
*author's initials_progressive number_title.ipynb*
For example:
*EF_01_Data Exploration.ipynb*
## Purpose
State the purpose of the notebook.
## Methodology
Quickly describe assumptions and processing steps.
## WIP - improvements
Use this section only if the notebook is not final.
Notable TODOs:
- todo 1;
- todo 2;
- todo 3.
## Results
Describe and comment the most important results.
## Suggested next steps
State suggested next steps, based on results obtained in this notebook.
# Setup
## Library import
We import all the required Python libraries
```
# Data manipulation
import pandas as pd
import numpy as np
# Options for pandas
pd.options.display.max_columns = 50
pd.options.display.max_rows = 30
import matplotlib.pyplot as plt
# Autoreload extension
if 'autoreload' not in get_ipython().extension_manager.loaded:
%load_ext autoreload
%autoreload 2
```
## Local library import
We import all the required local libraries libraries
```
# Include local library paths
import sys
# sys.path.append('path/to/local/lib') # uncomment and fill to import local libraries
# Import local libraries
```
# Parameter definition
We set all relevant parameters for our notebook. By convention, parameters are uppercase, while all the
other variables follow Python's guidelines.
# Data import
We retrieve all the required data for the analysis.
# Data processing
Put here the core of the notebook. Feel free di further split this section into subsections.
```
fp = r"C:\Users\sacchi_r\Documents\GitHub\carculator_bus\dev\EFA_HOT_Subsegm_HGV.XLSX"
df = pd.read_excel(fp)
df.head()
import matplotlib.pyplot as plt
from matplotlib import rcParams
import numpy as np
rcParams.update({'figure.autolayout': True})
#fig, axs = plt.subplots(figsize=(12,10),
# nrows=4, ncols=3, # fix as above
# gridspec_kw=dict(hspace=0.2),
# sharey=True) # Much control of gridspec
powertrains = [
#"diesel",
"CNG"
]
subsegments = [
"TT/AT CNG Euro-VI",
"TT/AT CNG Euro-IV",
"TT/AT CNG Euro-V",
]
components = df['Component'].unique().tolist()
euro_classes = df["EmConcept"].unique().tolist()
sizes = [
'RT ≤7,5t',
'RT >7,5-12t',
'RT >12t',
'RT >14-20t',
'RT >26-28t',
'TT/AT >40-50t',
'TT/AT >50-60t',
'TT/AT not specified'
]
idx=pd.IndexSlice
l_res = []
for sub in subsegments:
for component in components:
data = df.loc[(df["Subsegment"]==sub)
&(df["Component"]==component), ["V", "EFA"]]
ind = data["V"].values
vals = data["EFA"].values
if len(data)>0:
z = np.polyfit(ind, vals, 3)
f = np.poly1d(z)
l_res.append([sub, component, f[0], f[1], f[2], f[3]])
#print(component, f, z)
# calculate new x's and y's
x_new = np.linspace(0, 150, 50)
y_new = np.clip(f(x_new), 0, None)
#ax = plt.subplot(5, 3, components.index(component)+1)
#ax = plt.subplot(5, 3, components.index(component)+1)
#ax.plot(ind, vals, marker=".", linestyle="")
#ax.plot(x_new, y_new, linestyle="-", label=sub)
#ax.set_title(component)
#if components.index(component)==12:
# ax.legend(ncol=3)
#pd.DataFrame(l_res, columns=["powertrain", "euro_class", "size", "component", "intercept", "c", "b", "a"]).to_excel("HEM_factors_trucks.xlsx")
l_res
pd.DataFrame(l_res).to_excel("hbefa_32t_cng.xlsx")
pd.DataFrame(l_res).to_excel("CNG_factors.xlsx")
```
# References
We report here relevant references:
1. author1, article1, journal1, year1, url1
2. author2, article2, journal2, year2, url2
| github_jupyter |
<img src="https://github.com/pmservice/ai-openscale-tutorials/raw/master/notebooks/images/banner.png" align="left" alt="banner">
# Notebook for generating configuration for online subscriptions in IBM Watson OpenScale
This notebook shows how to generate the following artefacts:
1. Configuration JSON needed to configure an IBM Watson OpenScale online subscription. This JSON also contains information related to fairness configuration.
2. Drift Configuration Archive
In order to use this notebook you need to do the following:
1. Read the training data into a pandas dataframe called "data_df". There is sample code below to show how this can be done if the training data is in IBM Cloud Object Storage.
2. Edit the below cells and provide the training data and fairness configuration information.
3. Run the notebook. It will generate a JSON and a download link for the JSON will be present at the very end of the notebook.
4. Download the JSON by clicking on the link and upload it in the IBM AI OpenScale GUI.
If you have multiple models (deployments), you will have to repeat the above steps for each model (deployment).
**Note:** Please restart the kernel after executing below cell
```
!pip install pandas
!pip install ibm-cos-sdk
!pip install numpy
!pip install pyspark
!pip install lime
!pip install --upgrade ibm-watson-openscale
!pip install "ibm-wos-utils==4.0.24"
```
# Read training data into a pandas data frame
The first thing that you need to do is to read the training data into a pandas dataframe called "data_df". Given below is sample code for doing this if the training data is in IBM Cloud Object Storage. Please edit the below cell and make changes so that you can read your training data from the location where it is stored. Please ensure that the training data is present in a data frame called "data_df".
*Note: Pandas' read\_csv method converts the columns to its data types. If you want the column type to not be interpreted, specify the dtype param to read_csv method in this cell. More on this method [here](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html)*
*Note: By default NA values will be dropped while computing training data distribution and training the drift archive. Please ensure to handle the NA values during Pandas' read\_csv method*
```
# ----------------------------------------------------------------------------------------------------
# IBM Confidential
# OCO Source Materials
# 5900-A3Q, 5737-H76
# Copyright IBM Corp. 2018, 2021
# The source code for this Notebook is not published or other-wise divested of its trade
# secrets, irrespective of what has been deposited with the U.S.Copyright Office.
# ----------------------------------------------------------------------------------------------------
VERSION = "5.3.1"
# code to read file in COS to pandas dataframe object
import sys
import types
import pandas as pd
from ibm_botocore.client import Config
import ibm_boto3
def __iter__(self): return 0
api_key = "<API Key>"
resource_instance_id = "<COS Resource Instance ID>"
auth_endpoint = "https://iam.ng.bluemix.net/oidc/token"
service_endpoint = "<COS Service Endpoint>"
bucket = "<Bucket Name>"
file_name= "<File Name>"
cos_client = ibm_boto3.client(service_name="s3",
ibm_api_key_id=api_key,
ibm_auth_endpoint=auth_endpoint,
config=Config(signature_version="oauth"),
endpoint_url=service_endpoint)
body = cos_client.get_object(Bucket=bucket,Key=file_name)["Body"]
# add missing __iter__ method, so pandas accepts body as file-like object
if not hasattr(body, "__iter__"): body.__iter__ = types.MethodType( __iter__, body )
data_df = pd.read_csv(body)
data_df.head()
#Print columns from data frams
#print("column names:{}".format(list(data_df.columns.values)))
# Uncomment following 2 lines if you want to read training data from local CSV file when running through local Jupyter notebook
#data_df = pd.read_csv("<FULLPATH_TO_CSV_FILE>")
#data_df.head()
```
## Select the services for which configuration information needs to be generated
This notebook has support to generaton configuration information related to fairness , explainability and drift service. The below can be used by the user to control service specific configuration information.
Details of the service speicifc flags available:
- enable_fairness : Flag to allow generation of fairness specific data distribution needed for configuration
- enable_explainability : Flag to allow generation of explainability specific information
- enable_drift: Flag to allow generation of drift detection model needed by drift service
service_configuration_support = { <br>
"enable_fairness": True,
"enable_explainability": True,
"enable_drift": False
}
```
service_configuration_support = {
"enable_fairness": True,
"enable_explainability": True,
"enable_drift": True
}
```
## Training Data and Fairness Configuration Information
Please provide information about the training data which is used to train the model. In order to explain the configuration better, let us first consider an example of a Loan Processing Model which is trying to predict whether a person should get a loan or not. The training data for such a model will potentially contain the following columns: Credit_History, Monthly_salary, Applicant_Age, Loan_amount, Gender, Marital_status, Approval. The "Approval" column contains the target field (label column or class label) and it can have the following values: "Loan Granted", "Loan Denied" or "Loan Partially Granted". In this model we would like to ensure that the model is not biased against Gender=Female or Gender=Transgender. We would also like to ensure that the model is not biased against the age group 15 to 30 years or age group 61 to 120 years.
For the above model, the configuration information that we need to provide is:
- class_label: This is the name of the column in the training data dataframe (data_df) which contains the target field (also known as label column or the class label). For the Loan Processing Model it would be "Approval".
- feature_columns: This is a comma separated list of column names which contain the feature column names (in the training data dataframe data_df). For the Loan Processing model this would be: ["Credit_History", "Monthly_salary", "Applicant_Age", "Loan_amount", "Gender", "Marital_status"]
- categorical_columns: The list of column names (in data_df) which contain categorical values. This should also include those columns which originally contained categorical values and have now been converted to numeric values. E.g., in the Loan Processing Model, the Marital_status column originally could have values: Single, Married, Divorced, Separated, Widowed. These could have been converted to numeric values as follows: Single -> 0, Married -> 1, Divorced -> 2, Separated -> 3 and Widowed -> 4. Thus the training data will have numeric values. Please identify such columns as categorical. Thus the list of categorical columns for the Loan Processing Model will be Credit_History, Gender and Marital_status.
For the Loan Processing Model, this information will be provided as follows:
training_data_info = { <br>
"class_label": "Approval",
"feature_columns": ["Credit_History", "Monthly_salary", "Applicant_Age", "Loan_amount", "Gender", "Marital_status"],
"categorical_columns": ["Credit_History","Gender","Marital_status"]
}
**Note:** Please note that categorical columns selected should be subset of feature columns. If there are no categorical columns among the feature columns selected , please set "categorical_columns as [] or None"
Please edit the next cell and provide the above information for your model.
```
training_data_info = {
"class_label": "<EDIT THIS>",
"feature_columns": ["<EDIT THIS>"],
"categorical_columns": ["<EDIT THIS>"]
}
```
## Specify the Model Type
In the next cell, specify the type of your model. If your model is a binary classification model, then set the type to "binary". If it is a multi-class classifier then set the type to "multiclass". If it is a regression model (e.g., Linear Regression), then set it to "regression".
```
#Set model_type. Acceptable values are:["binary","multiclass","regression"]
model_type = "binary"
#model_type = "multiclass"
#model_type = "regression"
```
## Specify the Fairness Configuration
You need to provide the following information for the fairness configuration:
- fairness_attributes: These are the attributes on which you wish to monitor fairness. In the Loan Processing Model, we wanted to ensure that the model is not baised against people of specific age group and people belonging to a specific gender. Hence "Applicant_Age" and "Gender" will be the fairness attributes for the Loan Processing Model.
- With Indirect Bias support, you can also monitor protected attributes for fairness. The protected attributes are those attributes which are present in the training data but are not used to train the model. For example, sensitive attributes like gender, race, age may be present in training data but are not used for training. To check if there exists indirect bias with respect to some protected attribute due to possible correlation with some feature column, it can be specified in fairness configuration.
- type: The data type of the fairness attribute (e.g., float or int or double)
- minority: The minority group for which we want to ensure that the model is not biased. For the Loan Processing Model we wanted to ensure that the model is not biased against people in the age group 15 to 30 years & 61 to 120 years as well as people with Gender = Female or Gender = Transgender. Hence the minority group for the fairness attribute "Applicant_Age" will be [15,30] and [61,120] and the minority group for fairness attribute "Gender" will be: "Female", "Transgender".
- majority: The majority group for which the model might be biased towards. For the Loan Processing Model, the majority group for the fairness attribute "Applicant_Age" will be [31,60], i.e., all the ages except the minority group. For the fairness attribute "Gender" the majority group will be: "Male".
- threshold: The fairness threshold beyond which the Model is considered to be biased. For the Loan Processing Model, let us say that the Bank is willing to tolerate the fact that Female and Transgender applicants will get upto 20% lesser approved loans than Males. However, if the percentage is more than 20% then the Loan Processing Model will be considered biased. E.g., if the percentage of approved loans for Female or Transgender applicants is say 25% lesser than those approved for Male applicants then the Model is to be considered as acting in a biased manner. Thus for this scenario, the Fairness threshold will be 80 (100-20) (this is represented as a value normalized to 1, i.e., 0.8).
The fairness attributes for Loan Processing Model will be specified as:
fairness_attributes = [
{
"feature": "Applicant_Age",
"type" : "int",
"majority": [ [31,60] ],
"minority": [ [15, 30], [61,120] ],
"threshold" : 0.8
},
{
"feature": "Gender",
"type" : "string",
"majority": ["Male"],
"minority": ["Female", "Transgender"],
"threshold" : 0.8
}
]
Please edit the next cell and provide the fairness configuration for your model.
```
fairness_attributes = [{
"type" : "<DATA_TYPE>", #data type of the column eg: float or int or double
"feature": "<COLUMN_NAME>",
"majority": [
[X, Y] # range of values for column eg: [31, 45] for int or [31.4, 45.1] for float
],
"minority": [
[A, B], # range of values for column eg: [10, 15] for int or [10.5, 15.5] for float
[C, D] # range of values for column eg: [80, 100] for int or [80.0, 99.9] for float
],
"threshold": <VALUE> #such that 0<VALUE<=1. eg: 0.8
}]
```
## Specify the Favorable and Unfavorable class values
The second part of fairness configuration is about the favourable and unfavourable class values. Recall that in the case of Loan Processing Model, the target field (label column or class label) can have the following values: "Loan Granted", "Loan Denied" and "Loan Partially Granted". Out of these values "Loan Granted" and "Loan Partially Granted" can be considered as being favorable and "Loan Denied" is unfavorable. In other words in order to measure fairness, we need to know the target field values which can be considered as being favourable and those values which can be considered as unfavourable.
For the Loan Prediction Model, the values can be specified as follows:
parameters = {
"favourable_class" : [ "Loan Granted", "Loan Partially Granted" ],
"unfavourable_class": [ "Loan Denied" ]
}
In case of a regression models, the favourable and unfavourable classes will be ranges. For example, for a model which predicts medicine dosage, the favorable outcome could be between 80 ml to 120 ml or between 5 ml to 20 ml whereas unfavorable outcome will be values between 21 ml to 79ml. For such a model, the favorable and unfavorable values will be specified as follows:
parameters = {
"favourable_class" : [ [5, 20], [80, 120] ],
"unfavourable_class": [ [21, 79] ]
}
Please edit the next cell to provide information about your model.
```
# For classification models use the below.
parameters = {
"favourable_class" : [ "<EDIT THIS>", "<EDIT THIS>" ],
"unfavourable_class": [ "<EDIT THIS>" ]
}
# For regression models use the below. Delete the entry which is not required.
parameters = {
"favourable_class" : [ [<EDIT THIS>, <EDIT THIS>], [<EDIT THIS>,<EDIT THIS>] ],
"unfavourable_class": [ [<EDIT THIS>, <EDIT THIS>] ]
}
```
## Specify the number of records which should be processed for Fairness
The final piece of information that needs to be provided is the number of records (min_records) that should be used for computing the fairness. Fairness checks runs hourly. If min_records is set to 5000, then every hour fairness checking will pick up the last 5000 records which were sent to the model for scoring and compute the fairness on those 5000 records. Please note that fairness computation will not start till the time that 5000 records are sent to the model for scoring.
If we set the value of "min_records" to a small number, then fairness computation will get influenced by the scoring requests sent to the model in the recent past. In other words, the model might be flagged as being biased if it is acting in a biased manner on the last few records, but overall it might not be acting in a biased manner. On the other hand, if the "min_records" is set to a very large number, then we will not be able to catch model bias quickly. Hence the value of min_records should be set such that it is neither too small or too large.
Please updated the next cell to specify a value for min_records.
```
# min_records = <Minimum number of records to be considered for preforming scoring>
min_records = <EDIT THIS>
# max_records = <Maximum number of records to be considered while computing fairness> [OPTIONAL]
max_records = None
```
## End of Input
You need not edit anything beyond this point. Run the notebook and go to the very last cell. There will be a link to download the JSON file (called: "Download training data distribution JSON file"). Download the file and upload it using the IBM AI OpenScale GUI.
*Note: drop_na parameter of TrainingStats object should be set to 'False' if NA values are taken care while reading the training data in the above cells*
```
from ibm_watson_openscale.utils.training_stats import TrainingStats
enable_explainability = service_configuration_support.get("enable_explainability")
enable_fairness = service_configuration_support.get("enable_fairness")
if enable_explainability or enable_fairness:
fairness_inputs = None
if enable_fairness:
fairness_inputs = {
"fairness_attributes": fairness_attributes,
"min_records" : min_records,
"favourable_class" : parameters["favourable_class"],
"unfavourable_class": parameters["unfavourable_class"]
}
if max_records is not None:
fairness_inputs["max_records"] = max_records
input_parameters = {
"label_column": training_data_info["class_label"],
"feature_columns": training_data_info["feature_columns"],
"categorical_columns": training_data_info["categorical_columns"],
"fairness_inputs": fairness_inputs,
"problem_type" : model_type
}
training_stats = TrainingStats(data_df,input_parameters, explain=enable_explainability, fairness=enable_fairness, drop_na=True)
config_json = training_stats.get_training_statistics()
config_json["notebook_version"] = VERSION
#print(config_json)
```
### Indirect Bias
In case of Indirect bias i.e if protected attributes(the sensitive attributes like race, gender etc which are present in the training data but are not used to train the model) are being monitored for fairness:
- Bias service identifies correlations between the protected attribute and model features. Correlated attributes are also known as proxy features.
- Existence of correlations with model features can result in indirect bias w.r.t protected attribute even though it is not used to train the model.
- Highly correlated attributes based on their correlation strength are considered while computing bias for a given protected attribute.
The following cell identifies if user has configured protected attribute for fairness by checking the feature, non-feature columns and the fairness configuration. If protected attribute/s are configured then it identifies correlations and stores it in the fairness configuration.
```
# Checking if protected attributes are configured for fairness monitoring. If yes, then computing correlation information for each meta-field and updating it in the fairness configuration
if enable_fairness:
fairness_configuration = config_json.get("fairness_configuration")
training_columns = data_df.columns.tolist()
label_column = training_data_info.get("class_label")
training_columns.remove(label_column)
feature_columns = training_data_info.get("feature_columns")
non_feature_columns = list(set(training_columns) - set(feature_columns))
if non_feature_columns is not None and len(non_feature_columns) > 0:
protected_attributes = []
fairness_attributes_list = [attribute.get("feature") for attribute in fairness_attributes]
for col in non_feature_columns:
if col in fairness_attributes_list:
protected_attributes.append(col)
if len(protected_attributes) > 0:
from ibm_watson_openscale.utils.indirect_bias_processor import IndirectBiasProcessor
fairness_configuration = IndirectBiasProcessor().get_correlated_attributes(data_df, fairness_configuration, feature_columns, protected_attributes, label_column)
import json
print("Finished generating training distribution data")
# Create a file download link
import base64
from IPython.display import HTML
def create_download_link( title = "Download training data distribution JSON file", filename = "training_distribution.json"):
if enable_explainability or enable_fairness:
output_json = json.dumps(config_json, indent=2)
b64 = base64.b64encode(output_json.encode())
payload = b64.decode()
html = '<a download="{filename}" href="data:text/json;base64,{payload}" target="_blank">{title}</a>'
html = html.format(payload=payload,title=title,filename=filename)
return HTML(html)
else:
print("No download link generated as fairness/explainability services are disabled.")
create_download_link()
```
## Drift configuration archive generation
Please update the score function which will be used forgenerating drift detection model which will used for drift detection . This might take sometime to generate model and time taken depends on the training dataset size. The output of the score function should be a 2 arrays 1. Array of model prediction 2. Array of probabilities
- User is expected to make sure that the data type of the "class label" column selected and the prediction column are same . For eg : If class label is numeric , the prediction array should also be numeric
- Each entry of a probability array should have all the probabities of the unique class lable .
For eg: If the model_type=multiclass and unique class labels are A, B, C, D . Each entry in the probability array should be a array of size 4 . Eg : [ [0.50,0.30,0.10,0.10] ,[0.40,0.20,0.30,0.10]...]
**Note:**
- *User is expected to add "score" method , which should output prediction column array and probability column array.*
- *The data type of the label column and prediction column should be same . User needs to make sure that label column and prediction column array should have the same unique class labels*
- **Please update the score function below with the help of templates documented [here](https://github.com/IBM-Watson/aios-data-distribution/blob/master/Score%20function%20templates%20for%20drift%20detection.md)**
```
#Update score function
# def score(training_data_frame){
# <Fill in the template using the score function templates provided>
# }
#Generate drift detection model
from ibm_wos_utils.drift.drift_trainer import DriftTrainer
enable_drift = service_configuration_support.get("enable_drift")
if enable_drift:
drift_detection_input = {
"feature_columns":training_data_info.get("feature_columns"),
"categorical_columns":training_data_info.get("categorical_columns"),
"label_column": training_data_info.get("class_label"),
"problem_type": model_type
}
drift_trainer = DriftTrainer(data_df,drift_detection_input)
if model_type != "regression":
#Note: batch_size can be customized by user as per the training data size
drift_trainer.generate_drift_detection_model(score, batch_size=data_df.shape[0], check_for_ddm_quality=False)
#Note:
# - Two column constraints are not computed beyond two_column_learner_limit(default set to 200)
# - Categorical columns with large (determined by categorical_unique_threshold; default > 0.8) number of unique values relative to total rows in the column are discarded.
#User can adjust the value depending on the requirement
# user_overrides - Used to override drift constraint learning to selectively learn
# constraints on feature columns. Its a list of configuration, each specifying
# whether to learn distribution and/or range constraint on given set of columns.
# First configuration of a given column would take preference.
#
# "constraint_type" can have two possible values : single|double - signifying
# if this configuration is for single column or two column constraint learning.
#
# "learn_distribution_constraint" : True|False - signifying whether to learn
# distribution constraint for given config or not.
#
# "learn_range_constraint" : True|False - signifying whether to learn range
# constraint for given config or not. Only applicable to numerical feature columns.
#
# "features" : [] - provides either a list of feature columns to be governed by
# given configuration for constraint learning.
# Its a list of strings containing feature column names if "constraint_type" is "single".
# Its a list of list of strings containing feature column names if "constraint_type" if
# "double". If only one column name is provided, all of the two column constraints
# involving this column will be dictated by given configuration during constraint learning.
# This list is case-insensitive.
#
# In the example below, first config block says do not learn distribution and range single
# column constraints for features "MARITAL_STATUS", "PROFESSION", "IS_TENT" and "age".
# Second config block says do not learn distribution and range two column constraints
# where "IS_TENT", "PROFESSION", and "AGE" are one of the two columns. Whereas, specifically,
# do not learn two column distribution and range constraint on combination of "MARITAL_STATUS"
# and "PURCHASE_AMOUNT".
# "user_overrides"= [
# {
# "constraint_type": "single",
# "learn_distribution_constraint": False,
# "learn_range_constraint": False,
# "features": [
# "MARITAL_STATUS",
# "PROFESSION",
# "IS_TENT",
# "age"
# ]
# },
# {
# "constraint_type": "double",
# "learn_distribution_constraint": False,
# "learn_range_constraint": False,
# "features": [
# [
# "IS_TENT"
# ],
# [
# "MARITAL_STATUS"
# "PURCHASE_AMOUNT"
# ],
# [
# "PROFESSION"
# ],
# [
# "AGE"
# ]
# ]
# }
# ]
drift_trainer.learn_constraints(
two_column_learner_limit=200, categorical_unique_threshold=0.8, user_overrides=[])
drift_trainer.create_archive()
#Generate a download link for drift detection model
from IPython.display import HTML
import base64
import io
def create_download_link_for_ddm( title = "Download Drift detection model", filename = "drift_detection_model.tar.gz"):
#Retains stats information
if enable_drift:
with open(filename,"rb") as file:
ddm = file.read()
b64 = base64.b64encode(ddm)
payload = b64.decode()
html = '<a download="{filename}" href="data:text/json;base64,{payload}" target="_blank">{title}</a>'
html = html.format(payload=payload,title=title,filename=filename)
return HTML(html)
else:
print("Drift Detection is not enabled. Please enable and rerun the notebook")
create_download_link_for_ddm()
```
| github_jupyter |
<p><img alt="DataOwl" width=150 src="http://gwsolutions.cl/Images/dataowl.png", align="left", hspace=0, vspace=5></p>
<h1 align="center">Conceptos Básicos de Python</h1>
<h4 align="center">Comenzando con la programación</h4>
<pre><div align="center"> La idea de este notebook es que sirva como apoyo para iniciarse en la programación en Python.</div>
<div align="right"> En términos de código y estructura, este notebook esta basado en el BootCamp
<a href="https://github.com/Shekhar-rv/Python-for-Data-Science-and-Machine-Learning-Bootcamp">Python for Data Science and Machine Learning</a>.
.</div></pre>
<h2>Secciones</h2>
<div class="alert alert-danger" role="alert">
<ol>
<li><a href="#section1"> Tipos de Datos</a></li>
<li><a href="#section2"> Tipos de Operadores </a></li>
<li><a href="#section3"> Sentencias Condicionales </a></li>
<li><a href="#section4"> Ciclos</a></li>
<li><a href="#section5"> Funciones </a></li>
<li><a href="#section6"> Ejercicios</a></li>
</ol>
</div>
<hr>
<a id="section1"></a>
<h2>1. Tipos de Datos</h2>
<hr>
En Python tiene varios tipos de datos compuestos estándar disponibles por defecto en el interprete, como los tipos numéricos, booleanos, secuencias, conjuntos y diccionarios usados para agrupar otros valores.
Para el caso de las estructuras de datos se usan variables y constantes las cuales usan operadores para tratar los tipos de datos estándar.
```
#Datos númericos
x = 2 # Enteros
y = 2.0 # float (decimales)
z = 2 + 3j # Complejos
# Booleanos
x = True
y = False
# Secuencias
x = "Hola Mundo" # Strings
y = [1 , 2 , 3 , 4 , 5] # Listas
z = (1 , 2 , 3 , 4 , 5) # Tuplas
# Conjuntos
x = {1 , 2 , 3 , 4 , 5}
# Diccionarios
x = {"Ana" : "Profesora" , "Jorge" : "Director" , "Luis" : "Alumno/a" , "Marcela" : "Alumno/a" }
```
<a id="section2"></a>
<h2>2. Tipos de Operadores</h2>
<hr>
<h3>2.1 Operadores Aritméticos</h3>
Los valores numéricos pueden operarse mediante una serie de operadores aritméticos y matemáticos:
```
# Asignación de variables
x = 5
y = 2
# Suma
z = x + y
print(z)
# Resta
z = x - y
print(z)
# Multiplicación
z = x * y
print(z)
# División
z = x / y
print(z)
# División Entera
z = x // y
print(z)
# Resto Modular
z = x % y
print(z)
# Exponenciación
z = x**y
print(z)
```
<h3>2.2 Operadores Relacionales</h3>
Los operadores relacionales nos permiten ver si dos variables o constantes cumplen una relación de orden o de equivalencia. Los valores booleanos son además el resultado de expresiones que utilizan operadores relacionales:
```
# Asignación de variables
x = 5
y = 2
# Relación de igualdad
x == y
# Relación de desigualdad
x != y
# Relación de Mayor o igual
x >= y
```
<h3>2.3 Operadores de Asignación</h3>
Existe en Python todo un grupo de operadores los cuales le permiten básicamente asignar un valor a una variable, usando el operador “=”. Con estos operadores pueden aplicar la técnica denominada asignación aumentada.
```
# Asignación de variables
x = 5
# Asignación aumentada
x += 3
print(x)
```
<h3>2.4 Operadores de Pertenencia</h3>
Este tipo de operadores evalúan si un objeto pertenece a otro. Hay dos: in y not in, que es el complementario del anterior.
```
# Asignación de variables
x = "curso"
y = "Bienvenidos al curso"
n = 7
z = [ 1 , 2 , 3 , 4 , 5]
# Operador in
x in y
# Operador not in
n not in z
```
<h3>2.5 Operadores Lógicos</h3>
Python también incluye varios operadores lógicos: and, or y not. El primero devuelve True cuando ambos valores son True, el segundo devuelve True cuando algún valor es True, y el tercero devuelve True cuando el valor al que se aplica toma el valor False:
```
# Asignación de variables
x = (5 == 3)
y = (7 <= 12)
# Conjunción and
x and y
# Disyunción or
x or y
```
<a id="section3"></a>
<h2>3. Sentencias Condicionales</h2>
<hr>
La sentencia condicional if nos permite comprobar si se cumple una cierta condición con el fin de ejecutar un proceso u otro. En su forma más simple se utiliza la palabra reservada if seguida de la condición a probar y dos puntos.
Si queremos especificar qué código hay que ejecutar en el caso de que la condición no se cumpla podemos añadir la sentencia else, además podemos añadir tantas instrucciones elif como deseemos. Se evaluarán en orden y así que se cumpla una de las condiciones:
```
# Sentencia condicional
n = 0
if n > 0:
print("El número es positivo")
elif n < 0:
print("El número es negativo")
else:
print("El número es 0")
```
<a id="section4"></a>
<h2>4. Ciclos</h2>
<hr>
En Python se pueden utilizar dos tipos de ciclos, los for y while. Se utilizan los ciclos for cuando conoces la cantidad de repeticiones y los ciclos while cuando la cantidad de repeticiones depende de que se cumpla una condición.
<hr>
<h3>4.1 Ciclos for</h3>
El ciclo for nos permite ejecutar un proceso un número determinado de veces o para un conjunto determinado de valores:
```
L = [1 , 2 , 3 , 4 , 5]
for i in L:
print(i)
```
<h3>4.2 Ciclos while</h3>
Al contrario de lo que ocurre con los ciclos for, los bucles while no se repiten un número determinado de veces, sino mientras se cumpla una condición.
```
x = 1
while (x <= 5):
print(x)
x+=1
```
<a id="section5"></a>
<h2>5. Funciones</h2>
<hr>
Un elemento extremadamente útil es la posibilidad de definir y usar funciones. Si bien existen funciones definidas por Python y que nos ofrecen una funcionalidad concreta que no siempre va a adaptarse a nuestras necesidades, Python nos permite definir nuestra propias funciones, lo que tiene dos grandes ventajas:
* Nos evita tener que escribir el mismo código una y otra vez en nuestros programas. En lugar de esto movemos ese código a la función e invocamos ésta con los argumentos adecuados cada vez que queramos ejecutar esas instrucciones.
* También nos permiten sacar más provecho de nuestro trabajo al poder reutilizar estas funciones en nuestros programas.
<h3>5.1 Funciones predefinidas</h3>
Tenemos una serie de funciones predefinidas en Python que nos ayudan a expandir más lo que podemos hacer con este lenguaje:
* La entrada y salida de información se hacen con la función `print` y la función `input`:
* Tenemos algunas funciones matemáticas como: `abs`, `divmod`, `hex`, `max`, `min`,...
* Hay funciones que trabajan con caracteres y cadenas: `ascii`, `chr`, `format`, `repr`,...
* Además tenemos funciones que crean o convierten a determinados tipos de datos: `int`, `float`, `str`, `bool`, `range`, `dict`, `list`, ...
<h3>5.2 Creando nuestras propias funciones</h3>
En su forma más simple, la función es llamada sin ningún argumento y no devuelve ningún valor, simplemente ejecuta el código que contiene:
```
def saludar():
print("Hola")
saludar()
```
También podemos crear funciones que exijan uno o mas argumentos para ejecutarse:
```
def saludar(nombre):
print("Hola {}".format(nombre))
saludar("Jose")
```
<a id="section6"></a>
<h2>6. Ejercicios</h2>
<hr>
Ahora ejercitemos un poco con 6 ejercicios:
### 6.1 Ejercicio 1
Escriba un programa que encuentre todos los números divisibles por 7 pero que no son múltiplos de 5 entre el 2000 y el 3500 (Ambos incluidos). Debe mostrar los números separados por una coma.
<!-- The answer is below:
l=[]
for i in range(2000, 3201):
if (i%7==0) and (i%5!=0):
l.append(str(i))
print ','.join(l)
-->
```
L = [ ]
X = list(range(2000,3500))
for i in X:
if (i % 7 == 0) and (i % 5 != 0):
L.append(i)
else:
pass
print(L)
```
### 6.2 Ejercicio 2
Escriba un programa que encuentre todos los números entre 1000 y 3000 (Ambos incluidos) donde cada digito es un número par. Debe mostrar los números separados por una coma.
<!-- The answer is below:
values = []
for i in range(1000, 3001):
s = str(i)
if (int(s[0])%2==0) and (int(s[1])%2==0) and (int(s[2])%2==0) and (int(s[3])%2==0):
values.append(s)
print ",".join(values)
-->
```
L = []
X = list(range(1000,3000))
for i in X:
string = str(i)
if (int(string[0]) % 2 == 0) and (int(string[1]) % 2 == 0) and (int(string[2]) % 2 == 0) and (int(string[3]) % 2 == 0):
L.append(string)
else:
pass
print(L)
```
### 6.3 Ejercicio 3
Escribir un programa que calcule su balance en la cuenta de banco basado en su historial de transacciones entregado por un input. El historial de transacciones tiene el siguiente formato:
- D 100
- W 200
D significa depósito mientras W significa retiro.
Si suponemos que le ingresamos las siguientes transacciones al programa:
- D 300
- D 300
- W 200
- D 100
Entonces el programa debe mostrarnos
- Su balance es de 500
<!-- The answer is below:
netAmount = 0
while True:
s = raw_input()
if not s:
break
values = s.split(" ")
operation = values[0]
amount = int(values[1])
if operation=="D":
netAmount+=amount
elif operation=="W":
netAmount-=amount
else:
pass
print netAmount
-->
### 6.4 Ejercicio 4
Dado un número entero "n", escriba un programa que genere un diccionarion que contenga los pares (i, i**2), donde i es un número entero entre 1 y n (ambos incluidos), y que luego imprima el diccionario.
Supongamos que uno le entrega al programa el entero 8, entonces el programa debe mostrarnos:
{1: 1, 2: 4, 3: 9, 4: 16, 5: 25, 6: 36, 7: 49, 8: 64}
<!-- The answer is below:
n=int(raw_input())
d=dict()
for i in range(1,n+1):
d[i]=i*i
print d
-->
### 6.5 Ejercicio 5
Defina una función que reciba un número enter como argumento, y diga si el número es par o impar.
<!-- The answer is below:
def checkValue(n):
if n%2 == 0:
print "It is an even number"
else:
print "It is an odd number"
checkValue(7)
-->
### 6.6 Ejercicio 6
Dada una lista cualquiera de números enteros, escriba un programa que remueva todos los duplicados de la lista pero que conserve el orden original de los elementos:
<!-- The answer is below:
def removeDuplicate( li ):
newli=[]
seen = set()
for item in li:
if item not in seen:
seen.add( item )
newli.append(item)
return newli
li=[12,24,35,24,88,120,155,88,120,155]
print removeDuplicate(li)
-->
| github_jupyter |
Box Model from Sarmiento and Gruber "Ocean Biogeochemical Dynamics" page 11
```
import sys
import os
import copy
import numpy as np
import datetime
from matplotlib import pyplot as plt
BOXSIMU_PATH = '/home/aschi/Documents/MyPrivateRepo/boxsimu_project/'
if not BOXSIMU_PATH in sys.path:
sys.path.append(BOXSIMU_PATH)
from boxsimu.entities import Fluid, Variable
from boxsimu.box import Box
from boxsimu.transport import Flow, Flux
from boxsimu.condition import Condition
from boxsimu.system import BoxModelSystem
from boxsimu.process import Process, Reaction
from boxsimu.solver import Solver
from boxsimu import utils
from boxsimu import ur
def get_system():
#############################
# FLUIDS
#############################
seawater = Fluid('sea water', rho=1020*ur.kg/ur.meter**3)
#############################
# VARIABLES
#############################
po4 = Variable('po4')
#############################
# PROCESSES
#############################
#############################
# REACTIONS
#############################
#############################
# BOXES
#############################
inital_po4_masses = (6e-8*3e19*ur.kg + 6e-8*1e21*ur.kg)/2
upper_ocean = Box(
name='upper_ocean',
name_long='Upper Ocean Box',
fluid=seawater.q(3e19*ur.kg),
processes=[],
condition=Condition(),
variables=[po4.q(inital_po4_masses)],
reactions=[],
)
deep_ocean = Box(
name='deep_ocean',
name_long='Deep Ocean Box',
fluid=seawater.q(1e21*ur.kg),
condition=Condition(),
variables=[po4.q(inital_po4_masses)],
reactions=[],
)
#############################
# FLOWS
#############################
flow_downwelling = Flow(
name='Downwelling',
source_box=upper_ocean,
target_box=deep_ocean,
rate=6e17*ur.kg/ur.year,
tracer_transport=True,
)
flow_upwelling = Flow(
name='Upwelling',
source_box=deep_ocean,
target_box=upper_ocean,
rate=6e17*ur.kg/ur.year,
tracer_transport=True,
)
#############################
# FLUXES
#############################
biological_pump = Flux(
name='Biological Pump',
source_box=upper_ocean,
target_box=deep_ocean,
variable=po4,
rate=lambda t, c: 5.1e-8 * 6e17*ur.kg/ur.year
)
#############################
# SYSTEM
#############################
bmsystem = BoxModelSystem('Box Model from Sarmiento and Gruber "Ocean Biogeochemical Dynamics" page 11',
boxes=[upper_ocean, deep_ocean],
flows=[
flow_downwelling,
flow_upwelling,
],
fluxes=[
biological_pump,
],
global_condition=Condition(),
)
return bmsystem
system = get_system()
solver = Solver(system)
sol = solver.solve(250*ur.year, 0.5*ur.year)
sol.plot_variable_mass_of_all_boxes(system.variables.po4)
sol.plot_total_variable_masses()
```
| github_jupyter |
# Homework 3: Arrays, File I/O and Plotting
Please complete this homework assignment in code cells in the iPython notebook. Include comments in your code when necessary. Please rename the notebook as SIS ID_HW03.ipynb (your student ID number) and save the notebook once you have executed it as a PDF (note, that when saving as PDF you don't want to use the option with latex because it crashes, but rather the one to save it directly as a PDF).
**The homework should be submitted on bCourses under the Assignments tab (both the .ipynb and .pdf files). Please label it by your student ID number (SIS ID)**
## Problem 1: Sunspots
[Adapted from Newman, Exercise 3.1] At <a href="http://www-personal.umich.edu/~mejn/computational-physics/sunspots.txt">this link</a> (and also in the current directory on datahub) you will find a file called `sunspots.txt`, which contains the observed number of sunspots on the Sun for each month since January 1749. The file contains two columns of numbers, the first being the month and the second being the sunspot number.
a. Write a program that reads in the data and makes a graph of sunspots as a function of time. Adjust the $x$ axis so that the data fills the whole horizontal width of the graph.
b. Modify your code to display two subplots in a single figure: The plot from Part 1 with all the data, and a second subplot with the first 1000 data points on the graph.
c. Write a function `running_average(y, r)` that takes an array or list $y$ and calculates the running average of the data, defined by
$$ Y_k = \frac{1}{2r+1} \sum_{m=-r}^r y_{k+m},$$
where $y_k$ are the sunspot numbers in our case. Use this function and modify your second subplot (the one with the first 1000 data points) to plot both the original data and the running average on the same graph, again over the range covered by the first 1000 data points. Use $r=5$, but make sure your program allows the user to easily change $r$.
The next two parts may require you to google for how to do things. Make a strong effort to do these parts on your own without asking for help. If you do ask for help from a GSI or friend, first ask them to point you to the resource they used, and do your best to learn the necessary techniques from that resource yourself. Finding and learning from online documentation and forums is a very important skill. (Hint: Stack Exchange/Stack Overflow is often a great resource.)
d. Add legends to each of your subplots, but make them partially transparent, so that you can still see any data that they might overlap. *Note: In your program, you should only have to change $r$ for the running average in one place to adjust both the graph and the legend.*
e. Since the $x$ and $y$ axes in both subplots have the same units, add shared $x$ and $y$ labels to your plot that are centered on the horizontal and vertical dimensions of your figure, respectively. Also add a single title to your figure.
When your are finished, your plot should look something close to this:
```
# Don't rerun this snippet of code.
# If you accidentally do, uncomment the lines below and rerun
#from IPython.display import Image
#Image(filename="img/p1_output.png")
```
#### Hints
* The running average is not defined for the first and last few points that you're taking a running average over. (Why is that?) Notice, for instance, that the black curve in the plot above doesn't extend quite as far on either side as the red curve. For making your plot, it might be helpful if your `running_average` function returns an array of the $x$-values $x_k$ (or their corresponding indices $k$) along with an array of the $y$-values $Y_k$ that you compute for the running average.
* You can use the Latex code `$\pm$` for the $\pm$ symbol in the legend. You can also just write `+/-` if you prefer.
## Problem 2: Variety Plot
In this problem, you will reproduce the following as a single figure with four subplots, as best you can:
```
# Don't rerun this snippet of code.
# If you accidentally do, uncomment the lines below and rerun
#from IPython.display import Image
#Image(filename="img/p2_output.png")
```
Here are some hints and directions for each one:
**Upper-left:** This is an image of silicon taken with an electron microscope.
You can find the data file `stm.txt` [here](http://www-personal.umich.edu/~mejn/computational-physics/stm.txt) and in your datahub directory, among resources for the [Newman](http://www-personal.umich.edu/~mejn/computational-physics/) text.
You may assume that the upper-left of the array is indeed the upper-left of the image.
Both axes should run from 0 to 5.5.
This subplot uses the `gray` colormap.
**Upper-Right:** Matplotlib can plot any list of $(x,y)$ points you give it, including parametric or polar curves. The curve in this subplot is called a "deltoid", and is the result of the equations
$$ \begin{align*}
x &= 2\cos\theta + \cos2\theta \\
y &= 2\sin\theta - \sin2\theta
\end{align*} $$
over a range of $\theta$ from $0$ to $2\pi$.
To get the aspect ratio equal with nice spacing around the curve, try one of the following, depending on how you are making your subplots:
- if you're using `plt.subplot(...)` to get each subplot (the "state-machine" approach), add the `aspect='equal'` and `adjustable='datalim'` arguments to the deltoid subplot, so your command will look something like `plt.subplot(..., aspect='equal', adjustable='datalim')`.
- if you're using `... = plt.subplots(...)` (note the 's'!) or `ax = fig.add_subplot(...)` on a figure `fig` to get subplots with axes objects (the "object-oriented" approach), add the line `ax.set_aspect(aspect='equal', adjustable='datalim')`, where `ax` is the axes object you want to affect.
**Lower-Left:** This kind of plot is called a log-log plot, where both axes are on a logarithmic scale. Google or look in the matplotlib gallery to learn how to make this kind of plot.
The three curves are $y = x$, $y = x^2$, and $y = x^3$, where $x$ ranges over $10^{-1}$ to $10^1$. (Note: You can write powers of ten in python using the shorthand `1e-1` for $10^{-1}$, `1e1` for $10^1$, and so on.)
To make the pretty mathematical labels you see in the sample figure above, you can use
* `r'$y = x, x^2, x^3$'` for the title
* `r'$x$'` for the $x$-axis, and
* `r'$y$'` for the $y$-axis.
Just put these bits of code as you see them (with the **`r`** outside the quotes!) where you would normally put a string for the title or axes labels.
**Lower-Right:** Here you see a density plot with contours of the function
$$f(x,y) = \cos^2(\pi\,x\,y ) e^{-\frac{x^2 + 4 y}{8}},$$
over $x$ from -2 to 2 and $y$ from -3 to 0.2.
Use `meshgrid` to generate the $x$ and $y$ values. Be careful to make sure that the point $(-2,-3)$ is in the bottom left corner of the plot.
You'll need to use both `imshow` and `contour` to generate the density plot and then overlay it with contours. This plot uses the default contour spacing, so you don't need to worry about adjusting that. The colormap is `jet`, matplotlib's current default. (The default colormap will be changing to `viridis` in the next version.)
To get the ticks spaced out like you see here, use matplotlib's `xticks` or `set_xticks` functions for the $x$-axis (depending on how you're making your plots), and similar functions for the $y$-axis. You can pass each of these a single argument: a simple list or array of the numbers you want ticked on each axis.
**Spacing the subplots:** Once all is said and done and you run `plt.show()`, you may notice your plots are cramped and overlapping each other. Add the line `plt.tight_layout()` before `plt.show()`, and matplotlib will space things out in an attempt to avoid overlapping subplots.
| github_jupyter |
# DefinedAEpTandZ0 media example
```
%load_ext autoreload
%autoreload 2
import skrf as rf
import skrf.mathFunctions as mf
import numpy as np
from numpy import real, log, log10, sum, absolute, pi, sqrt
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator
from scipy.optimize import minimize
rf.stylely()
```
## Measurement of two CPWG lines with different lengths
The measurement where performed the 21th March 2017 on a Anritsu MS46524B 20GHz Vector Network Analyser. The setup is a linear frequency sweep from 1MHz to 10GHz with 10'000 points. Output power is 0dBm, IF bandwidth is 1kHz and neither averaging nor smoothing are used.
CPWGxxx is a L long, W wide, with a G wide gap to top ground, T thick copper coplanar waveguide on ground on a H height substrate with top and bottom ground plane. A closely spaced via wall is placed on both side of the line and the top and bottom ground planes are connected by many vias.
| Name | L (mm) | W (mm) | G (mm) | H (mm) | T (um) | Substrate |
| :--- | ---: | ---: | ---: | ---: | ---: | :--- |
| MSL100 | 100 | 1.70 | 0.50 | 1.55 | 50 | FR-4 |
| MSL200 | 200 | 1.70 | 0.50 | 1.55 | 50 | FR-4 |
The milling of the artwork is performed mechanically with a lateral wall of 45°.
The relative permittivity of the dielectric was assumed to be approximately 4.5 for design purpose.

```
# Load raw measurements
TL100 = rf.Network('CPWG100.s2p')
TL200 = rf.Network('CPWG200.s2p')
TL100_dc = TL100.extrapolate_to_dc(kind='linear')
TL200_dc = TL200.extrapolate_to_dc(kind='linear')
plt.figure()
plt.suptitle('Raw measurement')
TL100.plot_s_db()
TL200.plot_s_db()
plt.figure()
t0 = -2
t1 = 4
plt.suptitle('Time domain reflexion step response (DC extrapolation)')
ax = plt.subplot(1, 1, 1)
TL100_dc.s11.plot_z_time_step(pad=2000, window='hamming', z0=50, label='TL100', ax=ax, color='0.0')
TL200_dc.s11.plot_z_time_step(pad=2000, window='hamming', z0=50, label='TL200', ax=ax, color='0.2')
ax.set_xlim(t0, t1)
ax.xaxis.set_minor_locator(AutoMinorLocator(10))
ax.yaxis.set_minor_locator(AutoMinorLocator(5))
ax.patch.set_facecolor('1.0')
ax.grid(True, color='0.8', which='minor')
ax.grid(True, color='0.4', which='major')
plt.show()
```
Impedance from the line and from the connector section may be estimated on the step response.
The line section is not flat, there is some variation in the impedance which may be induced by manufacturing tolerances and dielectric inhomogeneity.
Note that the delay on the reflexion plot are twice the effective section delays because the wave travel back and forth on the line.
Connector discontinuity is about 50 ps long. TL100 line plateau (flat impedance part) is about 450 ps long.
```
Z_conn = 53.2 # ohm, connector impedance
Z_line = 51.4 # ohm, line plateau impedance
d_conn = 0.05e-9 # s, connector discontinuity delay
d_line = 0.45e-9 # s, line plateau delay, without connectors
```
## Dielectric effective relative permittivity extraction by multiline method
```
#Make the missing reflect measurement
#This is possible because we already have existing calibration
#and know what the open measurement would look like at the reference plane
#'refl_offset' needs to be set to -half_thru - connector_length.
reflect = TL100.copy()
reflect.s[:,0,0] = 1
reflect.s[:,1,1] = 1
reflect.s[:,1,0] = 0
reflect.s[:,0,1] = 0
# Perform NISTMultilineTRL algorithm. Reference plane is at the center of the thru.
cal = rf.NISTMultilineTRL([TL100, reflect, TL200], [1], [0, 100e-3], er_est=3.0, refl_offset=[-56e-3])
plt.figure()
plt.title('Corrected lines')
cal.apply_cal(TL100).plot_s_db()
cal.apply_cal(TL200).plot_s_db()
plt.show()
```
Calibration results shows a very low residual noise floor. The error model is well fitted.
```
from skrf.media import DefinedAEpTandZ0
freq = TL100.frequency
f = TL100.frequency.f
f_ghz = TL100.frequency.f/1e9
L = 0.1
A = 0.0
f_A = 1e9
ep_r0 = 2.0
tanD0 = 0.001
f_ep = 1e9
x0 = [ep_r0, tanD0]
ep_r_mea = cal.er_eff.real
A_mea = 20/log(10)*cal.gamma.real
def model(x, freq, ep_r_mea, A_mea, f_ep):
ep_r, tanD = x[0], x[1]
m = DefinedAEpTandZ0(frequency=freq, ep_r=ep_r, tanD=tanD, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
ep_r_mod = m.ep_r_f.real
A_mod = m.alpha * log(10)/20
return sum((ep_r_mod - ep_r_mea)**2) + 0.001*sum((20/log(10)*A_mod - A_mea)**2)
res = minimize(model, x0, args=(TL100.frequency, ep_r_mea, A_mea, f_ep),
bounds=[(2, 4), (0.001, 0.013)])
ep_r, tanD = res.x[0], res.x[1]
print('epr={:.3f}, tand={:.4f} at {:.1f} GHz.'.format(ep_r, tanD, f_ep * 1e-9))
m = DefinedAEpTandZ0(frequency=freq, ep_r=ep_r, tanD=tanD, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
plt.figure()
plt.suptitle('Effective relative permittivity and attenuation')
plt.subplot(2,1,1)
plt.ylabel('$\epsilon_{r,eff}$')
plt.plot(f_ghz, ep_r_mea, label='measured')
plt.plot(f_ghz, m.ep_r_f.real, label='model')
plt.legend()
plt.subplot(2,1,2)
plt.xlabel('Frequency [GHz]')
plt.ylabel('A (dB/m)')
plt.plot(f_ghz, A_mea, label='measured')
plt.plot(f_ghz, 20/log(10)*m.alpha, label='model')
plt.legend()
plt.show()
```
Relative permittivity $\epsilon_{e,eff}$ and attenuation $A$ shows a reasonable agreement.
A better agreement could be achieved by implementing the Kirschning and Jansen microstripline dispersion model or using a linear correction.
## Connectors effects estimation
```
# note: a half line is embedded in connector network
coefs = cal.coefs
r = mf.sqrt_phase_unwrap(coefs['forward reflection tracking'])
s1 = np.array([[coefs['forward directivity'],r],
[r, coefs['forward source match']]]).transpose()
conn = TL100.copy()
conn.name = 'Connector'
conn.s = s1
# delay estimation,
phi_conn = (np.angle(conn.s[:500,1,0]))
z = np.polyfit(f[:500], phi_conn, 1)
p = np.poly1d(z)
delay = -z[0]/(2*np.pi)
print('Connector + half thru delay: {:.0f} ps'.format(delay * 1e12))
print('TDR readed half thru delay: {:.0f} ps'.format(d_line/2 * 1e12))
d_conn_p = delay - d_line/2
print('Connector delay: {:.0f} ps'.format(d_conn_p * 1e12))
# connector model with guessed loss
half = m.line(d_line/2, 's', z0=Z_line)
mc = DefinedAEpTandZ0(m.frequency, ep_r=1, tanD=0.025, Z0=50,
f_low=1e3, f_high=1e18, f_ep=f_ep, model='djordjevicsvensson')
left = mc.line(d_conn_p, 's', z0=Z_conn)
right = left.flipped()
check = mc.thru() ** left ** half ** mc.thru()
plt.figure()
plt.suptitle('Connector + half thru comparison')
plt.subplot(2,1,1)
conn.plot_s_deg(1, 0, label='measured')
check.plot_s_deg(1, 0, label='model')
plt.ylabel('phase (rad)')
plt.legend()
plt.subplot(2,1,2)
conn.plot_s_db(1, 0, label='Measured')
check.plot_s_db(1, 0, label='Model')
plt.xlabel('Frequency (GHz)')
plt.ylabel('Insertion Loss (dB)')
plt.legend()
plt.show()
```
Connector + thru plots shows a reasonable agreement between calibration results and model.
## Final check
```
DUT = m.line(d_line, 's', Z_line)
DUT.name = 'model'
Check = m.thru() ** left ** DUT ** right ** m.thru()
Check.name = 'model with connectors'
plt.figure()
TL100.plot_s_db()
Check.plot_s_db(1,0, color='k')
Check.plot_s_db(0,0, color='k')
plt.show()
Check_dc = Check.extrapolate_to_dc(kind='linear')
plt.figure()
plt.suptitle('Time domain step-response')
ax = plt.subplot(1,1,1)
TL100_dc.s11.plot_z_time_step(pad=2000, window='hamming', label='Measured', ax=ax, color='k')
Check_dc.s11.plot_z_time_step(pad=2000, window='hamming', label='Model', ax=ax, color='b')
t0 = -2
t1 = 4
ax.set_xlim(t0, t1)
ax.xaxis.set_minor_locator(AutoMinorLocator(10))
ax.yaxis.set_minor_locator(AutoMinorLocator(5))
ax.patch.set_facecolor('1.0')
ax.grid(True, color='0.8', which='minor')
ax.grid(True, color='0.5', which='major')
```
The plots shows a reasonable agreement between model and measurement up to 4 GHz.
Further works may include implementing CPWG medium or modeling the line by more sections to account the impedance variation vs. position.
| github_jupyter |
Prerequisites: install [tensorflow 1.0](https://www.tensorflow.org/install/) and [scikit-image](http://scikit-image.org/docs/dev/api/skimage.html).
clone this fork of [tf-slim](https://github.com/marcotcr/tf-models) somewhere
download [the pretrained model](http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz) and put it in tf-models/slim/pretrained/
```
import tensorflow as tf
slim = tf.contrib.slim
import sys
sys.path.append('/Users/marcotcr/phd/tf-models/slim')
%load_ext autoreload
%autoreload 2
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
```
## Create a predict fn for inception v3, takes in a list of images and returns a matrix of prediction probabilities
```
from nets import inception
from preprocessing import inception_preprocessing
session = tf.Session()
image_size = inception.inception_v3.default_image_size
def transform_img_fn(path_list):
out = []
for f in path_list:
image_raw = tf.image.decode_jpeg(open(f).read(), channels=3)
image = inception_preprocessing.preprocess_image(image_raw, image_size, image_size, is_training=False)
out.append(image)
return session.run([out])[0]
from datasets import imagenet
names = imagenet.create_readable_names_for_imagenet_labels()
processed_images = tf.placeholder(tf.float32, shape=(None, 299, 299, 3))
import os
with slim.arg_scope(inception.inception_v3_arg_scope()):
logits, _ = inception.inception_v3(processed_images, num_classes=1001, is_training=False)
probabilities = tf.nn.softmax(logits)
checkpoints_dir = '/Users/marcotcr/phd/tf-models/slim/pretrained'
init_fn = slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v3.ckpt'),
slim.get_model_variables('InceptionV3'))
init_fn(session)
def predict_fn(images):
return session.run(probabilities, feed_dict={processed_images: images})
```
## Let's see the top 5 prediction for some image
```
images = transform_img_fn(['dogs.jpg'])
# I'm dividing by 2 and adding 0.5 because of how this Inception represents images
plt.imshow(images[0] / 2 + 0.5)
preds = predict_fn(images)
for x in preds.argsort()[0][-5:]:
print x, names[x], preds[0,x]
image = images[0]
## Now let's get an explanation
from lime import lime_image
import time
explainer = lime_image.LimeImageExplainer()
```
hide_color is the color for a superpixel turned OFF. Alternatively, if it is NONE, the superpixel will be replaced by the average of its pixels. Here, we set it to 0 (in the representation used by inception model, 0 means gray)
```
tmp = time.time()
# Hide color is the color for a superpixel turned OFF. Alternatively, if it is NONE, the superpixel will be replaced by the average of its pixels
explanation = explainer.explain_instance(image, predict_fn, top_labels=5, hide_color=0, num_samples=1000)
print time.time() - tmp
```
Image classifiers are a bit slow. Notice that an explanation in my macbookpro took 7.33 minutes
### Now let's see the explanation for the top class (Bernese mountain dog)
We can see the top 5 superpixels that are most positive towards the class with the rest of the image hidden
```
from skimage.segmentation import mark_boundaries
temp, mask = explanation.get_image_and_mask(240, positive_only=True, num_features=5, hide_rest=True)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
Or with the rest of the image present:
```
temp, mask = explanation.get_image_and_mask(240, positive_only=True, num_features=5, hide_rest=False)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
We can also see the 'pros and cons' (pros in green, cons in red)
```
temp, mask = explanation.get_image_and_mask(240, positive_only=False, num_features=10, hide_rest=False)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
Or the pros and cons that have weight at least 0.1
```
temp, mask = explanation.get_image_and_mask(240, positive_only=False, num_features=1000, hide_rest=False, min_weight=0.1)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
### Let's see the explanation for Egyptian cat
Most positive towards egyptian cat:
```
temp, mask = explanation.get_image_and_mask(286, positive_only=True, num_features=5, hide_rest=True)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
Pros and cons:
```
temp, mask = explanation.get_image_and_mask(286, positive_only=False, num_features=10, hide_rest=False)
plt.imshow(mark_boundaries(temp / 2 + 0.5, mask))
```
| github_jupyter |
# Packages
```
!pip install rouge --quiet
!pip install transformers --quiet
```
# Imports
```
import re
import json
import numpy as np
import pandas as pd
from rouge import Rouge
from bleu import list_bleu
from tqdm import tqdm_notebook
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
from torch.utils.data import Dataset, DataLoader
from torch.utils.data import Dataset, DataLoader, RandomSampler, SequentialSampler
from transformers import EncoderDecoderModel, AutoTokenizer, Seq2SeqTrainingArguments, Seq2SeqTrainer, EarlyStoppingCallback
from transformers import AdamW, get_linear_schedule_with_warmup, get_cosine_schedule_with_warmup, get_cosine_with_hard_restarts_schedule_with_warmup
from sklearn.model_selection import train_test_split
print("GPU Torch Available = {}".format(torch.cuda.is_available()))
print("Torch Version = {}".format(torch.__version__))
```
# Encoder-Decoder Model
```
# Model Selection
# Regular Models
bert_base_cased = 'bert-base-cased'
roberta_base = 'roberta-base'
gpt2 = 'gpt2'
electra = 'google/electra-small-discriminator'
t5_base = 't5-base'
bart = 'facebook/bart-base'
# Heavy Memory Dependant Models (For High RAM and High GPU Systems)
bert_large_cased = 'bert-large-cased'
roberta_large = 'roberta-large'
gpt2_medium = 'gpt2-medium'
t5_large = 't5-large'
bart_large = 'facebook/bart-large'
# Select Pretrained Weights
Pretrained_Weight = bert_base_cased # Select Pretrained Weights
# Encoder-Decoder
seq2seq = EncoderDecoderModel.from_encoder_decoder_pretrained(Pretrained_Weight, Pretrained_Weight)
# Tokenizer
tokenizer = AutoTokenizer.from_pretrained(Pretrained_Weight)
# Set Special Tokens
seq2seq.config.decoder_start_token_id = tokenizer.bos_token_id
seq2seq.config.eos_token_id = tokenizer.eos_token_id
seq2seq.config.pad_token_id = tokenizer.pad_token_id
# Parameters for Beam Search
seq2seq.config.vocab_size = seq2seq.config.decoder.vocab_size
seq2seq.config.max_length = 142
seq2seq.config.min_length = 56
seq2seq.config.no_repeat_ngram_size = 3
seq2seq.config.early_stopping = True
seq2seq.config.length_penalty = 2.0
seq2seq.config.num_beams = 6
```
# Dataset Loading and Preprocessing
```
from google.colab import drive
drive.mount('/content/drive')
# Loading Dataset
file = '/content/drive/MyDrive/Title Generation/Dataset/Dataset_Title_Summarization_of_Various_Summaries.xlsx'
df = pd.read_excel(file, names = ['ID','Combined Abstract', 'Title'])
df = df.drop(['ID'], axis=1)
df
# Train Test Split of Dataset
train_df, test_df = train_test_split(df, test_size = 0.25, random_state = 42)
print('Train Dataset Length = {}'.format(len(train_df)))
print('Test Dataset Length = {}'.format(len(test_df)))
# Data Preparation into Pandas Dataframe for Model Input
def get_data(dataframe):
abstract = list(dataframe['Combined Abstract'])
title = list(dataframe['Title'])
raw_data_train = {'Abstract': abstract, 'Title': title}
df = pd.DataFrame(raw_data_train, columns = ['Abstract','Title'])
return df
train_data = get_data(train_df)
test_data = get_data(test_df)
print('Training Data:')
print(train_data[0:3])
print('\nTesting Data:')
print(test_data[0:3])
# Data Preparation for Seq2Seq Model Input
class CustomDataset(Dataset):
def __init__(self, dataframe, tokenizer, max_len_enc, max_len_dec):
self.tokenizer = tokenizer
self.data = dataframe
self.abstract = dataframe.Abstract
self.title = dataframe.Title
self.encoder_max_len = max_len_enc
self.decoder_max_len = max_len_dec
def __len__(self):
return len(self.abstract)
def __getitem__(self, index):
# Abstract Tokenization
abstract_data = str(self.abstract[index])
inputs = self.tokenizer.encode_plus(abstract_data,
truncation=True,
add_special_tokens=True,
max_length = self.encoder_max_len,
padding = 'max_length',
return_token_type_ids = False)
input_ids = inputs['input_ids']
input_mask = inputs['attention_mask']
# Title Tokenization
title_data = str(self.title[index])
outputs = self.tokenizer.encode_plus(title_data,
truncation=True,
add_special_tokens=True,
max_length = self.decoder_max_len,
padding = 'max_length',
return_token_type_ids = False)
output_ids = outputs['input_ids']
output_mask = outputs['attention_mask']
return {'input_ids': torch.tensor(input_ids, dtype=torch.long),
'attention_mask': torch.tensor(input_mask, dtype=torch.long),
'decoder_input_ids': torch.tensor(output_ids, dtype=torch.long),
'decoder_attention_mask' : torch.tensor(output_mask, dtype=torch.long),
'labels': torch.tensor(output_ids, dtype=torch.long)}
ENCODER_MAX_LEN = 256 # Encoder Max Sequence Length (Change)
DECODER_MAX_LEN = 32 # Decoder Max Sequence Length
training_set = CustomDataset(train_data, tokenizer, ENCODER_MAX_LEN, DECODER_MAX_LEN) # Training Set
testing_set = CustomDataset(test_data, tokenizer, ENCODER_MAX_LEN, DECODER_MAX_LEN) # Testing Set
```
# Training
```
# Device Mapping Select (GPU or CPU)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
if (torch.cuda.is_available() == True):
print("Model Mapped CUDA::GPU")
seq2seq = seq2seq.cuda()
# Early Stopping Callback Setup
early_stop = EarlyStoppingCallback(early_stopping_patience = 3, early_stopping_threshold = 0.01)
# Learning Rate Schedulers
scheduler_options = ['linear', 'cosine', 'cosine_with_restarts', 'polynomial', 'constant', 'constant_with_warmup']
# Training Parameters (Should be Tuned)
training_args = Seq2SeqTrainingArguments(seed = 42,
output_dir="./models/model_name",
overwrite_output_dir = True,
evaluation_strategy = "epoch",
do_train = True,
do_eval = True,
learning_rate = 5e-5,
lr_scheduler_type = 'polynomial',
weight_decay = 0.01,
per_device_train_batch_size = 16,
per_device_eval_batch_size = 16,
predict_with_generate = False,
num_train_epochs = 20,
logging_steps = 2,
save_steps = 0,
warmup_steps = 16,
load_best_model_at_end = True)
# Instantiate Seq2Seq Trainer
trainer = Seq2SeqTrainer(model = seq2seq,
callbacks = [early_stop],
tokenizer = tokenizer,
args = training_args,
train_dataset = training_set,
eval_dataset = testing_set)
# Train Model
trainer.train()
```
# Testing
```
trainer.evaluate(testing_set)
```
# Model Save
```
# Model Save
model_save_path = '/content/drive/MyDrive/Title Generation/Model Weights/Seq2Seq_state_dict_Bert_Base_10000_Various_Summaries_Beam_6'
torch.save(seq2seq.state_dict(), model_save_path + '.pth')
```
# Model Load
```
# Model Load (Load Already Finetuned Model)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model_load_path = '/content/drive/MyDrive/Title Generation/Model Weights/Seq2Seq_state_dict_Bert_Base_10000_Various_Summaries_Beam_6.pth'
seq2seq.load_state_dict(torch.load(model_load_path, map_location = device))
```
# Rouge Score Calcultion
```
# Rouge Score Calculation
test_size = 2500
rouge = Rouge()
test_index_limit = test_size # Test Size for GPU Constraints
# Placeholders for Rouge Scores
rouge_1_f = []
rouge_1_p = []
rouge_1_r = []
rouge_2_f = []
rouge_2_p = []
rouge_2_r = []
rouge_l_f = []
rouge_l_p = []
rouge_l_r = []
# Calculation
for i in range(test_index_limit):
# Inference
data = test_data['Abstract'][i]
input_sentence_ids = torch.tensor(tokenizer.encode(data, add_special_tokens = True)).unsqueeze(0).cuda()
generated = seq2seq.generate(input_sentence_ids,
max_length = 20,
decoder_start_token_id = seq2seq.config.decoder.pad_token_id)
if (i % 50 == 0):
print("Inferene Done for Test ID = {}".format(i))
# Reference and Hypothesis for Rouge Score Calculation
hypothesis = tokenizer.batch_decode(generated, skip_special_tokens = True)[0] # Predicted Title
reference = test_data['Title'][i] # Reference Title
# Calculating Rouge Scores
score = rouge.get_scores(hypothesis, reference)
rouge_1_f.append(score[0]['rouge-1']['f'])
rouge_1_p.append(score[0]['rouge-1']['p'])
rouge_1_r.append(score[0]['rouge-1']['r'])
rouge_2_f.append(score[0]['rouge-2']['f'])
rouge_2_p.append(score[0]['rouge-2']['p'])
rouge_2_r.append(score[0]['rouge-2']['r'])
rouge_l_f.append(score[0]['rouge-l']['f'])
rouge_l_p.append(score[0]['rouge-l']['p'])
rouge_l_r.append(score[0]['rouge-l']['r'])
# Final Average Rouge Score Calculation
rouge_1_f_val = sum(rouge_1_f)/test_size
rouge_1_p_val = sum(rouge_1_p)/test_size
rouge_1_r_val = sum(rouge_1_r)/test_size
rouge_2_f_val = sum(rouge_2_f)/test_size
rouge_2_p_val = sum(rouge_2_p)/test_size
rouge_2_r_val = sum(rouge_2_r)/test_size
rouge_l_f_val = sum(rouge_l_f)/test_size
rouge_l_p_val = sum(rouge_l_p)/test_size
rouge_l_r_val = sum(rouge_l_r)/test_size
print('\n Scores:')
print('Avergae Rouge 1 F Score : {}'.format(rouge_1_f_val))
print('Avergae Rouge 1 Precision : {}'.format(rouge_1_p_val))
print('Avergae Rouge 1 Recall : {}'.format(rouge_1_r_val))
print('\n')
print('Avergae Rouge 2 F Score : {}'.format(rouge_2_f_val))
print('Avergae Rouge 2 Precision : {}'.format(rouge_2_p_val))
print('Avergae Rouge 2 Recall : {}'.format(rouge_2_r_val))
print('\n')
print('Avergae Rouge L F Score : {}'.format(rouge_l_f_val))
print('Avergae Rouge L Precision : {}'.format(rouge_l_p_val))
print('Avergae Rouge L Recall : {}'.format(rouge_l_r_val))
```
# Inference (Generation)
```
def inference(model, tokenizer, test_list, test_dataset, maximum_length, show_abstracts = True):
for test_index in test_list:
# Fetch Input and Reference from Test Dataset
data = test_dataset['Abstract'][test_index]
reference = test_dataset['Title'][test_index]
# Inference
input_sentence_ids = torch.tensor(tokenizer.encode(data, add_special_tokens=True)).unsqueeze(0).cuda()
generated = model.generate(input_sentence_ids,
max_length = maximum_length,
decoder_start_token_id = model.config.decoder.pad_token_id)
hypothesis = tokenizer.batch_decode(generated, skip_special_tokens=True)[0]
print('\nTest ID = {}'.format(test_index))
if (show_abstracts == True):
print('\nAbstract:')
print(data)
print('\nActual Title:')
print(reference)
print('\nPredicted Title:')
print(hypothesis)
test_list = [1,4,5,7,9,10,12,20,22,100,134,200] # Test Indices for Inference
inference(seq2seq, tokenizer, test_list, test_data, 24, show_abstracts = False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/TausifAnsari/Traffic-Sign-Recogizer/blob/master/Traffic_Sign_recognize.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# Run this cell and select the kaggle.json file downloaded
# from the Kaggle account settings page.
from google.colab import files
files.upload()
# Let's make sure the kaggle.json file is present.
!ls -lha kaggle.json
# Install the Kaggle API client.
!pip install -q kaggle
# The Kaggle API client expects this file to be in ~/.kaggle,
# so move it there.
!mkdir -p ~/.kaggle
!cp kaggle.json ~/.kaggle/
# This permissions change avoids a warning on Kaggle tool startup.
!chmod 600 ~/.kaggle/kaggle.json
!kaggle datasets download -d meowmeowmeowmeowmeow/gtsrb-german-traffic-sign
!mkdir dataset
!unzip /content/gtsrb-german-traffic-sign.zip -d /content/dataset
# import the necessary packages
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Dense
class TrafficSignNet:
@staticmethod
def build(width, height, depth, classes):
# initialize the model along with the input shape to be
# "channels last" and the channels dimension itself
model = Sequential()
inputShape = (height, width, depth)
chanDim = -1
# CONV => RELU => BN => POOL
model.add(Conv2D(8, (5, 5), padding="same",
input_shape=inputShape))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(MaxPooling2D(pool_size=(2, 2)))
# first set of (CONV => RELU => CONV => RELU) * 2 => POOL
model.add(Conv2D(16, (3, 3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(Conv2D(16, (3, 3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(MaxPooling2D(pool_size=(2, 2)))
# second set of (CONV => RELU => CONV => RELU) * 2 => POOL
model.add(Conv2D(32, (3, 3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(Conv2D(32, (3, 3), padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), padding="same", activation = "relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(Conv2D(64, (3, 3), padding="same", activation = "relu"))
model.add(BatchNormalization(axis=chanDim))
model.add(MaxPooling2D(pool_size=(2, 2)))
# first set of FC => RELU layers
model.add(Flatten())
model.add(Dense(128))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.5))
# second set of FC => RELU layers
model.add(Flatten())
model.add(Dense(128))
model.add(Activation("relu"))
model.add(BatchNormalization())
model.add(Dropout(0.5))
# softmax classifier
model.add(Dense(classes))
model.add(Activation("softmax"))
# return the constructed network architecture
return model
# import the necessary packages
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.utils import to_categorical
from sklearn.metrics import classification_report
from skimage import transform
from skimage import exposure
from skimage import io
import matplotlib.pyplot as plt
import numpy as np
import argparse
import random
import os
def load_split(basePath, csvPath):
# initialize the list of data and labels
data = []
labels = []
# load the contents of the CSV file, remove the first line (since
# it contains the CSV header), and shuffle the rows (otherwise
# all examples of a particular class will be in sequential order)
rows = open(csvPath).read().strip().split("\n")[1:]
random.shuffle(rows)
# loop over the rows of the CSV file
for (i, row) in enumerate(rows):
# check to see if we should show a status update
if i > 0 and i % 1000 == 0:
print("[INFO] processed {} total images".format(i))
# split the row into components and then grab the class ID
# and image path
(label, imagePath) = row.strip().split(",")[-2:]
# derive the full path to the image file and load it
imagePath = os.path.sep.join([basePath, imagePath])
image = io.imread(imagePath)
# resize the image to be 32x32 pixels, ignoring aspect ratio,
# and then perform Contrast Limited Adaptive Histogram
# Equalization (CLAHE)
image = transform.resize(image, (32, 32))
image = exposure.equalize_adapthist(image, clip_limit=0.1)
# update the list of data and labels, respectively
data.append(image)
labels.append(int(label))
# convert the data and labels to NumPy arrays
data = np.array(data)
labels = np.array(labels)
# return a tuple of the data and labels
return (data, labels)
# initialize the number of epochs to train for, base learning rate,
# and batch size
NUM_EPOCHS = 30
INIT_LR = 1e-3
BS = 64
# load the label names
labelNames = open("signnames.csv").read().strip().split("\n")[1:]
labelNames = [l.split(",")[1] for l in labelNames]
# derive the path to the training and testing CSV files
trainPath = os.path.sep.join(["/content/dataset", "Train.csv"])
testPath = os.path.sep.join(["/content/dataset", "Test.csv"])
# load the training and testing data
print("[INFO] loading training and testing data...")
(trainX, trainY) = load_split("/content/dataset", trainPath)
(testX, testY) = load_split("/content/dataset", testPath)
# scale data to the range of [0, 1]
trainX = trainX.astype("float32") / 255.0
testX = testX.astype("float32") / 255.0
# one-hot encode the training and testing labels
numLabels = len(np.unique(trainY))
trainY = to_categorical(trainY, numLabels)
testY = to_categorical(testY, numLabels)
# account for skew in the labeled data
classTotals = trainY.sum(axis=0)
classWeight = classTotals.max() / classTotals
# construct the image generator for data augmentation
aug = ImageDataGenerator(
rotation_range=10,
zoom_range=0.15,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.15,
horizontal_flip=False,
vertical_flip=False,
fill_mode="nearest")
# initialize the optimizer and model
print("[INFO] compiling model...")
opt = Adam(lr=INIT_LR, decay=INIT_LR / (NUM_EPOCHS * 0.5))
model = TrafficSignNet.build(width=32, height=32, depth=3,
classes=numLabels)
model.compile(loss="categorical_crossentropy", optimizer=opt,
metrics=["accuracy"])
# compile the model and train the network
print("[INFO] training network...")
H = model.fit_generator(
aug.flow(trainX, trainY, batch_size=BS),
validation_data=(testX, testY),
steps_per_epoch=trainX.shape[0] // BS,
epochs=NUM_EPOCHS,
class_weight=classWeight,
verbose=1)
# evaluate the network
print("[INFO] evaluating network...")
predictions = model.predict(testX, batch_size=BS)
print(classification_report(testY.argmax(axis=1),
predictions.argmax(axis=1), target_names=labelNames))
# save the network to disk
print("[INFO] serializing network to ...")
model.save("traffic.h5")
# plot the training loss and accuracy
N = np.arange(0, NUM_EPOCHS)
plt.style.use("ggplot")
plt.figure()
plt.plot(N, H.history["loss"], label="train_loss")
plt.plot(N, H.history["val_loss"], label="val_loss")
plt.plot(N, H.history["acc"], label="train_acc")
plt.plot(N, H.history["val_acc"], label="val_acc")
plt.title("Training Loss and Accuracy on Dataset")
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
plt.show()
# USAGE
# python predict.py --model output/trafficsignnet.model --images gtsrb-german-traffic-sign/Test --examples examples
# import the necessary packages
from tensorflow.keras.models import load_model
import matplotlib.pyplot as plt
from skimage import transform
from skimage import exposure
from skimage import io
from imutils import paths
import numpy as np
import argparse
import imutils
import random
import cv2
import os
# load the traffic sign recognizer model
print("[INFO] loading model...")
model = load_model("traffic.h5")
# load the label names
labelNames = open("signnames.csv").read().strip().split("\n")[1:]
labelNames = [l.split(",")[1] for l in labelNames]
# grab the paths to the input images, shuffle them, and grab a sample
print("[INFO] predicting...")
imagePaths = list(paths.list_images("/content/dataset/Train"))
random.shuffle(imagePaths)
imagePaths = imagePaths[:25]
fig = plt.figure(figsize = (15,15))
#fig.suptitle("Prediction",fontsize=16)
# loop over the image paths
for (i, imagePath) in enumerate(imagePaths):
# load the image, resize it to 32x32 pixels, and then apply
# Contrast Limited Adaptive Histogram Equalization (CLAHE),
# just like we did during training
image = io.imread(imagePath)
image = transform.resize(image, (32, 32))
image = exposure.equalize_adapthist(image, clip_limit=0.1)
# preprocess the image by scaling it to the range [0, 1]
image = image.astype("float32") / 255.0
image = np.expand_dims(image, axis=0)
# make predictions using the traffic sign recognizer CNN
preds = model.predict(image)
j = preds.argmax(axis=1)[0]
label = labelNames[j]
# load the image using OpenCV, resize it, and draw the label
# on it
image = cv2.imread(imagePath)
image = imutils.resize(image, width=128)
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
#plt.title("Sign :"+label)
plt.xlabel(label)
plt.imshow(image)
plt.show()
```
| github_jupyter |
# ImageJ in Python
This notebook shows how to use ImageJ as a Python library and uses the OMERO Python API to connect to an OMERO server.
In this setup, Fiji has already been installed.
We will use the [pyimagej](https://pypi.org/project/pyimagej/) library to access to the entire ImageJ API from Python.
## Packages
```
import numpy
import imagej
from omero.gateway import BlitzGateway
```
## Starting ImageJ from Python
To run the macro, we use ImageJ1 windows so it requires using ImageJ in GUI mode and requires handling the resulting windows. This is the reason the parameter `headless` is set to `False`.
You will also need to start the **[desktop](../../desktop)** if it is not already up.
The link should open in a different window. If you see an error message try refreshing the window.
If you do not run the macro, you do **not** need the desktop and do **not** need to set the `headless` (default is `True`).
```
ij = imagej.init('/srv/conda/vnc/Fiji.app', headless=False)
ij.getVersion()
```
## Connect to IDR
```
def connect(hostname, username, password):
"""
Connect to an OMERO server
:param hostname: Host name
:param username: User
:param password: Password
:return: Connected BlitzGateway
"""
conn = BlitzGateway(username, password,
host=hostname, secure=True)
conn.connect()
conn.c.enableKeepAlive(60)
return conn
conn = connect("ws://idr.openmicroscopy.org/omero-ws", "public", "public")
print("Connected as {}".format(conn.getUser().getName()))
```
## Load an image from IDR
```
image_id = 1884835
image = conn.getObject("Image", image_id)
```
## Load binary data from IDR
We load a plane as a 2D numpy. Using the Python API, allows us to load easily only the plane we need.
```
def load_numpy_array(image):
pixels = image.getPrimaryPixels()
return pixels.getPlane(0, 0, 0)
img = load_numpy_array(image)
print(img.shape)
```
## Display image using `ij.py.show()`
```
ij.py.show(img[:, :], cmap = 'gray')
```
## Process numpy arrays in ImageJ
We use the method `to_java()` to convert into ImageJ types.
```
result = numpy.zeros(img.shape)
sigma1 = 8
sigma2 = 2
# note the use of to_java on img and result to turn the numpy images into RAIs
ij.op().filter().dog(
ij.py.to_java(result),
ij.py.to_java(img),
sigma1,
sigma2)
# purple highlights the edges of the vessels, green highlights the centers
ij.py.show(result, cmap = 'PRGn')
```
## Run macro
Run a macro on the image. We use ImageJ1 windows so it requires using ImageJ in GUI mode and requires handling the resulting windows.
```
from jnius import autoclass
WindowManager = autoclass('ij.WindowManager')
ij.ui().show('Image', ij.py.to_java(img))
macro = """
run("8-bit");
"""
ij.py.run_macro(macro)
img = WindowManager.getCurrentImage()
img.changes = False
ij.py.show(result)
```
## Close the connection to IDR
```
conn.close()
```
### License (BSD 2-Clause)
Copyright (c) 2021, University of Dundee All rights reserved.
Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer. Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution. THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT OWNER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
| github_jupyter |
```
import pandas as pd
import time
import os.path
import warnings
warnings.filterwarnings('ignore')
# install DenMune clustering algorithm using pip command from the offecial Python repository, PyPi
# from https://pypi.org/project/denmune/
!pip install denmune
# then import it
from denmune import DenMune
# clone datasets from our repository datasets
if not os.path.exists('datasets'):
!git clone https://github.com/egy1st/datasets
```
You can get your validation results using 3 methods
- by showing the Analyzer
- extract values from the validity returned list from fit_predict function
- extract values from the Analyzer dictionary
The algorithm is associated with five built-in validity measures, which are:
- ACC, Accuracy
- F1 score
- NMI index (Normalized Mutual Information)
- AMI index (Adjusted Mutual Information)
- ARI index (Adjusted Rand Index)
```
# Let us show the analyzer by set show_analyzer to True, which is actually the default parameter's value
data_path = 'datasets/denmune/shapes/'
dataset = "aggregation"
knn = 6
data_file = data_path + dataset + '.csv'
X_train = pd.read_csv(data_file, sep=',', header=None)
y_train = X_train.iloc[:, -1]
X_train = X_train.drop(X_train.columns[-1], axis=1)
print ("Dataset:", dataset)
dm = DenMune(train_data=X_train,
train_truth=y_train,
k_nearest=knn,
rgn_tsne=False)
labels, validity = dm.fit_predict(show_noise=True, show_analyzer=True)
# secondly, we can extract validity returned list from fit_predict function
dm = DenMune(train_data=X_train, train_truth=y_train, k_nearest=knn, rgn_tsne=False)
labels, validity = dm.fit_predict(show_plots=False, show_noise=True, show_analyzer=False)
validity
Accuracy = validity['train']['ACC']
print ('Accuracy:',Accuracy, 'correctely identified points')
F1_score = validity['train']['F1']
print ('F1 score:', round(F1_score*100,2), '%')
NMI = validity['train']['NMI']
print ('NMI index:', round(NMI*100,2), '%')
AMI = validity['train']['AMI']
print ('AMI index:', round(AMI*100,2), '%')
ARI = validity['train']['ARI']
print ('ARI index:', round(ARI*100,2), '%')
# Third, we can extract extract values from the Analyzer dictionary
dm = DenMune(train_data=X_train, train_truth=y_train, k_nearest=knn, rgn_tsne=False)
labels, validity = dm.fit_predict(show_plots=False, show_noise=True, show_analyzer=False)
dm.analyzer
Accuracy = dm.analyzer['validity']['train']['ACC']
print ('Accuracy:',Accuracy, 'correctely identified points')
F1_score = dm.analyzer['validity']['train']['F1']
print ('F1 score:', round(F1_score*100,2), '%')
NMI = dm.analyzer['validity']['train']['NMI']
print ('NMI index:', round(NMI*100,2), '%')
AMI = dm.analyzer['validity']['train']['AMI']
print ('AMI index:', round(AMI*100,2), '%')
ARI = dm.analyzer['validity']['train']['ARI']
print ('ARI index:', round(ARI*100,2), '%')
```
| github_jupyter |
# pipda
A framework for data piping in python
Inspired by [siuba][1], [dfply][2], [plydata][3] and [dplython][4], but with simple yet powerful APIs to mimic the `dplyr` and `tidyr` packages in python
## Installation
```shell
pip install -U pipda
```
## Usage
Checkout [plyrda][6] for more detailed usages.
### Verbs
Verbs are functions next to the piping sign (i.e. `>>`) receiving the data directly.
```
try:
import pandas
except ImportError:
!pip install pandas
import pandas as pd
from pipda import (
register_verb,
register_func,
register_operator,
evaluate_expr,
Operator,
Symbolic,
Context
)
f = Symbolic()
df = pd.DataFrame({
'x': [0, 1, 2, 3],
'y': ['zero', 'one', 'two', 'three']
})
df
@register_verb(pd.DataFrame)
def head(data, n=5):
return data.head(n)
df >> head(2)
@register_verb(pd.DataFrame, context=Context.EVAL)
def mutate(data, **kwargs):
data = data.copy()
for key, val in kwargs.items():
data[key] = val
return data
df >> mutate(z=1)
df >> mutate(z=f.x)
# Verbs that don't compile f.a to data, but just the column name
@register_verb(pd.DataFrame, context=Context.SELECT)
def select(data, *columns):
return data.loc[:, columns]
# f.x won't be compiled as df.x but just 'x'
df >> mutate(z=2*f.x) >> select(f.x, f.z)
# Compile the args inside the verb
@register_verb(pd.DataFrame, context=Context.PENDING)
def mutate_existing(data, column, value):
column = evaluate_expr(column, data, Context.SELECT)
value = evaluate_expr(value, data, Context.EVAL)
data = data.copy()
data[column] = value
return data
# First f.x compiled as column name, and second as Series data
df2 = df >> mutate_existing(f.x, 10 * f.x)
df2
# Evaluate the arguments by yourself
@register_verb(pd.DataFrame, context=Context.PENDING)
def mutate_existing2(data, column, value):
column = evaluate_expr(column, data, Context.SELECT)
value = evaluate_expr(value, df2, Context.EVAL)
data = data.copy()
data[column] = value
return data
df >> mutate_existing2(f.x, 2 * f.x)
# register for multiple types
@register_verb(int, context=Context.EVAL)
def add(data, other):
return data + other
# add is actually a singledispatch generic function
@add.register(float, context=Context.EVAL)
def _(data, other):
return data * other
1 >> add(1)
1.1 >> add(1.0)
# As it's a singledispatch generic function, we can do it for multiple types
# with the same logic
@register_verb(context=Context.EVAL)
def mul(data, other):
raise NotImplementedError # not invalid until types registered
@mul.register(int, context=Context.EVAL)
@mul.register(float, context=Context.EVAL)
# or you could do @mul.register((int, float), context=Context.EVAL)
# context is also supported
def _(data, other):
return data * other
3 >> mul(2)
3.2 >> mul(2)
```
### Functions used in verb arguments
```
@register_func(context=Context.EVAL)
def if_else(data, cond, true, false):
cond.loc[cond.isin([True]), ] = true
cond.loc[cond.isin([False]), ] = false
return cond
# The function is then also a singledispatch generic function
df >> mutate(z=if_else(f.x>1, 20, 10))
# function without data argument
@register_func(None)
def length(strings):
return [len(s) for s in strings]
df >> mutate(z=length(f.y))
# register existing functions
from numpy import vectorize
len = register_func(None, context=Context.EVAL, func=vectorize(len))
# original function still works
print(len('abc'))
df >> mutate(z=len(f.y))
```
### Operators
You may also redefine the behavior of the operators
```
@register_operator
class MyOperators(Operator):
def xor(self, a, b):
"""Inteprete X ^ Y as pow(X, Y)."""
return a ** b
df >> mutate(z=f.x ^ 2)
```
### Context
The context defines how a reference (`f.A`, `f['A']`, `f.A.B` is evaluated)
```
from pipda import ContextBase
class MyContext(ContextBase):
name = 'my'
def getattr(self, parent, ref, is_direct):
# double it to distinguish getattr
return getattr(parent, ref)
def getitem(self, parent, ref, is_direct):
return parent[ref] * 2
@property
def ref(self):
# how we evaluate the ref in f[ref]
return self
@register_verb(context=MyContext())
def mutate_mycontext(data, **kwargs):
for key, val in kwargs.items():
data[key] = val
return data
df >> mutate_mycontext(z=f.x + f['x'])
# when ref in f[ref] is also needed to be evaluated
df = df >> mutate(zero=0, one=1, two=2, three=3)
df
df >> mutate_mycontext(m=f[f.y][:1].values[0])
# f.y returns ['zero', 'one', 'two', 'three']
# f[f.y] gets [[0, 2, 4, 6], [0, 2, 4, 6], [0, 2, 4, 6], [0, 2, 4, 6]]
# f[f.y][:1].values gets [[0, 4, 8, 16]]
# f[f.y][:1].values[0] returns [0, 8, 16, 32]
# Notes that each subscription ([]) will double the values
```
### Caveats
- You have to use `and_` and `or_` for bitwise and/or (`&`/`|`) operators, as `and` and `or` are python keywords.
- Limitations:
Any limitations apply to `executing` to detect the AST node will apply to `pipda`. It may not work in some circumstances where other AST magics apply.
- Calling registered verbs/functions regularly:
The piping syntax (`>>`) is recommended with `pipda`. Because everything is determined with this syntax.
However, `pipda` tries to support regular calling. The ambiguity can come from the situations where the arguments passed in can shift one position right (such that they fit the piping calling), and first value passed in can also be dispatched and fit in the second argument.
For example:
```python
@register_verb(int)
def add(a: int, b: int = 1):
return a + b
```
If you call it like this `add(2)`, then we have no idea if this is calling `add(2, b=1)`, or `add(b=2)` and it's waiting for the data (`a`) to be piped in. In such a case, the function is called in the former way, but a warning will be showing.
To avoid this, as it states in the warning message, according to the reasons of the ambiguity, we should make sure that the values passed in cannot be shifted one position right (given values for all arguments would do it):
```python
add(2, 1) # or add(2, b=1)
```
or try not to use optional arguments while defining the function;
or make sure the first value cannot be dispatched:
```python
@register_verb(int)
def add(a: int, b: float = 1.0):
return a + b
add(2.0)
```
In such a case, it is for sure that it is called like `add(b=2.0)` and wait for `a` to be piped in.
You can even have a different type annotation for the second argument, even the same value can be accepted:
```python
@register_verb(int)
def add(a: int, b: Optional[int] = 1):
return a + b
add(2)
```
This will force it to call `add(2, b=1)`, but this definitely has some side effects:
```python
verb(data, add(2))
```
Here `add(2)` is intended to be called like `add(b=2)`, but unexpectedly, it will call like `add(2, b=1)`. Using the piping syntax will perfectly solve this:
```python
data >> verb(add(2))
```
since we know the function called in a verb is supposed to wait for the data to be piped in.
See also: [Piping vs regular calling][7]
- Use another piping sign
```python
from pipda import register_piping
register_piping('^')
# register verbs and functions
df ^ verb1(...) ^ verb2(...)
```
Allowed signs are: `+`, `-`, `*`, `@`, `/`, `//`, `%`, `**`, `<<`, `>>`, `&`, `^` and `|`.
## How it works
### The verbs
```R
data %>% verb(arg1, ..., key1=kwarg1, ...)
```
The above is a typical `dplyr`/`tidyr` data piping syntax.
The counterpart R syntax we expect is:
```python
data >> verb(arg1, ..., key1=kwarg1, ...)
```
To implement that, we need to defer the execution of the `verb` by turning it into a `Verb` object, which holds all information of the function to be executed later. The `Verb` object won't be executed until the `data` is piped in. It all thanks to the [`executing`][5] package to let us determine the ast nodes where the function is called. So that we are able to determine whether the function is called in a piping mode.
If an argument is referring to a column of the data and the column will be involved in the later computation, the it also needs to be deferred. For example, with `dplyr` in `R`:
```R
data %>% mutate(z=a)
```
is trying add a column named `z` with the data from column `a`.
In python, we want to do the same with:
```python
data >> mutate(z=f.a)
```
where `f.a` is a `Reference` object that carries the column information without fetching the data while python sees it immmediately.
Here the trick is `f`. Like other packages, we introduced the `Symbolic` object, which will connect the parts in the argument and make the whole argument an `Expression` object. This object is holding the execution information, which we could use later when the piping is detected.
### The functions
Then what if we want to use some functions in the arguments of the `verb`?
For example:
```python
data >> select(starts_with('a'))
```
to select the columns with names start with `'a'`.
No doubt that we need to defer the execution of the function, too. The trick is that we let the function return a `function` object as well, and evaluate it as the argument of the verb.
### The operators
`pipda` also opens oppotunities to change the behavior of the operators in verb/function arguments. This allows us to mimic something like this:
```python
data >> select(-f.a) # select all columns but `a`
```
To do that, we turn it into an `Operator` object. Just like a `Verb` or a `Function` object, the execution is deferred. By default, the operators we used are from the python standard library `operator`. `operator.neg` in the above example.
You can also define you own by subclassing the `Operator` class, and then register it to replace the default one by decorating it with `register_operator`.
[1]: https://github.com/machow/siuba
[2]: https://github.com/kieferk/dfply
[3]: https://github.com/has2k1/plydata
[4]: https://github.com/dodger487/dplython
[5]: https://github.com/alexmojaki/executing
[6]: https://github.com/pwwang/plyrda
[7]: https://pwwang.github.io/datar/piping_vs_regular/
| github_jupyter |
# Load medical images
This notebook introduces how to easily load different formats of medical images in MONAI and execute many additional operations.
[](https://colab.research.google.com/github/Project-MONAI/tutorials/blob/master/modules/load_medical_images.ipynb)
## Setup environment
```
%pip install -q "monai[itk, nibabel, pillow]"
```
## Setup imports
```
# Copyright 2020 MONAI Consortium
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Copyright 2020 MONAI Consortium
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import os
import shutil
import numpy as np
import itk
from PIL import Image
import tempfile
from monai.data import ITKReader, NibabelReader, PILReader
from monai.transforms import LoadImage, LoadImaged, AddChanneld, Resized, ToTensord, Compose
from monai.config import print_config
print_config()
```
## Load Nifti image with default image reader
MONAI leverages `ITK` as the default image reader, it can support most of the common medical image formats.
More details, please check: https://github.com/InsightSoftwareConsortium/ITK/tree/master/Modules/IO
```
# generate 3D test images
tempdir = tempfile.mkdtemp()
test_image = np.random.rand(64, 128, 96)
filename = os.path.join(tempdir, "test_image.nii.gz")
itk_np_view = itk.image_view_from_array(test_image)
itk.imwrite(itk_np_view, filename)
data, meta = LoadImage()(filename)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load a list of Nifti images and stack as 1 training item
Loading a list of files, stack them together and add a new dimension as first dimension.
And use the meta data of the first image to represent the stacked result.
```
filenames = ["test_image.nii.gz", "test_image2.nii.gz", "test_image3.nii.gz"]
for i, name in enumerate(filenames):
filenames[i] = os.path.join(tempdir, name)
itk_np_view = itk.image_view_from_array(test_image)
itk.imwrite(itk_np_view, filenames[i])
data, meta = LoadImage()(filenames)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load 3D image in DICOM format
```
filename = os.path.join(tempdir, "test_image.dcm")
dcm_image = np.random.randint(256, size=(64, 128, 96)).astype(np.uint8())
itk_np_view = itk.image_view_from_array(dcm_image)
itk.imwrite(itk_np_view, filename)
data, meta = LoadImage()(filename)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load a list of DICOM images and stack as 1 training item
Loading a list of files, stack them together and add a new dimension as first dimension.
And use the meta data of the first image to represent the stacked result.
```
filenames = ["test_image.dcm", "test_image2.dcm", "test_image3.dcm"]
for i, name in enumerate(filenames):
filenames[i] = os.path.join(tempdir, name)
itk_np_view = itk.image_view_from_array(dcm_image)
itk.imwrite(itk_np_view, filenames[i])
data, meta = LoadImage()(filenames)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load 2D image in PNG format
```
test_image = np.random.randint(0, 256, size=[128, 256])
filename = os.path.join(tempdir, "test_image.png")
Image.fromarray(test_image.astype("uint8")).save(filename)
data, meta = LoadImage()(filename)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load image with specified image reader
The `LoadImage` transforms can automatically choose readers based on the supported subffixes and in below order:
- User specified reader at runtime when call this loader.
- Registered readers from the first to the last in list, user can register reader when initializing or at runtime.
- Default ITK reader.
And we can set additional parameters for the image readers, for example, set `c_order_axis_indexing=True` for `ITKReader`, this parameter will pass to ITK `read()` function later.
```
loader = LoadImage()
loader.register(ITKReader())
data, meta = loader(filename)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Load image and execute additional operations
Some image readers can support additional operations after reading the image from file.
For example, we can set a converter for PILReader: `PILReader(converter=lambda image: image.convert("LA"))`.
```
loader = LoadImage(PILReader(converter=lambda image: image.convert("LA")))
data, meta = loader(filename)
print(f"image data shape:{data.shape}")
print(f"meta data:{meta}")
```
## Connect `LoadImage` with other transforms
It's very easy to connect `LoadImage` transform with other transforms to construct a transform chain.
```
transform = Compose([
LoadImaged(keys="image"),
AddChanneld(keys="image"),
Resized(keys="image", spatial_size=[64, 64]),
ToTensord("image"),
])
test_data = {"image": filename}
result = transform(test_data)
print(f"image data shape:{result['image'].shape}")
print(f"meta data:{result['image_meta_dict']}")
```
## Cleanup data directory
Remove directory if a temporary was used.
```
shutil.rmtree(tempdir)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
_**Regression with Aml Compute**_
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Results](#Results)
1. [Test](#Test)
## Introduction
In this example we use an experimental feature, Model Proxy, to do a predict on the best generated model without downloading the model locally. The prediction will happen on same compute and environment that was used to train the model. This feature is currently in the experimental state, which means that the API is prone to changing, please make sure to run on the latest version of this notebook if you face any issues.
This notebook will also leverage MLFlow for saving models, allowing for more portability of the resulting models. See https://docs.microsoft.com/en-us/azure/machine-learning/how-to-use-mlflow for more details around MLFlow is AzureML.
If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../../../../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
In this notebook you will learn how to:
1. Create an `Experiment` in an existing `Workspace`.
2. Configure AutoML using `AutoMLConfig`.
3. Train the model using remote compute.
4. Explore the results.
5. Test the best fitted model.
## Setup
As part of the setup you have already created an Azure ML `Workspace` object. For Automated ML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
import json
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.25.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
ws = Workspace.from_config()
# Choose a name for the experiment.
experiment_name = 'automl-regression-model-proxy'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
output
```
### Using AmlCompute
You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you use `AmlCompute` as your training compute resource.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
# Try to ensure that the cluster name is unique across the notebooks
cpu_cluster_name = "reg-model-proxy"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Data
### Load Data
Load the hardware dataset from a csv file containing both training features and labels. The features are inputs to the model, while the training labels represent the expected output of the model. Next, we'll split the data using random_split and extract the training data for the model.
```
data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/machineData.csv"
dataset = Dataset.Tabular.from_delimited_files(data)
# Split the dataset into train and test datasets
train_data, test_data = dataset.random_split(percentage=0.8, seed=223)
label = "ERP"
```
## Train
Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification, regression or forecasting|
|**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics: <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|(sparse) array-like, shape = [n_samples, n_features]|
|**label_column_name**|(sparse) array-like, shape = [n_samples, ], targets values.|
**_You can find more information about primary metrics_** [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
```
automl_settings = {
"n_cross_validations": 3,
"primary_metric": 'r2_score',
"enable_early_stopping": True,
"experiment_timeout_hours": 0.3, #for real scenarios we recommend a timeout of at least one hour
"max_concurrent_iterations": 4,
"max_cores_per_iteration": -1,
"verbosity": logging.INFO,
"save_mlflow": True,
}
automl_config = AutoMLConfig(task = 'regression',
compute_target = compute_target,
training_data = train_data,
label_column_name = label,
**automl_settings
)
```
Call the `submit` method on the experiment object and pass the run configuration. Execution of remote runs is asynchronous. Depending on the data and the number of iterations this can run for a while. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output = False)
# If you need to retrieve a run that already started, use the following code
#from azureml.train.automl.run import AutoMLRun
#remote_run = AutoMLRun(experiment = experiment, run_id = '<replace with your run id>')
remote_run
```
## Results
```
remote_run.wait_for_completion(show_output=True)
```
### Retrieve the Best Child Run
Below we select the best pipeline from our iterations. The `get_best_child` method returns the best run. Overloads on `get_best_child` allow you to retrieve the best run for *any* logged metric.
```
best_run = remote_run.get_best_child()
print(best_run)
```
#### Show hyperparameters
Show the model pipeline used for the best run with its hyperparameters.
```
run_properties = json.loads(best_run.get_details()['properties']['pipeline_script'])
print(json.dumps(run_properties, indent = 1))
```
#### Best Child Run Based on Any Other Metric
Show the run and the model that has the smallest `root_mean_squared_error` value (which turned out to be the same as the one with largest `spearman_correlation` value):
```
lookup_metric = "root_mean_squared_error"
best_run = remote_run.get_best_child(metric = lookup_metric)
print(best_run)
y_test = test_data.keep_columns('ERP')
test_data = test_data.drop_columns('ERP')
y_train = train_data.keep_columns('ERP')
train_data = train_data.drop_columns('ERP')
```
#### Creating ModelProxy for submitting prediction runs to the training environment.
We will create a ModelProxy for the best child run, which will allow us to submit a run that does the prediction in the training environment. Unlike the local client, which can have different versions of some libraries, the training environment will have all the compatible libraries for the model already.
```
from azureml.train.automl.model_proxy import ModelProxy
best_model_proxy = ModelProxy(best_run)
y_pred_train = best_model_proxy.predict(train_data)
y_pred_test = best_model_proxy.predict(test_data)
```
#### Exploring results
```
y_pred_train = y_pred_train.to_pandas_dataframe().values.flatten()
y_train = y_train.to_pandas_dataframe().values.flatten()
y_residual_train = y_train - y_pred_train
y_pred_test = y_pred_test.to_pandas_dataframe().values.flatten()
y_test = y_test.to_pandas_dataframe().values.flatten()
y_residual_test = y_test - y_pred_test
print(y_residual_train)
print(y_residual_test)
```
| github_jupyter |
[](https://colab.research.google.com/github/ThomasAlbin/Astroniz-YT-Tutorials/blob/main/[ML1]-Asteroid-Spectra/14_dl_autoencoder_gmm_check.ipynb)
# Step 14: GMM - The right way?
Last time we finialized our project, using GMM to cluster the laten space of the Autoencoder's result.
Some would say that this is the final milestone of an un-supervised model. But wait! Do Gaussian describe our data really that good? How can we be sure? Let jump to [cell 15](#SessionStart), to get a FINAL answer!
(I promise, this is the very last session, no tricks!)
```
# Import standard libraries
import os
# Import installed libraries
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
# Scikit-Learn stuff
import sklearn
from sklearn import preprocessing
from sklearn.model_selection import StratifiedShuffleSplit
# Keras
import tensorflow.keras as keras
import tensorflow as tf
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# Set seeds to create reproducible experiments
np.random.seed(1)
tf.random.set_seed(1)
# Let's mount the Google Drive, where we store files and models (if applicable, otherwise work
# locally)
try:
from google.colab import drive
drive.mount('/gdrive')
core_path = "/gdrive/MyDrive/Colab/asteroid_taxonomy/"
except ModuleNotFoundError:
core_path = ""
# Load the level 2 asteroid data
asteroids_df = pd.read_pickle(os.path.join(core_path, "data/lvl2/", "asteroids.pkl"))
# Allocate the spectra to one array and the classes to another one
asteroids_X = np.array([k["Reflectance_norm550nm"].tolist() for k in asteroids_df["SpectrumDF"]])
asteroids_y = np.array(asteroids_df["Main_Group"].to_list())
asteroids_y_bus = np.array(asteroids_df["Bus_Class"].to_list())
```
## ShuffleSplit
Although we do not conduct a classification ML experiment, we still consider the distribution of the classes to train our network properly.
```
# In this example we create a single test-training split with a ratio of 0.8 / 0.2
sss = StratifiedShuffleSplit(n_splits=1, test_size=0.2)
# Create a simple, single train / test split
for train_index, test_index in sss.split(asteroids_X, asteroids_y):
X_train, X_test = asteroids_X[train_index], asteroids_X[test_index]
y_train, y_test = asteroids_y[train_index], asteroids_y[test_index]
y_train_bus, y_test_bus = asteroids_y_bus[train_index], asteroids_y_bus[test_index]
```
## Scaling
This time we are creating a scikit-learn scaler for our spectra data. The model's prediction signals need to be transformed inversely later on to display them correctly.
```
# Import the preprocessing module
from sklearn import preprocessing
# Instantiate the StandardScaler (mean 0, standard deviation 1) and use the training data to fit
# the scaler
scaler = preprocessing.StandardScaler().fit(X_train)
# Transform now the training data
X_train_scaled = scaler.transform(X_train)
# Scale the testing data ...
X_test_scaled = scaler.transform(X_test)
# And expanding the dimensionality for our ConvNet-based Autoencoder
X_train_scaled = np.expand_dims(X_train_scaled, axis=2)
X_test_scaled = np.expand_dims(X_test_scaled, axis=2)
```
## Building the Autoencoder
Now we create a ConvNet Autoencoder. Note that we are not using Keras-Tuner this time. Feel free to apply Keras-Tuner as a small coding exercise.
```
# Get the number of inputs
n_inputs = asteroids_X.shape[1]
# Let's create an autoencoder with a 2-D latent space
n_bottleneck = 5
def create_model():
# Input layer, this time without a normalisation layer
input_layer = keras.Input(shape=(n_inputs, 1))
# Conv Layers (we won't be using maxpooling, since the dimensionaliy is 49 and we do not
# alter the data in our example
hidden_layer = keras.layers.Conv1D(filters=16,
activation="relu",
kernel_size=3, padding="same")(input_layer)
hidden_layer = keras.layers.Conv1D(filters=32,
activation="relu",
kernel_size=3, padding="same")(hidden_layer)
# Encoder ("Bottleneck" of the Autoencoder)
bottleneck_lay = keras.layers.Flatten()(hidden_layer)
bottleneck_lay = keras.layers.Dense(n_bottleneck)(bottleneck_lay)
# The original shape must be restored and reshaped accordingly
reset_lay = keras.layers.Dense(49*32)(bottleneck_lay)
reshape_lay = keras.layers.Reshape((49, 32))(reset_lay)
# First and second hidden decoder layers
hidden_layer = keras.layers.Conv1DTranspose(filters=32,
kernel_size=3,
strides=1,
activation="relu",
padding="same")(reshape_lay)
hidden_layer = keras.layers.Conv1DTranspose(filters=16,
kernel_size=3,
strides=1,
activation="relu",
padding="same")(hidden_layer)
# Ouput layer (same size as input layer)
output_layer = keras.layers.Conv1D(1, 1, padding="same")(hidden_layer)
# Create model
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
# Create encoder model
encoder_model = keras.models.Model(inputs=input_layer, outputs=bottleneck_lay)
# We return the model and the encoder
return model, encoder_model
model, encoder_model = create_model()
# Compile the model and use a regression loss function
model.compile(optimizer='adam', loss='mse')
# Show the model summary
model.summary()
# Train the model
end_epoch = 500
batch_size = 32
# Early Stopping for our final model
es_callback = keras.callbacks.EarlyStopping(monitor='val_loss', patience=5)
history = model.fit(X_train_scaled, X_train_scaled,
epochs=end_epoch,
batch_size=batch_size,
verbose=0,
validation_split=0.25,
callbacks=[es_callback])
```
## The loss function
Let's show the loss of the training and test data. As you can see, the minimum-plateau is reached quite fast. The test data performs slightly better than the training data, since the loss results for the training data are based on an average of the batch size. The test results are based on all data.
```
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# plot the training and validation loss
plt.figure(figsize=(10,8))
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
# Add legend and labels
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("MSE Loss")
# ... aaaaand plot!
plt.show()
```
## Signal Reconstruction
... can be done by using the entire model to predict the output spectra. The following code snippet takes a signal (change the index value in the first line to use any other signal) and predicts the Autoencoder based spectrum.
As one can see, the results are "quite ok" but not perfect. The shape of the spectra can be reproduced. However, the signals are quite noisy.
```
# Which index shall be displayed?
index_val = 5
# Original signal
org_signal = scaler.inverse_transform(X_train_scaled[index_val].reshape(1, -1))[0]
# Reconstructed signal
rec_signal = scaler.inverse_transform(model.predict(X_train_scaled)[index_val].reshape(1, -1))[0]
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# plot the training and reconstructed data
plt.figure(figsize=(12,5))
plt.plot(org_signal, label='Original')
plt.plot(rec_signal, label='Reconstructed')
# Add legend and labels
plt.legend()
plt.xlabel("Spectra: array index")
plt.ylabel("Normalized Reflectance")
# ... aaaaand plot!
plt.show()
# Create dataframe that contains the encoder values and the corresponding class to see whether the
# autoencoder values cluster in a way
# Encode the spectra
X_train_encoded = encoder_model.predict(X_train_scaled)
X_test_encoded = encoder_model.predict(X_test_scaled)
# Merge the data
X_encoded = np.vstack((X_train_encoded, X_test_encoded))
# Instantiate the StandardScaler (mean 0, standard deviation 1) and use the encoded data to fit
# the scaler
enc_scaler = preprocessing.StandardScaler().fit(X_encoded)
# Transform now the encoded data (later used for our scikit-learn method)
X_encoded_scaled = enc_scaler.transform(X_encoded)
# Merge the classes
y_main = np.hstack((y_train, y_test))
y_bus = np.hstack((y_train_bus, y_test_bus))
# Create a column names array for the encoded space
encoder_space = [f"enc{enc_nr+1}" for enc_nr in range(n_bottleneck)]
encoder_space_cols = encoder_space.copy()
encoder_space.extend(["Main_Group", "Bus_Class"])
# Create the dataframe
encoded_df = pd.DataFrame(np.hstack((X_encoded_scaled,
y_main[np.newaxis].transpose(),
y_bus[np.newaxis].transpose())),
columns=encoder_space)
# Change the dtype to float
encoded_df.loc[:, encoder_space[:-2]] = encoded_df.loc[:, encoder_space[:-2]].astype(float)
```
## Gaussian Mixture Model
With the encoded values we will now conduct a clustering experiment using Gaussian Mixture Models (GMMs). The multi-dimensional data will be used to fit a varying number of Gaussians.
