text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork/Template/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
<h1>Logistic Regression and Bad Initialization Value</h1>
<h2>Objective</h2><ul><li> How bad initialization value can affects the accuracy of model. .</li></ul>
<h2>Table of Contents</h2>
<p>In this lab, you will see what happens when you use the root mean square error cost or total loss function and select a bad initialization value for the parameter values.</p>
<ul>
<li><a href="https://#Makeup_Data">Make Some Data</a></li>
<li><a href="https://#Model_Cost">Create the Model and Cost Function the PyTorch way</a></li>
<li><a href="https://#BGD">Train the Model:Batch Gradient Descent</a></li>
</ul>
<br>
<p>Estimated Time Needed: <strong>30 min</strong></p>
<hr>
<h2>Preparation</h2>
We'll need the following libraries:
```
# Import the libraries we need for this lab
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
import torch
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
```
Helper functions
The class <code>plot_error_surfaces</code> is just to help you visualize the data space and the Parameter space during training and has nothing to do with Pytorch.
```
# Create class for plotting and the function for plotting
class plot_error_surfaces(object):
# Construstor
def __init__(self, w_range, b_range, X, Y, n_samples = 30, go = True):
W = np.linspace(-w_range, w_range, n_samples)
B = np.linspace(-b_range, b_range, n_samples)
w, b = np.meshgrid(W, B)
Z = np.zeros((30, 30))
count1 = 0
self.y = Y.numpy()
self.x = X.numpy()
for w1, b1 in zip(w, b):
count2 = 0
for w2, b2 in zip(w1, b1):
Z[count1, count2] = np.mean((self.y - (1 / (1 + np.exp(-1*w2 * self.x - b2)))) ** 2)
count2 += 1
count1 += 1
self.Z = Z
self.w = w
self.b = b
self.W = []
self.B = []
self.LOSS = []
self.n = 0
if go == True:
plt.figure()
plt.figure(figsize=(7.5, 5))
plt.axes(projection='3d').plot_surface(self.w, self.b, self.Z, rstride=1, cstride=1, cmap='viridis', edgecolor='none')
plt.title('Loss Surface')
plt.xlabel('w')
plt.ylabel('b')
plt.show()
plt.figure()
plt.title('Loss Surface Contour')
plt.xlabel('w')
plt.ylabel('b')
plt.contour(self.w, self.b, self.Z)
plt.show()
# Setter
def set_para_loss(self, model, loss):
self.n = self.n + 1
self.W.append(list(model.parameters())[0].item())
self.B.append(list(model.parameters())[1].item())
self.LOSS.append(loss)
# Plot diagram
def final_plot(self):
ax = plt.axes(projection='3d')
ax.plot_wireframe(self.w, self.b, self.Z)
ax.scatter(self.W, self.B, self.LOSS, c='r', marker='x', s=200, alpha=1)
plt.figure()
plt.contour(self.w, self.b, self.Z)
plt.scatter(self.W, self.B, c='r', marker='x')
plt.xlabel('w')
plt.ylabel('b')
plt.show()
# Plot diagram
def plot_ps(self):
plt.subplot(121)
plt.ylim
plt.plot(self.x, self.y, 'ro', label="training points")
plt.plot(self.x, self.W[-1] * self.x + self.B[-1], label="estimated line")
plt.plot(self.x, 1 / (1 + np.exp(-1 * (self.W[-1] * self.x + self.B[-1]))), label='sigmoid')
plt.xlabel('x')
plt.ylabel('y')
plt.ylim((-0.1, 2))
plt.title('Data Space Iteration: ' + str(self.n))
plt.show()
plt.subplot(122)
plt.contour(self.w, self.b, self.Z)
plt.scatter(self.W, self.B, c='r', marker='x')
plt.title('Loss Surface Contour Iteration' + str(self.n))
plt.xlabel('w')
plt.ylabel('b')
# Plot the diagram
def PlotStuff(X, Y, model, epoch, leg=True):
plt.plot(X.numpy(), model(X).detach().numpy(), label=('epoch ' + str(epoch)))
plt.plot(X.numpy(), Y.numpy(), 'r')
if leg == True:
plt.legend()
else:
pass
```
Set the random seed:
```
# Set random seed
torch.manual_seed(0)
```
<!--Empty Space for separating topics-->
<h2 id="Makeup_Data">Get Some Data </h2>
Create the <code>Data</code> class
```
# Create the data class
class Data(Dataset):
# Constructor
def __init__(self):
self.x = torch.arange(-1, 1, 0.1).view(-1, 1)
self.y = torch.zeros(self.x.shape[0], 1)
self.y[self.x[:, 0] > 0.2] = 1
self.len = self.x.shape[0]
# Getter
def __getitem__(self, index):
return self.x[index], self.y[index]
# Get Length
def __len__(self):
return self.len
```
Make <code>Data</code> object
```
# Create Data object
data_set = Data()
```
<!--Empty Space for separating topics-->
<h2 id="Model_Cost">Create the Model and Total Loss Function (Cost)</h2>
Create a custom module for logistic regression:
```
# Create logistic_regression class
class logistic_regression(nn.Module):
# Constructor
def __init__(self, n_inputs):
super(logistic_regression, self).__init__()
self.linear = nn.Linear(n_inputs, 1)
# Prediction
def forward(self, x):
yhat = torch.sigmoid(self.linear(x))
return yhat
```
Create a logistic regression object or model:
```
# Create the logistic_regression result
model = logistic_regression(1)
```
Replace the random initialized variable values with some predetermined values that will not converge:
```
# Set the weight and bias
model.state_dict() ['linear.weight'].data[0] = torch.tensor([[-5]])
model.state_dict() ['linear.bias'].data[0] = torch.tensor([[-10]])
print("The parameters: ", model.state_dict())
```
Create a <code> plot_error_surfaces</code> object to visualize the data space and the parameter space during training:
```
# Create the plot_error_surfaces object
get_surface = plot_error_surfaces(15, 13, data_set[:][0], data_set[:][1], 30)
```
Define the dataloader, the cost or criterion function, the optimizer:
```
# Create dataloader object, crierion function and optimizer.
trainloader = DataLoader(dataset=data_set, batch_size=3)
criterion_rms = nn.MSELoss()
learning_rate = 2
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
```
<a id="ref2"></a>
<h2 align=center>Train the Model via Batch Gradient Descent </h2>
Train the model
```
# Train the model
def train_model(epochs):
for epoch in range(epochs):
for x, y in trainloader:
yhat = model(x)
loss = criterion_rms(yhat, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
get_surface.set_para_loss(model, loss.tolist())
if epoch % 20 == 0:
get_surface.plot_ps()
train_model(100)
```
Get the actual class of each sample and calculate the accuracy on the test data:
```
# Make the Prediction
yhat = model(data_set.x)
label = yhat > 0.5
print("The accuracy: ", torch.mean((label == data_set.y.type(torch.ByteTensor)).type(torch.float)))
```
Accuracy is 60% compared to 100% in the last lab using a good Initialization value.
<a href="https://dataplatform.cloud.ibm.com/registration/stepone?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01&context=cpdaas&apps=data_science_experience%2Cwatson_machine_learning"><img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork/Template/module%201/images/Watson_Studio.png"/></a>
<!--Empty Space for separating topics-->
<h2>About the Authors:</h2>
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Michelle Carey</a>, <a href="https://www.linkedin.com/in/jiahui-mavis-zhou-a4537814a?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Mavis Zhou</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ----------------------------------------------------------- |
| 2020-09-23 | 2.0 | Shubham | Migrated Lab to Markdown and added to course repo in GitLab |
<hr>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
# Introduction to Tableau
## Everyone Do Tableau Installation
https://public.tableau.com/
- Tableau Public
is a completely free version of the Tableau software that includes the majority of features included within Tableau Desktop.
https://www.tableau.com/products/desktop
- Tableau Desktop.
The main difference between the two versions is in working with multiple types of data sources (SQL databases, for example), and in working with data files that are not shared with the public.
## Activity 1 Loading Exploring Data
- Step 1: open up application

Not only is Tableau able to connect to data files - like CSV, XLS, and JSON - it is also able to connect to a multitude of servers - like MySQL, MongoDB, and Google Cloud.
- Step 2: Select "Excel" from the list of data sources available and load up GlobalSuperstoreOrders2016.xlsx within Tableau Inside Resource Folder.
After the data has been imported into Tableau you can see the individual sheets of data in excel
- Step 3: Once the data has been loaded a preview provided in the main area of the application.


Any and all changes made to the dataset within Tableau will not affect the original dataset. The purpose of Tableau is to create visualizations: manipulating data is not its strong-suit.
- Step 4 Filtering data is very simple, however, as all Tableau users need to do is click on the "Add" button beneath the Filters text in the top-right corner of the application and select what column they would like to filter by.

After selecting which column to filter by, the values to filter are then chosen manually or based upon some kind of condition.

Depending upon the data-type stored within a column, different filters may or may not be available. Selecting a column with a "Date" data-type, for example, allows users to filter rows based upon date ranges.

## Instructor Do Activity 2 - Building Basic Visuals
- Step 1: Once a dataset has been linked to a Tableau workbook, users can navigate into and edit individual worksheets at the bottom of the application.

- Step 2: Creating visualizations in Tableau is nearly identical to creating pivot tables in Excel. Users click and drag the headers of their original dataset into specific fields - Columns, Rows, Filters, etc. - in order to create a chart.

The difference between Dimensions and Measures
- Dimensions are categorical fields that data can be split up by.
- Measures are the metrics or numbers that users would like to analyze.
Drag the Category pill from the Dimensions panel into Rows to show the class how a small table containing the three categories within the dataset is created.
By dragging Segment into Rows and placing it after the Category pill, the table is made slightly more complex. Now each category within the visualization has been split into three distinct parts.

Dragging "Quantity" from the Measurements panel and placing it within Columns finally creates a true visualization: a bar chart showing the quantity of orders per segment per category.

The chart can then be made more detailed by adding more elements. By adding Market into Columns, for example, multiple charts are created to show the quantity of orders per segment per category within each geographic market.

if users would like to change what kind of visualization to employ, all they need to do is click the Show Me button at the top-right of the application and select the charting style desired.

Create a new worksheet within Tableau. Drag Sales into the Rows section.

The type of calculation performed on a Measures pill can be changed by clicking on the pill, selecting "Measure" from the drop-down menu, and then picking one of the calculation types present.

Now drag Order Date into the Columns field to create a very basic line chart.

Tableau has aggregated the dates at the year level. In order to expand this to include quarters, simply click on the plus symbol within the YEAR pill.

In order to compare how Q1 has performed over the years, simply move the QUARTER pill before YEAR.

# Students Turn Activity 2 Do Explore Data
# Data Exploration with Tableau
## Instructions
* Using the `GlobalSuperstoreOrders2016.xlsx` workbook, visualize the following:
1. The customers with the highest sales amount

2. The most profitable customers

3. The states with the highest average profit

4. A monthly timeline of sales

5. Profit by region and product category (in the United States).

# Review
The first visualization, of the customers with the highest sales, requires dragging the Customer Name pill to Rows, and the Sales pill to Columns.

To sort the data, click on the sort button:

In the next tab, in order to chart the most profitable customers, simply do the same as above, this time with the Profit pill:

To adjust the axis at the bottom, right click at the bottom along the axis, and select Edit Axis:

After filtering out negative profit figures, the chart should now look like this:

And to chart the states with the highest average profit, choose the Profit pill again, then Average under Measure:


The filter should be set to the United States:
Next, to display a monthly timeline, drag the Sales pill to Rows, then obtain its sum. Then drag Order Date to columns.



Finally, to visualize profit by region and product category, drag Category and Region pills to Columns, and create a sum of the Profit pill in Rows:


## Student Turn Activity 4
### No Shows

### Instructions
* Create a line chart that compares the ages of patients against the total number of appointments. Then split this graph based upon gender and whether the patient showed up to their appointment. For this first step, you'll need to convert `Age` from a measure to a dimension.
* Create a pair of bar charts that compare how many patients showed up to appointments versus how many were no-shows in different neighborhoods.
* Create a stacked bar chart that compares no-shows to those who made it to appointment based upon the day of the week.
* Create a pair of line graphs that compare age versus diabetes in both men and women.
* Create a pair of line graphs that compare age versus alcoholism in both men and women.
### Solutions
- Step 1 The first step for this activity is to drag Age to Columns, and Number of Records to Rows. Age must also be converted from measure into dimension by clicking on the arrow on the pill.

- Step 2 To split up the results by gender, drag Gender into Rows:

- Step 3 Finally, to stratify the results by no-show appointments, drag No-show to columns:

- Step 4 In the next visualization, students were asked to compare no-shows by neighborhood. This can be done in the following way:

No-show and Number of Records are dragged to Columns, and Neighbourhood to Rows.
- Step 4 It can also be visualized thus:

No-show is moved to Rows instead of Columns.
- Step 5 Visualize the number of no-show patients by the day of the week:

- Step 6 Since we're counting the number of no-show appointments, it makes sense to to drag No-show to Rows, and visualize this measure vertically. And since we're tallying the number of no-shows by the day of the week, to drag Scheduled Day into Columns:

- Step 6 We're shown results by year, instead of the day of the week. This can be selected by clicking on the arrow on the Scheduled Day pill, More, then Weekday.

- Step 8 To display a bar chart instead of a line chart, select Show Me, then the stacked bar chart option:

- Step 9 display the number of diabetics by gender and across age groups. One way to visualize this is by stacking Gender in Rows.

- Step 10 final visualization is very similar to the previous one, visualizing alcoholism instead of diabetes.

# Instructor Turn Activity 5
- In order to merge these two datasets together, click and drag the "People" sheet into to main area of Tableau alongside the "Orders" sheet.
- Tableau will automatically create an inner join on the columns that contain matching values. In this case, the join is on the "Region" columns.
- To change what type of join is used, simply click on the interlacing circles at the top of the application and select what form of join to use from the menu that appears. This same menu can be used to modify what columns to merge on.

It is also possible to create joins across data sources.

To do this, click on the "Add" button in the Connections panel and add the secondary data source desired. For the purposes of this demonstration, that is GlobalSuperstoreReturns2016.csv.
After the data source has been added, it can then be joined with the other data files desired using the method mentioned before.
Another interesting feature of Tableau is that columns containing text can be split so as to extract data.

Select what character to split the text on, whether to split from the beginning or end of the string, and then how many times the text should be split.
## Student Turn Activity 6 FIFA Analysis
### Instructions
* Create a join between each of the charts so that each player's data is matched up correctly.
* Create a pair of charts that compare the potential of a club's players to their overall ability (`Overall` column). Then sort them from best to worst.
* Create a chart that determines which soccer club is the most aggressive overall.
* Create a chart that determines which nationality has the greatest acceleration on average, making sure to note how many players are from each nation in a second chart.
* Create a chart that determines which nationality has the greatest long passing on average.
* Create a chart that marks the potential of a player over time as they age.
- Step 1 In order to join the two CSV files, drag them to the main pane in the Data Source tab, then, select an inner join:

- Step 2 The first visualization is of each player's potential, as well as overall ability, sorted in descending order:

- Step 3 The second visualization tallies the Aggression of each club.

- The next visualization is of average acceleration by country, as well as the number of records from each country.

- Step 4 The next visualization is of average long passing by country, as well as the number of players from that country.

- Step 5 The next visualization plots age against potential:

- Step 6 To be able to chart each age year as a discrete quantity, click on the Age pill and select Dimension.

| github_jupyter |
<a href="https://colab.research.google.com/github/AI4Finance-Foundation/FinRL/blob/master/FinRL_StockTrading_NeurIPS_2018.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Deep Reinforcement Learning for Stock Trading from Scratch: Multiple Stock Trading
* **Pytorch Version**
# Content
* [1. Problem Definition](#0)
* [2. Getting Started - Load Python packages](#1)
* [2.1. Install Packages](#1.1)
* [2.2. Check Additional Packages](#1.2)
* [2.3. Import Packages](#1.3)
* [2.4. Create Folders](#1.4)
* [3. Download Data](#2)
* [4. Preprocess Data](#3)
* [4.1. Technical Indicators](#3.1)
* [4.2. Perform Feature Engineering](#3.2)
* [5.Build Environment](#4)
* [5.1. Training & Trade Data Split](#4.1)
* [5.2. User-defined Environment](#4.2)
* [5.3. Initialize Environment](#4.3)
* [6.Implement DRL Algorithms](#5)
* [7.Backtesting Performance](#6)
* [7.1. BackTestStats](#6.1)
* [7.2. BackTestPlot](#6.2)
* [7.3. Baseline Stats](#6.3)
* [7.3. Compare to Stock Market Index](#6.4)
* [RLlib Section](#7)
<a id='0'></a>
# Part 1. Problem Definition
This problem is to design an automated trading solution for single stock trading. We model the stock trading process as a Markov Decision Process (MDP). We then formulate our trading goal as a maximization problem.
The algorithm is trained using Deep Reinforcement Learning (DRL) algorithms and the components of the reinforcement learning environment are:
* Action: The action space describes the allowed actions that the agent interacts with the
environment. Normally, a ∈ A includes three actions: a ∈ {−1, 0, 1}, where −1, 0, 1 represent
selling, holding, and buying one stock. Also, an action can be carried upon multiple shares. We use
an action space {−k, ..., −1, 0, 1, ..., k}, where k denotes the number of shares. For example, "Buy
10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or −10, respectively
* Reward function: r(s, a, s′) is the incentive mechanism for an agent to learn a better action. The change of the portfolio value when action a is taken at state s and arriving at new state s', i.e., r(s, a, s′) = v′ − v, where v′ and v represent the portfolio
values at state s′ and s, respectively
* State: The state space describes the observations that the agent receives from the environment. Just as a human trader needs to analyze various information before executing a trade, so
our trading agent observes many different features to better learn in an interactive environment.
* Environment: Dow 30 consituents
The data of the single stock that we will be using for this case study is obtained from Yahoo Finance API. The data contains Open-High-Low-Close price and volume.
<a id='1'></a>
# Part 2. Getting Started- Load Python Packages
<a id='1.1'></a>
## 2.1. Install all the packages through FinRL library
```
## install finrl library
!pip install git+https://github.com/AI4Finance-LLC/FinRL-Library.git
import os
if not os.path.exists("./" + config.DATA_SAVE_DIR):
os.makedirs("./" + config.DATA_SAVE_DIR)
if not os.path.exists("./" + config.TRAINED_MODEL_DIR):
os.makedirs("./" + config.TRAINED_MODEL_DIR)
if not os.path.exists("./" + config.TENSORBOARD_LOG_DIR):
os.makedirs("./" + config.TENSORBOARD_LOG_DIR)
if not os.path.exists("./" + config.RESULTS_DIR):
os.makedirs("./" + config.RESULTS_DIR)
```
<a id='1.2'></a>
## 2.2. Check if the additional packages needed are present, if not install them.
* Yahoo Finance API
* pandas
* numpy
* matplotlib
* stockstats
* OpenAI gym
* stable-baselines
* tensorflow
* pyfolio
<a id='1.3'></a>
## 2.3. Import Packages
```
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# matplotlib.use('Agg')
import datetime
%matplotlib inline
from finrl.apps import config
from finrl.finrl_meta.preprocessor.yahoodownloader import YahooDownloader
from finrl.finrl_meta.preprocessor.preprocessors import FeatureEngineer, data_split
from finrl.finrl_meta.env_stock_trading.env_stocktrading import StockTradingEnv
from finrl.drl_agents.stablebaselines3.models import DRLAgent
from finrl.finrl_meta.data_processor import DataProcessor
from finrl.plot import backtest_stats, backtest_plot, get_daily_return, get_baseline
from pprint import pprint
import sys
sys.path.append("../FinRL-Library")
import itertools
```
<a id='1.4'></a>
## 2.4. Create Folders
<a id='2'></a>
# Part 3. Download Data
Yahoo Finance is a website that provides stock data, financial news, financial reports, etc. All the data provided by Yahoo Finance is free.
* FinRL uses a class **YahooDownloader** to fetch data from Yahoo Finance API
* Call Limit: Using the Public API (without authentication), you are limited to 2,000 requests per hour per IP (or up to a total of 48,000 requests a day).
-----
class YahooDownloader:
Provides methods for retrieving daily stock data from
Yahoo Finance API
Attributes
----------
start_date : str
start date of the data (modified from config.py)
end_date : str
end date of the data (modified from config.py)
ticker_list : list
a list of stock tickers (modified from config.py)
Methods
-------
fetch_data()
Fetches data from yahoo API
```
# from config.py start_date is a string
config.START_DATE
# from config.py end_date is a string
config.END_DATE
df = YahooDownloader(start_date = '2009-01-01',
end_date = '2021-10-31',
ticker_list = config.DOW_30_TICKER).fetch_data()
print(config.DOW_30_TICKER)
df.shape
df.sort_values(['date','tic'],ignore_index=True).head()
```
# Part 4: Preprocess Data
Data preprocessing is a crucial step for training a high quality machine learning model. We need to check for missing data and do feature engineering in order to convert the data into a model-ready state.
* Add technical indicators. In practical trading, various information needs to be taken into account, for example the historical stock prices, current holding shares, technical indicators, etc. In this article, we demonstrate two trend-following technical indicators: MACD and RSI.
* Add turbulence index. Risk-aversion reflects whether an investor will choose to preserve the capital. It also influences one's trading strategy when facing different market volatility level. To control the risk in a worst-case scenario, such as financial crisis of 2007–2008, FinRL employs the financial turbulence index that measures extreme asset price fluctuation.
```
fe = FeatureEngineer(
use_technical_indicator=True,
tech_indicator_list = config.TECHNICAL_INDICATORS_LIST,
use_vix=True,
use_turbulence=True,
user_defined_feature = False)
processed = fe.preprocess_data(df)
list_ticker = processed["tic"].unique().tolist()
list_date = list(pd.date_range(processed['date'].min(),processed['date'].max()).astype(str))
combination = list(itertools.product(list_date,list_ticker))
processed_full = pd.DataFrame(combination,columns=["date","tic"]).merge(processed,on=["date","tic"],how="left")
processed_full = processed_full[processed_full['date'].isin(processed['date'])]
processed_full = processed_full.sort_values(['date','tic'])
processed_full = processed_full.fillna(0)
processed_full.sort_values(['date','tic'],ignore_index=True).head(10)
```
<a id='4'></a>
# Part 5. Design Environment
Considering the stochastic and interactive nature of the automated stock trading tasks, a financial task is modeled as a **Markov Decision Process (MDP)** problem. The training process involves observing stock price change, taking an action and reward's calculation to have the agent adjusting its strategy accordingly. By interacting with the environment, the trading agent will derive a trading strategy with the maximized rewards as time proceeds.
Our trading environments, based on OpenAI Gym framework, simulate live stock markets with real market data according to the principle of time-driven simulation.
The action space describes the allowed actions that the agent interacts with the environment. Normally, action a includes three actions: {-1, 0, 1}, where -1, 0, 1 represent selling, holding, and buying one share. Also, an action can be carried upon multiple shares. We use an action space {-k,…,-1, 0, 1, …, k}, where k denotes the number of shares to buy and -k denotes the number of shares to sell. For example, "Buy 10 shares of AAPL" or "Sell 10 shares of AAPL" are 10 or -10, respectively. The continuous action space needs to be normalized to [-1, 1], since the policy is defined on a Gaussian distribution, which needs to be normalized and symmetric.
## Training data split: 2009-01-01 to 2020-07-01
## Trade data split: 2020-07-01 to 2021-10-31
```
train = data_split(processed_full, '2009-01-01','2020-07-01')
trade = data_split(processed_full, '2020-07-01','2021-10-31')
print(len(train))
print(len(trade))
train.tail()
trade.head()
config.TECHNICAL_INDICATORS_LIST
stock_dimension = len(train.tic.unique())
state_space = 1 + 2*stock_dimension + len(config.TECHNICAL_INDICATORS_LIST)*stock_dimension
print(f"Stock Dimension: {stock_dimension}, State Space: {state_space}")
env_kwargs = {
"hmax": 100,
"initial_amount": 1000000,
"buy_cost_pct": 0.001,
"sell_cost_pct": 0.001,
"state_space": state_space,
"stock_dim": stock_dimension,
"tech_indicator_list": config.TECHNICAL_INDICATORS_LIST,
"action_space": stock_dimension,
"reward_scaling": 1e-4
}
e_train_gym = StockTradingEnv(df = train, **env_kwargs)
```
## Environment for Training
```
env_train, _ = e_train_gym.get_sb_env()
print(type(env_train))
```
<a id='5'></a>
# Part 6: Implement DRL Algorithms
* The implementation of the DRL algorithms are based on **OpenAI Baselines** and **Stable Baselines**. Stable Baselines is a fork of OpenAI Baselines, with a major structural refactoring, and code cleanups.
* FinRL library includes fine-tuned standard DRL algorithms, such as DQN, DDPG,
Multi-Agent DDPG, PPO, SAC, A2C and TD3. We also allow users to
design their own DRL algorithms by adapting these DRL algorithms.
```
agent = DRLAgent(env = env_train)
```
### Model Training: 5 models, A2C DDPG, PPO, TD3, SAC
### Model 1: A2C
```
agent = DRLAgent(env = env_train)
model_a2c = agent.get_model("a2c")
trained_a2c = agent.train_model(model=model_a2c,
tb_log_name='a2c',
total_timesteps=50000)
```
### Model 2: DDPG
```
agent = DRLAgent(env = env_train)
model_ddpg = agent.get_model("ddpg")
trained_ddpg = agent.train_model(model=model_ddpg,
tb_log_name='ddpg',
total_timesteps=50000)
```
### Model 3: PPO
```
agent = DRLAgent(env = env_train)
PPO_PARAMS = {
"n_steps": 2048,
"ent_coef": 0.01,
"learning_rate": 0.00025,
"batch_size": 128,
}
model_ppo = agent.get_model("ppo",model_kwargs = PPO_PARAMS)
trained_ppo = agent.train_model(model=model_ppo,
tb_log_name='ppo',
total_timesteps=50000)
```
### Model 4: TD3
```
agent = DRLAgent(env = env_train)
TD3_PARAMS = {"batch_size": 100,
"buffer_size": 1000000,
"learning_rate": 0.001}
model_td3 = agent.get_model("td3",model_kwargs = TD3_PARAMS)
trained_td3 = agent.train_model(model=model_td3,
tb_log_name='td3',
total_timesteps=30000)
```
### Model 5: SAC
```
agent = DRLAgent(env = env_train)
SAC_PARAMS = {
"batch_size": 128,
"buffer_size": 1000000,
"learning_rate": 0.0001,
"learning_starts": 100,
"ent_coef": "auto_0.1",
}
model_sac = agent.get_model("sac",model_kwargs = SAC_PARAMS)
trained_sac = agent.train_model(model=model_sac,
tb_log_name='sac',
total_timesteps=60000)
```
## Trading
Assume that we have $1,000,000 initial capital at 2020-07-01. We use the DDPG model to trade Dow jones 30 stocks.
### Set turbulence threshold
Set the turbulence threshold to be greater than the maximum of insample turbulence data, if current turbulence index is greater than the threshold, then we assume that the current market is volatile
```
data_risk_indicator = processed_full[(processed_full.date<'2020-07-01') & (processed_full.date>='2009-01-01')]
insample_risk_indicator = data_risk_indicator.drop_duplicates(subset=['date'])
insample_risk_indicator.vix.describe()
insample_risk_indicator.vix.quantile(0.996)
insample_risk_indicator.turbulence.describe()
insample_risk_indicator.turbulence.quantile(0.996)
```
### Trade
DRL model needs to update periodically in order to take full advantage of the data, ideally we need to retrain our model yearly, quarterly, or monthly. We also need to tune the parameters along the way, in this notebook I only use the in-sample data from 2009-01 to 2020-07 to tune the parameters once, so there is some alpha decay here as the length of trade date extends.
Numerous hyperparameters – e.g. the learning rate, the total number of samples to train on – influence the learning process and are usually determined by testing some variations.
```
#trade = data_split(processed_full, '2020-07-01','2021-10-31')
e_trade_gym = StockTradingEnv(df = trade, turbulence_threshold = 70,risk_indicator_col='vix', **env_kwargs)
# env_trade, obs_trade = e_trade_gym.get_sb_env()
trade.head()
df_account_value, df_actions = DRLAgent.DRL_prediction(
model=trained_sac,
environment = e_trade_gym)
df_account_value.shape
df_account_value.tail()
df_actions.head()
```
<a id='6'></a>
# Part 7: Backtest Our Strategy
Backtesting plays a key role in evaluating the performance of a trading strategy. Automated backtesting tool is preferred because it reduces the human error. We usually use the Quantopian pyfolio package to backtest our trading strategies. It is easy to use and consists of various individual plots that provide a comprehensive image of the performance of a trading strategy.
<a id='6.1'></a>
## 7.1 BackTestStats
pass in df_account_value, this information is stored in env class
```
print("==============Get Backtest Results===========")
now = datetime.datetime.now().strftime('%Y%m%d-%Hh%M')
perf_stats_all = backtest_stats(account_value=df_account_value)
perf_stats_all = pd.DataFrame(perf_stats_all)
perf_stats_all.to_csv("./"+config.RESULTS_DIR+"/perf_stats_all_"+now+'.csv')
#baseline stats
print("==============Get Baseline Stats===========")
baseline_df = get_baseline(
ticker="^DJI",
start = df_account_value.loc[0,'date'],
end = df_account_value.loc[len(df_account_value)-1,'date'])
stats = backtest_stats(baseline_df, value_col_name = 'close')
df_account_value.loc[0,'date']
df_account_value.loc[len(df_account_value)-1,'date']
```
<a id='6.2'></a>
## 7.2 BackTestPlot
```
print("==============Compare to DJIA===========")
%matplotlib inline
# S&P 500: ^GSPC
# Dow Jones Index: ^DJI
# NASDAQ 100: ^NDX
backtest_plot(df_account_value,
baseline_ticker = '^DJI',
baseline_start = df_account_value.loc[0,'date'],
baseline_end = df_account_value.loc[len(df_account_value)-1,'date'])
```
| github_jupyter |
# eQTL Analysis
```
import copy
import glob
import os
import subprocess
import cdpybio as cpb
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
pd.options.mode.chained_assignment = None # default='warn'
import pybedtools as pbt
import seaborn as sns
import socket
import statsmodels.stats.multitest as smm
import vcf as pyvcf
import cardipspy as cpy
import ciepy
%matplotlib inline
%load_ext rpy2.ipython
dy_name = 'eqtl_analysis'
import socket
if socket.gethostname() == 'fl-hn1' or socket.gethostname() == 'fl-hn2':
dy = os.path.join(ciepy.root, 'sandbox', dy_name)
cpy.makedir(dy)
pbt.set_tempdir(dy)
outdir = os.path.join(ciepy.root, 'output', dy_name)
cpy.makedir(outdir)
private_outdir = os.path.join(ciepy.root, 'private_output', dy_name)
cpy.makedir(private_outdir)
transcript_to_gene = pd.read_table(cpy.gencode_transcript_gene, header=None,
squeeze=True, index_col=0)
gene_info = pd.read_table(cpy.gencode_gene_info, index_col=0)
fn = os.path.join(ciepy.root, 'output', 'eqtl_input',
'tpm_log_filtered_phe_std_norm_peer_resid.tsv')
exp = pd.read_table(fn, index_col=0)
dy = os.path.join(ciepy.root, 'output/eqtl_processing/eqtls01')
fn = os.path.join(dy, 'qvalues.tsv')
qvalues = pd.read_table(fn, index_col=0)
fn = os.path.join(dy, 'lead_variants.tsv')
lead_vars = pd.read_table(fn, index_col=0)
fn = os.path.join(dy, 'lead_variants_single.tsv')
lead_vars_single = pd.read_table(fn, index_col=0)
fn = os.path.join(dy, 'gene_variant_pairs.tsv')
gene_variant = pd.read_table(fn, index_col=0)
dy = os.path.join(ciepy.root, 'output/eqtl_processing/eqtls01')
h2 = pd.read_table(os.path.join(dy, 'h2.tsv'), index_col=0, squeeze=True, header=None)
dy = os.path.join(ciepy.root, 'output/eqtl_processing/no_peer01')
#h2_no_peer = pd.read_table(os.path.join(dy, 'h2.tsv'), index_col=0, squeeze=True, header=None)
gold_eqtls = pd.read_table(
os.path.join(ciepy.root, 'output', 'eqtl_methods_exploration', 'gold_eqtls.tsv'),
index_col=0)
fn = os.path.join(ciepy.root, 'output/eqtl_processing/eqtls02', 'lead_variants.tsv')
lead_vars_second = pd.read_table(fn, index_col=0)
fn = os.path.join(ciepy.root, 'output/eqtl_processing/eqtls03', 'lead_variants.tsv')
lead_vars_third = pd.read_table(fn, index_col=0)
```
## Summary
```
print('We detected eQTLs for {:,.0f} of {:,} genes tested.'.format(qvalues.perm_sig.sum(),
qvalues.shape[0]))
for i in range(1, 4):
fn = os.path.join(ciepy.root, 'output/eqtl_processing/eqtls0{}/qvalues.tsv'.format(i))
tdf = pd.read_table(fn, index_col=0)
print('{:,.0f} of {:,} significant genes have eQTLs for analysis {}.'.format(
tdf.perm_sig.sum(), tdf.shape[0], i))
print('Number of genes with variant type as lead variant (ties allowed):')
s = set(lead_vars[lead_vars.perm_sig].gene_id + ':' + lead_vars[lead_vars.perm_sig].variant_type)
pd.Series([x.split(':')[1] for x in s]).value_counts()
761 + 615
print('Number of genes with lead variant from each caller (ties allowed):')
s = set(lead_vars[lead_vars.perm_sig].gene_id + ':' + lead_vars[lead_vars.perm_sig].variant_caller)
pd.Series([x.split(':')[1] for x in s]).value_counts()
n = sum(gene_variant.marker_id.apply(lambda x: 'CNV' in x))
print('Total number of CNV eQTLs: {:,}'.format(n))
sig = lead_vars[lead_vars.perm_sig]
sig_single = lead_vars[lead_vars.perm_sig]
a = gene_variant.shape[0]
b = len(set(gene_variant.location))
print('{:,} total variant-expression associations comprising {:,}'
' unique variants.'.format(a, b))
```
## Comparison to GTEx Multi-Tissue eQTLs
I want to visualize some of the permutation $p$-values versus the "real" $p$-value
for some of the GTEx multi-tissue eQTLs.
```
fig, axs = plt.subplots(2, 2, figsize=(10, 10))
axs = axs.flatten()
for i, g in enumerate(list(set(sig.gene_id) & set(gold_eqtls.index))[0:4]):
ax = axs[i]
fn = os.path.join(ciepy.root, 'private_output', 'run_eqtl_analysis', 'eqtls01',
'gene_results', g, '{}.tsv'.format(g))
min_fn = os.path.join(ciepy.root, 'private_output', 'run_eqtl_analysis', 'eqtls01',
'gene_results', g, 'minimum_pvalues.tsv')
if os.path.exists(min_fn):
res = ciepy.read_emmax_output(fn)
#t = res[res.MARKER_ID.apply(lambda x: gold_eqtls.ix[g, 'rsid'] in x)]
t = res[res.BEG == gold_eqtls.ix[g, 'start']]
min_pvals = pd.read_table(min_fn, header=None, squeeze=True)
ax = (-np.log10(min_pvals)).hist(ax=ax)
ax.set_ylabel('Number of $p$-values')
ax.set_xlabel('$-\log_{10}$ $p$-value')
ya, yb = ax.get_ylim()
ax.vlines(-np.log10(t.PVALUE), ya, yb, label='GTEx lead variant', alpha=0.5, linestyle='--')
ax.vlines(-np.log10(res.PVALUE.min()), ya, yb, colors=['red'], label='min $p$-value', alpha=0.5)
ax.set_title('{}'.format(gene_info.ix[g, 'gene_name']));
lgd = ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
fig.tight_layout()
fig.savefig(os.path.join(outdir, 'gtex_comparison.pdf'),
bbox_extra_artists=(lgd,), bbox_inches='tight')
```
The black line shows the $p$-value for the GTEx SNV. The red line shows the smallest $p$-value that
I observe for the gene. We can see that most of these genes are highly significant compared to the
permutation $p$-values.
## Distance to Nearest TSS
```
n,b,p = plt.hist(sig.tss_dist,
bins=np.arange(-500000, 510000, 10000),
normed=True, histtype='stepfilled',
linewidth=0)
plt.title('Distance from nearest TSS')
plt.ylabel('Fraction of eQTLs')
plt.xlabel('Distance in base pairs');
n,b,p = plt.hist(sig[sig.variant_type == 'ins'].tss_dist,
bins=np.arange(-1000000, 1010000, 10000),
normed=True, histtype='stepfilled', linewidth=0)
plt.title('Distance from nearest TSS')
plt.ylabel('Fraction of eQTLs')
plt.xlabel('Distance in base pairs');
n,b,p = plt.hist(sig[sig.variant_type == 'del'].tss_dist,
bins=np.arange(-1000000, 1010000, 10000),
normed=True, histtype='stepfilled', linewidth=0)
plt.title('Distance from nearest TSS')
plt.ylabel('Fraction of eQTLs')
plt.xlabel('Distance in base pairs');
n,b,p = plt.hist(sig[sig.variant_type == 'cnv'].tss_dist,
bins=np.arange(-1000000, 1010000, 10000),
normed=True, histtype='stepfilled', linewidth=0)
plt.title('Distance from nearest TSS')
plt.ylabel('Fraction of eQTLs')
plt.xlabel('Distance in base pairs');
```
The above two plots are the same data at different $x$-axis limits. The first plot
in particular is meant to be comparable to Fig. S9 from the GTEx paper.
```
ax = sns.jointplot(sig_single.tss_dist / 1000,
-np.log10(sig_single.pvalue),
stat_func=None, alpha=0.25)
ax.set_axis_labels(xlabel='Distance in kb',
ylabel='$-\log_{10}$ $p$-value')
plt.tight_layout()
plt.savefig(os.path.join(outdir, 'sig_pvalue_tss_dist.pdf'))
ax = sns.jointplot(sig_single.tss_dist / 1000,
sig_single.beta.abs(),
stat_func=None, alpha=0.25)
ax.set_axis_labels(xlabel='Distance in kb', ylabel='abs$(\\beta)$')
plt.tight_layout()
plt.savefig(os.path.join(outdir, 'sig_beta_tss_dist.pdf'))
dists = sig_single.tss_dist
a = dists[dists >= 0]
b = -dists[dists < 0]
r = plt.hist(list(np.log10(a + 1)) + list(-np.log10(b + 1)),
bins=100)
plt.title('Distance from nearest TSS')
plt.ylabel('Number of eQTLs')
plt.xlabel('Distance in $\log_{10}$ base pairs')
ya, yb = plt.ylim()
plt.vlines(np.log10(50000), ya, yb, linestyles='--')
plt.vlines(-np.log10(50000), ya, yb, linestyles='--')
plt.vlines(np.log10(3500), ya, yb, linestyles='--', color='red')
plt.vlines(-np.log10(3500), ya, yb, linestyles='--', color='red')
plt.tight_layout()
plt.savefig(os.path.join(outdir, 'sig_snv_tss_log_dist.pdf'))
```
Grubert et al. defined distal as greater than 50 kb away (outside of the black lines above).
They searched for *cis* QTLs within 2kb of peak boundaries or $\pm$3.5kb around the TSS (inside
of the red lines above). I'd like to go back and see what they thought about the intermediate region.
I'm somewhat surprised that there are more potential QTNs downstream of TSSs. In the plot, a positive
distance indicates that the SNV is downstream of the TSS (relative to the strand of the TSS) although
I believe this has been reported before (I think it was a PLOS Genetics paper that estimated where
causal variants were located and found that they were estimated to be slightly downstream of the TSS).
## Gencode Gene Type
```
t = pd.DataFrame(0, index=set(gene_info.gene_type), columns=['sig', 'not_sig'])
vc = gene_info.ix[qvalues[qvalues.perm_sig].index, 'gene_type'].value_counts()
t.ix[vc.index, 'sig'] = vc
vc = gene_info.ix[qvalues[qvalues.perm_sig == False].index, 'gene_type'].value_counts()
t.ix[vc.index, 'not_sig'] = vc
t = t[t.sum(axis=1) > 1]
t['total'] = t.sum(axis=1)
#t.sort_values(by='total', inplace=True, ascending=False)
t.sort_values(by='total', inplace=True)
t.drop('protein_coding')[['sig', 'not_sig']].plot(kind='barh')
plt.xlabel('Number of genes');
```
## Manhattan and Expression Plots
```
fn = os.path.join(ciepy.root, 'private_output', 'eqtl_input',
'filtered_all', '0000.vcf.gz')
vcf_reader = pyvcf.Reader(open(fn), compressed=True)
res_fns = glob.glob(os.path.join(ciepy.root, 'private_output', 'run_eqtl_analysis', 'eqtls01',
'gene_results', '*', 'ENS*.tsv'))
res_fns = pd.Series(res_fns,
index=[os.path.splitext(os.path.split(x)[1])[0] for x in res_fns])
qvalue_sig = qvalues[qvalues.perm_sig == 1]
qvalue_sig = qvalue_sig.sort_values('perm_qvalue')
def eqtl_summary(gene_id, left=None, right=None, fn_root=None):
res = ciepy.read_emmax_output(res_fns[gene_id])
res = res.sort_values('PVALUE')
ind = sig_single[sig_single.gene_id == gene_id].index[0]
if gene_info.ix[gene_id, 'strand'] == '+':
gene_left = gene_info.ix[gene_id, 'start']
gene_right = gene_info.ix[gene_id, 'end']
else:
gene_right = gene_info.ix[gene_id, 'start']
gene_left = gene_info.ix[gene_id, 'end']
if left:
res = res[res.BEG >= gene_left - left]
if right:
res = res[res.BEG <= gene_right + right]
plt.figure()
plt.scatter(res.BEG, -np.log10(res.PVALUE), color='grey', alpha=0.5)
plt.xlim(res.BEG.min(), res.BEG.max())
ya = 0
yb = (-np.log10(res.PVALUE)).max() + 1
plt.ylim(ya, yb)
# plt.vlines(most_sig_single.ix[g, 'tss_start'], ya, yb, color='blue',
# linestyles='--', alpha=0.5)
poly = plt.Polygon([[gene_left, 0.9 * yb], [gene_left, yb], [gene_right, 0.95 * yb]],
closed=True, facecolor='blue', edgecolor='none')
plt.gca().add_patch(poly)
#most_sig_single.ix[g, 'tss_start']
plt.xlabel('Genomic position')
plt.ylabel('$-\log_{10}$ $p$-value')
plt.title('Manhattan plot for {}'.format(gene_info.ix[gene_id, 'gene_name']));
if fn_root:
plt.tight_layout()
plt.savefig('{}_manhattan.pdf'.format(fn_root))
t = vcf_reader.fetch(res.CHROM.values[0],
res.BEG.values[0],
res.BEG.values[0] + 1)
r = t.next()
tdf = pd.DataFrame(exp.ix[gene_id])
tdf.columns = ['expression']
tdf['genotype'] = 0
hets = set(exp.columns) & set([s.sample for s in r.get_hets()])
tdf.ix[hets, 'genotype'] = 1
alts = set(exp.columns) & set([s.sample for s in r.get_hom_alts()])
tdf.ix[alts, 'genotype'] = 2
plt.figure()
#sns.lmplot(x='genotype', y='expression', data=tdf)
ax = sns.violinplot(x='genotype', y='expression', data=tdf, color='grey',
order=[0, 1, 2], scale='count')
sns.regplot(x='genotype', y='expression', data=tdf, scatter=False, color='red')
ya, yb = plt.ylim()
plt.text(0, yb, 'n={}'.format(sum(tdf.genotype == 0)),
verticalalignment='top', horizontalalignment='center')
plt.text(1, yb, 'n={}'.format(sum(tdf.genotype == 1)),
verticalalignment='top', horizontalalignment='center')
plt.text(2, yb, 'n={}'.format(sum(tdf.genotype == 2)),
verticalalignment='top', horizontalalignment='center')
plt.title('Expression vs. genotype for {}'.format(gene_info.ix[gene_id, 'gene_name']));
if fn_root:
plt.tight_layout()
plt.savefig('{}_violin.pdf'.format(fn_root))
eqtl_summary(sig.gene_id[0])
eqtl_summary(sig.gene_id[5])
eqtl_summary('ENSG00000100897.13')
eqtl_summary('ENSG00000173992.4')
```
## Pluripotency Genes
```
fn = os.path.join(ciepy.root, 'misc', 'stem_cell_population_maintenance.tsv')
a = pd.read_table(fn, header=None)
fn = os.path.join(ciepy.root, 'misc', 'regulation_of_stem_cell_population_maintenance.tsv')
b = pd.read_table(fn, header=None)
go_genes = set(a[2]) | set(b[2])
go_genes = go_genes & set(gene_info.gene_name)
n = len(set(a[2]) & set(gene_info.ix[qvalues[qvalues.perm_sig].index, 'gene_name']))
print('{} eGenes of {} from stem cell population maintenance.'.format(n, a.shape[0]))
n = len(set(b[2]) & set(gene_info.ix[qvalues[qvalues.perm_sig].index, 'gene_name']))
print('{} eGenes of {} from regulation of stem cell population maintenance.'.format(n, b.shape[0]))
n = len(set(a[2]) & set(b[2]) & set(gene_info.ix[qvalues[qvalues.perm_sig].index, 'gene_name']))
print('{} eGenes shared.'.format(n))
pgenes = ['LIN28A', 'POU5F1', 'SOX2', 'NANOG', 'MYC', 'KLF4', 'ZFP42']
for g in pgenes:
i = gene_info[gene_info.gene_name == g].index[0]
if i in sig.gene_id.values:
eqtl_summary(i, fn_root=os.path.join(outdir, '{}_summary'.format(g)))
# Markers of pluripotency from http://www.nature.com/nbt/journal/v33/n11/full/nbt.3387.html.
pgenes = ['CXCL5', 'IDO1', 'LCK', 'TRIM22', 'DNMT3B', 'HESX1', 'SOX2', 'POU5F1', 'NANOG']
for g in pgenes:
i = gene_info[gene_info.gene_name == g].index[0]
if i in sig.gene_id.values:
eqtl_summary(i, fn_root=os.path.join(outdir, '{}_summary'.format(g)))
genes = [u'BRIX1 ', u'CD9', u'COMMD3 ', u'CRABP2 ', u'CXCL5', u'DIAPH2',
u'DNMT3B', u'EDNRB ', u'FGF4 ', u'FGF5 ', u'FOXD3 ', u'GABRB3 ',
u'GAL ', u'GBX2', u'GDF3 ', u'GRB7 ', u'HCK ', u'HESX1', u'IDO1',
u'IFITM1 ', u'IFITM2', u'IGF2BP2', u'IL6ST ', u'KIT ', u'LCK', u'LIFR ',
u'LIN28A', u'NANOG', u'NODAL ', u'NOG', u'NR5A2', u'NR6A1', u'PODXL',
u'POU5F1', u'PTEN ', u'SEMA3A', u'SFRP2 ', u'SOX2', u'TDGF1', u'TERT',
u'TFCP2L1', u'TRIM22', u'UTF1', u'ZFP42']
for g in genes:
if g in gene_info.gene_name.values:
i = gene_info[gene_info.gene_name == g].index[0]
if i in sig.gene_id:
eqtl_summary(i)#, fn_root=os.path.join(outdir, '{}_summary'.format(g)))
pgenes = ['LIN28A', 'POU5F1', 'SOX2', 'NANOG', 'MYC', 'KLF4', 'ZFP42']
t = set(lead_vars_second.ix[lead_vars_second.perm_sig, 'gene_name'])
for g in pgenes:
if g in t:
print(g)
```
*POU5F1* has a second eQTL as well.
```
def ld_vs_pval(gene_id):
ind = lead_vars_single[lead_vars_single.gene_id == gene_id].index[0]
fn = '/publicdata/1KGP_20151103/LD/tabix/{}_EUR_ld_all.hap.ld.bed.gz'.format(sig_single.ix[ind, 'chrom'])
c = 'tabix {} {}:{}-{}'.format(fn, lead_vars_single.ix[ind].chrom,
sig_single.ix[ind].start - 1, sig_single.ix[ind].end)
res = subprocess.check_output(c, shell=True)
tdf = pd.DataFrame([x.split() for x in res.strip().split('\n')],
columns=['chrom', 'start', 'end', 'ld_info'])
se = pd.Series(tdf['ld_info'].apply(lambda x: x.split(':')).apply(lambda x: x[2]).values,
index=tdf['ld_info'].apply(lambda x: x.split(':')).apply(lambda x: x[1])).astype(float)
res = ciepy.read_emmax_output(res_fns[gene_id])
res.index = res.BEG.astype(str)
plt.scatter(se.values, -np.log10(res.ix[se.index, 'PVALUE'].values),
color='grey', alpha=0.5, s=100)
xa, xb = plt.xlim()
plt.hlines(-np.log10(lead_vars_single.ix[ind, 'pvalue']), xa, xb,
color='red', linestyle='--')
plt.xlim(xa, xb)
plt.ylabel('$-\log_{10}$ $p$-value')
plt.xlabel('LD with most significant SNV');
```
| github_jupyter |
### Nueral Machine Translation (NMT)
We are going to use the `Encoder` - `Decoder` achitecture. Think of it as an achitecture where an enocder maps the source-text to a "thought vector" that summarizes the text's contents, which is then input to the second part of the neural network that decodes the "thought vector" to the destination-text.
### Steps
* Consider the Danish text "der var engang" which translate to "once upon a time".
* We first convert the entire data-set to integer-tokens so the text "der var engang" becomes [12, 54, 1097].
* Each of these integer-tokens is then mapped to an embedding-vector with e.g. 128 elements, so the integer-token 12 could for example become [0.12, -0.56, ..., 1.19]
* These embedding vectoctors will be passed into a GRu with three layers.
The last GRU layer will output a single vector known as the "thought vector" that summarizes the contents of the source-text which is uses as the initial state of the GRU of the decoder.
* The destination-text "once upon a time" is padded with special markers "<sos>" and "<eos>" to indicate its beginning and end, so the sequence of integer-tokens becomes [2, 337, 640, 9, 79, 3].
* During training, the decoder will be given this entire sequence as input and the desired output sequence is [337, 640, 9, 79, 3] which is the same sequence but time-shifted one step.
* We are trying to teach the decoder to map the "thought vector" and the start-token "sos" (integer 2) to the next word "once" (integer 337), and then map the word "once" to the word "upon" (integer 640), and so forth.
### Imports
```
import os, math
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
from tensorflow import keras
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tf.__version__
```
### Data
We will be using the [Europarl dataset](http://www.statmt.org/europarl/) which contains pairs of most European laugauges. We are going to load the data as using the following helper function that was found [here](https://github.com/Hvass-Labs/TensorFlow-Tutorials/blob/master/europarl.py).
```
import urllib.request
import tarfile, zipfile, sys
class Download:
def __init__(self):
pass
def _print_download_progress(self, count, block_size, total_size):
# Percentage completion.
pct_complete = float(count * block_size) / total_size
# Limit it because rounding errors may cause it to exceed 100%.
pct_complete = min(1.0, pct_complete)
# Status-message. Note the \r which means the line should overwrite itself.
msg = "\r- Download progress: {0:.1%}".format(pct_complete)
# Print it.
sys.stdout.write(msg)
sys.stdout.flush()
def download(self, base_url, filename, download_dir):
save_path = os.path.join(download_dir, filename)
# Check if the file already exists, otherwise we need to download it now.
if not os.path.exists(save_path):
# Check if the download directory exists, otherwise create it.
if not os.path.exists(download_dir):
os.makedirs(download_dir)
print("Downloading", filename, "...")
# Download the file from the internet.
url = base_url + filename
file_path, _ = urllib.request.urlretrieve(url=url,
filename=save_path,
reporthook=self._print_download_progress)
print(" Done!")
def maybe_download_and_extract(self, url, download_dir):
filename = url.split('/')[-1]
file_path = os.path.join(download_dir, filename)
if not os.path.exists(file_path):
if not os.path.exists(download_dir):
os.makedirs(download_dir)
# Download the file from the internet.
file_path, _ = urllib.request.urlretrieve(url=url,
filename=file_path,
reporthook=self._print_download_progress)
print()
print("Download finished. Extracting files.")
if file_path.endswith(".zip"):
# Unpack the zip-file.
zipfile.ZipFile(file=file_path, mode="r").extractall(download_dir)
elif file_path.endswith((".tar.gz", ".tgz")):
# Unpack the tar-ball.
tarfile.open(name=file_path, mode="r:gz").extractall(download_dir)
print("Done.")
else:
print("Data has apparently already been downloaded and unpacked.")
download = Download()
class Europal:
data_dir = "data/europarl/"
data_url = "http://www.statmt.org/europarl/v7/"
def __init__(self):
pass
def maybe_download_and_extract(self, language_code="da"):
url = self.data_url + language_code + "-en.tgz"
download.maybe_download_and_extract(url=url, download_dir=self.data_dir)
def load_data(self, english=True, language_code="da", start="", end=""):
if english:
# Load the English data.
filename = "europarl-v7.{0}-en.en".format(language_code)
else:
# Load the other language.
filename = "europarl-v7.{0}-en.{0}".format(language_code)
path = os.path.join(self.data_dir, filename)
with open(path, encoding="utf-8") as file:
texts = [start + line.strip() + end for line in file]
return texts
europal = Europal()
language_code='da'
mark_start = 'sos '
mark_end = ' eos'
europal.maybe_download_and_extract(language_code=language_code)
```
### Now we can load the Sourc and the Target text from the dataset.
```
data_src = europal.load_data(english=False,
language_code=language_code)
data_trg = europal.load_data(english=True,
language_code=language_code,
start=mark_start,
end=mark_end
)
```
> We will be building a model that translate text from `danish` to `english`.
### Checking examples
```
data_src[0]
data_trg[0]
```
### Error in Data
The data-set contains about 2 million sentence-pairs. Some of the data is incorrect.
```
idx = 8002
data_src[idx]
data_trg[idx]
```
### Tokenizer
We need to converts text into numbers. We are going to do the following:
1. convert text-words into so-called integer-tokens
2. we are going to padd the tokens so that they will have the same length.
3. convert integer-tokens into vectors of floating-point numbers using a so-called embedding-layer.
> _We are going to set the maximum number of words in the vocabulary to `10_000`. which means any word that lies outside the top `10_000` will be automatically conveted to unknown_. This is very important because some of the words just apppeared once in the corpus so they may not be that important.
```
num_words = 10_000
class TokenizerWrap(Tokenizer):
def __init__(self, texts, padding, reverse=False, num_words =None):
Tokenizer.__init__(self, num_words= num_words)
self.fit_on_texts(texts)
self.index_to_word = dict(zip(self.word_index.values(),
self.word_index.keys()))
self.tokens = self.texts_to_sequences(texts)
if reverse:
self.tokens = [list(reversed(x)) for x in self.tokens]
truncating = 'pre'
else:
truncating = 'post'
self.num_tokens = [len(x) for x in self.tokens]
self.max_tokens = np.mean(self.num_tokens) + 2 * np.std(self.num_tokens)
self.max_tokens = int(self.max_tokens)
self.tokens_padded = pad_sequences(self.tokens,
maxlen=self.max_tokens,
padding=padding,
truncating=truncating)
def token_to_word(self, token):
return " " if token == 0 else self.index_to_word[token]
def tokens_to_string(self, tokens):
words = [self.index_to_word[token]
for token in tokens
if token != 0]
return " ".join(words)
def text_to_tokens(self, text, reverse=False, padding=False):
tokens = self.texts_to_sequences([text])
tokens = np.array(tokens)
if reverse:
tokens = np.flip(tokens, axis=1)
truncating = 'pre'
else:
truncating = 'post'
if padding:
return pad_sequences(tokens,
maxlen=self.max_tokens,
padding='pre',
truncating=truncating)
```
### SRC language tokenizer
Now we are going to create the tokenizer of the `src` language.
> Note that we pad zeros at the beginning ``('pre')`` of the sequences. We also reverse the sequences of tokens because the research literature suggests that this might improve performance, because the last words seen by the encoder match the first words produced by the decoder, so short-term dependencies are supposedly modelled more accurately.
```
tokenizer_src = TokenizerWrap(texts=data_src,
padding='pre',
reverse=True,
num_words=num_words)
```
### TRG tokenizer for the destination language.
> Note that this tokenizer does not reverse the sequences and it pads zeros at the end ``('post')`` of the arrays.
```
tokenizer_trg = TokenizerWrap(texts=data_trg,
padding='post',
reverse=False,
num_words=num_words)
```
> Note that the sequence-lengths are different for the source and destination languages. This is because texts with the same meaning may have different numbers of words in the two languages.
> Furthermore, we have made a compromise when tokenizing the original texts in order to save a lot of memory. This means we only truncate about 5% of the texts.
```
tokens_src = tokenizer_src.tokens_padded
tokens_trg = tokenizer_trg.tokens_padded
print(tokens_src.shape)
print(tokens_trg.shape)
```
### Start and end tokens
```
token_start = tokenizer_trg.word_index[mark_start.strip()]
token_end = tokenizer_trg.word_index[mark_end.strip()]
token_start, token_end
```
### Example of Token Sequences
```
tokens_src[0]
tokenizer_src.tokens_to_string(tokens_src[0])
tokens_trg[0]
tokenizer_trg.tokens_to_string(tokens_trg[0])
```
the original sentence in the dataset.
```
data_trg[0]
```
### Training data.
Now that the data-set has been converted to sequences of integer-tokens that are padded and truncated and saved in numpy arrays, we can easily prepare the data for use in training the neural network.
### Encoder inputs
The input to the encoder is merely the numpy array for the padded and truncated sequences of integer-tokens produced by the tokenizer:
```
encoder_input_data = tokens_src
```
### Decoder Inputs and Outputs
The input and output data for the decoder is identical, except shifted one time-step. We can use the same numpy array to save memory by slicing it, which merely creates different 'views' of the same data in memory.
```
tokens_trg
tokens_trg.shape
decoder_input_data = tokens_trg[:, :-1]
decoder_input_data.shape
decoder_output_data = tokens_trg[:, 1: ]
decoder_output_data.shape
```
> _These token-sequences are identical except they are shifted one time-step_.
```
decoder_input_data[0]
decoder_output_data[0]
```
If we use the tokenizer to convert these sequences back into text, we see that they are identical except for the first word which is 'sos' that marks the beginning of a text.
```
tokenizer_trg.tokens_to_string(decoder_input_data[idx])
tokenizer_trg.tokens_to_string(decoder_output_data[idx])
```
### Creating a NN
### Encoder
First we will create an encoder, it maps sequence of integers to a thought vector. We are going to use the functional API from keras which is more flexible than the sequential api.
```
encoder_input = keras.layers.Input(shape=(None, ), name='encoder_input')
embedding_size = 128
encoder_embedding = keras.layers.Embedding(input_dim=num_words,
output_dim=embedding_size,
name='encoder_embedding')
```
Next we are going to create 3 GRU's. Note that on the last GRU we are not returning `sequences` and the state size of all these GRU's will be 512.
```
state_size = 512
encoder_gru1 = keras.layers.GRU(state_size, name='encoder_gru1',
return_sequences=True)
encoder_gru2 = keras.layers.GRU(state_size, name='encoder_gru2',
return_sequences=True)
encoder_gru3 = keras.layers.GRU(state_size, name='encoder_gru3',
return_sequences=False)
```
Connecting te encoder layers.
... we are going to create a helper function that will connect the layers of the encoder using the Functional API approach.
```
def connect_encoder():
net = encoder_input
net = encoder_embedding(net)
net = encoder_gru1(net)
net = encoder_gru2(net)
net = encoder_gru3(net)
return net # encoder outputs
```
> _Note how the encoder uses the normal output from its last GRU-layer as the "thought vector". Research papers often use the internal state of the encoder's last recurrent layer as the "thought vector". But this makes the implementation more complicated and is not necessary when using the GRU. But if you were using the LSTM instead then it is necessary to use the LSTM's internal states as the "thought vector" because it actually has two internal vectors, which we would need to initialize the two internal states of the decoder's LSTM units._
```
encoder_output = connect_encoder()
```
### Decoder
Create the decoder-part which maps the "thought vector" to a sequence of integer-tokens.
The decoder takes two inputs. First it needs the "thought vector" produced by the encoder which summarizes the contents of the input-text.
```
decoder_initial_state = keras.layers.Input(shape=(state_size,),
name='decoder_initial_state')
```
The decoder also needs a sequence of integer-tokens as inputs. During training we will supply this with a full sequence of integer-tokens e.g. corresponding to the text "sos once upon a time eos"
During inference when we are translating new input-texts, we will start by feeding a sequence with just one integer-token for "esos" which marks the beginning of a text, and combined with the "thought vector" from the encoder, the decoder will hopefully be able to produce the correct next word e.g. "once".
```
decoder_input = keras.layers.Input(shape=(None, ), name='decoder_input')
```
... the decoder embedding layer convers integer tokens to floating values between -1 and 1.
```
decoder_embedding = keras.layers.Embedding(input_dim=num_words,
output_dim=embedding_size,
name='decoder_embedding')
```
We will then create 3 GRU layers of the decoder that will both return sequences.
> _Note that they all return sequences because we ultimately want to output a sequence of integer-tokens that can be converted into a text-sequence._
```
decoder_gru1 = keras.layers.GRU(state_size, name='decoder_gru1',
return_sequences=True)
decoder_gru2 = keras.layers.GRU(state_size, name='decoder_gru2',
return_sequences=True)
decoder_gru3 = keras.layers.GRU(state_size, name='decoder_gru3',
return_sequences=True)
```
The GRU layers output a tensor with shape ``[batch_size, sequence_length, state_size]``, where each "word" is encoded as a vector of length ``state_size``. We need to convert this into sequences of integer-tokens that can be interpreted as words from our vocabulary.
```
decoder_dense = keras.layers.Dense(num_words,
activation='softmax',
name='decoder_output')
```
The decoder is built using the functional API of Keras, which allows more flexibility in connecting the layers e.g. to route different inputs to the decoder. This is useful because we have to connect the decoder directly to the encoder, but we will also connect the decoder to another input so we can run it separately.
```
def connect_decoder(initial_state):
net = decoder_input
net = decoder_embedding(net)
net = decoder_gru1(net, initial_state=initial_state)
net = decoder_gru2(net, initial_state=initial_state)
net = decoder_gru3(net, initial_state=initial_state)
out = decoder_dense(net)
return out
```
### Connecting and creating a models
_First we connect the encoder directly to the decoder so it is one whole model that can be trained end-to-end. This means the initial-state of the decoder's GRU units are set to the output of the encoder_
```
decoder_output = connect_decoder(initial_state=encoder_output)
model_train = keras.Model(inputs=[encoder_input, decoder_input],
outputs=[decoder_output])
```
... then we create the encoder model serperately.
```
model_encoder = keras.Model(inputs=[encoder_input],
outputs=[encoder_output])
```
... then we create the model decoder alone as well.
```
decoder_output = connect_decoder(initial_state=decoder_initial_state)
model_decoder = keras.Model(inputs=[decoder_input, decoder_initial_state],
outputs=[decoder_output])
```
### Plotting the models
... train model
```
keras.utils.plot_model(model_train, dpi=64, show_shapes=True)
```
...encoder model
```
keras.utils.plot_model(model_encoder, dpi=64, show_shapes=True)
```
...decoder model
```
keras.utils.plot_model(model_decoder, dpi=64, show_shapes=True)
```
### Compile the Model
The output of the decoder is a sequence of one-hot encoded arrays. In order to train the decoder we need to supply the one-hot encoded arrays that we desire to see on the decoder's output, and then use a loss-function like cross-entropy to train the decoder to produce this desired output.
However, our data-set contains integer-tokens instead of one-hot encoded arrays. Each one-hot encoded array has 10000 elements so it would be extremely wasteful to convert the entire data-set to one-hot encoded arrays.
A better way is to use a so-called sparse cross-entropy loss-function, which does the conversion internally from integers to one-hot encoded arrays.
```
model_train.compile(optimizer=keras.optimizers.RMSprop(learning_rate=1e-3),
loss='sparse_categorical_crossentropy')
```
### Callback functions
1. checkpoint
```
path_checkpoint = 'best-model-checkpoint.keras'
callback_checkpoint = keras.callbacks.ModelCheckpoint(filepath=path_checkpoint,
monitor='val_loss',
verbose=1,
save_weights_only=True,
save_best_only=True
)
```
2. early stopping
```
callback_early_stopping = keras.callbacks.EarlyStopping(monitor='val_loss',
patience=3, verbose=1)
callbacks = [
callback_checkpoint,
callback_early_stopping
]
```
### Trainning the model.
We wrap the data in named dicts so we are sure the data is assigned correctly to the inputs and outputs of the model.
```
x_data = {
'encoder_input': encoder_input_data,
'decoder_input': decoder_input_data
}
y_data = {
'decoder_output': decoder_output_data
}
```
Training this model, as single epoch will take aproximately an hour.
```
model_train.fit(x=x_data,
y=y_data,
batch_size=128,
epochs=10,
validation_split=.005,
callbacks=callbacks)
```
### Model inference
```
def translate(input_text, true_output_text=None):
input_tokens = tokenizer_src.text_to_tokens(text=input_text,
reverse=True,
padding=True)
initial_state = model_encoder.predict(input_tokens)
max_tokens = tokenizer_trg.max_tokens
decoder_input_data = np.zeros(shape=(1, max_tokens), dtype=np.int)
token_int = token_start
output_text = ''
count_tokens = 0
while token_int != token_end and count_tokens < max_tokens:
decoder_input_data[0, count_tokens] = token_int
x_data = {
'decoder_initial_state': initial_state,
'decoder_input': decoder_input_data
}
decoder_output = model_decoder.predict(x_data)
token_onehot = decoder_output[0, count_tokens, :]
token_int = np.argmax(token_onehot)
sampled_word = tokenizer_trg.token_to_word(token_int)
output_text += " " + sampled_word
count_tokens += 1
output_tokens = decoder_input_data[0]
print("Input text:")
print(input_text)
print()
print("Translated text:")
print(output_text)
print()
if true_output_text is not None:
print("True output text:")
print(true_output_text)
print()
```
### Examples
We are going to use our train data.
```
idx = 3
translate(input_text=data_src[idx],
true_output_text=data_trg[idx])
idx = 4
translate(input_text=data_src[idx],
true_output_text=data_trg[idx])
```
from the user
```
translate(input_text="der var engang et land der hed Danmark",
true_output_text='Once there was a country named Denmark')
translate(input_text="Hvem spæner ud af en butik og tygger de stærkeste bolcher?",
true_output_text="Who runs out of a shop and chews the strongest bon-bons?")
```
### Conclusion
I did not train the model for even a single epoch, the model only train for aproximately 30 min but was able to pick up some words. To improve the model performance we may want to train the model for more than 10 epochs to get accurate results.
#### Credits.
* [Hvass-Labs](https://github.com/Hvass-Labs/TensorFlow-Tutorials/blob/master/21_Machine_Translation.ipynb)
```
```
| github_jupyter |
# T81-558: Applications of Deep Neural Networks
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), School of Engineering and Applied Science, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
**Module 4 Assignment: Regression Neural Network**
**Student Name: Your Name**
# Assignment Instructions
For this assignment you will use the **reg-30-spring-2018.csv** dataset. This is a dataset that I generated specifically for this semester. You can find the CSV file in the **data** directory of the class GitHub repository here: [reg-30-spring-2018.csv](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/data/reg-30-spring-2018.csv).
For this assignment you will train a neural network and return the predictions. You will submit these predictions to the **submit** function. See [Assignment #1](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class1.ipynb) for details on how to submit an assignment or check that one was submitted.
Complete the following tasks:
* Normalize all numeric to zscores and all text/categorical to dummies. Do not normalize the *target*.
* Your target (y) is the filed named *target*.
* If you find any missing values (NA's), replace them with the median values for that column.
* No need for any cross validation or holdout. Just train on the entire data set for 250 epochs.
* You might get a warning, such as **"Warning: The mean of column pred differs from the solution file by 2.39"**. Do not worry about small values, it would be very hard to get exactly the same result as I did.
* Your submitted dataframe will have these columns: id, pred.
# Helpful Functions
You will see these at the top of every module and assignment. These are simply a set of reusable functions that we will make use of. Each of them will be explained as the semester progresses. They are explained in greater detail as the course progresses. Class 4 contains a complete overview of these functions.
```
from sklearn import preprocessing
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import shutil
import os
import requests
import base64
# Encode text values to dummy variables(i.e. [1,0,0],[0,1,0],[0,0,1] for red,green,blue)
def encode_text_dummy(df, name):
dummies = pd.get_dummies(df[name])
for x in dummies.columns:
dummy_name = "{}-{}".format(name, x)
df[dummy_name] = dummies[x]
df.drop(name, axis=1, inplace=True)
# Encode text values to a single dummy variable. The new columns (which do not replace the old) will have a 1
# at every location where the original column (name) matches each of the target_values. One column is added for
# each target value.
def encode_text_single_dummy(df, name, target_values):
for tv in target_values:
l = list(df[name].astype(str))
l = [1 if str(x) == str(tv) else 0 for x in l]
name2 = "{}-{}".format(name, tv)
df[name2] = l
# Encode text values to indexes(i.e. [1],[2],[3] for red,green,blue).
def encode_text_index(df, name):
le = preprocessing.LabelEncoder()
df[name] = le.fit_transform(df[name])
return le.classes_
# Encode a numeric column as zscores
def encode_numeric_zscore(df, name, mean=None, sd=None):
if mean is None:
mean = df[name].mean()
if sd is None:
sd = df[name].std()
df[name] = (df[name] - mean) / sd
# Convert all missing values in the specified column to the median
def missing_median(df, name):
med = df[name].median()
df[name] = df[name].fillna(med)
# Convert all missing values in the specified column to the default
def missing_default(df, name, default_value):
df[name] = df[name].fillna(default_value)
# Convert a Pandas dataframe to the x,y inputs that TensorFlow needs
def to_xy(df, target):
result = []
for x in df.columns:
if x != target:
result.append(x)
# find out the type of the target column. Is it really this hard? :(
target_type = df[target].dtypes
target_type = target_type[0] if hasattr(target_type, '__iter__') else target_type
# Encode to int for classification, float otherwise. TensorFlow likes 32 bits.
if target_type in (np.int64, np.int32):
# Classification
dummies = pd.get_dummies(df[target])
return df.as_matrix(result).astype(np.float32), dummies.as_matrix().astype(np.float32)
else:
# Regression
return df.as_matrix(result).astype(np.float32), df.as_matrix([target]).astype(np.float32)
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
# Regression chart.
def chart_regression(pred,y,sort=True):
t = pd.DataFrame({'pred' : pred, 'y' : y.flatten()})
if sort:
t.sort_values(by=['y'],inplace=True)
a = plt.plot(t['y'].tolist(),label='expected')
b = plt.plot(t['pred'].tolist(),label='prediction')
plt.ylabel('output')
plt.legend()
plt.show()
# Remove all rows where the specified column is +/- sd standard deviations
def remove_outliers(df, name, sd):
drop_rows = df.index[(np.abs(df[name] - df[name].mean()) >= (sd * df[name].std()))]
df.drop(drop_rows, axis=0, inplace=True)
# Encode a column to a range between normalized_low and normalized_high.
def encode_numeric_range(df, name, normalized_low=-1, normalized_high=1,
data_low=None, data_high=None):
if data_low is None:
data_low = min(df[name])
data_high = max(df[name])
df[name] = ((df[name] - data_low) / (data_high - data_low)) \
* (normalized_high - normalized_low) + normalized_low
# This function submits an assignment. You can submit an assignment as much as you like, only the final
# submission counts. The paramaters are as follows:
# data - Pandas dataframe output.
# key - Your student key that was emailed to you.
# no - The assignment class number, should be 1 through 1.
# source_file - The full path to your Python or IPYNB file. This must have "_class1" as part of its name.
# . The number must match your assignment number. For example "_class2" for class assignment #2.
def submit(data,key,no,source_file=None):
if source_file is None and '__file__' not in globals(): raise Exception('Must specify a filename when a Jupyter notebook.')
if source_file is None: source_file = __file__
suffix = '_class{}'.format(no)
if suffix not in source_file: raise Exception('{} must be part of the filename.'.format(suffix))
with open(source_file, "rb") as image_file:
encoded_python = base64.b64encode(image_file.read()).decode('ascii')
ext = os.path.splitext(source_file)[-1].lower()
if ext not in ['.ipynb','.py']: raise Exception("Source file is {} must be .py or .ipynb".format(ext))
r = requests.post("https://api.heatonresearch.com/assignment-submit",
headers={'x-api-key':key}, json={'csv':base64.b64encode(data.to_csv(index=False).encode('ascii')).decode("ascii"),
'assignment': no, 'ext':ext, 'py':encoded_python})
if r.status_code == 200:
print("Success: {}".format(r.text))
else: print("Failure: {}".format(r.text))
```
# Assignment #4 Sample Code
The following code provides a starting point for this assignment.
```
import os
import pandas as pd
from scipy.stats import zscore
from keras.models import Sequential
from keras.layers.core import Dense, Activation
import pandas as pd
import io
import requests
import numpy as np
from sklearn import metrics
# This is your student key that I emailed to you at the beginnning of the semester.
key = "qgABjW9GKV1vvFSQNxZW9akByENTpTAo2T9qOjmh" # This is an example key and will not work.
# You must also identify your source file. (modify for your local setup)
# file='/resources/t81_558_deep_learning/assignment_yourname_class1.ipynb' # IBM Data Science Workbench
# file='C:\\Users\\jeffh\\projects\\t81_558_deep_learning\\t81_558_class1_intro_python.ipynb' # Windows
# file='/Users/jeff/projects/t81_558_deep_learning/assignment_yourname_class1.ipynb' # Mac/Linux
file = '...location of your source file...'
# Begin assignment
path = "./data/"
filename_read = os.path.join(path,"reg-30-spring-2018.csv")
df = pd.read_csv(filename_read)
# Encode the feature vector
ids = df['id']
# Save a copy, if you like
submit_df.to_csv('4.csv',index=False)
# Submit the assignment
submit(source_file=file,data=submit_df,key=key,no=4)
```
| github_jupyter |
# Realization of Non-Recursive Filters
*This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing. Please direct questions and suggestions to [Sascha.Spors@uni-rostock.de](mailto:Sascha.Spors@uni-rostock.de).*
## Segmented Convolution
In many applications one of the signals of a convolution is much longer than the other. For instance when filtering a speech signal $x_L[k]$ of length $L$ with a room impulse response $h_N[k]$ of length $N \ll L$. In such cases the [fast convolution](fast_convolution.ipynb), as introduced before, does not bring a benefit since both signals have to be zero-padded to a total length of at least $N+L-1$. Applying the fast convolution may then even be impossible in terms of memory requirements or overall delay. The filtering of a signal which is captured in real-time is also not possible by the fast convolution.
In order to overcome these limitations, various techniques have been developed that perform the filtering on limited portions of the signals. These portions are known as partitions, segments or blocks. The respective algorithms are termed as *segmented* or *block-based* algorithms. The following section introduces two techniques for the segmented convolution of signals. The basic concept of these is to divide the convolution $y[k] = x_L[k] * h_N[k]$ into multiple convolutions operating on (overlapping) segments of the signal $x_L[k]$.
### Overlap-Add Algorithm
The [overlap-add algorithm](https://en.wikipedia.org/wiki/Overlap%E2%80%93add_method) is based on splitting the signal $x_L[k]$ into non-overlapping segments $x_p[k]$ of length $P$
\begin{equation}
x_L[k] = \sum_{p = 0}^{L/P - 1} x_p[k - p \cdot P]
\end{equation}
where the segments $x_p[k]$ are defined as
\begin{equation}
x_p[k] = \begin{cases} x_L[k + p \cdot P] & \text{ for } k=0,1,\dots,P-1 \\ 0 & \text{ otherwise} \end{cases}
\end{equation}
Note that $x_L[k]$ might have to be zero-padded so that its total length is a multiple of the segment length $P$. Introducing the segmentation of $x_L[k]$ into the convolution yields
\begin{align}
y[k] &= x_L[k] * h_N[k] \\
&= \sum_{p = 0}^{L/P - 1} x_p[k - p \cdot P] * h[k] \\
&= \sum_{p = 0}^{L/P - 1} y_p[k - p \cdot P]
\end{align}
where $y_p[k] = x_p[k] * h_N[k]$. This result states that the convolution of $x_L[k] * h_N[k]$ can be split into a series of convolutions $y_p[k]$ operating on the samples of one segment only. The length of $y_p[k]$ is $N+P-1$. The result of the overall convolution is given by summing up the results from the segments shifted by multiples of the segment length $P$. This can be interpreted as an overlapped superposition of the results from the segments, as illustrated in the following diagram