The question is: How many Gaussians are needed to fit the data... "properly"?
For this, we will fit an increasing number of Gaussians and compute the corresponding, so-called *Bayesion Information Criterion* (BIC). This values ia a quantified version of Occam's Razor: more parameters may describe the data better, but more parameters will be punished. Goal: finding the perfect balance between "explainability" and "overfitting".
```
import sklearn.mixture
import tqdm
# Result dataframe that will contain the number of Gaussian components, the Bayesion Information
# Criterion (BIC) and the model itself
gmm_results_df = pd.DataFrame([], columns=["Nr_Comp", "BIC", "Model"])
# We iterate through a number of "component guesses"
max_gauss = 15
for index, gauss_components in tqdm.tqdm(enumerate(np.arange(1, max_gauss+1, 1))):
# Create and fit a temporary Gaussian Mixture Model
temp_gmm = sklearn.mixture.GaussianMixture(n_components=gauss_components,
covariance_type="full")
temp_gmm.fit(X_encoded_scaled)
# Store the number of components, the BIC and the model
gmm_results_df.loc[index, "Nr_Comp"] = gauss_components
gmm_results_df.loc[index, "BIC"] = temp_gmm.bic(X_encoded_scaled)
gmm_results_df.loc[index, "Model"] = temp_gmm
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# Plotting the BIC vs. the number of components
plt.figure(figsize=(10, 8))
plt.plot(gmm_results_df["Nr_Comp"],
gmm_results_df["BIC"],
linestyle="dashed",
marker="o",
markersize=5,
color="w",
alpha=0.7)
# Color the minimum value
gmm_results_best = gmm_results_df.loc[gmm_results_df["BIC"] == gmm_results_df["BIC"].min()]
plt.plot(gmm_results_best["Nr_Comp"],
gmm_results_best["BIC"],
marker="o",
markersize=15,
color="tab:green",
alpha=0.7)
# Some formatting
plt.xlabel("Number of Gaussian Components")
plt.ylabel("BIC")
plt.grid(linestyle="dashed", alpha=0.3)
plt.xlim(1, max_gauss)
# Let's take the best GMM!
best_gmm = gmm_results_best["Model"].iloc[0]
# Create a new dataframe column that labels the spectra based on our GMM model:
encoded_df.loc[:, "GMM_Class"] = best_gmm.predict(encoded_df[encoder_space_cols].values)
encoded_df
# Groupby the Main group and the GMM classification
encoded_grouped_df = pd.crosstab(index=encoded_df["Main_Group"],
columns=encoded_df["GMM_Class"],
values=encoded_df["enc1"],
aggfunc="count")
# Extract data, column and index names for plotting purposes
encoded_grouped_values = encoded_grouped_df.values
encoded_grouped_main = encoded_grouped_df.index.values
encoded_grouped_gmm = [f"C{k}" for k in encoded_grouped_df.columns.values]
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# Create a matrix-like plot of the results
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
cax = ax.matshow(encoded_grouped_values, cmap="afmhot")
fig.colorbar(cax, label="Number of spectra", fraction=0.05)
# Some plotting settings
ax.set_xticks(range(len(encoded_grouped_gmm)))
ax.set_yticks(range(len(encoded_grouped_main)))
ax.set_xticklabels(encoded_grouped_gmm)
ax.set_yticklabels(encoded_grouped_main)
ax.set_xlabel("GMM based classification")
ax.set_ylabel("Main Group")
ax.xaxis.set_label_position('top')
```
## GMM Check
<a id='SessionStart'></a>
Let's answer the question: are GMM the perfect choice? Let's focus on the S-types!
```
# Consider only the S-types and let's check the corresponding GMM class
encoded_main_s_df = encoded_df.loc[encoded_df["Main_Group"] == "S"].copy()
encoded_main_s_df.groupby("GMM_Class").count()
# We compute now the prediction probability of each laten vector
encoded_grouped_gmm_prob = [f"C{k}Prob" for k in encoded_grouped_df.columns.values]
encoded_main_s_df.loc[:, encoded_grouped_gmm_prob] = \
best_gmm.predict_proba(encoded_main_s_df[encoder_space_cols].values)
encoded_main_s_df
# In our case we use the very first Gaussian (please note: this is hardcoded and should be
# reproducible, considering the set seeds. If you change the seeds, check in cell 15 the "most
# precent" GMM Class and change the name here accordingly
gmm_cl = "C0Prob"
# Let's plot the probability distribution
plt.figure(figsize=(10, 8))
plt.hist(encoded_main_s_df[gmm_cl], bins=np.arange(0, 1.1, 0.05), color="tab:orange")
plt.xlim(0, 1)
plt.xlabel(f"Probability of class {gmm_cl}")
plt.ylabel("Number of spectra")
# In our case, over 250 spectra are assigned to Cluster 0 with a probability >= 95 %
# ... so we check only the spectra that are most likely linked to this Gaussian
encoded_main_s_df = encoded_main_s_df.loc[encoded_main_s_df[gmm_cl] >= 0.95]
encoded_main_s_df
# We'll use pingouin to ...
!pip install pingouin
import pingouin
# ... compute whether the data describe a Gaussian distribution
encoded_main_s = np.array(encoded_main_s_df[encoder_space_cols].values, dtype="float64")
pingouin.multivariate_normality(encoded_main_s)
```
# Summary & Outlook
Well, be aware what kind of clustering algorithm you choose! Sometimes, pitfalls are not always apparent, especially if the data is multi-dimensional. [scikit-learn](https://scikit-learn.org/stable/modules/clustering.html) provides a nice overview of other clustering algorithms. Try it out!
We conclude now our asteroid spectra part. Of course the journey is not finished and one could extend this project e.g., by considering the orbital elements, detection bias effects and other topics. But that's something for the future, maybe.
Our next project will focus on Near-Earth Objects (NEOs); and we'll answer the question how we could optimize our survey strategy to optimize the search for more NEOs! Stay tuned!
| github_jupyter |
```
import sklearn
import numpy as np
import sklearn.datasets as skd
import ast
from sklearn.feature_extraction import DictVectorizer
from sklearn import linear_model
from sklearn.svm import SVC
from sklearn.metrics import precision_recall_fscore_support
from sklearn.neighbors.nearest_centroid import NearestCentroid
from sklearn.metrics.pairwise import pairwise_distances
from sklearn.metrics import confusion_matrix
from scipy.sparse import vstack
import matplotlib.pyplot as plt
import itertools
import pickle
file = open("mr_train.obj",'rb')
mr_train = pickle.load(file)
file.close()
file = open("mr_test.obj",'rb')
mr_test = pickle.load(file)
file.close()
file = open("mr_cv.obj",'rb')
mr_cv = pickle.load(file)
file.close()
'''
file = open("b_train.obj",'rb')
b = pickle.load(file)
file.close()
file = open("c_cv.obj",'rb')
c = pickle.load(file)
file.close()
file = open("d_test.obj",'rb')
d = pickle.load(file)
file.close()
'''
file = open("x_train.obj",'rb')
x_train = pickle.load(file)
file.close()
file = open("x_test.obj",'rb')
x_test = pickle.load(file)
file.close()
file = open("x_cv.obj",'rb')
x_cv = pickle.load(file)
file.close()
print(len(mr_train.data))
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.figure(figsize=(20,10))
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45,fontsize=10)
plt.yticks(tick_marks, classes,fontsize=10)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
print("Training data has %d malware samples and %d features" % (x_train.shape[0], x_train.shape[1]))
print("Crossval data has %d malware samples and %d features" % (x_cv.shape[0], x_cv.shape[1]))
print("Test data has %d malware samples and %d features" % (x_test.shape[0], x_test.shape[1]))
print("Performing IG Feature selection...")
indices=np.argsort(np.asarray(x_train.sum(axis=0)).ravel(),axis=0)[::-1][:5000]
x_train_ig = x_train[:,indices]
x_cv_ig = x_cv[:,indices]
x_test_ig = x_test[:,indices]
ker='linear'
c=0.01
print("Training SVM Classifier with %s kernel and %.4f C ..." % (ker,c))
SVM = SVC(C=c,kernel=ker)
SVM.fit(x_train_ig,mr_train.target)
print("Obtaining predictions on test data...")
y_pred_cv=SVM.predict(x_cv_ig)
y_pred_test=SVM.predict(x_test_ig)
prec_cv, rec_cv, fsc_cv, sup_cv = precision_recall_fscore_support(mr_cv.target, y_pred_cv, average='weighted')
prec_test, rec_test, fsc_test, sup_test = precision_recall_fscore_support(mr_test.target, y_pred_test, average='weighted')
print("Precision on crossval data is %.4f" % prec_cv)
print("Recall on crossval data is %.4f" % rec_cv)
print("Precision on test data is %.4f" % prec_test)
print("Recall on test data is %.4f" % rec_test)
#Distance measure to hyperplanes
print("Computing mean distance of training samples of each class to their hyperplane")
t=np.array([])
#m=np.array([])
#std=np.array([])
alpha=1
'''
for i in range(len(mr_train.target_names)):
m= np.append(m,np.mean(SVM.decision_function(x_train_ig)[np.argmax(SVM.decision_function(x_train_ig),axis=1)==i][:,i]))
print('{} done'.format(mr_train.target_names[i]))
'''
# f1 = open("m_svm_IG.obj","wb")
# pickle.dump(m,f1)
# f1.close()
file = open("m_svm_IG.obj",'rb')
m = pickle.load(file)
file.close()
print(len(m))
'''
for i in range(len(mr_train.target_names)):
std = np.append(std,np.std(SVM.decision_function(x_train_ig)[np.argmax(SVM.decision_function(x_train_ig),axis=1)==i][:,i]))
print('{} done'.format(i))
'''
# f1 = open("std_svm_IG.obj","wb")
# pickle.dump(std,f1)
# f1.close()
file = open("std_svm_IG.obj",'rb')
std = pickle.load(file)
file.close()
print(len(std))
for i in range(len(mr_train.target_names)):
t=np.append(t,m[i]-alpha*std[i])
print('{} done'.format(i))
print("Finding and selecting drifted objects in test set based on thresholds")
ind_drift_hyp=np.array([])
for i in range(len(mr_train.target_names)):
ind_drift_hyp = np.append(ind_drift_hyp,np.flatnonzero(np.argmax(SVM.decision_function(x_test_ig),axis=1)==i)[np.flatnonzero([SVM.decision_function(x_test_ig)[np.where(np.argmax(SVM.decision_function(x_test_ig),axis=1)==i),i]<t[i]][0][0]==True)])
print('{} done'.format(i))
#f1 = open("ind_drift_hyp.obj","wb")
#pickle.dump(ind_drift_hyp,f1)
#f1.close()
"""
#Distance measure to hyperplanes
print("Computing mean distance of training samples of each class to their hyperplane")
t=np.array([])
m=np.array([])
std=np.array([])
alpha=1
for i in range(len(mr_train.target_names)):
m= np.append(m,np.mean(SVM.decision_function(x_train_ig)[np.argmax(SVM.decision_function(x_train_ig),axis=1)==i][:,i]))
std = np.append(std,np.std(SVM.decision_function(x_train_ig)[np.argmax(SVM.decision_function(x_train_ig),axis=1)==i][:,i]))
t=np.append(t,m[i]-alpha*std[i])
print('{} done'.format(mr_train.target_names[i]))
"""
"""
print("Finding and selecting drifted objects in test set based on thresholds")
ind_drift_hyp=np.array([])
for i in range(len(mr_train.target_names)):
ind_drift_hyp = np.append(ind_drift_hyp,np.flatnonzero(np.argmax(SVM.decision_function(x_test_ig),axis=1)==i)[np.flatnonzero([SVM.decision_function(x_test_ig)[np.where(np.argmax(SVM.decision_function(x_test_ig),axis=1)==i),i]<t[i]][0][0]==True)])
"""
ind_drift_hyp=ind_drift_hyp.astype(int)
print("Total drifted samples in test set: %d out of %d samples" % (ind_drift_hyp.size,len(mr_test.data)))
print("Percentage of drifted samples in test set: %.4f" % (np.double(ind_drift_hyp.size)/len(mr_test.data)))
print("Relabelling drifted samples, and retraining SVM...")
x_train_drift_hyp_drift = vstack([x_train_ig,x_test_ig[ind_drift_hyp]])
mr_train_drift_hyp_drift = np.append(mr_train.target,mr_test.target[ind_drift_hyp],axis=0)
SVM_drift_hyp = SVC(C=c,kernel=ker)
SVM_drift_hyp.fit(x_train_drift_hyp_drift,mr_train_drift_hyp_drift)
print("Computing predictions on test data with newly trained model...")
y_drift_hyp = SVM_drift_hyp.predict(x_test_ig)
prec_drift_hyp, rec_drift_hyp, fsc_drift_hyp, sup_drift_hyp = precision_recall_fscore_support(mr_test.target,y_drift_hyp, average='weighted')
print("Precision on test data with new classes with original model was %.4f" %prec_test)
print("Recall on test data with new classes with original model was %.4f" %rec_test)
print("Precision on test data with new classes with concept drift-aware model %.4f" %prec_drift_hyp)
print("Recall on test data with new classes with concept drift-aware model %.4f" %rec_drift_hyp)
#Distance measure to class centroids
print("Finding class centroids and computing distance of samples to centroids")
clf = NearestCentroid()
clf.fit(x_train_ig,mr_train.target)
dist_train = pairwise_distances(x_train_ig, clf.centroids_)
dist_test = pairwise_distances(x_test_ig, clf.centroids_)
print("Calculating drift_l2 thresholds...")
m = np.resize(np.array([]),8)
var = np.resize(np.array([]),8)
thresh = np.resize(np.array([]),8)
for i in range(8):
m[i] = np.mean(dist_train[np.where(np.argmin(dist_train,axis=1)==i)][:,i])
var[i] = np.sqrt(np.std(dist_train[np.where(np.argmin(dist_train,axis=1)==i)][:,i]))
thresh[i] = m[i]+var[i]
test_drift_l2 = np.resize(np.array([]),8)
test_total = np.resize(np.array([]),8)
test_d_per = np.resize(np.array([]),8)
print("Calculating drift_l2 on test data with new classes...")
for r in range(8):
test_drift_l2[r]=sum(dist_test[np.where(np.argmin(dist_test,axis=1)==r)][:,r] > thresh[r])
test_total[r]= sum(np.argmin(dist_test,axis=1)==r)
if test_total[r]!=0:
test_d_per[r]=test_drift_l2[r]/test_total[r]
else:
test_d_per[r]='nan'
print("In test set there are %d drift_l2ed malware of a total of %d samples, total drift_l2 percentage is %.4f" % (sum(test_drift_l2), sum(test_total), sum(test_drift_l2)/sum(test_total)))
print("Selecting drift_l2ed malware samples from test set...")
ind_array_test = np.array([])
indices_test = np.array([])
for i in range(8):
ind_array_test = np.where(np.argmin(dist_test,axis=1)==i)
indices_test = np.append(indices_test,ind_array_test[0][dist_test[np.where(np.argmin(dist_test,axis=1)==i)][:,i] > thresh[i]])
print("Appending drift_l2ed malware samples from test set to training set, and re-labelling...")
x_train_drift_l2 = vstack([x_train_ig,x_test_ig[indices_test.astype(int)]])
mr_train_drift_l2_target = np.append(mr_train.target,mr_test.target[indices_test.astype(int)],axis=0)
print("Training drift_l2-Aware SVM classifier with new training set...")
SVM_drift_l2 = SVC(C=c,kernel=ker)
SVM_drift_l2.fit(x_train_drift_l2,mr_train_drift_l2_target)
print("Computing predictions on test data with newly trained model...")
y_drift_l2 = SVM_drift_l2.predict(x_test_ig)
prec_drift_l2, rec_drift_l2, fsc_drift_l2, sup_drift_l2 = precision_recall_fscore_support(mr_test.target,y_drift_l2, average='weighted')
print("Precision on test data with new classes with original model was %.4f" %prec_test)
print("Recall on test data with new classes with original model was %.4f" %rec_test)
print("Precision on test data with new classes with concept drift_l2-aware model %.4f" %prec_drift_l2)
print("Recall on test data with new classes with concept drift_l2-aware model %.4f" %rec_drift_l2)
# Computing intersections of flagged samples and actual new class samples for hyperplane drift
intersection = np.intersect1d(np.sort(ind_drift_hyp),np.flatnonzero(mr_test.target > 7))
print(intersection.shape)
print("Precision with alpha = %d is %.4f, recall is %.4f, flagged samples are %d, percentage of samples flagged is %.4f and percentage of flagged samples that actually belong to new families is %.4f" % (alpha,prec_drift_hyp,rec_drift_hyp,len(ind_drift_hyp),np.double(len(ind_drift_hyp))/np.double(len(mr_test.data)),np.double(intersection.shape[0])/np.double(np.flatnonzero(mr_test.target > 7).size)))
#Confusion Matrices
print("Computing confusion matrices...")
cnf_matrix_cv = confusion_matrix(mr_cv.target, y_pred_cv)
cnf_matrix_test = confusion_matrix(mr_test.target,y_pred_test)
cnf_matrix_drift_hyp = confusion_matrix(mr_test.target,y_drift_hyp)
cnf_matrix_drift_l2 = confusion_matrix(mr_test.target,y_drift_l2)
print("Plotting confusion matrix for crossvalidation data")
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix_cv, classes=mr_cv.target_names,
title='Confusion matrix, without normalization, crossval data')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix_cv, classes=mr_cv.target_names, normalize=True,
title='Normalized confusion matrix, crossval data')
plt.show()
print("Plotting confusion matrix for test data in base model")
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix_test, classes=mr_test.target_names,
title='Confusion matrix, without normalization, test data')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix_test, classes=mr_test.target_names, normalize=True,
title='Normalized confusion matrix, test data')
plt.show()
print("Plotting confusion matrix for test data in drift-aware hyperplane model")
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix_drift_hyp, classes=mr_test.target_names,
title='Confusion matrix, without normalization, hyperplane model')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix_drift_hyp, classes=mr_test.target_names, normalize=True,
title='Normalized confusion matrix, hyperplane model')
plt.show()
print("Plotting confusion matrix for test data in drift-aware l2 model")
np.set_printoptions(precision=2)
plt.figure()
plot_confusion_matrix(cnf_matrix_drift_l2, classes=mr_test.target_names,
title='Confusion matrix, without normalization, l2 model')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix_drift_l2, classes=mr_test.target_names, normalize=True,
title='Normalized confusion matrix, l2 model')
plt.show()
```
| github_jupyter |
## Install and import dependencies
[TensorFlow Datasets](https://www.tensorflow.org/datasets/), an API that simplifies downloading and accessing datasets, and provides several sample datasets to work with.
```
!pip install -U tensorflow_datasets
import tensorflow as tf
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
import math
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
tf.enable_eager_execution()
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
```
## Import the Fashion MNIST dataset
```
dataset, metadata = tfds.load('fashion_mnist', as_supervised=True, with_info=True)
train_dataset, test_dataset = dataset['train'], dataset['test']
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
### Explore the data
```
num_train_examples = metadata.splits['train'].num_examples
num_test_examples = metadata.splits['test'].num_examples
print("Number of training examples: {}".format(num_train_examples))
print("Number of test examples: {}".format(num_test_examples))
```
## Preprocess the data
```
# Apply the normalization to each element in the train and test datasets
def normalize(images, labels):
return tf.cast(images, tf.float32) / 255, labels
train_dataset = train_dataset.map(normalize)
test_dataset = test_dataset.map(normalize)
```
### Explore the precessed data
```
# Take a single image, and remove the color dimension by reshaping
for image, label in test_dataset.take(1):
break
image = image.numpy().reshape((28, 28))
plt.figure()
plt.imshow(image, cmap=plt.cm.binary)
plt.colorbar()
plt.grid(False)
plt.show()
plt.figure(figsize=(10,10))
i = 0
for image, label in test_dataset.take(25):
image = image.numpy().reshape((28, 28))
plt.subplot(5, 5, i + 1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(image, cmap=plt.cm.binary)
plt.xlabel(class_names[label])
i += 1
plt.show()
```
## Build the model
Building the neural network requires configuring the layers of the model, then compiling the model.
### Setup the Layers
```
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
```
### Compile the model
Before the model is ready for traning, it needs a few more settings.
+ Loss function
+ Optimizer
+ Metrics - Used to monitor the traning and testing steps.
```
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
## Train the model
```
BATCH_SIZE = 32
train_dataset = train_dataset.repeat().shuffle(num_train_examples).batch(BATCH_SIZE)
test_dataset = test_dataset.batch(BATCH_SIZE)
model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
```
## Evaluate accuracy
```
test_loss, test_accuracy = model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
```
## Make predictions and explore
```
for test_images, test_labels in test_dataset.take(1):
test_images = test_images.numpy()
test_labels = test_labels.numpy()
predictions = model.predict(test_images)
predictions.shape
predictions[0]
print(np.argmax(predictions[0]) == test_labels[0])
def plot_image(i, predictions_array, true_labels, images):
predictions_array, true_label, img = predictions_array[i], true_labels[i], images[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img[..., 0], cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
color = 'blue' if predicted_label == true_label else 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100 * np.max(predictions_array),
class_names[true_label]),
color = color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
this_plot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
this_plot[predicted_label].set_color('red')
this_plot[true_label].set_color('blue')
i = 0
plt.figure(figsize=(6, 3))
plt.subplot(1, 2, 1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1, 2, 2)
plot_value_array(i, predictions, test_labels)
i = 12
plt.figure(figsize=(6, 3))
plt.subplot(1, 2, 1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1, 2, 2)
plot_value_array(i, predictions, test_labels)
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 5
num_cols = 3
num_images = num_rows * num_cols
plt.figure(figsize=(num_cols << 2, num_rows << 1))
for i in range(num_images):
plt.subplot(num_rows, num_cols << 1, i << 1 | 1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, num_cols << 1, (i << 1) + 2)
plot_value_array(i, predictions, test_labels)
```
Finally, we can use the trained model to make a prediction about a single image.
```
def show_single_image(img):
plt.figure()
plt.imshow(img.reshape((28, 28)), cmap=plt.cm.binary)
plt.colorbar()
plt.grid(False)
plt.show()
import random
idx = random.randint(0, 32 - 1) # Make a random selection
img = test_images[idx]
ans_label = test_labels[idx]
print(img.shape)
show_single_image(img)
print(class_names[ans_label])
# Add the image to a batch where it's the only member.
img = np.array([img])
print(img.shape)
```
Now predict the image
```
predictions_single = model.predict(img)
print(predictions_single)
ans = np.argmax(predictions_single[0])
print("Prediction:", class_names[ans], "\nLabel:", class_names[ans_label], "\nResult:", ans == ans_label)
plot_value_array(0, predictions_single, np.array([ans_label]))
_ = plt.xticks(range(10), class_names, rotation=45)
```
# Exercises
Experiment with different models and see how the accuracy results differ. In particular change the following parameters:
* Set training epochs set to 1
* Number of neurons in the Dense layer following the Flatten one. For example, go really low (e.g. 10) in ranges up to 512 and see how accuracy changes
* Add additional Dense layers between the Flatten and the final Dense(10, activation=tf.nn.softmax), experiment with different units in these layers
* Don't normalize the pixel values, and see the effect that has
Remember to enable GPU to make everything run faster (Runtime -> Change runtime type -> Hardware accelerator -> GPU).
Also, if you run into trouble, simply reset the entire environment and start from the beginning:
* Edit -> Clear all outputs
* Runtime -> Reset all runtimes
### Set training epochs set to 1
```
# Define a new model and train & test it
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=1, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
```
### Number of neurons in the Dense layer following the Flatten one.
For example, go really low (e.g. 10) in ranges up to 512 and see how accuracy changes
```
# Define a new model and train & test it
# 10
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(10, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
# Define a new model and train & test it
# 256
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
# Define a new model and train & test it
# 512
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
```
### Add additional Dense layers between the Flatten and the final Dense
```
# Define a new model and train & test it
# 1 Additional Dense layers
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
# Define a new model and train & test it
# 2 Additional Dense layers
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
# Define a new model and train & test it
# 3 Additional Dense layers
temporary_model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
temporary_model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
temporary_model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = temporary_model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
```
### Don't normalize the pixel values
```
dataset, metadata = tfds.load('fashion_mnist', as_supervised=True, with_info=True)
train_dataset, test_dataset = dataset['train'], dataset['test']
num_train_examples = metadata.splits['train'].num_examples
num_test_examples = metadata.splits['test'].num_examples
print("Number of training examples: {}".format(num_train_examples))
print("Number of test examples: {}".format(num_test_examples))
# Take a single image, and remove the color dimension by reshaping
for image, label in test_dataset.take(1):
break
image = image.numpy().reshape((28, 28))
plt.figure()
plt.imshow(image, cmap=plt.cm.binary)
plt.colorbar()
plt.grid(False)
plt.show()
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
BATCH_SIZE = 32
train_dataset = train_dataset.repeat().shuffle(num_train_examples).batch(BATCH_SIZE)
test_dataset = test_dataset.batch(BATCH_SIZE)
model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples / BATCH_SIZE))
test_loss, test_accuracy = model.evaluate(test_dataset, steps=math.ceil(num_test_examples / 32))
print("Accuracy on test dataset: ", test_accuracy)
```
| github_jupyter |
```
%matplotlib inline
```
4D interpolation
=============
Interpolation of a four-dimensional regular grid.
Quadrivariate
----------------
The
[quadrivariate](https://pangeo-pyinterp.readthedocs.io/en/latest/generated/pyinterp.quadrivariate.html#pyinterp.quadrivariate)
interpolation allows obtaining values at arbitrary points in a 4D space of a
function defined on a grid.
The distribution contains a 4D field `pres_temp_4D.nc` that will be used
in this help. This file is located in the `src/pyinterp/tests/dataset` directory
at the root of the project.
This method performs a bilinear interpolation in 2D space by considering the
axes of longitude and latitude of the grid, then performs a linear interpolation
in the third and fourth dimensions. Its interface is similar to the
[trivariate](https://pangeo-pyinterp.readthedocs.io/en/latest/generated/pyinterp.trivariate.html#pyinterp.trivariate)
class except for a fourth axis, which is handled by this object.
```
import cartopy.crs
import matplotlib
import matplotlib.pyplot
import numpy
import pyinterp
import pyinterp.backends.xarray
import pyinterp.tests
import xarray
```
The first step is to load the data into memory and create the
interpolator object:
```
ds = xarray.open_dataset(pyinterp.tests.grid4d_path())
interpolator = pyinterp.backends.xarray.Grid4D(ds.pressure)
```
We will build a new grid that will be used to build a new interpolated
grid.
```
mx, my, mz, mu = numpy.meshgrid(numpy.arange(-125, -70, 0.5),
numpy.arange(25, 50, 0.5),
numpy.datetime64("2000-01-01T12:00"),
0.5,
indexing="ij")
```
We interpolate our grid using a
[classical](https://pangeo-pyinterp.readthedocs.io/en/latest/generated/pyinterp.backends.xarray.Grid4D.quadrivariate.html#pyinterp.backends.xarray.Grid4D.quadrivariate):
```
quadrivariate = interpolator.quadrivariate(
dict(longitude=mx.flatten(),
latitude=my.flatten(),
time=mz.flatten(),
level=mu.flatten())).reshape(mx.shape)
```
Bicubic on 4D grid
----------------------
The grid used organizes the latitudes in descending order. We ask our
constructor to flip this axis in order to correctly evaluate the bicubic
interpolation from this 4D cube (only necessary to perform a bicubic
interpolation).
```
interpolator = pyinterp.backends.xarray.Grid4D(ds.pressure,
increasing_axes=True)
```
We interpolate our grid using a
[bicubic](https://pangeo-pyinterp.readthedocs.io/en/latest/generated/pyinterp.backends.xarray.Grid4D.quadrivariate.html#pyinterp.backends.xarray.Grid4D.bicubic)
interpolation in space followed by a linear interpolation in the temporal axis:
```
bicubic = interpolator.bicubic(dict(longitude=mx.flatten(),
latitude=my.flatten(),
time=mz.flatten(),
level=mu.flatten()),
nx=2,
ny=2).reshape(mx.shape)
```
We transform our result cubes into a matrix.
```
quadrivariate = quadrivariate.squeeze(axis=(2, 3))
bicubic = bicubic.squeeze(axis=(2, 3))
lons = mx[:, 0].squeeze()
lats = my[0, :].squeeze()
```
Let's visualize our results.
---
**Note**
The resolution of the example grid is very low (one pixel every one degree)
therefore the calculation window cannot find the required pixels at the edges to
calculate the interpolation correctly. See Chapter [Fill NaN
Values](https://pangeo-pyinterp.readthedocs.io/en/latest/auto_examples/ex_fill_undef.html)
to see how to address this issue.
---
```
fig = matplotlib.pyplot.figure(figsize=(5, 4))
ax1 = fig.add_subplot(
211, projection=cartopy.crs.PlateCarree(central_longitude=180))
pcm = ax1.pcolormesh(lons,
lats,
quadrivariate.T,
cmap='jet',
transform=cartopy.crs.PlateCarree())
ax1.coastlines()
ax1.set_title("Trilinear")
ax2 = fig.add_subplot(
212, projection=cartopy.crs.PlateCarree(central_longitude=180))
pcm = ax2.pcolormesh(lons,
lats,
bicubic.T,
cmap='jet',
transform=cartopy.crs.PlateCarree())
ax2.coastlines()
ax2.set_title("Spline & Linear in time")
fig.colorbar(pcm, ax=[ax1, ax2], shrink=0.8)
fig.show()
```
| github_jupyter |
<H2>MADATORY PYTHON LIBRARIES</H2>
```
import pandas as pd
```
<H2>POSIBLE SELECTION CRITERIA</H2>
If you had a look to the notebook dedicated to the index files (here), you will have detected several properties (raw & derived) that can be used to select netCDFs. Depending on the index file there will be certain criteria that will be missing or will not make any sense to use in order make some subsetting.
```
data = { 'Selection criteria': ['Bounding box','time range', 'time update', 'provider', 'data mode', 'parameters', 'data source', 'platform_category', 'timestamp'], 'index_history.txt': [' ✔', ' ✔',' ✔ (?)', ' ✔',' ✔', ' ✔',' ✔', ' ✔','-'], 'index_monthly.txt': [' ✔', ' ✔',' ✔ (?)', ' ✔',' ✔', ' ✔',' ✔', ' ✔',' ✔'],'index_latest.txt': [' ✔', ' ✔',' ✔', ' ✔',' ✔', ' ✔',' ✔', ' -',' ✔']}
pd.DataFrame(data=data)
```
<H2> SELECTION CRITERIA OVERVIEW </H2>
## 1. Bounding box
Thanks to the properities <i>'geospatial_lat_min','geospatial_lat_max','geospatial_lon_min' and 'geospatial_lon_max'</i> available on the index files as metadata for georeferenceing the content of every netCDF available on history, monthly and latest directories, it is posible to filter those and only keep the ones related to a specific area of interest.
This selection criteria is highly recomended for any netCDF in the latest directory (index_latest.txt), but its usefullness may decay when targeting certain moving platforms (vessels and gliders in particular) in monthly and history directories (index_monthly.txt and index_history.tx). These platforms can be released/deployed more than once (not the case of profilers which are released just once) and at very different locations, resulting in a very big bounding box that will not be any helpful when trying to filter datasets geographically.
```
data = { 'Selection criteria': ['geographical bounding box'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔ ✔ ✔']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_boundingbox.ipynb).
## 2. Time range
Thanks to the properities <i>'time_coverage_start' and 'time_coverage_end' </i> available on the index files as metadata for timereferenceing the content of every netCDF available on history, monthly and latest directories, it is posible to filter those and only keep the ones related to a specific time window of interest.
This selection criteria is highly recomended for any netCDF in the latest, monthly and history directories (index_latest.txt, index_monthly.txt and index_history.txt):
```
data = { 'Selection criteria': ['time range'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_timerange.ipynb)
## 3. Time update
Thanks to the properity <i>'time_update' </i> available on the index files as metadata for timereferenceing the content-update of every netCDF available on history, monthly and latest directories, it is posible to filter those and only keep the ones related to a specific time windown of interest.
This selection criteria is highly recomended for detecting changes in the content of the available neCDFs in the latest, monthly and history directories (index_latest.txt, index_monthly.txt & index_history.txt). Nevertheless, as the update rate of every file in these directories varies (hourly, daily and monthly basis respectively), in order to detect changes in real-time it might be a more effective selection criteria for netCDFs in the latest directory (index_latest.txt).
```
data = { 'Selection criteria': ['time update'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔ ✔']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_timeupdate.ipynb)
## 4. Provider
Thanks to the properity <i>'provider' </i> available on the index files as metadata for timereferenceing the content-update of every netCDF available on history, monthly and latest directories, it is posible to filter those and only keep the ones related to a specific institution.
This selection criteria is highly recomended for finding only those neCDFs beloging to a specific institution and can be apply with same efectivenness in the latest, monthly and history directories (index_latest.txt, index_monthly.txt & index_history.txt). Keep in mind though that the same provider can be referred by its longname and short name, so most institutions does not have a unique name but several.
```
data = { 'Selection criteria': ['provider'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔ ']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_provider.ipynb)
## 5. Parameters
Thanks to the properity <i>'parameters' </i> available on the index files as metadata for referenceing the content of every netCDF (in terms of variables measured) available on history, monthly and latest directories, it is posible to filter those and only keep the ones containing a certain parameter of interest.
This selection criteria is highly recomended for finding only those neCDFs including a certain parameter and can be apply with same efectivenness in the latest, monthly and history directories (index_latest.txt, index_monthly.txt & index_history.txt).
```
data = { 'Selection criteria': ['parameters'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔ ']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_parameter.ipynb)
## 6. Platform's data sources
Thanks to the properity <i>'file_name' </i> available on the index files for facilititating the ftp link to every netCDF available on history, monthly and latest directories, it is posible to filter those and only keep the ones beloging to a certain data source of interest.
The data source is always part of the file name by convention but, the location of such reference within the name structure changes (position 1 in file name for netCDFs in history directory and position 3 in files at monthly and latest directories), so you have to take this into account when filtering.
```
data = { 'Selection criteria': ['data source'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔ ']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_platform_data_source.ipynb)
## 7. Platform category
Thanks to the properity <i>'file_name' </i> available on the index files for facilititating the ftp link to every netCDF available on history, monthly and latest directories, it is posible in case of history and monhtly directories to filter those and only keep the ones beloging to a certain platform category; a much more wider concept than the data source.
```
data = { 'Selection criteria': ['platform category'], 'index_history.txt': [' ✔'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' - ']}
pd.DataFrame(data=data)
```
See example [here](https://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_platform_category.ipynb)
## 8. Timestamp
Thanks to the properity <i>'file_name' </i> available on the index files for facilitating the ftp link to every netCDF available on history, monthly and latest directories, it is posible, in case of monthly and latest directories, to filter those and only keep the ones matching a certain timestamp, explicited stated in the file name convention.
This selection criteria is highly recomended for finding only those neCDFs matching a certain timestamp in monthly and latest directories.
```
data = { 'Selection criteria': ['timestamp'], 'index_history.txt': [' -'], 'index_monthly.txt': [' ✔'],'index_latest.txt': [' ✔']}
pd.DataFrame(data=data)
```
See example [here](hhttps://github.com/CopernicusMarineInsitu/INSTACTraining-Phase2UPDATE/blob/master/PythonNotebooks/In_Situ_data_download_by_timestamp.ipynb)
| github_jupyter |
# MatPlotLib Basics
## Draw a line graph
```
%matplotlib inline
from scipy.stats import norm
import matplotlib.pyplot as plt
import numpy as np
x = np.arange(-3, 3, 0.001)
plt.plot(x, norm.pdf(x))
plt.show()
```
## Mutiple Plots on One Graph
```
plt.plot(x, norm.pdf(x))
plt.plot(x, norm.pdf(x, 1.0, 0.5))
plt.show()
```
## Save it to a File
```
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import norm
x = np.arange(-3, 3, 0.001)
plt.plot(x, norm.pdf(x))
plt.plot(x, norm.pdf(x, 1.0, 0.5))
plt.savefig('MyPlot.png', format='png')
```
## Adjust the Axes
```
axes = plt.axes()
axes.set_xlim([-5, 5])
axes.set_ylim([0, 1.0])
axes.set_xticks([-5, -4, -3, -2, -1, 0, 1, 2, 3, 4, 5])
axes.set_yticks([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])
plt.plot(x, norm.pdf(x))
plt.plot(x, norm.pdf(x, 1.0, 0.5))
plt.show()
```
## Add a Grid
```
axes = plt.axes()
axes.set_xlim([-5, 5])
axes.set_ylim([0, 1.0])
axes.set_xticks([-5, -4, -3, -2, -1, 0, 1, 2, 3, 4, 5])
axes.set_yticks([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])
axes.grid()
plt.plot(x, norm.pdf(x))
plt.plot(x, norm.pdf(x, 1.0, 0.5))
plt.show()
```
## Change Line Types and Colors
```
axes = plt.axes()
axes.set_xlim([-5, 5])
axes.set_ylim([0, 1.0])
axes.set_xticks([-5, -4, -3, -2, -1, 0, 1, 2, 3, 4, 5])
axes.set_yticks([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])
axes.grid()
plt.plot(x, norm.pdf(x), 'b-')
plt.plot(x, norm.pdf(x, 1.0, 0.5), 'r:,')
plt.show()
```
## Labeling Axes and Adding a Legend
```
axes = plt.axes()
axes.set_xlim([-5, 5])
axes.set_ylim([0, 1.0])
axes.set_xticks([-5, -4, -3, -2, -1, 0, 1, 2, 3, 4, 5])
axes.set_yticks([0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0])
axes.grid()
plt.xlabel('Greebles')
plt.ylabel('Probability')
plt.plot(x, norm.pdf(x), 'b-')
plt.plot(x, norm.pdf(x, 1.0, 0.5), 'r:')
plt.legend(['Sneetches', 'Gacks'], loc=4)
plt.show()
```
## XKCD Style :)
```
plt.xkcd()
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
plt.xticks([])
plt.yticks([])
ax.set_ylim([-30, 10])
data = np.ones(100)
data[70:] -= np.arange(30)
plt.annotate(
'THE DAY I REALIZED\nI COULD COOK BACON\nWHENEVER I WANTED',
xy=(70, 1), arrowprops=dict(arrowstyle='->'), xytext=(15, -10))
plt.plot(data)
plt.xlabel('time')
plt.ylabel('my overall health')
```
## Pie Chart
```
# Remove XKCD mode:
plt.rcdefaults()
values = [12, 55, 4, 32, 14]
colors = ['r', 'g', 'b', 'c', 'm']
explode = [0, 0, 0.2, 0, 0]
labels = ['India', 'United States', 'Russia', 'China', 'Europe']
plt.pie(values, colors= colors, labels=labels, explode = explode)
plt.title('Student Locations')
plt.show()
```
## Bar Chart
```
values = [12, 55, 4, 32, 14]
colors = ['r', 'g', 'b', 'c', 'm']
plt.bar(range(0,5), values, color= colors)
plt.show()
```
## Scatter Plot
```
from pylab import randn
X = randn(500)
Y = randn(500)
plt.scatter(X,Y)
plt.show()
```
## Histogram
```
incomes = np.random.normal(27000, 15000, 10000)
plt.hist(incomes, 50)
plt.show()
```
## Box & Whisker Plot
Useful for visualizing the spread & skew of data.
The red line represents the median of the data, and the box represents the bounds of the 1st and 3rd quartiles.
So, half of the data exists within the box.
The dotted-line "whiskers" indicate the range of the data - except for outliers, which are plotted outside the whiskers. Outliers are 1.5X or more the interquartile range.
This example below creates uniformly distributed random numbers between -40 and 60, plus a few outliers above 100 and below -100:
```
uniformSkewed = np.random.rand(100) * 100 - 40
high_outliers = np.random.rand(10) * 50 + 100
low_outliers = np.random.rand(10) * -50 - 100
data = np.concatenate((uniformSkewed, high_outliers, low_outliers))
plt.boxplot(data)
plt.show()
```
## Activity
Try creating a scatter plot representing random data on age vs. time spent watching TV. Label the axes.
| github_jupyter |
# 04장. 자료의 종류에는 어떤 것들이 있나요?
## 01. 파이썬에서 사용할 수 있는 자료의 종류
- 모든 프로그램은 자료(data)를 처리함.
- 자료는 종류에 따라 자료형(data type)으로 구분 지어 놓음
- 자료형: 정수, 실수, 문자열 등
```
x = 10
print('x=',x)
x = 3.14
print('x=',x)
x = 'Hello World!'
print('x=',x)
```
## 02. 정수형과 실수형
- 수치형 데이터(정수, 실수, 복소수)는 산술연산을 할 수 있음
```
x = 123
print(x)
print(type(x))
x = -123
print(x)
print(type(x))
x = 0
print(x)
print(type(x))
x = 3.14
print(x)
print(type(x))
x = -3.14
print(x)
print(type(x))
x = 0.0
print(x)
print(type(x))
```
## 03. 문자열이란?
- 문자열(string): 문자들의 나열(sequence of characters)
- 작은따옴표와 큰따옴표를 사용
```
'hello'
msg = "hello"
msg
print(msg)
```
## 04. 문자열을 만드는 방법
- 큰 따옴표나 작은 따옴표를 한 쌍으로 열고 닫음: “ “, ‘ ‘
```
msg = "hello'
msg="She said 'Hi'"
print(msg)
```
## 05. 100과 "100"을 구별해요
```
print(100+200)
print("100"+"200")
```
## 06. 문자열을 숫자로 변환
- input( )은 사용자가 입력한 데이터를 문자열 형태로 돌려줌
```
t = input("정수를 입력하시오: ")
x = int(t)
t = input("정수를 입력하시오: ")
y = int(t)
print(x+y)
x = float(input("실수를 입력하시오: "))
y = float(input("실수를 입력하시오: "))
print(x+y)
```
## 07. 숫자를 문자열로 변환
```
print("나는 현재" + 17 + "살이다.")
print("나는 현재 " + str(17) + "살이다.")
print("원주율은 " + str(3.14) + "입니다.")
```
## 08. 파이썬은 문자열 처리의 마술사
```
'Hello' + 'World'
fist_name = '길동'
lask_name = '홍'
name = lask_name+fist_name
print(name)
message = 'Congratulations!'
print(message * 3)
line = "="*30
print(line)
price = 1000
print("상품의 가격은 %s원 입니다" % price)
poem = "이렇게 정다운\n너 하나 나 하나는\n어디서 무엇이 되어\n다시 만나랴."
print(poem)
poem = '''이렇게 정다운
너 하나 나 하나는
어디서 무엇이 되어
다시 만나랴.'''
print(poem)
```
## 09. 필요한 문자열을 뽑아서 쓰자
- 문자열 변수 s[a : b]: 인덱스 a부터 b-1까지의 문자열 (a < b)
- s[a:b:c]:
- a < b이고 c > 0이면, a부터 b-1까지의 c간격의 문자열
- a > b이고 c < 0이면, a부터 b+1까지의 c간격의 문자열
- s[0: :1]: s[0]에서부터 문자열 끝까지 1간격의 문자열, s 문자열 전체
- s[-1: :-1]: s[-1]에서부터 문자열 처음까지 -1간격의 문자열, s 문자열 거꾸로 전체
- s[:]: s 문자열 전체
```
s = "Hello Python"
print(s[0])
print(s[1])
print(s[-1])
s = "Hello Python"
print(s[6:9])
print(s[-6:-2])
print(s[0:10:2])
print(s[-1:-7:-1])
```
## Lab. 소금물의 농도는?
- 소금물의 농도 = (소금의 양/소금물의 양) * 100(%)
```
print("소금물의 농도를 구하는 프로그램입니다")
salt = int(input("소금의 양은 몇 g입니까? "))
water = int(input("물의 양은 몇 g입니까? "))
density = salt / (salt+water) * 100
print("소금물의 농도: " + str(density) + "%")
```
## Lab. 간단한 챗봇(ChatBot) 프로그램
```
print("안녕하세요")
name = input('이름이 뭐예요? ')
print("만나서 반갑습니다. " + name + "님")
print(name + "님 이름의 길이는 다음과 같군요:", end=' ')
print(len(name))
age = int(input("나이가 어떻게 되요? "))
print("내년이면 "+ str(age+1)+ "세가 되시는 군요")
```
## Lab. 거북이와 인사해봐요
```
# import turtle
# t = turtle.Turtle()
# t.shape("turtle")
# s = turtle.textinput("", "이름을 입력하시오: ")
# t.write("안녕하세요?" + s +"씨, 터틀 인사드립니다.");
# #사각형 그리기
# t.left(90)
# t.forward(100)
# t.left(90)
# t.forward(100)
# t.left(90)
# t.forward(100)
# t.left(90)
# t.forward(100)
```
## Lab. 암호프로그램 만들기
```
P = "도서관에서 보자"
print("평문:",P)
print("암호문:", P[-1: -9 :-1])
```
## Lab. 2050년에 나는 몇살?
```
import time
now = time.time()
thisYear = int(1970 + now//(365*24*3600))
print("올해는 " + str(thisYear)+"년입니다.")
age = int(input("당신의 나이를 입력 하세요: "))
print("2050년에는 "+str(age + 2050-thisYear)+"살이군요.")
```
--- seYi
| github_jupyter |
# Coordinate descent
CuML library can implement lasso and elastic net algorithms. The lasso model extends LinearRegression with L2 regularization and elastic net extends LinearRegression with a combination of L1 and L2 regularizations.
We see tremendous speed up for datasets with large number of rows and less number of rows. Furthermore, the MSE value for the cuML implementation is much smaller than the scikit-learn implementation for very small datasets.
```
# Select a particular GPU to run the notebook (if needed)
# %env CUDA_VISIBLE_DEVICES=2
# Import the required libraries
import numpy as np
import pandas as pd
import cudf
import os
from cuml import Lasso as cuLasso
from sklearn.linear_model import Lasso
from sklearn.datasets import make_regression
from sklearn.metrics import mean_squared_error
from cuml.linear_model import ElasticNet as cuElasticNet
from sklearn.linear_model import ElasticNet
```
# Helper Functions
```
# Check if the mortgage dataset is present and then extract the data from it, else just create a random dataset for regression
import gzip
def load_data(nrows, ncols, cached = 'data/mortgage.npy.gz'):
# Split the dataset in a 80:20 split
train_rows = int(nrows*0.8)
if os.path.exists(cached):
print('use mortgage data')
with gzip.open(cached) as f:
X = np.load(f)
# The 4th column is 'adj_remaining_months_to_maturity'
# used as the label
X = X[:,[i for i in range(X.shape[1]) if i!=4]]
y = X[:,4:5]
rindices = np.random.randint(0,X.shape[0]-1,nrows)
X = X[rindices,:ncols]
y = y[rindices]
df_y_train = pd.DataFrame({'fea%d'%i:y[0:train_rows,i] for i in range(y.shape[1])})
df_y_test = pd.DataFrame({'fea%d'%i:y[train_rows:,i] for i in range(y.shape[1])})
else:
print('use random data')
X,y = make_regression(n_samples=nrows,n_features=ncols,n_informative=ncols, random_state=0)
df_y_train = pd.DataFrame({'fea0':y[0:train_rows,]})
df_y_test = pd.DataFrame({'fea0':y[train_rows:,]})
df_X_train = pd.DataFrame({'fea%d'%i:X[0:train_rows,i] for i in range(X.shape[1])})
df_X_test = pd.DataFrame({'fea%d'%i:X[train_rows:,i] for i in range(X.shape[1])})
return df_X_train, df_X_test, df_y_train, df_y_test
```
# Obtain and convert the dataset
```
%%time
# nrows = number of samples
# ncols = number of features of each sample
nrows = 2**21
ncols = 500
# Split the dataset into training and testing sets, in the ratio of 80:20 respectively
X_train, X_test, y_train, y_test = load_data(nrows,ncols)
print('training data',X_train.shape)
print('training label',y_train.shape)
print('testing data',X_test.shape)
print('testing label',y_test.shape)
print('label',y_test.shape)
%%time
# Convert the pandas dataframe to cudf format
X_cudf = cudf.DataFrame.from_pandas(X_train)
X_cudf_test = cudf.DataFrame.from_pandas(X_test)
y_cudf = y_train.values
y_cudf = y_cudf[:,0]
y_cudf = cudf.Series(y_cudf)
```
### Define the model parameters
```
# lr = learning rate
# algo = algorithm used in the model
lr = 0.001
algo = 'cyclic'
```
### Lasso
The lasso model implemented in cuml allows the user to change the following parameter values:
1. alpha: regularizing constant that is multiplied with L1 to control the extent of regularization. (default = 1)
2. normalize: variable decides if the predictors in X will be normalized or not. (default = False)
3. fit_intercept: if set to True the model tries to center the data. (default = True)
4. max_iter: maximum number of iterations for training (fitting) the data to the model. (default = 1000)
5. tol: the tolerance for optimization. (default = 1e-3)
3. algorithm: the user can set the algorithm value as 'cyclic' or 'random'
The model accepts only numpy arrays or cudf dataframes as the input. In order to convert your dataset to cudf format please read the cudf documentation on https://rapidsai.github.io/projects/cudf/en/latest/. For additional information on the lasso model please refer to the documentation on https://rapidsai.github.io/projects/cuml/en/latest/index.html
#### Scikit-learn model for lasso
```
%%time
# Use the sklearn lasso model to fit the training dataset
skols = Lasso(alpha=np.array([lr]), fit_intercept = True, normalize = False, max_iter = 1000, selection=algo, tol=1e-10)
skols.fit(X_train, y_train)
%%time
# Calculate the mean squared error for the sklearn lasso model on the testing dataset
sk_predict = skols.predict(X_test)
error_sk = mean_squared_error(y_test,sk_predict)
```
#### CuML model for lasso
```
%%time
# Run the cuml linear regression model to fit the training dataset
cuols = cuLasso(alpha=np.array([lr]), fit_intercept = True, normalize = False, max_iter = 1000, selection=algo, tol=1e-10)
cuols.fit(X_cudf, y_cudf)
%%time
# Calculate the mean squared error of the testing dataset using the cuml linear regression model
cu_predict = cuols.predict(X_cudf_test).to_array()
error_cu = mean_squared_error(y_test,cu_predict)
# Print the mean squared error of the sklearn and cuml model to compare the two
print("SKL MSE(y):")
print(error_sk)
print("CUML MSE(y):")
print(error_cu)
```
### Elastic Net
The elastic net model implemented in cuml contains the same parameters as the lasso model.
In addition to the variable values that can be altered in lasso, elastic net has another variable who's value can be changed
- l1_ratio: decides the ratio of amount of L1 and L2 regularization that would be applied to the model. When L1 ratio = 0, the model will have only L2 reqularization shall be applied to the model. (default = 0.5)
The model accepts only numpy arrays or cudf dataframes as the input. In order to convert your dataset to cudf format please read the cudf documentation on https://rapidsai.github.io/projects/cudf/en/latest/. For additional information on the lasso model please refer to the documentation on https://rapidsai.github.io/projects/cuml/en/latest/index.html
#### Scikit-learn model for elastic net
```
%%time
# Use the sklearn linear regression model to fit the training dataset
elastic_sk = ElasticNet(alpha=np.array([lr]), fit_intercept = True, normalize = False, max_iter = 1000, selection=algo, tol=1e-10)
elastic_sk.fit(X_train, y_train)
%%time
# Calculate the mean squared error of the sklearn linear regression model on the testing dataset
sk_predict_elas = elastic_sk.predict(X_test)
error_sk_elas = mean_squared_error(y_test,sk_predict_elas)
```
#### CuML model for elastic net
```
%%time
# Run the cuml linear regression model to fit the training dataset
elastic_cu = cuElasticNet(alpha=np.array([lr]), fit_intercept = True, normalize = False, max_iter = 1000, selection=algo, tol=1e-10)
elastic_cu.fit(X_cudf, y_cudf)
%%time
# Calculate the mean squared error of the testing dataset using the cuml linear regression model
cu_predict_elas = elastic_cu.predict(X_cudf_test).to_array()
error_cu_elas = mean_squared_error(y_test,cu_predict_elas)
# Print the mean squared error of the sklearn and cuml model to compare the two
print("SKL MSE(y):")
print(error_sk_elas)
print("CUML MSE(y):")
print(error_cu_elas)
```
| github_jupyter |
<p><font size="6"><b>Matplotlib: Introduction </b></font></p>
> *DS Data manipulation, analysis and visualisation in Python*
> *December, 2017*
> *© 2016, Joris Van den Bossche and Stijn Van Hoey (<mailto:jorisvandenbossche@gmail.com>, <mailto:stijnvanhoey@gmail.com>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
---
```
%matplotlib inline
```
# Matplotlib
[Matplotlib](http://matplotlib.org/) is a Python package used widely throughout the scientific Python community to produce high quality 2D publication graphics. It transparently supports a wide range of output formats including PNG (and other raster formats), PostScript/EPS, PDF and SVG and has interfaces for all of the major desktop GUI (graphical user interface) toolkits. It is a great package with lots of options.
However, matplotlib is...
> The 800-pound gorilla — and like most 800-pound gorillas, this one should probably be avoided unless you genuinely need its power, e.g., to make a **custom plot** or produce a **publication-ready** graphic.
> (As we’ll see, when it comes to statistical visualization, the preferred tack might be: “do as much as you easily can in your convenience layer of choice [nvdr e.g. directly from Pandas, or with seaborn], and then use matplotlib for the rest.”)
(quote used from [this](https://dansaber.wordpress.com/2016/10/02/a-dramatic-tour-through-pythons-data-visualization-landscape-including-ggplot-and-altair/) blogpost)
And that's we mostly did, just use the `.plot` function of Pandas. So, why do we learn matplotlib? Well, for the *...then use matplotlib for the rest.*; at some point, somehow!
Matplotlib comes with a convenience sub-package called ``pyplot`` which, for consistency with the wider matplotlib community, should always be imported as ``plt``:
```
import numpy as np
import matplotlib.pyplot as plt
```
## - dry stuff - The matplotlib `Figure`, `axes` and `axis`
At the heart of **every** plot is the figure object. The "Figure" object is the top level concept which can be drawn to one of the many output formats, or simply just to screen. Any object which can be drawn in this way is known as an "Artist" in matplotlib.
Lets create our first artist using pyplot, and then show it:
```
fig = plt.figure()
plt.show()
```
On its own, drawing the figure artist is uninteresting and will result in an empty piece of paper (that's why we didn't see anything above).
By far the most useful artist in matplotlib is the "Ax**e**s" artist. The Axes artist represents the "data space" of a typical plot, a rectangular axes (the most common, but not always the case, e.g. polar plots) will have 2 (confusingly named) Ax**i**s artists with tick labels and tick marks.
There is no limit on the number of Axes artists which can exist on a Figure artist. Let's go ahead and create a figure with a single Axes artist, and show it using pyplot:
```
ax = plt.axes()
```
Matplotlib's ``pyplot`` module makes the process of creating graphics easier by allowing us to skip some of the tedious Artist construction. For example, we did not need to manually create the Figure artist with ``plt.figure`` because it was implicit that we needed a figure when we created the Axes artist.
Under the hood matplotlib still had to create a Figure artist, its just we didn't need to capture it into a variable. We can access the created object with the "state" functions found in pyplot called **``gcf``** and **``gca``**.
## - essential stuff - `pyplot` versus Object based
Some example data:
```
x = np.linspace(0, 5, 10)
y = x ** 2
```
Observe the following difference:
**1. pyplot style: plt...** (you will see this a lot for code online!)
```
plt.plot(x, y, '-')
```
**2. creating objects**
```
fig, ax = plt.subplots()
ax.plot(x, y, '-')
```
Although a little bit more code is involved, the advantage is that we now have **full control** of where the plot axes are placed, and we can easily add more than one axis to the figure:
```
fig, ax1 = plt.subplots()
ax.plot(x, y, '-')
ax2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
ax1.plot(x, y, '-')
ax1.set_ylabel('y')
ax2.set_xlabel('x')
ax2.plot(x, y*2, 'r-')
```
<div class="alert alert-info" style="font-size:18px">
<b>REMEMBER</b>:
<ul>
<li>Use the **object oriented** power of Matplotlib!</li>
<li>Get yourself used to writing `fig, ax = plt.subplots()`</li>
</ul>
</div>
```
fig, ax = plt.subplots()
ax.plot(x, y, '-')
# ...
```
## An small cheat-sheet reference for some common elements
```
x = np.linspace(-1, 0, 100)
fig, ax = plt.subplots(figsize=(10, 7))
# Adjust the created axes so that its topmost extent is 0.8 of the figure.
fig.subplots_adjust(top=0.9)
ax.plot(x, x**2, color='0.4', label='power 2')
ax.plot(x, x**3, color='0.8', linestyle='--', label='power 3')
ax.vlines(x=-0.75, ymin=0., ymax=0.8, color='0.4', linestyle='-.')
ax.axhline(y=0.1, color='0.4', linestyle='-.')
ax.fill_between(x=[-1, 1.1], y1=[0.65], y2=[0.75], color='0.85')
fig.suptitle('Figure title', fontsize=18,
fontweight='bold')
ax.set_title('Axes title', fontsize=16)
ax.set_xlabel('The X axis')
ax.set_ylabel('The Y axis $y=f(x)$', fontsize=16)
ax.set_xlim(-1.0, 1.1)
ax.set_ylim(-0.1, 1.)
ax.text(0.5, 0.2, 'Text centered at (0.5, 0.2)\nin data coordinates.',
horizontalalignment='center', fontsize=14)
ax.text(0.5, 0.5, 'Text centered at (0.5, 0.5)\nin Figure coordinates.',
horizontalalignment='center', fontsize=14,
transform=ax.transAxes, color='grey')
ax.legend(loc='upper right', frameon=True, ncol=2)
```
For more information on legend positioning, check [this post](http://stackoverflow.com/questions/4700614/how-to-put-the-legend-out-of-the-plot) on stackoverflow!
## I do not like the style...
**...understandable**
Matplotlib had a bad reputation in terms of its default styling as figures created with earlier versions of Matplotlib were very Matlab-lookalike and mostly not really catchy.
Since Matplotlib 2.0, this has changed: https://matplotlib.org/users/dflt_style_changes.html!
However...
> *Des goûts et des couleurs, on ne discute pas...*
(check [this link](https://fr.wiktionary.org/wiki/des_go%C3%BBts_et_des_couleurs,_on_ne_discute_pas) if you're not french-speaking)
To account different tastes, Matplotlib provides a number of styles that can be used to quickly change a number of settings:
```
plt.style.available
x = np.linspace(0, 10)
with plt.style.context('default'): # 'seaborn', ggplot', 'bmh', 'grayscale', 'seaborn-whitegrid', 'seaborn-muted'
fig, ax = plt.subplots()
ax.plot(x, np.sin(x) + x + np.random.randn(50))
ax.plot(x, np.sin(x) + 0.5 * x + np.random.randn(50))
ax.plot(x, np.sin(x) + 2 * x + np.random.randn(50))
```
We should not start discussing about colors and styles, just pick **your favorite style**!
```
plt.style.use('seaborn-whitegrid')
```
<div class="alert alert-info">
<b>REMEMBER</b>:
<ul>
<li>If you just want **quickly a good-looking plot**, use one of the available styles (`plt.style.use('...')`)</li>
<li>Otherwise, the object-oriented way of working makes it possible to change everything!</li>
</ul>
</div>
## Interaction with Pandas
What we have been doing while plotting with Pandas:
```
import pandas as pd
flowdata = pd.read_csv('../data/vmm_flowdata.csv',
index_col='Time',
parse_dates=True)
flowdata.plot()
```
### Pandas versus matplotlib
#### Comparison 1: single plot
```
flowdata.plot(figsize=(16, 6)) # shift tab this!
```
Making this with matplotlib...
```
fig, ax = plt.subplots(figsize=(16, 6))
ax.plot(flowdata)
ax.legend(["L06_347", "LS06_347", "LS06_348"])
```
is still ok!
#### Comparison 2: with subplots
```
axs = flowdata.plot(subplots=True, sharex=True,
figsize=(16, 8), colormap='viridis', # Dark2
fontsize=15, rot=0)
```
Mimicking this in matplotlib (just as a reference):
```
from matplotlib import cm
import matplotlib.dates as mdates
colors = [cm.viridis(x) for x in np.linspace(0.0, 1.0, len(flowdata.columns))] # list comprehension to set up the colors
fig, axs = plt.subplots(3, 1, figsize=(16, 8))
for ax, col, station in zip(axs, colors, flowdata.columns):
ax.plot(flowdata.index, flowdata[station], label=station, color=col)
ax.legend()
if not ax.is_last_row():
ax.xaxis.set_ticklabels([])
ax.xaxis.set_major_locator(mdates.YearLocator())
else:
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.set_xlabel('Time')
ax.tick_params(labelsize=15)
```
Is already a bit harder ;-)
### Best of both worlds...
```
fig, ax = plt.subplots() #prepare a matplotlib figure
flowdata.plot(ax=ax) # use pandas for the plotting
# Provide further adaptations with matplotlib:
ax.set_xlabel("")
ax.tick_params(labelsize=15, pad=8)
fig.suptitle('Flow station time series', fontsize=15)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(16, 6)) #provide with matplotlib 2 axis
flowdata[["L06_347", "LS06_347"]].plot(ax=ax1) # plot the two timeseries of the same location on the first plot
flowdata["LS06_348"].plot(ax=ax2) # plot the other station on the second plot
# further adapt with matplotlib
ax1.set_ylabel("L06_347")
ax2.set_ylabel("LS06_348")
ax2.legend()
```
<div class="alert alert-info">
<b>Remember</b>:
<ul>
<li>You can do anything with matplotlib, but at a cost... [stackoverflow!!](http://stackoverflow.com/questions/tagged/matplotlib)</li>
<li>The preformatting of Pandas provides mostly enough flexibility for quick analysis and draft reporting. It is not for paper-proof figures or customization</li>
</ul>
<br>
If you take the time to make your perfect/spot-on/greatest-ever matplotlib-figure: Make it a **reusable function**!
</div>
An example of such a reusable function to plot data:
```
%%file plotter.py
#this writes a file in your directory, check it(!)
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from matplotlib import cm
from matplotlib.ticker import MaxNLocator
def vmm_station_plotter(flowdata, label="flow (m$^3$s$^{-1}$)"):
colors = [cm.viridis(x) for x in np.linspace(0.0, 1.0, len(flowdata.columns))] # list comprehension to set up the color sequence
fig, axs = plt.subplots(3, 1, figsize=(16, 8))
for ax, col, station in zip(axs, colors, flowdata.columns):
ax.plot(flowdata.index, flowdata[station], label=station, color=col) # this plots the data itself
ax.legend(fontsize=15)
ax.set_ylabel(label, size=15)
ax.yaxis.set_major_locator(MaxNLocator(4)) # smaller set of y-ticks for clarity
if not ax.is_last_row(): # hide the xticklabels from the none-lower row x-axis
ax.xaxis.set_ticklabels([])
ax.xaxis.set_major_locator(mdates.YearLocator())
else: # yearly xticklabels from the lower x-axis in the subplots
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.tick_params(axis='both', labelsize=15, pad=8) # enlarge the ticklabels and increase distance to axis (otherwise overlap)
return fig, axs
from plotter import vmm_station_plotter
# fig, axs = vmm_station_plotter(flowdata)
fig, axs = vmm_station_plotter(flowdata,
label="NO$_3$ (mg/l)")
fig.suptitle('Ammonium concentrations in the Maarkebeek', fontsize='17')
fig.savefig('ammonium_concentration.pdf')
```
<div class="alert alert-danger">
<b>NOTE</b>:
<ul>
<li>Let your hard work pay off, write your own custom functions!</li>
</ul>
</div>
<div class="alert alert-info" style="font-size:18px">
<b>Remember</b>:
`fig.savefig()` to save your Figure object!
</div>
# Seaborn
```
import seaborn as sns
```
* Built on top of Matplotlib, but providing
1. High level functions
2. Much cleaner default figures
* Works well with Pandas
## First example: `pairplot`
A scatterplot comparing the three stations with a color variation on the months:
```
flowdata["month"] = flowdata.index.month
sns.pairplot(flowdata["2009"].dropna(),
vars=["L06_347", "LS06_347", "LS06_348"],
diag_kind='kde', hue="month")
```
## Seaborn works well with Pandas & is built on top of Matplotlib
We will use the Titanic example again:
```
titanic = pd.read_csv('../data/titanic.csv')
titanic.head()
```
**Histogram**: Getting the univariaite distribution of the `Age`
```
fig, ax = plt.subplots()
sns.distplot(titanic["Age"].dropna(), ax=ax) # Seaborn does not like Nan values...
sns.rugplot(titanic["Age"].dropna(), color="g", ax=ax) # rugplot provides lines at the individual data point locations
ax.set_ylabel("Frequency")
```
<div class="alert alert-info">
<b>Remember</b>:
Similar to Pandas handling above, we can set up a `figure` and `axes` and add the seaborn output to it; adapt it afterwards
</div>
Compare two variables (**scatter-plot**):
```
g = sns.jointplot(x="Fare", y="Age",
data=titanic,
kind="scatter") #kde, hex
g = sns.jointplot(x="Fare", y="Age",
data=titanic,
kind="scatter") #kde, hex
# Adapt the properties with matplotlib by changing the available axes objects
g.ax_marg_x.set_ylabel("Frequency")
g.ax_joint.set_axis_bgcolor('0.1')
g.ax_marg_y.set_xlabel("Frequency")
```
<div class="alert alert-info">
<b>Remember</b>:
Adapting the output of a Seaborn `grid` of different axes can be done as well to adapt it with matplotlib
</div>
Who likes **regressions**?
```
fig, ax = plt.subplots()
sns.regplot(x="Fare", y="Age", data=titanic, ax=ax, lowess=False)
# adding the small lines to indicate individual data points
sns.rugplot(titanic["Fare"].dropna(), axis='x',
color="#6699cc", height=0.02, ax=ax)
sns.rugplot(titanic["Age"].dropna(), axis='y',
color="#6699cc", height=0.02, ax=ax)
```
# Need more matplotlib/seaborn inspiration?
For more in-depth material:
* http://www.labri.fr/perso/nrougier/teaching/matplotlib/
* notebooks in matplotlib section: http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/Index.ipynb#4.-Visualization-with-Matplotlib
* main reference: [matplotlib homepage](http://matplotlib.org/)
<div class="alert alert-info" style="font-size:18px">
<b>Remember</b>(!)
<ul>
<li>[matplotlib Gallery](http://matplotlib.org/gallery.html)</li>
<li>[seaborn gallery ](http://seaborn.pydata.org/examples/index.html)</li>
</ul>
<br>
Important resources to start from!
</div>
---
# Acknowledgement
> This notebook is partly based on material of the Met Office (Copyright (C) 2013 SciTools, GPL licensed): https://github.com/SciTools/courses
| github_jupyter |
<CENTER>
<img src="https://secure.meetupstatic.com/photos/theme_head/d/9/1/7/full_7435575.jpeg" width="100%">
<header>
<h1>Python Data Science</h1>
<h3>18 Octubre 2017</h3>
<h2>@victormartin</h2>
<p></p>
</header>
</CENTER>
```
%matplotlib inline
import pandas as pd
import numpy as np
import os
from datetime import datetime
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
pylab.rcParams['figure.figsize'] = (16, 6)
import seaborn as sns
sns.set_context("notebook", font_scale=1.4)
sns.set_style("whitegrid")
import warnings
warnings.filterwarnings('ignore')
```
# Contingut d'aquesta xerrada
- Data Science i Machine Learning
- Per què Python?
- Kaggle.com - Plataforma de competicions de data science
- Titanic - Getting Started competition
Per veure aquest notebook en format de presentació (Reveal.js):
```
$jupyter nbconvert datascience_introduction.ipynb --to slides --post serve
```
# Què vol dir *data science*?
> *Anàlisis de dades aplicant mètodes científics per tal d'extraure coneixement*
>
> [Definició de Wikipedia](https://en.wikipedia.org/wiki/Data_science])
Del reconeixement de patrons ha evolucionat cap al *machine learning* -> algorismes d'aprenentatge
# Què vol dir *machine learning*?
> *Camp d'estudi que dona a les computadores la habilitat d'aprendre sense haber estat explícitament programades*
>
> Arthur Samuel - 1959
# Diferents tipus de algoritmes de aprenentatge
- **[Supervised learning](https://en.wikipedia.org/wiki/Supervised_learning)**
- [Unsupervised learning](https://en.wikipedia.org/wiki/Unsupervised_learning)
- [Reinforcement learning](https://en.wikipedia.org/wiki/Reinforcement_learning)
- [Recommender systems](https://en.wikipedia.org/wiki/Recommender_system)
- (...)
## <font color="FireBrick ">Mega-links:</font>
- Curs [Introduction to Machine Learning](https://www.coursera.org/learn/machine-learning) de Andrew Ng a Coursera - Aquí **[un bon resum/apunts](http://www.holehouse.org/mlclass/)** pels més mandrosos ;)
# Aprenentatge supervisat
- Proveïr a un algorisme d'aprenentatge (**learning algorithm**) dades d'entrenament de les quals pugui aprendre.
- El **training data** està format per una sèrie d'atributs (**features**), i una columna amb la resposta correcte, la qual anomenem **target** (o **label**)
- Durant la fase de **training** el *learning algorithm* troba patrons sobre el *training data* que "mapejen" els *features* al *target* (resposta que volem predir). Retorna un **model** que captura aquests patrons.
- Per últim podem fer servir aquest *model* per fer prediccions amb dades que no sabem el *target*.
### Exemple de ML: Predicció de preu de venda de pisos
```
# TRAINING SET
dades = {
u'Superfície (m2)': [50,70,100,130,200],
u'Terrassa (m2)': [0,0,4,8,40],
u'Num banys': [1,1,2,2,4],
u'Plaçes Parking': [0,0,1,0,2],
u'Barri': ["Font de la Pólvora","Taialà","Eixample","Barri Vell","Palau"],
u'Preu (€)': [50000,75000,190000,280000,350000],
}
pd.DataFrame(dades).reindex(columns=[u"Barri", u"Superfície (m2)",u"Num banys",
u"Terrassa (m2)",u"Plaçes Parking",u"Preu (€)"])
```
** <small>Bon moment per recuperar els slides <a href="https://github.com/victormartingarcia/2017-pyGrn-intropandas" target="_blank">"Intro to Pandas"</a> :)</small>
```
# PREDICCIONS
dades = {
u'Superfície (m2)': [90,120],
u'Terrassa (m2)': [0,6],
u'Num banys': [1,2],
u'Plaçes Parking': [0,1],
u'Barri': ["Taialà","Palau"],
u'Preu (€)': ["???","???"],
}
pd.DataFrame(dades).reindex(columns=[u"Barri", u"Superfície (m2)",u"Num banys",
u"Terrassa (m2)",u"Plaçes Parking",u"Preu (€)"])
```
## Exemple visual ML
```
dades = {
u'Superfície (m2)': [50,70,100,130,200],
u'Preu (€)': [50000,55000,190000,220000,300000],
}
df_training = pd.DataFrame(dades)
df_training.reindex(columns= [u"Superfície (m2)",u"Preu (€)"])
# Dibuixem les dades d'entrenament
sns.lmplot(x=u'Superfície (m2)', y=u'Preu (€)',
data=df_training, # Data source
fit_reg=False, # Don't fix a regression line
size=6, aspect=2,
scatter_kws={"s": 250} ).set(xticks=np.arange(50,201,10));
```
**Quant puc demanar per un pis de 90m2?**
```
# Dibuixem les dades d'entrenament -> REGRESSIÓ LINEAL
sns.lmplot(x=u'Superfície (m2)', y=u'Preu (€)',
data=df_training, # Data source
fit_reg=True, # Fix a regression line
ci=0, size=6, aspect=2, scatter_kws={"s": 250}
).set(xticks=np.arange(50,201,10));
```
**Segons aquest model, per un pis de 90m2 puc demanar 130 000€**
```
# Dibuixem les dades d'entrenament -> Un altre algorisme predictiu
sns.lmplot(x=u'Superfície (m2)', y=u'Preu (€)',
data=df_training, # Data source
lowess=True, # Fix a logistic line
ci=0, size=6, aspect=2,
scatter_kws={"s": 250}).set(xticks=np.arange(50,201,10));
```
**WTF!!** Segons aquest altre model, per un pis de 90m2 puc demanar 150 000€ ( 20k € més!). Quin és el bò?
# Objectiu: Crear un model que *generalitzi*
Per *generalització* entenem que el model s'ajusti a TOTES les dades Out-sample, no només a les dades in-sample del training set.
<center>
<img src="https://www.netquest.com/hubfs/Imported_Blog_Media/data.png" width="60%"/>
<p align="center">Funció sin(2πx)</p>
</center>
# Underfitting i Overfitting
<center>
<img src="https://www.netquest.com/hubfs/Imported_Blog_Media/4modelos1.png" width="100%"/>
<p align="center">M = Ordre del polinomi</p>
</center>
# "La prueba del algodón": Cross-Validation
Per saber si un model generalitza, s'ha de probar amb dades noves (out-of-sample).
**Tècnica *Cross-Validation* **:
1. Separar el dataset en 2 conjunts (p.ex. 70%/30% de les dades)
1. Entrenarem el nostre model amb el primer conjunt (**training set**)
1. Aplicarem el model al segon conjunt (**validation set**)
1. Mesurarem el *performance* del nostre model amb qualsevol [métrica d'evaluació](http://scikit-learn.org/stable/modules/model_evaluation.html)
<center>
<img src="https://www.netquest.com/hubfs/Imported_Blog_Media/errorvsm.png" width="70%"/>
</center>
# Algoritmes d'aprenentatge
<center>
<img src="https://udarajay.com/content/images/2017/10/machinelearningalgorithms.png" width="100%"/>
</center>
# Random Forest
> When in doubt, use [Random Forest](https://en.wikipedia.org/wiki/Random_forest) — the go-to machine learning algorithm that is considered to be one of the most effective and versatile in solving almost any prediction task
>
> Rebecca Merrett, TechWorld Australia
## <font color="FireBrick ">Mega-links sobre introducció als algorismes de ML:</font>
- [Visualització/Exemple de arbres de decisió](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/)
- [Essentials of Machine Learning algorithms](https://www.analyticsvidhya.com/blog/2017/09/common-machine-learning-algorithms/)
# Worflow per abordar un problema de machine learning
<center>
<img src="https://udarajay.com/content/images/2017/10/17-038-ml-concept_0.jpg" width="90%"/>
</center>
## <font color="FireBrick ">Mega-links sobre aplicació pràctica de ML:</font>
- [Advice for applying Machine Learning](http://www.holehouse.org/mlclass/10_Advice_for_applying_machine_learning.html): Apunts del curs Introduction de Machine Learning de Coursera
- [Machine Learning Systems Design](http://www.holehouse.org/mlclass/11_Machine_Learning_System_Design.html): Apunts del curs Introduction de Machine Learning de Coursera
- [Best Practices for ML Engineering](http://martin.zinkevich.org/rules_of_ml/rules_of_ml.pdf): Escrit per un enginyer de Google (PDF)
# Per què Python?
* Históricament s'ha utilitzat llenguatges i eines com R, Matlab/Octave, Mathematica ...
La comunitat Python ha sapigut agafar el bò i millor d'aquestes eines:
* Anàlisis numèric Matlab -> **numpy**
* R dataframes -> **pandas**
* Matlab plotting -> **matplotlib**
* Mathematica notebooks -> **jupyter notebooks**
* R caret -> **scikit-learn**
* (...)
# [Kaggle.com](www.kaggle.com)
Competicions de datascience amb regles senzilles:
- Cada competició té 2 datasets públics: **training** (labeled) i **test**
- Els participants han de pujar un fitxer .CSV amb els camps ID (del test) i TARGET (la seva predicció). Aquest upload se'n diu **submission**
- Sobre un percentatge d'aquestes prediccions, s'evaluen l'accuracy mitjançant la [mètrica de la competició](https://www.kaggle.com/wiki/Metrics), i el submission apareix en el **public leaderboard**
- Quan acaba el termini de la competició, s'evaluen tots els submissions amb la resta de dades del **test set**, i es crea un **private leaderboard** que determina els guanyadors finals.
# Kaggle Tips and Tricks
- Fòrum de les competicions
- [Blog de Kaggle](http://blog.kaggle.com/) amb entrevistes a guanyadors de competicions concloses
- **Ensembles** -> Teaming
# Competició Getting Started: Titanic
- El 15 d'Abril de 1912 es va enfonsar el Titanic debut a la col.lisió amb un iceberg.
- Van morir 1502 de les 2224 persones que viatjaven a bord (entre passatgers i tripulació)
- Una de les raons es que no hi havia els suficients bots salvavides
- Tot i el component de sort, hi van haver grups amb més possibilitats de sobreviure que d'altres (dones, nens, passatgers de primera classe...)
** Problema de classificació binària:** Per cada passatger, hem de determinar si es va salvar o no
<center>
<br />
<img src="http://www.pearlanddean.com/sites/default/files/titanic_0.gif"/>
</center>
```
# data analysis and wrangling
import pandas as pd
import numpy as np
import random as rnd
# visualization
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
%matplotlib inline
pylab.rcParams['figure.figsize'] = 16, 12
import matplotlib
matplotlib.style.use('ggplot')
from sklearn.model_selection import cross_val_score
```
# 1/5 - Carregar les dades
Info pública sobre el dataset: https://www.kaggle.com/c/titanic/data
```
# Descripció de les dades
from IPython.display import IFrame
IFrame('http://rstudio-pubs-static.s3.amazonaws.com/24969_894d890964fd4308ab537bfde1f784d2.html', width=700, height=350)
# Training set
df_train = pd.read_csv('data/train.csv')
# 3 passatgers a l'atzar
df_train.sample(3)
```
# 2/5 - Preparar les dades
```
# eliminar columnes innecesàries
df_train = df_train.drop(["PassengerId", "Cabin", "Ticket","Embarked","Name","Sex"], axis=1)
# emplenar NaN
df_train = df_train.fillna(df_train.mean())
df_train.sample(3)
```
# 3/5 - Feature engineering
Ho guardem per més tard ;)
# 4/5 Creació i validació del model
```
# Benchmark Linear Regression
from sklearn.linear_model import LinearRegression
clf_LR = LinearRegression()
scores_LR = cross_val_score(clf_LR, df_train.drop("Survived", axis=1), df_train["Survived"], cv=4)
print("Linear Regression Accuracy: %0.5f (+/- %0.5f)" % (scores_LR.mean(), scores_LR.std() * 2))
# Benchmark Random Forest
from sklearn.ensemble import RandomForestClassifier
clf_RF = RandomForestClassifier(n_estimators=100)
scores_RF = cross_val_score(clf_RF, df_train.drop("Survived", axis=1), df_train["Survived"], cv=4)
print("Random Forest Accuracy: %0.5f (+/- %0.5f)" % (scores_RF.mean(), scores_RF.std() * 2))
# Entrenem el model amb totes les dades:
clf_RF.fit(df_train.drop("Survived", axis=1), df_train["Survived"]);
```
# 5/5 Submission
```
# LLegim el test dataset de disc
df_test = pd.read_csv('data/test.csv')
# eliminar columnes innecesàries
df_test = df_test.drop(["PassengerId", "Cabin", "Ticket","Embarked","Name","Sex"], axis=1)
# emplenar NaN
df_test = df_test.fillna(df_test.mean())
# Apliquem el model Random Forest que hem entrenat prèviament
output = clf_RF.predict(df_test)
df_output = pd.DataFrame()
df_output['PassengerId'] = pd.read_csv('data/test.csv')['PassengerId']
df_output['Survived'] = output
## Aquest serà el fitxer CSV del submission:
#df_output[['PassengerId','Survived']].to_csv('data/submission.csv',index=False)
df_output[['PassengerId','Survived']].sample(3)
```
# Feature Engineering: La mare dels ous
> Procés de transformar les dades "crues" (*raw data*) en atributs distintius (*features*) fent servir coneixement del domini (experiència), per tal de representar millor el problema de fons i amb l'objectiu de millorar l'*accuracy* de les prediccions en dades noves (*out of sample*)
**Exemple**: *"Les dones i els nens primer"*
```
df_train = pd.read_csv('data/train.csv')
# Percentatge homes/dones que es van salvar:
sns.factorplot('Sex', 'Survived', data = df_train, ci=None, kind = 'bar', size=6, aspect=2);
# Persones salvades segons edat
sns.set_style("whitegrid")
sns.factorplot('Age', 'Survived', data = df_train, hue_order = [1,0], orient="h", kind="violin", size=6, aspect=2);
# eliminar columnes innecesàries
df_train = df_train.drop(["PassengerId", "Cabin", "Ticket","Embarked","Name"], axis=1)
# emplenar NaN
df_train = df_train.fillna(df_train.mean())
# Creem un nou feature anomenat "AgeCategory"
df_train["AgeCategory"] = "Adult"
df_train.loc[df_train["Age"]<16, "AgeCategory"]="Child"
df_train.loc[df_train["Age"]>=60, "AgeCategory"]="Old"
df_train.sample(3)
# Transformem les features categòriques (Sex i AgeCategory) en booleanes!
df_train = pd.get_dummies(df_train, columns=["Sex", "AgeCategory"]).drop(["Age"], axis=1)
df_train.sample(3)
```
## Com millora l'algorisme ML amb aquestes noves columnes?
```
clf_RF = RandomForestClassifier(n_estimators=100)
scores_RF = cross_val_score(clf_RF, df_train.drop("Survived", axis=1), df_train["Survived"], cv=4)
print ("[Abans] Random Forest Accuracy: 0.68244 (+/- 0.04123)")
print ("[ ARA ] Random Forest Accuracy: %0.5f (+/- %0.5f)" % (scores_RF.mean(), scores_RF.std() * 2))
```
# Som capaços de millorar aquest 81% de accuracy entre tots?
```
from IPython.display import IFrame
IFrame('http://rstudio-pubs-static.s3.amazonaws.com/24969_894d890964fd4308ab537bfde1f784d2.html', width=700, height=350)
pd.read_csv('data/train.csv').sample(5)
# Idees Feature Engineering 1/3 : Categoritzar persones segons el títol de la columna Name (Mrs, Miss, Master, Officer, Royalty)
dictTitles = {
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir": "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess": "Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr": "Mr",
"Mrs": "Mrs",
"Miss": "Miss",
"Master": "Master",
"Lady": "Royalty"
}
X['Embarked'] = X['Embarked'].fillna('S')
X['title'] = X['Name'].apply(lambda x: x.split(',')[1].split('.')[0].strip())
X['title'] = X['title'].map(dictTitles)
X = X.drop(['Name'], axis=1)
X = pd.get_dummies(X, columns=['Embarked', 'title'])
# Idees Feature Engineering 2/3 : Crear feature amb el nombre de familiars que viatgen
X['family_size']= (X["SibSp"]+X["Parch"])+1
# Idees Feature Engineering 3/3 : Contar el número de persones amb el mateix "Ticket"
pd.read_csv('data/train.csv').groupby("Ticket")["Survived"].count().sort_values()
# CROSS-VALIDATION SAMPLE
df_train = pd.read_csv('data/train.csv')
# eliminar columnes innecesàries
df_train = df_train.drop(["PassengerId", "Cabin", "Ticket","Embarked","Name"], axis=1)
# emplenar NaN
df_train = df_train.fillna(df_train.mean())
# Creem un nou feature anomenat "AgeCategory"
df_train["AgeCategory"] = "Adult"
df_train.loc[df_train["Age"]<16, "AgeCategory"]="Child"
df_train.loc[df_train["Age"]>=60, "AgeCategory"]="Old"
# Transformem les features categòriques (Sex i AgeCategory) en booleanes!
df_train = pd.get_dummies(df_train, columns=["Sex", "AgeCategory"]).drop(["Age"], axis=1)
#########################################################
## PROBAR NOUS FEATURES AQUÍ!
#########################################################
#########################################################
#########################################################
clf_RF = RandomForestClassifier(n_estimators=100)
scores_RF = cross_val_score(clf_RF, df_train.drop("Survived", axis=1), df_train["Survived"], cv=4)
print ("Random Forest Accuracy: %0.5f (+/- %0.5f)" % (scores_RF.mean(), scores_RF.std() * 2))
```
## Gràcies per escoltar! Preguntes?
- https://github.com/victormartingarcia
- <a href="mailto:victor.martin.garcia@gmail.com">victor.martin.garcia@gmail.com</a>
- [@victormartin](https://twitter.com/victormartin)
Slides and data: Source: https://github.com/victormartingarcia/2017-pyGrn-introdatascience
Slides presented with 'live reveal' https://github.com/damianavila/RISE
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Visualization/nwi_wetlands_symbology.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/nwi_wetlands_symbology.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Visualization/nwi_wetlands_symbology.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/nwi_wetlands_symbology.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell. Uncomment these lines if you are running this notebook for the first time.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for the first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
# NWI legend: https://www.fws.gov/wetlands/Data/Mapper-Wetlands-Legend.html
def nwi_add_color(fc):
emergent = ee.FeatureCollection(
fc.filter(ee.Filter.eq('WETLAND_TY', 'Freshwater Emergent Wetland')))
emergent = emergent.map(lambda f: f.set(
'R', 127).set('G', 195).set('B', 28))
# print(emergent.first())
forested = fc.filter(ee.Filter.eq(
'WETLAND_TY', 'Freshwater Forested/Shrub Wetland'))
forested = forested.map(lambda f: f.set('R', 0).set('G', 136).set('B', 55))
pond = fc.filter(ee.Filter.eq('WETLAND_TY', 'Freshwater Pond'))
pond = pond.map(lambda f: f.set('R', 104).set('G', 140).set('B', 192))
lake = fc.filter(ee.Filter.eq('WETLAND_TY', 'Lake'))
lake = lake.map(lambda f: f.set('R', 19).set('G', 0).set('B', 124))
riverine = fc.filter(ee.Filter.eq('WETLAND_TY', 'Riverine'))
riverine = riverine.map(lambda f: f.set(
'R', 1).set('G', 144).set('B', 191))
fc = ee.FeatureCollection(emergent.merge(
forested).merge(pond).merge(lake).merge(riverine))
base = ee.Image(0).mask(0).toInt8()
img = base.paint(fc, 'R') \
.addBands(base.paint(fc, 'G')
.addBands(base.paint(fc, 'B')))
return img
fromFT = ee.FeatureCollection("users/wqs/Pipestem/Pipestem_HUC10")
Map.addLayer(ee.Image().paint(fromFT, 0, 2), {}, 'Watershed')
huc8_id = '10160002'
nwi_asset_path = 'users/wqs/NWI-HU8/HU8_' + huc8_id + '_Wetlands' # NWI wetlands for the clicked watershed
clicked_nwi_huc = ee.FeatureCollection(nwi_asset_path)
nwi_color = nwi_add_color(clicked_nwi_huc)
Map.centerObject(clicked_nwi_huc, 10)
Map.addLayer(nwi_color, {'gamma': 0.3, 'opacity': 0.7}, 'NWI Wetlands Color')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
# Lesson 8 - FastAI
## Collaborative Filtering Deep Dive
Collaborative filtering is a technique used by recommender systems. We will be taking a look at a movie reccomendation model.
```
#hide
!pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
#hide
from fastbook import *
```
## A First Look at the Data
```
from fastai.collab import *
from fastai.tabular.all import *
path = untar_data(URLs.ML_100k)
#lets grab the rating table
ratings = pd.read_csv(path/'u.data', delimiter='\t', header=None,
names=['user','movie','rating','timestamp'])
ratings.head()
```
### Lets simulate
Below we are simulating the reccomendation model. Here we assume know what the Latent Factors, but in reality we do not and need to determine them.
```
# Lets assume these are the values for a movie:
#Sci-fi, action, old
last_skywalker = np.array([0.98,0.9,-0.9])
#Lets assume there is a user who likes these categories
user1 = np.array([0.9,0.8,-0.6])
(user1*last_skywalker).sum()
```
> Pos val means that the user probably will like it
```
#another movie
casablanca = np.array([-0.99,-0.3,0.8])
(user1*casablanca).sum()
```
> Neg val means that the user might not like it
```
#Lets grab the movie table
movies = pd.read_csv(path/'u.item', delimiter='|', encoding='latin-1',
usecols=(0,1), names=('movie','title'), header=None)
movies.head()
```
### Lets merge our two tables
```
#we can merge the movie tabel w/ the rating tabel
ratings = ratings.merge(movies)
ratings.head()
```
## Creating dataloader
```
dls = CollabDataLoaders.from_df(ratings, user_name = 'user', item_name='title', bs=64) #must pass the correct columns
dls.show_batch()
dls.classes #We have the user and title class
len(dls.classes['user']),
```
## Initialize Parameters
```
n_users = len(dls.classes['user'])
n_movies = len(dls.classes['title'])
n_factors = 5 #Number of latent factors
user_factors = torch.randn(n_users, n_factors)
movie_factors = torch.randn(n_movies, n_factors)
```
## Sidebar: Indexing
It turns out we can represent looking up
an index as a matrix. See below
```
one_hot_3 = one_hot(3, n_users).float()
one_hot_3[:10]
user_factors[3] #parameter (latent factors) values at this index are
user_factors.t() @ one_hot_3
```
> Notice same values
## End Sidebar
## Collaborative Filtering from Scratch
Lets put what we did above into a class. This class will initialize the parameters for us and forward pass as well.
```
class DotProduct(Module): #extends Module class
def __init__(self, n_users, n_movies, n_factors):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
def forward(self, x): #Method called auto. anytime using Module class
users = self.user_factors(x[:,0]) #user ID's
movies = self.movie_factors(x[:,1]) #movie ID's
return (users * movies).sum(dim=1) #dim=0 is the minibatches, we want to sum over the other dim (1)
x,y = dls.one_batch()
x.shape
x[:3] #user ID, movie ID
y[:3] #ratings
```
## Training
```
model = DotProduct(n_users, n_movies, 50) #our model
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3)
```
> Not bad, but we can do better!
## Improving the model
We can improve our model by giving it a range as its a regression model. Passing it the range of (0-5.5) should improve the performance.
```
class DotProduct(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0,5.5)):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
self.y_range = y_range
def forward(self, x):
users = self.user_factors(x[:,0])
movies = self.movie_factors(x[:,1])
return sigmoid_range((users * movies).sum(dim=1), *self.y_range)
#now lets train again
model = DotProduct(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3)
```
> Didn't really improve, but thats ok
## Further improving the model
We should also add a bias as some rating may be skewing the data.
```
class DotProductBias(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0,5.5)):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
self.user_bias = Embedding(n_users, 1)
self.movie_bias = Embedding(n_movies, 1)
self.y_range = y_range
def forward(self, x):
users = self.user_factors(x[:,0])
movies = self.movie_factors(x[:,1])
res = (users * movies).sum(dim=1, keepdim=True)
#bias
res += self.user_bias(x[:,0]) + self.movie_bias(x[:,1])
return sigmoid_range(res, *self.y_range)
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3)
```
## Loss not improving
It seems like our loss is not improving regardless of the improvements: But if you take another look you should realise that it performs better during the earlier epochs (2 or 3). This means that we are overfitting the model. But how can we train it for more epochs without overfitting? This is where weight regularization comes in.
## Weight Decay
Weight decay is a regularization technique that penalizes the model for large weights. Overall, it prevents the model from overfitting during training.
```
x = np.linspace(-2,2,100)
a_s = [1,2,5,10,50]
ys = [a * x**2 for a in a_s]
_,ax = plt.subplots(figsize=(8,6))
for a,y in zip(a_s,ys): ax.plot(x,y, label=f'a={a}')
ax.set_ylim([0,5])
ax.legend();
```
> Graphic above demonstrates weight decay
## Train with weight decay
```
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.1) #Pass wd
```
> Nice our loss dropped down to .82! Also, notice that train_loss **increased**, this is because wd is preventing the model from overfitting
## Using FastAI ToolKit
Can also achieve same results using FastAI ToolKit. Notice we changed from **Learner** to **collab_learner**.
```
learn = collab_learner(dls, n_factors=50, y_range=(0, 5.5))
learn.fit_one_cycle(5, 5e-3, wd=0.1)
#We can also look inside model
movie_bias = learn.model.i_bias.weight.squeeze()
idxs = movie_bias.argsort(descending=True)[:5]
[dls.classes['title'][i] for i in idxs]
```
## Sidebar: Creating Our Own Embedding Module
So far we have been using the predefined Embedding, why don't we create our own?
```
class T(Module):
def __init__(self): self.a = torch.ones(3)
L(T().parameters())# calling method from Module class
class T(Module):
def __init__(self): self.a = nn.Parameter(torch.ones(3)) #Must wrap it with nn.Parameter()
L(T().parameters())
class T(Module):
def __init__(self): self.a = nn.Linear(1, 3, bias=False)
t = T()
L(t.parameters())
def create_params(size):
return nn.Parameter(torch.zeros(*size).normal_(0, 0.01))
```
> This is all we need to create our own embedding
```
class DotProductBias(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0,5.5)):
self.user_factors = create_params([n_users, n_factors])
self.user_bias = create_params([n_users])
self.movie_factors = create_params([n_movies, n_factors])
self.movie_bias = create_params([n_movies])
self.y_range = y_range
def forward(self, x):
users = self.user_factors[x[:,0]]
movies = self.movie_factors[x[:,1]]
res = (users*movies).sum(dim=1)
res += self.user_bias[x[:,0]] + self.movie_bias[x[:,1]]
return sigmoid_range(res, *self.y_range)
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.1)
```
> Notice similer performance
## End Sidebar
## Looking inside model
We can take a look inside our model by calling **learn.model**.
```
movie_bias = learn.model.movie_bias.squeeze() #Grab movie by bias
idxs = movie_bias.argsort()[:5] #Sort by least bias
[dls.classes['title'][i] for i in idxs]
```
> Least bias movies
```
idxs = movie_bias.argsort(descending=True)[:5] #Sort by most bias
[dls.classes['title'][i] for i in idxs]
```
> Most bias movies
```
g = ratings.groupby('title')['rating'].count()
top_movies = g.sort_values(ascending=False).index.values[:1000]
top_idxs = tensor([learn.dls.classes['title'].o2i[m] for m in top_movies])
movie_w = learn.model.movie_factors[top_idxs].cpu().detach()
movie_pca = movie_w.pca(3)
fac0,fac1,fac2 = movie_pca.t()
idxs = list(range(50))
X = fac0[idxs]
Y = fac2[idxs]
plt.figure(figsize=(12,12))
plt.scatter(X, Y)
for i, x, y in zip(top_movies[idxs], X, Y):
plt.text(x,y,i, color=np.random.rand(3)*0.7, fontsize=11)
plt.show()
```
> Notice that similer movies have been clumped togather
### Embedding Distance
We can also use simple math to find similer movies
```
movie_factors = learn.model.movie_factors
idx = dls.classes['title'].o2i['Forrest Gump (1994)']
distances = nn.CosineSimilarity(dim=1)(movie_factors, movie_factors[idx][None])
idx = distances.argsort(descending=True)[1]
dls.classes['title'][idx]
```
## Sidebar: Bootstrapping a Collaborative Filtering Model
Another approach to a Collaborative Filtering Model
```
embs = get_emb_sz(dls)
embs
class CollabNN(Module):
def __init__(self, user_sz, item_sz, y_range=(0,5.5), n_act=100):
self.user_factors = Embedding(*user_sz)
self.item_factors = Embedding(*item_sz)
self.layers = nn.Sequential(
nn.Linear(user_sz[1]+item_sz[1], n_act),
nn.ReLU(),
nn.Linear(n_act, 1))
self.y_range = y_range
def forward(self, x):
embs = self.user_factors(x[:,0]),self.item_factors(x[:,1])
x = self.layers(torch.cat(embs, dim=1))
return sigmoid_range(x, *self.y_range)
model = CollabNN(*embs)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.01)
```
### Using FastAI ToolKit
Can also achieve same results using FastAI ToolKit. Just enable **use_nn**
```
learn = collab_learner(dls, use_nn=True, y_range=(0, 5.5), layers=[100,50])
learn.fit_one_cycle(5, 5e-3, wd=0.1)
```
> Notice similer results
```
type(learn.model)
#This is what the class looks like
@delegates(TabularModel)
class EmbeddingNN(TabularModel):
def __init__(self, emb_szs, layers, **kwargs):
super().__init__(emb_szs, layers=layers, n_cont=0, out_sz=1, **kwargs)
```
## End sidebar
## Conclusion
Overall, I hope you learned how to create a reccomendation model. Some very important concepts were viewed at, such as latent factors and weight decay.
## Questionnaire
1. **What problem does collaborative filtering solve?**
Obtains latent factors needed to provided a good reccomendation.
1. **How does it solve it?**
It learns the latent factors via gradient descent and clumps up similer kind of factors.
1. **Why might a collaborative filtering predictive model fail to be a very useful recommendation system?**
If there is a lack of data from users to provide useful reccomendations.
1. **What does a crosstab representation of collaborative filtering data look like?**
Crosstab is where the colomn tabs are users, the row tabs are items, and values are filled out based on the user’s rating of the items.
1. **Write the code to create a crosstab representation of the MovieLens data (you might need to do some web searching!).**
1. **What is a latent factor? Why is it "latent"?**
Latent factos are the factors used to determine a prediction. They are latent as they are learned, NOT given to the model.
1. **What is a dot product? Calculate a dot product manually using pure Python with lists.**
The dot product is the sum of the products of the corresponding matrixs.
```python
a = [1,2,3]
b = [1,2,3]
sum(i[0]*i[1] for i in zip(a,b))
```
1. **What does `pandas.DataFrame.merge` do?**
Merges two DataFrame's togather
1. **What is an embedding matrix?**
It is what you multiply an embedding with.
1. **What is the relationship between an embedding and a matrix of one-hot-encoded vectors?**
The embedding is a matrix of a one-hot encoded vecotors, but is more computationally efficient.
1. **Why do we need `Embedding` if we could use one-hot-encoded vectors for the same thing?**
More computationally efficient and fastser.
1. **What does an embedding contain before we start training (assuming we're not using a pretained model)?**
It is randomly initialized.
1. **Create a class (without peeking, if possible!) and use it.**
```python
class Name:
def __init__def(self):
pass
def func_name(self):
pass
```
1. **What does `x[:,0]` return?**
User ids
1. **Rewrite the `DotProduct` class (without peeking, if possible!) and train a model with it.**
```python
class DotProduct(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0,5.5)):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
self.y_range = y_range
def forward(self, x):
users = self.user_factors(x[:,0])
movies = self.movie_factors(x[:,1])
return sigmoid_range((users * movies).sum(dim=1), *self.y_range)
```
1. **What is a good loss function to use for MovieLens? Why?**
Mean Squared Error Loss. Can use to compare how far the prediction is from the label.
1. **What would happen if we used cross-entropy loss with MovieLens? How would we need to change the model?**
We would need the model to output more predictions, only then can we pass it to the cross-entropy.
1. **What is the use of bias in a dot product model?**
Some rating may skeew the data, so a bias can help.
1. **What is another name for weight decay?**
L2 regularization
1. **Write the equation for weight decay (without peeking!).**
loss_with_wd = loss + wd * (parameters\**2).sum()
1. **Write the equation for the gradient of weight decay. Why does it help reduce weights?**
Add to the gradients 2*wd*parameters. Overall, this prevents overfitting by enforcing more evenly distributed weights.
1. **Why does reducing weights lead to better generalization?**
By enforcing more evenly distributed weights, the result is is more shallow, with less sharp surfaces.
1. **What does `argsort` do in PyTorch?**
Returns index values of the order of the original list.
1. **Does sorting the movie biases give the same result as averaging overall movie ratings by movie? Why/why not?**
No. It takes into account other factors which can influence the results.
1. **How do you print the names and details of the layers in a model?**
learn.model
1. **What is the "bootstrapping problem" in collaborative filtering?**
The model cannot make any recommendations without enough data
1. **How could you deal with the bootstrapping problem for new users? For new movies?**
Have them complete a questionair.
1. **How can feedback loops impact collaborative filtering systems?**
May cause the model to suffer from bias.
1. **When using a neural network in collaborative filtering, why can we have different numbers of factors for movies and users?**
Because we are not taking the dot product, and instead concatenating the embedding matrices, different number of factors is alright.
1. **Why is there an `nn.Sequential` in the `CollabNN` model?**
Can create nonlinearity as we can couple multiple layers together.
1. **What kind of model should we use if we want to add metadata about users and items, or information such as date and time, to a collaborative filtering model?**
Tabular model
### Further Research
1. **Take a look at all the differences between the `Embedding` version of `DotProductBias` and the `create_params` version, and try to understand why each of those changes is required. If you're not sure, try reverting each change to see what happens. (NB: even the type of brackets used in `forward` has changed!)**
1. **Find three other areas where collaborative filtering is being used, and find out what the pros and cons of this approach are in those areas.**
1. **Complete this notebook using the full MovieLens dataset, and compare your results to online benchmarks. See if you can improve your accuracy. Look on the book's website and the fast.ai forum for ideas. Note that there are more columns in the full dataset—see if you can use those too (the next chapter might give you ideas).**
1. **Create a model for MovieLens that works with cross-entropy loss, and compare it to the model in this chapter.**
Completed, see here https://usama280.github.io/PasteBlogs/
| github_jupyter |
## Setup
```
%matplotlib qt
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
from pathlib import Path
import os, sys
import random
sys.path.append( os.path.abspath('..') )
import utils
Path('Datasets').mkdir(exist_ok=True)
os.chdir('Datasets')
def plot_image_grid(imgs, labels, width, height, **kwargs):
fig = plt.figure()
for i in range(width * height):
ax = plt.subplot(height, width, i + 1)
plt.imshow(imgs[i], **kwargs)
plt.xlabel(labels[i])
plt.xticks([])
plt.yticks([])
plt.tight_layout()
plt.show()
return fig
```
## 1 MNIST
```
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
fig = plot_image_grid(x_train, y_train, width=8, height=5, cmap='gray_r', vmin=0, vmax=255)
fig.savefig('MNIST.pdf', bbox_inches='tight')
```
## 2 Fashion MNIST
```
(x_train, y_train), (x_test_, y_test) = tf.keras.datasets.fashion_mnist.load_data()
num_to_label = ['T-Shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
height = 5
width = 8
labels = [num_to_label[num] for num in y_train[0:width*height]]
fig = plot_image_grid(x_train, labels, width, height, cmap='gray_r', vmin=0, vmax=255)
fig.savefig('Fashion_MNIST.pdf', bbox_inches='tight')
```
## 3 CIFAR 10
```
(x_train, y_train), (x_test_, y_test) = tf.keras.datasets.cifar10.load_data()
num_to_label = ['Airplane', 'Automobile', 'Bird', 'Cat', 'Deer', 'Dog', 'Frog', 'Horse', 'Ship', 'Truck']
height = 5
width = 8
labels = [num_to_label[num] for num in y_train[0:width*height, 0]]
fig = plot_image_grid(x_train, labels, width, height, vmin=0, vmax=255)
fig.set_size_inches(8, 6)
fig.savefig('CIFAR10.pdf', bbox_inches='tight')
```
## 4 Flowers
This dataset must be downloaded separedly, you can find it in this link:
https://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html
After downloading, make sure the folder structure is as follows:
```
📂<PARENT>
┗ 📂flowers
┗ 📂imgs
┣ 📄image_00001.jpg
┣ 📄image_00002.jpg
┗ 📄 ...
```
### 4.1 Showing original dataset
```
flowers_parent_path = '<PATH TO FOLDER CONTAINING THE DATASET>'
flowers_dir = os.path.join(flowers_parent_path, 'flowers', 'imgs')
files = os.listdir(flowers_dir)
data = []
for f in files[1690:1690+12]:
img = tf.keras.preprocessing.image.load_img(os.path.join(flowers_dir, f))
data.append(tf.keras.preprocessing.image.img_to_array(img, dtype='uint8'))
fig = plt.figure()
for i in range(3 * 4):
ax = plt.subplot(3, 4, i + 1)
plt.imshow(data[i])
plt.xticks([])
plt.yticks([])
plt.tight_layout()
plt.show()
fig.savefig('Flowers.pdf', bbox_inches='tight')
```
### 4.2 Showing reduced dataset
Make sure to have created the reduced dataset first by running the code in the file `datasets_preprocess.ipynb`
```
data = np.load(os.path.join('..', '..', 'flowers.npy'))
SEED = 279923 # https://youtu.be/nWSFlqBOgl8?t=86 - I love this song
random.seed(SEED)
indexes = [*range(2, 2 + 60*30, 30)]
random.shuffle(indexes)
samples = data[indexes].astype('float32') / 255
grid = utils.fn.make_grid_image(samples, n_cols=10, border=1, pad=1)
fig = plt.figure()
plt.imshow(grid)
plt.axis(False)
plt.tight_layout(h_pad=0, w_pad=0)
fig.savefig('Flowers_reduced.pdf', bbox_inches='tight', pad_inches=0)
```
## 5 CelebA
This dataset must be downloaded separedly, you can find it in this link:
http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html
After downloading, make sure the folder structure is as follows:
```
📂<PARENT>
┗ 📂celeba
┗ 📂imgs
┣ 📄000001.jpg
┣ 📄000002.jpg
┗ 📄 ...
```
### 5.1 Showing original dataset
```
celeba_parent_path = '<PATH TO FOLDER CONTAINING THE DATASET>'
celeba_dir = os.path.join(celeba_parent_path, 'celeba', 'imgs')
files = os.listdir(celeba_dir)
data = []
for f in files[252:252+100]:
img = tf.keras.preprocessing.image.load_img(os.path.join(celeba_dir, f))
data.append(tf.keras.preprocessing.image.img_to_array(img, dtype='uint8'))
fig = plt.figure()
for i in range(3 * 4):
ax = plt.subplot(3, 4, i + 1)
plt.imshow(data[i])
plt.xticks([])
plt.yticks([])
plt.tight_layout()
plt.show()
fig.savefig('CelebA.pdf', bbox_inches='tight')
```
### 5.2 Showing reduced dataset
Make sure to have created the reduced dataset first by running the code in the file `datasets_preprocess.ipynb`
```
data = np.load(os.path.join('..', '..', 'celeba.npy'))
samples = data[68:68+60].astype('float32') / 255
grid = utils.fn.make_grid_image(samples, n_cols=10, border=1, pad=1)
fig = plt.figure()
plt.imshow(grid)
plt.axis(False)
plt.tight_layout(h_pad=0, w_pad=0)
fig.savefig('CelebA_reduced.pdf', bbox_inches='tight', pad_inches=0)
```
| github_jupyter |
To start, we need to import a bunch of libraries. Most of these should be included if you've installed the RDKit. The only libraries you should need to install are [minisom](https://github.com/JustGlowing/minisom) for the SOM and [tqdm](https://github.com/tqdm/tqdm) for progress bars.
```
pip install minisom tqdm
```
```
!pip install minisom
from collections import Counter
import pandas as pd
from matplotlib import pyplot as plt
from matplotlib.gridspec import GridSpec
from rdkit import Chem
from rdkit.Chem import AllChem, MACCSkeys, Draw
from rdkit import DataStructs
import numpy as np
from tqdm import tqdm
from minisom import MiniSom
import sys
from time import time
import math
```
Enable plots matplotlib plots in the this notebook
```
%matplotlib inline
```
A few functions to generate fingerprints. The first function generates 166-bit MACCS keys. The second generates Morgan fingerprints. While both will work for building a SOM, the process will be a bit faster with MACCS keys. I tend to like MACCS keys for generating SOMs. These fingerprints typically do a good job of grouping a set of molecules by scaffold. The third function takes a list of SMILES as input and returns as a list of fingerprints. If this function is called with one argument, it generates MACCS keys. We can also pass a function as a second argument to generate a different fingerprint type. For instance, we could call it like this to generate Morgan fingerprints.
```
generate_fps(my_smiles_list,morgan_as_np)
```
```
def maccs_as_np(mol):
"""
Generate MACCS fingerprints as a NumPy array
:param mol: input molecule
:return: fingerprint as a NumPy array
"""
bv = MACCSkeys.GenMACCSKeys(mol)
return np.array([int(x) for x in list(bv.ToBitString())], dtype=np.float32)
def morgan_as_np(mol):
"""
Generate a 1024 bit Morgan fingerprint as a NumPy array
:param mol: input molecule
:return: fingerprint as a NumPy array
"""
bv = AllChem.GetMorganFingerprintAsBitVect(mol, 3, nBits=1024)
arr = np.zeros((1,), dtype=np.float32)
DataStructs.ConvertToNumpyArray(bv, arr)
return arr
def generate_fps(smiles_list, fp_function=maccs_as_np):
"""
Take a list of SMILES as input and return a list of NumPy arrays
:param smiles_list: list of SMILES
:param fp_function: function to calculate fingerprints
:return: list of NumPy arrays containing fingerprints
"""
output_fp_list = []
for smiles in tqdm(smiles_list, desc="Generating Fingerprints"):
output_fp_list.append(fp_function(Chem.MolFromSmiles(smiles)))
return output_fp_list
```
A function to generate a grid of pie charts showing the distribution of active and inactive compounds in each grid cell.
```
# Adapted from the MiniSom example notebook
def depict_som(cluster_df, x_dim, y_dim, x_column="X", y_column="Y"):
"""
Draw a SOM with each cell depicted as a pie chart
:param cluster_df: data frame with SOM output, should have columns active, X, and Y
:param x_dim: X dimension of the SOM
:param y_dim: Y dimension of the SOM
:return:
"""
required_colums = [x_column, y_column, "active"]
for col in required_colums:
if col not in cluster_df.columns:
print(f"Error {col} not in dataframe columns", file=sys.stderr)
sys.exit(1)
cell_dict = {}
for k, v in [x for x in cluster_df.groupby([x_column, y_column])]:
cell_dict[k] = Counter(v["active"])
cell_names = cluster_df["active"].unique()
plt.figure(figsize=(x_dim, y_dim))
the_grid = GridSpec(x_dim, y_dim)
for position in cell_dict.keys():
label_fracs = [cell_dict[position][l] for l in cell_names]
plt.subplot(the_grid[(x_dim - 1) - position[1], position[0]], aspect=1)
patches, texts = plt.pie(label_fracs)
```
Build a SOM with minisom
```
def build_minisom_som(fp_list_in, x_dim=10, y_dim=10, num_iters=20000):
"""
Build a SOM with MiniSom
:param fp_list_in: input list of fingerprints as NumPy arrays
:param x_dim: X dimension of the SOM
:param y_dim: Y dimension of the SOM
:param num_iters: number of iterations when building the SOM
:return: lists with X and Y coordinates in the SOM
"""
print("Training SOM")
start_time = time()
som = MiniSom(x_dim, y_dim, len(fp_list_in[0]), sigma=0.3, learning_rate=0.5, random_seed=1)
som.train_random(fp_list_in, num_iters)
x = []
y = []
# find best matching units
print("Finding BMUs")
for row in fp_list_in:
x_val, y_val = som.winner(row)
x.append(x_val)
y.append(y_val)
elapsed_time = time()-start_time
print("Done\nElapsed time = %.2f sec" % elapsed_time)
return x, y
```
Now that we have the necessary functions in place, we can have some fun. We will read in a csv file containing SMILES, a molecule name, and 1 or 0 indicating that the molecule is active into a Pandas data frame. The first few lines of the file look like this.
```
SMILES,Name,active
Cn1ccnc1Sc2ccc(cc2Cl)Nc3c4cc(c(cc4ncc3C#N)OCCCN5CCOCC5)OC,168691,1
C[C@@]12[C@@H]([C@@H](CC(O1)n3c4ccccc4c5c3c6n2c7ccccc7c6c8c5C(=O)NC8)NC)OC,86358,1
Cc1cnc(nc1c2cc([nH]c2)C(=O)N[C@H](CO)c3cccc(c3)Cl)Nc4cccc5c4OC(O5)(F)F,575087,1
Cc1cnc(nc1c2cc([nH]c2)C(=O)N[C@H](CO)c3cccc(c3)Cl)Nc4cccc5c4OCO5,575065,1
```
We then use the function generate_fps to generate a list of fingerprints from the SMILES column in the dataframe. This list of fingerprints is then used to generate X and Y coordinates for each molecule in the grid. The x and y coordinates generated by build_minisom_som are then added as columns to the original dataframe. This dataframe, as well as the grid dimensions, are then passed to the depiction function to generate the plot below.
```
# read the input file into a dataframe
act_df = pd.read_csv("dude_erk2_mk01.csv")
# generate fingerprints
fp_list = generate_fps(act_df.SMILES)
# build the SOM
x_dim = 10
y_dim = 10
som_x, som_y = build_minisom_som(fp_list, x_dim, y_dim)
# set dataframe columns to SOM output
act_df["X"] = som_x
act_df["Y"] = som_y
# Draw the SOM
depict_som(act_df, x_dim, y_dim)
```
The dataset we used above is the [ERK2 (aka MK01)](http://dude.docking.org/targets/mk01) dataset that is part of the [DUD-E dataset](http://dude.docking.org/), which was designed for the evaluation of docking programs. The DUDE-E database consists of sets of active compounds, curated from the literature, and decoy compounds with similar calculated properties (molecular weight, LogP). The compound sets in the database were designed to evaluate the ability of a docking program to distinguish active compounds from decoys.
In the plot above the active compounds are shown in blue, while the decoy compounds are shown in orange. As we can see, our fingerprints do a reasonably good job of separating the active compounds from the decoys. In particular, we see that one cell at position 6,4 (we start counting from 0) is highly enriched in active compounds. Let's take a closer look at molecules in that cell.
We can use the Pandas query function to create a new dataframe containing only the molecules at position 6,4
```
my_cell = act_df.query("X==6 and Y==4")
```
To make it easier to visualize the molecules in cell 6,4 we will sort the SMILES so that the active molecules are first, followed by the decoy molecules.
```
my_smiles_list = [x[1] for x in sorted(zip(my_cell.active,my_cell.SMILES),reverse=True)]
```
Active and decoy are currently indicated by 1 and 0 in the dataframe. To have better labels, we will convert 1 and 0 to the strings "active" and decoy.
```
my_active_list = [x[0] for x in sorted(zip(my_cell.active,my_cell.SMILES),reverse=True)]
active_label_list = ["active" if a else "decoy" for a in my_active_list]
Draw.MolsToGridImage([Chem.MolFromSmiles(x) for x in my_smiles_list],molsPerRow=5,legends=active_label_list)
```
A quick look at the figure above indicates that the figure above shows that the active compounds are indeed very similar. The small number of decoy compounds in the same cell are somewhat similar to the actives but are clearly different chemotypes.
```
morgan_list = generate_fps(act_df.SMILES,morgan_as_np)
x_dim = 10
y_dim = 10
morgan_x, morgan_y = build_minisom_som(morgan_list, x_dim, y_dim)
act_df["morgan_X"] = morgan_x
act_df["morgan_Y"] = morgan_y
depict_som(act_df, x_dim, y_dim, x_column="morgan_X",y_column="morgan_Y")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sholtodouglas/learning_from_play/blob/master/experimental/Generate_dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
# TODO: Images in the latent space plot, and then images on test (need to supply them as goals)
import argparse
parser = argparse.ArgumentParser(description='LFP training arguments')
parser.add_argument('run_name')
parser.add_argument('--train_datasets', nargs='+', help='Training dataset names')
parser.add_argument('--test_datasets', nargs='+', help='Testing dataset names')
parser.add_argument('--bulk_datasets', nargs='+', help='data diversity dataset names')
parser.add_argument('--video_datasets', nargs='+', help='for contrastive learning')
parser.add_argument('-c', '--colab', default=False, action='store_true', help='Enable if using colab environment')
parser.add_argument('-s', '--data_source', default='DRIVE', help='Source of training data')
parser.add_argument('-tfr', '--from_tfrecords', default=False, action='store_true', help='Enable if using tfrecords format')
parser.add_argument('-d', '--device', default='TPU', help='Hardware device to train on')
parser.add_argument('-b', '--batch_size', default=512, type=int)
parser.add_argument('-wmax', '--window_size_max', default=50, type=int)
parser.add_argument('-wmin', '--window_size_min', default=20, type=int)
parser.add_argument('-la', '--actor_layer_size', default=2048, type=int, help='Layer size of actor, increases size of neural net')
parser.add_argument('-le', '--encoder_layer_size', default=512, type=int, help='Layer size of encoder, increases size of neural net')
parser.add_argument('-lp', '--planner_layer_size', default=2048, type=int, help='Layer size of planner, increases size of neural net')
parser.add_argument('-lg', '--goal_mapper_layer_size', default=512, type=int, help='Layer size of goal mapping networks from im and sent to goal space, increases size of neural net')
parser.add_argument('-embd', '--img_embedding_size', default=64, type=int, help='Embedding size of features,goal space')
parser.add_argument('-s_embd', '--sentence_embedding_size', default=512, type=int, help='Embedding size of MUSE sentence embeddings')
parser.add_argument('-g_embd', '--gripper_img_embedding_size', default=32, type=int, help='Embedding size of features,goal space')
parser.add_argument('-z', '--latent_dim', default=256, type=int, help='Size of the VAE latent space')
parser.add_argument('-zg', '--goal_space_dim', default=32, type=int, help='Size of the goal embedding space')
parser.add_argument('-g', '--gcbc', default=False, action='store_true', help='Enables GCBC, a simpler model with no encoder/planner')
parser.add_argument('-n', '--num_distribs', default=None, type=int, help='Number of distributions to use in logistic mixture model')
parser.add_argument('-q', '--qbits', default=None, type=int, help='Number of quantisation bits to discrete distributions into. Total quantisations = 2**qbits')
parser.add_argument('-lr', '--learning_rate', type=float, default=3e-4)
parser.add_argument('-t', '--train_steps', type=int, default=200000)
parser.add_argument('-r', '--resume', default=False, action='store_true')
parser.add_argument('-B', '--beta', type=float, default=0.00003)
parser.add_argument('-i', '--images', default=False, action='store_true')
parser.add_argument('-i2', '--images2', default=False, action='store_true')
parser.add_argument('-gi', '--gripper_images', default=False, action='store_true')
parser.add_argument('-cnn', '--cnn_type', type=str, default="spatial_softmax")
parser.add_argument('-sim', '--sim', default='Unity', help='Unity/Pybullet')
parser.add_argument('-vq', '--discrete', default=False, action='store_true')
parser.add_argument('-tmp', '--temperature', type=float, default=0.1)
parser.add_argument('--vq_tiles', type=int, default=5) # split into 5 tiles, must cleanly divide into max_seq_len
parser.add_argument('-nm', '--normalize', default=False, action='store_true')
parser.add_argument('-lang', '--use_language', default=False, action='store_true')
parser.add_argument('-cont', '--use_contrastive', default=False, action='store_true')
parser.add_argument('-enc_all', '--encode_all', default=False, action='store_true', help='encode_actions_and_proprio not just imgs')
parser.add_argument('-sub', '--sub_out_language_percent', type=float, default=0.25)
parser.add_argument('--fp16', default=False, action='store_true')
parser.add_argument('--bucket_name', help='GCS bucket name to stream data from')
parser.add_argument('--tpu_name', help='GCP TPU name') # Only used in the script on GCP
# Set these to split the dataset up so we control the proportion of lang vs bulk vs video etc - make them batch numbers
parser.add_argument('-ss', '--standard_split', type=int, default=0)
parser.add_argument('-bs', '--bulk_split', type=int, default=0)
parser.add_argument('-ls', '--lang_split', type=int, default=0)
parser.add_argument('-vs', '--video_split', type=int, default=0)
# args = parser.parse_args('''
# PROB0_02
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# -n 5
# '''.split())
# args = parser.parse_args('''
# IMB0_00003
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# -i
# -tfr
# -wmin 10
# -wmax 40
# '''.split())
args = parser.parse_args('''
PROB_B0_02
--train_dataset pybullet/UR5, pybullet/UR5_high_transition, pybullet/UR5_slow_gripper
--test_dataset pybullet/UR5_slow_gripper_test
-c
-d GPU
-b 2
-la 2048
-le 512
-lp 512
-z 256
-lr 3e-4
-tfr
-wmin 20
-wmax 40
-n 5
-sim Pybullet
--encode_all
'''.split())
# args = parser.parse_args('''
# GCSB0_0003
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# '''.split())
path = f"saved_models/{args.run_name}"
import argparse
parser = argparse.ArgumentParser(description='LFP training arguments')
parser.add_argument('run_name')
parser.add_argument('--train_datasets', nargs='+', help='Training dataset names')
parser.add_argument('--test_datasets', nargs='+', help='Testing dataset names')
parser.add_argument('--bulk_datasets', nargs='+', help='data diversity dataset names')
parser.add_argument('--video_datasets', nargs='+', help='for contrastive learning')
parser.add_argument('-c', '--colab', default=False, action='store_true', help='Enable if using colab environment')
parser.add_argument('-s', '--data_source', default='DRIVE', help='Source of training data')
parser.add_argument('-tfr', '--from_tfrecords', default=False, action='store_true', help='Enable if using tfrecords format')
parser.add_argument('-d', '--device', default='TPU', help='Hardware device to train on')
parser.add_argument('-b', '--batch_size', default=512, type=int)
parser.add_argument('-wmax', '--window_size_max', default=50, type=int)
parser.add_argument('-wmin', '--window_size_min', default=20, type=int)
parser.add_argument('-la', '--actor_layer_size', default=2048, type=int, help='Layer size of actor, increases size of neural net')
parser.add_argument('-le', '--encoder_layer_size', default=512, type=int, help='Layer size of encoder, increases size of neural net')
parser.add_argument('-lp', '--planner_layer_size', default=2048, type=int, help='Layer size of planner, increases size of neural net')
parser.add_argument('-lg', '--goal_mapper_layer_size', default=512, type=int, help='Layer size of goal mapping networks from im and sent to goal space, increases size of neural net')
parser.add_argument('-embd', '--img_embedding_size', default=64, type=int, help='Embedding size of features,goal space')
parser.add_argument('-s_embd', '--sentence_embedding_size', default=512, type=int, help='Embedding size of MUSE sentence embeddings')
parser.add_argument('-g_embd', '--gripper_img_embedding_size', default=32, type=int, help='Embedding size of features,goal space')
parser.add_argument('-z', '--latent_dim', default=256, type=int, help='Size of the VAE latent space')
parser.add_argument('-zg', '--goal_space_dim', default=32, type=int, help='Size of the goal embedding space')
parser.add_argument('-g', '--gcbc', default=False, action='store_true', help='Enables GCBC, a simpler model with no encoder/planner')
parser.add_argument('-n', '--num_distribs', default=None, type=int, help='Number of distributions to use in logistic mixture model')
parser.add_argument('-q', '--qbits', default=None, type=int, help='Number of quantisation bits to discrete distributions into. Total quantisations = 2**qbits')
parser.add_argument('-lr', '--learning_rate', type=float, default=3e-4)
parser.add_argument('-t', '--train_steps', type=int, default=200000)
parser.add_argument('-r', '--resume', default=False, action='store_true')
parser.add_argument('-B', '--beta', type=float, default=0.00003)
parser.add_argument('-i', '--images', default=False, action='store_true')
parser.add_argument('-i2', '--images2', default=False, action='store_true')
parser.add_argument('-gi', '--gripper_images', default=False, action='store_true')
parser.add_argument('-cnn', '--cnn_type', type=str, default="spatial_softmax")
parser.add_argument('-sim', '--sim', default='Unity', help='Unity/Pybullet')
parser.add_argument('-vq', '--discrete', default=False, action='store_true')
parser.add_argument('-tmp', '--temperature', type=float, default=0.1)
parser.add_argument('--vq_tiles', type=int, default=5) # split into 5 tiles, must cleanly divide into max_seq_len
parser.add_argument('-nm', '--normalize', default=False, action='store_true')
parser.add_argument('-lang', '--use_language', default=False, action='store_true')
parser.add_argument('-cont', '--use_contrastive', default=False, action='store_true')
parser.add_argument('-enc_all', '--encode_all', default=False, action='store_true', help='encode_actions_and_proprio not just imgs')
parser.add_argument('-sub', '--sub_out_language_percent', type=float, default=0.25)
parser.add_argument('--fp16', default=False, action='store_true')
parser.add_argument('--bucket_name', help='GCS bucket name to stream data from')
parser.add_argument('--tpu_name', help='GCP TPU name') # Only used in the script on GCP
# Set these to split the dataset up so we control the proportion of lang vs bulk vs video etc - make them batch numbers
parser.add_argument('-ss', '--standard_split', type=int, default=0)
parser.add_argument('-bs', '--bulk_split', type=int, default=0)
parser.add_argument('-ls', '--lang_split', type=int, default=0)
parser.add_argument('-vs', '--video_split', type=int, default=0)
# args = parser.parse_args('''
# PROB0_02
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# -n 5
# '''.split())
# args = parser.parse_args('''
# IMB0_00003
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# -i
# -tfr
# -wmin 10
# -wmax 40
# '''.split())
args = parser.parse_args('''
PROB_B0_02
--train_dataset pybullet/UR5, pybullet/UR5_high_transition, pybullet/UR5_slow_gripper
--test_dataset pybullet/UR5_slow_gripper_test
-c
-d GPU
-b 2
-la 2048
-le 512
-lp 512
-z 256
-lr 3e-4
-tfr
-wmin 20
-wmax 40
-n 5
-i
-sim Pybullet
--encode_all
'''.split())
# args = parser.parse_args('''
# GCSB0_0003
# --train_dataset UR5 UR5_slow_gripper UR5_high_transition
# --test_dataset UR5_slow_gripper_test
# -c
# -d GPU
# -b 512
# -la 2048
# -le 512
# -lp 512
# -z 256
# -lr 3e-4
# '''.split())
path = f"saved_models/{args.run_name}"
from importlib import reload
reload(lfp)
reload(lfp.data)
GLOBAL_BATCH_SIZE = 1
dl = lfp.data.PlayDataloader(normalize=args.normalize, include_imgs = args.images, include_imgs2 = args.images2, include_gripper_imgs = args.gripper_images, sim=args.sim, batch_size=GLOBAL_BATCH_SIZE, window_size=args.window_size_max, min_window_size=args.window_size_min, shuffle_size=1000)
valid_data = dl.extract(TEST_DATA_PATHS, from_tfrecords=args.from_tfrecords)
valid_dataset = dl.load(valid_data)
v_it = iter(valid_dataset)
import skimage.io as io
import matplotlib.pyplot as plt
from IPython.display import display, clear_output
from lfp.data import serialise_traj
def play_trajectory(images):
for i in range(0, len(images)):
if i % 5 == 0:
clear_output(wait=True)
fig = plt.imshow(images[i])
plt.show()
tag_matching = {
0: 'button',
1: 'cupboard door left',
2: 'cupboard door right',
3: 'drawer in',
4: 'drawer out',
5: 'block in/out drawer',
6: 'block in/out cupboard',
7: 'block',
8: 'block + shelf',
9: 'multi_object',
}
filename = 'labels'
tagging = True
if tagging: filename = 'tagged_labels'
path = TEST_DATA_PATHS[0]
path/filename
try:
data = list(np.load(path/f'{filename}.npz', allow_pickle=True)['data'])
except:
data = []
while(1):
batch = v_it.next()
play_trajectory(batch['imgs'][0])
val = input(f" {len(data)} Label: ")
while val in ['r']:
play_trajectory(images)
val = input("Label: ")
if val == 's':
pass
elif val == 'q':
print(f'Saving')
np.savez(path/filename, data=data, allow_objects=True)
break
else:
if tagging:
tag = input("Tag: ")
data.append({'batch':batch, 'label':val, 'tag':tag_matching[int(tag)]})
else:
data.append({'batch':batch, 'label':val, 'tag': ""})
np.savez(path/filename, data=data, allow_objects=True)
batch
import tensorflow_hub as hub
embed = hub.KerasLayer(str(STORAGE_PATH)+'/saved_models/universal_sentence_encoder')
save_path = str(path/'tf_records')+f"/{filename}.tfrecords"
save_path = str(path/'tf_records')+f"\\{filename}.tfrecords"
save_path
from tqdm import tqdm
with tf.io.TFRecordWriter(save_path) as file_writer:
for d in tqdm(data):
trajectory = {k:v[0] for k,v in d['batch'].items() if k not in ['dataset_path', 'tstep_idxs']}
trajectory['seq_lens'] = trajectory['obs'].shape[0]
trajectory['masks'] = tf.ones([ trajectory['obs'].shape[0]])
trajectory['goal_imgs'] = trajectory['imgs'][-1]
trajectory['label'] = d['label']
trajectory['label_embedding'] = np.squeeze(embed([d['label']])) # # 512
trajectory['tag'] = d['tag']
byte_stream = serialise_traj(trajectory)
file_writer.write(byte_stream)
record_paths = []
for p in [TRAIN_DATA_PATHS[0]]:
record_paths += tf.io.gfile.glob(str(p/'tf_records/*.tfrecords'))
d = extract_tfrecords(record_paths)
it = iter(d)
obs, acts, ags = [], [], []
for i in range(0,5000):
d1 = it.next()
obs.append(tf.io.parse_tensor(d1['obs'], tf.float32))
acts.append(tf.io.parse_tensor(d1['acts'], tf.float32))
ags.append(tf.io.parse_tensor(d1['achieved_goals'], tf.float32))
np.array(obs).shape
from tensorflow.train import BytesList, FloatList, Int64List
from tensorflow.train import Example, Features, Feature
def serialise(data):
obs, acts, achieved_goals, joint_poses, target_poses, acts_quat, acts_rpy_rel, velocities, obs_quat, sequence_index, sequence_id, img , proprioception = data['obs'], data['acts'], data['achieved_goals'], data['joint_poses'], data['target_poses'], data['acts_quat'], data['acts_rpy_rel'], data['velocities'], data['obs_quat'], data['sequence_index'], data['sequence_id'], data['img'], data['proprioception']
obs = Feature(bytes_list=BytesList(value=[obs.numpy(),]))
acts = Feature(bytes_list=BytesList(value=[acts.numpy(),]))
achieved_goals = Feature(bytes_list=BytesList(value=[achieved_goals.numpy(),]))
joint_poses = Feature(bytes_list=BytesList(value=[joint_poses.numpy(),]))
target_poses = Feature(bytes_list=BytesList(value=[target_poses.numpy(),]))
acts_quat = Feature(bytes_list=BytesList(value=[acts_quat.numpy(),]))
acts_rpy_rel = Feature(bytes_list=BytesList(value=[acts_rpy_rel.numpy(),]))
velocities = Feature(bytes_list=BytesList(value=[velocities.numpy(),]))
obs_quat = Feature(bytes_list=BytesList(value=[obs_quat.numpy(),]))
proprioception = Feature(bytes_list=BytesList(value=[proprioception.numpy(),]))
sequence_index = Feature(int64_list=Int64List(value=[sequence_index.numpy(),]))
sequence_id = Feature(int64_list=Int64List(value=[sequence_id.numpy(),]))
img = Feature(bytes_list=BytesList(value=[img.numpy(),]))
# img is already serialised because we never decode it!
features = Features(feature={
'obs': obs,
'acts': acts,
'achieved_goals': achieved_goals,
'joint_poses': joint_poses,
'target_poses': target_poses,
'acts_quat': acts_quat,
'acts_rpy_rel': acts_rpy_rel,
'velocities': velocities,
'obs_quat': obs_quat,
'proprioception': proprioception,
'sequence_index': sequence_index,
'sequence_id': sequence_id,
'img': img,
})
example = Example(features=features)
return example.SerializeToString()
# Write the records to a file.
from tqdm import tqdm_notebook
counter = 0
i = 0
path = str(STORAGE_PATH/'unified/tf_records')+f"/{counter}.tfrecords"
path
it = iter(d)
with tf.io.TFRecordWriter(path) as file_writer:
for data in tqdm_notebook(it):
byte_stream = serialise(data)
file_writer.write(byte_stream)
i += 1
if (i>0) and (i % 25000 == 0):
counter += 1
serialise(d1)
record_paths = tf.io.gfile.glob(str(STORAGE_PATH/'unified/tf_records/*.tfrecords'))
d2 = extract_tfrecords(record_paths)
it = iter(d2)
c= 0
for i in it:
c += 1
if c % 1000 == 0:
print(c, i['sequence_id'].numpy(), i['sequence_index'].numpy())
# d1_copy = it.next()
# d1_copy
for k,v in d1.items():
print(d1_copy[k].numpy() == v.numpy())
batch_size = 16
def load_tf_records(filenames, ordered=False):
# Read from TFRecords. For optimal performance, reading from multiple files at once and
# disregarding data order. Order does not matter since we will be shuffling the data anyway.
# check, does this ignore intra order or just inter order? Both are an issue!
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False # disable order, increase speed
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=60) # automatically interleaves reads from multiple files - keep it at 1 we need the order
dataset = dataset.with_options(ignore_order) # uses data as soon as it streams in, rather than in its original order
dataset = dataset.map(read_tfrecord, num_parallel_calls=600)
dataset = dataset.repeat()\
.batch(batch_size, drop_remainder=True)\
# .prefetch(dl.prefetch_size)
return dataset
data_paths = [str(STORAGE_PATH/'precompute')+f"/{x}.tfrecords" for x in range(0,13)]
d =load_tf_records(data_paths, ordered=False)#['/home/sholto/Desktop/AI/learning_from_play/data/UR5_high_transition/tf_records/21.tfrecords'], ordered=True) #record_paths
print(d)
d = iter(d)
x = d.next()
t = time.time()
for i in range(0,10):
# print(i)
x = d.next()
print(time.time()-t)
dl = lfp.data.PlayDataloader(include_imgs = args.images, batch_size=batch_size, window_size=args.window_size_max, min_window_size=args.window_size_min)
train_data = dl.extract(TRAIN_DATA_PATHS, from_tfrecords=args.from_tfrecords)
train_dataset = dl.load(train_data)
test_it = iter(train_dataset)
t = time.time()
for i in range(0,10):
test_it.next()
print(time.time()-t)
for i in range(0, 100):
example = t_it.next()
# np.frombuffer(example['goal_imgs'].numpy().tobytes(), np.uint8)
bstr = serialise(example)
d = read_tfrecord(bstr)
import matplotlib.pyplot as plt
plt.imshow(d['imgs'][0,15])
def load_tf_records(filenames, ordered=False):
# Read from TFRecords. For optimal performance, reading from multiple files at once and
# disregarding data order. Order does not matter since we will be shuffling the data anyway.
# check, does this ignore intra order or just inter order? Both are an issue!
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False # disable order, increase speed
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=tf.data.experimental.AUTOTUNE) # automatically interleaves reads from multiple files - keep it at 1 we need the order
dataset = dataset.with_options(ignore_order) # uses data as soon as it streams in, rather than in its original order
dataset = dataset.map(read_tfrecord, num_parallel_calls=tf.data.experimental.AUTOTUNE)
return dataset
d =load_dataset([record_paths], ordered=True)#['/home/sholto/Desktop/AI/learning_from_play/data/UR5_high_transition/tf_records/21.tfrecords'], ordered=True) #record_paths
print(d)
d = iter(d)
for k,v in example.items():
print(k, v.shape)
d
with tf.io.TFRecordWriter(str(path/'tf_records/')+f"/{demo}.tfrecords") as file_writer:
for data in it:
byte_stream = serialise(data)
file_writer.write(byte_stream)
import subprocess
ll = args.train_datasets
print(ll)
files = []
for l in ll :
files.append(tf.io.gfile.glob(f'gs://lfp_europe_west4_a/data/{l}/tf_records/*.tfrecords'))
print(len(files))
def done(files):
for l in files:
if len(l) > 0:
return False
return True
counter = 0
while not done(files):
for l in files:
if len(l) > 0:
r = l.pop()
list_files = subprocess.run(["gsutil", "cp", r, f"gs://lfp_europe_west4_a/data/combined/tf_records/{counter}.tfrecords"])
counter += 1
print(r)
```
| github_jupyter |
**Consider the following Python dictionary data and Python list labels:**
data = {'birds': ['Cranes', 'Cranes', 'plovers', 'spoonbills', 'spoonbills', 'Cranes', 'plovers', 'Cranes', 'spoonbills', 'spoonbills'],
'age': [3.5, 4, 1.5, np.nan, 6, 3, 5.5, np.nan, 8, 4],
'visits': [2, 4, 3, 4, 3, 4, 2, 2, 3, 2],
'priority': ['yes', 'yes', 'no', 'yes', 'no', 'no', 'no', 'yes', 'no', 'no']}
labels = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j']
```
import pandas as pd
import numpy as np
data = {'birds': ['Cranes', 'Cranes', 'plovers', 'spoonbills', 'spoonbills', 'Cranes', 'plovers', 'Cranes', 'spoonbills', 'spoonbills'], 'age': [3.5, 4, 1.5, np.nan, 6, 3, 5.5, np.nan, 8, 4], 'visits': [2, 4, 3, 4, 3, 4, 2, 2, 3, 2], 'priority': ['yes', 'yes', 'no', 'yes', 'no', 'no', 'no', 'yes', 'no', 'no']}
labels = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j']
```
**1. Create a DataFrame birds from this dictionary data which has the index labels.**
```
df = pd.DataFrame(data,index=labels)
df
```
**2. Display a summary of the basic information about birds DataFrame and its data.**
```
df.info()
df.describe()
```
**3. Print the first 2 rows of the birds dataframe **
```
df[:2]
df.head(2)
```
**4. Print all the rows with only 'birds' and 'age' columns from the dataframe**
```
df[['birds','age']]
print(df['birds'])
print(df['age'])
```
**5. select [2, 3, 7] rows and in columns ['birds', 'age', 'visits']**
```
df[['birds', 'age', 'visits']].iloc[[2,3,7]]
print(df['birds'].iloc[2],df['age'].iloc[2],df['visits'].iloc[2])
print(df['birds'].iloc[3],df['age'].iloc[3],df['visits'].iloc[3])
print(df['birds'].iloc[7],df['age'].iloc[7],df['visits'].iloc[7])
```
**6. select the rows where the number of visits is less than 4**
```
df[df['visits'] < 4]
```
**7. select the rows with columns ['birds', 'visits'] where the age is missing i.e NaN**
```
df[['birds', 'visits']][df['age'].isnull()]
```
**8. Select the rows where the birds is a Cranes and the age is less than 4**
```
df[(df['birds']=='Cranes') & (df['age']<4)]
```
**9. Select the rows the age is between 2 and 4(inclusive)**
```
df[(df['age']>=2)&(df['age']<=4)]
```
**10. Find the total number of visits of the bird Cranes**
```
df[df['birds']=='Cranes']['visits'].sum()
```
**11. Calculate the mean age for each different birds in dataframe.**
```
df[['birds','age']].groupby(['birds']).mean()
df['age'].mean()
```
**12. Append a new row 'k' to dataframe with your choice of values for each column. Then delete that row to return the original DataFrame.**
```
k_row = {'birds':'Parrot','age':2.5,'visits':2,'priority':'yes'}
df_k = pd.DataFrame(k_row,index=['k'])
df = df.append(df_k)
df
df.drop(index='k',inplace=True)
df
```
**13. Find the number of each type of birds in dataframe (Counts)**
```
df.groupby(['birds']).count()
df.groupby(['birds']).size()
```
**14. Sort dataframe (birds) first by the values in the 'age' in decending order, then by the value in the 'visits' column in ascending order.**
```
df.sort_values(['age','visits'],ascending=[False,True])
```
**15. Replace the priority column values with'yes' should be 1 and 'no' should be 0**
```
birds = df.copy()
birds['priority'] = birds.apply(lambda x: 1 if x['priority'] == 'yes' else 0 , axis=1)
birds
```
**16. In the 'birds' column, change the 'Cranes' entries to 'trumpeters'.**
```
birds = df.copy()
birds['birds'] = birds.apply(lambda x:'trumpeters' if x['birds']== 'Cranes' else x['birds'],axis=1)
birds
```
| github_jupyter |
## Analyzing full sweep: sub5, n=5
```
import os
import pickle
import numpy as np
import matplotlib.pyplot as plt
import gvar as gv
import scipy.sparse as sp
import scipy.sparse.linalg as sla
import lsqfit # WARNING: lsqfit 12.0 appears to be broken. `python3 -m pip install lsqfit==11.8`
from z2_sim.src.NumericsPython.Analysis.fourieranalysis import fourier_ana
from z2_sim.src.QuantumCircuits.Cirq_Code import io
%load_ext autoreload
%autoreload 2
```
### The data management is yucky.
But you already know that by now. See `noise_analysis_sub3`
```
noiseless_directory = "./results/noiseless"
flag = 'obc'
ns = [5]
# Load the noiseless data into a single dictionary that can be passed to `fourier_ana`
noiseless_data = {}
noiseless_tag = "control"
dts = [0.25]
for n in ns:
noiseless_subdirectory = os.path.join(noiseless_directory, f"{n}/")
noiseless_fname = f'noiseless_parameter_sweep_{noiseless_tag}_results_n{n}_{flag}.npy'
arr = np.load(os.path.join(noiseless_subdirectory, noiseless_fname))[:,:,:50,:,:]
noiseless_data[(flag, n)] = arr
# Submission card
dt = 0.25
j_sweep = np.asarray([0.714285, 0.625, .555556])
zeta_sweep = [0, 150000, 300000, 450000, 600000, 750000]
eps_sweep = [0, 0.0005, 0.001, 0.0015, 0.002, 0.0025, 0.003]
n_trajectories = 1000
tstart = 1
tstop = 51
nprocs = 1
# assert statements never hurt anyone
j_sweep_control = np.load( os.path.join(noiseless_directory, io.make_physical_params_sweep_fname(noiseless_tag, 'jcoup')))
assert np.allclose(j_sweep, j_sweep_control, atol=1e-5, rtol=1e-4)
noisy_directory = './results/noisy'
beta_sweep = 1 / j_sweep
assert np.allclose(beta_sweep, [1.4, 1.6, 1.8])
beta_sweep = [1.4, 1.6, 1.8]
beta_keys = np.round(beta_sweep, decimals=6)
# # Load the data into separate dictionaries that can be passed to `fourier_ana`
# # This way, any element of this data sequence can be treated on equal footing
# # with a noiseless data array.
noisy_data_grid = [[{} for _ in range(len(eps_sweep))] for _ in range(len(zeta_sweep))]
for n in ns:
noisy_subdirectory = os.path.join(noisy_directory, f"{n}/")
collector = io.CondorCollector(
path=noisy_subdirectory,
nprocs=nprocs,
n=n,
j_sweep=j_sweep,
dt_sweep=[dt],
trotter_intervals=[(tstart, tstop)],
zeta_sweep=zeta_sweep,
eps_sweep=eps_sweep, # don't look for eps=0
r=n_trajectories,
)
#!! epsilon dependence stored in the 4th index
#!! shot dependence stored in the 0th index
data_arr = collector.collect_htcondor_outputs()
for i in range(len(zeta_sweep)):
for j in range(len(eps_sweep)):
# (nprocs, len(j_sweep), len(dt_sweep), len(zeta_sweep), len(eps_sweep), trotter_steps, n, n)
noisy_data_grid[i][j][('obc', n)] = data_arr[:,:,:,i,j,:,:,:].mean(axis=0)
import pickle
with open(f'./final/noisy/{n}/noisy_n{n}_raw.pickle', 'w') as f:
pickle.dump(noisy_data_grid, f)
```
#### Verify that noisy results are seeded differently!
The fact that the plot below has many lines means that each noisy simulation was different. This was a concern of mine that seeds might get duplicated during the condor submission process.
### Perform the fourier analysis and mass analysis on each dataset
Noiseless datset followed by noisy dataset at each noise parameter.
```
bc_flags = ['obc']
noiseless_fourier = fourier_ana(
results=noiseless_data,
bc_flags=bc_flags,
grid_sizes=ns,
dts=dts,
betas=beta_sweep,
n_trotter=50,
samp_rate = 2048,
nboot = 200)
noisy_fourier_grid = [[None for _ in range(len(eps_sweep))] for _ in range(len(zeta_sweep))]
for i in range(len(zeta_sweep)):
for j in range(len(eps_sweep)):
noisy_data = noisy_data_grid[i][j]
noisy_fourier = fourier_ana(
results=noisy_data,
bc_flags=bc_flags,
grid_sizes=ns,
dts=dts,
betas=beta_sweep,
n_trotter=50,
samp_rate = 2048,
nboot = 200
)
noisy_fourier_grid[i][j] = noisy_fourier
noisy_data_grid[-1][2][('obc', n)][2, 0,:,1,1].real
import matplotlib.cm as cm
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(eps_sweep) )) ]))
k = 2
beta = beta_sweep[k]
trotter_steps = 50
for zeta_idx in [0, -1]:
fig, axes = plt.subplots(1, 2, figsize=(10, 5))
# Plot time series
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, eps in enumerate(eps_sweep):
y = noisy_data_grid[zeta_idx][i][('obc', n)][k, 0,:,1,1].real
axes[0].plot(range(trotter_steps), y, label=f'{eps}', c=colors[i], alpha=0.7)
# Plot spectrum
for i, eps in enumerate(eps_sweep):
y = noisy_fourier_grid[zeta_idx][i][0][('obc', n, 0.25, beta)]
axes[1].plot(range(trotter_steps), y, label=r"$\epsilon=$" + f'{eps}', c=colors[i], alpha=0.7)
axes[0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=2)
axes[1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=2)
axes[0].set_ylabel(r"$\langle O_s(t) \rangle$", size=20)
axes[0].set_xlabel(r"$t$", size=24)
axes[1].set_xlabel(r"$\omega$", size=24)
axes[1].set_ylabel(r"$F_O(\omega) $", size=20)
axes[1].legend(bbox_to_anchor=(1.0, 1), framealpha=1)
for ax in axes:
ax.grid()
# fig.savefig("../figures/sample_epsilon_timeseries.pdf")
import matplotlib.cm as cm
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(eps_sweep) -1 )) ]))
for k in range(3):
# k = 0
beta = beta_sweep[k]
trotter_steps = 50
zeta_idx = 0
fig, axes = plt.subplots(1, 2, figsize=(14, 5), constrained_layout=True)
# Plot time series
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_data_grid[zeta_idx][i][('obc', n)][k, 0,:,1,1].real
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[0].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# Plot spectrum
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_fourier_grid[zeta_idx][i][0][('obc', n, 0.25, beta)]
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[1].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# OVERLAY NOISELESS
axes[0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=1.5)
axes[1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=1.5)
axes[0].set_title(f"{n} x {n} grid, " + r"$(\beta, dt)= $" + f"({beta}, 0.25), " + r"$\zeta=0$", size=14)
axes[0].set_ylabel(r"$\mathcal{Re}\,\langle \mathcal{C}_{s,s}(t) \rangle$", size=20)
axes[0].set_xlabel(r"$t$", size=24)
axes[1].set_xlabel(r"$\omega$", size=24)
axes[1].set_ylabel(r"$A_{s,s}(\omega) $", size=20)
axes[1].legend(bbox_to_anchor=(1.0, 1), framealpha=1, prop={'size': 14})
for ax in axes:
ax.grid()
ax.set_xlim(0, 50)
# fig.savefig(f"../figures/sample_n{n}_zeta0_eps_timeseries.pdf")
```
## DEMO PLOTS
### $\zeta=0$, $\epsilon_2$ sweep
```
import matplotlib.cm as cm
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(eps_sweep) -1 )) ]))
k = 0
beta = beta_sweep[k]
trotter_steps = 50
zeta_idx = 0
fig, axes = plt.subplots(1, 2, figsize=(14, 5), constrained_layout=True)
# Plot time series
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_data_grid[zeta_idx][i][('obc', n)][k, 0,:,1,1].real
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[0].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# Plot spectrum
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_fourier_grid[zeta_idx][i][0][('obc', n, 0.25, beta)]
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[1].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# OVERLAY NOISELESS
axes[0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=1.5)
axes[1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=1.5)
axes[0].set_title(f"{n} x {n} grid, " + r"$(\beta, dt)= $" + f"({beta}, 0.25), " + r"$\zeta=0$", size=14)
axes[0].set_ylabel(r"$\mathcal{Re}\,\langle \mathcal{C}_{s,s}(t) \rangle$", size=20)
axes[0].set_xlabel(r"$t$", size=24)
axes[1].set_xlabel(r"$\omega$", size=24)
axes[1].set_ylabel(r"$A_{s,s}(\omega) $", size=20)
axes[1].legend(bbox_to_anchor=(1.0, 1), framealpha=1, prop={'size': 14})
for ax in axes:
ax.grid()
ax.set_xlim(0, 50)
fig.savefig(f"../figures/sample_n{n}_zeta0_eps_timeseries.pdf")
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(zeta_sweep)-1)) ]))
k = 0
beta = beta_sweep[k]
trotter_steps = 50
eps_idx = 0
fig, axes = plt.subplots(1, 2, figsize=(14, 5), constrained_layout=True)
# Plot time series
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, zeta in enumerate(zeta_sweep[1:]):
y = noisy_data_grid[i][eps_idx][('obc', n)][k, 0,:,1,1].real
lab = r"$\zeta=$" + f"{zeta / 10**5}" + r"$\times 10^5$"
axes[0].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# Plot spectrum
for i, zeta in enumerate(zeta_sweep[1:]):
y = noisy_fourier_grid[i][eps_idx][0][('obc', n, 0.25, beta)]
lab = r"$\zeta=$" + f"{zeta / 10**5}" + r"$\times 10^5$"
axes[1].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# OVERLAY NOISELESS
axes[0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=1.5)
axes[1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=1.5)
axes[0].set_title(f"{n} x {n} grid, " + r"$(\beta, dt)= $" + f"({beta}, 0.25), " + r"$\epsilon_2=0$", size=14)
axes[0].set_ylabel(r"$\mathcal{Re}\,\langle \mathcal{C}_{s,s}(t) \rangle$", size=20)
axes[0].set_xlabel(r"$t$", size=24)
axes[1].set_xlabel(r"$\omega$", size=24)
axes[1].set_ylabel(r"$A_{s,s}(\omega) $", size=20)
axes[1].legend(bbox_to_anchor=(1.0, 1), framealpha=1, prop={'size': 14})
for ax in axes:
ax.grid()
ax.set_xlim(0, 50)
fig.savefig(f"../figures/sample_n{n}_zeta_eps0_timeseries.pdf")
```
### Shared plot!
```
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(eps_sweep) -1 )) ]))
k = 0
beta = beta_sweep[k]
trotter_steps = 50
zeta_idx = 0
fig, axes = plt.subplots(2, 2, figsize=(14, 8), constrained_layout=True, sharex=True, sharey=False)
# Plot time series
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_data_grid[zeta_idx][i][('obc', n)][k, 0,:,1,1].real
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[0,0].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# Plot spectrum
for i, eps in enumerate(eps_sweep[1:]):
y = noisy_fourier_grid[zeta_idx][i][0][('obc', n, 0.25, beta)]
lab = r"$\epsilon_2=$" + f"{eps * 10**3}" + r"$\times 10^{-3}$"
axes[0,1].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# OVERLAY NOISELESS
axes[0,0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=1.5)
axes[0,1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=1.5)
axes[0,0].set_title(f"{n} x {n} grid, " + r"$(\beta_H, \delta t)= $" + f"({beta}, 0.25), " + r"$\zeta=0$", size=14)
axes[0,0].set_ylabel(r"$\mathcal{Re}\,\langle \mathcal{C}_{s,s}(t) \rangle$", size=20)
# axes[0,0].set_xlabel(r"$t$", size=24)
# axes[0,1].set_xlabel(r"$\omega$", size=24)
axes[0,1].set_ylabel(r"$A_{s,s}(\omega) $", size=20)
axes[0,1].legend(bbox_to_anchor=(1.0, 1), framealpha=1, prop={'size': 14})
colors = list(reversed([x for x in cm.rainbow(np.linspace(0, 1, len(zeta_sweep)-1)) ]))
# axes: [beta, dt, trotter_step, n, n]
# slice for dt=0.25, and keep only beta in [1.4, 1.6, 1.8]
for i, zeta in enumerate(zeta_sweep[1:]):
y = noisy_data_grid[i][eps_idx][('obc', n)][k, 0,:,1,1].real
lab = r"$\zeta=$" + f"{zeta / 10**5}" + r"$\times 10^5$"
axes[1,0].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# Plot spectrum
for i, zeta in enumerate(zeta_sweep[1:]):
y = noisy_fourier_grid[i][eps_idx][0][('obc', n, 0.25, beta)]
lab = r"$\zeta=$" + f"{zeta / 10**5}" + r"$\times 10^5$"
axes[1,1].plot(range(trotter_steps), y, label=lab, c=colors[i], alpha=1, lw=1.1)
# OVERLAY NOISELESS
axes[1,0].plot(noiseless_data[('obc', n)][k, 0,:,1,1].real, label="Noiseless", c='k', lw=1.5)
axes[1,1].plot(noiseless_fourier[0][('obc', n, 0.25, beta)], label="Noiseless", c='k', lw=1.5)
axes[1,0].set_ylabel(r"$\mathcal{Re}\,\langle \mathcal{C}_{s,s}(t) \rangle$", size=20)
axes[1,0].set_xlabel(r"$t$", size=24)
axes[1,1].set_xlabel(r"$\omega$", size=24)
axes[1,1].set_ylabel(r"$A_{s,s}(\omega) $", size=20)
axes[1,1].legend(bbox_to_anchor=(1.0, 1), framealpha=1, prop={'size': 14})
for ax in axes.flatten():
ax.grid()
ax.set_xlim(0, 50)
fig.savefig(f"../figures/sample_n{n}_zeta_eps_timeseries_combined.pdf")
```
## Bulk analysis
The primary analysis that we can apply en masse is to compare $\Delta E_{01}$ as a function of $dt$ across different noise strengths.
### Start by looking at mass versus dt
The following is less condensed but can maybe reveal trends in the noise behavior.
```
rel_err_grid = np.zeros((len(beta_sweep), len(zeta_sweep), len(eps_sweep)))
n = 5
dt = 0.25
for i, betakey in enumerate(beta_keys):
# Plot noiseless energy vs dt
ens = noiseless_fourier[1]
noiseless_y = np.abs(np.array(gv.mean(ens[('obc', n, dt, betakey)])))
for j, zeta in enumerate(zeta_sweep):
for k, eps in enumerate(eps_sweep):
ens_noisy = noisy_fourier_grid[j][k][1]
# Compute and plot absolute error
noisy_y = np.abs(np.array(gv.mean(ens_noisy[('obc', n, dt, betakey)])))
noisy_yerr = gv.sdev(ens_noisy[('obc', n, dt, betakey)])
# Plot relative error
rel_err = (noisy_y - noiseless_y) / noiseless_y
rel_err_grid[i,j,k] = rel_err
rel_err_grid[i,0,0] = 0
rel_err_grid[0]
rel_err_grid[1]
rel_err_grid[2]
for i, beta in enumerate(beta_sweep):
fig, ax = plt.subplots()
im = ax.imshow(rel_err_grid[i], cmap="seismic", vmin=-2, vmax=2, origin='lower')
ax.set_title(r"$\beta=$" + f"{beta}")
ax.set_xticks(range(len(eps_sweep)))
ax.set_xticklabels(eps_sweep)
ax.set_yticks(range(len(zeta_sweep)))
ax.set_yticklabels(zeta_sweep)
cbar_ax = fig.add_axes([0.85, 0.15, 0.05, 0.7])
fig.colorbar(im, cax=cbar_ax)
```
### Demo plot
```
import matplotlib.colors as colors
def truncate_colormap(cmap, minval=0.0, maxval=1.0, n=100):
new_cmap = colors.LinearSegmentedColormap.from_list(
'trunc({n},{a:.2f},{b:.2f})'.format(n=cmap.name, a=minval, b=maxval),
cmap(np.linspace(minval, maxval, n)))
return new_cmap
i = 1
beta = beta_sweep[i]
fig, ax = plt.subplots()
cmap = truncate_colormap(plt.get_cmap('seismic'), minval=0.5, maxval=1)
im = ax.imshow(abs(rel_err_grid[i]), cmap=cmap, vmin=0, origin='lower')
ax.set_xticks(range(len(eps_sweep)))
ax.set_xticklabels(eps_sweep)
ax.set_yticks(range(len(zeta_sweep)))
zeta_labs = ["0"] + [f"{zeta/(1e4 * np.pi * 2):3.2f}"+r"$\times 10^4$" for zeta in zeta_sweep[1:]]
ax.set_yticklabels(zeta_labs)
betastr = r"$\beta=$"+f"{beta}"
nstr = r"$n=$"+f"{n}"
ax.set_title(f"Glueball mass abs. rel. err. ({betastr}, {nstr})")
ax.set_xlabel(r"$\epsilon_2$", size=22)
ax.set_ylabel(r"$\zeta\,/\,2\pi$", size=22)
cbar_ax = fig.add_axes([0.85, 0.15, 0.05, 0.7])
fig.colorbar(im, cax=cbar_ax)
plt.savefig(f"../figures/glueball_err_demo_n{n}.pdf", bbox_inches='tight')
```
| github_jupyter |
# Variables
```
from __future__ import print_function
import torch
import numpy as np
import matplotlib
%matplotlib inline
import matplotlib.pyplot as plt
from datetime import date
date.today()
author = "kyubyong. https://github.com/Kyubyong/pytorch_exercises"
torch.__version__
np.__version__
```
NOTE on notation
_x, _y, _z, ...: NumPy 0-d or 1-d arrays
_X, _Y, _Z, ...: NumPy 2-d or higer dimensional arrays
x, y, z, ...: 0-d or 1-d tensors
X, Y, Z, ...: 2-d or higher dimensional tensors
```
from torch.autograd import Variable
```
## Variables
Q0. Create a variable `X` of the size (3, 2), filled with 1's.
```
X = Variable(torch.ones(3, 2))
print(X)
```
Q1. Get the tensor of Variable X.
```
X = Variable(torch.randn(3, 3))
Y = X.data
print(Y)
```
Q2. Complete the code.
```
# Create a trainable variable `w` of scalar 10.
w = Variable(torch.ones(1)*10, requires_grad=True)
gs, ws, grads = [], [], []
for i in range(10):
y = w ** 2 + 2
# apply backpropagation to y.
y.backward()
w.data -= 0.01 * w.grad.data
gs.append(i)
ws.append(w.data[0])
grads.append(w.grad.data[0])
plt.figure(figsize=(15,6))
ax=plt.subplot(1, 2, 1)
ax.scatter(gs, ws, c="b", label="w")
ax.legend(loc="upper right")
ax=plt.subplot(1, 2, 2)
plt.scatter(gs, grads, c="r", label="gradient")
plt.legend(loc="upper left")
plt.show()
```
Q3. Complete the code. <br>This is adapted from `http://pytorch.org/tutorials/beginner/examples_autograd/two_layer_net_autograd.html`.
```
# Untrainable variables
# Create untrainable variables X and Y.
X = Variable(torch.randn(64, 1000), requires_grad=False)
Y = Variable(torch.randn(64, 10), requires_grad=False)
# Trainable Variables
w1 = Variable(torch.randn(1000, 100), requires_grad=True)
w2 = Variable(torch.randn(100, 10), requires_grad=True)
# Parameters
n_epochs = 500
learning_rate = 1e-6
log_interval = 10
losses = []
for t in range(n_epochs):
Y_pred = X.matmul(w1).clamp(min=0).matmul(w2)
# Apply L2 loss to Y pred and Y.
loss = (Y_pred - Y).pow(2).sum()
losses.append(loss.data[0])
loss.backward()
w1.data -= learning_rate * w1.grad.data
w2.data -= learning_rate * w2.grad.data
# Manually zero the gradients after updating weights
w1.grad.data.zero_()
w2.grad.data.zero_()
if (t + 1) % log_interval == 0:
print("Epoch {:03d}/{:03d}: loss {:.5f}".format(
t + 1, n_epochs, loss.data[0]))
# plot
plt.figure()
plt.plot(losses, label="loss")
plt.legend()
plt.show()
```
| github_jupyter |
```
#all_slow
```
# Tutorial: Fine-Tuning a Language Model on Text Files with IMDB
> Tuning a base Language model on the IMDB dataset
## Introduction
In this tutorial we will be showing an end-to-end example of fine-tuning a Transformer language model on a custom dataset in text files format.
By the end of this you should be able to:
1. Build a dataset with the `LanguageModelDatasets` class, and their DataLoaders
2. Build a `LanguageModelTuner` quickly, find a good learning rate, and train with the One-Cycle Policy
3. Save that model away, to be used with deployment or other HuggingFace libraries
4. Apply inference using both the `Tuner` available function as well as with the `EasyTextGenerator` class within AdaptNLP
## Installing the Library
This tutorial utilizies the latest AdaptNLP version, as well as parts of the `fastai` library. Please run the below code to install them:
```python
!pip install adaptnlp -U
```
(or `pip3`)
```
#hide
from nbverbose.showdoc import *
```
## Getting the Dataset
First we need a dataset. We will use the `fastai` library to download the full `IMDB` Movie Reviews dataset
```
from fastai.data.external import URLs, untar_data
```
`URLs` holds a namespace of many data endpoints, and `untar_data` is a function that can download and extract any data from a given URL.
Combining both, we can download the data:
```
data_path = untar_data(URLs.IMDB)
```
If we look at what was downloaded, we will find a `train` and `test` folder:
```
data_path.ls()
```
In each are folders seperating each text file by class:
```
(data_path/'train').ls()
```
As a result, we can say the dataset follows the following format:
- `train`
- `class_a`
- `text1.txt`
- `text2.txt`
- ...
- `class_b`
- `text1.txt`
- ...
- `test` (or `valid`)
- `class_a`
- `text1.txt`
- ...
- `class_b`
- `text1.txt`
- ...
> Note: In this instance, test and validation have very similar meanings. Both are the dataset which is used to calculate the metrics during training (such as accuracy or F1Score)
Now that we have the dataset, and we know the format it is in, let's pick a viable model to train with
## Picking a Model with the Hub
AdaptNLP has a `HFModelHub` class that allows you to communicate with the HuggingFace Hub and pick a model from it, as well as a namespace `HF_TASKS` class with a list of valid tasks we can search by.
Let's try and find one suitable for sequence classification.
First we need to import the class and generate an instance of it:
```
from adaptnlp import HFModelHub, HF_TASKS
hub = HFModelHub()
```
Next we can search for a model:
```
models = hub.search_model_by_task(HF_TASKS.TEXT_GENERATION)
```
Let's look at a few:
```
models[:10]
```
These are models specifically tagged with the `text-generation` tag, so you may not see a few models you would expect such as `bert_base_cased`.
We'll use that first model, `distilgpt2`:
```
model = models[0]
model
```
Now that we have picked a model, let's use the data API to prepare our data
> Note: It should be mentioned that this is optional, you can always just pass in the string name of a model such as "bert-base-cased"
## Building `TaskDatasets` with `LanguageModelDatasets`
Each task has a high-level data wrapper around the `TaskDatasets` class. In our case this is the `LanguageModelDatasets` class:
```
from adaptnlp import LanguageModelDatasets
```
There are multiple different constructors for the `LanguageModelDatasets` class, and you should never call the main constructor directly.
We will be using `from_folders` method:
```
from adaptnlp import LanguageModelDatasets
show_doc(LanguageModelDatasets.from_folders)
```
Anything you would normally pass to the tokenizer call (such as `max_length`, `padding`) should go in `tokenize_kwargs`, and anything going to the `AutoTokenizer.from_pretrained` constructor should be passed to the `auto_kwargs`.
In our case we have a `train_path` and `valid_path`
```
from fastcore.basics import patch
```
Also, we will set a block_size of 128, and it is *not* a masked language model:
```
dsets = LanguageModelDatasets.from_folders(
train_path=data_path/'train',
valid_path=data_path/'test',
tokenizer_name=model.name,
block_size=128,
masked_lm=False
)
```
> Note: If you only have a training folder, just pass in a `split_func` or `split_pct` to either have it split the dataset in a custom way, or pass in a percentage to randomly split by
And finally turn it into some `AdaptiveDataLoaders`.
These are just fastai's `DataLoaders` class, but it overrides a few functions to have it work nicely with HuggingFace's `Dataset` class
```
show_doc(LanguageModelDatasets.dataloaders)
dls = dsets.dataloaders(batch_size=8)
```
Finally, let's view a batch of data with the `show_batch` function:
```
dls.show_batch()
```
When training a language model, the input and output are made to be the exact same, so there isn't a shown noticable difference here.
## Building `Tuner`
Next we need to build a compatible `Tuner` for our problem. These tuners contain good defaults for our problem space, including loss functions and metrics.
First let's import the `LanguageModelTuner` and view it's documentation
```
from adaptnlp import LanguageModelTuner
from adaptnlp import LanguageModelTuner
show_doc(LanguageModelTuner)
```
Next we'll pass in our `DataLoaders`, the name of our model, and the tokenizer:
> Note: If you are not using the data API (`TaskDatasets`, `SequenceClassificationDatasets`, etc), you need to pass in the tokenizer to the constructor as well with `tokenizer=tokenizer`
```
tuner = LanguageModelTuner(dls, model.name, dls.tokenizer)
```
By default we can see that it used `CrossEntropyLoss` as our loss function, and `Perplexity` as our metric
```
tuner.loss_func
_ = [print(m.name) for m in tuner.metrics]
```
Finally we just need to train our model!
## Fine-Tuning
And all that's left is to `tune`. There are only 4 or 5 functions you can call on our `tuner` currently, and this is by design to make it simplistic. In case you don't want to be boxed in however, if you pass in `expose_fastai_api=True` to our earlier call, it will expose the entirety of `Learner` to you, so you can call `fit_one_cycle`, `lr_find`, and everything else as `Tuner` uses `fastai` under the hood.
First, let's call `lr_find`, which uses fastai's Learning Rate Finder to help us pick a learning rate.
```
show_doc(LanguageModelTuner.lr_find)
tuner.lr_find()
```
It recommends a learning rate of around 5e-5, so we will use that.
```
lr = 5e-5
```
Let's look at the documentation for `tune`:
```
show_doc(LanguageModelTuner.tune)
```
We can pass in a number of epochs, a learning rate, a strategy, and additional fastai callbacks to call.
Valid strategies live in the `Strategy` namespace class, and consist of:
- OneCycle (Also called the [One-Cycle Policy](https://docs.fast.ai/callback.schedule.html#Learner.fit_one_cycle))
- [CosineAnnealing](https://docs.fast.ai/callback.schedule.html#Learner.fit_flat_cos)
- [SGDR](https://docs.fast.ai/callback.schedule.html#Learner.fit_sgdr)
```
from adaptnlp import Strategy
```
In this tutorial we will train with the One-Cycle policy, as currently it is one of the best schedulers to use.
```
tuner.tune(3, lr, strategy=Strategy.OneCycle)
```
## Saving Model
Now that we have a trained model, let's save those weights away.
Calling `tuner.save` will save both the model and the tokenizer in the same format as how HuggingFace does:
```
show_doc(LanguageModelTuner.save)
tuner.save('good_model')
```
## Performing Inference
There are two ways to get predictions, the first is with the `.predict` method in our `tuner`. This is great for if you just finished training and want to see how your model performs on some new data!
The other method is with AdaptNLP's inference API, which we will show afterwards
### In Tuner
First let's write a sentence to test with
```
sentence = "Hugh Jackman is a terrible"
```
And then predict with it:
```
show_doc(LanguageModelTuner.predict)
tuner.predict(sentence, num_tokens_to_produce=13)
```
### With the Inference API
Next we will use the `EasyTextGenerator` class, which AdaptNLP offers:
```
from adaptnlp import EasyTextGenerator
```
We simply construct the class:
```
classifier = EasyTextGenerator()
```
And call the `tag_text` method, passing in the sentence, the location of our saved model, and some names for our classes:
```
classifier.generate(
sentence,
model_name_or_path='good_model',
num_tokens_to_produce=13
)
```
And we got the exact same output!
| github_jupyter |
# Introduction
In this tutorial, we will learn about different environments, then implement dynamic programming (model-based) algorithms and study their performance.
# Notebook setup
## Instructions
- Import numpy, scipy and matplotlib
- Configure inline plots
```
% matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from pylab import *
```
# Tutorial
In order to learn about RL algorithms, we will explore different worlds (also called environments), which are listed below. For more details about each world, please look into their implementation in module RL_worlds.
## N-armed bandit task
In an N-armed bandit task, there is only a single state. Our implementation is a 4-armed bandit, which means there are 4 available actions, and each one returns a different reward.
```
from IPython.display import Image
Image(filename="fig/4ArmedBandit.png", width=600, height=225)
```
## Cheese world
In this one-dimensional grid there are 4 states and 2 possible actions: left and right. Arriving at the goal state gives you a reward of 1. Moving left from the start state (state 1 in the figure) stays in the same place, and moving anywhere from the goal state (state 4 in the figure) ends the episode.
```
from IPython.display import Image
Image(filename="fig/CheeseWorld.png", width=600, height=225)
```
## Cliff world
In this 4x10 grid there are 40 states and 4 possible actions: right, up, left and down. Falling into the cliff incurs a negative reward of -100 and ends the episode; moving into any other state incurs a reward of -1; moving into the world borders stays in the same place; moving anywhere from the goal state (state G in the figure) ends the episode.
```
from IPython.display import Image
Image(filename="fig/CliffWorld.png", width=600, height=225)
```
## Quentin's world
In this 10x10 grid there are 100 states and 4 possible actions: right, up, left and down. The start state is in green in the figure; moving into one of the red states incurs a reward of -1; moving into the world borders stays in the same place; moving into the goal state (yellow square in the upper right corner) gives you a reward of 1; and moving anywhere from the goal state ends the episode.
```
from IPython.display import Image
Image(filename="fig/QuentinsWorld.png", width=300, height=300)
```
## Multi-room windy gridworld with cliffs
In this 12x14 grid there are 168 states and 4 possible actions: right, up, left and down. The start state is marked with an S in the figure, and there are 2 goals states marked with a G. Each goal state is inside a room (the room walls are marked with darker lines). Moving into a goal states gives you a reward of 100. Moving into a wall or outside the world borders stays in the same place. The two rooms are windy, which means the resultant next states inside the rooms are shifted; the wind direction is indicated by a blue arrow, and the wind strength (size of the shift) in each column is indicated by a number between 0 and 2. There are also two cliffs marked in gray; falling into a cliff incurs a reward of -100 and ends the episode.
```
from IPython.display import Image
Image(filename="fig/gridworld.png", width=451.2, height=447.2)
```
## Helper functions
Please familiarize yourself with the code below, as it will help your write your own code to solve the exercises.
```
# Import definitions of the environments.
import RL_worlds as worlds
# Import helper functions for plotting.
from plot_util import *
def default_params(environment):
"""
Define the default parameters.
Args:
environment: an object corresponding to the environment.
Returns:
a dictionary containing the default parameters, where the keys
are strings (parameter names).
"""
params = dict()
params['environment'] = environment
params['alpha'] = 0.1 # learning rate
params['beta'] = 10 # inverse temperature
params['policy'] = 'epsilon_greedy'
params['epsilon'] = 0.05 # epsilon-greedy policy
params['learning_rule'] = 'q_learning'
params['epsilon_decay'] = 0.9
if environment.name == 'windy_cliff_grid':
params['gamma'] = 0.8 # temporal discount factor
elif environment.name == 'n_armed_bandit':
params['gamma'] = 0.9 # temporal discount factor
elif environment.name == 'cliff_world':
params['gamma'] = 1.0 # no discounting
elif environment.name == 'cheese_world':
params['gamma'] = 1.0 # no discounting
elif environment.name == 'quentins_world':
params['gamma'] = 0.9 # temporal discount factor
return params
```
## Exercise 1: Policy Evaluation
1. Write the policy evaluation algorithm.
Tip: the function should take the policy to be evaluated, the default parameters and a threshold for the stopping criterium as input and return the value function.
## Exercise 2: Policy Iteration
1. Write the policy iteration algorithm.
Tips: the function should take the default parameters and a threshold for the stopping criterium as input and return the optimal policy and the value function for the optimal policy; you should use the policy evaluation function written in Exercise 1.
## Exercise 3: Value Iteration
One drawback to policy iteration is that each of its iterations involves policy evaluation, which may be slow due to multiple sweeps through the state set. We can make this process faster by truncating the policy evaluation step of policy iteration, stopping it after just one sweep (one backup of each state). This modified algorithm is called value iteration. You can read more about it here: http://artint.info/html/ArtInt_227.html
1. Write the value iteration algorithm.
Tips: the function should take the default parameters and a threshold for the stopping criterium as input and return the policy and corresponding value function; you can write a helper function to calculate the one-step lookahead, i.e., the value for all actions in a given state using the current estimate of the value function.
## Exercise 4
1. Write code that allows you to test the performance of policy iteration and value iteration for a selected world (try cliff world, Quentin's world or windy cliff grid). Use the functions provided in the plot_util module to:
- Plot the action corresponding to the policy at each state;
- Plot the value associated with each state.
2. Experiment with different parameter values:
- Pick a range for the temporal discount factor $\gamma$ and look at how the results change.
3. Compare the results obtained using policy iteration to those obtained using value iteration. Do you notice any differences in their performance?
To make sure that your algorithms have been implemented correctly, compare your results to the ones shown below.
Cliff world using policy iteration and $\gamma$=0.9:
<img src="fig/tutorial1_ex4_policyimp_actions.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex4_policyimp_maxval.png",height="300",width="300">
Quentin's world using using value iteration and $\gamma$=0.9:
<img src="fig/tutorial1_ex4_valueit_actions.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex4_valueit_maxval.png",height="300",width="300">
| github_jupyter |
## Learning Rate Finder
```
%matplotlib inline
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai.callbacks import *
```
Learning rate finder plots lr vs loss relationship for a [`Learner`](/basic_train.html#Learner). The idea is to reduce the amount of guesswork on picking a good starting learning rate.
**Overview:**
1. First run lr_find `learn.lr_find()`
2. Plot the learning rate vs loss `learn.recorder.plot()`
3. Pick a learning rate before it diverges then start training
**Technical Details:** (first [described]('https://arxiv.org/abs/1506.01186') by Leslie Smith)
>Train [`Learner`](/basic_train.html#Learner) over a few iterations. Start with a very low `start_lr` and change it at each mini-batch until it reaches a very high `end_lr`. [`Recorder`](/basic_train.html#Recorder) will record the loss at each iteration. Plot those losses against the learning rate to find the optimal value before it diverges.
## Choosing a good learning rate
For a more intuitive explanation, please check out [Sylvain Gugger's post](https://sgugger.github.io/how-do-you-find-a-good-learning-rate.html)
```
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
def simple_learner(): return Learner(data, simple_cnn((3,16,16,2)), metrics=[accuracy])
learn = simple_learner()
```
First we run this command to launch the search:
```
show_doc(Learner.lr_find)
learn.lr_find(stop_div=False, num_it=200)
```
Then we plot the loss versus the learning rates. We're interested in finding a good order of magnitude of learning rate, so we plot with a log scale.
```
learn.recorder.plot()
min_grad_lr = learn.recorder.min_grad_lr
```
Then, we choose a value that is approximately in the middle of the sharpest downward slope. This is given as an indication by the LR Finder tool, so let's try 1e-2.
```
simple_learner().fit(2, 1e-2)
```
Don't just pick the minimum value from the plot!
```
learn = simple_learner()
simple_learner().fit(2, 1e-0)
```
Picking a value before the downward slope results in slow training:
```
learn = simple_learner()
simple_learner().fit(2, 1e-3)
```
#### Suggested LR
The red dot on the graph is the point with the minimum numerical gradient. We can use that point as a first guess for an LR
```
learn = simple_learner()
simple_learner().fit(2, min_grad_lr)
show_doc(LRFinder)
```
### Callback methods
You don't call these yourself - they're called by fastai's [`Callback`](/callback.html#Callback) system automatically to enable the class's functionality.
```
show_doc(LRFinder.on_train_begin)
show_doc(LRFinder.on_batch_end)
show_doc(LRFinder.on_epoch_end)
show_doc(LRFinder.on_train_end)
```
| github_jupyter |
```
import json
import pandas as pd
import numpy as np
import tqdm
import matplotlib.pyplot as plt
%matplotlib inline
import nltk
nltk.download('punkt')
```
# Stanford question answering dataset (SQuAD)
Today we are going to work with a popular NLP dataset.
Here is the description of the original problem:
```
Stanford Question Answering Dataset (SQuAD) is a new reading comprehension dataset, consisting of questions posed by crowdworkers on a set of Wikipedia articles, where the answer to every question is a segment of text, or span, from the corresponding reading passage. With 100,000+ question-answer pairs on 500+ articles, SQuAD is significantly larger than previous reading comprehension datasets.
```
We are not going to solve it :) Instead we will try to answer the question in a different way: given the question, we will find a **sentence** containing the answer, but not within the context, but in a **whole databank**
Just watch the hands
```
!wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v1.1.json
data = json.load(open('train-v1.1.json'))
data['data'][0]
```
The code here is very similar to `week5/`
```
from nltk.tokenize import RegexpTokenizer
from collections import Counter,defaultdict
tokenizer = RegexpTokenizer(r"\w+|\d+")
#Dictionary of tokens
token_counts = Counter()
def tokenize(value):
return tokenizer.tokenize(value.lower())
for q in tqdm.tqdm_notebook(data['data']):
for p in q['paragraphs']:
token_counts.update(tokenize(p['context']))
min_count = 4
tokens = [w for w, c in token_counts.items() if c > min_count]
dict_size = len(tokens)+2
id_to_word = dict()
word_to_id = dict()
token_to_id = {t:i+2 for i,t in enumerate(tokens)}
id_to_token = {i+2:t for i,t in enumerate(tokens)}
assert token_to_id['me'] != token_to_id['woods']
assert token_to_id[id_to_token[42]]==42
assert len(token_to_id)==len(tokens)
assert 0 not in id_to_token
from nltk.tokenize import sent_tokenize
def build_dataset(train_data):
'''Takes SQuAD data
Returns a list of tuples - a set of pairs (q, a_+)
'''
D = []
for q in tqdm.tqdm_notebook(train_data):
for p in q['paragraphs']:
offsets = []
curent_index = 0
for sent in sent_tokenize(p['context']):
curent_index+=len(sent)+2
offsets.append((curent_index, sent))
for qa in p['qas']:
answer = qa['answers'][0]
found = False
for o, sent in offsets:
if answer['answer_start']<o:
D.append((qa['question'], sent))
found = True
break
assert found
return D
from sklearn.model_selection import train_test_split
train_data, val_data = train_test_split(data['data'], test_size=0.1)
Dtrain = build_dataset(train_data)
Dval = build_dataset(val_data)
Dval[2]
def vectorize(strings, token_to_id, UNK):
'''This function gets a string array and transforms it to padded token matrix
Remember to:
- Transform a string to list of tokens
- Transform each token to it ids (if not in the dict, replace with UNK)
- Pad each line to max_len'''
max_len = 0
token_matrix = []
#<your code>\
num_tokens = []
for s in strings:
seq = []
for token in tokenize(s):
if token in token_to_id:
seq.append(token_to_id[token])
else:
seq.append(UNK)
token_matrix.append(seq)
max_len = max(max_len, len(token_matrix[-1]))
# empty batch plug
if max_len == 0:
max_len = 1
for i in range(len(token_matrix)):
num_tokens.append(len(token_matrix[i]))
while(len(token_matrix[i])<max_len):
token_matrix[i].append(0)
return np.array(token_matrix), np.array(num_tokens)
test = vectorize(["Hello, adshkjasdhkas, world", "data"], token_to_id, 1)[0]
assert test.shape==(2,3)
assert (test[:,1]==(1,0)).all()
print("Correct!")
```
# Deep Learning
The beginning is same as always
```
%env CUDA_VISIBLE_DEVICES = ""
import tensorflow as tf
tf.reset_default_graph()
target_space_dim = 50 # Here we define dimension of the target space
embeddings_size = 50
word_embeddings_matrix = <YOUR CODE>
def get_signle_input(name):
'''Returns a pair of inputs'''
return (tf.placeholder(dtype=tf.int32,shape=[None, None], name="word_ids_%s"%name),
tf.placeholder(dtype=tf.int32,shape=[None], name="num_words_%s"%name))
def encode(word_ids, num_words,name, reuse=False):
'''The function takes:
- word_ids - a matrix with word ids
- num_words - a vector, showing how many words is in each sample
- name - name for variables
- reuse - are weights reused
Returns:
- outputs - a matrix [batch_size, target_space_dim]
'''
<YOUR CODE>
return output
```
We are going to use a single `encode`, but with different weights. You can use different encode for anchor and negatives/positives.
Negative sampling can be either `in-graph` or `out-graph`. We start with out-graph. In the home assignment you are going to use in-graph.
```
def sample_semihard_outputs(anchor_output, positive_output):
"""Function samples negatives in-graph. Returns negative_output. Use it in home assignment"""
raise NotImplementedError
inputs = {name: get_signle_input(name) for name in ['anchor', 'positive', 'negative']}
margin = 0.1
anchor_output = encode(*inputs['anchor'], 'anchor')
positive_output = <YOUR CODE>
negative_output = <YOUR CODE>
positive_dot = <YOUR CODE>
negative_dot = <YOUR CODE>
loss = <YOUR CODE>
recall = tf.reduce_mean(tf.cast(tf.greater(positive_dot, negative_dot), tf.float32))
batch_size = 200
def iterate_batches(data, only_positives=False):
"""Takes a D
Returns a dict, containing pairs for each input type
only_positives indicates either we need to iterate over triplets vs only positive (needed for index)
"""
i = 0
while i < len(data):
batch = dict()
data_batch = data[i:i+batch_size]
batch['positive'] = vectorize([sample[1] for sample in data_batch], token_to_id, 1)
if not only_positives:
<YOUR CODE>
yield batch
i+=batch_size
optimizer = tf.train.AdamOptimizer() # <your code here>
global_step = tf.Variable(initial_value=0)
train_op = optimizer.minimize(
loss=loss,
global_step=global_step, var_list=tf.trainable_variables())
#list(iterate_batches(D))
def get_inputs(batch):
feed_dict = {}
for name, tensors in batch.items():
feed_dict[inputs[name][0]] = tensors[0]
feed_dict[inputs[name][1]] = tensors[1]
return feed_dict
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
def validate():
total_loss, total_recall = 0, 0
batches = 0
for batch in iterate_batches(Dval):
batches+=1
current_loss, current_recall = sess.run([loss, recall], get_inputs(batch))
total_loss+=current_loss
total_recall+=current_recall
total_loss/=batches
total_recall/=batches
if total_recall > 0.9:
print('Cool! If recall is right, you earned (3 pts)')
return (total_loss, total_recall)
num_epochs = 100
for j in range(num_epochs):
for i, (batch) in enumerate(iterate_batches(Dtrain)):
_, step, current_loss, current_recall = sess.run([train_op, global_step, loss, recall], get_inputs(batch))
print("Current step: %s. Current loss is %s, Current recall is %s" % (step, current_loss, current_recall))
if i%100==0:
print("Validation. Loss: %s, Recall: %s" %validate())
class Index(object):
"""Represents index of calculated embeddings"""
def __init__(self, D):
"""Class constructor takes a dataset and stores all unique sentences and their embeddings"""
<YOUR CODE>
def predict(self, query, top_size =1):
"""
Function takes:
- query is a string, containing question
Function returns:
- a list with len of top_size, containing the closet answer from the index
You may want to use np.argpartition
"""
<YOUR CODE>
def calculate_FHS(self, D):
"""Prototype for home assignment. Returns a float number"""
raise NotImplementedError
index = Index(Dval)
assert len(index.vectors) == len(index.sent)
assert type(index.sent[1])==str
assert index.vectors.shape == (len(index.sent), target_space_dim)
p = index.predict("Hey", top_size=3)
assert len(p) == 3
assert type(p[0])==str
assert index.predict("Hello", top_size=50)!=index.predict("Not Hello", top_size=50)
print("Ok (2 pts)")
index.predict('To show their strength in the international Communist movement, what did China do?', top_size=10)
Dval[np.random.randint(0, 100)]
```
# Home assignment
**Task 1.** (3 pts) Implement **semihard** sampling strategy. Use **in-graph** sampling. You have a prototype above
**Task 2.1.** (1 pt) Calculate a **FHS** (First Hit Success) metric on a whole validation dataset (over each query on whole `Dval` index). Prototype of the function in in `Index` class. Compare different model based on this metric. Add table with FHS values to your report.
**Task 2.2.** Add calculation of other representative metrics. You may want to calculate different recalls on a mini-batch, or some ranking metrics.
**Task 3.** (2 pt) Do experiments with deep architecture and find out the best one. Analyse your results and write a conclusion.
**describe your results here**
Bonus task 1. (2++ pts) Add manual negatives to the model. What can be a good manual negative in this case?
Bonus task 2. (2++ pts) Implement more efficient Nearest Neighbors Search method. How well it performs on our dataset?
| github_jupyter |
# High Level Python API
Up until now we have been using the low level Python API that Bifrost has to show the inner workings of the framework and how to build a pipeline. However, Bifrost also has a high level Python API that makes building blocks and pipelines easier with less code. In this section we will look at this interface.
The general idea of the high-level API is to allow a pipeline to be built out of processing blocks that are connected together like Lego. For example, here is a pipeline that reads from a sigproc file, performs a fast dispersion measure transform, and then writes out the data to disk:
```python
import bifrost as bf
import sys
filenames = sys.argv[1:]
print("Building pipeline")
data = bf.blocks.read_sigproc(filenames, gulp_nframe=128)
data = bf.blocks.copy(data, 'cuda')
data = bf.blocks.transpose(data, ['pol', 'freq', 'time'])
data = bf.blocks.fdmt(data, max_dm=100.)
data = bf.blocks.copy(data, 'cuda_host')
bf.blocks.write_sigproc(data)
print("Running pipeline")
bf.get_default_pipeline().run()
print("All done")
```
Here we are continually passing the block's output, `data`, to the next block in the pipeline. At runtime, the blocks are connected together (as a directed graph) with ring buffers in between.
### Creating a transform block
Many common processing operations have bifrost blocks, but it likely that you will need to make a custom block at some stage. To see how, let's start by revisiting the `CopyOp` block from the pipelines section:
```
class CopyOp(object):
def __init__(self, iring, oring, ntime_gulp=250, guarantee=True, core=-1):
self.iring = iring
self.oring = oring
self.ntime_gulp = ntime_gulp
self.guarantee = guarantee
self.core = core
def main(self):
with self.oring.begin_writing() as oring:
for iseq in self.iring.read(guarantee=self.guarantee):
ihdr = json.loads(iseq.header.tostring())
print("Copy: Start of new sequence:", str(ihdr))
time_tag = ihdr['time_tag']
navg = ihdr['navg']
nbeam = ihdr['nbeam']
chan0 = ihdr['chan0']
nchan = ihdr['nchan']
chan_bw = ihdr['bw'] / nchan
npol = ihdr['npol']
pols = ihdr['pols']
pols = pols.replace('CR', 'XY_real')
pols = pols.replace('CI', 'XY_imag')
igulp_size = self.ntime_gulp*nbeam*nchan*npol*4 # float32
ishape = (self.ntime_gulp,nbeam,nchan,npol)
self.iring.resize(igulp_size, igulp_size*5)
ogulp_size = igulp_size
oshape = ishape
self.oring.resize(ogulp_size)
ohdr = ihdr.copy()
ohdr_str = json.dumps(ohdr)
iseq_spans = iseq.read(igulp_size)
with oring.begin_sequence(time_tag=time_tag, header=ohdr_str) as oseq:
for ispan in iseq_spans:
if ispan.size < igulp_size:
continue # Ignore final gulp
with oseq.reserve(ogulp_size) as ospan:
idata = ispan.data_view(numpy.float32)
odata = ospan.data_view(numpy.float32)
odata[...] = idata
```
There is a lot of setup in here and iteration control that is common across many of the blocks that we have looked at. In the high level API much of this can be abstracted away using the classes defined in `bifrost.pipelines`:
```
import copy
from bifrost import pipeline
class NewCopyOp(pipeline.TransformBlock):
def __init__(self, iring, *args, **kwargs):
super(NewCopyOp, self).__init__(iring, *args, **kwargs)
def on_sequence(self, iseq):
ihdr = iseq.header
print("Copy: Start of new sequence:", str(ihdr))
ohdr = copy.deepcopy(iseq.header)
return ohdr
def on_data(self, ispan, ospan):
in_nframe = ispan.nframe
out_nframe = in_nframe
idata = ispan.data
odata = ospan.data
odata[...] = idata
return out_nframe
```
That is much more compact. The key things in this new class are:
1. the `on_sequence` method is called whenever a new sequence starts and is used to update the header for the output ring buffer and
2. the `on_data` method is called for each span/gulp that is processed.
Put another way, the `on_sequence` happens when there is new *metadata*, and `on_data` happens whenever there is new *data* to process. For example, `on_sequence` may be called when reading a new file, or starting an observation.
### Creating a source block
Similarly, we can translate the `GeneratorOp` and `WriterOp` blocks as well using sub-classes of `bifrost.pipeline.SourceBlock` and `bifrost.pipeline.SinkBlock`, respectively:
```
import os
import time
import numpy
class NewGeneratorOp(pipeline.SourceBlock):
def __init__(self, ntime_gulp, *args, **kwargs):
super(NewGeneratorOp, self).__init__(['generator',], 1,
*args, **kwargs)
self.ntime_gulp = ntime_gulp
self.ngulp_done = 0
self.ngulp_max = 10
self.navg = 24
tint = self.navg / 25e3
self.tgulp = tint * self.ntime_gulp
self.nbeam = 1
self.chan0 = 1234
self.nchan = 16*184
self.npol = 4
def create_reader(self, name):
self.ngulp_done = 0
class Random(object):
def __init__(self, name):
self.name = name
def __enter__(self):
return self
def __exit__(self, type, value, tb):
return True
def read(self, *args):
return numpy.random.randn(*args)
return Random(name)
def on_sequence(self, reader, name):
ohdr = {'time_tag': int(int(time.time())*196e6),
'seq0': 0,
'chan0': self.chan0,
'cfreq0': self.chan0*25e3,
'bw': self.nchan*25e3,
'navg': self.navg,
'nbeam': self.nbeam,
'nchan': self.nchan,
'npol': self.npol,
'pols': 'XX,YY,CR,CI',
}
ohdr['_tensor'] = {'dtype': 'f32',
'shape': [-1,
self.ntime_gulp,
self.nbeam,
self.nchan,
self.npol]
}
return [ohdr,]
def on_data(self, reader, ospans):
indata = reader.read(self.ntime_gulp, self.nbeam, self.nchan, self.npol)
time.sleep(self.tgulp)
if indata.shape[0] == self.ntime_gulp \
and self.ngulp_done < self.ngulp_max:
ospans[0].data[...] = indata
self.ngulp_done += 1
return [1]
else:
return [0]
```
For the `bifrost.pipeline.SourceBlock` we need have slightly different requirements on `on_sequence` and `on_data`. Plus, we also need to define a `create_reader` method that returns a context manager (a class with `__enter__` and `__exit__` methods).
For `on_sequence` we need to accept two arguments: a context manager created by `create_reader` and an identifying name (although it is not used here).
For `on_data` we also have two arguments now, the context manager and a list of output spans. In here we need to grab the data from `reader` and put it into the appropriate part of the output spans.
### The all-important bifrost `_tensor` dictionary
We also see in `on_sequence` here that the header dictionary has a new required `_tensor` key. This key is the key to automatically chaining blocks together into a pipeline since it defines the data type and dimensionality for the spans/gulps. At a minimum, the `_tensor` must define the data dtype and shape:
```python
'_tensor': {
'dtype': self.dtype,
'shape': [-1, self.gulp_size],
},
```
The first index of shape is -1, to indicate that this is a single gulp from the data stream.
However, most block require three additional keywords:
* `labels`, which give human-friendly names to each axis (e.g. 'time' or 'frequency').
* `scales`, which defines the start value and step size for each axis (e.g. `[1420, 0.1]` sets start value 1420, step size 0.1).
* `units`, which defines the units for each axis (e.g. 's' or 'MHz'). These are parsed using [Pint](https://pint.readthedocs.io/en/stable/), and can be used for consistency checking. For example, the `bf.views.merge_axes` view won't allow axes to merge if they have different units, say 'MHz' and 'Jy'. If you attempted to merge axes with 'MHz' and 'kHz' units, this would be allowed, and the corresponding `scales` are updated consistently.
Here is another example `_tensor` with all keywords:
```python
t0 = 1620192408.005579 # unix timestamp from when writing this tutorial
dt = 0.1 # 0.1 second step size
'_tensor': {
'dtype': 'cf32'
'shape': [-1, 1024, 4],
'labels': ['time', 'frequency', 'pol'],
'units': ['s', 'MHz', ''],
'scales': [[t0, dt], [1420, 0.1], [None, None]]
}
```
A transform block reads the tensor metadata, and must output a copy of the tensor with any required changes to shape/scale/units etc.
### Creating a sink block
We can also translate the original `WriterOp`:
```
class NewWriterOp(pipeline.SinkBlock):
def __init__(self, iring, *args, **kwargs):
super(NewWriterOp, self).__init__(iring, *args, **kwargs)
self.time_tag = None
self.navg = 0
self.nbeam = 0
self.nchan = 0
self.npol = 0
def on_sequence(self, iseq):
ihdr = iseq.header
print("Writer: Start of new sequence:", str(ihdr))
self.time_tag = iseq.time_tag
self.navg = ihdr['navg']
self.nbeam = ihdr['nbeam']
self.nchan = ihdr['nchan']
self.npol = ihdr['npol']
def on_data(self, ispan):
idata = ispan.data.view(numpy.float32)
idata = idata.reshape(-1, self.nbeam, self.nchan, self.npol)
with open(f"{self.time_tag}.dat", 'wb') as fh:
fh.write(idata.tobytes())
print(' ', fh.name, '@', os.path.getsize(fh.name))
self.time_tag += self.navg * idata.shape[0] * (int(196e6) // int(25e3))
```
Since this is a data sink we only have one argument for `on_data` which gives the block the current data span/gulp.
We then can put these new blocks all together and launch them under Bifrost's default pipeline with:
```
b_gen = NewGeneratorOp(250)
b_cpy = NewCopyOp(b_gen)
b_out = NewWriterOp(b_cpy)
p = pipeline.get_default_pipeline()
p.run()
del p
```
| github_jupyter |
In this notebook we show a basic script to reconstruct IMAT white beam data. We assume that users are familiar with basic framework concepts.
```
# imports
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import os
import scipy
# ccpi imports
from ccpi.framework import AcquisitionData, AcquisitionGeometry
from ccpi.framework import ImageData, ImageGeometry
from ccpi.processors import CenterOfRotationFinder, Resizer
from ccpi.astra.processors.FBP import FBP
# general parameters
imsize = 2048
# path
path = '/media/newhd/shared/Data/neutrondata/crab/TomoData/Fossils/'
## load flat-field
# path to flat-field
path_flat = path + 'Flat/'
# filename mask
filename_flat = 'IMAT00005147_openbeamafterf123_ob_{:03d}.tif'
# number of flats
n_flats = 40
# allocate average flat
flat = np.zeros((imsize, imsize), dtype=np.float32)
# loop through flats and calculate average flat
for i in range(n_flats):
# generate filename
filename = (path_flat + filename_flat).format(i + 1)
# load flat
try:
flat += np.transpose(np.asarray(Image.open(filename), dtype = np.float32))
except:
print('Error reading\n {}\n file.'.format(filename))
raise
flat /= n_flats
## load dark-field
# path to dark-field
path_dark = path + 'Dark/'
# filename mask
filename_dark = 'IMAT00005138_fossil1_darkbef_{:03d}.tif'
# number of darks
n_darks = 20
# allocate average dark
dark = np.zeros((imsize, imsize), dtype=np.float32)
# loop through flats and calculate average flat
for i in range(n_darks):
# generate filename
filename = (path_dark + filename_dark).format(i)
# load dark
try:
dark += np.transpose(np.asarray(Image.open(filename), dtype = np.float32))
except:
print('Error reading\n {}\n file.'.format(filename))
raise
dark /= n_darks
## load projections
# path to projections
path_projection = path + 'Sample/'
# filename mask
filename_projection = 'IMAT00005154_fossils_tomo_Sample_{:03d}.tif'
# number of projections
n_proj = 1049
# allocate array to store projections
proj = np.zeros((n_proj, imsize, imsize), dtype=np.float32)
# loop through projections
for i in range(n_proj):
print(i)
# generate filename
filename = (path_projection + filename_projection).format(i)
# load projection
try:
tmp = np.transpose(np.asarray(Image.open(filename), dtype = np.float32))
except:
print('Error reading\n {}\n file.'.format(filename))
raise
# apply flat/ dark filed correction (avoid dividion by 0) and take negative log
denom = flat - dark
nom = tmp - dark
mask = np.logical_and(nom > 0, denom > 0)
proj[i, mask] = -np.log(nom[mask] / denom[mask])
plt.subplot(121)
plt.imshow(proj[0, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('0 projection')
plt.subplot(122)
plt.imshow(proj[-1, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('last projection')
plt.show()
# create AcquisitionGeometry
# this data was acquired over 360 degree rotation,
# the first and the last projections are equal,
# therefore we skip the last projection
ag = AcquisitionGeometry(geom_type = 'parallel',
dimension = '3D',
angles = np.linspace(0, 2*np.pi, n_proj-1, endpoint = False, dtype=np.float32),
pixel_num_h = imsize,
pixel_num_v = imsize,
dimension_labels = ['angle', \
'vertical', \
'horizontal'])
# create AcquisitionData and pass actual data
# again we skip the last projection
ad = ag.allocate()
ad.fill(proj[:-1, :, :])
plt.subplot(121)
plt.imshow(ad.as_array()[0, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('0 projection')
plt.subplot(122)
plt.imshow(ad.as_array()[525, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('525 projection')
plt.show()
# there is quite a lot of empty space around ROI, we can crop data
# to reduce dataset size and speed-up reconstruction
# note, we will crop the data symmetrically to keep geometrical center
# centre of the detector and centre of the projection in the same point
# initialise the processsor
resizer = Resizer(roi=[-1, (200,imsize-200), (450,imsize-450)])
#set the input data
resizer.set_input(ad)
#get the output data
ad = resizer.get_output()
# update acquisition geometry
ag = ad.geometry
plt.subplot(121)
plt.imshow(ad.as_array()[0, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('cropped data, 0 projection')
plt.subplot(122)
plt.imshow(ad.as_array()[525, :, :], cmap = plt.cm.inferno)
plt.colorbar()
plt.title('cropped data, 525 projection')
plt.show()
## calculate centre of rotation
# we will use two levels to calculate for centre of rotation
# to compensate for misalignemnt between axis of rotation
# and detector plane
l1 = 400
l2 = 1200
# we will first reconstruct two slices without compensation for CoR offset
# create AcquisitionGeometry
ag_slice = AcquisitionGeometry(geom_type = 'parallel',
dimension = '2D',
angles = np.linspace(0, 2*np.pi, n_proj-1, endpoint = False, dtype=np.float32),
pixel_num_h = ag.pixel_num_h,
dimension_labels = ['angle', \
'horizontal'])
# Create Image Geometry
ig_slice = ImageGeometry(voxel_num_x=ag.pixel_num_h,
voxel_num_y=ag.pixel_num_h,
voxel_size_x=ag.pixel_size_h,
voxel_size_y=ag.pixel_size_h)
ad_slice_l1 = ag_slice.allocate()
ad_slice_l1.fill(ad.as_array()[:,l1,:])
ad_slice_l2 = ag_slice.allocate()
ad_slice_l2.fill(ad.as_array()[:,l2,:])
# initialise the processsor
fbp = FBP(ig_slice, ag_slice, device='cpu')
# set the input data
fbp.set_input(ad_slice_l1)
fbp.process()
# get the output data
FBP_l1 = fbp.get_output()
# set the input data
fbp.set_input(ad_slice_l2)
fbp.process()
# get the output data
FBP_l2 = fbp.get_output()
plt.subplot(121)
plt.imshow(FBP_l1.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, no CoR compensation'.format(l1))
plt.subplot(122)
plt.imshow(FBP_l2.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, no CoR compensation'.format(l2))
plt.show()
# use processor CenterOfRotationFinder to calculate cetre of rotation
# Note, the processor requires 0-180 degree acuqisition
ag_slice_180 = AcquisitionGeometry(geom_type = 'parallel',
dimension = '2D',
angles = np.linspace(0, 2*np.pi, (n_proj-1)//2, endpoint = False, dtype=np.float32),
pixel_num_h = ag.pixel_num_h,
dimension_labels = ['angle', \
'horizontal'])
ad_slice_180_l1 = ag_slice_180.allocate()
ad_slice_180_l1.fill(ad.as_array()[:524, l1, :])
ad_slice_180_l2 = ag_slice_180.allocate()
ad_slice_180_l2.fill(ad.as_array()[:524, l2, :])
cor = CenterOfRotationFinder()
cor.set_input(ad_slice_180_l1)
centre_l1 = cor.get_output()
cor = CenterOfRotationFinder()
cor.set_input(ad_slice_180_l2)
centre_l2 = cor.get_output()
# to compensate for misalignment, we will apply
# translation and rotation to the dataset
# first we will calculate necessary geometrical parameters
# calculate rotation angle
rot_angle = np.arcsin((centre_l2 - centre_l1) / (np.sqrt((centre_l2 - centre_l1) ** 2 + (l2 - l1) ** 2)))
# and offset
offset = centre_l1 - np.tan(-rot_angle) * (ag.pixel_num_v / 2 - l1) - ag.pixel_num_h / 2
# we will first translate axis of the rotation to have the pivot point in the geometrical centre of the detector
trans_ad = ag.allocate()
trans_ad.fill(scipy.ndimage.interpolation.shift(ad.as_array(), (0,0,-offset), order=1, mode='nearest'))
# and then rotate projections
rot_ad = ag.allocate()
rot_ad.fill(scipy.ndimage.interpolation.rotate(trans_ad.as_array(), -rot_angle*180/np.pi, reshape=False, axes=(1,2), order=1, mode='reflect'))
# to visualise effect of translation and rotation,
# we will reconstruct a couple of slices again
ad_slice_l1 = ag_slice.allocate()
ad_slice_l1.fill(trans_ad.as_array()[:,l1,:])
ad_slice_l2 = ag_slice.allocate()
ad_slice_l2.fill(trans_ad.as_array()[:,l2,:])
# initialise the processsor
fbp = FBP(ig_slice, ag_slice, device='cpu')
# set the input data
fbp.set_input(ad_slice_l1)
fbp.process()
# get the output data
FBP_l1 = fbp.get_output()
# set the input data
fbp.set_input(ad_slice_l2)
fbp.process()
# get the output data
FBP_l2 = fbp.get_output()
plt.subplot(121)
plt.imshow(FBP_l1.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, only translation'.format(l1))
plt.subplot(122)
plt.imshow(FBP_l2.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, only translation'.format(l2))
plt.show()
# to visualise effect of translation and rotation,
# we will reconstruct a couple of slices again
ad_slice_l1 = ag_slice.allocate()
ad_slice_l1.fill(rot_ad.as_array()[:,l1,:])
ad_slice_l2 = ag_slice.allocate()
ad_slice_l2.fill(rot_ad.as_array()[:,l2,:])
# initialise the processsor
fbp = FBP(ig_slice, ag_slice, device='cpu')
# set the input data
fbp.set_input(ad_slice_l1)
fbp.process()
# get the output data
FBP_l1 = fbp.get_output()
# set the input data
fbp.set_input(ad_slice_l2)
fbp.process()
# get the output data
FBP_l2 = fbp.get_output()
plt.subplot(121)
plt.imshow(FBP_l1.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, translation and rotation'.format(l1))
plt.subplot(122)
plt.imshow(FBP_l2.as_array(), cmap = plt.cm.inferno)
plt.colorbar()
plt.title('slice {}, translation and rotation'.format(l2))
plt.show()
# finally reconstruct full 3D volume
# reorder the data axes to prepare the data for the ASTRA operators.
data = rot_ad.subset(dimensions=['vertical','angle','horizontal'])
# Create Image Geometry
ig_full = ImageGeometry(voxel_num_x=ag.pixel_num_h,
voxel_num_y=ag.pixel_num_h,
voxel_num_z=ag.pixel_num_v,
voxel_size_x=ag.pixel_size_h,
voxel_size_y=ag.pixel_size_h,
voxel_size_z=ag.pixel_size_v)
# initialise the processsor
fbp = FBP(ig_full, ag, device='gpu')
# set the input data
fbp.set_input(data)
fbp.process()
# get the output data
FBP_output = fbp.get_output()
plt.subplot(221)
plt.imshow(FBP_output.as_array()[200, :, :])
plt.title('slice 200')
plt.subplot(222)
plt.imshow(FBP_output.as_array()[400, :, :])
plt.title('slice 400')
plt.subplot(223)
plt.imshow(FBP_output.as_array()[600, :, :])
plt.title('slice 600')
plt.subplot(224)
plt.imshow(FBP_output.as_array()[800, :, :])
plt.title('slice 800')
plt.show()
```
| github_jupyter |
# Deep Convolutional Generative Adversarial Network
## Import dependencies
```
import tensorflow as tf
import numpy as np
tf.__version__
np.__version__
```
## Load in data
```
from tensorflow.examples.tutorials.mnist import input_data
data = input_data.read_data_sets('data/MNIST/', one_hot=True)
print('Training: {:,}'.format(data.train.num_examples))
print('Testing: {:,}'.format(data.test.num_examples))
print('Validation: {:,}'.format(data.validation.num_examples))
```
## Define hyperparameters
```
# Inputs
image_size = 28
image_channel = 1
image_shape = (image_size, image_size, image_channel)
image_size_flat = image_size * image_size * image_channel
num_classes = 10
# Network
filter_size = 5
hidden1_filter = 32
hidden2_filter = 64
fc1_size = 1024
fc2_size = 1
dropout = 0.8
```
## Helper functions
### `weights` and `biases`
```
def weight(shape, name):
# initial = tf.truncated_normal(shape=shape, mean=0, stddev=0.02)
# return tf.Variable(initial, name='weight')
return tf.get_variable(name, [5, 5, 1, 32], initializer=tf.truncated_normal_initializer(stddev=0.02))
def bias(shape, name):
# initial = tf.zeros(shape=[shape])
# return tf.Variable(initial, name='bias')
return tf.get_variable(name, [32], initializer=tf.constant_initializer(0))
```
### `convolution` and `pooling`
```
def conv2d(X, W, strides=[1, 1, 1, 1]):
return tf.nn.conv2d(X, W, strides=strides, padding='SAME')
def max_pool(X):
return tf.nn.max_pool(X, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='SAME')
```
### `flatten` layer
```
def flatten(layer):
layer_shape = layer.get_shape()
num_features = np.array(layer_shape[1:4], dtype=int).prod()
layer_flat = tf.reshape(layer, [-1, num_features])
return layer_flat, num_features
```
### Layers
```
# convolutional layer
def conv_layer(prev_layer, prev_filter, layer_filter, layer_name,
strides=[1, 1, 1, 1], filter_size=5, use_pool=True, batch_norm=False):
# with tf.name_scope(layer_name):
W = weight(shape=[filter_size, filter_size, prev_filter, layer_filter], name=layer_name+'_W')
b = bias(shape=layer_filter, name=layer_name + '_b')
layer = conv2d(prev_layer, W, strides=strides) + b
if batch_norm:
layer = tf.contrib.layers.batch_norm(layer, epsilon=1e-3)
if use_pool:
layer = max_pool(layer)
layer = tf.nn.relu(layer)
return layer
# fully connected layer
def fc_layer(prev_layer, prev_size, layer_size, layer_name,
use_relu=True, dropout=False, batch_norm=False):
# with tf.name_scope(layer_name):
W = weight(shape=[prev_size, layer_size], name=layer_name + '_W')
b = bias(shape=layer_size, name=layer_name + '_b')
layer = tf.matmul(prev_layer, W) + b
# Use batch normalization
if batch_norm:
layer = tf.contrib.layers.batch_norm(layer, epsilon=1e-5)
# Use ReLU
if use_relu:
layer = tf.nn.relu(layer)
# Add dropout
if dropout:
layer = tf.nn.dropout(layer, keep_prob)
return layer
# Output layers
def output_layer(fc_layer, fc_size, num_classes):
# with tf.name_scope('output_layer'):
W = weight(shape=[fc_size, num_classes], name='output_layer_W')
b = bias(shape=num_classes, name='output_layer_b')
logits = tf.matmul(fc_layer, W) + b
y_pred = tf.nn.softmax(logits)
y_pred_true = tf.argmax(y_pred, axis=1)
return logits, y_pred_true
```
### Discrimitor (Convolutional NN)
```
def discriminator(X_image, reuse=False):
if reuse:
tf.get_variable_scope().reuse_variables()
# Convolutional blocks
hidden1 = conv_layer(X_image, image_channel, hidden1_filter, 'dis_hidden1', use_pool=True)
hidden2 = conv_layer(hidden1, hidden1_filter, hidden2_filter, 'dis_hidden2', use_pool=True)
# Fully connected blocks
hidden2_flat, hidden2_flat_filters = flatten(hidden2)
fc1_layer = fc_layer(hidden2_flat, hidden2_flat_filters, fc1_size, 'dis_fc1', use_relu=True)
fc2_layer = fc_layer(fc1_layer, fc1_size, fc2_size, 'dis_fc2', use_relu=False)
return fc2_layer
```
### Generator (Deconvolutional NN)
```
def generator(batch_size, z_dim, up_scale=56):
z = tf.truncated_normal(shape=[batch_size, z_dim], mean=0, stddev=1, name='z')
scale = up_scale * up_scale
# Fully connected block
fc1_layer = fc_layer(z, z_dim, scale, 'gen_fc1', use_relu=True, batch_norm=True)
fc1_layer = tf.reshape(fc1_layer, [-1, up_scale, up_scale, image_channel])
# Convolutional block
hidden1 = conv_layer(fc1_layer, image_channel, z_dim/2, 'gen_deconv1',
strides=[1, 2, 2, 1], filter_size=3, use_pool=False, batch_norm=True)
hidden1 = tf.image.resize_images(hidden1, size=[up_scale, up_scale])
hidden2 = conv_layer(hidden1, z_dim/2, z_dim/4, 'gen_deconv2',
strides=[1, 2, 2, 1], filter_size=3, use_pool=False, batch_norm=True)
hidden2 = tf.image.resize_images(hidden2, size=[up_scale, up_scale])
hidden3 = conv_layer(hidden2, z_dim/4, 1, 'gen_deconv3',
strides=[1, 2, 2, 1], filter_size=1, use_pool=False, batch_norm=False)
hidden3 = tf.nn.sigmoid(hidden3)
# No batch normalization in the final layer but we add sigmoid activation
# to make the generated images more compact/crisper. (batch_size, 28, 28, 1)
return hidden3
```
## Building the Network
```
sess = tf.Session()
batch_size = 50
z_dimensions = 100
X_placeholder = tf.placeholder('float', shape=[None, 28, 28, 1], name='X_placeholder')
Gz = generator(batch_size, z_dimensions) # the generated image
Dx = discriminator(X_placeholder) # classify the real image
Dg = discriminator(Gz, reuse=True) # classify the generated image
```
### Loss function
#### generator's loss
The generator wants to optimize the probability that **the generated image is rated highly** i.e _for every image it generates, the discriminator should say it's real_
#### discriminator's loss
The discriminator wants to:
* optimize the probability that **the real data is rated highly**
* optimize the probability that **the generated data is rated poorly**
That is, when it classifies images from the training data, it should say this is the real image and when it classifies images from the generated data, it should say this is the fake image
```
# generator's loss
g_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=Dg, labels=tf.ones_like(Dg)))
# discriminator's loss
d_loss_real = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=Dx, labels=tf.ones_like(Dx)))
d_loss_fake = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=Dg, labels=tf.zeros_like(Dg)))
d_loss = d_loss_real + d_loss_fake
```
| github_jupyter |
<div style="float: right; margin: 0px 0px 0px 0px"><img src="images/clusters.png" width="400px"></div>
# Clustering: Picking the 'K' hyperparameter
The unsupervised machine learning technique of clustering data into similar groups can be useful and fairly efficient in most cases. The big trick is often how you pick the number of clusters to make (the K hyperparameter).
The number of clusters may vary dramatically depending on the characteristics of the data, the different types of variables (numeric or categorical), how the data is normalized/encoded and the distance metric used.
<div style="float: left; margin: 20px 50px 20px 20px"><img src="images/picking.png" width="100px"></div>
**For this notebook we're going to focus specifically on the following:**
- Optimizing the number of clusters (K hyperparameter) using Silhouette Scoring
- Utilizing an algorithm (DBSCAN) that automatically determines the number of clusters
### Software
- Bro Analysis Tools (BAT): https://github.com/Kitware/bat
- Pandas: https://github.com/pandas-dev/pandas
- Scikit-Learn: http://scikit-learn.org/stable/index.html
<div style="float: right; margin: 20px 20px 20px 20px"><img src="images/scikit.png" width="200px"></div>
### Techniques
- One Hot Encoding: http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html
- t-SNE: https://distill.pub/2016/misread-tsne/
- Kmeans: http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html
- Silhouette Score: https://en.wikipedia.org/wiki/Silhouette_(clustering)
- DBSCAN: https://en.wikipedia.org/wiki/DBSCAN
```
# Third Party Imports
import pandas as pd
import numpy as np
import sklearn
from sklearn.manifold import TSNE
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.cluster import KMeans, DBSCAN
# Local imports
import bat
from bat.log_to_dataframe import LogToDataFrame
from bat.dataframe_to_matrix import DataFrameToMatrix
# Good to print out versions of stuff
print('BAT: {:s}'.format(bat.__version__))
print('Pandas: {:s}'.format(pd.__version__))
print('Scikit Learn Version:', sklearn.__version__)
# Create a Pandas dataframe from the Bro log
http_df = LogToDataFrame('../data/http.log')
# Print out the head of the dataframe
http_df.head()
```
# Our HTTP features are a mix of numeric and categorical data
When we look at the http records some of the data is numerical and some of it is categorical so we'll need a way of handling both data types in a generalized way. We have a DataFrameToMatrix class that handles a lot of the details and mechanics of combining numerical and categorical data, we'll use below.
<div style="float: right; margin: 10px 40px 10px 40px"><img src="images/transformers.png" width="200px"></div>
## Transformers
**We'll now use the Scikit-Learn tranformer class to convert the Pandas DataFrame to a numpy ndarray (matrix). The transformer class takes care of many low-level details**
* Applies 'one-hot' encoding for the Categorical fields
* Normalizes the Numeric fields
* The class can be serialized for use in training and evaluation
* The categorical mappings are saved during training and applied at evaluation
* The normalized field ranges are stored during training and applied at evaluation
```
# We're going to pick some features that might be interesting
# some of the features are numerical and some are categorical
features = ['id.resp_p', 'method', 'resp_mime_types', 'request_body_len']
# Use the DataframeToMatrix class (handles categorical data)
# You can see below it uses a heuristic to detect category data. When doing
# this for real we should explicitly convert before sending to the transformer.
to_matrix = DataFrameToMatrix()
http_feature_matrix = to_matrix.fit_transform(http_df[features], normalize=True)
print('\nNOTE: The resulting numpy matrix has 12 dimensions based on one-hot encoding')
print(http_feature_matrix.shape)
http_feature_matrix[:1]
# Plotting defaults
%matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams['font.size'] = 12.0
plt.rcParams['figure.figsize'] = 14.0, 7.0
```
<div style="float: right; margin: 10px 40px 10px 40px"><img src="images/silhouette.jpg" width="250px"></div>
# Silhouette Scoring
"The silhouette value is a measure of how similar an object is to its own cluster (cohesion) compared to other clusters (separation). The silhouette ranges from -1 to 1, where a high value indicates that the object is well matched to its own cluster and poorly matched to neighboring clusters. If most objects have a high value, then the clustering configuration is appropriate. If many points have a low or negative value, then the clustering configuration may have too many or too few clusters."