The overall procedure is denoted by the name *overlap-add* technique. The convolutions $y_p[k] = x_p[k] * h_N[k]$ can be realized efficiently by the [fast convolution](fast_convolution.ipynb) using zero-padding and fast Fourier transformations (FFTs) of length $M \geq P+N-1$.
A drawback of the overlap-add technique is that the next input segment is required to compute the result for the actual segment of the output. For real-time applications this introduces an algorithmic delay of one segment.
#### Example
The following example illustrates the overlap-add algorithm by showing the (convolved) segments and the overall result.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import scipy.signal as sig
L = 64 # length of input signal
N = 8 # length of impulse response
P = 16 # length of segments
# generate input signal
x = sig.triang(L)
# generate impulse response
h = sig.triang(N)
# overlap-add convolution
xp = np.zeros((L//P, P))
yp = np.zeros((L//P, N+P-1))
y = np.zeros(L+P-1)
for n in range(L//P):
xp[n, :] = x[n*P:(n+1)*P]
yp[n, :] = np.convolve(xp[n,:], h, mode='full')
y[n*P:(n+1)*P+N-1] += yp[n, :]
y = y[0:N+L]
# plot signals
plt.figure(figsize = (10,2))
plt.subplot(121)
plt.stem(x)
for n in np.arange(L//P)[::2]:
plt.axvspan(n*P, (n+1)*P-1, facecolor='g', alpha=0.5)
plt.title(r'Signal $x[k]$ and segments')
plt.xlabel(r'$k$')
plt.ylabel(r'$x[k]$')
plt.axis([0, L, 0, 1])
plt.subplot(122)
plt.stem(h)
plt.title(r'Impulse response $h[k]$')
plt.xlabel(r'$k$')
plt.ylabel(r'$h[k]$')
plt.axis([0, L, 0, 1])
for p in np.arange(L//P):
plt.figure(figsize = (10,2))
plt.stem(np.concatenate((np.zeros(p*P), yp[p, :])))
plt.title(r'Result of segment $p=%d$' %(p))
plt.xlabel(r'$k$')
plt.ylabel(r'$y_%d[k - %d P]$' %(p,p))
plt.axis([0, L+P, 0, 4])
plt.figure(figsize = (10,2))
plt.stem(y)
plt.title(r'Result $y[k] = x[k] * h[k]$')
plt.xlabel(r'$k$')
plt.ylabel(r'$y[k]$')
plt.axis([0, L+P, 0, 4]);
```
**Exercises**
* Change the length `N` of the impulse response and the length `P` of the segments. What changes?
* What influence have these two lengths on the numerical complexity of the overlap-add algorithm?
### Overlap-Save Algorithm
The [overlap-save](https://en.wikipedia.org/wiki/Overlap%E2%80%93save_method) algorithm, also known as *overlap-discard algorithm*, follows a different strategy as the overlap-add technique introduced above. It is based on an overlapping segmentation of the input $x_L[k]$ and application of the periodic convolution for the individual segments.
Lets take a closer look at the result of the periodic convolution $x_p[k] \circledast h_N[k]$, where $x_p[k]$ denotes a segment of length $P$ of the input signal and $h_N[k]$ the impulse response of length $N$. The result of a linear convolution $x_p[k]* h_N[k]$ would be of length $P + N -1$. The result of the periodic convolution of period $P$ for $P > N$ would suffer from a circular shift (time aliasing) and superposition of the last $N-1$ samples to the beginning. Hence, the first $N-1$ samples are not equal to the result of the linear convolution. However, the remaining $P- N + 1$ do so.
This motivates to split the input signal $x_L[k]$ into overlapping segments of length $P$ where the $p$-th segment overlaps its preceding $(p-1)$-th segment by $N-1$ samples
\begin{equation}
x_p[k] = \begin{cases}
x_L[k + p \cdot (P-N+1) - (N-1)] & \text{ for } k=0,1, \dots, P-1 \\
0 & \text{ otherwise}
\end{cases}
\end{equation}
The part of the circular convolution $x_p[k] \circledast h_N[k]$ of one segment $x_p[k]$ with the impulse response $h_N[k]$ that is equal to the linear convolution of both is given as
\begin{equation}
y_p[k] = \begin{cases}
x_p[k] \circledast h_N[k] & \text{ for } k=N-1, N, \dots, P-1 \\
0 & \text{ otherwise}
\end{cases}
\end{equation}
The output $y[k]$ is simply the concatenation of the $y_p[k]$
\begin{equation}
y[k] = \sum_{p=0}^{L/P - 1} y_p[k - p \cdot (P-N+1) + (N-1)]
\end{equation}
The overlap-save algorithm is illustrated in the following diagram

For the first segment $x_0[k]$, $N-1$ zeros have to be appended to the beginning of the input signal $x_L[k]$ for the overlapped segmentation. From the result of the periodic convolution $x_p[k] \circledast h_N[k]$ the first $N-1$ samples are discarded, the remaining $P - N + 1$ are copied to the output $y[k]$. This is indicated by the alternative notation *overlap-discard* used for the technique. The periodic convolution can be realized efficiently by a FFT/IFFT of length $P$.
#### Example
The following example illustrates the overlap-save algorithm by showing the results of the periodic convolutions of the segments. The discarded parts are indicated by the red background.
```
L = 64 # length of input signal
N = 8 # length of impulse response
P = 24 # length of segments
# generate input signal
x = sig.triang(L)
# generate impulse response
h = sig.triang(N)
# overlap-save convolution
nseg = (L+N-1)//(P-N+1) + 1
x = np.concatenate((np.zeros(N-1), x, np.zeros(P)))
xp = np.zeros((nseg, P))
yp = np.zeros((nseg, P))
y = np.zeros(nseg*(P-N+1))
for p in range(nseg):
xp[p, :] = x[p*(P-N+1):p*(P-N+1)+P]
yp[p, :] = np.fft.irfft(np.fft.rfft(xp[p, :]) * np.fft.rfft(h, P))
y[p*(P-N+1):p*(P-N+1)+P-N+1] = yp[p, N-1:]
y = y[0:N+L]
plt.figure(figsize = (10,2))
plt.subplot(121)
plt.stem(x[N-1:])
plt.title(r'Signal $x[k]$')
plt.xlabel(r'$k$')
plt.ylabel(r'$x[k]$')
plt.axis([0, L, 0, 1])
plt.subplot(122)
plt.stem(h)
plt.title(r'Impulse response $h[k]$')
plt.xlabel(r'$k$')
plt.ylabel(r'$h[k]$')
plt.axis([0, L, 0, 1])
for p in np.arange(nseg):
plt.figure(figsize = (10,2))
plt.stem(yp[p, :])
plt.axvspan(0, N-1+.5, facecolor='r', alpha=0.5)
plt.title(r'Result of periodic convolution of $x_%d[k]$ and $h_N[k]$' %(p))
plt.xlabel(r'$k$')
plt.axis([0, L+P, 0, 4])
plt.figure(figsize = (10,2))
plt.stem(y)
plt.title(r'Result $y[k] = x[k] * h[k]$')
plt.xlabel(r'$k$')
plt.ylabel(r'$y[k]$')
plt.axis([0, L+P, 0, 4]);
```
**Exercise**
* Change the length `N` of the impulse response and the length `P` of the segments. What changes?
* How many samples of the output signal $y[k]$ are computed per segment for a particular choice of these two values?
* What would be a good choice for the segment length `P` with respect to the length `N` of the impulse response?
### Practical Aspects and Extensions
* For both the overlap-add and overlap-save algorithm the length $P$ of the segments influences the lengths of the convolutions, FFTs and the number of output samples per segment. The segment length is often chosen as
* $P=N$ for overlap-add and
* $P = 2 N$ for overlap-save.
For both algorithms this requires FFTs of length $2 N$ to compute $P$ output samples. The overlap-add algorithm requires $P$ additional additions per segment in comparison to overlap-save.
* For real-valued signals $x_L[k]$ and impulse responses $h_N[k]$ real-valued FFTs lower the computational complexity significantly. As alternative, the $2 N$ samples in the FFT can be distributed into the real and complex part of a FFT of length $N$ [[Zölzer](../index.ipynb#Literature)].
* The impulse response can be changed in each segment in order to simulate time-variant linear systems. This is often combined with an overlapping computation of the output in order to avoid artifacts due to instationarities.
* For long impulse responses $h_N[k]$ or low-delay applications, algorithms have been developed which base on an additional segmentation of the impulse response. This is known as *partitioned convolution*.
**Copyright**
This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Sascha Spors, Digital Signal Processing - Lecture notes featuring computational examples, 2016-2017*.
| github_jupyter |
# Federated FastEstimator FGSM Tutorial
```
#Install dependencies if not already installed
!pip install tensorflow'>=2.3' torch'>=1.6' fastestimator
import fastestimator as fe
import tempfile
from fastestimator.architecture.pytorch import LeNet
from fastestimator.backend import to_tensor, argmax
from fastestimator.dataset.data import cifar10
from fastestimator.op.numpyop.meta import Sometimes
from fastestimator.op.numpyop.multivariate import HorizontalFlip
from fastestimator.op.numpyop.univariate import CoarseDropout, Normalize, Onehot
from fastestimator.op.tensorop import Average
from fastestimator.op.tensorop.gradient import Watch, FGSM
from fastestimator.op.tensorop.loss import CrossEntropy
from fastestimator.op.tensorop.model import ModelOp, UpdateOp
from fastestimator.trace.io import BestModelSaver
from fastestimator.trace.metric import Accuracy
from fastestimator.util import ImgData, to_number
from openfl.native.fastestimator import FederatedFastEstimator
from fastestimator.dataset.data import cifar10
from fastestimator.trace.adapt import LRScheduler,ReduceLROnPlateau
from fastestimator.op.numpyop.univariate import Normalize, ChannelTranspose
batch_size=128
train_data, eval_data = cifar10.load_data()
test_data = eval_data.split(0.5)
pipeline = fe.Pipeline(train_data=train_data,
eval_data=eval_data,
test_data=test_data,
batch_size=batch_size,
ops=[Normalize(inputs="x", outputs="x",
mean=(0.4914, 0.4822, 0.4465),
std=(0.2471, 0.2435, 0.2616)),
ChannelTranspose(inputs="x", outputs="x")])
model = fe.build(model_fn=lambda: LeNet(input_shape=(3, 32, 32)), \
optimizer_fn="adam", model_name="adv_model")
network = fe.Network(ops=[
Watch(inputs="x"),
ModelOp(model=model, inputs="x", outputs="y_pred"),
CrossEntropy(inputs=("y_pred", "y"), outputs="base_ce"),
FGSM(data="x", loss="base_ce", outputs="x_adverse", epsilon=0.04),
ModelOp(model=model, inputs="x_adverse", outputs="y_pred_adv"),
CrossEntropy(inputs=("y_pred_adv", "y"), outputs="adv_ce"),
Average(inputs=("base_ce", "adv_ce"), outputs="avg_ce"),
UpdateOp(model=model, loss_name="avg_ce")
])
estimator = fe.Estimator(pipeline=pipeline,
network=network,
epochs=1,
traces=[Accuracy(true_key="y", pred_key="y_pred", output_name="clean_accuracy"),
Accuracy(true_key="y", pred_key="y_pred_adv", output_name="adversarial_accuracy"),
ReduceLROnPlateau(model=model,metric='base_ce',patience=2),
BestModelSaver(model=model, save_dir=tempfile.mkdtemp(), metric="base_ce", save_best_mode="min",load_best_final=True),],
max_train_steps_per_epoch=None,
max_eval_steps_per_epoch=None,
monitor_names=["base_ce", "adv_ce"],
log_steps=50)
openfl_estimator = FederatedFastEstimator(estimator, override_config={'aggregator.settings.rounds_to_train':5})
model=openfl_estimator.fit()
model.state_dict()
```
| github_jupyter |
# Introducción a la visualización con matplotlib
_Después de estudiar la sintaxis de Python y empezar a manejar datos numéricos de manera un poco más profesional, ha llegado el momento de visualizarlos. Con la biblioteca **matplotlib** podemos crear gráficos de muy alta calidad y altamente personalizables._
_matplotlib es una biblioteca muy potente que requiere tiempo de práctica para dominarla. Vamos a empezar por lo más sencillo._
## ¿Qué es matplotlib?
* Estándar *de facto* para visualización en Python
* Pretende ser similar a las funciones de visualización de MATLAB
* Diferentes formas de usarla: interfaz `pyplot` y orientada a objetos
Lo primero que vamos a hacer es activar el modo *inline* - de esta manera las figuras aparecerán automáticamente incrustadas en el notebook.
```
%matplotlib inline
```
Importamos los paquetes necesarios:
```
import numpy as np
import matplotlib.pyplot as plt
```
La biblioteca matplotlib es gigantesca y es difícil hacerse una idea global de todas sus posibilidades en una primera toma de contacto. Es recomendable tener a mano la documentación y la galería (http://matplotlib.org/gallery.html#pylab_examples):
```
from IPython.display import HTML
HTML('<iframe src="http://matplotlib.org/gallery.html#pylab_examples" width="800" height="600"></iframe>')
```
Si hacemos clic en cualquiera de las imágenes, accedemos al código fuente que la ha generado (ejemplo: http://matplotlib.org/examples/pylab_examples/annotation_demo.html):
```
HTML('<iframe src="http://matplotlib.org/examples/pylab_examples/annotation_demo.html" width="800" height="600"></iframe>')
```
## Interfaz pyplot
La interfaz `pyplot` proporciona una serie de funciones que operan sobre un *estado global* - es decir, nosotros no especificamos sobre qué gráfica o ejes estamos actuando. Es una forma rápida y cómoda de crear gráficas pero perdemos parte del control.
### Función `plot`
El paquete `pyplot` se suele importar bajo el alias `plt`, de modo que todas las funciones se acceden a través de `plt.<funcion>`. La función más básica es la función `plot`:
```
plt
plt.plot([0.0, 0.1, 0.2, 0.7, 0.9], [1, -2, 3, 4, 1])
```
La función `plot` recibe una sola lista (si queremos especificar los valores *y*) o dos listas (si especificamos *x* e *y*). Naturalmente si especificamos dos listas ambas tienen que tener la misma longitud.
La tarea más habitual a la hora de trabajar con matplotlib es representar una función. Lo que tendremos que hacer es definir un dominio y evaluarla en dicho dominio. Por ejemplo:
$$ f(x) = e^{-x^2} $$
```
def f(x):
return np.exp(-x ** 2)
```
Definimos el dominio con la función `np.linspace`, que crea un vector de puntos equiespaciados:
```
x = np.linspace(-1, 3, 100)
```
Y representamos la función:
```
plt.plot(x, f(x), label="Función f(x)")
plt.xlabel("Eje $x$")
plt.ylabel("$f(x)$")
plt.legend()
plt.title("Función $f(x)$")
```
Notamos varias cosas:
* Con diversas llamadas a funciones dentro de `plt.` se actualiza el gráfico *actual*. Esa es la forma de trabajar con la interfaz pyplot.
* Podemos añadir etiquetas, y escribir $\LaTeX$ en ellas. Tan solo hay que encerrarlo entre signos de dólar $$.
* Añadiendo como argumento `label` podemos definir una leyenda.
### Personalización
La función `plot` acepta una serie de argumentos para personalizar el aspecto de la función. Con una letra podemos especificar el color, y con un símbolo el tipo de línea.
```
plt.plot(x, f(x), 'ro')
plt.plot(x, 1 - f(x), 'g--')
```
Esto en realidad son códigos abreviados, que se corresponden con argumentos de la función `plot`:
```
plt.plot(x, f(x), color='red', linestyle='', marker='o')
plt.plot(x, 1 - f(x), c='g', ls='--')
```
La lista de posibles argumentos y abreviaturas está disponible en la documentación de la función `plot` http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot.
### Más personalización, pero a lo loco
Desde matplotlib 1.4 se puede manipular fácilmente la apariencia de la gráfica usando **estilos**. Para ver qué estilos hay disponibles, escribiríamos `plt.style.available`.
```
plt.style.available
```
No hay muchos pero podemos crear los nuestros. Para activar uno de ellos, usamos `plt.style.use`. ¡Aquí va el que uso yo! https://gist.github.com/Juanlu001/edb2bf7b583e7d56468a
```
#plt.style.use("ggplot") # Afecta a todos los plots
```
<div class="alert alert-warning">No he sido capaz de encontrar una manera fácil de volver a la apariencia por defecto en el notebook. A ver qué dicen los desarrolladores (https://github.com/ipython/ipython/issues/6707) ¡pero de momento si quieres volver a como estaba antes toca reiniciar el notebook!</div>
Para emplear un estilo solo a una porción del código, creamos un bloque `with plt.style.context("STYLE")`:
```
with plt.style.context('ggplot'):
plt.plot(x, f(x))
plt.plot(x, 1 - f(x))
```
Y hay otro tipo de personalización más loca todavía:
```
with plt.xkcd():
plt.plot(x, f(x))
plt.plot(x, 1 - f(x))
plt.xlabel("Eje x")
plt.ylabel("Eje y")
```
¡Nunca imitar a XKCD fue tan fácil! http://xkcd.com/353/
### Otros tipo de gráficas
La función `scatter` muestra una nube de puntos, con posibilidad de variar también el tamaño y el color.
```
N = 100
x = np.random.randn(N)
y = np.random.randn(N)
plt.scatter(x, y)
```
Con `s` y `c` podemos modificar el tamaño y el color respectivamente. Para el color, a cada valor numérico se le asigna un color a través de un *mapa de colores*; ese mapa se puede cambiar con el argumento `cmap`. Esa correspondencia se puede visualizar llamando a la función `colorbar`.
```
s = np.abs(50 + 50 * np.random.randn(N))
c = np.random.randn(N)
plt.scatter(x, y, s=s, c=c, cmap=plt.cm.Blues)
plt.colorbar()
plt.scatter(x, y, s=s, c=c, cmap=plt.cm.Oranges)
plt.colorbar()
```
matplotlib trae por defecto muchos mapas de colores. En las SciPy Lecture Notes dan una lista de todos ellos (http://scipy-lectures.github.io/intro/matplotlib/matplotlib.html#colormaps)

La función `contour` se utiliza para visualizar las curvas de nivel de funciones de dos variables y está muy ligada a la función `np.meshgrid`. Veamos un ejemplo:
$$f(x) = x^2 - y^2$$
```
def f(x, y):
return x ** 2 - y ** 2
x = np.linspace(-2, 2)
y = np.linspace(-2, 2)
xx, yy = np.meshgrid(x, y)
zz = f(xx, yy)
plt.contour(xx, yy, zz)
plt.colorbar()
```
La función `contourf` es casi idéntica pero rellena el espacio entre niveles. Podemos especificar manualmente estos niveles usando el cuarto argumento:
```
plt.contourf(xx, yy, zz, np.linspace(-4, 4, 100))
plt.colorbar()
```
Para guardar las gráficas en archivos aparte podemos usar la función `plt.savefig`. matplotlib usará el tipo de archivo adecuado según la extensión que especifiquemos. Veremos esto con más detalle cuando hablemos de la interfaz orientada a objetos.
### Varias figuras
Podemos crear figuras con varios sistemas de ejes, pasando a `subplot` el número de filas y de columnas.
```
x = np.linspace(-1, 7, 1000)
fig = plt.figure()
plt.subplot(211)
plt.plot(x, np.sin(x))
plt.grid(False)
plt.title("Función seno")
plt.subplot(212)
plt.plot(x, np.cos(x))
plt.grid(False)
plt.title("Función coseno")
```
<div class="alert alert-info">¿Cómo se ajusta el espacio entre gráficas para que no se solapen los textos? Buscamos en Google "plt.subplot adjust" en el primer resultado tenemos la respuesta http://stackoverflow.com/a/9827848</div>
Como hemos guardado la figura en una variable, puedo recuperarla más adelate y seguir editándola.
```
fig.tight_layout()
fig
```
<div class="alert alert-warning">Si queremos manipular la figura una vez hemos abandonado la celda donde la hemos definido, tendríamos que utilizar la interfaz orientada a objetos de matplotlib. Es un poco lioso porque algunas funciones cambian de nombre, así que en este curso no la vamos a ver. Si te interesa puedes ver los notebooks de la primera edición, donde sí la introdujimos.
https://github.com/AeroPython/Curso_AeroPython/releases/tag/v1.0</div>
**Ejercicio**
Crear una función que represente gráficamente esta expresión:
$$\sin(2 \pi f_1 t) + \sin(2 \pi f_2 t)$$
Siendo $f_1$ y $f_2$ argumentos de entrada (por defecto $10$ y $100$) y $t \in [0, 0.5]$. Además, debe mostrar:
* leyenda,
* título "Dos frecuencias",
* eje x "Tiempo ($t$)"
y usar algún estilo de los disponibles.
```
def frecuencias(f1=10.0, f2=100.0):
max_time = 0.5
times = np.linspace(0, max_time, 1000)
signal = np.sin(2 * np.pi * f1 * times) + np.sin(2 * np.pi * f2 * times)
with plt.style.context("ggplot"):
plt.plot(signal, label="Señal")
plt.xlabel("Tiempo ($t$)")
plt.title("Dos frecuencias")
plt.legend()
frecuencias()
```
**Ejercicio**
Representar las curvas de nivel de esta función:
$$g(x, y) = \cos{x} + \sin^2{y}$$
Para obtener este resultado:

```
def g(x, y):
return np.cos(x) + np.sin(y) ** 2
# Necesitamos muchos puntos en la malla, para que cuando se
# crucen las líneas no se vean irregularidades
x = np.linspace(-2, 3, 1000)
y = np.linspace(-2, 3, 1000)
xx, yy = np.meshgrid(x, y)
zz = g(xx, yy)
# Podemos ajustar el tamaño de la figura con figsize
fig = plt.figure(figsize=(6, 6))
# Ajustamos para que tenga 13 niveles y que use el colormap Spectral
# Tenemos que asignar la salida a la variable cs para luego crear el colorbar
cs = plt.contourf(xx, yy, zz, np.linspace(-1, 2, 13), cmap=plt.cm.Spectral)
# Creamos la barra de colores
plt.colorbar()
# Con `colors='k'` dibujamos todas las líneas negras
# Asignamos la salida a la variable cs2 para crear las etiquetas
cs = plt.contour(xx, yy, zz, np.linspace(-1, 2, 13), colors='k')
# Creamos las etiquetas sobre las líneas
plt.clabel(cs)
# Ponemos las etiquetas de los ejes
plt.xlabel("Eje x")
plt.ylabel("Eje y")
plt.title(r"Función $g(x, y) = \cos{x} + \sin^2{y}$")
```
## Referencias
* Guía de matplotlib para principiantes http://matplotlib.org/users/beginner.html
* Tutorial de matplotlib en español http://pybonacci.org/tag/tutorial-matplotlib-pyplot/
* Referencia rápida de matplotlib http://scipy-lectures.github.io/intro/matplotlib/matplotlib.html#quick-references
---
<br/>
#### <h4 align="right">¡Síguenos en Twitter!
<br/>
###### <a href="https://twitter.com/AeroPython" class="twitter-follow-button" data-show-count="false">Follow @AeroPython</a> <script>!function(d,s,id){var js,fjs=d.getElementsByTagName(s)[0],p=/^http:/.test(d.location)?'http':'https';if(!d.getElementById(id)){js=d.createElement(s);js.id=id;js.src=p+'://platform.twitter.com/widgets.js';fjs.parentNode.insertBefore(js,fjs);}}(document, 'script', 'twitter-wjs');</script>
<br/>
###### Este notebook ha sido realizado por: Juan Luis Cano, y Álex Sáez
<br/>
##### <a rel="license" href="http://creativecommons.org/licenses/by/4.0/deed.es"><img alt="Licencia Creative Commons" style="border-width:0" src="http://i.creativecommons.org/l/by/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" property="dct:title">Curso AeroPython</span> por <span xmlns:cc="http://creativecommons.org/ns#" property="cc:attributionName">Juan Luis Cano Rodriguez y Alejandro Sáez Mollejo</span> se distribuye bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/deed.es">Licencia Creative Commons Atribución 4.0 Internacional</a>.
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Goal" data-toc-modified-id="Goal-1"><span class="toc-item-num">1 </span>Goal</a></span></li><li><span><a href="#Var" data-toc-modified-id="Var-2"><span class="toc-item-num">2 </span>Var</a></span></li><li><span><a href="#Init" data-toc-modified-id="Init-3"><span class="toc-item-num">3 </span>Init</a></span></li><li><span><a href="#Load" data-toc-modified-id="Load-4"><span class="toc-item-num">4 </span>Load</a></span></li><li><span><a href="#Get-SRA-runinfo-for-BioProjects" data-toc-modified-id="Get-SRA-runinfo-for-BioProjects-5"><span class="toc-item-num">5 </span>Get SRA runinfo for BioProjects</a></span></li><li><span><a href="#Get-sample-metadata" data-toc-modified-id="Get-sample-metadata-6"><span class="toc-item-num">6 </span>Get sample metadata</a></span></li><li><span><a href="#Dataset-summary" data-toc-modified-id="Dataset-summary-7"><span class="toc-item-num">7 </span>Dataset summary</a></span><ul class="toc-item"><li><span><a href="#per-bioproject" data-toc-modified-id="per-bioproject-7.1"><span class="toc-item-num">7.1 </span>per-bioproject</a></span></li></ul></li><li><span><a href="#Creating-sample-files-for-LLMGQC" data-toc-modified-id="Creating-sample-files-for-LLMGQC-8"><span class="toc-item-num">8 </span>Creating sample files for LLMGQC</a></span></li><li><span><a href="#sessionInfo" data-toc-modified-id="sessionInfo-9"><span class="toc-item-num">9 </span>sessionInfo</a></span></li></ul></div>
# Goal
* Adding to existing database of animal gut/feces metagenomes
* metagenomes from various studies
# Var
```
work_dir = '/ebio/abt3_projects/Georg_animal_feces/data/metagenome/multi-study/'
metadata_file = file.path(work_dir, 'metadata', 'study_metadata_v2.xlsx')
```
# Init
```
library(dplyr)
library(tidyr)
library(ggplot2)
library(readxl)
source('/ebio/abt3_projects/Georg_animal_feces/code/misc_r_functions/init.R')
```
# Load
```
metadata = readxl::read_xlsx(metadata_file, sheet='Round2') %>%
filter(! is.na(BioProject))
metadata %>% dfhead
```
# Get SRA runinfo for BioProjects
```
# function for getting files via entrez-direct
get_SRA_runinfo = function(bioproject, out_dir){
out_file = file.path(out_dir, paste0(bioproject, '_runinfo.csv'))
cmd = 'esearch -db sra -query {bioproject} | efetch --format runinfo > {out_file}'
cmd = glue::glue(cmd, bioproject = bioproject, out_file = out_file)
bash_job(cmd, conda_env='py3_genome')
return(bioproject)
}
# fetching data
out_dir = file.path(work_dir, 'SRA_runinfo')
make_dir(out_dir)
X = as.list(metadata$BioProject)
lapply(X, get_SRA_runinfo, out_dir = out_dir)
# listing output
sra_info_files = list.files(out_dir, '*_runinfo.csv', full.names=TRUE)
sra_info_files = data.frame(bioproject = gsub('_runinfo.csv', '', basename(as.character(sra_info_files))),
file_path = sra_info_files)
sra_info_files %>% dfhead
# which (if any) are missing?
metadata %>%
anti_join(sra_info_files, c('BioProject'='bioproject'))
# loading the runinfo
sra_runinfo = list()
for(F in sra_info_files$file_path){
z = file.info(F)
if(!is.na(z$size) && z$size > 20){
X = read.delim(F, sep=',') %>%
mutate(spots = as.Num(spots),
bases = as.Num(bases),
avgLength = as.Num(avgLength),
size_MB = as.Num(size_MB),
TaxID = as.Num(TaxID))
X$bioproject = gsub('_runinfo.csv', '', basename(F))
sra_runinfo[[F]] = X
}
}
sra_runinfo = do.call(rbind, sra_runinfo)
rownames(sra_runinfo) = 1:nrow(sra_runinfo)
sra_runinfo %>% dfhead
# summary
sra_runinfo %>%
dplyr::select(Run, spots, bases, avgLength, size_MB, LibraryStrategy,
LibraryLayout, Platform, Model, TaxID, ScientificName) %>%
summary
# filtering
sra_runinfo_f = sra_runinfo %>%
filter(spots > 1e6,
avgLength >= 100,
! LibraryStrategy %in% c('Hi-C', 'AMPLICON'),
LibraryLayout == 'PAIRED',
! Platform %in% c('ION_TORRENT'),
! TaxID %in% c(564, 315405, 1263854))
sra_runinfo_f %>% nrow %>% print
sra_runinfo_f$bioproject %>% unique %>% length %>% print
sra_runinfo_f %>%
dplyr::select(Run, spots, bases, avgLength, size_MB, LibraryStrategy,
LibraryLayout, Platform, Model, TaxID, ScientificName) %>%
summary
```
# Get sample metadata
* For each sample, get NCBI biosample data
* Using entrez + xml formatting script:
```
esearch -db sra -query <Sample> | elink -target biosample | efetch -format xml | cat | ./ncbi_biosample_info.py -
```
```
# creating commands
make_cmd = function(Sample, out_dir){
exe = '/ebio/abt3_projects/Georg_animal_feces/data/metagenome/multi-study/ncbi_biosample_info.py'
out_dir = file.path(out_dir, Sample)
make_dir(out_dir, quiet=TRUE)
out_file = file.path(out_dir,'biosample_info.tsv')
cmd = 'esearch -db sra -query {Sample} | elink -target biosample | efetch -format xml | cat | {exe} - {out_file}'
cmd = glue::glue(cmd, Sample = Sample, exe = exe, out_file = out_file)
return(cmd)
}
out_dir = file.path(work_dir, 'SRA_biosample_info')
make_dir(out_dir)
X = sra_runinfo_f$Sample %>% unique %>% as.list
cmds = lapply(X, make_cmd, out_dir=out_dir)
cmds %>% length %>% print
cmds[[1]]
conda_env = 'py3_genome'
ret = lapply(cmds, function(x) bash_job(x, conda_env=conda_env))
```
**Note:** I re-ran those failed commands 'manually' on the command line
```
# list output files
biosample_info_files = list.files(out_dir, 'biosample_info.tsv', full.names=TRUE, recursive=TRUE)
biosample_info_files %>% length %>% print
# combining output
biosample_info = list()
for(F in biosample_info_files){
df = read.delim(F, sep='\t', header=FALSE)
colnames(df) = c('Sample', 'BioSample', 'Tag', 'Attribute')
biosample_info[[F]] = df
}
biosample_info = do.call(rbind, biosample_info)
rownames(biosample_info) = 1:nrow(biosample_info)
biosample_info %>% dfhead
# checking that all samples are present in bioinfo table
setdiff(sra_runinfo_f$Sample, biosample_info$Sample) %>% length %>% print
setdiff(biosample_info$Sample, sra_runinfo_f$Sample) %>% length %>% print
```
# Dataset summary
```
# formatting and joining
tags = c('Id', 'Id__2', 'Title', 'Alias', 'Organism', 'Name', 'host', 'SRA_accession' ,
'collection_date', 'env_material', 'env_biome')
biosample_info_j = biosample_info %>%
filter(Tag %in% tags) %>%
spread(Tag, Attribute) %>%
inner_join(sra_runinfo_f %>%
dplyr::select(Sample, BioSample, Run, BioProject, spots, bases, avgLength, size_MB,
LibraryStrategy, LibraryLayout, Platform, Model, TaxID, ScientificName),
c('Sample', 'BioSample'))
biosample_info_j %>% dfhead
```
## per-bioproject
```
# env_material
p = biosample_info_j %>%
group_by(BioProject, env_material) %>%
summarize(n = n()) %>%
ungroup() %>%
ggplot(aes(BioProject, n, fill=env_material)) +
geom_bar(stat='identity', position='dodge') +
scale_y_log10() +
labs(y='No. of runs') +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(8, 3)
plot(p)
# env_biome
p = biosample_info_j %>%
group_by(BioProject, env_biome) %>%
summarize(n = n()) %>%
ungroup() %>%
ggplot(aes(BioProject, n, fill=env_biome)) +
geom_bar(stat='identity', position='dodge') +
scale_y_log10() +
labs(y='No. of runs') +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(8, 3)
plot(p)
# host
p = biosample_info_j %>%
group_by(BioProject, host) %>%
summarize(n = n()) %>%
ungroup() %>%
ggplot(aes(BioProject, n, fill=host)) +
geom_bar(stat='identity', position='dodge') +
scale_y_log10() +
labs(y='No. of runs') +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(8, 4)
plot(p)
```
# Creating sample files for LLMGQC
* separating by bioproject
```
# formatting table
sra_runinfo_f_f = sra_runinfo_f %>%
mutate(Lane = 1,
Remote = Run) %>%
rename('SampleID' = Sample,
'Sample' = SampleName) %>%
dplyr::select(Sample, Run, Lane, Remote,
spots, bases, avgLength, size_MB,
BioProject, bioproject, Study_Pubmed_id, BioSample,
SampleID, SampleType, TaxID, ScientificName,
LibraryName, LibraryStrategy, LibrarySelection, LibrarySource, LibraryLayout,
InsertSize, InsertDev, Platform, Model, source)
sra_runinfo_f_f %>% dfhead
# adding relevant biosample info
tags = c('Title', 'Alias', 'Name', 'host', 'SRA_accession' ,
'collection_date', 'env_material', 'env_biome')
sra_runinfo_f_f = sra_runinfo_f_f %>%
inner_join(biosample_info %>% filter(Tag %in% tags) %>% spread(Tag, Attribute),
c('SampleID'='Sample', 'BioSample'))
sra_runinfo_f_f %>% dfhead
# adding manually curated project metadata
sra_runinfo_f_f = sra_runinfo_f_f %>%
left_join(metadata, c('bioproject'='BioProject'))
sra_runinfo_f_f %>% dfhead
# number of runs per bioproject
sra_runinfo_f_f$BioProject %>% as.character %>% table
cat('-----\n')
# number of samples per bioproject
sra_runinfo_f_f %>% distinct(BioProject, Sample) %>% .$BioProject %>% as.character %>% table
# number of runs per bioproject
sra_runinfo_f_f %>%
group_by(BioProject, bioproject) %>%
summarize(n_runs = n()) %>%
ungroup() %>%
arrange(n_runs)
# read lengths per bioproject
p = sra_runinfo_f_f %>%
ggplot(aes(BioProject, avgLength)) +
geom_boxplot() +
theme_bw() +
theme(
axis.text.x = element_text(angle=45, hjust=1)
)
dims(6, 3)
plot(p)
D = file.path(work_dir, 'BioProjects_v2')
make_dir(D)
bioprojects = sra_runinfo_f_f$BioProject %>% unique
for(B in bioprojects){
df = sra_runinfo_f_f %>% filter(BioProject == B)
out_dir = file.path(work_dir, 'BioProjects_v2', B)
make_dir(out_dir)
out_file = file.path(out_dir, 'samples.txt')
write.table(df, out_file, sep='\t', quote=FALSE, row.names=FALSE)
cat('File written:', out_file, '\n')
}
```
# sessionInfo
```
sessionInfo()
```
| github_jupyter |
# Evaluate for sentiment analysis model
```
import sys
sys.path.append("..")
sys.path.append("../../")
import paddle
import paddlenlp
from paddlenlp.transformers import ErnieForSequenceClassification, ErnieTokenizer
```
Initialize the model and tokenizer
```
MODEL_NAME = "ernie-1.0"
model = ErnieForSequenceClassification.from_pretrained(MODEL_NAME, num_classes=2)
tokenizer = ErnieTokenizer.from_pretrained(MODEL_NAME)
```
load model paramerters
```
from paddlenlp.datasets import load_dataset
DATASET_NAME = 'chnsenticorp'
train_ds, dev_ds, test_ds = load_dataset(DATASET_NAME, splits=["train", "dev", "test"])
# Load the trained model.
!wget --no-check-certificate -c https://trustai.bj.bcebos.com/chnsenticorp-ernie-1.0.tar
!tar -xvf ./chnsenticorp-ernie-1.0.tar -C ../assets/
!rm ./chnsenticorp-ernie-1.0.tar
state_dict = paddle.load(f'../assets/{DATASET_NAME}-{MODEL_NAME}/model_state.pdparams')
model.set_dict(state_dict)
```
## Prepare for Interpretations
```
from trustai.interpretation.token_level import IntGradInterpreter
import numpy as np
from assets.utils import convert_example, load_data
from paddlenlp.data import Stack, Tuple, Pad
# preprocess data functions
def preprocess_fn(data):
examples = []
data_trans = []
for key in data:
data_trans.append(data[key])
for text in data_trans:
input_ids, segment_ids = convert_example(text, tokenizer, max_seq_length=128, is_test=True)
examples.append((input_ids, segment_ids))
batchify_fn = lambda samples, fn=Tuple(
Pad(axis=0, pad_val=tokenizer.pad_token_id), # input id
Pad(axis=0, pad_val=tokenizer.pad_token_id), # segment id
): fn(samples)
input_ids, segment_ids = batchify_fn(examples)
return paddle.to_tensor(input_ids, stop_gradient=False), paddle.to_tensor(segment_ids, stop_gradient=False)
```
download data for predict and evaluate
```
# download data
!wget --no-check-certificate -c https://trustai.bj.bcebos.com/data_samples/senti_ch_predict -P ../assets/
!wget --no-check-certificate -c https://trustai.bj.bcebos.com/data_samples/senti_ch_golden -P ../assets/
# predict data for predict
data = load_data("../assets/senti_ch_predict")
print("data:\n", list(data.values())[:2])
# golden data for evluate
goldens = load_data("../assets/senti_ch_golden")
print("goldens:\n", list(goldens.values())[:2])
from trustai.interpretation.token_level.common import get_word_offset
from trustai.interpretation.token_level.data_processor import VisualizationTextRecord, visualize_text
contexts = []
batch_words = []
for example in data.values():
contexts.append("[CLS]" + " " + example['context'] + " " + "[SEP]")
batch_words.append(["[CLS]"] + example['sent_token'] + ["[SEP]"])
word_offset_maps = []
subword_offset_maps = []
for i in range(len(contexts)):
word_offset_maps.append(get_word_offset(contexts[i], batch_words[i]))
subword_offset_maps.append(tokenizer.get_offset_mapping(contexts[i]))
```
## IG Interpreter
```
ig = IntGradInterpreter(model, device="gpu")
result = ig(preprocess_fn(data), steps=100)
align_res = ig.alignment(result, contexts, batch_words, word_offset_maps, subword_offset_maps, special_tokens=["[CLS]", '[SEP]'])
from trustai.interpretation.token_level.common import general_predict_fn
def prepare_eval_data(data, results, paddle_model):
res = {}
for data_id, inter_res in zip(data, results):
eval_data = {}
eval_data['id'] = data_id
eval_data['pred_label'] = inter_res.pred_label
eval_data['pred_proba'] = inter_res.pred_proba
eval_data['rationale'] = [inter_res.rationale]
eval_data['non_rationale'] = [inter_res.non_rationale]
eval_data['rationale_tokens'] = [inter_res.rationale_tokens]
eval_data['non_rationale_tokens'] = [inter_res.non_rationale_tokens]
rationale_context = "".join(inter_res.rationale_tokens)
non_rationale_context = "".join(inter_res.non_rationale_tokens)
input_data = {'rationale': {'text': rationale_context}, 'no_rationale': {'text': non_rationale_context}}
_, pred_probas = general_predict_fn(preprocess_fn(input_data), paddle_model)
eval_data['rationale_pred_proba'] = list(pred_probas[0])
eval_data['non_rationale_pred_proba'] = list(pred_probas[1])
res[data_id] = eval_data
return res
predicts = prepare_eval_data(data, align_res, model)
print(list(predicts.values())[0])
```
evaluate for interpretation result
```
from trustai.evaluation import Evaluator
evaluator = Evaluator()
result = evaluator.cal_map(goldens, predicts)
print("map score:",result)
result = evaluator.cal_f1(goldens, predicts)
print("plausibility f1:", result)
result = evaluator.calc_iou_f1(goldens, predicts)
print("plausibility iou f1:",result)
result = evaluator.cal_suf_com(goldens, predicts)
print("sufficency score:", result[0], "conciseness score:", result[1])
```
## Attention Interpreter
```
from trustai.interpretation.token_level.common import attention_predict_fn_on_paddlenlp
from trustai.interpretation.token_level import AttentionInterpreter
att = AttentionInterpreter(model, device="gpu", predict_fn=attention_predict_fn_on_paddlenlp)
result = att(preprocess_fn(data))
align_res = att.alignment(result, contexts, batch_words, word_offset_maps, subword_offset_maps, special_tokens=["[CLS]", '[SEP]'])
predicts = prepare_eval_data(data, align_res, model)
result = evaluator.cal_map(goldens, predicts)
print("map score:",result)
result = evaluator.cal_f1(goldens, predicts)
print("plausibility f1:", result)
result = evaluator.calc_iou_f1(goldens, predicts)
print("plausibility iou f1:", result)
result = evaluator.cal_suf_com(goldens, predicts)
print("sufficency score:", result[0], "conciseness score:", result[1])
```
## LIME Interpreter
```
from trustai.interpretation.token_level import LIMEInterpreter
lime = LIMEInterpreter(model, device="gpu",
unk_id=tokenizer.convert_tokens_to_ids('[UNK]'),
pad_id=tokenizer.convert_tokens_to_ids('[PAD]'))
result = lime(preprocess_fn(data), num_samples=1000)
align_res = lime.alignment(result, contexts, batch_words, word_offset_maps, subword_offset_maps, special_tokens=["[CLS]", '[SEP]'])
predicts = prepare_eval_data(data, align_res, model)
result = evaluator.cal_map(goldens, predicts)
print("map score:",result)
result = evaluator.cal_f1(goldens, predicts)
print("plausibility f1:", result)
result = evaluator.calc_iou_f1(goldens, predicts)
print("plausibility iou f1:",result)
result = evaluator.cal_suf_com(goldens, predicts)
print("sufficency score:", result[0], "conciseness score:", result[1])
```
| github_jupyter |
# Running Code
First and foremost, the Jupyter Notebook is an interactive environment for writing and running code. The notebook is capable of running code in a wide range of languages. However, each notebook is associated with a single kernel. This notebook is associated with the IPython kernel, therefore runs Python code.
## Code cells allow you to enter and run code
Run a code cell using `Shift-Enter` or pressing the <button class='btn btn-default btn-xs'><i class="icon-step-forward fa fa-step-forward"></i></button> button in the toolbar above:
```
a = 10
print(a)
```
There are two other keyboard shortcuts for running code:
* `Alt-Enter` runs the current cell and inserts a new one below.
* `Ctrl-Enter` run the current cell and enters command mode.
Note that the `In []:` symbol next to each cell tells you about the state of the cell
* `In [ ]:` means that the cell has not been run yet.
* `In [*]:` means that the cell is currently running.
* `In [1]:` means that the cell has finished running, and was the 1st cell run.
The number tells you the order that the cells have been run in. This is important as some cells require the previous cells to have been run. This is the case above: attempting to `print` the variable `a` before it's defined will fail. You can try this by restarting the kernel and running the second cell before the first if you're interested (see below for instructions on restarting the kernel).
## Managing the Kernel
Code is run in a separate process called the Kernel. The Kernel can be interrupted or restarted. You may wish to do this if you think there's an issue with your code and you want to rerun it. Try running the following cell and then hit the <button class='btn btn-default btn-xs'><i class='icon-stop fa fa-stop'></i></button> button in the toolbar above.
```
import time
time.sleep(10)
```
If the Kernel dies you will be prompted to restart it. Here we call the low-level system libc.time routine with the wrong argument via
ctypes to segfault the Python interpreter. You'll need to uncomment the last three lines of code to see the segfault. You can do this by highlighting multiple lines and pressing `Ctrl+/` to uncomment them.
```
import sys
from ctypes import CDLL
# This will crash a Linux or Mac system
# equivalent calls can be made on Windows
# Uncomment these lines if you would like to see the segfault
# dll = 'dylib' if sys.platform == 'darwin' else 'so.6'
# libc = CDLL("libc.%s" % dll)
# libc.time(-1) # BOOM!!
```
## Cell menu
The "Cell" menu has a number of menu items for running code in different ways. These include:
* Run and Select Below
* Run and Insert Below
* Run All
* Run All Above
* Run All Below
## Restarting the kernels
The kernel maintains the state of a notebook's computations. You can reset this state by restarting the kernel. This is done by clicking on the <button class='btn btn-default btn-xs'><i class='fa fa-repeat icon-repeat'></i></button> in the toolbar above. You will be asked to confirm that you wish to restart the kernal.
## sys.stdout and sys.stderr
The stdout and stderr streams are displayed as text in the output area. Errors appear on a red background.
```
print("hi, stdout")
print('hi, stderr', file=sys.stderr)
```
## Output is asynchronous
All output is displayed as it is generated in the kernel. If you execute the next cell, you will see the output one piece at a time, not all at the end.
```
import time, sys
for i in range(8):
print(i)
time.sleep(0.5)
```
## Large outputs
To better handle large outputs, the output area can be collapsed. Run the following cell and then single- or double- click on the active area to the left of the output:
```
for i in range(50):
print(i)
```
Beyond a certain point, output will scroll automatically:
```
for i in range(500):
print(2**i - 1)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 深層畳み込み敵対的生成ネットワーク(DCGAN)
<table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/tutorials/generative/dcgan"> <img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org で表示</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/tutorials/generative/dcgan.ipynb"> <img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab で実行</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/generative/dcgan.ipynb"> <img src="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/tutorials/generative/dcgan.ipynb"> GitHub でソースを表示</a></td>
<td> <img src="https://www.tensorflow.org/images/download_logo_32px.png"><a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/generative/dcgan.ipynb">ノートブックをダウンロード</a> </td>
</table>
このチュートリアルでは、[深層畳み込み敵対的生成ネットワーク](https://arxiv.org/pdf/1511.06434.pdf) (DCGAN) を使用して手書きの数字の画像を生成する方法を実演します。このコードは、`tf.GradientTape` トレーニングループを伴う [Keras Sequential API](https://www.tensorflow.org/guide/keras) を使用して記述されています。
## GAN とは?
[敵対的生成ネットワーク](https://arxiv.org/abs/1406.2661) (GAN) は現在コンピュータサイエンス分野で最も興味深い構想です。2 つのモデルが敵対的なプロセスにより同時にトレーニングされます。*ジェネレータ*(「芸術家」)が本物のような画像の制作を学習する一方で、*ディスクリミネータ*(「芸術評論家」)は本物の画像を偽物と見分けることを学習します。

トレーニング中、*ジェネレータ*では、本物に見える画像の作成が徐々に上達し、*ディスクリミネータ*では、本物と偽物の区別が上達します。このプロセスは、*ディスクリミネータ*が本物と偽物の画像を区別できなくなった時点で平衡に達します。

このノートブックでは、このプロセスを MNIST データセットで実演しています。以下のアニメーションは、50 エポックでトレーニングする過程で*ジェネレータ*が生成した一連の画像を示しています。画像は、ランダムノイズとして始まり、徐々に手書きの数字へと似ていきます。

GAN についてさらに学習するには、MIT の「[Intro to Deep Learning](http://introtodeeplearning.com/)」コースをご覧ください。
### TensorFlow とその他のライブラリをインポートする
```
import tensorflow as tf
tf.__version__
# To generate GIFs
!pip install imageio
import glob
import imageio
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
from tensorflow.keras import layers
import time
from IPython import display
```
### データセットを読み込んで準備する
ジェネレータとディスクリミネータのトレーニングには、MNIST データセットを使用します。ジェネレータは、MNIST データに似た手書きの数字を生成するようになります。
```
(train_images, train_labels), (_, _) = tf.keras.datasets.mnist.load_data()
train_images = train_images.reshape(train_images.shape[0], 28, 28, 1).astype('float32')
train_images = (train_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
BUFFER_SIZE = 60000
BATCH_SIZE = 256
# Batch and shuffle the data
train_dataset = tf.data.Dataset.from_tensor_slices(train_images).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
```
## モデルを作成する
ジェネレータとディスクリミネータの定義には、[Keras Sequential API](https://www.tensorflow.org/guide/keras#sequential_model) を使用します。
### ジェネレータ
ジェネレータは、`tf.keras.layers.Conv2DTranspose` (アップサンプリング) レイヤーを使用して、シード (ランダムノイズ) から画像を生成します。このシードを入力として取る `Dense` レイヤーから始め、期待する画像サイズ (28x28x1) に到達するまで何度もアップサンプリングします。tanh を使用する出力レイヤーを除き、各レイヤーに `tf.keras.layers.LeakyReLU` アクティベーションが使用されています。
```
def make_generator_model():
model = tf.keras.Sequential()
model.add(layers.Dense(7*7*256, use_bias=False, input_shape=(100,)))
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Reshape((7, 7, 256)))
assert model.output_shape == (None, 7, 7, 256) # Note: None is the batch size
model.add(layers.Conv2DTranspose(128, (5, 5), strides=(1, 1), padding='same', use_bias=False))
assert model.output_shape == (None, 7, 7, 128)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(64, (5, 5), strides=(2, 2), padding='same', use_bias=False))
assert model.output_shape == (None, 14, 14, 64)
model.add(layers.BatchNormalization())
model.add(layers.LeakyReLU())
model.add(layers.Conv2DTranspose(1, (5, 5), strides=(2, 2), padding='same', use_bias=False, activation='tanh'))
assert model.output_shape == (None, 28, 28, 1)
return model
```
(まだトレーニングされていない)ジェネレータを使用して画像を作成します。
```
generator = make_generator_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
plt.imshow(generated_image[0, :, :, 0], cmap='gray')
```
### ディスクリミネータ
ディスクリミネータは CNN ベースの画像分類子です。
```
def make_discriminator_model():
model = tf.keras.Sequential()
model.add(layers.Conv2D(64, (5, 5), strides=(2, 2), padding='same',
input_shape=[28, 28, 1]))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Conv2D(128, (5, 5), strides=(2, 2), padding='same'))
model.add(layers.LeakyReLU())
model.add(layers.Dropout(0.3))
model.add(layers.Flatten())
model.add(layers.Dense(1))
return model
```
(まだトレーニングされていない)ディスクリミネータを使用して、生成された画像を本物と偽物に分類します。モデルは、本物の画像に対して正の値を出力し、偽物の画像には負の値を出力するようにトレーニングされます。
```
discriminator = make_discriminator_model()
decision = discriminator(generated_image)
print (decision)
```
## 損失とオプティマイザを定義する
両方のモデルに損失関数とオプティマイザを定義します。
```
# This method returns a helper function to compute cross entropy loss
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
```
### ディスクリミネータの損失
このメソッドは、ディスクリミネータが本物と偽物の画像をどれくらいうまく区別できるかを数値化します。本物の画像に対するディスクリミネータの予測を 1 の配列に比較し、(生成された)偽物の画像に対するディスクリミネータの予測を 0 の配列に比較します。
```
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
total_loss = real_loss + fake_loss
return total_loss
```
### ジェネレータの損失
ジェネレータの損失は、ディスクリミネータをどれくらいうまく騙せたかを数値化します。直感的に、ジェネレータがうまく機能しているのであれば、ディスクリミネータはその偽物の画像を本物(または 1)として分類します。ここでは、生成された画像に対するディスクリミネータの判定を 1 の配列に比較します。
```
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
```
2 つのネットワークを個別にトレーニングするため、ディスクリミネータオプティマイザとジェネレータオプティマイザは異なります。
```
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
```
### チェックポイントを保存する
このノートブックでは、モデルの保存と復元方法も実演します。これは長時間実行するトレーニングタスクが中断された場合に役立ちます。
```
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
```
## トレーニングループを定義する
```
EPOCHS = 50
noise_dim = 100
num_examples_to_generate = 16
# We will reuse this seed overtime (so it's easier)
# to visualize progress in the animated GIF)
seed = tf.random.normal([num_examples_to_generate, noise_dim])
```
トレーニングループは、ランダムシードを入力として受け取っているジェネレータから始まります。そのシードを使って画像が生成されると、ディスクリミネータを使って本物の画像(トレーニングセットから取り出された画像)と偽物の画像(ジェネレータが生成した画像)が分類されます。これらの各モデルに対して損失が計算されると、勾配を使用してジェネレータとディスクリミネータが更新されます。
```
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
# Produce images for the GIF as we go
display.clear_output(wait=True)
generate_and_save_images(generator,
epoch + 1,
seed)
# Save the model every 15 epochs
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# Generate after the final epoch
display.clear_output(wait=True)
generate_and_save_images(generator,
epochs,
seed)
```
**画像を生成して保存する**
```
def generate_and_save_images(model, epoch, test_input):
# Notice `training` is set to False.
# This is so all layers run in inference mode (batchnorm).
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(4,4))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
```
## モデルをトレーニングする
上記で定義した `train()` メソッドを呼び出し、ジェネレータとディスクリミネータを同時にトレーニングします。GAN のトレーニングには注意が必要です。ジェネレータとディスクリミネータが互いを押さえつけることのないようにすることが重要です (同じようなレートでトレーニングするなど)。
トレーニング開始時に生成された画像はランダムノイズのように見えます。トレーニングが進行するにつれ、生成された数字は徐々に本物に見えるようになります。約 50 エポック後には、これらは MNIST 数字に似たものになります。Colab でのこの過程には、デフォルトの設定でエポック当たり約 1 分がかかります。
```
train(train_dataset, EPOCHS)
```
最新のチェックポイントを復元します。
```
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
```
## GIF を作成する
```
# Display a single image using the epoch number
def display_image(epoch_no):
return PIL.Image.open('image_at_epoch_{:04d}.png'.format(epoch_no))
display_image(EPOCHS)
```
トレーニング中に保存した画像を使用して、アニメーション GIF を作成するには、`imageio` を使用します。
```
anim_file = 'dcgan.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('image*.png')
filenames = sorted(filenames)
last = -1
for i,filename in enumerate(filenames):
frame = 2*(i**0.5)
if round(frame) > round(last):
last = frame
else:
continue
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
import IPython
if IPython.version_info > (6,2,0,''):
display.Image(filename=anim_file)
import tensorflow_docs.vis.embed as embed
embed.embed_file(anim_file)
```
## 次のステップ
このチュートリアルでは、GAN を記述してトレーニングするために必要となる完全なコードを紹介しました。次のステップでは、[Kaggle で提供されている](https://www.kaggle.com/jessicali9530/celeba-dataset) Large-scale Celeb Faces Attributes (CelebA) データセットなどの別のデータセットを使って実験してみるとよいでしょう。GAN についてさらに学習する場合は、[NIPS 2016 Tutorial: Generative Adversarial Networks](https://arxiv.org/abs/1701.00160) をお勧めします。
| github_jupyter |
## Based on data generated on Apr 9, 2019
# General setup
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
%matplotlib inline
pd.options.display.max_columns = 150
involved_hebrew = pd.read_csv('involved_hebrew.csv')
```
# General info
```
involved_hebrew.info()
involved_hebrew.describe()
involved_hebrew.head()
print('Number unique values')
involved_hebrew.nunique(axis=0)
```
# Provider_and_id, unique number of accidents, number of people involved in accident
Although there are 1,678,965 involved people, every involved person appears as a separate line, there are actually 710,423 accidents
```
# Observations grouped by provider_and_id - actual accidents and number of observations per provider_and_id
print('Number of involved per accident - head of accidents')
grouped_by_provider_id = involved_hebrew.groupby('provider_and_id')
grouped_by_provider_id['provider_and_id'].value_counts().head(25)
# Number of times same provider_and_id appears and how many times it appears
print('Number of accidents with each number of involved per accident')
same_provider_id_appears = grouped_by_provider_id['provider_and_id'].value_counts().value_counts()
same_provider_id_appears
# Just the top 10 of most occuring number of people involved in the accident
print('Number of accidents with each number of involved per accident - top 10 most frequent number of involved')
same_provider_id_appears[:10]
fig = plt.figure(figsize=(25,5))
ax = sns.boxplot(grouped_by_provider_id.count()['accident_id'])
ax.set_xlabel('Number of involved, most 2-3, 1 and 4 also common, sometimes up to ~80 involved')
num_accidents = sum(same_provider_id_appears)
num_involved = len(involved_hebrew)
print(f'Number of accidents in the table: {num_accidents}, number involved in these accidents: {num_involved}')
percents_num_involved = pd.DataFrame({'Num people involved': same_provider_id_appears.index, 'Num accidents': same_provider_id_appears})
percents_num_involved['% accidents'] = percents_num_involved['Num accidents'] / num_accidents * 100
percents_num_involved = percents_num_involved.sort_index()
percents_num_involved['% accidents accumulated'] = percents_num_involved['% accidents'].cumsum()
print('Percents of people involved in an accident')
percents_num_involved.head(10)
ax = sns.scatterplot(x='Num people involved', y='% accidents', color='red', data=percents_num_involved.head(6))
ax.set_title('Percent of accidents per number of people involved in an accident')
ax = sns.lineplot(x='Num people involved', y='% accidents accumulated', data=percents_num_involved.head(6))
ax.set_title('Accumulative percentage of incidents per number of people involved in an accident')
```
Small insight: 72% have up to 2 people involved, 90% have 3 or less, 95% have 4 or less, 99% have 6 or less
```
print('Example of 6 people involved')
involved_hebrew[involved_hebrew['provider_and_id'] == 12008000037]
```
# provider_code
```
provider_code_counts = involved_hebrew['provider_code'].value_counts()
print('Counts of different values of provider_code')
provider_code_counts
print(f'Small: About {(100 * provider_code_counts[1]/(provider_code_counts[1] + provider_code_counts[3])).round(0)} % of accidents get investigated by the police.')
```
# involved_type
```
print('Counts of different values of involved_type:')
involved_hebrew['involved_type'].value_counts()
print('Counts of different values of involved_type_hebrew:')
involved_hebrew['involved_type_hebrew'].value_counts()
```
This is a table of all involved, an involved can be a passanger/pedastrian that was injured. But he can also be a driver. There are 2 types of drivers - injured and not injured
```
driver_or_injured = involved_hebrew.copy()
driver_or_injured['driver'] = driver_or_injured['involved_type'].apply(lambda involved_type: 1 if involved_type in [1,2] else 0)
driver_or_injured['injured'] = driver_or_injured['involved_type'].apply(lambda involved_type: 1 if involved_type in [2,3] else 0)
print(f"Drivers: {driver_or_injured['driver'].sum()}, injured: {driver_or_injured['injured'].sum()}")
print(f"Small: Drivers are around {(100 * driver_or_injured['driver'].value_counts()[1] / len(involved_hebrew)).round()}% of involved in accidents")
print('Number of injured and not injured:')
driver_or_injured['injured'].value_counts()
print(f"Small: Injured are around {(100 * driver_or_injured['injured'].value_counts()[1] / len(involved_hebrew)).round()}% of involved in accidents where there are injuries")
fig = plt.figure(figsize=(20,3))
ax = sns.countplot(driver_or_injured.groupby(by='provider_and_id')['injured'].sum())
ax.set_title('Number of injured per accident')
print('Numbers of accidents per number of injured:')
driver_or_injured.groupby(by='provider_and_id')['injured'].sum().value_counts()
injured_nums = pd.DataFrame({'Num accidents': driver_or_injured.groupby(by='provider_and_id')['injured'].sum().value_counts().sort_index()})
injured_nums.index.names = ['Num injured']
injured_nums['% accidents'] = injured_nums['Num accidents'] / injured_nums['Num accidents'].sum() * 100
injured_nums['acc % accidents'] = injured_nums['% accidents'].cumsum()
print('Number of accidents with given number of injured ')
injured_nums
tmp = driver_or_injured.groupby(by='provider_and_id')[['injured','provider_code']].max()
tmp['provider_code'] = tmp['provider_code'].map({1:1, 3:0})
print("Seems that there is no accident without any injured (we only have data with injuries), so can't do correlation between whether police opened investigation for injured")
#tmp.corr()
```
# license_acquiring_date
```
involved_hebrew['license_acquiring_date'].value_counts().sort_index()
```
TODO: Years 1911, 3055, 4444 don't make sense and need to be cleaned out
Need to understand if all the values of 752295 without license_acquiring_date are because they were not drivers
Seems that small part of Injured drivers don't have a license date, but a very large part (>50%) of the Drivers don't have!!!
```
num_years_before_accident = involved_hebrew.copy()
num_years_before_accident = involved_hebrew[((involved_hebrew['involved_type'] == 1) | (involved_hebrew['involved_type'] == 2))
& (involved_hebrew['license_acquiring_date'] != 0)
& (involved_hebrew['license_acquiring_date'] >= 1948)
& (involved_hebrew['license_acquiring_date'] <= 2018)
& (involved_hebrew['license_acquiring_date'] <= involved_hebrew['accident_year'])]
num_years_before_accident['years_on_road'] = num_years_before_accident.apply(lambda driver: driver['accident_year'] - driver['license_acquiring_date'], axis=1)
print(f'Number of accidents: {len(num_years_before_accident)}')
plt.figure(figsize=(20,5)).tight_layout()
ax = sns.countplot(num_years_before_accident['years_on_road'])
ax.set_ylabel('Number of accidents')
ax.set_title('Number of accidents a driver is involved in based on number of years on the road')
ax = sns.distplot(num_years_before_accident['years_on_road'], bins=10)
ax.set_title('KDE of years on the road vs chance of being involved in an accident')
print(f"Number of accidents in first 10 years of driving\n{num_years_before_accident['years_on_road'].value_counts().sort_index().head(10)}")
per_num_years_before_accident = pd.DataFrame({'Years on road': num_years_before_accident['years_on_road'].value_counts().sort_index().index,
'Num accidents': num_years_before_accident['years_on_road'].value_counts().sort_index()})
per_num_years_before_accident['% accidents'] = (per_num_years_before_accident['Num accidents'] /
per_num_years_before_accident['Num accidents'].sum() * 100)
per_num_years_before_accident['% accidents accumulated'] = per_num_years_before_accident['% accidents'].cumsum()
print('Number of accidents with numerous years with license of the driver:')
per_num_years_before_accident.head(12)
```
Small: 9% of accidents happen with drivers in their first 2 years after getting a license, 20% in their first 5 years, and a third of accidents happen with drivers in their first 9 years
How come many values are missing for 'license_acquiring_date'?
```
license_year_exists = involved_hebrew.copy()
license_year_exists = license_year_exists[(license_year_exists['involved_type'] == 1) | (license_year_exists['involved_type'] == 2)]
license_year_exists['License year given'] = (license_year_exists['license_acquiring_date'] != 0).astype(int)
ax = sns.countplot(x='accident_year', data=license_year_exists, hue='License year given')
ax.set_title('Number of drivers with given license year per year of accident')
```
Seems not dependent on year - around 1/3 of accidents, there is no license date for the driver
```
ax = sns.countplot(x='License year given', hue='provider_code', data=license_year_exists)
ax.set_title('Number of drivers with a given license year based on whether a police report was open (provider_code 1)')
ax = sns.countplot(hue='License year given', x='provider_code', data=license_year_exists)
ax.set_title('Number of drivers with a given license year based on whether a police report was open (provider_code 1)')
```
Conclusion: when the accident doesn't get investigated, there is usually no information when the license was issued
# age_group
```
print('Number of involved in every age_group:')
involved_hebrew['age_group'].value_counts()
print('Number of involved in every age_group_hebrew:')
involved_hebrew['age_group_hebrew'].value_counts()
print(f"Percentage doesn't have age group: {involved_hebrew['age_group_hebrew'].value_counts()['לא ידוע'] / len(involved_hebrew) * 100}")
print(f"Number of missing age group: total: {involved_hebrew['age_group_hebrew'].value_counts()['לא ידוע']}, investigated by police: {involved_hebrew[involved_hebrew['provider_code'] == 1]['age_group_hebrew'].value_counts()['לא ידוע']}, not investigated by police: {involved_hebrew[involved_hebrew['provider_code'] == 3]['age_group_hebrew'].value_counts()['לא ידוע']}")
print('Something strange happens with age_group 2 and 3:')
involved_hebrew['age_group'].value_counts()[12:14]
print('age_group 2 and 3 is split up into 4 groups in age_group_hebrew:')
involved_hebrew['age_group_hebrew'].value_counts()[['05-09', '10-14', '05-ספטמבר', 'אוקטובר-14']]
print('Adding pairs of age_group_hebrew categories adds up to age_group 2 and 3:')
counts = involved_hebrew['age_group_hebrew'].value_counts()
print(counts['05-09'] + counts['05-ספטמבר'])
print(counts['10-14'] + counts['אוקטובר-14'])
print('Years where age_group_hebrew categories are screwed up for 05-..:')
involved_hebrew[involved_hebrew['age_group_hebrew'] == '05-ספטמבר'].groupby(by='accident_year')['provider_and_id'].count()
print('Years where age_group_hebrew categories are screwed up for 14-..:')
involved_hebrew[involved_hebrew['age_group_hebrew'] == 'אוקטובר-14'].groupby(by='accident_year')['provider_and_id'].count()
```
There is indeed a problem in the field *age_group_hebrew* that 2 values of *age_group* are split into 2 values each of *age_group_hebrew*
Fixing in the original table:
```
fixed = driver_or_injured.copy() # to have also info of whether it's a driver or injured
fixed['age_group_hebrew'].replace({'05-ספטמבר': '05-09', 'אוקטובר-14':'10-14'}, inplace=True)
print('Involved age groups after fix:')
fixed['age_group_hebrew'].value_counts().sort_index()
print('Ages of involved drivers')
fixed[fixed['driver'] == 1]['age_group_hebrew'].value_counts().sort_index()
```
Seems there are some really minor drivers ages 0-4 etc.
```
known_age_groups = fixed[fixed['age_group_hebrew'] != 'לא ידוע'].sort_values(by='age_group_hebrew')
plt.figure(figsize=(20,5))
ax = sns.countplot(x='age_group_hebrew',data=known_age_groups)
ax.set_title('Number of people involved in accidents based on age group')
plt.figure(figsize=(20,5))
ax = sns.countplot(x='age_group_hebrew',data=known_age_groups[known_age_groups['driver'] == 1])
ax.set_title('Number of drivers involved in accidents based on age group')
plt.figure(figsize=(20,5))
ax = sns.countplot(x='age_group_hebrew',data=known_age_groups[known_age_groups['injured'] == 1])
ax.set_title('Number of injured in accidents based on age group')
age_acc = pd.DataFrame({'Num injured': known_age_groups[known_age_groups['injured'] == 1]['age_group_hebrew'].value_counts().sort_index()})
age_acc['% injured'] = age_acc['Num injured'] / age_acc['Num injured'].sum() * 100
age_acc['Acc % injured'] = age_acc['% injured'].cumsum()
print('Number of injured based on age groups:')
age_acc
```
Small: Almost 10% of injured are 14 and younger, 7% are younger than 19
# Sex
```
print('Split of involved into Male and Female:')
fixed['sex_hebrew'].value_counts()
ax = sns.countplot(x='sex_hebrew', data=fixed)
ax.set_title('Males and females involved in accidents')
ax = sns.countplot(hue='sex_hebrew', x='driver', data=fixed)
ax.set_title('Split involved into drivers/not drivers involved in accidents per male/female')
ax = sns.countplot(hue='sex_hebrew', x='injured', data=fixed)
ax.set_title('Split involved into drivers/not drivers involved in accidents per male/female')
```
Small:
- there are many more men involved than women (almost twice as much)
- most majority of men that are involved are driving, women are slightly more drivers than non-drivers - perhaps many of men are drivers that are not injured
- Larger part of women are injured than men - perhaps because there are many more men drivers (will be involved even if not injured), women are less drivers, and will appear in the statistics more if injured
# vehicle_type
```
print('Different values of vehicle_type_hebrew with number of involved in this type of vehicle:')
fixed['vehicle_type_hebrew'].value_counts()
print('Different values of vehicle_type_hebrew with number of involved in this type of vehicle (ordered alphabetically):')
fixed['vehicle_type_hebrew'].value_counts().sort_index()
```
TODO: do some more analysis about this category
# safety_measures_use
```
print('Different values of safety_measures_use_hebrew with number of involved:')
fixed['safety_measures_use_hebrew'].value_counts(dropna=False)
print('Different values of safety_measures_use with number of involved:')
fixed['safety_measures_use'].value_counts(dropna=False)
```
Mostly not usefull since in vast majority empty (~98%)
```
fixed.groupby(by='accident_year')['safety_measures_use'].value_counts(dropna=False)
```
Don't have this information more than before also in 2018
# involve_yishuv_symbol and involve_yishuv_name
```
print('Number of involved per involve_yishuv_symbol:')
fixed['involve_yishuv_symbol'].value_counts(dropna=False).head()
print('Numer of different values of involve_yishuv_symbol:')
fixed['involve_yishuv_symbol'].value_counts(dropna=False).shape[0]
print('Numer of different values of involve_yishuv_name:')
fixed['involve_yishuv_name'].value_counts(dropna=False).shape[0]
```
We can see that involve_yishuv_name and involve_yishuv_symbol don't match - even looking at the number of unique values
```
print('Most frequent Yishuv names (involve_yishuv_name):')
fixed['involve_yishuv_name'].value_counts(dropna=False).head(20)
print('Number of involved without Yishuv name:')
fixed['involve_yishuv_name'].isnull().sum()
print(f"Out of all involved, there are {fixed['involve_yishuv_symbol'].value_counts(dropna=False).shape[0]} involve_yishuv_symbol unique values, out of which NaN: {fixed['involve_yishuv_symbol'].isnull().sum()}")
print(f"Out of all involved, there are {fixed['involve_yishuv_name'].value_counts(dropna=False).shape[0]} involve_yishuv_name unique values, out of which NaN: {fixed['involve_yishuv_name'].isnull().sum()}")
```
# injury_severity
```
print('Different number of values for injury_severity:')
fixed['injury_severity'].value_counts(dropna=False)
print('Different number of values for injury_severity_hebrew:')
fixed['injury_severity_hebrew'].value_counts(dropna=False)
print('Different number of values for injury_severity_hebrew out of actually injured:')
fixed[fixed['injured'] == 1]['injury_severity_hebrew'].value_counts(dropna=False)
print('Different number of values for injury_severity_hebrew out of NOT injured:')
fixed[fixed['injured'] == 0]['injury_severity_hebrew'].value_counts(dropna=False)
```
This is great, since it shows that every time someone is classified as "injured", it has severity, and when the person is not injured, there is no injury_severity
```
ax = sns.countplot(fixed[fixed['injured'] == 1]['injury_severity_hebrew'])
ax.set_title('Number of injured per injury_serverity_hebrew')
num_acc_with_killed = fixed[fixed['injury_severity'] == 1].groupby(by='provider_and_id')['provider_and_id'].count().shape[0]
print(f"% of accidents with a killed person: {round(num_acc_with_killed * 100 / num_accidents,1)}%")
num_injured = fixed[fixed['injured'] == 1].shape[0]
num_killed = fixed[fixed['injury_severity'] == 1].shape[0]
print(f"% of killed from all injured: {round(num_killed * 100 / num_injured, 2)}%")
ax = sns.countplot(data=fixed[fixed['injured'] == 1], x='accident_year')
ax.set_title('Number of injured per year')
ax = sns.countplot(data=fixed[fixed['injury_severity'] == 1], x='accident_year')
ax.set_title('Number of killed per year')
ax = sns.countplot(data=fixed[fixed['injury_severity'] == 2], x='accident_year')
ax.set_title('Number of injured severly per year')
ax = sns.countplot(data=fixed[fixed['injury_severity'] == 3], x='accident_year')
ax.set_title('Number of lightly injured per year')
```
- 80,000 get injured every year in traffic accidents, but there is a trend going down, it was 120,000 10 years ago!!!
- 2000 get seriously injured per year, in the last 10 years, first there was a drop, but since 2011 it again climbs up
- More than 300 per year get killed, with a big drop in 2012 and climbing since with a drop again in 2017/2018
- Number of lightly injured - seems that got dropped seriously in last 10 years, but maybe it’s more how injured are being counted
# injured_type
```
print('Number of injured per injured_type:')
fixed[fixed['injured'] == 1]['injured_type'].value_counts(dropna=False)
print('Number of injured per injured_type_hebrew:')
fixed[fixed['injured'] == 1]['injured_type_hebrew'].value_counts(dropna=False)
injured_type = pd.DataFrame({'Num injured': fixed[fixed['injured'] == 1]['injured_type_hebrew'].value_counts(dropna=False)})
injured_type['% injured'] = injured_type['Num injured'] / injured_type['Num injured'].sum() * 100
print('Number of injured per injured_type_hebrew and % from all injured:')
injured_type
fig = plt.figure(figsize=(25,5))
ax = sns.countplot(x='injured_type_hebrew', data=fixed[fixed['injury_severity']==1])
ax.set_title('Number of injured per injured_type_hebrew')
injured_type_killed = pd.DataFrame({'Num killed': fixed[fixed['injury_severity']==1]['injured_type_hebrew'].value_counts()})
injured_type_killed['% injured'] = injured_type_killed['Num killed'] / injured_type_killed['Num killed'].sum() * 100
print('Number of killed per injured_type_hebrew:')
injured_type_killed
injured_type_serious = pd.DataFrame({'Num seriously injured': fixed[fixed['injury_severity']==2]['injured_type_hebrew'].value_counts()})
injured_type_serious['% injured'] = injured_type_serious['Num seriously injured'] / injured_type_serious['Num seriously injured'].sum() * 100
print('Number of seriously injured per injured_type_hebrew:')
injured_type_serious
```
# injured_position
```
print('Number of injured per injured_position:')
fixed[fixed['injured'] == 1]['injured_position'].value_counts(dropna=False)
print('Number of injured per injured_position_hebrew:')
fixed[fixed['injured'] == 1]['injured_position_hebrew'].value_counts(dropna=False)
```
# population_type
```
print('Number of involved per population_type:')
fixed['population_type'].value_counts(dropna=False)
print('Number of involved per population_type_hebrew:')
fixed['population_type_hebrew'].value_counts(dropna=False)
```
# home_region
```
print('Number of involved per home_region:')
fixed['home_region'].value_counts(dropna=False)
print('Number of involved per home_region_hebrew:')
fixed['home_region_hebrew'].value_counts(dropna=False)
```
# home_district
```
print('Number of involved per home_district:')
fixed['home_district'].value_counts(dropna=False)
print('Number of involved per home_district_hebrew:')
fixed['home_district_hebrew'].value_counts(dropna=False)
```
# home_natural_area
```
print('Number of different values for home_natural_area:')
fixed['home_natural_area'].value_counts(dropna=False).shape[0]
print('Number of different values for home_natural_area_hebrew:')
fixed['home_natural_area_hebrew'].value_counts(dropna=False).shape[0]
print('Different numbers of involved per home_natural_area:')
fixed['home_natural_area'].value_counts(dropna=False)
print('Different numbers of involved per home_natural_area_hebrew:')
fixed['home_natural_area_hebrew'].value_counts(dropna=False)
fixed['home_natural_area_hebrew'].value_counts(dropna=False).values - fixed['home_natural_area'].value_counts(dropna=False).values
```
The values don't completely match between home_natural_area_hebrew and home_natural_area!!!
# home_municipal_status
```
print('Different numbers of involved per home_municipal_status:')
fixed['home_municipal_status'].value_counts(dropna=False).shape[0]
print('Different numbers of involved per home_municipal_status_hebrew:')
fixed['home_municipal_status_hebrew'].value_counts(dropna=False).shape[0]
print('Most frequent home_municipal_status values with number of involved:')
fixed['home_municipal_status'].value_counts(dropna=False).head()
print('Most frequent home_municipal_status_hebrew values with number of involved:')
fixed['home_municipal_status_hebrew'].value_counts(dropna=False)
```
Home_municipal_status - not useful - missing in majority of the fields, and not matching home_municipal_status_hebrew
# home_residence_type
```
print('Different unique values of home_residence_type:')
fixed['home_residence_type'].value_counts(dropna=False).shape[0]
print('Different unique values of home_residence_type_hebrew:')
fixed['home_residence_type_hebrew'].value_counts(dropna=False).shape[0]
```
We see that *home_residence_type* and *home_residence_type_hebrew* don't match!
```
print('Most frequent values of home_residence_type with number of involved')
fixed['home_residence_type'].value_counts(dropna=False).head()
print('Number of involved with police investigation with null value for home_residence_type:')
fixed[fixed['provider_code'] == 1]['home_residence_type'].isnull().sum()
print('Number of involved without police investigation with null value for home_residence_type:')
fixed[fixed['provider_code'] == 3]['home_residence_type'].isnull().sum()
print('Different values of home_residence_type_hebrew with number of involved:')
fixed['home_residence_type_hebrew'].value_counts(dropna=False)
```
# hospital_time
```
print('Different values of hospital_time with number of involved:')
fixed['hospital_time'].value_counts(dropna=False)
print('Different values of hospital_time_hebrew with number of involved:')
fixed['hospital_time_hebrew'].value_counts(dropna=False)
```
Not very useful since doesn't appear in vast majority of cases
# medical_type
```
print('Different values of medical_type with number of involved:')
fixed['medical_type'].value_counts(dropna=False)
print('Different values of medical_type_hebrew with number of involved:')
fixed['medical_type_hebrew'].value_counts(dropna=False)
```
Not extemely useful since missing in vast majority of cases
# release_dest
```
print('Different values of release_dest with number of involved:')
fixed['release_dest'].value_counts(dropna=False)
print('Different values of release_dest_hebrew with number of involved:')
fixed['release_dest_hebrew'].value_counts(dropna=False)
```
Possibly not useful since missing a lot of values
# safety_measures_use
```
print('Different values of safety_measures_use with number of involved:')
fixed['safety_measures_use'].value_counts(dropna=False)
print('Different values of safety_measures_use_hebrew with number of involved:')
fixed['safety_measures_use_hebrew'].value_counts(dropna=False)
```
Not very useful since missing in vast majority of cases
# late_deceased
```
print('Number of involved with different late_deceased values:')
fixed['late_deceased'].value_counts(dropna=False)
print('Number of involved with different late_deceased_hebrew values:')
fixed['late_deceased_hebrew'].value_counts(dropna=False)
print('Number of involved with different injury_severity values:')
fixed['injury_severity'].value_counts(dropna=False)
```
Seems to be same number 3826 in both lists, but it seems in injury_severity doesn't take into account late death
# car_id, involve_id
```
print('Number of involved per different car_id values:')
fixed['car_id'].value_counts(dropna=False)
print('Number of involved per different involve_id values:')
fixed['involve_id'].value_counts(dropna=False)
```
Seems that were both are deleted (we don't get the original values) and values are meaningless ?
# accident_year, accident_month
```
print('Number of involved in accidents per year')
fixed['accident_year'].value_counts(dropna=False)
print('Number of involved in accidents per month')
fixed['accident_month'].value_counts(dropna=False).sort_index()
ax = sns.countplot(fixed['accident_month'])
ax.set_title('Number of involved in accidents per month')
ax = sns.countplot(data=fixed[fixed['injury_severity']==1], x='accident_month')
ax.set_title('Number of killed in accidents per month')
```
# Connection between police investigation and injury severity
```
print('Number killed per provider_code:')
fixed[fixed['injury_severity'] == 1].groupby(['provider_code'])['provider_code'].count()
print('Number severely injured per provider_code:')
fixed[fixed['injury_severity'] == 2].groupby(['provider_code'])['provider_code'].count()
print('Number lightly injured per provider_code:')
fixed[fixed['injury_severity'] == 3].groupby(['provider_code'])['provider_code'].count()
```
Seems that every time there is a serious injury, police investigates
# Numbers and basic conclusions
- 1,678,965 involved
- 710,423 accidents
- Number of involved per accident:
- most 2-3, 1 and 4 also common, sometimes up to ~80 involved
- 60% have 2 people involved, 90% have 3 or less, 95% have 4 or less, 99% have 6 or less
- About 25.0 % of accidents get investigated by the police
- Drivers are round 75.0% of involved in accidents, Injured are round 62.0% of involved in accidents
- Number of injured - 73% - 1 injured, 16% - 2 injured, 6% - 3 injured, 90% 1 or 2, 95% 2 or less, 98% 3 or less, 99% 5 or less
- Seems that every accident has an injured, we don’t get other data
- 9% of accidents happen with drivers in their first 2 years after getting a license, 20% in their first 5 years, and a third of accidents happen with drivers in their first 9 years
- Age group
- 21% don’t have age group info, vast majority when not investigated by police Number of missing age group: total: 356515, investigated by police: 16366, not investigated by police: 340149
- Almost 10% of injured are 14 and younger, 7% are younger than 19
- Sex
- there are many more men involved than women (almost twice as much)
- most majority of men that are involved are driving, women are slightly more drivers than non-drivers - perhaps many of them are drivers that are not injured
- Larger part of women are injured than men - perhaps because there are many more men drivers (will be involved even if not injured), women are less drivers, and will appear in the statistics more if injured
- Safety information appears only in 2.7% of cases - not useful
- Yishuv
- Out of all involved, there are 1557 involve_yishuv_symbol unique values, out of which NaN: 368345
- Out of all involved, there are 1265 involve_yishuv_name unique values, out of which NaN: 370302
- When someone is injured, there is always severity, so can count on this value
- Number of injured/killed:
- 80,000 get injured every year in traffic accidents, but there is a trend going down, it was 120,000 10 years ago!!!
- 2000 get seriously injured per year, in the last 10 years, first there was a drop, but since 2011 it again climbs up
- More than 300 per year get killed, with a big drop in 2012 and climbing since with a drop again in 2017/2018
- Number of lightly injured - seems that got dropped seriously in last 10 years, but maybe it’s more how injured are being counted
- Out of people killed
- 32% are pedenstrians (9% of injured)
- 12% are motorcycle riders (8% of injured)
- 4% are bicycle riders (1.5% of injured)
- **Total:** almost 40% are not in cars
- Out of people seriously injured
- 31% are pedenstrians (9% of injured)
- 19% are motorcycle riders (8% of injured)
- 4% are bicycle riders (1.5% of injured)
- **Total:** almost 45% are not in cars
- Seems that a month is not a good predictor of people being injured, seems that summer months are slightly worse, also when looking at number of killed per month
- Police investigated all accidents with killed or severity injured, but only ~25% of accidents with light injuries
# Conclusions about usefulness of different fields
- To be removed
- Safety_measures - missing in vast majority (98%)
- Home_municipal_status - missing in majority of the fields, and not matching home_municipal_status_hebrew
- Home_residence_type - split into a lot of values, missing when no police report, home_residence_type and home_residence_type_hebrew don’t match ?
- Hospital_time - NaN in vast majority of cases ?
- Medical_type - NaN in vast majority of cases ?
- Release_dest - NaN in vast majority of cases ?
- Safety_measures_use - missing in vast majority of cases
- To be careful with
- license_acquiring_date - missing in ½ the cases - mostly not investigated
- age_group - missing 21% of the time, most when not investigated
- involve_yishuv_symbol / involve_yishuv_name - seem not to fully match (see numbers)
- population_type - a little NaNs
- Home_region - a lot of NaNs - mostly when no police report
- Home_district - a lot of NaNs - mostly when no police report
- home_natural_area and home_natural_area_hebrew don’t fully match
Late_deceased - has some more info than injury_severity=1
- To be cleaned / fixed
- license_acquiring_date - Years 1911, 3055, 4444 don't make sense and need to be cleaned out
- Age_group_hebrew - 2 values of age_group are split into 2 values each of age_group_hebrew
| github_jupyter |
# Modeling Source-to-Sink systems using FastScape: 9. Cyclic variations in climate/precipitation

```
import xsimlab as xs
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
#plt.style.use('dark_background')
%load_ext xsimlab.ipython
import hvplot.xarray
```
In this experiment, we will focus on climatic/precipitation variations that only affect the basin, not the source. In this way we can appreciate the effect of increased/decreased precipitation on the transport capacity of the depositional/transit system.
```
from fastscape.models import marine_model
from fastscape.processes import (BlockUplift)
transit_model = (marine_model.
drop_processes('diffusion').
drop_processes('init_topography').
drop_processes('uplift').
drop_processes('marine').
drop_processes('sea').
update_processes({'uplift': BlockUplift}))
transit_model.visualize(show_inputs=True)
xl = 100e3
yl = 100e3
nx = 101
ny = 101
X = np.linspace(0,xl,nx)
Y = np.linspace(0,yl,ny)
x,y = np.meshgrid(X, Y)
u0 = 3e-2
u1 = -1e-4
u = np.zeros((ny,nx))
ylim = 2*yl/(nx-1)
u = np.where(y<ylim, u0, u1*(yl-y)/(yl-ylim))
```
To make sure that the source area is not affected by the precipitation, we will keep $K_f$ in the source constant. For this we need to create a 3D array for $K_f$ that contains both a spatial and temporal variations.
```
nstep = 201 # total number of steps
neq = 101 # number of steps to reach steady-state
teq = 1e7 # time to reach steady-state
period = 1e6 # period of climatic forcing
tfinal = teq + 5*period # final time
# Here we build the time array (note that not all time steps are of the same length)
tim1 = np.linspace(0,teq,101)
tim2 = np.linspace(teq + period/10, tfinal, 100)
tim = np.concatenate((tim1,tim2))
# build precipitation array
precip = np.where(tim>teq, 1 + 0.5*np.sin(2*np.pi*(tim-teq)/period), 1)
# build Kf array and transform it into an xarray of dimension 'time' and 'space'
Kf = 1e-5
m = 0.4
Kf_tim = np.where(tim>teq, Kf*precip**m, Kf)
Kf_tim_space = np.broadcast_to(Kf_tim,(ny,nx,len(tim))).copy()
Kf_tim_space[:,:2,:] = Kf
G_tim = np.where(tim>teq, 0.5/precip, 0.5)
# create xarrays to provide adequate dimensions for FastScape
Kf_xr = xr.DataArray(data=Kf_tim_space.transpose(), dims=['time','y', 'x'])
G_xr = xr.DataArray(data=G_tim, dims=['time'])
# plots the variations of $G$ with time
fig, ax = plt.subplots(nrows = 1, ncols = 1, sharex=False, sharey=True, figsize=(12,7))
ax.plot(tim, G_tim)
# %create_setup transit_model --default --verbose
import xsimlab as xs
ds_in = xs.create_setup(
model=transit_model,
clocks={'time': tim,
'strati': tim[::10]},
master_clock='time',
input_vars={
# nb. of grid nodes in (y, x)
'grid__shape': [ny,nx],
# total grid length in (y, x)
'grid__length': [yl,xl],
# node status at borders
'boundary__status': ['looped','looped','fixed_value','core'],
'uplift__rate': u,
# MFD partioner slope exponent
'flow__slope_exp': 1,
# drainage area exponent
'spl__area_exp': m,
# slope exponent
'spl__slope_exp': 1,
# bedrock channel incision coefficient
'spl__k_coef_bedrock': Kf_xr,
# soil (sediment) channel incision coefficient
'spl__k_coef_soil': Kf_xr,
# detached bedrock transport/deposition coefficient
'spl__g_coef_bedrock': G_xr,
# soil (sediment) transport/deposition coefficient
'spl__g_coef_soil': G_xr,
# surface topography elevation
'topography__elevation': np.random.random((ny,nx)),
# horizon freezing (deactivation) time
'strati__freeze_time': tim,
},
output_vars={'topography__elevation': 'time',
'drainage__area': 'time',
'strati__elevation': 'strati'}
)
with xs.monitoring.ProgressBar():
ds_out = ds_in.xsimlab.run(model=transit_model)
from ipyfastscape import TopoViz3d
app = TopoViz3d(ds_out, canvas_height=600, time_dim="time")
app.components['background_color'].set_color('lightgray')
app.components['vertical_exaggeration'].set_factor(5)
app.components['timestepper'].go_to_time(ds_out.time[99])
app.show()
```
We see that the climatic variations are imprinted in the stratigraphy but not everywhere in the basin: the variations are largest in the vicinity of the main gorge coming out of the mountain where the principal fan forms. Away from the fan, the perturbations are not stored.
Also th ebasin needs to be relatively filled for the climatic signal to be stored (i.e., the amplitude of the sugnal is much smaller in the early stages of development of the basin).
```
fig, ax = plt.subplots(figsize=(12,8))
nout = 101
for iout in range(nout-1, -1, -1):
ds_out.strati__elevation.isel(strati=-1).isel(horizon=iout).sel(x=xl/2)[ds_out.y>ylim].plot()
fig, ax = plt.subplots(figsize=(12,8))
nout = 101
for iout in range(nout-1, -1, -1):
ds_out.strati__elevation.isel(strati=-1).isel(horizon=iout).sel(y=ylim*3).plot()
```
We can also compute the flux coming out of the basin (i.e. across the model boundary) to estimate, by comparison with the precipitation signal, how the climatic signal has been "filtered" or not by the basin/depositional system.
```
nstep = len(ds_out.time)
flux = [0]
sumtop0 = ds_out.topography__elevation.isel(time=0).where(ds_out.y>=ylim).sum()
for step in range(1,nstep):
sumtop = ds_out.topography__elevation.isel(time=step).where(ds_out.y>=ylim).sum()
flux.append(
(sumtop0 - sumtop)/
(ds_out.time.values[step] - ds_out.time.values[step-1])
)
sumtop0 = sumtop
total_area = ds_out.grid__shape[0].values*ds_out.grid__shape[1].values
flux0 = ds_out.uplift__rate.mean().values*total_area
flux = flux/flux0
```
Here we plot the normalized excess/default of the out-going flux (at steady-state it should be equal to 0)
```
fig, ax = plt.subplots(nrows = 1, ncols = 1, sharex=False, sharey=True, figsize=(12,7))
ax.plot(tim, flux, label='flux')
ax.plot(tim, precip, label='precip')
ax.legend()
```
Finally, we can compute the gain (ratio of relative amplitudes between response and forcing) and time lag.
```
mid = 101
amp_flux = flux[mid:].max() - flux[mid:].min()
amp_forcing = precip[mid:].max() - precip[mid:].min()
print('forcing:',amp_forcing,'response:', amp_flux)
print('time lag:',(tim[np.argmax(precip[180:])+180] - tim[np.argmax(flux[180:])+180])/period)
```
Analyze this result; do not hesitate to run the model for different values of the forcing period.
| github_jupyter |
# XLA in Python
[](https://colab.research.google.com/github/google/jax/blob/master/docs/notebooks/XLA_in_Python.ipynb)
<img style="height:100px;" src="https://raw.githubusercontent.com/tensorflow/tensorflow/master/tensorflow/compiler/xla/g3doc/images/xlalogo.png"> <img style="height:100px;" src="https://upload.wikimedia.org/wikipedia/commons/c/c3/Python-logo-notext.svg">
_Anselm Levskaya_, _Qiao Zhang_
XLA is the compiler that JAX uses, and the compiler that TF uses for TPUs and will soon use for all devices, so it's worth some study. However, it's not exactly easy to play with XLA computations directly using the raw C++ interface. JAX exposes the underlying XLA computation builder API through a python wrapper, and makes interacting with the XLA compute model accessible for messing around and prototyping.
XLA computations are built as computation graphs in HLO IR, which is then lowered to LLO that is device specific (CPU, GPU, TPU, etc.).
As end users we interact with the computational primitives offered to us by the HLO spec.
**Caution: This is a pedagogical notebook covering some low level XLA details, the APIs herein are neither public nor stable!**
## References
__xla__: the doc that defines what's in HLO - but note that the doc is incomplete and omits some ops.
https://www.tensorflow.org/xla/operation_semantics
more details on ops in the source code.
https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/client/xla_builder.h
__python xla client__: this is the XLA python client for JAX, and what we're using here.
https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/python/xla_client.py
https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/python/xla_client_test.py
__jax__: you can see how jax interacts with the XLA compute layer for execution and JITing in these files.
https://github.com/google/jax/blob/master/jax/lax.py
https://github.com/google/jax/blob/master/jax/lib/xla_bridge.py
https://github.com/google/jax/blob/master/jax/interpreters/xla.py
## Colab Setup and Imports
```
import numpy as np
# We only need to import JAX's xla_client, not all of JAX.
from jax.lib import xla_client as xc
xops = xc.ops
# Plotting
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib import gridspec
from matplotlib import rcParams
rcParams['image.interpolation'] = 'nearest'
rcParams['image.cmap'] = 'viridis'
rcParams['axes.grid'] = False
```
## Simple Computations
```
# make a computation builder
c = xc.XlaBuilder("simple_scalar")
# define a parameter shape and parameter
param_shape = xc.Shape.array_shape(np.dtype(np.float32), ())
x = xops.Parameter(c, 0, param_shape)
# define computation graph
y = xops.Sin(x)
# build computation graph
# Keep in mind that incorrectly constructed graphs can cause
# your notebook kernel to crash!
computation = c.Build()
# get a cpu backend
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(computation)
# define a host variable with above parameter shape
host_input = np.array(3.0, dtype=np.float32)
# place host variable on device and execute
device_input = cpu_backend.buffer_from_pyval(host_input)
device_out = compiled_computation.execute([device_input ,])
# retrive the result
device_out[0].to_py()
# same as above with vector type:
c = xc.XlaBuilder("simple_vector")
param_shape = xc.Shape.array_shape(np.dtype(np.float32), (3,))
x = xops.Parameter(c, 0, param_shape)
# chain steps by reference:
y = xops.Sin(x)
z = xops.Abs(y)
computation = c.Build()
# get a cpu backend
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(computation)
host_input = np.array([3.0, 4.0, 5.0], dtype=np.float32)
device_input = cpu_backend.buffer_from_pyval(host_input)
device_out = compiled_computation.execute([device_input ,])
# retrive the result
device_out[0].to_py()
```
## Simple While Loop
```
# trivial while loop, decrement until 0
# x = 5
# while x > 0:
# x = x - 1
#
in_shape = xc.Shape.array_shape(np.dtype(np.int32), ())
# body computation:
bcb = xc.XlaBuilder("bodycomp")
x = xops.Parameter(bcb, 0, in_shape)
const1 = xops.Constant(bcb, np.int32(1))
y = xops.Sub(x, const1)
body_computation = bcb.Build()
# test computation:
tcb = xc.XlaBuilder("testcomp")
x = xops.Parameter(tcb, 0, in_shape)
const0 = xops.Constant(tcb, np.int32(0))
y = xops.Gt(x, const0)
test_computation = tcb.Build()
# while computation:
wcb = xc.XlaBuilder("whilecomp")
x = xops.Parameter(wcb, 0, in_shape)
xops.While(test_computation, body_computation, x)
while_computation = wcb.Build()
# Now compile and execute:
# get a cpu backend
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(while_computation)
host_input = np.array(5, dtype=np.int32)
device_input = cpu_backend.buffer_from_pyval(host_input)
device_out = compiled_computation.execute([device_input ,])
# retrive the result
device_out[0].to_py()
```
## While loops w/ Tuples - Newton's Method for sqrt
```
Xsqr = 2
guess = 1.0
converged_delta = 0.001
maxit = 1000
in_shape_0 = xc.Shape.array_shape(np.dtype(np.float32), ())
in_shape_1 = xc.Shape.array_shape(np.dtype(np.float32), ())
in_shape_2 = xc.Shape.array_shape(np.dtype(np.int32), ())
in_tuple_shape = xc.Shape.tuple_shape([in_shape_0, in_shape_1, in_shape_2])
# body computation:
# x_{i+1} = x_i - (x_i**2 - y) / (2 * x_i)
bcb = xc.XlaBuilder("bodycomp")
intuple = xops.Parameter(bcb, 0, in_tuple_shape)
y = xops.GetTupleElement(intuple, 0)
x = xops.GetTupleElement(intuple, 1)
guard_cntr = xops.GetTupleElement(intuple, 2)
new_x = xops.Sub(x, xops.Div(xops.Sub(xops.Mul(x, x), y), xops.Add(x, x)))
result = xops.Tuple(bcb, [y, new_x, xops.Sub(guard_cntr, xops.Constant(bcb, np.int32(1)))])
body_computation = bcb.Build()
# test computation -- convergence and max iteration test
tcb = xc.XlaBuilder("testcomp")
intuple = xops.Parameter(tcb, 0, in_tuple_shape)
y = xops.GetTupleElement(intuple, 0)
x = xops.GetTupleElement(intuple, 1)
guard_cntr = xops.GetTupleElement(intuple, 2)
criterion = xops.Abs(xops.Sub(xops.Mul(x, x), y))
# stop at convergence criteria or too many iterations
test = xops.And(xops.Gt(criterion, xops.Constant(tcb, np.float32(converged_delta))),
xops.Gt(guard_cntr, xops.Constant(tcb, np.int32(0))))
test_computation = tcb.Build()
# while computation:
# since jax does not allow users to create a tuple input directly, we need to
# take multiple parameters and make a intermediate tuple before feeding it as
# an initial carry to while loop
wcb = xc.XlaBuilder("whilecomp")
y = xops.Parameter(wcb, 0, in_shape_0)
x = xops.Parameter(wcb, 1, in_shape_1)
guard_cntr = xops.Parameter(wcb, 2, in_shape_2)
tuple_init_carry = xops.Tuple(wcb, [y, x, guard_cntr])
xops.While(test_computation, body_computation, tuple_init_carry)
while_computation = wcb.Build()
# Now compile and execute:
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(while_computation)
y = np.array(Xsqr, dtype=np.float32)
x = np.array(guess, dtype=np.float32)
maxit = np.array(maxit, dtype=np.int32)
device_input_y = cpu_backend.buffer_from_pyval(y)
device_input_x = cpu_backend.buffer_from_pyval(x)
device_input_maxit = cpu_backend.buffer_from_pyval(maxit)
device_out = compiled_computation.execute([device_input_y, device_input_x, device_input_maxit])
# retrive the result
print("square root of {y} is {x}".format(y=y, x=device_out[1].to_py()))
```
## Calculate Symm Eigenvalues
Let's exploit the XLA QR implementation to solve some eigenvalues for symmetric matrices.
This is the naive QR algorithm, without acceleration for closely-spaced eigenvalue convergence, nor any permutation to sort eigenvalues by magnitude.
```
Niter = 200
matrix_shape = (10, 10)
in_shape_0 = xc.Shape.array_shape(np.dtype(np.float32), matrix_shape)
in_shape_1 = xc.Shape.array_shape(np.dtype(np.int32), ())
in_tuple_shape = xc.Shape.tuple_shape([in_shape_0, in_shape_1])
# body computation -- QR loop: X_i = Q R , X_{i+1} = R Q
bcb = xc.XlaBuilder("bodycomp")
intuple = xops.Parameter(bcb, 0, in_tuple_shape)
x = xops.GetTupleElement(intuple, 0)
cntr = xops.GetTupleElement(intuple, 1)
Q, R = xops.QR(x, True)
RQ = xops.Dot(R, Q)
xops.Tuple(bcb, [RQ, xops.Sub(cntr, xops.Constant(bcb, np.int32(1)))])
body_computation = bcb.Build()
# test computation -- just a for loop condition
tcb = xc.XlaBuilder("testcomp")
intuple = xops.Parameter(tcb, 0, in_tuple_shape)
cntr = xops.GetTupleElement(intuple, 1)
test = xops.Gt(cntr, xops.Constant(tcb, np.int32(0)))
test_computation = tcb.Build()
# while computation:
wcb = xc.XlaBuilder("whilecomp")
x = xops.Parameter(wcb, 0, in_shape_0)
cntr = xops.Parameter(wcb, 1, in_shape_1)
tuple_init_carry = xops.Tuple(wcb, [x, cntr])
xops.While(test_computation, body_computation, tuple_init_carry)
while_computation = wcb.Build()
# Now compile and execute:
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(while_computation)
X = np.random.random(matrix_shape).astype(np.float32)
X = (X + X.T) / 2.0
it = np.array(Niter, dtype=np.int32)
device_input_x = cpu_backend.buffer_from_pyval(X)
device_input_it = cpu_backend.buffer_from_pyval(it)
device_out = compiled_computation.execute([device_input_x, device_input_it])
host_out = device_out[0].to_py()
eigh_vals = host_out.diagonal()
plt.title('D')
plt.imshow(host_out)
print('sorted eigenvalues')
print(np.sort(eigh_vals))
print('sorted eigenvalues from numpy')
print(np.sort(np.linalg.eigh(X)[0]))
print('sorted error')
print(np.sort(eigh_vals) - np.sort(np.linalg.eigh(X)[0]))
```
## Calculate Full Symm Eigensystem
We can also calculate the eigenbasis by accumulating the Qs.
```
Niter = 100
matrix_shape = (10, 10)
in_shape_0 = xc.Shape.array_shape(np.dtype(np.float32), matrix_shape)
in_shape_1 = xc.Shape.array_shape(np.dtype(np.float32), matrix_shape)
in_shape_2 = xc.Shape.array_shape(np.dtype(np.int32), ())
in_tuple_shape = xc.Shape.tuple_shape([in_shape_0, in_shape_1, in_shape_2])
# body computation -- QR loop: X_i = Q R , X_{i+1} = R Q
bcb = xc.XlaBuilder("bodycomp")
intuple = xops.Parameter(bcb, 0, in_tuple_shape)
X = xops.GetTupleElement(intuple, 0)
O = xops.GetTupleElement(intuple, 1)
cntr = xops.GetTupleElement(intuple, 2)
Q, R = xops.QR(X, True)
RQ = xops.Dot(R, Q)
Onew = xops.Dot(O, Q)
xops.Tuple(bcb, [RQ, Onew, xops.Sub(cntr, xops.Constant(bcb, np.int32(1)))])
body_computation = bcb.Build()
# test computation -- just a for loop condition
tcb = xc.XlaBuilder("testcomp")
intuple = xops.Parameter(tcb, 0, in_tuple_shape)
cntr = xops.GetTupleElement(intuple, 2)
test = xops.Gt(cntr, xops.Constant(tcb, np.int32(0)))
test_computation = tcb.Build()
# while computation:
wcb = xc.XlaBuilder("whilecomp")
X = xops.Parameter(wcb, 0, in_shape_0)
O = xops.Parameter(wcb, 1, in_shape_1)
cntr = xops.Parameter(wcb, 2, in_shape_2)
tuple_init_carry = xops.Tuple(wcb, [X, O, cntr])
xops.While(test_computation, body_computation, tuple_init_carry)
while_computation = wcb.Build()
# Now compile and execute:
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(while_computation)
X = np.random.random(matrix_shape).astype(np.float32)
X = (X + X.T) / 2.0
Omat = np.eye(matrix_shape[0], dtype=np.float32)
it = np.array(Niter, dtype=np.int32)
device_input_X = cpu_backend.buffer_from_pyval(X)
device_input_Omat = cpu_backend.buffer_from_pyval(Omat)
device_input_it = cpu_backend.buffer_from_pyval(it)
device_out = compiled_computation.execute([device_input_X, device_input_Omat, device_input_it])
host_out = device_out[0].to_py()
eigh_vals = host_out.diagonal()
eigh_mat = device_out[1].to_py()
plt.title('D')
plt.imshow(host_out)
plt.figure()
plt.title('U')
plt.imshow(eigh_mat)
plt.figure()
plt.title('U^T A U')
plt.imshow(np.dot(np.dot(eigh_mat.T, X), eigh_mat))
print('sorted eigenvalues')
print(np.sort(eigh_vals))
print('sorted eigenvalues from numpy')
print(np.sort(np.linalg.eigh(X)[0]))
print('sorted error')
print(np.sort(eigh_vals) - np.sort(np.linalg.eigh(X)[0]))
```
## Convolutions
I keep hearing from the AGI folks that we can use convolutions to build artificial life. Let's try it out.
```
# Here we borrow convenience functions from LAX to handle conv dimension numbers.
from typing import NamedTuple, Sequence
class ConvDimensionNumbers(NamedTuple):
"""Describes batch, spatial, and feature dimensions of a convolution.
Args:
lhs_spec: a tuple of nonnegative integer dimension numbers containing
`(batch dimension, feature dimension, spatial dimensions...)`.
rhs_spec: a tuple of nonnegative integer dimension numbers containing
`(out feature dimension, in feature dimension, spatial dimensions...)`.
out_spec: a tuple of nonnegative integer dimension numbers containing
`(batch dimension, feature dimension, spatial dimensions...)`.
"""
lhs_spec: Sequence[int]
rhs_spec: Sequence[int]
out_spec: Sequence[int]
def _conv_general_proto(dimension_numbers):
assert type(dimension_numbers) is ConvDimensionNumbers
lhs_spec, rhs_spec, out_spec = dimension_numbers
proto = xc.ConvolutionDimensionNumbers()
proto.input_batch_dimension = lhs_spec[0]
proto.input_feature_dimension = lhs_spec[1]
proto.output_batch_dimension = out_spec[0]
proto.output_feature_dimension = out_spec[1]
proto.kernel_output_feature_dimension = rhs_spec[0]
proto.kernel_input_feature_dimension = rhs_spec[1]
proto.input_spatial_dimensions.extend(lhs_spec[2:])
proto.kernel_spatial_dimensions.extend(rhs_spec[2:])
proto.output_spatial_dimensions.extend(out_spec[2:])
return proto
Niter=13
matrix_shape = (1, 1, 20, 20)
in_shape_0 = xc.Shape.array_shape(np.dtype(np.int32), matrix_shape)
in_shape_1 = xc.Shape.array_shape(np.dtype(np.int32), ())
in_tuple_shape = xc.Shape.tuple_shape([in_shape_0, in_shape_1])
# Body computation -- Conway Update
bcb = xc.XlaBuilder("bodycomp")
intuple = xops.Parameter(bcb, 0, in_tuple_shape)
x = xops.GetTupleElement(intuple, 0)
cntr = xops.GetTupleElement(intuple, 1)
# convs require floating-point type
xf = xops.ConvertElementType(x, xc.DTYPE_TO_XLA_ELEMENT_TYPE['float32'])
stamp = xops.Constant(bcb, np.ones((1,1,3,3), dtype=np.float32))
conv_dim_num_proto = _conv_general_proto(ConvDimensionNumbers(lhs_spec=(0,1,2,3), rhs_spec=(0,1,2,3), out_spec=(0,1,2,3)))
convd = xops.ConvGeneralDilated(xf, stamp, [1, 1], [(1, 1), (1, 1)], (), (), conv_dim_num_proto)
# # logic ops require integer types
convd = xops.ConvertElementType(convd, xc.DTYPE_TO_XLA_ELEMENT_TYPE['int32'])
bool_x = xops.Eq(x, xops.Constant(bcb, np.int32(1)))
# core update rule
res = xops.Or(
# birth rule
xops.And(xops.Not(bool_x), xops.Eq(convd, xops.Constant(bcb, np.int32(3)))),
# survival rule
xops.And(bool_x, xops.Or(
# these are +1 the normal numbers since conv-sum counts self
xops.Eq(convd, xops.Constant(bcb, np.int32(4))),
xops.Eq(convd, xops.Constant(bcb, np.int32(3))))
)
)
# Convert output back to int type for type constancy
int_res = xops.ConvertElementType(res, xc.DTYPE_TO_XLA_ELEMENT_TYPE['int32'])
xops.Tuple(bcb, [int_res, xops.Sub(cntr, xops.Constant(bcb, np.int32(1)))])
body_computation = bcb.Build()
# Test computation -- just a for loop condition
tcb = xc.XlaBuilder("testcomp")
intuple = xops.Parameter(tcb, 0, in_tuple_shape)
cntr = xops.GetTupleElement(intuple, 1)
test = xops.Gt(cntr, xops.Constant(tcb, np.int32(0)))
test_computation = tcb.Build()
# While computation:
wcb = xc.XlaBuilder("whilecomp")
x = xops.Parameter(wcb, 0, in_shape_0)
cntr = xops.Parameter(wcb, 1, in_shape_1)
tuple_init_carry = xops.Tuple(wcb, [x, cntr])
xops.While(test_computation, body_computation, tuple_init_carry)
while_computation = wcb.Build()
# Now compile and execute:
cpu_backend = xc.get_local_backend("cpu")
# compile graph based on shape
compiled_computation = cpu_backend.compile(while_computation)
# Set up initial state
X = np.zeros(matrix_shape, dtype=np.int32)
X[0,0, 5:8, 5:8] = np.array([[0,1,0],[0,0,1],[1,1,1]])
# Evolve
movie = np.zeros((Niter,)+matrix_shape[-2:], dtype=np.int32)
for it in range(Niter):
itr = np.array(it, dtype=np.int32)
device_input_x = cpu_backend.buffer_from_pyval(X)
device_input_it = cpu_backend.buffer_from_pyval(itr)
device_out = compiled_computation.execute([device_input_x, device_input_it])
movie[it] = device_out[0].to_py()[0,0]
# Plot
fig = plt.figure(figsize=(15,2))
gs = gridspec.GridSpec(1,Niter)
for i in range(Niter):
ax1 = plt.subplot(gs[:, i])
ax1.axis('off')
ax1.imshow(movie[i])
plt.subplots_adjust(left=0.0, right=1.0, top=1.0, bottom=0.0, hspace=0.0, wspace=0.05)
```
## Fin
There's much more to XLA, but this hopefully highlights how easy it is to play with via the python client!
| github_jupyter |
# Getting started with Bruges
This notebook accompanies [a blog post on agilegeoscience.com](http://www.agilegeoscience.com/blog/).
You need to install Bruges to run this notebook. You can clone the repository on GitHub or if you have pip installed, in a terminal you can do,
pip install bruges
This notebook also requires Striplog, which you can install like so:
pip install striplog
```
import bruges as b
b.__version__
```
<hr />
## AVO calculations
| | Vp [m/s] | Vs [m/s] | Density [kg/m3] |
|-------------|----------|----------|----------|
| upper layer | 3300 | 1500 | 2400 |
| lower layer | 3050 | 1400 | 2075 |
```
# Upper layer rock properties
vp1 = 3300.0
vs1 = 1500.0
rho1 = 2400.0
# Lower layer rock properties
vp2 = 3050.0
vs2 = 1400.0
rho2 = 2075.0
rc_0 = b.reflection.akirichards(vp1, vs1, rho1, vp2, vs2, rho2, theta1=0)
rc_0
rc_30 = b.reflection.akirichards(vp1, vs1, rho1, vp2, vs2, rho2, theta1=30)
rc_30
```
We can also get the individual terms of Shuey's linear approximation:
```
rc_terms = b.reflection.shuey2(vp1, vs1, rho1, vp2, vs2, rho2, theta1=30, terms=True)
rc_terms
```
The second term is the product of gradient and $sin^2 \theta$. So we can compute the gradient from this:
```
import numpy as np
intercept = rc_terms[0]
gradient = rc_terms[1] / np.sin(np.radians(30))**2
intercept, gradient
```
We can pass in a list of angles,
```
theta_list = [0, 10, 20, 30]
rc_list = b.reflection.akirichards(vp1, vs1, rho1, vp2, vs2, rho2, theta1=theta_list)
rc_list
```
Create an array of angles from 0 to 70, incremented by 1,
```
theta_range = np.arange(0, 70)
```
Create some variables for our data so we don't have to type so much
```
rc_range = b.reflection.akirichards(vp1, vs1, rho1, vp2, vs2, rho2, theta_range)
```
Compare the two-term Aki-Richards approximation with the full Zoeprittz equation for a interface between two rocks:
```
rc_z = b.reflection.zoeppritz(vp1, vs1, rho1, vp2, vs2, rho2, theta_range)
```
Put all this data on an AVO plot:
```
import matplotlib.pyplot as plt
%matplotlib inline
style = {'color': 'blue',
'fontsize': 10,
'ha':'left',
'va':'top',}
fig = plt.figure(figsize=(12,5))
# AVO plot
ax1 = fig.add_subplot(121)
ax1.plot(theta_range, rc_z, 'k', lw=3, alpha=0.25, label='Zoeppritz')
ax1.plot(theta_range, rc_range, 'k.', lw=3, alpha=0.5, label='Aki-Richards')
# We'll also add the four angles...
ax1.plot(theta_list, rc_list, 'bo', ms=10, alpha = 0.5)
# Putting some annotations on the plot.
for theta, rc in zip(theta_list, rc_list):
ax1.text(theta, rc-0.004, '{:.3f}'.format(rc), **style)
ax1.legend()
ax1.set_ylim((-0.15, -0.05))
ax1.set_xlabel('Angle (degrees)')
ax1.set_ylabel('Amplitude')
ax1.grid()
# Intercept-Gradient crossplot.
ax2 = fig.add_subplot(122)
ax2.plot(intercept, gradient, 'bo', ms=10, alpha = 0.5)
# Put spines for x and y axis.
ax2.axvline(0, color='k')
ax2.axhline(0, color='k')
# Set square axes limits.
mx = 0.25
ax2.set_xlim((-mx, mx))
ax2.set_ylim((-mx, mx))
# Label the axes and add gridlines.
ax2.set_xlabel('Intercept')
ax2.set_ylabel('Gradient')
ax2.grid()
plt.show()
```
<hr />
## Elastic moduli calculations
Say I want to compute the Lamé parameters λ and µ, from V<sub>P</sub>, V<sub>S</sub>, and Density. As long as my inputs are in SI units, I can insert these values directly:
| | Vp [m/s] | Vs [m/s] | Density [kg/m3] |
|-------------|----------|----------|----------|
| upper layer | 3300 | 1500 | 2400 |
| lower layer | 3050 | 1400 | 2075 |
```
# Upper layer only
b.rockphysics.lam(vp1, vs1, rho1), b.rockphysics.mu(vp1, vs1, rho1)
```
We can print all the values in terms of GPa ($\times 10^9$ Pa)
```
print('Upper layer')
print('lambda [GPa]: {:.2f}'.format(b.rockphysics.lam(vp1, vs1, rho1) / 1e9))
print(' mu [GPa]: {:.2f}'.format(b.rockphysics.mu(vp1, vs1, rho1) / 1e9))
print('\nLower layer')
print('lambda [GPa]: {:.2f}'.format(b.rockphysics.lam(vp2, vs2, rho2) / 1e9))
print(' mu [GPa]: {:.2f}'.format(b.rockphysics.mu(vp2, vs2, rho2) / 1e9))
```
<hr />
## Rock property analysis and crossplotting
```
import striplog
striplog.__version__
from striplog import Well
well = Well('data/P-132_synth.las', null_subs=np.nan)
# Crop the log to first and last RHOB samples, where the data is:
rhob, dt, z = b.util.top_and_tail(well.data['RHOB_despiked'], well.data['Sonic_despiked'], well.data['DEPT'])
```
Convert everything to SI units:
```
vp = 1e6 / (3.28084 * dt)
vs = (vp - 1360)/1.16 # Castanga model
rhob *= 1000
lm_log = b.rockphysics.lam(vp, vs, rhob)
mu_log = b.rockphysics.mu(vp, vs, rhob)
```
Create a crossplot:
```
plt.figure(figsize=(7,6))
plt.scatter(lm_log*rhob, mu_log*rhob, s=30, c=z, cmap="gist_earth",
edgecolor='none', # marker edge color
alpha = 0.05)
# Give the plot a colorbar.
cb = plt.colorbar(shrink=0.75)
cb.ax.invert_yaxis() # put shallow colors at the top
cb.set_ticks([500,1000,1500,2000,2500,3000])
cb.set_label("Depth [m]")
# Give the plot some annotation.
plt.xlabel(r'$\lambda \rho$', size=18)
plt.ylabel(r'$\mu \rho$', size=18)
plt.grid()
```
<hr />
## Backus averaging
```
lb = 60 # Backus averaging length in metres.
dz = 0.1524 # Sample interval of the log in metres.
vp0, vs0 = b.rockphysics.backus(vp, vs, rhob, lb, dz)
vp0
fs = 8 # control the fontsize
fig = plt.figure(figsize=(3,8))
ax1 = plt.subplot(111)
ax1.plot(vp, z, 'k', alpha=0.25, label='Vp')
ax1.plot(vp0, z, 'b', label='Backus average')
# Do some plot annotations
ax1.set_title('P-velocity')
ax1.set_ylabel(r'depth $m$', size=12)
ax1.invert_yaxis()
ax1.set_xlim((4000,6200))
ax1.set_xlabel(r'Vp $m/s$')
ax1.grid()
ax1.legend()
labels = ax1.get_xticklabels()
for label in labels:
label.set_rotation(90)
label.set_fontsize(fs)
plt.show()
```
<hr />
<img src="https://avatars1.githubusercontent.com/u/1692321?v=3&s=200" style="float:right;" width="40px" /><p style="color:gray; float:right;">© 2015 <a href="http://www.agilegeoscience.com/">Agile Geoscience</a> — <a href="https://creativecommons.org/licenses/by/4.0/">CC-BY</a> — Have fun! </p>
| github_jupyter |
```
import heapq, random, itertools, operator, functools, math
import matplotlib.pyplot as plt
from sympy import IndexedBase
def take(iterable, n):
for _ in range(n):
yield next(iterable)
a = IndexedBase('a')
actions = [a[i] for i in range(10)]
actions
q_star = { a: random.gauss(0, 1) for a in actions }
def rewarder(q_star, process, *, stationarity):
reward = None
Q = {a: v for a, v in q_star.items()}
while True:
action, q, n, t, by = process.send(reward)
qs = Q[action]
reward = random.gauss(qs, 1)
yield action, q, qs, by, reward
Q = {a:stationarity(a, q) for a, q in Q.items()}
def k_armed_bandit(actions, epsilon, init=lambda action: 0, *, update,measure):
Q = [(-q, -q, 0, i, a) for i, a in enumerate(actions) for q in [init(a)]]
heapq.heapify(Q)
for t in itertools.count():
by = None
if random.random() > epsilon:
_, q, n, i, a = heapq.heappop(Q)
by = 'e' # exploiting flag
else:
j = random.randrange(0, len(Q))
_, q, n, i, a = Q.pop(j)
heapq.heapify(Q) # takes linear time to maintain the heap invariant.
by = 's' # search flag
q = -q # to restore "max-heap" behavior
n_next = n + 1
tup = (a, q, n_next, t, by)
reward = yield tup
q_updated = update(*(tup + (reward,)))
q_measured = measure(a, q_updated, n_next, t, by, reward)
heapq.heappush(Q, (-q_measured, -q_updated, n_next, i, a))
def optimistic_k_armed_bandit(init):
return functools.partial(k_armed_bandit, init=init, measure=lambda *args: args[1])
def sample_average(action, q, n, t, by, reward):
return q + ((reward - q)/n)
def exponential_recency_weighted_average(alpha):
return lambda action, q, n, t, by, reward: q + alpha*(reward - q)
def upper_confidence_bound(c):
def M(action, q, n, t, by, reward):
#return q if by == 's' else (q + c * math.sqrt(math.log(t+1)/n))
return q + c * math.sqrt(math.log(t+1)/n)
return functools.partial(k_armed_bandit, measure=M)
def unbiased_exponential_recency_weighted_average(alpha):
o = 0.0
def unbiased(action, q, n, t, by, reward):
nonlocal o
o = o + alpha*(1-o)
step_size = alpha / o
return q + step_size*(reward-q)
return unbiased
def stationary(a, q):
return q
def gauss_error_non_stationary(sd):
return lambda a, q: q + random.gauss(0, sd)
def avg(*args):
return sum(args)/len(args)
R = rewarder(q_star,
process=k_armed_bandit(actions, epsilon=0.1, update=sample_average),
stationarity=stationary)
plt.plot(list(take(map(lambda tup: tup[-1], R), 1000)));
R = rewarder(q_star,
process=optimistic_k_armed_bandit(actions, epsilon=0.1, update=sample_average),
stationarity=stationary)
plt.plot(list(take(map(lambda tup: tup[-1], R), 1000)));
list(take(R, 100))
R = rewarder(q_star,
process=k_armed_bandit(actions,
epsilon=0.1,
update=exponential_recency_weighted_average(alpha=0.1)),
stationarity=gauss_error_non_stationary(sd=0.01))
plt.plot(list(take(map(lambda tup: tup[-1], R), 1000)));
R = rewarder(q_star,
process=k_armed_bandit(actions,
epsilon=0.1,
update=unbiased_exponential_recency_weighted_average(alpha=0.1)),
stationarity=gauss_error_non_stationary(sd=0.01))
plt.plot(list(take(map(lambda tup: tup[-1], R), 1000)));
def benchmark(confs, nprocesses=2000, length=10000, filename=None):
plt.ylim(-0.5, 2)
avgs = []
for conf in confs:
process = conf['process']
runs = [map(lambda tup: tup[-1],
rewarder(q_star,
process= process(actions,
epsilon=conf['epsilon'],
update=conf['update']),
stationarity=conf['stationarity']))
for i in range(nprocesses)]
R = list(take(map(avg, *runs), length))
avgs.append(R)
plt.plot(R, label=conf['label'])
ret = plt.legend()
if filename:
plt.savefig(filename)
return ret, avgs
confs = [
{
'epsilon':0.1,
'update':sample_average,
'stationarity':stationary,
'label': 'stationary, eps=0.1, sample average'
},
{
'epsilon':0.1,
'update':exponential_recency_weighted_average(alpha=0.1),
'stationarity':stationary,
'label': 'stationary, eps=0.1, ERW=0.1 average'
},
{
'epsilon':0.1,
'update':unbiased_exponential_recency_weighted_average(alpha=0.1),
'stationarity':stationary,
'label': 'stationary, eps=0.1, ERW=0.1 unbiased average'
},
{
'epsilon':0.1,
'update':sample_average,
'stationarity':gauss_error_non_stationary(sd=0.01),
'label': 'non stationary with gaussian errs, eps=0.1, sample average'
},
{
'epsilon':0.1,
'update':exponential_recency_weighted_average(alpha=0.1),
'stationarity':gauss_error_non_stationary(sd=0.01),
'label': 'non stationary with gaussian errs, eps=0.1, ERW=0.1 average'
},
{
'epsilon':0.1,
'update':unbiased_exponential_recency_weighted_average(alpha=0.1),
'stationarity':gauss_error_non_stationary(sd=0.01),
'label': 'non stationary with gaussian errs, eps=0.1, ERW=0.1 unbiased average'
},
]
# bench
fig, avgs = benchmark(confs, filename='non-stationary-benchmark.svg', length=10_000)
fig
confs = [
{
'epsilon':0,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.0, sample average, stationary'
},
{
'epsilon':0.01,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.01, sample average, stationary'
},
{
'epsilon':0.1,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.1, sample average, stationary'
},
{
'epsilon':0.5,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.5, sample average, stationary'
},
]
# bench
fig, avgs = benchmark(confs, filename='stationary-benchmark.svg', length=10_000)
fig;
confs = [
{
'epsilon':0,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.0, sample average, stationary'
},
{
'epsilon':0.01,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.01, sample average, stationary'
},
{
'epsilon':0.1,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.1, sample average, stationary'
},
{
'epsilon':0.5,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy, eps=0.5, sample average, stationary'
},
]
# bench
fig, avgs = benchmark(confs, filename='stationary-benchmark.svg', length=10_000)
fig;
confs = [
{
'process': functools.partial(k_armed_bandit, measure=lambda *args: args[1]),
'epsilon':0.1,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy realistic, eps=0.0, sample average, stationary'
},
{
'process': optimistic_k_armed_bandit(init=lambda action: 100),
'epsilon':0.1,
'update':sample_average,
'stationarity':stationary,
'label': 'greedy optimistic, eps=0.01, sample average, stationary'
},
{
'process': upper_confidence_bound(c=2),
'epsilon':0,
'update':sample_average,
'stationarity':stationary,
'label': 'upper confidence bound, eps=0.0, sample average, stationary'
},
]
# bench
fig, avgs = benchmark(confs, filename='stationary-benchmark.svg', nprocesses=2000, length=500)
fig;
```
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
#export
from nb_005b import *
```
# Carvana
## Setup
(See final section of notebook for one-time data processing steps.)
```
PATH = Path('../../data/carvana')
PATH_PNG = PATH/'train_masks_png'
PATH_X_FULL = PATH/'train'
PATH_X_128 = PATH/'train-128'
PATH_Y_FULL = PATH_PNG
PATH_Y_128 = PATH/'train_masks-128'
# start with the 128x128 images
PATH_X = PATH_X_128
PATH_Y = PATH_Y_128
img_f = next(PATH_X.iterdir())
open_image(img_f).show()
#export
class ImageMask(Image):
"Class for image segmentation target"
def clone(self)->'ImageBase':
"Clones this item"
return self.__class__(self.px.clone())
def lighting(self, func:LightingFunc, *args:Any, **kwargs:Any)->'Image': return self
def refresh(self):
self.sample_kwargs['mode'] = 'nearest'
return super().refresh()
@property
def data(self)->TensorImage:
"Returns this images pixels as a tensor"
return self.px.long()
def open_mask(fn:PathOrStr) -> ImageMask:
"Return `ImageMask` object create from mask in file `fn`"
return ImageMask(pil2tensor(PIL.Image.open(fn)).float())
def get_y_fn(x_fn): return PATH_Y/f'{x_fn.name[:-4]}_mask.png'
img_y_f = get_y_fn(img_f)
y = open_mask(img_y_f)
y.show()
#export
# Same as `show_image`, but renamed with _ prefix
def _show_image(img:Image, ax:plt.Axes=None, figsize:tuple=(3,3), hide_axis:bool=True, cmap:str='binary',
alpha:float=None) -> plt.Axes:
if ax is None: fig,ax = plt.subplots(figsize=figsize)
ax.imshow(image2np(img), cmap=cmap, alpha=alpha)
if hide_axis: ax.axis('off')
return ax
def show_image(x:Image, y:Image=None, ax:plt.Axes=None, figsize:tuple=(3,3), alpha:float=0.5,
hide_axis:bool=True, cmap:str='viridis'):
ax1 = _show_image(x, ax=ax, hide_axis=hide_axis, cmap=cmap)
if y is not None: _show_image(y, ax=ax1, alpha=alpha, hide_axis=hide_axis, cmap=cmap)
if hide_axis: ax1.axis('off')
def _show(self:Image, ax:plt.Axes=None, y:Image=None, **kwargs):
if y is not None: y=y.data
return show_image(self.data, ax=ax, y=y, **kwargs)
Image.show = _show
x = open_image(img_f)
x.show(y=y)
x.shape
y.shape
```
## Dataset
- data types: regr, class, seg, bbox, polygon, generative (s/res, color), custom
```
#export
class DatasetTfm(Dataset):
"`Dataset` that applies a list of transforms to every item drawn"
def __init__(self, ds:Dataset, tfms:TfmList=None, tfm_y:bool=False, **kwargs:Any):
"this dataset will apply `tfms` to `ds`"
self.ds,self.tfms,self.kwargs,self.tfm_y = ds,tfms,kwargs,tfm_y
self.y_kwargs = {**self.kwargs, 'do_resolve':False}
def __len__(self)->int: return len(self.ds)
def __getitem__(self,idx:int)->Tuple[Image,Any]:
"returns tfms(x),y"
x,y = self.ds[idx]
x = apply_tfms(self.tfms, x, **self.kwargs)
if self.tfm_y: y = apply_tfms(self.tfms, y, **self.y_kwargs)
return x, y
def __getattr__(self,k):
"passthrough access to wrapped dataset attributes"
return getattr(self.ds, k)
import nb_002b
nb_002b.DatasetTfm = DatasetTfm
#export
class SegmentationDataset(DatasetBase):
"A dataset for segmentation task"
def __init__(self, x:Collection[PathOrStr], y:Collection[PathOrStr]):
assert len(x)==len(y)
self.x,self.y = np.array(x),np.array(y)
def __getitem__(self, i:int) -> Tuple[Image,ImageMask]:
return open_image(self.x[i]), open_mask(self.y[i])
def get_datasets(path):
x_fns = [o for o in path.iterdir() if o.is_file()]
y_fns = [get_y_fn(o) for o in x_fns]
mask = [o>=1008 for o in range(len(x_fns))]
arrs = arrays_split(mask, x_fns, y_fns)
return [SegmentationDataset(*o) for o in arrs]
train_ds,valid_ds = get_datasets(PATH_X_128)
train_ds,valid_ds
x,y = next(iter(train_ds))
x.shape, y.shape, type(x), type(y)
size=128
def get_tfm_datasets(size):
datasets = get_datasets(PATH_X_128 if size<=128 else PATH_X_FULL)
tfms = get_transforms(do_flip=True, max_rotate=4, max_lighting=0.2)
return transform_datasets(train_ds, valid_ds, tfms=tfms, tfm_y=True, size=size, padding_mode='border')
transform_datasets
train_tds,*_ = get_tfm_datasets(size)
_,axes = plt.subplots(1,4, figsize=(12,6))
for i, ax in enumerate(axes.flat):
imgx,imgy = train_tds[i]
imgx.show(ax, y=imgy)
default_norm,default_denorm = normalize_funcs(*imagenet_stats)
bs = 64
def get_data(size, bs):
return DataBunch.create(*get_tfm_datasets(size), bs=bs, tfms=default_norm)
data = get_data(size, bs)
#export
def show_xy_images(x:Tensor,y:Tensor,rows:int,figsize:tuple=(9,9)):
"Shows a selection of images and targets from a given batch."
fig, axs = plt.subplots(rows,rows,figsize=figsize)
for i, ax in enumerate(axs.flatten()): show_image(x[i], y=y[i], ax=ax)
plt.tight_layout()
x,y = next(iter(data.train_dl))
x,y = x.cpu(),y.cpu()
x = default_denorm(x)
show_xy_images(x,y,4, figsize=(9,9))
x.shape, y.shape
```
## Model
```
#export
class Debugger(nn.Module):
"A module to debug inside a model"
def forward(self,x:Tensor) -> Tensor:
set_trace()
return x
class StdUpsample(nn.Module):
"Standard upsample module"
def __init__(self, n_in:int, n_out:int):
super().__init__()
self.conv = conv2d_trans(n_in, n_out)
self.bn = nn.BatchNorm2d(n_out)
def forward(self, x:Tensor) -> Tensor:
return self.bn(F.relu(self.conv(x)))
def std_upsample_head(c, *nfs:Collection[int]) -> Model:
"Creates a sequence of upsample layers"
return nn.Sequential(
nn.ReLU(),
*(StdUpsample(nfs[i],nfs[i+1]) for i in range(4)),
conv2d_trans(nfs[-1], c)
)
head = std_upsample_head(2, 512,256,256,256,256)
head
#export
def dice(input:Tensor, targs:Tensor) -> Rank0Tensor:
"Dice coefficient metric for binary target"
n = targs.shape[0]
input = input.argmax(dim=1).view(n,-1)
targs = targs.view(n,-1)
intersect = (input*targs).sum().float()
union = (input+targs).sum().float()
return 2. * intersect / union
def accuracy(input:Tensor, targs:Tensor) -> Rank0Tensor:
"Accuracy"
n = targs.shape[0]
input = input.argmax(dim=1).view(n,-1)
targs = targs.view(n,-1)
return (input==targs).float().mean()
class CrossEntropyFlat(nn.CrossEntropyLoss):
"Same as `nn.CrossEntropyLoss`, but flattens input and target"
def forward(self, input:Tensor, target:Tensor) -> Rank0Tensor:
n,c,*_ = input.shape
return super().forward(input.view(n, c, -1), target.view(n, -1))
metrics=[accuracy, dice]
learn = ConvLearner(data, tvm.resnet34, 2, custom_head=head,
metrics=metrics, loss_fn=CrossEntropyFlat())
lr_find(learn)
learn.recorder.plot()
lr = 1e-1
learn.fit_one_cycle(10, slice(lr))
learn.unfreeze()
learn.save('0')
learn.load('0')
lr = 2e-2
learn.fit_one_cycle(10, slice(lr/100,lr))
x,y,py = learn.pred_batch()
py = py.argmax(dim=1).unsqueeze(1)
for i, ax in enumerate(plt.subplots(4,4,figsize=(10,10))[1].flat):
show_image(default_denorm(x[i].cpu()), py[i], ax=ax)
learn.save('1')
size=512
bs = 8
data = get_data(size, bs)
learn.data = data
learn.load('1')
learn.freeze()
lr = 2e-2
learn.fit_one_cycle(5, slice(lr))
learn.save('2')
learn.load('2')
lr = 2e-2
learn.unfreeze()
learn.fit_one_cycle(8, slice(lr/100,lr))
learn.save('3')
x,py = learn.pred_batch()
for i, ax in enumerate(plt.subplots(4,4,figsize=(10,10))[1].flat):
show_image(default_denorm(x[i].cpu()), py[i]>0, ax=ax)
```
## Preprocessing steps
```
def convert_img(fn): Image.open(fn).save(PATH_PNG/f'{fn.name[:-4]}.png')
def resize_img(fn, dirname):
Image.open(fn).resize((128,128)).save((fn.parent.parent)/dirname/fn.name)
def do_conversion():
PATH_PNG.mkdir(exist_ok=True)
PATH_X.mkdir(exist_ok=True)
PATH_Y.mkdir(exist_ok=True)
files = list((PATH/'train_masks').iterdir())
with ThreadPoolExecutor(8) as e: e.map(convert_img, files)
files = list((PATH_PNG).iterdir())
with ThreadPoolExecutor(8) as e: e.map(partial(resize_img, dirname='train_masks-128'), files)
files = list((PATH/'train').iterdir())
with ThreadPoolExecutor(8) as e: e.map(partial(resize_img, dirname='train-128'), files)
```
| github_jupyter |
# Large scale text analysis with deep learning (3 points)
Today we're gonna apply the newly learned tools for the task of predicting job salary.
<img src="https://storage.googleapis.com/kaggle-competitions/kaggle/3342/media/salary%20prediction%20engine%20v2.png" width=400px>
_Special thanks to [Oleg Vasilev](https://github.com/Omrigan/) for the core assignment idea._
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
```
### About the challenge
For starters, let's download and unpack the data from [here].
You can also get it from [yadisk url](https://yadi.sk/d/vVEOWPFY3NruT7) the competition [page](https://www.kaggle.com/c/job-salary-prediction/data) (pick `Train_rev1.*`).
```
!wget https://ysda-seminars.s3.eu-central-1.amazonaws.com/Train_rev1.zip
!unzip Train_rev1.zip
data = pd.read_csv("./Train_rev1.csv", index_col=None)
data.shape
data.head()
```
One problem with salary prediction is that it's oddly distributed: there are many people who are paid standard salaries and a few that get tons o money. The distribution is fat-tailed on the right side, which is inconvenient for MSE minimization.
There are several techniques to combat this: using a different loss function, predicting log-target instead of raw target or even replacing targets with their percentiles among all salaries in the training set. We gonna use logarithm for now.
_You can read more [in the official description](https://www.kaggle.com/c/job-salary-prediction#description)._
```
data['Log1pSalary'] = np.log1p(data['SalaryNormalized']).astype('float32')
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.hist(data["SalaryNormalized"], bins=20);
plt.subplot(1, 2, 2)
plt.hist(data['Log1pSalary'], bins=20);
```
Our task is to predict one number, __Log1pSalary__.
To do so, our model can access a number of features:
* Free text: __`Title`__ and __`FullDescription`__
* Categorical: __`Category`__, __`Company`__, __`LocationNormalized`__, __`ContractType`__, and __`ContractTime`__.
```
text_columns = ["Title", "FullDescription"]
categorical_columns = ["Category", "Company", "LocationNormalized", "ContractType", "ContractTime"]
TARGET_COLUMN = "Log1pSalary"
data[categorical_columns] = data[categorical_columns].fillna('NaN') # cast missing values to string "NaN"
data.sample(3)
```
### Preprocessing text data
Just like last week, applying NLP to a problem begins from tokenization: splitting raw text into sequences of tokens (words, punctuation, etc).
__Your task__ is to lowercase and tokenize all texts under `Title` and `FullDescription` columns. Store the tokenized data as a __space-separated__ string of tokens for performance reasons.
It's okay to use nltk tokenizers. Assertions were designed for WordPunctTokenizer, slight deviations are okay.
```
print("Raw text:")
print(data["FullDescription"][2::100000])
import nltk
#TODO YOUR CODE HERE
tokenizer = nltk.tokenize.WordPunctTokenizer()
# YOUR CODE HERE
```
Now we can assume that our text is a space-separated list of tokens:
```
print("Tokenized:")
print(data["FullDescription"][2::100000])
assert data["FullDescription"][2][:50] == 'mathematical modeller / simulation analyst / opera'
assert data["Title"][54321] == 'international digital account manager ( german )'
```
Not all words are equally useful. Some of them are typos or rare words that are only present a few times.
Let's count how many times is each word present in the data so that we can build a "white list" of known words.
```
from collections import Counter
token_counts = Counter()
# Count how many times does each token occur in both "Title" and "FullDescription" in total
#TODO <YOUR CODE>
print("Total unique tokens :", len(token_counts))
print('\n'.join(map(str, token_counts.most_common(n=5))))
print('...')
print('\n'.join(map(str, token_counts.most_common()[-3:])))
assert token_counts.most_common(1)[0][1] in range(2600000, 2700000)
assert len(token_counts) in range(200000, 210000)
print('Correct!')
# Let's see how many words are there for each count
plt.hist(list(token_counts.values()), range=[0, 10**4], bins=50, log=True)
plt.xlabel("Word counts");
```
__Task 1.1__ Get a list of all tokens that occur at least 10 times.
```
min_count = 10
# tokens from token_counts keys that had at least min_count occurrences throughout the dataset
tokens = sorted(t for t, c in token_counts.items() if c >= min_count)#TODO<YOUR CODE HERE>
# Add a special tokens for unknown and empty words
UNK, PAD = "UNK", "PAD"
tokens = [UNK, PAD] + tokens
print("Vocabulary size:", len(tokens))
assert type(tokens) == list
assert len(tokens) in range(32000, 35000)
assert 'me' in tokens
assert UNK in tokens
print("Correct!")
```
__Task 1.2__ Build an inverse token index: a dictionary from token(string) to it's index in `tokens` (int)
```
token_to_id = {t: i for i, t in enumerate(tokens)}#TODO<your code here>
assert isinstance(token_to_id, dict)
assert len(token_to_id) == len(tokens)
for tok in tokens:
assert tokens[token_to_id[tok]] == tok
print("Correct!")
```
And finally, let's use the vocabulary you've built to map text lines into neural network-digestible matrices.
```
UNK_IX, PAD_IX = map(token_to_id.get, [UNK, PAD])
def as_matrix(sequences, max_len=None):
""" Convert a list of tokens into a matrix with padding """
if isinstance(sequences[0], str):
sequences = list(map(str.split, sequences))
max_len = min(max(map(len, sequences)), max_len or float('inf'))
matrix = np.full((len(sequences), max_len), np.int32(PAD_IX))
for i,seq in enumerate(sequences):
row_ix = [token_to_id.get(word, UNK_IX) for word in seq[:max_len]]
matrix[i, :len(row_ix)] = row_ix
return matrix
print("Lines:")
print('\n'.join(data["Title"][::100000].values), end='\n\n')
print("Matrix:")
print(as_matrix(data["Title"][::100000]))
```
Now let's encode the categirical data we have.
As usual, we shall use one-hot encoding for simplicity. Kudos if you implement more advanced encodings: tf-idf, pseudo-time-series, etc.
```
from sklearn.feature_extraction import DictVectorizer
# we only consider top-1k most frequent companies to minimize memory usage
top_companies, top_counts = zip(*Counter(data['Company']).most_common(1000))
recognized_companies = set(top_companies)
data["Company"] = data["Company"].apply(lambda comp: comp if comp in recognized_companies else "Other")
categorical_vectorizer = DictVectorizer(dtype=np.float32, sparse=False)
categorical_vectorizer.fit(data[categorical_columns].apply(dict, axis=1))
```
### The deep learning part
Once we've learned to tokenize the data, let's design a machine learning experiment.
As before, we won't focus too much on validation, opting for a simple train-test split.
__To be completely rigorous,__ we've comitted a small crime here: we used the whole data for tokenization and vocabulary building. A more strict way would be to do that part on training set only. You may want to do that and measure the magnitude of changes.
```
from sklearn.model_selection import train_test_split
data_train, data_val = train_test_split(data, test_size=0.2, random_state=42)
data_train.index = range(len(data_train))
data_val.index = range(len(data_val))
print("Train size = ", len(data_train))
print("Validation size = ", len(data_val))
import torch
def to_tensors(batch, device):
batch_tensors = dict()
for key, arr in batch.items():
if key in ["FullDescription", "Title"]:
batch_tensors[key] = torch.tensor(arr, device=device, dtype=torch.int64)
else:
batch_tensors[key] = torch.tensor(arr, device=device)
return batch_tensors
def make_batch(data, max_len=None, word_dropout=0, device=torch.device('cpu')):
"""
Creates a keras-friendly dict from the batch data.
:param word_dropout: replaces token index with UNK_IX with this probability
:returns: a dict with {'title' : int64[batch, title_max_len]
"""
batch = {}
batch["Title"] = as_matrix(data["Title"].values, max_len)
batch["FullDescription"] = as_matrix(data["FullDescription"].values, max_len)
batch['Categorical'] = categorical_vectorizer.transform(data[categorical_columns].apply(dict, axis=1))
if word_dropout != 0:
batch["FullDescription"] = apply_word_dropout(batch["FullDescription"], 1. - word_dropout)
if TARGET_COLUMN in data.columns:
batch[target_column] = data[TARGET_COLUMN].values
return to_tensors(batch, device)
def apply_word_dropout(matrix, keep_prop, replace_with=UNK_IX, pad_ix=PAD_IX,):
dropout_mask = np.random.choice(2, np.shape(matrix), p=[keep_prop, 1 - keep_prop])
dropout_mask &= matrix != pad_ix
return np.choose(dropout_mask, [matrix, np.full_like(matrix, replace_with)])
make_batch(data_train[:3], max_len=10)
```
#### Architecture
Our basic model consists of three branches:
* Title encoder
* Description encoder
* Categorical features encoder
We will then feed all 3 branches into one common network that predicts salary.

This clearly doesn't fit into keras' __Sequential__ interface. To build such a network, one will have to use PyTorch.
```
import torch
import torch.nn as nn
import torch.functional as F
class SalaryPredictor(nn.Module):
def __init__(self, n_tokens=len(tokens), n_cat_features=len(categorical_vectorizer.vocabulary_), hid_size=64):
super().__init__()
# YOUR CODE HERE
def forward(self, batch):
# YOUR CODE HERE
model = SalaryPredictor()
model = SalaryPredictor()
batch = make_batch(data_train[:100])
criterion = nn.MSELoss()
dummy_pred = model(batch)
dummy_loss = criterion(dummy_pred, batch[TARGET_COLUMN])
assert dummy_pred.shape == torch.Size([100])
assert len(torch.unique(dummy_pred)) > 20, "model returns suspiciously few unique outputs. Check your initialization"
assert dummy_loss.ndim == 0 and 0. <= dummy_loss <= 250., "make sure you minimize MSE"
```
#### Training and evaluation
As usual, we gonna feed our monster with random minibatches of data.
As we train, we want to monitor not only loss function, which is computed in log-space, but also the actual error measured in dollars.
```
def iterate_minibatches(data, batch_size=256, shuffle=True, cycle=False, device=torch.device('cpu'), **kwargs):
""" iterates minibatches of data in random order """
while True:
indices = np.arange(len(data))
if shuffle:
indices = np.random.permutation(indices)
for start in range(0, len(indices), batch_size):
batch = make_batch(data.iloc[indices[start : start + batch_size]], **kwargs)
yield batch
if not cycle: break
```
### Model training
We can now fit our model the usual minibatch way. The interesting part is that we train on an infinite stream of minibatches, produced by `iterate_minibatches` function.
```
import tqdm
BATCH_SIZE = 16
EPOCHS = 5
DEVICE = torch.device('cpu')
def print_metrics(model, data, batch_size=BATCH_SIZE, name="", **kw):
squared_error = abs_error = num_samples = 0.0
model.eval()
with torch.no_grad():
for batch in iterate_minibatches(data, batch_size=batch_size, shuffle=False, **kw):
batch_pred = model(batch)
squared_error += torch.sum(torch.square(batch_pred - batch[TARGET_COLUMN]))
abs_error += torch.sum(torch.abs(batch_pred - batch[TARGET_COLUMN]))
num_samples += len(batch_y)
mse = squared_error.detach().cpu().numpy() / num_samples
mae = abs_error.detach().cpu().numpy() / num_samples
print("%s results:" % (name or ""))
print("Mean square error: %.5f" % mse)
print("Mean absolute error: %.5f" % mae)
return mse, mae
model = SalaryPredictor().to(DEVICE)
criterion = nn.MSELoss(reduction='sum')
optimizer = torch.optim.SGD(model.parameters(), lr=1e-4)
for epoch in range(EPOCHS):
print(f"epoch: {epoch}")
model.train()
for i, batch in tqdm.tqdm_notebook(enumerate(
iterate_minibatches(data_train, batch_size=BATCH_SIZE, device=DEVICE)),
total=len(data_train) // BATCH_SIZE
):
pred = model(batch)
loss = criterion(pred, batch[TARGET_COLUMN])
optimizer.zero_grad()
loss.backward()
optimizer.step()
print_metrics(model, data_val)
```
### Bonus part: explaining model predictions
It's usually a good idea to understand how your model works before you let it make actual decisions. It's simple for linear models: just see which words learned positive or negative weights. However, its much harder for neural networks that learn complex nonlinear dependencies.
There are, however, some ways to look inside the black box:
* Seeing how model responds to input perturbations
* Finding inputs that maximize/minimize activation of some chosen neurons (_read more [on distill.pub](https://distill.pub/2018/building-blocks/)_)
* Building local linear approximations to your neural network: [article](https://arxiv.org/abs/1602.04938), [eli5 library](https://github.com/TeamHG-Memex/eli5/tree/master/eli5/formatters)
Today we gonna try the first method just because it's the simplest one.
```
def explain(model, sample, col_name='Title'):
""" Computes the effect each word had on model predictions """
sample = dict(sample)
sample_col_tokens = [tokens[token_to_id.get(tok, 0)] for tok in sample[col_name].split()]
data_drop_one_token = pd.DataFrame([sample] * (len(sample_col_tokens) + 1))
for drop_i in range(len(sample_col_tokens)):
data_drop_one_token.loc[drop_i, col_name] = ' '.join(UNK if i == drop_i else tok
for i, tok in enumerate(sample_col_tokens))
*predictions_drop_one_token, baseline_pred = model.predict(make_batch(data_drop_one_token))[:, 0]
diffs = baseline_pred - predictions_drop_one_token
return list(zip(sample_col_tokens, diffs))
from IPython.display import HTML, display_html
def draw_html(tokens_and_weights, cmap=plt.get_cmap("bwr"), display=True,
token_template="""<span style="background-color: {color_hex}">{token}</span>""",
font_style="font-size:14px;"
):
def get_color_hex(weight):
rgba = cmap(1. / (1 + np.exp(weight)), bytes=True)
return '#%02X%02X%02X' % rgba[:3]
tokens_html = [
token_template.format(token=token, color_hex=get_color_hex(weight))
for token, weight in tokens_and_weights
]
raw_html = """<p style="{}">{}</p>""".format(font_style, ' '.join(tokens_html))
if display:
display_html(HTML(raw_html))
return raw_html
i = 36605
tokens_and_weights = explain(model, data.loc[i], "Title")
draw_html([(tok, weight * 5) for tok, weight in tokens_and_weights], font_style='font-size:20px;');
tokens_and_weights = explain(model, data.loc[i], "FullDescription")
draw_html([(tok, weight * 10) for tok, weight in tokens_and_weights]);
i = 12077
tokens_and_weights = explain(model, data.loc[i], "Title")
draw_html([(tok, weight * 5) for tok, weight in tokens_and_weights], font_style='font-size:20px;');
tokens_and_weights = explain(model, data.loc[i], "FullDescription")
draw_html([(tok, weight * 10) for tok, weight in tokens_and_weights]);
i = np.random.randint(len(data))
print("Index:", i)
print("Salary (gbp):", np.expm1(model.predict(make_batch(data.iloc[i: i+1]))[0, 0]))
tokens_and_weights = explain(model, data.loc[i], "Title")
draw_html([(tok, weight * 5) for tok, weight in tokens_and_weights], font_style='font-size:20px;');
tokens_and_weights = explain(model, data.loc[i], "FullDescription")
draw_html([(tok, weight * 10) for tok, weight in tokens_and_weights]);
```
__Terrible start-up idea #1962:__ make a tool that automaticaly rephrases your job description (or CV) to meet salary expectations :)
| github_jupyter |
# Estimator(Custom)
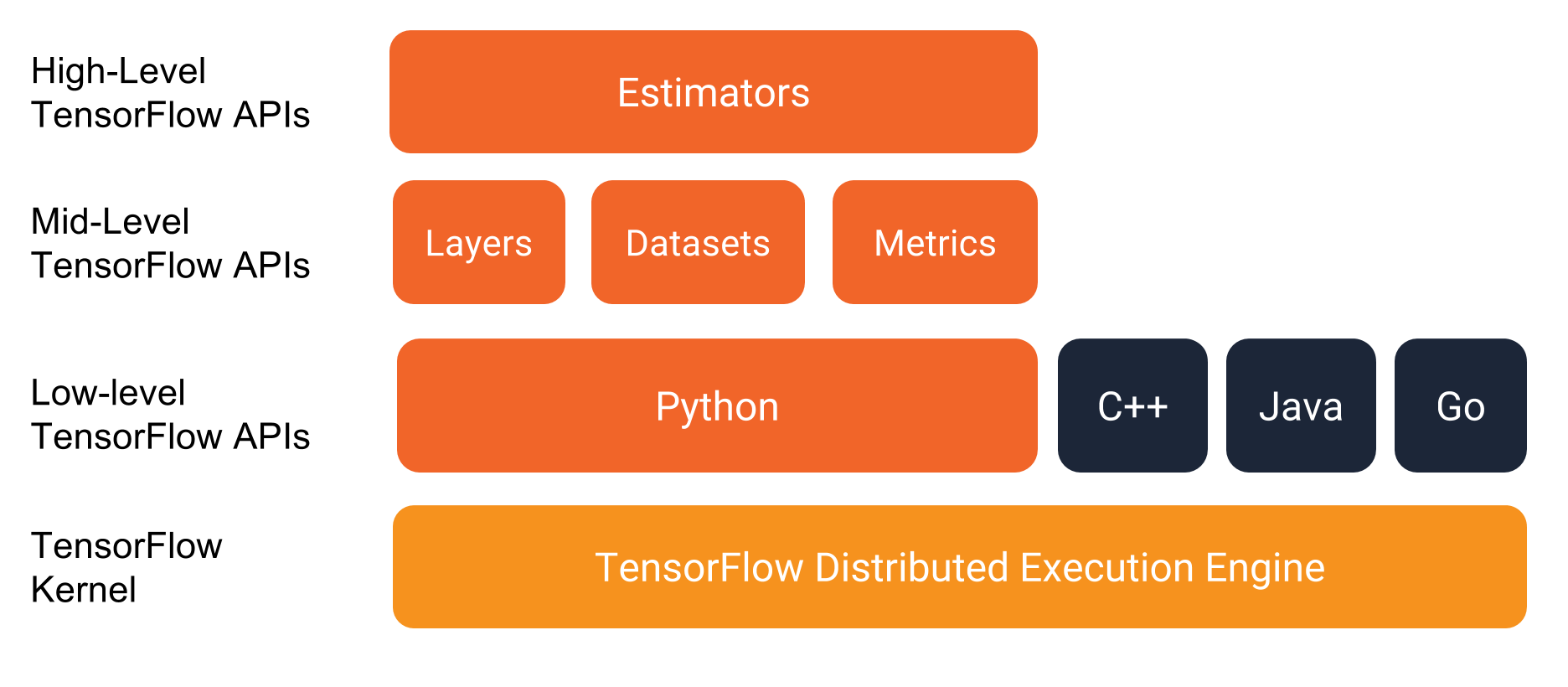
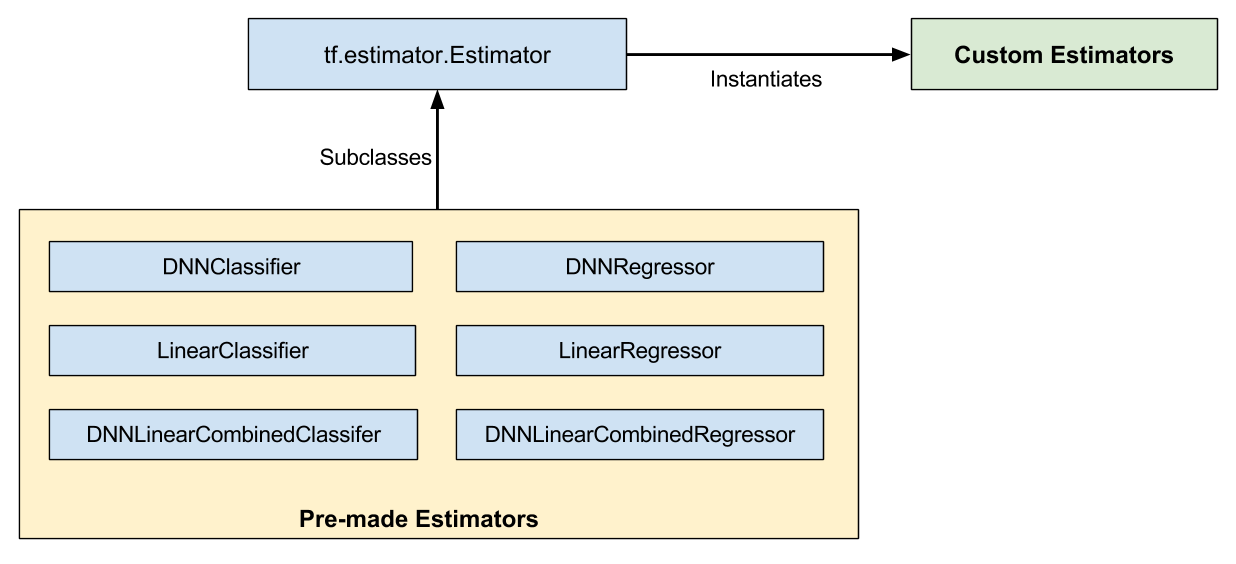
- Tensorflow High Level API
- [Tensorflow 공식 문서](https://www.tensorflow.org/get_started/custom_estimators)
- 미리 정의된 모델(pre-made) 말고도 custom하게 estimator 사용 가능
- tf.Session을 따로 관리할 필요 없으며, ```tf.global_variables_initializer()``` ```tf.local_variables_initializer()```도 필요없음
- 이 글에선 custom estimator에 대해 이야기함
## 구성 요소
- ```input_fn()``` : feature, label return, feature는 dict으로!
- ```model_fn(features, labels, mode)``` : mode별로 분기 => train은 loss, op, evaluate는 pred, accuracy, pred는 prob, class
- ```est = tf.estimator.Estimator(model_fn)```
- ```est.train(input_fn, steps=500)```
- ```est.evaluate(input_fn, steps=10)```
- ```est.predict(pred_input_fn = tf.estimator.inputs.numpy_input_fn({'feature': data}))```
## 참고 자료
- [이찬우님 유튜브](https://www.youtube.com/watch?v=4vJ_2NtsTVg&list=PL1H8jIvbSo1piZJRnp9bIww8Fp2ddIpeR&index=4)
---
```
import tensorflow as tf
import numpy as np
BATCH_SIZE = 100
```
## input_fn
```
def input_fn():
'''
data load하고 feature, label을 return
단, feature는 dict 형식으로 넣어서 predict때도 사용할 수 있도록 함
'''
dataset = tf.data.TextLineDataset("./test_data.csv")\
.batch(2)\
.repeat(999999)\
.make_one_shot_iterator()\
.get_next()
lines = tf.decode_csv(dataset, record_defaults=[[0]]*10)
feature = tf.stack(lines[1:], axis=1)
label = tf.expand_dims(lines[0], axis=-1)
feature = tf.cast(feature, tf.float32)
label = tf.cast(label, tf.float32)
return {'feature': feature}, label
```
## Model
```
def model_fn(features, labels, mode):
'''
mode별로 분기 => train은 loss, op, evaluate는 pred, accuracy
'''
TRAIN = mode == tf.estimator.ModeKeys.TRAIN
EVAL = mode == tf.estimator.ModeKeys.EVAL
PRED = mode == tf.estimator.ModeKeys.PREDICT
layer1 = tf.layers.dense(features["feature"], units=9, activation=tf.nn.relu)
layer2 = tf.layers.dense(layer1, units=9, activation=tf.nn.relu)
layer3 = tf.layers.dense(layer2, units=9, activation=tf.nn.relu)
layer4 = tf.layers.dense(layer3, units=9, activation=tf.nn.relu)
out = tf.layers.dense(layer4, units=1)
if TRAIN:
global_step = tf.train.get_global_step()
loss = tf.losses.sigmoid_cross_entropy(labels, out)
train_op = tf.train.GradientDescentOptimizer(1e-2).minimize(loss, global_step=global_step)
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
elif EVAL:
loss = tf.losses.sigmoid_cross_entropy(labels, out) # test loss
pred = tf.nn.sigmoid(out)
accuracy = tf.metrics.accuracy(labels, tf.round(pred))
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, eval_metric_ops={'acc': accuracy})
elif PRED:
prob = tf.nn.sigmoid(out)
_class = tf.round(prob)
return tf.estimator.EstimatorSpec(mode=mode, predictions={'prob': prob, 'class': _class})
if __name__ == '__main__':
tf.logging.set_verbosity(tf.logging.INFO)
est = tf.estimator.Estimator(model_fn)
est.train(input_fn, steps=500)
est.evaluate(input_fn, steps=10)
data1 = np.array([1,2,3,4,5,6,7,8,9], np.float32)
data2 = np.array([5,5,5,5,5,5,5,5,5], np.float32)
data3 = np.array([9-i for i in range(9)], np.float32)
data = np.stack([data1, data2, data3]) # 여러 데이터 input
pred_input_fn = tf.estimator.inputs.numpy_input_fn({'feature': data}, shuffle=False)
for d, pred in zip(data, est.predict(pred_input_fn)):
print('feature: {}, prob: {}, class: {}'.format(d, pred['prob'], pred['class']))
```
| github_jupyter |
```
import pandas as pd, numpy as np
import matplotlib as mpl, matplotlib.pyplot as plt, seaborn as sns
from tqdm import tqdm_notebook as tqdm
# from help_functions import *
%matplotlib inline
sns.set_style('whitegrid')
mpl.rcParams['savefig.dpi'] = 300
```
# Helper Functions
```
def pd_tickround(df, tick_size=1):
return np.round(df.astype(float).divide(tick_size)).multiply(tick_size).astype(int)
def tickround(x, tick_size=5):
return int(tick_size * round(float(x)/tick_size))
def calc_n_ticks(side, inverse, target_price, tick_size, maker_fee, taker_fee):
if side == 'long':
if inverse:
n_ticks = -target_price * (taker_fee - maker_fee) / (tick_size * (taker_fee - maker_fee - 1)) + 1
else:
n_ticks = target_price * (taker_fee - maker_fee) / tick_size + 1
elif side == 'short':
if inverse:
n_ticks = (target_price/tick_size) * (1 / (maker_fee - taker_fee -1) + 1) + 1
else:
n_ticks = target_price * (taker_fee - maker_fee) / tick_size + 1
return abs(n_ticks)
def calc_exec_price(side, inverse, target_price, tick_size, maker_fee, taker_fee, thresh=5):
n_ticks = calc_n_ticks(side, inverse, target_price, tick_size, maker_fee, taker_fee)
n_ticks = np.max([0, n_ticks-thresh])
if side == 'long':
exec_price = target_price + n_ticks * tick_size
elif side == 'short':
exec_price = target_price - n_ticks * tick_size
return tickround(exec_price, tick_size)
```
# Load Data
```
settings_path = 'data/symbol_settings.csv'
symbol_settings = pd.read_csv(settings_path, index_col=0)
symbol_settings
```
# Bitmex-Specific Idiosyncracies
## Plot Break-Evens
The code in this section isn't the cleanest; you're gonna have to forgive me on that one
```
symbol = 'XBTUSD'
# Load symbol-specific parameters
ind = symbol_settings.loc[symbol]
side = 'long'
inverse = ind['inverse']
tick_size = ind['tick_size']
maker_fee = ind['maker_fee']
taker_fee = ind['taker_fee']
target_price = ind['target_price']
lo_ticks = 0
hi_ticks = 30
ticks = np.arange(-lo_ticks, hi_ticks)
prices = pd_tickround(pd.Series(target_price + ticks*tick_size, index=ticks))
# Calculate passive costs
if side == 'long':
if inverse:
limit_fees = 1 - target_price/(prices-tick_size) + maker_fee
else:
limit_fees = (prices-tick_size)/target_price - 1 + maker_fee
elif side == 'short':
if inverse:
limit_fees = maker_fee - (1 - target_price/(prices+tick_size))
else:
limit_fees = maker_fee - (prices+tick_size)/target_price + 1
n_ticks = calc_n_ticks(side=side, inverse=inverse, maker_fee=maker_fee, taker_fee=taker_fee, target_price=target_price, tick_size=tick_size)
break_even = target_price + n_ticks * tick_size
# Plot Break-Evens
fig, ax = plt.subplots(figsize=(6,4))
ax.plot(prices, limit_fees)
ax.axvline(target_price, c='k', lw=0.5)
ax.axhline(0, c='k', lw=0.5)
ax.axhline(taker_fee, c='r', lw=0.5, linestyle=':')
ax.scatter(break_even, taker_fee, marker='o', color='r')
ax.annotate('# ticks: {:.3}'.format(n_ticks), xy=(break_even, taker_fee),
xytext=(break_even*0.9996, taker_fee*1.25),
arrowprops=dict(facecolor='black', shrink=0.05),
)
ax.set_xticks(ax.get_xticks()[::1])
xvals = ax.get_xticks()
ax.set_xticklabels([int(x/100) for x in xvals])
yvals = ax.get_yticks()
ax.set_yticklabels(['{:,.2%}'.format(x) for x in yvals])
ax.set_xlabel('Price (USD)')
ax.set_ylabel('Total Costs (% of Order Value)')
ax.set_title('XBTUSD Break-Evens', fontsize=13)
fig.tight_layout()
fig.savefig('XBTUSD Break-Evens')
symbol = 'TRXXBT'
# Load symbol-specific parameters
ind = symbol_settings.loc[symbol]
side = 'long'
inverse = ind['inverse']
tick_size = ind['tick_size']
maker_fee = ind['maker_fee']
taker_fee = ind['taker_fee']
target_price = ind['target_price']
lo_ticks = 0
hi_ticks = 5
ticks = np.arange(-lo_ticks, hi_ticks)
prices = pd_tickround(pd.Series(target_price + ticks*tick_size, index=ticks))
# Calculate passive costs
if side == 'long':
if inverse:
limit_fees = 1 - target_price/(prices-tick_size) + maker_fee
else:
limit_fees = (prices-tick_size)/target_price - 1 + maker_fee
elif side == 'short':
if inverse:
limit_fees = maker_fee - (1 - target_price/(prices+tick_size))
else:
limit_fees = maker_fee - (prices+tick_size)/target_price + 1
n_ticks = calc_n_ticks(side=side, inverse=inverse, maker_fee=maker_fee, taker_fee=taker_fee, target_price=target_price, tick_size=tick_size)
break_even = target_price + n_ticks * tick_size
# Plot Break-Evens
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(prices, limit_fees)
ax.axvline(target_price, c='k', lw=0.5)
ax.axhline(0, c='k', lw=0.5)
ax.axhline(taker_fee, c='r', lw=0.5, linestyle=':')
ax.scatter(break_even, taker_fee, marker='o', color='r')
# ax.annotate('# ticks: {}'.format(n_ticks), xy=(2,1))
ax.annotate('# ticks: {:,.2}'.format(n_ticks), xy=(break_even, taker_fee),
xytext=(break_even*.993, taker_fee*1.55),
arrowprops=dict(facecolor='black', shrink=0.05),
)
ax.set_xlim(166, 172)
ax.set_xticks(ax.get_xticks()[::1])
yvals = ax.get_yticks()
ax.set_yticklabels(['{:,.2%}'.format(x) for x in yvals])
ax.set_xlabel('Price (satoshis)')
ax.set_ylabel('Total Costs (% of Order Value)')
ax.set_title('TRXU19 Cost Break-Evens', fontsize=13)
fig.tight_layout()
fig.savefig('TRXU19 Break-Evens')
```
## Calculate Break-Evens for all symbols
```
symbols = symbol_settings.index
t = tqdm(symbols, leave=True)
for side in ['long', 'short']:
print(side)
for symbol in t:
t.set_description(symbol)
t.refresh()
ind = symbol_settings.loc[symbol] # individual settings
inverse = ind['inverse']
tick_size = ind['tick_size']
maker_fee = ind['maker_fee']
taker_fee = ind['taker_fee']
target_price = ind['target_price']
n_ticks = calc_n_ticks(side=side, inverse=inverse, maker_fee=maker_fee, taker_fee=taker_fee, target_price=target_price, tick_size=tick_size)
exec_price = calc_exec_price(side=side, inverse=inverse, maker_fee=maker_fee, taker_fee=taker_fee, target_price=target_price, tick_size=tick_size)
print(symbol, int(n_ticks), target_price, exec_price)
```
# Estimating Timing Risk
This is going to take a while.. go grab a cup of coffee
```
n_trials = 50000
auto_correlations = [0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65]
results = pd.DataFrame(columns=auto_correlations, index=np.arange(n_trials))
for auto_correlation in tqdm(auto_correlations):
for trial in range(n_trials):
# print('\n Trial #{}'.format(trial+1))
# Reset the count
down_ticks = 0
up_ticks = 0
total_excursion = 0 # tracks the total drift away from the start
# Choose the first tick randomly
tick = np.random.randint(0,2)
while down_ticks < 2 and total_excursion < 25:
# Set probability of next tick based on prev tick
prob_up = auto_correlation if tick > 0 else (1 - auto_correlation)
prob_down = 1 - prob_up
tick = np.random.choice([0,1], p=[prob_down, prob_up])
# print('tick: {}1'.format('+' if tick>0 else '-'))
# downtick
if tick == 0:
down_ticks += 1
total_excursion -= 1
elif tick == 1:
# up_ticks += 1
down_ticks = 0 # reset the count; looking for 2 consecutive
total_excursion += 1
fill = total_excursion + 1
# print('Down Ticks: {}, Total Excursion: {}'.format(down_ticks, total_excursion))
# print('Fill: {}'.format(fill))
results.loc[trial, auto_correlation] = fill
fig, ax = plt.subplots(figsize=(10,7))
for col in results.columns:
(results[col].value_counts() / len(results)).sort_index().cumsum().plot(label=col, ax=ax)
plt.legend(title='Auto-Cor')
plt.axhline(0.9, c='k', lw=0.55)
# plt.xticks(np)
plt.xlim(-2, 5)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Fishbone-Moncrief Initial Data
## Author: Zach Etienne
### Formatting improvements courtesy Brandon Clark
[comment]: <> (Abstract: TODO)
[comment]: <> (Notebook Status and Validation Notes: TODO)
### NRPy+ Source Code for this module: [FishboneMoncriefID/FishboneMoncriefID.py](../edit/FishboneMoncriefID/FishboneMoncriefID.py)
## Introduction:
This goal of this module will be to construct Fishbone-Moncrief initial data for GRMHD simulations in a format suitable for the Einstein Toolkit (ETK). We will be using the equations as derived in [the original paper](http://articles.adsabs.harvard.edu/cgi-bin/nph-iarticle_query?1976ApJ...207..962F&data_type=PDF_HIGH&whole_paper=YES&type=PRINTER&filetype=.pdf), which will hereafter be called "***the FM paper***". Since we want to use this with the ETK, our final result will be in Cartesian coordinates. The natural coordinate system for these data is spherical, however, so we will use [reference_metric.py](../edit/reference_metric.py) ([**Tutorial**](Tutorial-Reference_Metric.ipynb)) to help with the coordinate transformation.
This notebook documents the equations in the NRPy+ module [FishboneMoncrief.py](../edit/FishboneMoncriefID/FishboneMoncriefID.py). Then, we will build an Einstein Toolkit [thorn](Tutorial-ETK_thorn-FishboneMoncriefID.ipynb) to set this initial data.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Initialize core Python/NRPy+ modules
1. [Step 2](#fishbonemoncrief): Implementing Fishbone-Moncrief initial data within NRPy+
1. [Step 2.a](#registergridfunctions): Register within NRPy+ needed gridfunctions and initial parameters
1. [Step 2.b](#l_of_r): Specific angular momentum $l(r)$
1. [Step 2.c](#enthalpy): Specific enthalpy $h$
1. [Step 2.d](#pressure_density): Pressure and density, from the specific enthalpy
1. [Step 2.e](#covariant_velocity): Nonzero covariant velocity components $u_\mu$
1. [Step 2.f](#inverse_bl_metric): Inverse metric $g^{\mu\nu}$ for the black hole in Boyer-Lindquist coordinates
1. [Step 2.g](#xform_to_ks): Transform components of four-veloicty $u^\mu$ to Kerr-Schild
1. [Step 2.h](#ks_metric): Define Kerr-Schild metric $g_{\mu\nu}$ and extrinsic curvature $K_{ij}$
1. [Step 2.i](#magnetic_field): Seed poloidal magnetic field $B^i$
1. [Step 2.j](#adm_metric): Set the ADM quantities $\alpha$, $\beta^i$, and $\gamma_{ij}$ from the spacetime metric $g_{\mu\nu}$
1. [Step 2.k](#magnetic_field_comoving_frame): Set the magnetic field components in the comoving frame $b^\mu$, and $b^2$, which is twice the magnetic pressure
1. [Step 2.l](#lorentz_fac_valencia): Lorentz factor $\Gamma = \alpha u^0$ and Valencia 3-velocity $v^i_{(n)}$
1. [Step 3](#output_to_c): Output SymPy expressions to C code, using NRPy+
1. [Step 4](#code_validation): Code Validation against Code Validation against `FishboneMoncriefID.FishboneMoncriefID` NRPy+ module NRPy+ module
1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Initialize core Python/NRPy+ modules \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
We begin by importing the packages and NRPy+ modules that we will need. We will also set some of the most commonly used parameters.
```
# Step 1a: Import needed NRPy+ core modules:
import NRPy_param_funcs as par
import indexedexp as ixp
import grid as gri
import finite_difference as fin
from outputC import *
import loop
import reference_metric as rfm
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
#Set the spatial dimension parameter to 3.
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
thismodule = "FishboneMoncriefID"
```
<a id='fishbonemoncrief'></a>
# Step 2: The Fishbone-Moncrief Initial Data Prescription \[Back to [top](#toc)\]
$$\label{fishbonemoncrief}$$
With NRPy's most important functions now available to us, we can start to set up the rest of the tools we will need to build the initial data.
<a id='registergridfunctions'></a>
## Step 2.a: Register within NRPy+ needed gridfunctions and initial parameters \[Back to [top](#toc)\]
$$\label{registergridfunctions}$$
We will now register the gridfunctions we expect to use. Critically, we register the physical metric and extrinsic curvature tensors.
```
gPhys4UU = ixp.register_gridfunctions_for_single_rank2("AUX","gPhys4UU", "sym01", DIM=4)
KDD = ixp.register_gridfunctions_for_single_rank2("EVOL","KDD", "sym01")
# Variables needed for initial data given in spherical basis
r, th, ph = gri.register_gridfunctions("AUX",["r","th","ph"])
r_in,r_at_max_density,a,M = par.Cparameters("REAL",thismodule,
["r_in","r_at_max_density", "a","M"],
[ 6.0, 12.0, 0.9375,1.0])
kappa,gamma = par.Cparameters("REAL",thismodule,["kappa","gamma"], [1.0e-3, 4.0/3.0])
LorentzFactor = gri.register_gridfunctions("AUX","LorentzFactor")
```
<a id='l_of_r'></a>
## Step 2.b: Specific angular momentum $l(r)$ \[Back to [top](#toc)\]
$$\label{l_of_r}$$
Now, we can begin actually building the ID equations. We will start with the value of the angular momentum $l$ at the position $r \equiv$`r_at_max_density` where the density is at a maximum, as in equation 3.8 of the FM paper:
\begin{align}
l(r) &= \pm \left( \frac{M}{r^3} \right) ^{1/2}
\left[ \frac{r^4+r^2a^2-2Mra^2 \mp a(Mr)^{1/2}(r^2-a^2)}
{r^2 -3Mr \pm 2a(Mr)^{1/2}} \right].
\end{align}
```
def calculate_l_at_r(r):
l = sp.sqrt(M/r**3) * (r**4 + r**2*a**2 - 2*M*r*a**2 - a*sp.sqrt(M*r)*(r**2-a**2))
l /= r**2 - 3*M*r + 2*a*sp.sqrt(M*r)
return l
# First compute angular momentum at r_at_max_density, TAKING POSITIVE ROOT. This way disk is co-rotating with black hole
# Eq 3.8:
l = calculate_l_at_r(r_at_max_density)
```
<a id='enthalpy'></a>
## Step 2.c: Specific enthalpy $h$ \[Back to [top](#toc)\]
$$\label{enthalpy}$$
Next, we will follow equation 3.6 of the FM paper to compute the enthalpy $h$ by first finding its logarithm $\ln h$. Fortunately, we can make this process quite a bit simpler by first identifying the common subexpressions. Let
\begin{align}
\Delta &= r^2 - 2Mr + a^2 \\
\Sigma &= r^2 + a^2 \cos^2 (\theta) \\
A &= (r^2+a^2)^2 - \Delta a^2 \sin^2(\theta);
\end{align}
furthermore, let
\begin{align}
\text{tmp3} &= \sqrt{\frac{1 + 4 l^2 \Sigma^2 \Delta}{A \sin^2 (\theta)}}. \\
\end{align}
(These terms reflect the radially-independent part of the log of the enthalpy, `ln_h_const`.)
So,
$$
{\rm ln\_h\_const} = \frac{1}{2} * \log \left( \frac{1+\text{tmp3}}{\Sigma \Delta/A} \right) - \frac{1}{2} \text{tmp3} - \frac{2aMrl}{A}
$$
```
# Eq 3.6:
# First compute the radially-independent part of the log of the enthalpy, ln_h_const
Delta = r**2 - 2*M*r + a**2
Sigma = r**2 + a**2*sp.cos(th)**2
A = (r**2 + a**2)**2 - Delta*a**2*sp.sin(th)**2
# Next compute the radially-dependent part of log(enthalpy), ln_h
tmp3 = sp.sqrt(1 + 4*l**2*Sigma**2*Delta/(A*sp.sin(th))**2)
# Term 1 of Eq 3.6
ln_h = sp.Rational(1,2)*sp.log( ( 1 + tmp3) / (Sigma*Delta/A))
# Term 2 of Eq 3.6
ln_h -= sp.Rational(1,2)*tmp3
# Term 3 of Eq 3.6
ln_h -= 2*a*M*r*l/A
```
Additionally, let
\begin{align}
\Delta_{\rm in} &= r_{\rm in}^2 - 2Mr_{\rm in} + a^2 \\
\Sigma_{\rm in} &= r_{\rm in}^2 + a^2 \cos^2 (\pi/2) \\
A_{\rm in} &= (r_{\rm in}^2+a^2)^2 - \Delta_{\rm in} a^2 \sin^2(\pi/2)
\end{align}
and
\begin{align}
\text{tmp3in} &= \sqrt{\frac{1 + 4 l^2 \Sigma_{\rm in}^2 \Delta_{\rm in}}{A_{\rm in} \sin^2 (\theta)}}, \\
\end{align}
corresponding to the radially Independent part of log(enthalpy), $\ln h$:
\begin{align}
{\rm mln\_h\_in} = -\frac{1}{2} * \log \left( \frac{1+\text{tmp3in}}{\Sigma_{\rm in} \Delta_{\rm in}/A_{\rm in}} \right) + \frac{1}{2} \text{tmp3in} + \frac{2aMr_{\rm in}l}{A_{\rm in}}. \\
\end{align}
(Note that there is some typo in the expression for these terms given in Eq 3.6, so we opt to just evaluate negative of the first three terms at r=`r_in` and th=pi/2 (the integration constant), as described in the text below Eq. 3.6.)
So, then, we exponentiate:
\begin{align}
\text{hm1} \equiv h-1 &= e^{{\rm ln\_h}+{\rm mln\_h\_in}}-1. \\
\end{align}
```
# Next compute the radially-INdependent part of log(enthalpy), ln_h
# Note that there is some typo in the expression for these terms given in Eq 3.6, so we opt to just evaluate
# negative of the first three terms at r=r_in and th=pi/2 (the integration constant), as described in
# the text below Eq. 3.6, basically just copying the above lines of code.
# Delin = Delta_in ; Sigin = Sigma_in ; Ain = A_in .
Delin = r_in**2 - 2*M*r_in + a**2
Sigin = r_in**2 + a**2*sp.cos(sp.pi/2)**2
Ain = (r_in**2 + a**2)**2 - Delin*a**2*sp.sin(sp.pi/2)**2
tmp3in = sp.sqrt(1 + 4*l**2*Sigin**2*Delin/(Ain*sp.sin(sp.pi/2))**2)
# Term 4 of Eq 3.6
mln_h_in = -sp.Rational(1,2)*sp.log( ( 1 + tmp3in) / (Sigin*Delin/Ain))
# Term 5 of Eq 3.6
mln_h_in += sp.Rational(1,2)*tmp3in
# Term 6 of Eq 3.6
mln_h_in += 2*a*M*r_in*l/Ain
hm1 = sp.exp(ln_h + mln_h_in) - 1
```
<a id='pressure_density'></a>
## Step 2.d: Pressure and density, from the specific enthalpy \[Back to [top](#toc)\]
$$\label{pressure_density}$$
Python 3.4 + SymPy 1.0.0 has a serious problem taking the power here; it hangs forever, so instead we use the identity $x^{1/y} = \exp(\frac{1}{y} * \log(x))$. Thus, our expression for density becomes (in Python 2.7 + SymPy 0.7.4.1):
\begin{align}
\rho_0 &= \left( \frac{(h-1)(\gamma-1)}{\kappa \gamma} \right)^{1/(\gamma-1)} \\
&= \exp \left[ {\frac{1}{\gamma-1} \log \left( \frac{(h-1)(\gamma-1)}{\kappa \gamma}\right)} \right]
\end{align}
Additionally, the pressure $P_0 = \kappa \rho_0^\gamma$
```
rho_initial,Pressure_initial = gri.register_gridfunctions("AUX",["rho_initial","Pressure_initial"])
# Python 3.4 + sympy 1.0.0 has a serious problem taking the power here, hangs forever.
# so instead we use the identity x^{1/y} = exp( [1/y] * log(x) )
# Original expression (works with Python 2.7 + sympy 0.7.4.1):
# rho_initial = ( hm1*(gamma-1)/(kappa*gamma) )**(1/(gamma - 1))
# New expression (workaround):
rho_initial = sp.exp( (1/(gamma-1)) * sp.log( hm1*(gamma-1)/(kappa*gamma) ))
Pressure_initial = kappa * rho_initial**gamma
```
<a id='covariant_velocity'></a>
## Step 2.e: Nonzero covariant velocity components $u_\mu$ \[Back to [top](#toc)\]
$$\label{covariant_velocity}$$
We now want to compute eq 3.3; we will start by finding $e^{-2 \chi}$ in Boyer-Lindquist (BL) coordinates. By eq 2.16, $\chi = \psi - \nu$, so, by eqs. 3.5,
\begin{align}
e^{2 \nu} &= \frac{\Sigma \Delta}{A} \\
e^{2 \psi} &= \frac{A \sin^2 \theta}{\Sigma} \\
e^{-2 \chi} &= e^{2 \nu} / e^{2 \psi} = e^{2(\nu - \psi)}.
\end{align}
Next, we will calculate the 4-velocity $u_i$ of the fluid disk in BL coordinates. We start with eqs. 3.3 and 2.13, finding
\begin{align}
u_{(r)} = u_{(\theta)} &= 0 \\
u_{(\phi)} &= \sqrt{-1+ \frac{1}{2}\sqrt{1 + 4l^2e^{-2 \chi}}} \\
u_{(t)} &= - \sqrt{1 + u_{(\phi)}^2}.
\end{align}
Given that $\omega = 2aMr/A$, we then find that, in BL coordinates,
\begin{align}
u_r = u_{\theta} &= 0 \\
u_{\phi} &= u_{(\phi)} \sqrt{e^{2 \psi}} \\
u_t &= u_{(t)} \sqrt{e^{2 \nu}} - \omega u_{\phi},
\end{align}
using eq. 2.13 to get the last relation.
```
# Eq 3.3: First compute exp(-2 chi), assuming Boyer-Lindquist coordinates
# Eq 2.16: chi = psi - nu, so
# Eq 3.5 -> exp(-2 chi) = exp(-2 (psi - nu)) = exp(2 nu)/exp(2 psi)
exp2nu = Sigma*Delta / A
exp2psi = A*sp.sin(th)**2 / Sigma
expm2chi = exp2nu / exp2psi
# Eq 3.3: Next compute u_(phi).
u_pphip = sp.sqrt((-1 + sp.sqrt(1 + 4*l**2*expm2chi))/2)
# Eq 2.13: Compute u_(t)
u_ptp = -sp.sqrt(1 + u_pphip**2)
# Next compute spatial components of 4-velocity in Boyer-Lindquist coordinates:
uBL4D = ixp.zerorank1(DIM=4) # Components 1 and 2: u_r = u_theta = 0
# Eq 2.12 (typo): u_(phi) = e^(-psi) u_phi -> u_phi = e^(psi) u_(phi)
uBL4D[3] = sp.sqrt(exp2psi)*u_pphip
# Assumes Boyer-Lindquist coordinates:
omega = 2*a*M*r/A
# Eq 2.13: u_(t) = 1/sqrt(exp2nu) * ( u_t + omega*u_phi )
# --> u_t = u_(t) * sqrt(exp2nu) - omega*u_phi
# --> u_t = u_ptp * sqrt(exp2nu) - omega*uBL4D[3]
uBL4D[0] = u_ptp*sp.sqrt(exp2nu) - omega*uBL4D[3]
```
<a id='inverse_bl_metric'></a>
## Step 2.f: Inverse metric $g^{\mu\nu}$ for the black hole in Boyer-Lindquist coordinates \[Back to [top](#toc)\]
$$\label{inverse_bl_metric}$$
Next, we will use eq. 2.1 to find the inverse physical (as opposed to conformal) metric in BL coordinates, using the shorthands defined in eq. 3.5:
\begin{align}
g_{tt} &= - \frac{\Sigma \Delta}{A} + \omega^2 \sin^2 \theta \frac{A}{\Sigma} \\
g_{t \phi} = g_{\phi t} &= - \omega \sin^2 \theta \frac{A}{\Sigma} \\
g_{\phi \phi} &= \sin^2 \theta \frac{A}{\Sigma},
\end{align}
which can be inverted to show that
\begin{align}
g^{tt} &= - \frac{A}{\Delta \Sigma} \\
g^{t \phi} = g^{\phi t} &= \frac{2aMr}{\Delta \Sigma} \\
g^{\phi \phi} &= - \frac{4a^2M^2r^2}{\Delta A \Sigma} + \frac{\Sigma^2}{A \Sigma \sin^2 \theta}.
\end{align}
With this, we will now be able to raise the index on the BL $u_i$: $u^i = g^{ij} u_j$
```
# Eq. 3.5:
# w = 2*a*M*r/A;
# Eqs. 3.5 & 2.1:
# gtt = -Sig*Del/A + w^2*Sin[th]^2*A/Sig;
# gtp = w*Sin[th]^2*A/Sig;
# gpp = Sin[th]^2*A/Sig;
# FullSimplify[Inverse[{{gtt,gtp},{gtp,gpp}}]]
gPhys4BLUU = ixp.zerorank2(DIM=4)
gPhys4BLUU[0][0] = -A/(Delta*Sigma)
# DO NOT NEED TO SET gPhys4BLUU[1][1] or gPhys4BLUU[2][2]!
gPhys4BLUU[0][3] = gPhys4BLUU[3][0] = -2*a*M*r/(Delta*Sigma)
gPhys4BLUU[3][3] = -4*a**2*M**2*r**2/(Delta*A*Sigma) + Sigma**2/(A*Sigma*sp.sin(th)**2)
uBL4U = ixp.zerorank1(DIM=4)
for i in range(4):
for j in range(4):
uBL4U[i] += gPhys4BLUU[i][j]*uBL4D[j]
```
<a id='xform_to_ks'></a>
## Step 2.g: Transform components of four-velocity $u^\mu$ to Kerr-Schild \[Back to [top](#toc)\]
$$\label{xform_to_ks}$$
Now, we will transform the 4-velocity from the Boyer-Lindquist to the Kerr-Schild basis. This algorithm is adapted from [HARM](https://github.com/atchekho/harmpi/blob/master/init.c). This definees the tensor `transformBLtoKS`, where the diagonal elements are $1$, and the non-zero off-diagonal elements are
\begin{align}
\text{transformBLtoKS}_{tr} &= \frac{2r}{r^2-2r+a^2} \\
\text{transformBLtoKS}_{\phi r} &= \frac{a}{r^2-2r+a^2} \\
\end{align}
```
# https://github.com/atchekho/harmpi/blob/master/init.c
# Next transform Boyer-Lindquist velocity to Kerr-Schild basis:
transformBLtoKS = ixp.zerorank2(DIM=4)
for i in range(4):
transformBLtoKS[i][i] = 1
transformBLtoKS[0][1] = 2*r/(r**2 - 2*r + a*a)
transformBLtoKS[3][1] = a/(r**2 - 2*r + a*a)
#uBL4U = ixp.declarerank1("UBL4U",DIM=4)
# After the xform below, print(uKS4U) outputs:
# [UBL4U0 + 2*UBL4U1*r/(a**2 + r**2 - 2*r), UBL4U1, UBL4U2, UBL4U1*a/(a**2 + r**2 - 2*r) + UBL4U3]
uKS4U = ixp.zerorank1(DIM=4)
for i in range(4):
for j in range(4):
uKS4U[i] += transformBLtoKS[i][j]*uBL4U[j]
```
<a id='ks_metric'></a>
## Step 2.h: Define Kerr-Schild metric $g_{\mu\nu}$ and extrinsic curvature $K_{ij}$ \[Back to [top](#toc)\]
$$\label{ks_metric}$$
We will also adopt the Kerr-Schild metric for Fishbone-Moncrief disks. Further details can be found in [Cook's Living Review](http://gravity.psu.edu/numrel/jclub/jc/Cook___LivRev_2000-5.pdf) article on initial data, or in the appendix of [this](https://arxiv.org/pdf/1704.00599.pdf) article. So, in KS coordinates,
\begin{align}
\rho^2 &= r^2 + a^2 \cos^2 \theta \\
\Delta &= r^2 - 2Mr + a^2 \\
\alpha &= \left(1 + \frac{2Mr}{\rho^2}\right)^{-2} \\
\beta^0 &= \frac{2 \alpha^2 Mr}{\rho^2} \\
\gamma_{00} &= 1 + \frac{2Mr}{\rho^2} \\
\gamma_{02} = \gamma_{20} &= -\left(1+\frac{2Mr}{\rho^2}\right) a \sin^2 \theta \\
\gamma_{11} &= \rho^2 \\
\gamma_{22} &= \left(r^2+a^2+\frac{2Mr}{\rho^2} a^2 \sin^2 \theta\right) \sin^2 \theta.
\end{align}
(Note that only the non-zero components of $\beta^i$ and $\gamma_{ij}$ are defined here.)
```
# Adopt the Kerr-Schild metric for Fishbone-Moncrief disks
# http://gravity.psu.edu/numrel/jclub/jc/Cook___LivRev_2000-5.pdf
# Alternatively, Appendix of https://arxiv.org/pdf/1704.00599.pdf
rhoKS2 = r**2 + a**2*sp.cos(th)**2 # Eq 79 of Cook's Living Review article
DeltaKS = r**2 - 2*M*r + a**2 # Eq 79 of Cook's Living Review article
alphaKS = 1/sp.sqrt(1 + 2*M*r/rhoKS2)
betaKSU = ixp.zerorank1()
betaKSU[0] = alphaKS**2*2*M*r/rhoKS2
gammaKSDD = ixp.zerorank2()
gammaKSDD[0][0] = 1 + 2*M*r/rhoKS2
gammaKSDD[0][2] = gammaKSDD[2][0] = -(1 + 2*M*r/rhoKS2)*a*sp.sin(th)**2
gammaKSDD[1][1] = rhoKS2
gammaKSDD[2][2] = (r**2 + a**2 + 2*M*r/rhoKS2 * a**2*sp.sin(th)**2) * sp.sin(th)**2
```
We can also define the following useful quantities, continuing in KS coordinates:
\begin{align}
A &= a^2 \cos (2 \theta) + a^2 +2r^2 \\
B &= A + 4Mr \\
D &= \sqrt{\frac{2Mr}{a^2 \cos^2 \theta +r^2}+1};
\end{align}
we will also define the extrinsic curvature:
\begin{align}
K_{00} &= D\frac{A+2Mr}{A^2 B} (4M(a^2 \cos(2 \theta)+a^s-2r^2)) \\
K_{01} = K_{10} &= \frac{D}{AB} (8a^2Mr\sin \theta \cos \theta) \\
K_{02} = K_{20} &= \frac{D}{A^2} (-2aM \sin^2 \theta (a^2\cos(2 \theta)+a^2-2r^2)) \\
K_{11} &= \frac{D}{B} (4Mr^2) \\
K_{12} = K_{21} &= \frac{D}{AB} (-8a^3Mr \sin^3 \theta \cos \theta) \\
K_{22} &= \frac{D}{A^2 B} (2Mr \sin^2 \theta (a^4(r-M) \cos(4 \theta) + a^4 (M+3r) + 4a^2 r^2 (2r-M) + 4a^2 r \cos(2 \theta) (a^2 + r(M+2r)) + 8r^5)). \\
\end{align}
Note that the indexing for extrinsic curvature only runs from 0 to 2, since there are no time components to the tensor.
```
AA = a**2 * sp.cos(2*th) + a**2 + 2*r**2
BB = AA + 4*M*r
DD = sp.sqrt(2*M*r / (a**2 * sp.cos(th)**2 + r**2) + 1)
KDD[0][0] = DD*(AA + 2*M*r)/(AA**2*BB) * (4*M*(a**2 * sp.cos(2*th) + a**2 - 2*r**2))
KDD[0][1] = KDD[1][0] = DD/(AA*BB) * 8*a**2*M*r*sp.sin(th)*sp.cos(th)
KDD[0][2] = KDD[2][0] = DD/AA**2 * (-2*a*M*sp.sin(th)**2 * (a**2 * sp.cos(2*th) + a**2 - 2*r**2))
KDD[1][1] = DD/BB * 4*M*r**2
KDD[1][2] = KDD[2][1] = DD/(AA*BB) * (-8*a**3*M*r*sp.sin(th)**3*sp.cos(th))
KDD[2][2] = DD/(AA**2*BB) * \
(2*M*r*sp.sin(th)**2 * (a**4*(r-M)*sp.cos(4*th) + a**4*(M+3*r) +
4*a**2*r**2*(2*r-M) + 4*a**2*r*sp.cos(2*th)*(a**2 + r*(M+2*r)) + 8*r**5))
```
We must also compute the inverse and determinant of the KS metric. We can use the NRPy+ [indexedexp.py](../edit/indexedexp.py) function to do this easily for the inverse physical 3-metric $\gamma^{ij}$, and then use the lapse $\alpha$ and the shift $\beta^i$ to find the full, inverse 4-dimensional metric, $g^{ij}$. We use the general form relating the 3- and 4- metric from (B&S 2.122)
\begin{equation}
g_{\mu\nu} = \begin{pmatrix}
-\alpha^2 + \beta\cdot\beta & \beta_i \\
\beta_j & \gamma_{ij}
\end{pmatrix},
\end{equation}
and invert it. That is,
\begin{align}
g^{00} &= -\frac{1}{\alpha^2} \\
g^{0i} = g^{i0} &= \frac{\beta^{i-1}}{\alpha^2} \\
g^{ij} = g^{ji} &= \gamma^{(i-1) (j-1)} - \frac{\beta^{i-1} \beta^{j-1}}{\alpha^2},
\end{align}
keeping careful track of the differences in the indexing conventions for 3-dimensional quantities and 4-dimensional quantities (Python always indexes lists from 0, but in four dimensions, the 0 direction corresponds to time, while in 3+1, the connection to time is handled by other variables).
```
# For compatibility, we must compute gPhys4UU
gammaKSUU,gammaKSDET = ixp.symm_matrix_inverter3x3(gammaKSDD)
# See, e.g., Eq. 4.49 of https://arxiv.org/pdf/gr-qc/0703035.pdf , where N = alpha
gPhys4UU[0][0] = -1 / alphaKS**2
for i in range(1,4):
if i>0:
# if the quantity does not have a "4", then it is assumed to be a 3D quantity.
# E.g., betaKSU[] is a spatial vector, with indices ranging from 0 to 2:
gPhys4UU[0][i] = gPhys4UU[i][0] = betaKSU[i-1]/alphaKS**2
for i in range(1,4):
for j in range(1,4):
# if the quantity does not have a "4", then it is assumed to be a 3D quantity.
# E.g., betaKSU[] is a spatial vector, with indices ranging from 0 to 2,
# and gammaKSUU[][] is a spatial tensor, with indices again ranging from 0 to 2.
gPhys4UU[i][j] = gPhys4UU[j][i] = gammaKSUU[i-1][j-1] - betaKSU[i-1]*betaKSU[j-1]/alphaKS**2
```
<a id='magnetic_field'></a>
## Step 2.i: Seed poloidal magnetic field $B^i$ \[Back to [top](#toc)\]
$$\label{magnetic_field}$$
The original Fishbone-Moncrief initial data prescription describes a non-self-gravitating accretion disk in hydrodynamical equilibrium about a black hole. The following assumes that a very weak magnetic field seeded into this disk will not significantly disturb this equilibrium, at least on a dynamical (free-fall) timescale.
Now, we will set up the magnetic field that, when simulated with a GRMHD code, will give us insight into the electromagnetic emission from the disk. We define the vector potential $A_i$ to be proportional to $\rho_0$, and, as usual, let the magnetic field $B^i$ be the curl of the vector potential.
```
A_b = par.Cparameters("REAL",thismodule,"A_b",1.0)
A_3vecpotentialD = ixp.zerorank1()
# Set A_phi = A_b*rho_initial FIXME: why is there a sign error?
A_3vecpotentialD[2] = -A_b * rho_initial
BtildeU = ixp.register_gridfunctions_for_single_rank1("EVOL","BtildeU")
# Eq 15 of https://arxiv.org/pdf/1501.07276.pdf:
# B = curl A -> B^r = d_th A_ph - d_ph A_th
BtildeU[0] = sp.diff(A_3vecpotentialD[2],th) - sp.diff(A_3vecpotentialD[1],ph)
# B = curl A -> B^th = d_ph A_r - d_r A_ph
BtildeU[1] = sp.diff(A_3vecpotentialD[0],ph) - sp.diff(A_3vecpotentialD[2],r)
# B = curl A -> B^ph = d_r A_th - d_th A_r
BtildeU[2] = sp.diff(A_3vecpotentialD[1],r) - sp.diff(A_3vecpotentialD[0],th)
```
<a id='adm_metric'></a>
## Step 2.j: Set the ADM quantities $\alpha$, $\beta^i$, and $\gamma_{ij}$ from the spacetime metric $g_{\mu\nu}$ \[Back to [top](#toc)\]
$$\label{adm_metric}$$
Now, we wish to build the 3+1-dimensional variables in terms of the inverse 4-dimensional spacetime metric $g^{ij},$ as demonstrated in eq. 4.49 of [Gourgoulhon's lecture notes on 3+1 formalisms](https://arxiv.org/pdf/gr-qc/0703035.pdf) (letting $N=\alpha$). So,
\begin{align}
\alpha &= \sqrt{-\frac{1}{g^{00}}} \\
\beta^i &= \alpha^2 g^{0 (i+1)} \\
\gamma^{ij} &= g^{(i+1) (j+1)} + \frac{\beta^i \beta_j}{\alpha^2},
\end{align}
again keeping careful track of the differences in the indexing conventions for 3-dimensional quantities and 4-dimensional quantities. We will also take the inverse of $\gamma^{ij}$, obtaining (naturally) $\gamma_{ij}$ and its determinant $|\gamma|$. (Note that the function we use gives the determinant of $\gamma^{ij}$, which is the reciprocal of $|\gamma|$.)
```
# Construct spacetime metric in 3+1 form:
# See, e.g., Eq. 4.49 of https://arxiv.org/pdf/gr-qc/0703035.pdf , where N = alpha
alpha = gri.register_gridfunctions("EVOL",["alpha"])
betaU = ixp.register_gridfunctions_for_single_rank1("EVOL","betaU")
alpha = sp.sqrt(1/(-gPhys4UU[0][0]))
betaU = ixp.zerorank1()
for i in range(3):
betaU[i] = alpha**2 * gPhys4UU[0][i+1]
gammaUU = ixp.zerorank2()
for i in range(3):
for j in range(3):
gammaUU[i][j] = gPhys4UU[i+1][j+1] + betaU[i]*betaU[j]/alpha**2
gammaDD = ixp.register_gridfunctions_for_single_rank2("EVOL","gammaDD","sym01")
gammaDD,igammaDET = ixp.symm_matrix_inverter3x3(gammaUU)
gammaDET = 1/igammaDET
```
Now, we will lower the index on the shift vector $\beta_j = \gamma_{ij} \beta^i$ and use that to calculate the 4-dimensional metric tensor, $g_{ij}$. So, we have
\begin{align}
g_{00} &= -\alpha^2 + \beta^2 \\
g_{0 (i+1)} = g_{(i+1) 0} &= \beta_i \\
g_{(i+1) (j+1)} &= \gamma_{ij},
\end{align}
where $\beta^2 \equiv \beta^i \beta_i$.
```
###############
# Next compute g_{\alpha \beta} from lower 3-metric, using
# Eq 4.47 of https://arxiv.org/pdf/gr-qc/0703035.pdf
betaD = ixp.zerorank1()
for i in range(3):
for j in range(3):
betaD[i] += gammaDD[i][j]*betaU[j]
beta2 = sp.sympify(0)
for i in range(3):
beta2 += betaU[i]*betaD[i]
gPhys4DD = ixp.zerorank2(DIM=4)
gPhys4DD[0][0] = -alpha**2 + beta2
for i in range(3):
gPhys4DD[0][i+1] = gPhys4DD[i+1][0] = betaD[i]
for j in range(3):
gPhys4DD[i+1][j+1] = gammaDD[i][j]
```
<a id='magnetic_field_comoving_frame'></a>
## Step 2.k: Set the magnetic field components in the comoving frame $b^\mu$, and $b^2$, which is twice the magnetic pressure \[Back to [top](#toc)\]
$$\label{magnetic_field_comoving_frame}$$
Next compute $b^{\mu}$ using Eqs 23, 24, 27 and 31 of [this paper](https://arxiv.org/pdf/astro-ph/0503420.pdf):
\begin{align}
B^i &= \frac{\tilde{B}}{\sqrt{|\gamma|}} \\
B^0_{(u)} &= \frac{u_{i+1} B^i}{\alpha} \\
b^0 &= \frac{B^0_{(u)}}{\sqrt{4 \pi}} \\
b^{i+1} &= \frac{\frac{B^i}{\alpha} + B^0_{(u)} u^{i+1}}{u^0 \sqrt{4 \pi}}
\end{align}
```
###############
# Next compute b^{\mu} using Eqs 23 and 31 of https://arxiv.org/pdf/astro-ph/0503420.pdf
uKS4D = ixp.zerorank1(DIM=4)
for i in range(4):
for j in range(4):
uKS4D[i] += gPhys4DD[i][j] * uKS4U[j]
# Eq 27 of https://arxiv.org/pdf/astro-ph/0503420.pdf
BU = ixp.zerorank1()
for i in range(3):
BU[i] = BtildeU[i]/sp.sqrt(gammaDET)
# Eq 23 of https://arxiv.org/pdf/astro-ph/0503420.pdf
BU0_u = sp.sympify(0)
for i in range(3):
BU0_u += uKS4D[i+1]*BU[i]/alpha
smallbU = ixp.zerorank1(DIM=4)
smallbU[0] = BU0_u / sp.sqrt(4 * sp.pi)
# Eqs 24 and 31 of https://arxiv.org/pdf/astro-ph/0503420.pdf
for i in range(3):
smallbU[i+1] = (BU[i]/alpha + BU0_u*uKS4U[i+1])/(sp.sqrt(4*sp.pi)*uKS4U[0])
smallbD = ixp.zerorank1(DIM=4)
for i in range(4):
for j in range(4):
smallbD[i] += gPhys4DD[i][j]*smallbU[j]
smallb2 = sp.sympify(0)
for i in range(4):
smallb2 += smallbU[i]*smallbD[i]
```
<a id='lorentz_fac_valencia'></a>
## Step 2.l: Lorentz factor $\Gamma = \alpha u^0$ and Valencia 3-velocity $v^i_{(n)}$ \[Back to [top](#toc)\]
$$\label{lorentz_fac_valencia}$$
Now, we will define the Lorentz factor ($= \alpha u^0$) and the Valencia 3-velocity $v^i_{(n)}$, which sets the 3-velocity as measured by normal observers to the spatial slice:
\begin{align}
v^i_{(n)} &= \frac{u^i}{u^0 \alpha} + \frac{\beta^i}{\alpha}, \\
\end{align}
as shown in eq 11 of [this](https://arxiv.org/pdf/1501.07276.pdf) paper. We will also compute the product of the square root of the determinant of the 3-metric with the lapse.
```
###############
LorentzFactor = alpha * uKS4U[0]
# Define Valencia 3-velocity v^i_(n), which sets the 3-velocity as measured by normal observers to the spatial slice:
# v^i_(n) = u^i/(u^0*alpha) + beta^i/alpha. See eq 11 of https://arxiv.org/pdf/1501.07276.pdf
Valencia3velocityU = ixp.zerorank1()
for i in range(3):
Valencia3velocityU[i] = uKS4U[i + 1] / (alpha * uKS4U[0]) + betaU[i] / alpha
sqrtgamma4DET = sp.symbols("sqrtgamma4DET")
sqrtgamma4DET = sp.sqrt(gammaDET)*alpha
```
<a id='output_to_c'></a>
## Step 3: Output above-generated expressions to C code, using NRPy+ \[Back to [top](#toc)\]
$$\label{output_to_c}$$
Finally, we have contructed the underlying expressions necessary for the Fishbone-Moncrief initial data. By means of demonstration, we will use NRPy+'s `FD_outputC()` to print the expressions. (The actual output statements are commented out right now, to save time in testing.)
```
KerrSchild_CKernel = [\
lhrh(lhs=gri.gfaccess("out_gfs","alpha"),rhs=alpha),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU0"),rhs=betaU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU1"),rhs=betaU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","betaU2"),rhs=betaU[2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD00"),rhs=gammaDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD01"),rhs=gammaDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD02"),rhs=gammaDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD11"),rhs=gammaDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD12"),rhs=gammaDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","gammaDD22"),rhs=gammaDD[2][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD00"),rhs=KDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD01"),rhs=KDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD02"),rhs=KDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD11"),rhs=KDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD12"),rhs=KDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","KDD22"),rhs=KDD[2][2]),\
]
#fin.FD_outputC("stdout",KerrSchild_CKernel)
FMdisk_Lorentz_uUs_CKernel = [\
lhrh(lhs=gri.gfaccess("out_gfs","LorentzFactor"),rhs=LorentzFactor),\
# lhrh(lhs=gri.gfaccess("out_gfs","uKS4U1"),rhs=uKS4U[1]),\
# lhrh(lhs=gri.gfaccess("out_gfs","uKS4U2"),rhs=uKS4U[2]),\
# lhrh(lhs=gri.gfaccess("out_gfs","uKS4U3"),rhs=uKS4U[3]),\
]
#fin.FD_outputC("stdout",FMdisk_Lorentz_uUs_CKernel)
FMdisk_hm1_rho_P_CKernel = [\
# lhrh(lhs=gri.gfaccess("out_gfs","hm1"),rhs=hm1),\
lhrh(lhs=gri.gfaccess("out_gfs","rho_initial"),rhs=rho_initial),\
lhrh(lhs=gri.gfaccess("out_gfs","Pressure_initial"),rhs=Pressure_initial),\
]
#fin.FD_outputC("stdout",FMdisk_hm1_rho_P_CKernel)
udotu = sp.sympify(0)
for i in range(4):
udotu += uKS4U[i]*uKS4D[i]
#NRPy_file_output(OUTDIR+"/standalone-spherical_coords/NRPy_codegen/FMdisk_Btildes.h", [],[],[],
# ID_protected_variables + ["r","th","ph"],
# [],[uKS4U[0], "uKS4Ut", uKS4U[1],"uKS4Ur", uKS4U[2],"uKS4Uth", uKS4U[3],"uKS4Uph",
# uKS4D[0], "uKS4Dt", uKS4D[1],"uKS4Dr", uKS4D[2],"uKS4Dth", uKS4D[3],"uKS4Dph",
# uKS4D[1] * BU[0] / alpha, "Bur", uKS4D[2] * BU[1] / alpha, "Buth", uKS4D[3] * BU[2] / alpha, "Buph",
# gPhys4DD[0][0], "g4DD00", gPhys4DD[0][1], "g4DD01",gPhys4DD[0][2], "g4DD02",gPhys4DD[0][3], "g4DD03",
# BtildeU[0], "BtildeUr", BtildeU[1], "BtildeUth",BtildeU[2], "BtildeUph",
# smallbU[0], "smallbUt", smallbU[1], "smallbUr", smallbU[2], "smallbUth",smallbU[3], "smallbUph",
# smallb2,"smallb2",udotu,"udotu"])
FMdisk_Btildes_CKernel = [\
lhrh(lhs=gri.gfaccess("out_gfs","BtildeU0"),rhs=BtildeU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","BtildeU1"),rhs=BtildeU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","BtildeU2"),rhs=BtildeU[2]),\
]
#fin.FD_outputC("stdout",FMdisk_Btildes_CKernel)
```
We will now use the relationships between coordinate systems provided by [reference_metric.py](../edit/reference_metric.py) to convert our expressions to Cartesian coordinates. See [Tutorial-Reference_Metric](Tutorial-Reference_Metric.ipynb) for more detail.
```
# Now that all derivatives of ghat and gbar have been computed,
# we may now substitute the definitions r = rfm.xxSph[0], th=rfm.xxSph[1],...
# WARNING: Substitution only works when the variable is not an integer. Hence the if not isinstance(...,...) stuff.
# If the variable isn't an integer, we revert transcendental functions inside to normal variables. E.g., sin(x2) -> sinx2
# Reverting to normal variables in this way makes expressions simpler in NRPy, and enables transcendental functions
# to be pre-computed in SENR.
alpha = alpha.subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
for i in range(DIM):
betaU[i] = betaU[i].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
for j in range(DIM):
gammaDD[i][j] = gammaDD[i][j].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
KDD[i][j] = KDD[i][j].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
# GRMHD variables:
# Density and pressure:
hm1 = hm1.subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
rho_initial = rho_initial.subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
Pressure_initial = Pressure_initial.subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
LorentzFactor = LorentzFactor.subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
# "Valencia" three-velocity
for i in range(DIM):
BtildeU[i] = BtildeU[i].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
uKS4U[i+1] = uKS4U[i+1].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
uBL4U[i+1] = uBL4U[i+1].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
Valencia3velocityU[i] = Valencia3velocityU[i].subs(r,rfm.xxSph[0]).subs(th,rfm.xxSph[1]).subs(ph,rfm.xxSph[2])
```
At last, we will use our reference metric formalism and the Jacobian associated with the two coordinate systems to convert the spherical initial data to Cartesian coordinates. The module reference_metric.py provides us with the definition of $r, \theta, \phi$ in Cartesian coordinates. To find dthe Jacobian to then transform from spherical to Cartesian, we must find the tensor \begin{equation} \frac{\partial x_i}{\partial y_j}, \end{equation} where $x_i \in \{r,\theta,\phi\}$ and $y_i \in \{x,y,z\}$. We will also compute its inverse.
```
# uUphi = uKS4U[3]
# uUphi = sympify_integers__replace_rthph(uUphi,r,th,ph,rfm.xxSph[0],rfm.xxSph[1],rfm.xxSph[2])
# uUt = uKS4U[0]
# uUt = sympify_integers__replace_rthph(uUt,r,th,ph,rfm.xxSph[0],rfm.xxSph[1],rfm.xxSph[2])
# Transform initial data to our coordinate system:
# First compute Jacobian and its inverse
drrefmetric__dx_0UDmatrix = sp.Matrix([[sp.diff(rfm.xxSph[0],rfm.xx[0]), sp.diff( rfm.xxSph[0],rfm.xx[1]), sp.diff( rfm.xxSph[0],rfm.xx[2])],
[sp.diff(rfm.xxSph[1],rfm.xx[0]), sp.diff(rfm.xxSph[1],rfm.xx[1]), sp.diff(rfm.xxSph[1],rfm.xx[2])],
[sp.diff(rfm.xxSph[2],rfm.xx[0]), sp.diff(rfm.xxSph[2],rfm.xx[1]), sp.diff(rfm.xxSph[2],rfm.xx[2])]])
dx__drrefmetric_0UDmatrix = drrefmetric__dx_0UDmatrix.inv()
# Declare as gridfunctions the final quantities we will output for the initial data
IDalpha = gri.register_gridfunctions("EVOL","IDalpha")
IDgammaDD = ixp.register_gridfunctions_for_single_rank2("EVOL","IDgammaDD","sym01")
IDKDD = ixp.register_gridfunctions_for_single_rank2("EVOL","IDKDD","sym01")
IDbetaU = ixp.register_gridfunctions_for_single_rank1("EVOL","IDbetaU")
IDValencia3velocityU = ixp.register_gridfunctions_for_single_rank1("EVOL","IDValencia3velocityU")
IDalpha = alpha
for i in range(3):
IDbetaU[i] = 0
IDValencia3velocityU[i] = 0
for j in range(3):
# Matrices are stored in row, column format, so (i,j) <-> (row,column)
IDbetaU[i] += dx__drrefmetric_0UDmatrix[(i,j)]*betaU[j]
IDValencia3velocityU[i] += dx__drrefmetric_0UDmatrix[(i,j)]*Valencia3velocityU[j]
IDgammaDD[i][j] = 0
IDKDD[i][j] = 0
for k in range(3):
for l in range(3):
IDgammaDD[i][j] += drrefmetric__dx_0UDmatrix[(k,i)]*drrefmetric__dx_0UDmatrix[(l,j)]*gammaDD[k][l]
IDKDD[i][j] += drrefmetric__dx_0UDmatrix[(k,i)]*drrefmetric__dx_0UDmatrix[(l,j)]* KDD[k][l]
# -={ Spacetime quantities: Generate C code from expressions and output to file }=-
KerrSchild_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","IDalpha"),rhs=IDalpha),\
lhrh(lhs=gri.gfaccess("out_gfs","IDbetaU0"),rhs=IDbetaU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDbetaU1"),rhs=IDbetaU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDbetaU2"),rhs=IDbetaU[2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD00"),rhs=IDgammaDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD01"),rhs=IDgammaDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD02"),rhs=IDgammaDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD11"),rhs=IDgammaDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD12"),rhs=IDgammaDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDgammaDD22"),rhs=IDgammaDD[2][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD00"),rhs=IDKDD[0][0]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD01"),rhs=IDKDD[0][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD02"),rhs=IDKDD[0][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD11"),rhs=IDKDD[1][1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD12"),rhs=IDKDD[1][2]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDKDD22"),rhs=IDKDD[2][2]),\
]
# -={ GRMHD quantities: Generate C code from expressions and output to file }=-
FMdisk_GRHD_hm1_to_print = [lhrh(lhs=gri.gfaccess("out_gfs","rho_initial"),rhs=rho_initial)]
FMdisk_GRHD_velocities_to_print = [\
lhrh(lhs=gri.gfaccess("out_gfs","IDValencia3velocityU0"),rhs=IDValencia3velocityU[0]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDValencia3velocityU1"),rhs=IDValencia3velocityU[1]),\
lhrh(lhs=gri.gfaccess("out_gfs","IDValencia3velocityU2"),rhs=IDValencia3velocityU[2]),\
]
```
To verify this against the old version of FishboneMoncriefID from the old version of NRPy, we use the `mathematica_code()` output function.
```
# Comment out debug code for now, to reduce this file's size.
#from mathematica_output import *
# print("ID1alpha = " + sp.mathematica_code(IDalpha) + ";")
# print("ID1beta0 = " + sp.mathematica_code(IDbetaU[0]) + ";")
# print("ID1beta1 = " + sp.mathematica_code(IDbetaU[1]) + ";")
# print("ID1beta2 = " + sp.mathematica_code(IDbetaU[2]) + ";")
# print("ID1gamma00 = " + sp.mathematica_code(IDgammaDD[0][0]) + ";")
# print("ID1gamma01 = " + sp.mathematica_code(IDgammaDD[0][1]) + ";")
# print("ID1gamma02 = " + sp.mathematica_code(IDgammaDD[0][2]) + ";")
# print("ID1gamma11 = " + sp.mathematica_code(IDgammaDD[1][1]) + ";")
# print("ID1gamma12 = " + sp.mathematica_code(IDgammaDD[1][2]) + ";")
# print("ID1gamma22 = " + sp.mathematica_code(IDgammaDD[2][2]) + ";")
# print("ID1K00 = " + sp.mathematica_code(IDKDD[0][0]) + ";")
# print("ID1K01 = " + sp.mathematica_code(IDKDD[0][1]) + ";")
# print("ID1K02 = " + sp.mathematica_code(IDKDD[0][2]) + ";")
# print("ID1K11 = " + sp.mathematica_code(IDKDD[1][1]) + ";")
# print("ID1K12 = " + sp.mathematica_code(IDKDD[1][2]) + ";")
# print("ID1K22 = " + sp.mathematica_code(IDKDD[2][2]) + ";")
# print("hm11 = " + sp.mathematica_code(hm1) + ";")
# print("ID1Valencia3velocityU0 = " + sp.mathematica_code(IDValencia3velocityU[0]) + ";")
# print("ID1Valencia3velocityU1 = " + sp.mathematica_code(IDValencia3velocityU[1]) + ";")
# print("ID1Valencia3velocityU2 = " + sp.mathematica_code(IDValencia3velocityU[2]) + ";")
```
<a id='code_validation'></a>
# Step 4: Code Validation against `FishboneMoncriefID.FishboneMoncriefID` NRPy+ module \[Back to [top](#toc)\]
$$\label{code_validation}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for these Fishbone-Moncrief initial data between
1. this tutorial and
2. the NRPy+ [FishboneMoncriefID.FishboneMoncriefID](../edit/FishboneMoncriefID/FishboneMoncriefID.py) module.
```
gri.glb_gridfcs_list = []
import FishboneMoncriefID.FishboneMoncriefID as fmid
fmid.FishboneMoncriefID()
print("IDalpha - fmid.IDalpha = " + str(IDalpha - fmid.IDalpha))
print("rho_initial - fmid.rho_initial = " + str(rho_initial - fmid.rho_initial))
print("hm1 - fmid.hm1 = " + str(hm1 - fmid.hm1))
for i in range(DIM):
print("IDbetaU["+str(i)+"] - fmid.IDbetaU["+str(i)+"] = " + str(IDbetaU[i] - fmid.IDbetaU[i]))
print("IDValencia3velocityU["+str(i)+"] - fmid.IDValencia3velocityU["+str(i)+"] = "\
+ str(IDValencia3velocityU[i] - fmid.IDValencia3velocityU[i]))
for j in range(DIM):
print("IDgammaDD["+str(i)+"]["+str(j)+"] - fmid.IDgammaDD["+str(i)+"]["+str(j)+"] = "
+ str(IDgammaDD[i][j] - fmid.IDgammaDD[i][j]))
print("IDKDD["+str(i)+"]["+str(j)+"] - fmid.IDKDD["+str(i)+"]["+str(j)+"] = "
+ str(IDKDD[i][j] - fmid.IDKDD[i][j]))
```
<a id='latex_pdf_output'></a>
# Step 5: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename [Tutorial-FishboneMoncriefID.pdf](Tutorial-FishboneMoncriefID.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
!jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-FishboneMoncriefID.ipynb
!pdflatex -interaction=batchmode Tutorial-FishboneMoncriefID.tex
!pdflatex -interaction=batchmode Tutorial-FishboneMoncriefID.tex
!pdflatex -interaction=batchmode Tutorial-FishboneMoncriefID.tex
!rm -f Tut*.out Tut*.aux Tut*.log
```
| github_jupyter |
## Import Library
```
# import the library
from apyori import apriori
from mlxtend.frequent_patterns import apriori, association_rules
from mlxtend.preprocessing import TransactionEncoder
import networkx as nx
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
# read the datasets
movie_data = pd.read_csv('my_movies.csv')
movie_data.head()
```
## Data Cleaning
```
# selecting specific rows
m_data = movie_data.iloc[ :, 5:]
m_data
m_data.columns.value_counts()
m_data.info()
m_data.isnull().sum()
```
- No missing values are found
- Data is in numeric format
## Basic Visualization
```
# basic counts for movies
fig, ax = plt.subplots(4, 3, figsize=(15, 15))
sns.countplot(m_data['Sixth Sense'], ax = ax[0, 0])
sns.countplot(m_data['Gladiator'], ax = ax[0, 1])
sns.countplot(m_data['LOTR1'], ax = ax[0, 2])
sns.countplot(m_data['Harry Potter1'], ax = ax[1, 0])
sns.countplot(m_data['Patriot'], ax = ax[1, 1])
sns.countplot(m_data['LOTR2'], ax = ax[1, 2])
sns.countplot(m_data['LOTR'], ax = ax[2, 0])
sns.countplot(m_data['Braveheart'], ax = ax[2, 1])
sns.countplot(m_data['Green Mile'], ax = ax[2, 2])
sns.countplot(m_data['Harry Potter2'], ax = ax[3, 0])
plt.subplots_adjust(hspace = 0.5, wspace = 0.5)
```
## Analysis Part (Association)
##### A) Try different values of support and confidence. Observe the change in number of rules for different support,confidence values
##### B) Change the minimum length in apriori algorithm
##### C) Visulize the obtained rules using different plots
1) Combination 1:
min support = 20 %
max_len = 1
2) Combination 2:
min support = 20 %
max_len = 2
3) Combination 3:
min support = 20 %
max_len = 3
4) Combination 4:
min support = 10 %
max_len = 1
5) Combination 5:
min support = 30 %
max_len = 5
##### Combination 1
```
# apriori algorithm
frequent_movieSets_1 = apriori(m_data, min_support = 0.5, use_colnames=True, max_len = 1)
frequent_movieSets_1
# association rule criteria
rule_1 = association_rules(frequent_movieSets_1, metric='lift', min_threshold = 0.8)
rule_1
```
- With this combination, association rule is not work
- But we get the max support fot movie (Gladiator) about 0.7
##### Combination 2
```
# apriori algorithm
frequent_movieSets_2 = apriori(m_data, min_support = 0.5, use_colnames=True, max_len = 2)
frequent_movieSets_2
```
##### lift
```
# association rule criteria
rule_2L = association_rules(frequent_movieSets_2, metric='lift', min_threshold = 0.8)
rule_2L
```
##### confidence
```
rule_2C = association_rules(frequent_movieSets_2, metric='confidence', min_threshold = 0.8)
rule_2C
```
- With lift consideration, pair (Gladiator)-(Sixth Sense) is visible but with confidence consideration, pair is not exist
- With both analysis, values for the support + confidence + lift + leaverge + conviction are remains same
- conviction value is high for (Patriot)-(Gladiator) which means there is strong dependency exist
```
# support v/s confidence
support = rule_2L['support']
confidence = rule_2L['confidence']
plt.scatter(support, confidence, c = 'r', alpha = 0.5)
plt.xlabel('support')
plt.ylabel('confidence')
plt.show()
# graph visualization with node form
fig, ax = plt.subplots(figsize=(10,5))
GA = nx.from_pandas_edgelist(rule_2C , source = 'antecedents', target = 'consequents')
nx.draw(GA, with_labels = True)
plt.show()
```
- From above node diagram, if people watch the movie (Gladiator) then they also like to watch movies (Sixth Sense) and (Patriot)
##### Combination 3
```
# apriori algorithm
frequent_movieSets_3 = apriori(m_data, min_support = 0.5, use_colnames=True, max_len = 3)
frequent_movieSets_3
```
- With this combination, we have same result as previous case
##### Combination 4
```
# apriori algorithm
frequent_movieSets_4 = apriori(m_data, min_support = 0.1, use_colnames=True, max_len = 1)
frequent_movieSets_4
# association rule criteria
rule_4 = association_rules(frequent_movieSets_4, metric='lift', min_threshold = 1)
rule_4
```
- With this combination, no association rule is work
##### Combination 5
```
# apriori algorithm
frequent_movieSets_5 = apriori(m_data, min_support = 0.03, use_colnames=True, max_len = 5)
frequent_movieSets_5
```
##### lift
```
# association rule criteria
rule_5L = association_rules(frequent_movieSets_5, metric='lift', min_threshold = 1)
rule_5L.sort_values('lift', ascending=False)
```
##### confidence
```
# association rule criteria
rule_5C = association_rules(frequent_movieSets_5, metric='confidence', min_threshold = 1)
rule_5C.sort_values('confidence', ascending=False)
# add condition: lift should be greater than 8 and confidence should be greater than 1
rule_5L = rule_5L[(rule_5L['lift'] >= 8) & (rule_5L['confidence'] >= 1)].sort_values('lift', ascending=False)
rule_5L_top7 = rule_5L.head(7)
rule_5L
# add condition: lift should be greater than 8 and confidence should be greater than 1
rule_5C = rule_5C[(rule_5C['lift'] >= 8) & (rule_5C['confidence'] >= 1)].sort_values('lift', ascending=False)
rule_5C_top7 = rule_5C.head(7)
rule_5C
# support v/s confidence
support = rule_5L['support']
confidence = rule_5L['confidence']
plt.scatter(support, confidence, c = 'r', alpha = 0.5)
plt.xlabel('support')
plt.ylabel('confidence')
plt.show()
# graph visualization with node form
fig, ax = plt.subplots(figsize=(20,20))
GA = nx.from_pandas_edgelist(rule_5L_top7 , source = 'antecedents', target = 'consequents')
nx.draw(GA, with_labels = True)
plt.show()
```
- From above node diagram, there are 4 conclusions
- If people watch the movie (LOTR2, Green Mile) then they also like to watch movies (LOTR1, Harry Potter1) and (LOTR1, Sixth Sence, Harry Potter1)
- If people watch the movie (LOTR2, Green Mile) then they also like to watch movies (LOTR2, Harry Potter1) and (LOTR2, Sixth Sence, Harry Potter1)
- If people watch the movie (Harry Potter1, Green Mile) then they also like to watch movies (LOTR1, LOTR2, Sixth Sence)
- If people watch the movie (LOTR) then they also like to watch movies (Gladiator, Green Mile)
| github_jupyter |
# Modeling and Simulation in Python
Case study.
Copyright 2017 Allen Downey
License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
```
### Unrolling
Let's simulate a kitten unrolling toilet paper. As reference material, see [this video](http://modsimpy.com/kitten).
The interactions of the kitten and the paper roll are complex. To keep things simple, let's assume that the kitten pulls down on the free end of the roll with constant force. Also, we will neglect the friction between the roll and the axle.