- https://en.wikipedia.org/wiki/Silhouette_(clustering)
```
from sklearn.metrics import silhouette_score
scores = []
clusters = range(2,20)
for K in clusters:
clusterer = KMeans(n_clusters=K)
cluster_labels = clusterer.fit_predict(http_feature_matrix)
score = silhouette_score(http_feature_matrix, cluster_labels)
scores.append(score)
# Plot it out
pd.DataFrame({'Num Clusters':clusters, 'score':scores}).plot(x='Num Clusters', y='score')
```
## Silhouette graphs shows that 10 is the 'optimal' number of clusters
- 'Optimal': Human intuition and clustering involves interpretation/pattern finding and is often partially subjective :)
- For large datasets running an exhaustive search can be time consuming
- For large datasets you can often get a large K using max score, so pick the 'knee' of the graph as your K
```
# So we know that the highest (closest to 1) silhouette score is at 10 clusters
kmeans = KMeans(n_clusters=10).fit_predict(http_feature_matrix)
# TSNE is a great projection algorithm. In this case we're going from 12 dimensions to 2
projection = TSNE().fit_transform(http_feature_matrix)
# Now we can put our ML results back onto our dataframe!
http_df['cluster'] = kmeans
http_df['x'] = projection[:, 0] # Projection X Column
http_df['y'] = projection[:, 1] # Projection Y Column
# Now use dataframe group by cluster
cluster_groups = http_df.groupby('cluster')
# Plot the Machine Learning results
colors = {-1:'black', 0:'green', 1:'blue', 2:'red', 3:'orange', 4:'purple', 5:'brown', 6:'pink', 7:'lightblue', 8:'grey', 9:'yellow'}
fig, ax = plt.subplots()
for key, group in cluster_groups:
group.plot(ax=ax, kind='scatter', x='x', y='y', alpha=0.5, s=250,
label='Cluster: {:d}'.format(key), color=colors[key])
# Now print out the details for each cluster
pd.set_option('display.width', 1000)
for key, group in cluster_groups:
print('\nCluster {:d}: {:d} observations'.format(key, len(group)))
print(group[features].head(3))
```
<div style="float: right; margin: 30px 20px 20px 20px"><img src="images/no_hands.jpg" width="250px"></div>
# Look Ma... No K!
### DBSCAN
Density-based spatial clustering is a data clustering algorithm that given a set of points in space, groups points that are closely packed together and marking low-density regions as outliers.
- You don't have to pick K
- There are other hyperparameters (eps and min_samples) but defaults often work well
- https://en.wikipedia.org/wiki/DBSCAN
- Hierarchical version: https://github.com/scikit-learn-contrib/hdbscan
```
# Now try DBScan
http_df['cluster_db'] = DBSCAN().fit_predict(http_feature_matrix)
print('Number of Clusters: {:d}'.format(http_df['cluster_db'].nunique()))
# Now use dataframe group by cluster
cluster_groups = http_df.groupby('cluster_db')
# Plot the Machine Learning results
fig, ax = plt.subplots()
for key, group in cluster_groups:
group.plot(ax=ax, kind='scatter', x='x', y='y', alpha=0.5, s=250,
label='Cluster: {:d}'.format(key), color=colors[key])
```
<div style="float: right; margin: 20px 20px 20px 20px"><img src="images/magic.jpg" width="300px"></div>
# DBSCAN automagically determined 10 clusters!
So obviously we got a bit lucky here and for different datasets with different feature distributions DBSCAN may not give you the optimal number of clusters right off the bat. There are two hyperparameters that can be tweeked but like we said the defaults often work well. See the DBSCAN and Hierarchical DBSCAN links for more information.
- https://en.wikipedia.org/wiki/DBSCAN
- Hierarchical version: https://github.com/scikit-learn-contrib/hdbscan
<div style="float: right; margin: 50px 0px 0px 0px"><img src="https://www.kitware.com/img/small_logo_over.png"></div>
## Wrap Up
Well that's it for this notebook, given the usefulness and relatively efficiency of clustering it a good technique to include in your toolset. Understanding the K hyperparameter and how to determine optimal K (or not if you're using DBSCAN) is a good trick to know.
If you liked this notebook please visit the [bat](https://github.com/Kitware/bat) project for more notebooks and examples.
| github_jupyter |
```
import json
import os
import glob
import shutil
# 画像関係
import numpy as np
import cv2
from PIL import Image
# 画像表示
import matplotlib.pyplot as plt
IMAGE_SIZE = 256
```
# データの確認
```
# データのリスト
json_list = glob.glob('seg_dogs/*.json')
img_list = [f.replace('json', 'jpg') for f in json_list]
print(len(json_list))
no = 1
# アノテーションデータ読み込み
with open(json_list[no]) as f:
data = json.loads(f.read())
# 1つだけ取り出す
shape = data['shapes'][0]
label = shape['label']
points = shape['points']
shape_type = shape['shape_type']
print('[label]', label)
print('[shape_type]', shape_type)
print('[points]', points)
```
# データの変換
```
# 画像読み込み
img = cv2.imread(img_list[no])
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# アノテーション部分
mask = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
mask = cv2.fillPoly(mask, np.int32([points]), 1)
# 横並びに表示
fig = plt.figure(figsize=(12, 6))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
ax1.imshow(img)
ax2.imshow(mask, cmap='gray')
```
# データの保存
```
# フォルダ作成 trainとvalにデータを分けます
train_dir = 'train'
val_dir = 'val'
if not os.path.exists(train_dir):
os.mkdir(train_dir)
os.mkdir(train_dir + '/images')
os.mkdir(train_dir + '/masks')
if not os.path.exists(val_dir):
os.mkdir(val_dir)
os.mkdir(val_dir + '/images')
os.mkdir(val_dir + '/masks')
# 114個のデータを用意したので 100 と 14 に分けます
for ind, file in enumerate(json_list):
points = []
with open(file) as f:
data = json.loads(f.read())
for s in data['shapes']:
points.append(s['points'])
if points:
# 画像データを読み込み画像サイズ取得
img_path = file.replace('json', 'jpg')
img = cv2.imread(img_path)
# ファイル名
file_name = os.path.basename(img_path)
# jsonのアノテーションデータ
# 犬:1
# 背景:0
mask = np.zeros((img.shape[0], img.shape[1]), dtype=np.uint8)
for p in points:
mask = cv2.fillPoly(mask, np.int32([p]), 1)
# リサイズ
img = cv2.resize(img, (IMAGE_SIZE, IMAGE_SIZE), interpolation=cv2.INTER_NEAREST)
mask = cv2.resize(mask, (IMAGE_SIZE, IMAGE_SIZE), interpolation=cv2.INTER_NEAREST)
# 保存
file_name = file_name.replace('jpg', 'png')
if ind<100:
maskim = Image.fromarray(np.uint8(mask))
maskim.save(f'train/masks/{file_name}')
cv2.imwrite(f'train/images/{file_name}', img)
else:
maskim = Image.fromarray(np.uint8(mask))
maskim.save(f'val/masks/{file_name}')
cv2.imwrite(f'val/images/{file_name}', img)
```
| github_jupyter |
# Dragon Real Estate-Price Predictator
```
import pandas as pd
housing = pd.read_csv("data.csv")
housing.head()
housing.info()
housing['CHAS'].value_counts()
housing.describe()
housing.corr()
%matplotlib inline
#for ploting histogram
# import matplotlib.pyplot as plt
# housing.hist(bins=50,figsize=(20,15))
```
# #Train-Test Splitting
```
#for learning purpose
# import numpy as np
# def split_train_test(data,test_ratio):
# np.random.seed(42)
# shuffled = np.random.permutation(len(data))
# print(shuffled)
# test_set_size = int(len(data)*test_ratio)
# test_indices=shuffled[:test_set_size]
# train_indices=shuffled[test_set_size:]
# return data.iloc[train_indices],data.iloc[test_indices]
# train_set, test_set =split_train_test(housing,0.2)
# print(f"Rows in train set: {len(train_set)}\nRows in test set: {len(test_set)}\n")
from sklearn.model_selection import train_test_split
train_set,test_set = train_test_split(housing,test_size=0.2,random_state=42)
print(f"Rows in train set: {len(train_set)}\nRows in test set: {len(test_set)}\n")
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1,test_size=0.2,random_state=42)
for train_index,test_index in split.split(housing,housing['CHAS']):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
strat_test_set.describe()
strat_train_set['CHAS'].value_counts()
strat_test_set['CHAS'].value_counts()
# 376/28
# 95/7
housing = strat_train_set.copy()
```
# Lokking For Correlation
```
corr_matrix = housing.corr()
corr_matrix['MEDV'].sort_values(ascending=False)
from pandas.plotting import scatter_matrix
attributes = ["MEDV","RM","ZN","LSTAT"]
scatter_matrix(housing[attributes],figsize =(12,8))
housing.plot(kind="scatter",x="RM",y="MEDV",alpha=0.8)
# housing.plot(kind="scatter",x="LSTAT",y="MEDV",alpha=0.8)
# housing.plot(kind="scatter",x="LSTAT",y="RM",alpha=0.8)
```
# Trying out ATTRIBUTES COMBINATION
```
housing["TAXRM"] = housing["TAX"]/housing["RM"]
housing["TAXRM"]
housing.head()
corr_matrix = housing.corr()
corr_matrix['MEDV'].sort_values(ascending=False)
housing.plot(kind="scatter",x="TAXRM",y="MEDV",alpha=0.8)
housing = strat_train_set.drop("MEDV",axis=1)
housing_labels = strat_train_set["MEDV"].copy()
```
## missing attributes
```
# To take a missing attributes,you have three option:
# 1. Get rid of the missing data points
# 2. Get rid of the all atribute
# 3. Set the value to some value(0,mean,median)
a = housing.dropna(subset=["RM"])#option 1
a.shape
# Note that their is no RM missing data points and also note That and the orignal dataframe will remain unchanged
# a
housing.drop("RM",axis=1).shape#option2
# Note that their is no RM column and also note That and the orignal dataframe will remain unchanged
median=housing["RM"].median() # Compute median for option 3
housing["RM"].fillna(median)
housing.shape
housing.describe()# before we started filling missing attributes
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy = "median")
imputer.fit(housing)
imputer.statistics_
x = imputer.transform(housing)
housing_tr = pd.DataFrame(x,columns=housing.columns)
housing_tr.describe()
```
# Scikit_learn Design
Primarily,three types of objects
1.Estimators-It estimates some parameter based on a dataset. Eg .Importer, It has a fit method and Transform method.
fit method - Fits the data sets and calcultes the internal parameters
2.Transformers - transform method take input and returns output based on the learnings from fit ().It also has convience
function called fit_transform(),Which fits and then transforms.
3.Predictors - linearRegression model is an example of predictor. fit() and predict() are two common functions. It also gives
score() function which will evaluate the predictions.
# Feature Scaling
```
# # Primarlly , two types of feature Scalling methods:
# 1.Min-max scalling (Normalisation)
# (value-min)/(max-min)
# sklearn provides a class called MinMaxScaler for this
# 2.Standardization
# (value-mean)/std
# sklearn provides a class called Standard Scaler for this
```
# Creating a pipline
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
my_pipeline = Pipeline([
('imputer',SimpleImputer(strategy='median')),
# .....add as many as you want in your pipeline
('std_scaler', StandardScaler()),
])
housing_num_tr = my_pipeline.fit_transform(housing)
housing_num_tr.shape
```
# Selecting a desired model for dragon Real Estates
```
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
# model= LinearRegression()
# model = DecisionTreeRegressor()
model = RandomForestRegressor()
model.fit(housing_num_tr,housing_labels)
some_data = housing.iloc[:5]
some_labels = housing_labels.iloc[:5]
prepared_data = my_pipeline.transform(some_data)
model.predict(prepared_data)
list(some_labels)
```
# Evaluating the model
```
import numpy as np
from sklearn.metrics import mean_squared_error
housing_predictions = model.predict(housing_num_tr)
mse = mean_squared_error(housing_labels,housing_predictions)
rmse = np.sqrt(mse)
rmse
```
# Using better evaluation technique-Cross validation
```
# 1 2 3 4 5 6 7 8 9 10
from sklearn.model_selection import cross_val_score
scores = cross_val_score(model,housing_num_tr,housing_labels,scoring="neg_mean_squared_error",cv=10)
rmse_scores = np.sqrt(-scores)
rmse_scores
def print_scores(scores):
print("Scores", scores)
print("Mean: ", scores.mean())
print("Standard deviation: ", scores.std())
print_scores(rmse_scores)
```
# Saving The Model on test data
```
from joblib import dump, load
dump(model,'Dragon.joblib')
```
## Testing the model on test data
```
X_test = strat_test_set.drop("MEDV",axis=1)
Y_test = strat_test_set["MEDV"].copy()
X_test_prepared = my_pipeline.transform(X_test)
final_predictions = model.predict(X_test_prepared)
final_mse = mean_squared_error(Y_test,final_predictions)
final_rmse = np.sqrt(final_mse)
# print(final_predictions,list(Y_test))
final_rmse
prepared_data[0]
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/reservoir-engineering/blob/master/Reservoir%20Simulation%20Ertekin/Unit%205%20Finite-Difference%20Approximation%20to%20Linear-Flow%20Equations/fd1d_linearflow_explicit.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Unit 5 Finite-Difference Approximations to Linear-Flow Problems**
## Explicit Formulation (*Forward-Difference Approximation*)
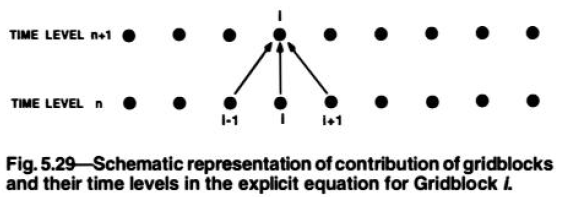
$$p_{i}^{n+1} = p^i_n + (\frac{\alpha_c B_l \Delta t}{V_b \phi c_l})_i \cdot q_{lsc_i} + (\frac{\alpha_c B_l \Delta t}{V_b \phi c_l})_i \cdot [T_{lx_{i+1/2}}^{n} p_{i+1}^{n}-(T_{lx_{i+1/2}}^{n} + T_{lx_{i-1/2}}^{n})p_{i}^{n}+T_{lx_{i-1/2}}p_{i-1}^{n}]$$
In the following codes, $(\frac{\alpha_c B_l \Delta t}{V_b \phi c_l})_i$ will be written as `factor`
Transmissibility of coupling cells, $T_{lx_{i\pm1/2}}^{n}$, written as `T_min_half` or `T_plus_half`
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
!git clone https://github.com/yohanesnuwara/reservoir-engineering
```
## Example 5.8 Five reservoir gridblocks with no flow at both boundaries
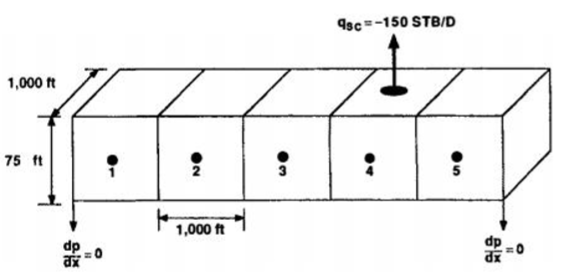
### For timestep 10 days
```
"Task. determine pressure distribution during first year of production with timestep 10 days"
# known
p_initial = 6000 # initial pressure, in psia
delta_x = 1000 # ft
delta_y = 1000
delta_z = 75
ngrid = 5
grid_loc = 4 # grid location where production well is located
B = 1 # phase FVF, assumed constant over pressure, rb/stb
c = 3.5E-06 # phase compressibility, psi^-1
k_x = 15 # perm in x direction, md
poro = 0.18
mu = 10 # phase viscosity, cp
delta_t = 10 # days
qsc = -150 # minus means production, stb/d
# conversion
k_x = 15 * 1E-03 # darcy to mD
# calculate factor
Vb = delta_x * delta_y * delta_z
alpha = 5.615 # volume conversion factor, is a constant
factor = (alpha * B * delta_t) / (Vb * poro * c)
factor
```
Because of uniform gridblocks, the equation for `tr_plus_coupling` and `tr_min_coupling` become:
$$T_{lx_{i+1/2}}^{n} = T_{lx_{i-1/2}}^{n} = (\beta_c \frac{A_x k_x}{\mu_l B_l \Delta x})_{i+1/2}^{n} = (\beta_c \frac{A_x k_x}{\mu_l B_l \Delta x})_{i-1/2}^{n}$$
```
# calculate transmissibility of coupling cells
beta = 1.127 # transmissibility conversion factor, is a constant
Ax = delta_y * delta_z
T_plus_half = beta * ((Ax * k_x) / (mu * B * delta_x))
T_min_half = T_plus_half
T_min_half, T_plus_half
```
There are 5 grids (`grid_1`, `grid_2`, `grid_3`, `grid_4`, `grid_5`), so the transmissibility coefficients will be: $$T_{lx_{i+1/2}}^{n}=(T_{lx_{1+1/2}}^{n}, T_{lx_{2+1/2}}^{n}, T_{lx_{3+1/2}}^{n}, T_{lx_{4+1/2}}^{n}, T_{lx_{5+1/2}}^{n})$$
and
$$T_{lx_{i-1/2}}^{n}=(T_{lx_{1/2}}^{n}, T_{lx_{1+1/2}}^{n}, T_{lx_{2+1/2}}^{n}, T_{lx_{3+1/2}}^{n}, T_{lx_{4+1/2}}^{n})$$
The values of $T_{lx_{5+1/2}}^{n}=0$ and $T_{lx_{1/2}}^{n}=0$ (the edge of gridblocks), so the values of each are:
$$T_{lx_{i+1/2}}^{n}=(0.127, 0.127, 0.127, 0.127, 0)$$
and
$$T_{lx_{i-1/2}}^{n}=(0, 0.127, 0.127, 0.127, 0.127)$$
```
q = np.full(ngrid-1, T_min_half)
Ti_plus_halves = np.append(q, [0])
print(Ti_plus_halves)
p = np.full(ngrid-1, T_plus_half)
Ti_min_halves = np.append([0], p)
print(Ti_min_halves)
print("At grid 1, the coupling transmissibility coeffs are:", Ti_min_halves[0], "for T_min_half and:", Ti_plus_halves[0], "for T_plus_half.")
print("At grid 3, the coupling transmissibility coeffs are:", Ti_min_halves[2], "for T_min_half and:", Ti_plus_halves[2], "for T_plus_half.")
print("At grid 5, the coupling transmissibility coeffs are:", Ti_min_halves[4], "for T_min_half and:", Ti_plus_halves[4], "for T_plus_half.")
```
Requires array for $q_{sc}$
```
qsc = [0, 0, 0, -150, 0] # production well in grid 4
```
Calculate $p_{i}^{n+1}$ for each grid in each time
```
pi = np.full(ngrid, p_initial) # array of pressure in each grid [6000, 6000, 6000, 6000, 6000]
time = np.arange(15, 370, delta_t)
pi_arr = []
min_arr = []
plus_arr = []
for j in range(len(time)):
pnew_arr = []
minus = pi[0]
plus = pi[-1]
minus_arr = []
plusus_arr = []
for i, obj in enumerate(pi):
if i > 0:
minus = pi[i-1]
if i < (len(pi) - 1):
plus = pi[i+1]
pnew = pi[i] + (factor * ((Ti_plus_halves[i] * plus) - ((Ti_plus_halves[i] + Ti_min_halves[i]) * pi[i]) + (Ti_min_halves[i] * minus))) + (factor * qsc[i])
pnew_arr.append(float(pnew))
minus_arr.append(float(minus))
plusus_arr.append(float(plus))
pi = pnew_arr
min_arr.append(minus_arr)
plus_arr.append(plusus_arr)
pi_arr.append(pi)
df = pd.DataFrame.from_records(pi_arr)
df = pd.DataFrame(pd.np.column_stack([time, df]), columns=['time', 'grid 1', 'grid 2', 'grid 3', 'grid 4', 'grid 5'])
df
```
### For timestep 15 days
```
delta_t = 15 # days
# calculate factor
Vb = delta_x * delta_y * delta_z
alpha = 5.615 # volume conversion factor, is a constant
factor = (alpha * B * delta_t) / (Vb * poro * c)
pi = np.full(ngrid, 6000) # array of pressure in each grid [6000, 6000, 6000, 6000, 6000]
time = np.arange(15, 370, delta_t)
pi_arr = []
min_arr = []
plus_arr = []
for j in range(len(time)):
pnew_arr = []
minus = pi[0]
plus = pi[-1]
minus_arr = []
plusus_arr = []
for i, obj in enumerate(pi):
if i > 0:
minus = pi[i-1]
if i < (len(pi) - 1):
plus = pi[i+1]
pnew = pi[i] + (factor * ((Ti_plus_halves[i] * plus) - ((Ti_plus_halves[i] + Ti_min_halves[i]) * pi[i]) + (Ti_min_halves[i] * minus))) + (factor * qsc[i])
pnew_arr.append(float(pnew))
minus_arr.append(float(minus))
plusus_arr.append(float(plus))
pi = pnew_arr
min_arr.append(minus_arr)
plus_arr.append(plusus_arr)
pi_arr.append(pi)
df = pd.DataFrame.from_records(pi_arr)
df = pd.DataFrame(pd.np.column_stack([time, df]), columns=['time', 'grid 1', 'grid 2', 'grid 3', 'grid 4', 'grid 5'])
df
```
## Example 5.9 Similar to Ex 5.8, but **constant pressure at left boundary of reservoir**
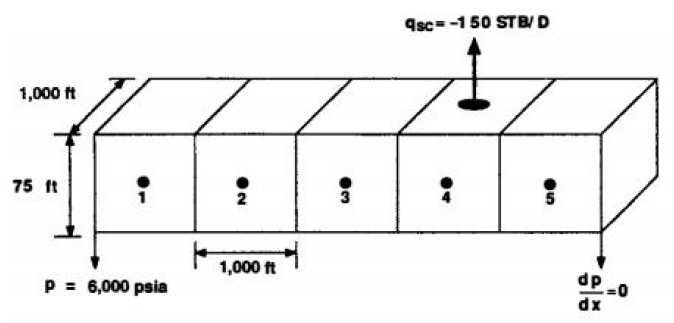
The only modification is in the line:
```
pnew = pi[i] + (factor * ((Ti_plus_halves[i] * plus) - ((Ti_plus_halves[i] + Ti_min_halves[i]) * pi[i]) + (Ti_min_halves[i] * minus))) + (factor * qsc[i])
pnew_arr.append(float(pnew))
pnew_arr[0] = pi[0]
```
Where `pnew_arr[0] = pi[0]` means the pressure in the first gridblock is kept constant, equals to initial pressure `pi` 6000 psi.
### For timestep 15 days
```
pi = np.full(ngrid, p_initial) # array of pressure in each grid [6000, 6000, 6000, 6000, 6000]
time = np.arange(15, 370, delta_t)
pi_arr = []
min_arr = []
plus_arr = []
for j in range(len(time)):
pnew_arr = []
plus = pi[-1]
minus_arr = []
plusus_arr = []
for i, obj in enumerate(pi):
if i > 0:
minus = pi[i-1]
if i < (len(pi) - 1):
plus = pi[i+1]
pnew = pi[i] + (factor * ((Ti_plus_halves[i] * plus) - ((Ti_plus_halves[i] + Ti_min_halves[i]) * pi[i]) + (Ti_min_halves[i] * minus))) + (factor * qsc[i])
pnew_arr.append(float(pnew))
pnew_arr[0] = pi[0]
minus_arr.append(float(minus))
plusus_arr.append(float(plus))
pi = pnew_arr
min_arr.append(minus_arr)
plus_arr.append(plusus_arr)
pi_arr.append(pi)
df = pd.DataFrame.from_records(pi_arr)
df = pd.DataFrame(pd.np.column_stack([time, df]), columns=['time', 'grid 1', 'grid 2', 'grid 3', 'grid 4', 'grid 5'])
df
```
## Example 5.10 Halving the grid spacing of Ex 5.9 (more refined grids)
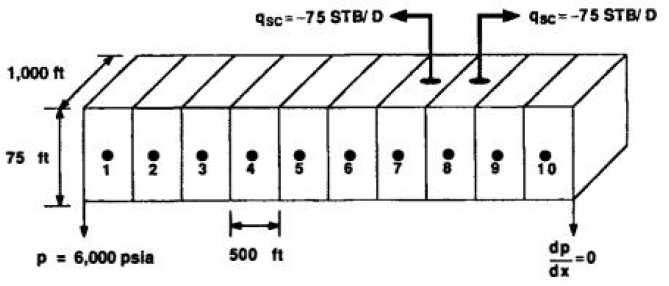
### For timestep 15 days
```
# known
p_initial = 6000 # initial pressure, in psia
delta_x = 500 # ft, half the previous one
delta_y = 1000
delta_z = 75
ngrid = 10 # now twice the number of previous gridblocks
grid_loc = 4 # grid location where production well is located
B = 1 # phase FVF, assumed constant over pressure, rb/stb
c = 3.5E-06 # phase compressibility, psi^-1
k_x = 15 # perm in x direction, md
poro = 0.18
mu = 10 # phase viscosity, cp
delta_t = 15 # days
qsc = -150 # minus means production, stb/d
# conversion
k_x = 15 * 1E-03 # darcy to mD
# calculate factor
Vb = delta_x * delta_y * delta_z
alpha = 5.615 # volume conversion factor, is a constant
factor = (alpha * B * delta_t) / (Vb * poro * c)
factor
# calculate transmissibility of coupling cells
beta = 1.127 # transmissibility conversion factor, is a constant
Ax = delta_y * delta_z
T_plus_half = beta * ((Ax * k_x) / (mu * B * delta_x))
T_min_half = T_plus_half
T_min_half, T_plus_half
q = np.full(ngrid-1, T_min_half)
Ti_plus_halves = np.append(q, [0])
print(Ti_plus_halves)
p = np.full(ngrid-1, T_plus_half)
Ti_min_halves = np.append([0], p)
print(Ti_min_halves)
print("At grid 1, the coupling transmissibility coeffs are:", Ti_min_halves[0], "for T_min_half and:", Ti_plus_halves[0], "for T_plus_half.")
print("At grid 3, the coupling transmissibility coeffs are:", Ti_min_halves[2], "for T_min_half and:", Ti_plus_halves[2], "for T_plus_half.")
print("At grid 5, the coupling transmissibility coeffs are:", Ti_min_halves[4], "for T_min_half and:", Ti_plus_halves[4], "for T_plus_half.")
qsc = [0, 0, 0, 0, 0, 0, -75, -75, 0, 0] # production well in grid 7 and 8
pi = np.full(ngrid, p_initial) # array of pressure in each grid [6000, 6000, 6000, 6000, 6000]
time = np.arange(15, 370, delta_t)
pi_arr = []
min_arr = []
plus_arr = []
for j in range(len(time)):
pnew_arr = []
plus = pi[-1]
minus_arr = []
plusus_arr = []
for i, obj in enumerate(pi):
if i > 0:
minus = pi[i-1]
if i < (len(pi) - 1):
plus = pi[i+1]
pnew = pi[i] + (factor * ((Ti_plus_halves[i] * plus) - ((Ti_plus_halves[i] + Ti_min_halves[i]) * pi[i]) + (Ti_min_halves[i] * minus))) + (factor * qsc[i])
pnew_arr.append(float(pnew))
pnew_arr[0] = pi[0]
minus_arr.append(float(minus))
plusus_arr.append(float(plus))
pi = pnew_arr
min_arr.append(minus_arr)
plus_arr.append(plusus_arr)
pi_arr.append(pi)
df = pd.DataFrame.from_records(pi_arr)
df = pd.DataFrame(pd.np.column_stack([time, df]), columns=['time', 'grid 1', 'grid 2', 'grid 3', 'grid 4', 'grid 5', 'grid 6', 'grid 7', 'grid 8', 'grid 9', 'grid 10'])
df
```
| github_jupyter |
# Overview of Cryptographic Primitives
In this section we cover the basics of the cryptographic primitives that can be used for cryptocurrencies. We specially cover hashing algorithms and the asymmetric cryptography that is used for signatures.
## Hashing Algorithms
The goal is to have a long message as input and produce an output which is much shorter called the hash or message digest. Furthermore, we want it to have certain properties.
- **Input**: long message
- **Output**: short fixed size block (called hash or message digest)
*We want the same input always produces the same output (deterministic)*
## *<font color=" #6495ED">Exercise</font>*
- A *non* security related example of hash functions?
- A security related example of hash functions?
*** There is a difference between hash function and cryptographic hash functions ***
### Desired properties
- Pre-image: Given a hash *h* it is computationally infeasible to find a message *m* that produces *h*
- Second preimage: Given message m, it is computationally infeasible to find a message m’, (m ≠ m’) such that, h(m) = h(m’)
- Collisions: It is computationally difficult to find any two messages m, m’ (m ≠ m’) such that, h(m) = h(m’)
**Examples**:
- Recommended Hash Algorithm (SHA-2, SHA-3) by NIST
- RIPEMD
- <strike>SHA-1</strike>: output 160 bits being phased out (shattered.io)
- <strike>MD2, MD4, and MD5</strike> by Ron Rivest [RFC1319, 1320, 1321]
### SHA Family
Secure Hash Algorithm (SHA) family, is a series of hashing algorithms. Ranging from SHA-0 to SHA-3. SHA-0 should never be used, it's advised to move from SHA-1 to SHA-2. SHA-3 is the most recent version, published in 2015.
* SHA-1: Digest size (160), Block size (512)
* SHA-2: Digest size (224, 256, 384, or 512), Block size (512, 1024)
* SHA-3: Digest size (224, 256, 384, 512), Block size (1600)
### RIPEMD (RACE Integrity Primitives Evaluation Message Digest)
Another family of hashing functions. Based on the design principles of MD4. Bitcoin used the RIPEMD-160, the 160 bit version of the hashing family.
```
from cryptography.hazmat.backends import default_backend
from cryptography.hazmat.primitives import hashes
import base64 # to produce human readable encoding of the bytes
digest = hashes.Hash(hashes.SHA256(), backend=default_backend())
digest.update(b"PyCon")
digest.update(b"2018")
msg_digest = digest.finalize()
# Notice the output size of the digest
print ("msg_digest:", len(msg_digest), len(msg_digest) * 8)
print ("base64 encoding:", base64.b64encode(msg_digest))
print()
digest = hashes.Hash(hashes.SHA256(), backend=default_backend())
digest.update(b"PyCon2018")
msg_digest = digest.finalize()
# Notice the output size of the digest
print ("msg_digest:", len(msg_digest), len(msg_digest) * 8)
print ("base64 encoding:", base64.b64encode(msg_digest))
import hashlib
print(hashlib.algorithms_available)
ripemd160 = hashlib.new('ripemd160')
ripemd160.update(b"PyCon 2018")
msg_digest = ripemd160.digest()
print ("msg_digest:", len(msg_digest), len(msg_digest) * 8)
print ("base64 encoding:", base64.b64encode(msg_digest))
```
## *<font color=" #6495ED">Exercise</font> *
- Change "PyCon2018", to "PyCon 2018" and see check the result
```
## SOLUTION
```
Since the hashes are random, we can predict another hash of a value based on previous outputs. Therefore, to find a specific hash we need to bruteforce all possible input values, to find our desired output. In other words, the previous work that we have done, won't help us in the future problems.
## *<font color=" #6495ED">Exercise</font> *
- Find a has value that starts with two consecutive zeros ('00')
- Plot the number of tries it takes for a hash with n zeros (n = {1,2,3,4,5}), based on the average of 5 runs.
```
## SOLUTION
## SOLUTION
## SOLUTION
```
## Different Hashing Algorithms
Bitcoin uses SHA-256 and RIPEMD-160. However, there many are other hashing families, some of which are used by other cryptocurrencies. At the core all cryptographically secure hashing functions have the same three features that we mentioned before, some variant have extra features. For example, there are variant that are slower/faster, use more memory.
- scrypt (Litecoin, Dogecoin)
- CryptoNight (Monero)
- X11 (DASH)
## Asymmetric Encryption Simplified
Reminder: In the asymmetric encryption schemes the parties use ***different*** keys, that are mathematically ***related*** to each other.
## *<font color=" #6495ED">Exercise</font>*
- Why asymmetric encryption is useful?
- Give a few examples where it can be used?
## Elliptic-curve cryptography (ECC)
An approach to public key cryptography using elliptical curves over finite fields. Believed to provide the same level of security by having smaller key size.
Points on an elliptical curve over finite field that satisfy the following equation.
$y^2 = x^3+a.x+b$
### SECCP256k1
Standardized by Standards for Efficient Cryptography (SEC). Became significantly popular after being used in Bitcoin. $y^2 = x^3 + 7$
<img src="include/Secp256k1.png" alt="Secp256k1" style="width: 450px;"/>
source: https://en.bitcoin.it/wiki/Secp256k1
### Digital Signature Algorithm (DSA)
A signature algorithm, based on ElGamal. Private key is a large random number $x$, and public key in $y = g^x$ mod $p$.
To sign a message $m$, with parameters (p, q, g):
- Generate random value $k$, $1 < k < q$
- r = $(g ^ k$ mod $p$) mod $q$
- s = $ k ^ {-1} (H(m) + xr)$ mod $q$
- Signature is the tuple $(r,s)$
To verify a signature:
- $w = s ^ {−1}$ mod $q$
- $u_1 = H (m) \cdot w$ mod $q$
- $u_2 = r \cdot w$ mod $q$
- $v = ( g ^ {u_1} y^{u_2}$ mod $p$ ) mod $q$
- $v = r$, $iff$ siganture is valid
### Elliptic Curve Digital Signature Algorithm (ECDSA)
DSA just with the EC. Replace exponentiation with scalar point multiplication (over simplification)
```
from cryptography.hazmat.backends import default_backend
from cryptography.hazmat.primitives import hashes
from cryptography.hazmat.primitives.asymmetric import ec
private_key = ec.generate_private_key(ec.SECP256K1(), default_backend())
data = b"data to be signed"
signature = private_key.sign(data, ec.ECDSA(hashes.SHA256()))
public_key = private_key.public_key()
public_key.verify(signature, data, ec.ECDSA(hashes.SHA256()))
```
## *<font color=" #6495ED">Exercise</font>*
- Change the signature to some random value
- Change data to some random value
- Change the hashing value
```
# SOLUTION
# SOLUTION
# SOLUTION
# SOLUTION
```
| github_jupyter |
# Geochronology Calculations
```
import matplotlib.pyplot as plt
from bokeh.plotting import figure, output_notebook, show
from bokeh.layouts import column
from bokeh.models import Range1d, LinearAxis, ColumnDataSource, LabelSet, Span, Slope, Label, Legend
from scipy.interpolate import CubicSpline
import pandas as pd
import numpy as np
from IPython.core.display import display, HTML
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
output_notebook()
import geochron_apps as gc
```
<center><img src="images/geochronology.png" align="center">
https://www.explainxkcd.com/wiki/index.php/1829:_Geochronology
</center>
The following presentation shows some of the geochronology calculations learned in the Advanced Geochronology class at University of Saskatchewan taught by Bruce Eglington and Camille Partin, 2021. Some of the images in this presentation are taken from lectures given by the instructor.
This notebook contains sample calculations typically used in geochronology. It can be obtained at https://git.cs.usask.ca/msv275/advanced-geochronology.
It can be cloned through the git command:
* git clone https://git.cs.usask.ca/msv275/advanced-geochronology.git
## U-Pb Calculations
* Calculate and graph a Wetherill Concordia diagram assuming the 1975 IUGS constants
* Calculate and graph a Tera-Wasserburg Concordia diagram assuming the 1975 IUGS constants
* Calculate the 206Pb/238U, 207Pb/235U and 207Pb/206Pb ratios for a zircon formed at 3000 Ma, 2000 Ma, 1000 Ma, 500 Ma, 200 Ma and 100 Ma and plot these on both of the above graphs
* Graph Concordia curves on both Wetherill and Tera-Wasserburg plots using the most recently published decay constants and ratios for 238U and 235U decay.
* Compare the old and new decay constant and ratio results
First we need to calculate the ratios needed for each plot. The latest decay rate I could get were derived from the half life of 235U = 703.05(+-0.58) Ma from Mattinson(2010). For 238U = 4.4683(+-0.0096) Ga from Villa et al. (2016).
The equation used to calculate this decay constant is: t<sub>1/2</sub> = 0.693/*k*
```
lambda238=1.55125*10**-10,
lambda235=9.8485*10**-10
lambda238_2016=gc.convert_halflife(4469.30)
lambda235_2010=gc.convert_halflife(703.05)
print("The decay rate for U238 in 2016 was measured at {}.".format(lambda238_2016))
print("The decay rate for U235 in 2010 was measured at {}.".format(lambda235_2010))
```
In order to plot both the Wetherill and Tera-Wasserburg plots we need to calculate some U and Pb isotope ratios.
* For the Wetherill Concordia we need 206Pb/235U and 207Pb/238U.
* For the Tera-Wasserburg we need 207Pb/206Pb and 238U/206Pb
The equation used to calculate ratio of daughter to parent is *e*<sup>λ x t</sup> - 1. This is used for the 207Pb/235U and 206Pb/238U ratios.
The inverse of this equation is used to calculate 238U/206Pb.
Finally the 207Pb/206Pb is already known from the previous calculations, however it must be multiplied by 1/137.88 to account for the assumed 238U to 235 ratio in all rocks.
This will be done using the 1975 and the most recent data seperately.
```
df_1975 = gc.get_U_ratio_data()
df_2020 = gc.get_U_ratio_data(lambda238_2016,lambda235_2010)
```
As a sanity check, we will print off the first 5 calculations from each set.
```
df_1975.head()
df_2020.head()
age_list = [100,200,500,1000,2000,3000,4000,4500]
ages = gc.get_age_df(df_2020,age_list)
ages
```
The next plot uses the bokeh library and is interactive. It will allow us to zoom and pan and look at the lines in more detail.
```
figure1 = gc.get_figure("Wetherill Concordia Diagram", "207Pb/235U", "206Pb/238U", [0,100], [0,1.2])
figure1 = gc.plot_wetherill(figure1, df_1975, "green", "U Decay Rate 1975", 6)
figure1 = gc.plot_wetherill(figure1, df_2020, "red", "U Decay Rate Recent", 2)
figure1 = gc.plot_ages_wetherill(figure1, ages)
figure1.legend.location = "bottom_right"
show(figure1)
```
This plot shows that there is very little differentiation, at a large scale. If we zoom in, we can see some seperation. Now we will do the same with the Tera-Wasserburg plot.
```
figure2 = gc.get_figure("Tera-Wasserburg Concordia Diagram", "238U/206Pb", "207Pb/206Pb", [0,75], [0,0.62])
figure2 = gc.plot_tera_wasserburg(figure2, df_1975, "green", "U Decay Rate 1975", 6)
figure2 = gc.plot_tera_wasserburg(figure2, df_2020, "red", "U Decay Rate Recent", 2)
figure2 = gc.plot_ages_tera_wasserburg(figure2, ages)
figure2.legend.location = "top_right"
show(figure2)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
%matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark= False
m = 100
```
# Generate dataset
```
np.random.seed(12)
y = np.random.randint(0,3,500)
idx= []
for i in range(3):
print(i,sum(y==i))
idx.append(y==i)
x = np.zeros((500,))
np.random.seed(12)
x[idx[0]] = np.random.uniform(low =-1,high =0,size= sum(idx[0]))
x[idx[1]] = np.random.uniform(low =0,high =1,size= sum(idx[1]))
x[idx[2]] = np.random.uniform(low =2,high =3,size= sum(idx[2]))
x[idx[0]][0], x[idx[2]][5]
print(x.shape,y.shape)
idx= []
for i in range(3):
idx.append(y==i)
for i in range(3):
y= np.zeros(x[idx[i]].shape[0])
plt.scatter(x[idx[i]],y,label="class_"+str(i))
plt.legend()
bg_idx = [ np.where(idx[2] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
np.unique(bg_idx).shape
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
for i in range(3):
y= np.zeros(x[idx[i]].shape[0])
plt.scatter(x[idx[i]],y,label="class_"+str(i))
plt.legend()
foreground_classes = {'class_0','class_1' }
background_classes = {'class_2'}
fg_class = np.random.randint(0,2)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(2,3)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
a.shape
np.reshape(a,(m,1))
desired_num = 2000
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,2)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(2,3)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(np.reshape(a,(m,1)))
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
mosaic_list_of_images.shape
mosaic_list_of_images.shape, mosaic_list_of_images[0]
for j in range(m):
print(mosaic_list_of_images[0][j])
mosaic_list_of_images[0:2], mosaic_list_of_images[1000:1002]
0.2*-2.32955771 + 0.2*0.86577398 + 0.2*0.79067386 + 0.2*0.65150581 + 0.2*0.79065145
np.zeros(5)
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number, m):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
cnt = 0
counter = np.zeros(m)
for i in range(len(mosaic_dataset)):
img = torch.zeros([1], dtype=torch.float64)
np.random.seed(int(dataset_number*10000 + i))
give_pref = foreground_index[i] #np.random.randint(0,9)
# print("outside", give_pref,foreground_index[i])
for j in range(m):
if j == give_pref:
img = img + mosaic_dataset[i][j]*dataset_number/m #2 is data dim
else :
img = img + mosaic_dataset[i][j]*(m-dataset_number)/((m-1)*m)
if give_pref == foreground_index[i] :
# print("equal are", give_pref,foreground_index[i])
cnt += 1
counter[give_pref] += 1
else :
counter[give_pref] += 1
avg_image_dataset.append(img)
print("number of correct averaging happened for dataset "+str(dataset_number)+" is "+str(cnt))
print("the averaging are done as ", counter)
return avg_image_dataset , labels , foreground_index
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:1000], mosaic_label[0:1000], fore_idx[0:1000] , 1, m)
test_dataset , labels , fg_index = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[1000:2000], mosaic_label[1000:2000], fore_idx[1000:2000] , m, m)
avg_image_dataset_1 = torch.stack(avg_image_dataset_1, axis = 0)
# mean = torch.mean(avg_image_dataset_1, keepdims= True, axis = 0)
# std = torch.std(avg_image_dataset_1, keepdims= True, axis = 0)
# avg_image_dataset_1 = (avg_image_dataset_1 - mean) / std
# print(torch.mean(avg_image_dataset_1, keepdims= True, axis = 0))
# print(torch.std(avg_image_dataset_1, keepdims= True, axis = 0))
# print("=="*40)
test_dataset = torch.stack(test_dataset, axis = 0)
# mean = torch.mean(test_dataset, keepdims= True, axis = 0)
# std = torch.std(test_dataset, keepdims= True, axis = 0)
# test_dataset = (test_dataset - mean) / std
# print(torch.mean(test_dataset, keepdims= True, axis = 0))
# print(torch.std(test_dataset, keepdims= True, axis = 0))
# print("=="*40)
x1 = (avg_image_dataset_1).numpy()
y1 = np.array(labels_1)
# idx1 = []
# for i in range(3):
# idx1.append(y1 == i)
# for i in range(3):
# z = np.zeros(x1[idx1[i]].shape[0])
# plt.scatter(x1[idx1[i]],z,label="class_"+str(i))
# plt.legend()
plt.scatter(x1[y1==0], y1[y1==0]*0, label='class 0')
plt.scatter(x1[y1==1], y1[y1==1]*0, label='class 1')
# plt.scatter(x1[y1==2], y1[y1==2]*0, label='class 2')
plt.legend()
plt.title("dataset1 CIN with alpha = 1/"+str(m))
x1 = (avg_image_dataset_1).numpy()
y1 = np.array(labels_1)
idx_1 = y1==0
idx_2 = np.where(idx_1==True)[0]
idx_3 = np.where(idx_1==False)[0]
color = ['#1F77B4','orange', 'brown']
true_point = len(idx_2)
plt.scatter(x1[idx_2[:25]], y1[idx_2[:25]]*0, label='class 0', c= color[0], marker='o')
plt.scatter(x1[idx_3[:25]], y1[idx_3[:25]]*0, label='class 1', c= color[1], marker='o')
plt.scatter(x1[idx_3[50:75]], y1[idx_3[50:75]]*0, c= color[1], marker='o')
plt.scatter(x1[idx_2[50:75]], y1[idx_2[50:75]]*0, c= color[0], marker='o')
plt.legend()
plt.xticks( fontsize=14, fontweight = 'bold')
plt.yticks( fontsize=14, fontweight = 'bold')
plt.xlabel("X", fontsize=14, fontweight = 'bold')
# plt.savefig(fp_cin+"ds1_alpha_04.png", bbox_inches="tight")
# plt.savefig(fp_cin+"ds1_alpha_04.pdf", bbox_inches="tight")
avg_image_dataset_1[0:10]
x1 = (test_dataset).numpy()/m
y1 = np.array(labels)
# idx1 = []
# for i in range(3):
# idx1.append(y1 == i)
# for i in range(3):
# z = np.zeros(x1[idx1[i]].shape[0])
# plt.scatter(x1[idx1[i]],z,label="class_"+str(i))
# plt.legend()
plt.scatter(x1[y1==0], y1[y1==0]*0, label='class 0')
plt.scatter(x1[y1==1], y1[y1==1]*0, label='class 1')
# plt.scatter(x1[y1==2], y1[y1==2]*0, label='class 2')
plt.legend()
plt.title("test dataset1 ")
test_dataset.numpy()[0:10]/m
test_dataset = test_dataset/m
test_dataset.numpy()[0:10]
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
avg_image_dataset_1[0].shape, avg_image_dataset_1[0]
batch = 200
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
testdata_1 = MosaicDataset(test_dataset, labels )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet,self).__init__()
self.linear1 = nn.Linear(1,2)
# self.linear2 = nn.Linear(50,10)
# self.linear3 = nn.Linear(10,3)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
def forward(self,x):
# x = F.relu(self.linear1(x))
# x = F.relu(self.linear2(x))
x = (self.linear1(x))
return x
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/i
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("unique out: ", np.unique(out), "unique pred: ", np.unique(pred) )
print("correct: ", correct, "total ", total)
print('Accuracy of the network on the 1000 test dataset %d: %.2f %%' % (number , 100 * correct / total))
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
net = Whatnet().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.Adam(net.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 1500
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.05:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %.2f %%' % ( 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
return loss_curi, net
train_loss_all=[]
testloader_list= [ testloader_1 ]
loss, net = train_all(trainloader_1, 1, testloader_list)
train_loss_all.append(loss)
net.linear1.weight, net.linear1.bias
%matplotlib inline
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
```
| github_jupyter |
### Merging data
```
# import modules
import numpy as np
import pandas as pd
# import zipcodes
zipcode = pd.read_csv("data/zipfips.csv")
zipcode.head()
# function for data files
def imp_acsdf(num, samp ):
dfname = "df" + num
print("Dataframe Name is: %s" % dfname)
if samp == False:
dfname = pd.read_csv("./data/ACS_16_5YR_DP" + num + "_with_ann.csv",skiprows=[1], low_memory=False)
else:
dfname = pd.read_csv("./data/ACS_16_5YR_DP" + num + "_with_ann.csv", nrows = 1000, skiprows=[1] )
print (dfname.shape)
return dfname
# function for meta files
def imp_acsmeta(num ):
metaname = "meta" + num
print("MetaData Name is: %s" % metaname)
metaname = pd.read_csv("./data/ACS_16_5YR_DP" + num + "_metadata.csv", header=None, index_col = 0 )
print(metaname.shape)
return metaname
dict_df = {}
dict_meta={}
for i in range(4):
numstr = "0" + str(i+2)
key_namedf = "df" + numstr
dict_df[key_namedf] = imp_acsdf(num = numstr, samp = False) # change to false to import all data
key_namemeta = "meta" + numstr
dict_meta[key_namemeta] = imp_acsmeta(num = numstr)
dict_df.keys()
# add df and meta to dict
df02 = dict_df['df02']
df03 = dict_df['df03']
df04 = dict_df['df04']
df05 = dict_df['df05']
meta02 = dict_meta['meta02'].to_dict()[1]
meta03 = dict_meta['meta03'].to_dict()[1]
meta04 = dict_meta['meta04'].to_dict()[1]
meta05 = dict_meta['meta05'].to_dict()[1]
# function to extact estimates
def extract_Est(df):
df = df[df.columns.drop(list(df.filter(regex='HC02')))]
df = df[df.columns.drop(list(df.filter(regex='HC03')))]
df = df[df.columns.drop(list(df.filter(regex='HC04')))]
return df
# Extract only the estimates in df which all start with 'HCO1'
df02 = extract_Est(df02)
df03 = extract_Est(df03)
df04 = extract_Est(df04)
df05 = extract_Est(df05)
# Function to extract meaningful categories from variable names
def extract_catdf(df, dname, dno):
df = pd.DataFrame.from_dict(df, orient='index')
df = df.rename_axis('v').reset_index().rename(columns={0:'detail'})
df =df.iloc[3:]
df['type'] = df.detail.str.split(';').str[0]
df['cat0'] = dname
df['cat1'] = df.detail.str.split(';').str[1].str.split('-',1).str[0].str.strip()
df['cat2'] = df.detail.str.split(';').str[1].str.split('-',1).str[1].str.strip()
df['vname'] = dno + df.v
return df
# name categories extracted from variable names
m02 = extract_catdf(meta02, "Social", "02")
m03 = extract_catdf(meta03, "Economic","03")
m04 = extract_catdf(meta04, "Housing","04")
m05 = extract_catdf(meta05 , "Demographic","05")
# concatenate meta categories
meta = pd.concat([m02, m03, m04, m05], ignore_index=True)
print(meta.type.unique())
print(meta.cat0.unique())
print(meta.cat1.unique())
print(meta.cat2.unique())
print(meta.cat3.unique())
meta = meta[meta.type == 'Estimate']
def rename_cols(df, num):
cols = df.columns[~df.columns.str.contains('GEO')]
df.rename(columns = dict(zip(cols, num + cols )), inplace=True)
return df
df02 = rename_cols(df02,'02')
df03 = rename_cols(df03,'03')
df04 = rename_cols(df04,'04')
df05 = rename_cols(df05,'05')
# merge data files with zipcodes
print(zipcode.shape)
df = pd.merge(left=zipcode, right=df02, right_on='GEO.id2', left_on='zipcode', how = 'left')
print(df.shape)
df = pd.merge(left=df, right=df03, right_on='GEO.id2', left_on='zipcode', how = 'left')
print(df.shape)
df = pd.merge(left=df, right=df04, right_on='GEO.id2', left_on='zipcode', how = 'left')
print(df.shape)
df = pd.merge(left=df, right=df05, right_on='GEO.id2', left_on='zipcode', how = 'left')
print(df.shape)
df.head()
meta.head()
# save df and meta to csv
df.to_csv("./data/df.csv")
meta.to_csv("./data/meta.csv")
```
### Cleaning data
```
# function to group data
def group_data(df, geo = 'country'):
if geo == 'country':
df
elif geo == 'state':
df = df.groupby("state").sum()
elif geo == 'county':
df = df.groupby(["state", "county"]).sum()
elif geo == 'city':
df = df.groupby(["state", "county", "city"]).sum()
return df
grouped_data = group_data(df, geo = 'city')
grouped_data.head()
meta['level1'] = meta.cat2.str.split('-',1).str[0].str.strip()
meta['level2'] = meta.cat2.str.split('-',1).str[1].str.strip()
meta['level3'] = meta.level2.str.split('-',1).str[1].str.strip()
meta.head()
df.describe()
df.select_dtypes(include=['object']).columns
columns = ['02HC01_VC21', '02HC01_VC22', '02HC01_VC52', '02HC01_VC53',
'02HC01_VC54', '02HC01_VC55', '02HC01_VC56', '02HC01_VC57',
'02HC01_VC58', '02HC01_VC95', '02HC01_VC96', '02HC01_VC216',
'02HC01_VC217', '02HC01_VC218', '03HC01_VC12', '03HC01_VC36',
'03HC01_VC85', '03HC01_VC86', '03HC01_VC90', '03HC01_VC92',
'03HC01_VC94', '03HC01_VC98', '03HC01_VC100', '03HC01_VC114',
'03HC01_VC115', '03HC01_VC118', '03HC01_VC121', '03HC01_VC122',
'03HC01_VC124', '03HC01_VC125', '03HC01_VC126', '03HC01_VC161',
'03HC01_VC162', '03HC01_VC163', '03HC01_VC164', '03HC01_VC165',
'03HC01_VC166', '03HC01_VC167', '03HC01_VC168', '03HC01_VC169',
'03HC01_VC171', '03HC01_VC172', '03HC01_VC173', '03HC01_VC174',
'03HC01_VC175', '03HC01_VC176', '03HC01_VC177', '03HC01_VC178',
'03HC01_VC179', '03HC01_VC180', '04HC01_VC08', '04HC01_VC09',
'04HC01_VC50', '04HC01_VC69', '04HC01_VC70', '04HC01_VC106',
'04HC01_VC108', '04HC01_VC128', '04HC01_VC146', '04HC01_VC155',
'04HC01_VC191', '05HC01_VC23']
df[columns] = df[columns].apply(pd.to_numeric, errors='coerce')
# see column types
df.dtypes
df.select_dtypes(include=['object']).columns
# get info on data
print(df.info())
# # DataFrame method to check for missing values in a column
# # The .all() method returns True if all values are True
# # if using it on a DataFrame, you need to chain another .all()
# # method so that you return only one True or False value.
# assert pd.notnull(df['GEO.id_x']).all().all()
# assert pd.notnull(df['HC01_VC150']).all().all()
# assert (df.HC01_VC150 >=0).all().all()
df.tail(20)
# # save df and meta to csv
# df.to_csv("./data/df1.csv")
# meta.to_csv("./data/meta1.csv")
df.head()
meta.head()
social = meta[meta['cat0'] == 'Social']
total_household = social[social['level1'] == 'Total households']
total_household
total_household.shape
total_household = total_household[: -3]
total_household
household_list = total_household["vname"].tolist()
household_list
new_cols = ['zipcode', 'latitude', 'longitude', 'city', 'state', 'county', 'fips', '02HC01_VC03', '02HC01_VC04', '02HC01_VC05']
df_th = df[new_cols].fillna(0)
df_th = df_th[(df_th != 0).all(1)]
df_th.head()
# save df to csv
df_th.to_csv("./data/df_th.csv")
df_th.iloc[0:20]
household_list1 = total_household["cat2"].tolist()
household_list1
```
| github_jupyter |
<figure>
<IMG SRC="https://mamba-python.nl/images/logo_basis.png" WIDTH=125 ALIGN="right">
</figure>
# Common pitfalls
<br>
This notebook is created for the MAMBA Python course to explain the common pitfalls using Python.
<br><br>
<div style="text-align: right"> developed by MAMBA </div>
Table of content:<a class="anchor" id="0"></a>
1. [variable names](#1)
1. [reusing variable names](#11)
2. [built-in functions](#12)
2. [nested loops and statements](#2)
3. [copy-paste](#4)
4. [answers to exercises](#answers)
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
%matplotlib inline
```
[back to TOC](#0)
## 1. variable names<a class="anchor" id="1"></a>
Naming your variables may sound trivial but it is actually quite important for the readability of your code. Some naming conventions are written down in PEP-8. Keeping to these allows for the next reader to easily understand which are function names, class names, packages etc.
An example of a widely adopted naming convention is shown below. The convention is that <b>variables and function names</b> should be <b>lowercase</b>, with words separated by <b>underscores</b> as necessary to improve readability.
```
# according to conventions (PEP-8)
list_of_animals = ['dog', 'cat', 'mouse', 'duck']
# not according to conventions (PEP-8)
ListOfAnimals = ['dog','cat','mouse','duck']
```
### A. reusing variable names<a class="anchor" id="11"></a>
#### Exercise 1
#### answer
### B. built-in functions<a class="anchor" id="12"></a>
There are a number of built-in functions that are available without importing them via a package. These functions can be shown with the command below.
```
dir(__builtin__)
```
If you use a built-in function as a variable name, you overwrite the built-in function and cannot use it anymore. See the two examples below. You have to restart the kernel <button class='btn btn-default btn-xs'><i class='fa fa-repeat icon-repeat'></i></button> in order to retrieve the built-in function.
Note: after you understood the examples and did the exercise it may be helpful to comment out the cells below. Otherwise you constantly have to restart the kernel if you accidentally run these cells. You can comment out a cell by selecting its code and press `[ctrl][/]`.
```
# example 1
range(0,3) #The built-in function range generates the integer numbers between the given start integer to the stop integer
print([x for x in range(0,3)])
range = (3,6) #this overwrites the built-in function 'range'
print([x for x in range])
range(0,3) #now you cannot call this function anymore!
# example 2
list = [1,2,3] #this overwrites the built-in function 'list'
a = (5,4,7) #if you want to convert this tuple to a list you get an error
list(a)
```
#### Exercise 2
You have an integer, a float and a list. You want to divide the integer by the float and use the remainder as an index for the list. The code below gives a TypeError. Try to find out what causes the error and change the code to avoid it.
```
int = 2
float = 2.0
a_list = [1, 2, 3, 4]
index_a = int/float
print(a[int(index_a)])
```
#### answer
[back to TOC](#0)
## 2. loops<a class="anchor" id="2"></a>
loops are a very powerful tool but can make code slow and hard to read. The common approach is to avoid loops, especially nested loops, if possible.
### A. avoid loops with pandas<a class="anchor" id="21"></a>
The example below is about a pandas DataFrame with data from a whatsapp conversation. Every row has a datetime, user and message (the text is ommited because of privacy issues).
```
df = pd.read_csv(r'..\..\..\..\Practical_examples\whatsapp data\data\_chat_df.csv')
df['datetime'] = pd.to_datetime(df['Unnamed: 0'])
del df['Unnamed: 0']
df.head()
```
Let's say you want to get a new column with the hour of the day of each row. You could use a for-loop to loop over every row and get the hour of the day from the datetime column. See the example below. This will take some time. Using the `%%time` command you can see how much time.
```
%%time
df['hour'] = 0
for row_no, row in df.iterrows():
df.loc[row_no]['hour'] = row['datetime'].hour
print(df.head())
```
There is an alternative for the for-loop, this is in general referred to as a vectorized solution. The operation is not executed row-by-row but on the whole vector (column) at once. The example below shows a vectorized solution.
```
df['hour'] = df['datetime'].dt.hour
df.head()
```
#### exercise 3
Create a new colum 'user_int' that only contains the number of the user as an integer. You can check the dtype of the column with `df['user_int'].dtype`.
#### answer
### B. avoid nested for loops<a class="anchor" id="22"></a>
Consider the code below in which an image of a ship is blurred by taking the average gray value of a 10x10 pixel patch.
source: https://realpython.com/numpy-array-programming/
```
from skimage import io
url = ('https://www.history.navy.mil/bin/imageDownload?image=/'
'content/dam/nhhc/our-collections/photography/images/'
'80-G-410000/80-G-416362&rendition=cq5dam.thumbnail.319.319.png')
img = io.imread(url, as_grey=True)
fig, ax = plt.subplots()
ax.imshow(img, cmap='gray')
# using a nested for loop
m, n = img.shape
mm, nn = m - size + 1, n - size + 1
patch_means = np.empty((mm, nn))
for i in range(mm):
for j in range(nn):
patch_means[i, j] = img[i: i+size, j: j+size].mean()
fig, ax = plt.subplots()
ax.imshow(patch_means, cmap='gray')
ax.grid(False)
shape = (img.shape[0] - size + 1, img.shape[1] - size + 1, size, size)
patches = stride_tricks.as_strided(img, shape=shape, strides=img.strides*2)
veclen = 10**2
patches.reshape(*patches.shape[:2], veclen).mean(axis=-1).shape
strided_means = patches.mean(axis=(-1, -2))
fig, ax = plt.subplots()
ax.imshow(strided_means, cmap='gray')
ax.grid(False)
```
#### Exercise 4
Below you see some code to calculate the sum of the rows and the columns of a numpy array. The code contains a nested for loop. Find a way to calculate the sum of the rows and columns without using a nested for-loop.
Bonus: find a way to calculate the sum of the rows and columns without using a for-loop at all.
```
rand_arr = np.random.random(size=(3,4))
rand_arr
row_sum_list = []
col_sum_list = []
for i in range(len(rand_arr)):
row_sum = 0
for j in range(len(rand_arr[i])):
row_sum += rand_arr[i][j]
if i==0:
col_sum_list.append(np.sum(rand_arr[:,j]))
row_sum_list.append(row_sum)
print('sum of rows',row_sum_list)
print('sum of columns',col_sum_list)
```
#### answer
[back to TOC](#0)
## 3. copy-paste <a class="anchor" id="3"></a>
copy and paste is both a blessing and a curse for programming. It may look obvious to copy-paste a piece of code if you need it again. However this may turn out to be very time consuming in the end.
```
df2 = pd.DataFrame({ 'A' : 1.,
'B' : pd.Timestamp('20130102'),
'C' : pd.Series(1,index=list(range(4)),dtype='float32'),
'D' : np.array([3] * 4,dtype='int32'),
'E' : pd.Categorical(["test","train","test","train"]),
'F' : 'foo' })
```
#### Exercise 5
#### answer
[back to TOC](#0)
## 4. answers to exercises <a class="anchor" id="answers"></a>
#### answer to exercise 1
#### answer to exercise 2
You need to rename the variables int and float to avoid overriding the built-in functions `int()` and `float()`.
Note: if you used `int` as a variable name you need to restart the kernel <button class='btn btn-default btn-xs'><i class='fa fa-repeat icon-repeat'></i></button> to be able to use the `int()` built-in function.
```
no_int = 2
no_float = 2.0
a_list = ['item 0','item 1', 'item 2', 'item 3']
index_a = no_int/no_float
print(a_list[int(index_a)])
```
#### answer to exercise 3
```
df['user_int'] = df.user.str[-1].astype('int')
df.head()
```
#### answer to exercise 4
```
# answer 1 without nested for-loops
row_sum_list = []
for i in range(len(rand_arr)):
row_sum_list.append(np.sum(rand_arr[i,:]))
col_sum_list = []
for j in range(len(rand_arr[0])):
col_sum_list.append(np.sum(rand_arr[:,j]))
print('sum of rows',row_sum_list)
print('sum of columns',col_sum_list)
# answer 2 without for-loops
print('sum of rows', rand_arr.sum(axis=1))
print('sum of columns', rand_arr.sum(axis=0))
# answer 3 without for-loops
print('sum of rows', np.sum(rand_arr, axis=1))
print('sum of columns', np.sum(rand_arr, axis=0))
```
#### answer to exercise 5
| github_jupyter |
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import re
# ./parsed.json is the stdout of the scraper tool in this directory
df = pd.read_json("./parsed.json", lines=True)
df
def parse_ds(entity):
m = re.search(r"(?P<dataset>[^@#]*)([@#].+)?", entity)
return m.group("dataset")
def parse_cmd(row):
cmd = row.Cmd
binary, verb, *tail = re.split(r"\s+", cmd) # NOTE whitespace in dataset names => don't use comp
dataset = None
if binary == "zfs":
if verb == "send":
if len(tail) == 0:
verb = "send-feature-test"
else:
dataset = parse_ds(tail[-1])
if "-n" in tail:
verb = "send-dry"
elif verb == "recv" or verb == "receive":
verb = "receive"
if len(tail) > 0:
dataset = parse_ds(tail[-1])
else:
verb = "receive-CLI-test"
elif verb == "get":
dataset = parse_ds(tail[-1])
elif verb == "list":
if "-r" in tail and "-d" in tail and "1" in tail:
dataset = parse_ds(tail[-1])
verb = "list-single-dataset"
else:
dataset = "!ALL_POOLS!"
verb = "list-all-filesystems"
elif verb == "bookmark":
dataset = parse_ds(tail[-2])
elif verb == "hold":
dataset = parse_ds(tail[-1])
elif verb == "snapshot":
dataset = parse_ds(tail[-1])
elif verb == "release":
dss = tail[-1].split(",")
if len(dss) > 1:
raise Exception("cannot handle batch-release")
dataset = parse_ds(dss[0])
elif verb == "holds" and "-H" in tail:
dss = tail[-1].split(",")
if len(dss) > 1:
raise Exception("cannot handle batch-holds")
dataset = parse_ds(dss[0])
elif verb == "destroy":
dss = tail[-1].split(",")
if len(dss) > 1:
raise Exception("cannot handle batch-holds")
dataset = parse_ds(dss[0])
return {'action':binary + "-" + verb, 'dataset': dataset }
res = df.apply(parse_cmd, axis='columns', result_type='expand')
res = pd.concat([df, res], axis='columns')
for cat in ["action", "dataset"]:
res[cat] = res[cat].astype('category')
res["LogTimeUnix"] = pd.to_datetime(res.LogTime)
res["OtherTime"] = res.TotalTime - res.Usertime - res.Systime
x = res.melt(id_vars=["action", "dataset"], value_vars=["TotalTime", "OtherTime", "Usertime", "Systime"])
x
print("commands with NaN values")
set(x[x.isna().any(axis=1)].action.values)
# (~x.action.astype('str').isin(["zfs-send", "zfs-recv"]))
totaltimes = x[(x.variable == "TotalTime")].groupby(["action", "dataset"]).sum().reset_index()
display(totaltimes)
totaltimes_by_action = totaltimes.groupby("action").sum().sort_values(by="value")
totaltimes_by_action.plot.barh()
totaltimes.groupby("dataset").sum().plot.barh(fontsize=5)
most_expensive_action = totaltimes_by_action.idxmax().value
display(most_expensive_action)
most_expensive_action_by_dataset = totaltimes[totaltimes.action == most_expensive_action].groupby("dataset").sum().sort_values(by="value")
most_expensive_action_by_dataset.plot.barh(rot=50, fontsize=5, figsize=(10, 20))
plt.savefig('most-expensive-command.pdf')
res
# %matplotlib notebook
# res.index = res.LogTimeUnix
# resampled = res.pivot(columns='action', values='TotalTime').resample("1s").sum()
# resampled.cumsum().plot()
# res["BeginTime"] = res.LogTimeUnix.dt.total_seconds()
# holds = res[res.action == "zfs-holds"]
# sns.stripplot(x="LogTimeUnix", y="action", data=res)
# res["LogTimeUnix"].resample("20min").sum()
# res[res.action == "zfs-holds"].plot.scatter(x="LogTimeUnix", y="TotalTime")
#res[res.action == "zfs-holds"].pivot(columns='action', values=['TotalTime', 'Systime', "Usertime"]).resample("1s").sum().cumsum().plot()
pivoted = res.reset_index(drop=True).pivot_table(values=['TotalTime', 'Systime', "Usertime"], index="LogTimeUnix", columns="action")
pivoted
pivoted.cumsum()[[("TotalTime", "zfs-holds"),("Systime", "zfs-holds"),("Usertime", "zfs-holds")]].plot()
pivoted = res.reset_index(drop=True).pivot_table(values=['TotalTime'], index="LogTimeUnix", columns="action")
cum_invocation_counts_per_action = pivoted.isna().astype(int).cumsum()
cum_invocation_counts_per_action
# zfs-get as reference value
cum_invocation_counts_per_action[[("TotalTime","zfs-holds"),("TotalTime","zfs-get")]].plot()
```
| github_jupyter |
This notebook compares the email activities and draft activites of an IETF working group.
Import the BigBang modules as needed. These should be in your Python environment if you've installed BigBang correctly.
```
import bigbang.mailman as mailman
from bigbang.parse import get_date
#from bigbang.functions import *
from bigbang.archive import Archive
from ietfdata.datatracker import *
```
Also, let's import a number of other dependencies we'll use later.
```
import pandas as pd
import datetime
import matplotlib.pyplot as plt
import numpy as np
import math
import pytz
import pickle
import os
```
## Load the HRPC Mailing List
Now let's load the email data for analysis.
```
wg = "httpbisa"
urls = [wg]
archives = [Archive(url,mbox=True) for url in urls]
activities = [arx.get_activity(resolved=False) for arx in archives]
activity = activities[0]
```
## Load IETF Draft Data
Next, we will use the `ietfdata` tracker to look at the frequency of drafts for this working group.
```
from ietfdata.datatracker import *
from ietfdata.datatracker_ext import *
import pandas as pd
dt = DataTracker()
g = dt.group_from_acronym("httpbisa")
drafts = [draft for draft in dt.documents(group=g, doctype=dt.document_type_from_slug("draft"))]
draft_df = pd.DataFrame.from_records([
{'time' : draft.time, 'title' : draft.title, 'id' : draft.id} for draft in drafts]
)
```
We will want to use the data of the drafts. Time resolution is too small.
```
draft_df['date'] = draft_df['time'].dt.date
draft_df
```
## Gender score and tendency measures
This notebook uses the (notably imperfect) method of using first names to guess the gender of each draft author.
```
from gender_detector import gender_detector as gd
detector = gd.GenderDetector('us')
def gender_score(name):
"""
Takes a full name and returns a score for the guessed
gender.
1 - male
0 - female
.5 - unknown
"""
try:
first_name = name.split(" ")[0]
guess = detector.guess(first_name)
score = 0
if guess == "male":
return 1.0
elif guess == "female":
return 0.0
else:
# name does not have confidence to guesss
return 0.5
except:
# Some error, "unknown"
return .5
```
## Gender guesses on mailing list activity
Now to use the gender guesser to track the contributions by differently gendered participants over time.
```
from bigbang.parse import clean_name
gender_activity = activity.groupby(
by=lambda x: gender_score(clean_name(x)),
axis=1).sum().rename({0.0 : "women", 0.5 : "unknown", 1.0 : "men"},
axis="columns")
```
Note that our gender scoring method currently is unable to get a clear guess for a large percentage of the emails!
```
print("%f.2 percent of emails are from an unknown gender." \
% (gender_activity["unknown"].sum() / gender_activity.sum().sum()))
plt.bar(["women","unknown","men"],gender_activity.sum())
plt.title("Total emails sent by guessed gender")
```
## Plotting
Some preprocessing is necessary to get the drafts data ready for plotting.
```
from matplotlib import cm
viridis = cm.get_cmap('viridis')
drafts_per_day = draft_df.groupby('date').count()['title']
dpd_log = drafts_per_day.apply(lambda x: np.log1p(x))
```
For each of the mailing lists we are looking at, plot the rolling average (over `window`) of number of emails sent per day.
Then plot a vertical line with the height of the drafts count and colored by the gender tendency.
```
window = 100
plt.figure(figsize=(12, 6))
for i, gender in enumerate(gender_activity.columns):
colors = [viridis(0), viridis(.5), viridis(.99)]
ta = gender_activity[gender]
rmta = ta.rolling(window).mean()
rmtadna = rmta.dropna()
plt.plot_date(np.array(rmtadna.index),
np.array(rmtadna.values),
color = colors[i],
linestyle = '-', marker = None,
label='%s email activity - %s' % (wg, gender),
xdate=True)
vax = plt.vlines(drafts_per_day.index,
0,
drafts_per_day,
colors = 'r', # draft_gt_per_day,
cmap = 'viridis',
label=f'{wg} drafts ({drafts_per_day.sum()} total)'
)
plt.legend()
plt.title("%s working group emails and drafts" % (wg))
#plt.colorbar(vax, label = "more womanly <-- Gender Tendency --> more manly")
#plt.savefig("activites-marked.png")
#plt.show()
```
### Is gender diversity correlated with draft output?
```
from scipy.stats import pearsonr
import pandas as pd
def calculate_pvalues(df):
df = df.dropna()._get_numeric_data()
dfcols = pd.DataFrame(columns=df.columns)
pvalues = dfcols.transpose().join(dfcols, how='outer')
for r in df.columns:
for c in df.columns:
pvalues[r][c] = round(pearsonr(df[r], df[c])[1], 4)
return pvalues
drafts_per_ordinal_day = pd.Series({x[0].toordinal(): x[1] for x in drafts_per_day.items()})
drafts_per_ordinal_day
ta.rolling(window).mean()
garm = np.log1p(gender_activity.rolling(window).mean())
garm['diversity'] = (garm['unknown'] + garm['women']) / garm['men']
garm['drafts'] = drafts_per_ordinal_day
garm['drafts'] = garm['drafts'].fillna(0)
garm.corr(method='pearson')
calculate_pvalues(garm)
```
Some variations...
```
garm_dna = garm.dropna(subset=['drafts'])
```
| github_jupyter |
# Intel® SSD Data Center Tool Connector
This notebook demonstrates some of the quick analysis that can be done using the TOKIO connector for the Intel SSD Data Center Tool (ISDCT). The format of the aggregated ISDCT outputs is specific to a tool developed at NERSC by David Paul and is therefore site-specific to NERSC, but the individual parsers for each ISDCT output file are generic.
```
%matplotlib inline
import os
import datetime
import numpy as np
import matplotlib
matplotlib.rcParams.update({'font.size': 18})
import matplotlib.pyplot as plt
import tokio.config
import tokio.connectors.nersc_isdct
import tokio.tools.common
TARGET_DATE = datetime.datetime(year=2018, month=4, day=13)
GENERATE_PLOTS = True
PLOT_SUFFIX = "png" # or pdf, gif, jpeg...
print "Generating report for %s" % TARGET_DATE.strftime('%c')
isdct_file = tokio.tools.common.enumerate_dated_files(start=TARGET_DATE,
end=TARGET_DATE,
template=tokio.config.ISDCT_FILES)
print "Using input file: %s" % isdct_file[0]
isdct_data = tokio.connectors.nersc_isdct.NerscIsdct(isdct_file[0])
isdct_df = isdct_data.to_dataframe()
```
## Distribution of Lifetime Read/Write Loads
The following histograms demonstrate how many bytes have been written to and read from the SSD device _by applications_ over the entire service life of the SSD.
```
for rw, column in ('read','data_units_read_bytes'), ('write', 'data_units_written_bytes'):
fig, ax = matplotlib.pyplot.subplots()
fig.set_size_inches(10, 6)
fig.suptitle("%s Volume Distribution" % rw.title())
ax.set_axisbelow(True)
ax.grid(True)
ax.set_xlabel("TiB %s" % rw.title())
ax.set_ylabel("Number of SSDs")
(isdct_df[column] / 2.0**40).hist(ax=ax, edgecolor='black')
if GENERATE_PLOTS:
output_file = 'histogram_%s_%s_%s.%s' % (rw, column, TARGET_DATE.strftime("%Y-%m-%d"), PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
```
The read/write ratio from our applications should ideally match the read/write performance balance of the NVMe drives. Writes are typically slower than reads on flash.
```
fig, ax = matplotlib.pyplot.subplots(figsize=(10, 6))
ax.set_axisbelow(True)
ax.grid(True)
ax.set_xlabel("Read/Write Ratio")
ax.set_ylabel("Number of SSDs")
(isdct_df['data_units_read_bytes'] / isdct_df['data_units_written_bytes']).hist(ax=ax, edgecolor='black')
if GENERATE_PLOTS:
output_file = 'histogram_readwrite_ratio.%s' % (PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
```
## Write Amplification Distribution
Write amplification factor (WAF) is the ratio of bytes written to the device _by applications_ to the bytes written to the physical NAND chips, which includes both application-generated writes as well as writes caused by garbage collection.
A WAF of 1.0 is ideal; 2.0 is a normal level for the Intel SSDs we have in production. High WAF is usually indicative of either
1. very new SSDs which have not seen much application-generated I/O; in these cases, the constant background load of the NVMe controller bubbles up to the surface
2. workloads which are very SSD-unfriendly. These typically include writes that are not 4K aligned. With DataWarp, the only non-4K aligned writes are those which are smaller than 4K.
```
fig, ax = matplotlib.pyplot.subplots()
fig.set_size_inches(10, 6)
fig.suptitle("WAF Distribution")
ax.set_axisbelow(True)
ax.grid(True)
ax.set_xlabel("Write Amplification Factor")
ax.set_ylabel("Number of SSDs")
isdct_df['write_amplification_factor'].hist(ax=ax, edgecolor='black')
if GENERATE_PLOTS:
output_file = 'histogram_waf_%s.%s' % (TARGET_DATE.strftime("%Y-%m-%d"), PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
```
## Drive Writes per Day
Our Intel P3608 SSDs have a warranty of 5.0 drive writes per day (DWPD) when provisioned at 1.6 TB capacity for the five-year service life of the drive.
We have the option of reformatting the drives as 2.0 TB drives, which reduces the warranted endurance rating to 1.0 DWPD.
```
fig, ax = matplotlib.pyplot.subplots()
fig.set_size_inches(10, 6)
fig.suptitle("DWPD Distribution")
ax.set_axisbelow(True)
ax.grid(True)
ax.set_xlabel("Drive Writes per Day")
ax.set_ylabel("Number of SSDs")
drive_writes = isdct_df['data_units_written_bytes'] / isdct_df['physical_size']
dwpd = drive_writes / isdct_df['power_on_hours'] * 24.0
dwpd.hist(ax=ax, edgecolor='black')
if GENERATE_PLOTS:
output_file = 'histogram_dwpd_%s.%s' % (TARGET_DATE.strftime("%Y-%m-%d"), PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
```
## Correlation Scatter Plots
Because many of the health metrics are ratios that get skewed when SSDs have seen very light use, it is sometimes helpful to correlate these health metrics with the number of hours the drives have been in service.
We expect the total volume of I/O to each drive to increase over time, and the WAF should decrease over time as each drive reaches steady state.
```
scatter_plots = [
('power_on_hours', 'data_units_written_bytes'),
('power_on_hours', 'data_units_read_bytes'),
('power_on_hours', 'write_amplification_factor'),
]
def scatter_and_fit_plot(df, x_key, y_key, fit=True):
fig, ax = matplotlib.pyplot.subplots()
fig.set_size_inches(10, 6)
x = df[x_key].values
y = df[y_key].values
ax.plot(x, y, 'o', alpha=0.5)
if fit:
### attempt a linear fit to generate a visual aid
m, b = np.polyfit(x, y, 1)
ax.plot(x, m*x+b, "-")
ax.set_xlabel(x_key.replace('_', ' ').title())
ax.set_ylabel(y_key.replace('_', ' ').title())
plt.grid(True)
if GENERATE_PLOTS:
output_file = 'correlate_%s-%s_%s.%s' % (x_key, y_key, TARGET_DATE.strftime("%Y-%m-%d"), PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
for (x_key, y_key) in scatter_plots:
scatter_and_fit_plot(isdct_df, x_key, y_key)
```
## Identify faulty node power sources
The "PLI Lock Loss" counter was originally thought to be an indicator of unhealthy drives. It turns out that this metric is really a PLL (phase-locked loop) lock loss count, which increments when the PCIe timing signal falls irreparably out of sync with the internal clock on the SSD. This is __not__ an indicator of bad drive health as originally thought; it is an indicator of unclean power to the host node.
```
pli_lock_losses = isdct_df[isdct_df['smart_pli_lock_loss_count_raw'] > 0]
pli_lock_losses[['node_name', 'smart_pli_lock_loss_count_raw', 'power_on_hours']]\
.sort_values('smart_pli_lock_loss_count_raw', ascending=False)
x_key = 'power_on_hours'
y_key = 'smart_pli_lock_loss_count_raw'
fig, ax = matplotlib.pyplot.subplots()
fig.set_size_inches(10, 6)
ax.plot(isdct_df[x_key].values,
isdct_df[y_key].values,
marker='o',
linestyle='none',
alpha=0.5,
label="All SSDs")
ax.plot(pli_lock_losses[x_key],
pli_lock_losses[y_key],
marker='o',
linestyle='none',
alpha=0.5,
color='red',
markersize=10,
markerfacecolor='none',
label="Nonzero PLI Lock Loss")
ax.legend()
ax.set_xlabel(x_key.replace('_', ' ').title())
ax.set_ylabel(y_key.replace('_', ' ').title())
plt.grid(True)
if GENERATE_PLOTS:
output_file = 'lockloss_vs_poweron_%s.%s' % (TARGET_DATE.strftime("%Y-%m-%d"), PLOT_SUFFIX)
fig.savefig(output_file, bbox_inches='tight')
print "Saved figure to", output_file
```
| github_jupyter |
# IST 718: Big Data Analytics
- Professor: Willard Williamson <wewillia@syr.edu>
- Faculty Assistant: Palaniappan Muthukkaruppan
## General instructions:
- You are welcome to discuss the problems with your classmates but __you are not allowed to copy any part of your answers from your classmates. Short code snippets are allowed from the internet. Any code is allowed from the class text books or class provided code.__
- Please do not change the file names. The FAs and the professor use these names to grade your homework.
- Remove or comment out code that contains `raise NotImplementedError`. This is mainly to make the `assert` statement fail if nothing is submitted.
- The tests shown in some cells (i.e., `assert` and `np.testing.` statements) are used to grade your answers. **However, the professor and FAs will use __additional__ test for your answer. Think about cases where your code should run even if it passess all the tests you see.**
- Before submitting your work through Blackboard, remember to save and press `Validate` (or go to
`Kernel`$\rightarrow$`Restart and Run All`).
```
# load these packages
from pyspark.ml import feature
from pyspark.ml import clustering
from pyspark.ml import Pipeline
from pyspark.sql import functions as fn
import numpy as np
import seaborn as sns
from pyspark.sql import SparkSession
from pyspark.ml import feature, regression, evaluation, Pipeline
from pyspark.sql import functions as fn, Row
import matplotlib.pyplot as plt
from pyspark.ml.feature import PCA
spark = SparkSession.builder.getOrCreate()
sc = spark.sparkContext
import pandas as pd
import os
```
The following cell is used to determine if the environment is databricks or personal computer and load the csv file accordingly.
```
def get_training_dataframe():
data_file_name = "colleges_data_science_programs.csv"
# get the databricks runtime version
db_env = os.getenv("DATABRICKS_RUNTIME_VERSION")
grading_env = os.getenv("GRADING_RUNTIME_ENV")
# if the databricks env var exists
if db_env != None:
full_path_name = "/FileStore/tables/%s" % data_file_name
elif grading_env != None:
full_path_name = "C:/Users/Will/Desktop/SU/datasets/%s" % data_file_name
else:
full_path_name = data_file_name
return spark.read.csv(full_path_name, inferSchema=True, header=True).fillna('').orderBy('id')
ds_programs_df = get_training_dataframe
```
# Unsupervised learning
It is recommended to follow the notebook `unsupervised_learning.ipynb`.
The following dataset contains information about dozens of "data science" programs across the US.
```
ds_programs_df = get_training_dataframe()
ds_programs_df.head()
```
## Question 1: (10 pts)
This dataset contains many columns that we can use to understand how these data science programs differ from one another. In this question, you will create a dataframe `ds_programs_text_df` which simply adds a column `text` to the dataframe `ds_programs_df`. This column will have the concatenation of the following columns separated by a space: `program`, `degree` and `department` (find the appropriate function in the `fn` package)
```
# (10 pts) Create ds_programs_text_df here
from pyspark.sql.functions import concat, col, lit
ds_programs_text_df = ds_programs_df.withColumn('text',concat(col("program"), lit(" "), col("degree"), lit(" "), col("department")))
#raise NotImplementedError()
```
An example of the `ds_programs_text_df` should give you:
```python
ds_programs_text_df.orderBy('id').first().text
```
```console
'Data Science Masters Mathematics and Statistics'
```
```
# (10 pts)
np.testing.assert_equal(ds_programs_text_df.count(), 222)
np.testing.assert_equal(set(ds_programs_text_df.columns), {'admit_reqs',
'business',
'capstone',
'cost',
'country',
'courses',
'created_at',
'databases',
'degree',
'department',
'ethics',
'id',
'machine learning',
'mapreduce',
'name',
'notes',
'oncampus',
'online',
'part-time',
'program',
'program_size',
'programminglanguages',
'state',
'text',
'university_count',
'updated_at',
'url',
'visualization',
'year_founded'})
np.testing.assert_array_equal(ds_programs_text_df.orderBy('id').rdd.map(lambda x: x.text).take(5),
['Data Science Masters Mathematics and Statistics',
'Analytics Masters Business and Information Systems',
'Data Science Masters Computer Science',
'Business Intelligence & Analytics Masters Business',
'Advanced Computer Science(Data Analytics) Masters Computer Science'])
```
# Question 2: (10 pts)
The following code creates a dataframe `ds_features_df` which adds a column `features` to `ds_programs_text_df` that contains the `tfidf` of the column `text`:
```
# read-only
pipe_features = \
Pipeline(stages=[
feature.Tokenizer(inputCol='text', outputCol='words'),
feature.CountVectorizer(inputCol='words', outputCol='tf'),
feature.IDF(inputCol='tf', outputCol='tfidf'),
feature.StandardScaler(withStd=False, withMean=True, inputCol='tfidf', outputCol='features')]).\
fit(ds_programs_text_df)
```
Create a pipeline model `pipe_pca` that computes the two first principal components of `features` as computed by `pipe_features` and outputs a column `scores`. Use that pipeline to create a dataframe `ds_features_df` with the columns `id`, `name`, `url`, and `scores`.
```
# create the pipe_pca PipelineModel below (10 pts)
pca = PCA(k=2, inputCol="features", outputCol="scores")
pipe_pca = Pipeline(stages=[pipe_features, pca]).fit(ds_programs_text_df)
ds_features_df = pipe_pca.transform(ds_programs_text_df).select('id', 'name', 'url','scores')
#raise NotImplementedError()
# Tests for (10 pts)
np.testing.assert_equal(pipe_pca.stages[0], pipe_features)
np.testing.assert_equal(type(pipe_pca.stages[1]), feature.PCAModel)
np.testing.assert_equal(set(ds_features_df.columns), {'id', 'name', 'scores', 'url'})
np.testing.assert_equal(ds_features_df.first().scores.shape, (2, ))
```
# Question 3: (10 pts)
Create a scatter plot with the x axis containing the first principal component (loading) and the y axis containing the second principal component (loading) of `ds_features_df`
```
# below perform the appropriate
loadings = pipe_pca.stages[-1].pc.toArray()
plt.figure(figsize=(8,8))
plot01 = plt.scatter(loadings[:,0], loadings[:,1], edgecolor='', alpha=0.5)
plt.title("PCA Scatter Plot")
plt.xlabel("Principal Component 1")
plt.ylabel("Principal Component 2")
display(plot01.figure)
#raise NotImplementedError()
```
# Question 4 (10 pts)
Create two Pandas dataframes `pc1_pd` and `pc2_pd` with the columns `word` and `abs_loading` that contain the top 5 words in absolute loading for the principal components 1 and 2, respetively. You can extract the vocabulary from the stage that contains the count vectorizer in `pipe_features`:
```
# your code here
pc1_pd = pd.DataFrame({'word' : pipe_features.stages[1].vocabulary,
'abs_loading' : np.abs(loadings[:,0])}).sort_values('abs_loading', ascending=False).head()
pc2_pd = pd.DataFrame({'word' : pipe_features.stages[1].vocabulary,
'abs_loading' : np.abs(loadings[:,1])}).sort_values('abs_loading', ascending=False).head()
#raise NotImplementedError()
pc1_pd
pc2_pd
# (10 pts)
assert type(pc1_pd) == pd.core.frame.DataFrame
assert type(pc2_pd) == pd.core.frame.DataFrame
np.testing.assert_array_equal(pc1_pd.shape, (5, 2))
np.testing.assert_array_equal(pc2_pd.shape, (5, 2))
np.testing.assert_equal(set(pc1_pd.columns), {'abs_loading', 'word'})
np.testing.assert_equal(set(pc2_pd.columns), {'abs_loading', 'word'})
```
# Question 5: (10 pts)
Create a new pipeline for PCA called `pipe_pca2` where you fit 50 principal components. Extract the the `PCAModel` from the stages of this pipeline, and assign to a variable `explainedVariance` the variance explained by components of such model. Finally, assign to a variable `best_k` the value $k$ such that ($k+1$)-th component is not able to explain more than 0.01 variance. You can use a for-loop to find such best k.
```
# your code here
pca1 = PCA(k=50, inputCol="features", outputCol="scores")
pipe_pca2 = Pipeline(stages=[pipe_features, pca1]).fit(ds_programs_text_df)
explainedVariance = pipe_pca2.stages[-1].explainedVariance
#raise NotImplementedError()
k = 0
for k in range(len(explainedVariance)):
if explainedVariance[k] < 0.01:
print(k)
break
best_k = k
best_k
# Tests for (10 pts)
np.testing.assert_equal(pipe_pca2.stages[0], pipe_features)
np.testing.assert_equal(type(pipe_pca2.stages[1]), feature.PCAModel)
np.testing.assert_equal(len(explainedVariance), 50)
np.testing.assert_array_less(5, best_k)
```
# Question 6: (10 pts)
Create a new pipeline for PCA called pipe_pca3 where you fit all possible principal components for this dataset. Extract the the PCAModel from the stages of this pipeline, and use the object property named explainedVariance to create 2 separate plots: A scree plot and a plot of cumulative variance explained. Use the plots to check your code in the above question, is you answer from the above question believable?
```
# your code here
pca2 = PCA(k=118, inputCol="features", outputCol="scores")
pipe_pca3 = Pipeline(stages=[pipe_features, pca2]).fit(ds_programs_text_df)
#raise NotImplementedError()
plt.figure()
explained_var = pipe_pca3.stages[-1].explainedVariance
plot02 = plt.scatter(np.arange(1, len(explained_var)+1), explained_var)
plt.title("DS Programs Explained Variance")
plt.xlabel('Number of components')
plt.ylabel('Eexplained variance')
display(plot02.figure)
cum_sum = np.cumsum(explained_var)
plt.figure()
cumsum_plot = plt.scatter(np.arange(1, len(explained_var)+1), cum_sum)
plt.title("DS Programs Cumulative Sum of Explained Variance")
plt.xlabel("Cumulative Components")
plt.ylabel("Cumulative Sum of Variance Explained")
display(cumsum_plot.figure)
```
# Question 7: (10 pts)
Create a pipeline named k_means_pipe below by adding a feature.Normalizer and a clustering.KMeans object onto the end of the pipe_features pipeline object. Set the seed of the KMeans clustering object to 2 and the number of clusters K to 5. Transform the new pipe on the ds_programs_text_df dataframe creating a new dataframe named k_means_df.
```
# your code here
norm = feature.Normalizer(inputCol="features", outputCol="norm_features", p=2.0)
kmeans = clustering.KMeans(k=5, seed=2, featuresCol='norm_features', predictionCol='kmeans_clust')
k_means_pipe = Pipeline(stages=[pipe_features, norm, kmeans]).fit(ds_programs_text_df)
k_means_df = k_means_pipe.transform(ds_programs_text_df)
#raise NotImplementedError()
```
The `kmeans_clust` col in the resulting datafame contains the cluster assignment. It's hard to visualize the clusters because we trained on all of the features in ds_programs_text_df.
```
# Show the head of the dataframe from above
display(k_means_df.toPandas().head())
np.testing.assert_equal(len(pd.unique(k_means_df.toPandas()['kmeans_clust'])), 5)
```
# Question 8: (10 pts)
The goal of this question is to make it easier to visualize the clusters by reducing the dimensionality of ds_programs_text_df to 2 dimensions using PCA. Replace the Normalizer in the k_means_pipe above with a PCA object that reduces dimensionality to 2 dimensions. Save the PCA output scores in a column named `kmeans_clust`. Transform the pipeline on ds_programs_text_df creating a new dataframe named k_means_pca_df. <br>
Define a function named get_kmeans_df which takes a parameter num_clusters, builds the pipeline described above using the parameter num_clusters to determine the number of KMeans clusters, and returns a pandas dataframe by transforming the pipeline on ds_programs_text_df.
```
# your code here
kmeans1 = clustering.KMeans(k=5, seed=2, featuresCol='scores', predictionCol='kmeans_clust')
k_means_pipe = Pipeline(stages=[pipe_features, pca, kmeans1]).fit(ds_programs_text_df)
k_means_pca_df = k_means_pipe.transform(ds_programs_text_df)
#raise NotImplementedError()
def get_kmeans_df(num_clusters):
kmeans = clustering.KMeans(k=num_clusters, seed=2, featuresCol='scores', predictionCol='kmeans_clust')
k_means_pipe = Pipeline(stages=[pipe_features, pca, kmeans]).fit(ds_programs_text_df)
df = k_means_pipe.transform(ds_programs_text_df).toPandas()
return df
# test
k_means_df = get_kmeans_df(3)
np.testing.assert_equal(k_means_df.shape[0], 222)
np.testing.assert_equal(k_means_df.shape[1], 35)
np.testing.assert_equal(len(pd.unique(k_means_df['kmeans_clust'])), 3)
```
# Question 9: (10 pts)
In the cell below, define a function named plot_kmeans which takes the dataframe returned by the get_kmeans_df function defined above as a single argument. The plot_kmeans function shall creates a scatter plot of the PCA scores from the input dataframe where each score point is colored by the cluster assignment. The plot title shall be the number of clusters in the dataframe. The X and Y axes shall have descriptive labels. The plot shall include a legend mapping colors to cluster number. Feel free to use whatever plotting technique you like but the seaborn scatterplot function is one easy way of creating this function.
```
# your code here
def plot_kmeans(df):
score1=[]
score2=[]
for i in range(len(df.scores)):
score1.append(df.scores[i][0])
score2.append(df.scores[i][1])
df['score1'] = score1
df['score2'] = score2
a = len(df["kmeans_clust"].unique())
colors = df["kmeans_clust"]
plt.figure(figsize=(8,8))
plot01 = sns.scatterplot(df['score1'], df['score2'], hue=colors, size=colors, palette="Set1")
plt.title("PCA Scatter Plot for " + str(a) + " Clusters")
plt.xlabel("Principal Component 1")
plt.ylabel("Principal Component 2")
return display(plot01.figure)
#raise NotImplementedError()
print(sns.__version__)
# test your plot function here
# The plot should have 3 clusters, a title indicating that K=3, a legend, and labeled X and Y axes
plot_kmeans(get_kmeans_df(3))
```
Create 5 cluster plots picking various K of your choice below using get_kmeans and plot_kmeans.
```
# your code here
plot_kmeans(get_kmeans_df(4))
#raise NotImplementedError()
plot_kmeans(get_kmeans_df(2))
plot_kmeans(get_kmeans_df(6))
plot_kmeans(get_kmeans_df(9))
```
# Question 10: (10 pts)
Think about the cumulative variance and the scree plot from the question above. What is one problem related to variance which might make the above KMeans not be an optimal choice using only 2 principal components.
Your Text Answer Here:
The above KMeans cannot be an optimal choice because of the amount of variance being captured by the first 2 principal components. Though the first two components posses the highest amount of variance in the data, we are still preserving less than 10% of the total variance of the data. In essence, we should preserve 90-95% of the total variance in data by choosing the number of components to be 50 so that we can preserve the variables that contribute most to the “cluster separation”.
```
```
| github_jupyter |
## OpenMC Mini Workshop at ASEMOT

**Place**: IIT Bombay
**Time**: Dec 15, 2018 from 1:30 to 3:30 pm
**Facilitators**: Jeremy Roberts and Richard Reed
## The Plan
| Topic | Duration |
|---------------------|----------|
| Ensure/Debug Remote Access | 15 min |
| Overview of OpenMC (key features) | 15 min |
| A basic pin cell (materials, geometries, settings, and statepoints) | 30 min |
| A basic assembly (universes, lattices, and tallies) | 45 min |
## Remote Access
Download `asemot.pem` from `https://github.com/corps-g/asemot_openmc`.
In `bash`, navigate to where you downloaded `asemot.pem` and enter
```
ssh -i "asemot.pem" ubuntu@ec2-54-197-16-236.compute-1.amazonaws.com
```
Everyone has the same username `ubuntu` but each registrant has a folder named based on email address. If your email is
```
jeremy@host.com
```
then your folder name is `jeremy`. Type `ls` to see all folders. Use `cd` to enter yours, which contains the two Python files we'll use as examples.
## Caveats and Acknowledgements
I am *not* an OpenMC developer, nor am I an *expert* user!
Much of the material included draws directly from a workshop provided by Paul Romano to the Canadian Nuclear Laboratories and the various examples provided by the OpenMC documentation.
## The Monte Carlo Method and the Life of a Neutron

### Tallying Along The Way
Often, analysis requires some quantity $X = \int d\mathbf{r} \int d\mathbf{\Omega} \int dE \;
f(\mathbf{r},\mathbf{\Omega},E) \psi (\mathbf{r},\mathbf{\Omega},E)$.
During simulation, estimate $\hat{X} = \frac{1}{N} \sum\limits_{i \in T} w_i \ell_i f_i$
With several histories and $\hat{X}_1, \hat{X}_2, \ldots$, compute
$$
\bar{X} = \frac{1}{N} {\sum\limits_{i=1}^N \hat{X}_i} \qquad \text{and} \qquad
s^2_X = \frac{1}{N-1} \left ( \frac{1}{N} {\sum\limits_{i=1}^N \hat{X}_i^2} -
\bar{X}^2 \right )
$$
## Monte Carlo and Criticality Calculations

## Quick Overview of OpenMC
### History and Status
- Development started in 2011 because
1. parallel scaling of existing codes was not great
2. export control made research continuity difficult
- DOE/NNSA approved open release in 2012
- Formally funded as of 2016
- 14+ active developers with contributions from at least 61 individuals
- Roughly 100k lines of C++, Fortran, and Python
- OpenMC is still considered to be in *beta*!!
### Where Do I Get Help?
User's Group: [openmc-users@googlegroups.com](openmc-users@googlegroups.com)
Developer's Group: [openmc-dev@googlegroups.com](openmc-dev@googlegroups.com)
Documentation: [http://openmc.readthedocs.org/en/latest/](http://openmc.readthedocs.org/en/latest/)
### OpenMC Inputs
Execution of OpenMC requires three XML files:
- `settings.xml`
- `materials.xml`
- `geometry.xml`
with optional features enabled by other files:
- `tallies.xml`
- `plots.xml`
### Python API
XML is good, but scripted inputs are better:
- All XML files automatically generated via Python scripts
- Rich interface to output data (tallies, etc.)
- Developmental interface to C API for in-memory, coupled-physics calculations
### Scalable Parallelism

(courtesy P. Romano)
### Handling Temperature Dependence
Because $\Sigma$ depends on $T$, and $T$ varies with $\mathbf{r}$, need 100's of GB's of data using a *brute force* approach.
**Alternatives**: polynomial fits (Yesilyurt and Martin), target motion sampling (Viitanen and Leppanen), and the **windowed-multipole** (WMP) approach (Hwang; revisited by Forget and Josey in OpenMC), based on the following:
$$
\begin{align*}
\sigma(u,0) &= \frac{1}{u^2} \sum\limits_j \Re \left [ \frac{r_j}{p_j - u}
\right ] \\ \sigma(u,T) &= \frac{1}{2u^2\sqrt{\xi}} \sum\limits_j \Re
\left [ ir_j \sqrt{\pi} W(z_j^0) - \frac{r_j}{\pi} C\left (
\frac{p_j}{\sqrt{\xi}}, \frac{u}{2\sqrt{\xi}} \right ) \right ]
\end{align*}
$$
Now, (relatively) little data required, and the problem becomes FLOP dominated (good for large, parallel machines with little memory! (See details in [Josey et al., J. Comp. Phys. **307** (2016)](https://dspace.mit.edu/openaccess-disseminate/1721.1/114236))
**Example**: Use WMP data to show impact of Doppler broadening on the absorption cross section of U-238 at 6.67 eV.
Data paths should be set up on the remote server.
For folks with laptops, show the cross_sections.xml file and how to append WMP data.
Note, WMP is not synced by default, nor does it handle thermal scattering. Hence, stick with 294 K for the rest of the workshop.
```
import openmc
import numpy as np
import matplotlib.pyplot as plt
path_u238_h5 = '/home/robertsj/Research/openmc/scripts/nndc_hdf5/U238.h5'
path_u238_mp = '/home/robertsj/Research/openmc/scripts/nndc_hdf5/wmp/092238.h5'
u238_h5 = openmc.data.IncidentNeutron.from_hdf5(path_u238_h5)
u238_mp = openmc.data.WindowedMultipole.from_hdf5(path_u238_mp)
E = np.linspace(5.5, 7.5, 1000)
plt.semilogy(E, u238_mp(E, 0)[1], 'k') # the "1" indexes sigma_a
plt.semilogy(E, u238_h5[102].xs['294K'](E), 'r') # mt=102 is the ENDF mt for sigma_a
plt.semilogy(E, u238_mp(E, 294)[1], 'g--')
plt.semilogy(E, u238_mp(E, 900)[1], 'b-.')
plt.legend(['0 K', '294 K (from ACE)', '294 K (from WMP)', '900 K'])
plt.xlabel('E (eV)')
plt.ylabel('$\sigma_a(E)$')
plt.show()
```
## Diving In: A Single Pin Cell
**Example**: compute $k_{\infty}$ for this fuel pin:

where the densities for fuel, cladding, and water are 10 g/cm$^3$, 6.6 g/cm$^3$, and 1 g/cm$^3$, respectively.
### Defining `materials.xml`
First, import `openmc`:
```
import openmc
```
Then, define a variable to represent the fuel material:
```
uo2 = openmc.Material(1, "uo2")
uo2.add_nuclide('U235', 0.04)
uo2.add_nuclide('U238', 0.96)
uo2.add_nuclide('O16', 2.0)
uo2.set_density('g/cm3', 10.0)
uo2
uo2
```
Now, do the same for Zr and water:
```
zirconium = openmc.Material(2, "zirconium")
zirconium.add_element('Zr', 1.0)
zirconium.set_density('g/cm3', 6.6)
zirconium # note automated elemental composition
h2o = openmc.Material(3, "h2o")
h2o.add_nuclide('H1', 2.0)
h2o.add_nuclide('O16', 1.0)
h2o.set_density('g/cm3', 1.0)
h2o.add_s_alpha_beta('c_H_in_H2O')
```
Now, add these materials to a material library and export the required XML file:
```
materials = openmc.Materials()
materials += [uo2, zirconium, h2o]
materials.export_to_xml()
!cat materials.xml # this calls "cat materials.xml" in bash
```
### Defining `geometry.xml`
OpenMC uses a combinatorial geometry similar to MCNP and Serpent.
Start by defining the surfaces needed to represent the fuel pin. First, the four sides of the box:
```
pitch = 1.26 # all dimensions in cm
left = openmc.XPlane(x0=-pitch/2, boundary_type='reflective')
right = openmc.XPlane(x0=pitch/2, boundary_type='reflective')
bottom = openmc.YPlane(y0=-pitch/2, boundary_type='reflective')
top = openmc.YPlane(y0=pitch/2, boundary_type='reflective')
```
Now, define the two cylinders:
```
fuel_outer = openmc.ZCylinder(R=0.41)
clad_outer = openmc.ZCylinder(R=0.48)
```
Now, define *cells* comprised of a region enclosed by surfaces and materials. For example, the fuel region is defined by everything "inside of" the `fuel_outer` surface and is filled with `uo2`:
```
fuel = openmc.Cell(1, 'fuel')
fuel.region = -fuel_outer # the "-" indicates "inside of"
fuel.fill = uo2
```
Similarly, cells for the cladding and coolant can be defined as follows:
```
cladding = openmc.Cell(2, 'cladding')
cladding.region = +fuel_outer & -clad_outer # + means "outside of" and "&" represents intersection
cladding.fill = zirconium
coolant = openmc.Cell(3, 'coolant')
coolant.region = +left & -right & +bottom & -top & +clad_outer
coolant.fill = h2o
```
Finally, define a root *universe* comprised of our cells, define a geometry with that root universe, and export to XML via:
```
root = openmc.Universe(cells=(fuel, cladding, coolant))
geom = openmc.Geometry(root)
geom.export_to_xml()
!cat geometry.xml
```
### Defining `settings.xml`
At a minimum, one must specific the number of (active) batches and particle histories per batch, but other settings include an initial source of neutrons, number of computational threads, etc.
```
settings = openmc.Settings()
settings.batches = 100
settings.inactive = 10
settings.particles = 1000
settings.export_to_xml()
!cat settings.xml
```
Now, we can execute OpenMC. In the terminal, run, e.g.,
```bash
robertsj@sampo ~/path/to/xmlfiles/ $ openmc
```
Or, we can run it right in Python:
```
!rm tallies.xml
!rm *h5
openmc.run()
```
### Looking at the Output
So, what is produced?
```
ls -al
```
The file `statepoint.100.h5` is an HDF5 file containing data associated with the first 100 batches. One can use `h5dump` to see more of it (the output is long). Rather, some useful information includes the following (shown by example):
```
import h5py
f = h5py.File('statepoint.100.h5', 'r') # open the file in read mode
print(list(f['/k_combined']))
f.close()
```
We can do the same thing with the Python API directly:
```
sp = openmc.StatePoint('statepoint.100.h5')
sp.k_combined.nominal_value, sp.k_combined.std_dev
```
### Visualizing the Geometry
We can define one additional XML file that OpenMC can use to generate a bitmap image of the geometry:
```
p = openmc.Plot()
p.filename = 'pin_cell'
p.width = (1.26, 1.26)
p.pixels = (100, 100)
p.color_by = 'material' # or 'cell'
plots = openmc.Plots([p])
plots.export_to_xml()
openmc.plot_geometry()
openmc.plot_inline(p)
```
### Additional Exercises
1. Write a Python script that uses the pin-cell model just created to produce a plot of $k_{\infty}$ as a function of the pitch-to-diameter ratio. For simplicity, keep the fuel and cladding radii constant while adjusting the pin pitch.
2. Plot `k_generations` from `statepoint.100.h5` as a function of generation. Also plot the running average value of `k_generations` as a function of generation. Does the final value match `k_combined`?
## Moving On: An Assembly of Fuel Pins
Let us produce a $4 \times 4$ assembly that uses the same basic fuel pin and looks like the following:

Then, we'll attempt two things:
1. Compute the pin power distribution
2. Compute two-group cross sections for this assembly
The materials are the *same* as for the pin cell, so the same `materials.xml` can be used.
Some of the geometry *does* change. In summary, these are needed:
- surfaces and cell to contain the entire $4 \times 4$ lattice
- surfaces, cells, and universes to define each unique lattice element (i.e., fuel pin or empty coolant channel)
- a $4\times 4$ lattice filled with these two universes
First, define the surfaces. Note that the `left`, `right`, `bottom`, and `top` surfaces will enclose the $4 \times 4$ lattice (centered at $(x, y) = (0, 0)$) with *reflective* conditions.
```
pitch = 1.26
left = openmc.XPlane(x0=-pitch*2, boundary_type='reflective')
right = openmc.XPlane(x0=pitch*2, boundary_type='reflective')
bottom = openmc.YPlane(y0=-pitch*2, boundary_type='reflective')
top = openmc.YPlane(y0=pitch*2, boundary_type='reflective')
fuel_outer = openmc.ZCylinder(R=0.41)
clad_outer = openmc.ZCylinder(R=0.48)
```
Now, define those cells needed for the fuel pin. Note, the `coolant` cell is *infinite* in extent, but that's okay: the lattice will impose boundaries!
```
fuel = openmc.Cell(1, 'fuel')
fuel.fill = uo2
fuel.region = -fuel_outer
cladding = openmc.Cell(2, 'cladding')
cladding.fill = zirconium #
cladding.region = +fuel_outer & -clad_outer
coolant = openmc.Cell(3, 'coolant')
coolant.fill = h2o
coolant.region = +clad_outer
```
Next, define the single cell needed for the empty coolant channel. Note, this one is *finite*, but its bounds are outside the area defined by one lattice element, so it's okay: the lattice will define the correct boundaries.
```
coolant_channel = openmc.Cell(4, 'coolant_channel')
coolant_channel.fill = h2o
coolant_channel.region = +left & -right & +bottom & -top # *first* use of these surfaces
```
Define the fuel and coolant universes:
```
univ_f = openmc.Universe(cells=[fuel, cladding, coolant], universe_id=1)
univ_c = openmc.Universe(cells=[coolant_channel], universe_id=2)
```
### Defining Lattices of Repeated Units
Next, define the $4\times 4$ lattice:
```
lattice2d = openmc.RectLattice(lattice_id=5)
lattice2d.lower_left = [-2*pitch, -2*pitch]
lattice2d.pitch = [pitch, pitch]
lattice2d.universes = [[univ_f, univ_f, univ_f, univ_f],
[univ_f, univ_c, univ_f, univ_f],
[univ_f, univ_f, univ_f, univ_f],
[univ_f, univ_f, univ_f, univ_f]]
```
Finally, create one cell to hold the lattice, set that as the cell for the root universe, and export the resulting geometry to XML:
```
lattice_cell = openmc.Cell(cell_id=999)
lattice_cell.region = +left & -right & +bottom & -top # second use of these surfaces
lattice_cell.fill = lattice2d
root = openmc.Universe(universe_id=0, name='root universe')
root.add_cells([lattice_cell])
geom = openmc.Geometry(root)
geom.export_to_xml()
```
It's always wise to check the geometry by visual inspection:
```
p = openmc.Plot()
p.filename = 'lattice_plot'
p.width = (4*pitch, 4*pitch)
p.pixels = (300, 300)
p.color_by = 'material'
plots = openmc.Plots([p])
plots.export_to_xml()
openmc.plot_geometry()
openmc.plot_inline(p)
```
### Defining `tallies.xml`
Although $k_{\text{eff}}$ is often important, most analysis requires determination of a quantity $X$ that can be represented as
$$
X = \underbrace{\int d\mathbf{r} \int d\mathbf{\Omega} \int dE}_{\text{filters}}
\underbrace{f(\mathbf{r}, \mathbf{\Omega} E)}_{\text{scores}} \psi(\mathbf{r}, \mathbf{\Omega}, E)\, .
$$
Possible **filters** include, among others, the selective integration over particular geometry cells and energy bins.
Possible **scores** (i.e., weighting functions) include the unit weight (for computing integral fluxes), various cross sections, and neutron/energy production from fission.
Now, define a tally for the two-group flux (boundary at 0.625 eV) integrated averaged over the entire lattice volume. First, an energy filter is needed:
```
energy_filter = openmc.EnergyFilter([0.0, 0.625, 2.0e7])
```
Now, create a new tally, add the filter, and define what to score:
```
flux_tally = openmc.Tally(tally_id=1)
flux_tally.filters = [energy_filter]
flux_tally.scores = ['flux']
```
Next, define a tally to compute the pin powers (as approximated by the fission density). This is most easily accomplished using a *mesh tally*, which is a tally whose spatial integration is defined by an arbitrary Cartesian mesh.