This figure shows the paper roll with $r$, $F$, and $\tau$. As a vector quantity, the direction of $\tau$ is into the page, but we only care about its magnitude for now.
We'll start by loading the units we need.
```
radian = UNITS.radian
m = UNITS.meter
s = UNITS.second
kg = UNITS.kilogram
N = UNITS.newton
```
And a few more parameters in the `Params` object.
```
params = Params(Rmin = 0.02 * m,
Rmax = 0.055 * m,
Mcore = 15e-3 * kg,
Mroll = 215e-3 * kg,
L = 47 * m,
tension = 2e-4 * N,
t_end = 120 * s)
```
`make_system` computes `rho_h`, which we'll need to compute moment of inertia, and `k`, which we'll use to compute `r`.
```
def make_system(params):
"""Make a system object.
params: Params with Rmin, Rmax, Mcore, Mroll,
L, tension, and t_end
returns: System with init, k, rho_h, Rmin, Rmax,
Mcore, Mroll, ts
"""
unpack(params)
init = State(theta = 0 * radian,
omega = 0 * radian/s,
y = L)
area = pi * (Rmax**2 - Rmin**2)
rho_h = Mroll / area
k = (Rmax**2 - Rmin**2) / 2 / L / radian
return System(init=init, k=k, rho_h=rho_h,
Rmin=Rmin, Rmax=Rmax,
Mcore=Mcore, Mroll=Mroll,
t_end=t_end)
```
Testing `make_system`
```
system = make_system(params)
system.init
```
Here's how we compute `I` as a function of `r`:
```
def moment_of_inertia(r, system):
"""Moment of inertia for a roll of toilet paper.
r: current radius of roll in meters
system: System object with Mcore, rho, Rmin, Rmax
returns: moment of inertia in kg m**2
"""
unpack(system)
Icore = Mcore * Rmin**2
Iroll = pi * rho_h / 2 * (r**4 - Rmin**4)
return Icore + Iroll
```
When `r` is `Rmin`, `I` is small.
```
moment_of_inertia(system.Rmin, system)
```
As `r` increases, so does `I`.
```
moment_of_inertia(system.Rmax, system)
```
## Exercises
Write a slope function we can use to simulate this system. Here are some suggestions and hints:
* `r` is no longer part of the `State` object. Instead, we compute `r` at each time step, based on the current value of `y`, using
$y = \frac{1}{2k} (r^2 - R_{min}^2)$
* Angular velocity, `omega`, is no longer constant. Instead, we compute torque, `tau`, and angular acceleration, `alpha`, at each time step.
* I changed the definition of `theta` so positive values correspond to clockwise rotation, so `dydt = -r * omega`; that is, positive values of `omega` yield decreasing values of `y`, the amount of paper still on the roll.
* Your slope function should return `omega`, `alpha`, and `dydt`, which are the derivatives of `theta`, `omega`, and `y`, respectively.
* Because `r` changes over time, we have to compute moment of inertia, `I`, at each time step.
That last point might be more of a problem than I have made it seem. In the same way that $F = m a$ only applies when $m$ is constant, $\tau = I \alpha$ only applies when $I$ is constant. When $I$ varies, we usually have to use a more general version of Newton's law. However, I believe that in this example, mass and moment of inertia vary together in a way that makes the simple approach work out. Not all of my collegues are convinced.
```
# Solution goes here
```
Test `slope_func` with the initial conditions.
```
# Solution goes here
```
Run the simulation.
```
# Solution goes here
```
And look at the results.
```
results.tail()
```
Check the results to see if they seem plausible:
* The final value of `theta` should be about 220 radians.
* The final value of `omega` should be near 4 radians/second, which is less one revolution per second, so that seems plausible.
* The final value of `y` should be about 35 meters of paper left on the roll, which means the kitten pulls off 12 meters in two minutes. That doesn't seem impossible, although it is based on a level of consistency and focus that is unlikely in a kitten.
* Angular velocity, `omega`, should increase almost linearly at first, as constant force yields almost constant torque. Then, as the radius decreases, the lever arm decreases, yielding lower torque, but moment of inertia decreases even more, yielding higher angular acceleration.
Plot `theta`
```
def plot_theta(results):
plot(results.theta, color='C0', label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
plot_theta(results)
```
Plot `omega`
```
def plot_omega(results):
plot(results.omega, color='C2', label='omega')
decorate(xlabel='Time (s)',
ylabel='Angular velocity (rad/s)')
plot_omega(results)
```
Plot `y`
```
def plot_y(results):
plot(results.y, color='C1', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
plot_y(results)
```
| github_jupyter |
```
%load_ext autoreload
%matplotlib inline
import logging
import sys
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import HTML
from matplotlib import animation
from scipy.spatial import procrustes
from scipy.linalg import orthogonal_procrustes
from sklearn.decomposition import PCA
from sklearn.preprocessing import MinMaxScaler
from sklearn.tree import DecisionTreeRegressor, export_graphviz
import pydotplus
from sklearn.externals.six import StringIO
from IPython.display import Image
import hyperhyper as hy
import dataset
import math
import logging
# logging.basicConfig(stream=sys.stdout, level=logging.DEBUG)
def get_best(year):
b = hy.Bunch(f'/mnt/data2/ptf/bunches/bi_{year}_decay_4')
db = dataset.connect(f'sqlite:////mnt/data2/ptf/bunches/bi_{year}_decay_4/results.db')
statement = 'SELECT * FROM experiments where pair_args__dynamic_window="decay" ORDER BY micro_results DESC LIMIT 1'
for best in list(db.query(statement)):
oov = True if best['pair_args__delete_oov'] == 1 else False
window = int(best['pair_args__window'])
if not isinstance(window, int):
window = int.from_bytes(window, "little")
neg = float(best['neg'])
if neg.is_integer():
neg = int(neg)
dim = int(best['dim'])
print(oov, best)
try:
print(best['neg'])
kv, res = b.svd(impl='scipy', evaluate=True, pair_args={'subsample': 'deter', 'subsample_factor': best['pair_args__subsample_factor'], 'delete_oov': True, 'decay_rate': best['pair_args__decay_rate'], 'window': window, 'dynamic_window': 'decay'}, neg=neg, eig=best['eig'], dim=dim, keyed_vector=True)
print(res)
print(best)
except Exception as e:
print(e)
return kv
def draw_image(w, kv):
pca = PCA(n_components=2)
scaler = MinMaxScaler()
vecs = [(w, kv[w])]
for name, _ in kv.most_similar(w):
vecs.append((name, kv[name]))
principalComponents = pca.fit_transform([x[1] for x in vecs])
principalComponents = scaler.fit_transform(principalComponents)
fig, ax = plt.subplots()
ax.scatter(principalComponents[:, 0], principalComponents[:, 1])
for i, txt in enumerate([x[0] for x in vecs]):
x = principalComponents[i][0] + 0.02
y = principalComponents[i][1] - 0.02
ax.annotate(txt, (x, y))
def get_most_similar(w, kv, num=10):
names = [w]
vecs = [kv[w]]
for name, _ in kv.most_similar(w, topn=num + 10):
names.append(name)
vecs.append(kv[name])
for idx, n in enumerate(names):
for n2 in names:
if n == n2:
continue
if n[:-1] == n2 and n[-1] in ('s', 'n'):
del names[idx]
del vecs[idx]
return names[:num], vecs[:num]
def to_2d(vecs):
pca = PCA(n_components=2)
scaler = MinMaxScaler((-1, 1))
points = pca.fit_transform(vecs)
points = scaler.fit_transform(points)
print(len(points))
return points, [pca, scaler]
kvs = [get_best(year) for year in [2010, 2012, 2014, 2016, 2018]]
# kvs = [kv1, kv2, kv3, kv4]
def align(m1, m2, names1, names2):
m1_reordered = []
for idx2, n2 in enumerate(names2):
if n2 in names1:
idx1 = names1.index(n2)
m1_reordered.append(m1[idx1])
else:
m1_reordered.append(np.zeros(2))
_,m, score = procrustes(m1_reordered, m2)
sc = MinMaxScaler((-1, 1))
m = sc.fit_transform(m)
print(score)
return m
def align_orth(m1, m2, names1, names2):
m1_reordered = []
m2_reordered = []
for idx2, n2 in enumerate(names2):
if n2 in names1:
idx1 = names1.index(n2)
m1_reordered.append(m1[idx1])
m2_reordered.append(m2[idx2])
transformation_matrix, score = orthogonal_procrustes(m2_reordered, m1_reordered)
# print(score)
m = m2.dot(transformation_matrix)
sc = MinMaxScaler((-1, 1))
m = sc.fit_transform(m)
return m
def to_length(a):
# push points out
a = np.array(a)
cof = 3/np.linalg.norm(a)
return a * cof
# kvs = kvs[:3]
def get_location_for_missing_next():
pass
def get_all(words, kv):
return words, [kv[w] for w in words]
def get_settings(w, words=None):
if not words is None:
ps = [get_all(words, kv) for kv in kvs]
else:
# raw
ps = [get_most_similar(w, kv, 10) for kv in kvs]
transformed_points = []
next_points = []
transformers = []
names = []
for i in range(len(ps)):
p = ps[i]
names.append(p[0])
tp, tr = to_2d(p[1])
if i > 0:
tp = align_orth(transformed_points[-1], tp, names[i - 1], names[i])
transformed_points.append(tp)
transformers.append(tr)
for i in range(len(ps) - 1):
set_net_p = []
# for name in names[i]:
# if name in kvs[i + 1]:
# value = kvs[i + 1][name]
# pca, scaler = transformers[i + 1]
# value = scaler.transform(pca.transform([value]))[0]
# else:
# value = np.array([-2, -2])
# set_net_p.append(value)
# next_points.append(set_net_p)
for n_idx, name in enumerate(names[i]):
if name in names[i + 1]:
value = transformed_points[i + 1][names[i + 1].index(name)]
else:
value = to_length(transformed_points[i][n_idx])
set_net_p.append(value)
next_points.append(set_net_p)
next_points.append(transformed_points[-1])
# insert the points that will appear in the next scene to current scne
# because we are iterating over the names, add the names after the main loops
names_to_add = []
for i in range(1, len(ps)):
new_names = []
for n_idx, n in enumerate(names[i]):
if not n in names[i - 1]:
new_names.append(n)
transformed_points[i - 1] = np.concatenate(
(
transformed_points[i - 1],
np.asarray([to_length(transformed_points[i][n_idx])]),
),
axis=0,
)
next_points[i - 1] = np.concatenate(
(next_points[i - 1], np.asarray([transformed_points[i][n_idx]])),
axis=0,
)
names_to_add.append(new_names)
# add the new names here
for i, n in enumerate(names_to_add):
names[i] += n
return names, transformed_points, next_points
def get_anim(names, cur_points, next_points):
num_settings = len(names)
fig, ax = plt.subplots()
ax.set_xlim((-1.2, 1.2))
ax.set_ylim((-1.2, 1.2))
fig.set_size_inches(5, 5)
scat = ax.scatter([], [])
ans = []
for n in names[0]:
ans.append(ax.annotate("", xy=(0, 0)))
year = ax.annotate("", xy=(0.7, -1.2))
def init():
print("init")
scat.set_offsets(cur_points[0])
for n in names[0]:
ans.append(ax.annotate("", xy=(0, 0)))
return (scat, year, *ans)
# animation function. This is called sequentially
def animate(t):
# t /= 2
cur_set = math.floor(t)
t -= cur_set
if t > 0.5:
t -= 0.5
t *= 2
interpolation = (
np.array(cur_points[cur_set]) * (1 - t)
+ np.array(next_points[cur_set]) * t
)
else:
interpolation = cur_points[cur_set]
year.set_text(str(2010 + cur_set * 2) + '/' + str(2011 + cur_set * 2))
# clear old scatter points
interpolation = np.concatenate((interpolation, np.ones((20, 2)) + 10), axis=0)
scat.set_offsets(interpolation)
updated_idx = 0
for idx, n in enumerate(names[cur_set]):
updated_idx += 1
ans[idx].set_position(interpolation[idx] + (0.03, -0.025))
ans[idx].set_text(n)
# clear old annotations
while updated_idx < len(ans):
ans[updated_idx].set_text("")
updated_idx += 1
# del ans[updated_idx]
return (scat, year, *ans)
# call the animator. blit=True means only re-draw the parts that have changed.
anim = animation.FuncAnimation(
fig,
animate,
init_func=init,
frames=np.arange(0, num_settings, 0.01),
interval=60,
blit=True,
)
return anim
# s = get_settings('migrant', words=['deutscher', 'syrier', 'flüchtling', 'migrant', 'asylant', 'faulenzer', 'straftäter', 'ausländer', 'vergewaltiger', 'mörder', 'feigling'])
# s = get_settings('migrant', words=['merkel', 'mutti', 'abschiebung', 'grenze', 'zaun', 'angie', 'verbrecher'] + ['deutscher', 'syrier', 'flüchtling', 'migrant', 'asylant', 'faulenzer', 'straftäter', 'ausländer', 'vergewaltiger', 'mörder', 'feigling'])
# s = get_settings('grüne', words=['berlin', 'london', 'paris', 'washington', 'athen', 'madrid', 'wien', 'kopenhagen', 'oslo'])
# s = get_settings('grüne', words=['merkel', 'seehofer', 'gabriel', 'roth', 'trittin', 'wagenknecht', 'gysi'])
s = get_settings('mittelmeer')
a = get_anim(*s)
# a.save('basic_animation.mp4', fps=30) # extra_args=['-vcodec', 'libx264']
a
HTML(a.to_html5_video())
```
| github_jupyter |
Para entrar no modo apresentação, execute a seguinte célula e pressione `-`
```
%reload_ext slide
```
<span class="notebook-slide-start"/>
# APIs do GitHub (v3)
Este notebook apresenta os seguintes tópicos:
- [APIs do GitHub](#APIs-do-GitHub)
- [Autenticação](#Autentica%C3%A7%C3%A3o)
- [API v3](#API-v3)
- [Exercício 6](#Exerc%C3%ADcio-6)
- [Exercício 7](#Exerc%C3%ADcio-7)
- [Exercício 8](#Exerc%C3%ADcio-8)
## APIs do GitHub
Como o GitHub oferece APIs para obter informações de repositórios, usá-las em geral é melhor do que fazer crawling.
O GitHub possui duas versões estáveis de APIs:
- REST API v3: https://developer.github.com/v3/
- GraphQL API v4: https://developer.github.com/v4/
A forma de usar cada API é diferente e a taxa de requisições permitidas também é. Neste minicurso, usaremos requests para acessar ambas as APIs, mas existem bibliotecas prontas (como a PyGitHub para a v3) que fazem o acesso.
### Autenticação
Para usar qualquer uma das APIs, é necessário gerar um token de autenticação no GitHub seguindo os seguintes passos.
Primeiro, vá em configurações da conta.
<img src="images/github1.png" alt="Página inicial do GitHub" width="auto"/>
Em seguida, abra configurações de desenvolvedor.
<img src="images/github2.png" alt="Página de Configurações do Usuário" width="auto"/>
Abra "Personal access tokens" e clique em "Generate new token".
<img src="images/github3.png" alt="Página de Tokens de Acesso Pessoal" wi3dth="auto"/>
Escolha as permissões que você deseja no token.
<img src="images/github4.png" alt="Página de Criação de Token de Acesso Pessoal" width="auto"/>
Copie o token gerado para algum lugar seguro. Para o minicurso, eu copiei o meu token para `~/githubtoken.txt` e vou carregá-lo para a variável `token` a seguir. <span class="notebook-slide-extra" data-count="1"/>
```
from ipywidgets import FileUpload, interact
@interact(files=FileUpload())
def set_token(files={}):
global token
if files:
for key, values in files.items():
token = values['content'].decode("utf-8").strip()
print("Token Loaded!")
```
## API v3
Com o token em mãos, podemos começar a usa a API v3. O acesso a API do GitHub é feito a https://api.github.com. Portanto, precisamos mudar o site de nosso servidor de proxy. Para isso, podemos fechar e reiniciar da seguinte forma:
```bash
python proxy.py https://api.github.com/
```
Inicialmente, vamos fazer uma requisição para verificar se a autenticação funciona e para vermos nosso limite de requisições. <span class="notebook-slide-extra" data-count="1"/>
```
import requests
SITE = "http://localhost:5000/" # ou https://api.github.com
def token_auth(request):
request.headers["User-Agent"] = "Minicurso" # Necessário
request.headers["Authorization"] = "token {}".format(token)
return request
response = requests.get(SITE, auth=token_auth)
response.status_code
```
Resultado 200 - a autenticação funcionou. <span class="notebook-slide-scroll" data-position="-1"/>
O limite de acesso vem definido no header. <span class="notebook-slide-extra" data-count="3"/>
```
response.headers["X-RateLimit-Limit"]
response.headers["X-RateLimit-Remaining"]
response.headers["X-RateLimit-Reset"]
```
O retorno da API v3 é sempre um JSON. O acesso a https://api.github.com retorna as URLS válidas da API. <span class="notebook-slide-extra" data-count="1"/>
```
response.json()
```
Vamos ver o que a API tem sobre algum repositório.
Primeiro precisamos ver qual URL usar. <span class="notebook-slide-extra" data-count="1"/>
```
_['repository_url']
```
Em seguida, fazemos a requisição para saber o que tem no repositorio `gems-uff/sapos`. <span class="notebook-slide-extra" data-count="2"/>
```
response = requests.get(SITE + "repos/gems-uff/sapos", auth=token_auth)
response.status_code
data = response.json()
data
```
O resultado tem diversos resultados e URLs para pegar mais informações. Vamos pegar algumas informações diretas interessantes. <span class="notebook-slide-extra" data-count="1"/>
```
print("Estrelas:", data["stargazers_count"])
print("Forks:", data["forks"])
print("Watchers:", data["subscribers_count"])
print("Issues abertas:", data["open_issues"])
print("Linguagem:", data["language"])
```
Se quisermos saber quem são os colaboradores do projeto e quais são as issues existentes, podemos obter as respectivas URLs. <span class="notebook-slide-extra" data-count="1"/>
```
print("Colaboradores:", data["contributors_url"])
print("Issues:", data["issues_url"])
```
Agora podemos obter a lista de colaboradores. <span class="notebook-slide-extra" data-count="2"/>
```
response = requests.get(SITE + "repos/gems-uff/sapos/contributors", auth=token_auth)
response.status_code
data = response.json()
data
```
### Gráfico de Barras
A partir desta lista, podemos fazer um gráfico de barras de contribuições. <span class="notebook-slide-extra" data-count="2"/>
```
contributions = {x["login"]: x["contributions"] for x in data}
contributions
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
labels, values = zip(*contributions.items())
indexes = np.arange(len(labels))
width = 1
plt.barh(indexes, values, width)
plt.yticks(indexes, labels)
plt.show()
```
Nesse código:
- Importamos `matplotlib` e `numpy` para gerar o gráfico
- Chamamos `%matplotlib inline` para permitir a visualização da figura
- Separamos o dicionário `contributions` em duas listas de `labels` e `values`
- Criamos um `arange` de índices
- Criamos o gráfico de barras horizontal, usando `barh`
- Definimos os ticks de y como sendo os `labels` extraídos anteriormente
Podemos acessar também a url de issues. <span class="notebook-slide-extra" data-count="3"/>
```
response = requests.get(SITE + "repos/gems-uff/sapos/issues", auth=token_auth)
response.status_code
data = response.json()
data
len(data)
```
Por padrão, a API retorna 30 itens por página. Dessa forma. a lista retornou apenas a primeira página de issues. <span class="notebook-slide-position" data-count="-1"/>
Podemos acessar a segunda página com o parâmetro `?page=2`. <span class="notebook-slide-extra" data-count="3"/>
```
response = requests.get(SITE + "repos/gems-uff/sapos/issues?page=2", auth=token_auth)
response.status_code
data2 = response.json()
data2
len(data2)
```
Podemos formar uma lista com todas as issues abertas. <span class="notebook-slide-extra" data-count="1"/>
```
open_issues = data + data2
```
Essas são apenas as issues abertas. Para pegarmos as issues fechadas, precisamos definir `state=closed`. Podemos aproveitar e definir também `per_page=100` (limite máximo) e fazer um código para pegar todas as páginas. <span class="notebook-slide-extra" data-count="1"/>
```
should_continue = True
page = 1
closed_issues = []
while should_continue:
response = requests.get(SITE + "repos/gems-uff/sapos/issues?page={}&per_page=100&state=closed".format(page), auth=token_auth)
if response.status_code != 200:
print("Fail:", response.status_code)
break
data = response.json()
closed_issues += data
if len(data) < 100:
should_continue = False
page += 1
len(closed_issues), page - 1
```
Foram encontradas 262 issues em 3 páginas. <span class="notebook-slide-scroll" data-position="-1"/>
Agora podemos fazer um gráfico que mostre a evolução de issues abertas ao longo do tempo.
Para fazer esse gráfico, primeiro precisamos combinar as issues e descobrir qual foi a data da issue mais antiga. <span class="notebook-slide-extra" data-count="1"/>
```
import dateutil.parser
all_issues = open_issues + closed_issues
oldest_issue = min(
all_issues,
key=lambda x: dateutil.parser.parse(x["created_at"])
)
oldest_date = dateutil.parser.parse(oldest_issue["created_at"])
oldest_date
```
A partir desta data, podemos criar um range de dias até hoje para ser o nosso índice do gráfico e um array de zeros do `numpy` para acumularmos a quantidade de issues abertas. <span class="notebook-slide-extra" data-count="1"/>
```
from datetime import datetime, timezone
today = datetime.now(timezone.utc)
delta = today - oldest_date
days = delta.days
print(days)
indexes = np.arange(days)
values = np.zeros(days)
```
Podemos percorrer todas as issues abertas, incrementando `values` do período em que elas foram abertas até hoje. E podemos percorrer todas as issues fechadas incrementando `values` do período em que elas foram abertas até o período em que elas foram fechadas. <span class="notebook-slide-extra" data-count="1"/>
```
for issue in open_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
values[created_at_index:] += 1
for issue in closed_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
closed_at = dateutil.parser.parse(issue["closed_at"])
closed_at_index = (closed_at - oldest_date).days
values[created_at_index:closed_at_index] += 1
```
Já é possível plotar o gráfico desta forma, mas o entendimento dos eixos ainda não é o ideal. <span class="notebook-slide-extra" data-count="1"/>
```
plt.plot(indexes, values)
```
Precisamos definir quais são os anos no eixo x. <span class="notebook-slide-extra" data-count="1"/>
```
from math import ceil
labels = [datetime(2013 + i, 1, 1, tzinfo=timezone.utc) for i in range(ceil(delta.days / 365))]
label_indexes = [(label - oldest_date).days for label in labels]
label_years = [label.year for label in labels]
plt.xticks(label_indexes, label_years)
plt.plot(indexes, values)
plt.show()
```
Também podemos definir o que é cada eixo. <span class="notebook-slide-extra" data-count="1"/>
```
plt.xticks(label_indexes, label_years)
plt.xlabel("Time")
plt.ylabel("Open Issues")
plt.plot(indexes, values)
plt.show()
```
Issues podem ter diversos labels. Agora vamos fazer um gráfico que mostre barras estacadas com a evolução de cada tipo de issue. <span class="notebook-slide-extra" data-count="1"/>
```
from collections import defaultdict
values = defaultdict(lambda: np.zeros(days))
for issue in open_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
for label in issue["labels"]:
values[label["name"]][created_at_index:] += 1
if not issue["labels"]:
values["no-label"][created_at_index:] += 1
for issue in closed_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
closed_at = dateutil.parser.parse(issue["closed_at"])
closed_at_index = (closed_at - oldest_date).days
for label in issue["labels"]:
values[label["name"]][created_at_index:closed_at_index] += 1
if not issue["labels"]:
values["no-label"][created_at_index:closed_at_index] += 1
bottom = np.zeros(days)
legend_color = []
legend_text = []
for label, yvalues in values.items():
if not label[0].isdigit(): # Exclui tags de versões
ax = plt.bar(indexes, yvalues, 1,
bottom=bottom)
legend_color.append(ax[0])
bottom += yvalues
legend_text.append(label)
plt.xticks(label_indexes, label_years)
plt.xlabel("Time")
plt.ylabel("Open Issues By Type")
plt.legend(legend_color, legend_text)
plt.show()
```
## Exercício 6
Crie um gráfico de linhas que mostre apenas issues do tipo bug. <span class="notebook-slide-extra" data-count="1"/>
```
plt.plot(indexes, values["bug"])
plt.xticks(label_indexes, label_years)
plt.xlabel("Time")
plt.ylabel("Open Bug Issues")
plt.show()
```
## Exercício 7
Crie um gráfico de barras para mostrar a participação de usuários em cada issue. Considere o atributo `user`. <span class="notebook-slide-extra" data-count="2"/>
```
values = defaultdict(lambda: np.zeros(days))
for issue in open_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
values[issue["user"]["login"]][created_at_index:] += 1
for issue in closed_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - oldest_date).days
closed_at = dateutil.parser.parse(issue["closed_at"])
closed_at_index = (closed_at - oldest_date).days
values[issue["user"]["login"]][created_at_index:closed_at_index] += 1
bottom = np.zeros(days)
legend_color = []
legend_text = []
for label, yvalues in values.items():
if not label[0].isdigit(): # Exclui tags de versões
ax = plt.bar(indexes, yvalues, 1,
bottom=bottom)
legend_color.append(ax[0])
bottom += yvalues
legend_text.append(label)
plt.xticks(label_indexes, label_years)
plt.xlabel("Time")
plt.ylabel("Open Issues By User")
plt.legend(
legend_color, legend_text,
bbox_to_anchor=(0,1.02,1,0.2), loc="lower left",
mode="expand", borderaxespad=0, ncol=2
)
plt.show()
```
## Exercício 8
Filtre o gráfico do total de issues abertas para mostrar apenas o ano 2014. <span class="notebook-slide-extra" data-count="2"/>
```
yfirst = datetime(2014, 1, 1, tzinfo=timezone.utc)
ylast = datetime(2015, 1, 1, tzinfo=timezone.utc)
yfirst_index = (yfirst - oldest_date).days
ylast_index = (ylast - oldest_date).days
deltadays = (ylast - yfirst).days
values = np.zeros(deltadays)
indexes = np.arange(deltadays)
for issue in open_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - yfirst).days
if created_at < ylast:
if created_at < yfirst:
created_at_index = 0
values[created_at_index:] += 1
for issue in closed_issues:
created_at = dateutil.parser.parse(issue["created_at"])
created_at_index = (created_at - yfirst).days
if created_at < ylast:
if created_at < yfirst:
created_at_index = 0
closed_at = dateutil.parser.parse(issue["closed_at"])
closed_at_index = (closed_at - yfirst).days
if closed_at >= ylast:
closed_at_index = None
if closed_at >= yfirst:
values[created_at_index:closed_at_index] += 1
labels = [datetime(2014, i + 1, 1, tzinfo=timezone.utc) for i in range(12)]
label_indexes = [(label - yfirst).days for label in labels]
label_years = [label.month for label in labels]
plt.xticks(label_indexes, label_years)
plt.plot(indexes, values)
plt.show()
```
Continua: [7.API.v4.ipynb](7.API.v4.ipynb)
| github_jupyter |
# Measurement Error Mitigation
```
from qiskit import *
```
### Introduction
The effect of noise is to give us outputs that are not quite correct. The effect of noise that occurs throughout a computation will be quite complex in general, as one would have to consider how each gate transforms the effect of each error.
A simpler form of noise is that occuring during final measurement. At this point, the only job remaining in the circuit is to extract a bit string as an output. For an $n$ qubit final measurement, this means extracting one of the $2^n$ possible $n$ bit strings. As a simple model of the noise in this process, we can imagine that the measurement first selects one of these outputs in a perfect and noiseless manner, and then noise subsequently causes this perfect output to be randomly perturbed before it is returned to the user.
Given this model, it is very easy to determine exactly what the effects of measurement errors are. We can simply prepare each of the $2^n$ possible basis states, immediately measure them, and see what probability exists for each outcome.
As an example, we will first create a simple noise model, which randomly flips each bit in an output with probability $p$.
```
from qiskit.providers.aer.noise import NoiseModel
from qiskit.providers.aer.noise.errors import pauli_error, depolarizing_error
def get_noise(p):
error_meas = pauli_error([('X',p), ('I', 1 - p)])
noise_model = NoiseModel()
noise_model.add_all_qubit_quantum_error(error_meas, "measure") # measurement error is applied to measurements
return noise_model
```
Let's start with an instance of this in which each bit is flipped $1\%$ of the time.
```
noise_model = get_noise(0.01)
```
Now we can test out its effects. Specifically, let's define a two qubit circuit and prepare the states $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$. Without noise, these would lead to the definite outputs `'00'`, `'01'`, `'10'` and `'11'`, respectively. Let's see what happens with noise. Here, and in the rest of this section, the number of samples taken for each circuit will be `shots=10000`.
```
for state in ['00','01','10','11']:
qc = QuantumCircuit(2,2)
if state[0]=='1':
qc.x(1)
if state[1]=='1':
qc.x(0)
qc.measure(qc.qregs[0],qc.cregs[0])
print(state+' becomes',
execute(qc,Aer.get_backend('qasm_simulator'),noise_model=noise_model,shots=10000).result().get_counts())
```
Here we find that the correct output is certainly the most dominant. Ones that differ on only a single bit (such as `'01'`, `'10'` in the case that the correct output is `'00'` or `'11'`), occur around $1\%$ of the time. Those than differ on two bits occur only a handful of times in 10000 samples, if at all.
So what about if we ran a circuit with this same noise model, and got an result like the following?
```
{'10': 98, '11': 4884, '01': 111, '00': 4907}
```
Here `'01'` and `'10'` occur for around $1\%$ of all samples. We know from our analysis of the basis states that such a result can be expected when these outcomes should in fact never occur, but instead the result should be something that differs from them by only one bit: `'00'` or `'11'`. When we look at the results for those two outcomes, we can see that they occur with roughly equal probability. We can therefore conclude that the initial state was not simply $\left|00\right\rangle$, or $\left|11\right\rangle$, but an equal superposition of the two. If true, this means that the result should have been something along the lines of.
```
{'11': 4977, '00': 5023}
```
Here is a circuit that produces results like this (up to statistical fluctuations).
```
qc = QuantumCircuit(2,2)
qc.h(0)
qc.cx(0,1)
qc.measure(qc.qregs[0],qc.cregs[0])
print(execute(qc,Aer.get_backend('qasm_simulator'),noise_model=noise_model,shots=10000).result().get_counts())
```
In this example we first looked at results for each of the definite basis states, and used these results to mitigate the effects of errors for a more general form of state. This is the basic principle behind measurement error mitigation.
### Error mitigation in with linear algebra
Now we just need to find a way to perform the mitigation algorithmically rather than manually. We will do this by describing the random process using matrices. For this we need to rewrite our counts dictionaries as column vectors. For example, the dictionary `{'10': 96, '11': 1, '01': 95, '00': 9808}` describing would be rewritten as
$$
C =
\begin{pmatrix}
9808 \\
95 \\
96 \\
1
\end{pmatrix}.
$$
Here the first element is that for `'00'`, the next is that for `'01'`, and so on.
The information gathered from the basis states $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$ can then be used to define a matrix, which rotates from an ideal set of counts to one affected by measurement noise. This is done by simply taking the counts dictionary for $\left|00\right\rangle$, normalizing it it so that all elements sum to one, and then using it as the first column of the matrix. The next column is similarly defined by the counts dictionary obtained for $\left|00\right\rangle$, and so on.
There will be statistical variations each time the circuit for each basis state is run. In the following, we will use the data obtained when this section was written, which was as follows.
```
00 becomes {'10': 96, '11': 1, '01': 95, '00': 9808}
01 becomes {'10': 2, '11': 103, '01': 9788, '00': 107}
10 becomes {'10': 9814, '11': 90, '01': 1, '00': 95}
11 becomes {'10': 87, '11': 9805, '01': 107, '00': 1}
```
This gives us the following matrix.
$$
M =
\begin{pmatrix}
0.9808&0.0107&0.0095&0.0001 \\
0.0095&0.9788&0.0001&0.0107 \\
0.0096&0.0002&0.9814&0.0087 \\
0.0001&0.0103&0.0090&0.9805
\end{pmatrix}
$$
If we now take the vector describing the perfect results for a given state, applying this matrix gives us a good approximation of the results when measurement noise is present.
$$ C_{noisy} = M ~ C_{ideal}$$.
As an example, let's apply this process for the state $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$,
$$
\begin{pmatrix}
0.9808&0.0107&0.0095&0.0001 \\
0.0095&0.9788&0.0001&0.0107 \\
0.0096&0.0002&0.9814&0.0087 \\
0.0001&0.0103&0.0090&0.9805
\end{pmatrix}
\begin{pmatrix}
0 \\
5000 \\
5000 \\
0
\end{pmatrix}
=
\begin{pmatrix}
101 \\
4895.5 \\
4908 \\
96.5
\end{pmatrix}.
$$
In code, we can express this as follows.
```
import numpy as np
M = [[0.9808,0.0107,0.0095,0.0001],
[0.0095,0.9788,0.0001,0.0107],
[0.0096,0.0002,0.9814,0.0087],
[0.0001,0.0103,0.0090,0.9805]]
Cideal = [[0],
[5000],
[5000],
[0]]
Cnoisy = np.dot( M, Cideal)
print('C_noisy =\n',Cnoisy)
```
Either way, the resulting counts found in $C_{noisy}$, for measuring the $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$ with measurement noise, come out quite close to the actual data we found earlier. So this matrix method is indeed a good way of predicting noisy results given a knowledge of what the results should be.
Unfortunately, this is the exact opposite of what we need. Instead of a way to transform ideal counts data into noisy data, we need a way to transform noisy data into ideal data. In linear algebra, we do this for a matrix $M$ by finding the inverse matrix $M^{-1}$,
$$C_{ideal} = M^{-1} C_{noisy}.$$
```
import scipy.linalg as la
M = [[0.9808,0.0107,0.0095,0.0001],
[0.0095,0.9788,0.0001,0.0107],
[0.0096,0.0002,0.9814,0.0087],
[0.0001,0.0103,0.0090,0.9805]]
Minv = la.inv(M)
print(Minv)
```
Applying this inverse to $C_{noisy}$, we can obtain an approximation of the true counts.
```
Cmitigated = np.dot( Minv, Cnoisy)
print('C_mitigated =\n',Cmitigated)
```
Of course, counts should be integers, and so these values need to be rounded. This gives us a very nice result.
$$
C_{mitigated} =
\begin{pmatrix}
0 \\
5000 \\
5000 \\
0
\end{pmatrix}
$$
This is exactly the true result we desire. Our mitigation worked extremely well!
### Error mitigation in Qiskit
```
from qiskit.ignis.mitigation.measurement import (complete_meas_cal,CompleteMeasFitter)
```
The process of measurement error mitigation can also be done using tools from Qiskit. This handles the collection of data for the basis states, the construction of the matrices and the calculation of the inverse. The latter can be done using the pseudo inverse, as we saw above. However, the default is an even more sophisticated method using least squares fitting.
As an example, let's stick with doing error mitigation for a pair of qubits. For this we define a two qubit quantum register, and feed it into the function `complete_meas_cal`.
```
qr = qiskit.QuantumRegister(2)
meas_calibs, state_labels = complete_meas_cal(qr=qr, circlabel='mcal')
```
This creates a set of circuits to take measurements for each of the four basis states for two qubits: $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$.
```
for circuit in meas_calibs:
print('Circuit',circuit.name)
print(circuit)
print()
```
Let's now run these circuits without any noise present.
```
# Execute the calibration circuits without noise
backend = qiskit.Aer.get_backend('qasm_simulator')
job = qiskit.execute(meas_calibs, backend=backend, shots=1000)
cal_results = job.result()
```
With the results we can construct the calibration matrix, which we have been calling $M$.
```
meas_fitter = CompleteMeasFitter(cal_results, state_labels, circlabel='mcal')
print(meas_fitter.cal_matrix)
```
With no noise present, this is simply the identity matrix.
Now let's create a noise model. And to make things interesting, let's have the errors be ten times more likely than before.
```
noise_model = get_noise(0.1)
```
Again we can run the circuits, and look at the calibration matrix, $M$.
```
backend = qiskit.Aer.get_backend('qasm_simulator')
job = qiskit.execute(meas_calibs, backend=backend, shots=1000, noise_model=noise_model)
cal_results = job.result()
meas_fitter = CompleteMeasFitter(cal_results, state_labels, circlabel='mcal')
print(meas_fitter.cal_matrix)
```
This time we find a more interesting matrix, and one that is not invertible. Let's see how well we can mitigate for this noise. Again, let's use the Bell state $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$ for our test.
```
qc = QuantumCircuit(2,2)
qc.h(0)
qc.cx(0,1)
qc.measure(qc.qregs[0],qc.cregs[0])
results = qiskit.execute(qc, backend=backend, shots=10000, noise_model=noise_model).result()
noisy_counts = results.get_counts()
print(noisy_counts)
```
In Qiskit we mitigate for the noise by creating a measurement filter object. Then, taking the results from above, we use this to calulate a mitigated set of counts. Qiskit returns this as a dictionary, so that the user doesn't need to use vectors themselves to get the result.
```
# Get the filter object
meas_filter = meas_fitter.filter
# Results with mitigation
mitigated_results = meas_filter.apply(results)
mitigated_counts = mitigated_results.get_counts(0)
```
To see the results most clearly, let's plot both the noisy and mitigated results.
```
from qiskit.visualization import *
plot_histogram([noisy_counts, mitigated_counts], legend=['noisy', 'mitigated'])
```
Here we have taken results for which almost $20\%$ of samples are in the wrong state, and turned it into an exact representation of what the true results should be. However, this example does have just two qubits with a simple noise model. For more qubits, and more complex noise models or data from real devices, the mitigation will have more of a challenge. Perhaps you might find methods that are better than those Qiskit uses!
| github_jupyter |
```
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import numpy as np
from sklearn.model_selection import train_test_split
api_emb = np.load('api_emb.npy')
app_emb = np.load('app_tr_emb.npy') # (# of apps, 64)
bm_tr = np.load('bm_tr.npy') # (# of tr apps, 1000 apis)
counts_tr = np.load('counts_tr.npy')
counts_tst = np.load('counts_tst.npy')
ls
bm_tr.shape, counts_tr.shape, api_emb.shape
X_train, X_val, y_train, y_val = train_test_split(counts_tr, app_emb)
multihot_tr = np.where(counts_tr == 0, counts_tr, 1)
X_train, X_val, y_train, y_val = train_test_split(multihot_tr, app_emb)
X_train, X_val, y_train, y_val = train_test_split(bm_tr, app_emb)
device = torch.device('cuda:0')
X_train = torch.tensor(X_train, dtype=torch.float).to(device)
X_val = torch.tensor(X_val, dtype=torch.float).to(device)
y_train = torch.tensor(y_train, dtype=torch.float).to(device)
y_val = torch.tensor(y_val, dtype=torch.float).to(device)
api_emb.shape
class CustomNet(torch.nn.Module):
def __init__(self):
"""
In the constructor we instantiate two nn.Linear modules and assign them as
member variables.
"""
super(CustomNet, self).__init__()
self.w = torch.nn.Linear(64, 64)
self.api_emb = torch.tensor(api_emb, dtype=torch.float, device=device)
def forward(self, x):
"""
In the forward function we accept a Tensor of input data and we must return
a Tensor of output data. We can use Modules defined in the constructor as
well as arbitrary operators on Tensors.
"""
# app (1, 1000) 0 0 -10 15, 0 0 ...
masked_apis = x.matmul(self.api_emb).float() # (1, 1000) * (1000, 64)
y_pred = self.w(masked_apis) # (1, 64) (64, 64) - > (1, 64)
return y_pred
nn.Embedding
class CustomNet(torch.nn.Module):
def __init__(self):
"""
In the constructor we instantiate two nn.Linear modules and assign them as
member variables.
"""
super(CustomNet, self).__init__()
self.w1 = torch.nn.Linear(64, 128)
self.w2 = torch.nn.Linear(128, 64)
self.api_emb = torch.tensor(api_emb, dtype=torch.float, device=device, requires_grad=False)
def forward(self, x):
"""
In the forward function we accept a Tensor of input data and we must return
a Tensor of output data. We can use Modules defined in the constructor as
well as arbitrary operators on Tensors.
"""
# app (1, 1000) 0 0 -10 15, 0 0 ...
masked_apis = x.matmul(self.api_emb).float() # (1, 1000) * (1000, 64)
l1 = F.leaky_relu(self.w1(masked_apis))
y_pred = self.w2(l1)
return y_pred
model = CustomNet().to(device)
criterion = torch.nn.MSELoss(reduction='mean')
learning_rate = 1e-4
optimizer = torch.optim.Adamax(model.parameters(), lr=learning_rate)
for t in range(20000):
y_pred = model(X_train)
# Compute and print loss
loss = criterion(y_pred, y_train)
# if t % 100 == 99:
# print(t, loss.item())
# Zero gradients, perform a backward pass, and update the weights.
optimizer.zero_grad()
loss.backward()
optimizer.step()
if t % 1000 == 999 or t == 0:
with torch.no_grad():
valid_loss = criterion(model(X_val), y_val)
print(t, loss.item(), valid_loss.item())
# add svm here and print score
0.05 0.07
multihot_tst = np.where(counts_tst == 0, counts_tst, 1)
multihot_tst = torch.tensor(multihot_tst, dtype=torch.float).to(device)
app_tst_emb = model.forward(multihot_tst).cpu().detach().numpy()
bm_tst = np.load('bm_tst.npy')
bm_tst = torch.tensor(bm_tst, dtype=torch.float).to(device)
app_tst_emb = model.forward(bm_tst).cpu().detach().numpy()
counts_tst = np.load('counts_tst.npy')
counts_tst = torch.tensor(counts_tst, dtype=torch.float).to(device)
app_tst_emb = model.forward(counts_tst).cpu().detach().numpy()
import pandas as pd
meta_tr = pd.read_csv('../data/processed/meta_tr.csv', index_col=0)
meta_tst = pd.read_csv('../data/processed/meta_tst.csv', index_col=0)
y_tr = meta_tr.label == 'class1'
y_tst = meta_tst.label == 'class1'
from sklearn.svm import SVC
svm = SVC(kernel='rbf')
svm.fit(app_emb, y_tr)
svm.score(app_emb, y_tr)
svm.score(app_tst_emb, y_tst)
```
| github_jupyter |
# Astronomy 8824 - Numerical and Statistical Methods in Astrophysics
## Statistical Methods Topic II. Bayesian Parameter Estimation
These notes are for the course Astronomy 8824: Numerical and Statistical Methods in Astrophysics. It is based on notes from David Weinberg with modifications and additions by Paul Martini.
David's original notes are available from his website: http://www.astronomy.ohio-state.edu/~dhw/A8824/index.html
#### Background reading:
- Statistics, Data Mining, and Machine Learning in Astronomy, Chapter 5
```
import math
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
from scipy import optimize
from scipy.optimize import minimize
# matplotlib settings
SMALL_SIZE = 14
MEDIUM_SIZE = 16
BIGGER_SIZE = 18
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=BIGGER_SIZE) # fontsize of the x and y labels
plt.rc('lines', linewidth=2)
plt.rc('axes', linewidth=2)
plt.rc('xtick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=MEDIUM_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
```
LaTex macros hidden here --
$\newcommand{\expect}[1]{{\left\langle #1 \right\rangle}}$
$\newcommand{\intinf}{\int_{-\infty}^{\infty}}$
$\newcommand{\xbar}{\overline{x}}$
$\newcommand{\ybar}{\overline{y}}$
$\newcommand{\like}{{\cal L}}$
$\newcommand{\llike}{{\rm ln}{\cal L}}$
$\newcommand{\xhat}{\hat{x}}$
$\newcommand{\yhat}{\hat{y}}$
$\newcommand{\xhati}{\hat{x}_i}$
$\newcommand{\yhati}{\hat{y}_i}$
$\newcommand{\sigxi}{\sigma_{x,i}}$
$\newcommand{\sigyi}{\sigma_{y,i}}$
### Bayesian Parameter Estimation
In Bayesian parameter estimation, one has a model that is assumed to describe the data, and the task is to determine its parameters.
The hypothesis is that the "true value of parameter is $\theta_{\rm true}=\theta$" (discrete) or the "true value of parameter is $\theta \leq \theta_{\rm true} \leq \theta+d\theta$" (continuous). The parameter(s) indicated by $\theta$ are the hypothesis.
$$
p(\theta|DI) = p(\theta|I) {p(D|\theta I) \over p(D|I)}.
$$
If $\theta$ is continuous, then technically the posterior $p(\theta|DI)$ and the prior $p(\theta|I)$ are multiplied by $d\theta$.
A Bayesian searches for the parameter value with **maximum posterior probability** $p(\theta|DI)$.
If $p(\theta|I)$ is flat (a "flat prior"), then this is also the value with **maximum likelihood** $p(D|\theta I)$.
Maximum likelihood estimators play a major role in both Bayesian and classical approaches.
### A simple example: mean of data
The parameter (denoted $\theta$ above) we want to measure is the mean $\mu$. We will determine this from some data $D$, which in this case is $N$ measurements $x_i$. The dispersion $\sigma$ is known, and $x_i$ are Gaussian distributed and independent. (Following Loredo, $\S 5.2.2$; see also Ivezic et al. $\S 5.6.1$)
Assume a flat prior: $p(\mu|I) = (\mu_{\rm max}-\mu_{\rm min})^{-1}$. A flat prior is sometimes also called the "least informative prior." It effectively says that based on prior information, the parameter could be anywhere on the interval with equal probability.
The likelihood (term in numerator) is:
$$\eqalign{
p(\{x_i\}|\mu I) &=
\prod_i (2\pi \sigma^2)^{-1/2} \exp\left[-{(x_i-\mu)^2 \over 2\sigma^2}\right]
\cr
&=
(2\pi \sigma^2)^{-N/2} \exp\left[-{1\over 2\sigma^2}\sum_i (x_i-\mu)^2\right]\cr
&=
(2\pi \sigma^2)^{-N/2} \exp\left[-{Ns^2 \over 2\sigma^2}\right]
\exp\left[-{N\over 2\sigma^2}(\xbar-\mu)^2\right],
}
$$
where $\xbar = {1\over N} \sum x_i$ is sample mean and $s^2 = {1\over N}\sum (x_i-\xbar)^2$ is sample variance. Note that this is not a properly normalized pdf, and in particularly it can be very small for large $N$. It may be useful to think of it as a function of the data.
The global likelihood (term in denominator) is:
$$
p(\{x_i\}|I) = \int_{\mu_{\rm min}}^{\mu_{\rm max}} p(\{x_i\}|\mu I) d\mu
$$
The final result is
$$
p(\mu|\{x_i\} I) d\mu = K \left({N \over 2\pi \sigma^2}\right)^{1/2}
\exp\left[-{N \over 2\sigma^2} (\xbar-\mu)^2\right], \quad
\mu_{\rm min} \leq \mu \leq \mu_{\rm max},
$$
a Gaussian with mean $\xbar$ and dispersion $\sigma/\sqrt{N}$, truncated at $\mu_{\rm min}$ and $\mu_{\rm max}$,
with $K$ a normalization constant such that the probability integrates to one.
The logarithm of the posterior pdf is:
$$
L_p = \ln p(\mu|\{x_i\} I) = \mathrm{constant} - \sum^N_{i=1} \frac{(x_i - \mu)^2}{2 \sigma_i^2}
$$
The maximum likelihood will be at $(dLp/d\mu)_{\mu=\mu_0}= 0$. When we solve for this we find:
$$
0 = 2\mu_0 \sum^N_{i=1} \frac{x_i - \mu}{2 \sigma_i^2} = \mu_0 \sum \frac{x_i}{\sigma_i^2} - \mu_0^2 \sum \frac{1}{\sigma_i}
$$
$$
\mu_0 \sum \frac{x_i}{\sigma_i^2} = \sum \frac{1}{\sigma_i}
$$
or
$$
\mu_0 = \frac{ \sum \frac{x_i}{\sigma_i^2} }{\sum \frac{1}{\sigma_i^2} }
$$
This shows that $\mu_0$ is just the weighted, arithmetic mean of the measurements.
### Some comments on priors
In the above example, as long as the prior range is big compared to $\sigma/\sqrt{N}$, the prior doesn't matter.
Otherwise it does, by truncation and normalization $K>1$.
If new measurements come in, they can be incorporated by taking the output of this result as a _prior_ for new analysis.
At least at an informal level, this is often done, e.g., $H_0$ priors on CMB analyses.
To have the posterior probability $p(\theta|DI) \propto p(D|\theta I)$ (the likelihood), we need the prior $p(\theta|I)$ to be flat in the range allowed by the data, not universally.
For example, we may know that $\mu > 0$ on physical grounds. If $\bar{x} \gg \sigma/\sqrt{N}$ (the sample mean is large compared to the uncertainty in the mean), then $p(\mu|I)$ is approximately flat in the allowable range if it is "broad" compared to $\sigma/\sqrt{N}$. But if $\bar{x} \sim \sigma/\sqrt{N}$, then a flat prior cannot be a good approximation.
For a positive-definite parameter where we have essentially no prior knowledge about its value, a common choice of prior is $p(\theta|I) \propto 1/\theta$, i.e., flat in $\ln\theta$
instead of $\theta$ itself.
```
fig, axarr = plt.subplots(1, 2, figsize=(12, 5))
sig = 1
x = np.random.normal(5, sig, 1000)
sig_x = sig*np.ones(len(x))
sig2 = np.power(sig_x, 2)
# weighted mean
meanx = np.sum(x/sig2)/np.sum(1/sig2)
axarr[0].hist(x)
axarr[0].set_xlim(-3, 10)
axarr[0].plot([0, 0], [0, 300], 'k--')
axarr[0].plot([meanx, meanx], [0, 300], 'k:')
y = np.random.normal(0.2, 1, 1000)
# weighted mean
meany = np.sum(y/sig2)/np.sum(1/sig2)
axarr[1].hist(y)
axarr[1].set_xlim(-3, 10)
axarr[1].plot([0, 0], [0, 300], 'k--')
axarr[1].plot([meany, meany], [0, 300], 'k:')
# Note that in the plot on the right, the uncertainty on the mean is such that plausible values of the mean
# would conflict with a positive-definite prior
```
### Maximum Posterior vs. Maximum Likelihood
From a Bayesian point of view, the end result of a parameter estimation calculation _is_ the posterior probability distribution $p(\theta | DI)$. Recall that
$$
p(\theta|DI) = p(\theta|I) {p(D|\theta I) \over p(D|I)}.
$$
For a flat prior $p(\theta|I)$, the posterior is proportional to the likelihood $P(D|\theta I)$.
Frequentist parameter estimation methods often focus on maximum likelihood estimators, so there is much in common
between frequentist and Bayesian approaches. Parameter estimates based on $p(\theta |DI)$ or $p(D|\theta I)$ will
be similar if $p(\theta | I)$ is flat in the region of parameter space allowed by the data.
If you give an expression for, table of, or plot of the likelihood function, then you have presented all of the evidence of the data, and others can apply prior probabilities or frequentist assessments
as they wish. Thus, if statistics are important to your answer, there is much to be said for presenting things this way if you can.
### A maximum likelihood example: weighted mean
Suppose that we are estimating the mean from $N$ data points that have different errors ("heteroscedastic" data):
The likelihood is
$$
p(\{x_i\}|\mu I) =
\prod_i (2\pi \sigma_i^2)^{-1/2}
\exp\left[-{(x_i-\mu)^2 \over 2\sigma_i^2}\right]~.
$$
Now take the log and set $d\ln L(\mu)/d\mu = 0$.
A few-line derivation shows that the maximum likelihood estimator is
$$
\hat{\mu}_w = {1 \over \sum_i 1/\sigma_i^2}\sum_i {x_i \over \sigma_i^2}~.
$$
The contribution of each data point is weighted by its inverse variance.
The variance of this estimator is
$$
{\rm Var}(\hat{\mu}_w) = {1 \over \sum_i 1/\sigma_i^2}~.
$$
### Maximum likelihood vs. maximum posterior example: mean of Poisson data
(From Bailer-Jones, _Practical Bayesian Inference_, $\S 4.4.5$)
Recall that a Poisson distribution describes the probability of observing $k$ events in some interval, given some expected average number $\lambda$:
$$
P(k) = \frac{\lambda^k \mathrm{e}^{-\lambda}}{k!}
$$
Suppose we have $N$ datapoints ${y_i}$ drawn from a Poisson distribution with unknown $\lambda$. For example, we might have a few X-ray photons detected from an astronomical source, and we want to estimate its flux.
We have (for some reason) a prior $P(\lambda) = \exp(-\lambda/a)$ where $a$ is known.
First we will compute the Maximum Likelihood:
$$
L(\lambda) =
\prod_i {e^{-\lambda} \lambda^{y_i} \over y_i!}~.
$$
Thus
$$
\ln L(\lambda) = \sum_i [y_i\ln\lambda - \lambda - \ln(y_i!)] ~.
$$
Setting $d\ln L(\mu)/d\lambda = 0$ gives
$$
\sum_i \left({y_i \over \lambda}-1\right) = 0
\quad \Longrightarrow \quad
\lambda = {1 \over N} \sum_i y_i.
$$
The maximum likelihood estimate is simply the mean of the data.
Now we will compute the maximum posterior estimate. To do this we maximize:
$$
\ln P(\lambda | \{y_i\}) = \ln L(\lambda) + \ln P(\lambda) + {\rm const}
$$
$$
\frac{d}{d\lambda} \left[ \sum y_i \ln \lambda - \lambda - \ln y! + \ln P(\lambda) + {\rm const} \right] = 0
$$
with the normalized prior
$$
P(\lambda) = {1 \over a} e^{-\lambda/a} \quad \Longrightarrow \quad
{d \ln P(\lambda) \over d\lambda} = -{1\over a}~.
$$
Differentiating and setting to zero gives
$$
\sum_i \left({y_i \over \lambda}-1\right) - {1 \over a} = 0
\quad \Longrightarrow \quad
\lambda = {1 \over N+1/a} \sum_i y_i.
$$
The prior causes the maximum posterior estimate to yield a smaller $\lambda$ relative to the maximum likelihood estimate.
As the number of data points increases, the influence of the prior decreases because $1/a$ must become smaller relative to $N$.
In the limit $a \rightarrow \infty$ the prior is flat and the solution goes to the maximum likelihood solution.
### Confidence intervals
We often _summarize_ the results of a calculation with an estimate and a confidence interval. Bayesians seem to prefer the term "credible region" to "confidence interval," but they appear to be nearly interchangeable, although they are based on different concepts of probability.
Typically, one would quote the maximum likelihood (or maximum posterior probability) value as the estimate, though if the likelihood function is far from Gaussian, people sometimes quote the likelihood weighted mean.
The confidence interval is a region of highest likelihood (or posterior probability) and is characterized by the fraction of the probability that it contains.
For a 1-dimensional problem (1 parameter), this is usually straightforward, though even here a complicated likelihood function may have multiple maxima.
For a Gaussian likelihood function,
$$
\llike = \llike_{\rm max} - {1\over 2}\Delta\chi^2,
\qquad
\like = \like_{\rm max} e^{-\Delta\chi^2/2}.
$$
(We will delve further into $\chi^2$ and $\Delta \chi^2$ in the next classes, but for now you can regard $\Delta \chi^2$ as a measure of the deviation of $p(D|\theta I)$ from its maximum value.)
The regions $\Delta\chi^2 \leq 1$, $\Delta\chi^2 \leq 4$, and $\Delta\chi^2 \leq 9$ contain 68.3\%, 95.4\%, and 99.73\% of the probability. Since a Gaussian is $(2\pi\sigma^2)^{-1/2} e^{-x^2/2\sigma^2}$, these values of $\Delta\chi^2$ correspond to $1\sigma$, $2\sigma$, and $3\sigma$ deviations.
For a non-Gaussian likelihood function, it can be useful instead to quote the values where $\like$ falls to some fraction of its maximum value, say 0.1, in which case the parameter value is 10 times less probable than its most probable value. This particular fraction corresponds in the Gaussian case to $2.14\sigma$, since $e^{-2.14^2/2}=0.1$.
If there are multiple parameters, errors on different parameters may be correlated.
Confidence intervals are defined by contours in a multi-dimensional parameter space.
If the likelihood function is a multi-variate Gaussian, then the confidence contours are ellipses, with the direction of the ellipse axes depending on the covariance of the errors in the parameters.
For the 2-d case, the contours $\Delta\chi^2 = 2.30,$ 6.17, and 11.80 enclose 68.3\%, 95.4\%, and 99.73\% of the probability. ($\Delta\chi^2 = 0.21$ contains 99\%.) See the _Numerical Recipes_ chapter on "Modeling of Data" for higher dimensions and more discussion.
These are sometimes referred to as "$1\sigma$", "$2\sigma$", and "$3\sigma$" regions, when there are multiple dimensions this usage is loose at best and can be misleading.
In some cases, a sensible choice of parameters will eliminate or minimize covariance, making results easier to interpret. An obvious case is the slope and intercept of a linear fit. These are usually highly correlated, but the covariance can be eliminated by defining the intercept at an appropriate "pivot point," fitting $y=a(x-x_p)+b$ instead of $y=ax+b$.
```
# Line with N observed values
np.random.seed(1216)
Npts = 20
x = np.linspace(5, 10, Npts)
m = 2
b = 0
y = m*x + b # True values y_i
# Determine a unique sigma for each point
sig_y = np.random.normal(0., 1., Npts)
# Scatter each point based on that standard deviation
yhat = y + sig_y # Observed values yhat_i
p = np.polyfit(x, yhat, 1, cov=True)
fig, axarr = plt.subplots(1, 2, figsize=(12,6))
axarr[0].errorbar(x, yhat, yerr=np.ones(len(x)), fmt='bo', capsize=4, label="Data")
axarr[0].plot(x, p[0][0]*x + p[0][1], 'k:', label="Fit")
axarr[0].set_ylabel("Y")
axarr[0].set_xlabel("X")
axarr[0].set_xlim(-5, 11)
axarr[0].legend(loc='upper left')
print("Covariance Matrix for left panel ")
print(p[1])
print("Note the significant covariance between the slope and intercept (non-zero off-diagonal elements) ")
print("")
yhat = y + sig_y # Observed values yhat_i
mu = np.mean(x)
p = np.polyfit(x-mu, yhat, 1, cov=True)
axarr[1].errorbar(x-mu, yhat, yerr=np.ones(len(x)), fmt='bo', capsize=4, label="Data")
axarr[1].plot(x-mu, p[0][0]*(x-mu) + p[0][1], 'k:', label="Fit")
axarr[1].set_xlim(-5, 11)
axarr[1].set_xlabel(r"X-$\mu$")
print("Covariance Matrix for right panel ")
print(p[1])
print("There is no longer any covariance")
```
### Marginalization
Suppose that we are simultaneously fitting multiple parameters $\theta_i$ but that we would like to know the confidence interval for one of them in particular, e.g., $\theta_1$.
One of the strengths of Bayesian statistics is that it offers a clear way of doing this:
$$
p(\theta_1|DI) = \int p(\{\theta_i\}|DI) d\theta_2 d\theta_3 ...d\theta_n.
$$
This procedure of integrating over "nuisance parameters" is called "marginalization." (The above expression is the marginal pdf of $\theta_1$.)
_The approach doesn't make sense in the frequentist framework because one cannot talk about the
probability of a parameter value._
**Example:** Suppose we have data that we are using to estimate the slope $a$, intercept $b$, and intrinsic scatter $\sigma$ of a linear relation between $x$ and $y$.
If we just want to know the posterior distribution for the slope, we can find it from
$$
p(a|DI) = \intinf db \int_0^\infty d\sigma p(ab\sigma|DI).
$$
We don't have to go down to a single dimension, e.g., if we don't care about the dispersion $\sigma$ but would like to know the joint distribution of $a$ and $b$:
$$
p(ab|DI) = \int_0^\infty d\sigma p(ab\sigma|DI)d\sigma.
$$
Marginalization plays a crucial role in, for example, cosmological analyses of CMB and large scale structure data,
where the cosmological model being fit typically has 6-10 free parameters but we are often interested in learning
about constraints on specific ones, such as $H_0$ or the effective number of neutrino species.
Systematic uncertainties in the measurements can often be treated by introducing a nuisance parameter that describes
them, such as a calibration offset, imposing some prior, and then marginalizing over these nuisance parameters
when fitting for other parameters of physical interest.
Of course, sometimes one astronomer's nuisance is another astronomer's science, and vice versa.
### Straight line fitting: the "standard" case
Determine maximum likelihood values of $a$ and $b$ in a linear fit $y=ax + b$, given data points with known errors on $y$, assuming Gaussian error distribution:
$$
p(\yhat_i | y_i) = (2\pi\sigma^2_{y,i})^{-1/2}
\exp\left[-(\yhat_i-y_i)^2 \over 2\sigma^2_{y,i}\right],
$$
where $y_i$ is the true value, $\sigma_{y, i}$ is the error on $y_i$, and $\yhati$ is the observed value.
Likelihood
$$\eqalign{
\like &= p(\{\yhat_i\}|a,b) = \prod_i p(\yhat_i | ax_i + b) \cr
&=
\prod_i (2\pi\sigyi^2)^{-1/2} \exp\left[-
{(\yhat_i-ax_i-b)^2 \over 2\sigyi^2}\right].
}
$$
It is often convenient to work with the logarithm of the likelihood
$$
\llike = -{1\over 2} \sum {(\yhat_i-ax_i-b)^2 \over 2\sigyi^2} + C,
$$
where $C$ depends on the (known) errors $\sigyi$ but is independent of $a$ and $b$. (In this case C has the term $(2 \pi \sigma^2_{y,i})^{-1/2}$).
The maximum likelihood solution is thus the solution with minimum
$$
\chi^2 = \sum {(\yhat_i-ax_i-b)^2 \over \sigyi^2},
$$
and $\llike = -\chi^2/2 + C$. Note that maximizing the likelihood is the same as minimizing $\chi^2$ **if** the errors are Gaussian.
For this problem, one can find standard analytic expressions (e.g., Numerical Recipes $\S 15.2$) for $a$ and $b$ in terms of the data and error bars by solving the equations that define the maximum of the likelihood function,
$$
{\partial \llike \over \partial a} = 0, \qquad
{\partial \llike \over \partial b} = 0.
$$
For the case of equal and independent errors on each data point, the results can be written:
$$
\hat{a} = {\bar{xy}-\bar{x}\bar{y} \over \bar{x^2}-\bar{x}^2}
$$
and
$$
\hat{b} = \bar{y}-\hat{a}\bar{x}.
$$
We will get formulas for the general case (more general than given in NR) when we discuss general linear models.
### Straight line fitting: a non-standard but very useful case
Now consider a more complicated variation of this problem: fit $\ybar=ax+b$, with measurement errors in $x$ and $y$ _and_ intrinsic scatter in the relation between $y$ and $x$.
A model with intrinsic scatter (here assumed constant from point to point and denoted $\sigma$) is usually more realistic than the commonly adopted, perfect correlation model.
*If* all of the scatters are Gaussian distributed, we have the following probability distributions for the true values, and the observed values of both x and y. There is a distribution for the true values due to the intrinsic scatter, and there are distributions for both $\xhati$ and $\yhati$ due to measurement errors.
$$\eqalign{
p(y_i | x_i) &= (2\pi\sigma^2)^{-1/2}
\exp\left[-(y_i-ax_i-b)^2 \over 2\sigma^2\right] \cr
p(\yhat_i | y_i) &= (2\pi\sigma^2_{y,i})^{-1/2}
\exp\left[-(\yhat_i-y_i)^2 \over 2\sigma^2_{y,i}\right] \cr
p(\xhat_i | x_i) &= (2\pi\sigma^2_{x,i})^{-1/2}
\exp\left[-(\xhat_i-x_i)^2 \over 2\sigma^2_{x,i}\right].
}
$$
In this case we want to maximize
$$
\like = \prod_i p(\yhati|\xhati) \qquad \Longrightarrow \qquad
\llike = \sum_i \ln p(\yhati|\xhati).
$$
So we need the expression for $p(\yhati|\xhati)$. This is an expression for the observed $\yhati$ given the observed $\xhati$.
$$\eqalign{
p(\yhati|\xhati) &= \intinf dy_i \,p(\yhati|y_i)\, p(y_i|\xhati) \cr
&= \intinf dy_i\, p(\yhati|y_i) \intinf dx_i\, p(y_i|x_i)\, p(x_i|\xhati). \cr
}
$$
$p(x_i|\xhati)$ is the probability of the real $x_i$ given the observed $\xhati$.
Now assume a flat prior on $x_i$, $p(x_i)=$const., so that $p(x_i|\xhati) = p(\xhati|x_i)$ (by Bayes' theorem and the requirement that probabilities integrate to one). This assumption is non-trivial, but usually OK because we only require flatness over the range allowed by $\xhati$.
We can now substitute our expressions for the probabilities, and several pages of algebra and integrals lead eventually to the expression
$$
p(\yhat_i | \xhati) = (2\pi)^{-1/2} (\sigma^2 + \sigyi^2 + a^2\sigxi^2)^{-1/2}
\exp\left[-{(\yhati-a\xhati-b)^2 \over 2(\sigma^2+\sigyi^2+a^2\sigxi^2)}
\right].
$$
This expression looks eminently sensible. For $\sigxi=0$, we get a Gaussian whose width is the quadrature sum of the intrinsic and observationalscatter in $y$. Non-zero $\sigxi$ increases the probability of larger
deviation between observed and predicted $y_i$ by allowing the true value of $ax_i+b$ to be closer to $\yhati$
than $a\xhat_i+b$.
A deviation $\Delta y_i/\sigyi$ has similar weight to a deviation $a\Delta x_i/\sigxi$.
If you think of $x$ and $y$ as having different units, then it is obvious that a factor of $a$ is needed to give $\sigyi$ and $a\sigxi$ the same dimensions.
The maximum likelihood solution requires maximizing
$$\eqalign{
\sum_i \ln p(\yhati|\xhati) = &
-{1\over 2} \sum_i \ln(\sigma^2 + \sigyi^2 + a^2\sigxi^2) \cr
& - \sum_i {(\yhati-a\xhati-b)^2 \over 2(\sigma^2 + \sigyi^2 + a^2\sigxi^2)}
+ {\mathrm constant},
}
$$
and thus solving the equations
$$
{\partial \llike \over \partial a} = 0, \qquad
{\partial \llike \over \partial b} = 0, \qquad
{\partial \llike \over \partial \sigma} = 0.
$$
There is a straightforward algebraic solution for $b$,
$$
b = {-\sum_i (a\xhati - \yhati) W_i \over \sum_i W_i},
$$
where the weights are
$$
W_i = {1 \over 2(\sigma^2 + \sigyi^2 + a^2\sigxi^2)}~.
$$
This is just an inverse-variance weighted average of the individual estimates of $b$.
I couldn't find algebraic solutions for $a$ and $\sigma$, but it is straightforward to search a grid of $(a,\sigma)$, finding the best $b$ for each $(a,\sigma)$ from the above equation and evaluating
the overall likelihood.
There are a couple of points worth noting about the likelihood expression.
First, you might naively have thought that with intrinsic scatter as a free parameter, the maximum likelihood solution would be to have a very large intrinsic scatter, since then each deviation would contribute very little to $\chi^2$.
However, while the second term in the likelihood always rewards large $\sigma^2$, the first term penalizes it, basically because the prediction $ax+b$ is diluted by being spread over a large range, so it doesn't get much "credit" when it is close.
If a significant fraction of points have deviations that put them on the exponential tail of the Gaussian, then raising $\sigma$ will increase the likelihood, but once the typical deviation falls to $\sim 1\sigma$, raising $\sigma$ will decrease the likelihood.
This is, of course, what ought to happen. If the prediction is a scatterplot (as happens in the limit of large intrinsic scatter), then it is unlikely to actually have the points lie close to a line.
Second, if we reverse the roles of $y$ and $x$, letting the intrinsic scatter be on $x$ rather than $y$, then the solution for $a$ and $b$ (especially $a$) will be different.
Intrinsic scatter on $y$ is a _different_ hypothesis from intrinsic scatter on $x$, and the corresponding best
fit slopes and intercepts are different.
The difference goes away if $\sigma$ is small compared to the observational errors.
### Example implementation of fitting a straight line to data with heteroscedastic errors and intrinsic scatter
```
### Example of increasing intrinsic scatter
np.random.seed(1216)
Npts = 10
x_i = np.linspace(0, 10, Npts)
a = 1.
b = 0.
sig_int = 1.
y_i = a*x_i + b # True values y_i
sig_x = 0.5
sig_y = sig_x
# Heteroscedastic errors
sig_x_i = np.random.uniform(0.5*sig_x, sig_x, Npts)
sig_y_i = np.random.uniform(0.5*sig_y, sig_y, Npts)
# Homoscedastic errors
# sig_x_i = 0.5*np.ones(Npts) # np.random.uniform(0.25, 0.5, Npts)
# sig_y_i = 0.5*np.ones(Npts) # np.random.uniform(0.25, 0.5, Npts)
def getline(x_i, y_i, sig_x_i, sig_y_i, sig):
xhat_i = x_i.copy()
yhat_i = y_i.copy()
# Apply the scatter to x and y
for i in range(Npts):
# apply the error to each point, and add intrinsic scatter to yhat_i
err_y = np.random.normal(0, sig_y_i[i])
xhat_i[i] += np.random.normal(0, sig_x_i[i])
yhat_i[i] += np.random.normal(0, np.sqrt(err_y*err_y + sig*sig) )
return xhat_i, yhat_i
sig_int = 0.1
lab = "{:.1f}".format(sig_int)
xhat_i, yhat_i = getline(x_i, y_i, sig_x_i, sig_y_i, sig=sig_int)
fig, axarr = plt.subplots(1, 3, figsize=(14,5), sharey=True)
axarr[0].errorbar(xhat_i, yhat_i, xerr=sig_x_i, yerr=sig_y_i, fmt='bo', capsize=4, label=lab) #".format(sig))
axarr[0].plot(x_i, y_i, 'k:')
axarr[0].set_ylabel("Y")
axarr[0].set_xlabel("X")
axarr[0].legend(loc='upper left')
sig_int = 2
lab = "{:.1f}".format(sig_int)
xhat_i, yhat_i = getline(x_i, y_i, sig_x_i, sig_y_i, sig=sig_int)
axarr[1].errorbar(xhat_i, yhat_i, xerr=sig_x_i, yerr=sig_y_i, fmt='bo', capsize=4, label=lab)
axarr[1].plot(x_i, y_i, 'k:')
axarr[1].set_xlabel("X")
axarr[1].legend(loc='upper left')
sig_int = 5
lab = "{:.1f}".format(sig_int)
xhat_i, yhat_i = getline(x_i, y_i, sig_x_i, sig_y_i, sig=sig_int)
axarr[2].errorbar(xhat_i, yhat_i, xerr=sig_x_i, yerr=sig_y_i, fmt='bo', capsize=4, label=lab)
axarr[2].plot(x_i, y_i, 'k:')
axarr[2].set_xlabel("X")
axarr[2].legend(loc='upper left')
plt.tight_layout()
### Example of fitting a line to data with heteroscedastic errors and intrinsic scatter
np.random.seed(1216)
Npts = 100
x_i = np.linspace(0, 10, Npts)
a = 1.
b = 2.
sig_int = 1.0
y_i = a*x_i + b # True values y_i
sig_x = 0.5
sig_y = sig_x
# Heteroscedastic errors
sig_x_i = np.random.uniform(0.5*sig_x, sig_x, Npts)
sig_y_i = np.random.uniform(0.5*sig_y, sig_y, Npts)
# Homoscedastic errors
# sig_x_i = 0.5*np.ones(Npts) # np.random.uniform(0.25, 0.5, Npts)
# sig_y_i = 0.5*np.ones(Npts) # np.random.uniform(0.25, 0.5, Npts)
lab = "{:.1f}".format(sig_int)
xhat_i, yhat_i = getline(x_i, y_i, sig_x_i, sig_y_i, sig=sig_int)
def lnprob(theta, xhat_i, yhat_i, sig_x_i, sig_y_i):
'''
observations: xhat_i, yhat_i
uncertainties: sig_x_i, sig_y_i
model parameters:
am * x + bm with intrinsic scatter sigma
'''
am, bm, sigm = theta
lprob = -0.5*np.log(sigm*sigm + sig_y_i*sig_y_i + am*am*sig_x_i*sig_x_i) - \
0.5*np.power( yhat_i - am*xhat_i - bm, 2)/(sigm*sigm + sig_y_i*sig_y_i + am*am*sig_x_i*sig_x_i)
return np.sum(lprob)
func = lambda *args: -lnprob(*args)
x0 = np.array([a, b, sig_int]) + np.random.randn(3)
print("Starting guess ", x0)
# BFGS is the Broyden, Fletcher, Goldfarb, and Shanno algorithm. It is a second-order method.
# It approximates the second derivative (Hessian)
result = minimize(func, x0, args=(xhat_i, yhat_i, sig_x_i, sig_y_i), method="BFGS")
errs = np.sqrt(np.diag(result.hess_inv)) # The inverse of the Hessian matrix is the covariance matrix
print("Result for slope, intercept, and intrinsic scatter: ", result.x)
print("Uncertainties on the result: {0:.2f} {1:.2f} {2:.2f}".format(errs[0], errs[1], errs[2]))
print("Input values: ", a, b, sig_int)
# Plot the result and compare to the input
plt.figure(figsize=(8, 8))
plt.errorbar(xhat_i, yhat_i, xerr=sig_x_i, yerr=sig_y_i, fmt='bo')
ydraw = result.x[0]*x_i + result.x[1]
plt.plot(x_i, ydraw, 'b', label="Fit")
plt.plot(x_i, y_i, 'r', label="Input")
plt.xlabel("X")
plt.ylabel("Y")
plt.legend()
result
```
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import tensorflow as tf
print(tf.__version__)
import numpy as np
import matplotlib.pyplot as plt
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/Sunspots.csv \
-O /tmp/sunspots.csv
import csv
time_step = []
sunspots = []
with open('/tmp/sunspots.csv') as csvfile:
reader = csv.reader(csvfile, delimiter=',')
next(reader)
for row in reader:
sunspots.append(float(row[2]))
time_step.append(int(row[0]))
series = np.array(sunspots)
time = np.array(time_step)
plt.figure(figsize=(10, 6))
plot_series(time, series)
split_time = 3000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
window_size = 60
batch_size = 32
shuffle_buffer_size = 1000
def windowed_dataset(series, window_size, batch_size, shuffle_buffer):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(window_size + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(window_size + 1))
dataset = dataset.shuffle(shuffle_buffer).map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
dataset = windowed_dataset(x_train, window_size, batch_size, shuffle_buffer_size)
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(20, input_shape=[window_size], activation="relu"),
tf.keras.layers.Dense(10, activation="relu"),
tf.keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=tf.keras.optimizers.SGD(lr=1e-7, momentum=0.9))
model.fit(dataset,epochs=100,verbose=0)
forecast=[]
for time in range(len(series) - window_size):
forecast.append(model.predict(series[time:time + window_size][np.newaxis]))
forecast = forecast[split_time-window_size:]
results = np.array(forecast)[:, 0, 0]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, results)
tf.keras.metrics.mean_absolute_error(x_valid, results).numpy()
```
| github_jupyter |
#### Q1(a) Import the necessary packages.
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import scipy.io
from sklearn.preprocessing import StandardScaler
import tensorflow
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
```
#### (b) Import the reuters dataset with the following line of code:
```
from tensorflow.keras.datasets import reuters
```
#### (c) Split the datasets into training and test sets with the following line of code. Specify the num_words = 500, and size of test set = 0.2.(num_words: the most common words appearing in the reuters news.)
```
(x_train, y_train), (x_test, y_test) = reuters.load_data(num_words=500, test_split=0.2)
x_train
```
#### (d) We don’t need to create word tokens since the words in the news are already converted into word indices. Pad the sequences so the all sequences have the same length. Set the length = 100.
```
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
#word_index = reuters.word_index
sequences = np.array(x_train)
padded = pad_sequences(sequences,maxlen=100,padding="pre")
test_sequences = np.array(x_test)
test_padded = pad_sequences(test_sequences,maxlen=100,padding="pre")
```
#### (e) One hot encode y_train and y_test. There are 46 categories.
```
from tensorflow.keras.utils import to_categorical
y_train_final = to_categorical(y_train, 46)
y_test_final = to_categorical(y_test, 46)
```
#### (f) (i) Build a Sequential neural network.
```
model = keras.Sequential([
layers.Embedding(500, 32, input_length=100),
layers.Flatten(),
layers.Dense(512, activation='relu'),
layers.Dense(46, activation='softmax'),
])
```
#### (ii) compile the neural network.
```
model.compile(optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['accuracy'])
```
#### (iii) Fit the model on training set. Use x_test and y_test_final as your validation_data. epochs = 10, batch_size = 32
```
history = model.fit(
padded, y_train_final,
validation_data=(test_padded,y_test_final),
epochs=10,
batch_size=32,
)
test_loss, test_acc = model.evaluate(test_padded, y_test_final, verbose=2)
```
#### (iv) Plot the values of training loss and validation loss over epochs. Use another graph to plot the values of training accuracy and validation accuracy over epochs.
```
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
epochs = range(1, len(loss_values) + 1)
plt.plot(epochs, loss_values, 'bo', label='Training Loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
plt.plot(epochs, acc, 'bo', label='Training Accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
#### (g) (i) Build a Sequential neural network.
```
model = keras.Sequential([
layers.Embedding(500, 32, input_length=100),
layers.LSTM(32),
layers.Dense(512, activation='relu'),
layers.Dense(46, activation='softmax'),
])
```
#### (ii) compile the neural network.
```
model.compile(optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['accuracy'])
```
#### (iii) Fit the model on training set. Use x_test and y_test as your validation_data. epochs = 30, batch_size = 32
```
history = model.fit(
padded, y_train_final,
validation_data=(test_padded,y_test_final),
epochs=30,
batch_size=32,
)
test_loss, test_acc = model.evaluate(test_padded, y_test_final, verbose=2)
```
#### (iv) Plot the values of training loss and validation loss over epochs. Use another graph to plot the values of training accuracy and validation accuracy over epochs.
```
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
epochs = range(1, len(loss_values) + 1)
plt.plot(epochs, loss_values, 'bo', label='Training Loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
plt.plot(epochs, acc, 'bo', label='Training Accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Exploring-and-understanding-data" data-toc-modified-id="Exploring-and-understanding-data-1"><span class="toc-item-num">1 </span>Exploring and understanding data</a></div><div class="lev1 toc-item"><a href="#What-is-tabular-data?" data-toc-modified-id="What-is-tabular-data?-2"><span class="toc-item-num">2 </span>What is tabular data?</a></div><div class="lev1 toc-item"><a href="#Pandas" data-toc-modified-id="Pandas-3"><span class="toc-item-num">3 </span>Pandas</a></div><div class="lev2 toc-item"><a href="#The-Series-and-DataFrame" data-toc-modified-id="The-Series-and-DataFrame-31"><span class="toc-item-num">3.1 </span>The Series and DataFrame</a></div><div class="lev1 toc-item"><a href="#The-process-of-exploratory-data-analysis" data-toc-modified-id="The-process-of-exploratory-data-analysis-4"><span class="toc-item-num">4 </span>The process of exploratory data analysis</a></div><div class="lev2 toc-item"><a href="#The-10,000-foot-view" data-toc-modified-id="The-10,000-foot-view-41"><span class="toc-item-num">4.1 </span>The 10,000-foot view</a></div><div class="lev2 toc-item"><a href="#Univariate" data-toc-modified-id="Univariate-42"><span class="toc-item-num">4.2 </span>Univariate</a></div><div class="lev3 toc-item"><a href="#An-aside:-the-difference-between-standard-deviation-and-sample-deviation" data-toc-modified-id="An-aside:-the-difference-between-standard-deviation-and-sample-deviation-421"><span class="toc-item-num">4.2.1 </span>An aside: the difference between standard deviation and sample deviation</a></div><div class="lev3 toc-item"><a href="#Visualizing-a-DataFrame" data-toc-modified-id="Visualizing-a-DataFrame-422"><span class="toc-item-num">4.2.2 </span>Visualizing a DataFrame</a></div><div class="lev2 toc-item"><a href="#Multivariate" data-toc-modified-id="Multivariate-43"><span class="toc-item-num">4.3 </span>Multivariate</a></div><div class="lev1 toc-item"><a href="#More-advanced-Pandas" data-toc-modified-id="More-advanced-Pandas-5"><span class="toc-item-num">5 </span>More advanced Pandas</a></div><div class="lev2 toc-item"><a href="#Missing-data" data-toc-modified-id="Missing-data-51"><span class="toc-item-num">5.1 </span>Missing data</a></div><div class="lev2 toc-item"><a href="#Chaining-and-grouping" data-toc-modified-id="Chaining-and-grouping-52"><span class="toc-item-num">5.2 </span>Chaining and grouping</a></div>
```
%matplotlib inline
```
# Exploring and understanding data
Today we're going to use a tabular dataset to get hands-on experience with two other core Python data science libraries: pandas and matplotlib. We're going to use these libraries to explore, do statistics on, and visualize different parts of our dataset to get a handle on what's there.
# What is tabular data?
The tabular format is fundamental to data science. We got a taste of tabular data last week when we loaded CSV and JSON data, but let's take a closer look now.
Most people are familiar with tabular data from working with spreadsheet software like Excel. In a table, "records" or "samples" are stored in rows, and "features" or "attributes" are stored in columns. For example, in the `good_movies.csv` dataset that we took a look at last week, there were 7 columns representing the fields `title`, `year`, `oscar_nominations`, `short_summary`, `star_1`, `star_2`, and `star_3` and 4 rows representing the movies *La La Land*, *Moonlight*, *Argo*, and *Gone Girl*.