First, define the mesh. Each mesh element aligns exactly with a lattice element:
```
mesh = openmc.Mesh(mesh_id=1)
mesh.type = 'regular'
mesh.dimension = [4, 4, 1]
mesh.lower_left = [-2*pitch, -2*pitch, -1e7]
mesh.width = [pitch, pitch, 2e7]
```
Now, create the mesh filter and the mesh tally:
```
mesh_filter = openmc.MeshFilter(mesh)
mesh_tally = openmc.Tally(tally_id=2)
mesh_tally.filters = [mesh_filter]
mesh_tally.scores = ['fission']
```
With the tallies defined, export to `tallies.xml`:
```
tallies_file = openmc.Tallies([flux_tally, mesh_tally])
tallies_file.export_to_xml()
```
Update `settings.xml` to include *inactive* generations:
```
settings = openmc.Settings()
settings.batches = 100
settings.inactive = 10
settings.particles = 1000
settings.export_to_xml()
```
Run the simulation:
```
!rm *.h5 # clean up any existing output
openmc.run(threads=4)
```
Now, view `tallies.out`:
```
!cat tallies.out
```
## The End
Next steps: visit the OpenMC documentation, join the user group, and do the examples!
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Deploying a web service to Azure Kubernetes Service (AKS)
In this notebook, we show the following steps for deploying a web service using AzureML:
- Provision an AKS cluster (one time action)
- Deploy the service
- Test the web service
- Scale up the service
```
import json
import os
import subprocess
import numpy as np
import pandas as pd
import requests
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.webservice import Webservice, AksWebservice
from azureml.core.workspace import Workspace
from dotenv import set_key, get_key, find_dotenv
from utilities import text_to_json, get_auth
env_path = find_dotenv(raise_error_if_not_found=True)
```
AML will use the following information to create an image, provision a cluster and deploy a service. Replace the values in the following cell with your information.
```
image_name = get_key(env_path, 'image_name')
aks_service_name = "YOUR_AKS_SERVICE_NAME"
aks_name = "YOUR_AKS_NAME"
aks_location = "YOUR_AKS_LOCATION"
set_key(env_path, "aks_service_name", aks_service_name)
set_key(env_path, "aks_name", aks_name)
set_key(env_path, "aks_location", aks_location)
```
## Get workspace
Load existing workspace from the config file.
```
ws = Workspace.from_config(auth=get_auth(env_path))
print(ws.name, ws.resource_group, ws.location, sep="\n")
image = ws.images[image_name]
```
## Provision the AKS Cluster
This is a one time setup. You can reuse this cluster for multiple deployments after it has been created. If you delete the cluster or the resource group that contains it, then you would have to recreate it. Let's first check if there are enough cores in the subscription for the cluster .
```
vm_dict = {
"Dv2": {
"size": "Standard_D4_v2",
"cores": 8
}
}
vm_family = "Dv2"
node_count = 4
requested_cores = node_count * vm_dict[vm_family]["cores"]
results = subprocess.run([
"az", "vm", "list-usage",
"--location", get_key(env_path, "aks_location"),
"--query", "[?contains(localName, '%s')].{max:limit, current:currentValue}" % (vm_family)
], stdout=subprocess.PIPE)
quota = json.loads(''.join(results.stdout.decode('utf-8')))
diff = int(quota[0]['max']) - int(quota[0]['current'])
if diff <= requested_cores:
print("Not enough cores in region, asking for {} but have {}".format(requested_cores, diff))
raise Exception("Core Limit", "Note enough cores to satisfy request")
print("There are enough cores, you may continue...")
prov_config = AksCompute.provisioning_configuration(
agent_count=4, vm_size="Standard_D4_v2", location=aks_location
)
# Create the cluster
aks_target = ComputeTarget.create(
workspace=ws, name=aks_name, provisioning_configuration=prov_config
)
%%time
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
```
Let's check that the cluster is created successfully.
```
aks_status = aks_target.get_status()
assert aks_status == 'Succeeded', 'AKS failed to create'
```
## Deploy web service to AKS
Next, we deploy the web service. We deploy two pods with 1 CPU core each.
```
#Set the web service configuration
aks_config = AksWebservice.deploy_configuration(num_replicas=2, cpu_cores=1)
aks_service = Webservice.deploy_from_image(
workspace=ws,
name=aks_service_name,
image=image,
deployment_config=aks_config,
deployment_target=aks_target,
)
%%time
aks_service.wait_for_deployment(show_output=True)
print(aks_service.state)
```
You can check the logs of the web service with the below.
```
aks_service.get_logs()
```
## Test the web service
We now test the web sevice.
```
dupes_test_path = './data_folder/dupes_test.tsv'
dupes_test = pd.read_csv(dupes_test_path, sep='\t', encoding='latin1')
text_to_score = dupes_test.iloc[0,4]
text_to_score
jsontext = text_to_json(text_to_score)
%%time
prediction = aks_service.run(input_data = jsontext)
print(prediction)
```
Let's try a few more duplicate questions and display their top 3 original matches. Let's first get the scoring URL and and API key for the web service.
```
scoring_url = aks_service.scoring_uri
api_key = aks_service.get_keys()[0]
headers = {'content-type': 'application/json', 'Authorization':('Bearer '+ api_key)}
r = requests.post(scoring_url, data=jsontext, headers=headers) # Run the request twice since the first time takes a
%time r = requests.post(scoring_url, data=jsontext, headers=headers) # little longer due to the loading of the model
print(r)
r.json()
dupes_to_score = dupes_test.iloc[:5,4]
results = [
requests.post(scoring_url, data=text_to_json(text), headers=headers)
for text in dupes_to_score
]
```
Let's print top 3 matches for each duplicate question.
```
[eval(results[i].json())[0:3] for i in range(0, len(results))]
```
Next let's quickly check what the request response performance is for the deployed model on AKS cluster.
```
text_data = list(map(text_to_json, dupes_to_score)) # Retrieve the text data
timer_results = list()
for text in text_data:
res=%timeit -r 1 -o -q requests.post(scoring_url, data=text, headers=headers)
timer_results.append(res.best)
timer_results
print("Average time taken: {0:4.2f} ms".format(10 ** 3 * np.mean(timer_results)))
```
## Scaling
In this part, we scale the number of pods to make sure we fully utilize the AKS cluster. To connect to the Kubernetes cluster, we will use kubectl, the Kubernetes command-line client. To install, run the following:
```
!sudo az aks install-cli
```
Next, we will get the credentials to connect to the cluster.
```
os.makedirs(os.path.join(os.path.expanduser('~'),'.kube'), exist_ok=True)
config_path = os.path.join(os.path.expanduser('~'),'.kube/config')
with open(config_path, 'a') as f:
f.write(aks_target.get_credentials()['userKubeConfig'])
```
Let's check the nodes and pods of the cluster.
```
!kubectl get nodes
!kubectl get pods --all-namespaces
!kubectl get events
```
We can now scale up the number of pods.
```
!kubectl scale --current-replicas=2 --replicas=10 {"deployment/" + aks_service_name}
!kubectl get pods --all-namespaces
!kubectl get deployment
```
Next, we will test the [throughput of the web service](06_SpeedTestWebApp.ipynb).
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/1.4.Biomedical_NER_SparkNLP_paper_reproduce.ipynb)
# Biomedical Named Entity Recognition (NER) at Scale
official paper : https://arxiv.org/abs/2011.06315
NER in Spark NLP Colab notebooks: https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Public/4.NERDL_Training.ipynb
## Blogposts and videos:
https://towardsdatascience.com/named-entity-recognition-ner-with-bert-in-spark-nlp-874df20d1d77
NerDL worksghop (90 min): https://www.youtube.com/watch?v=YM-e4eOiQ34
https://medium.com/spark-nlp/named-entity-recognition-for-healthcare-with-sparknlp-nerdl-and-nercrf-a7751b6ad571
https://medium.com/atlas-research/ner-for-clinical-text-7c73caddd180
```
import sparknlp
spark = sparknlp.start() # for GPU training >> sparknlp.start(gpu = True)
from sparknlp.base import *
from sparknlp.annotator import *
print("Spark NLP version", sparknlp.version())
print("Apache Spark version:", spark.version)
spark
```
## CoNLL Data Prep
We are going to use NCBI-Disease dataset in this notebook. All the other datasets used in the official paper arw prepared using the datasets at https://github.com/cambridgeltl/MTL-Bioinformatics-2016/tree/master/data. Since the data preparation steps from official tsv dfiles to conll format we utilize here is a time consuming and proprietary process, we share only one of them (**NCBI-Disease**) as an example.
```
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NCBI_disease_official_test.conll
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/NCBI_disease_official_train_dev.conll
with open("NCBI_disease_official_test.conll") as f:
train_txt =f.read()
print (train_txt[:500])
from sparknlp.training import CoNLL
training_data = CoNLL().readDataset(spark, 'NCBI_disease_official_train_dev.conll')
training_data.show(3)
training_data.count()
import pyspark.sql.functions as F
training_data.select(F.explode(F.arrays_zip('token.result', 'pos.result', 'label.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("pos"),
F.expr("cols['2']").alias("ner_label")).show(truncate=50)
training_data.select(F.explode(F.arrays_zip('token.result','label.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth")).groupBy('ground_truth').count().orderBy('count', ascending=False).show(100,truncate=False)
test_data = CoNLL().readDataset(spark, 'NCBI_disease_official_test.conll')
test_data.select(F.explode(F.arrays_zip('token.result','label.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth")).groupBy('ground_truth').count().orderBy('count', ascending=False).show(100,truncate=False)
# You can use any word embeddings you want (Glove, Elmo, Bert, custom etc.)
glove_embeddings = WordEmbeddingsModel.pretrained('glove_6B_300','xx')\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
```
**We will get the embeddings for test set and write to disk as a parquet to be able to get metrics after each epoch during training.**
```
glove_embeddings.transform(test_data).write.parquet('test_data_with_embeddings.parquet')
!mkdir ner_logs
nerTagger = NerDLApproach()\
.setInputCols(["sentence", "token", "embeddings"])\
.setLabelColumn("label")\
.setOutputCol("ner")\
.setMaxEpochs(10)\
.setLr(0.003)\
.setBatchSize(8)\
.setRandomSeed(0)\
.setVerbose(1)\
.setEvaluationLogExtended(True) \
.setEnableOutputLogs(True)\
.setIncludeConfidence(True)\
.setTestDataset('test_data_with_embeddings.parquet')\
.setOutputLogsPath('ner_logs') # if not set, logs will be written to ~/annotator_logs
# .setGraphFolder('graphs') >> put your graph file (pb) under this folder if you are using a custom graph generated thru 4.1 NerDL-Graph.ipynb notebook
# .setEnableMemoryOptimizer() >> if you have a limited memory and a large conll file, you can set this True to train batch by batch
# do hyperparameter by tuning the params above (max epoch, LR, dropout etc.) to get better results
ner_pipeline = Pipeline(stages=[
glove_embeddings,
nerTagger
])
```
### Fitting
```
%%time
ner_model = ner_pipeline.fit(training_data)
# 1 epoch takes around 3 min with batch size=8
# if you get an error for incompatible TF graph, use 4.1 NerDL-Graph.ipynb notebook to create a graph
#! cd ~/annotator_logs && ls -lt
!cd ner_logs && ls -lt
!tail -n 10 ner_logs/NerDLApproach_af2879065348.log
```
### Test set evaluation
```
import pyspark.sql.functions as F
predictions = ner_model.transform(test_data)
predictions.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).show(truncate=False)
```
Licensed user will have an access to internal NERDLMetrics module to do this more efficient and easily without going out of Spark. But open source users need to use sklearn.metrics or any other equivalent module to do the same.
```
from sklearn.metrics import classification_report
preds_df = predictions.select(F.explode(F.arrays_zip('token.result','label.result','ner.result')).alias("cols")) \
.select(F.expr("cols['0']").alias("token"),
F.expr("cols['1']").alias("ground_truth"),
F.expr("cols['2']").alias("prediction")).toPandas()
print (classification_report(preds_df['ground_truth'], preds_df['prediction']))
```
### Entity level evaluation (strict eval)
```
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Public/utils/conll_eval.py
import conll_eval
metrics = conll_eval.evaluate(preds_df['ground_truth'].values, preds_df['prediction'].values)
# micro, macro, avg
metrics[0]
import pandas as pd
pd.DataFrame(metrics[1], columns=['entity','precision','recall','f1','support'])
```
### Ner log parser
```
!wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Public/utils/ner_log_parser.py
import ner_log_parser
%matplotlib inline
ner_log_parser.get_charts('/content/ner_logs/NerDLApproach_af2879065348.log')
ner_log_parser.loss_plot('/content/ner_logs/NerDLApproach_af2879065348.log')
```
### Saving the trained model
```
ner_model.stages
ner_model.stages[1].write().overwrite().save('NER_glove_e10_b8')
!ls -lt
```
## Prediction Pipeline
```
document = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')
token = Tokenizer()\
.setInputCols(['sentence'])\
.setOutputCol('token')
glove_embeddings = WordEmbeddingsModel.pretrained('glove_6B_300','xx')\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
loaded_ner_model = NerDLModel.load("NER_glove_e10_b8")\
.setInputCols(["sentence", "token", "embeddings"])\
.setOutputCol("ner")
converter = NerConverter()\
.setInputCols(["sentence", "token", "ner"])\
.setOutputCol("ner_span")
ner_prediction_pipeline = Pipeline(stages = [
document,
sentence,
token,
glove_embeddings,
loaded_ner_model,
converter
])
empty_data = spark.createDataFrame([['']]).toDF("text")
prediction_model = ner_prediction_pipeline.fit(empty_data)
text = '''
A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and subsequent type two diabetes mellitus ( T2DM ), one prior episode of HTG-induced pancreatitis three years prior to presentation , associated with an acute hepatitis , and obesity with a body mass index ( BMI ) of 33.5 kg/m2 , presented with a one-week history of polyuria , polydipsia , poor appetite , and vomiting .
'''
sample_data = spark.createDataFrame([[text]]).toDF("text")
sample_data.show(truncate=100)
preds = prediction_model.transform(sample_data)
preds.select(F.explode(F.arrays_zip("ner_span.result","ner_span.metadata")).alias("entities")) \
.select(F.expr("entities['0']").alias("chunk"),
F.expr("entities['1'].entity").alias("entity")).show(truncate=False)
from sparknlp.base import LightPipeline
light_model = LightPipeline(prediction_model)
result = light_model.annotate(text)
list(zip(result['token'], result['ner']))
import pandas as pd
result = light_model.fullAnnotate(text)
ner_df= pd.DataFrame([(int(x.metadata['sentence']), x.result, x.begin, x.end, y.result) for x,y in zip(result[0]["token"], result[0]["ner"])],
columns=['sent_id','token','start','end','ner'])
ner_df.head(20)
```
### Highlight Entities
```
ann_text = light_model.fullAnnotate(text)[0]
ann_text.keys()
from sparknlp_display import NerVisualizer
visualiser = NerVisualizer()
print ('Standard Output')
visualiser.display(ann_text, label_col='ner_span', document_col='document')
```
| github_jupyter |
# Programming Assignment:
## Готовим LDA по рецептам
Как вы уже знаете, в тематическом моделировании делается предположение о том, что для определения тематики порядок слов в документе не важен; об этом гласит гипотеза «мешка слов». Сегодня мы будем работать с несколько нестандартной для тематического моделирования коллекцией, которую можно назвать «мешком ингредиентов», потому что она состоит из рецептов блюд разных кухонь. Тематические модели ищут слова, которые часто вместе встречаются в документах, и составляют из них темы. Мы попробуем применить эту идею к рецептам и найти кулинарные «темы». Эта коллекция хороша тем, что не требует предобработки. Кроме того, эта задача достаточно наглядно иллюстрирует принцип работы тематических моделей.
Для выполнения заданий, помимо часто используемых в курсе библиотек, потребуются модули *json* и *gensim*. Первый входит в дистрибутив Anaconda, второй можно поставить командой
*pip install gensim*
Построение модели занимает некоторое время. На ноутбуке с процессором Intel Core i7 и тактовой частотой 2400 МГц на построение одной модели уходит менее 10 минут.
### Загрузка данных
Коллекция дана в json-формате: для каждого рецепта известны его id, кухня (cuisine) и список ингредиентов, в него входящих. Загрузить данные можно с помощью модуля json (он входит в дистрибутив Anaconda):
```
import json
with open("data/07_recipes.json") as f:
recipes = json.load(f)
print recipes[0]
```
### Составление корпуса
```
from gensim import corpora, models
import numpy as np
```
Наша коллекция небольшая, и целиком помещается в оперативную память. Gensim может работать с такими данными и не требует их сохранения на диск в специальном формате. Для этого коллекция должна быть представлена в виде списка списков, каждый внутренний список соответствует отдельному документу и состоит из его слов. Пример коллекции из двух документов:
[["hello", "world"], ["programming", "in", "python"]]
Преобразуем наши данные в такой формат, а затем создадим объекты corpus и dictionary, с которыми будет работать модель.
```
texts = [recipe["ingredients"] for recipe in recipes]
dictionary = corpora.Dictionary(texts) # составляем словарь
corpus = [dictionary.doc2bow(text) for text in texts] # составляем корпус документов
print texts[0]
print corpus[0]
```
У объекта dictionary есть полезная переменная dictionary.token2id, позволяющая находить соответствие между ингредиентами и их индексами.
### Обучение модели
Вам может понадобиться [документация](https://radimrehurek.com/gensim/models/ldamodel.html) LDA в gensim.
__Задание 1.__ Обучите модель LDA с 40 темами, установив количество проходов по коллекции 5 и оставив остальные параметры по умолчанию.
Затем вызовите метод модели *show_topics*, указав количество тем 40 и количество токенов 10, и сохраните результат (топы ингредиентов в темах) в отдельную переменную. Если при вызове метода *show_topics* указать параметр *formatted=True*, то топы ингредиентов будет удобно выводить на печать, если *formatted=False*, будет удобно работать со списком программно. Выведите топы на печать, рассмотрите темы, а затем ответьте на вопрос:
Сколько раз ингредиенты "salt", "sugar", "water", "mushrooms", "chicken", "eggs" встретились среди топов-10 всех 40 тем? При ответе __не нужно__ учитывать составные ингредиенты, например, "hot water".
Передайте 6 чисел в функцию save_answers1 и загрузите сгенерированный файл в форму.
У gensim нет возможности фиксировать случайное приближение через параметры метода, но библиотека использует numpy для инициализации матриц. Поэтому, по утверждению автора библиотеки, фиксировать случайное приближение нужно командой, которая написана в следующей ячейке. __Перед строкой кода с построением модели обязательно вставляйте указанную строку фиксации random.seed.__
```
np.random.seed(76543)
# здесь код для построения модели:
%time ldamodel = models.ldamodel.LdaModel(corpus=corpus, id2word=dictionary, num_topics=40, passes=5)
top_ingredients = ldamodel.show_topics(num_topics=40, num_words=10, formatted=False)
print ldamodel.show_topics(num_topics=40, num_words=10, formatted=True)[0]
top_ingredients[0][1][2]
from collections import Counter
ingredients = ["salt", "sugar", "water", "mushrooms", "chicken", "eggs"]
occurrences = [
ingr
for _, topic_top in top_ingredients
for topic_ingr, prob in topic_top
for ingr in ingredients
if ingr == topic_ingr
]
counter = Counter(occurrences)
result = [counter[ingr] for ingr in ingredients]
print counter
print result
def save_answers1(c_salt, c_sugar, c_water, c_mushrooms, c_chicken, c_eggs):
with open("out/07_cooking_lda_pa_01.txt", "w") as fout:
fout.write(" ".join([str(el) for el in [c_salt, c_sugar, c_water, c_mushrooms, c_chicken, c_eggs]]))
save_answers1(*result)
```
### Фильтрация словаря
В топах тем гораздо чаще встречаются первые три рассмотренных ингредиента, чем последние три. При этом наличие в рецепте курицы, яиц и грибов яснее дает понять, что мы будем готовить, чем наличие соли, сахара и воды. Таким образом, даже в рецептах есть слова, часто встречающиеся в текстах и не несущие смысловой нагрузки, и поэтому их не желательно видеть в темах. Наиболее простой прием борьбы с такими фоновыми элементами — фильтрация словаря по частоте. Обычно словарь фильтруют с двух сторон: убирают очень редкие слова (в целях экономии памяти) и очень частые слова (в целях повышения интерпретируемости тем). Мы уберем только частые слова.
```
import copy
dictionary2 = copy.deepcopy(dictionary)
```
__Задание 2.__ У объекта dictionary2 есть переменная *dfs* — это словарь, ключами которого являются id токена, а элементами — число раз, сколько слово встретилось во всей коллекции. Сохраните в отдельный список ингредиенты, которые встретились в коллекции больше 4000 раз. Вызовите метод словаря *filter_tokens*, подав в качестве первого аргумента полученный список популярных ингредиентов. Вычислите две величины: dict_size_before и dict_size_after — размер словаря до и после фильтрации.
Затем, используя новый словарь, создайте новый корпус документов, corpus2, по аналогии с тем, как это сделано в начале ноутбука. Вычислите две величины: corpus_size_before и corpus_size_after — суммарное количество ингредиентов в корпусе (для каждого документа вычислите число различных ингредиентов в нем и просуммируйте по всем документам) до и после фильтрации.
Передайте величины dict_size_before, dict_size_after, corpus_size_before, corpus_size_after в функцию save_answers2 и загрузите сгенерированный файл в форму.
```
freq_ingredients = [token_id for token_id, occur in dictionary2.dfs.items() if occur > 4000]
print len(freq_ingredients)
removed_ingr = [dictionary.id2token[ingr] for ingr in freq_ingredients]
dict_size_before = len(dictionary)
print dict_size_before
dictionary2.filter_tokens(bad_ids=freq_ingredients)
dict_size_after = len(dictionary2)
print dict_size_after
corpus2 = [dictionary2.doc2bow(text) for text in texts]
print texts[0]
print corpus2[0]
def count_corpus_size(corpus):
return sum([len(recipe) for recipe in corpus])
corpus_size_before = count_corpus_size(corpus)
corpus_size_after = count_corpus_size(corpus2)
def save_answers2(dict_size_before, dict_size_after, corpus_size_before, corpus_size_after):
with open("out/07_cooking_lda_pa_02.txt", "w") as fout:
fout.write(" ".join([str(el) for el in [dict_size_before, dict_size_after, corpus_size_before, corpus_size_after]]))
save_answers2(dict_size_before, dict_size_after, corpus_size_before, corpus_size_after)
```
### Сравнение когерентностей
__Задание 3.__ Постройте еще одну модель по корпусу corpus2 и словарю dictionary2, остальные параметры оставьте такими же, как при первом построении модели. Сохраните новую модель в другую переменную (не перезаписывайте предыдущую модель). Не забудьте про фиксирование seed!
Затем воспользуйтесь методом *top_topics* модели, чтобы вычислить ее когерентность. Передайте в качестве аргумента соответствующий модели корпус. Метод вернет список кортежей (топ токенов, когерентность), отсортированных по убыванию последней. Вычислите среднюю по всем темам когерентность для каждой из двух моделей и передайте в функцию save_answers3.
```
np.random.seed(76543)
# здесь код для построения модели:
%time ldamodel2 = models.ldamodel.LdaModel(corpus=corpus2, id2word=dictionary2, num_topics=40, passes=5)
coherence = ldamodel.top_topics(corpus)
coherence2 = ldamodel2.top_topics(corpus2)
print coherence2[2]
np.array([coh for _, coh in coherence])
def get_avg_coherence(coherence):
return np.mean(np.array([coh for _, coh in coherence]))
avg_coherence = get_avg_coherence(coherence)
avg_coherence2 = get_avg_coherence(coherence2)
def save_answers3(coherence, coherence2):
with open("out/07_cooking_lda_pa_03.txt", "w") as fout:
fout.write(" ".join(["%3f"%el for el in [coherence, coherence2]]))
save_answers3(avg_coherence, avg_coherence2)
```
Считается, что когерентность хорошо соотносится с человеческими оценками интерпретируемости тем. Поэтому на больших текстовых коллекциях когерентность обычно повышается, если убрать фоновую лексику. Однако в нашем случае этого не произошло.
### Изучение влияния гиперпараметра alpha
В этом разделе мы будем работать со второй моделью, то есть той, которая построена по сокращенному корпусу.
Пока что мы посмотрели только на матрицу темы-слова, теперь давайте посмотрим на матрицу темы-документы. Выведите темы для нулевого (или любого другого) документа из корпуса, воспользовавшись методом *get_document_topics* второй модели:
```
texts[:1]
list(ldamodel2.get_document_topics(corpus2[:3]))
```
Также выведите содержимое переменной *.alpha* второй модели:
```
ldamodel2.alpha
```
У вас должно получиться, что документ характеризуется небольшим числом тем. Попробуем поменять гиперпараметр alpha, задающий априорное распределение Дирихле для распределений тем в документах.
__Задание 4.__ Обучите третью модель: используйте сокращенный корпус (corpus2 и dictionary2) и установите параметр __alpha=1__, passes=5. Не забудьте про фиксацию seed! Выведите темы новой модели для нулевого документа; должно получиться, что распределение над множеством тем практически равномерное. Чтобы убедиться в том, что во второй модели документы описываются гораздо более разреженными распределениями, чем в третьей, посчитайте суммарное количество элементов, __превосходящих 0.01__, в матрицах темы-документы обеих моделей. Другими словами, запросите темы модели для каждого документа с параметром *minimum_probability=0.01* и просуммируйте число элементов в получаемых массивах. Передайте две суммы (сначала для модели с alpha по умолчанию, затем для модели в alpha=1) в функцию save_answers4.
```
np.random.seed(76543)
# здесь код для построения модели:
%time ldamodel3 = models.ldamodel.LdaModel(corpus=corpus2, id2word=dictionary2, num_topics=40, passes=5, alpha=1)
list(ldamodel3.get_document_topics(corpus2[0]))[:5]
def get_document_topics_count(model, corpus, min_prob=0.01):
probs = model.get_document_topics(corpus, minimum_probability=min_prob)
return sum([len(lst) for lst in probs])
def save_answers4(count_model2, count_model3):
with open("out/07_cooking_lda_pa_04.txt", "w") as fout:
fout.write(" ".join([str(el) for el in [count_model2, count_model3]]))
save_answers4(
get_document_topics_count(ldamodel2, corpus2),
get_document_topics_count(ldamodel3, corpus2)
)
```
Таким образом, гиперпараметр __alpha__ влияет на разреженность распределений тем в документах. Аналогично гиперпараметр __eta__ влияет на разреженность распределений слов в темах.
### LDA как способ понижения размерности
Иногда, распределения над темами, найденные с помощью LDA, добавляют в матрицу объекты-признаки как дополнительные, семантические, признаки, и это может улучшить качество решения задачи. Для простоты давайте просто обучим классификатор рецептов на кухни на признаках, полученных из LDA, и измерим точность (accuracy).
__Задание 5.__ Используйте модель, построенную по сокращенной выборке с alpha по умолчанию (вторую модель). Составьте матрицу $\Theta = p(t|d)$ вероятностей тем в документах; вы можете использовать тот же метод get_document_topics, а также вектор правильных ответов y (в том же порядке, в котором рецепты идут в переменной recipes). Создайте объект RandomForestClassifier со 100 деревьями, с помощью функции cross_val_score вычислите среднюю accuracy по трем фолдам (перемешивать данные не нужно) и передайте в функцию save_answers5.
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
doc_topics = ldamodel2.get_document_topics(corpus2)
len(corpus2)
np.tile(3, 4)
ndocs = len(corpus2)
i = [None] * ndocs
j = [None] * ndocs
d = [None] * ndocs
for ind, doc in enumerate(doc_topics):
i[ind] = [ind] * len(doc)
j[ind] = [jj for jj, _ in doc]
d[ind] = [prob for _, prob in doc]
i = np.hstack(i)
j = np.hstack(j)
d = np.hstack(d)
for ind in range(5):
print i[ind], j[ind], d[ind]
from scipy.sparse import coo_matrix
X = coo_matrix((d, (i, j)))
cuisines = {r['cuisine'] for r in recipes}
cuisine2id = {c:i for i, c in enumerate(cuisines)}
id2cuisine = {v:k for k, v in cuisine2id.items()}
print cuisine2id
y = [cuisine2id[r['cuisine']] for r in recipes]
y[:10]
rfclass = RandomForestClassifier(n_estimators=100)
%time result = cross_val_score(rfclass, X, y, cv=3, n_jobs=3, error_score='accuracy')
result
def save_answers5(accuracy):
with open("out/07_cooking_lda_pa_05.txt", "w") as fout:
fout.write(str(accuracy))
save_answers5(result.mean())
```
Для такого большого количества классов это неплохая точность. Вы можете попроовать обучать RandomForest на исходной матрице частот слов, имеющей значительно большую размерность, и увидеть, что accuracy увеличивается на 10–15%. Таким образом, LDA собрал не всю, но достаточно большую часть информации из выборки, в матрице низкого ранга.
### LDA — вероятностная модель
Матричное разложение, использующееся в LDA, интерпретируется как следующий процесс генерации документов.
Для документа $d$ длины $n_d$:
1. Из априорного распределения Дирихле с параметром alpha сгенерировать распределение над множеством тем: $\theta_d \sim Dirichlet(\alpha)$
1. Для каждого слова $w = 1, \dots, n_d$:
1. Сгенерировать тему из дискретного распределения $t \sim \theta_{d}$
1. Сгенерировать слово из дискретного распределения $w \sim \phi_{t}$.
Подробнее об этом в [Википедии](https://en.wikipedia.org/wiki/Latent_Dirichlet_allocation).
В контексте нашей задачи получается, что, используя данный генеративный процесс, можно создавать новые рецепты. Вы можете передать в функцию модель и число ингредиентов и сгенерировать рецепт :)
```
def generate_recipe(model, num_ingredients):
theta = np.random.dirichlet(model.alpha)
for i in range(num_ingredients):
t = np.random.choice(np.arange(model.num_topics), p=theta)
topic = model.show_topic(t, topn=model.num_terms)
topic_distr = [x[1] for x in topic]
terms = [x[0] for x in topic]
w = np.random.choice(terms, p=topic_distr)
print w
generate_recipe(ldamodel2, 10)
```
### Интерпретация построенной модели
Вы можете рассмотреть топы ингредиентов каждой темы. Большиснтво тем сами по себе похожи на рецепты; в некоторых собираются продукты одного вида, например, свежие фрукты или разные виды сыра.
Попробуем эмпирически соотнести наши темы с национальными кухнями (cuisine). Построим матрицу $A$ размера темы $x$ кухни, ее элементы $a_{tc}$ — суммы $p(t|d)$ по всем документам $d$, которые отнесены к кухне $c$. Нормируем матрицу на частоты рецептов по разным кухням, чтобы избежать дисбаланса между кухнями. Следующая функция получает на вход объект модели, объект корпуса и исходные данные и возвращает нормированную матрицу $A$. Ее удобно визуализировать с помощью seaborn.
```
import pandas
import seaborn
from matplotlib import pyplot as plt
%matplotlib inline
def compute_topic_cuisine_matrix(model, corpus, recipes):
# составляем вектор целевых признаков
targets = list(set([recipe["cuisine"] for recipe in recipes]))
# составляем матрицу
tc_matrix = pandas.DataFrame(data=np.zeros((model.num_topics, len(targets))), columns=targets)
for recipe, bow in zip(recipes, corpus):
recipe_topic = model.get_document_topics(bow)
for t, prob in recipe_topic:
tc_matrix[recipe["cuisine"]][t] += prob
# нормируем матрицу
target_sums = pandas.DataFrame(data=np.zeros((1, len(targets))), columns=targets)
for recipe in recipes:
target_sums[recipe["cuisine"]] += 1
return pandas.DataFrame(tc_matrix.values/target_sums.values, columns=tc_matrix.columns)
def plot_matrix(tc_matrix):
plt.figure(figsize=(10, 10))
seaborn.heatmap(tc_matrix, square=True)
# Визуализируйте матрицу
plot_matrix(compute_topic_cuisine_matrix(ldamodel2, corpus2, recipes))
```
Чем темнее квадрат в матрице, тем больше связь этой темы с данной кухней. Мы видим, что у нас есть темы, которые связаны с несколькими кухнями. Такие темы показывают набор ингредиентов, которые популярны в кухнях нескольких народов, то есть указывают на схожесть кухонь этих народов. Некоторые темы распределены по всем кухням равномерно, они показывают наборы продуктов, которые часто используются в кулинарии всех стран.
Жаль, что в датасете нет названий рецептов, иначе темы было бы проще интерпретировать...
### Заключение
В этом задании вы построили несколько моделей LDA, посмотрели, на что влияют гиперпараметры модели и как можно использовать построенную модель.
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import pycountry
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_enable_multi_backend_download = \
os.environ.get("RADARCOVID_REPORT__ENABLE_MULTI_BACKEND_DOWNLOAD")
if environment_enable_multi_backend_download:
report_backend_identifiers = None
else:
report_backend_identifiers = [report_backend_identifier]
report_backend_identifiers
environment_invalid_shared_diagnoses_dates = \
os.environ.get("RADARCOVID_REPORT__INVALID_SHARED_DIAGNOSES_DATES")
if environment_invalid_shared_diagnoses_dates:
invalid_shared_diagnoses_dates = environment_invalid_shared_diagnoses_dates.split(",")
else:
invalid_shared_diagnoses_dates = []
invalid_shared_diagnoses_dates
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe():
return pd.read_csv("https://raw.githubusercontent.com/owid/covid-19-data/master/public/data/owid-covid-data.csv")
confirmed_df_ = download_cases_dataframe()
confirmed_df_.iloc[0]
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["date", "new_cases", "iso_code"]]
confirmed_df.rename(
columns={
"date": "sample_date",
"iso_code": "country_code",
},
inplace=True)
def convert_iso_alpha_3_to_alpha_2(x):
try:
return pycountry.countries.get(alpha_3=x).alpha_2
except Exception as e:
logging.info(f"Error converting country ISO Alpha 3 code '{x}': {repr(e)}")
return None
confirmed_df["country_code"] = confirmed_df.country_code.apply(convert_iso_alpha_3_to_alpha_2)
confirmed_df.dropna(inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
confirmed_days_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
confirmed_days_df["sample_date_string"] = \
confirmed_days_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_days_df.tail()
def sort_source_regions_for_display(source_regions: list) -> list:
if report_backend_identifier in source_regions:
source_regions = [report_backend_identifier] + \
list(sorted(set(source_regions).difference([report_backend_identifier])))
else:
source_regions = list(sorted(source_regions))
return source_regions
report_source_regions = report_backend_client.source_regions_for_date(
date=extraction_datetime.date())
report_source_regions = sort_source_regions_for_display(
source_regions=report_source_regions)
report_source_regions
def get_cases_dataframe(source_regions_for_date_function, columns_suffix=None):
source_regions_at_date_df = confirmed_days_df.copy()
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: source_regions_for_date_function(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df["_source_regions_group"] = source_regions_at_date_df. \
source_regions_at_date.apply(lambda x: ",".join(sort_source_regions_for_display(x)))
source_regions_at_date_df.tail()
#%%
source_regions_for_summary_df_ = \
source_regions_at_date_df[["sample_date", "_source_regions_group"]].copy()
source_regions_for_summary_df_.rename(columns={"_source_regions_group": "source_regions"}, inplace=True)
source_regions_for_summary_df_.tail()
#%%
confirmed_output_columns = ["sample_date", "new_cases", "covid_cases"]
confirmed_output_df = pd.DataFrame(columns=confirmed_output_columns)
for source_regions_group, source_regions_group_series in \
source_regions_at_date_df.groupby("_source_regions_group"):
source_regions_set = set(source_regions_group.split(","))
confirmed_source_regions_set_df = \
confirmed_df[confirmed_df.country_code.isin(source_regions_set)].copy()
confirmed_source_regions_group_df = \
confirmed_source_regions_set_df.groupby("sample_date").new_cases.sum() \
.reset_index().sort_values("sample_date")
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df.merge(
confirmed_days_df[["sample_date_string"]].rename(
columns={"sample_date_string": "sample_date"}),
how="right")
confirmed_source_regions_group_df["new_cases"] = \
confirmed_source_regions_group_df["new_cases"].clip(lower=0)
confirmed_source_regions_group_df["covid_cases"] = \
confirmed_source_regions_group_df.new_cases.rolling(7, min_periods=0).mean().round()
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[confirmed_output_columns]
confirmed_source_regions_group_df = confirmed_source_regions_group_df.replace(0, np.nan)
confirmed_source_regions_group_df.fillna(method="ffill", inplace=True)
confirmed_source_regions_group_df = \
confirmed_source_regions_group_df[
confirmed_source_regions_group_df.sample_date.isin(
source_regions_group_series.sample_date_string)]
confirmed_output_df = confirmed_output_df.append(confirmed_source_regions_group_df)
result_df = confirmed_output_df.copy()
result_df.tail()
#%%
result_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
result_df = confirmed_days_df[["sample_date_string"]].merge(result_df, how="left")
result_df.sort_values("sample_date_string", inplace=True)
result_df.fillna(method="ffill", inplace=True)
result_df.tail()
#%%
result_df[["new_cases", "covid_cases"]].plot()
if columns_suffix:
result_df.rename(
columns={
"new_cases": "new_cases_" + columns_suffix,
"covid_cases": "covid_cases_" + columns_suffix},
inplace=True)
return result_df, source_regions_for_summary_df_
confirmed_eu_df, source_regions_for_summary_df = get_cases_dataframe(
report_backend_client.source_regions_for_date)
confirmed_es_df, _ = get_cases_dataframe(
lambda date: [spain_region_country_code],
columns_suffix=spain_region_country_code.lower())
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
base_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=base_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_base_df = tek_list_df[tek_list_df.region == report_backend_identifier]
tek_list_base_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_base_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_base_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_base_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df \
.groupby(["upload_date"]).shared_teks.max().reset_index() \
.sort_values(["upload_date"], ascending=False) \
.rename(columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
invalid_shared_diagnoses_dates_mask = \
estimated_shared_diagnoses_df.sample_date_string.isin(invalid_shared_diagnoses_dates)
estimated_shared_diagnoses_df[invalid_shared_diagnoses_dates_mask] = 0
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Official Statistics
```
import requests
import pandas.io.json
official_stats_response = requests.get("https://radarcovid.covid19.gob.es/kpi/statistics/basics")
official_stats_response.raise_for_status()
official_stats_df_ = pandas.io.json.json_normalize(official_stats_response.json())
official_stats_df = official_stats_df_.copy()
official_stats_df["date"] = pd.to_datetime(official_stats_df["date"], dayfirst=True)
official_stats_df.head()
official_stats_column_map = {
"date": "sample_date",
"applicationsDownloads.totalAcummulated": "app_downloads_es_accumulated",
"communicatedContagions.totalAcummulated": "shared_diagnoses_es_accumulated",
}
accumulated_suffix = "_accumulated"
accumulated_values_columns = \
list(filter(lambda x: x.endswith(accumulated_suffix), official_stats_column_map.values()))
interpolated_values_columns = \
list(map(lambda x: x[:-len(accumulated_suffix)], accumulated_values_columns))
official_stats_df = \
official_stats_df[official_stats_column_map.keys()] \
.rename(columns=official_stats_column_map)
official_stats_df["extraction_date"] = extraction_date
official_stats_df.head()
official_stats_path = "Data/Statistics/Current/RadarCOVID-Statistics.json"
previous_official_stats_df = pd.read_json(official_stats_path, orient="records", lines=True)
previous_official_stats_df["sample_date"] = pd.to_datetime(previous_official_stats_df["sample_date"], dayfirst=True)
official_stats_df = official_stats_df.append(previous_official_stats_df)
official_stats_df.head()
official_stats_df = official_stats_df[~(official_stats_df.shared_diagnoses_es_accumulated == 0)]
official_stats_df.sort_values("extraction_date", ascending=False, inplace=True)
official_stats_df.drop_duplicates(subset=["sample_date"], keep="first", inplace=True)
official_stats_df.head()
official_stats_stored_df = official_stats_df.copy()
official_stats_stored_df["sample_date"] = official_stats_stored_df.sample_date.dt.strftime("%Y-%m-%d")
official_stats_stored_df.to_json(official_stats_path, orient="records", lines=True)
official_stats_df.drop(columns=["extraction_date"], inplace=True)
official_stats_df = confirmed_days_df.merge(official_stats_df, how="left")
official_stats_df.sort_values("sample_date", ascending=False, inplace=True)
official_stats_df.head()
official_stats_df[accumulated_values_columns] = \
official_stats_df[accumulated_values_columns] \
.astype(float).interpolate(limit_area="inside")
official_stats_df[interpolated_values_columns] = \
official_stats_df[accumulated_values_columns].diff(periods=-1)
official_stats_df.drop(columns="sample_date", inplace=True)
official_stats_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
official_stats_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_eu_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df = confirmed_es_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df = result_summary_df.merge(source_regions_for_summary_df, how="left")
result_summary_df.set_index(["sample_date", "source_regions"], inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(result_summary_df.shared_diagnoses_es / result_summary_df.covid_cases_es).fillna(0)
result_summary_df.head(daily_plot_days)
def compute_aggregated_results_summary(days) -> pd.DataFrame:
aggregated_result_summary_df = result_summary_df.copy()
aggregated_result_summary_df["covid_cases_for_ratio"] = \
aggregated_result_summary_df.covid_cases.mask(
aggregated_result_summary_df.shared_diagnoses == 0, 0)
aggregated_result_summary_df["covid_cases_for_ratio_es"] = \
aggregated_result_summary_df.covid_cases_es.mask(
aggregated_result_summary_df.shared_diagnoses_es == 0, 0)
aggregated_result_summary_df = aggregated_result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(days).agg({
"covid_cases": "sum",
"covid_cases_es": "sum",
"covid_cases_for_ratio": "sum",
"covid_cases_for_ratio_es": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum",
"shared_diagnoses_es": "sum",
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
aggregated_result_summary_df = aggregated_result_summary_df.fillna(0).astype(int)
aggregated_result_summary_df["teks_per_shared_diagnosis"] = \
(aggregated_result_summary_df.shared_teks_by_upload_date /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case"] = \
(aggregated_result_summary_df.shared_diagnoses /
aggregated_result_summary_df.covid_cases_for_ratio).fillna(0)
aggregated_result_summary_df["shared_diagnoses_per_covid_case_es"] = \
(aggregated_result_summary_df.shared_diagnoses_es /
aggregated_result_summary_df.covid_cases_for_ratio_es).fillna(0)
return aggregated_result_summary_df
aggregated_result_with_7_days_window_summary_df = compute_aggregated_results_summary(days=7)
aggregated_result_with_7_days_window_summary_df.head()
last_7_days_summary = aggregated_result_with_7_days_window_summary_df.to_dict(orient="records")[1]
last_7_days_summary
aggregated_result_with_14_days_window_summary_df = compute_aggregated_results_summary(days=13)
last_14_days_summary = aggregated_result_with_14_days_window_summary_df.to_dict(orient="records")[1]
last_14_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"source_regions": "Source Countries",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases (Source Countries)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date (Source Countries)",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date (Source Countries)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date (Source Countries)",
"shared_diagnoses": "Shared Diagnoses (Source Countries – Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis (Source Countries)",
"shared_diagnoses_per_covid_case": "Usage Ratio (Source Countries)",
"covid_cases_es": "COVID-19 Cases (Spain)",
"app_downloads_es": "App Downloads (Spain – Official)",
"shared_diagnoses_es": "Shared Diagnoses (Spain – Official)",
"shared_diagnoses_per_covid_case_es": "Usage Ratio (Spain)",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
"covid_cases_es",
"app_downloads_es",
"shared_diagnoses_es",
"shared_diagnoses_per_covid_case_es",
]
summary_percentage_columns= [
"shared_diagnoses_per_covid_case_es",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.droplevel(level=["source_regions"]) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 30), legend=False)
ax_ = summary_ax_list[0]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
for percentage_column in summary_percentage_columns:
percentage_column_index = summary_columns.index(percentage_column)
summary_ax_list[percentage_column_index].yaxis \
.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(12, 1 + 0.6 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}" if x != 0 else "",
display_column_name_mapping["shared_diagnoses_per_covid_case_es"]: lambda x: f"{x:.2%}" if x != 0 else "",
}
general_columns = \
list(filter(lambda x: x not in display_formatters, display_column_name_mapping.values()))
general_formatter = lambda x: f"{x}" if x != 0 else ""
display_formatters.update(dict(map(lambda x: (x, general_formatter), general_columns)))
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index.get_level_values("sample_date") == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.item()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.item()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.item()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.item()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.item()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.item()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
if len(report_source_regions) == 1:
display_brief_source_regions = report_source_regions[0]
else:
display_brief_source_regions = f"{len(report_source_regions)} 🇪🇺"
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
df = df.copy()
df_styler = df.style.format(display_formatters)
media_path = get_temporary_image_path()
dfi.export(df_styler, media_path)
return media_path
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
def generate_summary_api_results(df: pd.DataFrame) -> list:
api_df = df.reset_index().copy()
api_df["sample_date_string"] = \
api_df["sample_date"].dt.strftime("%Y-%m-%d")
api_df["source_regions"] = \
api_df["source_regions"].apply(lambda x: x.split(","))
return api_df.to_dict(orient="records")
summary_api_results = \
generate_summary_api_results(df=result_summary_df)
today_summary_api_results = \
generate_summary_api_results(df=extraction_date_result_summary_df)[0]
summary_results = dict(
backend_identifier=report_backend_identifier,
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=today_summary_api_results,
last_7_days=last_7_days_summary,
last_14_days=last_14_days_summary,
daily_results=summary_api_results)
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
def format_shared_diagnoses_per_covid_case(value) -> str:
if value == 0:
return "–"
return f"≤{value:.2%}"
display_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=shared_diagnoses_per_covid_case)
display_last_14_days_shared_diagnoses_per_covid_case = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case"])
display_last_14_days_shared_diagnoses_per_covid_case_es = \
format_shared_diagnoses_per_covid_case(value=last_14_days_summary["shared_diagnoses_per_covid_case_es"])
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: {display_shared_diagnoses_per_covid_case}
Last 14 Days:
- Usage Ratio (Estimation): {display_last_14_days_shared_diagnoses_per_covid_case}
- Usage Ratio (Official): {display_last_14_days_shared_diagnoses_per_covid_case_es}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
**Beer Production Forecasting**
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Evaluate](#Evaluate)
## Introduction
This notebook demonstrates demand forecasting for Beer Production Dataset using AutoML.
AutoML highlights here include using Deep Learning forecasts, Arima, Prophet, Remote Execution and Remote Inferencing, and working with the `forecast` function. Please also look at the additional forecasting notebooks, which document lagging, rolling windows, forecast quantiles, other ways to use the forecast function, and forecaster deployment.
Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.
Notebook synopsis:
1. Creating an Experiment in an existing Workspace
2. Configuration and remote run of AutoML for a time-series model exploring Regression learners, Arima, Prophet and DNNs
4. Evaluating the fitted model using a rolling test
## Setup
```
import os
import azureml.core
import pandas as pd
import numpy as np
import logging
import warnings
from pandas.tseries.frequencies import to_offset
# Squash warning messages for cleaner output in the notebook
warnings.showwarning = lambda *args, **kwargs: None
from azureml.core.workspace import Workspace
from azureml.core.experiment import Experiment
from azureml.train.automl import AutoMLConfig
from matplotlib import pyplot as plt
from sklearn.metrics import mean_absolute_error, mean_squared_error
from azureml.train.estimator import Estimator
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.35.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
```
As part of the setup you have already created a <b>Workspace</b>. To run AutoML, you also need to create an <b>Experiment</b>. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.
```
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'beer-remote-cpu'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
### Using AmlCompute
You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you use `AmlCompute` as your training compute resource.
> Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "beer-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS12_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Data
Read Beer demand data from file, and preview data.
Let's set up what we know about the dataset.
**Target column** is what we want to forecast.
**Time column** is the time axis along which to predict.
**Time series identifier columns** are identified by values of the columns listed `time_series_id_column_names`, for example "store" and "item" if your data has multiple time series of sales, one series for each combination of store and item sold.
**Forecast frequency (freq)** This optional parameter represents the period with which the forecast is desired, for example, daily, weekly, yearly, etc. Use this parameter for the correction of time series containing irregular data points or for padding of short time series. The frequency needs to be a pandas offset alias. Please refer to [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) for more information.
This dataset has only one time series. Please see the [orange juice notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales) for an example of a multi-time series dataset.
```
import pandas as pd
from pandas import DataFrame
from pandas import Grouper
from pandas import concat
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
plt.figure(figsize=(20, 10))
plt.tight_layout()
plt.subplot(2, 1, 1)
plt.title('Beer Production By Year')
df = pd.read_csv("Beer_no_valid_split_train.csv", parse_dates=True, index_col= 'DATE').drop(columns='grain')
test_df = pd.read_csv("Beer_no_valid_split_test.csv", parse_dates=True, index_col= 'DATE').drop(columns='grain')
plt.plot(df)
plt.subplot(2, 1, 2)
plt.title('Beer Production By Month')
groups = df.groupby(df.index.month)
months = concat([DataFrame(x[1].values) for x in groups], axis=1)
months = DataFrame(months)
months.columns = range(1,13)
months.boxplot()
plt.show()
target_column_name = 'BeerProduction'
time_column_name = 'DATE'
time_series_id_column_names = []
freq = 'M' #Monthly data
```
### Split Training data into Train and Validation set and Upload to Datastores
```
from helper import split_fraction_by_grain
from helper import split_full_for_forecasting
train, valid = split_full_for_forecasting(df, time_column_name)
train.to_csv("train.csv")
valid.to_csv("valid.csv")
test_df.to_csv("test.csv")
datastore = ws.get_default_datastore()
datastore.upload_files(files = ['./train.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
datastore.upload_files(files = ['./valid.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
datastore.upload_files(files = ['./test.csv'], target_path = 'beer-dataset/tabular/', overwrite = True,show_progress = True)
from azureml.core import Dataset
train_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/train.csv')])
valid_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/valid.csv')])
test_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/test.csv')])
```
### Setting forecaster maximum horizon
The forecast horizon is the number of periods into the future that the model should predict. Here, we set the horizon to 12 periods (i.e. 12 months). Notice that this is much shorter than the number of months in the test set; we will need to use a rolling test to evaluate the performance on the whole test set. For more discussion of forecast horizons and guiding principles for setting them, please see the [energy demand notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand).
```
forecast_horizon = 12
```
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|forecasting|
|**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>
|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
|**enable_dnn**|Enable Forecasting DNNs|
```
from azureml.automl.core.forecasting_parameters import ForecastingParameters
forecasting_parameters = ForecastingParameters(
time_column_name=time_column_name,
forecast_horizon=forecast_horizon,
freq='MS' # Set the forecast frequency to be monthly (start of the month)
)
# We will disable the enable_early_stopping flag to ensure the DNN model is recommended for demonstration purpose.
automl_config = AutoMLConfig(task='forecasting',
primary_metric='normalized_root_mean_squared_error',
experiment_timeout_hours = 1,
training_data=train_dataset,
label_column_name=target_column_name,
validation_data=valid_dataset,
verbosity=logging.INFO,
compute_target=compute_target,
max_concurrent_iterations=4,
max_cores_per_iteration=-1,
enable_dnn=True,
enable_early_stopping=False,
forecasting_parameters=forecasting_parameters)
```
We will now run the experiment, starting with 10 iterations of model search. The experiment can be continued for more iterations if more accurate results are required. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output= True)
# If you need to retrieve a run that already started, use the following code
# from azureml.train.automl.run import AutoMLRun
# remote_run = AutoMLRun(experiment = experiment, run_id = '<replace with your run id>')
```
Displaying the run objects gives you links to the visual tools in the Azure Portal. Go try them!
### Retrieve the Best Model for Each Algorithm
Below we select the best pipeline from our iterations. The get_output method on automl_classifier returns the best run and the fitted model for the last fit invocation. There are overloads on get_output that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration.
```
from helper import get_result_df
summary_df = get_result_df(remote_run)
summary_df
from azureml.core.run import Run
from azureml.widgets import RunDetails
forecast_model = 'TCNForecaster'
if not forecast_model in summary_df['run_id']:
forecast_model = 'ForecastTCN'
best_dnn_run_id = summary_df['run_id'][forecast_model]
best_dnn_run = Run(experiment, best_dnn_run_id)
best_dnn_run.parent
RunDetails(best_dnn_run.parent).show()
best_dnn_run
RunDetails(best_dnn_run).show()
```
## Evaluate on Test Data
We now use the best fitted model from the AutoML Run to make forecasts for the test set.
We always score on the original dataset whose schema matches the training set schema.
```
from azureml.core import Dataset
test_dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'beer-dataset/tabular/test.csv')])
# preview the first 3 rows of the dataset
test_dataset.take(5).to_pandas_dataframe()
compute_target = ws.compute_targets['beer-cluster']
test_experiment = Experiment(ws, experiment_name + "_test")
import os
import shutil
script_folder = os.path.join(os.getcwd(), 'inference')
os.makedirs(script_folder, exist_ok=True)
shutil.copy('infer.py', script_folder)
from helper import run_inference
test_run = run_inference(test_experiment, compute_target, script_folder, best_dnn_run, test_dataset, valid_dataset, forecast_horizon,
target_column_name, time_column_name, freq)
RunDetails(test_run).show()
from helper import run_multiple_inferences
summary_df = run_multiple_inferences(summary_df, experiment, test_experiment, compute_target, script_folder, test_dataset,
valid_dataset, forecast_horizon, target_column_name, time_column_name, freq)
for run_name, run_summary in summary_df.iterrows():
print(run_name)
print(run_summary)
run_id = run_summary.run_id
test_run_id = run_summary.test_run_id
test_run = Run(test_experiment, test_run_id)
test_run.wait_for_completion()
test_score = test_run.get_metrics()[run_summary.primary_metric]
summary_df.loc[summary_df.run_id == run_id, 'Test Score'] = test_score
print("Test Score: ", test_score)
summary_df
```
| github_jupyter |
## Click Prediction
This is a toy demo to show the overall process. This should not be expected to product accurate click predictions.
```
import pandas as pd
import torch
import torch.nn as nn
from torch.autograd import Variable
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.metrics import accuracy_score, confusion_matrix
import sys
sys.path.insert(0,'../..')
from api import *
```
## Services available:
* gscservice: Google Search Console
* gaservice: Google Analytics. A wrapper around https://github.com/debrouwere/google-analytics.
* semrushservice: SEM Rush. A port of https://github.com/storerjeremy/python-semrush.
* watsonservice: IBM Watson API. Pulls keywords and entities for a given html
## 1) Get GSC Data
```
gsc_profile = 'https://www.domain.com'
days_back = 180
# Combine the dataframe by nquery and page.
def combine_gsc(df):
df = df.groupby(['query'], as_index=False).agg({"impressions": "sum","clicks":"sum","position": "mean"})
df['ctr'] = df['clicks']/df['impressions']
df['position'] = df['position'].astype(int)
df = df.round({'ctr': 2})
return df
df = gscservice.get_site_data(gsc_profile, days_back, output_fn="demo3.csv")
df = combine_gsc(df).reset_index()
#Reduce to top queries by clicks
df = df[df.clicks > 5]
df.shape
```
## 2) Set Up DataLoader
```
import dataset
features = df[['position','impressions']]
labels = df[['clicks']]
def apply_embed(row):
embed = row['embedding']
for i, e in enumerate(embed):
row['e_'+str(i)] = e
return row
data_loader_bert, df_bert = dataset.load_bert_df(input_df=df, input_row="query")
df_bert_embed = df_bert.apply(apply_embed,axis=1).drop(columns=['embedding','linex_index','tokens'])
features = pd.concat([features.reset_index(drop =True), df_bert_embed.reset_index(drop =True)], axis=1)
data_loader = dataset.load_pandas(features, labels, batch_size=32, shuffle=True, drop_last=True)
```
## 3) Set up Model
```
torch.manual_seed(123)
# Make it
model = torch.nn.Sequential(
torch.nn.Linear(770, 100),
torch.nn.ReLU(),
torch.nn.Linear(100, 1),
)
# Optimizing options
loss_function = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr = 0.00001)
#optimizer = torch.optim.SGD(model.parameters(), lr=0.0001)
# Number of Epochs
n_epochs = 500
```
## 4) Train
```
loss_list = []
for epoch in range(n_epochs):
epoch +=1
epoch_loss = 0
optimizer.zero_grad()
for x, y in data_loader:
p_y = model(x)
loss = loss_function(p_y, y)
loss.backward()
optimizer.step()
epoch_loss += loss.data[0]
#print('epoch {}, loss {}'.format(epoch,epoch_loss))
loss_list.append(epoch_loss)
x_train = data_loader.dataset.tensors[0]
predicted =model(x_train).data.numpy()
y_correct = data_loader.dataset.tensors[1].data.numpy()
plt.plot(y_correct, label = 'from data', alpha = .5)
plt.plot(predicted, label = 'prediction', alpha = 0.5)
plt.legend()
plt.show()
#print(model.state_dict())
plt.plot(loss_list, label = 'train')
plt.legend();
predictions = [{'pred': float(x[0]), 'act':float(act_list[i][0])} for i, x in enumerate(pred_list)]
# Click predictions vs actual if prediction >= 10.
predictions
```
| github_jupyter |
## Importing and prepping data
```
import pandas as pd
import numpy as np
import diff_classifier.aws as aws
import diff_classifier.pca as pca
features = []
remote_folder = '9_28_18_Gel_Interface_Vids' #Folder in AWS S3 containing files to be analyzed
bucket = 'mckenna.data'
vids = 5
combos = [['0_4', '0_6'], ['0_6', '0_8'], ['0_8', '1_0'], ['1_0', '1_2']]
gels = [0.4, 0.6, 0.8, 1.0, 1.2]
counter2 = 0
counter = 0
for combo in combos:
for num in range(1, vids+1):
filename = 'features_100x_{}_{}_gel_{}_bulk_vid_{}.csv'.format(combo[0], combo[1], combo[0], num)
print(filename)
aws.download_s3('{}/{}'.format(remote_folder, filename), filename, bucket_name=bucket)
fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
print('{} size: {}'.format(filename, fstats.shape))
fstats['Interface'] = pd.Series(fstats.shape[0]*['{}/{}'.format(combo[0], combo[1])], index=fstats.index)
fstats['Bulk Agarose'] = pd.Series(fstats.shape[0]*[combo[0]], index=fstats.index)
fstats['Bulk Agarose Int'] = pd.Series(fstats.shape[0]*[gels[counter2]], index=fstats.index)
if gels[counter2] < 1:
fstats['Bulk Agarose Bin'] = pd.Series(fstats.shape[0]*['low'], index=fstats.index)
else:
fstats['Bulk Agarose Bin'] = pd.Series(fstats.shape[0]*['hi'], index=fstats.index)
fstats['Gel Code'] = pd.Series(fstats.shape[0]*['{}/{} {}'.format(combo[0], combo[1], combo[0])], index=fstats.index)
fstats['Video Number'] = pd.Series(fstats.shape[0]*[num], index=fstats.index)
counter = counter + 1
if counter == 1:
fstats_tot = fstats
else:
fstats_tot = fstats_tot.append(fstats, ignore_index=True)
counter2 = counter2 + 1
counter = 0
combo = combos[-1]
for num in range(1, vids+1):
try:
filename = 'features_100x_{}_{}_gel_{}_bulk_vid_{}.csv'.format(combo[0], combo[1], combo[1], num)
print(filename)
aws.download_s3('{}/{}'.format(remote_folder, filename), filename, bucket_name=bucket)
fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
print('{} size: {}'.format(filename, fstats.shape))
fstats['Interface'] = pd.Series(fstats.shape[0]*['{}/{}'.format(combo[0], combo[1])], index=fstats.index)
fstats['Bulk Agarose'] = pd.Series(fstats.shape[0]*[combo[1]], index=fstats.index)
fstats['Bulk Agarose Int'] = pd.Series(fstats.shape[0]*[1.2], index=fstats.index)
fstats['Gel Code'] = pd.Series(fstats.shape[0]*['{}/{} {}'.format(combo[0], combo[1], combo[1])], index=fstats.index)
fstats['Video Number'] = pd.Series(fstats.shape[0]*[num], index=fstats.index)
counter = counter + 1
fstats_tot = fstats_tot.append(fstats, ignore_index=True)
except:
print('Filename missing: '.format(filename))
fstats_tot.to_csv('features.csv')
#PCA analyses with too many datapoints fail. You get rows with lots of NAs. I'm going to try making a subset of the data first
#and then do a PCA analysis on that.
#include all in analysis
import random
subset = np.sort(np.array(random.sample(range(fstats_tot.shape[0]), 500000)))
fstats_sub = fstats_tot.loc[subset, :].reset_index(drop=True)
fstats_tot['Gel Code'].unique()
for typ in fstats_tot['Bulk Agarose'].unique():
fstats_type = fstats_tot[fstats_tot['Bulk Agarose']==typ].reset_index(drop=True)
print(fstats_type.shape)
#with equal sample sizes for each particle type
import random
counter = 0
for typ in fstats_tot['Bulk Agarose'].unique():
fstats_type = fstats_tot[fstats_tot['Bulk Agarose']==typ].reset_index(drop=True)
print(fstats_type.shape)
subset = np.sort(np.array(random.sample(range(fstats_type.shape[0]), 32000)))
if counter == 0:
fstats_sub = fstats_type.loc[subset, :].reset_index(drop=True)
else:
fstats_sub = fstats_sub.append(fstats_type.loc[subset, :].reset_index(drop=True), ignore_index=True)
counter = counter + 1
#fstats = pd.read_csv(filename, encoding = "ISO-8859-1", index_col='Unnamed: 0')
nonnum = ['Interface', 'Bulk Agarose', 'Bulk Agarose Int', 'Gel Code', 'Video Number', 'Track_ID', 'Mean Mean_Intensity', 'Std Mean_Intensity', 'X', 'Y',
'Mean X', 'Mean Y', 'Std X', 'Std Y', 'Bulk Agarose Bin']
fstats_num = fstats_sub.drop(nonnum, axis=1)
fstats_raw = fstats_num.as_matrix()
#fstats
```
## PCA analysis
The pca.pca_analysis function provides a completely contained PCA analysis of the input trajectory features dataset. It includes options to impute NaN values (fill in with average values or drop them), and to scale features. Read the docstring for more information.
```
ncomp = 16
pcadataset = pca.pca_analysis(fstats_tot, dropcols=nonnum, n_components=ncomp)
pcadataset.components.to_csv('components.csv')
pcadataset.prcomps
```
The pca.kmo function calculates the Kaiser-Meyer-Olkin statistic, a measure of sampling adequacy. Check the docstring for more information.
```
kmostat = pca.kmo(pcadataset.scaled)
import scipy.stats as stat
stat.bartlett(pcadataset.scaled[0, :], pcadataset.scaled[1, :], pcadataset.scaled[2, :], pcadataset.scaled[3, :])
newstr = ''
for i in range(pcadataset.scaled.shape[0]-1):
newstr = newstr + 'pcadataset.scaled[{}, :], '.format(i)
newstr = 'stat.bartlett(' + newstr + 'pcadataset.scaled[{}, :])'.format(i+1)
test = eval(newstr)
test
```
## Visualization
Users can then compare average principle component values between subgroups of the data. In this case, all particles were taken from the same sample, so there are no experimental subgroups. I chose to compare short trajectories to long trajectories, as I would expect differences between the two groups.
```
fstats_tot['Bulk Agarose'].unique()
import numpy as np
#ncomp = 14
dicti = {}
#test = np.exp(np.nanmean(np.log(pcadataset.final[pcadataset.final['Particle Size']==200].as_matrix()), axis=0))[-6:]
#test1 = np.exp(np.nanmean(np.log(pcadataset.final[pcadataset.final['Particle Size']==500].as_matrix()), axis=0))[-6:]
dicti[0] = np.nanmean(pcadataset.final[pcadataset.final['Bulk Agarose']=='0_4'].values[:, -ncomp:], axis=0)
dicti[1] = np.nanmean(pcadataset.final[pcadataset.final['Bulk Agarose']=='0_6'].values[:, -ncomp:], axis=0)
dicti[2] = np.nanmean(pcadataset.final[pcadataset.final['Bulk Agarose']=='0_8'].values[:, -ncomp:], axis=0)
dicti[3] = np.nanmean(pcadataset.final[pcadataset.final['Bulk Agarose']=='1_0'].values[:, -ncomp:], axis=0)
dicti[4] = np.nanmean(pcadataset.final[pcadataset.final['Bulk Agarose']=='1_2'].values[:, -ncomp:], axis=0)
labels = ['0_4', '0_6', '0_8', '1_0', '1_2']
pca.plot_pca(dicti, savefig=True, labels=labels, rticks=np.linspace(-4, 4, 9))
```
The variable pcadataset.prcomps shows the user the major contributions to each of the new principle components. When observing the graph above, users can see that there are some differences between short trajectories and long trajectories in component 0 (asymmetry1 being the major contributor) and component 1 (elongation being the major contributor).
```
#labels=['10K', '5K', '1K', 'COOH']
feats = pca.feature_violin(pcadataset.final, label='Bulk Agarose', lvals=labels, fsubset=ncomp, yrange=[-12, 12])
fstats1 = pca.feature_plot_3D(pcadataset.final, label='Bulk Agarose', lvals=labels, randcount=400, ylim=[-12, 12],
xlim=[-12, 12], zlim=[-12, 12], features=[0, 1, 3])
#ncomp = 14
trainp = np.array([])
testp = np.array([])
labels3 = ['low', 'hi']
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcadataset.final, 'Bulk Agarose Bin', labels3, equal_sampling=True,
tsize=1000, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcadataset.final.values[:, -ncomp:]
y2 = pcadataset.final['Bulk Agarose Bin'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
#ncomp = 14
trainp = np.array([])
testp = np.array([])
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcadataset.final, 'Bulk Agarose', labels, equal_sampling=True,
tsize=1000, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcadataset.final.values[:, -ncomp:]
y2 = pcadataset.final['Bulk Agarose'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
#ncomp = 14
trainp = np.array([])
testp = np.array([])
labelshort = ['0_4', '1_2']
pcasub = pcadataset.final[pcadataset.final['Bulk Agarose'].isin(labelshort)]
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcasub, 'Bulk Agarose', labelshort, equal_sampling=True,
tsize=500, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcasub.values[:, -ncomp:]
y2 = pcasub['Bulk Agarose'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
#ncomp = 14
trainp = np.array([])
testp = np.array([])
labelshort = ['0_4', '1_0']
pcasub = pcadataset.final[pcadataset.final['Bulk Agarose'].isin(labelshort)]
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcasub, 'Bulk Agarose', labelshort, equal_sampling=True,
tsize=500, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcasub.values[:, -ncomp:]
y2 = pcasub['Bulk Agarose'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
#ncomp = 14
trainp = np.array([])
testp = np.array([])
labelshort = ['0_4', '0_8']
pcasub = pcadataset.final[pcadataset.final['Bulk Agarose'].isin(labelshort)]
for i in range(0, 20):
KNNmod, X, y = pca.build_model(pcasub, 'Bulk Agarose', labelshort, equal_sampling=True,
tsize=500, input_cols=ncomp, model='MLP', NNhidden_layer=(6, 2))
trainp = np.append(trainp, pca.predict_model(KNNmod, X, y))
X2 = pcasub.values[:, -ncomp:]
y2 = pcasub['Bulk Agarose'].values
testp = np.append(testp, pca.predict_model(KNNmod, X2, y2))
print('Run {}: {}'.format(i, testp[i]))
print('{} +/ {}'.format(np.mean(trainp), np.std(trainp)))
print('{} +/ {}'.format(np.mean(testp), np.std(testp)))
from sklearn.neural_network import MLPRegressor
def build_modelR(rawdata, feature, featvals, equal_sampling=True,
tsize=20, from_end=True, input_cols=6, model='KNN',
**kwargs):
"""Builds a K-nearest neighbor model using an input dataset.
Parameters
----------
rawdata : pandas.core.frames.DataFrame
Raw dataset of n samples and p features.
feature : string or int
Feature in rawdata containing output values on which KNN
model is to be based.
featvals : string or int
All values that feature can take.
equal_sampling : bool
If True, training dataset will contain an equal number
of samples that take each value of featvals. If false,
each sample in training dataset will be taken randomly
from rawdata.
tsize : int
Size of training dataset. If equal_sampling is False,
training dataset will be exactly this size. If True,
training dataset will contain N x tsize where N is the
number of unique values in featvals.
n_neighbors : int
Number of nearest neighbors to be used in KNN
algorithm.
from_end : int
If True, in_cols will select features to be used as
training data defined end of rawdata e.g.
rawdata[:, -6:]. If False, input_cols will be read
as a tuple e.g. rawdata[:, 10:15].
input_col : int or tuple
Defined in from_end above.
Returns
-------
clf : sklearn.neighbors.classification.KNeighborsClassifier
KNN model
X : numpy.ndarray
training input dataset used to create clf
y : numpy.ndarray
training output dataset used to create clf
"""
defaults = {'NNsolver': 'adam', 'NNalpha': 1e-5,
'NNhidden_layer': (5, 2), 'NNrandom_state': 1,
'beta_1': 0.9, 'beta_2': 0.999, 'epsilon': 1e-8,
}
for defkey in defaults.keys():
if defkey not in kwargs.keys():
kwargs[defkey] = defaults[defkey]
if equal_sampling:
for featval in featvals:
if from_end:
test = rawdata[rawdata[feature] == featval
].values[:, -input_cols:]
else:
test = rawdata[rawdata[feature] == featval
].values[:, input_cols[0]:input_cols[1]]
to_plot = np.array(random.sample(range(0, test.shape[0]
), tsize))
if featval == featvals[0]:
X = test[to_plot, :]
y = rawdata[rawdata[feature] == featval
][feature].values[to_plot]
else:
X = np.append(X, test[to_plot, :], axis=0)
y = np.append(y, rawdata[rawdata[feature] == featval
][feature].values[to_plot], axis=0)
else:
if from_end:
test = rawdata.values[:, -input_cols:]
else:
test = rawdata.values[:, input_cols[0]:input_cols[1]]
to_plot = np.array(random.sample(range(0, test.shape[0]), tsize))
X = test[to_plot, :]
y = rawdata[feature].values[to_plot]
if model is 'MLP':
clf = MLPRegressor(solver=kwargs['NNsolver'], alpha=kwargs['NNalpha'],
hidden_layer_sizes=kwargs['NNhidden_layer'],
random_state=kwargs['NNrandom_state'],
beta_1=kwargs['beta_1'], beta_2=kwargs['beta_2'],
epsilon=kwargs['epsilon'])
clf.fit(X, y)
return clf, X, y
labels2 = [0.4, 0.6, 0.8, 1.0, 1.2]
trainp = np.array([])
testp = np.array([])
for i in range(0, 20):
MLPmod, X, y = build_modelR(pcadataset.final, 'Bulk Agarose Int', labels2, equal_sampling=True,
tsize=1000, input_cols=ncomp, model='MLP', NNhidden_layer=(13, 11, 9, 7, 5, 3))
trainp = np.append(trainp, MLPmod.score(X, y))
X2 = pcadataset.final.values[:, -ncomp:]
y2 = pcadataset.final['Bulk Agarose Int'].values
testp = np.append(testp, MLPmod.score(X2, y2))
print(testp[i])
import matplotlib.pyplot as plt
plt.scatter(y2, MLPmod.predict(X2), alpha=0.002)
x = np.linspace(0, 10, 11)
plt.plot(x, x, c='b', linewidth=1)
plt.ylim(0.4, 1.2)
plt.xlim(0.35, 1.25)
ticks = [0, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, 1.4, 1.6, 1.8]
plt.xticks(labels2)
plt.yticks(ticks)
import matplotlib.pyplot as plt
plt.scatter(y, MLPmod.predict(X), alpha=0.006)
x = np.linspace(0, 10, 11)
plt.plot(x, x, c='b', linewidth=1)
plt.ylim(0.4, 1.2)
plt.xlim(0.35, 1.25)
ticks = [0, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, 1.4, 1.6, 1.8]
plt.xticks(labels2)
plt.yticks(ticks)
```
| github_jupyter |
```
import os
import os.path as pth
import json
import shutil
import pandas as pd
from tqdm import tqdm
import tensorflow as tf
import tensorflow.keras as keras
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
BASE_MODEL_NAME = 'MobileNetV2-for-upload'
my_model_base = keras.applications.mobilenet_v2
my_model = my_model_base.MobileNetV2
config = {
'is_zscore':True,
# 'input_shape': (540, 960, 3),
'aug': {
'resize': (270, 480), # just resize
#'resize': (405, 720), # resize 75% and random crop.
},
#'input_shape': (224, 360, 3),
'input_shape': (270, 480, 3),
'output_activation': 'softmax',
'num_class': 1049,
'output_size': 1049,
'conv':{
'conv_num': (0), # (3,5,3),
'base_channel': 0, # 4,
'kernel_size': 0, # 3,
'padding':'same',
'stride':'X'
},
'pool':{
'type':'X',
'size':'X',
'stride':'X',
'padding':'same'
},
'fc':{
'fc_num': 0,
},
'activation':'relu',
'between_type': 'avg',
'is_batchnorm': True,
'is_dropout': False,
'dropout_rate': 0.5,
'batch_size': 64,
'buffer_size': 256,
'loss': 'CategoricalCrossentropy',
'num_epoch': 10000,
'learning_rate': 1e-3,
'random_state': 7777
}
image_feature_description = {
'image_raw': tf.io.FixedLenFeature([], tf.string),
'randmark_id': tf.io.FixedLenFeature([], tf.int64),
# 'id': tf.io.FixedLenFeature([], tf.string),
}
def _parse_image_function(example_proto):
return tf.io.parse_single_example(example_proto, image_feature_description)
def map_func(target_record):
img = target_record['image_raw']
label = target_record['randmark_id']
img = tf.image.decode_jpeg(img, channels=3)
img = tf.dtypes.cast(img, tf.float32)
return img, label
def resize_and_crop_func(image, label):
result_image = tf.image.resize(image, config['aug']['resize'])
result_image = tf.image.random_crop(image, size=config['input_shape'], seed=7777) # crop revived.
return result_image, label
def image_aug_func(image, label):
pass
return image, label
def post_process_func(image, label):
# result_image = result_image / 255
result_image = my_model_base.preprocess_input(image)
onehot_label = tf.one_hot(label, depth=config['num_class'])
return result_image, onehot_label
data_base_path = pth.join('data', 'public')
os.makedirs(data_base_path, exist_ok=True)
category_csv_name = 'category.csv'
category_json_name = 'category.json'
submission_csv_name = 'sample_submisstion.csv'
train_csv_name = 'train.csv'
# train_zip_name = 'train.zip'
train_tfrecord_name = 'all_train.tfrecords'
train_tfrecord_path = pth.join(data_base_path, train_tfrecord_name)
val_tfrecord_name = 'all_val.tfrecords'
val_tfrecord_path = pth.join(data_base_path, val_tfrecord_name)
# test_zip_name = 'test.zip'
test_tfrecord_name = 'test.tfrecords'
test_tfrecord_path = pth.join(data_base_path, test_tfrecord_name)
train_csv_path = pth.join(data_base_path, train_csv_name)
train_df = pd.read_csv(train_csv_path)
train_dict = {k:v for k, v in train_df.values}
submission_csv_path = pth.join(data_base_path, submission_csv_name)
submission_df = pd.read_csv(submission_csv_path)
# submission_df.head()
category_csv_path = pth.join(data_base_path, category_csv_name)
category_df = pd.read_csv(category_csv_path)
category_dict = {k:v for k, v in category_df.values}
# category_df.head()
train_tfrecord_path
```
### Model
```
import tensorflow as tf
from tensorflow.keras.preprocessing import image
import cv2
import matplotlib.pyplot as plt
from PIL import Image
from sklearn.model_selection import train_test_split, KFold, RepeatedKFold, GroupKFold, RepeatedStratifiedKFold
from sklearn.utils import shuffle
import numpy as np
import pandas as pd
import os
import os.path as pth
import shutil
import time
from tqdm import tqdm
import itertools
from itertools import product, combinations
import numpy as np
from PIL import Image
from IPython.display import clear_output
from multiprocessing import Process, Queue
import datetime
import tensorflow.keras as keras
from tensorflow.keras.utils import to_categorical, Sequence
from tensorflow.keras.layers import Input, Dense, Activation, BatchNormalization, \
Flatten, Conv3D, AveragePooling3D, MaxPooling3D, Dropout, \
Concatenate, GlobalMaxPool3D, GlobalAvgPool3D
from tensorflow.keras.models import Sequential, Model, load_model
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras.callbacks import ModelCheckpoint,LearningRateScheduler, \
EarlyStopping
from tensorflow.keras.losses import mean_squared_error, mean_absolute_error
from tensorflow.keras import backend as K
from tensorflow.keras.constraints import max_norm
conv_comb_list = []
conv_comb_list += [(0,)]
base_channel_list = [0]
fc_list = [0] # 128, 0
# between_type_list = [None, 'avg', 'max']
between_type_list = ['avg']
batch_size_list = [80]
activation_list = ['relu']
# len(conv_comb_list), conv_comb_list
def build_cnn(config):
input_layer = Input(shape=config['input_shape'], name='input_layer')
pret_model = my_model(
input_tensor=input_layer, include_top=False, weights='imagenet',
input_shape=config['input_shape'], pooling=config['between_type'],
classes=config['output_size']
)
pret_model.trainable = False
x = pret_model.output
if config['between_type'] == None:
x = Flatten(name='flatten_layer')(x)
if config['is_dropout']:
x = Dropout(config['dropout_rate'], name='output_dropout')(x)
x = Dense(config['output_size'], activation=config['output_activation'],
name='output_fc')(x)
# x = Activation(activation=config['output_activation'], name='output_activation')(x)
model = Model(inputs=input_layer, outputs=x, name='{}'.format(BASE_MODEL_NAME))
return model
model = build_cnn(config)
model.summary(line_length=150)
del model
origin_train_len = len(train_df) / 5 * 4
origin_val_len = len(train_df) / 5 * 1
train_num_steps = int(np.ceil((origin_train_len)/config['batch_size']))
val_num_steps = int(np.ceil((origin_val_len)/config['batch_size']))
model_base_path = data_base_path
model_checkpoint_path = pth.join(model_base_path, 'checkpoint')
for conv_comb, activation, base_channel, \
between_type, fc_num, batch_size \
in itertools.product(conv_comb_list, activation_list,
base_channel_list, between_type_list, fc_list,
batch_size_list):
config['conv']['conv_num'] = conv_comb
config['conv']['base_channel'] = base_channel
config['activation'] = activation
config['between_type'] = between_type
config['fc']['fc_num'] = fc_num
config['batch_size'] = batch_size
base = BASE_MODEL_NAME
base += '_resize_{}'.format(config['aug']['resize'][0])
base += '_conv_{}'.format('-'.join(map(lambda x:str(x),config['conv']['conv_num'])))
base += '_basech_{}'.format(config['conv']['base_channel'])
base += '_act_{}'.format(config['activation'])
base += '_pool_{}'.format(config['pool']['type'])
base += '_betw_{}'.format(config['between_type'])
base += '_fc_{}'.format(config['fc']['fc_num'])
base += '_zscore_{}'.format(config['is_zscore'])
base += '_batch_{}'.format(config['batch_size'])
if config['is_dropout']:
base += '_DO_'+str(config['dropout_rate']).replace('.', '')
if config['is_batchnorm']:
base += '_BN'+'_O'
else:
base += '_BN'+'_X'
model_name = base
print(model_name)
### Define dataset
dataset = tf.data.TFRecordDataset(train_tfrecord_path, compression_type='GZIP')
dataset = dataset.map(_parse_image_function, num_parallel_calls=tf.data.experimental.AUTOTUNE)
# dataset = dataset.cache()
dataset = dataset.map(map_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
dataset = dataset.map(resize_and_crop_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
dataset = dataset.map(image_aug_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
dataset = dataset.shuffle(config['buffer_size'])
dataset = dataset.batch(config['batch_size'])
dataset = dataset.map(post_process_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
dataset = dataset.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
val_dataset = tf.data.TFRecordDataset(val_tfrecord_path, compression_type='GZIP')
val_dataset = val_dataset.map(_parse_image_function, num_parallel_calls=tf.data.experimental.AUTOTUNE)
val_dataset = val_dataset.map(map_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
val_dataset = val_dataset.map(resize_and_crop_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
# val_dataset = val_dataset.map(image_aug_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
# val_dataset = val_dataset.shuffle(config['buffer_size'])
val_dataset = val_dataset.batch(config['batch_size'])
val_dataset = val_dataset.map(post_process_func, num_parallel_calls=tf.data.experimental.AUTOTUNE)
# val_dataset = val_dataset.cache()
val_dataset = val_dataset.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
model_path = pth.join(
model_checkpoint_path, model_name,
)
model = build_cnn(config)
# model.summary()
# model.compile(loss=config['loss'], optimizer=Adam(lr=config['learning_rate']),
# metrics=['acc', 'Precision', 'Recall', 'AUC'])
initial_epoch = 0
if pth.isdir(model_path) and len([_ for _ in os.listdir(model_path) if _.endswith('hdf5')]) >= 1:
model.compile(loss=config['loss'], optimizer=Adam(lr=config['learning_rate']),
metrics=['acc', 'Precision', 'Recall', 'AUC'])
model_chk_name = sorted(os.listdir(model_path))[-1]
initial_epoch = int(model_chk_name.split('-')[0])
model.load_weights(pth.join(model_path, model_chk_name))
else:
model.compile(optimizer='rmsprop', loss='categorical_crossentropy',
metrics=['acc', 'Precision', 'Recall', 'AUC'])
model.fit(
x=dataset, epochs=16, # train only top layers for just a few epochs.
validation_data=val_dataset, shuffle=True,
#callbacks = [checkpointer, es], #batch_size=config['batch_size']
initial_epoch=initial_epoch,
# steps_per_epoch=train_num_steps, validation_steps=val_num_steps,
verbose=1)
for i, layer in enumerate(model.layers):
print(i, layer.name)
for layer in model.layers[:135]:
layer.trainable = False
for layer in model.layers[135:]:
layer.trainable = True
model.compile(loss=config['loss'], optimizer=Adam(lr=config['learning_rate']),
metrics=['acc', 'Precision', 'Recall', 'AUC'])
# ### Freeze first layer
# conv_list = [layer for layer in model.layers if isinstance(layer, keras.layers.Conv2D)]
# conv_list[0].trainable = False
# # conv_list[1].trainable = False
os.makedirs(model_path, exist_ok=True)
model_filename = pth.join(model_path, '{epoch:06d}-{val_loss:0.6f}-{loss:0.6f}.hdf5')
checkpointer = ModelCheckpoint(
filepath=model_filename, verbose=1,
period=1, save_best_only=True,
monitor='val_loss'
)
es = EarlyStopping(monitor='val_loss', verbose=1, patience=6)
hist = model.fit(
x=dataset, epochs=config['num_epoch'],
validation_data=val_dataset, shuffle=True,
callbacks = [checkpointer, es], #batch_size=config['batch_size']
initial_epoch=initial_epoch,
# steps_per_epoch=train_num_steps, validation_steps=val_num_steps,
verbose=1
)
model_analysis_path = model_path.replace('checkpoint', 'analysis')
visualization_path = pth.join(model_analysis_path,'visualization')
os.makedirs(visualization_path, exist_ok=True)
print()
# clear_output()
for each_label in ['loss', 'acc', 'precision', 'recall', 'auc']:
fig, ax = plt.subplots()
ax.plot(hist.history[each_label], 'g', label='train_{}'.format(each_label))
ax.plot(hist.history['val_{}'.format(each_label)], 'r', label='val_{}'.format(each_label))
ax.set_xlabel('epoch')
ax.set_ylabel('loss')
ax.legend(loc='upper left')
if not each_label == 'loss':
plt.ylim(0, 1)
plt.show()
filename = 'learning_curve_{}'.format(each_label)
# fig.savefig(pth.join(visualization_path, filename), transparent=True)
plt.cla()
plt.clf()
plt.close('all')
np.savez_compressed(pth.join(visualization_path, 'learning_curve'),
loss=hist.history['loss'],
val_loss=hist.history['val_loss'],
acc=hist.history['acc'],
val_acc=hist.history['val_acc'],
precision=hist.history['precision'],
vaval_precisionl_mae=hist.history['val_precision'],
recall=hist.history['recall'],
val_recall=hist.history['val_recall'],
auc=hist.history['auc'],
val_auc=hist.history['val_auc']
)
model.save(pth.join(model_path, '000000_last.hdf5'))
K.clear_session()
del(model)
model_analysis_base_path = pth.join(model_base_path, 'analysis', model_name)
with open(pth.join(model_analysis_base_path, 'config.json'), 'w') as f:
json.dump(config, f)
chk_name_list = sorted([name for name in os.listdir(model_path) if name != '000000_last.hdf5'])
for chk_name in chk_name_list[:-2]:
os.remove(pth.join(model_path, chk_name))
# clear_output()
```
### Inference
```
image_feature_description_for_test = {
'image_raw': tf.io.FixedLenFeature([], tf.string),
# 'randmark_id': tf.io.FixedLenFeature([], tf.int64),
# 'id': tf.io.FixedLenFeature([], tf.string),
}
def _parse_image_function_for_test(example_proto):
return tf.io.parse_single_example(example_proto, image_feature_description_for_test)
def map_func_for_test(target_record):
img = target_record['image_raw']
img = tf.image.decode_jpeg(img, channels=3)
img = tf.dtypes.cast(img, tf.float32)
return img
def resize_and_crop_func_for_test(image):
result_image = tf.image.resize(image, config['aug']['resize'])
result_image = tf.image.random_crop(image, size=config['input_shape'], seed=7777) # revive
return result_image
def post_process_func_for_test(image):
# result_image = result_image / 255
result_image = my_model_base.preprocess_input(image)
return result_image
submission_base_path = pth.join(data_base_path, 'submission')
os.makedirs(submission_base_path, exist_ok=True)
for conv_comb, activation, base_channel, \
between_type, fc_num, batch_size \
in itertools.product(conv_comb_list, activation_list,
base_channel_list, between_type_list, fc_list,
batch_size_list):
config['conv']['conv_num'] = conv_comb
config['conv']['base_channel'] = base_channel
config['activation'] = activation
config['between_type'] = between_type
config['fc']['fc_num'] = fc_num
config['batch_size'] = batch_size
base = BASE_MODEL_NAME
base += '_resize_{}'.format(config['aug']['resize'][0])
base += '_conv_{}'.format('-'.join(map(lambda x:str(x),config['conv']['conv_num'])))
base += '_basech_{}'.format(config['conv']['base_channel'])
base += '_act_{}'.format(config['activation'])
base += '_pool_{}'.format(config['pool']['type'])
base += '_betw_{}'.format(config['between_type'])
base += '_fc_{}'.format(config['fc']['fc_num'])
base += '_zscore_{}'.format(config['is_zscore'])
base += '_batch_{}'.format(config['batch_size'])
if config['is_dropout']:
base += '_DO_'+str(config['dropout_rate']).replace('.', '')
if config['is_batchnorm']:
base += '_BN'+'_O'
else:
base += '_BN'+'_X'
model_name = base
print(model_name)
### Define dataset
test_dataset = tf.data.TFRecordDataset(test_tfrecord_path, compression_type='GZIP')
test_dataset = test_dataset.map(_parse_image_function_for_test, num_parallel_calls=tf.data.experimental.AUTOTUNE)
test_dataset = test_dataset.map(map_func_for_test, num_parallel_calls=tf.data.experimental.AUTOTUNE)
test_dataset = test_dataset.map(resize_and_crop_func_for_test, num_parallel_calls=tf.data.experimental.AUTOTUNE)
test_dataset = test_dataset.batch(config['batch_size'])
test_dataset = test_dataset.map(post_process_func_for_test, num_parallel_calls=tf.data.experimental.AUTOTUNE)
test_dataset = test_dataset.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
model_path = pth.join(
model_checkpoint_path, model_name,
)
model = build_cnn(config)
# model.summary()
model.compile(loss=config['loss'], optimizer=Adam(lr=config['learning_rate']),
metrics=['acc', 'Precision', 'Recall', 'AUC'])
initial_epoch = 0
model_chk_name = sorted(os.listdir(model_path))[-1]
initial_epoch = int(model_chk_name.split('-')[0])
model.load_weights(pth.join(model_path, model_chk_name))
preds = model.predict(test_dataset, verbose=1)
pred_labels = np.argmax(preds, axis=1)
pred_probs = np.array([pred[indice] for pred, indice in zip(preds, pred_labels)])
submission_csv_path = pth.join(data_base_path, submission_csv_name)
submission_df = pd.read_csv(submission_csv_path)
submission_df['landmark_id'] = pred_labels
submission_df['conf'] = pred_probs
today_str = datetime.date.today().strftime('%Y%m%d')
result_filename = '{}.csv'.format(model_name)
submission_csv_fileaname = pth.join(submission_base_path, '_'.join([today_str, result_filename]))
submission_df.to_csv(submission_csv_fileaname, index=False)
```
| github_jupyter |
# Get Started
*by [Longqi@Cornell](http://www.cs.cornell.edu/~ylongqi/) licensed under [Creative Commons Attribution 4.0 International License](https://creativecommons.org/licenses/by/4.0/)*
This tutorial demonstrates the process of training and evaluating recommendation algorithms using OpenRec (tutorial on implementing new recommendation algorithm: [tutorial]()):
* Prepare training and evaluation datasets.
* Instantiate a recommender.
* Instantiate a sampler.
* Instantiate evaluators.
* Instantiate a model trainer.
* TRAIN AND EVALUATE!
### Prepare training and evaluation datasets
* Download your favorite dataset from the web. In this tutorial, we use [a relatively small citeulike dataset](http://www.wanghao.in/CDL.htm) for demonstration purpose (It requires `unrar` package to unpack the data).
```
import os
try:
from urllib.request import urlretrieve
except ImportError:
from urllib import urlretrieve
urlretrieve('http://www.wanghao.in/data/ctrsr_datasets.rar', 'ctrsr_datasets.rar')
os.system('unrar x ctrsr_datasets.rar')
```
* Convert raw data into [numpy structured array](https://docs.scipy.org/doc/numpy-1.13.0/user/basics.rec.html). As required by the **ImplicitDataset** class, two keys `user_id` and `item_id` are required. Each row in the converted numpy array represents an interaction. The array might contain additional keys based on the use cases.
```
import numpy as np
import random
users_count = 0
interactions_count = 0
with open('ctrsr_datasets/citeulike-a/users.dat', 'r') as fin:
for line in fin:
interactions_count += int(line.split()[0])
users_count += 1
# radomly hold out an item per user for validation and testing respectively.
val_structured_arr = np.zeros(users_count, dtype=[('user_id', np.int32), ('item_id', np.int32)])
test_structured_arr = np.zeros(users_count, dtype=[('user_id', np.int32), ('item_id', np.int32)])
train_structured_arr = np.zeros(interactions_count-11102, dtype=[('user_id', np.int32), ('item_id', np.int32)])
interaction_ind = 0
next_user_id = 0
next_item_id = 0
map_to_item_id = dict() # Map item id from 0 to len(items)-1
with open('ctrsr_datasets/citeulike-a/users.dat', 'r') as fin:
for line in fin:
item_list = line.split()[1:]
random.shuffle(item_list)
for ind, item in enumerate(item_list):
if item not in map_to_item_id:
map_to_item_id[item] = next_item_id
next_item_id += 1
if ind == 0:
val_structured_arr[next_user_id] = (next_user_id, map_to_item_id[item])
elif ind == 1:
test_structured_arr[next_user_id] = (next_user_id, map_to_item_id[item])
else:
train_structured_arr[interaction_ind] = (next_user_id, map_to_item_id[item])
interaction_ind += 1
next_user_id += 1
```
* Instantiate training, validation, and testing datasets. As the data is from users' implicit feedback, we choose the **ImplicitDataset** class, as opposed to the general **Dataset** class.
```
from openrec.tf1.legacy.utils import ImplicitDataset
train_dataset = ImplicitDataset(raw_data=train_structured_arr,
max_user=users_count,
max_item=len(map_to_item_id), name='Train')
val_dataset = ImplicitDataset(raw_data=val_structured_arr,
max_user=users_count,
max_item=len(map_to_item_id), name='Val')
test_dataset = ImplicitDataset(raw_data=test_structured_arr,
max_user=users_count,
max_item=len(map_to_item_id), name='Test')
```
### Instantiate a recommender
We use the [BPR recommender](http://openrec.tf1.readthedocs.io/en/latest/recommenders/openrec.tf1.recommenders.bpr.html) that implements the pure Baysian Personalized Ranking (BPR) algorithm
```
from openrec.tf1.legacy.recommenders import BPR
bpr_model = BPR(batch_size=1000,
max_user=train_dataset.max_user(),
max_item=train_dataset.max_item(),
dim_embed=20,
opt='Adam')
```
### Instantiate a sampler
A basic [pairwise sampler](http://openrec.tf1.readthedocs.io/en/latest/utils/openrec.tf1.utils.samplers.html) is used, i.e., each instance contains an user, an item that the user interacts, and an item that the user did NOT interact.
```
from openrec.tf1.legacy.utils.samplers import PairwiseSampler
sampler = PairwiseSampler(batch_size=1000, dataset=train_dataset, num_process=1)
```
### Instantiate evaluators
Define evaluators that you plan to use. This tutorial evaluate the recommender against Area Under Curve (AUC).
```
from openrec.tf1.legacy.utils.evaluators import AUC
auc_evaluator = AUC()
```
### Instantiate a model trainer
The **implicit model trainer** drives the training and evaluation of the recommender using defined *implicit feedback datasets*, sampler, model, and evaluators.
```
from openrec.tf1.legacy import ImplicitModelTrainer
model_trainer = ImplicitModelTrainer(batch_size=1000,
test_batch_size=100,
train_dataset=train_dataset,
model=bpr_model,
sampler=sampler)
```
### TRAIN AND EVALUATE!
```
model_trainer.train(num_itr=10000,
display_itr=1000,
eval_datasets=[val_dataset, test_dataset],
evaluators=[auc_evaluator])
```
| github_jupyter |
## CrypTen - Training an Encrypted Neural Network across Workers using Plans
We will train an encrypted neural network across different PySyft workers (deployed as [Grid Nodes](https://github.com/OpenMined/PyGrid/tree/dev/apps/node)). For this we will be using Plans and we will be using CrypTen as a backend for SMPC.
Authors:
- George Muraru - Twitter: [@gmuraru](https://twitter.com/georgemuraru)
- Ayoub Benaissa - Twitter: [@y0uben11](https://twitter.com/y0uben11)
## Training Overivew
* In this tutorial we will use the [MNIST dataset](http://yann.lecun.com/exdb/mnist/)
* The features we need for training the network are split accross two workers (we will name them *alice* and *bob*)
## Setup
### Download/install needed repos
* Clone the [PyGrid repo](https://github.com/OpenMined/PyGrid)
* we need this because *alice* and *bob* are two different Nodes in our network
* install the PyGrid node component using *poetry*
### Bring up the PyGridNodes
* In the *PyGrid* repo:
1. install *poetry* (```pip install poetry```)
2. go to *apps/nodes*
3. run ```poetry install``` (those steps are also in the README from the PyGrid repo)
4. start *bob* and *alice* using:
```
./run.sh --id alice --port 3000 --start_local_db
./run.sh --id bob --port 3001 --start_local_db
```
This will start two workers, *alice* and *bob* and we will connect to them using the port 3000 and 3001.
### Dataset preparation
* Run the cell bellow to download a script from the CrypTen repository
* It will be used to split the features between the workers
* Each party will get only a subset of features.
* We will use only 100 entries from the dataset
* We will use binary classification (0 vs [1-9] digits)
```
!wget "https://raw.githubusercontent.com/facebookresearch/CrypTen/b1466440bde4db3e6e1fcb1740584d35a16eda9e/tutorials/mnist_utils.py" -O "mnist_utils.py"
!python "mnist_utils.py" --option features --reduced 100 --binary
```
## Prepare the ground
```
import pytest
import crypten
import torch
import torch.nn as nn
import torch.nn.functional as F
from time import time
import syft as sy
from syft.frameworks.crypten.context import run_multiworkers
from syft.grid.clients.data_centric_fl_client import DataCentricFLClient
torch.manual_seed(0)
torch.set_num_threads(1)
hook = sy.TorchHook(torch)
```
## Neural network to train
```
class ExampleNet(nn.Module):
def __init__(self):
super(ExampleNet, self).__init__()
self.conv1 = nn.Conv2d(1, 16, kernel_size=5, padding=0)
self.fc1 = nn.Linear(16 * 12 * 12, 100)
self.fc2 = nn.Linear(100, 2)
def forward(self, x):
out = self.conv1(x)
out = F.relu(out)
out = F.max_pool2d(out, 2)
out = out.view(-1, 16 * 12 * 12)
out = self.fc1(out)
out = F.relu(out)
out = self.fc2(out)
return out
```
## Setup the workers and send them the data
We need to have the two GridNodes workers running.
```
# Syft workers
print("[%] Connecting to workers ...")
alice = DataCentricFLClient(hook, "ws://localhost:3000")
bob = DataCentricFLClient(hook, "ws://localhost:3001")
print("[+] Connected to workers")
print("[%] Sending training data ...")
# Prepare the labels
label_eye = torch.eye(2)
labels = torch.load("/tmp/train_labels.pth")
labels = labels.long()
labels_one_hot = label_eye[labels]
# Prepare and send training data
alice_train = torch.load("/tmp/alice_train.pth").tag("alice_train")
alice_ptr = alice_train.send(alice)
bob_train = torch.load("/tmp/bob_train.pth").tag("bob_train")
bob_ptr = bob_train.send(bob)
print("[+] Data ready")
```
## Check the data shape
One entry from the MNIST dataset contains 28x28 features. Those are splitted accross our workers.
We can check it out by running the next cell!
```
print(f"Alice data shape {alice_train.shape}")
print(f"Bob data shape {bob_train.shape}")
```
## Initialize a dummy model
Instanciate a model and create a dummy input that could be forwarded through it. This is needed to build the CrypTen model.
```
dummy_input = torch.empty(1, 1, 28, 28)
pytorch_model = ExampleNet()
```
### Define the CrypTen computation
We need to specify for the ```run_multiworkers``` decorator:
* the workers that will take part in the computation
* the master address, this will be used for their synchronization
* the instantiated model that will be sent
* a dummy input for the model
We will use the ```func2plan``` decorator to:
* trace the operations from our function
* sending the plan operations to *alice* and *bob* - the plans operations will act as the function
* run the plans operations on both workers
```
ALICE = 0 # Alice rank in CrypTen
BOB = 1 # Bob rank in CrypTen
@run_multiworkers(
[alice, bob], master_addr="127.0.0.1", model=pytorch_model, dummy_input=dummy_input
)
@sy.func2plan()
def run_encrypted_training(
model=None,
learning_rate=0.001,
num_epochs=2,
batch_size=10,
num_batches=bob_ptr.shape[0]//10,
labels_one_hot=labels_one_hot,
crypten=crypten,
torch=torch,
):
x_alice_enc = crypten.load("alice_train", ALICE)
x_bob_enc = crypten.load("bob_train", BOB)
x_combined_enc = crypten.cat([x_alice_enc, x_bob_enc], dim=2)
x_combined_enc = x_combined_enc.unsqueeze(1)
model.encrypt()
model.train()
loss = crypten.nn.MSELoss()
l_values = []
for i in range(num_epochs):
for batch in range(num_batches):
start, end = batch * batch_size, (batch + 1) * batch_size
x_train = x_combined_enc[start:end]
y_batch = labels_one_hot[start:end]
y_train = crypten.cryptensor(y_batch, requires_grad=True)
# perform forward pass:
output = model(x_train)
loss_value = loss(output, y_train)
# set gradients to "zero"
model.zero_grad()
# perform backward pass:
loss_value.backward()
# update parameters
model.update_parameters(learning_rate)
# Print progress every batch:
batch_loss = loss_value.get_plain_text()
l_values.append(batch_loss)
model.decrypt()
return (l_values, model)
```
## Run the CrypTen computation
Now let's run the computation defined above
```
# Get the returned values
# key 0 - return values for alice
# key 1 - return values for bob
print("[%] Starting computation")
func_ts = time()
*losses, model = run_encrypted_training()[0]
func_te = time()
print(f"[+] run_encrypted_training() took {int(func_te - func_ts)}s")
losses_per_epoch = len(losses) // 2
for i in range(2):
print(f"Epoch {i}:")
for batch, loss in enumerate(losses[i * losses_per_epoch:(i+1) * losses_per_epoch]):
print(f"\tBatch {(batch+1)} of 10 Loss: {loss:.4f}")
```
The model returned is a CrypTen model, but we can always run the usual PySyft methods to share the parameters and so on, as far as the model in not encrypted.
```
cp = sy.VirtualWorker(hook=hook, id="cp")
model.fix_prec()
model.share(alice, bob, crypto_provider=cp)
print(model)
print(list(model.parameters())[0])
```
## CleanUp
```
# CleanUp portion taken from the CrypTen project
import os
filenames = ['/tmp/alice_train.pth',
'/tmp/bob_train.pth',
'/tmp/alice_test.pth',
'/tmp/bob_test.pth',
'/tmp/train_labels.pth',
'/tmp/test_labels.pth',
'mnist_utils.py']
for fn in filenames:
if os.path.exists(fn): os.remove(fn)
```
# Congratulations!!! - Time to Join the Community!
Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
### Star PySyft on GitHub
The easiest way to help our community is just by starring the GitHub repos! This helps raise awareness of the cool tools we're building.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org)
### Join a Code Project!
The best way to contribute to our community is to become a code contributor! At any time you can go to PySyft GitHub Issues page and filter for "Projects". This will show you all the top level Tickets giving an overview of what projects you can join! If you don't want to join a project, but you would like to do a bit of coding, you can also look for more "one off" mini-projects by searching for GitHub issues marked "good first issue".
- [PySyft Projects](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3AProject)
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22Good+first+issue+%3Amortar_board%3A%22)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
[OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
## 4. Visualización de Datos
[Playlist de Ciencia de Datos en castellano](https://www.youtube.com/playlist?list=PLjyvn6Y1kpbEmRY4-ELeRA80ZywV7Xd67)
[](https://www.youtube.com/watch?v=GHZIkEwlu7M&list=PLjyvn6Y1kpbEmRY4-ELeRA80ZywV7Xd67&index=5)
Además de los estadísticos de resumen, la visualización de datos ayuda a comprender las características de los datos y cómo se relacionan las diferentes variables.

Hay muchos ejemplos de visualización de datos con [Matplotlib](https://matplotlib.org/gallery/index.html), [Seaborn](https://seaborn.pydata.org/examples/index.html), y [Plotly](https://plot.ly/python/). En este tutorial, revisamos algunos ejemplos para visualizar:
- series de tiempo: línea
- variables correlacionadas: dispersión, diagrama de pares
- distribuciones de datos: barra, caja, violín, distribución, diagrama conjunto
Cada gráfico se muestra con una de las librerías de gráficos. Matplotlib es una librería de nivel base de Python, Seaborn utiliza matplotlib y automatiza gráficos más complejos, y Plotly crea gráficos interactivos.
```
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
import plotly.express as px
```

### Generar Datos
Ejecuta la siguiente celda para:
- Generar `n` valores espaciados linealmente entre `0` y `n-1` con `np.linspace(start,end,count)`
- Selecciona muestras aleatorias de una distribución uniforme entre 0 y 1 con `np.random.rand(count)`
- Selecciona muestras aleatorias de una distribución normal (gaussiana) con `np.random.normal(mean,std,count)`
- Crea una serie de tiempo que cambie según `y[i]*0.1` manteniendose en el rango de `-3` a `3`
- Combina `tt`, `x`, `y`, y `z` con una columna vertical `np.vstack` y transponer `.T` para datos orientados a columnas.
- Crear una DataFrame pandas con columnas `tt`, `x`, `y`, y `z`
```
import numpy as np
import pandas as pd
np.random.seed(0) # cambiar "seed" para una respuesta diferente
n = 1000
tt = np.linspace(0,n-1,n)
x = np.random.rand(n)+tt/500
y = np.random.normal(0,x,n)
z = [0]
for i in range(1,n):
z.append(min(max(-3,z[i-1]+y[i]*0.1),3))
data = pd.DataFrame(np.vstack((tt,x,y,z)).T,\
columns=['time','x','y','z'])
data['w'] = '0-499'
for i in range(int(n/2),n):
data.at[i,'w'] = '500-999'
data.head()
```

### Gráfico
El gráfico "plot" es el más básico. Hay un tutorial introductorio a gráficos en el [curso introductorio a Python en inglés, Lección 12](https://github.com/APMonitor/begin_python/blob/master/12.%20Plotting.ipynb). Visita este curso si necesitas más información sobre gráficos básicos como `plt.plot()`
```
plt.plot(tt,z)
plt.show()
```
La línea del gráfico puede ser cambiada y mejorada utilizando estilos seleccionados. A continuación se muestra un ejemplo con opciones comunes.
**c=Colors** (Color)
============= ===============================
Carácter Color
============= ===============================
``'b'`` azul
``'g'`` verde
``'r'`` rojo
``'y'`` amarillo
``'k'`` negro
============= ===============================
**m=Markers** (Marca)
============= ===============================
Carácter Descripción
============= ===============================
``'.'`` punto
``'o'`` círculo
``'s'`` cuadrado
``'^'`` triángulo
``'*'`` estrella
============= ===============================
**ln=Line Styles** (Estilo de línea)
============= ===============================
Carácter Descripción
============= ===============================
``'-'`` línea contínua
``'--'`` línea discontínua
``'-.'`` línea de puntos y guiones
``':'`` línea de puntos
============= ===============================
```
plt.figure(1,figsize=(10,6)) # ajustar el tamaño de la figura
ax=plt.subplot(2,1,1) # sub-gráfico 1
plt.plot(tt,z,'r-',linewidth=3,label='z') # gráfico con línea roja
ax.grid() # agregar cuadrícula
plt.ylabel('z'); plt.legend() # agregar ylabel (nombre en eje y), leyenda
plt.subplot(2,1,2) # sub-gráfico 2
plt.plot(tt,x,'b.',label='x') # gráfico de puntos azules
plt.plot(tt,y,color='orange',label='y',alpha=0.7) # gráfico anaranjado
plt.xlabel('time'); plt.legend() # leyenda
plt.savefig('04-myFig.png',transparent=True,dpi=600) # guardar el gráfico
plt.show() # indicar el gráfico
```

### Actividad de Gráficos
Crea una gráfica con los datos:
```python
xt = [0,0.1,0.2,0.3,0.5,0.8,1.0]
yt = [1.0,2.1,3.5,6.5,7.2,5.9,6.3]
```

### Gráfico de Dispersión
Los gráficos de dispersión son similares a los gráficos regulares, pero muestran puntos individuales en lugar de valores conectados en serie. Matplotlib y Plotly se utilizan en este ejemplo. Matplotlib es rápido y sencillo mientras Plotly tiene características para gráficos interactivos!
```
# matplotlib
plt.scatter(x,y)
plt.show()
# plotly
fig = px.scatter(data,x='x',y='y',color='w',size='x',hover_data=['w'])
fig.show()
```

### Actividad con Gráfico de Dispersión
Crear un gráfico de dispersión con `matplotlib` o `plotly` que ilustre `xt` combinado con `yt` y `zt`:
```python
xt = np.array([0,0.1,0.2,0.3,0.5,0.8,1.0])
yt = np.array([1.0,2.1,3.5,6.5,7.2,5.9,6.3])
zt = xt*yt
```
Cambia la forma de los puntos a un cuadrado para `yt` y un triángulo para `zt`. Agregua una etiqueta para indicar qué puntos son `yt` y `zt`.

### Gráfico de Barras
Los gráficos de barras ilustran un histograma en un rango de intervalo. La opción `alpha` es la transparencia entre `0` y `1`. Un valor de `0.7` es un buen valor para mostrar los datos subyacentes y superpuestos.
```
bins = np.linspace(-3,3,31)
plt.hist(y,bins,label='y')
plt.hist(z,bins,alpha=0.7,label='z')
plt.legend()
plt.show()
```

### Actividad con Gráfico de Barras
Genera un diagrama de barras que muestre la distribución de `xt`, `yt`, y `zt`:
```python
nt = 1000
xt = np.random.rand(nt)
yt = np.random.normal(0,1,nt)
zt = xt*yt
```
Utiliza `bins = np.linspace(-3,3,31)` para crear el histograma.

### Gráfico de Pares
Un gráfico de pares o pair plot muestra la correlación entre variables. Tiene distribuciones en la diagonal y diagramas de dispersión en el resto de lugares. Un gráfico de pares también muestra un color diferente (`hue`) por categoría `w`. Los gráficos de pares muestran correlaciones entre pares de variables que pueden estar relacionadas y dan una buena indicación de las características (entradas explicativas) que se utilizan para clasificación o regresión.
```
sns.pairplot(data[['x','y','z','w']],hue=('w'))
plt.show()
```

### Actividad con Gráfico de Pares
Crea un gráfico de pares que muestra la correlación entre `xt`, `yt`, y `zt` Entre los primeros 500 y segundos 500 valores aleatorios que están categorizados como `Dist`. Crea un DataFrame de `pandas` con:
```python
nt = 100
xt = np.random.rand(nt)
yt = np.random.normal(0,1,nt)
zt = xt*yt
dt = pd.DataFrame(np.column_stack([xt,yt,zt]),columns=['xt','yt','zt'])
dt['Dist'] = 'First'
for i in range(int(nt/2),nt):
dt.at[i,'Dist'] = 'Second'
```

### Diagrama de Caja
Un diagrama de caja muestra los cuartiles de datos. En este caso, compararemos los primeros 500 puntos con los últimos 500 puntos.
```
sns.boxplot(x='w',y='x',data=data)
plt.show()
```

### Actividad con Diagrama de Caja
Crea un diagrama de caja que muestre los cuartiles de `yt` por la primera y segunda serie como se indica en `Dist`.

### Diagrama de Violín
Un diagrama de violín combina los cuartiles del diagrama de caja con la distribución.
```
sns.violinplot(x='w',y='x',data=data,size=6)
plt.show()
```

### Actividad con Diagrama de Violín
Crea un diagrama de violín que muestre los cuartiles y la distribución de `zt` por primera y segunda serie como se indica en `Dist` en la DataFrame `dt`.

### Gráfico Conjunto
Un gráfico conjunto muestra dos variables, con las distribuciones univariadas y conjuntas. Intenta `kind='reg'`, `'kde'`, y `'hex'` para ver diferentes estilos de gráficos conjuntos.
```
sns.jointplot('x','z',data=data,kind="reg") #hex, kde, reg
plt.show()
```

### Actividad con Gráfico Conjunto
Crea una gráfica conjunta que muestre la distribución conjunta de `yt` y `zt` en el DataFrame `dt`.
### Actividad con el TCLab

### Generar o Recuperar Datos

Carga un archivo de datos de muestra si no cuentas con un TCLab. De lo contrario, genera un archivo a partir de los datos del TCLab con segundos (`t`), niveles de calentador (`Q1` y `Q2`), y temperaturas (`lab.T1` y `lab.T2`). Registra los datos cada segundo durante 120 segundos y cambia los niveles del calentador cada 30 segundos a un número aleatorio entre 0 y 80 con `np.random.randint()`. No es necesario cambiar este programa, solo ejecútalo para recolectar los datos durante 2 minutos.
```
import tclab, time, csv
import numpy as np
try:
n = 120
with open('04-tclab.csv',mode='w',newline='') as f:
cw = csv.writer(f)
cw.writerow(['Time','Q1','Q2','T1','T2'])
with tclab.TCLab() as lab:
print('t Q1 Q2 T1 T2')
for t in range(n):
if t%30==0:
Q1 = np.random.randint(0,81)
Q2 = np.random.randint(0,81)
lab.Q1(Q1); lab.Q2(Q2)
cw.writerow([t,Q1,Q2,lab.T1,lab.T2])
if t%5==0:
print(t,Q1,Q2,lab.T1,lab.T2)
time.sleep(1)
data4=pd.read_csv('04-tclab.csv')
except:
print('Conectar al TCLab para generar datos')
url = 'http://apmonitor.com/do/uploads/Main/tclab_dyn_data2.txt'
data4=pd.read_csv(url)
data4.columns = ['Time','Q1','Q2','T1','T2']
data4.head()
```
### Análisis Gráfico
Analizar `Q1`, `Q2`, `T1`, y `T2` gráficamente con un gráfico de series de tiempo y un gráfico de pares. La serie de tiempo debe mostrar `Q1` y `Q2` en la subgráfica superior, y `T1` y `T2` en la subgráfica inferior. El diagrama de pares debe ser un gráfico de cuadrícula `2x2` que muestra los pares calentador / temperatura como `Q1`/`T1`, `Q2`/`T2`.
| github_jupyter |
# 文章要約アプリの作成
## 1週目
「文章要約 API」と検索していただくと、株式会社リクルートテクノロジーズが開発した`Summpy`というAPIがヒットするかと思います。いろいろと調べてみたのですが、一般向けに公開している文章要約APIは`Summpy`が良いようなのでこちらを使っていきたいと思います。
実際は、`Summpy`とは別の`Text Summarization API`というこれまたリクルートが公開しているAPIもあるのですが、こちらは **「1文の最大文字数は200文字、且つ最大文章数は10」** という制限があるため、Summpy使用の方向で進めていきたいと思います。
※最大文章数10はなかなか厳しいです
`Text Summarization API`に興味のある方は[こちら](https://a3rt.recruit-tech.co.jp/product/TextSummarizationAPI/)を参考に実装してみてください。そこまで難しくないと思います。
### Summpy使用にあたり
以下の記事にてSummpyが初めて公開されました。
[自動要約APIを作ったので公開します](https://recruit-tech.co.jp/blog/2015/10/30/summpy-released/)
記事のリンク先にGithubへのリンクがあったため、そちらを参考に進めていこうと考えました。
しかし、以下のGithubに公開されているものが`python2`で書かれており、python3で使用できない形となっております。
https://github.com/recruit-tech/summpy
python3バージョンの`Summpy`のコードがないかをいろいろと検索してみたら、python3に変更してくれた方がいました。
そちらを使用してみようと試みてみましたが、多々エラーが起こってしまいうまくいかなかったです。
https://github.com/boarnasia/summpy
そこで、かなり大変ではありますがpython2用の、`summpy`を用いてpython3に対応できるように修正していきます。
### gitからクローンする
まず、作業ディレクトリとして、`text_summary`というディレクトリを作成し、そのディレクトリ内で作業を進めていきます。
`jupyter`のホームから`terminal`を開いてください。 以下のコマンドでGithubからクローンしてきましょう。
```
git clone https://github.com/recruit-tech/summpy.git
```
`summpy`というフォルダ内に`summpy`というフォルダがあるのでややこしいですが、必要なファイルはすべて`summpy/summpy`の中にあります。
不要なファイル類を削除するために以下のコマンドを打ちましょう。
まず、`summpy`を`_summpy`に変更します。
`mv`: ファイルを移動する際に使用するコマンド
```
mv summpy _summpy
```
次に`_summpy/summpy`の中の全てのファイルを`text_summary`に移動します。
```
mv ./_summpy/summpy ./
```
そして、不要なディレクトリである`_summpy`を削除しましょう。
```
rm -r -f _summpy
```
`ls`で確認し、`summpy`ディレクトリのみになっていたら成功です。
### python2 -> python3 に変換
python2とpython3の違いはいくつかありますが、このコードがpython2で書かれているのものだなと気づくときの多くが`print文`を見たときです。
[Python 2.7.x と 3.x の決定的な違いを例とともに](https://postd.cc/the-key-differences-between-python-2-7-x-and-python-3-x-with-examples/)
- python2 : print 'Hello World'
- python3 : print('Hello World' )
上記のように、python2は`()`が存在しません。このような違いを1つずつ修正していくのは面倒だなと思ったので、
**「python2 to python3」**と調べてみたところ`2to3`というpytho2をpython3に書き換えてくれるものが見つかったのでこちらを使っていきましょう。
以下のサイトを参考に3系にコードを変換します。
https://docs.python.jp/3/library/2to3.html
以下のコードで`summpy`のフォルダ内のpythonファイルを全て3系に変換しましょう。
既存の2系ファイルが全て3系ファイルに書き換えられます。
- -w : ファイルを変換結果に置き換える場合
- -n : バックアップファイルが不要な場合
- -f all : 全ての変換を適用する場合
```
2to3 -f all -n -W ./summpy/
```

これでPython3への変換が完了しました。
わかりやすい目安としては、`print`文を確認すると、3系に変換されたことが確認できます。
- python2:print text
- python3:print(text)
それでは、`summpy`フォルダと同階層に`python_summpy`というjupyter notebookを作成しましょう。
Githubの`README.md`のやり方に沿って文章要約をしていきます!
### 使用するテキストの読み込み
まずは、今回要約に用いるテキストファイルを以下からダウンロードしましょう。
[kikagaku.txt](kikagaku.txt)
このテキストは弊社の[こちら](https://www.kikagaku.co.jp/column/kikagaku-1st/)のブログ内容を使用しております。
それでは早速、テキストファイルを読み込みましょう。
```
with open('kikagaku.txt') as f:
text = ''.join(f.readlines())
```
こちらは**自然言語**の講義にて記事ファイルを読み込む際にお伝えしました。
```
text
len(text)
```
7953文字と長めの文章です。
### 文章要約の実装
文章要約のメインが`summpy/lexrank.py`の中の`summarize`にあたるので、そちらを読み込みましょう。
```
from summpy.lexrank import summarize
```
上記のように読み込もうとするとエラーが起きてしまいますが、これは `networkx`をインストールしていないことが原因のため、`networkx`をインストールしましょう。
```
!pip install networkx
```
それでは、もう一度読み込みます。
```
from summpy.lexrank import summarize
```
特にエラーがおきていなければ成功です。では、引き続き`README.md`通りに進めて文章要約を行います。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
python3への変換を完了したのでしっかり実装できるかと思ったのですが、そう簡単には実装できないようです。
エラー文をみてみると、`MeCab`のエラーのようです。
Google検索で`「in method 'Tagger_parseToNode', argument 2 of type 'char const *'」`と調べると、MeCabで`Tagger`を読み込んだ後に一度空文字でパースしてあげると良いようです。
【参考記事】
https://shogo82148.github.io/blog/2015/12/20/mecab-in-python3-final/
https://teratail.com/questions/88592
以下画像の赤下線のファイルでエラーが起きているので、上から順にたどっていきます。上から3番目の`text_summary/summpy/misc/mecab_segmenter.py`ファイル`67行目`の関数を実行したタイミングで`MeCab`のエラーが起きていることを確認できます。

そこで、`text_summary/summpy/misc/mecab_segmenter.py`ファイルを開くと、8行目でTaggerを読み込んでいる部分があるので、9行目に以下の1行を入れて保存しましょう。
```
_mecab = MeCab.Tagger()
_mecab.parse('') #追記
```
それでは、もう一度実行したいところなのですが、ここで1点注意点があります。
pythonファイルを書き換えた後、JupyterNotebook上でもう一度`import`してもうまく反映されないことが多々あります。そのため少し面倒ですが、`python_summpy`を一度シャットダウンしてから再度開き、実行しましょう。
※上記のシャットダウン→実行の流れは今後、ファイル内のコード変更のたびに行って下さい。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
変更してもなお同じエラーが起きてしまいます。どの記事もたいてい同じような解決策が書いてあり、参考にならなそうです。
そこで、大変ではありますが原因追求のため、試行錯誤していきたいと思います。
※私がどのような手順で解決していったのか流れを書いていきます。
#### 試行錯誤1
`parseToNode(self, *args)`というMeCabの関数を呼び出したタイミングでエラーが起きているので、関数実行時に渡している引数がなにかおかしいではないかと考えてみます。つまり、`mecab_segmenter.py`68行目にある`sent`という変数に問題がないかを確認していきます。
以下のように`for文`の前後で`sent`の方を確認します。
`mecab_segmenter.py`

それでは、再起動し実行しましょう。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
出力を確認してみると、`if文`の前が`str型` でif文内にてエンコードされた関係で、`bytes型`になっています。
```
<class 'str'>
<class 'bytes'>
```
pythonの`str型`と`bytes型`の違いについては以下の記事をご覧ください。
https://www.kannon.link/fuku/index.php/2017/02/22/01-34/
#### 試行錯誤2
ここで私は「`str型`のままではダメなのかな?」と疑問に思ったので、`if文`をコメントアウトしてみます。

もう一度実行してみます。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
#### 試行錯誤3
実行してみると違うエラーが出てきました。もしかしたら一歩前進したかもしれません。今度のエラー`'str' object has no attribute 'decode'`は「`str型`は`decode`できませんよ」というエラー内容です。
そのため、`mecab_segmenter.py`23行目と26行目の`.decode()`の部分を消してしまいましょう。
先程の`print文`2つも削除しておきましょう。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
#### 試行錯誤4
また違うエラーが出てきました。`「AttributeError: __next__」`とGoogleで調べていただくと、
pythonには、**generatorクラス**というものがあり、その中のメソッドに`next`があるみたいです。
【参考記事】
https://qiita.com/yuragawa/items/b2022b534633114207e9 <br/>
https://stackoverflow.com/questions/12274606/theres-no-next-function-in-a-yield-generator-in-python-3
上記の記事などを参考に以下のパターンを試してみました。
- .next()
- .\__next\__()
- next(node)
- .next
今回は`node = node.__next__`ではなく、`node = node.next`にしたほうが良いみたいです。
`mecab_segmenter.py`29行目を変更しましょう。

```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
#### 試行錯誤5
また違うエラーが出てきました。。。
先程までと同様にGoogleで調べてみると[こちら](https://github.com/ewels/MultiQC/issues/592)の記事に以下のような記述がありました。
```
pip install networkx==1.11
pip install multiqc==1.2
```
もしかしたら`networkx`と`multiqc`というライブラリのバージョンが違うのではないかと思い、確認してみました。
```
import networkx
import multiqc
networkx.__version__
```
`networkx`のバージョンは記事とは異なり、`multiqc`はインストールされていなかったです。
そこで`networkx`をアンインストール後、バージョン指定で再インストール、`multiqc`はバージョン指定でインストールしてみます。
以下のコマンドをターミナル打ちましょう。
```bash
pip uninstall networkx
pip install networkx==1.11
pip install multiqc==1.2
```
シャットダウンし、もう一度実行してみましょう。
```
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
```
ようやくエラーが起こることなく通りました!!
最後まで実装しましょう!
```
for sent in sentences:
print(sent.strip())
```
実際に5つの文章で要約されましたが、いかがでしょうか。 個人的には今回の記事の中でも重要な文章をいくつか抜粋してきてくれているなと感じています。
すべての文章がうまく抜粋できているわけではないですが、記事の概要をざっとつかめるような結果となっています。
5つの文ではなく、もう少し文章数を増やしたり減らしたりしたら結果がどう変わるかも考察してみてください!
もしかしたら要約に適切な文章数が見つけられるかもしれません。
## 2週目
### 英文要約を行おう
先週は日本語要約として`summpy`についてお伝えしました。今週最初は英文における文章要約です。英語の文章要約は`summa-textrank`というものを使用しましょう。以下のGithubを参考に進めていきます。
https://github.com/summanlp/textrank
#### ライブラリのインストール
```
!pip3 install summa --user
```
#### 使用する英文
今回は以下の記事を使用していきましょう。
https://www.japantimes.co.jp/sports/2018/06/20/soccer/early-present-helps-japan-claim-perfect-start-world-cup-campaign/
[こちら](english.txt)が上記の英文をテキストファイルにしたものになります。
```
with open('english.txt', encoding='utf-8') as f:
text = ''.join(f.readlines())
print(text)
```
まずは、インポートしましょう!`summa.summarizer`の中の`summarize`という関数を使用します。
```
from summa.summarizer import summarize
```
`summa`は`summpy`と違い、`〇文`で文章要約するのではなく、`○単語`以内で要約する形となっております。今回は**50単語**以内で要約してみましょう。
```
#50単語以内での要約
summarize(text, words=50)
```
デフォルトで英語の要約となっていますが、引数`language='english'`の部分を`spanish`に変更すればスペイン後の要約も可能。
また、以下のように文ごとに`split`することも可能です。
```
#50単語以内での要約
summarize(text, words=50, split=True)
```
英語なのでしっかりと要約できているのか定かではないですが、上記の様に非常に簡単に英語の文章要約をすることが可能です。
#### 補足(キーワード抽出)
`summa`を用いると以下のようにキーワード抽出も行ってくれます。
```
from summa import keywords
print(keywords.keywords(text))
```
#### 問題
今回の`summa`を用いた文章要約では以下の言語に対応しておりますが、下記の**対応言語**はどこを探せば見つかるでしょうか。
##### 対応言語
- danish(デンマーク語)
- dutch(オランダ語)
- english(英語)
- finnish(フィンランド語)
- french(フランス語)
- german(ドイツ語)
- hungarian(ハンガリー語)
- italian(イタリア語)
- norwegian(ノルウェー語)
- porter(不明)
- portuguese(ポルトガル語)
- romanian(ルーマニア語)
- russian(ロシア語)
- spanish(スペイン語)
- swedish(スウェーデン語)
#### 回答
`README.md`を確認すれば簡単に見つかりました。。。
私はてっきり書いていないものだと思いこんでいたため以下のように探しました。ご参考までに。
まずは、`summarize()`関数にて`language`という引数を取っているため、`summa > summarizer > summarize`を確認しましょう。その次に、以下の順番で引数`language`が使用されている関数をたどっていくと必要とする情報にたどり着きます。
- preprocessing > textcleaner > clean_text_by_sentences
- preprocessing > textcleaner > init_textcleanner
- preprocessing > textcleaner > set_stemmer_language
`summa > preprocessing > textcleaner.py`の`43〜49行目`に以下のような記述部分があります。以下を確認することで、先程の**対応言語**を見つけることができました。
```python
def set_stemmer_language(language):
global STEMMER
if not language in LANGUAGES:
raise ValueError("Valid languages are danish, dutch, english, finnish," +
" french, german, hungarian, italian, norwegian, porter, portuguese," +
"romanian, russian, spanish, swedish")
STEMMER = SnowballStemmer(language)
```
### 日本語文と英文に対応できる関数
次に、入ってくるテキストが**日本語**か**英語**かによって場合分けするような関数を作成しましょう。
#### 英語と日本語の場合分け
まずは、英語か日本語かを場合分けするような`if文`を作成する必要があります。先程読み込んだ英文を`text_english`、先週使用した`kikagkau.txt`を`text_japananese`という変数にしておきましょう。
```
text_english = text
text_english
```
日本語は前回使用したテキストを使用します。
```
with open('kikagaku.txt', encoding='utf-8') as f:
text_japanese = ''.join(f.readlines())
```
日本語かどうかの判定を`Unicode` の違いによって条件分岐を行っています。
```
import unicodedata
def is_japanese(text):
for _text in text:
name = unicodedata.name(_text)
if "CJK UNIFIED" in name \
or "HIRAGANA" in name \
or "KATAKANA" in name:
return True
return False
is_japanese(text_japanese)
```
日本語の文章であるため、しっかりと`True`と判定されました。同様に`text_english`でも行ってみましょう。
```
is_japanese(text_english)
```
なぜか英語の文だとうまくいかないみたいなので、このエラーを解決していきましょう。エラーは`name = unicodedata.name(_text) `の部分で起きているため、引数にとっている`_text`が原因ではないかと考えられます。以下のように`print文`で`_text`を出力してみましょう。
```
import unicodedata
def is_japanese(text):
for _text in text:
print(_text)
name = unicodedata.name(_text)
if "CJK UNIFIED" in name \
or "HIRAGANA" in name \
or "KATAKANA" in name:
return True
return False
is_japanese(text_english)
```
出力を確認してみると、`.`もしくは、`空白`の影響でエラーが起きている気がします。もう少し詳しくみてみましょう。
```
import unicodedata
def is_japanese(text):
for _text in text:
print(_text+'\n-----')
name = unicodedata.name(_text)
if "CJK UNIFIED" in name \
or "HIRAGANA" in name \
or "KATAKANA" in name:
return True
return False
is_japanese(text_english)
```
確認してみると、`.`ではなく、`空白`が入ったタイミングでエラーが起きています。実際の文章のどの部分でエラーが起きているのかを確認してみましょう。
```
text_english
```
文章を確認してみると、以下の**改行**を表す`\n`でエラーが起きているのがわかります。

この`\n`がエラーの原因である可能性が高いため、こちらを**空文字**で置き換えてあげましょう。
※おそらくエスケープシーケンス(`\n`)にUnicodeネームがないためのエラー
```
def is_japanese(text):
text = text.replace('\n', '')
for _text in text:
name = unicodedata.name(_text)
if "CJK UNIFIED" in name \
or "HIRAGANA" in name \
or "KATAKANA" in name:
return True
return False
is_japanese(text_english)
```
しっかりと`False`と判定されました!こちらの関数をもとに英語、日本語の両方に対応した要約を行いましょう。
```
from summpy.lexrank import summarize
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
```
エラーが起きてしまいました。こちらは前回最後にデバッグした部分のエラーです。前回の対処法として、`networkx`のバージョンを`1.11`で指定してインストールし直しました。、英語要約の`summa`の裏側では`networkx`が`2.0`で動いているのに対し、日本語要約の`summpy`では、`1.11`で動いている状況です。これをどちらかに揃えないといけないのが今回の課題です。
```
import networkx
networkx.__version__
```
日本語要約の`summpy`側を`networkx==2.0`で動かすことができないかを考えていきましょう。前回バージョンを下げて対応した**エラー文**と** エラー箇所**を再度確認してみます。

エラー内容である`add_edge() takes 3 positional arguments but 4 were given`は「`add_edge()`関数は`3つの引数`を取るはずなのに、`4つ`与えられてしまっているよ」という内容のエラーです。`add_edge()`の引数は、`i`, `j`, `{'weight': weight}`の3つのはずですが、エラーが起きてしまっています。
実際に`networkx`の`add_edge`という関数はどのように引数を取るのが正解であるかがわからないため、`「networkx add_edge」`とGoogleで検索してみましょう。
そうすると、以下の記事がヒットするかと思います。
[networkx.Graph.add_edge — NetworkX 2.1 documentation](https://networkx.github.io/documentation/stable/reference/classes/generated/networkx.Graph.add_edge.html)

このドキュメント内の下方に`add_edge()`における引数(`weight`)の指定の仕方が書かれています。

確認してみると`{'weight': weight}`ではなく、`weight=3`のように指定しているのがわかります。そこで`summpy/lexrank.py`の85行目引数の部分を以下のように変更し、保存しましょう。

一度`Jupyter notebook`をシャットダウンし、もう一度実行してみてください。
```
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
```
エラーが起きずに実行されたかと思います。要約された文章がしっかり入っているか確認してみましょう。
```
for sent in sentences:
print(sent.strip())
```
しっかりと要約できているのが確認できました。それでは、本題である**「英語と日本語に対応する文章要約」**に戻りましょう。
一旦、引数のパラメータの値は固定して実装していきます。
#### 日本語の場合
```
text = text_japanese
if is_japanese(text):
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
print(sentences)
```
#### 英語の場合
```
text = text_english
# 日本語の場合
if is_japanese(text):
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
print(sentences)
#英語の場合
else:
summarize(text, words=50, split=True)
print(sentences)
```
先程と同様に行ったのですが、エラーが起きてしまいました。今回の原因は、日本語要約`summpy`側の関数名が`summarize`であり、英語要約`summa`側の関数も`summarize`であることが原因です。そのため、以下のように修正しましょう。
```
# from summa.summarizer import summarize
from summa import summarizer
text = text_english
# 日本語の場合
if is_japanese(text):
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
print(sentences)
#英語の場合
else:
sentences = summarizer.summarize(text, words=50, split=True)
print(sentences)
```
これで日本語と英語の両方に対応させることができたため、関数化しましょう。
```
def get_summary(text):
# 日本語の場合
if is_japanese(text):
sentences, debug_info = summarize(
text_japanese, sent_limit=5, continuous=True, debug=True
)
#英語の場合
else:
sentences = summarizer.summarize(text, words=50, split=True)
return sentences
get_summary(text_japanese)
get_summary(text_english)
```
### Pythonファイルからの読み込み
上記で作成した`get_summary`を別のPythonファイルに移し、そのPythonファイルを読みこむことで、関数を呼び出してください。Pythonファイルの名前は、`summary.py`にしましょう。
```
from summary import get_summary
get_summary(text_japanese)
get_summary(text_english)
```
上記のようにインポートできたら成功です。`summary.py`のコードは以下になります。
```python
import unicodedata
from summpy.lexrank import summarize
from summa import summarizer
def is_japanese(text):
text = text.replace('\n', '')
for _text in text:
name = unicodedata.name(_text)
if "CJK UNIFIED" in name
or "HIRAGANA" in name
or "KATAKANA" in name:
return True
return False
def get_summary(text):
# 日本語の場合
if is_japanese(text):
sentences, debug_info = summarize(
text, sent_limit=5, continuous=True, debug=True
)
#英語の場合
else:
sentences = summarizer.summarize(text, words=50, split=True)
return sentences
```
## 3週目
### 文章要約モデルをアプリに埋め込む
#### 必要なページ
今回は大きく分けて以下の2つのページで構成されています。
- TOPページ
- 要約後のページ
##### TOPページ(index.html)

##### 要約後のページ(summary.html)

##### 扱うライブラリ
`Materialize.css`を使用してレイアウトなどを整えていきます。
[Materialize.css](https://materializecss.com/)
これは個人の好みの問題なので、`Bootstrap`等を使用しても大丈夫です。
前回同様Dockerを使用して作成していくのがスムーズかもしれません。
### TOPページの作成
#### index.html
```html
<!DOCTYPE html>
<html>
<head>
<!--Import Google Icon Font-->
<link href="https://fonts.googleapis.com/icon?family=Material+Icons" rel="stylesheet">
<!--Import materialize.css-->
<!-- Compiled and minified CSS -->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/materialize/1.0.0-rc.2/css/materialize.min.css">
<meta charset="utf-8">
<title>文章要約</title>
<link rel="stylesheet" href="static/css/index.css">
</head>
<body>
<div class="row">
<!-- header -->
<header class="col s12 teal lighten-1">
<div class="col s12 offset-s1">
<a href="/"><p class="white-text">文章要約アプリ</p></a>
</div>
</header>
<div class="input_text input-field col s5 offset-s1">
<form id="summarize" action="/" method=post enctype=multipart/form-data>
<div class="input_text input-field">
<textarea name="text" class="materialize-textarea"></textarea>
<label for="textarea1">Type text you want to summarize</label>
</div>
<div class="col s2 offset-s9">
<button class="btn waves-effect waves-light" form="summarize" type="submit" name="summarize" value=Summarize>Summarize
</button>
</div>
</form>
</div>
<div class="output_summary input-field col s5">
<div class="input_text input-field">
<textarea id="textarea1" class="materialize-textarea" disabled>
</textarea>
<label for="textarea1">Summary</label>
</div>
</div>
<div class="col s1">
</div>
</div>
</body>
</html>
```
```html
<!-- Compiled and minified CSS -->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/materialize/1.0.0-rc.2/css/materialize.min.css">
```
上記の部分で、`materialize.min.css`をCDN経由で読み込んでいます。
レイアウトは、以下の`Grid`の部分を参考に行っています。基本的には大枠に`class="row"`を付与し、その中身に`class="col s6"`などと書き、**12カラム**に分割し考えています。
[](https://materializecss.com/grid.html)
```html
<header class="col s12 teal lighten-1">
<div class="col s12 offset-s1">
<a href="/"><p class="white-text">文章要約アプリ</p></a>
</div>
</header>
```
こちらが`header`部分になります。ヘッダー部分をクリックすると、**TOPページ**に移行します。
```html
<div class="input_text input-field col s5 offset-s1">
<form id="summarize" action="/" method=post enctype=multipart/form-data>
<div class="input_text input-field">
<textarea name="text" class="materialize-textarea"></textarea>
<label for="textarea1">Type text you want to summarize</label>
</div>
<div class="col s2 offset-s9">
<button class="btn waves-effect waves-light" form="summarize" type="submit" name="summarize" value=Summarize>Summarize
</button>
</div>
</form>
</div>
```
こちらが要約したいテキストを表示する部分になります。以下のサイトを参考にしつつ、フォームを作成しました。
https://materializecss.com/text-inputs.html
```html
<div class="output_summary input-field col s5">
<div class="input_text input-field">
<textarea id="textarea1" class="materialize-textarea" disabled>
</textarea>
<label for="textarea1">Summary</label>
</div>
</div>
<div class="col s1">
</div>
```
上記は、要約した文章を入れる部分になります。まだ記述していないですが、後々、`app.py`から渡される要約結果を表示します。
#### app.py
```python
# coding: utf-8
from flask import Flask, render_template, request, redirect, url_for
# app という名前でFlaskオブジェクトをインスタンス化
app = Flask(__name__)
# rootディレクトリにアクセスした場合の挙動
@app.route('/', methods=['GET', 'POST'])
def index():
return render_template("index.html")
# --- メインで実行される関数 ---
if __name__ == "__main__":
app.run(host="0.0.0.0", port=5000, debug=True)
```
こちらはいつもどおりFlaskの基本的な部分となっております。
### 要約後のページ
#### summary.html
```html
<!DOCTYPE html>
<html>
<head>
<!--Import Google Icon Font-->
<link href="https://fonts.googleapis.com/icon?family=Material+Icons" rel="stylesheet">
<!--Import materialize.css-->
<!-- Compiled and minified CSS -->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/materialize/1.0.0-rc.2/css/materialize.min.css">
<meta charset="utf-8">
<title>文章要約</title>
</head>
<body>
<div class="row">
<!-- header -->
<header class="col s12 teal lighten-1">
<div class="col s12 offset-s1">
<a href="/"><p class="white-text">文章要約アプリ</p></a>
</div>
</header>
<div class="input_text input-field col s5 offset-s1">
<form id="summarize" action="/" method=post enctype=multipart/form-data>
<div class="input_text input-field">
<textarea name="text" class="materialize-textarea"></textarea>
<label for="textarea1">Type text you want to summarize</label>
</div>
<div class="col s2 offset-s9">
<button class="btn waves-effect waves-light" form="summarize" type="submit" name="summarize" value=Summarize>Summarize
</button>
</div>
</form>
</div>
<div class="output_summary input-field col s5">
<div class="input_text input-field">
<textarea id="textarea1" class="materialize-textarea" disabled>
{% for summary in summaries %}
{{summary.replace('\r', '').replace('\n', '')}}
{% endfor %}
</textarea>
<label for="textarea1">Summary</label>
</div>
</div>
<div class="col s1">
</div>
</div>
</body>
</html>
```
`index.html`に加えて変更した点は最後の以下の部分です。`app.py`から引き渡される変数`summaries`をfor文で1文ずつ取得しています。その際に、無駄なエスケープシーケンスが多く含まれているので`replace`を用いて除去します。
```html
<div class="output_summary input-field col s5">
<div class="input_text input-field">
<textarea id="textarea1" class="materialize-textarea" disabled>
{% for summary in summaries %}
{{summary.replace('\r', '').replace('\n', '')}}
{% endfor %}
</textarea>
<label for="textarea1">Summary</label>
</div>
</div>
```
#### ファイル階層
文章要約に必要な`summary.py(前回作成済み)`と`summpy`フォルダを`app.py`と同じ階層に配置します。
```bash
.
├── __pycache__
│ └── summary.cpython-36.pyc
├── app.py
├── static
├── summary.py
├── summpy
│ ├── __init__.py
│ ├── __pycache__
│ │ ├── __init__.cpython-36.pyc
│ │ ├── lexrank.cpython-36.pyc
│ │ └── tools.cpython-36.pyc
│ ├── lexrank.py
│ ├── mcp_summ.py
│ ├── misc
│ │ ├── __init__.py
│ │ ├── __pycache__
│ │ │ ├── __init__.cpython-36.pyc
│ │ │ ├── divrank.cpython-36.pyc
│ │ │ └── mecab_segmenter.cpython-36.pyc
│ │ ├── divrank.py
│ │ ├── janome_segmenter.py
│ │ └── mecab_segmenter.py
│ ├── server.py
│ ├── server_data
│ │ └── test.html
│ └── tools.py
└── templates
├── index.html
└── summary.html
```
#### app.py
```python
# coding: utf-8
from flask import Flask, render_template, request, redirect, url_for
# 追加
from summary import get_summary
# app という名前でFlaskオブジェクトをインスタンス化
app = Flask(__name__)
# rootディレクトリにアクセスした場合の挙動
@app.route('/', methods=['GET', 'POST'])
def index():
if request.method == 'POST':
text = request.form['text']
summaries = get_summary(text)
return render_template("summary.html", summaries=summaries)
return render_template("index.html")
# --- メインで実行される関数 ---
if __name__ == "__main__":
app.run(host="0.0.0.0", port=5000, debug=True)
```
最初に追加した部分は、`summary`のimport部分です。
```python
from summary import get_summary
```
```python
def index():
if request.method == 'POST':
text = request.form['text']
summaries = get_summary(text)
return render_template("summary.html", summaries=summaries)
return render_template("index.html")
```
`index.html`からテキストが`POST`されたら、そのテキストを読み込み`text`という変数に代入しています。その`text`を`get_summary()`に渡し、その要約結果`summaries`を`summary.html`に渡しています。
### 発展課題
上記を終えた方は以下のような課題に取り組んでみるのも面白いと思います。
- レイアウトのブラッシュアップ
- 多言語対応
- 要約時のパラメータも調整可能
| github_jupyter |
# Using AXI GPIO with PYNQ
## Goal
The aim of this notebook is to show how to use AXI GPIO from PYNQ.
Multiple AXI GPIO controllers can be implemented in the programmable logic and used to control internal or external GPIO signals.
## Hardware design
This example uses a bitstream that connects three AXI GPIO controllers to the LEDs, buttons, and switches and can be used with the PYNQ-Z1 or PYNQ-Z2 board. (Each AXI GPIO controller has 2 channels, so multiple peripherals could be controlled from one AXI GPIO IP, but for simplicity and demonstration purposes, separate AXI GPIO controllers are used.

### 1. Download the tutorial overlay
The `axi_gpio.bit` and `axi_gpio.tcl` files can be found in the bitstreams directory local to this folder.
The bitstream can be downloaded by passing the relative path to the Overlay class.
* Check the bitstream and .tcl exists in the bitstream directory
```
!dir ./bitstream
```
* Download the bitstream
```
from pynq import Overlay
axi_gpio_design = Overlay("./bitstream/axi_gpio.bit")
```
Check the IP Dictionary for this design. The IP dictionary lists AXI IP in the design, and for this example will list the AXI GPIO controllers for the buttons, leds, and switches. The Physical address, the address range and IP type will be listed. If any interrupts, or GPIO were connected to the PS, they would also be reported.
```
axi_gpio_design.ip_dict
```
## AxiGPIO class
The PYNQ AxiGPIO class will be used to access the AXI GPIO controllers.
### 1. Controlling the switches and push-buttons
The instances can be found and referenced from the IP dictionary.
```
from pynq.lib import AxiGPIO
buttons_instance = axi_gpio_design.ip_dict['buttons']
buttons = AxiGPIO(buttons_instance).channel1
buttons.read()
```
The buttons controller is connected to all four user push-buttons on the board (BTN0 to BTN3). Try pressing any combination of the buttons and rerunning the cell above.
The AXI GPIO controller for the switches can be used in a similar way:
```
switches_instance = axi_gpio_design.ip_dict['switches']
switches = AxiGPIO(switches_instance).channel1
print(f"Switches: {switches.read()}")
```
### 2. Controlling the LEDs
The LEDs can be used in a similar way.
```
from pynq.lib import AxiGPIO
led_instance = axi_gpio_design.ip_dict['leds']
led = AxiGPIO(led_instance).channel1
```
The outputs can be addressed using a slice.
```
led[0:4].write(0x1)
from time import sleep
led[0:4].write(0x3)
sleep(1)
led[0:4].write(0x7)
sleep(1)
led[0:4].write(0xf)
```
* Reset the LEDs
```
led[0:4].off()
```
### 3 Putting it together
Run a loop to set the LEDs to the value of the pushbuttons.
Before executing the next cell, make sure Switch 0 (SW0) is "on". While the loop is running, press a push-button and notice the corresponding LED turns on. To exist the loop, change Switch 0 to off.
```
while(switches.read() is 1):
led[0:4].write(buttons.read())
```
| github_jupyter |
## The Oren-Nayar (1994) reflectance model
In this notebook we derive and validate our approximation to the Oren-Nayar scattering model.
```
%matplotlib inline
%run notebook_setup.py
import matplotlib.pyplot as plt
import numpy as np
np.seterr(invalid="ignore");
```
Here's the standard Lambertian reflectance as a function of `b` and `theta` at a point `(x, y)` on the surface, as derived in the Appendix of the `starry` paper. Recall that `b` is the semi-minor axis of the day/night terminator and `theta` is the orientation of the terminator with respect to the `x` axis:
```
@np.vectorize
def Lambertian(b, theta, x, y):
"""Lambertian intensity at a point on the surface."""
ct = np.cos(theta)
st = np.sin(theta)
bc = np.sqrt(1 - b ** 2)
z = np.sqrt(1 - x ** 2 - y ** 2)
ci = -bc * st * x + bc * ct * y - b * z
return np.maximum(0, ci)
```
And here's the Oren-Nayar (1994) reflectance from their Equation (30), expressed as a function of the same `starry` parameters:
```
def R(axis=[0, 1, 0], theta=0):
"""Axis-angle rotation matrix in 3D."""
axis = np.array(axis) / np.sqrt(np.sum(np.array(axis) ** 2))
cost = np.cos(theta)
sint = np.sin(theta)
return np.reshape(
[
cost + axis[0] * axis[0] * (1 - cost),
axis[0] * axis[1] * (1 - cost) - axis[2] * sint,
axis[0] * axis[2] * (1 - cost) + axis[1] * sint,
axis[1] * axis[0] * (1 - cost) + axis[2] * sint,
cost + axis[1] * axis[1] * (1 - cost),
axis[1] * axis[2] * (1 - cost) - axis[0] * sint,
axis[2] * axis[0] * (1 - cost) - axis[1] * sint,
axis[2] * axis[1] * (1 - cost) + axis[0] * sint,
cost + axis[2] * axis[2] * (1 - cost),
],
[3, 3],
)
@np.vectorize
def OrenNayarExplicit(b, theta, sig, x, y):
"""Oren-Nayar intensity at a point on the surface from Equation (30) in Oren & Nayar (1994)."""
# Compute the three vectors in the observer (sky) frame
bc = np.sqrt(1 - b ** 2)
s0 = np.array([-bc * np.sin(theta), bc * np.cos(theta), -b]) # source vector
v0 = np.array([0, 0, 1.0]) # observer vector
n0 = np.array([x, y, np.sqrt(1 - x ** 2 - y ** 2)]) # normal vector
# Transform to the surface normal frame (in which the Oren-Nayar model is defined)
n = np.array([0, 0, 1.0]) # normal vector
# Find the rotation matrix that transforms to
# the normal frame
axis = np.cross(n0, n)
costhet = np.dot(n0, n)
thet = np.arccos(costhet)
Rn = R(axis, -thet)
# Check that the sign is right
# Rn.dot(n0) should be equal to n
# (if it's not, we need to rotate
# by `thet`, not `-thet`)
Rnp = R(axis, thet)
if Rnp.dot(n0)[2] > Rn.dot(n0)[2]:
Rn = Rnp
# Rotate s and v
s = Rn.dot(s0)
v = Rn.dot(v0)
# Compute the angles
theta_r = np.arccos(np.dot(v, n))
theta_i = np.arccos(np.dot(s, n))
if theta_i >= np.pi / 2:
# Unilluminated!
return 0
phi_r = np.arctan2(v[1], v[0])
phi_i = np.arctan2(s[1], s[0])
del_phi = phi_i - phi_r
# Now compute the intensity
alpha = max(theta_r, theta_i)
beta = min(theta_r, theta_i)
f = max(0, np.cos(del_phi)) * np.sin(alpha) * np.tan(beta)
sig2 = sig ** 2
A = 1 - 0.5 * sig2 / (sig2 + 0.33)
B = 0.45 * sig2 / (sig2 + 0.09)
S = A + B * f
return Lambertian(b, theta, x, y) * S
```
First, we're going to show that the above model is equivalent to the following, much simpler, equation:
```
@np.vectorize
def OrenNayar(b, theta, sig, x, y):
"""Oren-Nayar intensity at a point on the surface (simplified)."""
ct = np.cos(theta)
st = np.sin(theta)
bc = np.sqrt(1 - b ** 2)
z = np.sqrt(1 - x ** 2 - y ** 2)
ci = -bc * st * x + bc * ct * y - b * z
f1 = -b / z - ci
f2 = -b / ci - z
sig2 = sig ** 2
A = 1 - 0.5 * sig2 / (sig2 + 0.33)
B = 0.45 * sig2 / (sig2 + 0.09)
S = A + B * np.maximum(0, np.minimum(f1, f2))
return Lambertian(b, theta, x, y) * S
```
Let's plot the two models for a variety of roughnesses and compare the output images:
```
# Params
res = 100
theta = np.pi / 3
b = -0.5
sig = 1.0
# Grid up the surface
grid = np.linspace(-1, 1, res)
x, y = np.meshgrid(grid, grid)
# Plot
fig, ax = plt.subplots(3, 6)
for i, sig in enumerate(np.array([0, 10, 20, 40, 60, 90]) * np.pi / 180):
# Explicit
IExplicit = OrenNayarExplicit(b, theta, sig, x, y)
ax[0, i].imshow(
IExplicit, origin="lower", extent=(-1, 1, -1, 1), cmap="plasma", vmax=1, vmin=0
)
ax[0, i].set(frame_on=False, xticks=[], yticks=[])
# Re-parametrized
IReparam = OrenNayar(b, theta, sig, x, y)
ax[1, i].imshow(
IReparam, origin="lower", extent=(-1, 1, -1, 1), cmap="plasma", vmax=1, vmin=0
)
ax[1, i].set(frame_on=False, xticks=[], yticks=[])
# Difference
ax[2, i].imshow(
np.abs(IExplicit - IReparam),
origin="lower",
extent=(-1, 1, -1, 1),
cmap="plasma",
vmax=1e-15,
vmin=0,
)
ax[2, i].set(frame_on=False, xticks=[], yticks=[])
ax[0, i].set_title(r"$\sigma = {:.0f}^\circ$".format(sig * 180 / np.pi))
ax[0, 0].set_ylabel("explicit")
ax[1, 0].set_ylabel("reparam")
ax[2, 0].set_ylabel(r"diff $\times 10^{15}$");
```
Now, because of the fact that the Oren-Nayar model involves terms like $\frac{1}{z}$ as well as $\text{max}()$ and $\text{min}()$ piecewise functions, there's no way to express it exactly in terms of spherical harmonics (which we need in order for the `starry` algorithm to work). So we need to approximate it with a spherical harmonic expansion. Normally we would use a Taylor expansion, but because of the piecewise nature of the function, we can't get an expansion that's good everywhere. So we'll perform a (regularized) least-squares fit. We'll fit a polynomial in the Cartesian coordinates $x$, $y$, and $z \equiv \sqrt{1 - x^2 - y^2}$ on the projected disk, which has a 1-to-1 correspondence to the spherical harmonics (via the change-of-basis matrix $\mathbf{A_1}$ (c.f. [Luger et al. 2019](https://ui.adsabs.harvard.edu/abs/2019AJ....157...64L/abstract)). Our fit will also include a polynomial in $b$ and $b_c \equiv \sqrt{1 - b^2}$.
Specifically, we are going to fit the non-Lambertian component of the model, which is the function
$$
g \equiv \cos\theta_i f
$$
where
$$
f \equiv \mathrm{max}\left(0, \cos(\phi_r - \phi_i)\right) \sin\alpha \tan\beta
$$
(see the paper for details on what each of the variables mean). We will fit a polynomial to $f$ in a frame in which the illumination source is along the $y-z$ plane, so the terminator angle $\theta = 0$ and the model should be symmetric about the $x$-axis (for different source orientations, we can always rotate the solution in the $x-y$ plane). We will then multiply $f$ by
$$
\cos\theta_i = b_c y - b z
$$
to get $g$. The end result will be the coefficients $w$ of the polynomial fit to $g$,
$$
p(x, y, b) = \sum_{i,j,k,p,q} w_{ijkpq} x^i y^j z^k b^p b_c^q \quad.
$$
for all $i, j, k, p, q$ satisfying
$$
0 \le i < 5, \, i \, \text{even}
\\
0 \le j \le 5
\\
0 \le k \le 1
\\
i + j + k \le 5
\\
1 \le p \le 5
\\
0 \le q \le 4
$$
Note that the lowest power of $b$ is one, since we require that $p(x, y, b = 0) = 0$ everywhere; the simplified Oren-Nayar model (Equation 30 in their paper) is Lambertian at half-phase.
```
def get_f_exact(x, y, z, b):
r"""
Return the expression
.. math::
f \equiv
\mathrm{max}\left(0, \cos(\phi_r - \phi_i)\right)
\sin\alpha \tan\beta
from Equation (30) in Oren & Nayar (1994) as a function
of the Cartesian coordinates on the sky-projected sphere
seen at a phase where the semi-minor axis of the terminator
is `b`.
"""
bc = np.sqrt(1 - b ** 2)
ci = bc * y - b * z
f1 = -b / z - ci
f2 = -b / ci - z
f = np.maximum(0, np.minimum(f1, f2))
return f
def get_ijk(n):
"""Get the exponents of x, y, z i the nth term of the polynomial basis."""
l = int(np.floor(np.sqrt(n)))
m = n - l * l - l
mu = l - m
nu = l + m
if mu % 2 == 0:
i = mu // 2
j = nu // 2
k = 0
else:
i = (mu - 1) // 2
j = (nu - 1) // 2
k = 1
return i, j, k
def poly_basis(x, y, z, deg):
"""Return the polynomial basis evaluated at `x`, `y`, `z`."""
N = (deg + 1) ** 2
B = np.zeros((len(x * y * z), N))
for n in range(N):
i, j, k = get_ijk(n)
B[:, n] = x ** i * y ** j * z ** k
return B
def design_matrix(x, y, z, b, deg, Nb):
"""
Return the x-y-z-b-bc Vandermonde design matrix.
NOTE: The lowest power of `b` is *ONE*, since
we need `f = 0` eveywhere when `b = 0` for
a smooth transition to Lambertian at crescent
phase.
"""
N = (deg + 1) ** 2
u = 0
X = np.zeros((len(y * z * b), N * Nb ** 2))
bc = np.sqrt(1 - b ** 2)
B = poly_basis(x, y, z, deg)
for n in range(N):
for p in range(1, Nb + 1):
for q in range(Nb):
X[:, u] = B[:, n] * b ** p * bc ** q
u += 1
return X
def index_of(i, j, k, p, q, deg, Nb):
"""
Return the index in `w` corresponding to a certain term.
NOTE: Not at all optimized!
"""
idx = 0
for n in range((deg + 1) ** 2):
i0, j0, k0 = get_ijk(n)
for p0 in range(1, Nb + 1):
for q0 in range(Nb):
if (
(i0 == i)
and (j0 == j)
and (k0 == k)
and (p0 == p)
and (q0 == q)
):
return idx
idx += 1
raise IndexError("Invalid polynomial index!")
def get_w(deg=5, Nb=4, res=100, prior_var=1e2):
"""
Return the coefficients of the 5D fit to `g`
in `x`, `y`, `z`, `b`, and `bc`.
We fit the function `f` (see above) with a
polynomial of order `deg0 = deg - 1` in `x`, `y`, and `z`
and `Nb0 = Nb - 1` in `b` and `bc`. We then multiply the
result by `cos(theta_i)` to get the function `g`,
which is just a polynomial of
order 1 in `y`, `z`, `b`, and `bc`.
The final polynomial has order `deg` and `Nb` in the
Cartesian and terminator coordinates, respectively, and is a fit
to the function
.. math::
g \equiv
\cos\theta_i
\mathrm{max}\left(0, \cos(\phi_r - \phi_i)\right)
\sin\alpha \tan\beta
from Equation (30) in Oren & Nayar (1994).
"""
# Degrees before multiplying by cos(theta_i)
deg0 = deg - 1
Nb0 = Nb - 1
# Construct a 3D grid in (x, y, b)
bgrid = np.linspace(-1, 0, res)
xygrid = np.linspace(-1, 1, res)
x, y, b = np.meshgrid(xygrid, xygrid, bgrid)
z = np.sqrt(1 - x ** 2 - y ** 2)
idx = np.isfinite(z) & (y > b * np.sqrt(1 - x ** 2))
x = x[idx]
y = y[idx]
z = z[idx]
b = b[idx]
# Compute the exact `f` function on this grid
f = get_f_exact(x, y, z, b)
# Construct the design matrix for fitting
X = design_matrix(x, y, z, b, deg=deg0, Nb=Nb0)
# "Data" inverse covariance. Make the errorbars large
# when cos(theta_i) is small, since the intensity there
# is small anyways.
cinv = np.ones_like(f)
bc = np.sqrt(1 - b ** 2)
ci = bc * y - b * z
cinv *= ci ** 2
# Prior inverse covariance. Make odd powers of x have
# *very* narrow priors centered on zero, since the
# function should be symmetric about the y axis.
# We'll explicitly zero out these coefficients below.
PInv = np.eye((deg0 + 1) ** 2 * Nb0 ** 2)
u = 0
for n in range((deg0 + 1) ** 2):
i, _, _ = get_ijk(n)
if (i % 2) != 0:
inv_var = 1e15
else:
inv_var = 1 / prior_var
for p in range(1, Nb0 + 1):
for q in range(Nb0):
PInv[u, u] = inv_var
u += 1
# Solve the L2 problem
XTCInv = X.T * cinv
w0 = np.linalg.solve(XTCInv.dot(X) + PInv, XTCInv.dot(f),)
# Zero out really tiny values.
w0[np.abs(w0) < 1e-10] = 0.0
# Now multiply by cos_thetai = bc * y - b * z.
# The powers of x, y, z, b, bc are
# i, j, k, p, q, respectively
w = np.zeros((deg + 1) ** 2 * Nb ** 2)
for n in range((deg0 + 1) ** 2):
i, j, k = get_ijk(n)
for p in range(1, Nb0 + 1):
for q in range(Nb0):
idx = index_of(i, j, k, p, q, deg0, Nb0)
w[index_of(i, j + 1, k, p, q + 1, deg, Nb)] += w0[idx]
if k == 0:
w[index_of(i, j, k + 1, p + 1, q, deg, Nb)] -= w0[idx]
else:
# transform z^2 --> 1 - x^2 - y^2
w[index_of(i, j, 0, p + 1, q, deg, Nb)] -= w0[idx]
w[index_of(i + 2, j, 0, p + 1, q, deg, Nb)] += w0[idx]
w[index_of(i, j + 2, 0, p + 1, q, deg, Nb)] += w0[idx]
return w
```
We can now test the accuracy of our polynomial fit.
```
# Get the polynomial coefficients
deg = 5
Nb = 4
w = get_w(deg=deg, Nb=Nb)
# Grid the surface
res = 300
xygrid = np.linspace(-1, 1, res)
x, y = np.meshgrid(xygrid, xygrid)
x = x.reshape(-1)
y = y.reshape(-1)
z = np.sqrt(1 - x ** 2 - y ** 2)
# Compare for several values of b
nimg = 6
fig, ax = plt.subplots(nimg, 4, figsize=(6, 8))
for axis in ax[:, :3].flatten():
axis.set(frame_on=False, xticks=[], yticks=[])
for axis in ax[:, 3].flatten():
axis.set(frame_on=False, yticks=[])
axis.tick_params(labelsize=8)
for i, b in enumerate(np.linspace(-1, 0, nimg, endpoint=False)):
# Illumination profile
bc = np.sqrt(1 - b ** 2)
ci = bc * y - b * z
# Compute the exact `f` function on this grid
f = ci * get_f_exact(x, y, z, b)
# Get our approximation
X = design_matrix(x, y, z, b, deg=deg, Nb=Nb)
fapprox = X.dot(w)
# Mask the nightside & reshape
idx = np.isfinite(z) & (y < b * np.sqrt(1 - x ** 2))
f[idx] = 0
fapprox[idx] = 0
f = f.reshape(res, res)
fapprox = fapprox.reshape(res, res)
# Plot
vmin = 0
vmax = 1
ax[i, 0].imshow(
f, origin="lower", extent=(-1, 1, -1, 1), vmin=vmin, vmax=vmax
)
ax[i, 1].imshow(
fapprox,
origin="lower",
extent=(-1, 1, -1, 1),
vmin=vmin,
vmax=vmax,
)
ax[i, 2].imshow(
f - fapprox,
origin="lower",
extent=(-1, 1, -1, 1),
vmin=-0.1,
vmax=0.1,
cmap="RdBu",
)
bins = np.linspace(-0.1, 0.1, 50)
ax[i, 3].hist((f - fapprox).flatten(), bins=bins)
ax[i, 3].set_xlim(-0.1, 0.1)
ax[i, 0].set_ylabel("b = {:.1f}".format(b), fontsize=10)
ax[0, 0].set_title("Oren-Nayar", fontsize=10)
ax[0, 1].set_title("fit", fontsize=10)
ax[0, 2].set_title("diff", fontsize=10)
ax[0, 3].set_title("diff", fontsize=10);
```
The figure above shows the non-Lambertian component of the observed intensity on the sphere at different illumination phases under the Oren-Nayar model (first column) and our polynomial approximation (second column), normalized so that dark blue = 0 and yellow = 1. The third column shows the difference between the two models, normalized to the range $[-0.1, 0.1]$, and the last column shows a histogram of the differences. In all cases, the polynomial approximation is good to within a few percent of the Oren-Nayar model. At full phase the maximum difference approaches 10% at the very edges, but the contribution of this region to the total flux is tiny.
In `starry`, users may increase the polynomial degree of the approximation by setting the `STARRY_OREN_NAYAR_DEG` macro at compile-time; this will result in better accuracy.
Finally, let's take a look at what the *actual* intensity looks like for a body under the Oren-Nayar model, using `starry`.
```
import starry
starry.config.lazy = False
```
Let's instantiate a map:
```
map = starry.Map(reflected = True)
```
We'll visualize the map with a varying illumination phase, specified via the source position parameters:
```
phase = np.linspace(0, 2 * np.pi, 100)
xs = np.cos(phase)
zs = np.sin(phase)
ys = 0
```
We now set the `roughness` parameter, which corresponds to the $\sigma$ in the Oren-Nayar model. Note that by default we specify `roughness` in degrees. Let's look at a case with zero roughness (the Lambertian case):
```
map.roughness = 0
map.show(xs=xs, ys=ys, zs=zs)
flux0 = map.flux(xs=xs, ys=ys, zs=zs)
```
Now let's look at a *very* rough surface:
```
map.roughness = 90
map.show(xs=xs, ys=ys, zs=zs)
flux90 = map.flux(xs=xs, ys=ys, zs=zs)
```
The difference is most pronounced at full phase, where it is evident that the limb is *much* brighter in the rough case. We can also look at the phase curves of each sphere:
```
plt.plot(phase / (2 * np.pi), flux0, label="Lambertian")
plt.plot(phase / (2 * np.pi), flux90, label="Rough surface")
plt.legend()
plt.xlabel("phase")
plt.ylabel("flux");
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import torch
from UnarySim.sw.kernel.div import UnaryDiv
from UnarySim.sw.stream.gen import RNG, SourceGen, BSGen
from UnarySim.sw.metric.metric import ProgressiveError
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import ticker, cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import time
import math
import numpy as np
import seaborn as sns
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
def test(mode="unipolar",
depth_abs=4,
depth_kernel=2,
depth_sync=2,
shiftreg=False,
rng="Sobol",
rng_dim=4,
bitwidth=8,
total_cnt=100,
savepdf=False):
stype = torch.float
btype = torch.float
rtype = torch.float
print("========================================================")
print(mode)
print("========================================================")
if mode is "unipolar":
# all values in unipolar are non-negative
# dividend is always non greater than divisor
# divisor is non-zero
low_bound = 0
up_bound = 2**bitwidth
elif mode is "bipolar":
# values in bipolar are arbitrarily positive or negative
# abs of dividend is always non greater than abs of divisor
# abs of divisor is non-zero
low_bound = -2**(bitwidth-1)
up_bound = 2**(bitwidth-1)
divisor_list = []
dividend_list = []
for divisor_val in range(up_bound, low_bound-1, -1):
divisor_list.append([])
dividend_list.append([])
for dividend_val in range(low_bound, up_bound+1, 1):
divisor_list[up_bound-divisor_val].append(divisor_val)
dividend_list[up_bound-divisor_val].append(dividend_val)
dividend = torch.tensor(dividend_list).type(torch.float).div(up_bound).to(device)
divisor = torch.tensor(divisor_list).type(torch.float).div(up_bound).to(device)
quotient = dividend.div(divisor)
# find the invalid postions in quotient
quotient_nan = torch.isnan(quotient)
quotient_inf = torch.isinf(quotient)
quotient_mask = quotient_nan + quotient_inf
quotient[quotient_mask] = 0
quotient = quotient.clamp(-1, 1)
result_pe_total = []
for rand_idx in range(1, total_cnt+1):
quotientPE = ProgressiveError(quotient, mode=mode).to(device)
dividendPE = ProgressiveError(dividend, mode=mode).to(device)
dividendSRC = SourceGen(dividend, bitwidth, mode=mode, rtype=rtype)().to(device)
divisorPE = ProgressiveError(divisor, mode=mode).to(device)
divisorSRC = SourceGen(divisor, bitwidth, mode=mode, rtype=rtype)().to(device)
dut_div = UnaryDiv(depth_abs=depth_abs,
depth_kernel=depth_kernel,
depth_sync=depth_sync,
shiftreg_abs=shiftreg,
mode=mode,
rng=rng,
rng_dim=rng_dim,
stype=stype,
btype=btype).to(device)
dividendRNG = RNG(bitwidth, rand_idx, rng, rtype)().to(device)
dividendBS = BSGen(dividendSRC, dividendRNG, stype).to(device)
divisorRNG = RNG(bitwidth, rand_idx+1, rng, rtype)().to(device)
divisorBS = BSGen(divisorSRC, divisorRNG, stype).to(device)
with torch.no_grad():
start_time = time.time()
for i in range(2**bitwidth):
dividend_bs = dividendBS(torch.tensor([i]))
dividendPE.Monitor(dividend_bs)
divisor_bs = divisorBS(torch.tensor([i]))
divisorPE.Monitor(divisor_bs)
quotient_bs = dut_div(dividend_bs, divisor_bs)
quotientPE.Monitor(quotient_bs)
# get the result for different rng
result_pe = quotientPE()[1].cpu().numpy()
result_pe[quotient_mask.cpu().numpy()] = np.nan
result_pe_total.append(result_pe)
# get the result for different rng
result_pe_total = np.array(result_pe_total)
#######################################################################
# check the error of all simulation
#######################################################################
result_pe_total_no_nan = result_pe_total[~np.isnan(result_pe_total)]
print("RMSE:{:1.4}".format(math.sqrt(np.mean(result_pe_total_no_nan**2))))
print("MAE: {:1.4}".format(np.mean(np.abs(result_pe_total_no_nan))))
print("bias:{:1.4}".format(np.mean(result_pe_total_no_nan)))
print("max: {:1.4}".format(np.max(result_pe_total_no_nan)))
print("min: {:1.4}".format(np.min(result_pe_total_no_nan)))
#######################################################################
# check the error according to input value
#######################################################################
avg_total = np.mean(result_pe_total, axis=0)
avg_total[quotient_mask.cpu().numpy()] = 0
fig, ax = plt.subplots()
fig.set_size_inches(5.5, 4)
axis_len = quotientPE()[1].size()[0]
divisor_y_axis = []
dividend_x_axis = []
for axis_index in range(axis_len):
divisor_y_axis.append((up_bound-axis_index/(axis_len-1)*(up_bound-low_bound))/up_bound)
dividend_x_axis.append((axis_index/(axis_len-1)*(up_bound-low_bound)+low_bound)/up_bound)
X, Y = np.meshgrid(dividend_x_axis, divisor_y_axis)
Z = avg_total
levels = [-0.09, -0.06, -0.03, 0.00, 0.03, 0.06, 0.09]
cs = plt.contourf(X, Y, Z, levels, cmap=cm.RdBu, extend="both")
cbar = fig.colorbar(cs)
# plt.tight_layout()
plt.xticks(np.arange(low_bound/up_bound, up_bound/up_bound+0.1, step=0.5))
# ax.xaxis.set_ticklabels([])
plt.yticks(np.arange(low_bound/up_bound, up_bound/up_bound+0.1, step=0.5))
# ax.yaxis.set_ticklabels([])
if savepdf is True:
plt.savefig("div-"+mode+"-bw"+str(bitwidth)+"-k"+str(depth_kernel)+"-ISCB"+".pdf",
dpi=300,
bbox_inches='tight')
plt.show()
plt.close()
test(mode="unipolar", depth_abs=3, depth_kernel=2, depth_sync=2, shiftreg=False, rng="Sobol", rng_dim=4, total_cnt=100, savepdf=False)
test(mode="bipolar", depth_abs=3, depth_kernel=2, depth_sync=2, shiftreg=False, rng="Sobol", rng_dim=4, total_cnt=100, savepdf=False)
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 회귀: 자동차 연비 예측하기
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/keras/basic_regression"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ko/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ko/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
</td>
</table>
Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도
불구하고 [공식 영문 문서](https://www.tensorflow.org/?hl=en)의 내용과 일치하지 않을 수 있습니다.
이 번역에 개선할 부분이 있다면
[tensorflow/docs](https://github.com/tensorflow/docs) 깃헙 저장소로 풀 리퀘스트를 보내주시기 바랍니다.
문서 번역이나 리뷰에 지원하려면 [이 양식](https://bit.ly/tf-translate)을
작성하거나
[docs@tensorflow.org](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs)로
메일을 보내주시기 바랍니다.
*회귀*(regression)는 가격이나 확률 같이 연속된 출력 값을 예측하는 것이 목적입니다. 이와는 달리 *분류*(classification)는 여러개의 클래스 중 하나의 클래스를 선택하는 것이 목적입니다(예를 들어, 사진에 사과 또는 오렌지가 포함되어 있을 때 어떤 과일인지 인식하는 것).
이 노트북은 [Auto MPG](https://archive.ics.uci.edu/ml/datasets/auto+mpg) 데이터셋을 사용하여 1970년대 후반과 1980년대 초반의 자동차 연비를 예측하는 모델을 만듭니다. 이 기간에 출시된 자동차 정보를 모델에 제공하겠습니다. 이 정보에는 실린더 수, 배기량, 마력(horsepower), 공차 중량 같은 속성이 포함됩니다.
이 예제는 `tf.keras` API를 사용합니다. 자세한 내용은 [케라스 가이드](https://www.tensorflow.org/guide/keras)를 참고하세요.
```
# 산점도 행렬을 그리기 위해 seaborn 패키지를 설치합니다
!pip install seaborn
from __future__ import absolute_import, division, print_function, unicode_literals
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
```
## Auto MPG 데이터셋
이 데이터셋은 [UCI 머신 러닝 저장소](https://archive.ics.uci.edu/ml/)에서 다운로드할 수 있습니다.
### 데이터 구하기
먼저 데이터셋을 다운로드합니다.
```
dataset_path = keras.utils.get_file("auto-mpg.data", "https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
```
판다스를 사용하여 데이터를 읽습니다.
```
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(dataset_path, names=column_names,
na_values = "?", comment='\t',
sep=" ", skipinitialspace=True)
dataset = raw_dataset.copy()
dataset.tail()
```
### 데이터 정제하기
이 데이터셋은 일부 데이터가 누락되어 있습니다.
```
dataset.isna().sum()
```
문제를 간단하게 만들기 위해서 누락된 행을 삭제하겠습니다.
```
dataset = dataset.dropna()
```
`"Origin"` 열은 수치형이 아니고 범주형이므로 원-핫 인코딩(one-hot encoding)으로 변환하겠습니다:
```
origin = dataset.pop('Origin')
dataset['USA'] = (origin == 1)*1.0
dataset['Europe'] = (origin == 2)*1.0
dataset['Japan'] = (origin == 3)*1.0
dataset.tail()
```
### 데이터셋을 훈련 세트와 테스트 세트로 분할하기
이제 데이터를 훈련 세트와 테스트 세트로 분할합니다.
테스트 세트는 모델을 최종적으로 평가할 때 사용합니다.
```
train_dataset = dataset.sample(frac=0.8,random_state=0)
test_dataset = dataset.drop(train_dataset.index)
```
### 데이터 조사하기
훈련 세트에서 몇 개의 열을 선택해 산점도 행렬을 만들어 살펴 보겠습니다.
```
sns.pairplot(train_dataset[["MPG", "Cylinders", "Displacement", "Weight"]], diag_kind="kde")
```
전반적인 통계도 확인해 보죠:
```
train_stats = train_dataset.describe()
train_stats.pop("MPG")
train_stats = train_stats.transpose()
train_stats
```
### 특성과 레이블 분리하기
특성에서 타깃 값 또는 "레이블"을 분리합니다. 이 레이블을 예측하기 위해 모델을 훈련시킬 것입니다.
```
train_labels = train_dataset.pop('MPG')
test_labels = test_dataset.pop('MPG')
```
### 데이터 정규화
위 `train_stats` 통계를 다시 살펴보고 각 특성의 범위가 얼마나 다른지 확인해 보죠.
특성의 스케일과 범위가 다르면 정규화(normalization)하는 것이 권장됩니다. 특성을 정규화하지 않아도 모델이 *수렴할 수 있지만*, 훈련시키기 어렵고 입력 단위에 의존적인 모델이 만들어집니다.
노트: 의도적으로 훈련 세트만 사용하여 통계치를 생성했습니다. 이 통계는 테스트 세트를 정규화할 때에도 사용됩니다. 이는 테스트 세트를 모델이 훈련에 사용했던 것과 동일한 분포로 투영하기 위해서입니다.
```
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
```
정규화된 데이터를 사용하여 모델을 훈련합니다.
주의: 여기에서 입력 데이터를 정규화하기 위해 사용한 통계치(평균과 표준편차)는 원-핫 인코딩과 마찬가지로 모델에 주입되는 모든 데이터에 적용되어야 합니다. 여기에는 테스트 세트는 물론 모델이 실전에 투입되어 얻은 라이브 데이터도 포함됩니다.
## 모델
### 모델 만들기
모델을 구성해 보죠. 여기에서는 두 개의 완전 연결(densely connected) 은닉층으로 `Sequential` 모델을 만들겠습니다. 출력 층은 하나의 연속적인 값을 반환합니다. 나중에 두 번째 모델을 만들기 쉽도록 `build_model` 함수로 모델 구성 단계를 감싸겠습니다.
```
def build_model():
model = keras.Sequential([
layers.Dense(64, activation=tf.nn.relu, input_shape=[9]),
layers.Dense(64, activation=tf.nn.relu),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=['mean_absolute_error', 'mean_squared_error'])
return model
model = build_model()
```
### 모델 확인
`.summary` 메서드를 사용해 모델에 대한 간단한 정보를 출력합니다.
```
model.summary()
```
모델을 한번 실행해 보죠. 훈련 세트에서 `10` 샘플을 하나의 배치로 만들어 `model.predict` 메서드를 호출해 보겠습니다.
```
example_batch = normed_train_data[:10]
example_result = model.predict(example_batch)
example_result
```
제대로 작동하는 것 같네요. 결괏값의 크기와 타입이 기대했던 대로입니다.
### 모델 훈련
이 모델을 1,000번의 에포크(epoch) 동안 훈련합니다. 훈련 정확도와 검증 정확도는 `history` 객체에 기록됩니다.
```
# 에포크가 끝날 때마다 점(.)을 출력해 훈련 진행 과정을 표시합니다
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
```
`history` 객체에 저장된 통계치를 사용해 모델의 훈련 과정을 시각화해 보죠.
```
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
import matplotlib.pyplot as plt
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure(figsize=(8,12))
plt.subplot(2,1,1)
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mean_absolute_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_absolute_error'],
label = 'Val Error')
plt.ylim([0,5])
plt.legend()
plt.subplot(2,1,2)
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mean_squared_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_squared_error'],
label = 'Val Error')
plt.ylim([0,20])
plt.legend()
plt.show()
plot_history(history)
```
이 그래프를 보면 수 백번 에포크를 진행한 이후에는 모델이 거의 향상되지 않는 것 같습니다. `model.fit` 메서드를 수정하여 검증 점수가 향상되지 않으면 자동으로 훈련을 멈추도록 만들어 보죠. 에포크마다 훈련 상태를 점검하기 위해 *EarlyStopping 콜백(callback)*을 사용하겠습니다. 지정된 에포크 횟수 동안 성능 향상이 없으면 자동으로 훈련이 멈춥니다.
이 콜백에 대해 더 자세한 내용은 [여기](https://www.tensorflow.org/versions/master/api_docs/python/tf/keras/callbacks/EarlyStopping)를 참고하세요.
```
model = build_model()
# patience 매개변수는 성능 향상을 체크할 에포크 횟수입니다
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
```
이 그래프를 보면 검증 세트의 평균 오차가 약 +/- 2 MPG입니다. 좋은 결과인가요? 이에 대한 평가는 여러분에게 맡기겠습니다.
모델을 훈련할 때 사용하지 않았던 **테스트 세트**에서 모델의 성능을 확인해 보죠. 이를 통해 모델이 실전에 투입되었을 때 모델의 성능을 짐작할 수 있습니다:
```
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=0)
print("테스트 세트의 평균 절대 오차: {:5.2f} MPG".format(mae))
```
## 예측
마지막으로 테스트 세트에 있는 샘플을 사용해 MPG 값을 예측해 보겠습니다:
```
test_predictions = model.predict(normed_test_data).flatten()
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
```
모델이 꽤 잘 예측한 것 같습니다. 오차의 분포를 살펴 보죠.
```
error = test_predictions - test_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
```
가우시안 분포가 아니지만 아마도 훈련 샘플의 수가 매우 작기 때문일 것입니다.
## 결론
이 노트북은 회귀 문제를 위한 기법을 소개합니다.
* 평균 제곱 오차(MSE)는 회귀 문제에서 자주 사용하는 손실 함수입니다(분류 문제에서 사용하는 손실 함수와 다릅니다).
* 비슷하게 회귀에서 사용되는 평가 지표도 분류와 다릅니다. 많이 사용하는 회귀 지표는 평균 절댓값 오차(MAE)입니다.
* 수치 입력 데이터의 특성이 여러 가지 범위를 가질 때 동일한 범위가 되도록 각 특성의 스케일을 독립적으로 조정해야 합니다.
* 훈련 데이터가 많지 않다면 과대적합을 피하기 위해 은닉층의 개수가 적은 소규모 네트워크를 선택하는 방법이 좋습니다.
* 조기 종료(Early stopping)은 과대적합을 방지하기 위한 좋은 방법입니다.
| github_jupyter |
```
# Q: Why do I used Elon Musk's OpenAI?
# A: There are three APIs we tried: 1) IBM's watson (need money for large data procesing)
# 2) text blob (did not ffind a correlation) 3) Sentiment Neuron (current, some correlation found)
# Q: What is the format of data to be matched:
# A: In weeks.
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
from scipy.stats import pearsonr
import seaborn as sns
sns.set_style('whitegrid')
plt.rcParams['figure.figsize'] = (10,5)
# 1) AMAZON
# This is a data with Sentiment neuron's analysis on 8000 headlines scrapped from financial content
freq_data = pd.read_json("Amazon_8000_newsSentiment.json")
print(freq_data)
# a function that helps catagorize dates into weeks
def weekNum(t):
if t.month == 12 and t.week == 1:
return t.week + (t.year + 1) * 100
elif t.month == 1 and t.week == 53:
return 1 + t.year*100
else:
return t.week + t.year * 100
# This block reform the data, making it separated in weeks.
# Convert the date data to timeStamp data
freq_data['WeekNum'] = pd.to_datetime(freq_data['Date'])
# Assign weeknum to each date: (e.g. the first week of 2016 -> 201601)
freq_data['WeekNum'] = freq_data['WeekNum'].apply(weekNum)
# Create a column for the sum of numeric sentiment of the week (for later analysis)
weekly_freq_data = freq_data.groupby(freq_data['WeekNum']).sum()
# Create a column for the mean of numeric sentiment of the week (for later analysis)
weekly_freq_data['freqAvg'] = freq_data.groupby(freq_data['WeekNum']).mean()
# A price data from Yahoo (formatted in weeks)
weekly_price_data = pd.read_json("AMZN2yrInWeeks.json")
# combine two sets of data into one dataframe named "dataNeeded"
weekly_price_data['WeekNum'] = weekly_price_data.index
dataNeeded = (weekly_price_data.join(weekly_freq_data))
dataNeeded.dropna(inplace=True)
# Create a column for the change of weekly's sentiment (for later analysis)
dataNeeded['frequencyChange'] = dataNeeded['freqeuency'].diff()
# Correlation testing: see "Find a correlation2 (weekly, Amazon, OpenAI).ipynb"
# Only meaningful correlation are shown below
# "last week's WeeklyChangeTotal vs. frequencyChange"
# WeeklyChangeTotal: The sum of daily stock price changes in the week
# frequency: the sum of sentiment values in the week
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:-1],dataNeeded['frequencyChange'][2:])
print("corr val: ", stat)
print('p val: ', pval)
# for alpha = 0.1
# "Having an increase of stock price in the last week"
# is positively correlated (0.31) to
# "having more satisfying headlines this week"
# (only) some visualizations
dataNeeded.index = range(len(dataNeeded)) # make 201732 the week 1, so the horizontal data is continuous
# Scatter
fig = plt.figure(1)
plt.scatter(dataNeeded.index[1:-1]+1,dataNeeded['frequencyChange'][2:], c='green', label="frequencyChange")
plt.scatter(dataNeeded.index[1:-1],dataNeeded['WeeklyChangeAvg'][1:-1], c='red', label="WeeklyChangeAvg")
plt.title("last week's WeeklyChangeTotal and frequencyChange")
plt.xlabel('# of weeks from 201732', fontsize=12)
# line graph
fig,ax = plt.subplots()
ax.plot(dataNeeded.index[1:-1]+1,dataNeeded['WeeklyChangeAvg'][1:-1])
ax.plot(dataNeeded.index[2:],dataNeeded['frequencyChange'][2:])
# Another meaningful correlation for AMAZON:
# "last week's WeeklyChangeAvg vs. frequencyChange correlated"
# WeeklyChangeAvg: The mean of daily stock price changes in the week
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:-1],dataNeeded['frequencyChange'][2:])
print("corr val: ", stat)
print('p val: ', pval)
# for alpha = 0.1
# "Having more increases than decreases of stock price in the last week"
# is positively correlated (0.31) to
# "having more satisfying headlines this week"
#2) NIKE
# This is a data with Sentiment neuron's analysis on 8000 headlines scrapped from financial content
freq_data = pd.read_json("Nike_8000_newsSentiment.json")
# a function that helps catagorize dates into weeks
def weekNum(t):
if t.month == 12 and t.week == 1:
return t.week + (t.year + 1) * 100
elif t.month == 1 and t.week == 53:
return 1 + t.year*100
else:
return t.week + t.year * 100
# This block reform the data, making it separated in weeks.
# Convert the date data to timeStamp data
freq_data['WeekNum'] = pd.to_datetime(freq_data['Date'])
# Assign weeknum to each date: (e.g. the first week of 2016 -> 201601)
freq_data['WeekNum'] = freq_data['WeekNum'].apply(weekNum)
# Create a column for the sum of numeric sentiment of the week (for later analysis)
weekly_freq_data = freq_data.groupby(freq_data['WeekNum']).sum()
# Create a column for the mean of numeric sentiment of the week (for later analysis)
weekly_freq_data['freqAvg'] = freq_data.groupby(freq_data['WeekNum']).mean()
# A price data from Yahoo (formatted in weeks)
weekly_price_data = pd.read_json("NIKE7yrsInWeeks.json")
# combine two sets of data into one dataframe named "dataNeeded"
weekly_price_data['WeekNum'] = weekly_price_data.index
dataNeeded = (weekly_price_data.join(weekly_freq_data))
dataNeeded.dropna(inplace=True)
# Create a column for the change of weekly's sentiment (for later analysis)
dataNeeded['frequencyChange'] = dataNeeded['freqeuency'].diff()
# Most correlated:
# "WeeklyChangeTotal vs. frequency correlated (same week)"
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'],dataNeeded['freqeuency'])
print("corr val: ", stat)
print('p val: ', pval)
# for alpha = 0.01
# "Having an increase of stock price "
# is positively correlated (0.16) to
# "having satisfying headlines this week"
# (only) some visualizations
dataNeeded.index = range(len(dataNeeded)) # make 201108 the week 1, so the horizontal data is continuous
# Scatter
fig = plt.figure(1)
plt.scatter(dataNeeded.index,dataNeeded['freqeuency'], c='green', label="freqeuency")
plt.scatter(dataNeeded.index,dataNeeded['WeeklyChangeTotal'], c='red', label="WeeklyChangeTotal")
plt.title("WeeklyChangeTotal and frequency")
plt.xlabel('# of weeks from 201108', fontsize=12)
# line graph
fig,ax = plt.subplots()
ax.plot(dataNeeded.index,dataNeeded['freqeuency'])
ax.plot(dataNeeded.index,dataNeeded['WeeklyChangeTotal'])
# Other meaningful correlations for Nike:
# 2. "WeeklyChangeAvg vs. freqeuency "
# WeeklyChangeAvg: The mean of daily stock price changes in the week
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'],dataNeeded['freqeuency'])
print("corr val: ", stat)
print('p val: ', pval)
# 3. "WeeklyChangeAvg vs. frequencyChange "
stat, pval = pearsonr(dataNeeded['WeeklyChangeAvg'][1:],dataNeeded['frequencyChange'][1:])
print("corr val: ", stat)
print('p val: ', pval)
# 4. "WeeklyChangeTotal vs. frequencyChange "
stat, pval = pearsonr(dataNeeded['WeeklyChangeTotal'][1:],dataNeeded['frequencyChange'][1:])
print("corr val: ", stat)
print('p val: ', pval)
```
| github_jupyter |
## Questionário 73 (Q73)
Orientações:
- Registre suas respostas no questionário de mesmo nome no SIGAA.
- O tempo de registro das respostas no questionário será de 10 minutos. Portanto, resolva primeiro as questões e depois registre-as.
- Haverá apenas 1 (uma) tentativa de resposta.
- Submeta seu arquivo-fonte (utilizado para resolver as questões) em formato _.ipynb_ pelo SIGAA anexando-o à Tarefa denominada "Envio de arquivo" correspondente ao questionário.
*Nota:* o arquivo-fonte será utilizado apenas como prova de execução da tarefa. Nenhuma avaliação será feita quanto ao estilo de programação.
<hr>
Para responder às questões, leia o texto introdutório a seguir.
>Diversos países firmam acordos bilaterais com o intuito de fortalecer interesses mútuos. Uma rede multinacional da qual o Brasil faz parte começou a ser modelada por cientistas de dados a partir de um grafo não dirigido em que os _nós_ do grafo representam os países, renomeados segundo o código Alpha-3 do padrão [IBAN](https://www.iban.com/country-codes), e as _arestas_ representam a existência de um acordo bilateral.
> A figura abaixo mostra, por exemplo, um subgrafo dessa rede formado por Áustria (AUT), Bélgica (BEL), Brasil (BRA), Emirados Árabes Unidos (ARE) e Estados Unidos (USA).
```{figure} ../figs/q/q73.png
---
width: 660px
name: rede
---
Exemplo de rede de países que mantêm acordos bilaterais.
```
> O arquivo `paises-acordo-bilateral.txt` contém, implicitamente, a lista de conexões que formam o grafo da rede inteira, as quais são determinadas por pares do tipo `x,y`, onde `x` e `y` são nomes de países não padronizados. Por exemplo, o par `China,Norway` indica que há um acordo bilateral entre China e Noruega.
>*Obs.:* acesse o arquivo [aqui](https://github.com/gcpeixoto/ICD/tree/main/database/paises-acordo-bilateral.txt).
**Questão 1.** Faça a raspagem da tabela de códigos de países disponíveis na página [IBAN](https://www.iban.com/country-codes) para recuperar os códigos Alpha-3 para cada país contido na lista de arestas e crie um segundo arquivo chamado `paises-acordo-bilateral-IBAN.txt`. Use o módulo `networkx` e a função `read_edgelist` para construir o grafo da rede multinacional. Em seguida, assinale a alternativa correta para a tupla (número de nós, número de arestas) que você encontrou. Sugestão: use as funções `get_table_head` e `get_table_body` criadas no capítulo do livro de ICD sobre _Raspagem de dados_.
A. (14, 28)
B. (16, 30)
C. (12, 36)
D. (14, 38)
## GABARITO
Alternativa **D**.
## Geração de arquivo de acordo bilateral
```
import numpy as np
np.random.seed(3)
countries = ('Argentina','Austria','Belgium','Brazil','China',
'United Arab Emirates (the)',
'United States of America (the)','Germany',
'India','Israel','Netherlands (the)',
'Norway','Russian Federation (the)','South Africa')
adj = np.random.randint(0,2,(len(countries),len(countries)))
adj[np.diag_indices_from(adj)] = 0
adj = np.tril(adj)
adj = np.tril(adj) + np.tril(adj).T
f = open('../database/paises-acordo-bilateral.txt','w')
for i in range(len(countries)):
for j in range(i+1,len(countries)):
if adj[i,j] == 1:
s = countries[i] + ',' + countries[j] + '\n'
f.write(s)
f.close()
```
## Raspagem do IBAN
```
from urllib.request import urlopen
from bs4 import BeautifulSoup
import pandas as pd
import networkx as nx
html = urlopen('https://www.iban.com/country-codes')
bs = BeautifulSoup(html.read(),'html.parser')
# extrai cabeçalho
def get_table_head(t):
'''Lê objeto tabela e extrai header para lista'''
res = []
thead = t.find('thead')
th = thead.find_all('th')
for f in th:
res.append(f.getText().strip())
return res
t_header = get_table_head(bs.body)
# extrai linhas
def get_table_body(t):
res = []
tbody = t.find('tbody')
tr = tbody.find_all('tr')
for row in tr:
this_row = []
row_fields = row.find_all('td')
for f in row_fields:
this_row.append(f.getText().strip())
res.append(this_row)
return res
r = get_table_body(bs.body)
# DataFrame
iban = pd.DataFrame(r,columns=t_header).drop_duplicates().drop(columns=['Alpha-2 code','Numeric']).reset_index(drop=True)
```
## Escreve edgelist
```
f = open('../database/paises-acordo-bilateral.txt','r')
g = open('../database/paises-acordo-bilateral-IBAN.txt','w')
for i in f.readlines():
p1,p2 = i.strip().split(',')
p1 = iban[iban['Country'] == p1]['Alpha-3 code'].values
p2 = iban[iban['Country'] == p2]['Alpha-3 code'].values
p1,p2 = p1[0],p2[0]
s = p1 + ',' + p2 + '\n'
g.write(s)
f.close()
g.close()
G = nx.read_edgelist('../database/paises-acordo-bilateral-IBAN.txt',delimiter=',')
Gsub = nx.subgraph(G,['AUT','ARE','USA','BRA','BEL',''])
nx.draw_networkx(Gsub,with_labels=True)
G.number_of_nodes(),G.number_of_edges()
```
**Questão 2.** A _centralidade de grau_ `deg`, calculada para cada nó do grafo completo pelo módulo `networkx`, pode ser interpretada, para este estudo de caso, como uma medida relativa da pré-disposição de um país para se abrir à globalização. Neste sentido, calcule `deg` e assinale a opção cujo país é o mais **fechado** ao fenômeno da globalização.
A. CHN
B. BRA
C. ARG
D. NLD
## GABARITO
Alternativa **D**.
```
# minima deg é NLD
deg = nx.degree_centrality(G)
deg = {k: v for k, v in sorted(deg.items(), key=lambda item: item[1],reverse=True)}
deg
```
**Questão 3.** Semelhantemente à interpretação da questão anterior, a _centralidade de intermediação_ `bet` fornece uma medida relativa de quão boa é a confiança e respeitabilidade diplomática de um país para a concretização de acordos. Calcule `bet` e assinale a opção cujo país é o mais respeitado para intermediar acordos.
A. AUT
B. ZAF
C. DEU
D. ISR
## GABARITO
Alternativa **C**.
```
# maxima bet é DEU
bet = nx.betweenness_centrality(G)
bet = {k: v for k, v in sorted(bet.items(), key=lambda item: item[1],reverse=True)}
bet
!rm ../database/paises-acordo-bilateral-IBAN.txt
```
| github_jupyter |
### Import packages
```
library(data.table)
library(Matrix)
library(proxy)
library(Rtsne)
library(densityClust)
library(data.table)
library(irlba)
library(umap)
library(ggplot2)
library(RColorBrewer)
```
### Load Data
```
load('../../run_methods/Cusanovich2018/Cusanovich2018_buenrostro2018.RData')
```
### Following tutorial at https://shendurelab.github.io/fly-atac/docs/#usecase2
```
#desired number of clusters
nClusters = length(levels(as.factor(metadata$label)))
nClusters
#To identify clusters of cells, we use the density peak algorithm.
tsnedist = dist(tsnetfidf$Y)
set.seed(0)
dclust = densityClust(tsnedist,gaussian=T)
dclust = findClusters(dclust, rho = 20, delta = 9)
#number of clusters produced
nClusters_produced = length(levels(as.factor(dclust$clusters)))
nClusters_produced
levels(as.factor(dclust$clusters))
#plot from tutorial
#The density peak algorithm requires you to set two parameters - “delta” and “rho”. For each data point, the algorithm calculates a local density of other points within some set distance and the minimum distance to the next point that has a higher local density. On the basis of these two values, you can choose a set of points that are outliers both in local density and the distance to another point with a higher density, which become cluster “peaks”. Below, we show you the distribution of these two values in our data set and where we decided to draw the cutoff. You can read more about this algorithm here.
options(repr.plot.width=6, repr.plot.height=6)
plot(dclust$rho,dclust$delta,pch=20,cex=0.6)
points(dclust$rho[dclust$peaks],dclust$delta[dclust$peaks],col="red",pch=20,cex=0.8)
text(dclust$rho[dclust$peaks]-2,dclust$delta[dclust$peaks]+1.5,labels=dclust$clusters[dclust$peaks])
abline(v=20)
abline(h=9)
#plot from tutorial
tsnecols = c("#E31A1C","#FFD700","#771122","#777711","#1F78B4","#68228B","#AAAA44",
"#60CC52","#771155","#DDDD77","#774411","#AA7744","#AA4455","#117744",
"#000080","#44AA77","#AA4488","#DDAA77")
plot(tsnetfidf$Y,pch=20,col=tsnecols[as.factor(dclust$clusters)],main="Density Peak Clusters",cex=0.25)
text(tsnetfidf$Y[dclust$peaks,1],tsnetfidf$Y[dclust$peaks,2],labels=dclust$clusters[dclust$peaks],cex=2.5)
str(tsnetfidf)
plot.tsne(tsnetfidf$Y,as.factor(dclust$clusters))
plot.tsne(tsnetfidf$Y,as.factor(metadata[,'label']))
## results from pca and tsne reduction types are the same...
#result = data.frame("cusanovich2018.tsne" = seurat_obj.tsne_clustering@ident)
#result2 = data.frame("cusanovich2018.pca" = seurat_obj@ident)
#m = merge(result,result2,by='row.names')
df_out = data.frame("cusanovich2018" = dclust$clusters)
rownames(df_out) = rownames(metadata)
write.table(df_out,file="clusteringSolution.tsv", quote=FALSE, sep='\t', col.names = NA)
head(df_out)
```
| github_jupyter |
# Pretty Plotting (literally)
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import hotstepper as hs
from hotstepper import Bases, Basis, Step, Steps
import hotstepper.samples as samples
```
## Pretty Plot
Somtimes, we just want a pretty plot, inparticular, with step data, the formal plotting method should indicate the nature of the step end points, for example, filled markers indicate set membership, open markers we are plotting right up to, but technically not including that last point.
```
st1 = Step(5,15,2)
st2 = Step(end=6)
st3 = Step(4,weight=3)
sts = Steps().add([st1,st2,st3])
ax = sts.plot(method='pretty')
sts.plot(ax=ax,method='smooth',smooth_factor=0.3)
```
## Steps are funny
One of the things makes step functions different to regular functions is that they can be evaluated differently depending on the rule used to determine their membership to the range set. That all sounds fancy and stuff, but what it really means is where do we decide to make the step value change, before the step key, on the step key or after.
For those needing something abit more mathematically concrete, please see the full explanation of step functions and their properties in the [HotStepper Documentation](https://hotstepper.readthedocs.io/step_functions.html).
The short answer is, the step function will look different depending on the rule we use to evaluate where step changes occur.
HotStepper by default uses the where='post' evaluation location, as this conforms with the signal processing definition used for the [Heaviside Step Function](https://en.wikipedia.org/wiki/Heaviside_step_function) which is the default basis.
An easy way to see the differences and how they look, we can pass different where= parameter values to the plot method. We use the pretty plot method for the default (where='post') to clearly see the step change locations, changing the where parameter to either 'pre' or 'mid', we see how the changes occur relative to this fixed locations.
```
ax = sts.plot(method='pretty')
sts.plot(ax=ax,where='mid',linestyle='--',label='evaluation at mid')
sts.plot(ax=ax,where='pre',linestyle='-.',label='evaluation at pre')
ax.legend();
```
## Smooth Plotting Multiple Basis (kernel)
Changing the basis to use for smooth plotting is pretty simple, just pick your nwe Base from the pre-baked selection, wrap it in a Basis, which allows you to specify a smoothing parameter to be used with that basis and give it to the plot method.
A few items to point out here, as you can take your next **steps** in multiple ways with HotStepper.
- Can smooth plot by calling smooth_plot() on the steps
- Can smooth plot by using method='smooth' in the call to plot() on the steps
```
ax = sts.plot(method='function',label='funcion',linewidth=3,linestyle='-.')
sts.plot(ax=ax,method='smooth',label='smooth (default)',smooth_factor=0.5)
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.arctan,0.5),label='smooth (arctan)')
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.exponential,0.5),label='smooth (exponential)')
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.sigmoid),smooth_factor=2,label='smooth (sigmoid)')
sts.plot(ax=ax,method='smooth',smooth_factor=3,label='smooth (default - strong)')
ax.legend();
```
Hostepper has a couple of basis functions that don't do smoothing in the same way the other bases do. The short answer is that these basis functions don't have a limit representation of the Heaviside function, unlike the other bases. For a full explanation of what this means, please see the [Basis and Bases](https://hotstepper.readthedocs.io/basis.html) documentation. So the normal and sinc basis represent a class of basis functions ([kernel functions](https://en.wikipedia.org/wiki/Kernel_smoother)) that are centered around zero and provide a means to smooth the step **changes** instead of the step function **values**. This can be very helpful when analysing the nature of the transitions that occur within the steps data.
```
ax = sts.plot(method='smooth', smooth_basis=Basis(Bases.normal),smooth_factor=0.5,label='smooth (normal)',linestyle='--')
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.sinc),smooth_factor=0.5,label='smooth (sinc)',linestyle='-.')
x = np.arange(-1,sts.last()*1.5,0.01)
for s in [st1,st2,st3]:
ax.step(x,s(x),linewidth=3)
ax.legend();
```
If you create a new Basis and specify a smoothing parameter for that Basis, then you specify a smooth_factor in the call to smooth plot or plot with method='smooth', the parameter in the Basis object will be overridden by the provided smooth_factor.
```
ax = sts.plot(label='function',method='function')
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.sigmoid,2),label='smooth (sigmoid)',linestyle='--')
sts.plot(ax=ax,method='smooth', smooth_basis=Basis(Bases.sigmoid,2),smooth_factor=0.5,label='smooth (sigmoid - override)',linestyle='-.')
ax.legend();
```
| github_jupyter |
# Scheduled Integration of ClinGen Gene-Disease Validity Data into WikiData
ClinGen (Clinical Genome Resource) develops curated data of genetic associations <br>
CC0 https://clinicalgenome.org/docs/terms-of-use/
This scheduled bot operates through WDI to integrate ClinGen Gene-Disease Validity Data <br>
https://search.clinicalgenome.org/kb/gene-validity/ <br>
https://github.com/SuLab/GeneWikiCentral/issues/116 <br>
http://jenkins.sulab.org/ <br>
Python script contributions, in order: Sabah Ul-Hasan, Andra Waagmeester, Andrew Su, Ginger Tsueng
## Checks
- Login automatically aligns with given environment
- For loop checks for both HGNC Qid and MONDO Qid per each row (ie if HGNC absent or multiple, then checks MONDO)
- For loop works on multiple Qid option, tested using A2ML1 as pseudo example
- For loop puts correct Qid for either HGNC or MONDO, if available </br>
- For loop only writes 'complete' in output if written to Wikidata
## Issues
- create_reference() and update_retrieved_if_new_multiple_refs functions adds and/or updates ref to existing HGNC or MONDO value in genetic association statement within 180 days (doesn't overwrite URLs from non-ClinGen sources)
- Updated, not updated, skipped - not definitive or mapping error..
- Maybe get of Definitive column, but keep Gene and Disease QID
```
# Relevant Modules and Libraries
## Installations by shell
!pip install --upgrade pip # Installs pip, ensures it's up-to-date
!pip3 install tqdm # Visualizes installation progress (progress bar)
!pip3 install wikidataintegrator # For wikidata
## Installations by python
from wikidataintegrator import wdi_core, wdi_login # Core and login from wikidataintegrator module
from wikidataintegrator.ref_handlers import update_retrieved_if_new_multiple_refs # For retrieving references
from datetime import datetime # For identifying the current date and time
import copy # Copies references needed in the .csv for uploading to wikidata
import time # For keeping track of total for loop run time
import os # OS package to ensure interaction between the modules (ie WDI) and current OS being used
import pandas as pd # Pandas for data organization, then abbreviated to pd
import numpy as np # Another general purpose package
from termcolor import colored # Imports colored package from termcolor
# import ssl
# ssl._create_default_https_context = ssl._create_unverified_context
# Login for running WDI
print("Logging in...")
## **remove lines when scheduling to Jenkins** Enter your own username and password
os.environ["WDUSER"] = "username" # Uses os package to call and set the environment for wikidata username
os.environ["WDPASS"] = "password"
## Conditional that outputs error command if not in the local python environment
if "WDUSER" in os.environ and "WDPASS" in os.environ:
WDUSER = os.environ['WDUSER']
WDPASS = os.environ['WDPASS']
else:
raise ValueError("WDUSER and WDPASS must be specified in local.py or as environment variables")
## Sets attributed username and password as 'login'
login = wdi_login.WDLogin(WDUSER, WDPASS)
# ClinGen gene-disease validity data
## Read as csv
df = pd.read_csv('https://search.clinicalgenome.org/kb/gene-validity.csv', skiprows=6, header=None)
## Label column headings
df.columns = ['Gene', 'HGNC Gene ID', 'Disease', 'MONDO Disease ID','SOP','Classification','Report Reference URL','Report Date']
## Create time stamp of when downloaded (error if isoformat() used)
timeStringNow = datetime.now().strftime("+%Y-%m-%dT00:00:00Z")
## Create empty columns for output file (ignore warnings)
df['Status'] = "pending" # "Status" column with 'pending' for all cells: 'error', 'complete', 'skipped' (meaning previously logged within 180 days)
df['Definitive'] = "" # Empty cell to be replaced with 'yes' or 'no' string
df['Gene QID'] = "" # To be replaced with 'absent' or 'multiple'
df['Disease QID'] = "" # To be replaced with 'absent' or 'multiple'
df.head(6)
# Create a function for adding references to then be iterated in the loop "create_reference()"
def create_reference(): # Indicates a parameter included before running rest of function (otherwise may not recognize)
refStatedIn = wdi_core.WDItemID(value="Q64403342", prop_nr="P248", is_reference=True) # ClinGen Qid = Q64403342, 'stated in' Pid = P248
timeStringNow = datetime.now().strftime("+%Y-%m-%dT00:00:00Z") # Create time stamp of when downloaded (error if isoformat() used)
refRetrieved = wdi_core.WDTime(timeStringNow, prop_nr="P813", is_reference=True) # Calls on previous 'timeStringNow' string, 'retrieved' Pid = P813
refURL = wdi_core.WDUrl((df.loc[index, 'Report Reference URL']), prop_nr="P854", is_reference=True) # 'reference URL' Pid = P854
return [refStatedIn, refRetrieved, refURL]
# For loop that executes the following through each row of the dataframe
start_time = time.time() # Keep track of how long it takes loop to run
for index, row in df.iterrows(): # Index is a row number, row is all variables and values for that row
# Identify the string in the Gene or Disease column for a given row
HGNC = df.loc[index, 'HGNC Gene ID'].replace("HGNC:", "") # .replace() changes HGNC: to space for SparQL query
MONDO = df.loc[index, 'MONDO Disease ID'].replace("_", ":")
# SparQL query to search for Gene or Disease in Wikidata based on HGNC ID (P354) or MonDO ID (P5270)
sparqlQuery_HGNC = "SELECT * WHERE {?gene wdt:P354 \""+HGNC+"\"}"
result_HGNC = wdi_core.WDItemEngine.execute_sparql_query(sparqlQuery_HGNC) # Resultant query
sparqlQuery_MONDO = "SELECT * WHERE {?disease wdt:P5270 \""+MONDO+"\"}"
result_MONDO = wdi_core.WDItemEngine.execute_sparql_query(sparqlQuery_MONDO)
# Assign resultant length of dictionary for either Gene or Disease (number of Qid)
HGNC_qlength = len(result_HGNC["results"]["bindings"])
MONDO_qlength = len(result_MONDO["results"]["bindings"])
# Conditional utilizing length value for output table, accounts for absent/present combos
if HGNC_qlength == 1:
HGNC_qid = result_HGNC["results"]["bindings"][0]["gene"]["value"].replace("http://www.wikidata.org/entity/", "")
df.at[index, 'Gene QID'] = HGNC_qid # Input HGNC Qid in 'Gene QID' cell
if HGNC_qlength < 1: # If no Qid
df.at[index, 'Status'] = "error"
df.at[index, 'Gene QID'] = "absent"
if HGNC_qlength > 1: # If multiple Qid
df.at[index, 'Status'] = "error"
df.at[index, 'Gene QID'] = "multiple"
if MONDO_qlength == 1:
MONDO_qid = result_MONDO["results"]["bindings"][0]["disease"]["value"].replace("http://www.wikidata.org/entity/", "")
df.at[index, 'Disease QID'] = MONDO_qid
if MONDO_qlength < 1:
df.at[index, 'Status'] = "error"
df.at[index, 'Disease QID'] = "absent"
if MONDO_qlength > 1:
df.at[index, 'Status'] = "error"
df.at[index, 'Disease QID'] = "multiple"
# Conditional inputs error such that only rows are written for where Classification = 'Definitive'
if row['Classification']!='Definitive': # If the string is NOT 'Definitive' for the Classification column
df.at[index, 'Status'] = "error" # Then input "error" in the Status column
df.at[index, 'Definitive'] = "no" # And'no' for Definitive column
continue # Skips rest and goes to next row
else: # Otherwise
df.at[index, 'Definitive'] = "yes" # Input 'yes' for Definitive column, go to next step
# Conditional continues to write into WikiData only if 1 Qid for each + Definitive classification
if HGNC_qlength == 1 & MONDO_qlength == 1:
# Call upon create_reference() function created
reference = create_reference()
# Add disease value to gene item page, and gene value to disease item page (symmetry)
# Creates 'gene assocation' statement (P2293) whether or not it's already there, and includes the references
statement_HGNC = [wdi_core.WDItemID(value=MONDO_qid, prop_nr="P2293", references=[copy.deepcopy(reference)])]
wikidata_HGNCitem = wdi_core.WDItemEngine(wd_item_id=HGNC_qid,
data=statement_HGNC,
global_ref_mode='CUSTOM', # parameter that looks within 180 days
ref_handler=update_retrieved_if_new_multiple_refs,
append_value=["P2293"])
wikidata_HGNCitem.get_wd_json_representation() # Gives json structure that submitted to API, helpful for debugging
wikidata_HGNCitem.write(login)
statement_MONDO = [wdi_core.WDItemID(value=HGNC_qid, prop_nr="P2293", references=[copy.deepcopy(reference)])]
wikidata_MONDOitem = wdi_core.WDItemEngine(wd_item_id=MONDO_qid,
data=statement_MONDO,
global_ref_mode='CUSTOM',
ref_handler=update_retrieved_if_new_multiple_refs,
append_value=["P2293"])
wikidata_MONDOitem.get_wd_json_representation()
wikidata_MONDOitem.write(login)
HGNC_name = df.loc[index, 'Gene'] # To output gene name > HGNC ID
MONDO_name = df.loc[index, 'Disease']
df.at[index, 'Status'] = "complete"
end_time = time.time() # Captures when loop run ends
print("The total time of this loop is:", end_time - start_time, "seconds, or", (end_time - start_time)/60, "minutes")
# Write output to a .csv file
now = datetime.now() # Retrieves current time and saves it as 'now'
# Includes hour:minute:second_dd-mm-yyyy time stamp (https://en.wikipedia.org/wiki/ISO_8601)
df.to_csv("ClinGenBot_Status-Output_" + now.isoformat() + ".csv") # isoformat
```
| github_jupyter |
```
'''
Multiplexed Convolutional Neural Network Implementation on MNIST
Vin Ay
Input : [Image, Control Signal] -> Image = B/W or W/B Digits, Control Signal = 0 (for B/W), 1 (for W/B)
Output : [Digit Index]
'''
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Softmax
from keras.layers import Conv2D, MaxPooling2D, Input
from keras import backend as K
from keras.models import Model
from multiplexer import Multiplexer
import numpy as np
import random
batch_size = 128
num_classes = 10
epochs = 10
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(x_train.shape[0], 28, 28,1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
# Create Invert Color dataset for other branch of the network
x_train_inv = 1 - x_train
x_test_inv = 1 - x_test
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# Build data batch generator for Training and Testing
def getTrainBatch(batch_size):
while True:
batch_x = []
batch_c = []
batch_y = []
for i in range(batch_size):
ch = random.randint(0,1)
ch2 = random.randint(0, 60000-1)
if ch == 0:
batch_x.append(x_train[ch2])
batch_c.append(ch)
batch_y.append(y_train[ch2])
else:
batch_x.append(x_train_inv[ch2])
batch_c.append(ch)
batch_y.append(y_train[ch2])
yield([np.array(batch_x), np.array(batch_c)], np.array(batch_y))
def getTestBatch(batch_size):
while True:
batch_x = []
batch_c = []
batch_y = []
for i in range(batch_size):
ch = random.randint(0,1)
ch2 = random.randint(0, 10000-1)
if ch == 0:
batch_x.append(x_test[ch2])
batch_c.append(ch)
batch_y.append(y_test[ch2])
else:
batch_x.append(x_test_inv[ch2])
batch_c.append(ch)
batch_y.append(y_test[ch2])
yield([np.array(batch_x), np.array(batch_c)], np.array(batch_y))
# Build Multiplexed Model for training both types of data separately
input_shape = (28, 28,1)
input_im = Input(shape = input_shape)
control = Input(shape = (1,), dtype = 'int32')
x = Conv2D(32, (3,3), activation = 'relu')(input_im)
x = MaxPooling2D(pool_size = (2,2))(x)
x = Conv2D(64, (3,3), activation = 'relu')(x)
x = MaxPooling2D(pool_size = (2,2))(x)
x = Dropout(0.25)(x)
x = Flatten()(x)
x = Dense(128, activation = 'relu')(x)
x = Dropout(0.5)(x)
x = Dense(2*num_classes)(x)
output = Multiplexer(num_classes, 2)([x, control])
output = Softmax()(output)
model = Model([input_im, control], output)
# Save model image on disk
from keras.utils import plot_model
plot_model(model, to_file='model.png')
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
# Start training the model
model.fit_generator(getTrainBatch(batch_size),
epochs=epochs,
verbose=1,
steps_per_epoch=len(x_train)/batch_size,
validation_data=getTestBatch(batch_size),
validation_steps = len(x_test)/batch_size
)
# Save model on disk
model.save("mnist_10ep.h5")
'''
Plotting images to visualize output of the network
'''
import matplotlib.pyplot as plt
from PIL import Image
import tensorflow as tf
import time
for i in range(10):
plt.figure(figsize=(3, 3))
plt.subplot(1, 3, 1)
im = tf.keras.preprocessing.image.array_to_img( x_test[i], data_format=None, scale=True,dtype=None)
plt.imshow(im, cmap="gray")
plt.subplot(1, 3, 3)
im2 = tf.keras.preprocessing.image.array_to_img( x_test_inv[i], data_format=None, scale=True,dtype=None)
plt.imshow(im2, cmap="gray")
plt.show()
black_res = model.predict([x_test[i].reshape(1,28,28,1),np.array([0])])
white_res = model.predict([x_test_inv[i].reshape(1,28,28,1),np.array([1])])
b_ind = np.argmax(black_res)
w_ind = np.argmax(white_res)
print(b_ind,":",black_res[0][b_ind]," ", w_ind,":",white_res[0][w_ind])
```
| github_jupyter |
```
# Import dependencies.
import pandas as pd
from collections import Counter
```
# Runs
### Import 1905 and 1969 csvs and isolate the incidence of average Run values for World Series winners and non-World Series Winners.