In other words, the rows are individual movies, and the columns represent pieces of information that we know about each movie.
# Pandas
Pandas is the Swiss Army Knife of data analysis in Python. Built on top of NumPy, Pandas wraps arrays with additional functions and metadata to create data frames, a paradigm for storing tabular data borrowed from R.
## The Series and DataFrame
Let's use Pandas to read and explore the `good_movies.csv` dataset again:
```
import pandas as pd
good_movies = pd.read_csv('data/good_movies.csv')
good_movies.head()
```
Pandas automatically gives each row an integer index that guarantees the row can be uniquely identified, but otherwise, the data is exactly the same. The `good_movies.head()` method prints out a few rows from the "head" (top) of the dataset. Since there were only 4 rows in this dataset, `head` prints them all.
There are lots of other ways of reading in data as well (we won't cover these):
- `read_json`
- `read_excel`
- `read_sql`
- `read_html`
- `read_clipboard`
- `read_pickle`
How exactly does Pandas hold the data?
```
type(good_movies)
```
One of the fundamental data structures in Pandas is the DataFrame, which stores 2-dimensional (i.e. tabular/matrix) data. The Pandas DataFrame is basically an ordered collection of heterogeneous *Series* of 1-dimensional data.
There are a few core methods for understanding DataFrames that will be important to understand. We already saw `head()` for print the first several rows of a DataFrame. Some others are:
```
good_movies.shape # dataset has 4 rows and 7 columns
good_movies.info() # tell me some basic info about what's in each column
good_movies.describe() # give some summary statistics for the numeric columns
```
Indexing Pandas DataFrames is a bit different than NumPy. In particular, you can index by the *name* of rows and columns with `loc` or by their *index* with `iloc`. For example, if we wanted to see the summary of *Gone Girl*, we could use:
```
gone_girl_summary = good_movies.loc[3, 'short_summary']
print(gone_girl_summary)
gone_girl_summary = good_movies.iloc[3, 3]
print(gone_girl_summary)
```
**Question**: Why did the first value in the index argument not change?
Or, instead, we can retrieve *all* of the summaries:
```
summaries = good_movies.loc[:, 'short_summary']
print(summaries)
summaries = good_movies.iloc[:, 3] # short_summaries is the third column
print(summaries)
```
Speaking of columns, how are these columns from our dataset being stored after we extract them from our original DataFrame?
```
type(summaries)
```
The `Series` object is another fundamental data type in Pandas. `Series` objects store 1-dimensional (i.e. vector) data, like a single column of a `DataFrame`.
For dealing with tabular data, `Series` and `DataFrames` are much more powerful than pure NumPy arrays. For example, we're not forced to index rows by integers; we can specify a column (as long as it contains unique elements) to use as an index:
```
good_movies = pd.read_csv('data/good_movies.csv', index_col='title')
good_movies.head()
good_movies.loc['Gone Girl', 'short_summary']
summaries = good_movies.loc[:, 'short_summary']
summaries.loc['Gone Girl']
```
How is data stored in each of these series?
```
type(summaries.values)
```
It's also easy to filter rows on certain conditions:
```
good_movies[good_movies['oscar_nominations'] > 5]
```
It's also very convenient to do arithmetic and summary statistics on the data:
```
good_movies['oscar_nominations'].count()
good_movies['oscar_nominations'].sum()
good_movies['oscar_nominations'].mean()
good_movies['oscar_nominations'].median()
good_movies['oscar_nominations'].std()
```
**Your Turn**

For the rest of the class, we're going to dive deep into the passengers of the *RMS Titanic*. The file `data/titanic.csv` contains the following data on each of the passengers:
- pclass: Passenger class (1 = first; 2 = second; 3 = third)
- survived: Survival (0 = no; 1 = yes)
- name: Name
- sex: Sex
- age: Age
- sibsp: Number of siblings/spouses aboard
- parch: Number of parents/children aboard
- ticket: Ticket number
- fare: Passenger fare
- cabin: Cabin number
- embarked: Port of embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
- boat: Lifeboat (if survived)
- body: Body number (if did not survive and body was recovered)
Questions to answer:
- Read `data/titanic.csv` into a Pandas DataFrame. What is the best index for rows?
- What was the average and median ages of passengers?
- What was the overall survival rate?
- What was the price of the most expensive ticket? Whose tickets were they?
- How many passengers had no family on board?
```
titanic = pd.read_csv('data/titanic.csv', index_col=None)
```
# The process of exploratory data analysis
When we get a new tabular dataset that we don't know anything about, there is a fairly standard process of exploration that we can do on that dataset to get a better handle on it:
- Get some really general information about the dataset as a whole
- Dig into each column individually
- Try to understand the relationships *between* columns
To illustrate the process, we'll use a mystery dataset that we don't know anything about:
```
mystery_data = pd.read_csv('data/mystery.csv', index_col=None)
```
## The 10,000-foot view
A few questions we should ask:
- How many rows are there?
- How many columns are there?
- What are the datatypes of each column?
- How many missing values (if any) are there?
The Pandas built-in `.info()` method gives a great, quick answer to these questions:
```
mystery_data.info()
```
And we can use head to look at some actual values in the columns:
```
mystery_data.head()
```
## Univariate
For each column, we should then ask:
- What are the *central tendancies* of the column?
- "Central tendancies" = "typical" value(s)
- Especially - Mean, median, mode
- What is the *spread* of the data?
- Often, variance/standard deviation
- But can also be: quartiles (or other percentiles), min/max values, etc
- What the distribution *look like*?
Again, Pandas provides a convenient way of looking at the usual univariate statistics:
```
mystery_data.describe()
for col in mystery_data.columns:
print('median of col {}: {}'.format(col, mystery_data.loc[:, col].median()))
```
What does it mean for a column to have a median and mean that are different?
### An aside: the difference between standard deviation and sample deviation
Sometimes, you'll see two slightly different definitions for the "spread" of a column of data - the standard deviation:
$\sigma_X = \sqrt{ \frac{1}{n}\sum_i^n (x_i - \mu_X) }$
and the sample deviation:
$\sigma_X = \sqrt{ \frac{1}{n-1}\sum_i^n (x_i - \mu_X) }$
(where $\mu$ is the mean of the column $X$ and $x_i$ is the $i$th value of $X$). What's the difference between dividing by $n$ and by $n-1$?
The *standard deviation* is used when $X$ contains the *entire population* that you're estimating the width of. So, for example, if you want to know the spread of test scores in a class and you have the score of each and every student, use the standard deviation.
The *sample deviation* is used to correct for bias when you're trying to estimate the width of a population that you only have a sample of data points from. For example, if you are trying to extrapolate the spread of test scores on the SAT by looking at how 1,000 students performed, you should use the *sample deviation*.
In practice, however, standard deviation and sample deviation are very similar if the number of samples is large.
### Visualizing a DataFrame
Summary statistics can only take us so far in understanding the data in a new dataset. Visualizations like histograms are another great way of getting a high-level overview for what the data looks like.
Plotting functionality is built into Pandas. In particular, the built-in plot functions are a thin wrapper around a very powerful data visualization library called matplotlib. We won't directly use matplotlib in this course; we'll do all of our plotting by calling the Pandas wrapper functions. The pure matplotlib functions are very similar, however.
```
# {dataframe_name}.{column_name}.plot.{plot_type}(plotting options)
mystery_data.A.plot.hist(bins=30)
print('mean: {}'.format(mystery_data.A.mean()))
print('median: {}'.format(mystery_data.A.median()))
print('sample deviation: {}'.format(mystery_data.A.std()))
mystery_data.B.plot.hist(bins=30)
print('mean: {}'.format(mystery_data.B.mean()))
print('median: {}'.format(mystery_data.B.median()))
print('sample deviation: {}'.format(mystery_data.B.std()))
mystery_data.C.plot.hist(bins=30)
print('mean: {}'.format(mystery_data.C.mean()))
print('median: {}'.format(mystery_data.C.median()))
print('sample deviation: {}'.format(mystery_data.C.std()))
```
**Your turn**
Try following the exploratory data analysis steps on the Titanic dataset (i.e. get the 10,000-foot view, then get some column-by-column insights). What interesting things do you find?
## Multivariate
After understanding what is contained in *each individual column*, it's important to understand how each column is related to the others. The related ideas of *correlation* and *covariance* are common ways of understanding pairwise dependencies between columns.
The *covariance* measures, unsurprisingly, the extent to which two columns co-vary. In other words, how often do values with a large distance from the mean in one column correspond to a large distance from the mean in another column?
The covariance between columns $X$ and $Y$ is defined as:
$ cov(X, Y) = \frac{1}{n} \sum_{i=1}^n (x_i - \mu_X)(y_i - \mu_Y) $
Think about what it means for covariance between 2 columns to be large vs small.
One problem with covariance is that comparing covariances between different columns can be tricky - columns that are naturally wider will tend to have larger covariances. The *correlation* between $X$ and $Y$ is a similar concept to covariance, but that corrects for the difference in widths:
$ corr(X, Y) = \frac{ cov(X, Y) }{ \sigma_X \sigma_Y} $
The correlation is always a number between -1 and 1, making it easy to interpret.
```
mystery_data.corr()
```
**Your turn**
Calculate the correlation between pairs of columns in the Titanic dataset. Which columns correlate most strongly with the passengers who survived? What other relatively strong correlations do you find?
# More advanced Pandas
## Missing data
Unfortunately, missing data is a fact of life, so being able to easily deal with blank values in our datasets is crucial. Pandas treats missing values are true "first class citizens" in datasets:
```
titanic.head()
```
Missing values are represented by `NaN`:
```
titanic.loc[0, 'body']
type(titanic.loc[0, 'body'])
```
And "nothingness" carries forward if you do mathematical operations on missing values:
```
titanic.loc[0, 'body'] + 5
```
...which matches intuition.
One way of dealing with missing values is to just ignore it!
```
titanic_none_missing = titanic.dropna()
titanic_none_missing.info()
```
By default, `dropna` drops a row from the dataset if *any value* is missing. This turned out to be a problem, since it looks like there weren't any rows that didn't have missing values!
Instead, maybe we just want to drop the rows where `body` is missing:
```
# the `subset` argument allows us to only consider certain columns
titanic_body_not_missing = titanic.dropna(subset=['body'])
titanic_body_not_missing.info()
```
By doing this, we discover that none of the bodies recovered had been assigned to boats.
Sometimes, though, just ignoring missing data is a bit too aggressive. Instead, sometimes we just want to fill in the missing data with other data:
```
# `inplace` argument lets us change the data without creating a new object
titanic['body'].fillna('Person either survived or their body was not recovered.', inplace=True)
titanic.head()
```
**Your turn**
Using the Pandas `notnull` method, make a histogram of the values in the `age` column that aren't missing. Then, try filling the missing values in the `age` column with the column's average and median. Plot a histogram of each filled column - how do the histograms differ?
The bottom line is - no matter how you choose to deal with your missing data, there will be trade-offs. Proceed with caution.
## Chaining and grouping
https://github.com/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/03.08-Aggregation-and-Grouping.ipynb
https://blog.socialcops.com/engineering/machine-learning-python/
The idea of *method chaining* is prominently featured in Pandas. Method chaining allows to perform several operations on a dataset in one fell swoop, making it much cleaner to code up multi-step analysis that we might want to perform. For example, instead of writing:
```
titanic_temp = titanic.dropna(subset=['body'])
titanic_temp.describe()
```
...we can write simply:
```
titanic.dropna(subset=['body']).describe()
```
The result is the same, but we avoided using ugly placeholder variables and extra lines of code.
Often, instead of looking at statistics of an entire dataset, we want to look at statistics of groups within the dataset. For example, we can easily calculate the survival rate of *all* passengers on the Titanic (recall `survived` = 0 if the person did not survive and 1 if they survived),
```
titanic['survived'].mean()
```
...but it might also be interesting to see the survival rate broken out by gender. If you've used SQL before, this type of command might look familiar:
```
titanic.groupby('sex')['survived'].mean()
```
What happened here? The `groupby` method groups all of the data in our dataset into groups based on which gender each person was. We then select only the `survived` column and calculate the mean.
**Your turn**
Use grouping, selecting, and aggregating to answer the following questions:
- What was the survival rate of first class passengers? What about third class?
- What percentage of the total fares paid was paid by passengers in first class?
- What port of embarkation had the highest average ticket price?
| github_jupyter |
# Regression Task with Late Data Integration
```
# Import pandas and numpy libraries for data analysis
import numpy as np
import pandas as pd
from sklearn.preprocessing import StandardScaler
# Import the metric to calculate mean squared error
from sklearn.metrics import mean_squared_error
# Import Tensorflow libraries for deep learning
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense, Dropout, Input, Concatenate
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.constraints import max_norm
# Define the neural network model to use with only one view
def init_model(input_dim, learning_rate, epochs, momentum, neurons, trainable=True):
input = Input(shape=(input_dim,))
layer = Dense(neurons, activation='sigmoid',
kernel_constraint=max_norm(3)) (input)
layer = Dropout(rate=0.6) (layer)
layer = Dense(neurons, activation='sigmoid',
kernel_constraint=max_norm(3)) (layer)
layer = Dropout(rate=0.6) (layer)
predictions = Dense(1, activation='linear') (layer)
model = Model(inputs=input, outputs=predictions)
rms = SGD(lr=learning_rate, decay=learning_rate / epochs,
momentum=momentum)
model.trainable = trainable
if (trainable) :
model.compile(loss='mean_squared_error', optimizer=rms, metrics=['mean_squared_error'])
return model
# Define the neural network to use in a multi=view fashion
def init_multi_model(input_dim,input_dim2, learning_rate, epochs, momentum, neurons, metabolic_fluxes_layer,
gene_expression_layer):
metabolic_fluxes_input = Input(shape=(input_dim,))
gene_expression_input = Input(shape=(input_dim2,))
comb_layer = Concatenate()([metabolic_fluxes_layer(metabolic_fluxes_input),
gene_expression_layer(gene_expression_input)])
comb_layer = Dense(neurons, activation='sigmoid', kernel_constraint=max_norm(3)) (comb_layer)
predictions = Dense(1, activation='linear') (comb_layer)
model = Model(inputs=[metabolic_fluxes_input, gene_expression_input], outputs=predictions)
rms = SGD(lr=learning_rate, decay=learning_rate / epochs, momentum=momentum)
model.compile(loss='mean_squared_error', optimizer=rms, metrics=['mean_squared_error'])
return model
# Specify the proportion of data to be used as the test set
percent_test = 0.3
# Import flux data
metabolic_data = pd.read_csv('fluxes.csv', encoding='utf-8')
# Disregard null fluxes
metabolic_data = metabolic_data.loc[:, (metabolic_data.abs() >= 1e-7).any(axis=0)]
# Import gene expression data
gene_expression_data = pd.read_csv('gene_expression_data.csv', encoding='utf-8')
X = gene_expression_data[gene_expression_data.columns[:-1]]
Y = gene_expression_data[gene_expression_data.columns[-1]]
# Split gene expression data into training and test sets
gene_expression_train, gene_expression_test, Y_train, Y_test = train_test_split(X, Y, test_size=percent_test, shuffle=False)
# Split flux data into training and test sets
metabolic_fluxes_train, metabolic_fluxes_test = train_test_split(X, test_size=percent_test, shuffle=False)
# Perform feature scaling to normalize the training data
stdscaler_f = StandardScaler()
stdscaler_g = StandardScaler()
metabolic_fluxes_train = stdscaler_f.fit_transform(metabolic_fluxes_train)
gene_expression_train = stdscaler_g.fit_transform(gene_expression_train)
# Normalize the test sets with the same parametric values as the training sets
metabolic_fluxes_test = stdscaler_f.transform(metabolic_fluxes_test)
gene_expression_test = stdscaler_g.transform(gene_expression_test)
# Define the number of epochs, batches, learning rate and validation set split size
epochs = 6000
batches = 256
lrate = 0.005
validation = 0.2
lrate2 = 0.05
epochs2 = 500
# Define the stochastic gradient descent algorithm and the early stopping strategy to prevent overfitting
rms = SGD(lr=lrate , decay=lrate / epochs, momentum=0.75)
earlyStopping=EarlyStopping(monitor='val_loss', patience=15000, verbose=0, mode='auto')
# Initialize separate single=view models for the gene expression and flux datasets
model_gene_expression = init_model(gene_expression_train.shape[1], lrate, 3000, 0.75, 1000)
model_metabolic_fluxes = init_model(metabolic_fluxes_train.shape[1], lrate, 3000, 0.75,1000)
# Fit both the learning models on the training data
model_gene_expression.fit(x=gene_expression_train, y=Y_train, epochs=epochs, batch_size=batches,
validation_split=validation, callbacks=[earlyStopping], verbose=0)
model_metabolic_fluxes.fit(x=metabolic_fluxes_train, y=Y_train, epochs=epochs, batch_size=batches,
validation_split=validation, callbacks=[earlyStopping], verbose=0)
# Remove the last layer from each single view model
model_gene_expression.layers.pop()
model_gene_expression.layers.pop()
model_gene_expression.outputs = [model_gene_expression.layers[-1].output]
model_metabolic_fluxes.layers.pop()
model_metabolic_fluxes.layers.pop()
model_metabolic_fluxes.outputs = [model_metabolic_fluxes.layers[-1].output]
# Initialize the multi=modal model
multi_modal_model = init_multi_model(metabolic_fluxes_train.shape[1], gene_expression_train.shape[1], lrate2, epochs2,
0.75, 15, model_metabolic_fluxes, model_gene_expression)
# Fit the multi=modal model using training samples
multi_modal_model.fit(x=[metabolic_fluxes_train, gene_expression_train], y=Y_train, epochs=epochs2, batch_size=batches,
validation_split=validation, verbose=0)
# Generate predictions for the test set samples
predictions = multi_modal_model.predict([metabolic_fluxes_test, gene_expression_test])
# Print mean squared error
print('MSE: ', mean_squared_error(predictions, Y_test))
```
| github_jupyter |
<table> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by <a href="http://abu.lu.lv" target="_blank">Abuzer Yakaryilmaz</a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
<h2> <font color="blue"> Solution for </font>Freivalds</h2>
<a id="task3"></a>
<h3> Task 3 </h3>
Freivalds reads 50 random strings of length 40.
Find the final probabilistic state for each string.
Is there any relation between
<ul>
<li>the numbers of $ a $'s and $ b $'s, say $ N_a $ and $ N_b $, and </li>
<li>the probabilities of the first bit being in zero and one, say $ p_0 $ and $ p_1 $?</li>
</ul>
More specifically:
<ul>
<li> When $ N_a > N_b $, is $ p_0 < p_1 $ or $ p_0 > p_1 $? </li>
<li> When $ N_a < N_b $, is $ p_0 < p_1 $ or $ p_0 > p_1 $? </li>
</ul>
Or simply check the signs of $ (N_a - N_b) $ and $ (p_0-p_1) $ for each string.
<i>Hint: The multiplication of two numbers with the same signs is a positive number, and the multiplication of two numbers with the opposite signs gives a negative number.</i>
<h3>Solution</h3>
```
# for random number generation
from random import randrange
# we will use evolve function
def evolve(Op,state):
newstate=[]
for i in range(len(Op)): # for each row
newstate.append(0)
for j in range(len(state)): # for each element in state
newstate[i] = newstate[i] + Op[i][j] * state[j] # summation of pairwise multiplications
return newstate # return the new probabilistic state
# the initial state
state = [0.5, 0, 0.5, 0]
# probabilistic operator for symbol a
A = [
[0.5, 0, 0, 0],
[0.25, 1, 0, 0],
[0, 0, 1, 0],
[0.25, 0, 0, 1]
]
# probabilistic operator for symbol b
B = [
[1, 0, 0, 0],
[0, 1, 0.25, 0],
[0, 0, 0.5, 0],
[0, 0, 0.25, 1]
]
#
# your solution is here
#
length = 40
total = 50
# total = 1000 # we will also test our code for 1000 strings
# we will check 5 cases
# let's use a list
cases = [0,0,0,0,0]
for i in range(total): # total number of strings
Na = 0
Nb = 0
string = ""
state = [0.5, 0, 0.5, 0]
for j in range(length): # generate random string
if randrange(2) == 0:
Na = Na + 1 # new symbol is a
string = string + "a"
state = evolve(A,state) # update the probabilistic state by A
else:
Nb = Nb + 1 # new symbol is b
string = string + "b"
state = evolve(B,state) # update the probabilistic state by B
# now we have the final state
p0 = state[0] + state[1] # the probabilities of being in 00 and 01
p1 = state[2] + state[3] # the probabilities of being in 10 and 11
print() # print an empty line
print("(Na-Nb) is",Na-Nb,"and","(p0-p1) is",p0-p1)
# let's check possible different cases
# start with the case in which both are nonzero
# then their multiplication is nonzero
# let's check the sign of their multiplication
if (Na-Nb) * (p0-p1) < 0:
print("they have opposite sign")
cases[0] = cases[0] + 1
elif (Na-Nb) * (p0-p1) > 0:
print("they have the same sign")
cases[1] = cases[1] + 1
# one of them should be zero
elif (Na-Nb) == 0:
if (p0-p1) == 0:
print("both are zero")
cases[2] = cases[2] + 1
else:
print("(Na-Nb) is zero, but (p0-p1) is nonzero")
cases[3] = cases[3] + 1
elif (p0-p1) == 0:
print("(Na-Nb) is nonzero, while (p0-p1) is zero")
cases[4] = cases[4] + 1
# check the case(s) that are observed and the case(s) that are not observed
print() # print an empty line
print(cases)
```
<b> Interpretation </b>
There are five different cases about $ (N_a-N_b) $ and $ (p_0-p_1) $:
<ul>
<li> $ cases[0] $: they have opposite sign </li>
<li> $ cases[1] $: they have the same sign </li>
<li> $ cases[2] $: both are zero </li>
<li> $ cases[3] $: $ (N_a-N_b) $ is zero, but $ (p_0-p_1) $ is nonzero </li>
<li> $ cases[4] $: $ (N_a-N_b) $ is nonzero, while $ (p_0-p_1) $ is zero </li>
</ul>
<b>However</b>, we observed only two cases: $ cases[0] $ and $ cases[2] $.
(1) If the numbers of $ a $'s and $ b $'s are the same, then the probability of observing $ 0 $ in the first bit (i.e., $\mathbf{0}0$ or $\mathbf{0}1$) and the probability of observing 1 in the first bit (i.e., $\mathbf{1}0$ or $\mathbf{1}1$) are the same.
$$
N_a = N_b \longleftrightarrow p_0 = p_1.
$$
(2) If the numbers of $ a $'s and $ b $'s are not the same, then we have only $ (N_a - N_b) \cdot (p_0-p_1) < 0 $.
(2.a) If the number of $ a $'s is greater than the number of $ b $'s, then the probability of observing $ 0 $ in the first bit (i.e., $\mathbf{0}0$ or $\mathbf{0}1$) is less than the probability of observing 1 in the first bit (i.e., $\mathbf{1}0$ or $\mathbf{1}1$).
$$
N_a > N_b \longrightarrow p_0 < p_1.
$$
(2.b) If the number of $ a $'s is less than the number of $ b $'s, then the probability of observing $ 0 $ in the first bit (i.e., $\mathbf{0}0$ or $\mathbf{0}1$) is greater than the probability of observing 1 in the first bit (i.e., $\mathbf{1}0$ or $\mathbf{1}1$).
$$
N_a < N_b \longrightarrow p_0 > p_1.
$$
<hr>
If we have more $ a $'s, we expect to observe $ 10 $ and $ 11 $ more than $ 00 $ and $ 01 $.
If we have more $ b $'s, we expect to observe $ 00 $ and $ 01 $ more than $ 10 $ and $ 11 $.
If we have equal numbers of $a$'s and $b$'s, we expect to observe $ 0 $ and $ 1 $ in the first bit equally often.
| github_jupyter |
```
#12/29/20
#runnign synthetic benchmark graphs for synthetic OR datasets generated
#making benchmark images
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
from keras.utils.generic_utils import Progbar
from keras.layers.merge import _Merge
import keras.losses
from keras.datasets import mnist
from functools import partial
from collections import defaultdict
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import os
import pickle
import numpy as np
import isolearn.io as isoio
import isolearn.keras as isol
import pandas as pd
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
from sequence_logo_helper import dna_letter_at, plot_dna_logo
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class EpochVariableCallback(Callback) :
def __init__(self, my_variable, my_func) :
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_begin(self, epoch, logs={}) :
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#ONLY RUN THIS CELL ONCE
from tensorflow.python.framework import ops
#Stochastic Binarized Neuron helper functions (Tensorflow)
#ST Estimator code adopted from https://r2rt.com/beyond-binary-ternary-and-one-hot-neurons.html
#See Github https://github.com/spitis/
def st_sampled_softmax(logits):
with ops.name_scope("STSampledSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.squeeze(tf.multinomial(logits, 1), 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
def st_hardmax_softmax(logits):
with ops.name_scope("STHardmaxSoftmax") as namescope :
nt_probs = tf.nn.softmax(logits)
onehot_dim = logits.get_shape().as_list()[1]
sampled_onehot = tf.one_hot(tf.argmax(nt_probs, 1), onehot_dim, 1.0, 0.0)
with tf.get_default_graph().gradient_override_map({'Ceil': 'Identity', 'Mul': 'STMul'}):
return tf.ceil(sampled_onehot * nt_probs)
@ops.RegisterGradient("STMul")
def st_mul(op, grad):
return [grad, grad]
#Gumbel Distribution Sampler
def gumbel_softmax(logits, temperature=0.5) :
gumbel_dist = tf.contrib.distributions.RelaxedOneHotCategorical(temperature, logits=logits)
batch_dim = logits.get_shape().as_list()[0]
onehot_dim = logits.get_shape().as_list()[1]
return gumbel_dist.sample()
#model functions for loading optimus scramblers
import keras.backend as K
def mask_dropout_multi_scale(mask, drop_scales=[1, 2, 4, 7], min_drop_rate=0.0, max_drop_rate=0.5) :
rates = K.random_uniform(shape=(K.shape(mask)[0], 1, 1, 1), minval=min_drop_rate, maxval=max_drop_rate)
scale_logits = K.random_uniform(shape=(K.shape(mask)[0], len(drop_scales), 1, 1, 1), minval=-5., maxval=5.)
scale_probs = K.softmax(scale_logits, axis=1)
ret_mask = mask
for drop_scale_ix, drop_scale in enumerate(drop_scales) :
ret_mask = mask_dropout(ret_mask, rates * scale_probs[:, drop_scale_ix, ...], drop_scale=drop_scale)
return K.switch(K.learning_phase(), ret_mask, mask)
def mask_dropout(mask, drop_rates, drop_scale=1) :
random_tensor_downsampled = K.random_uniform(shape=(
K.shape(mask)[0],
1,
K.cast(K.shape(mask)[2] / drop_scale, dtype=tf.int32),
K.shape(mask)[3]
), minval=0.0, maxval=1.0)
keep_mask_downsampled = random_tensor_downsampled >= drop_rates
keep_mask = K.repeat_elements(keep_mask_downsampled, rep=drop_scale, axis=2)
ret_mask = mask * K.cast(keep_mask, dtype=tf.float32)
return ret_mask
def mask_dropout_single_scale(mask, drop_scale=1, min_drop_rate=0.0, max_drop_rate=0.5) :
rates = K.random_uniform(shape=(K.shape(mask)[0], 1, 1, 1), minval=min_drop_rate, maxval=max_drop_rate)
random_tensor_downsampled = K.random_uniform(shape=(
K.shape(mask)[0],
1,
K.cast(K.shape(mask)[2] / drop_scale, dtype=tf.int32),
K.shape(mask)[3]
), minval=0.0, maxval=1.0)
keep_mask_downsampled = random_tensor_downsampled >= rates
keep_mask = K.repeat_elements(keep_mask_downsampled, rep=drop_scale, axis=2)
ret_mask = mask * K.cast(keep_mask, dtype=tf.float32)
return K.switch(K.learning_phase(), ret_mask, mask)
#PWM Masking and Sampling helper functions
def mask_pwm(inputs) :
pwm, onehot_template, onehot_mask = inputs
return pwm * onehot_mask + onehot_template
def sample_pwm_st(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = st_sampled_softmax(flat_pwm)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
def sample_pwm_gumbel(pwm_logits) :
n_sequences = K.shape(pwm_logits)[0]
seq_length = K.shape(pwm_logits)[2]
flat_pwm = K.reshape(pwm_logits, (n_sequences * seq_length, 4))
sampled_pwm = gumbel_softmax(flat_pwm, temperature=0.5)
return K.reshape(sampled_pwm, (n_sequences, 1, seq_length, 4))
#Generator helper functions
def initialize_sequence_templates(generator, sequence_templates, background_matrices) :
embedding_templates = []
embedding_masks = []
embedding_backgrounds = []
for k in range(len(sequence_templates)) :
sequence_template = sequence_templates[k]
onehot_template = iso.OneHotEncoder(seq_length=len(sequence_template))(sequence_template).reshape((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] not in ['N', 'X'] :
nt_ix = np.argmax(onehot_template[0, j, :])
onehot_template[:, j, :] = -4.0
onehot_template[:, j, nt_ix] = 10.0
elif sequence_template[j] == 'X' :
onehot_template[:, j, :] = -1.0
onehot_mask = np.zeros((1, len(sequence_template), 4))
for j in range(len(sequence_template)) :
if sequence_template[j] == 'N' :
onehot_mask[:, j, :] = 1.0
embedding_templates.append(onehot_template.reshape(1, -1))
embedding_masks.append(onehot_mask.reshape(1, -1))
embedding_backgrounds.append(background_matrices[k].reshape(1, -1))
embedding_templates = np.concatenate(embedding_templates, axis=0)
embedding_masks = np.concatenate(embedding_masks, axis=0)
embedding_backgrounds = np.concatenate(embedding_backgrounds, axis=0)
generator.get_layer('template_dense').set_weights([embedding_templates])
generator.get_layer('template_dense').trainable = False
generator.get_layer('mask_dense').set_weights([embedding_masks])
generator.get_layer('mask_dense').trainable = False
generator.get_layer('background_dense').set_weights([embedding_backgrounds])
generator.get_layer('background_dense').trainable = False
#Generator construction function
def build_sampler(batch_size, seq_length, n_classes=1, n_samples=1, sample_mode='st') :
#Initialize Reshape layer
reshape_layer = Reshape((1, seq_length, 4))
#Initialize background matrix
onehot_background_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='background_dense')
#Initialize template and mask matrices
onehot_template_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='zeros', name='template_dense')
onehot_mask_dense = Embedding(n_classes, seq_length * 4, embeddings_initializer='ones', name='mask_dense')
#Initialize Templating and Masking Lambda layer
masking_layer = Lambda(mask_pwm, output_shape = (1, seq_length, 4), name='masking_layer')
background_layer = Lambda(lambda x: x[0] + x[1], name='background_layer')
#Initialize PWM normalization layer
pwm_layer = Softmax(axis=-1, name='pwm')
#Initialize sampling layers
sample_func = None
if sample_mode == 'st' :
sample_func = sample_pwm_st
elif sample_mode == 'gumbel' :
sample_func = sample_pwm_gumbel
upsampling_layer = Lambda(lambda x: K.tile(x, [n_samples, 1, 1, 1]), name='upsampling_layer')
sampling_layer = Lambda(sample_func, name='pwm_sampler')
permute_layer = Lambda(lambda x: K.permute_dimensions(K.reshape(x, (n_samples, batch_size, 1, seq_length, 4)), (1, 0, 2, 3, 4)), name='permute_layer')
def _sampler_func(class_input, raw_logits) :
#Get Template and Mask
onehot_background = reshape_layer(onehot_background_dense(class_input))
onehot_template = reshape_layer(onehot_template_dense(class_input))
onehot_mask = reshape_layer(onehot_mask_dense(class_input))
#Add Template and Multiply Mask
pwm_logits = masking_layer([background_layer([raw_logits, onehot_background]), onehot_template, onehot_mask])
#Compute PWM (Nucleotide-wise Softmax)
pwm = pwm_layer(pwm_logits)
#Tile each PWM to sample from and create sample axis
pwm_logits_upsampled = upsampling_layer(pwm_logits)
sampled_pwm = sampling_layer(pwm_logits_upsampled)
sampled_pwm = permute_layer(sampled_pwm)
sampled_mask = permute_layer(upsampling_layer(onehot_mask))
return pwm_logits, pwm, sampled_pwm, onehot_mask, sampled_mask
return _sampler_func
#for formulation 2 graphing
def returnXMeanLogits(e_train):
#returns x mean logits for displayign the pwm difference for the version 2 networks
#Visualize background sequence distribution
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 1)) + pseudo_count) / (x_train.shape[0] + 4. * pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
return x_mean_logits, x_mean
#loading testing dataset
from optimusFunctions import *
import pandas as pd
csv_to_open = "optimus5_synthetic_random_insert_if_uorf_1_start_2_stop_variable_loc_512.csv"
dataset_name = csv_to_open.replace(".csv", "")
print (dataset_name)
data_df = pd.read_csv("./" + csv_to_open) #open from scores folder
seq_e_test = one_hot_encode(data_df, seq_len=50)
benchmarkSet_seqs = seq_e_test
x_test = np.reshape(benchmarkSet_seqs, (benchmarkSet_seqs.shape[0], 1, benchmarkSet_seqs.shape[1], benchmarkSet_seqs.shape[2]))
print (x_test.shape)
e_train = pd.read_csv("bottom5KIFuAUGTop5KIFuAUG.csv")
print ("training: ", e_train.shape[0], " testing: ", x_test.shape[0])
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_mean_logits, x_mean = returnXMeanLogits(e_train)
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
#background
def returnXMeanLogits(e_train):
#returns x mean logits for displayign the pwm difference for the version 2 networks
#Visualize background sequence distribution
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 1)) + pseudo_count) / (x_train.shape[0] + 4. * pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
return x_mean_logits, x_mean
e_train = pd.read_csv("bottom5KIFuAUGTop5KIFuAUG.csv")
print ("training: ", e_train.shape[0], " testing: ", x_test.shape[0])
#one hot encode with optimus encoders
seq_e_train = one_hot_encode(e_train,seq_len=50)
x_mean_logits, x_mean = returnXMeanLogits(e_train)
x_train = seq_e_train
x_train = np.reshape(x_train, (x_train.shape[0], 1, x_train.shape[1], x_train.shape[2]))
#Define sequence template for optimus
sequence_template = 'N'*50
sequence_mask = np.array([1 if sequence_template[j] == 'N' else 0 for j in range(len(sequence_template))])
#Visualize background sequence distribution
save_figs = True
plot_dna_logo(np.copy(x_mean), sequence_template=sequence_template, figsize=(14, 0.65), logo_height=1.0, plot_start=0, plot_end=50)
#Calculate mean training set conservation
entropy = np.sum(x_mean * -np.log(x_mean), axis=-1) / np.log(2.0)
conservation = 2.0 - entropy
x_mean_conservation = np.sum(conservation) / np.sum(sequence_mask)
print("Mean conservation (bits) = " + str(x_mean_conservation))
#Calculate mean training set kl-divergence against background
x_train_clipped = np.clip(np.copy(x_train[:, 0, :, :]), 1e-8, 1. - 1e-8)
kl_divs = np.sum(x_train_clipped * np.log(x_train_clipped / np.tile(np.expand_dims(x_mean, axis=0), (x_train_clipped.shape[0], 1, 1))), axis=-1) / np.log(2.0)
x_mean_kl_divs = np.sum(kl_divs * sequence_mask, axis=-1) / np.sum(sequence_mask)
x_mean_kl_div = np.mean(x_mean_kl_divs)
print("Mean KL Div against background (bits) = " + str(x_mean_kl_div))
#Initialize Encoder and Decoder networks
batch_size = 32
seq_length = 50
n_samples = 128
sample_mode = 'st'
#sample_mode = 'gumbel'
#Load sampler
sampler = build_sampler(batch_size, seq_length, n_classes=1, n_samples=n_samples, sample_mode=sample_mode)
#Load Predictor
predictor_path = 'optimusRetrainedMain.hdf5'
predictor = load_model(predictor_path)
predictor.trainable = False
predictor.compile(optimizer=keras.optimizers.SGD(lr=0.1), loss='mean_squared_error')
#Build scrambler model
dummy_class = Input(shape=(1,), name='dummy_class')
input_logits = Input(shape=(1, seq_length, 4), name='input_logits')
pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask = sampler(dummy_class, input_logits)
scrambler_model = Model([input_logits, dummy_class], [pwm_logits, pwm, sampled_pwm, pwm_mask, sampled_mask])
#Initialize Sequence Templates and Masks
initialize_sequence_templates(scrambler_model, [sequence_template], [x_mean_logits])
scrambler_model.trainable = False
scrambler_model.compile(
optimizer=keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999),
loss='mean_squared_error'
)
#open all score and reshape as needed
file_names = [
"l2x_" + dataset_name + "_importance_scores_test.npy",
"invase_" + dataset_name + "_conv_importance_scores_test.npy",
"l2x_" + dataset_name + "_full_data_importance_scores_test.npy",
"invase_" + dataset_name + "_conv_full_data_importance_scores_test.npy",
]
#deepexplain_optimus_utr_OR_logic_synth_1_start_2_stops_method_integrated_gradients_importance_scores_test.npy
model_names =[
"l2x",
"invase",
"l2x_full_data",
"invase_full_data",
]
model_importance_scores_test = [np.load("./" + file_name) for file_name in file_names]
for scores in model_importance_scores_test:
print (scores.shape)
for model_i in range(len(model_names)) :
if model_importance_scores_test[model_i].shape[-1] > 1 :
model_importance_scores_test[model_i] = np.sum(model_importance_scores_test[model_i], axis=-1, keepdims=True)
for scores in model_importance_scores_test:
print (scores.shape)
#reshape for mse script -> if not (3008, 1, 50, 1) make it that shape
idealShape = model_importance_scores_test[0].shape
print (idealShape)
for model_i in range(len(model_names)) :
if model_importance_scores_test[model_i].shape != idealShape:
model_importance_scores_test[model_i] = np.expand_dims(model_importance_scores_test[model_i], 1)
for scores in model_importance_scores_test:
print (scores.shape)
on_state_logit_val = 50.
print (x_test.shape)
dummy_test = np.zeros((x_test.shape[0], 1))
x_test_logits = 2. * x_test - 1.
print (x_test_logits.shape)
print (dummy_test.shape)
x_test_squeezed = np.squeeze(x_test)
y_pred_ref = predictor.predict([x_test_squeezed], batch_size=32, verbose=True)[0]
_, _, _, pwm_mask, sampled_mask = scrambler_model.predict([x_test_logits, dummy_test], batch_size=batch_size)
feature_quantiles = [0.76, 0.82, 0.88]
for name in model_names:
for quantile in feature_quantiles:
totalName = name + "_" + str(quantile).replace(".","_") + "_quantile_MSE"
data_df[totalName] = None
print (data_df.columns)
feature_quantiles = [0.76, 0.82, 0.88]
#batch_size = 128
from sklearn import metrics
model_mses = []
for model_i in range(len(model_names)) :
print("Benchmarking model '" + str(model_names[model_i]) + "'...")
feature_quantile_mses = []
for feature_quantile_i, feature_quantile in enumerate(feature_quantiles) :
print("Feature quantile = " + str(feature_quantile))
if len(model_importance_scores_test[model_i].shape) >= 5 :
importance_scores_test = np.abs(model_importance_scores_test[model_i][feature_quantile_i, ...])
else :
importance_scores_test = np.abs(model_importance_scores_test[model_i])
n_to_test = importance_scores_test.shape[0] // batch_size * batch_size
importance_scores_test = importance_scores_test[:n_to_test]
importance_scores_test *= np.expand_dims(np.max(pwm_mask[:n_to_test], axis=-1), axis=-1)
quantile_vals = np.quantile(importance_scores_test, axis=(1, 2, 3), q=feature_quantile, keepdims=True)
quantile_vals = np.tile(quantile_vals, (1, importance_scores_test.shape[1], importance_scores_test.shape[2], importance_scores_test.shape[3]))
top_logits_test = np.zeros(importance_scores_test.shape)
top_logits_test[importance_scores_test > quantile_vals] = on_state_logit_val
top_logits_test = np.tile(top_logits_test, (1, 1, 1, 4)) * x_test_logits[:n_to_test]
_, _, samples_test, _, _ = scrambler_model.predict([top_logits_test, dummy_test[:n_to_test]], batch_size=batch_size)
print (samples_test.shape)
msesPerPoint = []
for data_ix in range(samples_test.shape[0]) :
#for each sample, look at kl divergence for the 128 size batch generated
#for MSE, just track the pred vs original pred
if data_ix % 1000 == 0 :
print("Processing example " + str(data_ix) + "...")
#from optimus R^2, MSE, Pearson R script
justPred = np.expand_dims(np.expand_dims(x_test[data_ix, 0, :, :], axis=0), axis=-1)
justPredReshape = np.reshape(justPred, (1,50,4))
expanded = np.expand_dims(samples_test[data_ix, :, 0, :, :], axis=-1) #batch size is 128
expandedReshape = np.reshape(expanded, (n_samples, 50,4))
y_test_hat_ref = predictor.predict(x=justPredReshape, batch_size=1)[0][0]
y_test_hat = predictor.predict(x=[expandedReshape], batch_size=32)
pwmGenerated = y_test_hat.tolist()
tempOriginals = [y_test_hat_ref]*y_test_hat.shape[0]
asArrayOrig = np.array(tempOriginals)
asArrayGen = np.array(pwmGenerated)
squeezed = np.squeeze(asArrayGen)
mse = metrics.mean_squared_error(asArrayOrig, squeezed)
#msesPerPoint.append(mse)
totalName = model_names[model_i] + "_" + str(feature_quantile).replace(".","_") + "_quantile_MSE"
data_df.at[data_ix, totalName] = mse
msesPerPoint.append(mse)
msesPerPoint = np.array(msesPerPoint)
feature_quantile_mses.append(msesPerPoint)
model_mses.append(feature_quantile_mses)
#Store benchmark results as tables
save_figs = False
mse_table = np.zeros((len(model_mses), len(model_mses[0])))
for i, model_name in enumerate(model_names) :
for j, feature_quantile in enumerate(feature_quantiles) :
mse_table[i, j] = np.mean(model_mses[i][j])
#Plot and store mse table
f = plt.figure(figsize = (4, 6))
cells = np.round(mse_table, 3).tolist()
print("--- MSEs ---")
max_len = np.max([len(model_name.upper().replace("\n", " ")) for model_name in model_names])
print(("-" * max_len) + " " + " ".join([(str(feature_quantile) + "0")[:4] for feature_quantile in feature_quantiles]))
for i in range(len(cells)) :
curr_len = len([model_name.upper().replace("\n", " ") for model_name in model_names][i])
row_str = [model_name.upper().replace("\n", " ") for model_name in model_names][i] + (" " * (max_len - curr_len))
for j in range(len(cells[i])) :
cells[i][j] = (str(cells[i][j]) + "00000")[:4]
row_str += " " + cells[i][j]
print(row_str)
print("")
table = plt.table(cellText=cells, rowLabels=[model_name.upper().replace("\n", " ") for model_name in model_names], colLabels=feature_quantiles, loc='center')
ax = plt.gca()
#f.patch.set_visible(False)
ax.axis('off')
ax.axis('tight')
plt.tight_layout()
if save_figs :
plt.savefig(dataset_name + "_l2x_and_invase_full_data" + "_mse_table.png", dpi=300, transparent=True)
plt.savefig(dataset_name + "_l2x_and_invase_full_data" + "_mse_table.eps")
plt.show()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import glob
import nibabel as nib
import os
import time
import pandas as pd
import numpy as np
import cv2
from skimage.transform import resize
from mricode.utils import log_textfile, createPath, data_generator
from mricode.utils import copy_colab
from mricode.utils import return_iter
from mricode.utils import return_csv
from mricode.config import config
from mricode.models.SimpleCNN_small import SimpleCNN
from mricode.models.DenseNet_NoDict import MyDenseNet
import tensorflow as tf
from tensorflow.keras.layers import Conv3D
from tensorflow import nn
from tensorflow.python.ops import nn_ops
from tensorflow.python.framework import tensor_shape
from tensorflow.python.keras.engine.base_layer import InputSpec
from tensorflow.python.keras.utils import conv_utils
tf.__version__
tf.test.is_gpu_available()
path_output = './output/'
path_tfrecords = '/data2/res64/down/'
path_csv = '/data2/csv/'
filename_res = {'train': 'intell_residual_train.csv', 'val': 'intell_residual_valid.csv', 'test': 'intell_residual_test.csv'}
filename_final = filename_res
sample_size = 'site16_allimages'
batch_size = 8
onlyt1 = False
Model = SimpleCNN
#Model = MyDenseNet
versionkey = 'down256' #down256, cropped128, cropped64, down64
modelname = 'simplecnnsmall__allimages_' + versionkey
createPath(path_output + modelname)
train_df, val_df, test_df, norm_dict = return_csv(path_csv, filename_final, False)
train_iter = config[versionkey]['iter_train']
val_iter = config[versionkey]['iter_val']
test_iter = config[versionkey]['iter_test']
t1_mean = config[versionkey]['norm']['t1'][0]
t1_std= config[versionkey]['norm']['t1'][1]
t2_mean=config[versionkey]['norm']['t2'][0]
t2_std=config[versionkey]['norm']['t2'][1]
ad_mean=config[versionkey]['norm']['ad'][0]
ad_std=config[versionkey]['norm']['ad'][1]
fa_mean=config[versionkey]['norm']['fa'][0]
fa_std=config[versionkey]['norm']['fa'][1]
md_mean=config[versionkey]['norm']['md'][0]
md_std=config[versionkey]['norm']['md'][1]
rd_mean=config[versionkey]['norm']['rd'][0]
rd_std=config[versionkey]['norm']['rd'][1]
norm_dict
cat_cols = {'female': 2, 'race.ethnicity': 5, 'high.educ_group': 4, 'income_group': 8, 'married': 6}
num_cols = [x for x in list(val_df.columns) if '_norm' in x]
def calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict):
for col in num_cols:
tmp_col = col
tmp_std = norm_dict[tmp_col.replace('_norm','')]['std']
tmp_y_true = tf.cast(y_true[col], tf.float32).numpy()
tmp_y_pred = np.squeeze(y_pred[col].numpy())
if not(tmp_col in out_loss):
out_loss[tmp_col] = np.sum(np.square(tmp_y_true-tmp_y_pred))
else:
out_loss[tmp_col] += np.sum(np.square(tmp_y_true-tmp_y_pred))
if not(tmp_col in out_acc):
out_acc[tmp_col] = np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
else:
out_acc[tmp_col] += np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
for col in list(cat_cols.keys()):
tmp_col = col
if not(tmp_col in out_loss):
out_loss[tmp_col] = tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
else:
out_loss[tmp_col] += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
if not(tmp_col in out_acc):
out_acc[tmp_col] = tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
else:
out_acc[tmp_col] += tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
return(out_loss, out_acc)
def format_output(out_loss, out_acc, n, cols, print_bl=False):
loss = 0
acc = 0
output = []
for col in cols:
output.append([col, out_loss[col]/n, out_acc[col]/n])
loss += out_loss[col]/n
acc += out_acc[col]/n
df = pd.DataFrame(output)
df.columns = ['name', 'loss', 'acc']
if print_bl:
print(df)
return(loss, acc, df)
@tf.function
def train_step(X, y, model, optimizer, cat_cols, num_cols):
with tf.GradientTape() as tape:
predictions = model(X)
i = 0
loss = tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for i in range(1,len(num_cols)):
loss += tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for col in list(cat_cols.keys()):
loss += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y[col]), tf.squeeze(predictions[col]))
gradients = tape.gradient(loss, model.trainable_variables)
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
with tf.control_dependencies(mean_std):
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
return(y, predictions, loss)
@tf.function
def test_step(X, y, model):
predictions = model(X)
return(y, predictions)
def epoch(data_iter, df, model, optimizer, cat_cols, num_cols, norm_dict):
out_loss = {}
out_acc = {}
n = 0.
n_batch = 0.
total_time_dataload = 0.
total_time_model = 0.
start_time = time.time()
for batch in data_iter:
total_time_dataload += time.time() - start_time
start_time = time.time()
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
if False:
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
subjectid = decoder(batch['subjectid'])
y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
#X = tf.concat([t1], axis=4)
X = tf.concat([t1, t2], axis=4)
if optimizer != None:
y_true, y_pred, loss = train_step(X, y, model, optimizer, cat_cols, num_cols)
else:
y_true, y_pred = test_step(X, y, model)
out_loss, out_acc = calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict)
n += X.shape[0]
n_batch += 1
if (n_batch % 10) == 0:
log_textfile(path_output + modelname + '/log' + '.log', str(n_batch))
total_time_model += time.time() - start_time
start_time = time.time()
return (out_loss, out_acc, n, total_time_model, total_time_dataload)
def get_labels(df, subjectid, cols = ['nihtbx_fluidcomp_uncorrected_norm']):
subjects_df = pd.DataFrame(subjectid)
result_df = pd.merge(subjects_df, df, left_on=0, right_on='subjectkey', how='left')
output = {}
for col in cols:
output[col] = np.asarray(result_df[col].values)
return output
def best_val(df_best, df_val, df_test, e):
df_best = pd.merge(df_best, df_val, how='left', left_on='name', right_on='name')
df_best = pd.merge(df_best, df_test, how='left', left_on='name', right_on='name')
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_epochs'] = e
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_epochs'] = e
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_test'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_test']
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_val'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_val']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best = df_best.drop(['cur_loss_val', 'cur_acc_val', 'cur_loss_test', 'cur_acc_test'], axis=1)
return(df_best)
decoder = np.vectorize(lambda x: x.decode('UTF-8'))
template = 'Epoch {0}, Loss: {1:.3f}, Accuracy: {2:.3f}, Val Loss: {3:.3f}, Val Accuracy: {4:.3f}, Time Model: {5:.3f}, Time Data: {6:.3f}'
for col in [0]:
log_textfile(path_output + modelname + '/log' + '.log', cat_cols)
log_textfile(path_output + modelname + '/log' + '.log', num_cols)
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam(lr = 0.001)
model = Model(cat_cols, num_cols)
df_best = None
for e in range(20):
log_textfile(path_output + modelname + '/log' + '.log', 'Epochs: ' + str(e))
loss = tf.Variable(0.)
acc = tf.Variable(0.)
val_loss = tf.Variable(0.)
val_acc = tf.Variable(0.)
test_loss = tf.Variable(0.)
test_acc = tf.Variable(0.)
tf.keras.backend.set_learning_phase(True)
train_out_loss, train_out_acc, n, time_model, time_data = epoch(train_iter, train_df, model, optimizer, cat_cols, num_cols, norm_dict)
tf.keras.backend.set_learning_phase(False)
val_out_loss, val_out_acc, n, _, _ = epoch(val_iter, val_df, model, None, cat_cols, num_cols, norm_dict)
test_out_loss, test_out_acc, n, _, _ = epoch(test_iter, test_df, model, None, cat_cols, num_cols, norm_dict)
loss, acc, _ = format_output(train_out_loss, train_out_acc, n, list(cat_cols.keys())+num_cols)
val_loss, val_acc, df_val = format_output(val_out_loss, val_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_val.columns = ['name', 'cur_loss_val', 'cur_acc_val']
df_test.columns = ['name', 'cur_loss_test', 'cur_acc_test']
if e == 0:
df_best = pd.merge(df_test, df_val, how='left', left_on='name', right_on='name')
df_best['best_acc_epochs'] = 0
df_best['best_loss_epochs'] = 0
df_best.columns = ['name', 'best_loss_test', 'best_acc_test', 'best_loss_val', 'best_acc_val', 'best_acc_epochs', 'best_loss_epochs']
df_best = best_val(df_best, df_val, df_test, e)
print(df_best[['name', 'best_loss_test', 'best_acc_test']])
print(df_best[['name', 'best_loss_val', 'best_acc_val']])
log_textfile(path_output + modelname + '/log' + '.log', template.format(e, loss, acc, val_loss, val_acc, time_model, time_data))
if e in [10, 15]:
optimizer.lr = optimizer.lr/3
log_textfile(path_output + modelname + '/log' + '.log', 'Learning rate: ' + str(optimizer.lr))
df_best.to_csv(path_output + modelname + '/df_best' + str(e) + '.csv')
df_best.to_csv(path_output + modelname + '/df_best' + '.csv')
model.save_weights(path_output + modelname + '/checkpoints/' + str(e) + '/')
error
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_test.to_csv('final_output_all.csv')
inputs = tf.keras.Input(shape=(64,64,64,2), name='inputlayer123')
a = model(inputs)['female']
mm = tf.keras.models.Model(inputs=inputs, outputs=a)
from tf_explain.core.smoothgrad import SmoothGrad
import pickle
explainer = SmoothGrad()
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
X_i = X[i]
X_i = tf.expand_dims(X_i, axis=0)
y_i = y_list[i]
grid = explainer.explain((X_i, _), mm, y_i, 20, 1.)
output_grid[y_i] += grid
output_n[y_i] += 1
pickle.dump([output_grid, output_n], open( "smoothgrad_female_all.p", "wb" ) )
#output_grid, output_n = pickle.load(open( "smoothgrad_female.p", "rb" ))
def apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size):
"""
Replace a part of the image with a grey patch.
Args:
image (numpy.ndarray): Input image
top_left_x (int): Top Left X position of the applied box
top_left_y (int): Top Left Y position of the applied box
patch_size (int): Size of patch to apply
Returns:
numpy.ndarray: Patched image
"""
patched_image = np.array(image, copy=True)
patched_image[
top_left_x : top_left_x + patch_size, top_left_y : top_left_y + patch_size, top_left_z : top_left_z + patch_size, :
] = 0
return patched_image
import math
def get_sensgrid(image, mm, class_index, patch_size):
sensitivity_map = np.zeros((
math.ceil(image.shape[0] / patch_size),
math.ceil(image.shape[1] / patch_size),
math.ceil(image.shape[2] / patch_size)
))
for index_z, top_left_z in enumerate(range(0, image.shape[2], patch_size)):
patches = [
apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size)
for index_x, top_left_x in enumerate(range(0, image.shape[0], patch_size))
for index_y, top_left_y in enumerate(range(0, image.shape[1], patch_size))
]
coordinates = [
(index_y, index_x)
for index_x, _ in enumerate(range(0, image.shape[0], patch_size))
for index_y, _ in enumerate(range(0, image.shape[1], patch_size))
]
predictions = mm.predict(np.array(patches), batch_size=1)
target_class_predictions = [prediction[class_index] for prediction in predictions]
for (index_y, index_x), confidence in zip(coordinates, target_class_predictions):
sensitivity_map[index_y, index_x, index_z] = 1 - confidence
sm = resize(sensitivity_map, (64,64,64))
heatmap = (sm - np.min(sm)) / (sm.max() - sm.min())
return(heatmap)
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
print(i)
X_i = X[i]
y_i = y_list[i]
grid = get_sensgrid(X_i, mm, y_i, 4)
output_grid[y_i] += grid
output_n[y_i] += 1
if counter==6:
break
pickle.dump([output_grid, output_n], open( "heatmap_female_all.p", "wb" ) )
error
batch = next(iter(train_iter))
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
#subjectid = decoder(batch['subjectid'])
#y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
#X = tf.concat([t1, t2, ad, fa, md, rd], axis=4)
X = tf.concat([t1, t2], axis=4)
tf.keras.backend.set_learning_phase(True)
model(X)['female']
tf.keras.backend.set_learning_phase(False)
model(X)['female']
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
model = Model(cat_cols, num_cols)
model.non_trainable_variables
```
| github_jupyter |
# <a title="Activity Recognition" href="https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition" > LSTMs for Human Activity Recognition</a>
Human Activity Recognition (HAR) using smartphones dataset and an LSTM RNN. Classifying the type of movement amongst six categories:
- WALKING,
- WALKING_UPSTAIRS,
- WALKING_DOWNSTAIRS,
- SITTING,
- STANDING,
- LAYING.
Compared to a classical approach, using a Recurrent Neural Networks (RNN) with Long Short-Term Memory cells (LSTMs) require no or almost no feature engineering. Data can be fed directly into the neural network who acts like a black box, modeling the problem correctly. [Other research](https://archive.ics.uci.edu/ml/machine-learning-databases/00240/UCI%20HAR%20Dataset.names) on the activity recognition dataset can use a big amount of feature engineering, which is rather a signal processing approach combined with classical data science techniques. The approach here is rather very simple in terms of how much was the data preprocessed.
Let's use Google's neat Deep Learning library, TensorFlow, demonstrating the usage of an LSTM, a type of Artificial Neural Network that can process sequential data / time series.
## Video dataset overview
Follow this link to see a video of the 6 activities recorded in the experiment with one of the participants:
<p align="center">
<a href="http://www.youtube.com/watch?feature=player_embedded&v=XOEN9W05_4A
" target="_blank"><img src="http://img.youtube.com/vi/XOEN9W05_4A/0.jpg"
alt="Video of the experiment" width="400" height="300" border="10" /></a>
<a href="https://youtu.be/XOEN9W05_4A"><center>[Watch video]</center></a>
</p>
## Details about the input data
I will be using an LSTM on the data to learn (as a cellphone attached on the waist) to recognise the type of activity that the user is doing. The dataset's description goes like this:
> The sensor signals (accelerometer and gyroscope) were pre-processed by applying noise filters and then sampled in fixed-width sliding windows of 2.56 sec and 50% overlap (128 readings/window). The sensor acceleration signal, which has gravitational and body motion components, was separated using a Butterworth low-pass filter into body acceleration and gravity. The gravitational force is assumed to have only low frequency components, therefore a filter with 0.3 Hz cutoff frequency was used.
That said, I will use the almost raw data: only the gravity effect has been filtered out of the accelerometer as a preprocessing step for another 3D feature as an input to help learning. If you'd ever want to extract the gravity by yourself, you could fork my code on using a [Butterworth Low-Pass Filter (LPF) in Python](https://github.com/guillaume-chevalier/filtering-stft-and-laplace-transform) and edit it to have the right cutoff frequency of 0.3 Hz which is a good frequency for activity recognition from body sensors.
## What is an RNN?
As explained in [this article](http://karpathy.github.io/2015/05/21/rnn-effectiveness/), an RNN takes many input vectors to process them and output other vectors. It can be roughly pictured like in the image below, imagining each rectangle has a vectorial depth and other special hidden quirks in the image below. **In our case, the "many to one" architecture is used**: we accept time series of [feature vectors](https://www.quora.com/What-do-samples-features-time-steps-mean-in-LSTM/answer/Guillaume-Chevalier-2) (one vector per [time step](https://www.quora.com/What-do-samples-features-time-steps-mean-in-LSTM/answer/Guillaume-Chevalier-2)) to convert them to a probability vector at the output for classification. Note that a "one to one" architecture would be a standard feedforward neural network.
> <a href="http://karpathy.github.io/2015/05/21/rnn-effectiveness/" ><img src="http://karpathy.github.io/assets/rnn/diags.jpeg" /></a>
> http://karpathy.github.io/2015/05/21/rnn-effectiveness/
## What is an LSTM?
An LSTM is an improved RNN. It is more complex, but easier to train, avoiding what is called the vanishing gradient problem. I recommend [this article](http://colah.github.io/posts/2015-08-Understanding-LSTMs/) for you to learn more on LSTMs.
## Results
Scroll on! Nice visuals awaits.
```
# All Includes
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import tensorflow as tf # Version 1.0.0 (some previous versions are used in past commits)
from sklearn import metrics
import os
# Useful Constants
# Those are separate normalised input features for the neural network
INPUT_SIGNAL_TYPES = [
"body_acc_x_",
"body_acc_y_",
"body_acc_z_",
"body_gyro_x_",
"body_gyro_y_",
"body_gyro_z_",
"total_acc_x_",
"total_acc_y_",
"total_acc_z_"
]
# Output classes to learn how to classify
LABELS = [
"WALKING",
"WALKING_UPSTAIRS",
"WALKING_DOWNSTAIRS",
"SITTING",
"STANDING",
"LAYING"
]
```
## Let's start by downloading the data:
```
## Note: Linux bash commands start with a "!" inside those "ipython notebook" cells
#
DATA_PATH = "data/"
#
#!pwd && ls
#os.chdir(DATA_PATH)
#!pwd && ls
#
#!python download_dataset.py
#
#!pwd && ls
#os.chdir("..")
#!pwd && ls
#
DATASET_PATH = DATA_PATH + "UCI HAR Dataset/"
print("\n" + "Dataset is now located at: " + DATASET_PATH)
#
```
## Preparing dataset:
```
TRAIN = "train/"
TEST = "test/"
X_train_signals_paths = [DATASET_PATH + TRAIN + "Inertial Signals/" + signal + "train.txt" for signal in INPUT_SIGNAL_TYPES]
'''
['data/UCI HAR Dataset/train/Inertial Signals/body_acc_x_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/body_acc_y_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/body_acc_z_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/body_gyro_x_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/body_gyro_y_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/body_gyro_z_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/total_acc_x_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/total_acc_y_train.txt',
'data/UCI HAR Dataset/train/Inertial Signals/total_acc_z_train.txt']
'''
X_test_signals_paths = [DATASET_PATH + TEST + "Inertial Signals/" + signal + "test.txt" for signal in INPUT_SIGNAL_TYPES]
'''
['data/UCI HAR Dataset/test/Inertial Signals/body_acc_x_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/body_acc_y_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/body_acc_z_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/body_gyro_x_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/body_gyro_y_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/body_gyro_z_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/total_acc_x_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/total_acc_y_test.txt',
'data/UCI HAR Dataset/test/Inertial Signals/total_acc_z_test.txt']
'''
##################################################
# Load "X" (the neural network's training and testing inputs)
##################################################
def load_X(X_signals_paths):
X_signals = []
for signal_type_path in X_signals_paths:
file = open(signal_type_path, 'r')
# Read dataset from disk, dealing with text files' syntax
X_signals.append([np.array(serie, dtype=np.float32) for serie in [ row.replace(' ', ' ').strip().split(' ') for row in file]])
file.close()
return np.transpose(np.array(X_signals), (1, 2, 0))
X_train = load_X(X_train_signals_paths) # (7352, 128, 9)
X_test = load_X(X_test_signals_paths) # (2947, 128, 9)
##################################################
# Load "y" (the neural network's training and testing outputs)
##################################################
def load_y(y_path):
file = open(y_path, 'r')
# Read dataset from disk, dealing with text file's syntax
y_ = np.array([elem for elem in [row.replace(' ', ' ').strip().split(' ') for row in file]], dtype=np.int32)
file.close()
# Substract 1 to each output class for friendly 0-based indexing
return y_ - 1
y_train_path = DATASET_PATH + TRAIN + "y_train.txt"
''' data/UCI HAR Dataset/train/y_train.txt '''
y_test_path = DATASET_PATH + TEST + "y_test.txt"
''' data/UCI HAR Dataset/test/y_test.txt '''
y_train = load_y(y_train_path) # (7352, 1)
y_test = load_y(y_test_path) # (2947, 1)
```
## Additionnal Parameters:
Here are some core parameter definitions for the training.
For example, the whole neural network's structure could be summarised by enumerating those parameters and the fact that two LSTM are used one on top of another (stacked) output-to-input as hidden layers through time steps.
```
# Input Data
training_data_count = len(X_train) # 7352 training series (with 50% overlap between each serie)
test_data_count = len(X_test) # 2947 testing series
n_steps = len(X_train[0]) # 128 timesteps per series
n_input = len(X_train[0][0]) # 9 input parameters per timestep
# LSTM Neural Network's internal structure
n_hidden = 32 # Hidden layer num of features
n_classes = 6 # Total classes (should go up, or should go down)
# Training
learning_rate = 0.0025
lambda_loss_amount = 0.0015
training_iters = training_data_count * 300 # Loop 300 times on the dataset
batch_size = 1500
display_iter = 30000 # To show test set accuracy during training
# Some debugging info
print("Some useful info to get an insight on dataset's shape and normalisation:")
print("(X shape, y shape, every X's mean, every X's standard deviation)")
print(X_test.shape, y_test.shape, np.mean(X_test), np.std(X_test))
print("The dataset is therefore properly normalised, as expected, but not yet one-hot encoded.")
print(('X_train: {}').format(X_train.shape))
print(('X_test: {}').format(X_test.shape))
print(('y_train: {}').format(y_train.shape))
print(('y_test: {}').format(y_test.shape))
##128: readings/window
##[acc_x", "acc_y", "acc_z", "gyro_x", "gyro_y", "gyro_z", "total_acc_x", "total_acc_y" , "total_acc_z"]
##
##["WALKING", "WALKING_UPSTAIRS", "WALKING_DOWNSTAIRS", "SITTING", "STANDING", "LAYING"]
X_train[0][0]
X_train[0][1]
X_train[0][0]
```
## Utility functions for training:
```
def LSTM_RNN(_X, _weights, _biases):
# Function returns a tensorflow LSTM (RNN) artificial neural network from given parameters.
# Moreover, two LSTM cells are stacked which adds deepness to the neural network.
# Note, some code of this notebook is inspired from an slightly different
# RNN architecture used on another dataset, some of the credits goes to
# "aymericdamien" under the MIT license.
# (NOTE: This step could be greatly optimised by shaping the dataset once
# input shape: (batch_size, n_steps, n_input)
_X = tf.transpose(_X, [1, 0, 2]) # permute n_steps and batch_size
# Reshape to prepare input to hidden activation
_X = tf.reshape(_X, [-1, n_input])
# new shape: (n_steps*batch_size, n_input)
# ReLU activation, thanks to Yu Zhao for adding this improvement here:
_X = tf.nn.relu(tf.matmul(_X, _weights['hidden']) + _biases['hidden'])
# Split data because rnn cell needs a list of inputs for the RNN inner loop
_X = tf.split(_X, n_steps, 0)
# new shape: n_steps * (batch_size, n_hidden)
# Define two stacked LSTM cells (two recurrent layers deep) with tensorflow
lstm_cell_1 = tf.contrib.rnn.BasicLSTMCell(n_hidden, forget_bias=1.0, state_is_tuple=True)
lstm_cell_2 = tf.contrib.rnn.BasicLSTMCell(n_hidden, forget_bias=1.0, state_is_tuple=True)
lstm_cells = tf.contrib.rnn.MultiRNNCell([lstm_cell_1, lstm_cell_2], state_is_tuple=True)
# Get LSTM cell output
outputs, states = tf.contrib.rnn.static_rnn(lstm_cells, _X, dtype=tf.float32)
# Get last time step's output feature for a "many-to-one" style classifier,
# as in the image describing RNNs at the top of this page
lstm_last_output = outputs[-1]
# Linear activation
return tf.matmul(lstm_last_output, _weights['out']) + _biases['out']
def extract_batch_size(_train, step, batch_size):
# Function to fetch a "batch_size" amount of data from "(X|y)_train" data.
shape = list(_train.shape)
shape[0] = batch_size
batch_s = np.empty(shape)
for i in range(batch_size):
# Loop index
index = ((step-1)*batch_size + i) % len(_train)
batch_s[i] = _train[index]
return batch_s
def one_hot(y_, n_classes=n_classes):
# Function to encode neural one-hot output labels from number indexes
# e.g.:
# one_hot(y_=[[5], [0], [3]], n_classes=6):
# return [[0, 0, 0, 0, 0, 1], [1, 0, 0, 0, 0, 0], [0, 0, 0, 1, 0, 0]]
y_ = y_.reshape(len(y_))
return np.eye(n_classes)[np.array(y_, dtype=np.int32)] # Returns FLOATS
```
## Let's get serious and build the neural network:
```
################
# n_steps: 128 readings / window
# n_input: 9 [acc_x", "acc_y", "acc_z", "gyro_x", "gyro_y", "gyro_z", "total_acc_x", "total_acc_y" , "total_acc_z"]
# n_classes: 6 ["WALKING", "WALKING_UPSTAIRS", "WALKING_DOWNSTAIRS", "SITTING", "STANDING", "LAYING"]
# n_hidden: 32
#training_data_count: 7352
#test_data_count: 2947
#learning_rate: 0.0025
#lambda_loss_amount: 0.0015
# training_iters: 2205600
#batch_size: 1500
#display_iter: 30000
################
# Graph input/output
x = tf.placeholder(tf.float32, [None, n_steps, n_input])
y = tf.placeholder(tf.float32, [None, n_classes])
# Graph weights
weights = {
'hidden': tf.Variable(tf.random_normal([n_input, n_hidden])), # Hidden layer weights
'out': tf.Variable(tf.random_normal([n_hidden, n_classes], mean=1.0))
}
biases = {
'hidden': tf.Variable(tf.random_normal([n_hidden])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
weights
# prediction
pred = LSTM_RNN(x, weights, biases)
# Loss, optimizer and evaluation:
#################################
# L2 loss prevents this overkill neural network to overfit the data
l2 = lambda_loss_amount * sum(tf.nn.l2_loss(tf_var) for tf_var in tf.trainable_variables())
# Softmax loss
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y, logits=pred)) + l2
# Adam Optimizer
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
correct_pred = tf.equal(tf.argmax(pred,1), tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
```
## Hooray, now train the neural network:
```
# To keep track of training's performance
test_losses = []
test_accuracies = []
train_losses = []
train_accuracies = []
# Launch the graph
sess = tf.InteractiveSession(config=tf.ConfigProto(log_device_placement=True))
init = tf.global_variables_initializer()
sess.run(init)
# Perform Training steps with "batch_size" amount of example data at each loop
step = 1
while step * batch_size <= training_iters:
batch_xs = extract_batch_size(X_train, step, batch_size)
batch_ys = one_hot(extract_batch_size(y_train, step, batch_size))
# Fit training using batch data
_, loss, acc = sess.run(
[optimizer, cost, accuracy],
feed_dict={
x: batch_xs,
y: batch_ys})
train_losses.append(loss)
train_accuracies.append(acc)
# Evaluate network only at some steps for faster training:
if (step*batch_size % display_iter == 0) or (step == 1) or (step * batch_size > training_iters):
# To not spam console, show training accuracy/loss in this "if"
print("Training iter #" + str(step*batch_size) + \
": Batch Loss = " + "{:.6f}".format(loss) + \
", Accuracy = {}".format(acc))
# Evaluation on the test set (no learning made here - just evaluation for diagnosis)
loss, acc = sess.run(
[cost, accuracy],
feed_dict={
x: X_test,
y: one_hot(y_test)
}
)
test_losses.append(loss)
test_accuracies.append(acc)
print("PERFORMANCE ON TEST SET: " + \
"Batch Loss = {}".format(loss) + \
", Accuracy = {}".format(acc))
step += 1
print("Optimization Finished!")
# Accuracy for test data
one_hot_predictions, accuracy, final_loss = sess.run(
[pred, accuracy, cost],
feed_dict={
x: X_test,
y: one_hot(y_test)
}
)
test_losses.append(final_loss)
test_accuracies.append(accuracy)
print("FINAL RESULT: " + \
"Batch Loss = {}".format(final_loss) + \
", Accuracy = {}".format(accuracy))
```
## Training is good, but having visual insight is even better:
Okay, let's plot this simply in the notebook for now.
```
# (Inline plots: )
%matplotlib inline
font = {
'family' : 'Bitstream Vera Sans',
'weight' : 'bold',
'size' : 18
}
matplotlib.rc('font', **font)
width = 12
height = 12
plt.figure(figsize=(width, height))
indep_train_axis = np.array(range(batch_size, (len(train_losses)+1)*batch_size, batch_size))
plt.plot(indep_train_axis, np.array(train_losses), "b--", label="Train losses")
plt.plot(indep_train_axis, np.array(train_accuracies), "g--", label="Train accuracies")
indep_test_axis = np.append(
np.array(range(batch_size, len(test_losses)*display_iter, display_iter)[:-1]),
[training_iters]
)
plt.plot(indep_test_axis, np.array(test_losses), "b-", label="Test losses")
plt.plot(indep_test_axis, np.array(test_accuracies), "g-", label="Test accuracies")
plt.title("Training session's progress over iterations")
plt.legend(loc='upper right', shadow=True)
plt.ylabel('Training Progress (Loss or Accuracy values)')
plt.xlabel('Training iteration')
plt.show()
```
## And finally, the multi-class confusion matrix and metrics!
```
# Results
predictions = one_hot_predictions.argmax(1)
print("Testing Accuracy: {}%".format(100*accuracy))
print("")
print("Precision: {}%".format(100*metrics.precision_score(y_test, predictions, average="weighted")))
print("Recall: {}%".format(100*metrics.recall_score(y_test, predictions, average="weighted")))
print("f1_score: {}%".format(100*metrics.f1_score(y_test, predictions, average="weighted")))
print("")
print("Confusion Matrix:")
confusion_matrix = metrics.confusion_matrix(y_test, predictions)
print(confusion_matrix)
normalised_confusion_matrix = np.array(confusion_matrix, dtype=np.float32)/np.sum(confusion_matrix)*100
print("")
print("Confusion matrix (normalised to % of total test data):")
print(normalised_confusion_matrix)
print("Note: training and testing data is not equally distributed amongst classes, ")
print("so it is normal that more than a 6th of the data is correctly classifier in the last category.")
# Plot Results:
width = 12
height = 12
plt.figure(figsize=(width, height))
plt.imshow(
normalised_confusion_matrix,
interpolation='nearest',
cmap=plt.cm.rainbow
)
plt.title("Confusion matrix \n(normalised to % of total test data)")
plt.colorbar()
tick_marks = np.arange(n_classes)
plt.xticks(tick_marks, LABELS, rotation=90)
plt.yticks(tick_marks, LABELS)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
sess.close()
```
## Conclusion
Outstandingly, **the final accuracy is of 91%**! And it can peak to values such as 93.25%, at some moments of luck during the training, depending on how the neural network's weights got initialized at the start of the training, randomly.
This means that the neural networks is almost always able to correctly identify the movement type! Remember, the phone is attached on the waist and each series to classify has just a 128 sample window of two internal sensors (a.k.a. 2.56 seconds at 50 FPS), so it amazes me how those predictions are extremely accurate given this small window of context and raw data. I've validated and re-validated that there is no important bug, and the community used and tried this code a lot. (Note: be sure to report something in the issue tab if you find bugs, otherwise [Quora](https://www.quora.com/), [StackOverflow](https://stackoverflow.com/questions/tagged/tensorflow?sort=votes&pageSize=50), and other [StackExchange](https://stackexchange.com/sites#science) sites are the places for asking questions.)
I specially did not expect such good results for guessing between the labels "SITTING" and "STANDING". Those are seemingly almost the same thing from the point of view of a device placed at waist level according to how the dataset was originally gathered. Thought, it is still possible to see a little cluster on the matrix between those classes, which drifts away just a bit from the identity. This is great.
It is also possible to see that there was a slight difficulty in doing the difference between "WALKING", "WALKING_UPSTAIRS" and "WALKING_DOWNSTAIRS". Obviously, those activities are quite similar in terms of movements.
I also tried my code without the gyroscope, using only the 3D accelerometer's 6 features (and not changing the training hyperparameters), and got an accuracy of 87%. In general, gyroscopes consumes more power than accelerometers, so it is preferable to turn them off.
## Improvements
In [another open-source repository of mine](https://github.com/guillaume-chevalier/HAR-stacked-residual-bidir-LSTMs), the accuracy is pushed up to nearly 94% using a special deep LSTM architecture which combines the concepts of bidirectional RNNs, residual connections, and stacked cells. This architecture is also tested on another similar activity dataset. It resembles the nice architecture used in "[Google’s Neural Machine Translation System: Bridging the Gap between Human and Machine Translation](https://arxiv.org/pdf/1609.08144.pdf)", without an attention mechanism, and with just the encoder part - as a "many to one" architecture instead of a "many to many" to be adapted to the Human Activity Recognition (HAR) problem. I also worked more on the problem and came up with the [LARNN](https://github.com/guillaume-chevalier/Linear-Attention-Recurrent-Neural-Network), however it's complicated for just a little gain. Thus the current, original activity recognition project is simply better to use for its outstanding simplicity.
If you want to learn more about deep learning, I have also built a list of the learning ressources for deep learning which have revealed to be the most useful to me [here](https://github.com/guillaume-chevalier/Awesome-Deep-Learning-Resources).
## References
The [dataset](https://archive.ics.uci.edu/ml/datasets/Human+Activity+Recognition+Using+Smartphones) can be found on the UCI Machine Learning Repository:
> Davide Anguita, Alessandro Ghio, Luca Oneto, Xavier Parra and Jorge L. Reyes-Ortiz. A Public Domain Dataset for Human Activity Recognition Using Smartphones. 21th European Symposium on Artificial Neural Networks, Computational Intelligence and Machine Learning, ESANN 2013. Bruges, Belgium 24-26 April 2013.
The RNN image for "many-to-one" is taken from Karpathy's post:
> Andrej Karpathy, The Unreasonable Effectiveness of Recurrent Neural Networks, 2015,
> http://karpathy.github.io/2015/05/21/rnn-effectiveness/
## Citation
Copyright (c) 2016 Guillaume Chevalier. To cite my code, you can point to the URL of the GitHub repository, for example:
> Guillaume Chevalier, LSTMs for Human Activity Recognition, 2016,
> https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition
My code is available for free and even for private usage for anyone under the [MIT License](https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition/blob/master/LICENSE), however I ask to cite for using the code.
## Extra links
### Connect with me
- [LinkedIn](https://ca.linkedin.com/in/chevalierg)
- [Twitter](https://twitter.com/guillaume_che)
- [GitHub](https://github.com/guillaume-chevalier/)
- [Quora](https://www.quora.com/profile/Guillaume-Chevalier-2)
- [YouTube](https://www.youtube.com/c/GuillaumeChevalier)
- [Dev/Consulting](http://www.neuraxio.com/en/)
### Liked this project? Did it help you? Leave a [star](https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition/stargazers), [fork](https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition/network/members) and share the love!
This activity recognition project has been seen in:
- [Hacker News 1st page](https://news.ycombinator.com/item?id=13049143)
- [Awesome TensorFlow](https://github.com/jtoy/awesome-tensorflow#tutorials)
- [TensorFlow World](https://github.com/astorfi/TensorFlow-World#some-useful-tutorials)
- And more.
---
```
# Let's convert this notebook to a README automatically for the GitHub project's title page:
!jupyter nbconvert --to markdown LSTM.ipynb
!mv LSTM.md README.md
```
| github_jupyter |
```
# Install default libraries
import pathlib
import sys
# Import installed modules
import pandas as pd
import numpy as np
# Import the Python script from the auxiliary folder
sys.path.insert(1, "../auxiliary")
import data_fetch
# Set a local download path and the URL to the 67P shape model data set
dl_path = "../kernels/dsk/"
# Set dictionary with 2 different resolutions
comet_models = {"low": "ROS_CG_M001_OSPCLPS_N_V1.OBJ",
"high": "ROS_CG_M004_OSPGDLR_N_V1.OBJ"}
# Which model?
model_type = "high"
# Shape model DL
dl_url = f"https://naif.jpl.nasa.gov/pub/naif/ROSETTA/kernels/dsk/{comet_models[model_type]}"
# If file not present: download it!
if not pathlib.Path(f"../kernels/dsk/{comet_models[model_type]}").is_file():
# Download the shape model, create (if needed) the download path and store the data set
data_fetch.download_file(dl_path, dl_url)
# Load the shape model. The first column lists whether the row is a vertex or face. The second,
# third and fourth column list the coordiantes (vertex) and vertex indices (faces)
comet_67p_shape_obj = pd.read_csv(f"../kernels/dsk/{comet_models[model_type]}", \
delim_whitespace=True, \
names=["TYPE", "X1", "X2", "X3"])
# Assign the vertices and faces
vertices = comet_67p_shape_obj.loc[comet_67p_shape_obj["TYPE"] == "v"][["X1", "X2", "X3"]].values \
.tolist()
faces = comet_67p_shape_obj.loc[comet_67p_shape_obj["TYPE"] == "f"][["X1", "X2", "X3"]].values
# The index in the faces sub set starts at 1. For Python, it needs to start at 0.
faces = faces - 1
# Convert the indices to integer
faces = faces.astype(int)
# Convert the numpy array to a Python list
faces = faces.tolist()
# Doing some SPICE magic here!
import spiceypy
# Load the time kernel
spiceypy.furnsh("../kernels/lsk/naif0012.tls")
# Load the CG related FIXED reference frame
spiceypy.furnsh("../kernels/fk/ROS_CHURYUMOV_V01.TF")
# Load in the "rebound" kernel
spiceypy.furnsh("../kernels/spk/LORB_C_G_FIXED_RBD_7_V2_0.BSP")
# Get the kernel, check the cardinality and determine the startin and end times
rebound_kernel = spiceypy.spkcov(spk="../kernels/spk/LORB_C_G_FIXED_RBD_7_V2_0.BSP",
idcode=-226800)
rebound_kernel_card = spiceypy.wncard(rebound_kernel)
kernel_start_et, kernel_end_et = spiceypy.wnfetd(rebound_kernel,
rebound_kernel_card-1)
# List that will store the Philae trajectory
philae_pos_vec = []
# Iterate trough the time steps where Philae jumped around
for et_step in np.arange(kernel_start_et+100, kernel_end_et-100, 10):
# Compute the position of Philae w.r.t. the center of 67P
philae_pos_vec.append(spiceypy.spkgps(targ=-226800,
et=et_step,
ref="67P/C-G_CK",
obs=1000012)[0])
# Convert to numpy array
philae_pos_vec = np.array(philae_pos_vec)
# Let's take a look
print(philae_pos_vec)
# Now we want to plot the comet (as last time), but with Philae's trajectory!
import visvis as vv
from visvis import Point, Pointset
# Convert the philae position vector to a visvis Pointset
pp = Pointset(philae_pos_vec)
# Create visvis application
app = vv.use()
# Create the 3 D shape model as a mesh. verticesPerFace equals 3 since triangles define the
# mesh"s surface in this case
shape_obj = vv.mesh(vertices=vertices, faces=faces, verticesPerFace=3)
shape_obj.specular = 0.0
shape_obj.diffuse = 0.9
# Get axes objects
axes = vv.gca()
# Set a black background
axes.bgcolor = "black"
# Deactivate the grid and make the x, y, z axes invisible
axes.axis.showGrid = False
axes.axis.visible = False
# Set some camera settings
# Please note: if you want to "fly" arond the comet with w, a, s, d (translation) and i, j, k, l
# (tilt) replace "3d" with "fly"
axes.camera = "3d"
# Field of view in degrees
axes.camera.fov = 60
# Set default azmiuth and elevation angle in degrees
axes.camera.azimuth = 120
axes.camera.elevation = 25
# Turn off the main light
axes.light0.Off()
# Create a fixed light source
light_obj = axes.lights[1]
light_obj.On()
light_obj.position = (5.0, 5.0, 5.0, 0.0)
# The stuff above was basically from last time. Now, add Philae's trajectory
vv.plot(pp, ls='-', lc="w", lw=5)
# ... and run the application!
app.Run()
# Tasks:
#
# 1. Plot also Philae descent phase
# 2. Create an animation of the descent phase + the Sun's rays
```
| github_jupyter |
# Emukit Bayesian Optimization Benchmark
This notebook uses the `emukit.benchmarking` package to compare two Bayesian optimization methods against each other, using the Branin test function.
```
import emukit
import numpy as np
```
## Set up test function
We use the Branin function which is already included in Emukit, both the function and the appropriate input domain are ready made for us.
```
from emukit.test_functions.branin import branin_function
branin_fcn, parameter_space = branin_function()
```
## Set up methods to test
We compare Bayesian optimization using different models. All the methods collect points one at a time in a sequential fashion and use the expected improvement acquisition function. The models we test are:
- A Gaussian process with Matern52 covariance function
- Random forest using the pyrfr package
We choose to create lambda functions with a consistent interface that return an instance of a loop with a given initial data set.
```
from emukit.examples.enums import ModelType, AcquisitionType
from emukit.examples.optimization_loops import create_bayesian_optimization_loop
from emukit.examples.single_objective_bayesian_optimization import GPBayesianOptimization
loops = [
('Random Forest', lambda x, y: create_bayesian_optimization_loop(x, y, parameter_space, AcquisitionType.EI,
ModelType.RandomForest)),
('Gaussian Process', lambda x, y: GPBayesianOptimization(parameter_space.parameters, x, y,
acquisition_type=AcquisitionType.EI, noiseless=True))
]
```
# Run benchmark
A total of 10 initial data sets are generated of 5 observations that are randomly sampled from the input domain. For every intial data set, each method is run for 30 optimization iterations. The Gaussian process model has its hyper-parameters optimized after each function observation whereas the other models have fixed hyper-parameters.
```
from emukit.benchmarking.benchmarker import Benchmarker
from emukit.benchmarking.metrics import MinimumObservedValueMetric, TimeMetric
n_repeats = 30
n_initial_data = 5
n_iterations = 50
metrics = [MinimumObservedValueMetric(), TimeMetric()]
benchmarkers = Benchmarker(loops, branin_fcn, parameter_space, metrics=metrics)
benchmark_results = benchmarkers.run_benchmark(n_iterations=n_iterations, n_initial_data=n_initial_data,
n_repeats=n_repeats)
```
# Plot results
Plot the results of each method against each other. The plot shows the average value and standard deviation of the lowest observed value up to the given iteration.
```
from emukit.benchmarking.benchmark_plot import BenchmarkPlot
colours = ['m', 'c']
line_styles = ['-', '--']
metrics_to_plot = ['minimum_observed_value']
plots = BenchmarkPlot(benchmark_results, loop_colours=colours, loop_line_styles=line_styles,
metrics_to_plot=metrics_to_plot)
plots.make_plot()
```
# Plot results against time
Using the `TimeMetric` object above, the time taken to complete each iteration of the loops was recorded. Here we plot the minimum observed value against the time taken.
```
# Plot against time
plots = BenchmarkPlot(benchmark_results, loop_colours=colours, loop_line_styles=line_styles, x_axis='time',
metrics_to_plot=metrics_to_plot)
plots.make_plot()
```
# Conclusion
We have shown how to use Emukit to benchmark different methods against each other for Bayesian optimziation. This methodology can easily be expanded to more loops using different models and acquisition functions.
| github_jupyter |
# CNN for Classification
---
In this notebook, we define **and train** an CNN to classify images from the [Fashion-MNIST database](https://github.com/zalandoresearch/fashion-mnist).
### Load the [data](http://pytorch.org/docs/master/torchvision/datasets.html)
In this cell, we load in both **training and test** datasets from the FashionMNIST class.
```
# our basic libraries
import torch
import torchvision
# data loading and transforming
from torchvision.datasets import FashionMNIST
from torch.utils.data import DataLoader
from torchvision import transforms
# The output of torchvision datasets are PILImage images of range [0, 1].
# We transform them to Tensors for input into a CNN
## Define a transform to read the data in as a tensor
data_transform = transforms.ToTensor()
# choose the training and test datasets
train_data = FashionMNIST(root='./data', train=True,
download=True, transform=data_transform)
test_data = FashionMNIST(root='./data', train=False,
download=True, transform=data_transform)
# Print out some stats about the training and test data
print('Train data, number of images: ', len(train_data))
print('Test data, number of images: ', len(test_data))
# prepare data loaders, set the batch_size
## TODO: you can try changing the batch_size to be larger or smaller
## when you get to training your network, see how batch_size affects the loss
batch_size = 20
train_loader = DataLoader(train_data, batch_size=batch_size, shuffle=True)
test_loader = DataLoader(test_data, batch_size=batch_size, shuffle=True)
# specify the image classes
classes = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
### Visualize some training data
This cell iterates over the training dataset, loading a random batch of image/label data, using `dataiter.next()`. It then plots the batch of images and labels in a `2 x batch_size/2` grid.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(batch_size):
ax = fig.add_subplot(2, batch_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title(classes[labels[idx]])
```
### Define the network architecture
The various layers that make up any neural network are documented, [here](http://pytorch.org/docs/master/nn.html). For a convolutional neural network, we'll use a simple series of layers:
* Convolutional layers
* Maxpooling layers
* Fully-connected (linear) layers
You are also encouraged to look at adding [dropout layers](http://pytorch.org/docs/stable/nn.html#dropout) to avoid overfitting this data.
---
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the feedforward behavior of a network that employs those initialized layers in the function `forward`, which takes in an input image tensor, `x`. The structure of this Net class is shown below and left for you to fill in.
Note: During training, PyTorch will be able to perform backpropagation by keeping track of the network's feedforward behavior and using autograd to calculate the update to the weights in the network.
#### Define the Layers in ` __init__`
As a reminder, a conv/pool layer may be defined like this (in `__init__`):
```
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
```
#### Refer to Layers in `forward`
Then referred to in the `forward` function like this, in which the conv1 layer has a ReLu activation applied to it before maxpooling is applied:
```
x = self.pool(F.relu(self.conv1(x)))
```
You must place any layers with trainable weights, such as convolutional layers, in the `__init__` function and refer to them in the `forward` function; any layers or functions that always behave in the same way, such as a pre-defined activation function, may appear *only* in the `forward` function. In practice, you'll often see conv/pool layers defined in `__init__` and activations defined in `forward`.
#### Convolutional layer
The first convolution layer has been defined for you, it takes in a 1 channel (grayscale) image and outputs 10 feature maps as output, after convolving the image with 3x3 filters.
#### Flattening
Recall that to move from the output of a convolutional/pooling layer to a linear layer, you must first flatten your extracted features into a vector. If you've used the deep learning library, Keras, you may have seen this done by `Flatten()`, and in PyTorch you can flatten an input `x` with `x = x.view(x.size(0), -1)`.
### TODO: Define the rest of the layers
It will be up to you to define the other layers in this network; we have some recommendations, but you may change the architecture and parameters as you see fit.
Recommendations/tips:
* Use at least two convolutional layers
* Your output must be a linear layer with 10 outputs (for the 10 classes of clothing)
* Use a dropout layer to avoid overfitting
```
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# 1 input image channel (grayscale), 10 output channels/feature maps
# 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 10, 3)
## TODO: Define the rest of the layers:
# include another conv layer, maxpooling layers, and linear layers
# also consider adding a dropout layer to avoid overfitting
## TODO: define the feedforward behavior
def forward(self, x):
# one activated conv layer
x = F.relu(self.conv1(x))
# final output
return x
# instantiate and print your Net
net = Net()
print(net)
```
### TODO: Specify the loss function and optimizer
Learn more about [loss functions](http://pytorch.org/docs/master/nn.html#loss-functions) and [optimizers](http://pytorch.org/docs/master/optim.html) in the online documentation.
Note that for a classification problem like this, one typically uses cross entropy loss, which can be defined in code like: `criterion = nn.CrossEntropyLoss()`. PyTorch also includes some standard stochastic optimizers like stochastic gradient descent and Adam. You're encouraged to try different optimizers and see how your model responds to these choices as it trains.
```
import torch.optim as optim
## TODO: specify loss function (try categorical cross-entropy)
criterion = None
## TODO: specify optimizer
optimizer = None
```
### A note on accuracy
It's interesting to look at the accuracy of your network **before and after** training. This way you can really see that your network has learned something. In the next cell, let's see what the accuracy of an untrained network is (we expect it to be around 10% which is the same accuracy as just guessing for all 10 classes).
```
# Calculate accuracy before training
correct = 0
total = 0
# Iterate through test dataset
for images, labels in test_loader:
# forward pass to get outputs
# the outputs are a series of class scores
outputs = net(images)
# get the predicted class from the maximum value in the output-list of class scores
_, predicted = torch.max(outputs.data, 1)
# count up total number of correct labels
# for which the predicted and true labels are equal
total += labels.size(0)
correct += (predicted == labels).sum()
# calculate the accuracy
accuracy = 100 * correct / total
# print it out!
print('Accuracy before training: ', accuracy)
```
### Train the Network
Below, we've defined a `train` function that takes in a number of epochs to train for. The number of epochs is how many times a network will cycle through the training dataset.
Here are the steps that this training function performs as it iterates over the training dataset:
1. Zero's the gradients to prepare for a forward pass
2. Passes the input through the network (forward pass)
3. Computes the loss (how far is the predicted classes are from the correct labels)
4. Propagates gradients back into the network’s parameters (backward pass)
5. Updates the weights (parameter update)
6. Prints out the calculated loss
```
def train(n_epochs):
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
inputs, labels = data
# zero the parameter (weight) gradients
optimizer.zero_grad()
# forward pass to get outputs
outputs = net(inputs)
# calculate the loss
loss = criterion(outputs, labels)
# backward pass to calculate the parameter gradients
loss.backward()
# update the parameters
optimizer.step()
# print loss statistics
# to convert loss into a scalar and add it to running_loss, we use .item()
running_loss += loss.item()
if batch_i % 1000 == 999: # print every 1000 mini-batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/1000))
running_loss = 0.0
print('Finished Training')
# define the number of epochs to train for
n_epochs = 5 # start small to see if your model works, initially
# call train
train(n_epochs)
```
### Test the Trained Network
Once you are satisfied with how the loss of your model has decreased, there is one last step: test!
You must test your trained model on a previously unseen dataset to see if it generalizes well and can accurately classify this new dataset. For FashionMNIST, which contains many pre-processed training images, a good model should reach **greater than 85% accuracy** on this test dataset. If you are not reaching this value, try training for a larger number of epochs, tweaking your hyperparameters, or adding/subtracting layers from your CNN.
```
# initialize tensor and lists to monitor test loss and accuracy
test_loss = torch.zeros(1)
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
# set the module to evaluation mode
net.eval()
for batch_i, data in enumerate(test_loader):
# get the input images and their corresponding labels
inputs, labels = data
# forward pass to get outputs
outputs = net(inputs)
# calculate the loss
loss = criterion(outputs, labels)
# update average test loss
test_loss = test_loss + ((torch.ones(1) / (batch_i + 1)) * (loss.data - test_loss))
# get the predicted class from the maximum value in the output-list of class scores
_, predicted = torch.max(outputs.data, 1)
# compare predictions to true label
correct = np.squeeze(predicted.eq(labels.data.view_as(predicted)))
# calculate test accuracy for *each* object class
# we get the scalar value of correct items for a class, by calling `correct[i].item()`
for i in range(batch_size):
label = labels.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
print('Test Loss: {:.6f}\n'.format(test_loss.numpy()[0]))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize sample test results
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get predictions
preds = np.squeeze(net(images).data.max(1, keepdim=True)[1].numpy())
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(batch_size):
ax = fig.add_subplot(2, batch_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx] else "red"))
```
### Question: What are some weaknesses of your model? (And how might you improve these in future iterations.)
**Answer**: Double-click and write answer, here.
### Save Your Best Model
Once you've decided on a network architecture and are satisfied with the test accuracy of your model after training, it's time to save this so that you can refer back to this model, and use it at a later data for comparison of for another classification task!
```
## TODO: change the model_name to something uniqe for any new model
## you wish to save, this will save it in the saved_models directory
model_dir = 'saved_models/'
model_name = 'model_1.pt'
# after training, save your model parameters in the dir 'saved_models'
# when you're ready, un-comment the line below
# torch.save(net.state_dict(), model_dir+model_name)
```
### Load a Trained, Saved Model
To instantiate a trained model, you'll first instantiate a new `Net()` and then initialize it with a saved dictionary of parameters (from the save step above).
```
# instantiate your Net
# this refers to your Net class defined above
net = Net()
# load the net parameters by name
# uncomment and write the name of a saved model
#net.load_state_dict(torch.load('saved_models/model_1.pt'))
print(net)
# Once you've loaded a specific model in, you can then
# us it or further analyze it!
# This will be especialy useful for feature visualization
```
| github_jupyter |
# Visualizing a Gensim model
To illustrate how to use [`pyLDAvis`](https://github.com/bmabey/pyLDAvis)'s gensim [helper funtions](https://pyldavis.readthedocs.org/en/latest/modules/API.html#module-pyLDAvis.gensim) we will create a model from the [20 Newsgroup corpus](http://qwone.com/~jason/20Newsgroups/). Minimal preprocessing is done and so the model is not the best, the goal of this notebook is to demonstrate the the helper functions.
## Downloading the data
```
%%bash
mkdir -p data
pushd data
if [ -d "20news-bydate-train" ]
then
echo "The data has already been downloaded..."
else
wget http://qwone.com/%7Ejason/20Newsgroups/20news-bydate.tar.gz
tar xfv 20news-bydate.tar.gz
rm 20news-bydate.tar.gz
fi
echo "Lets take a look at the groups..."
ls 20news-bydate-train/
popd
```
## Exploring the dataset
Each group dir has a set of files:
```
ls -lah data/20news-bydate-train/sci.space | tail -n 5
```
Lets take a peak at one email:
```
!head data/20news-bydate-train/sci.space/61422 -n 20
```
## Loading the tokenizing the corpus
```
from glob import glob
import re
import string
import funcy as fp
from gensim import models
from gensim.corpora import Dictionary, MmCorpus
import nltk
import pandas as pd
# quick and dirty....
EMAIL_REGEX = re.compile(r"[a-z0-9\.\+_-]+@[a-z0-9\._-]+\.[a-z]*")
FILTER_REGEX = re.compile(r"[^a-z '#]")
TOKEN_MAPPINGS = [(EMAIL_REGEX, "#email"), (FILTER_REGEX, ' ')]
def tokenize_line(line):
res = line.lower()
for regexp, replacement in TOKEN_MAPPINGS:
res = regexp.sub(replacement, res)
return res.split()
def tokenize(lines, token_size_filter=2):
tokens = fp.mapcat(tokenize_line, lines)
return [t for t in tokens if len(t) > token_size_filter]
def load_doc(filename):
group, doc_id = filename.split('/')[-2:]
with open(filename, errors='ignore') as f:
doc = f.readlines()
return {'group': group,
'doc': doc,
'tokens': tokenize(doc),
'id': doc_id}
docs = pd.DataFrame(list(map(load_doc, glob('data/20news-bydate-train/*/*')))).set_index(['group','id'])
docs.head()
```
## Creating the dictionary, and bag of words corpus
```
def nltk_stopwords():
return set(nltk.corpus.stopwords.words('english'))
def prep_corpus(docs, additional_stopwords=set(), no_below=5, no_above=0.5):
print('Building dictionary...')
dictionary = Dictionary(docs)
stopwords = nltk_stopwords().union(additional_stopwords)
stopword_ids = map(dictionary.token2id.get, stopwords)
dictionary.filter_tokens(stopword_ids)
dictionary.compactify()
dictionary.filter_extremes(no_below=no_below, no_above=no_above, keep_n=None)
dictionary.compactify()
print('Building corpus...')
corpus = [dictionary.doc2bow(doc) for doc in docs]
return dictionary, corpus
dictionary, corpus = prep_corpus(docs['tokens'])
MmCorpus.serialize('newsgroups.mm', corpus)
dictionary.save('newsgroups.dict')
```
## Fitting the LDA model
```
%%time
lda = models.ldamodel.LdaModel(corpus=corpus, id2word=dictionary, num_topics=50, passes=10)
lda.save('newsgroups_50_lda.model')
```
## Visualizing the model with pyLDAvis
Okay, the moment we have all been waiting for is finally here! You'll notice in the visualizaiton that we have a few junk topics that would probably disappear after better preprocessing of the corpus. This is left as an exercises to the reader. :)
```
import pyLDAvis.gensim as gensimvis
import pyLDAvis
vis_data = gensimvis.prepare(lda, corpus, dictionary)
pyLDAvis.display(vis_data)
```
## Fitting the HDP model
We could visualize the LDA model with pyLDAvis, in the same maner, we can also visualize gensim HDP models with pyLDAvis.
The difference between HDP and LDA is that HDP is a non-parametric method. Which means you don't need to specify the number of topics, HDP will fit as many topics as it can and find the optimal number of topics by itself.
```
%%time
# The optional parameter T here indicates that HDP should find no more than 50 topics
# if there exists any.
hdp = models.hdpmodel.HdpModel(corpus, dictionary, T=50)
hdp.save('newsgroups_hdp.model')
```
## Visualizing the HDP model with pyLDAvis
As for the LDA model, you only need to give your model, the corpus and the dictionary associated to prepare the visualization.
```
vis_data = gensimvis.prepare(hdp, corpus, dictionary)
pyLDAvis.display(vis_data)
```
| github_jupyter |
# Using submodels in PyBaMM
In this notebook we show how to modify existing models by swapping out submodels, and how to build your own model from scratch using existing submodels. To see all of the models and submodels available in PyBaMM, please take a look at the documentation [here](https://pybamm.readthedocs.io/en/latest/source/models/index.html).
## Changing a submodel in an exisiting battery model
PyBaMM is designed to be a flexible modelling package that allows users to easily compare different models and numerical techniques within a common framework. Battery models within PyBaMM are built up using a number of submodels that describe different physics included within the model, such as mass conservation in the electrolyte or charge conservation in the solid. For ease of use, a number of popular battery models are pre-configured in PyBaMM. As an example, we look at the Single Particle Model (for more information see [here](./models/SPM.ipynb)).
First we import pybamm
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
```
Then we load the SPM
```
model = pybamm.lithium_ion.SPM()
```
We can look at the submodels that make up the SPM by accessing `model.submodels`, which is a dictionary of the submodel's name (i.e. the physics it represents) and the submodel that is selected
```
for name, submodel in model.submodels.items():
print(name, submodel)
```
When you load a model in PyBaMM it builds by default. Building the model sets all of the model variables and sets up any variables which are coupled between different submodels: this is the process which couples the submodels together and allows one submodel to access variables from another. If you would like to swap out a submodel in an existing battery model you need to load it without building it by passing the keyword `build=False`
```
model = pybamm.lithium_ion.SPM(build=False)
```
This collects all of the submodels which make up the SPM, but doesn't build the model. Now you are free to swap out one submodel for another. For instance, you may want to assume that diffusion within the negative particles is infinitely fast, so that the PDE describing diffusion is replaced with an ODE for the uniform particle concentration. To change a submodel you simply update the dictionary entry, in this case to the `XAveragedPolynomialProfile` submodel
```
model.submodels["negative particle"] = pybamm.particle.no_distribution.XAveragedPolynomialProfile(model.param, "Negative","uniform profile", options=model.options)
```
where we pass in the model parameters, the electrode (negative or positive) the submodel corresponds to, and the name of the polynomial we want to use. In the example we assume uniform concentration within the particle, corresponding to a zero-order polynomial.
Now if we look at the submodels again we see that the model for the negative particle has been changed
```
for name, submodel in model.submodels.items():
print(name, submodel)
```
Building the model also sets up the equations, boundary and initial conditions for the model. For example, if we look at `model.rhs` before building we see that it is empty
```
model.rhs
```
If we try to use this empty model, PyBaMM will give an error. So, before proceeding we must build the model
```
model.build_model()
```
Now if we look at `model.rhs` we see that it contains an entry relating to the concentration in each particle, as expected for the SPM
```
model.rhs
```
Now the model can be used in a simulation and solved in the usual way, and we still have access to model defaults such as the default geometry and default spatial methods which are used in the simulation
```
simulation = pybamm.Simulation(model)
simulation.solve([0, 3600])
simulation.plot()
```
## Building a custom model from submodels
Instead of editing a pre-existing model, you may wish to build your own model from scratch by combining existing submodels of you choice. In this section, we build a Single Particle Model in which the diffusion is assumed infinitely fast in both particles.
To begin, we load a base lithium-ion model. This sets up the basic model structure behind the scenes, and also sets the default parameters to be those corresponding to a lithium-ion battery. Note that the base model does not select any default submodels, so there is no need to pass `build=False`.
```
model = pybamm.lithium_ion.BaseModel()
```
Submodels can be added to the `model.submodels` dictionary in the same way that we changed the submodels earlier.
We use the simplest model for the external circuit, which is the "current control" submodel
```
model.submodels["external circuit"] = pybamm.external_circuit.CurrentControl(model.param)
```
We want to build a 1D model, so select the `Uniform` current collector model (if the current collectors are behaving uniformly, then a 1D model is appropriate). We also want the model to be isothermal, so select the thermal model accordingly. Further, we assume that the porosity and active material are constant in space and time.
```
model.submodels["current collector"] = pybamm.current_collector.Uniform(model.param)
model.submodels["thermal"] = pybamm.thermal.isothermal.Isothermal(model.param)
model.submodels["porosity"] = pybamm.porosity.Constant(model.param, model.options)
model.submodels["negative active material"] = pybamm.active_material.Constant(
model.param, "Negative", model.options
)
model.submodels["positive active material"] = pybamm.active_material.Constant(
model.param, "Positive", model.options
)
```
We assume that the current density varies linearly in the electrodes. This corresponds the the leading-order terms in Ohm's law in the limit in which the SPM is derived in [[3]](#References)
```
model.submodels["negative electrode potentials"] = pybamm.electrode.ohm.LeadingOrder(
model.param, "Negative"
)
model.submodels["positive electrode potentials"] = pybamm.electrode.ohm.LeadingOrder(
model.param, "Positive"
)
```
We assume uniform concentration in both the negative and positive particles
```
model.submodels["negative particle"] = pybamm.particle.no_distribution.PolynomialProfile(
model.param, "Negative", "uniform profile", options=model.options
)
model.submodels["positive particle"] = pybamm.particle.no_distribution.PolynomialProfile(
model.param, "Positive", "uniform profile", options=model.options
)
```
In the Single Particle Model, the overpotential can be obtianed by inverting the Butler-Volmer relation, so we choose the `InverseButlerVolmer` submodel for the interface, with the "main" lithium-ion reaction (and default lithium ion options). Because of how the current is implemented, we also need to separately specify the `CurrentForInverseButlerVolmer` submodel
```
model.submodels[
"negative interface"
] = pybamm.interface.InverseButlerVolmer(
model.param, "Negative", "lithium-ion main", options=model.options
)
model.submodels[
"positive interface"
] = pybamm.interface.InverseButlerVolmer(
model.param, "Positive", "lithium-ion main", options=model.options
)
model.submodels[
"negative interface current"
] = pybamm.interface.CurrentForInverseButlerVolmer(
model.param, "Negative", "lithium-ion main"
)
model.submodels[
"positive interface current"
] = pybamm.interface.CurrentForInverseButlerVolmer(
model.param, "Positive", "lithium-ion main"
)
model.submodels["negative interface utilisation"] = pybamm.interface_utilisation.Full(
model.param, "Negative", model.options
)
model.submodels["positive interface utilisation"] = pybamm.interface_utilisation.Full(
model.param, "Positive", model.options
)
```
We don't want any SEI formation or lithium plating in this model
```
model.submodels["sei"] = pybamm.sei.NoSEI(model.param)
model.submodels["lithium plating"] = pybamm.lithium_plating.NoPlating(model.param)
```
Finally, for the electrolyte we assume that diffusion is infinitely fast so that the concentration is uniform, and also use the leading-order model for charge conservation, which leads to a linear variation in ionic current in the electrodes
```
model.submodels["electrolyte diffusion"] = pybamm.electrolyte_diffusion.ConstantConcentration(
model.param
)
model.submodels["electrolyte conductivity"] = pybamm.electrolyte_conductivity.LeadingOrder(
model.param
)
```
Now that we have set all of the submodels we can build the model
```
model.build_model()
```
We can then use the model in a simulation in the usual way
```
simulation = pybamm.Simulation(model)
simulation.solve([0, 3600])
simulation.plot()
```
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
# 基于 GraphSage 的无监督学习
GraphScope提供了处理学习任务的功能。本次教程,我们将会展示GraphScope如何使用GraphSage算法训练一个无监督学习模型。
本次教程的学习任务是链接预测,通过计算在图中顶点之间存在边的概率来预测链接。
在这一任务中,我们使用GraphScope内置的GraphSage算法在[PPI](https://humgenomics.biomedcentral.com/articles/10.1186/1479-7364-3-3-291)数据集上训练一个模型,这一训练模型可以用来预测蛋白质结构之间的链接。这一任务可以被看作在一个异构链接网络上的无监督训练任务。
在这一任务中,GraphSage算法会将图中的结构信息和属性信息压缩为每个节点上的低维嵌入向量,这些嵌入和表征可以进一步用来预测节点间的链接。
这一教程将会分为以下几个步骤:
- 建立会话和载图
- 启动GraphScope的学习引擎,并将图关联到引擎上
- 使用内置的GCN模型定义训练过程,并定义相关的超参
- 开始训练
首先,我们要新建一个会话,并载入数据
```
import os
import graphscope
k8s_volumes = {
"data": {
"type": "hostPath",
"field": {
"path": "/testingdata",
"type": "Directory"
},
"mounts": {
"mountPath": "/home/jovyan/datasets",
"readOnly": True
}
}
}
# 建立会话
graphscope.set_option(show_log=True)
sess = graphscope.session(k8s_volumes=k8s_volumes)
# 加载PPI图数据
graph = sess.g()
graph = graph.add_vertices("/home/jovyan/datasets/ppi/node.csv", "protein")
graph = graph.add_edges("/home/jovyan/datasets/ppi/edge.csv", "link")
```
## Launch learning engine
然后,我们需要定义一个特征列表用于图的训练。训练特征集合必须从点的属性集合中选取。在这个例子中,我们选择了属性集合中所有以"feat-"为前缀的属性作为训练特征集,这一特征集也是PPI数据中点的特征集。
借助定义的特征列表,接下来,我们使用会话的`learning`方法来开启一个学习引擎。(`learning`方法的文档可参考[Session](https://graphscope.io/docs/reference/session.html))
在这个例子中,我们在`learning`方法中,指定在数据中`protein`类型的顶点和`link`类型边上进行模型训练。
利用`gen_labels`参数,我们将`protein`点数据集作为训练集。
```
# define the features for learning
paper_features = []
for i in range(50):
paper_features.append("feat-" + str(i))
# launch a learning engine.
lg = sess.learning(graph, nodes=[("protein", paper_features)],
edges=[("protein", "link", "protein")],
gen_labels=[
("train", "protein", 100, (0, 100)),
])
```
这里我们使用内置的GraphSage模型定义训练过程。你可以在[Graph Learning Model](https://graphscope.io/docs/learning_engine.html#data-model)获取更多内置学习模型的信息。
在本次示例中,我们使用tensorflow作为NN后端训练器。
```
import numpy as np
from graphscope.learning.examples import GraphSage
from graphscope.learning.graphlearn.python.model.tf.trainer import LocalTFTrainer
from graphscope.learning.graphlearn.python.model.tf.optimizer import get_tf_optimizer
# unsupervised GraphSage.
def train(config, graph):
def model_fn():
return GraphSage(
graph,
config["class_num"],
config["features_num"],
config["batch_size"],
categorical_attrs_desc=config["categorical_attrs_desc"],
hidden_dim=config["hidden_dim"],
in_drop_rate=config["in_drop_rate"],
neighs_num=config["neighs_num"],
hops_num=config["hops_num"],
node_type=config["node_type"],
edge_type=config["edge_type"],
full_graph_mode=config["full_graph_mode"],
unsupervised=config['unsupervised'],
)
trainer = LocalTFTrainer(
model_fn,
epoch=config["epoch"],
optimizer=get_tf_optimizer(
config["learning_algo"], config["learning_rate"], config["weight_decay"]
),
)
trainer.train()
embs = trainer.get_node_embedding()
np.save(config['emb_save_dir'], embs)
# define hyperparameters
config = {
"class_num": 128, # output dimension
"features_num": 50,
"batch_size": 512,
"categorical_attrs_desc": "",
"hidden_dim": 128,
"in_drop_rate": 0.5,
"hops_num": 2,
"neighs_num": [5, 5],
"full_graph_mode": False,
"agg_type": "gcn", # mean, sum
"learning_algo": "adam",
"learning_rate": 0.01,
"weight_decay": 0.0005,
'unsupervised': True,
"epoch": 1,
'emb_save_dir': './id_emb',
"node_type": "protein",
"edge_type": "link",
}
```
## Run training process
在定义完训练过程和超参后,现在我们可以使用学习引擎和定义的超参开始训练过程。
```
train(config, lg)
```
训练完毕后,需要关掉会话
```
sess.close()
```
| github_jupyter |
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Accuracy-Calculation-COST-323" data-toc-modified-id="Accuracy-Calculation-COST-323-1"><span class="toc-item-num">1 </span>Accuracy Calculation COST 323</a></div><div class="lev2 toc-item"><a href="#Description-of-the-accuracy-calculation-sheet" data-toc-modified-id="Description-of-the-accuracy-calculation-sheet-11"><span class="toc-item-num">1.1 </span>Description of the accuracy calculation sheet</a></div><div class="lev2 toc-item"><a href="#Source" data-toc-modified-id="Source-12"><span class="toc-item-num">1.2 </span>Source</a></div><div class="lev3 toc-item"><a href="#Tolarance-classes" data-toc-modified-id="Tolarance-classes-121"><span class="toc-item-num">1.2.1 </span>Tolarance classes</a></div><div class="lev2 toc-item"><a href="#Test" data-toc-modified-id="Test-13"><span class="toc-item-num">1.3 </span>Test</a></div><div class="lev1 toc-item"><a href="#References" data-toc-modified-id="References-2"><span class="toc-item-num">2 </span>References</a></div>
# Accuracy Calculation COST 323
The accuracy calculation can be consulting in \cite{jacob1998european}. In the
accuracy calculation sheet provided by COST 323, we can extract the follow
information:
## Description of the accuracy calculation sheet
Initial verification: (Yes=1, No=0)
Sheet Fields (the header columns stay in line 8 and 9 into the sheet):
* SYSTEM/Entity (column A/Index),
* Number (column B) [Input],
* Identified (column C) [Input],
* Mean (column D) [Input],
* Std deviat (column E) [Input],
* p_o (column F),
* Class (column G),
* d (column H),
* d_min (column I) [Input/Minimization Solver Output],
* d_c (column J),
* class (column K),
* p (column L) - related to column I,
* p (column M) - related to column H,
* p' (column N),
* Accepted (column O)
Test plan
* "r1"=full repeatability
* "r2"=extended repeatability
* "rr1"=limited reproducibility
* "rr2"=full reproducibility
Env
* "I"=environmental repeatability
* "II"=environmental limited reproducibility
* "III"=environmental full reproducibility
Description:
* Number: number of measured vehicles/axles/etc. which are valid and kept for the
test (may be less than those of the test plan)
* Identified: percentage of the total number of measured vehicles/axles/etc.
which are valid and kept for the test (enter by hand)
* Mean, Std Deviation: mean and standard deviation of the individual relative
errors with respect to the static reference loads
* p_o: minimum required level of confidence within the class tolerance d, i.e.
of the confidence interval [-d;d]
* d_min: for each criterion, tolerance (half width of the confidence interval)
which exactly corresponds to the minimum required level of confidence p_o
* d_c: "best acceptable interpolated class for the given criterion", i.e.
tolerance on the GW which exactly corresponds to the d_min of the given
criterion
## Source
```
from matplotlib import pyplot as plt
from scipy.stats import norm
from scipy.optimize import root, fsolve
from scipy import stats
import numba as nb
import numpy as np
import pandas as pd
%matplotlib inline
def calc_min_confidence(
data: pd.DataFrame, test_plan: str, env_condition: str
):
"""
=100*(2*NORMDIST(\\
IF(\$B\$5="r1";2,675/IF(\$D\$5="III";1,1;IF(\$D\$5="II";1,05;1));\\
IF(\$B\$5="r2";2,36/IF(\$D\$5="III";1,1;IF(\$D\$5="II";1,05;1));\\
IF(\$B\$5="RR1";2,155/IF(\$D\$5="III";1,1;IF(\$D\$5="II";1,05;1));\\
IF(\$B\$5="RR2";2/IF(\$D\$5="III";1,1;IF(\$D\$5="II";1,05;1));0))\\
))-TINV(0,05;B9-1)/SQRT(B9);0;1;TRUE())-1)
"""
# data = data.copy()
def _calc(v: float):
if env_condition=='I':
_v = 1
elif env_condition=='II':
_v = 1.05
elif env_condition=='III':
_v = 1.1
else:
raise Exception('INVALID_ENV_CONDITION')
if test_plan=='R1':
_v = 2.675/_v
elif test_plan=='R2':
_v = 2.36/_v
elif test_plan=='RR1':
_v = 2.155/_v
elif test_plan=='RR2':
_v = 2/_v
else:
raise Exception('INVALID_TEST_PLAN')
# in isf, the probabity is divided by 2
# because in excel TINV is 2-side tail
return 100 * (
2 * norm.cdf(_v-stats.t.isf(0.05/2, v-1)/np.sqrt(v), 0, 1)-1
)
test_plan = test_plan.upper()
env_condition = env_condition.upper()
data['min_confidence'] = data.number.apply(_calc)
return data
def calc_best_acceptable_class(
data: pd.DataFrame, initial_verification: bool
) -> pd.DataFrame:
"""
"""
# data = data.copy()
# IF(M$4=0;1;1,25)
factor = 1.25 if initial_verification else 1
best_acceptable_class = []
# gwv
# =I9∗IF(M$4=0;1;1,25)
best_acceptable_class.append(data.loc['gwv', 'min_tolerance'] * factor)
# group of axles
# =IF(I10<10; 0,7∗I10∗IF(M$4=0;1;1,25); I10∗IF(M$4=0;1;1,25)−3)
v = data.loc['group_axles', 'min_tolerance']
v = v*0.7*factor if v < 10 else v*factor-3
best_acceptable_class.append(v)
# single axle
# =IF(I11<15;
# I11∗(I11∗IF(M$4=0;1;1,25)+97)∗IF(M$4=0;1;1,25)/168;
# I11∗IF(M$4=0;1;1,25)−5)
v = data.loc['single_axle', 'min_tolerance']
v = v*(v*factor+97)*factor/168 if v < 15 else v*factor-5
best_acceptable_class.append(v)
# axle of a group
# =IF(I12<20;I12∗IF(M$4=0;1;1,25)/2;(I12∗IF(M$4=0;1;1,25)−10))
v = data.loc['axle_group', 'min_tolerance']
v = v*factor/2 if v < 20 else v*factor-10
best_acceptable_class.append(v)
data['best_acceptable_class'] = best_acceptable_class
return data
def calc_classification(data: pd.DataFrame) -> pd.DataFrame:
"""
=IF(OR(J9<=5;J9>7);ROUNDUP((J9/5);0)*5;7)
"""
def _calc(v: float):
# =IF(OR(J9<=5;J9>7);ROUNDUP((J9/5);0)*5;7)
return np.ceil(v/5)*5 if v<=5 or v>7 else 7
data['class_value'] = data.best_acceptable_class.apply(_calc)
return data
def resolve_class_name(data: pd.DataFrame) -> pd.DataFrame:
"""
=IF(K9<=5;CONCATENATE("A(";TEXT(K9;"0");")");\\
IF(K9<=7;CONCATENATE("B+(";TEXT(K9;"0");")");\\
IF(K9<=10;CONCATENATE("B(";TEXT(K9;"0");")");\\
IF(K9<=15;CONCATENATE("C(";TEXT(K9;"0");")");\\
IF(K9<=20;CONCATENATE("D+(";TEXT(K9;"0");")");\\
IF(K9<=25;CONCATENATE("D(";TEXT(K9;"0");")");\\
CONCATENATE("E(";TEXT(K9;"0");")")))))))
"""
def _resolve(v: int):
c = (
'A' if v<=5 else
'B+' if v<=7 else
'B' if v<=10 else
'C' if v<=15 else
'D+' if v<=20 else
'D' if v<=25 else
'E'
)
return '%s(%s)' % (c, int(v))
data['class_name'] = data.class_value.apply(_resolve)
return data
def calc_delta(data: pd.DataFrame, initial_verification: bool):
"""
"""
d = []
# factor
# IF(M$4=0;1;0,8)
factor = 0.8 if initial_verification else 1
# gwv
# =K9*IF(M$4=0;1;0,8)
d.append(data.loc['gwv', 'class_value']*factor)
# group of axles
# =IF(K10<7;K10/0,7;IF(K10<30;K10+3;K10*1,1))*IF(M$4=0;1;0,8)
v = data.loc['group_axles', 'class_value']
v = v/0.7 if v<7 else v+3 if v<30 else v*1.1
d.append(v*factor)
# single axle
# =IF(K11<10;K11*(85-K11)/50;IF(K11<25;K11+5;6*K11/5))*IF(M$4=0;1;0,8)
v = data.loc['single_axle', 'class_value']
v = v*(85-v)/50 if v<10 else v+5 if v<25 else 6*v/5
d.append(v*factor)
# axle of group
# =IF(K12<10;2*K12;IF(K12<25;K12+10;6*K12/5+5))*IF(M$4=0;1;0,8)
v = data.loc['axle_group', 'class_value']
v = 2*v if v<10 else v+10 if v<25 else 6*v/5+5
d.append(v*factor)
data['d'] = d
return d
def calc_confidence_level(data: pd.DataFrame) -> pd.DataFrame:
"""
* Number (column B) [Input],
* Identified (column C) [Input],
* Mean (column D) [Input],
* Std deviat (column E) [Input],
* p_o (column F),
* Class (column G),
* d (column H),
* d_min (column I) [Input/Minimization Solver Output],
* d_c (column J),
* class (column K),
* p (column L) - related to column I,
* p (column M) - related to column H,
* Accepted (column O)
=100*(
1-TDIST((H9/E9-D9/E9)-TINV(0,05;B9-1)/SQRT(B9);B9-1;1)-
TDIST((H9/E9+D9/E9)-TINV(0,05;B9-1)/SQRT(B9);B9-1;1)
)
"""
def _calc(v: pd.Series) -> pd.Series:
return 100*(
1-stats.t.sf(
(v.d/v['std']-v['mean']/v['std'])-stats.t.isf(0.05/2, v.number-1)/
np.sqrt(v.number),
v.number-1
)-stats.t.sf(
(v.d/v['std']+v['mean']/v['std'])-stats.t.isf(0.05/2, v.number-1)/
np.sqrt(v.number),v.number-1
)
)
data['confidence_level'] = data.T.apply(_calc)
return data
def resolve_accepted_class(data: pd.DataFrame) -> str:
"""
O12 = MAX(K11:K12)
=IF(O12<=5;CONCATENATE("A(";TEXT(O12;"0");")");\\
IF(O12<=7;CONCATENATE("B+(";TEXT(O12;"0");")");\\
IF(O12<=10;CONCATENATE("B(";TEXT(O12;"0");")");\\
IF(O12<=15;CONCATENATE("C(";TEXT(O12;"0");")");\\
IF(O12<=20;CONCATENATE("D+(";TEXT(O12;"0");")");\\
IF(O12<=25;CONCATENATE("D(";TEXT(O12;"0");")");\\
CONCATENATE("E(";TEXT(O12;"0");")")))))))
"""
v = data['class_value'].max()
c = (
'A' if v<=5 else
'B+' if v<=7 else
'B' if v<=10 else
'C' if v<=15 else
'D+' if v<=20 else
'D' if v<=25 else
'E'
)
return '%s(%s)' % (c, int(v))
def solver_min_tolerance(data: pd.DataFrame) -> pd.DataFrame:
"""
* Number (column B) [Input],
* Identified (column C) [Input],
* Mean (column D) [Input],
* Std deviat (column E) [Input],
* p_o (column F),
* Class (column G),
* d (column H),
* d_min (column I) [Input/Minimization Solver Output],
* d_c (column J),
* class (column K),
* p (column L) - related to column I,
* p (column M) - related to column H,
* Accepted (column O)
=100*(
1-
TDIST((I9/E9-D9/E9)-TINV(0,05;B9-1)/SQRT(B9);B9-1;1)-
TDIST((I9/E9+D9/E9)-TINV(0,05;B9-1)/SQRT(B9);B9-1;1)
)
"""
for i in data.index:
s = data.loc[i, :]
_number = s['number']
_mean = s['mean']
_std = s['std']
_min_confidence = s['min_confidence']
_factor = stats.t.isf(0.05/2, _number-1)/np.sqrt(_number)
_dof = _number-1
func = lambda _min_tolerance: _min_confidence-100*(
1-
stats.t.sf(
(_min_tolerance/_std-_mean/_std)-_factor, _dof)-
stats.t.sf(
(_min_tolerance/_std+_mean/_std)-_factor, _dof)
)
try:
data.loc[i, 'min_tolerance'] = fsolve(func, [1])[0]
except:
data.loc[i, 'min_tolerance'] = np.nan
return data
```
### Tolarance classes
```
# 'A', 'B+', 'B', 'C', 'D+', 'D', 'E'
tolerance_table = pd.DataFrame(
[], columns=('A', 'B+', 'B', 'C', 'D+', 'D', 'E'),
index=(
'gwv', 'load_group_axles', 'load_single_axle', 'load_axle_group'
)
)
tolerance_table.loc['gwv',:] = [5, 7, 10, 15, 20, 25, np.inf]
tolerance_table.loc['load_group_axles',:] = [7, 10, 13, 18, 23, 28, np.inf]
tolerance_table.loc['load_single_axle',:] = [8, 11, 15, 20, 25, 30, np.inf]
tolerance_table.loc['load_axle_group',:] = [10, 14, 20, 25, 30, 35, np.inf]
tolerance_table
```
## Test
```
test_plan = 'RR2'
env_condition = 'I'
initial_verification = False
cols = ['number', 'valid_measures', 'mean', 'std']
indx = ['gwv', 'group_axles', 'single_axle', 'axle_group']
data = pd.DataFrame(
[[86, 96.15, -2.27, 6.09],
[66, 96.15, 1.00, 8.44],
[197, 95.50, -3.92, 7.66],
[169, 96.00, -0.19, 10.07]],
columns=cols, index=indx
)
data
print('Minimum confidence calcuation Test', end=' ... ')
calc_min_confidence(
data=data,
test_plan=test_plan,
env_condition=env_condition
)
np.testing.assert_allclose(
data['min_confidence'].values,
np.array([92.584, 92.060, 93.704, 93.542]),
atol=0.001 # becouse round function
)
print('SUCCESS')
print('Best acceptable class Test', end=' ... ')
data['min_tolerance'] = [
13.0293783341772,
17.2176428723888,
17.0913086307026,
20.2663837816549,
]
calc_best_acceptable_class(
data=data, initial_verification=initial_verification
)
np.testing.assert_allclose(
data['best_acceptable_class'].values,
np.array([13.029, 14.218, 12.091, 10.266]),
atol=0.001 # because round function
)
print('SUCCESS')
print('Class Value Test', end=' ... ')
calc_classification(data=data)
np.testing.assert_allclose(
data['class_value'].values,
np.array([15, 15, 15, 15])
)
print('SUCCESS')
print('Class Name Test', end=' ... ')
resolve_class_name(data=data)
np.testing.assert_array_equal(
data['class_name'].values,
np.array(['C(15)', 'C(15)', 'C(15)', 'C(15)'])
)
print('SUCCESS')
print('delta Test', end=' ... ')
calc_delta(data=data, initial_verification=initial_verification)
np.testing.assert_allclose(
data['d'].values,
np.array([15, 18, 20, 25])
)
print('SUCCESS')
data
print('Confidence Test', end=' ... ')
calc_confidence_level(data=data)
np.testing.assert_allclose(
data['confidence_level'].values,
np.array([96.276, 93.461, 97.260, 97.902]),
atol=0.001
)
print('SUCCESS')
print('Accepted Class Test', end=' ... ')
np.testing.assert_equal(
resolve_accepted_class(data=data),
'C(15)'
)
print('SUCCESS')
print('Minimum Tolerance Solver Test', end=' ... ')
data.min_tolerance = 0
%time solver_min_tolerance(data=data)
np.testing.assert_allclose(
data.min_tolerance.values,
np.array([
13.0293783341772,
17.2176428723888,
17.0913086307026,
20.2663837816549
]), atol=1e-5
)
print('SUCCESS')
# show graphical visualization about the solver function to achieve the
# mininum tolerance
# data sheet value
data_y = [
13.0293783341772,
17.2176428723888,
17.0913086307026,
20.2663837816549
]
_delta = 1e-5
lims = [
(data_y[0]-_delta, data_y[0]+_delta),
(data_y[1]-_delta, data_y[1]+_delta),
(data_y[2]-_delta, data_y[2]+_delta),
(data_y[3]-_delta, data_y[3]+_delta),
]
for k, s in data.T.items():
_number = s['number']
_mean = s['mean']
_std = s['std']
_min_confidence = s['min_confidence']
_factor = stats.t.isf(0.05/2, _number-1)/np.sqrt(_number)
_dof = _number-1
func = lambda _min_tolerance: _min_confidence-100*(
1-
stats.t.sf(
(_min_tolerance/_std-_mean/_std)-_factor, _dof)-
stats.t.sf(
(_min_tolerance/_std+_mean/_std)-_factor, _dof)
)
xlim = lims.pop(0)
x = np.linspace(xlim[0], xlim[1], 1000)
ax = plt.figure().gca()
data_func = pd.DataFrame({k: [func(xi) for xi in x]}, index=x)
data_func.plot(ax=ax)
ax.plot(data_y.pop(0), 0, 'o', label='excel', color='black')
ax.plot(fsolve(func, [1]), 0, 'o', label='fsolve', color='green')
ax.legend()
locs, labels = plt.xticks()
plt.setp(labels, rotation=45)
plt.grid(True)
plt.show()
```
# References
(<a id="cit-jacob1998european" href="#call-jacob1998european">JACOB and O'Brien, 1998</a>) B. JACOB and E.J. O'Brien, ``_European Specification on Weigh-In-Motion of Road Vehicles (COST323)_'', SECOND EUROPEAN CONFERENCE ON WEIGH-IN-MOTION OF ROAD VEHICLES, HELD LISBON, PORTUGAL 14-16 SEPTEMBER 1998, 1998.
| github_jupyter |
```
# imports
import openai
import git
import yaml
import pathlib
import json
# for all of the directories under certified-operators/operators, go through their subdirectories and look at each manifests/*.clusterserviceversion.yaml file to extract the fields in .spec.install.spec.Deployments, and return those as a list
def get_operator_deployments(operator_dir: str) -> list[dict]:
deployments = []
# go through each directory under certified-operators/operators
for dir in pathlib.Path(operator_dir).iterdir():
# skip if not a directory
if not dir.is_dir():
continue
# go through each subdirectory under the current directory
for subdir in dir.iterdir():
# skip if not a directory
if not subdir.is_dir():
continue
# look at each manifests/*.clusterserviceversion.yaml file
for file in subdir.glob('manifests/*.clusterserviceversion.yaml'):
# parse the yaml file
with open(file, 'r') as f:
yaml_data = yaml.load(f, Loader=yaml.FullLoader)
# extract the spec.install.spec.Deployments fields
if 'spec' in yaml_data and 'install' in yaml_data['spec'] and 'spec' in yaml_data['spec']['install'] and 'deployments' in yaml_data['spec']['install']['spec']:
deploy = {
'operatorName': yaml_data['metadata']['name'],
'deployments': yaml_data['spec']['install']['spec']['deployments']
}
# return the list of deployments
deployments.append(deploy)
return deployments
deployments = get_operator_deployments('certified-operators/operators')
# deployments
# dump all of the deployments to a file titled deployments.yaml
with open('deployments.yaml', 'w') as f:
yaml.dump(deployments, f)
# create a sample set of training data for OpenAI in a JSONL format like this:
completions = [
{
"prompt": "# deploy a ConfigMap named my-config-map\nKind: ConfigMap",
"completion": "\nmetadata:\n\tname: my-config-map\nspec:\n\tmy-data: this is my data\n"
},
{
"prompt": "# deploy a Service named my-service\nKind: Service",
"completion": "\nmetadata:\n\tname: my-service\nspec:\n\tports:\n\t- port: 80\n\t- port: 443\n"
}
]
training_data = []
# go through each directory in completions/ and obtain the representation for completion.yml and prompt.yml, then place them into a dict and append it to the training data list
for dir in pathlib.Path('completions/').iterdir():
# skip if not a directory
if not dir.is_dir():
continue
sample = {
'prompt': '',
'completion': ''
}
# read the contents of completion.yml into a string
with open(dir / 'completion.yml', 'r') as f:
sample['completion'] = f.read()
# read the contents of prompt.yml into a string
with open(dir / 'prompt.yml', 'r') as f:
sample['prompt'] = f.read()
# append the dict to the training data list
training_data.append(sample)
# output the training data into a json file
with open('training_data.jsonl', 'w') as f:
json.dump(training_data, f)
# create a set of training data for openai based on the deployments extracted
training_data = []
# process the list of deployments in deployments.yaml
with open('deployments.yaml', 'r') as f:
deployments = yaml.load(f, Loader=yaml.FullLoader)
# for each deployment, extract the operatorName and the deployments fields
for deployment in deployments:
prompt = '# deploy a Deployment named ' + deployment['operatorName'] + '\nKind: Deployment'
# format each deployment dict in the deployments field into a string
deployment_strs = [yaml.dump(d, default_flow_style=False) for d in deployment['deployments']]
# join the deployments fields into a string separated by '---'
completion = '\n---\n'.join(deployment_strs)
# append the test data into our training_data set
training_data.append({
'prompt': prompt,
'completion': completion
})
# dump the training data into a json file
with open('training_data.json', 'w') as f:
json.dump(training_data, f)
```
| github_jupyter |
## SFR package example
Demonstrates functionality of Flopy SFR module using the example documented by [Prudic and others (2004)](http://pubs.er.usgs.gov/publication/ofr20041042):
#### Problem description:
* Grid dimensions: 1 Layer, 15 Rows, 10 Columns
* Stress periods: 1 steady
* Flow package: LPF
* Stress packages: SFR, GHB, EVT, RCH
* Solver: SIP
<img src="./img/Prudic2004_fig6.png" width="400" height="500"/>
```
import sys
import platform
import os
import numpy as np
import glob
import shutil
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
import flopy.utils.binaryfile as bf
from flopy.utils.sfroutputfile import SfrFile
mpl.rcParams['figure.figsize'] = (11, 8.5)
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('pandas version: {}'.format(pd.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
#Set name of MODFLOW exe
# assumes executable is in users path statement
exe_name = 'mf2005'
if platform.system() == 'Windows':
exe_name += '.exe'
```
#### copy over the example files to the working directory
```
path = 'data'
if os.path.isfile(path):
os.remove(path)
elif os.path.isdir(path):
shutil.rmtree(path)
os.mkdir(path)
gpth = os.path.join('..', 'data', 'mf2005_test', 'test1ss.*')
for f in glob.glob(gpth):
shutil.copy(f, path)
gpth = os.path.join('..', 'data', 'mf2005_test', 'test1tr.*')
for f in glob.glob(gpth):
shutil.copy(f, path)
```
### Load example dataset, skipping the SFR package
```
m = flopy.modflow.Modflow.load('test1ss.nam', version='mf2005', exe_name=exe_name,
model_ws=path, load_only=['ghb', 'evt', 'rch', 'dis', 'bas6', 'oc', 'sip', 'lpf'])
oc = m.oc
oc.stress_period_data
```
### Read pre-prepared reach and segment data into numpy recarrays using numpy.genfromtxt()
Reach data (Item 2 in the SFR input instructions), are input and stored in a numpy record array
http://docs.scipy.org/doc/numpy/reference/generated/numpy.recarray.html
This allows for reach data to be indexed by their variable names, as described in the SFR input instructions.
For more information on Item 2, see the Online Guide to MODFLOW:
<http://water.usgs.gov/nrp/gwsoftware/modflow2000/MFDOC/index.html?sfr.htm>
```
rpth = os.path.join('..', 'data', 'sfr_examples', 'test1ss_reach_data.csv')
reach_data = np.genfromtxt(rpth, delimiter=',', names=True)
reach_data
```
### Segment Data structure
Segment data are input and stored in a dictionary of record arrays, which
```
spth = os.path.join('..', 'data', 'sfr_examples', 'test1ss_segment_data.csv')
ss_segment_data = np.genfromtxt(spth, delimiter=',', names=True)
segment_data = {0: ss_segment_data}
segment_data[0][0:1]['width1']
```
### define dataset 6e (channel flow data) for segment 1
dataset 6e is stored in a nested dictionary keyed by stress period and segment,
with a list of the following lists defined for each segment with icalc == 4
FLOWTAB(1) FLOWTAB(2) ... FLOWTAB(NSTRPTS)
DPTHTAB(1) DPTHTAB(2) ... DPTHTAB(NSTRPTS)
WDTHTAB(1) WDTHTAB(2) ... WDTHTAB(NSTRPTS)
```
channel_flow_data = {0: {1: [[0.5, 1.0, 2.0, 4.0, 7.0, 10.0, 20.0, 30.0, 50.0, 75.0, 100.0],
[0.25, 0.4, 0.55, 0.7, 0.8, 0.9, 1.1, 1.25, 1.4, 1.7, 2.6],
[3.0, 3.5, 4.2, 5.3, 7.0, 8.5, 12.0, 14.0, 17.0, 20.0, 22.0]]}}
```
### define dataset 6d (channel geometry data) for segments 7 and 8
dataset 6d is stored in a nested dictionary keyed by stress period and segment,
with a list of the following lists defined for each segment with icalc == 4
FLOWTAB(1) FLOWTAB(2) ... FLOWTAB(NSTRPTS)
DPTHTAB(1) DPTHTAB(2) ... DPTHTAB(NSTRPTS)
WDTHTAB(1) WDTHTAB(2) ... WDTHTAB(NSTRPTS)
```
channel_geometry_data = {0: {7: [[0.0, 10.0, 80.0, 100.0, 150.0, 170.0, 240.0, 250.0],
[20.0, 13.0, 10.0, 2.0, 0.0, 10.0, 13.0, 20.0]],
8: [[0.0, 10.0, 80.0, 100.0, 150.0, 170.0, 240.0, 250.0],
[25.0, 17.0, 13.0, 4.0, 0.0, 10.0, 16.0, 20.0]]}}
```
### Define SFR package variables
```
nstrm = len(reach_data) # number of reaches
nss = len(segment_data[0]) # number of segments
nsfrpar = 0 # number of parameters (not supported)
nparseg = 0
const = 1.486 # constant for manning's equation, units of cfs
dleak = 0.0001 # closure tolerance for stream stage computation
ipakcb = 53 # flag for writing SFR output to cell-by-cell budget (on unit 53)
istcb2 = 81 # flag for writing SFR output to text file
dataset_5 = {0: [nss, 0, 0]} # dataset 5 (see online guide)
```
### Instantiate SFR package
Input arguments generally follow the variable names defined in the Online Guide to MODFLOW
```
sfr = flopy.modflow.ModflowSfr2(m, nstrm=nstrm, nss=nss, const=const, dleak=dleak, ipakcb=ipakcb, istcb2=istcb2,
reach_data=reach_data,
segment_data=segment_data,
channel_geometry_data=channel_geometry_data,
channel_flow_data=channel_flow_data,
dataset_5=dataset_5, unit_number=15)
sfr.reach_data[0:1]
```
### Plot the SFR segments
any column in the reach_data array can be plotted using the ```key``` argument
```
sfr.plot(key='iseg');
```
### Check the SFR dataset for errors
```
chk = sfr.check()
m.external_fnames = [os.path.split(f)[1] for f in m.external_fnames]
m.external_fnames
m.write_input()
m.run_model()
```
### Load SFR formated water balance output into pandas dataframe using the `SfrFile` class
* requires the **pandas** library
```
sfr_outfile = os.path.join('..', 'data', 'sfr_examples', 'test1ss.flw')
sfrout = SfrFile(sfr_outfile)
df = sfrout.get_dataframe()
df.head()
```
#### Plot streamflow and stream/aquifer interactions for a segment
```
inds = df.segment == 3
print(df.reach[inds].astype(str))
#ax = df.ix[inds, ['Qin', 'Qaquifer', 'Qout']].plot(x=df.reach[inds])
ax = df.loc[inds, ['reach', 'Qin', 'Qaquifer', 'Qout']].plot(x='reach')
ax.set_ylabel('Flow, in cubic feet per second')
ax.set_xlabel('SFR reach');
```
### Look at stage, model top, and streambed top
```
streambed_top = m.sfr.segment_data[0][m.sfr.segment_data[0].nseg == 3][['elevup', 'elevdn']][0]
streambed_top
df['model_top'] = m.dis.top.array[df.row.values - 1, df.column.values -1]
fig, ax = plt.subplots()
plt.plot([1, 6], list(streambed_top), label='streambed top')
#ax = df.loc[inds, ['stage', 'model_top']].plot(ax=ax, x=df.reach[inds])
ax = df.loc[inds, ['reach', 'stage', 'model_top']].plot(ax=ax, x='reach')
ax.set_ylabel('Elevation, in feet')
plt.legend();
```
### Get SFR leakage results from cell budget file
```
bpth = os.path.join('data', 'test1ss.cbc')
cbbobj = bf.CellBudgetFile(bpth)
cbbobj.list_records()
sfrleak = cbbobj.get_data(text=' STREAM LEAKAGE')[0]
sfrleak[sfrleak == 0] = np.nan # remove zero values
```
### Plot leakage in plan view
```
im = plt.imshow(sfrleak[0], interpolation='none', cmap='coolwarm', vmin = -3, vmax=3)
cb = plt.colorbar(im, label='SFR Leakage, in cubic feet per second');
```
### Plot total streamflow
```
sfrQ = sfrleak[0].copy()
sfrQ[sfrQ == 0] = np.nan
sfrQ[df.row.values-1, df.column.values-1] = df[['Qin', 'Qout']].mean(axis=1).values
im = plt.imshow(sfrQ, interpolation='none')
plt.colorbar(im, label='Streamflow, in cubic feet per second');
```
## Reading transient SFR formatted output
the `SfrFile` class handles this the same way
files for the transient version of the above example were already copied to the `data` folder in the third cell above
first run the transient model to get the output:
```
>mf2005 test1tr.nam
```
```
flopy.run_model(exe_name, 'test1tr.nam', model_ws=path, silent=True)
sfrout_tr = SfrFile(os.path.join('data', 'test1tr.flw'))
dftr = sfrout_tr.get_dataframe()
dftr.head()
```
### plot a hydrograph
plot `Qout` (simulated streamflow) and `Qaquifer` (simulated stream leakage) through time
```
fig, axes = plt.subplots(2, 1, sharex=True)
dftr8 = dftr.loc[(dftr.segment == 8) & (dftr.reach == 5)]
dftr8.Qout.plot(ax=axes[0])
axes[0].set_ylabel('Simulated streamflow, cfs')
dftr8.Qaquifer.plot(ax=axes[1])
axes[1].set_ylabel('Leakage to aquifer, cfs');
```
| github_jupyter |
# Multi-GPU training with `tf.keras` or Estimators and `tf.data`
We can train on multiple GPUs directly via `tf.keras`'s distributed strategy scope.
TensorFlow's [Estimators](https://www.tensorflow.org/programmers_guide/estimators) API is another useful way to training models in a distributed environment such as on nodes with multiple GPUs or on many nodes with GPUs. This is particularly useful when training on huge datasets especially when used with the `tf.keras` API.
Here we will first present the `tf.keras` API for the tiny Fashion-MNIST dataset and then show a practical usecase in the end via Estimators.
**TL;DR**: Essentially what we want to remember is that a `tf.keras.Model` can be trained with `tf.estimator` API by converting it to an `tf.estimator.Estimator` object via the `tf.keras.estimator.model_to_estimator` method. Once converted we can apply the machinery that `Estimator` provides to train on different hardware configurations.
```
import os
import time
import tensorflow as tf
from tensorflow.python.ops import lookup_ops
import numpy as np
```
## Import the Fashion-MNIST dataset
We will use the [Fashion-MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset, a drop-in replacement of MNIST, which contains thousands of grayscale images of [Zalando](https://www.zalando.de/) fashion articles. Getting the training and test data is as simple as:
```
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.fashion_mnist.load_data()
```
We want to convert the pixel values of these images from a number between 0 and 255 to a number between 0 and 1 and convert the dataset to the `[B, H, W, C]` format where `B` is the number of images, `H` and `W` are the height and width and `C` the number of channels (1 for grayscale) of our images:
```
TRAINING_SIZE = len(train_images)
TEST_SIZE = len(test_images)
train_images = np.asarray(train_images, dtype=np.float32) / 255
# Convert the train images and add channels
train_images = train_images.reshape((TRAINING_SIZE, 28, 28, 1))
test_images = np.asarray(test_images, dtype=np.float32) / 255
# Convert the train images and add channels
test_images = test_images.reshape((TEST_SIZE, 28, 28, 1))
```
Next, we want to convert the labels from an integer format (e.g., `2` or `Pullover`), to a [one hot encoding](https://en.wikipedia.org/wiki/One-hot) (e.g., `0, 0, 1, 0, 0, 0, 0, 0, 0, 0`). To do so, we'll use the `tf.keras.utils.to_categorical` [function](https://www.tensorflow.org/api_docs/python/tf/keras/utils/to_categorical) function:
```
# How many categories we are predicting from (0-9)
LABEL_DIMENSIONS = 10
train_labels = tf.keras.utils.to_categorical(train_labels, LABEL_DIMENSIONS)
test_labels = tf.keras.utils.to_categorical(test_labels, LABEL_DIMENSIONS)
# Cast the labels to floats, needed later
train_labels = train_labels.astype(np.float32)
test_labels = test_labels.astype(np.float32)
```
## Distribution strategy
So how do we go about training a `tf.keras` model to use multi-GPUs? We can use the `tf.distribute.MirroredStrategy` paradigm which does in-graph replication with synchronous training. See this talk on [Distributed TensorFlow training](https://www.youtube.com/watch?v=bRMGoPqsn20) for more information about this strategy.
Essentially each worker GPU has a copy of the graph and gets a subset of the data on which it computes the local gradients and then waits for all the workers to finish in a synchronous manner. Then the workers communicate their local gradients to each other via a ring Allreduce operation which is typically optimized to reduce network bandwidth and increase through-put. Once all the gradients have arrived each worker averages them and updates its parameter and the next step begins. This is ideal in situations where you have multiple GPUs on a single node connected via some high-speed interconnect.
To create a `MirroredStrategy` just instantiate it via:
```
strategy = tf.distribute.MirroredStrategy()
```
## Build a `tf.keras` model
We will create our neural network using the [Keras Functional API](https://www.tensorflow.org/guide/keras#functional_api). Keras is a high-level API to build and train deep learning models and is user friendly, modular and easy to extend. `tf.keras` is TensorFlow's implementation of this API and it supports such things as [Eager Execution](https://www.tensorflow.org/guide/eager), `tf.data` pipelines and Estimators.
In terms of the architecture we will use ConvNets. On a very high level ConvNets are stacks of Convolutional layers (`Conv2D`) and Pooling layers (`MaxPooling2D`). But most importantly they will take for each training example a 3D tensors of shape (`height`, `width`, `channels`) where for the case of grayscale images `channels=1` and return a 3D tensor.
Therefore after the ConvNet part we will need to `Flatten` the tensor and add `Dense` layers, the last one returning the `LABEL_DIMENSIONS` outputs with the `softmax` activation.
To allow this model to train on multiple GPUs via the strategy we defined above, we need to create and compile the `tf.keras` model in our `strategy.scope`:
```
with strategy.scope():
inputs = tf.keras.Input(shape=(28,28,1)) # Returns a placeholder tensor
x = tf.keras.layers.Conv2D(filters=32, kernel_size=(3, 3), activation=tf.nn.relu)(inputs)
x = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=2)(x)
x = tf.keras.layers.Conv2D(filters=64, kernel_size=(3, 3), activation=tf.nn.relu)(x)
x = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=2)(x)
x = tf.keras.layers.Conv2D(filters=64, kernel_size=(3, 3), activation=tf.nn.relu)(x)
x = tf.keras.layers.Flatten()(x)
x = tf.keras.layers.Dense(64, activation=tf.nn.relu)(x)
predictions = tf.keras.layers.Dense(LABEL_DIMENSIONS, activation='sigmoid')(x)
model = tf.keras.Model(inputs=inputs, outputs=predictions)
model.compile(loss='categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
metrics=['accuracy'])
```
## Create an `tf.data` input function
Next we define a data importing function which returns a `tf.data` dataset of `(images, labels)` batches of our data. The function below takes in `numpy` arrays and returns the dataset via an ETL process.
Note that in the end we are also calling the `prefetch` method which will buffer the data to the GPUs while they are training so that the next batch is ready and waiting for the GPUs rather than having the GPUs wait for the data at each iteration. The GPU might still not be fully utilized and to improve this we can use fused versions of the transformation operations like `shuffle_and_repeat` instead of two separate operations, but I have kept the simple case here.
```
def input_fn(images, labels, epochs, batch_size):
# Convert the inputs to a Dataset. (E)
dataset = tf.data.Dataset.from_tensor_slices((images, labels))
# Shuffle, repeat, and batch the examples. (T)
SHUFFLE_SIZE = 5000
dataset = dataset.shuffle(SHUFFLE_SIZE).repeat(epochs).batch(batch_size)
dataset = dataset.prefetch(None)
# Return the dataset. (L)
return dataset
```
## Training
```
BATCH_SIZE = 512
EPOCHS = 10
steps_per_epoch = int(np.ceil(60000 / float(BATCH_SIZE)))
model.fit(input_fn(train_images, train_labels,
epochs=EPOCHS,
batch_size=BATCH_SIZE), epochs=EPOCHS, steps_per_epoch=steps_per_epoch)
```
## Create an Estimator
To create an Estimator from the compiled Keras model we call the `model_to_estimator` method. Note that the initial model state of the Keras model is preserved in the created Estimator.
So what's so good about Estimators? Well to start off with:
* you can run Estimator-based models on a local host or an a distributed multi-GPU environment without changing your model;
* Estimators simplify sharing implementations between model developers;
* Estimators build the graph for you, so a bit like Eager Execution, there is no explicit session.
```
inputs = tf.keras.Input(shape=(28,28,1)) # Returns a placeholder tensor
x = tf.keras.layers.Conv2D(filters=32, kernel_size=(3, 3), activation=tf.nn.relu)(inputs)
x = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=2)(x)
x = tf.keras.layers.Conv2D(filters=64, kernel_size=(3, 3), activation=tf.nn.relu)(x)
x = tf.keras.layers.MaxPooling2D(pool_size=(2, 2), strides=2)(x)
x = tf.keras.layers.Conv2D(filters=64, kernel_size=(3, 3), activation=tf.nn.relu)(x)
x = tf.keras.layers.Flatten()(x)
x = tf.keras.layers.Dense(64, activation=tf.nn.relu)(x)
predictions = tf.keras.layers.Dense(LABEL_DIMENSIONS, activation='sigmoid')(x)
model = tf.keras.Model(inputs=inputs, outputs=predictions)
model.compile(loss='categorical_crossentropy',
optimizer=tf.compat.v1.train.AdamOptimizer(learning_rate=0.001),
metrics=['accuracy'])
config = tf.estimator.RunConfig(train_distribute=strategy)
estimator = tf.keras.estimator.model_to_estimator(model, config=config)
```
## Train the Estimator
Lets first define a `SessionRunHook` class for recording the times of each iteration of stochastic gradient descent:
```
class TimeHistory(tf.estimator.SessionRunHook):
def begin(self):
self.times = []
def before_run(self, run_context):
self.iter_time_start = time.time()
def after_run(self, run_context, run_values):
self.times.append(time.time() - self.iter_time_start)
```
Now the good part! We can call the `train` function on our Estimator giving it the `input_fn` we defined (with the batch size and the number of epochs we wish to train for) and a `TimeHistory` instance via it's `hooks` argument:
```
BATCH_SIZE = 512
EPOCHS = 5
time_hist = TimeHistory()
estimator.train(input_fn=lambda:input_fn(train_images,
train_labels,
epochs=EPOCHS,
batch_size=BATCH_SIZE),
hooks=[time_hist])
```
## Performance
Since we have our timing hook we can now use it to calculate the total time of training as well as the number of images we train on per second:
```
NUM_GPUS = 2
total_time = sum(time_hist.times)
print(f"total time with {NUM_GPUS} GPUs: {total_time} seconds")
avg_time_per_batch = np.mean(time_hist.times)
print(f"{BATCH_SIZE*NUM_GPUS/avg_time_per_batch} images/second with {NUM_GPUS} GPUs")
```
## Evaluate the Estimator
In order to check the performance of our model we then call the `evaluate` method on our Estimator:
```
estimator.evaluate(lambda:input_fn(test_images,
test_labels,
epochs=1,
batch_size=BATCH_SIZE))
```
## Retinal OCT (optical coherence tomography) images example
To test the scaling performance on some bigger dataset we can use the [Retinal OCT images](https://www.kaggle.com/paultimothymooney/kermany2018) dataset, on of the many great datasets from [Kaggle](https://www.kaggle.com/datasets). This dataset consists of cross sections of the retinas of living patients grouped into four categories: NORMAL, CNV, DME and DRUSEN.

The dataset has a total of 84,495 X-Ray JPEG images, typically `512x496`, and can be downloaded via the `kaggle` CLI:
```
#!pip install kaggle
#!kaggle datasets download -d paultimothymooney/kermany2018
```
Once downloaded the training and test set image classes are in their own respective folder so we can define a pattern as:
```
train_folder = os.path.join('OCT2017', 'train', '**', '*.jpeg')
test_folder = os.path.join('OCT2017', 'test', '**', '*.jpeg')
labels = ['CNV', 'DME', 'DRUSEN', 'NORMAL']
```
Next we have our Estimator's input function which takes any file pattern and returns resized images and one hot encoded labels as a `tf.data.Dataset`. Here we follow the best practices from the [Input Pipeline Performance Guide](https://www.tensorflow.org/performance/datasets_performance). Note in particular that if the `prefetch_buffer_size` is `None` then TensorFlow will use an optimal prefetch buffer size automatically:
```
def input_fn(file_pattern, labels,
image_size=(224,224),
shuffle=False,
batch_size=64,
num_epochs=None,
buffer_size=4096,
prefetch_buffer_size=None):
table = lookup_ops.index_table_from_tensor(tf.constant(labels))
num_classes = len(labels)
def _map_func(filename):
label = tf.string_split([filename], delimiter=os.sep).values[-2]
image = tf.image.decode_jpeg(tf.io.read_file(filename), channels=3)
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
image = tf.image.resize(image, size=image_size)
return (image, tf.one_hot(table.lookup(label), num_classes))
dataset = tf.data.Dataset.list_files(file_pattern, shuffle=shuffle)
if num_epochs is not None and shuffle:
dataset = dataset.apply(tf.data.experimental.shuffle_and_repeat(buffer_size, num_epochs))
elif shuffle:
dataset = dataset.shuffle(buffer_size)
elif num_epochs is not None:
dataset = dataset.repeat(num_epochs)
dataset = dataset.apply(
tf.data.experimental.map_and_batch(map_func=_map_func,
batch_size=batch_size,
num_parallel_calls=os.cpu_count()))
dataset = dataset.prefetch(buffer_size=prefetch_buffer_size)
return dataset
```
In order to train this we will use a pretrained VGG16 and train just it's last 5 layers:
```
with strategy.scope():
keras_vgg16 = tf.keras.applications.VGG16(input_shape=(224,224,3),
include_top=False)
output = keras_vgg16.output
output = tf.keras.layers.Flatten()(output)
predictions = tf.keras.layers.Dense(len(labels), activation=tf.nn.softmax)(output)
model = tf.keras.Model(inputs=keras_vgg16.input, outputs=predictions)
for layer in keras_vgg16.layers[:-4]:
layer.trainable = False
model.compile(loss='categorical_crossentropy',
optimizer=tf.compat.v1.train.AdamOptimizer(learning_rate=0.001),
metrics=['accuracy'])
```
Now we have all we need and can proceed as above and train our model in a few minutes using `NUM_GPUS` GPUs:
```
config = tf.estimator.RunConfig(train_distribute=strategy)
estimator = tf.keras.estimator.model_to_estimator(model, config=config)
BATCH_SIZE = 32
EPOCHS = 2
time_hist = TimeHistory()
estimator.train(input_fn=lambda:input_fn(train_folder,
labels,
shuffle=True,
batch_size=BATCH_SIZE,
buffer_size=2048,
num_epochs=EPOCHS,
prefetch_buffer_size=4),
hooks=[time_hist])
NUM_GPUS = 2
total_time = sum(time_hist.times)
print(f"total time with {NUM_GPUS} GPUs: {total_time} seconds")
avg_time_per_batch = np.mean(time_hist.times)
print(f"{BATCH_SIZE*NUM_GPUS/avg_time_per_batch} images/second with {NUM_GPUS} GPUs")
```
Once trained we can evaluate the accuracy on the test set, which should be around 95% (not bad for an initial baseline!):
```
estimator.evaluate(input_fn=lambda:input_fn(test_folder,
labels,
shuffle=True,
batch_size=BATCH_SIZE,
buffer_size=2048,
num_epochs=1))
```
| github_jupyter |
#### Metode i primjena vjestacke inteligencije
#### Laboratorijska vjezba 5
#### Student: Masovic Haris
#### Index: 1689/17993
## 0. Dependencies
```
import sys
!{sys.executable} -m pip install matplotlib
```
## 2. Dataset: Reuters
### 2.1 Ucitavanje dataset-a
```
from keras.datasets import reuters
(train_data, train_labels), (test_data, test_labels) = reuters.load_data(num_words=10000)
```
#### 2.1.1 Koliko novosti ima u `train_data`, a koliko u `test_data`?
```
print(*['Broj novosti u train_data', len(train_data)])
print(*['Broj novosti u test_data', len(test_data)])
```
#### 2.1.2 Izlistajte vrijednosti iz `train_data[0]`.
```
print(train_data[0])
```
#### 2.1.3 Za prvih 15 sekvenci isprintati: a) broj elemenata u sekvenci, b) kojoj klasi pripada, c) maksimalan i d) minimalan indeks u sekvenci. U izvjestaju napisati kod koji ste koristili za ovaj zadatak, kao i rezultate. Primjer rezultata dat je u ispisu ispod. (Koristiti for petlju.)
```
import numpy as np
for i in range(0, 15):
print('Broj elemenata u sekvenci train_data[{0}] je: ({1},), a klasa je: {2}'.format(i, len(train_data[i]), train_labels[i]))
print(*['Najveci index u sekvenci je:', np.max(train_data[i])])
print(*['Najmanji index u sekvenci je:', np.min(train_data[i])])
```
#### 2.1.4 Koji opseg vrijednosti se nalazi u `train_labels` (min, max), a koji u `test_labels`?
```
import numpy as np
print('Opseg vrijednosti za trazene labele:')
print('')
print(*['train_labels opseg (min,max):', '({0},{1})'.format(np.min(train_labels), np.max(train_labels))])
print(*['test_labels opseg (min,max):', '({0},{1})'.format(np.min(test_labels), np.max(test_labels))])
```
#### 2.1.5 Koja sekvenca ima najveci indeks u datom dataset-u? To ne mora biti broj vezan za num words. Korisiti numpy paket (import numpy as np).
```
import numpy as np
print(*['Najveci index unutar liste lista:', np.max(np.max(train_data))])
```
#### 2.1.6 Na stanici `https://keras.io/datasets/` mozete naci sve parametre koje mozete korisiti u pozivu funkcije `load_data()`. Objasnite svaki od parametara (9).
- `path` - putanja gdje ce se kesirati podaci (u odnosu na relativnu putanju `~/.keras/dataset`).
- `num_words` - predstavlja broj najcesce koristenih rijeci koje ce biti uzete u razmatranje, a ako se ne posalje broj, defaultna vrijednost je da uzme sve rijeci.
- `skip_top` - parametar koji govori koliko top N najvise frekventnih rijeci da se preskoci u uzimanju, default je nula.
- `maxlen` - predstavlja maximalnu duzinu sekvence koja da se koristi, ako sekvenca postoji bice skracena, ukoliko se ne posalje nista, nema skracivanja.
- `test_split` - decimalni broj izmedju 0 i 1 koji govori kolika frakcija da se uzme za testne podatke, po defaultu je 20-80 ratio tj. 20% da se uzimaju kao testni podaci.
- `seed` - parametar koji se koristi za mijesanje ponavljajucih podataka.
- `start_char` - parametar koji odredjuje karakter kojim je definisan pocetak sekvence.
- `oov_char` - parametar kojim odredjuje karakter kojim ce biti zamijenjene rijeci koje su izbacene na osnovu `num_words` ili `skip_top` opcija.
- `index_from` - parametar koji odredjuje rijeci kojecće se ideksirati, tj. rijeci vece ili jednake indeksu `index_from` ce biti indeksirane.
### 2.2 Rjecnik Reuters dataset-a
```
word_index = reuters.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[0]])
print(decoded_review)
```
#### 2.2.1 Za prvih 15 sekvenci u `train_data` dekodirati novosti i odrediti kojoj kategoriji pripadaju.
```
def return_formatted_review(string, width=100):
if len(string) > width:
string = string[:width-3] + '...'
return "{0:<{1}s}".format(string, width)
for i in range(0, 15):
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[i]])
print(*['Novost', i, 'je u kategoriji:', train_labels[i]])
print(return_formatted_review(decoded_review))
```
#### 2.2.2 Za 10 nasumicnih (random) novosti iz citavog dataset-a izvrsiti dekodiranje, te ih isprintati na ekran, po uzoru na kod dat ispod, zajedno sa informacijom o grupi kojoj pripadaju. U ispisu su koristene "..." radi zauzimanja manjeg prostora. Koristiti naredbu randint(0, nekibroj), pri cemu je nekibroj maksimalan broj sekvenci u skupu `train_data`.
```
import numpy as np
random_numbers = np.random.randint(0, len(train_data), size = 10)
# u slucaju da ne zelite skracenu varijantu stringa, samo ispisati `decoded_review`
for i in range(0, len(random_numbers)):
decoded_review = ' '.join([reverse_word_index.get(i - 3, '?') for i in train_data[i]])
print('Novost ID {0} je definisana kao: \"{1}\". Rijec je o novosti iz grupe {2}.'
.format(random_numbers[i], return_formatted_review(decoded_review, 40), train_labels[random_numbers[i]]))
```
### 2.3 Organizacija podataka
```
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
def to_one_hot(labels, dimension=46):
results = np.zeros((len(labels), dimension))
for i, label in enumerate(labels):
results[i, label] = 1.
return results
```
#### 2.3.1 Detaljno objasnite kako navedeni kodovi rade i da li postoje razlike medu njima.
- Funkcija `vectorize_sequences` definise sekvencu od `dimension` broja elemenata, koji imaju vrijednost ili `1.` ili `0.`. Ukoliko je vrijednost `1.` onda se indeks nalazi u odredjenoj sekveni koja je proslijedjena kroz prvi parametar, u protivnom se ne nalazi.
- Prvo se u varijablu `results` definisu sve nule kroz uredjeni par `(len(sequences), dimension)`, onda prolaskom kroz sve sekvence, dodijeljuje se za svaku sekvencu enumeracija i postavlja se vrijednost na `1.`. Na kraju se vraca matrica `results`.
- Kada je u pitanju funkcija `to_one_hot` mozemo primijeniti isti opis i za nju, samo umjesto `sequences` koristimo varijablu `labels`.
- Razlike izmedju ove 2 funkcije nema osim defaultnog `dimension` parametra.
#### 2.3.2 Primijenite funkciju `vectorize_sequences()` na podatke iz skupa za treniranje i testiranje. Rezultate funkcije snimite u varijable `x_train` i `x_test`. Ovdje je dovoljno napisati koristeni kod.
```
data_size = 10000
x_train = vectorize_sequences(train_data, data_size)
x_test = vectorize_sequences(test_data, data_size)
y_train = np.asarray(train_labels).astype('float32')
y_test = np.asarray(test_labels).astype('float32')
```
#### 2.3.3 Utvrdite koja vrsta elemenata je snimljena u `x_train`, a koja u `y_train` (vektor, skalar, int, float i td). Napisite naredbe koje ste koristili.
- Koristene naredbe su naredba `type` i naredna `.dtype`.
- Vidimo, na osnovu koda ispod, da su i `x_train` i `y_train` tipa numpy array `numpy.ndarray`, a elementi unutar array-a su `float64` u slucaju `x_train`, odnosno `float32` u slucaju `float32`.
```
print(*['x_train', type(x_train), 'tip elemenata', x_train.dtype])
print(*['y_train', type(y_train), 'tip elemenata', y_train.dtype])
```
#### 2.3.4 Za prvih 15 sekvenci isprintati: a) broj elemenata u sekvenci, b) vrijednosti elemenata (skraceni broj elemenata je dovoljan).
```
for i in range(0, 15):
print(*['Za x_train[{0}], broj elemenata u sekvenci je:'.format(i), len(x_train[i])])
print(*['Elementi u ovoj sekvenci:', x_train[i]])
print(*['y_train, broj elemenata:', len(y_train), 'elementi:', y_train])
```
#### 2.3.5 Primijenite funkciju `to_one_hot()` na podatke iz labela za treniranje i testiranje. Rezultate funkcije snimite u varijable `one_hot_train_labels` i `one_hot_test_labels`.
```
dimension_size = 46
one_hot_train_labels = to_one_hot(train_labels, dimension_size)
one_hot_test_labels = to_one_hot(test_labels, dimension_size)
```
#### 2.3.6 Za 15 nasumicnih (random) sekvenci isprintati: a) staru labelu, b) zatim provjeriti da li je na tom indeksu zapisana ‘1’, kao i to da li je u dva okolna elementa u vektora napisana ‘0’.
```
import numpy as np
def random_15_elements(new_labels, old_labels):
random_numbers = np.random.randint(0, len(train_data), size = 15)
for i in range(0, len(random_numbers)):
index = random_numbers[i]
class_value = int(old_labels[index])
array_for_index = new_labels[index]
binary_value = array_for_index[class_value]
print(*['Za novost id:', index, 'stara labela je:', old_labels[index]])
print(*['Nova labela je:', binary_value, 'i ona je jednaka 1. za novost id:', binary_value == 1.])
length = len(array_for_index)
if class_value != 0:
index_before = array_for_index[class_value - 1]
print('Element prije je jednak 0.:', index_before == 0.)
if class_value != length - 1:
index_after = array_for_index[class_value + 1]
print('Element poslije je jednak 0.:', index_after == 0.)
print('')
random_15_elements(one_hot_train_labels, y_train)
```
#### 2.3.7 Zamijeniti funkciju to one hot sa funkcijom `to_categorical` u uraditi opet prethodni zadatak.
```
from keras.utils import to_categorical
to_categorical_train_labels = to_categorical(train_labels)
to_categorical_test_labels = to_categorical(test_labels)
random_15_elements(to_categorical_train_labels, y_train)
y_train = to_categorical_train_labels
y_test = to_categorical_test_labels
```
## 3. Definisanje neuronske mreze
### 3.1 Podesavanje slojeva
#### 3.1.1 Ako ulazni vektor ima 10,000 elemenata, i takvih vektora ima 200, a sloj ima 64 izlaza, koja je dimenzija matrice W1? Koristiti se informacijama sa predavanja.
- Dimenzija matrice W druge ce biti `10000x64` odnosno `640k`.
### 3.2 Arhitektura mreze
#### 3.2.1 Napisati kod za definisanje arhitekture mreze.
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
```
#### 3.2.2 Napisati kod za definisanje optimizatora, funkcije gubitka i mjere tacnosti.
```
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
```
#### 3.2.3 Objasnite kako radi `categorical_crossentropy` i zasto se koristi za ovaj primjer. (Koristi Keras dokumentaciju za pronalazenje odgovora na ovo pitanje ili preporucenu knjigu.)
`categorical_crossentropy` racuna funkciju gubitka na sljedeci nacin:
\begin{equation*}
Gubitak = \sum_{i=1}^k{y_i * log(y_i)}
\end{equation*}
- `y` predstavlja niz skalara, dok `k` predstavlja koliko izlaza imamo za nas problem.
- Ovakav nacin racunanja gubitka omogucava dobru mjeru koliko su dvije diskretne distribucije vjerovatnoce razlicite za dva elementa, sto znaci da u slucaju da specifican `i` unutar `y` niza je jednak 1, to znaci da samo taj skalar unutar niza se "desava" odnosno ima najvecu vrijednost.
- U nasem primjeru ovo je veoma pogodno jer imamo `46` klasifikacija, pri cemu nam je na izlazu potreban samo jedan tip od tih `46`.
#### 3.2.4 Objasnite zasto koristimo softmax. Objasnite dimenzionalnost izlaznog nivoa.
- Posto je u pitanju `multiclass` problem, tj. 1 entry moze imati `46` varijanti izlaza, da bi se na odredjeni nacin znalo koji ulaz pripada kojem izlazu, to se moze rijesiti vjerovatnocom. Softmax daje na svom izlazu vjerovatnoce da odredjeni entry pripada odredjenim klasama, a suma mora biti jednaka `1`. Zbog ovog se koristi softmax.
- Dimenzionalnost izlaznog nivoa je `46`, jer imamo toliko klasifikacija.
### 3.3 Validacija modela
#### 3.3.1 Koje naredbe cete koristiti za izdvajanje 1,000 uzoraka iz originalnog skupa za treniranje?
```
x_val = x_train[:1000]
partial_x_train = x_train[1000:]
```
#### 3.3.2 Koje naredbe cete koristiti za kodiranje labela za novonastale podatke?
```
y_val = y_train[:1000]
partial_y_train = y_train[1000:]
```
### 3.4 Treniranje modela
```
reuters_model = model.fit(partial_x_train, partial_y_train, epochs=20, batch_size=512, validation_data=(x_val, y_val))
```
#### 3.4.1 Nacrtajte grafikone na kojima ce biti prikazani podaci o gubitku kod treniranja i validacije u odnosu na broj epoha.
```
import matplotlib.pyplot as plt
history_dict = reuters_model.history
print(*['Dict keys:', history_dict.keys()])
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc = history_dict['accuracy']
val_acc_values = history_dict['val_accuracy']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, loss_values, 'bo', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
```
#### 3.4.2 Nacrtajte grafikone na kojima ce biti prikazani podaci o tacnosti kod treniranja i validacije u odnosu na broj epoha.
```
plt.clf()
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc_values, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
#### 3.4.3 Nakon koje epohe model ulazi u overfitting?
- Nakon 7 epohe model ulazi u overfitting.
#### 3.4.4 Ponovo pokrenuti treniranje mreze u odnosu na prethodni odgovor i ponovo izvrsiti mjerenje.
```
reuters_model_7_epoch = model.fit(partial_x_train, partial_y_train, epochs=7, batch_size=512, validation_data=(x_val, y_val))
```
#### 3.4.5 Zatim izvrsiti mjerenja na testnom skupu koristeci funkciju `model.evaluate()`. Objasnite razliku izmedu funkcije `model.evaluate()` i `model.predict()`.
- Razlika izmedju `model.evaluate` i `model.predict` je sto `model.evaluate` testira vrijednosti i racuna funkciju gubitka za sve klase koje su trazene, i vraca listu gubitaka i metrika u jednoj vrijendosti. `model.predict` samo predvidja odnosno testira izlaz modela u zavisnosti od ulaznih podataka. Izlazne vrijednosti ove dvije funkcije su razlicite jer ne racunaju iste stvari.
```
val_loss, val_acc = model.evaluate(x_val, y_val)
print(*['Validacija - tacnost: ', val_acc, 'a gubitak:', val_loss])
```
#### 3.4.6 Ponovo izmjeriti tacnost i funkciju gubitka i uporediti sa prethodnim mjerenjima.
- Poredeci sa prethodnim graficima, vidimo da najbolju tacnost i najmanji gubitak ostvarujemo pri kraju `7` epohe.
```
import matplotlib.pyplot as plt
history_dict = reuters_model_7_epoch.history
print(*['Dict keys:', history_dict.keys()])
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
acc = history_dict['accuracy']
val_acc_values = history_dict['val_accuracy']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, loss_values, 'bo', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.clf()
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc_values, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
```
### 3.5 Testiranje modela na testnim podacima
```
predictions = model.predict(x_test)
```
#### 3.5.1 Koja je dimenzionalnost jedne predikcije (bilo koje)?
- Dimenzionalnost je 46.
#### 3.5.2 Koja je suma vrijednosti iz jedne predikcije (bilo koje)? Da li je to slucaj i sa ostalim predikcijama? Zasto je to tako?
- Suma vrijednosti bilo koje predikcije je 1.
- To je slucaj sa svim predikcijama.
- Zato sto softmax na svom izlazu za svaku klasu daje 46 vrijednosti koje sumarno predstavljaju jedinicu, ali za svaku predstavljaju pripadnost te predikcije za tu klasu.
#### 3.5.3 Za prvih 10 predikcija iz skupa `x_test` naci indeks sa najvecoj vjerovatnocom (korisiti numpy i argmax). Da li je to u skladu sa dostupnim oznakama (labelama)?
```
import numpy as np
prvih_deset = predictions[:10]
for i, item in enumerate(prvih_deset):
max_index = np.argmax(item)
predicted_label = np.argmax(predictions[max_index])
real_label = np.argmax(y_test[max_index])
decoded_max = ' '.join([reverse_word_index.get(i - 3, '?') for i in test_data[max_index]])
print(*['Novost ID:', i, 'stvarna labela:', real_label, 'predicted labela:', predicted_label])
print(return_formatted_review(decoded_max, 100))
print('')
```
#### 3.5.4 Koliko novosti ima maksimalnu vjerovatnocu vecu od 0.5?
```
import numpy as np
print(*['Broj novosti sa vjerovatnocom vecom od 0.5:', np.count_nonzero(predictions > 0.5)])
```
#### 3.5.5 Koja je tacnost nad testnim skupom podataka, a koliki je gubitak?
```
test_loss, test_acc = model.evaluate(x_test, y_test)
print(*['Test - tacnost: ', test_acc, 'a gubitak:', test_loss])
```
## 4. Dalja eksperimentiranja
```
print(*['Validacija - tacnost: ', val_acc, 'a gubitak:', val_loss])
print(*['Test - tacnost: ', test_acc, 'a gubitak:', test_loss])
```
#### 4.0.1 Modifikovati postojeci model mreze na nacin da u drugom sloju koristite 4 skrivene procesne jedinice. Koja je tacnost ovog modela i gubitak? Uporedite sa originalnim modelom.
- Poredeci sa orginalnim modelom, tacnost je smanjena, a gubitak povecan bilo validacijski dataset ili testni dataset. Kod je dat ispod.
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(4, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
reuters_model = model.fit(partial_x_train, partial_y_train, epochs=7, batch_size=512, validation_data=(x_val, y_val))
val_loss, val_acc = model.evaluate(x_val, y_val)
test_loss, test_acc = model.evaluate(x_test, y_test)
print('')
print(*['Validacija - tacnost: ', val_acc, 'a gubitak:', val_loss])
print(*['Test - tacnost: ', test_acc, 'a gubitak:', test_loss])
```
#### 4.0.2 U modelu ste koristili dva skrivena sloja. Pokusajte koristiti jedan ili tri skrivena sloja i utvrdite kako to utice na tacnost kod validacijskog i testnog skupa.
- Koristena su tri skrivena sloja, tacnost je povecana, a gubitak je smanjen i za validacijski i za testni dataset. Kod je dat ispod.
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
reuters_model = model.fit(partial_x_train, partial_y_train, epochs=7, batch_size=512, validation_data=(x_val, y_val))
val_loss, val_acc = model.evaluate(x_val, y_val)
test_loss, test_acc = model.evaluate(x_test, y_test)
print('')
print(*['Validacija - tacnost: ', val_acc, 'a gubitak:', val_loss])
print(*['Test - tacnost: ', test_acc, 'a gubitak:', test_loss])
```
#### 4.0.3 Pokusajte koristiti slojeve s vise skrivenih jedinica ili manje skrivenih jedinica, na primjer 32, 128 i 256 jedinice. Utvrdite kako to utice na tacnost kod validacijskog i testnog skupa.
- Povecan je broj skrivenih jedinica u drugom sloju, tacnost je povecana, a gubitak je smanjen i za validacijski i za testni dataset. Kod je dat ispod.
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
reuters_model = model.fit(partial_x_train, partial_y_train, epochs=7, batch_size=512, validation_data=(x_val, y_val))
val_loss, val_acc = model.evaluate(x_val, y_val)
test_loss, test_acc = model.evaluate(x_test, y_test)
print('')
print(*['Validacija - tacnost: ', val_acc, 'a gubitak:', val_loss])
print(*['Test - tacnost: ', test_acc, 'a gubitak:', test_loss])
```
| github_jupyter |
# Natural language inference: Task and datasets
```
__author__ = "Christopher Potts"
__version__ = "CS224u, Stanford, Spring 2019"
```
## Contents
1. [Overview](#Overview)
1. [Our version of the task](#Our-version-of-the-task)
1. [Primary resources](#Primary-resources)
1. [NLI model landscape](#NLI-model-landscape)
1. [Set-up](#Set-up)
1. [Properties of the corpora](#Properties-of-the-corpora)
1. [SNLI properties](#SNLI-properties)
1. [MultiNLI properties](#MultiNLI-properties)
1. [Working with SNLI and MultiNLI](#Working-with-SNLI-and-MultiNLI)
1. [Readers](#Readers)
1. [The NLIExample class](#The-NLIExample-class)
1. [Labels](#Labels)
1. [Tree representations](#Tree-representations)
1. [Annotated MultiNLI subsets](#Annotated-MultiNLI-subsets)
1. [Other NLI datasets](#Other-NLI-datasets)
## Overview
Natural Language Inference (NLI) is the task of predicting the logical relationships between words, phrases, sentences, (paragraphs, documents, ...). Such relationships are crucial for all kinds of reasoning in natural language: arguing, debating, problem solving, summarization, and so forth.
[Dagan et al. (2006)](https://link.springer.com/chapter/10.1007%2F11736790_9), one of the foundational papers on NLI (also called Recognizing Textual Entailment; RTE), make a case for the generality of this task in NLU:
> It seems that major inferences, as needed by multiple applications, can indeed be cast in terms of textual entailment. For example, __a QA system__ has to identify texts that entail a hypothesized answer. [...] Similarly, for certain __Information Retrieval__ queries the combination of semantic concepts and relations denoted by the query should be entailed from relevant retrieved documents. [...] In __multi-document summarization__ a redundant sentence, to be omitted from the summary, should be entailed from other sentences in the summary. And in __MT evaluation__ a correct translation should be semantically equivalent to the gold standard translation, and thus both translations should entail each other. Consequently, we hypothesize that textual entailment recognition is a suitable generic task for evaluating and comparing applied semantic inference models. Eventually, such efforts can promote the development of entailment recognition "engines" which may provide useful generic modules across applications.
## Our version of the task
Our NLI data will look like this:
| Premise | Relation | Hypothesis |
|---------|---------------|------------|
| turtle | contradiction | linguist |
| A turtled danced | entails | A turtle moved |
| Every reptile danced | entails | Every turtle moved |
| Some turtles walk | contradicts | No turtles move |
| James Byron Dean refused to move without blue jeans | entails | James Dean didn't dance without pants |
In the [word-entailment bakeoff](nli_wordentail_bakeoff.ipynb), we looked at a special case of this where the premise and hypothesis are single words. This notebook begins to introduce the problem of NLI more fully.
## Primary resources
We're going to focus on two large, human-labeled, relatively naturalistic entailment corpora:
* [The Stanford Natural Language Inference corpus (SNLI)](https://nlp.stanford.edu/projects/snli/)
* [The Multi-Genre NLI Corpus (MultiNLI)](https://www.nyu.edu/projects/bowman/multinli/)
The first was collected by a group at Stanford, led by [Sam Bowman](https://www.nyu.edu/projects/bowman/), and the second was collected by a group at NYU, also led by [Sam Bowman](https://www.nyu.edu/projects/bowman/). They have the same format and were crowdsourced using the same basic methods. However, SNLI is entirely focused on image captions, whereas MultiNLI includes a greater range of contexts.
This notebook presents tools for working with these corpora. The [second notebook in the unit](nli_02_models.ipynb) concerns models of NLI.
## NLI model landscape
<img src="fig/nli-model-landscape.png" width=800 />
## Set-up
* As usual, you need to be fully set up to work with [the CS224u repository](https://github.com/cgpotts/cs224u/).
* If you haven't already, download [the course data](http://web.stanford.edu/class/cs224u/data/data.zip), unpack it, and place it in the directory containing the course repository – the same directory as this notebook. (If you want to put it somewhere else, change `DATA_HOME` below.)
```
import nli
import os
import pandas as pd
import random
DATA_HOME = os.path.join("data", "nlidata")
SNLI_HOME = os.path.join(DATA_HOME, "snli_1.0")
MULTINLI_HOME = os.path.join(DATA_HOME, "multinli_1.0")
ANNOTATIONS_HOME = os.path.join(DATA_HOME, "multinli_1.0_annotations")
```
## Properties of the corpora
For both SNLI and MultiNLI, MTurk annotators were presented with premise sentences and asked to produce new sentences that entailed, contradicted, or were neutral with respect to the premise. A subset of the examples were then validated by an additional four MTurk annotators.
### SNLI properties
* All the premises are captions from the [Flickr30K corpus](http://shannon.cs.illinois.edu/DenotationGraph/).
* Some of the sentences rather depressingly reflect stereotypes ([Rudinger et al. 2017](https://aclanthology.coli.uni-saarland.de/papers/W17-1609/w17-1609)).
* 550,152 train examples; 10K dev; 10K test
* Mean length in tokens:
* Premise: 14.1
* Hypothesis: 8.3
* Clause-types
* Premise S-rooted: 74%
* Hypothesis S-rooted: 88.9%
* Vocab size: 37,026
* 56,951 examples validated by four additional annotators
* 58.3% examples with unanimous gold label
* 91.2% of gold labels match the author's label
* 0.70 overall Fleiss kappa
* Top scores currently around 89%.
### MultiNLI properties
* Train premises drawn from five genres:
1. Fiction: works from 1912–2010 spanning many genres
1. Government: reports, letters, speeches, etc., from government websites
1. The _Slate_ website
1. Telephone: the Switchboard corpus
1. Travel: Berlitz travel guides
* Additional genres just for dev and test (the __mismatched__ condition):
1. The 9/11 report
1. Face-to-face: The Charlotte Narrative and Conversation Collection
1. Fundraising letters
1. Non-fiction from Oxford University Press
1. _Verbatim_ articles about linguistics
* 392,702 train examples; 20K dev; 20K test
* 19,647 examples validated by four additional annotators
* 58.2% examples with unanimous gold label
* 92.6% of gold labels match the author's label
* Test-set labels available as a Kaggle competition.
* Top matched scores currently around 0.81.
* Top mismatched scores currently around 0.83.
## Working with SNLI and MultiNLI
### Readers
The following readers should make it easy to work with these corpora:
* `nli.SNLITrainReader`
* `nli.SNLIDevReader`
* `nli.MultiNLITrainReader`
* `nli.MultiNLIMatchedDevReader`
* `nli.MultiNLIMismatchedDevReader`
The base class is `nli.NLIReader`, which should be easy to use to define additional readers.
If you did change `data_home`, `snli_home`, or `multinli_home` above, then you'll need to call these readers with `dirname` as an argument, where `dirname` is your `snli_home` or `multinli_home`, as appropriate.
Because the datasets are so large, it is often useful to be able to randomly sample from them. All of the reader classes allow this with their keyword argument `samp_percentage`. For example, the following samples approximately 10% of the examples from the SNLI training set:
```
nli.SNLITrainReader(SNLI_HOME, samp_percentage=0.10)
```
The precise number of examples will vary somewhat because of the way the sampling is done. (Here, we trade efficiency for precision in the number of cases we return; see the implementation for details.)
### The NLIExample class
All of the readers have a `read` method that yields `NLIExample` example instances, which have the following attributes:
* __annotator_labels__: `list of str`
* __captionID__: `str`
* __gold_label__: `str`
* __pairID__: `str`
* __sentence1__: `str`
* __sentence1_binary_parse__: `nltk.tree.Tree`
* __sentence1_parse__: `nltk.tree.Tree`
* __sentence2__: `str`
* __sentence2_binary_parse__: `nltk.tree.Tree`
* __sentence2_parse__: `nltk.tree.Tree`
```
snli_iterator = iter(nli.SNLITrainReader(SNLI_HOME).read())
snli_ex = next(snli_iterator)
print(snli_ex)
snli_ex
```
### Labels
```
snli_labels = pd.Series(
[ex.gold_label for ex in nli.SNLITrainReader(SNLI_HOME, filter_unlabeled=False).read()])
snli_labels.value_counts()
multinli_labels = pd.Series(
[ex.gold_label for ex in nli.MultiNLITrainReader(MULTINLI_HOME, filter_unlabeled=False).read()])
multinli_labels.value_counts()
```
### Tree representations
Both corpora contain __three versions__ of the premise and hypothesis sentences:
1. Regular string representations of the data
1. Unlabeled binary parses
1. Labeled parses
```
snli_ex.sentence1
```
The binary parses lack node labels; so that we can use `nltk.tree.Tree` with them, the label `X` is added to all of them:
```
snli_ex.sentence1_binary_parse
```
Here's the full parse tree with syntactic categories:
```
snli_ex.sentence1_parse
```
The leaves of either tree are a tokenized version of the example:
```
snli_ex.sentence1_parse.leaves()
```
## Annotated MultiNLI subsets
MultiNLI includes additional annotations for a subset of the dev examples. The goal is to help people understand how well their models are doing on crucial NLI-related linguistic phenomena.
```
matched_ann_filename = os.path.join(
ANNOTATIONS_HOME,
"multinli_1.0_matched_annotations.txt")
mismatched_ann_filename = os.path.join(
ANNOTATIONS_HOME,
"multinli_1.0_mismatched_annotations.txt")
def view_random_example(annotations):
ann_ex = random.choice(list(annotations.items()))
pairid, ann_ex = ann_ex
ex = ann_ex['example']
print("pairID: {}".format(pairid))
print(ann_ex['annotations'])
print(ex.sentence1)
print(ex.gold_label)
print(ex.sentence2)
matched_ann = nli.read_annotated_subset(matched_ann_filename, MULTINLI_HOME)
view_random_example(matched_ann)
```
## Other NLI datasets
* [The FraCaS textual inference test suite](http://www-nlp.stanford.edu/~wcmac/downloads/) is a smaller, hand-built dataset that is great for evaluating a model's ability to handle complex logical patterns.
* [SemEval 2013](https://www.cs.york.ac.uk/semeval-2013/) had a wide range of interesting data sets for NLI and related tasks.
* [The SemEval 2014 semantic relatedness shared task](http://alt.qcri.org/semeval2014/task1/) used an NLI dataset called [Sentences Involving Compositional Knowledge (SICK)](http://alt.qcri.org/semeval2014/task1/index.php?id=data-and-tools). `data/nlidata` contains a parsed version of SICK created by [Sam Bowman](https://www.nyu.edu/projects/bowman/).
* [MedNLI](https://physionet.org/physiotools/mimic-code/mednli/) is specialized to the medical domain, using data derived from [MIMIC III](https://mimic.physionet.org).
* [XNLI](https://github.com/facebookresearch/XNLI) is a multilingual NLI dataset derived from MultiNLI.
* [Diverse Natural Language Inference Collection (DNC)](http://decomp.io/projects/diverse-natural-language-inference/) transforms existing annotations from other tasks into NLI problems for a diverse range of reasoning challenges.
* [SciTail](http://data.allenai.org/scitail/) is an NLI dataset derived from multiple-choice science exam questions and Web text.
* Models for NLI might be adapted for use with [the 30M Factoid Question-Answer Corpus](http://agarciaduran.org/).
* Models for NLI might be adapted for use with [the Penn Paraphrase Database](http://paraphrase.org/).
| github_jupyter |
```
# !wget https://storage.googleapis.com/bert_models/2018_11_23/multi_cased_L-12_H-768_A-12.zip
# !unzip multi_cased_L-12_H-768_A-12.zip
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from bert import modeling
import numpy as np
import tensorflow as tf
import pandas as pd
from tqdm import tqdm
import json
with open('dataset.json') as fopen:
data = json.load(fopen)
train_X = data['train_X']
train_Y = data['train_Y']
test_X = data['test_X']
test_Y = data['test_Y']
BERT_VOCAB = 'multi_cased_L-12_H-768_A-12/vocab.txt'
BERT_INIT_CHKPNT = 'multi_cased_L-12_H-768_A-12/bert_model.ckpt'
BERT_CONFIG = 'multi_cased_L-12_H-768_A-12/bert_config.json'
tokenizer = tokenization.FullTokenizer(
vocab_file=BERT_VOCAB, do_lower_case=False)
GO = 101
EOS = 102
from unidecode import unidecode
def get_inputs(x, y):
input_ids, input_masks, segment_ids, ys = [], [], [], []
for i in tqdm(range(len(x))):
tokens_a = tokenizer.tokenize(unidecode(x[i]))
tokens_b = tokenizer.tokenize(unidecode(y[i]))
tokens = ["[CLS]"] + tokens_a + ["[SEP]"]
segment_id = [0] * len(tokens)
input_id = tokenizer.convert_tokens_to_ids(tokens)
input_mask = [1] * len(input_id)
input_ids.append(input_id)
input_masks.append(input_mask)
segment_ids.append(segment_id)
r = tokenizer.convert_tokens_to_ids(tokens_b + ["[SEP]"])
if len([k for k in r if k == 0]):
print(y[i], i)
break
ys.append(r)
return input_ids, input_masks, segment_ids, ys
train_input_ids, train_input_masks, train_segment_ids, train_Y = get_inputs(train_X, train_Y)
test_input_ids, test_input_masks, test_segment_ids, test_Y = get_inputs(test_X, test_Y)
bert_config = modeling.BertConfig.from_json_file(BERT_CONFIG)
epoch = 20
batch_size = 16
warmup_proportion = 0.1
num_train_steps = len(train_input_ids)
num_warmup_steps = int(num_train_steps * warmup_proportion)
class Model:
def __init__(
self,
size_layer,
num_layers,
learning_rate = 2e-5,
training = True,
):
self.X = tf.placeholder(tf.int32, [None, None])
self.segment_ids = tf.placeholder(tf.int32, [None, None])
self.input_masks = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.count_nonzero(self.X, 1, dtype=tf.int32)
self.Y_seq_len = tf.count_nonzero(self.Y, 1, dtype=tf.int32)
batch_size = tf.shape(self.X)[0]
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
model = modeling.BertModel(
config=bert_config,
is_training=training,
input_ids=self.X,
input_mask=self.input_masks,
token_type_ids=self.segment_ids,
use_one_hot_embeddings=False)
output_layer = model.get_sequence_output()
pooled_output = model.get_pooled_output()
embedding = model.get_embedding_table()
dense = tf.layers.Dense(bert_config.vocab_size)
def cells(size_layer=size_layer, reuse=False):
return tf.nn.rnn_cell.LSTMCell(size_layer,initializer=tf.orthogonal_initializer(),reuse=reuse)
lstm_state = tf.nn.rnn_cell.LSTMStateTuple(c=pooled_output, h=pooled_output)
encoder_state = tuple([lstm_state] * num_layers)
decoder_cells = tf.nn.rnn_cell.MultiRNNCell([cells(size_layer) for _ in range(num_layers)])
training_helper = tf.contrib.seq2seq.TrainingHelper(
inputs = tf.nn.embedding_lookup(embedding, decoder_input),
sequence_length = self.Y_seq_len,
time_major = False)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cells,
helper = training_helper,
initial_state = encoder_state,
output_layer = dense)
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
self.training_logits = training_decoder_output.rnn_output
predicting_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(
embedding = embedding,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS)
predicting_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cells,
helper = predicting_helper,
initial_state = encoder_state,
output_layer = dense)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = True,
maximum_iterations = 2 * tf.reduce_max(self.X_seq_len))
self.fast_result = predicting_decoder_output.sample_id
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate).minimize(self.cost)
y_t = tf.argmax(self.training_logits,axis=2)
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(y_t, masks)
mask_label = tf.boolean_mask(self.Y, masks)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(bert_config.hidden_size, 2)
sess.run(tf.global_variables_initializer())
import collections
import re
def get_assignment_map_from_checkpoint(tvars, init_checkpoint):
"""Compute the union of the current variables and checkpoint variables."""
assignment_map = {}
initialized_variable_names = {}
name_to_variable = collections.OrderedDict()
for var in tvars:
name = var.name
m = re.match('^(.*):\\d+$', name)
if m is not None:
name = m.group(1)
name_to_variable[name] = var
init_vars = tf.train.list_variables(init_checkpoint)
assignment_map = collections.OrderedDict()
for x in init_vars:
(name, var) = (x[0], x[1])
if 'bert/' + name in name_to_variable:
assignment_map[name] = name_to_variable['bert/' + name]
initialized_variable_names[name] = 1
initialized_variable_names[name + ':0'] = 1
elif name in name_to_variable:
assignment_map[name] = name_to_variable[name]
initialized_variable_names[name] = 1
initialized_variable_names[name + ':0'] = 1
return (assignment_map, initialized_variable_names)
tvars = tf.trainable_variables()
checkpoint = BERT_INIT_CHKPNT
assignment_map, initialized_variable_names = get_assignment_map_from_checkpoint(tvars,
checkpoint)
saver = tf.train.Saver(var_list = assignment_map)
saver.restore(sess, checkpoint)
pad_sequences = tf.keras.preprocessing.sequence.pad_sequences
from tqdm import tqdm
import time
for EPOCH in range(epoch):
train_acc, train_loss, test_acc, test_loss = [], [], [], []
pbar = tqdm(
range(0, len(train_input_ids), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_input_ids))
batch_x = train_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post')
batch_mask = train_input_masks[i: index]
batch_mask = pad_sequences(batch_mask, padding='post')
batch_segment = train_segment_ids[i: index]
batch_segment = pad_sequences(batch_segment, padding='post')
batch_y = pad_sequences(train_Y[i: index], padding='post')
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.input_masks: batch_mask,
model.segment_ids: batch_segment
},
)
train_loss.append(cost)
train_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_input_ids), batch_size), desc = 'test minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post')
batch_y = pad_sequences(test_Y[i: index], padding='post')
batch_mask = test_input_masks[i: index]
batch_mask = pad_sequences(batch_mask, padding='post')
batch_segment = test_segment_ids[i: index]
batch_segment = pad_sequences(batch_segment, padding='post')
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.input_masks: batch_mask,
model.segment_ids: batch_segment
},
)
test_loss.append(cost)
test_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss = np.mean(train_loss)
train_acc = np.mean(train_acc)
test_loss = np.mean(test_loss)
test_acc = np.mean(test_acc)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
from tensor2tensor.utils import bleu_hook
results = []
for i in tqdm(range(0, len(test_X), batch_size)):
index = min(i + batch_size, len(test_X))
batch_x = test_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post')
batch_y = pad_sequences(test_Y[i: index], padding='post')
batch_mask = test_input_masks[i: index]
batch_mask = pad_sequences(batch_mask, padding='post')
batch_segment = test_segment_ids[i: index]
batch_segment = pad_sequences(batch_segment, padding='post')
feed = {
model.X: batch_x,
model.input_masks: batch_mask,
model.segment_ids: batch_segment
}
p = sess.run(model.fast_result,feed_dict = feed)
result = []
for row in p:
result.append([i for i in row if i > 3 and i not in [101, 102]])
results.extend(result)
rights = []
for r in test_Y:
rights.append([i for i in r if i > 3 and i not in [101, 102]])
bleu_hook.compute_bleu(reference_corpus = rights,
translation_corpus = results)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W2D5_GenerativeModels/W2D5_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a> <a href="https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W2D5_GenerativeModels/W2D5_Tutorial4.ipynb" target="_parent"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" alt="Open in Kaggle"/></a>
# (Bonus) Tutorial 4: Deploying Neural Networks on the Web
**Week 2, Day 5: Generative models**
**By Neuromatch Academy**
__Content creators:__ Sam Ray, Vladimir Haltakov, Konrad Kording
__Production editors:__ Spiros Chavlis
---
# Tutorial Objectives
In this tutorial, you will learn the basics of how to deploy your deep learning models as web applications using some modern frameworks and libraries. In this tutorial, you will learn to:
- Serve web pages with Flask
- Apply the MVVM design pattern to write maintainable code
- Create an interactive UI for your service
- Deploy your deep learning models as a REST API
- Deploying your application on Heroku
```
# @title Tutorial slides
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/p6wty/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
These are the slides for the videos in this tutorial. If you want to locally download the slides, click [here](https://osf.io/p6wty/download).
---
# Setup
Run the following cells to install and include important dependencies.
```
# @title Install dependencies
!pip install --upgrade jupyter-client --quiet
!pip install flask-ngrok Flask-RESTful flasgger --quiet
# Imports
import io
import platform
from urllib.request import urlopen
import flasgger
from flask_restful import Api
from flask_ngrok import run_with_ngrok
from flask_restful import Resource, fields, marshal
from flask import Flask, render_template_string, request, redirect
import torch
import numpy as np
from PIL import Image
from torchvision import models
import torchvision.transforms as transforms
```
---
# Section 1: Introduction
```
# @title Video 1: Deploying Neural Networks on the Web
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1754y1E7Qf", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"yQtPGtz4jDI", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
We will start by building a simple web application in Flask, which we'll keep extending throughout the tutorial. In the end, you will have a web app where you can upload an image and have it classified automatically by a neural network model.
---
# Section 2: Flask
Flask is a web application micro-framework built with Python. Flask is popular because it's lightweight, easy to use, scalable, and has tons of great extensions. Nowadays, Falsk is used for many different applications like web applications, REST APIs, socket-based services, and by companies like LinkedIn or Pinterest.
In this section, you will learn to create simple Flask websites.
## Section 2.1: Your First Flask App
```
# @title Video 2: Flask
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1sA411P7Rq", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"uVqu-9IBIRg", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Creating a minimal Flask app is very simple. You need to create a `Flask` object and define the handler for the root URL returning the HTML response. You need to provide the applications module or package, but we can use `__name__` as a convenient shortcut.
We need one small trick because the app will be running in a notebook. If you just run the app, it will be accessible at `http://127.0.0.1:5000`. The problem is that this is a local address to the server where the notebook is running, so you can't access it. This is where `ngrok` helps - it creates a tunnel from the notebook server to the outside world. Make sure you use the ngrok URL when testing your app.
> Note: the call to `app.run()` will not return on its own. Make sure to stop the running cell when you want to move to the next one.
```
# Create a Flask app object
app = Flask(__name__)
# Define a function to be called when the user accesses the root URL (/)
@app.route("/")
def home():
# Return a very simple HTML response
return "<h1>Welcome to Neuromatch</h1>"
# You need ngrok to expose the Flask app outside of the notebook
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
## Section 2.2: Using Jinja2 Templates
```
# @title Video 3: Jinja Templates
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Mb4y167eg", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"u25FfNIAKsg", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
The default template engine used by Flask is Jinja2. Jinja2 offers features that help you write clean and reusable templates such as inheritance, humanizing, and formatting data (there's an extension for this), dividing components into sub-modules, etc.
In this section, we are going to add Jinja2 templates to the app. WIth Jinja2 you can use variables and control flow commands, like ifs and loops in your HTML code. Then you can pass data from your Python code to the template when it is rendered.
Let's first define the template of a simple web page showing some platform properties. We are going to loop over the `platform` dictionary containing property keys and their corresponding values:
```
{% for key, value in platform.items() %}
```
One row of the HTML table is created for every element of the dictionary. You can display the content of the variables like that: `{{ value }}`
```
# Jinja2 HTML template
template_str = '''
<html lang="en">
<body>
<div style="width: 50%; margin: 100px auto">
<h1>Platform Info</h1>
<table>
<tr style="width:">
<th style="width: 200px">Property</th>
<th>Value</th>
</tr>
{% for key, value in platform.items() %}
<tr>
<td style="width: 200px">{{ key }}</td>
<td>{{ value }}</td>
</tr>
{% endfor %}
</table>
</div>
</body>
</html>
'''
```
You can then render the template passing the platform properties that you can retrieve using the `platform` package.
```
app = Flask(__name__)
@app.route("/")
def home():
# Get the platform properties as a dict
properties = platform.uname()._asdict()
# Render the Jinja2 template
return render_template_string(template_str, platform=properties)
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
## Section 2.3: Apply the MVVM Design Pattern
```
# @title Video 4: Using the MVVM Design Pattern
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1YA411P766", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"13bFN4L6c9I", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Design patterns provide a way of writing reusable, adaptable, and extendable code. Design patterns are not libraries, but rather a set of best practices to follow when designing your software.
Model View View-Model (MVVM) is a powerful design pattern commonly used in web applications (and other GUI applications).
* **View** - this is that part of your code the user interacts with (the web page)
* **Model** - this is the representation of the data that you want to interact with.
* **View-Model** - this is the part that handles the application state and that transforms the data from the Model to a representation suitable for display and back.
<img src="https://raw.githubusercontent.com/NeuromatchAcademy/course-content-dl/main/tutorials/W2D5_GenerativeModels/static/deploy.png">
Let's implement the MVVM pattern in Flask. You will first create classes for each of the 3 parts of the MVVM pattern used to display information about 2D points.
```
# Model - simple point Model storing a 2D point
class PointModel:
# Initialize a 2D point
def __init__(self, x, y):
self.x = x
self.y = y
# View - simple View displaying the information about a 2D point
class PointView(Resource):
def get(self):
point = PointViewModel.get_sample_data()
return f"Point: (x={point.x}, y={point.y})"
# ViewModel - simple ViewModel retrieving the data and passing it to the view
class PointViewModel:
# Create some sample data
@classmethod
def get_sample_data(cls):
return PointModel(2, 5)
# Register a handler for "/" in the API calling the PointView
def setup(self, api):
api.add_resource(PointView, '/')
```
You can now create your Falsk app and use the `Api` object where you can register your ViewModel.
```
# Create a Flask app
app = Flask(__name__)
# Create an Api object where different ViewModels can be registered
api = Api(app)
# Create a PointViewModel and register it to the API
pvm = PointViewModel()
pvm.setup(api)
# Run with ngrok
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
## Section 2.4: Creating a REST API
```
# @title Video 5: REST API
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1A64y1z74c", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"cIjaEE6tKpk", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
REST (Representational State Transfer) is a set of rules according to which APIs are designed to enable your service to interact with other services. If HTML pages are interfaces designed for humans, you can think about REST APIs as interfaces made for computers.
A common way to implement a REST API is for your application to respond to certain requests by returning a JSON string containing the required data.
Let's now create a new View and ViewModel that will provide the platform properties in JSON format.
```
# New View displaying some platform properties
class PlatformView(Resource):
def get(self):
"""
This examples uses PlatformView Resource
It works also with swag_from, schemas and spec_dict
---
responses:
200:
description: A single Machine item
schema:
id: Machine
properties:
machine:
type: string
description: The type of the processor
default: None
node:
type: string
description: The name of the current virtual machine
default: None
processor:
type: string
description: The type of the processor arch
default: None
system:
type: string
description: The name of the user
default: None
"""
# Specification of the returned data
resource_fields = {
'system': fields.String,
'machine': fields.String,
'processor': fields.String,
'node': fields.String
}
# Serialize the data according to the specification
return marshal(platform.uname()._asdict(), resource_fields)
# A simple ViewModel that displays the PlatformView at /platform
class PlatformViewModel:
def setup(self, api):
api.add_resource(PlatformView, '/platform')
```
Note the documentation in the `get` method. We can use the `flasgger` package to automatically create documentation of your REST API. You can access it at `/apidocs`.
```
# Create the Flask app and register the ViewModel
app = Flask(__name__)
api = Api(app)
pvm = PlatformViewModel()
pvm.setup(api)
# Redirect / to /platform for convenience
@app.route('/')
def redirect_platform():
return redirect("/platform", code=302)
# Register Swagger to create API documentation at /apidocs
swg = flasgger.Swagger(app)
# Run the app
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
---
# Section 3: Vue.js
```
# @title Video 6: Vue.js
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Yv411K7GS", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"PD6l9pkjw-c", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
We already talked about the MVVM pattern and implemented it using a back end framework - Flask. Applying the same pattern on the front end can also be beneficial when creating dynamic applications.
Vue.js is a great front end library that implements the MVVM design pattern. It is widely used for creating user interfaces and single-page applications.
In this section, you will learn how to implement a simple Vue.js front end that fetches data from the platform REST API we created in the previous section.
## Section 3.1: Defining the Vue Template
We define our HTML template similarly to Jinja. The big difference is that the Vue template is rendered dynamically after the page is loaded, while the Jinja templates are rendered in the backend before the page is served.
Using variables is very similar to Jinja, but now you also have some JavaScript code handling the state of the application and the binding to the data. You can use the `axois` package to fetch data from our platform REST API, when Vue is initialized (mounted).
```
vue_template = """
<html>
<head>
<script src="https://cdn.jsdelivr.net/npm/vue/dist/vue.js"></script>
<script src="https://cdnjs.cloudflare.com/ajax/libs/axios/0.21.1/axios.min.js"></script>
</head>
<body>
<div id="app">
<ul>
<li><strong>System: </strong>{{ platform.system }}</li>
<li><strong>Machine: </strong>{{ platform.machine }}</li>
<li><strong>Processor: </strong>{{ platform.processor }}</li>
<li><strong>Node: </strong>{{ platform.node }}</li>
</ul>
</div>
<script >
var app = new Vue({
el: '#app',
data() {
return {
platform: null
}
},
mounted () {
axios.get('/platform')
.then(response => (this.platform = response.data))
}
});
</script>
</body>
</html>
"""
```
## Section 3.2: Serving the Vue.js App
You can now again run your Flask app and serve the Vue template from the root URL. We are still using Flask to implement our logic on the backend and provide the platform REST API.
This way of doing things may seem more complicated, but it has the advantage that it is dynamic. In the next section, we will add some dynamic functionality.
```
# Create the Flask app with the previously defined platform API
app = Flask(__name__)
api = Api(app)
# Serve the Platform REST API
pvm = PlatformViewModel()
pvm.setup(api)
swg = flasgger.Swagger(app)
# Serve the Vue template page at /
@app.route("/")
def home():
return vue_template
# Run the app
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
---
# Section 4: Model Presentation
```
# @title Video 7: Deploying a PyTorch model
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Zb4y1z7aT", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"6UCLk37XWDs", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Now (finally) we have all the tools we need to deploy our neural network! We are going to use a pre-trained DenseNet mode. In the first step, we are going to create an API entry point that accepts an image as input and classifies it. After that, we will create a dynamic UI for easier interaction.
## Section 4.1: Image Classification API
```
# @title Video 8: Classification with a Pre-trained Model
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Dq4y1n7ks", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"UGByJ-_0whk", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
First, we need to load a pre-trained DenseNet trained on ImageNet. You can use `torchvision.models` to quickly get a pre-trained model for many popular neural network architectures.
```
# Load a pre-trainied DenseNet model from torchvision.models
model = models.densenet121(pretrained=True)
# Switch the model to evaluation mode
model.eval()
# Load the class labels from a file
class_labels_url = "https://raw.githubusercontent.com/pytorch/hub/master/imagenet_classes.txt"
class_labels = urlopen(class_labels_url).read().decode("utf-8").split("\n")
# Define the transformation of the input image
transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]),
])
```
Define a function to predict the most likely class using the model. Note that we need to pass a batch of images to the model. Since we have only 1, we can just use `unsqueese(0)` to add an additional dimension.
```
def predict(model, transform, image, class_labels):
# Transform the image and convert it to a tensor
image_tensor = transform(image).unsqueeze(0)
# Pass the image through the model
with torch.no_grad():
output = model(image_tensor)
# Select the class with the higherst probability and look up the name
class_id = torch.argmax(output).item()
class_name = class_labels[class_id]
# Return the class name
return class_name
```
Let's test the `predict` function using an image of a [dog](https://unsplash.com/photos/2l0CWTpcChI/download?force=true&w=640).
```
# Load and display the image
dog_image = Image.open(io.BytesIO(urlopen("https://unsplash.com/photos/2l0CWTpcChI/download?force=true&w=480").read()))
display(dog_image)
# Classify the image
display(predict(model, transform, dog_image, class_labels))
```
## Section 4.2: Create a Dynamic Application
```
# @title Video 9: Create a Dynamic Application
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Vy4y1L7V9", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"DJsK2bc9wuk", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
We will now create a Flask app that receives an image at `/predict` and passes it through the model. We will also implement an interactive UI to upload the image and call the API.
The UI consists of a file upload field, a classify button, and an image displaying the uploaded file.
```
index_template = """
<html>
<head>
<!-- Load vue.js and axois.js -->
<script src="https://cdn.jsdelivr.net/npm/vue/dist/vue.js"></script>
<script src="https://cdnjs.cloudflare.com/ajax/libs/axios/0.21.1/axios.min.js"></script>
</head>
<body>
<!-- The APP UI -->
<div id="app" style="width: 50%; margin: 200px auto">
<form id="imageForm" enctype="multipart/form-data" method="POST" style="text-align: center; display: block">
<label for="imageFile">Select image to classify:</label
><input id="imageFile" name="file" type="file" style="margin-left: 10px" />
<img v-if="image" :src="image" style="width: 250px; display: block; margin: 50px auto 10px" />
<div v-if="prediction" style="font-size: 32px; font-weight: bold; text-align: center">
{{ prediction }}
</div>
<input v-if="image" type="submit" value="Classify Image" style="margin: 20px 20px" />
</form>
</div>
<script>
<!-- The Vue application -->
var app = new Vue({
el: "#app",
data() {
return {
image: null,
prediction: null,
};
},
});
<!-- Calling the predict API when the form is submitted -->
document.getElementById("imageForm").addEventListener("submit", (e) => {
axios
.post("/predict", new FormData(document.getElementById("imageForm")), {
headers: {
"Content-Type": "multipart/form-data",
},
})
.then((response) => (app.prediction = response.data));
e.preventDefault();
});
<!-- Display the selected image -->
document.getElementById("imageFile").addEventListener("change", (e) => {
const [file] = document.getElementById("imageFile").files;
if (file) {
app.image = URL.createObjectURL(file);
}
});
</script>
</body>
</html>
"""
```
The application has two entry points:
* `/` - serve the Vue template with the interactive UI
* `/predict` - a REST API classifying the image received as input
```
app = Flask(__name__)
# Serve the Vue template with the interactive UI
@app.route("/")
def home():
return index_template
# Classification API
@app.route("/predict", methods=['POST'])
def predict_api():
# Fetch the image from the request and convert it to a Pillow image
image_file = request.files['file']
image_bytes = image_file.read()
image = Image.open(io.BytesIO(image_bytes))
# Predict the class from the image
class_name = predict(model, transform, image, class_labels)
# Return the result
return class_name
# Run the app
run_with_ngrok(app)
## Uncomment below to run the app
# app.run()
```
---
# Section 5: Deploy a Flask app on Heroku
```
# @title Video 10: Deploy on Heroku
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1oo4y1S77Z", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"kaf6z-tAxCY", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Now you are going to deploy your application as a real web server outside of the notebook. We are going to use Heroku for this. Heroku is a PaaS (Platform-as-a-Service) that offers pre-configured environments so you can deploy an application easily and quickly. They also offer a free tier which is enough for deploying simple apps.
But first, you need to test your application locally.
## Section 5.1: Preparing Your Environment
```
# @title Video 11: Prepare Python Environment
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1bv411K7dP", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"IMd_sRm4fJM", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
You need to do all the steps from here on on your own machine and not in the notebook. You need to make sure that you have Python 3 installed and some code editor (for example VS Code). You will also be using the terminal a lot in this section.
First, you need to prepare your Python environment and install all required dependencies. You should first create an empty folder where you will store your application and do the following steps.
**1. Create a new virtual environment**
Run the following code in the terminal to create a new Python virtual environment:
```
python -m venv .venv
```
**2. Activate the virtual environment**
Now, you need to activate the environment, which is a bit different on Linux/macOS and Windows.
For Linux and macOS:
```
source .venv/bin/activate
```
For Windows:
```
.venv\Scripts\activate.bat
```
**3. Install dependencies**
You need to install some packages that you will need using `pip`:
```
pip install flask Pillow gunicorn
```
> Note: the package `gunicorn` is a web server that is needed later when your code runs on Heroku.
**4. Install PyTorch**
Depending on your system, there are different ways to install `torch` and `torchvision`. Refer to the [Installation page](https://pytorch.org/get-started/locally/) for the exact command. We recommend using `pip`.
On macOS and Windows for example this is straightforward:
```
pip install torch torchvision
```
> Note: avoid installing `torchaudio` since it is not needed, but may cause problems with the package size when deploying on Heroku later.
## Section 5.2: Create Your Application
```
# @title Video 12: Creating a Local Application
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1bM4y157xK", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"fF1fmIXz5NQ", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
You are now ready to create the files needed for your application. For now, you need just 2 files.
**`app.py`**
This is the main file of your application. Inside, you will put the code for running your PyTorch model as well as the code of your Flask application.
```python
import os
import io
import torch
from urllib.request import urlopen
from PIL import Image
from torchvision import models
import torchvision.transforms as transforms
from flask import Flask, request, send_from_directory
# Load a pre-trainied DenseNet model from torchvision.models
model = models.densenet121(pretrained=True)
model.eval()
# Load the class labels from a file
class_labels_url = (
"https://raw.githubusercontent.com/pytorch/hub/master/imagenet_classes.txt"
)
class_labels = urlopen(class_labels_url).read().decode("utf-8").split("\n")
# Define the transofrmation of the input image
transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]),
])
def predict(model, transform, image, class_labels):
# Transform the image and convert it to a tensor
image_tensor = transform(image).unsqueeze(0)
# Pass the image through the model
with torch.no_grad():
output = model(image_tensor)
# Select the class with the higherst probability
class_id = torch.argmax(output).item()
class_name = class_labels[class_id]
return class_name
app = Flask(__name__)
@app.route("/")
def home():
return send_from_directory("static", "index.html")
@app.route("/predict", methods=["POST"])
def predict_api():
# Fetch the image from the request and convert it
image_file = request.files["file"]
image_bytes = image_file.read()
image = Image.open(io.BytesIO(image_bytes))
# Predict the class from the image
class_name = predict(model, transform, image, class_labels)
# Write result as JSON
return class_name
# Run the app
if __name__ == "__main__":
app.run(debug=False, threaded=True, port=os.getenv("PORT", 5000))
```
**static/index.html**
This file should contain the HTML code of your Vue template - exactly as it is.
```html
<html>
<head>
<!-- Load vue.js and axois.js -->
<script src="https://cdn.jsdelivr.net/npm/vue/dist/vue.js"></script>
<script src="https://cdnjs.cloudflare.com/ajax/libs/axios/0.21.1/axios.min.js"></script>
</head>
<body>
<!-- The APP UI -->
<div id="app" style="width: 50%; margin: 200px auto">
<form id="imageForm" enctype="multipart/form-data" method="POST" style="text-align: center; display: block">
<label for="imageFile">Select image to classify:</label
><input id="imageFile" name="file" type="file" style="margin-left: 10px" />
<img v-if="image" :src="image" style="width: 250px; display: block; margin: 50px auto 10px" />
<div v-if="prediction" style="font-size: 32px; font-weight: bold; text-align: center">
{{ prediction }}
</div>
<input v-if="image" type="submit" value="Classify Image" style="margin: 20px 20px" />
</form>
</div>
<script>
<!-- The Vue application -->
var app = new Vue({
el: "#app",
data() {
return {
image: null,
prediction: null,
};
},
});
<!-- Calling the predict API when the form is submitted -->
document.getElementById("imageForm").addEventListener("submit", (e) => {
axios
.post("/predict", new FormData(document.getElementById("imageForm")), {
headers: {
"Content-Type": "multipart/form-data",
},
})
.then((response) => (app.prediction = response.data));
e.preventDefault();
});
<!-- Display the selected image -->
document.getElementById("imageFile").addEventListener("change", (e) => {
const [file] = document.getElementById("imageFile").files;
if (file) {
app.image = URL.createObjectURL(file);
}
});
</script>
</body>
</html>
```
## Section 5.3: Testing Your Application Locally
You are now ready to test your application. Once you have your environment set up correctly and your application files created you can just start it:
```
python app.py
```
You can now access your application at http://127.0.0.1:5000. No `ngrok` needed anymore!
## Section 5.4: Preparing for Deployment on Heroku
```
# @title Video 13: Preparing for Heroku
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Nq4y1Q71H", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"L2W0C7nMttI", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
Before we can deploy on Heroku there are a couple of things we need to prepare.
**Create `Procfile`**
Every application running on Heroku needs a `Procfile` where you need to specify how the app should be run. In our case, it is quite easy, because we can use `gunicorn` as a web server. Create a file named `Procfile` in the root folder of your application and put the following code inside:
```
web: gunicorn app:app
```
**Create `requirements.txt`**
We also need to tell Heroku which Python packages need to be installed. Heroku uses the standard way of defining Python dependencies - a `requirements.txt` file. You can create it with the following command:
```
pip freeze > requirements.txt
```
**Fix the `torch` version**
Now, this should be enough in theory, but we need one small change. The problem is that by default `torch` comes with both the CPU and GPU code, which creates a package that exceeds the maximum size limit on the Heroku free tier. Therefore we need to make sure that we only specify the CPU version of `torch`.
You need to open the `requirements.txt` file and modify it as follows:
1. Add the following line in the beginning, telling Heroku where to look for the packages `-f https://download.pytorch.org/whl/torch_stable.html
`
2. Find the line defining the torch dependency and change it to `torch==1.9.0+cpu`
3. Do the same with `torchvision` by changing it to `torchvision==0.10.0+cpu
`
## Section 5.5: Deploying on Heroku
```
# @title Video 14: Deploying on Heroku
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1P64y1z7cU", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"Ni9YKotZUQk", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
You are now finally ready to deploy to Heroku! There are just a couple of steps needed.
**1. Create a Heroku account**
Create a free account on Heroku at https://www.heroku.com/
**2. Install the Heroku CLI**
Use [this guide](https://devcenter.heroku.com/articles/heroku-cli) to install the Heroku CLI for your system. After installation, you should be able to run the `heroku` command in your terminal.
**3. Login to Heroku**
Run the following command in the terminal and log in:
```
heroku login
```
**4. Create a new Heroku App**
Run the following command to create a new application. When choosing the application name, you need to make sure that it doesn't exist yet. It may be a good idea to add your name to it (I chose `vladimir-classifier-app`).
```
heroku create <application name>
```
**5. Initialize a Git repository**
The deployment on Heroku is done using `git`. If you don't have it installed already, check [this guide](https://git-scm.com/book/en/v2/Getting-Started-Installing-Git). You need to initialize a new repository, commit your files and push it to Heroku:
```
git init
git add app.py Procfile requirements.txt static
git commit -m "Initial commit"
heroku git:remote -a <application name>
git push heroku master
```
Your application will now be packaged and uploaded to Heroku. This may take a couple of minutes, but when done you will be able to access your application at https://<application name>.herokuapp.com.
You can also go to your dashboard and see your application there: https://dashboard.heroku.com/apps.
---
# Summary
```
# @title Video 15: Summary
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Qg411L7RM", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"pDLdNOuUtKk", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
In this tutorial you learned the basics of some modern tools for creating dynamic web applications and REST APIs. You also learned how you can deploy your neural network model as a web app.
You can now build on top of that and create more sophisticated and awesome applications and make them available to millions of people!
| github_jupyter |
# End-to-end Reusable ML Pipeline with Seldon and Kubeflow
In this example we showcase how to build re-usable components to build an ML pipeline that can be trained and deployed at scale.
We will automate content moderation on the Reddit comments in /r/science building a machine learning NLP model with the following components:

This tutorial will break down in the following sections:
1) Run all the services (Kubeflow and Seldon)
2) Test and build all our reusable pipeline steps
3) Use Kubeflow to Train the Pipeline and Deploy to Seldon
5) Test Seldon Deployed ML REST Endpoints
6) Visualise Seldon's Production ML Pipelines
## Before you start
Make sure you install the following dependencies, as they are critical for this example to work:
* Helm v3.0.0+
* A Kubernetes cluster running v1.13 or above (minkube / docker-for-windows work well if enough RAM)
* kubectl v1.14+
* ksonnet v0.13.1+
* kfctl 0.5.1 - Please use this exact version as there are major changes every few months
* Python 3.6+
* Python DEV requirements (we'll install them below)
Let's get started! 🚀🔥 We will be building the end-to-end pipeline below:

```
!cat requirements-dev.txt
!pip install -r requirements-dev.txt
```
## 1) Run all the services (Kubeflow and Seldon)
Kubeflow's CLI allows us to create a project which will allow us to build the configuration we need to deploy our kubeflow and seldon clusters.
```
!kfctl init kubeflow-seldon
!ls kubeflow-seldon
```
Now we run the following commands to basically launch our Kubeflow cluster with all its components.
It may take a while to download all the images for Kubeflow so feel free to make yourself a cup of ☕.
If you have a terminal you can see how the containers are created in real-time by running `kubectl get pods -n kubeflow -w`.
```
%%bash
cd kubeflow-seldon
kfctl generate all -V
kfctl apply all -V
```
## Setup Seldon Core
Use the setup notebook to [Install Seldon Core](../../seldon_core_setup.ipynb#Install-Seldon-Core). Instructions [also online](./seldon_core_setup.html).
### Temporary fix for Argo image
At the time of writing we need to make some updates in the Argo images with the following commands below.
(This basically changes the images to the latest ones, otherwise we will get an error when we attach the volume)
```
!kubectl -n kubeflow patch deployments. workflow-controller --patch '{"spec": {"template": {"spec": {"containers": [{"name": "workflow-controller", "image": "argoproj/workflow-controller:v2.3.0-rc3"}]}}}}'
!kubectl -n kubeflow patch deployments. ml-pipeline --patch '{"spec": {"template": {"spec": {"containers": [{"name": "ml-pipeline-api-server", "image": "elikatsis/ml-pipeline-api-server:0.1.18-pick-1289"}]}}}}'
# !kubectl -n kubeflow patch configmaps workflow-controller-configmap --patch '{"data": {"config": "{ executorImage: argoproj/argoexec:v2.3.0-rc3,artifactRepository:{s3: {bucket: mlpipeline,keyPrefix: artifacts,endpoint: minio-service.kubeflow:9000,insecure: true,accessKeySecret: {name: mlpipeline-minio-artifact,key: accesskey},secretKeySecret: {name: mlpipeline-minio-artifact,key: secretkey}}}}" }}'
```
The last command you need to run actually needs to be manual as the patch cannot change configmap contents directly
You need to run the edit commad and change the executorImage to: `argoproj/argoexec:v2.3.0-rc3`
The command should be run from a terminal:
```
kubectl edit configmaps workflow-controller-configmap -n kubeflow
```
## 2) Test and build all our reusable pipeline steps
We will start by building each of the components in our ML pipeline.