```
# Open up the 1905.csv and inspect.
df2 = pd.read_csv("../clean_data/1905.csv")
df2 = df2.drop("Unnamed: 0", axis=1)
df2
# Open up the 1969.csv and inspect.
df3 = pd.read_csv("../clean_data/1969.csv")
df3 = df3.drop("Unnamed: 0", axis=1)
df3
```
### Gather the winners of the World Series.
# Collect data in list for ws_win Runs.
# Collect data in list for nonws_win Runs.
# Collect data for all Runss.
# Turn data into dataframes.
```
# Make a list of World Series winners and non-World Series winners Ruhs for 1905 onwards.
ws_win_Runs_1905 = []
ws_nowin_Runs_1905 = []
for entry in range(len(df2)):
if df2["WSWin"][entry] == "Y":
Runs = int(df2["R"][entry])
ws_win_Runs_1905.append(Runs)
elif df2["WSWin"][entry] == "N":
Runs = str(round((df2["R"][entry]), 1))
ws_nowin_Runs_1905.append(Runs)
# Make a list of World Series and non-World Series winners Runs for 1969 onwards.
ws_win_Runs_1969 = []
ws_nowin_Runs_1969 = []
for entry in range(len(df3)):
if df3["WSWin"][entry] == "Y":
Runs = int(df3["R"][entry])
ws_win_Runs_1969.append(Runs)
elif df3["WSWin"][entry] == "N":
Runs = int(df3["R"][entry])
ws_nowin_Runs_1969.append(Runs)
# Make a list of all Runs for 1905 onwards.
ws_Runs_tot_1905 = []
for entry in range(len(df2)):
Runs = int(df2["R"][entry])
ws_Runs_tot_1905.append(Runs)
# Make a list of all Runs for 1969 onwards.
ws_Runs_tot_1969 = []
for entry in range(len(df3)):
Runs = int(df3["R"][entry])
ws_Runs_tot_1969.append(Runs)
# Find percent Runs frequency of World Series winners 1905 onwards. Set up dataframe.
ws_win_counter_1905 = Counter(ws_win_Runs_1905)
wswin_1905 = pd.Series(ws_win_counter_1905, index = ws_win_counter_1905.keys())
wswin_1905 = wswin_1905.reset_index()
wswin_1905 = wswin_1905.rename(columns = {"index": "Runs", 0: "Count"})
# Find percent Runs frequency of non-World Series winners 1905 onwards. Set up dataframe.
ws_nowin_counter_1905 = Counter(ws_nowin_Runs_1905)
wsnowin_1905 = pd.Series(ws_nowin_counter_1905, index = ws_nowin_counter_1905.keys())
wsnowin_1905 = wsnowin_1905.reset_index()
wsnowin_1905 = wsnowin_1905.rename(columns = {"index": "Runs", 0: "Count"})
# Find percent Runs frequency of World Series winners 1969 onwards. Set up dataframe.
ws_win_counter_1969 = Counter(ws_win_Runs_1969)
wswin_1969 = pd.Series(ws_win_counter_1969, index = ws_win_counter_1969.keys())
wswin_1969 = wswin_1969.reset_index()
wswin_1969 = wswin_1969.rename(columns = {"index": "Runs", 0: "Count"})
# Find percent Runs frequency of non-World Series winners 1969 onwards. Set up dataframe.
ws_nowin_counter_1969 = Counter(ws_nowin_Runs_1969)
wsnowin_1969 = pd.Series(ws_nowin_counter_1969, index = ws_nowin_counter_1969.keys())
wsnowin_1969 = wsnowin_1969.reset_index()
wsnowin_1969 = wsnowin_1969.rename(columns = {"index": "Runs", 0: "Count"})
# Find Runs frequency of all teams 1905 onwards. Set up dataframe.
wins_Runs_counter_1905 = Counter(ws_Runs_tot_1905)
tot_Runs_1905 = pd.Series(wins_Runs_counter_1905, index = wins_Runs_counter_1905.keys())
tot_Runs_1905 = tot_Runs_1905.reset_index()
tot_Runs_1905 = tot_Runs_1905.rename(columns = {"index": "Runs", 0: "Count"})
# Find Runs frequency of all teams 1969 onwards. Set up dataframe.
wins_Runs_counter_1969 = Counter(ws_Runs_tot_1969)
tot_Runs_1969 = pd.Series(wins_Runs_counter_1969, index = wins_Runs_counter_1969.keys())
tot_Runs_1969 = tot_Runs_1969.reset_index()
tot_Runs_1969 = tot_Runs_1969.rename(columns = {"index": "Runs", 0: "Count"})
```
### Create another column for Frequency.
# Turn the Count number into a frequency of the sample population.
# Add the information to the dataframe.
```
# Find frequency of World Series winners of 1905 onwards.
total1 = wswin_1905["Count"].sum()
frequency1 = [x/total1 for x in wswin_1905["Count"]]
wswin_1905["Frequency"] = frequency1
wswin_1905
# Find frequency of non-World Series winners of 1905 onwards.
total2 = wsnowin_1905["Count"].sum()
frequency2 = [x/total2 for x in wsnowin_1905["Count"]]
wsnowin_1905["Frequency"] = frequency2
wsnowin_1905
# Find frequency of World Series winners of 1969 onwards.
total1 = wswin_1969["Count"].sum()
frequency1 = [x/total1 for x in wswin_1969["Count"]]
wswin_1969["Frequency"] = frequency1
wswin_1969
# Find frequency of non-World Series winners of 1969 onwards.
total2 = wsnowin_1969["Count"].sum()
frequency2 = [x/total2 for x in wsnowin_1969["Count"]]
wsnowin_1969["Frequency"] = frequency2
wsnowin_1969
# Find frequency of all from 1905 onwards.
total5 = tot_Runs_1905["Count"].sum()
frequency5 = [x/total5 for x in tot_Runs_1905["Count"]]
tot_Runs_1905["Frequency"] = frequency5
tot_Runs_1905
# Find frequency of all from 1969 onwards.
total5 = tot_Runs_1969["Count"].sum()
frequency5 = [x/total5 for x in tot_Runs_1969["Count"]]
tot_Runs_1969["Frequency"] = frequency5
tot_Runs_1969
```
### Export data for later analysis by bar chart.
```
wswin_1905.to_csv("../clean_data/wswin_1905Runs.csv")
wsnowin_1905.to_csv("../clean_data/nowswin_1905Runs.csv")
wswin_1969.to_csv("../clean_data/wswin_1969Runs.csv")
wsnowin_1969.to_csv("../clean_data/nowswin_1969Runs.csv")
tot_Runs_1905.to_csv("../clean_data/totwins_1905Runs.csv")
tot_Runs_1969.to_csv("../clean_data/totwins_1969Runs.csv")
```
| github_jupyter |
```
import os
import stocklab
import stocklab_twse.data_bundle
import stocklab_twse.analysis_bundle
config_file = 'config.yml'
stocklab.configure(config_file)
```
## Ad-hoc node
```
"""Ad-hoc node template"""
from stocklab.node import *
from stocklab import DataIdentifier as DI
from stocklab_twse.utils.datetime import Date
from stocklab_twse.utils import date_range
class FooNode(Node):
args = Args(
stock = Arg(),
date = Arg(type=Date),
)
def evaluate(stock, date):
raise NotImplementedError()
"""Installation"""
from stocklab.core import bundle
from stocklab.core.node import flush_cache
bundle.register(FooNode, allow_overwrite=True)
flush_cache()
"""Usage"""
raise NotImplementedError()
```
## Analysis feature
```
"""Plot stuff"""
#http://mrjbq7.github.io/ta-lib/
#https://ta-lib.org/d_api/ta_setunstableperiod.html
import numpy as np
from stocklab import DataIdentifier as DI
from stocklab_twse.utils import date_range
from stocklab_twse.utils.plot import plot
n = 160
sid = '2330'
dates = list(date_range(end=20181201, window=n))
dates.sort()
ohlcs = [DI('DailyData')(stock=sid, date=d) for d in dates]
ohlcs = [(inf['open'], inf['max'], inf['min'], inf['close'], inf['delta_price_share']) for inf in ohlcs]
ohlcs = np.asarray(ohlcs)
import talib
#talib.set_unstable_period('EMA', 70)
talib.set_unstable_period('ALL', 0)
patterns = talib.get_function_groups()['Pattern Recognition']
signs = np.array([getattr(talib, p)(ohlcs[:,0], ohlcs[:,1], ohlcs[:,2], ohlcs[:,3]) for p in patterns])
foo = talib.CDL3BLACKCROWS(ohlcs[:,0], ohlcs[:,1], ohlcs[:,2], ohlcs[:,3])
mom = talib.MOM(ohlcs[:,-2], timeperiod=5)
inphase, quadrature = talib.HT_PHASOR(ohlcs[:,-2])
#aux = {'UB': upperband, 'MB': middleband, 'LB': lowerband}
aux = {'foo': foo}
taux = {'vol': ohlcs[:,-1]}
taux = {'inphase': inphase, 'quad': quadrature}
#sgn = np.zeros(n)
#sgn = signs[9]
golden = np.array([
DI('GoldenSign')(stock=sid, date=d, window=15, gain=1.02, mode='time_first')
for d in dates
])
plot(dates, ohlcs[:,0:4], signs=golden, aux=aux)
#plot(dates, ohlcs[:,0:4], signs=sgn, aux=aux, top_aux=taux)
#print([(s>0).sum() for s in signs])
#print([(s<0).sum() for s in signs])
"""Performance evaluation (backtracing)"""
from stocklab_twse.utils import Simulator
signs = np.array([getattr(talib, p)(ohlcs[:,0], ohlcs[:,1], ohlcs[:,2], ohlcs[:,3]) for p in patterns])
sim = Simulator()
hold = sim(dates, golden, ohlcs[:,-2])
plot(dates, ohlcs[:,0:4], signs=golden, top_aux={'sim': hold})
```
| github_jupyter |
# ResNet CIFAR-10 with tensorboard
This notebook shows how to use TensorBoard, and how the training job writes checkpoints to a external bucket.
The model used for this notebook is a ResNet model, trained with the CIFAR-10 dataset.
See the following papers for more background:
[Deep Residual Learning for Image Recognition](https://arxiv.org/pdf/1512.03385.pdf) by Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun, Dec 2015.
[Identity Mappings in Deep Residual Networks](https://arxiv.org/pdf/1603.05027.pdf) by Kaiming He, Xiangyu Zhang, Shaoqing Ren, and Jian Sun, Jul 2016.
### Set up the environment
```
import os
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
role = get_execution_role()
```
### Download the CIFAR-10 dataset
Downloading the test and training data will take around 5 minutes.
```
import utils
utils.cifar10_download()
```
### Upload the data to a S3 bucket
```
inputs = sagemaker_session.upload_data(path='/tmp/cifar10_data', key_prefix='data/DEMO-cifar10')
```
**sagemaker_session.upload_data** will upload the CIFAR-10 dataset from your machine to a bucket named **sagemaker-{region}-{*your aws account number*}**, if you don't have this bucket yet, sagemaker_session will create it for you.
### Complete source code
- [source_dir/resnet_model.py](source_dir/resnet_model.py): ResNet model
- [source_dir/resnet_cifar_10.py](source_dir/resnet_cifar_10.py): main script used for training and hosting
## Create a training job using the sagemaker.TensorFlow estimator
```
from sagemaker.tensorflow import TensorFlow
source_dir = os.path.join(os.getcwd(), 'source_dir')
estimator = TensorFlow(entry_point='resnet_cifar_10.py',
source_dir=source_dir,
role=role,
hyperparameters={'min_eval_frequency': 10},
training_steps=1000, evaluation_steps=100,
train_instance_count=2, train_instance_type='ml.c4.xlarge',
base_job_name='tensorboard-example')
estimator.fit(inputs, run_tensorboard_locally=True)
```
The **```fit```** method will create a training job named **```tensorboard-example-{unique identifier}```** in two **ml.c4.xlarge** instances. These instances will write checkpoints to the s3 bucket **```sagemaker-{your aws account number}```**.
If you don't have this bucket yet, **```sagemaker_session```** will create it for you. These checkpoints can be used for restoring the training job, and to analyze training job metrics using **TensorBoard**.
The parameter **```run_tensorboard_locally=True```** will run **TensorBoard** in the machine that this notebook is running. Everytime a new checkpoint is created by the training job in the S3 bucket, **```fit```** will download the checkpoint to the temp folder that **TensorBoard** is pointing to.
When the **```fit```** method starts the training, it will log the port that **TensorBoard** is using to display the metrics. The default port is **6006**, but another port can be choosen depending on its availability. The port number will increase until finds an available port. After that the port number will printed in stdout.
It takes a few minutes to provision containers and start the training job.**TensorBoard** will start to display metrics shortly after that.
You can access **TensorBoard** locally at [http://localhost:6006](http://localhost:6006) or using your SageMaker notebook instance [proxy/6006/](/proxy/6006/)(TensorBoard will not work if forget to put the slash, '/', in end of the url). If TensorBoard started on a different port, adjust these URLs to match.This example uses the optional hyperparameter **```min_eval_frequency```** to generate training evaluations more often, allowing to visualize **TensorBoard** scalar data faster. You can find the available optional hyperparameters [here](https://github.com/aws/sagemaker-python-sdk#optional-hyperparameters)**.
# Deploy the trained model to prepare for predictions
The deploy() method creates an endpoint which serves prediction requests in real-time.
```
predictor = estimator.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
```
# Make a prediction with fake data to verify the endpoint is up
Prediction is not the focus of this notebook, so to verify the endpoint's functionality, we'll simply generate random data in the correct shape and make a prediction.
```
import numpy as np
random_image_data = np.random.rand(32, 32, 3)
predictor.predict(random_image_data)
```
# Cleaning up
To avoid incurring charges to your AWS account for the resources used in this tutorial you need to delete the **SageMaker Endpoint:**
```
sagemaker.Session().delete_endpoint(predictor.endpoint)
```
| github_jupyter |
# Building Dense Vectors Using Transformers
We will be using the [`sentence-transformers/stsb-distilbert-base`](https://huggingface.co/sentence-transformers/stsb-distilbert-base) model to build our dense vectors.
```
from transformers import AutoTokenizer, AutoModel
import torch
```
First we initialize our model and tokenizer:
```
tokenizer = AutoTokenizer.from_pretrained('sentence-transformers/stsb-distilbert-base')
model = AutoModel.from_pretrained('sentence-transformers/stsb-distilbert-base')
```
Then we tokenize a sentence just as we have been doing before:
```
text = "hello world what a time to be alive!"
tokens = tokenizer.encode_plus(text, max_length=128,
truncation=True, padding='max_length',
return_tensors='pt')
```
We process these tokens through our model:
```
outputs = model(**tokens)
outputs
```
The dense vector representations of our `text` are contained within the `outputs` **'last_hidden_state'** tensor, which we access like so:
```
embeddings = outputs.last_hidden_state
embeddings
embeddings.shape
```
After we have produced our dense vectors `embeddings`, we need to perform a *mean pooling* operation on them to create a single vector encoding (the **sentence embedding**). To do this mean pooling operation we will need to multiply each value in our `embeddings` tensor by it's respective `attention_mask` value - so that we ignore non-real tokens.
To perform this operation, we first resize our `attention_mask` tensor:
```
attention_mask = tokens['attention_mask']
attention_mask.shape
mask = attention_mask.unsqueeze(-1).expand(embeddings.size()).float()
mask.shape
attention_mask
mask[0][0].shape
mask
```
Each vector above represents a single token attention mask - each token now has a vector of size 768 representing it's *attention_mask* status. Then we multiply the two tensors to apply the attention mask:
```
masked_embeddings = embeddings * mask
masked_embeddings.shape
masked_embeddings
```
Then we sum the remained of the embeddings along axis `1`:
```
summed = torch.sum(masked_embeddings, 1)
summed.shape
```
Then sum the number of values that must be given attention in each position of the tensor:
```
summed_mask = torch.clamp(mask.sum(1), min=1e-9)
summed_mask.shape
summed_mask
```
Finally, we calculate the mean as the sum of the embedding activations `summed` divided by the number of values that should be given attention in each position `summed_mask`:
```
mean_pooled = summed / summed_mask
mean_pooled
```
And that is how we calculate our dense similarity vector.
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.gridspec as gridspec
from matplotlib.ticker import MaxNLocator
from scipy import stats
import matplotlib.pyplot as plt
import math
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_absolute_error, mean_squared_error
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import GridSearchCV, KFold
import plotly.express as px
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
data = pd.read_csv('/kaggle/input/car-purchase-data/Car_Purchasing_Data.csv',encoding='latin-1')
data
data.isnull().sum()
def plot_3chart(df, feature):
# Creating a customized chart. and giving in figsize and everything.
fig = plt.figure(constrained_layout=True, figsize=(27, 10))
# creating a grid of 3 cols and 3 rows.
grid = gridspec.GridSpec(ncols=3, nrows=3, figure=fig)
# Customizing the histogram grid.
ax1 = fig.add_subplot(grid[0, :2])
# Set the title.
ax1.set_title('Histogram')
# plot the histogram.
sns.distplot(df.loc[:, feature],
hist=True,
kde=True,
ax=ax1,
color='Red')
ax1.legend(labels=['Normal', 'Actual'])
# customizing the QQ_plot.
ax2 = fig.add_subplot(grid[1, :2])
# Set the title.
ax2.set_title('Probability Plot')
# Plotting the QQ_Plot.
stats.probplot(df.loc[:, feature].fillna(np.mean(df.loc[:, feature])),
plot=ax2)
ax2.get_lines()[0].set_markerfacecolor('Blue')
ax2.get_lines()[0].set_markersize(12.0)
# Customizing the Box Plot.
ax3 = fig.add_subplot(grid[:, 2])
# Set title.
ax3.set_title('Box Plot')
# Plotting the box plot.
sns.boxplot(df.loc[:, feature], orient='v', ax=ax3, color='Green')
ax3.yaxis.set_major_locator(MaxNLocator(nbins=24))
plt.suptitle(f'{feature}', fontsize=24)
plot_3chart(data, 'Age')
plot_3chart(data,'Annual Salary')
plot_3chart(data,'Net Worth')
plot_3chart(data,'Credit Card Debt')
sns.countplot(data['Gender'])
fig = px.treemap(data, path=['Country'], values='Annual Salary',
color='Net Worth', hover_data=['Country'],
color_continuous_scale='dense', title='Countries with different annual salaries ')
fig.show()
lable = LabelEncoder()
data.Country = lable.fit_transform(data.Country)
X = data.drop(["Customer Name", 'Country',"Customer e-mail", "Car Purchase Amount",'Credit Card Debt'], axis=1)
y = data["Car Purchase Amount"]
X.head()
sns.set(font_scale=1.1)
correlation_train = data.corr()
mask = np.triu(correlation_train.corr())
plt.figure(figsize=(18, 15))
sns.heatmap(correlation_train,
annot=True,
fmt='.1f',
cmap='coolwarm',
square=True,
mask=mask,
linewidths=1)
plt.show()
y1 = y
y1=y1.values.reshape(-1,1)
from sklearn.preprocessing import MinMaxScaler,StandardScaler
scaler=MinMaxScaler()
#scaler = StandardScaler()
y1 = scaler.fit_transform(y1)
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.15)
X_train
scores = []
n = 100
model1 = RandomForestRegressor(n_estimators = n)
model1.fit(X_train, y_train)
scores.append(model1.score(X_test, y_test))
y_pred1 = model1.predict(X_test)
RFerror = mean_absolute_error(y_test, y_pred1)
from sklearn import datasets, ensemble
from sklearn.inspection import permutation_importance
params = {'n_estimators': 1000,
'max_depth': 4,
'min_samples_split': 5,
'learning_rate': 0.01,
'loss': 'ls'}
reg = ensemble.GradientBoostingRegressor(**params)
reg.fit(X_train, y_train)
GBRerror = mean_absolute_error(y_test, reg.predict(X_test))
regpred = reg.staged_predict(X_test)
test_score = np.zeros((params['n_estimators'],), dtype=np.float64)
for i, y_pred in enumerate(reg.staged_predict(X_test)):
test_score[i] = reg.loss_(y_test, y_pred)
fig = plt.figure(figsize=(18, 10))
plt.subplot(1, 1, 1)
plt.title('Deviance')
plt.plot(np.arange(params['n_estimators']) + 1, reg.train_score_, 'b-',
label='Training Set Deviance')
plt.plot(np.arange(params['n_estimators']) + 1, test_score, 'r-',
label='Test Set Deviance')
plt.legend(loc='upper right')
plt.xlabel('Boosting Iterations')
plt.ylabel('Deviance')
fig.tight_layout()
plt.show()
from xgboost import XGBRegressor
xgb=XGBRegressor()
from sklearn.model_selection import cross_val_score
cv = 10
performance=cross_val_score(xgb,X,y,cv=cv,scoring="neg_mean_absolute_error",n_jobs=-1)
mae=-performance
xgb.fit(X,y)
y_pred3=xgb.predict(X_test)
print(mae)
XGBerror = mae
print("Mean Absolute Errors by: Random Forest =",RFerror)
print("Gradient Boost = ",GBRerror)
print("XGB regressor = ",XGBerror.mean())
```
# TRAINING ANN MODEL
```
X.shape
X=scaler.fit_transform(X)
X_train,X_test,y_train,y_test=train_test_split(X,y1,test_size=0.15)
import tensorflow.keras
from keras.models import Sequential
from keras.layers import Dense
model=Sequential()
model.add(Dense(80,input_dim=4,activation='relu'))
model.add(Dense(40,activation='relu'))
model.add(Dense(1,activation='linear'))
model.compile(optimizer='adam',loss='mean_squared_error')
model.summary()
epochs_hist=model.fit(X_train,y_train,epochs=200,batch_size=50,verbose=1,validation_split=0.2)
y_predict=model.predict(X_test)
y_predict.shape
mae = mean_absolute_error(y_test,y_predict)
mse = mean_squared_error(y_test,y_predict)
print(f'MAE = {mae}')
print(f'RMSE = {mse}')
ANNpredictions = pd.DataFrame(y_predict)
RFpredictions = pd.DataFrame(y_pred1)
XGBpredictions = pd.DataFrame(y_pred3)
GBRprediction = pd.DataFrame(regpred)
Xdata = pd.DataFrame(X)
ydata = pd.DataFrame(y)
```
# Mean Absolute Error of our Neural Network is far better than the Regressor Models.
```
Xdata.to_csv('Xdata.csv', index=False)
ydata.to_csv('ydata.csv', index=False)
ANNpredictions.to_csv('Predictedcarprices', index = False)
```
| github_jupyter |
```
"""Evaluate RPMNet. Also contains functionality to compute evaluation metrics given transforms
Example Usages:
1. Evaluate RPMNet
python eval.py --noise_type crop --resume [path-to-model.pth]
2. Evaluate precomputed transforms (.npy file containing np.array of size (B, 3, 4) or (B, n_iter, 3, 4))
python eval.py --noise_type crop --transform_file [path-to-transforms.npy]
"""
from collections import defaultdict
import json
import os
import pickle
import time
from typing import Dict, List
import numpy as np
import open3d # Need to import before torch
import pandas as pd
from scipy import sparse
from tqdm import tqdm
import torch
from arguments import rpmnet_eval_arguments
from common.misc import prepare_logger
from common.torch import dict_all_to_device, CheckPointManager, to_numpy
from common.math import se3
from common.math_torch import se3
from common.math.so3 import dcm2euler
from data_loader.datasets import get_test_datasets
import models.rpmnet
def compute_metrics(data: Dict, pred_transforms) -> Dict:
"""Compute metrics required in the paper
"""
def square_distance(src, dst):
return torch.sum((src[:, :, None, :] - dst[:, None, :, :]) ** 2, dim=-1)
with torch.no_grad():
pred_transforms = pred_transforms
gt_transforms = data['transform_gt']
points_src = data['points_src'][..., :3]
points_ref = data['points_ref'][..., :3]
points_raw = data['points_raw'][..., :3]
# Euler angles, Individual translation errors (Deep Closest Point convention)
# TODO Change rotation to torch operations
r_gt_euler_deg = dcm2euler(gt_transforms[:, :3, :3].detach().cpu().numpy(), seq='xyz')
r_pred_euler_deg = dcm2euler(pred_transforms[:, :3, :3].detach().cpu().numpy(), seq='xyz')
t_gt = gt_transforms[:, :3, 3]
t_pred = pred_transforms[:, :3, 3]
r_mse = np.mean((r_gt_euler_deg - r_pred_euler_deg) ** 2, axis=1)
r_mae = np.mean(np.abs(r_gt_euler_deg - r_pred_euler_deg), axis=1)
t_mse = torch.mean((t_gt - t_pred) ** 2, dim=1)
t_mae = torch.mean(torch.abs(t_gt - t_pred), dim=1)
# Rotation, translation errors (isotropic, i.e. doesn't depend on error
# direction, which is more representative of the actual error)
concatenated = se3.concatenate(se3.inverse(gt_transforms), pred_transforms)
rot_trace = concatenated[:, 0, 0] + concatenated[:, 1, 1] + concatenated[:, 2, 2]
residual_rotdeg = torch.acos(torch.clamp(0.5 * (rot_trace - 1), min=-1.0, max=1.0)) * 180.0 / np.pi
residual_transmag = concatenated[:, :, 3].norm(dim=-1)
# Modified Chamfer distance
src_transformed = se3.transform(pred_transforms, points_src)
ref_clean = points_raw
src_clean = se3.transform(se3.concatenate(pred_transforms, se3.inverse(gt_transforms)), points_raw)
dist_src = torch.min(square_distance(src_transformed, ref_clean), dim=-1)[0]
dist_ref = torch.min(square_distance(points_ref, src_clean), dim=-1)[0]
chamfer_dist = torch.mean(dist_src, dim=1) + torch.mean(dist_ref, dim=1)
metrics = {
'r_mse': r_mse,
'r_mae': r_mae,
't_mse': to_numpy(t_mse),
't_mae': to_numpy(t_mae),
'err_r_deg': to_numpy(residual_rotdeg),
'err_t': to_numpy(residual_transmag),
'chamfer_dist': to_numpy(chamfer_dist)
}
return metrics
def summarize_metrics(metrics):
"""Summaries computed metrices by taking mean over all data instances"""
summarized = {}
for k in metrics:
if k.endswith('mse'):
summarized[k[:-3] + 'rmse'] = np.sqrt(np.mean(metrics[k]))
elif k.startswith('err'):
summarized[k + '_mean'] = np.mean(metrics[k])
summarized[k + '_rmse'] = np.sqrt(np.mean(metrics[k]**2))
else:
summarized[k] = np.mean(metrics[k])
return summarized
def print_metrics(logger, summary_metrics: Dict, losses_by_iteration: List = None,
title: str = 'Metrics'):
"""Prints out formated metrics to logger"""
logger.info(title + ':')
logger.info('=' * (len(title) + 1))
if losses_by_iteration is not None:
losses_all_str = ' | '.join(['{:.5f}'.format(c) for c in losses_by_iteration])
logger.info('Losses by iteration: {}'.format(losses_all_str))
logger.info('DeepCP metrics:{:.4f}(rot-rmse) | {:.4f}(rot-mae) | {:.4g}(trans-rmse) | {:.4g}(trans-mae)'.format(
summary_metrics['r_rmse'], summary_metrics['r_mae'],
summary_metrics['t_rmse'], summary_metrics['t_mae'],
))
logger.info('Rotation error {:.4f}(deg, mean) | {:.4f}(deg, rmse)'.format(
summary_metrics['err_r_deg_mean'], summary_metrics['err_r_deg_rmse']))
logger.info('Translation error {:.4g}(mean) | {:.4g}(rmse)'.format(
summary_metrics['err_t_mean'], summary_metrics['err_t_rmse']))
logger.info('Chamfer error: {:.7f}(mean-sq)'.format(
summary_metrics['chamfer_dist']
))
def inference(data_loader, model: torch.nn.Module):
"""Runs inference over entire dataset
Args:
data_loader (torch.utils.data.DataLoader): Dataset loader
model (model.nn.Module): Network model to evaluate
Returns:
pred_transforms_all: predicted transforms (B, n_iter, 3, 4) where B is total number of instances
endpoints_out (Dict): Network endpoints
"""
_logger.info('Starting inference...')
model.eval()
pred_transforms_all = []
all_betas, all_alphas = [], []
total_time = 0.0
endpoints_out = defaultdict(list)
total_rotation = []
with torch.no_grad():
for val_data in tqdm(data_loader):
rot_trace = val_data['transform_gt'][:, 0, 0] + val_data['transform_gt'][:, 1, 1] + \
val_data['transform_gt'][:, 2, 2]
rotdeg = torch.acos(torch.clamp(0.5 * (rot_trace - 1), min=-1.0, max=1.0)) * 180.0 / np.pi
total_rotation.append(np.abs(to_numpy(rotdeg)))
dict_all_to_device(val_data, _device)
time_before = time.time()
pred_transforms, endpoints = model(val_data, _args.num_reg_iter)
total_time += time.time() - time_before
if _args.method == 'rpmnet':
all_betas.append(endpoints['beta'])
all_alphas.append(endpoints['alpha'])
if isinstance(pred_transforms[-1], torch.Tensor):
pred_transforms_all.append(to_numpy(torch.stack(pred_transforms, dim=1)))
else:
pred_transforms_all.append(np.stack(pred_transforms, axis=1))
# Saves match matrix. We only save the top matches to save storage/time.
# However, this still takes quite a bit of time to save. Comment out if not needed.
if 'perm_matrices' in endpoints:
perm_matrices = to_numpy(torch.stack(endpoints['perm_matrices'], dim=1))
thresh = np.percentile(perm_matrices, 99.9, axis=[2, 3]) # Only retain top 0.1% of entries
below_thresh_mask = perm_matrices < thresh[:, :, None, None]
perm_matrices[below_thresh_mask] = 0.0
for i_data in range(perm_matrices.shape[0]):
sparse_perm_matrices = []
for i_iter in range(perm_matrices.shape[1]):
sparse_perm_matrices.append(sparse.coo_matrix(perm_matrices[i_data, i_iter, :, :]))
endpoints_out['perm_matrices'].append(sparse_perm_matrices)
_logger.info('Total inference time: {}s'.format(total_time))
total_rotation = np.concatenate(total_rotation, axis=0)
_logger.info('Rotation range in data: {}(avg), {}(max)'.format(np.mean(total_rotation), np.max(total_rotation)))
pred_transforms_all = np.concatenate(pred_transforms_all, axis=0)
return pred_transforms_all, endpoints_out
def evaluate(pred_transforms, data_loader: torch.utils.data.dataloader.DataLoader):
""" Evaluates the computed transforms against the groundtruth
Args:
pred_transforms: Predicted transforms (B, [iter], 3/4, 4)
data_loader: Loader for dataset.
Returns:
Computed metrics (List of dicts), and summary metrics (only for last iter)
"""
_logger.info('Evaluating transforms...')
num_processed, num_total = 0, len(pred_transforms)
if pred_transforms.ndim == 4:
pred_transforms = torch.from_numpy(pred_transforms).to(_device)
else:
assert pred_transforms.ndim == 3 and \
(pred_transforms.shape[1:] == (4, 4) or pred_transforms.shape[1:] == (3, 4))
pred_transforms = torch.from_numpy(pred_transforms[:, None, :, :]).to(_device)
metrics_for_iter = [defaultdict(list) for _ in range(pred_transforms.shape[1])]
for data in tqdm(data_loader, leave=False):
dict_all_to_device(data, _device)
batch_size = 0
for i_iter in range(pred_transforms.shape[1]):
batch_size = data['points_src'].shape[0]
cur_pred_transforms = pred_transforms[num_processed:num_processed+batch_size, i_iter, :, :]
metrics = compute_metrics(data, cur_pred_transforms)
for k in metrics:
metrics_for_iter[i_iter][k].append(metrics[k])
num_processed += batch_size
for i_iter in range(len(metrics_for_iter)):
metrics_for_iter[i_iter] = {k: np.concatenate(metrics_for_iter[i_iter][k], axis=0)
for k in metrics_for_iter[i_iter]}
summary_metrics = summarize_metrics(metrics_for_iter[i_iter])
print_metrics(_logger, summary_metrics, title='Evaluation result (iter {})'.format(i_iter))
return metrics_for_iter, summary_metrics
def save_eval_data(pred_transforms, endpoints, metrics, summary_metrics, save_pathdic):
"""Saves out the computed transforms
"""
# Save transforms
np.save(os.path.join(save_path, 'pred_transforms.npy'), pred_transforms)
# Save endpoints if any
for k in endpoints:
if isinstance(endpoints[k], np.ndarray):
np.save(os.path.join(save_path, '{}.npy'.format(k)), endpoints[k])
else:
with open(os.path.join(save_path, '{}.pickle'.format(k)), 'wb') as fid:
pickle.dump(endpoints[k], fid)
# Save metrics: Write each iteration to a different worksheet.
writer = pd.ExcelWriter(os.path.join(save_path, 'metrics.xlsx'))
for i_iter in range(len(metrics)):
metrics[i_iter]['r_rmse'] = np.sqrt(metrics[i_iter]['r_mse'])
metrics[i_iter]['t_rmse'] = np.sqrt(metrics[i_iter]['t_mse'])
metrics[i_iter].pop('r_mse')
metrics[i_iter].pop('t_mse')
metrics_df = pd.DataFrame.from_dict(metrics[i_iter])
metrics_df.to_excel(writer, sheet_name='Iter_{}'.format(i_iter+1))
writer.close()
# Save summary metrics
summary_metrics_float = {k: float(summary_metrics[k]) for k in summary_metrics}
with open(os.path.join(save_path, 'summary_metrics.json'), 'w') as json_out:
json.dump(summary_metrics_float, json_out)
_logger.info('Saved evaluation results to {}'.format(save_path))
def get_model():
_logger.info('Computing transforms using {}'.format(_args.method))
if _args.method == 'rpmnet':
assert _args.resume is not None
model = models.rpmnet.get_model(_args)
model.to(_device)
saver = CheckPointManager(os.path.join(_log_path, 'ckpt', 'models'))
saver.load(_args.resume, model)
else:
raise NotImplementedError
return model
def main():
# Load data_loader
test_dataset = get_test_datasets(_args)
test_loader = torch.utils.data.DataLoader(test_dataset,
batch_size=_args.val_batch_size, shuffle=False)
if _args.transform_file is not None:
_logger.info('Loading from precomputed transforms: {}'.format(_args.transform_file))
pred_transforms = np.load(_args.transform_file)
endpoints = {}
else:
model = get_model()
pred_transforms, endpoints = inference(test_loader, model) # Feedforward transforms
# Compute evaluation matrices
eval_metrics, summary_metrics = evaluate(pred_transforms, data_loader=test_loader)
save_eval_data(pred_transforms, endpoints, eval_metrics, summary_metrics, _args.eval_save_path)
_logger.info('Finished')
parser = rpmnet_eval_arguments()
_args = parser.parse_args(args=["--resume=../logs/clean-trained.pth", "--num_reg_iter=10"])
# print(args)
_logger, _log_path = prepare_logger(_args, log_path=_args.eval_save_path)
os.environ['CUDA_VISIBLE_DEVICES'] = str(_args.gpu)
if _args.gpu >= 0 and (_args.method == 'rpm' or _args.method == 'rpmnet'):
os.environ['CUDA_VISIBLE_DEVICES'] = str(_args.gpu)
_device = torch.device('cuda:0') if torch.cuda.is_available() else torch.device('cpu')
else:
_device = torch.device('cpu')
model = get_model()
model.eval()
```
## official test set
```
# main()
test_dataset = get_test_datasets(_args)
test_loader = torch.utils.data.DataLoader(test_dataset,
batch_size=_args.val_batch_size, shuffle=False)
i = 0
val_data = ''
for data in test_loader:
if i==1:
val_data = data
i += 1
# help(dict_all_to_device)
dict_all_to_device(val_data, _device)
model.eval()
with torch.no_grad():
dict_all_to_device(val_data, _device)
pred_transforms, endpoints = model(val_data, _args.num_reg_iter)
val_data['points_ref'].shape, val_data['points_src'].shape
```
## load LiDAR cloud and compute transform
```
import open3d as o3d
# pcd1 = o3d.io.read_point_cloud("/home/li/Documents/pcl_tutorial/room_scan1_sub.pcd")
# pcd2 = o3d.io.read_point_cloud("/home/li/Documents/pcl_tutorial/room_scan2_sub.pcd")
# pcd1 = o3d.io.read_point_cloud("/home/li/Documents/pcl_tutorial/remaining_cloud011_sub.pcd")
# pcd2 = o3d.io.read_point_cloud("/home/li/Documents/pcl_tutorial/remaining_cloud012_sub.pcd")
pcd1 = o3d.io.read_point_cloud("/home/li/car1_map_sub.pcd")
pcd2 = o3d.io.read_point_cloud("/home/li/car2_map_sub.pcd")
pc1 = np.asarray(pcd1.points)
pc2 = np.asarray(pcd2.points)
center1 = pc1.mean(axis=0)
center2 = pc2.mean(axis=0)
# np.asarray(pcd1.normals)
val_data = {}
val_data['points_ref'] = torch.Tensor([np.concatenate(
[np.asarray(pcd1.points)-center1, np.asarray(pcd1.normals)], axis=1)])
val_data['points_src'] = torch.Tensor([np.concatenate(
[np.asarray(pcd2.points)-center2, np.asarray(pcd2.normals)], axis=1)])
val_data['points_ref'].shape, val_data['points_src'].shape
model.eval()
with torch.no_grad():
dict_all_to_device(val_data, _device)
pred_transforms, endpoints = model(val_data, _args.num_reg_iter)
pred_transforms
transform = pred_transforms[-1][0].tolist()
T_ab_str = ''
for i in range(3):
for j in range(4):
T_ab_str += str(transform[i][j]) + ' '
T_ab_str += '\n'
T_ab_str += '0 0 0 1'
# print(l)
# T_ab_str += str(rotation_ab_pred_list[i][0]) + ' ' + str(rotation_ab_pred_list[i][1]) \
# + ' ' + str(rotation_ab_pred_list[i][2]) + ' ' \
# + str(translation_ab_pred_list[i]) + '\n'
print(T_ab_str)
```
## Load single point cloud and manually transform it as a source cloud
```
import data_loader.transforms as Transforms
import common.math.se3 as se3
SE3_Z = Transforms.RandomRotatorZ()
transform_mat = SE3_Z.generate_transform()
print(transform_mat)
import open3d as o3d
pcd = o3d.io.read_point_cloud("/home/li/Lille_street_small_sub.pcd")
pc1 = np.asarray(pcd.points)
pc2 = se3.transform(transform_mat, pc1[:, :3])
# pc1.shape, pc2.shape
center1 = pc1.mean(axis=0)
center2 = pc2.mean(axis=0)
# center1, center2
pc1 = pc1 - center1
pc2 = pc2 - center2
normals1 = np.asarray(pcd.normals)
normals2 = se3.transform(transform_mat, normals1)
# normals1, normals2
center1, center2
val_data = {}
val_data['points_ref'] = torch.Tensor([np.concatenate(
[pc1, normals1], axis=1)])
val_data['points_src'] = torch.Tensor([np.concatenate(
[pc2, normals2], axis=1)])
val_data['points_ref'].shape, val_data['points_src'].shape
model.eval()
with torch.no_grad():
dict_all_to_device(val_data, _device)
pred_transforms, endpoints = model(val_data, _args.num_reg_iter)
pred_transforms
transform = pred_transforms[-1][0].tolist()
T_ab_str = ''
for i in range(3):
for j in range(4):
T_ab_str += str(transform[i][j]) + ' '
T_ab_str += '\n'
T_ab_str += '0 0 0 1'
print(T_ab_str)
pcd.points = o3d.utility.Vector3dVector(pc1)
o3d.io.write_point_cloud("/home/li/Lille_street_small_sub.pcd", pcd)
pcd2 = o3d.geometry.PointCloud()
pcd2.points = o3d.utility.Vector3dVector(pc2)
o3d.io.write_point_cloud("/home/li/Lille_street_small_rotated.pcd", pcd2)
""" Train RPMNet
Example usage:
python train.py --noise_type crop
python train.py --noise_type jitter --train_batch_size 4
"""
from collections import defaultdict
import os
import random
from typing import Dict, List
from matplotlib.pyplot import cm as colormap
import numpy as np
import open3d # Ensure this is imported before pytorch
from tensorboardX import SummaryWriter
import torch
import torch.nn as nn
import torch.utils.data
from tqdm import tqdm
from arguments import rpmnet_train_arguments
from common.colors import BLUE, ORANGE
from common.misc import prepare_logger
from common.torch import dict_all_to_device, CheckPointManager, TorchDebugger
from common.math_torch import se3
from data_loader.datasets import get_train_datasets
from eval import compute_metrics, summarize_metrics, print_metrics
from models.rpmnet import get_model
# Set up arguments and logging
parser = rpmnet_train_arguments()
_args = parser.parse_args()
_logger, _log_path = prepare_logger(_args)
if _args.gpu >= 0:
os.environ['CUDA_VISIBLE_DEVICES'] = str(_args.gpu)
_device = torch.device('cuda:0') if torch.cuda.is_available() else torch.device('cpu')
else:
_device = torch.device('cpu')
def main():
train_set, val_set = get_train_datasets(_args)
run(train_set, val_set)
def compute_losses(data: Dict, pred_transforms: List, endpoints: Dict,
loss_type: str = 'mae', reduction: str = 'mean') -> Dict:
"""Compute losses
Args:
data: Current mini-batch data
pred_transforms: Predicted transform, to compute main registration loss
endpoints: Endpoints for training. For computing outlier penalty
loss_type: Registration loss type, either 'mae' (Mean absolute error, used in paper) or 'mse'
reduction: Either 'mean' or 'none'. Use 'none' to accumulate losses outside
(useful for accumulating losses for entire validation dataset)
Returns:
losses: Dict containing various fields. Total loss to be optimized is in losses['total']
"""
losses = {}
num_iter = len(pred_transforms)
# Compute losses
gt_src_transformed = se3.transform(data['transform_gt'], data['points_src'][..., :3])
if loss_type == 'mse':
# MSE loss to the groundtruth (does not take into account possible symmetries)
criterion = nn.MSELoss(reduction=reduction)
for i in range(num_iter):
pred_src_transformed = se3.transform(pred_transforms[i], data['points_src'][..., :3])
if reduction.lower() == 'mean':
losses['mse_{}'.format(i)] = criterion(pred_src_transformed, gt_src_transformed)
elif reduction.lower() == 'none':
losses['mse_{}'.format(i)] = torch.mean(criterion(pred_src_transformed, gt_src_transformed),
dim=[-1, -2])
elif loss_type == 'mae':
# MSE loss to the groundtruth (does not take into account possible symmetries)
criterion = nn.L1Loss(reduction=reduction)
for i in range(num_iter):
pred_src_transformed = se3.transform(pred_transforms[i], data['points_src'][..., :3])
if reduction.lower() == 'mean':
losses['mae_{}'.format(i)] = criterion(pred_src_transformed, gt_src_transformed)
elif reduction.lower() == 'none':
losses['mae_{}'.format(i)] = torch.mean(criterion(pred_src_transformed, gt_src_transformed),
dim=[-1, -2])
else:
raise NotImplementedError
# Penalize outliers
for i in range(num_iter):
ref_outliers_strength = (1.0 - torch.sum(endpoints['perm_matrices'][i], dim=1)) * _args.wt_inliers
src_outliers_strength = (1.0 - torch.sum(endpoints['perm_matrices'][i], dim=2)) * _args.wt_inliers
if reduction.lower() == 'mean':
losses['outlier_{}'.format(i)] = torch.mean(ref_outliers_strength) + torch.mean(src_outliers_strength)
elif reduction.lower() == 'none':
losses['outlier_{}'.format(i)] = torch.mean(ref_outliers_strength, dim=1) + \
torch.mean(src_outliers_strength, dim=1)
discount_factor = 0.5 # Early iterations will be discounted
total_losses = []
for k in losses:
discount = discount_factor ** (num_iter - int(k[k.rfind('_')+1:]) - 1)
total_losses.append(losses[k] * discount)
losses['total'] = torch.sum(torch.stack(total_losses), dim=0)
return losses
def save_summaries(writer: SummaryWriter, data: Dict, predicted: List, endpoints: Dict = None,
losses: Dict = None, metrics: Dict = None, step: int = 0):
"""Save tensorboard summaries"""
subset = [0, 1]
with torch.no_grad():
# Save clouds
if 'points_src' in data:
points_src = data['points_src'][subset, ..., :3]
points_ref = data['points_ref'][subset, ..., :3]
colors = torch.from_numpy(
np.concatenate([np.tile(ORANGE, (*points_src.shape[0:2], 1)),
np.tile(BLUE, (*points_ref.shape[0:2], 1))], axis=1))
iters_to_save = [0, len(predicted)-1] if len(predicted) > 1 else [0]
# Save point cloud at iter0, iter1 and after last iter
concat_cloud_input = torch.cat((points_src, points_ref), dim=1)
writer.add_mesh('iter_0', vertices=concat_cloud_input, colors=colors, global_step=step)
for i_iter in iters_to_save:
src_transformed_first = se3.transform(predicted[i_iter][subset, ...], points_src)
concat_cloud_first = torch.cat((src_transformed_first, points_ref), dim=1)
writer.add_mesh('iter_{}'.format(i_iter+1), vertices=concat_cloud_first, colors=colors, global_step=step)
if endpoints is not None and 'perm_matrices' in endpoints:
color_mapper = colormap.ScalarMappable(norm=None, cmap=colormap.get_cmap('coolwarm'))
for i_iter in iters_to_save:
ref_weights = torch.sum(endpoints['perm_matrices'][i_iter][subset, ...], dim=1)
ref_colors = color_mapper.to_rgba(ref_weights.detach().cpu().numpy())[..., :3]
writer.add_mesh('ref_weights_{}'.format(i_iter), vertices=points_ref,
colors=torch.from_numpy(ref_colors) * 255, global_step=step)
if endpoints is not None:
if 'perm_matrices' in endpoints:
for i_iter in range(len(endpoints['perm_matrices'])):
src_weights = torch.sum(endpoints['perm_matrices'][i_iter], dim=2)
ref_weights = torch.sum(endpoints['perm_matrices'][i_iter], dim=1)
writer.add_histogram('src_weights_{}'.format(i_iter), src_weights, global_step=step)
writer.add_histogram('ref_weights_{}'.format(i_iter), ref_weights, global_step=step)
# Write losses and metrics
if losses is not None:
for l in losses:
writer.add_scalar('losses/{}'.format(l), losses[l], step)
if metrics is not None:
for m in metrics:
writer.add_scalar('metrics/{}'.format(m), metrics[m], step)
writer.flush()
def validate(data_loader, model: torch.nn.Module, summary_writer: SummaryWriter, step: int):
"""Perform a single validation run, and saves results into tensorboard summaries"""
_logger.info('Starting validation run...')
with torch.no_grad():
all_val_losses = defaultdict(list)
all_val_metrics_np = defaultdict(list)
for val_data in data_loader:
dict_all_to_device(val_data, _device)
pred_test_transforms, endpoints = model(val_data, _args.num_reg_iter)
val_losses = compute_losses(val_data, pred_test_transforms, endpoints,
loss_type=_args.loss_type, reduction='none')
val_metrics = compute_metrics(val_data, pred_test_transforms[-1])
for k in val_losses:
all_val_losses[k].append(val_losses[k])
for k in val_metrics:
all_val_metrics_np[k].append(val_metrics[k])
all_val_losses = {k: torch.cat(all_val_losses[k]) for k in all_val_losses}
all_val_metrics_np = {k: np.concatenate(all_val_metrics_np[k]) for k in all_val_metrics_np}
mean_val_losses = {k: torch.mean(all_val_losses[k]) for k in all_val_losses}
# Rerun on random and worst data instances and save to summary
rand_idx = random.randint(0, all_val_losses['total'].shape[0] - 1)
worst_idx = torch.argmax(all_val_losses['{}_{}'.format(_args.loss_type, _args.num_reg_iter - 1)]).cpu().item()
indices_to_rerun = [rand_idx, worst_idx]
data_to_rerun = defaultdict(list)
for i in indices_to_rerun:
cur = data_loader.dataset[i]
for k in cur:
data_to_rerun[k].append(cur[k])
for k in data_to_rerun:
data_to_rerun[k] = torch.from_numpy(np.stack(data_to_rerun[k], axis=0))
dict_all_to_device(data_to_rerun, _device)
pred_transforms, endpoints = model(data_to_rerun, _args.num_reg_iter)
summary_metrics = summarize_metrics(all_val_metrics_np)
losses_by_iteration = torch.stack([mean_val_losses['{}_{}'.format(_args.loss_type, k)]
for k in range(_args.num_reg_iter)]).cpu().numpy()
print_metrics(_logger, summary_metrics, losses_by_iteration, 'Validation results')
save_summaries(summary_writer, data=data_to_rerun, predicted=pred_transforms, endpoints=endpoints,
losses=mean_val_losses, metrics=summary_metrics, step=step)
score = -summary_metrics['chamfer_dist']
return score
def run(train_set, val_set):
"""Main train/val loop"""
_logger.debug('Trainer (PID=%d), %s', os.getpid(), _args)
model = get_model(_args)
model.to(_device)
global_step = 0
# dataloaders
train_loader = torch.utils.data.DataLoader(train_set,
batch_size=_args.train_batch_size, shuffle=True, num_workers=_args.num_workers)
val_loader = torch.utils.data.DataLoader(val_set,
batch_size=_args.val_batch_size, shuffle=False, num_workers=_args.num_workers)
# optimizer
optimizer = torch.optim.Adam(model.parameters(), lr=_args.lr)
# Summary writer and Checkpoint manager
train_writer = SummaryWriter(os.path.join(_log_path, 'train'), flush_secs=10)
val_writer = SummaryWriter(os.path.join(_log_path, 'val'), flush_secs=10)
saver = CheckPointManager(os.path.join(_log_path, 'ckpt', 'model'), keep_checkpoint_every_n_hours=0.5)
if _args.resume is not None:
global_step = saver.load(_args.resume, model, optimizer)
# trainings
torch.autograd.set_detect_anomaly(_args.debug)
model.train()
steps_per_epoch = len(train_loader)
if _args.summary_every < 0:
_args.summary_every = abs(_args.summary_every) * steps_per_epoch
if _args.validate_every < 0:
_args.validate_every = abs(_args.validate_every) * steps_per_epoch
for epoch in range(0, _args.epochs):
_logger.info('Begin epoch {} (steps {} - {})'.format(epoch, global_step, global_step + len(train_loader)))
tbar = tqdm(total=len(train_loader), ncols=100)
for train_data in train_loader:
global_step += 1
optimizer.zero_grad()
# Forward through neural network
dict_all_to_device(train_data, _device)
pred_transforms, endpoints = model(train_data, _args.num_train_reg_iter) # Use less iter during training
# Compute loss, and optimize
train_losses = compute_losses(train_data, pred_transforms, endpoints,
loss_type=_args.loss_type, reduction='mean')
if _args.debug:
with TorchDebugger():
train_losses['total'].backward()
else:
train_losses['total'].backward()
optimizer.step()
tbar.set_description('Loss:{:.3g}'.format(train_losses['total']))
tbar.update(1)
if global_step % _args.summary_every == 0: # Save tensorboard logs
save_summaries(train_writer, data=train_data, predicted=pred_transforms, endpoints=endpoints,
losses=train_losses, step=global_step)
if global_step % _args.validate_every == 0: # Validation loop. Also saves checkpoints
model.eval()
val_score = validate(val_loader, model, val_writer, global_step)
saver.save(model, optimizer, step=global_step, score=val_score)
model.train()
tbar.close()
_logger.info('Ending training. Number of steps = {}.'.format(global_step))
if __name__ == '__main__':
main()
```
| github_jupyter |
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import itertools
import os
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.preprocessing import LabelBinarizer, LabelEncoder
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import confusion_matrix
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout
from keras.preprocessing import text, sequence
from keras import utils
# This code was tested with TensorFlow v1.4
print("You have TensorFlow version", tf.__version__)
data = pd.read_csv("C:/Users/Adithya/Desktop/train.csv")
data.head()
data['tags'].value_counts()
train_size = int(len(data) * .8)
print ("Train size: %d" % train_size)
print ("Test size: %d" % (len(data) - train_size))
train_posts = data['title'][:train_size]
train_tags = data['tags'][:train_size]
test_posts = data['title'][train_size:]
test_tags = data['tags'][train_size:]
max_words = 15000
tokenize = text.Tokenizer(num_words=max_words, char_level=False)
tokenize.fit_on_texts(train_posts) # only fit on train
x_test = tokenize.texts_to_matrix(test_posts)
x_train = tokenize.texts_to_matrix(train_posts)
print(x_test.shape)
encoder = LabelEncoder()
encoder.fit(train_tags)
test_tags = test_tags.map(lambda s: '<unknown>' if s not in encoder.classes_ else s)
encoder.classes_ = np.append(encoder.classes_, '<unknown>')
y_train = encoder.transform(train_tags)
y_test = encoder.transform(test_tags)
num_classes = np.max(y_train)+2
y_train = utils.to_categorical(y_train, num_classes)
y_test = utils.to_categorical(y_test, num_classes)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print('y_train shape:', y_train.shape)
print('y_test shape:', y_test.shape)
# This model trains very quickly and 2 epochs are already more than enough
# Training for more epochs will likely lead to overfitting on this dataset
# You can try tweaking these hyperparamaters when using this model with your own data
batch_size = 32
epochs = 2
model = Sequential()
model.add(Dense(512, input_shape=(max_words,)))
model.add(Activation('relu'))
model.add(Dropout(0.3))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# model.fit trains the model
# The validation_split param tells Keras what % of our training data should be used in the validation set
# You can see the validation loss decreasing slowly when you run this
# Because val_loss is no longer decreasing we stop training to prevent overfitting
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_split=0.1)
score = model.evaluate(x_test, y_test,
batch_size=batch_size, verbose=1)
print('Test score:', score[0])
print('Test accuracy:', score[1])
text_labels = encoder.classes_
for i in range(30):
prediction = model.predict(np.array([x_test[i]]))
predicted_label = text_labels[np.argmax(prediction)]
print(test_posts.iloc[i][:50], "...")
print('Actual label:' + test_tags.iloc[i])
print("Predicted label: " + predicted_label + "\n")
y_softmax = model.predict(x_test)
y_test_1d = []
y_pred_1d = []
for i in range(len(y_test)):
probs = y_test[i]
index_arr = np.nonzero(probs)
one_hot_index = index_arr[0].item(0)
y_test_1d.append(one_hot_index)
for i in range(0, len(y_softmax)):
probs = y_softmax[i]
predicted_index = np.argmax(probs)
y_pred_1d.append(predicted_index)
def plot_confusion_matrix(cm, classes,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title, fontsize=30)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45, fontsize=22)
plt.yticks(tick_marks, classes, fontsize=22)
fmt = '.2f'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label', fontsize=25)
plt.xlabel('Predicted label', fontsize=25)
text=['elif using dictionary but without excecute the values']
yes = tokenize.texts_to_matrix(text)
prediction = model.predict(np.array(yes))
predicted_label = text_labels[np.argmax(prediction)]
print(predicted_label)
from keras.models import load_model
model.save('/home/adithya/Desktop/my_model1.h5')
```
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Public/5.1_Text_classification_examples_in_SparkML_SparkNLP.ipynb)
# Text Classification with Spark NLP
```
import os
import sys
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
from sparknlp.annotator import *
from sparknlp.common import *
from sparknlp.base import *
import pandas as pd
import sparknlp
spark = sparknlp.start()
print("Spark NLP version: ", sparknlp.version())
print("Apache Spark version: ", spark.version)
! wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Public/data/news_category_train.csv
! wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Public/data/news_category_test.csv
# newsDF = spark.read.parquet("data/news_category.parquet") >> if it is a parquet
newsDF = spark.read \
.option("header", True) \
.csv("news_category_train.csv")
newsDF.show(truncate=50)
newsDF.show(truncate=50)
newsDF.take(2)
from pyspark.sql.functions import col
newsDF.groupBy("category") \
.count() \
.orderBy(col("count").desc()) \
.show()
```
## Building Classification Pipeline
### LogReg with CountVectorizer
Tokenizer: Tokenization
stopwordsRemover: Remove Stop Words
countVectors: Count vectors (“document-term vectors”)
```
from pyspark.ml.feature import CountVectorizer, HashingTF, IDF, OneHotEncoder, StringIndexer, VectorAssembler, SQLTransformer
%%time
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
stemmer = Stemmer() \
.setInputCols(["cleanTokens"]) \
.setOutputCol("stem")
finisher = Finisher() \
.setInputCols(["stem"]) \
.setOutputCols(["token_features"]) \
.setOutputAsArray(True) \
.setCleanAnnotations(False)
countVectors = CountVectorizer(inputCol="token_features", outputCol="features", vocabSize=10000, minDF=5)
label_stringIdx = StringIndexer(inputCol = "category", outputCol = "label")
nlp_pipeline = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
stemmer,
finisher,
countVectors,
label_stringIdx])
nlp_model = nlp_pipeline.fit(newsDF)
processed = nlp_model.transform(newsDF)
processed.count()
processed.select('description','token_features').show(truncate=50)
processed.select('token_features').take(2)
processed.select('features').take(2)
processed.select('description','features','label').show()
# set seed for reproducibility
(trainingData, testData) = processed.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
trainingData.printSchema()
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression(maxIter=10, regParam=0.3, elasticNetParam=0)
lrModel = lr.fit(trainingData)
predictions = lrModel.transform(testData)
predictions.filter(predictions['prediction'] == 0) \
.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
evaluator = MulticlassClassificationEvaluator(predictionCol="prediction")
evaluator.evaluate(predictions)
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
y_true = predictions_tf.select("label")
y_true = y_true.toPandas()
y_pred = predictions_tf.select("prediction")
y_pred = y_pred.toPandas()
y_pred.prediction.value_counts()
cnf_matrix = confusion_matrix(list(y_true.label.astype(int)), list(y_pred.prediction.astype(int)))
cnf_matrix
print(classification_report(y_true.label, y_pred.prediction))
print(accuracy_score(y_true.label, y_pred.prediction))
```
### LogReg with TFIDF
```
from pyspark.ml.feature import HashingTF, IDF
hashingTF = HashingTF(inputCol="token_features", outputCol="rawFeatures", numFeatures=10000)
idf = IDF(inputCol="rawFeatures", outputCol="features", minDocFreq=5) #minDocFreq: remove sparse terms
nlp_pipeline_tf = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
stemmer,
finisher,
hashingTF,
idf,
label_stringIdx])
nlp_model_tf = nlp_pipeline_tf.fit(newsDF)
processed_tf = nlp_model_tf.transform(newsDF)
processed_tf.count()
# set seed for reproducibility
processed_tf.select('description','features','label').show()
(trainingData, testData) = processed_tf.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
lrModel_tf = lr.fit(trainingData)
predictions_tf = lrModel_tf.transform(testData)
predictions_tf.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
y_true = predictions_tf.select("label")
y_true = y_true.toPandas()
y_pred = predictions_tf.select("prediction")
y_pred = y_pred.toPandas()
print(classification_report(y_true.label, y_pred.prediction))
print(accuracy_score(y_true.label, y_pred.prediction))
```
### Random Forest with TFIDF
```
from pyspark.ml.classification import RandomForestClassifier
rf = RandomForestClassifier(labelCol="label", \
featuresCol="features", \
numTrees = 100, \
maxDepth = 4, \
maxBins = 32)
# Train model with Training Data
rfModel = rf.fit(trainingData)
predictions_rf = rfModel.transform(testData)
predictions_rf.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
y_true = predictions_rf.select("label")
y_true = y_true.toPandas()
y_pred = predictions_rf.select("prediction")
y_pred = y_pred.toPandas()
print(classification_report(y_true.label, y_pred.prediction))
print(accuracy_score(y_true.label, y_pred.prediction))
```
## LogReg with Spark NLP Glove Word Embeddings
```
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
glove_embeddings = WordEmbeddingsModel().pretrained() \
.setInputCols(["document",'cleanTokens'])\
.setOutputCol("embeddings")\
.setCaseSensitive(False)
embeddingsSentence = SentenceEmbeddings() \
.setInputCols(["document", "embeddings"]) \
.setOutputCol("sentence_embeddings") \
.setPoolingStrategy("AVERAGE")
embeddings_finisher = EmbeddingsFinisher() \
.setInputCols(["sentence_embeddings"]) \
.setOutputCols(["finished_sentence_embeddings"]) \
.setOutputAsVector(True)\
.setCleanAnnotations(False)
explodeVectors = SQLTransformer(statement=
"SELECT EXPLODE(finished_sentence_embeddings) AS features, * FROM __THIS__")
label_stringIdx = StringIndexer(inputCol = "category", outputCol = "label")
nlp_pipeline_w2v = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
glove_embeddings,
embeddingsSentence,
embeddings_finisher,
explodeVectors,
label_stringIdx])
nlp_model_w2v = nlp_pipeline_w2v.fit(newsDF)
processed_w2v = nlp_model_w2v.transform(newsDF)
processed_w2v.count()
processed_w2v.select('finished_embeddings').take(1)
processed_w2v.select("finished_embeddings").show(1)
processed_w2v.select('finished_sentence_embeddings').take(1)
# IF SQLTransformer IS NOT USED INSIDE THE PIPELINE, WE CAN EXPLODE OUTSIDE
from pyspark.sql.functions import explode
# processed_w2v= processed_w2v.withColumn("features", explode(processed_w2v.finished_sentence_embeddings))
processed_w2v.select("features").take(1)
processed_w2v.select("features").take(1)
processed_w2v.select('description','features','label').show()
# set seed for reproducibility
(trainingData, testData) = processed_w2v.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
from pyspark.sql.functions import udf
@udf("long")
def num_nonzeros(v):
return v.numNonzeros()
testData = testData.where(num_nonzeros("features") != 0)
testData.count()
lrModel_w2v = lr.fit(trainingData)
predictions_w2v = lrModel_w2v.transform(testData)
predictions_w2v.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
y_true = predictions_w2v.select("label")
y_true = y_true.toPandas()
y_pred = predictions_w2v.select("prediction")
y_pred = y_pred.toPandas()
print(classification_report(y_true.label, y_pred.prediction))
print(accuracy_score(y_true.label, y_pred.prediction))
processed_w2v.select('description','cleanTokens.result').show(truncate=50)
```
## LogReg with Spark NLP Bert Embeddings
```
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
bert_embeddings = BertEmbeddings\
.pretrained('bert_base_cased', 'en') \
.setInputCols(["document",'cleanTokens'])\
.setOutputCol("bert")\
.setCaseSensitive(False)\
.setPoolingLayer(0)
embeddingsSentence = SentenceEmbeddings() \
.setInputCols(["document", "bert"]) \
.setOutputCol("sentence_embeddings") \
.setPoolingStrategy("AVERAGE")
embeddings_finisher = EmbeddingsFinisher() \
.setInputCols(["sentence_embeddings"]) \
.setOutputCols(["finished_sentence_embeddings"]) \
.setOutputAsVector(True)\
.setCleanAnnotations(False)
label_stringIdx = StringIndexer(inputCol = "category", outputCol = "label")
nlp_pipeline_bert = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
bert_embeddings,
embeddingsSentence,
embeddings_finisher,
label_stringIdx])
nlp_model_bert = nlp_pipeline_bert.fit(newsDF)
processed_bert = nlp_model_bert.transform(newsDF)
processed_bert.count()
from pyspark.sql.functions import explode
processed_bert= processed_bert.withColumn("features", explode(processed_bert.finished_sentence_embeddings))
processed_bert.select('description','features','label').show()
# set seed for reproducibility
(trainingData, testData) = processed_bert.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0)
lrModel = lr.fit(trainingData)
from pyspark.sql.functions import udf
@udf("long")
def num_nonzeros(v):
return v.numNonzeros()
testData = testData.where(num_nonzeros("features") != 0)
predictions = lrModel.transform(testData)
predictions.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
import pandas as pd
df = predictions.select('description','category','label','prediction').toPandas()
print(classification_report(df.label, df.prediction))
print(accuracy_score(df.label, df.prediction))
```
## LogReg with ELMO Embeddings
```
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
elmo_embeddings = ElmoEmbeddings.load('/Users/vkocaman/cache_pretrained/elmo_en_2.4.0_2.4_1580488815299')\
.setPoolingLayer("word_emb")\
.setInputCols(["document",'cleanTokens'])\
.setOutputCol("elmo")
embeddingsSentence = SentenceEmbeddings() \
.setInputCols(["document", "elmo"]) \
.setOutputCol("sentence_embeddings") \
.setPoolingStrategy("AVERAGE")
embeddings_finisher = EmbeddingsFinisher() \
.setInputCols(["sentence_embeddings"]) \
.setOutputCols(["finished_sentence_embeddings"]) \
.setOutputAsVector(True)\
.setCleanAnnotations(False)
label_stringIdx = StringIndexer(inputCol = "category", outputCol = "label")
nlp_pipeline_elmo = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
elmo_embeddings,
embeddingsSentence,
embeddings_finisher,
label_stringIdx])
nlp_model_elmo = nlp_pipeline_elmo.fit(newsDF)
processed_elmo = nlp_model_elmo.transform(newsDF)
processed_elmo.count()
(trainingData, testData) = newsDF.randomSplit([0.7, 0.3], seed = 100)
processed_trainingData = nlp_model_elmo.transform(trainingData)
processed_trainingData.count()
processed_testData = nlp_model_elmo.transform(testData)
processed_testData.count()
processed_trainingData.columns
processed_testData= processed_testData.withColumn("features", explode(processed_testData.finished_sentence_embeddings))
processed_trainingData= processed_trainingData.withColumn("features", explode(processed_trainingData.finished_sentence_embeddings))
from pyspark.sql.functions import udf
@udf("long")
def num_nonzeros(v):
return v.numNonzeros()
processed_testData = processed_testData.where(num_nonzeros("features") != 0)
%%time
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0)
lrModel = lr.fit(processed_trainingData)
processed_trainingData.columns
predictions = lrModel.transform(processed_testData)
predictions.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
df = predictions.select('description','category','label','prediction').toPandas()
df.shape
df.head()
from sklearn.metrics import classification_report, accuracy_score
print(classification_report(df.label, df.prediction))
print(accuracy_score(df.label, df.prediction))
```
## LogReg with Universal Sentence Encoder
```
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
useEmbeddings = UniversalSentenceEncoder.load('/Users/vkocaman/cache_pretrained/tfhub_use_en_2.4.0_2.4_1580582893733')\
.setInputCols("document")\
.setOutputCol("use_embeddings")
embeddings_finisher = EmbeddingsFinisher() \
.setInputCols(["use_embeddings"]) \
.setOutputCols(["finished_use_embeddings"]) \
.setOutputAsVector(True)\
.setCleanAnnotations(False)
label_stringIdx = StringIndexer(inputCol = "category", outputCol = "label")
use_pipeline = Pipeline(
stages=[
document_assembler,
useEmbeddings,
embeddings_finisher,
label_stringIdx]
)
use_df = use_pipeline.fit(newsDF).transform(newsDF)
use_df.select('finished_use_embeddings').show(3)
from pyspark.sql.functions import explode
use_df= use_df.withColumn("features", explode(use_df.finished_use_embeddings))
use_df.show(2)
# set seed for reproducibility
(trainingData, testData) = use_df.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
import pandas as pd
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0)
lrModel = lr.fit(trainingData)
predictions = lrModel.transform(testData)
predictions.filter(predictions['prediction'] == 0) \
.select("description","category","probability","label","prediction") \
.orderBy("probability", ascending=False) \
.show(n = 10, truncate = 30)
df = predictions.select('description','category','label','prediction').toPandas()
#df['result'] = df['result'].apply(lambda x: x[0])
df.head()
print(classification_report(df.label, df.prediction))
print(accuracy_score(df.label, df.prediction))
```
### train on entire dataset
```
lr = LogisticRegression(maxIter=20, regParam=0.3, elasticNetParam=0)
lrModel = lr.fit(use_df)
test_df = spark.read.parquet("data/news_category_test.parquet")
test_df = use_pipeline.fit(test_df).transform(test_df)
test_df= test_df.withColumn("features", explode(test_df.finished_use_embeddings))
test_df.show(2)
predictions = lrModel.transform(test_df)
df = predictions.select('description','category','label','prediction').toPandas()
df['label'] = df.category.replace({'World':2.0,
'Sports':3.0,
'Business':0.0,
'Sci/Tech':1.0})
df.head()
print(classification_report(df.label, df.prediction))
print(accuracy_score(df.label, df.prediction))
```
## Spark NLP Licensed DocClassifier
```
from sparknlp_jsl.annotator import *
# set seed for reproducibility
(trainingData, testData) = newsDF.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
stemmer = Stemmer() \
.setInputCols(["cleanTokens"]) \
.setOutputCol("stem")
logreg = DocumentLogRegClassifierApproach()\
.setInputCols(["stem"])\
.setLabelCol("category")\
.setOutputCol("prediction")
nlp_pipeline = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
stemmer,
logreg])
nlp_model = nlp_pipeline.fit(trainingData)
processed = nlp_model.transform(testData)
processed.count()
processed.select('description','category','prediction.result').show(truncate=50)
processed.select('description','prediction.result').show(truncate=50)
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
import pandas as pd
df = processed.select('description','category','prediction.result').toPandas()
df.head()
df.result[0][0]
df = processed.select('description','category','prediction.result').toPandas()
df['result'] = df['result'].apply(lambda x: x[0])
df.head()
df = processed.select('description','category','prediction.result').toPandas()
df['result'] = df['result'].apply(lambda x: x[0])
print(classification_report(df.category, df.result))
print(accuracy_score(df.category, df.result))
```
# ClassifierDL
```
# actual content is inside description column
document = DocumentAssembler()\
.setInputCol("description")\
.setOutputCol("document")
use = UniversalSentenceEncoder.load('/Users/vkocaman/cache_pretrained/tfhub_use_en_2.4.4_2.4_1583158595769')\
.setInputCols(["document"])\
.setOutputCol("sentence_embeddings")
# the classes/labels/categories are in category column
classsifierdl = ClassifierDLApproach()\
.setInputCols(["sentence_embeddings"])\
.setOutputCol("class")\
.setLabelColumn("category")\
.setMaxEpochs(5)\
.setEnableOutputLogs(True)
pipeline = Pipeline(
stages = [
document,
use,
classsifierdl
])
# set seed for reproducibility
(trainingData, testData) = newsDF.randomSplit([0.7, 0.3], seed = 100)
print("Training Dataset Count: " + str(trainingData.count()))
print("Test Dataset Count: " + str(testData.count()))
pipelineModel = pipeline.fit(trainingData)
from sklearn.metrics import classification_report, accuracy_score
df = pipelineModel.transform(testDataset).select('category','description',"class.result").toPandas()
df['result'] = df['result'].apply(lambda x: x[0])
print(classification_report(df.category, df.result))
print(accuracy_score(df.category, df.result))
```
## Loading the trained classifier from disk
```
classsifierdlmodel = ClassifierDLModel.load('classifierDL_model_20200317_5e')
import sparknlp
sparknlp.__path__
.setInputCols(["sentence_embeddings"])\
.setOutputCol("class")\
.setLabelColumn("category")\
.setMaxEpochs(5)\
.setEnableOutputLogs(True)
trainDataset = spark.read \
.option("header", True) \
.csv("data/news_category_train.csv")
trainDataset.count()
trainingData.count()
document = DocumentAssembler()\
.setInputCol("description")\
.setOutputCol("document")
sentence = SentenceDetector()\
.setInputCols(['document'])\
.setOutputCol('sentence')
use = UniversalSentenceEncoder.load('/Users/vkocaman/cache_pretrained/tfhub_use_en_2.4.4_2.4_1583158595769')\
.setInputCols(["sentence"])\
.setOutputCol("sentence_embeddings")
classsifierdlmodel = ClassifierDLModel.load('classifierDL_model_20200317_5e')
pipeline = Pipeline(
stages = [
document,
sentence,
use,
classsifierdlmodel
])
pipeline.fit(testData.limit(1)).transform(testData.limit(10)).select('category','description',"class.result").show(10, truncate=50)
lm = LightPipeline(pipeline.fit(testDataset.limit(1)))
lm.annotate('In its first two years, the UK dedicated card companies have surge')
text='''
Fearing the fate of Italy, the centre-right government has threatened to be merciless with those who flout tough restrictions. As of Wednesday it will also include all shops being closed across Greece, with the exception of supermarkets. Banks, pharmacies, pet-stores, mobile phone stores, opticians, bakers, mini-markets, couriers and food delivery outlets are among the few that will also be allowed to remain open.
'''
lm = LightPipeline(pipeline.fit(testDataset.limit(1)))
lm.annotate(text)
```
# Classifier DL + Glove + Basic text processing
```
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
lemma = LemmatizerModel.pretrained('lemma_antbnc') \
.setInputCols(["token"]) \
.setOutputCol("lemma")
lemma_pipeline = Pipeline(
stages=[document_assembler,
tokenizer,
lemma,
glove_embeddings])
lemma_pipeline.fit(trainingData.limit(1000)).transform(trainingData.limit(1000)).show(truncate=30)
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
lemma = LemmatizerModel.pretrained('lemma_antbnc') \
.setInputCols(["cleanTokens"]) \
.setOutputCol("lemma")
glove_embeddings = WordEmbeddingsModel().pretrained() \
.setInputCols(["document",'lemma'])\
.setOutputCol("embeddings")\
.setCaseSensitive(False)
embeddingsSentence = SentenceEmbeddings() \
.setInputCols(["document", "embeddings"]) \
.setOutputCol("sentence_embeddings") \
.setPoolingStrategy("AVERAGE")
classsifierdl = ClassifierDLApproach()\
.setInputCols(["sentence_embeddings"])\
.setOutputCol("class")\
.setLabelColumn("category")\
.setMaxEpochs(10)\
.setEnableOutputLogs(True)
clf_pipeline = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
lemma,
glove_embeddings,
embeddingsSentence,
classsifierdl])
!rm -rf classifier_dl_pipeline_glove
clf_pipelineModel.save('classifier_dl_pipeline_glove')
clf_pipelineModel = clf_pipeline.fit(trainingData)
df = clf_pipelineModel.transform(testDataset).select('category','description',"class.result").toPandas()
df['result'] = df['result'].apply(lambda x: x[0])
print(classification_report(df.category, df.result))
print(accuracy_score(df.category, df.result))
!cd data && ls -l
import pandas as pd
import
news_df = newsDF.toPandas()
news_df.head()
news_df.to_csv('data/news_dataset.csv', index=False)
document_assembler = DocumentAssembler() \
.setInputCol("description") \
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
normalizer = Normalizer() \
.setInputCols(["token"]) \
.setOutputCol("normalized")
stopwords_cleaner = StopWordsCleaner()\
.setInputCols("normalized")\
.setOutputCol("cleanTokens")\
.setCaseSensitive(False)
lemma = LemmatizerModel.pretrained('lemma_antbnc') \
.setInputCols(["cleanTokens"]) \
.setOutputCol("lemma")
glove_embeddings = WordEmbeddingsModel().pretrained() \
.setInputCols(["document",'lemma'])\
.setOutputCol("embeddings")\
.setCaseSensitive(False)
txt_pipeline = Pipeline(
stages=[document_assembler,
tokenizer,
normalizer,
stopwords_cleaner,
lemma,
glove_embeddings,
embeddingsSentence])
txt_pipelineModel = txt_pipeline.fit(testData.limit(1))
txt_pipelineModel.save('text_prep_pipeline_glove')
df.head()
```
| github_jupyter |
## State of Washington - Licensee - Darling Growers
* UBI: 603513879
We'll be using the [`cannapy`](https://github.com/CannabisData/cannapy) library to access the portal data. `cannapy` aims to provide an abstract interface for accessing and working with *Cannabis* data from around the world. It utilizes [xmunoz](https://github.com/xmunoz)'s [`sodapy`](https://github.com/xmunoz/sodapy) client to access Socrata-based open data portals and can return data loaded into [Pandas DataFrames](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html).
### Dataset: Licensed Businesses
* Canonical Dataset ID: **bhbp-x4eb**
* Detail screen on the WSLCB Portal: https://data.lcb.wa.gov/Licensing/Licensed-Businesses/u3zh-ri66
* Detail screen on Socrata's Open Data Foundry: https://dev.socrata.com/foundry/data.lcb.wa.gov/bhbp-x4eb
```
import time
import cannapy.us.wa.wslcb.portal as wslcb
import pandas as pd
# Specify your own Socrata App Token if you plan to experiment
app_token = 'XaB9MBqc81C3KT4Vps6Wh5LZt'
# Instantiate a cannapy interface to the WSLCB open data portal
portal = wslcb.WSLCBPortal(app_token)
# We'll be using the Licensed Businesses dataset
dataset_id = 'bhbp-x4eb'
# And we're looking for data on a particular licensee
licensee_ubi = '603513879'
# Check when the dataset was last updated
last_updated = portal.dataset_last_updated(dataset_id)
print('Last updated: {}'.format(time.strftime('%c', last_updated)))
# Retrieve the dataset preloaded into a Pandas DataFrame
licenses = portal.get_dataframe(dataset_id)
# The UBI column uniquely identifies each licensee, but obscures ownership of multiple licenses by the same entity.
# Let's break that column apart into its constituent parts:
# Unified Business Identifier (UBI): first nine digits
# Business ID Number: next three digits
# Location Number: last four digits
df_v2 = licenses.rename(columns={'ubi': 'ubi_source'})
df_v2['ubi'] = df_v2.ubi_source.str[0:9]
df_v2['ubi_business_id'] = df_v2.ubi_source.str[9:12]
df_v2['ubi_location'] = df_v2.ubi_source.str[12:]
licensee_licenses = df_v2.loc[df_v2['ubi'] == licensee_ubi]
licensee_licenses
```
### Dataset: Enforcement Visits
* Canonical Dataset ID: **w7wg-8m52**
* Detail screen on the WSLCB Portal: https://data.lcb.wa.gov/dataset/Enforcement-Visits-Dataset/jizx-thwg
* Detail screen on Socrata's Open Data Foundry: https://dev.socrata.com/foundry/data.lcb.wa.gov/w7wg-8m52
```
# Let's see how many enforcement visits the licensee has hosted
dataset_id = 'w7wg-8m52'
# Select the licensee's license number
# TODO: find a way to do this without hardcoding the row number
licensee_license_number = licensee_licenses.loc[753, 'license']
# Check when the dataset was last updated
last_updated = portal.dataset_last_updated(dataset_id)
print('Last updated: {}'.format(time.strftime('%c', last_updated)))
# Retrieve the dataset preloaded into a Pandas DataFrame
enforcement_visits = portal.get_dataframe(dataset_id)
# Suppress the chained assignment warning: https://stackoverflow.com/a/20627316/7622699
pd.options.mode.chained_assignment = None
# Pull aside the enforcement visits by the selected licensee
licensee_enforcement_visits = enforcement_visits.loc[enforcement_visits['license_number'] == licensee_license_number]
# Sort the DataFrame by 'date'
licensee_enforcement_visits.sort_values(by='date', inplace=True)
licensee_enforcement_visits
```
### Dataset: Violations
* Canonical Dataset ID: **dgm4-3cm6**
* Detail screen on the WSLCB Portal: https://data.lcb.wa.gov/dataset/Violations-Dataset/dx3i-tzh2
* Detail screen on Socrata's Open Data Foundry: https://dev.socrata.com/foundry/data.lcb.wa.gov/dgm4-3cm6
```
# Let's pull up all of the licensee's violations
dataset_id = 'dgm4-3cm6'
# Check when the dataset was last updated
last_updated = portal.dataset_last_updated(dataset_id)
print('Last updated: {}'.format(time.strftime('%c', last_updated)))
# Retrieve the dataset preloaded into a Pandas DataFrame
violations = portal.get_dataframe(dataset_id)
# Pull aside the violations by the selected licensee
licensee_violations = violations.loc[violations['license_number'] == licensee_license_number]
# Sort the DataFrame by 'visit_date'
licensee_violations.sort_values(by='visit_date', inplace=True)
licensee_violations
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jaisal1311/Deep-Learning/blob/master/Computer%20Vision/FER/facial_expression_recognizer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# The Notebook file where I am going to create the model and train it
The plan is to create a human visual system like replica for computer to look through camera working as their eyes and understand the mood by reading on facial expression.
I will load the data here, make it ready or the model, then create the model and train it here. After it is done I will save the trained parameters to use in a python script which can understand the facial expression with the help of camera and the OpenCV module.
```
# performing the imports we may need to use later in the notebook
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import torch, os
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader, random_split
from torchvision import transforms, utils
from torchvision.transforms import ToTensor
from torchvision.utils import make_grid
```
## 1. Getting the Data and Understanding it
After looking around the internet I found a dataset in csv format known as FER2013 containing pixel values of the photos and their respective classes.
```
# Let us first load the dataset
df = pd.read_csv('../input/fer2013/fer2013.csv') # This is because I used the notebook to train on kaggle
# df = pd.read_csv('/data/fer2013.csv')
# and then take a look at the csv dataframe
df.head()
```
After taking a look at the data frame we can see clearly that there are 3 columns in the
frame first has the index number of the classes that is 0 for happy face and 1 for sad but we do not know which is what yet so we will categorize it ourselves a little later
The second column contains the pixel values for the photo of the faces with the respective reactions and usage column states which data rows are for testing which are for training.
Let us try to take out someinsights from the dataset
```
print('Total lenth of dataset: ', len(df.Usage), '\n') # Which is actually also the lenth of total dataframe
print('Total Categories we have: ', df.Usage.unique())
print('Number of data in each category: ', '{ Training: ', len(df[df.Usage == 'Training']), ' PublicTest: ', len(df[df.Usage == 'PublicTest']), ' PrivateTest: ', len(df[df.Usage == 'PrivateTest']), '}\n')
print('Total Expression Classes we have: ', df.emotion.unique(), '\n')
print('Type of the pixel data is: ', type(df.pixels[0]))
```
Now let us try to understand which clas of the expression represents which expression ourselves!
```
# since the pixels are in a single long presentation this function will turn it to a matrix of 48x48
def makemtrx(lst, n=48):
for i in range(0, 48*48, n):
yield lst[i:i + n]
# This function will help to show the images
def showimg(data):
pixel = [int(i) for i in data[1].split(' ')]
pixel = np.array(list(makemtrx(pixel)))
plt.imshow(pixel, cmap='gray')
plt.xlabel(f'Expression Class: {data[0]}')
plt.plot()
for i in range(7):
plt.figure()
showimg(df[df.emotion == i].values[0])
```
After looking at the expressionin the photos i can conclude the classes for the expression below.
```
classes = {
0: 'Angry', 1: 'Disgust', 2: 'Fear', 3: 'Happy', 4: 'Sad', 5: 'Surprise', 6: 'Neutral'
}
```
Now let me get the data ready for the model!!
```
# getting the dataset ready
df_train = pd.concat([df[(df.Usage == 'Training')], df[df.Usage == 'PublicTest']], ignore_index=True).drop(['Usage'], axis=1)
df_test = df[df.Usage == 'PrivateTest'].drop(['Usage'], axis=1).reset_index().drop(['index'], 1)
# differentiating between labels and images
train_images = df_train.iloc[:, 1]
train_labels = df_train.iloc[:, 0]
test_images = df_test.iloc[:, 1]
test_labels = df_test.iloc[:, 0]
```
Now we will create a class for the dataset!
```
# this is for the transforms
train_trfm = transforms.Compose(
[
transforms.ToPILImage(),
transforms.Grayscale(num_output_channels=1),
transforms.RandomCrop(48, padding=4, padding_mode='reflect'),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5), inplace=True)
])
val_trfm = transforms.Compose(
[
transforms.ToPILImage(),
transforms.Grayscale(num_output_channels=1),
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))
])
# Creating the class for our dataset for the FER
class FERDataset(Dataset):
def __init__(self, images, labels, transforms):
self.X = images
self.y = labels
self.transforms = transforms
def __len__(self):
return len(self.X)
def __getitem__(self, i):
data = [int(m) for m in self.X[i].split(' ')]
data = np.asarray(data).astype(np.uint8).reshape(48,48,1)
data = self.transforms(data)
label = self.y[i]
return (data, label)
# assigning the transformed data
train_data = FERDataset(train_images, train_labels, train_trfm)
val_data = FERDataset(test_images, test_labels, val_trfm)
```
I'll be using the test data as validation data since the test data is also labeled,that way I get more data.
Getting the data ready is done, now lets just inspect if everything is all right or not.
```
def showimg(data):
img, lbl = data
plt.figure()
plt.imshow(torch.squeeze(img), cmap='gray')
plt.xlabel(f'{classes[lbl]} ({lbl})')
plt.plot()
showimg(train_data[5])
```
Now when the data is ready we will have a validation dataset and dataloader for them with batch sizes
```
random_seed = 42
torch.manual_seed(random_seed);
batch_num = 400
train_dl = DataLoader(train_data, batch_num, shuffle=True, num_workers=4, pin_memory=True)
val_dl = DataLoader(val_data, batch_num*2, num_workers=4, pin_memory=True)
```
Since the dataloader and eevrything is ready now, lets take a look at the photos
```
def show_batch(dl):
for images, labels in dl:
fig, ax = plt.subplots(figsize=(12, 6))
ax.set_xticks([]); ax.set_yticks([])
ax.imshow(make_grid(images, nrow=16).permute(1, 2, 0))
break
show_batch(train_dl)
```
Everything is completed with the data processing! Now we can begin creating our model!!
## Creating The Model
We will craete a base class for the image classification about the expression which will later be encapsuled in the class for the model
```
def accuracy(outputs, labels):
_, preds = torch.max(outputs, dim=1)
return torch.tensor(torch.sum(preds==labels).item()/len(preds))
class FERBase(nn.Module):
# this takes is batch from training dl
def training_step(self, batch):
images, labels = batch
out = self(images) # calls the training model and generates predictions
loss = F.cross_entropy(out, labels) # calculates loss compare to real labels using cross entropy
return loss
# this takes in batch from validation dl
def validation_step(self, batch):
images, labels = batch
out = self(images)
loss = F.cross_entropy(out, labels)
acc = accuracy(out, labels) # calls the accuracy function to measure the accuracy
return {'val_loss': loss.detach(), 'val_acc': acc}
def validation_epoch_end(self, outputs):
batch_losses = [x['val_loss'] for x in outputs]
epoch_loss = torch.stack(batch_losses).mean() # finds out the mean loss of the epoch batch
batch_accs = [x['val_acc'] for x in outputs]
epoch_acc = torch.stack(batch_accs).mean() # finds out the mean acc of the epoch batch
return {'val_loss': epoch_loss.item(), 'val_acc': epoch_acc.item()}
def epoch_end(self, epoch, result):
print("Epoch [{}], last_lr: {:.5f}, train_loss: {:.4f}, val_loss: {:.4f}, val_acc: {:.4f}".format(
epoch, result['lrs'][-1], result['train_loss'], result['val_loss'], result['val_acc']))
```
Now we will create our model for the Facial Expression Recognition
```
def conv_block(in_chnl, out_chnl, pool=False, padding=1):
layers = [
nn.Conv2d(in_chnl, out_chnl, kernel_size=3, padding=padding),
nn.BatchNorm2d(out_chnl),
nn.ReLU(inplace=True)]
if pool: layers.append(nn.MaxPool2d(2))
return nn.Sequential(*layers)
class FERModel(FERBase):
def __init__(self, in_chnls, num_cls):
super().__init__()
self.conv1 = conv_block(in_chnls, 64, pool=True) # 64x24x24
self.conv2 = conv_block(64, 128, pool=True) # 128x12x12
self.resnet1 = nn.Sequential(conv_block(128, 128), conv_block(128, 128)) # Resnet layer 1: includes 2 conv2d
self.conv3 = conv_block(128, 256, pool=True) # 256x6x6
self.conv4 = conv_block(256, 512, pool=True) # 512x3x3
self.resnet2 = nn.Sequential(conv_block(512, 512), conv_block(512, 512)) # Resnet layer 2: includes 2 conv2d
self.classifier = nn.Sequential(nn.MaxPool2d(3),
nn.Flatten(),
nn.Linear(512, num_cls)) # num_cls
def forward(self, xb):
out = self.conv1(xb)
out = self.conv2(out)
out = self.resnet1(out) + out
out = self.conv3(out)
out = self.conv4(out)
out = self.resnet2(out) + out
return self.classifier(out)
model = FERModel(1, 7)
```
Let us nowlook if the model gives us the output array as I am hoping for. The output array must contain 7 values, that is 7 probabilities of what class could it be from the expression classes.
```
for images, lbl in train_dl:
print('shape of image: ', images.shape)
out = model(images)
print('shape of output: ', out.shape)
print('Output: ', out[0])
break
```
Now let us code for to utilise the GPU for the training!
```
def get_default_device():
"""Pick GPU if available, else CPU"""
if torch.cuda.is_available():
return torch.device('cuda')
else:
return torch.device('cpu')
def to_device(data, device):
"""Move tensor(s) to chosen device"""
if isinstance(data, (list,tuple)):
return [to_device(x, device) for x in data]
return data.to(device, non_blocking=True)
class DeviceDataLoader():
"""Wrap a dataloader to move data to a device"""
def __init__(self, dl, device):
self.dl = dl
self.device = device
def __iter__(self):
"""Yield a batch of data after moving it to device"""
for b in self.dl:
yield to_device(b, self.device)
def __len__(self):
"""Number of batches"""
return len(self.dl)
device = get_default_device()
device
```
Now let us wrap our datas to the available device
```
train_dl = DeviceDataLoader(train_dl, device)
val_dl = DeviceDataLoader(val_dl, device)
to_device(model, device);
```
## Training The Model
Since the model itself is ready, there is not any more headache left to create some functions for the training start training.
```
@torch.no_grad() # this is for stopping the model from keeping track ofold parameters
def evaluate(model, val_loader):
# This function will evaluate the model and give back the val acc and loss
model.eval()
outputs = [model.validation_step(batch) for batch in val_loader]
return model.validation_epoch_end(outputs)
# getting the current learning rate
def get_lr(optimizer):
for param_group in optimizer.param_groups:
return param_group['lr']
# this fit function follows the intuition of 1cycle lr
def fit(epochs, max_lr, model, train_loader=train_dl, val_loader=val_dl, weight_decay=0, grad_clip=None, opt_func=torch.optim.Adam):
torch.cuda.empty_cache()
history = [] #keep track of the evaluation results
# setting upcustom optimizer including weight decay
optimizer = opt_func(model.parameters(), max_lr, weight_decay=weight_decay)
# setting up 1cycle lr scheduler
sched = torch.optim.lr_scheduler.OneCycleLR(optimizer, max_lr, epochs=epochs, steps_per_epoch=len(train_loader))
for epoch in range(epochs):
# training
model.train()
train_losses = []
lrs = []
for batch in train_loader:
loss = model.training_step(batch)
train_losses.append(loss)
loss.backward()
# gradient clipping
if grad_clip:
nn.utils.clip_grad_value_(model.parameters(), grad_clip)
optimizer.step()
optimizer.zero_grad()
# record the lr
lrs.append(get_lr(optimizer))
sched.step()
#validation
result = evaluate(model, val_loader)
result['train_loss'] = torch.stack(train_losses).mean().item()
result['lrs'] = lrs
model.epoch_end(epoch, result)
history.append(result)
return history
```
We also need to load our model in the device too
```
model = to_device(FERModel(1, 7), device)
evaluate(model, val_dl)
```
Now we will train
```
max_lr = 0.001
grad_clip = 0.1
weight_decay = 1e-4
%%time
history = fit(30, max_lr, model, weight_decay=weight_decay, grad_clip=grad_clip)
def plot_losses(history):
train_losses = [x.get('train_loss') for x in history]
val_losses = [x['val_loss'] for x in history]
plt.plot(train_losses, '-bx')
plt.plot(val_losses, '-rx')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['Training', 'Validation'])
plt.title('Loss vs. No. of epochs');
def plot_lrs(history):
lrs = np.concatenate([x.get('lrs', []) for x in history])
plt.plot(lrs)
plt.xlabel('Batch no.')
plt.ylabel('Learning rate')
plt.title('Learning Rate vs. Batch no.');
plot_losses(history)
plt.figure()
plot_lrs(history)
```
**History log**
* model1: 1>16>32>64>128>256>256 cnn then, 256*6*6>1024>512>7 fnn, 10epoch 0.001lr, 3epoch 0.0001lr, 55%, overfit
* model2: 1>64>128>256>512>1024 cnn then 1024*6*6>1024>512>128>7 fnn, 10epoch, 0.0001lr, 56%, minimum overfitting
* model3: 1>128>256>600>1200>2048>1024 cnn then 1024*3*3>1024>512>7 fnn, 5epochs 1e-4lr, 6epochs 1e-5lr,5epochs, 1e-6lr, 58%, minimum overfit
* resnet9: maxlr 0.001, epochs 20-30, 69%, minimum overfit
* resnet9: maxlr 0.01, epoch 20, 71%
* resnet9: maxlr 0.001, 30 epcohs, 72%
## Saving the Model
Now since the training is done we can succesfully save our model so that we can use it in our facial recognition script!
```
torch.save(model.state_dict(), 'FER2013-Resnet9.pth')
!pip install jovian
import jovian
jovian.commit(project='facial-expression-recognizer')
```
| github_jupyter |
```
pip install mljar-supervised
pip install openml
from sklearn.model_selection import train_test_split
import pandas as pd
df = pd.read_csv("undersample_2.csv")
X = df.iloc[:,2:32]
y = df.iloc[:,1]
from sklearn.preprocessing import LabelEncoder
l = LabelEncoder()
y = l.fit_transform(y)
X_train, X_tmp, y_train, y_tmp = train_test_split(X, y, test_size=0.2, random_state=42)
X_test, X_valid, y_test, y_valid = train_test_split(X_tmp, y_tmp, test_size=0.2, random_state=42)
from supervised.automl import AutoML
import os
a_ens_3 = AutoML(total_time_limit=10000,mode="Perform",
explain_level=2, algorithms = ['Decision Tree', 'Random Forest','Neural Network', 'Nearest Neighbors','Xgboost'],
validation_strategy={"validation_type": "kfold","k_folds": 5, "shuffle": True, "stratify": True,
"random_seed": 1230})
from sklearn.ensemble import RandomForestClassifier
from supervised.automl import AutoML
import os
import numpy as np
import pandas as pd
import sklearn.model_selection
from sklearn.metrics import log_loss, f1_score
from sklearn.datasets import make_gaussian_quantiles
from sklearn.ensemble import AdaBoostClassifier
from sklearn.metrics import accuracy_score
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import confusion_matrix
import numpy as np
np.random.seed(1337)
# for reproducibility import h5pyfromkeras.modelsimportSequential
from keras.optimizers import Adam
from keras.initializers import TruncatedNormal
from keras.layers import Input, Dense, Dropout, Flatten, Conv2D, MaxPooling2D
from keras.callbacks import ReduceLROnPlateau
from sklearn.metrics import roc_curve, auc
a_ens_3.fit(X,y)
predictions= a_ens_3.predict(X_test)
predictions
pip install plot-metric
import matplotlib.pyplot as plt
from plot_metric.functions import BinaryClassification
from sklearn.metrics import confusion_matrix
# Visualisation with plot_metric
bc = BinaryClassification(predictions, y_test, labels=["Class 1", "Class 2"])
print(bc)# Figures
plt.figure(figsize=(5,5))
bc.plot_roc_curve()
plt.show()
tn, fp, fn, tp=confusion_matrix(y_test,predictions).ravel()
accuracy=(tn+tp)/(tn+tp+fn+fp)
prec=(tp)/(tp+fp)
rec=tp/(tp+fn)
F1=2*(prec*rec)/(prec+rec)
confusion_matrix(y_test, predictions)
print("F1 Score: ",format(F1))
from sklearn.metrics import roc_auc_score
from sklearn.metrics import precision_recall_curve
from matplotlib import pyplot
print("ROC:")
roc_auc=roc_auc_score(y_test, predictions)
print(roc_auc)
!zip -r /content/AutoML_US2.zip /content/AutoML_US2/
from google.colab import files
files.download("/content/AutoML_US2.zip")
```
| github_jupyter |
```
import numpy as np
from matplotlib import pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
from torch.optim import lr_scheduler
```
## R, 真实数据
- 这是生成真实的数据, 被用于模仿
```
def get_distribution_sampler(mu, sigma, batchSize, FeatureNum):
"""
Generate Target Data, Gaussian
Input
- mu: 均值
- sugma: 方差
Output
"""
return Variable(torch.Tensor(np.random.normal(mu, sigma, (batchSize, FeatureNum))))
data_mean = 4
data_stddev = 1.25
batch_size = 1
featureNum = 500
d_real_data = get_distribution_sampler(data_mean, data_stddev, batch_size, featureNum)
# ----------------------
# 计算每个范围的数据个数
# ----------------------
binRange = np.arange(0,8,0.5)
hist1,_ = np.histogram(d_real_data.numpy(), bins=binRange)
# --------
# 绘制图像
# --------
fig, ax1 = plt.subplots()
fig.set_size_inches(20, 10)
plt.set_cmap('RdBu')
x = np.arange(len(binRange)-1)
w=0.3
# 绘制多个bar在同一个图中, 这里需要控制width
plt.bar(x, hist1, width=w*3, align='center')
# 设置坐标轴的标签
ax1.yaxis.set_tick_params(labelsize=15) # 设置y轴的字体的大小
ax1.set_xticks(x) # 设置xticks出现的位置
# 创建xticks
xticksName = []
for i in range(len(binRange)-1):
xticksName = xticksName + ['{}<x<{}'.format(str(np.round(binRange[i],1)), str(np.round(binRange[i+1],1)))]
ax1.set_xticklabels(xticksName)
# 设置坐标轴名称
ax1.set_ylabel("Count", fontsize='xx-large')
plt.show()
```
## I, Input Data
- 这是输出到模型中的数据
```
def get_generator_input_sampler(m, n):
"""
Uniform-dist data into generator, _NOT_ Gaussian
Input
- m: 表示batchsize
- n: 表示feature count
Output
- 返回的是生成数据的分布
"""
return torch.rand(m, n)
g_input_size = 3
minibatch_size = 2
d_gen_input = get_generator_input_sampler(minibatch_size, g_input_size)
print(d_gen_input)
```
## G, generator
- generator是一个普通的前向传播网络
- generator使用的是tanh激活函数
- generator输入是均值分布(来自I的数据)
- generator的任务是模仿R中的数据, 即希望模仿正态分布的数据(generator是没有见过R的)
```
class Generator(nn.Module):
def __init__(self, input_size, hidden_size, output_size, f):
super(Generator, self).__init__()
self.map1 = nn.Linear(input_size, hidden_size)
self.map2 = nn.Linear(hidden_size, hidden_size)
self.map3 = nn.Linear(hidden_size, output_size)
self.relu = nn.ReLU()
self.f = f # 激活函数
def forward(self, x):
x = self.map1(x)
x = self.relu(x)
x = self.map2(x)
x = self.relu(x)
x = self.map3(x)
return x
# 测试generator
g_input_size = 1 # Random noise dimension coming into generator, per output vector
g_hidden_size = 5 # Generator complexity
g_output_size = 1 # Size of generated output vector
generator_activation_function = torch.tanh
G = Generator(input_size=g_input_size,
hidden_size=g_hidden_size,
output_size=g_output_size,
f=generator_activation_function)
G(torch.tensor([[0.5], [0.4]]))
```
## D, discriminator
- discriminator和generator一样, 也是简单的前向传播网络
- discriminator使用的是sigmoid函数
- discriminator使用R和G产生的数据
- 最后的label是0与1, 从R中的数据label是1, 从G中产生的数据label是0
```
class Discriminator(nn.Module):
def __init__(self, input_size, hidden_size, output_size, f):
super(Discriminator, self).__init__()
self.map1 = nn.Linear(input_size, hidden_size)
self.map2 = nn.Linear(hidden_size, hidden_size)
self.map3 = nn.Linear(hidden_size, output_size)
self.relu = nn.ReLU()
self.f = f
def forward(self, x):
x = self.relu(self.map1(x))
x = self.relu(self.map2(x))
x = self.f(self.map3(x))# 最后生成的是概率
return x
d_input_size = 1 # Minibatch size - cardinality of distributions
d_hidden_size = 10 # Discriminator complexity
d_output_size = 1 # Single dimension for 'real' vs. 'fake' classification
discriminator_activation_function = torch.sigmoid
D = Discriminator(input_size=d_input_size,
hidden_size=d_hidden_size,
output_size=d_output_size,
f=discriminator_activation_function)
D(torch.tensor([0.5]))
```
## 辅助函数
```
def get_moments(ds):
"""
- Return the first 4 moments of the data provided
- 返回一个数据的四个指标, 分别是均值, 方差, 偏度, 峰读
- 我们希望通过这四个指标, 来判断我们生成的数据是否是需要的数据
"""
finals = []
for d in ds:
mean = torch.mean(d) # d的均值
diffs = d - mean
var = torch.mean(torch.pow(diffs, 2.0))
std = torch.pow(var, 0.5) # d的方差
zscores = diffs / (std+0.001) # 对原始数据 zscores = (d-mean)/std
skews = torch.mean(torch.pow(zscores, 3.0)) # 峰度
kurtoses = torch.mean(torch.pow(zscores, 4.0)) - 3.0 # excess kurtosis, should be 0 for Gaussian
final = torch.cat((mean.reshape(1,), std.reshape(1,), skews.reshape(1,), kurtoses.reshape(1,)))
# 这里返回的是高斯分布的四个特征
finals.append(final)
return torch.stack(finals)
a = get_moments(d_real_data)
a
```
## Training
- 首先训练D, 我们使用real data vs. fake data, with accurate labels
- 接着我们训练G来使得其生成的数据无法被D正确分辨(这时候fix住D的参数, 使得G产生数据, 使得D预测的label是1)
- 接着重复上面的两个操作
```
d_input_size = 4
d_hidden_size = 10
d_output_size = 1
discriminator_activation_function = torch.sigmoid
g_input_size = 50
g_output_size = 200
g_output_size = 500
generator_activation_function = torch.tanh
featureNum = g_output_size # 一组样本有500个服从正太分布的数据
minibatch_size = 10 # batch_size的大小
num_epochs = 2001
d_steps = 20 # discriminator的训练轮数
g_steps = 20 # generator的训练轮数
# ----------
# 初始化网络
# ----------
D = Discriminator(input_size=d_input_size,
hidden_size=d_hidden_size,
output_size=d_output_size,
f=discriminator_activation_function)
G = Generator(input_size=g_input_size,
hidden_size=g_hidden_size,
output_size=g_output_size,
f=generator_activation_function)
# ----------------------
# 初始化优化器和损失函数
# ----------------------
d_learning_rate = 0.0001
g_learning_rate = 0.0001
criterion = nn.BCELoss() # Binary cross entropy: http://pytorch.org/docs/nn.html#bceloss
d_optimizer = optim.Adam(D.parameters(), lr=d_learning_rate)
g_optimizer = optim.Adam(G.parameters(), lr=g_learning_rate)
d_exp_lr_scheduler = optim.lr_scheduler.CosineAnnealingLR(d_optimizer, T_max = d_steps*5, eta_min=0.00001)
g_exp_lr_scheduler = optim.lr_scheduler.CosineAnnealingLR(g_optimizer, T_max = g_steps*5, eta_min=0.00001)
G_mean = [] # 生成器生成的数据的均值
G_std = [] # 生成器生成的数据的方差
for epoch in range(num_epochs):
# -------------------
# Train the Detective
# -------------------
for d_index in range(d_steps):
# Train D on real+fake
d_exp_lr_scheduler.step()
D.zero_grad()
# Train D on real, 这里的label是1
d_real_data = get_distribution_sampler(data_mean, data_stddev, minibatch_size, featureNum) # 真实的样本
d_real_decision = D(get_moments(d_real_data)) # 求出数据的四个重要特征
d_real_error = criterion(d_real_decision, Variable(torch.ones([minibatch_size, 1]))) # 计算error
d_real_error.backward() # 进行反向传播
# Train D on fake, 这里的label是0
d_gen_input = get_generator_input_sampler(minibatch_size, g_input_size)
d_fake_data = G(d_gen_input)
d_fake_decision = D(get_moments(d_fake_data))
d_fake_error = criterion(d_fake_decision, Variable(torch.zeros([minibatch_size, 1])))
d_fake_error.backward()
# Optimizer
d_optimizer.step()
# -------------------
# Train the Generator
# -------------------
for g_index in range(g_steps):
# Train G on D's response(使得G生成的x让D判断为1)
g_exp_lr_scheduler.step()
G.zero_grad()
gen_input = get_generator_input_sampler(minibatch_size, g_input_size)
g_fake_data = G(gen_input) # 使得generator生成样本
dg_fake_decision = D(get_moments(g_fake_data)) # D来做的判断
g_error = criterion(dg_fake_decision, Variable(torch.ones([minibatch_size, 1])))
G_mean.append(g_fake_data.mean().item())
G_std.append(g_fake_data.std().item())
g_error.backward()
g_optimizer.step()
if epoch%10==0:
print("Epoch: {}, G data's Mean: {}, G data's Std: {}".format(epoch, G_mean[-1], G_std[-1]))
print("Epoch: {}, Real data's Mean: {}, Real data's Std: {}".format(epoch, d_real_data.mean().item(), d_real_data.std().item()))
print('-'*10)
```
## 最终的结果分析
- 绘制出G产生数据的分布
- 绘制出mean的变化曲线
- 绘制出std的变化曲线
```
# ----------------------
# 计算每个范围的数据个数
# ----------------------
binRange = np.arange(0,8,0.5)
hist1,_ = np.histogram(g_fake_data.squeeze().detach().numpy(), bins=binRange)
# --------
# 绘制图像
# --------
fig, ax1 = plt.subplots()
fig.set_size_inches(20, 10)
plt.set_cmap('RdBu')
x = np.arange(len(binRange)-1)
w=0.3
# 绘制多个bar在同一个图中, 这里需要控制width
plt.bar(x, hist1, width=w*3, align='center')
# 设置坐标轴的标签
ax1.yaxis.set_tick_params(labelsize=15) # 设置y轴的字体的大小
ax1.set_xticks(x) # 设置xticks出现的位置
# 创建xticks
xticksName = []
for i in range(len(binRange)-1):
xticksName = xticksName + ['{}<x<{}'.format(str(np.round(binRange[i],1)), str(np.round(binRange[i+1],1)))]
ax1.set_xticklabels(xticksName)
# 设置坐标轴名称
ax1.set_ylabel("Count", fontsize='xx-large')
plt.show()
fig, ax1 = plt.subplots()
fig.set_size_inches(20, 10)
ax1.plot(G_mean)
plt.show()
fig, ax1 = plt.subplots()
fig.set_size_inches(20, 10)
ax1.plot(G_std)
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
# Sample plots in Matplotlib
Here you'll find a host of example plots with the code that
generated them.
Line Plot
=========
Here's how to create a line plot with text labels using
:func:`~matplotlib.pyplot.plot`.
.. figure:: ../../gallery/lines_bars_and_markers/images/sphx_glr_simple_plot_001.png
:target: ../../gallery/lines_bars_and_markers/simple_plot.html
:align: center
:scale: 50
Simple Plot
Multiple subplots in one figure
===============================
Multiple axes (i.e. subplots) are created with the
:func:`~matplotlib.pyplot.subplot` function:
.. figure:: ../../gallery/subplots_axes_and_figures/images/sphx_glr_subplot_001.png
:target: ../../gallery/subplots_axes_and_figures/subplot.html
:align: center
:scale: 50
Subplot
Images
======
Matplotlib can display images (assuming equally spaced
horizontal dimensions) using the :func:`~matplotlib.pyplot.imshow` function.
.. figure:: ../../gallery/images_contours_and_fields/images/sphx_glr_image_demo_003.png
:target: ../../gallery/images_contours_and_fields/image_demo.html
:align: center
:scale: 50
Example of using :func:`~matplotlib.pyplot.imshow` to display a CT scan
Contouring and pseudocolor
==========================
The :func:`~matplotlib.pyplot.pcolormesh` function can make a colored
representation of a two-dimensional array, even if the horizontal dimensions
are unevenly spaced. The
:func:`~matplotlib.pyplot.contour` function is another way to represent
the same data:
.. figure:: ../../gallery/images_contours_and_fields/images/sphx_glr_pcolormesh_levels_001.png
:target: ../../gallery/images_contours_and_fields/pcolormesh_levels.html
:align: center
:scale: 50
Example comparing :func:`~matplotlib.pyplot.pcolormesh` and :func:`~matplotlib.pyplot.contour` for plotting two-dimensional data
Histograms
==========
The :func:`~matplotlib.pyplot.hist` function automatically generates
histograms and returns the bin counts or probabilities:
.. figure:: ../../gallery/statistics/images/sphx_glr_histogram_features_001.png
:target: ../../gallery/statistics/histogram_features.html
:align: center
:scale: 50
Histogram Features
Paths
=====
You can add arbitrary paths in Matplotlib using the
:mod:`matplotlib.path` module:
.. figure:: ../../gallery/shapes_and_collections/images/sphx_glr_path_patch_001.png
:target: ../../gallery/shapes_and_collections/path_patch.html
:align: center
:scale: 50
Path Patch
Three-dimensional plotting
==========================
The mplot3d toolkit (see `toolkit_mplot3d-tutorial` and
`mplot3d-examples-index`) has support for simple 3d graphs
including surface, wireframe, scatter, and bar charts.
.. figure:: ../../gallery/mplot3d/images/sphx_glr_surface3d_001.png
:target: ../../gallery/mplot3d/surface3d.html
:align: center
:scale: 50
Surface3d
Thanks to John Porter, Jonathon Taylor, Reinier Heeres, and Ben Root for
the `.mplot3d` toolkit. This toolkit is included with all standard Matplotlib
installs.
Streamplot
==========
The :meth:`~matplotlib.pyplot.streamplot` function plots the streamlines of
a vector field. In addition to simply plotting the streamlines, it allows you
to map the colors and/or line widths of streamlines to a separate parameter,
such as the speed or local intensity of the vector field.
.. figure:: ../../gallery/images_contours_and_fields/images/sphx_glr_plot_streamplot_001.png
:target: ../../gallery/images_contours_and_fields/plot_streamplot.html
:align: center
:scale: 50
Streamplot with various plotting options.
This feature complements the :meth:`~matplotlib.pyplot.quiver` function for
plotting vector fields. Thanks to Tom Flannaghan and Tony Yu for adding the
streamplot function.
Ellipses
========
In support of the `Phoenix <http://www.jpl.nasa.gov/news/phoenix/main.php>`_
mission to Mars (which used Matplotlib to display ground tracking of
spacecraft), Michael Droettboom built on work by Charlie Moad to provide
an extremely accurate 8-spline approximation to elliptical arcs (see
:class:`~matplotlib.patches.Arc`), which are insensitive to zoom level.
.. figure:: ../../gallery/shapes_and_collections/images/sphx_glr_ellipse_demo_001.png
:target: ../../gallery/shapes_and_collections/ellipse_demo.html
:align: center
:scale: 50
Ellipse Demo
Bar charts
==========
Use the :func:`~matplotlib.pyplot.bar` function to make bar charts, which
includes customizations such as error bars:
.. figure:: ../../gallery/statistics/images/sphx_glr_barchart_demo_001.png
:target: ../../gallery/statistics/barchart_demo.html
:align: center
:scale: 50
Barchart Demo
You can also create stacked bars
(`bar_stacked.py <../../gallery/lines_bars_and_markers/bar_stacked.html>`_),
or horizontal bar charts
(`barh.py <../../gallery/lines_bars_and_markers/barh.html>`_).
Pie charts
==========
The :func:`~matplotlib.pyplot.pie` function allows you to create pie
charts. Optional features include auto-labeling the percentage of area,
exploding one or more wedges from the center of the pie, and a shadow effect.
Take a close look at the attached code, which generates this figure in just
a few lines of code.
.. figure:: ../../gallery/pie_and_polar_charts/images/sphx_glr_pie_features_001.png
:target: ../../gallery/pie_and_polar_charts/pie_features.html
:align: center
:scale: 50
Pie Features
Tables
======
The :func:`~matplotlib.pyplot.table` function adds a text table
to an axes.
.. figure:: ../../gallery/misc/images/sphx_glr_table_demo_001.png
:target: ../../gallery/misc/table_demo.html
:align: center
:scale: 50
Table Demo
Scatter plots
=============
The :func:`~matplotlib.pyplot.scatter` function makes a scatter plot
with (optional) size and color arguments. This example plots changes
in Google's stock price, with marker sizes reflecting the
trading volume and colors varying with time. Here, the
alpha attribute is used to make semitransparent circle markers.
.. figure:: ../../gallery/lines_bars_and_markers/images/sphx_glr_scatter_demo2_001.png
:target: ../../gallery/lines_bars_and_markers/scatter_demo2.html
:align: center
:scale: 50
Scatter Demo2
GUI widgets
===========
Matplotlib has basic GUI widgets that are independent of the graphical
user interface you are using, allowing you to write cross GUI figures
and widgets. See :mod:`matplotlib.widgets` and the
`widget examples <../../gallery/index.html>`_.
.. figure:: ../../gallery/widgets/images/sphx_glr_slider_demo_001.png
:target: ../../gallery/widgets/slider_demo.html
:align: center
:scale: 50
Slider and radio-button GUI.
Filled curves
=============
The :func:`~matplotlib.pyplot.fill` function lets you
plot filled curves and polygons:
.. figure:: ../../gallery/lines_bars_and_markers/images/sphx_glr_fill_001.png
:target: ../../gallery/lines_bars_and_markers/fill.html
:align: center
:scale: 50
Fill
Thanks to Andrew Straw for adding this function.
Date handling
=============
You can plot timeseries data with major and minor ticks and custom
tick formatters for both.
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_date_001.png
:target: ../../gallery/text_labels_and_annotations/date.html
:align: center
:scale: 50
Date
See :mod:`matplotlib.ticker` and :mod:`matplotlib.dates` for details and usage.
Log plots
=========
The :func:`~matplotlib.pyplot.semilogx`,
:func:`~matplotlib.pyplot.semilogy` and
:func:`~matplotlib.pyplot.loglog` functions simplify the creation of
logarithmic plots.
.. figure:: ../../gallery/scales/images/sphx_glr_log_demo_001.png
:target: ../../gallery/scales/log_demo.html
:align: center
:scale: 50
Log Demo
Thanks to Andrew Straw, Darren Dale and Gregory Lielens for contributions
log-scaling infrastructure.
Polar plots
===========
The :func:`~matplotlib.pyplot.polar` function generates polar plots.
.. figure:: ../../gallery/pie_and_polar_charts/images/sphx_glr_polar_demo_001.png
:target: ../../gallery/pie_and_polar_charts/polar_demo.html
:align: center
:scale: 50
Polar Demo
Legends
=======
The :func:`~matplotlib.pyplot.legend` function automatically
generates figure legends, with MATLAB-compatible legend-placement
functions.
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_legend_001.png
:target: ../../gallery/text_labels_and_annotations/legend.html
:align: center
:scale: 50
Legend
Thanks to Charles Twardy for input on the legend function.
TeX-notation for text objects
=============================
Below is a sampling of the many TeX expressions now supported by Matplotlib's
internal mathtext engine. The mathtext module provides TeX style mathematical
expressions using `FreeType <https://www.freetype.org/>`_
and the DejaVu, BaKoMa computer modern, or `STIX <http://www.stixfonts.org>`_
fonts. See the :mod:`matplotlib.mathtext` module for additional details.
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_mathtext_examples_001.png
:target: ../../gallery/text_labels_and_annotations/mathtext_examples.html
:align: center
:scale: 50
Mathtext Examples
Matplotlib's mathtext infrastructure is an independent implementation and
does not require TeX or any external packages installed on your computer. See
the tutorial at :doc:`/tutorials/text/mathtext`.
Native TeX rendering
====================
Although Matplotlib's internal math rendering engine is quite
powerful, sometimes you need TeX. Matplotlib supports external TeX
rendering of strings with the *usetex* option.
.. figure:: ../../gallery/text_labels_and_annotations/images/sphx_glr_tex_demo_001.png
:target: ../../gallery/text_labels_and_annotations/tex_demo.html
:align: center
:scale: 50
Tex Demo
EEG GUI
=======
You can embed Matplotlib into pygtk, wx, Tk, or Qt applications.
Here is a screenshot of an EEG viewer called `pbrain
<https://github.com/nipy/pbrain>`__.

The lower axes uses :func:`~matplotlib.pyplot.specgram`
to plot the spectrogram of one of the EEG channels.
For examples of how to embed Matplotlib in different toolkits, see:
* :doc:`/gallery/user_interfaces/embedding_in_gtk3_sgskip`
* :doc:`/gallery/user_interfaces/embedding_in_wx2_sgskip`
* :doc:`/gallery/user_interfaces/mpl_with_glade3_sgskip`
* :doc:`/gallery/user_interfaces/embedding_in_qt_sgskip`
* :doc:`/gallery/user_interfaces/embedding_in_tk_sgskip`
XKCD-style sketch plots
=======================
Just for fun, Matplotlib supports plotting in the style of `xkcd
<https://www.xkcd.com/>`_.
.. figure:: ../../gallery/showcase/images/sphx_glr_xkcd_001.png
:target: ../../gallery/showcase/xkcd.html
:align: center
:scale: 50
xkcd
Subplot example
===============
Many plot types can be combined in one figure to create
powerful and flexible representations of data.
```
import matplotlib.pyplot as plt
import numpy as np
np.random.seed(19680801)
data = np.random.randn(2, 100)
fig, axs = plt.subplots(2, 2, figsize=(5, 5))
axs[0, 0].hist(data[0])
axs[1, 0].scatter(data[0], data[1])
axs[0, 1].plot(data[0], data[1])
axs[1, 1].hist2d(data[0], data[1])
plt.show()
```
| github_jupyter |
---
description: Few words about PyTorch Datasets
---
# Preamble: PyTorch Datasets
This short preamble will briefly go through the basic notions of Dataset offered natively by PyTorch. A solid grasp of these notions are needed to understand:
1. How PyTorch data loading works in general
2. How AvalancheDatasets differs from PyTorch Datasets
## 📚 Dataset: general definition
In PyTorch, **a `Dataset` is a class** exposing two methods:
- `__len__()`, which returns the amount of instances in the dataset (as an `int`).
- `__getitem__(idx)`, which returns the data point at index `idx`.
In other words, a Dataset instance is just an object for which, similarly to a list, one can simply:
- Obtain its length using the Python `len(dataset)` function.
- Obtain a single data point using the `x, y = dataset[idx]` syntax.
The content of the dataset can be either loaded in memory when the dataset is instantiated (like the torchvision MNIST dataset does) or, for big datasets like ImageNet, the content is kept on disk, with the dataset keeping the list of files in an internal field. In this case, data is loaded from the storage on-the-fly when `__getitem__(idx)` is called. The way those things are managed is specific to each dataset implementation.
## PyTorch Datasets
The PyTorch library offers 4 Dataset implementations:
- `Dataset`: an interface defining the `__len__` and `__getitem__` methods.
- `TensorDataset`: instantiated by passing X and Y tensors. Each row of the X and Y tensors is interpreted as a data point. The `__getitem__(idx)` method will simply return the `idx`-th row of X and Y tensors.
- `ConcatDataset`: instantiated by passing a list of datasets. The resulting dataset is a concatenation of those datasets.
- `Subset`: instantiated by passing a dataset and a list of indices. The resulting dataset will only contain the data points described by that list of indices.
As explained in the mini *How-To*s, Avalanche offers a customized version for all these 4 datasets.
## Transformations
Most datasets from the *torchvision* libraries (as well as datasets found "in the wild") allow for a `transformation` function to be passed to the dataset constructor. The support for transformations is not mandatory for a dataset, but it is quite common to support them. The transformation is used to process the X value of a data point before returning it. This is used to normalize values, apply augmentations, etcetera.
As explained in the mini *How-To*s, the `AvalancheDataset` class implements a very rich and powerful set of functionalities for managing transformations.
## Quick note on the IterableDataset class
A variation of the standard `Dataset` exist in PyTorch: the [IterableDataset](https://pytorch.org/docs/stable/data.html#iterable-style-datasets). When using an `IterableDataset`, one can load the data points in a sequential way only (by using a tape-alike approach). The `dataset[idx]` syntax and `len(dataset)` function are not allowed. **Avalanche does NOT support `IterableDataset`s.** You shouldn't worry about this because, realistically, you will never encounter such datasets.
## DataLoader
The `Dataset` is a very simple object that only returns one data point given its index. In order to create minibatches and speed-up the data loading process, a `DataLoader` is required.
The PyTorch `DataLoader` class is a very efficient mechanism that, given a `Dataset`, will return **minibatches** by optonally **shuffling** data brefore each epoch and by **loading data in parallel** by using multiple workers.
## Preamble wrap-up
To wrap-up, let's see how the native, *non-Avalanche*, PyTorch components work in practice. In the following code we create a `TensorDataset` and then we load it in minibatches using a `DataLoader`.
```
import torch
from torch.utils.data.dataset import TensorDataset
from torch.utils.data.dataloader import DataLoader
# Create a dataset of 100 data points described by 22 features + 1 class label
x_data = torch.rand(100, 22)
y_data = torch.randint(0, 5, (100,))
# Create the Dataset
my_dataset = TensorDataset(x_data, y_data)
# Create the DataLoader
my_dataloader = DataLoader(my_dataset, batch_size=10, shuffle=True, num_workers=4)
# Run one epoch
for x_minibatch, y_minibatch in my_dataloader:
print('Loaded minibatch of', len(x_minibatch), 'instances')
# Output: "Loaded minibatch of 10 instances" x10 times
```
## Next steps
With these notions in mind, you can start start your journey on understanding the functionalities offered by the AvalancheDatasets by going through the *Mini How-To*s.
Please refer to the [list of the *Mini How-To*s regarding AvalancheDatasets](https://avalanche.continualai.org/how-tos/avalanchedataset) for a complete list. It is recommended to start with the **"Creating AvalancheDatasets"** *Mini How-To*.
## 🤝 Run it on Google Colab
You can run _this chapter_ and play with it on Google Colaboratory by clicking here: [](https://colab.research.google.com/github/ContinualAI/avalanche/blob/master/notebooks/how-tos/avalanchedataset/preamble-pytorch-datasets.ipynb)
| github_jupyter |
# Fixing predict
```
import numpy as np
import pandas as pd
import sys
embeddings = pd.read_pickle("embeddings.pkl")
whales = np.load('raw_predictions.npy')
# get array showing for each class where started it's embeddings
ids = embeddings['Id'].values
last_id, starts = ids[0], [0]
for ind, curr_id in enumerate(ids):
if last_id != curr_id:
starts.append(ind)
last_id = curr_id
starts.append(len(ids))
starts = np.array(starts)
embeddings = embeddings.drop(['Id'], axis=1).values
mean_dist = np.empty((whales.shape[0], len(starts)), dtype=float)
mean_dist[:, 0] = sys.maxsize # new_whale class: constant
#there were computation mistakes before this
mean_emb = np.mean(np.concatenate((embeddings, whales), axis=0), axis = 0)
whales -= mean_emb
embeddings -= mean_emb
```
--- predict we want to fix ---
```
# get 2D array showing mean dist between val embedding and embeddings of group
# using stepped calculation to prevent RAM OOM
class_offset = 1 # to compensate new_whale (class 0) - first column in mean_dist
embeddings_offset = 0
splitted_starts = [starts[:len(starts) // 2 + 1], starts[len(starts) // 2:]]
for starts in [splitted_starts[0]]:
#for starts in splitted_starts:
starts -= embeddings_offset
curr_embeddings = embeddings[starts[0]:starts[-1]]
concat = np.concatenate((curr_embeddings, whales), axis=0)
prod = np.dot(concat, np.transpose(concat))
sq_norms = np.reshape(np.diag(prod), (-1, 1))
dist = sq_norms - 2.0 * prod + np.transpose(sq_norms)
dist = dist[curr_embeddings.shape[0]:, :curr_embeddings.shape[0]]
dist = np.sqrt(np.maximum(dist, 0.0))
print(curr_embeddings.shape)
print(concat.shape)
print(prod.shape)
print(sq_norms.shape)
print(dist.shape)
i = 0
print(np.mean(dist[:, starts[i]:starts[i + 1]], axis=1))
res1 = np.mean(dist[:, starts[i]:starts[i + 1]], axis=1)
```
--- same predict but only for class 1 ---
```
p = 0
for starts in [np.array([splitted_starts[0][p], splitted_starts[0][p+1]]) ]:
curr_embeddings = embeddings[starts[0]:starts[-1]]
concat = np.concatenate((curr_embeddings, whales), axis=0)
prod = np.dot(concat, np.transpose(concat))
sq_norms = np.reshape(np.diag(prod), (-1, 1))
dist = sq_norms - 2.0 * prod + np.transpose(sq_norms)
dist = dist[curr_embeddings.shape[0]:, :curr_embeddings.shape[0]]
dist = np.sqrt(np.maximum(dist, 0.0))
print(curr_embeddings.shape)
print(concat.shape)
print(prod.shape)
print(sq_norms.shape)
print(dist.shape)
print(np.mean(dist, axis=1))
res2 = np.mean(dist, axis=1)
```
--- same but with scipy ---
```
p = 0
for starts in [np.array([splitted_starts[0][p], splitted_starts[0][p+1]]) ]:
curr_embeddings = embeddings[starts[0]:starts[-1]]
res3 = np.linalg.norm(whales - curr_embeddings[0], axis = 1)
res3
res1
np.where((res1 - res2 > 0.00001))[0].shape[0]
np.where((res1 - res3 > 0.00001))[0].shape[0]
from scipy.spatial import distance
dist = distance.cdist(whales, embeddings[:10], 'euclidean')
dist[:, 0]
```
It's ok now!
# Checking embeddings are collapsed
```
import numpy as np
import pandas as pd
import sys
embeddings = pd.read_pickle("embeddings.pkl")
embeddings = embeddings.drop(['Id'], axis=1).values
whales = np.load('raw_predictions.npy')
mean_emb = np.mean(np.concatenate((embeddings, whales), axis=0), axis=0)
mean_emb
np.where(embeddings - mean_emb > 0.05)[0].shape
np.where(whales - mean_emb > 0.05)[0].shape
```
| github_jupyter |
# Data encoding
In this page, we will introduce the problem of data encoding for quantum machine learning, then describe and implement various data encoding methods.
## Introduction
Data representation is crucial for the success of machine learning models. For classical machine learning, the problem is how to represent the data numerically, so that it can be best processed by a classical machine learning algorithm.
For quantum machine learning, this question is similar, but more fundamental: how to represent and efficiently input the data into a quantum system, so that it can be processed by a quantum machine learning algorithm. This is usually referred to as data *encoding*, but is also called data *embedding* or *loading*.
This process is a critical part of quantum machine learning algorithms and directly affects their computational power.
## Methods
Let's consider a classical dataset $\mathscr{X}$ consisting of $M$ samples, each with $N$ [features](gloss:features):
$$\class{script-x}{\mathscr{X}} = \class{brace}{\{}x^{(1)},\class{ellipsis}{\dots},\cssId{_x-lil-m}{x^{(m)}},\dots,x^{(M)}\class{brace}{\}}$$
where $x^{(m)}$ is an $N$ dimensional vector for $m = 1, ..., M$. To represent this dataset in a qubit system, we can use various embedding techniques, some of which are briefly explained and implemented below, as per References [1](#References) and [2](#References).
### Basis encoding
Basis encoding associates a classical $N$-bit string with a [computational basis state](gloss:computational-basis-state) of a $N$-qubit system. For example, if $x = 5$, this can be represented as a $4$-bit string as $0101$, and by a $4$-qubit system as the quantum state $|0101\rangle$. More generally, for an $N$-bit string: $x = (b_1, b_2, ... , b_N)$, the corresponding $N$-qubit state is $\cssId{ket-x}{| x \rangle} = | b_1, b_2, ... , b_N \rangle$ with $b_n \class{in}{\in} \{0,1\}$ for $n = 1 , \dots , N$.
For the classical dataset $\mathscr{X}$ described above, to use basis encoding, each datapoint must be a $N$-bit string: $x^{(m)} = (b_1, b_2, ... , b_N)$, which then can be mapped directly to the quantum state $|x^{m}\rangle = |b_1, b_2, ... , b_N \rangle$ with $b_n \in \{0, 1 \} $ for $n = 1, ..., N$ and $m = 1, ..., M$. We can represent the entire dataset as superpositions of computational basis states:
$$\cssId{_ket-dataset}{| \mathscr{X} \rangle} = \frac{1}{\sqrt{\cssId{_m}{M}}}\cssId{_sum-m}{\sum_{m=1}^{M}|x^{m} \rangle} $$
<!-- ::: q-block -->
### Basis encoding
q-statevector-binary-encoding
p Add/remove bit strings to/from our input dataset on the left to see how basis encoding encodes this in the state vector on the right.
<!-- ::: -->
In Qiskit, once we calculate what state will encode our dataset, we can use the `initialize` function to prepare it. For example, the dataset $\mathscr{X} = \{x^{(1)}=101, x^{(2)}=111\}$ is encoded as the state $|\mathscr{X}\rangle= \frac{1}{\sqrt{2}}(|101\rangle+|111\rangle)$:
```
import math
from qiskit import QuantumCircuit
desired_state = [
0,
0,
0,
0,
0,
1 / math.sqrt(2),
0,
1 / math.sqrt(2)]
qc = QuantumCircuit(3)
qc.initialize(desired_state, [0,1,2])
qc.decompose().decompose().decompose().decompose().decompose().draw()
```
This example illustrates a couple of disadvantages of basis encoding. While it is simple to understand, the state vectors can become quite sparse, and schemes to implement it are usually not efficient.
### Amplitude encoding
Amplitude encoding encodes data into the amplitudes of a quantum state. It represents a normalised classical $N$-dimensional datapoint, $x$, as the amplitudes of a $n$-qubit quantum state, $|\psi_x\rangle$:
$$|\psi_x\rangle = \sum_{i=1}^N x_i |i\rangle$$
where $N = 2^n$, $x_i$ is the $i^{th}$ element of $x$ and $|i\rangle$ is the $i^{th}$ computational basis state.
To encode the classical dataset $\mathscr{X}$ described above, we concatenate all $M$ $N$-dimensional datapoints into one amplitude vector, of length $N \times M$:
$$\alpha=\cssId{_a-norm}{A_{\text{norm}}}(x_{1}^{(1)},...,x_{N}^{(1)},...,x_{1}^{(m)},...,x_{N}^{(m)},...,x_{1}^{(M)},...,x_{N}^{(M)})$$
where $A_{\text{norm}}$ is a normalisation constant, such that $|\alpha|^2 = 1$. The dataset $\mathscr{X}$ can now be represented in the computational basis as:
$$|\mathscr{X}\rangle = \sum_{i=1}^N \alpha_i |i\rangle$$
where $\alpha_i$ are elements of the amplitude vector and $|i\rangle$ are the computational basis states. The number of amplitudes to be encoded is $N \times M$. As a system of $n$ qubits provides $2^n$ amplitudes, amplitude embedding requires $n \ge \mathrm{log}_2(NM)$ qubits.
<!-- ::: q-block -->
### Amplitude encoding
q-statevector-amplitude-encoding
p Change the values of the datapoints on the left, and see how amplitude encoding encodes these as a state vector on the right.
<!-- ::: -->
As an example, let's encode the dataset $\mathscr{X}= \{x^{(1)}=(1.5,0), x^{(2)}=(-2,3)\}$ using amplitude encoding. Concatenating both data points and normalizing the resulting vector, we get:
$$\alpha = \frac{1}{\sqrt{15.25}}(1.5,0,-2,3)$$
and the resulting 2-qubit quantum state would be:
$$|\mathscr{X}\rangle = \frac{1}{\sqrt{15.25}}(1.5|00\rangle-2|10\rangle+3|11\rangle)$$
In the example above, the total number of elements of the amplitude vector, $N \times M$, is a power of 2. When $N \times M$ is not a power of 2, we can simply choose a value for $n$ such that $2^n\geq MN$ and pad the amplitude vector with uninformative constants.
Like in basis encoding, once we calculate what state will encode our dataset, in Qiskit we can use the `initialize` function to prepare it:
```
desired_state = [
1 / math.sqrt(15.25) * 1.5,
0,
1 / math.sqrt(15.25) * -2,
1 / math.sqrt(15.25) * 3]
qc = QuantumCircuit(2)
qc.initialize(desired_state, [0,1])
qc.decompose().decompose().decompose().decompose().decompose().draw()
```
The advantage of amplitude encoding is that it only requires $\mathrm{log}_2(NM)$ qubits to encode. However, subsequent algorithms must operate on the amplitudes of a quantum state, and methods to prepare and measure the quantum states tend not to be efficient.
### Angle encoding
Angle encoding encodes $N$ features into the rotation angles of $n$ qubits, where $N \le n$. For example, the datapoint $x = (x_1,...,x_N)$ can be encoded as:
$$\cssId{_}{|x\rangle} = \class{_big-o-times-n}{\bigotimes^N_{i=1}} \cos(x_i)|0\rangle + \sin(x_i)|1\rangle$$
This is different from the previous two encoding methods, as it only encodes one datapoint at a time, rather than a whole dataset. It does however, only use $N$ qubits and a constant depth quantum circuit, making it amenable to current quantum hardware.
We can specify angle encoding as a [unitary](gloss:unitary):
$$ S_{x_j} = \class{_big-o-times-n}{\bigotimes_{i=1}^N} U(x_j^{(i)}) $$
where:
$$
U(x_j^{(i)}) = \begin{bmatrix} \cos(x_j^{(i)}) & -\sin(x_j^{(i)}) \\ \sin(x_j^{(i)}) & \cos(x_j^{(i)}) \\ \end{bmatrix}
$$
Remembering that a single-qubit rotation around the $Y$-axis is:
$$RY(\theta) = \exp(-i \frac{\theta}{2} Y) = \begin{pmatrix} \cos{\frac{\theta}{2}} & -\sin{\frac{\theta}{2}} \\ \sin{\frac{\theta}{2}} & \cos{\frac{\theta}{2}} \end{pmatrix} $$
We note that $U(x_j^{(i)}) = RY(2x_j^{(i)})$, and as an example, encode the datapoint $x = (0, \pi/4, \pi/2)$ using qiskit:
```
qc = QuantumCircuit(3)
qc.ry(0, 0)
qc.ry(math.pi/4, 1)
qc.ry(math.pi/2, 2)
qc.draw()
```
Dense angle encoding is a slight generalization of angle encoding, that encodes two features per qubit using the relative phase, where the datapoint $x = (x_1,...,x_N)$ can be encoded as:
$$|x\rangle = \class{_big-o-times-n2}{\bigotimes_{i=1}^{N/2}} \cos(x_{2i-1})|0\rangle + e^{i x_{2i}}\sin(x_{2i-1})|1\rangle$$
Although the angle and dense angle encoding use sinuosoids and exponentials, there is nothing special about these functions, and we can easily abstract these to a general class of qubit encodings that use arbitrary functions, or define the encodings as arbitrary unitaries, implemented as parameterized quantum circuits.
### Arbitrary encoding
Arbitrary encoding encodes $N$ features as rotations on $N$ parameterized gates on $n$ qubits, where $n \leq N$. Like angle encoding, it only encodes one datapoint at a time, rather than a whole dataset. It also uses a constant depth quantum circuit and $n \leq N$ qubits, meaning it can be run on current quantum hardware.
For example, to use the Qiskit [`EfficientSU2`](https://qiskit.org/documentation/stubs/qiskit.circuit.library.EfficientSU2.html) circuit to encode 12 features, would only use 3 qubits:
```
from qiskit.circuit.library import EfficientSU2
circuit = EfficientSU2(num_qubits=3, reps=1, insert_barriers=True)
circuit.decompose().draw()
```
Here we encode the datapoint $x = [0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0,1.1,1.2]$ with 12 features, using each of the parameterized gates to encode a different feature.
```
x = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1.0, 1.1, 1.2]
encode = circuit.bind_parameters(x)
encode.decompose().draw()
```
The Qiskit [`ZZFeatureMap`](https://qiskit.org/documentation/stubs/qiskit.circuit.library.ZZFeatureMap.html) circuit with 3 qubits, only encodes a datapoint of 3 features, despite having 6 parameterized gates:
```
from qiskit.circuit.library import ZZFeatureMap
circuit = ZZFeatureMap(3, reps=1, insert_barriers=True)
circuit.decompose().draw()
x = [0.1, 0.2, 0.3]
encode = circuit.bind_parameters(x)
encode.decompose().draw()
```
<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-encoding-0") -->
<!-- ::: .question -->
A parameterized quantum circuit has 16 parameters. What is the largest number of features it can encode?
<!-- ::: -->
<!-- ::: .option -->
1. 4
<!-- ::: -->
<!-- ::: .option -->
2. 8
<!-- ::: -->
<!-- ::: .option(correct) -->
3. 16
<!-- ::: -->
<!-- ::: .option -->
3. 32
<!-- ::: -->
<!-- ::: -->
<!-- ::: -->
The performance of different parameterized quantum circuits on different types of data is an active area of investigation.
<div style="display: none">
$$\cssId{big-o-times}{\bigotimes}$$
</div>
## References
1. Maria Schuld and Francesco Petruccione, *Supervised Learning with Quantum Computers*, Springer 2018, [doi:10.1007/978-3-319-96424-9](https://www.springer.com/gp/book/9783319964232).
2. Ryan LaRose and Brian Coyle, *Robust data encodings for quantum classifiers*, Physical Review A 102, 032420 (2020), [doi:10.1103/PhysRevA.102.032420](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.102.032420), [arXiv:2003.01695](https://arxiv.org/abs/2003.01695).
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
```
# default_exp interpret
#export
from fastai2.data.all import *
from fastai2.optimizer import *
from fastai2.learner import *
import sklearn.metrics as skm
#hide
from fastai2.test_utils import *
```
# Interpretation
> Classes to build objects to better interpret predictions of a model
```
#export
@typedispatch
def plot_top_losses(x, y, *args, **kwargs):
raise Exception(f"plot_top_losses is not implemented for {type(x)},{type(y)}")
#export
_all_ = ["plot_top_losses"]
#export
class Interpretation():
"Interpretation base class, can be inherited for task specific Interpretation classes"
def __init__(self, dl, inputs, preds, targs, decoded, losses):
store_attr(self, "dl,inputs,preds,targs,decoded,losses")
@classmethod
def from_learner(cls, learn, ds_idx=1, dl=None, act=None):
"Construct interpretatio object from a learner"
if dl is None: dl = learn.dls[ds_idx]
return cls(dl, *learn.get_preds(dl=dl, with_input=True, with_loss=True, with_decoded=True, act=None))
def top_losses(self, k=None, largest=True):
"`k` largest(/smallest) losses and indexes, defaulting to all losses (sorted by `largest`)."
return self.losses.topk(ifnone(k, len(self.losses)), largest=largest)
def plot_top_losses(self, k, largest=True, **kwargs):
losses,idx = self.top_losses(k, largest)
if not isinstance(self.inputs, tuple): self.inputs = (self.inputs,)
if isinstance(self.inputs[0], Tensor): inps = tuple(o[idx] for o in self.inputs)
else: inps = self.dl.create_batch(self.dl.before_batch([tuple(o[i] for o in self.inputs) for i in idx]))
b = inps + tuple(o[idx] for o in (self.targs if is_listy(self.targs) else (self.targs,)))
x,y,its = self.dl._pre_show_batch(b, max_n=k)
b_out = inps + tuple(o[idx] for o in (self.decoded if is_listy(self.decoded) else (self.decoded,)))
x1,y1,outs = self.dl._pre_show_batch(b_out, max_n=k)
if its is not None:
plot_top_losses(x, y, its, outs.itemgot(slice(len(inps), None)), self.preds[idx], losses, **kwargs)
#TODO: figure out if this is needed
#its None means that a batch knos how to show itself as a whole, so we pass x, x1
#else: show_results(x, x1, its, ctxs=ctxs, max_n=max_n, **kwargs)
learn = synth_learner()
interp = Interpretation.from_learner(learn)
x,y = learn.dls.valid_ds.tensors
test_eq(interp.inputs, x)
test_eq(interp.targs, y)
out = learn.model.a * x + learn.model.b
test_eq(interp.preds, out)
test_eq(interp.losses, (out-y)[:,0]**2)
#export
class ClassificationInterpretation(Interpretation):
"Interpretation methods for classification models."
def __init__(self, dl, inputs, preds, targs, decoded, losses):
super().__init__(dl, inputs, preds, targs, decoded, losses)
self.vocab = self.dl.vocab
if is_listy(self.vocab): self.vocab = self.vocab[-1]
def confusion_matrix(self):
"Confusion matrix as an `np.ndarray`."
x = torch.arange(0, len(self.vocab))
d,t = flatten_check(self.decoded, self.targs)
cm = ((d==x[:,None]) & (t==x[:,None,None])).long().sum(2)
return to_np(cm)
def plot_confusion_matrix(self, normalize=False, title='Confusion matrix', cmap="Blues", norm_dec=2,
plot_txt=True, **kwargs):
"Plot the confusion matrix, with `title` and using `cmap`."
# This function is mainly copied from the sklearn docs
cm = self.confusion_matrix()
if normalize: cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
fig = plt.figure(**kwargs)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
tick_marks = np.arange(len(self.vocab))
plt.xticks(tick_marks, self.vocab, rotation=90)
plt.yticks(tick_marks, self.vocab, rotation=0)
if plot_txt:
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
coeff = f'{cm[i, j]:.{norm_dec}f}' if normalize else f'{cm[i, j]}'
plt.text(j, i, coeff, horizontalalignment="center", verticalalignment="center", color="white" if cm[i, j] > thresh else "black")
ax = fig.gca()
ax.set_ylim(len(self.vocab)-.5,-.5)
plt.tight_layout()
plt.ylabel('Actual')
plt.xlabel('Predicted')
plt.grid(False)
def most_confused(self, min_val=1):
"Sorted descending list of largest non-diagonal entries of confusion matrix, presented as actual, predicted, number of occurrences."
cm = self.confusion_matrix()
np.fill_diagonal(cm, 0)
res = [(self.vocab[i],self.vocab[j],cm[i,j])
for i,j in zip(*np.where(cm>=min_val))]
return sorted(res, key=itemgetter(2), reverse=True)
def print_classification_report(self):
"Print scikit-learn classification report"
d,t = flatten_check(self.decoded, self.targs)
print(skm.classification_report(t, d, target_names=[str(v) for v in self.vocab]))
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# Custom derivative rules for JAX-transformable Python functions
[](https://colab.research.google.com/github/google/jax/blob/main/docs/notebooks/Custom_derivative_rules_for_Python_code.ipynb)
*mattjj@ Mar 19 2020, last updated Oct 14 2020*
There are two ways to define differentiation rules in JAX:
1. using `jax.custom_jvp` and `jax.custom_vjp` to define custom differentiation rules for Python functions that are already JAX-transformable; and
2. defining new `core.Primitive` instances along with all their transformation rules, for example to call into functions from other systems like solvers, simulators, or general numerical computing systems.
This notebook is about #1. To read instead about #2, see the [notebook on adding primitives](https://jax.readthedocs.io/en/latest/notebooks/How_JAX_primitives_work.html).
For an introduction to JAX's automatic differentiation API, see [The Autodiff Cookbook](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html). This notebook assumes some familiarity with [jax.jvp](https://jax.readthedocs.io/en/latest/jax.html#jax.jvp) and [jax.grad](https://jax.readthedocs.io/en/latest/jax.html#jax.grad), and the mathematical meaning of JVPs and VJPs.
## TL;DR
### Custom JVPs with `jax.custom_jvp`
```
import jax.numpy as jnp
from jax import custom_jvp
@custom_jvp
def f(x, y):
return jnp.sin(x) * y
@f.defjvp
def f_jvp(primals, tangents):
x, y = primals
x_dot, y_dot = tangents
primal_out = f(x, y)
tangent_out = jnp.cos(x) * x_dot * y + jnp.sin(x) * y_dot
return primal_out, tangent_out
from jax import jvp, grad
print(f(2., 3.))
y, y_dot = jvp(f, (2., 3.), (1., 0.))
print(y)
print(y_dot)
print(grad(f)(2., 3.))
# Equivalent alternative using the defjvps convenience wrapper
@custom_jvp
def f(x, y):
return jnp.sin(x) * y
f.defjvps(lambda x_dot, primal_out, x, y: jnp.cos(x) * x_dot * y,
lambda y_dot, primal_out, x, y: jnp.sin(x) * y_dot)
print(f(2., 3.))
y, y_dot = jvp(f, (2., 3.), (1., 0.))
print(y)
print(y_dot)
print(grad(f)(2., 3.))
```
### Custom VJPs with `jax.custom_vjp`
```
from jax import custom_vjp
@custom_vjp
def f(x, y):
return jnp.sin(x) * y
def f_fwd(x, y):
# Returns primal output and residuals to be used in backward pass by f_bwd.
return f(x, y), (jnp.cos(x), jnp.sin(x), y)
def f_bwd(res, g):
cos_x, sin_x, y = res # Gets residuals computed in f_fwd
return (cos_x * g * y, sin_x * g)
f.defvjp(f_fwd, f_bwd)
print(grad(f)(2., 3.))
```
## Example problems
To get an idea of what problems `jax.custom_jvp` and `jax.custom_vjp` are meant to solve, let's go over a few examples. A more thorough introduction to the `jax.custom_jvp` and `jax.custom_vjp` APIs is in [the next section](#scrollTo=Dr0aNkBslfQf).
### Numerical stability
One application of `jax.custom_jvp` is to improve the numerical stability of differentiation.
Say we want to write a function called `log1pexp`, which computes $x \mapsto \log ( 1 + e^x )$. We can write that using `jax.numpy`:
```
import jax.numpy as jnp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
log1pexp(3.)
```
Since it's written in terms of `jax.numpy`, it's JAX-transformable:
```
from jax import jit, grad, vmap
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
But there's a numerical stability problem lurking here:
```
print(grad(log1pexp)(100.))
```
That doesn't seem right! After all, the derivative of $x \mapsto \log (1 + e^x)$ is $x \mapsto \frac{e^x}{1 + e^x}$, and so for large values of $x$ we'd expect the value to be about 1.
We can get a bit more insight into what's going on by looking at the jaxpr for the gradient computation:
```
from jax import make_jaxpr
make_jaxpr(grad(log1pexp))(100.)
```
Stepping through how the jaxpr would be evaluated, we can see that the last line would involve multiplying values that floating point math will round to 0 and $\infty$, respectively, which is never a good idea. That is, we're effectively evaluating `lambda x: (1 / (1 + jnp.exp(x))) * jnp.exp(x)` for large `x`, which effectively turns into `0. * jnp.inf`.
Instead of generating such large and small values, hoping for a cancellation that floats can't always provide, we'd rather just express the derivative function as a more numerically stable program. In particular, we can write a program that more closely evaluates the equal mathematical expression $1 - \frac{1}{1 + e^x}$, with no cancellation in sight.
This problem is interesting because even though our definition of `log1pexp` could already be JAX-differentiated (and transformed with `jit`, `vmap`, ...), we're not happy with the result of applying standard autodiff rules to the primitives comprising `log1pexp` and composing the result. Instead, we'd like to specify how the whole function `log1pexp` should be differentiated, as a unit, and thus arrange those exponentials better.
This is one application of custom derivative rules for Python functions that are already JAX transformable: specifying how a composite function should be differentiated, while still using its original Python definition for other transformations (like `jit`, `vmap`, ...).
Here's a solution using `jax.custom_jvp`:
```
from jax import custom_jvp
@custom_jvp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
@log1pexp.defjvp
def log1pexp_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = log1pexp(x)
ans_dot = (1 - 1/(1 + jnp.exp(x))) * x_dot
return ans, ans_dot
print(grad(log1pexp)(100.))
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
Here's a `defjvps` convenience wrapper to express the same thing:
```
@custom_jvp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
log1pexp.defjvps(lambda t, ans, x: (1 - 1/(1 + jnp.exp(x))) * t)
print(grad(log1pexp)(100.))
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
### Enforcing a differentiation convention
A related application is to enforce a differentiation convention, perhaps at a boundary.
Consider the function $f : \mathbb{R}_+ \mapsto \mathbb{R}_+$ with $f(x) = \frac{x}{1 + \sqrt{x}}$, where we take $\mathbb{R}_+ = [0, \infty)$. We might implement $f$ as a program like this:
```
def f(x):
return x / (1 + jnp.sqrt(x))
```
As a mathematical function on $\mathbb{R}$ (the full real line), $f$ is not differentiable at zero (because the limit defining the derivative doesn't exist from the left). Correspondingly, autodiff produces a `nan` value:
```
print(grad(f)(0.))
```
But mathematically if we think of $f$ as a function on $\mathbb{R}_+$ then it is differentiable at 0 [Rudin's Principles of Mathematical Analysis Definition 5.1, or Tao's Analysis I 3rd ed. Definition 10.1.1 and Example 10.1.6]. Alternatively, we might say as a convention we want to consider the directional derivative from the right. So there is a sensible value for the Python function `grad(f)` to return at `0.0`, namely `1.0`. By default, JAX's machinery for differentiation assumes all functions are defined over $\mathbb{R}$ and thus doesn't produce `1.0` here.
We can use a custom JVP rule! In particular, we can define the JVP rule in terms of the derivative function $x \mapsto \frac{\sqrt{x} + 2}{2(\sqrt{x} + 1)^2}$ on $\mathbb{R}_+$,
```
@custom_jvp
def f(x):
return x / (1 + jnp.sqrt(x))
@f.defjvp
def f_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = f(x)
ans_dot = ((jnp.sqrt(x) + 2) / (2 * (jnp.sqrt(x) + 1)**2)) * x_dot
return ans, ans_dot
print(grad(f)(0.))
```
Here's the convenience wrapper version:
```
@custom_jvp
def f(x):
return x / (1 + jnp.sqrt(x))
f.defjvps(lambda t, ans, x: ((jnp.sqrt(x) + 2) / (2 * (jnp.sqrt(x) + 1)**2)) * t)
print(grad(f)(0.))
```
### Gradient clipping
While in some cases we want to express a mathematical differentiation computation, in other cases we may even want to take a step away from mathematics to adjust the computation autodiff performs. One canonical example is reverse-mode gradient clipping.
For gradient clipping, we can use `jnp.clip` together with a `jax.custom_vjp` reverse-mode-only rule:
```
from functools import partial
from jax import custom_vjp
@custom_vjp
def clip_gradient(lo, hi, x):
return x # identity function
def clip_gradient_fwd(lo, hi, x):
return x, (lo, hi) # save bounds as residuals
def clip_gradient_bwd(res, g):
lo, hi = res
return (None, None, jnp.clip(g, lo, hi)) # use None to indicate zero cotangents for lo and hi
clip_gradient.defvjp(clip_gradient_fwd, clip_gradient_bwd)
import matplotlib.pyplot as plt
from jax import vmap
t = jnp.linspace(0, 10, 1000)
plt.plot(jnp.sin(t))
plt.plot(vmap(grad(jnp.sin))(t))
def clip_sin(x):
x = clip_gradient(-0.75, 0.75, x)
return jnp.sin(x)
plt.plot(clip_sin(t))
plt.plot(vmap(grad(clip_sin))(t))
```
### Python debugging
Another application that is motivated by development workflow rather than numerics is to set a `pdb` debugger trace in the backward pass of reverse-mode autodiff.
When trying to track down the source of a `nan` runtime error, or just examine carefully the cotangent (gradient) values being propagated, it can be useful to insert a debugger at a point in the backward pass that corresponds to a specific point in the primal computation. You can do that with `jax.custom_vjp`.
We'll defer an example until the next section.
### Implicit function differentiation of iterative implementations
This example gets pretty deep in the mathematical weeds!
Another application for `jax.custom_vjp` is reverse-mode differentiation of functions that are JAX-transformable (by `jit`, `vmap`, ...) but not efficiently JAX-differentiable for some reason, perhaps because they involve `lax.while_loop`. (It's not possible to produce an XLA HLO program that efficiently computes the reverse-mode derivative of an XLA HLO While loop because that would require a program with unbounded memory use, which isn't possible to express in XLA HLO, at least without side-effecting interactions through infeed/outfeed.)
For example, consider this `fixed_point` routine which computes a fixed point by iteratively applying a function in a `while_loop`:
```
from jax.lax import while_loop
def fixed_point(f, a, x_guess):
def cond_fun(carry):
x_prev, x = carry
return jnp.abs(x_prev - x) > 1e-6
def body_fun(carry):
_, x = carry
return x, f(a, x)
_, x_star = while_loop(cond_fun, body_fun, (x_guess, f(a, x_guess)))
return x_star
```
This is an iterative procedure for numerically solving the equation $x = f(a, x)$ for $x$, by iterating $x_{t+1} = f(a, x_t)$ until $x_{t+1}$ is sufficiently close to $x_t$. The result $x^*$ depends on the parameters $a$, and so we can think of there being a function $a \mapsto x^*(a)$ that is implicitly defined by equation $x = f(a, x)$.
We can use `fixed_point` to run iterative procedures to convergence, for example running Newton's method to calculate square roots while only executing adds, multiplies, and divides:
```
def newton_sqrt(a):
update = lambda a, x: 0.5 * (x + a / x)
return fixed_point(update, a, a)
print(newton_sqrt(2.))
```
We can `vmap` or `jit` the function as well:
```
print(jit(vmap(newton_sqrt))(jnp.array([1., 2., 3., 4.])))
```
We can't apply reverse-mode automatic differentiation because of the `while_loop`, but it turns out we wouldn't want to anyway: instead of differentiating through the implementation of `fixed_point` and all its iterations, we can exploit the mathematical structure to do something that is much more memory-efficient (and FLOP-efficient in this case, too!). We can instead use the implicit function theorem [Prop A.25 of Bertsekas's Nonlinear Programming, 2nd ed.], which guarantees (under some conditions) the existence of the mathematical objects we're about to use. In essence, we linearize at the solution and solve those linear equations iteratively to compute the derivatives we want.
Consider again the equation $x = f(a, x)$ and the function $x^*$. We want to evaluate vector-Jacobian products like $v^\mathsf{T} \mapsto v^\mathsf{T} \partial x^*(a_0)$.
At least in an open neighborhood around the point $a_0$ at which we want to differentiate, let's assume that the equation $x^*(a) = f(a, x^*(a))$ holds for all $a$. Since the two sides are equal as functions of $a$, their derivatives must be equal as well, so let's differentiate both sides:
$\qquad \partial x^*(a) = \partial_0 f(a, x^*(a)) + \partial_1 f(a, x^*(a)) \partial x^*(a)$.
Setting $A = \partial_1 f(a_0, x^*(a_0))$ and $B = \partial_0 f(a_0, x^*(a_0))$, we can write the quantity we're after more simply as
$\qquad \partial x^*(a_0) = B + A \partial x^*(a_0)$,
or, by rearranging,
$\qquad \partial x^*(a_0) = (I - A)^{-1} B$.
That means we can evaluate vector-Jacobian products like
$\qquad v^\mathsf{T} \partial x^*(a_0) = v^\mathsf{T} (I - A)^{-1} B = w^\mathsf{T} B$,
where $w^\mathsf{T} = v^\mathsf{T} (I - A)^{-1}$, or equivalently $w^\mathsf{T} = v^\mathsf{T} + w^\mathsf{T} A$, or equivalently $w^\mathsf{T}$ is the fixed point of the map $u^\mathsf{T} \mapsto v^\mathsf{T} + u^\mathsf{T} A$. That last characterization gives us a way to write the VJP for `fixed_point` in terms of a call to `fixed_point`! Moreover, after expanding $A$ and $B$ back out, we can see we need only to evaluate VJPs of $f$ at $(a_0, x^*(a_0))$.
Here's the upshot:
```
from jax import vjp
@partial(custom_vjp, nondiff_argnums=(0,))
def fixed_point(f, a, x_guess):
def cond_fun(carry):
x_prev, x = carry
return jnp.abs(x_prev - x) > 1e-6
def body_fun(carry):
_, x = carry
return x, f(a, x)
_, x_star = while_loop(cond_fun, body_fun, (x_guess, f(a, x_guess)))
return x_star
def fixed_point_fwd(f, a, x_init):
x_star = fixed_point(f, a, x_init)
return x_star, (a, x_star)
def fixed_point_rev(f, res, x_star_bar):
a, x_star = res
_, vjp_a = vjp(lambda a: f(a, x_star), a)
a_bar, = vjp_a(fixed_point(partial(rev_iter, f),
(a, x_star, x_star_bar),
x_star_bar))
return a_bar, jnp.zeros_like(x_star)
def rev_iter(f, packed, u):
a, x_star, x_star_bar = packed
_, vjp_x = vjp(lambda x: f(a, x), x_star)
return x_star_bar + vjp_x(u)[0]
fixed_point.defvjp(fixed_point_fwd, fixed_point_rev)
print(newton_sqrt(2.))
print(grad(newton_sqrt)(2.))
print(grad(grad(newton_sqrt))(2.))
```
We can check our answers by differentiating `jnp.sqrt`, which uses a totally different implementation:
```
print(grad(jnp.sqrt)(2.))
print(grad(grad(jnp.sqrt))(2.))
```
A limitation to this approach is that the argument `f` can't close over any values involved in differentiation. That is, you might notice that we kept the parameter `a` explicit in the argument list of `fixed_point`. While other JAX mechanisms can handle closed-over transformation-traced values in the arguments to higher-order functions (as is done for the control flow primitives like `lax.cond`, `lax.scan`, and `lax.while_loop` itself), `jax.custom_vjp` used as above cannot. A `fixed_point` routine that used a bit more of JAX's internals could have a more convenient and robust API.
## Basic usage of `jax.custom_jvp` and `jax.custom_vjp` APIs
### Use `jax.custom_jvp` to define forward-mode (and, indirectly, reverse-mode) rules
Here's a canonical basic example of using `jax.custom_jvp`:
```
from jax import custom_jvp
import jax.numpy as jnp
# f :: a -> b
@custom_jvp
def f(x):
return jnp.sin(x)
# f_jvp :: (a, T a) -> (b, T b)
def f_jvp(primals, tangents):
x, = primals
t, = tangents
return f(x), jnp.cos(x) * t
f.defjvp(f_jvp)
from jax import jvp
print(f(3.))
y, y_dot = jvp(f, (3.,), (1.,))
print(y)
print(y_dot)
```
In words, we start with a primal function `f` that takes inputs of type `a` and produces outputs of type `b`. We associate with it a JVP rule function `f_jvp` that takes a pair of inputs representing the primal inputs of type `a` and the corresponding tangent inputs of type `T a`, and produces a pair of outputs representing the primal outputs of type `b` and tangent outputs of type `T b`. The tangent outputs should be a linear function of the tangent inputs.
You can also use `f.defjvp` as a decorator, as in
```python
@custom_jvp
def f(x):
...
@f.defjvp
def f_jvp(primals, tangents):
...
```
Even though we defined only a JVP rule and no VJP rule, we can use both forward- and reverse-mode differentiation on `f`. JAX will automatically transpose the linear computation on tangent values from our custom JVP rule, computing the VJP as efficiently as if we had written the rule by hand:
```
from jax import grad
print(grad(f)(3.))
print(grad(grad(f))(3.))
```
For automatic transposition to work, the JVP rule's output tangents must be linear as a function of the input tangents. Otherwise a transposition error is raised.
Multiple arguments work like this:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
@f.defjvp
def f_jvp(primals, tangents):
x, y = primals
x_dot, y_dot = tangents
primal_out = f(x, y)
tangent_out = 2 * x * y * x_dot + x ** 2 * y_dot
return primal_out, tangent_out
print(grad(f)(2., 3.))
```
The `defjvps` convenience wrapper lets us define a JVP for each argument separately, and the results are computed separately then summed:
```
@custom_jvp
def f(x):
return jnp.sin(x)
f.defjvps(lambda t, ans, x: jnp.cos(x) * t)
print(grad(f)(3.))
```
Here's a `defjvps` example with multiple arguments:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
f.defjvps(lambda x_dot, primal_out, x, y: 2 * x * y * x_dot,
lambda y_dot, primal_out, x, y: x ** 2 * y_dot)
print(grad(f)(2., 3.))
print(grad(f, 0)(2., 3.)) # same as above
print(grad(f, 1)(2., 3.))
```
As a shorthand, with `defjvps` you can pass a `None` value to indicate that the JVP for a particular argument is zero:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
f.defjvps(lambda x_dot, primal_out, x, y: 2 * x * y * x_dot,
None)
print(grad(f)(2., 3.))
print(grad(f, 0)(2., 3.)) # same as above
print(grad(f, 1)(2., 3.))
```
Calling a `jax.custom_jvp` function with keyword arguments, or writing a `jax.custom_jvp` function definition with default arguments, are both allowed so long as they can be unambiguously mapped to positional arguments based on the function signature retrieved by the standard library `inspect.signature` mechanism.
When you're not performing differentiation, the function `f` is called just as if it weren't decorated by `jax.custom_jvp`:
```
@custom_jvp
def f(x):
print('called f!') # a harmless side-effect
return jnp.sin(x)
@f.defjvp
def f_jvp(primals, tangents):
print('called f_jvp!') # a harmless side-effect
x, = primals
t, = tangents
return f(x), jnp.cos(x) * t
from jax import vmap, jit
print(f(3.))
print(vmap(f)(jnp.arange(3.)))
print(jit(f)(3.))
```
The custom JVP rule is invoked during differentiation, whether forward or reverse:
```
y, y_dot = jvp(f, (3.,), (1.,))
print(y_dot)
print(grad(f)(3.))
```
Notice that `f_jvp` calls `f` to compute the primal outputs. In the context of higher-order differentiation, each application of a differentiation transform will use the custom JVP rule if and only if the rule calls the original `f` to compute the primal outputs. (This represents a kind of fundamental tradeoff, where we can't make use of intermediate values from the evaluation of `f` in our rule _and also_ have the rule apply in all orders of higher-order differentiation.)
```
grad(grad(f))(3.)
```
You can use Python control flow with `jax.custom_jvp`:
```
@custom_jvp
def f(x):
if x > 0:
return jnp.sin(x)
else:
return jnp.cos(x)
@f.defjvp
def f_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = f(x)
if x > 0:
return ans, 2 * x_dot
else:
return ans, 3 * x_dot
print(grad(f)(1.))
print(grad(f)(-1.))
```
### Use `jax.custom_vjp` to define custom reverse-mode-only rules
While `jax.custom_jvp` suffices for controlling both forward- and, via JAX's automatic transposition, reverse-mode differentiation behavior, in some cases we may want to directly control a VJP rule, for example in the latter two example problems presented above. We can do that with `jax.custom_vjp`:
```
from jax import custom_vjp
import jax.numpy as jnp
# f :: a -> b
@custom_vjp
def f(x):
return jnp.sin(x)
# f_fwd :: a -> (b, c)
def f_fwd(x):
return f(x), jnp.cos(x)
# f_bwd :: (c, CT b) -> CT a
def f_bwd(cos_x, y_bar):
return (cos_x * y_bar,)
f.defvjp(f_fwd, f_bwd)
from jax import grad
print(f(3.))
print(grad(f)(3.))
```
In words, we again start with a primal function `f` that takes inputs of type `a` and produces outputs of type `b`. We associate with it two functions, `f_fwd` and `f_bwd`, which describe how to perform the forward- and backward-passes of reverse-mode autodiff, respectively.
The function `f_fwd` describes the forward pass, not only the primal computation but also what values to save for use on the backward pass. Its input signature is just like that of the primal function `f`, in that it takes a primal input of type `a`. But as output it produces a pair, where the first element is the primal output `b` and the second element is any "residual" data of type `c` to be stored for use by the backward pass. (This second output is analogous to [PyTorch's save_for_backward mechanism](https://pytorch.org/tutorials/beginner/examples_autograd/two_layer_net_custom_function.html).)
The function `f_bwd` describes the backward pass. It takes two inputs, where the first is the residual data of type `c` produced by `f_fwd` and the second is the output cotangents of type `CT b` corresponding to the output of the primal function. It produces an output of type `CT a` representing the cotangents corresponding to the input of the primal function. In particular, the output of `f_bwd` must be a sequence (e.g. a tuple) of length equal to the number of arguments to the primal function.
So multiple arguments work like this:
```
from jax import custom_vjp
@custom_vjp
def f(x, y):
return jnp.sin(x) * y
def f_fwd(x, y):
return f(x, y), (jnp.cos(x), jnp.sin(x), y)
def f_bwd(res, g):
cos_x, sin_x, y = res
return (cos_x * g * y, -sin_x * g)
f.defvjp(f_fwd, f_bwd)
print(grad(f)(2., 3.))
```
Calling a `jax.custom_vjp` function with keyword arguments, or writing a `jax.custom_vjp` function definition with default arguments, are both allowed so long as they can be unambiguously mapped to positional arguments based on the function signature retrieved by the standard library `inspect.signature` mechanism.
As with `jax.custom_jvp`, the custom VJP rule comprised by `f_fwd` and `f_bwd` is not invoked if differentiation is not applied. If function is evaluated, or transformed with `jit`, `vmap`, or other non-differentiation transformations, then only `f` is called.
```
@custom_vjp
def f(x):
print("called f!")
return jnp.sin(x)
def f_fwd(x):
print("called f_fwd!")
return f(x), jnp.cos(x)
def f_bwd(cos_x, y_bar):
print("called f_bwd!")
return (cos_x * y_bar,)
f.defvjp(f_fwd, f_bwd)
print(f(3.))
print(grad(f)(3.))
from jax import vjp
y, f_vjp = vjp(f, 3.)
print(y)
print(f_vjp(1.))
```
**Forward-mode autodiff cannot be used on the** `jax.custom_vjp` **function** and will raise an error:
```
from jax import jvp
try:
jvp(f, (3.,), (1.,))
except TypeError as e:
print('ERROR! {}'.format(e))
```
If you want to use both forward- and reverse-mode, use `jax.custom_jvp` instead.
We can use `jax.custom_vjp` together with `pdb` to insert a debugger trace in the backward pass:
```
import pdb
@custom_vjp
def debug(x):
return x # acts like identity
def debug_fwd(x):
return x, x
def debug_bwd(x, g):
import pdb; pdb.set_trace()
return g
debug.defvjp(debug_fwd, debug_bwd)
def foo(x):
y = x ** 2
y = debug(y) # insert pdb in corresponding backward pass step
return jnp.sin(y)
```
```python
jax.grad(foo)(3.)
> <ipython-input-113-b19a2dc1abf7>(12)debug_bwd()
-> return g
(Pdb) p x
DeviceArray(9., dtype=float32)
(Pdb) p g
DeviceArray(-0.91113025, dtype=float32)
(Pdb) q
```
## More features and details
### Working with `list` / `tuple` / `dict` containers (and other pytrees)
You should expect standard Python containers like lists, tuples, namedtuples, and dicts to just work, along with nested versions of those. In general, any [pytrees](https://jax.readthedocs.io/en/latest/pytrees.html) are permissible, so long as their structures are consistent according to the type constraints.
Here's a contrived example with `jax.custom_jvp`:
```
from collections import namedtuple
Point = namedtuple("Point", ["x", "y"])
@custom_jvp
def f(pt):
x, y = pt.x, pt.y
return {'a': x ** 2,
'b': (jnp.sin(x), jnp.cos(y))}
@f.defjvp
def f_jvp(primals, tangents):
pt, = primals
pt_dot, = tangents
ans = f(pt)
ans_dot = {'a': 2 * pt.x * pt_dot.x,
'b': (jnp.cos(pt.x) * pt_dot.x, -jnp.sin(pt.y) * pt_dot.y)}
return ans, ans_dot
def fun(pt):
dct = f(pt)
return dct['a'] + dct['b'][0]
pt = Point(1., 2.)
print(f(pt))
print(grad(fun)(pt))
```
And an analogous contrived example with `jax.custom_vjp`:
```
@custom_vjp
def f(pt):
x, y = pt.x, pt.y
return {'a': x ** 2,
'b': (jnp.sin(x), jnp.cos(y))}
def f_fwd(pt):
return f(pt), pt
def f_bwd(pt, g):
a_bar, (b0_bar, b1_bar) = g['a'], g['b']
x_bar = 2 * pt.x * a_bar + jnp.cos(pt.x) * b0_bar
y_bar = -jnp.sin(pt.y) * b1_bar
return (Point(x_bar, y_bar),)
f.defvjp(f_fwd, f_bwd)
def fun(pt):
dct = f(pt)
return dct['a'] + dct['b'][0]
pt = Point(1., 2.)
print(f(pt))
print(grad(fun)(pt))
```
### Handling non-differentiable arguments
Some use cases, like the final example problem, call for non-differentiable arguments like function-valued arguments to be passed to functions with custom differentiation rules, and for those arguments to also be passed to the rules themselves. In the case of `fixed_point`, the function argument `f` was such a non-differentiable argument. A similar situation arises with `jax.experimental.odeint`.
#### `jax.custom_jvp` with `nondiff_argnums`
Use the optional `nondiff_argnums` parameter to `jax.custom_jvp` to indicate arguments like these. Here's an example with `jax.custom_jvp`:
```
from functools import partial
@partial(custom_jvp, nondiff_argnums=(0,))
def app(f, x):
return f(x)
@app.defjvp
def app_jvp(f, primals, tangents):
x, = primals
x_dot, = tangents
return f(x), 2. * x_dot
print(app(lambda x: x ** 3, 3.))
print(grad(app, 1)(lambda x: x ** 3, 3.))
```
Notice the gotcha here: no matter where in the argument list these parameters appear, they're placed at the *start* of the signature of the corresponding JVP rule. Here's another example:
```
@partial(custom_jvp, nondiff_argnums=(0, 2))
def app2(f, x, g):
return f(g((x)))
@app2.defjvp
def app2_jvp(f, g, primals, tangents):
x, = primals
x_dot, = tangents
return f(g(x)), 3. * x_dot
print(app2(lambda x: x ** 3, 3., lambda y: 5 * y))
print(grad(app2, 1)(lambda x: x ** 3, 3., lambda y: 5 * y))
```
#### `jax.custom_vjp` with `nondiff_argnums`
A similar option exists for `jax.custom_vjp`, and, similarly, the convention is that the non-differentiable arguments are passed as the first arguments to the `_bwd` rule, no matter where they appear in the signature of the original function. The signature of the `_fwd` rule remains unchanged - it is the same as the signature of the primal function. Here's an example:
```
@partial(custom_vjp, nondiff_argnums=(0,))
def app(f, x):
return f(x)
def app_fwd(f, x):
return f(x), x
def app_bwd(f, x, g):
return (5 * g,)
app.defvjp(app_fwd, app_bwd)
print(app(lambda x: x ** 2, 4.))
print(grad(app, 1)(lambda x: x ** 2, 4.))
```
See `fixed_point` above for another usage example.
**You don't need to use** `nondiff_argnums` **with array-valued arguments**, for example ones with integer dtype. Instead, `nondiff_argnums` should only be used for argument values that don't correspond to JAX types (essentially don't correspond to array types), like Python callables or strings. If JAX detects that an argument indicated by `nondiff_argnums` contains a JAX Tracer, then an error is raised. The `clip_gradient` function above is a good example of not using `nondiff_argnums` for integer-dtype array arguments.
| github_jupyter |
# Introduction to Matplotlib
- Different kinds of figures
- line plots
- scatter plots
- lines
- histogram
- heatmap
- 3D (using a 3d Numpy array)
- multiple plots on the same axis
- adding legend
- Cutomizing the figures
- changing color (including defining your own color using rgb values)
- changing line style
- changing marker
- grid()
- changing the background
- The anatomy of matplotlib (figures and axes)
- subplots
- `
- Extra (for fun)
- xkcd
### Introduction
Humans are very visual creatures: we understand things better when we see things visualized. Matplotlib is a flexible python library which can help you visualize your results. You don’t need much to get started: you need to make the necessary imports, prepare some data, and you can start plotting with the help of the `plot()` function! Once you created your plot, you might need to explicitly write a command to show it, `show()`.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Let's start by creating a Numpy array
```
plt.plot([0, 10, 5, 7, 2, 1, 8, 8, 4, 4]);
```
Note that this results in a figure that contains the values (y axis) and their correspondign index number (x axis)
```
y = [0, 10, 5, 7, 2, 1, 8, 8, 4, 4]
x = [.1, .2, .3, .4, .5, .6, .7, .8, .9, 1]
plt.plot(x, y);
x = np.arange(0, 360, .01)
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y);
```
### Playing around with the line properties
- color
- style
- width
```
plt.plot(x, y, color='r');
```
Colors can be specified in different ways in matplotlib:
- an RGB or RGBA tuple of float values defined between 0 and 1 (0 to 255 normalized)
- a string representation of a float value defined between 0 and 1. This is used for grayscale
- one of the characters below
| Character | Color |
|-----------|-------|
| 'b' | blue |
| 'g' | green |
| 'r' | red |
| 'c' | cyan |
| 'm' | magenta |
| 'y' | yellow |
| 'k' | black |
| 'w' | white |
```
plt.plot(x, y, color=(1, 0, 0));
plt.plot(x, y, color='.7');
```
There are mainly four line styles available in matplotlib:
- "dotted"
- "dashdot"
- "dashed"
- "solid"
you can use the following line to get a docstring:
``` python
plt.Line2D.set_linestyle?
```

```
plt.plot(x, y, color='r', linestyle='dotted');
plt.plot(x, y, color='r', linestyle=':');
```
There is a shortcut!!
```
plt.plot(x, y, 'r:');
plt.plot(x, y, color='r', linestyle='dashdot', linewidth=10);
```
### Playing around with the axes
- axes range
- labels
- fontsize
- tick labels
- labels
- number of labels
- fontsize
We can label our axis using `xlabel()` and `ylabel()` methods (try adding labels)
```
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y);
```
We can limit the range (make is smaller or bigger) of numbers for each axis
```
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
plt.xlim(-100, 460)
plt.ylim(-10, 10);
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
plt.xlabel('X Label')
plt.ylabel('Y Label');
```
alright. Let's now play with the axis values. we can change the resolution at which we are showing values in each axes.
```
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
plt.xlabel('X Label')
plt.ylabel('Y Label')
plt.locator_params(axis='both', nbins=16)
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
# plt.xlabel('X Label')
plt.ylabel('Y Label')
plt.xticks([100, 200, 300], ['Python', 'Is', 'Awesome']);
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
# plt.xlabel('X Label')
plt.ylabel('Y Label', fontsize=30)
plt.yticks(fontsize=30)
plt.xticks([90, 180, 300], ['Python', 'Is', 'Awesome'], fontsize=30);
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y)
ax = plt.gca()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# ax.spines['bottom'].set_visible(True)
# ax.spines['left'].set_visible(True)
# plt.axis('off')
plt.xlim(-10, 370)
plt.ylim(-2, 2)
# plt.xlabel('X Label')
plt.ylabel('Y Label', fontsize=30)
plt.yticks(fontsize=30)
plt.xticks([90, 180, 300], ['Python', 'Is', 'Awesome'], fontsize=30);
```
Adding legends, colorbar, and grid
```
x_rad = np.deg2rad(x)
y = np.sin(x_rad)
plt.plot(x, y, label='Our first line')
ax = plt.gca()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# ax.spines['bottom'].set_visible(True)
# ax.spines['left'].set_visible(True)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
plt.xlabel('')
plt.ylabel('Y Label', fontsize=30)
plt.yticks(fontsize=30)
plt.xticks([90, 180, 300], ['Python', 'Is', 'Awesome'], fontsize=30)
plt.grid()
plt.legend(loc='upper left');
x_rad = np.deg2rad(x)
y1 = np.sin(x_rad)
y2 = np.cos(x_rad)
plt.plot(x, y1, label='sin', color='r', linestyle='-.', linewidth=5)
plt.plot(x, y2, label='cos', color='g', linestyle='-.', linewidth=5)
ax = plt.gca()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
# ax.spines['bottom'].set_visible(True)
# ax.spines['left'].set_visible(True)
plt.xlim(-10, 370)
plt.ylim(-2, 2)
plt.xlabel('')
plt.ylabel('Y Label', fontsize=30)
plt.locator_params(axis='y', nbins=5)
plt.yticks(fontsize=30)
plt.xticks([90, 180, 300], ['Python', 'Is', 'Awesome'], fontsize=30)
plt.grid()
plt.legend(loc='upper left');
plt.suptitle('Title', fontsize=30);
```
### Anatomy of matplotlib plot
In essence, there are two big components that you need to take into account:
- The **Figure** is the overall window or page that everything is drawn on. It’s the top-level component of all the ones that you will consider in the following points. You can create multiple independent Figures. A Figure can have several other things in it, such as a **suptitle**, which is a centered title to the figure. You’ll also find that you can add a **legend** and **color bar**, for example, to your Figure.
- To the figure you add Axes. The Axes is the area on which the data is plotted with functions such as `plot()` and `scatter()` and that can have ticks, labels, etc. associated with it. This explains why Figures can contain multiple Axes.
<div style="text-align:center">
<a href="https://matplotlib.org/1.5.1/faq/usage_faq.html">
<img src ="https://matplotlib.org/1.5.1/_images/fig_map.png" />
</div>
Pretty much whatever we did above can be done with the axis objec as well. In other words, the methods used to manipulate the properties of a plot, for example `plt.xlabel()` all exist for an axes object as well. For instacne, instead of `plt.xlabel()` we can also use `ax.set_xlabel()`.
```
fig, ax = plt.subplots() # figsize=(10, 5)
# remove axis boarders
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.plot(x, y1, label='sin', color='r', linestyle='-.', linewidth=5)
ax.plot(x, y2, label='cos', color='g', linestyle='-.', linewidth=5)
# setting labels
ax.set_xlabel('', fontsize=30) # plt.xlabel('X Label')
ax.set_ylabel('Y Label', fontsize=30); # plt.ylabel('Y Label')
# changing the values on the axes
ax.locator_params(axis='both', nbins=4)
ax.set_xticks([90, 180, 300])
ax.set_xticklabels(['Python', 'Is', 'Awesome'])
ax.tick_params(axis='both',
direction='in',
labelsize=30)
ax.legend(loc='lower left')
ax.grid()
```
### Subplots
```
fig, axes = plt.subplots(nrows=2, ncols=1)
axes
fig, axes = plt.subplots(nrows=2, ncols=1)
axes[0].plot(x, y1)
axes[1].plot(x, y2);
(ax1, ax2) = axes
fig, (ax1, ax2) = plt.subplots(nrows=2, ncols=1)
ax1.plot(x, y1)
ax2.plot(x, y2);
```
What if we have more subplots?
```
fig, axes = plt.subplots(nrows=3, ncols=3)
for ax in axes.flat:
ax.plot(np.random.random(100))
fig, axes = plt.subplots(nrows=3, ncols=3)
for ax in axes.flat:
ax.plot(np.random.random(100))
fig.tight_layout()
```
If we have several plots that share the same axis we can share the numbers on the axis (i.e., ticklabels)
```
fig, axes = plt.subplots(nrows=3, ncols=3, sharex=True, sharey=True)
for ax in axes.flat:
ax.plot(np.random.random(100))
x = np.arange(0, 360, 5)
x_rad = np.deg2rad(x)
y1 = np.sin(x_rad)
y2 = np.cos(x_rad)
fig, axes = plt.subplots(nrows=3, ncols=3, sharex=True, sharey=True, figsize=(15, 10))
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'w', 'b']
for ind, (col, ax) in enumerate(zip(colors, axes.flat)):
ax.plot(x, y1 + np.random.random(x.shape[0]), color=col, linewidth=ind+1, label= 'Line width = ' + str(ind))
# changing the values on the axes
ax.locator_params(axis='both', nbins=4)
ax.tick_params(axis='both', labelsize=30)
ax.legend(loc='lower left', fontsize=15)
ax.grid()
# Note that we are setting the labels for specific axes
axes[2, 1].set_xlabel('X Label', fontsize=30)
axes[1, 0].set_ylabel('Y Label', fontsize=30)
fig.tight_layout()
from matplotlib.gridspec import GridSpec
fig = plt.figure(figsize=(15, 10))
gs = GridSpec(3, 3)
ax1 = plt.subplot(gs[0, :])
ax2 = plt.subplot(gs[1, :-1])
ax3 = plt.subplot(gs[1:, -1])
ax4 = plt.subplot(gs[2, 0])
ax5 = plt.subplot(gs[2, 1])
axes = (ax1, ax2, ax3, ax4, ax5)
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'w', 'b']
for ind, (col, ax) in enumerate(zip(colors, axes)):
ax.plot(x, y1 + np.random.random(x.shape[0]), color=col, linewidth=ind+1, label= 'Line width = ' + str(ind))
# changing the values on the axes
ax.locator_params(axis='both', nbins=4)
ax.tick_params(axis='both', labelsize=30)
ax.legend(loc='lower left', fontsize=15)
ax.grid()
fig.tight_layout()
```
### Save the figure
- resolution
- transparent background
```
fig.savefig("NameOfTheFile.png", transparent=True)
```
---
### Bonus: Fun with Matplotlib
```
with plt.xkcd():
plt.plot(np.sin(np.linspace(0, 10)))
plt.title('Whoo Hoo!!!');
with plt.xkcd():
fig, axes = plt.subplots(nrows=3, ncols=3, sharex=True, sharey=True, figsize=(15, 10))
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'w', 'b']
for ind, (col, ax) in enumerate(zip(colors, axes.flat)):
ax.plot(x, y1 + np.random.random(x.shape[0]), color=col, linewidth=ind+1)
ax.grid()
fig.tight_layout()
```
#### **Installing new font for your matplotlib library**
1. Download the font
2. Install the font in the system
3. Rebuilt the font cache
Downlaod a font is as easy as searching the name and add "download" in google.<br>
Downlaod links:
- xkcd: https://github.com/ipython/xkcd-font/blob/master/xkcd-script/font/xkcd-script.ttf
- Humor Sans: https://github.com/shreyankg/xkcd-desktop/blob/master/Humor-Sans.ttf
- Comic Sans MS: https://www.wfonts.com/download/data/2014/06/05/comic-sans-ms/comic-sans-ms.zip
To get the path to the fonts directory of the system,a s well as getting a list of available fonts, you cand o the following:
``` Python
import matplotlib.font_manager
paths = matplotlib.font_manager.findSystemFonts(fontpaths=None, fontext='ttf')
names = [path.split('/')[-1].split('.')[0] for path in paths]
names
```
once you have the path, and the fonts. you can install the fonts, and check again if the fonts are available (using the above code snippet).
But you will probably still get the same warning as before. So rebult the matplotlib font cache and (hopefully) the warning is gone and your figure will be using the font of your choice. Rebuilt as follow:
``` python
import matplotlib
matplotlib.font_manager._rebuild()
```
```
import matplotlib.font_manager
paths = matplotlib.font_manager.findSystemFonts(fontpaths=None, fontext='ttf')
names = [path.split('/')[-1].split('.')[0] for path in paths]
# names
"Humor-Sans" in names
matplotlib.font_manager._rebuild()
matplotlib.font_manager._rebuild
```
### References:
- https://www.datacamp.com/courses/introduction-to-data-visualization-with-python (first two blocks)
- https://matplotlib.org/gallery.html
- https://github.com/matplotlib/AnatomyOfMatplotlib
- https://github.com/jbmouret/matplotlib_for_papers
- https://jakevdp.github.io/blog/2013/07/10/XKCD-plots-in-matplotlib/
- https://jakevdp.github.io/blog/2012/10/07/xkcd-style-plots-in-matplotlib/
- the [cheat sheet](https://s3.amazonaws.com/assets.datacamp.com/blog_assets/Python_Matplotlib_Cheat_Sheet.pdf)
| github_jupyter |
#### This notebook is used to visualise and compare the results of the experiments conducted in this paper. This notebook specifically illustrates the said results for the credibility threshold of 0.01.
The results are stored in the 'results' folder. We had parallelized the process of running the experiments into batches of 4 (4900 tweets divided into 4 contiguous groups). Hence, the file names have the following nomenclature : _rev2-cold-tau_threshold-attack_typetweetIndexStart_to_tweetIndexEnd.pickle_
```
import pickle
import matplotlib.pyplot as plt
from collections import Counter
with open('results\\rev2-cold-001-replacement0_to_1225.pickle', 'rb') as handle:
replacement0_to_1225_001 = pickle.load(handle)
with open('results\\rev2-cold-001-default0_to_1225.pickle', 'rb') as handle:
default0_to_1225_001 = pickle.load(handle)
with open('results\\rev2-cold-001-insertion0_to_1225.pickle', 'rb') as handle:
insertion0_to_1225_001 = pickle.load(handle)
with open('results\\rev2-cold-001-replacement1225_to_2450.pickle', 'rb') as handle:
replacement1225_to_2450_001 = pickle.load(handle)
with open('results\\rev2-cold-001-default1225_to_2450.pickle', 'rb') as handle:
default1225_to_2450_001 = pickle.load(handle)
with open('results\\rev2-cold-001-insertion1225_to_2450.pickle', 'rb') as handle:
insertion1225_to_2450_001 = pickle.load(handle)
with open('results\\rev2-cold-001-replacement2450_to_3675.pickle', 'rb') as handle:
replacement2450_to_3675_001 = pickle.load(handle)
with open('results\\rev2-cold-001-default2450_to_3675.pickle', 'rb') as handle:
default2450_to_3675_001 = pickle.load(handle)
with open('results\\rev2-cold-001-insertion2450_to_3675.pickle', 'rb') as handle:
insertion2450_to_3675_001 = pickle.load(handle)
with open('results\\rev2-cold-001-replacement3675_to_4900.pickle', 'rb') as handle:
replacement3675_to_4900_001 = pickle.load(handle)
with open('results\\rev2-cold-001-default3675_to_4900.pickle', 'rb') as handle:
default3675_to_4900_001 = pickle.load(handle)
with open('results\\rev2-cold-001-insertion3675_to_4900.pickle', 'rb') as handle:
insertion3675_to_4900_001 = pickle.load(handle)
default_lst = [default0_to_1225_001,default1225_to_2450_001,default2450_to_3675_001,default3675_to_4900_001]
default = set().union(*default_lst)
print(len(default))
```
We combine the 4 dictionaries of the insertion attack type results into 1 dictionary and visualize the result.
```
insertion = {**insertion0_to_1225_001,**insertion1225_to_2450_001,**insertion2450_to_3675_001,**insertion3675_to_4900_001}
plt.hist([val for val in insertion.values()],bins=10) # density=False would make counts
plt.ylabel('Frequency')
plt.xlabel('Data');
plt.title('Insertion Plot')
plt.show()
counter1 = Counter(insertion.values())
print(counter1)
print(sum(counter1.values()))
import seaborn as sns
hist = sns.histplot(data=[val for val in insertion.values()],bins=10,color='blue',discrete=True)
hist.set_xlabel("Number of Fake Accounts Needed",fontsize=16)
hist.set_ylabel("Number of Tweets",fontsize=16)
hist.tick_params(labelsize=16)
hist.set_yticks([0,500,1000,1500,2000,2500,3000,3500,4000]) # <--- set the ticks first
hist.set_yticklabels([0,500,100,1500,2000,2500,3000,3500,4000])
hist.set_xticks([1,2,3,4,5,6,7,8,9,10]) # <--- set the ticks first
hist.set_xticklabels([1,2,3,4,5,6,7,8,9,">=10"])
plt.ylim(0,4000)
```
We combine the 4 dictionaries of the replacement attack type results into 1 dictionary and visualize the result.
```
replacement = {**replacement0_to_1225_001,**replacement1225_to_2450_001,**replacement2450_to_3675_001,**replacement3675_to_4900_001}
plt.hist([val for val in replacement.values()],bins=10) # density=False would make counts
plt.ylabel('Frequency')
plt.xlabel('Data');
plt.title('Replacement Plot')
plt.show()
counter2 = Counter(replacement.values())
print(counter2)
print(sum(counter2.values()))
import seaborn as sns
sns.color_palette("tab10")
hist = sns.histplot(data=[val for val in replacement.values()],bins=10,color='blue',discrete=True)
hist.set_xlabel("Number of Fake Accounts Needed",fontsize=16)
hist.set_ylabel("Number of Tweets",fontsize=16)
hist.tick_params(labelsize=16)
hist.set_yticks([0,10,20,30,40,50,60,70]) # <--- set the ticks first
hist.set_yticklabels([0,10,20,30,40,50,60,70])
hist.set_xticks([1,2,3,4,5,6,7,8,9,10]) # <--- set the ticks first
hist.set_xticklabels([1,2,3,4,5,6,7,8,9,'>=10'])
plt.ylim(0,75)
```
We similarly load results for the birdwatch data and visualise it. The results for the birdwatch data are obtained by running the birdwatch-metric-process-tweets.py script.
```
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm
from time import time
import plotly.express as px
import plotly.graph_objects as go
import numpy as np
import random,string
import math
from collections import Counter
from scipy import stats
from collections import defaultdict
with open('results/bw-insertion.pickle', 'rb') as handle:
insertion_bw = pickle.load(handle)
with open('results/bw-replacement.pickle', 'rb') as handle:
replacement_bw = pickle.load(handle)
with open('results/bw-default.pickle', 'rb') as handle:
default_bw = pickle.load(handle)
plt.hist([val for val in insertion_bw.values()],bins=10,ec='black',color='green') # density=False would make counts
plt.ylabel('Number of Tweets')
plt.xlabel('Number of Fake Accounts')
plt.title('Insertion Plot')
plt.show()
counter1 = Counter(insertion_bw.values())
print(counter1)
print(sum(counter1.values()))
plt.hist([val for val in replacement_bw.values()],bins=10) # density=False would make counts
plt.ylabel('Number of Tweets')
plt.xlabel('Number of Fake Accounts')
plt.title('Replacement Plot')
plt.show()
counter2 = Counter(replacement_bw.values())
print(counter2)
print(sum(counter2.values()))
import seaborn as sns
sns.histplot(data=[val for val in insertion_bw.values()],bins=10,color='blue',kde=True)
plt.ylabel('Number of Tweets')
plt.xlabel('Number of Fake Accounts')
import seaborn as sns
sns.histplot(data=[val for val in replacement_bw.values()],bins=10,color='blue',kde=True)
plt.ylabel('Number of Tweets')
plt.xlabel('Number of Fake Accounts')
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-deep')
x = [val for val in insertion_bw.values()]
y = [val for val in insertion.values()]
bins = np.linspace(0,10,10)
fig, ax1 = plt.subplots()
ax1.hist([x, y], color=['gray','red'],label=['Birdwatch', 'HawkEye'])
ax1.set_xlabel("Number of Fake Accounts Needed",fontsize=12)
ax1.set_ylabel("Number of Tweets",fontsize=14)
plt.legend(loc='upper right')
plt.xticks([1.4,2.3,3.3,4.1,5,6,6.9,7.8,8.7,9.5],[1,2,3,4,5,6,7,8,9,10])
plt.locator_params(axis="x", nbins=10)
plt.locator_params(axis="y", nbins=20)
plt.show()
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-deep')
x = [val for val in replacement_bw.values()]
y = [val for val in replacement.values()]
bins = np.linspace(0,10,10)
fig, ax1 = plt.subplots()
ax1.hist([x, y], color=['gray','blue'],label=['Birdwatch', 'HawkEye'])
ax1.set_xlabel("Number of Fake Accounts Needed",fontsize=16)
ax1.set_ylabel("Number of Tweets",fontsize=16)
plt.legend(loc='upper left',fontsize=14)
plt.xticks([1.4,2.3,3.3,4.1,5,6,6.9,7.8,8.7,9.5],[1,2,3,4,5,6,7,8,9,">=10"])
plt.xticks(fontsize=16)
hist.set_yticks([10,20,30,40,50,60,70]) # <--- set the ticks first
hist.set_yticklabels([10,20,30,40,50,60,70])
plt.yticks(fontsize=16)
plt.locator_params(axis="x", nbins=10)
plt.show()
```
We perform a two tailed t-test to compare the birdwatch and hawkeye results.
```
insertion_common_keys = list(insertion.keys() & insertion_bw.keys())
replacement_common_keys = list(replacement.keys() & replacement_bw.keys())
insertion_bw_analysis = [insertion_bw[key] for key in insertion_common_keys]
insertion_rev2_analysis = [insertion[key] for key in insertion_common_keys]
replacement_bw_analysis = [replacement_bw[key] for key in replacement_common_keys]
replacement_rev2_analysis = [replacement[key] for key in replacement_common_keys]
print(stats.ttest_rel(insertion_rev2_analysis,insertion_bw_analysis,alternative='greater'))
print(stats.ttest_rel(replacement_rev2_analysis,replacement_bw_analysis,alternative='greater'))
```
| github_jupyter |
# Project: The Movie DB (TMDB) Data Analysis
## Table of Contents
<ul>
<li><a href="#intro">Introduction</a></li>
<li><a href="#wrangling">Data Wrangling</a></li>
<li><a href="#eda">Exploratory Data Analysis</a></li>
<li><a href="#conclusions">Conclusions</a></li>
</ul>
<a id='intro'></a>
## Introduction
>I will be investigating The Movie DB (TMDB) data set which is a popular tool used to obtain an array of information on over 10,000 films released from 1960 to 2015. I will analyse the active production companies, genre popularity and genre profitability.
>In particular, the questions I will be exploring include:
> 1) Which Primary Genres are the most popular (i.e. highest average popularity score)?
> 2) Which production companies have produced the most films since the inception of the data set and what is the percentage breakdown of major production company activity?
> 3) Which Primary Genre had the highest average profit (inflation-adjusted) and which film was the most profitable overall (inflation-adjusted)?
```
# Use this cell to set up import statements for all of the packages that you
# plan to use.
import pandas as pd
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
# Remember to include a 'magic word' so that your visualizations are plotted
# inline with the notebook. See this page for more:
# http://ipython.readthedocs.io/en/stable/interactive/magics.html
```
<a id='wrangling'></a>
## Data Wrangling
We will read the data, have a quick look at the first few rows and get a picture of the data types and the number of data points.
### General Properties
```
# Load your data and print out a few lines. Perform operations to inspect data
# types and look for instances of missing or possibly errant data.
df = pd.read_csv('tmdb-movies.csv')
df.head()
df.info()
```
### Data Cleaning
>First we will check for any duplicate rows (i.e. each column holds the same value) and remove them.
>Subsequently, we will review the data types to ensure that the columns are holding appropriate data types (i.e. string types are displayed as objects and numerical values are either integer or floats.)
>As we will be conducting further analysis and exploration specifically on Genres, it will be important to clean the data with respect to this column (remove nulls and add a Primary Genre column after separating the multivalues)
```
# After discussing the structure of the data and any problems that need to be
# cleaned, perform those cleaning steps in the second part of this section.
df.duplicated().sum() #Check number of duplicates
df.drop_duplicates(inplace = True) #There is only one duplicate row so we have dropped this from the data set.
df.dtypes #Check data types
```
>The data types appear to hold appropriate data types and do not require any conversions.
>Now we will look at the genres column. We will need to drop any nulls from the dataframe.
```
df['genres'].isnull().sum() #count null genres (i.e. genre not assigned)
df = df.dropna(subset=['genres']) #drop NaN in genres column from df
df['genres'].isnull().sum() #Check 0 nulls for genre
```
>We should separate the genres and print them in a list to ensure they are readable for further analysis.
```
genre_com = list(map(str,(df['genres']))) #List of genre combinations
genre = []
for i in genre_com:
split_genre = list(map(str, i.split('|'))) #splitting genres for movies with multi value genres
for j in split_genre:
if j not in genre:
genre.append(j)#add single all single genre movies to df_genre
# printing list of seperated genres.
print(genre)
```
> There are several multi-value elements which appear in the genres column. After reviewing the data, it appears that the first element listed in the multi-value rows under genres represents the "Primary Genre" of the film. I have made the decision to add a new column called "p_genre" to conduct a more targeted analysis on genres.
>I feel this allows the analysis on genre popularity and genre profitability to be targeted towards the film's primary genre and gives less weight to a more common reappearing genre (such as Action, Drama or Comedy which appear very frequently as secondary genres).
>Another potential issue which may arise is with closely linked genres such as Thriller/Mystery/Crime where there is potential cross-over in the genre specification. Should these genres be separated as the genres may share highly similar features?
>I feel that the genre specification can be considered as a limitation of the analysis with either applying approach and has the potential to skew the resulting conclusions.
```
df_gen = df.genres.str.split('|') #splitting genres out of dataframe
df.loc[:,'p_genre'] = df_gen.str[0] #adding a new Primary Genre column to the dataframe
df.groupby('p_genre')['id'].nunique() #Checking that the data sample is reasonable for analysis
df.head() #checking if p_genre has been added appropriately
```
>As we want to analyse profit, we will need to create a new column. A better measure to compare the profitabilty of films over timeframe from 1960 to 2015 is inflation_adjusted profit which subtracts the 2015 equivalent inflation-adjusted budget from the 2015 equivalent inflation-adjusted revenue.
>Here I made a new 'profit_adj' column
```
df.loc[:,'profit_adj'] = df['revenue_adj'] - df['budget_adj'];
df.head()
```
> As we will be conducting further exploratory analysis on production companies, it might be worthwhile to get a picture of how many production companies are presented in the list.
> Here I created a list of the separated production companies. It appears that there a lot more than I initially expected and highlights the presence of thousands of smaller film producers (independent). This will require further consideration in our exploratory analysis.
```
# Obtaining a list of production companies
prd_details = list(map(str,(df['production_companies'])))
prd = []
for i in prd_details:
split_prd = list(map(str, i.split('|')))
for j in split_prd:
if j not in prd:
prd.append(j)
# count elements in list of seperated genres.
len(prd)
```
<a id='eda'></a>
## Exploratory Data Analysis
We will now compute statistics and create visualizations with the goal of addressing the research questions posed in the Introduction section. We will apply a systematic approach, looking at one variable at a time, and then following it up by looking at relationships between variables.
### Question 1: Which Primary Genres are the most popular (i.e. highest average popularity score)?
```
# Use this, and more code cells, to explore your data. Don't forget to add
# Markdown cells to document your observations and findings.
pg_pop = df.groupby('p_genre')['popularity'].mean()
popt = pg_pop.plot.bar();
popt.set_xlabel("Primary Genre");
popt.set_ylabel("Mean Popularity");
```
> From the barplot above it can be seen that the Adventure genre appears to have the highest average popularity score Primary Genre from the data set. Please note that this is a more targeted analysis of the primary genre of a film as opposed to analysing the multiple of genres listed per film.
### Question 2: Which production companies have produced the most films since the inception of the data set and what is the percentage breakdown of major production company activity?
I made an assumption here that a 'major production company' is one that has produced at least 100 films in the data set.
```
split_prd = df['production_companies'].str.cat(sep = '|') #separating out "|" in the production_companies column
split_prd = pd.Series(split_prd.split('|')) #converting split data to a series so we can filter
prd_cnt = split_prd.value_counts(ascending = False) #sorting list in descending order by count.
m_prd_c = prd_cnt[prd_cnt >= 100] #defining major production companies as production companies that have produced at least 100 films in the dataset.
m_prd_c
```
>It can be seen that Universal Pictures has produced the most films over from 1960 to 2015, closely followed by Warner Bros. It should be noted that the top 3 (Universal, Warner Bros. and Paramount) have been significantly more active movie producers than the rest of the data set.
```
label = list(map(str,m_prd_c[0:8].keys()))
label.append('Others')
mpc = m_prd_c[0:8]
sum = 0
for i in m_prd_c[8:]:
sum += i
mpc['sum'] = sum
fig1, ax1 = plt.subplots()
ax1.pie(mpc,labels = label, autopct = '%1.1f%%', startangle = 90)
ax1.axis('equal')
plt.title("Percentage breakdown of major production companies")
plt.show()
```
>Above I have plotted a pie chart with the top 8 major production companies (i.e. produced more than 100 films since 1960) and grouped the remaining producers together in Others. It can be seen that the production companies are broken down in clusters with the top 3 producers ahead by a significant margin from the rest and the remaining producers are quite closely grouped.
### Question 3: Which Primary Genre had the highest average profit (inflation-adjusted) and which film had the highest profit overall (inflation-adjusted)?
```
# Continue to explore the data to address your additional research
# questions. Add more headers as needed if you have more questions to
# investigate.
pg_pro = df.groupby('p_genre')['profit_adj'].mean()
pg_pr = pg_pro.plot.bar();
pg_pr.set_xlabel("Primary Genre");
pg_pr.set_ylabel("Mean Profit");
```
>The most profitable (inflation-adjusted) Primary Genre in the data set is Adventure by a significant margin.
```
def max_obs(col_name):
#maximum
#taking the index (row) of the highest number in any specified column, passing col_name
max_pt = df[col_name].idxmax()
#calling by index number above,storing row details to a variable, max_data
max_data = pd.DataFrame(df.loc[max_pt])
return max_data
max_obs('profit_adj')
```
>The most profitable (inflation-adjusted) film in the data set was the original Star Wars released in 1977, generating an infl-adj. profit of $2.75b
<a id='conclusions'></a>
## Conclusions
>From our data wrangling we have cleaned the data, modified the genre column to specify and create a Primary Genre column, added an inflation-adjusted profit column and split out the production companies to conduct our EDA and answer the questions around genre, profitability and production companies.
>In our EDA we found that the Adventure genre as a primary genre had the highest average popularity score. This was visualised in a bar plot. I chose to use a single Primary Genre to ensure that the analysis was more targeted towards the main genre that a film depicts (rather than the popularity of common genres such as Action/Comedy/Drama being skewed towards the overall mean popularity) and receives the full weight of the popularity score. In contrast, the limitation to using the Primary Genre is that it may not adequately capture the popularity for movies with clear multiple genres.
>Additionally, we defined a major production company as production companies that have produced over 100 films from 1960 to 2015 (the timeframe of the data set). We found that Universal Pictures was the most active production company, closely followed by Warner Bros and this was presented in a pie chart. I was also surprised by the high presence of small production companies which produced a very few number of films.
> Finally we found that the most profitable (inflation-adjusted) Primary Genre was Adventure and the most profitable (inflation-adjusted) film was Star Wars. I chose to use the inflation-adjusted measure to adequately compare the budget and revenue figures from different time periods.
```
from subprocess import call
call(['python', '-m', 'nbconvert', 'Investigate_a_Dataset.ipynb'])
```
| github_jupyter |
# Structured Sparsity Example
It is well-known that neural networks have redundant filters, thus one would like
to reduce the number of filters, then the number of output feature maps
in convolutions and the number of output dimensions in affine are reduced,
which not only reduces the memory space but also the computational cost.
For example, in 2D Convolution case, one can get a slim network by reducing unnecessary 3D kernels,
$w_m \in \mathcal{R}^{N \times K_h \times K_w}$ where
$w_m$ denotes one convolution 3d filter,
$N$ is the number of input maps,
$K_h$ is the kernel height, and
$K_w$ is the kernel width.
This can be achieved by sparsing the filters. It is induced by using
`Structured Sparsity Learning` called `SSL` in the following paper,
```
WeiWen, et al.,
"Learning Structured Sparsity in Deep Neural Networks",
https://arxiv.org/abs/1608.03665
```
Literally, `SSL` includes the filters which have many zeros elements but it has structure,
in this case, $w_m$ might become zero, thus one can ignore such filters. Thus,
one get a slim network.
Mathematically, there are two regularization to induce sparsity; $R_f(W)$ and $R_c(W)$, Each of which are denoted by
$$R_f(W) = \sum_{m=1}^{M}\sqrt{\sum_{n,k_{h},k_{w}=1}^{N,K_{h},K_{w}}w_{m,n,k_{h}, k{w}}^{2}},$$
$$R_c(W) = \sum_{n=1}^{N}\sqrt{\sum_{m,k_{h},k_{w}=1}^{M,K_{h},K_{w}}w_{m,n,k_{h}, k{w}}^{2}} .$$
where $R_f(W)$ induces the *filter-wise* sparsity and $R_c(W)$ does the *channel-wise* sparsity. Note that $R_c(W)$ also induces the *filter-wise* sparsity since in Neural Network context, an input map is the result of a preceding layer so that including the *channel-wise* sparsity corresponds to the *filter-wise* sparsity in the preceding layer.
Usually, $R_f(W)$ and $R_c(W)$ are used together for each layer,
$$\lambda_f \sum_{l=1}^{L} R_f(W^{l}) + \lambda_c \sum_{l=1}^{L} R_c(W^{l}),$$
where $\lambda_f$ and $\lambda_c$ are the hyper parameters.
One follows the steps for using `SSL`,
1. Train a reference network with Structured Sparsity induced regularization
2. Finetune a reference network without unnecessary filters
For using this example, first train a network,
```sh
python classification.py -c "cudnn" \
--monitor-path "monitor.filter.lambda-5e4" \
--model-save-path "monitor.filter.lambda-5e4" \
--filter-decay 5e-4 \
--channel-decay 5e-4 \
-d 0
```
Then, finetune that network,
```sh
python finetuning.py -c "cudnn" \
--monitor-path "monitor.finetune.filter.lambda-5e4.rrate-025" \
--model-save-path "monitor.finetune.filter.lambda-5e4.rrate-025" \
--model-load-path "monitor.filter.lambda-5e4/${the best result}.h5" \
--reduction-rate 0.25 \
--val-interval 1000 \
-d 0
```
## References
1. Wen Wei, Wu Chunpeng, Wang Yandan, Chen Yiran, and Li Hai "Learning Structured Sparsity in Deep Neural Networks", arXiv:1608.03665
| github_jupyter |
**Initialization**
* I use these 3 lines of code on top of my each Notebooks because it will help to prevent any problems while reloading and reworking on a same Project or Problem. And the third line of code helps to make visualization within the Notebook.
```
#@ Initialization:
%reload_ext autoreload
%autoreload 2
%matplotlib inline
```
**Downloading the Dependencies**
* I have downloaded all the Libraries and Dependencies required for this Project in one particular cell.
```
#@ Downloading the Libraries and Dependencies:
import os, glob
from random import shuffle
from IPython.display import display
import numpy as np # Module to work with Arrays.
from keras.preprocessing import sequence # Module to handle Padding Input.
from keras.models import Sequential # Base Keras Neural Network Model.
from keras.layers import Dense, Dropout, Flatten # Layers Objects to pile into Model.
from keras.layers import LSTM # Convolutional Layer and MaxPooling.
from nltk.tokenize import TreebankWordTokenizer # Module for Tokenization.
from gensim.models.keyedvectors import KeyedVectors
```
**Getting the Data**
* I have used Google Colab for this Project so the process of downloading and reading the Data might be different in other platforms. I have used [**Large Moview Review Dataset**](https://ai.stanford.edu/~amaas/data/sentiment/) for this Project. This is a dataset for binary sentiment classification containing substantially more data. The Dataset has a set of 25,000 highly polar movie reviews for training and 25,000 for testing. There is additional unlabeled data for use as well. Raw text and already processed bag of words formats are provided.
```
#@ Getting the Data:
def preprocess_data(filepath):
positive_path = os.path.join(filepath, "pos")
negative_path = os.path.join(filepath, "neg")
pos_label = 1
neg_label = 0
dataset = []
for filename in glob.glob(os.path.join(positive_path, '*.txt')): # Positive Sentiment Dataset.
with open(filename, "r") as f:
dataset.append((pos_label, f.read()))
for filename in glob.glob(os.path.join(negative_path, '*.txt')): # Negative Sentiment Dataset.
with open(filename, "r") as f:
dataset.append((neg_label, f.read()))
shuffle(dataset) # Shuffling the Dataset.
return dataset
```
**Processing the Dataset**
* I have manually downloaded the Dataset from [**Large Moview Review Dataset**](https://ai.stanford.edu/~amaas/data/sentiment/). I have used the small subset of Data.
```
#@ Processing the Dataset:
PATH = "/content/drive/My Drive/Colab Notebooks/Data/Smalltrain" # Path to the Dataset.
dataset = preprocess_data(PATH) # Processing the Dataset.
#@ Inspecting the Dataset:
dataset[:3] # Inspecting the Dataset.
```
**Tokenization and Vectorization**
* The next step is to perform the Tokenization and Vectorization of the Dataset. I will use Google news pretrained Model Vectors for the process of Vectorization. The Google News Word2vec Vocabulary includes some stopwords as well.
```
#@ Tokenization and Vectorization:
# !wget -c "https://s3.amazonaws.com/dl4j-distribution/GoogleNews-vectors-negative300.bin.gz" # Pretrained Word2vec Model.
word_vectors = KeyedVectors.load_word2vec_format("/content/GoogleNews-vectors-negative300.bin.gz", # Word2vec Model Vectors.
binary=True, limit=100000)
#@ Function for Tokenization and Vectorization:
def tokenize_and_vectorize(dataset):
tokenizer = TreebankWordTokenizer() # Instantiating the Tokenizer.
vectorized_data = []
for sample in dataset:
tokens = tokenizer.tokenize(sample[1]) # Process for Tokenization.
sample_vecs = []
for token in tokens:
try:
sample_vecs.append(word_vectors[token]) # Process for Vectorization.
except KeyError:
pass
vectorized_data.append(sample_vecs)
return vectorized_data # Returning the Vectorized Data.
#@ Function for Collecting the Target Labels:
def collect_expected(dataset):
""" Collecting the Target Labels: 0 for Negative Review and 1 for Positive Review. """
expected=[]
for sample in dataset:
expected.append(sample[0])
return expected
#@ Tokenization and Vectorization:
vectorized_data = tokenize_and_vectorize(dataset)
expected = collect_expected(dataset)
```
**Splitting into Training and Testing.**
* Now, I will split the above obtained Dataset into Training set and a Test set. I will split the Dataset into 80% for Training and 20% for Test set. The next code will bucket the Data into Training set X train along with correct labels y train and similarly into Test set X test along with correct labels y test.
```
#@ Splitting the Dataset into Training set and Test set:
split_part = int(len(vectorized_data) * 0.8)
#@ Training set:
X_train = vectorized_data[:split_part]
y_train = expected[:split_part]
#@ Test set:
X_test = vectorized_data[split_part:]
y_test = expected[split_part:]
```
### **Long Short Term Memory**
* Long Short Term Memory or LSTM is an Artificial Recurrent Neural Network or RNN architecture used in the field of Deep Learning. Unlike standard Feedforward Neural Networks, LSTM has Feedback connections. It can not only process single data points, but also entire sequences of data such as Speech or Video.
```
#@ Parameters of LSTM Neural Network:
maxlen = 500 # Maximum review length.
batch_size = 32 # Number of samples shown to the network before updating the weights.
embedding_dims = 300 # Length of token vectors for passing in RNN.
epochs = 10 # Number of times for passing the training dataset.
num_neurons = 50
```
**Padding and Truncating the Sequence**
* **Keras** has the preprocessing helper method called pad_sequences which is used to pad the input Data. But it works only on the sequence of scalars and sequence of vectors. Now, I will write the helper function to pad the input Data.
```
#@ Padding and Truncating the Token Sequence:
def pad_trunc(data, maxlen):
""" Padding the Dataset with zero Vectors. """
new_data = []
# Creating zeros vectors of length of Word vectors.
zero_vector = []
for _ in range(len(data[0][0])):
zero_vector.append(0.0)
for sample in data:
if len(sample) > maxlen:
temp = sample[:maxlen]
elif len(sample) < maxlen:
temp = sample
# Append the appropriate number of 0 vectors to the list.
additional_elems = maxlen - len(sample)
for _ in range(additional_elems):
temp.append(zero_vector)
else:
temp = sample
new_data.append(temp)
return new_data
#@ Gathering the Truncated and Augmented Data:
X_train = pad_trunc(X_train, maxlen)
X_test = pad_trunc(X_test, maxlen)
#@ Converting the Data into Numpy Arrays:
X_train = np.reshape(X_train, (len(X_train), maxlen, embedding_dims))
y_train = np.array(y_train)
X_test = np.reshape(X_test, (len(X_test), maxlen, embedding_dims))
y_test = np.array(y_test)
#@ Inspecting the shape of the Data:
display(f"Shape of Training Data {X_train.shape, y_train.shape}")
display(f"Shape of Testing Data {X_test.shape, y_test.shape}")
```
**Long Short Term Memory**
* Now, The Dataset is ready to build the Neural Network.
```
#@ Long Short Term Memory or LSTM:
model = Sequential() # Standard Model Definition for Keras.
model.add(LSTM( # Adding the LSTM Layer.
num_neurons, return_sequences=True,
input_shape=(maxlen, embedding_dims)
))
model.add(Dropout(0.2)) # Adding the Dropout Layer.
model.add(Flatten()) # Flatten the output of LSTM.
model.add(Dense(1, activation="sigmoid")) # Output Layer.
#@ Compiling the LSTM Neural Network:
model.compile(
loss="binary_crossentropy",
optimizer="rmsprop",
metrics=["accuracy"]
)
#@ Training the LSTM Neural Network:
model.fit(
X_train, y_train, # Training Dataset.
batch_size=batch_size,
epochs=epochs,
validation_data=(X_test, y_test) # Validation Dataset.
)
#@ Inspecting the Summary of the Model:
print("\n")
model.summary() # Summary of the Model.
```
**Saving the LSTM Model**
```
#@ Saving the Recurrent Neural Network:
model_structure = model.to_json()
with open("lstm.json", "w") as json_file:
json_file.write(model_structure)
model.save_weights("lstm.h5")
print("Model saved!!")
```
**Model Evaluation**
* Now, I have trained a Model. I will make a sentence with Positive Sentiment and I will predict the Sentiment of the sentence using the Neural Network.
```
#@ Model Evaluation:
sample_1 = """ I hate that the dismal weather had me down for so long, \
when will it break! Ugh, when does happiness return? The sun is \
blinding and the puffy clouds are too thin. I can't wait for the weekend."""
#@ Making Predictions:
vec_list = tokenize_and_vectorize([(1, sample_1)])
test_vec_list = pad_trunc(vec_list, maxlen)
test_vec = np.reshape(test_vec_list, (len(test_vec_list), maxlen, embedding_dims))
#@ Inspecting the Prediction:
f"The predicted sentiment by the Model is: {model.predict_classes(test_vec)}"
```
**Optimizing the Vector Size**
* Padding and Truncating each sample to 400 Tokens was important for Convolutional Neural Nets so that the filters could scan a vector with a consistent length.
```
#@ Optimizing the Vector Size:
def test_len(data, maxlen):
total_len = truncated = exact = padded = 0
for sample in data:
total_len = len(sample)
if len(sample) > maxlen:
truncated += 1
elif len(sample) < maxlen:
padded += 1
else:
exact += 1
print(f"Padded: {padded}")
print(f"Equal: {exact}")
print(f"Truncated: {truncated}")
print(f"Average length: {total_len/len(data)}")
#@ Applying in the Dataset:
test_len(vectorized_data, 500)
```
**Optimized Long Short Term Memory**
```
#@ Parameters of LSTM Neural Network:
maxlen = 200 # Maximum review length.
batch_size = 32 # Number of samples shown to the network before updating the weights.
embedding_dims = 300 # Length of token vectors for passing in RNN.
epochs = 10 # Number of times for passing the training dataset.
num_neurons = 50
#@ Gathering the Truncated and Augmented Data:
X_train = pad_trunc(X_train, maxlen)
X_test = pad_trunc(X_test, maxlen)
#@ Converting the Data into Numpy Arrays:
X_train = np.reshape(X_train, (len(X_train), maxlen, embedding_dims))
y_train = np.array(y_train)
X_test = np.reshape(X_test, (len(X_test), maxlen, embedding_dims))
y_test = np.array(y_test)
#@ Long Short Term Memory or LSTM:
model = Sequential() # Standard Model Definition for Keras.
model.add(LSTM( # Adding the LSTM Layer.
num_neurons, return_sequences=True,
input_shape=(maxlen, embedding_dims)
))
model.add(Dropout(0.2)) # Adding the Dropout Layer.
model.add(Flatten()) # Flatten the output of LSTM.
model.add(Dense(1, activation="sigmoid")) # Output Layer.
#@ Compiling the LSTM Neural Network:
model.compile(
loss="binary_crossentropy",
optimizer="rmsprop",
metrics=["accuracy"]
)
#@ Training the LSTM Neural Network:
model.fit(
X_train, y_train, # Training Dataset.
batch_size=batch_size,
epochs=epochs,
validation_data=(X_test, y_test) # Validation Dataset.
)
#@ Inspecting the Summary of the Model:
print("\n")
model.summary() # Summary of the Model.
```
| github_jupyter |
# Random Forest Classification
## Importing the libraries
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
## Importing the dataset
```
train = pd.read_csv("train.csv", engine='python',nrows=100000)
features = [col for col in list(train.columns) if 'feature' in col]
train = train[train['weight'] != 0]
train['action'] = (train['resp'].values > 0).astype(int)
train = train.dropna()
X = train.loc[:, features]
y = train.loc[:, 'action']
del train
X = np.array(X)
y = np.array(y)
print(len(X), len(y))
```
## Splitting the dataset into the Training set and Test set
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1, random_state = 0)
```
## Feature Scaling
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
```
## Training the Random Forest Classification model on the Training set
```
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(n_estimators = 20, criterion = 'entropy', random_state = 0)
classifier.fit(X_train, y_train)
```
## Predicting a new result
```
# print(classifier.predict(sc.transform([[30,87000]])))
```
## Predicting the Test set results
```
y_pred = classifier.predict(X_test)
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
```
## Making the Confusion Matrix
```
from sklearn.metrics import confusion_matrix, accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
accuracy_score(y_test, y_pred)
```
## Visualising the Training set results
```
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(X_train), y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(X1, X2, classifier.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Random Forest Classification (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
## Visualising the Test set results
```
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(X_test), y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(X1, X2, classifier.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Random Forest Classification (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
| github_jupyter |
# Convert Tensorflow model to ONNX
Tensorflow and ONNX both define their own graph format to represent to model. You can use [tensorflow-onnx](https://github.com/onnx/tensorflow-onnx "Title") to export a Tensorflow model to ONNX.
We divide the guide into 2 parts: part 1 covers basic conversion and part 2 advanced topics. The following content will be covered in order:
1. Procedures to convert tensorflow model
- get tensorflow model
- convert to ONNX
- validate
2. Key conceptions
- opset
- data format
## Step 1 - Get Tensorflow model
Tensorflow uses several file formats to represent a model, such as checkpoint files, graph with weight(called `frozen graph` next) and saved_model, and it has APIs to generate these files, you can find the code snippets in the script [tensorflow_to_onnx_example.py](./assets/tensorflow_to_onnx_example.py)
And `tensorflow-onnx` can accept all the three formats to represent a Tensorflow model, **the format "saved_model" should be the preference** since it doesn't require the user to specify input and output names of graph.
we will cover it in this section and cover the other two in the last section. And also, you could get more detail from `tensorflow-onnx`'s [README](https://github.com/onnx/tensorflow-onnx/blob/master/README.md "Title") file.
```
import os
import shutil
import tensorflow as tf
from assets.tensorflow_to_onnx_example import create_and_train_mnist
def save_model_to_saved_model(sess, input_tensor, output_tensor):
from tensorflow.saved_model import simple_save
save_path = r"./output/saved_model"
if os.path.exists(save_path):
shutil.rmtree(save_path)
simple_save(sess, save_path, {input_tensor.name: input_tensor}, {output_tensor.name: output_tensor})
print("please wait for a while, because the script will train MNIST from scratch")
tf.reset_default_graph()
sess_tf, saver, input_tensor, output_tensor = create_and_train_mnist()
print("save tensorflow in format \"saved_model\"")
save_model_to_saved_model(sess_tf, input_tensor, output_tensor)
```
## Step 2 - Convert to ONNX
`tensorflow-onnx` has several entries to convert tensorflow model with different tensorflow formats, this section will cover "saved_model" only, "frozen graph" and "checkpoint" will be covered in [part 2](./TensorflowToOnnx-2.ipynb).
Also, `tensorflow-onnx` has exported related python APIs, so users can call them directly on their script instead of command line, also the detail will be covered in [part 2](./TensorflowToOnnx-2.ipynb).
```
# generating mnist.onnx using saved_model
!python -m tf2onnx.convert \
--saved-model ./output/saved_model \
--output ./output/mnist1.onnx \
--opset 7
```
## Step 3 - Validate
There are several framework can run model in ONNX format, here [ONNXRuntime](https://github.com/microsoft/onnxruntime "Title") , opensourced by `Microsoft`, is used to make sure the generated ONNX graph behaves well.
The input "image.npz" is an image of handwritten "7", so the expected classification result of model should be "7".
```
import numpy as np
import onnxruntime as ort
img = np.load("./assets/image.npz").reshape([1, 784])
sess_ort = ort.InferenceSession("./output/mnist1.onnx")
res = sess_ort.run(output_names=[output_tensor.name], input_feed={input_tensor.name: img})
print("the expected result is \"7\"")
print("the digit is classified as \"%s\" in ONNXRruntime"%np.argmax(res))
```
## Key conceptions
This command line should work for most tensorflow models if they are available a saved_model. In some cases you might encounter issues that require extra options.
The most important concept is "**opset** version": ONNX is an evolving standard, for example it will add more new operations and enhance existing operations, so different opset version will contain different operations and operations may have different behavior. The default version "tensorflow-onnx" used is 7 and ONNX supports version 10 now, so if the conversion failed, you may try different version, by command line option "--opset", to see if it works.
Continue with [part 2](./TensorflowToOnnx-2.ipynb) that explains advanced topics.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.