### Let's first have a look at our clean_text step:
```
!ls pipeline/pipeline_steps
```
Like in this step, all of the other steps can be found in the `pipeline/pipeline_steps/` folder, and all have the following structure:
* `pipeline_step.py` which exposes the functionality through a CLI
* `Transformer.py` which transforms the data accordingly
* `requirements.txt` which states the python dependencies to run
* `build_image.sh` which uses `s2i` to build the image with one line
### Let's check out the CLI for clean_text
The pipeline_step CLI is the entry point for the kubeflow image as it will be able to pass any relevant parameters
```
!python pipeline/pipeline_steps/clean_text/pipeline_step.py --help
```
This is actually a very simple file, as we are using the click library to define the commands:
```
!cat pipeline/pipeline_steps/clean_text/pipeline_step.py
```
The Transformer is where the data munging and transformation stage comes in, which will be wrapped by the container and exposed through the Seldon Engine to ensure our pipeline can be used in production.
Seldon provides multiple different features, such as abilities to send custom metrics, pre-process / post-process data and more. In this example we will only be exposing the `predict` step.
```
!cat pipeline/pipeline_steps/clean_text/Transformer.py
```
If you want to understand how the CLI pipeline talks to each other, have a look at the end to end test in `pipeline/pipeline_tests/`:
```
!pytest ./pipeline/pipeline_tests/. --disable-pytest-warnings
```
To build the image we provide a build script in each of the steps that contains the instructions:
```
!cat pipeline/pipeline_steps/clean_text/build_image.sh
```
The only thing you need to make sure is that Seldon knows how to wrap the right model and file.
This can be achieved with the s2i/environment file.
As you can see, here we just tell it we want it to use our `Transformer.py` file:
```
!cat pipeline/pipeline_steps/clean_text/.s2i/environment
```
Once this is defined, the only thing we need to do is to run the `build_image.sh` for all the reusable components.
Here we show the manual way to do it:
```
%%bash
# we must be in the same directory
cd pipeline/pipeline_steps/clean_text/ && ./build_image.sh
cd ../data_downloader && ./build_image.sh
cd ../lr_text_classifier && ./build_image.sh
cd ../spacy_tokenize && ./build_image.sh
cd ../tfidf_vectorizer && ./build_image.sh
```
## 3) Train our NLP Pipeline through the Kubeflow UI
We can access the Kubeflow dashboard to train our ML pipeline via http://localhost/_/pipeline-dashboard
If you can't edit this, you need to make sure that the ambassador gateway service is accessible:
```
!kubectl get svc ambassador -n kubeflow
```
In my case, I need to change the kind from `NodePort` into `LoadBalancer` which can be done with the following command:
```
!kubectl patch svc ambassador --type='json' -p '[{"op":"replace","path":"/spec/type","value":"LoadBalancer"}]' -n kubeflow
```
Now that I've changed it to a loadbalancer, it has allocated the external IP as my localhost so I can access it at http://localhost/_/pipeline-dashboard
```
!kubectl get svc ambassador -n kubeflow
```
If this was successfull, you should be able to access the dashboard

### Define the pipeline
Now we want to generate the pipeline. For this we can use the DSL provided by kubeflow to define the actual steps required.
The pipeline will look as follows:

```
!cat train_pipeline/nlp_pipeline.py
```
### Breaking down the code
As you can see in the DSL, we have the ContainerOp - each of those is a step in the Kubeflow pipeline.
At the end we can see the `seldondeploy` step which basically deploys the trained pipeline
The definition of the SeldonDeployment graph is provided in the `deploy_pipeline/seldon_production_pipeline.yaml` file.
The seldondeployment file defines our production execution graph using the same reusable components.
```
!cat deploy_pipeline/seldon_production_pipeline.yaml
```
### Seldon Production pipeline contents
If we look at the file we'll be using to deploy our pipeline, we can see that it has the following key points:
1) Reusable components definitions as containerSpecs: cleantext, spacytokenizer, tfidfvectorizer & lrclassifier
2) DAG (directed acyclic graph) definition for REST pipeline: cleantext -> spacytokenizer -> tfidfvectorizer -> lrclassifier
This graph in our production deployment looks as follows:

### Generate the pipeline files to upload to Kubeflow
To generate the pipeline we just have to run the pipeline file, which will output the `tar.gz` file that will be uploaded.
```
%%bash
# Generating graph definition
python train_pipeline/nlp_pipeline.py
ls train_pipeline/
```
### Run the pipeline
We now need to upload the resulting `nlp_pipeline.py.tar.gz` file generated.
This can be done through the "Upload PIpeline" button in the UI at http://localhost/_/pipeline-dashboard.
Once it's uploaded, we want to create and trigger a run! You should now be able to see how each step is executed:

### Inspecting the data created in the Persistent Volume
The pipeline saves the output of the pipeline together with the trained model in the persistent volume claim.
The persistent volume claim is the same name as the argo workflow:
```
!kubectl get workflow -n kubeflow
```
Our workflow is there! So we can actually access it by running
```
!kubectl get workflow -n kubeflow -o jsonpath='{.items[0].metadata.name}'
```
And we can use good old `sed` to insert this workflow name in our PVC-Access controler which we can use to inspect the contents of the volume:
```
!sed "s/PVC_NAME/"$(kubectl get workflow -n kubeflow -o jsonpath='{.items[0].metadata.name}')"-my-pvc/g" deploy_pipeline/pvc-access.yaml
```
We just need to apply this container with our kubectl command, and we can use it to inspect the mounted folder:
```
!sed "s/PVC_NAME/"$(kubectl get workflow -n kubeflow -o jsonpath='{.items[0].metadata.name}')"-my-pvc/g" deploy_pipeline/pvc-access.yaml | kubectl -n kubeflow apply -f -
!kubectl get pods -n kubeflow pvc-access-container
```
Now we can run an `ls` command to see what's inside:
```
!kubectl -n kubeflow exec -it pvc-access-container ls /mnt
!kubectl delete -f deploy_pipeline/pvc-access.yaml -n kubeflow
```
## 5) Test Deployed ML REST Endpoints
Now that it's running we have a production ML text pipeline that we can Query using REST and GRPC
First we can check if our Seldon deployment is running with
```
!kubectl -n kubeflow get seldondeployment
```
We will need the Seldon Pipeline Deployment name to reach the API, so we can get it using:
```
!kubectl -n kubeflow get seldondeployment -o jsonpath='{.items[0].metadata.name}'
```
Now we can interact with our API in two ways:
1) Using CURL or any client like PostMan
2) Using the Python SeldonClient
### Using CURL from the terminal
When using CURL, the only thing we need to provide is the data in JSON format, as well as the url, which is of the format:
```
http://<ENDPOINT>/seldon/kubeflow/<PIPELINE_NAME>/api/v0.1/predictions
```
```
%%bash
curl -X POST -H 'Content-Type: application/json' \
-d "{'data': {'names': ['text'], 'ndarray': ['Hello world this is a test']}}" \
http://127.0.0.1/seldon/kubeflow/$(kubectl -n kubeflow get seldondeployment -o jsonpath='{.items[0].metadata.name}')/api/v0.1/predictions
```
### Using the SeldonClient
We can also use the Python SeldonClient to interact with the pipeline we just deployed
```
from seldon_core.seldon_client import SeldonClient
import numpy as np
import subprocess
host = "localhost"
port = "80" # Make sure you use the port above
batch = np.array(["Hello world this is a test"])
payload_type = "ndarray"
# Get the deployment name
deployment_name = subprocess.getoutput("kubectl -n kubeflow get seldondeployment -o jsonpath='{.items[0].metadata.name}'")
transport="rest"
namespace="kubeflow"
sc = SeldonClient(
gateway="ambassador",
ambassador_endpoint=host + ":" + port,
namespace=namespace)
client_prediction = sc.predict(
data=batch,
deployment_name=deployment_name,
names=["text"],
payload_type=payload_type,
transport="rest")
print(client_prediction)
```
## 6) Visualise Seldon's Production ML Pipelines
We can visualise the performance using the SeldonAnalytics package, which we can deploy using:
```
!helm install seldon-core-analytics --repo https://storage.googleapis.com/seldon-charts --namespace kubeflow
```
In my case, similar to what I did with Ambassador, I need to make sure the the service is a LoadBalancer instead of a NodePort
```
!kubectl patch svc grafana-prom --type='json' -p '[{"op":"replace","path":"/spec/type","value":"LoadBalancer"}]' -n kubeflow
!kubectl get svc grafana-prom -n kubeflow
```
Now we can access it at the port provided, in my case it is http://localhost:32445/d/3swM2iGWz/prediction-analytics?refresh=5s&orgId=1
(initial username is admin and password is password, which will be requested to be changed on the first login)
Generate a bunch of requests and visualise:
```
while True:
client_prediction = sc.predict(
data=batch,
deployment_name=deployment_name,
names=["text"],
payload_type=payload_type,
transport="rest")
```
## You now have a full end-to-end training and production NLP pipeline 😎

| github_jupyter |
```
!pip install transformers
!pip install torchvision
!pip install SentencePiece
import os
import math
import torch
from torch.nn import BCEWithLogitsLoss
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from transformers import AdamW, XLNetTokenizer, XLNetModel, XLNetLMHeadModel, XLNetConfig
from keras.preprocessing.sequence import pad_sequences
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
from tqdm import tqdm, trange
import matplotlib.pyplot as plt
%matplotlib inline
from google.colab import drive
drive.mount('/content/drive')
```
## Tokenzation and Attention
```
tokenizer = XLNetTokenizer.from_pretrained('xlnet-base-cased', do_lower_case=True)
def tokenize_inputs(text_list, tokenizer, num_embeddings=512):
"""
Tokenizes the input text input into ids. Appends the appropriate special
characters to the end of the text to denote end of sentence. Truncate or pad
the appropriate sequence length.
"""
# tokenize the text, then truncate sequence to the desired length minus 2 for
# the 2 special characters
tokenized_texts = list(map(lambda t: tokenizer.tokenize(t)[:num_embeddings-2], text_list))
# convert tokenized text into numeric ids for the appropriate LM
input_ids = [tokenizer.convert_tokens_to_ids(x) for x in tokenized_texts]
# append special token "<s>" and </s> to end of sentence
input_ids = [tokenizer.build_inputs_with_special_tokens(x) for x in input_ids]
# pad sequences
input_ids = pad_sequences(input_ids, maxlen=num_embeddings, dtype="long", truncating="post", padding="post")
return input_ids
def create_attn_masks(input_ids):
"""
Create attention masks to tell model whether attention should be applied to
the input id tokens. Do not want to perform attention on padding tokens.
"""
# Create attention masks
attention_masks = []
# Create a mask of 1s for each token followed by 0s for padding
for seq in input_ids:
seq_mask = [float(i>0) for i in seq]
attention_masks.append(seq_mask)
return attention_masks
```
## Load fintuned model
```
def load_model(save_path):
"""
Load the model from the path directory provided
"""
checkpoint = torch.load(save_path)
model_state_dict = checkpoint['state_dict']
model = XLNetForMultiLabelSequenceClassification(num_labels=model_state_dict["classifier.weight"].size()[0])
model.load_state_dict(model_state_dict)
epochs = checkpoint["epochs"]
lowest_eval_loss = checkpoint["lowest_eval_loss"]
train_loss_hist = checkpoint["train_loss_hist"]
valid_loss_hist = checkpoint["valid_loss_hist"]
return model, epochs, lowest_eval_loss, train_loss_hist, valid_loss_hist
torch.cuda.empty_cache()
config = XLNetConfig()
class XLNetForMultiLabelSequenceClassification(torch.nn.Module):
def __init__(self, num_labels=2):
super(XLNetForMultiLabelSequenceClassification, self).__init__()
self.num_labels = num_labels
self.xlnet = XLNetModel.from_pretrained('xlnet-base-cased')
self.classifier = torch.nn.Linear(768, num_labels)
torch.nn.init.xavier_normal_(self.classifier.weight)
def forward(self, input_ids, token_type_ids=None,\
attention_mask=None, labels=None):
# last hidden layer
last_hidden_state = self.xlnet(input_ids=input_ids,\
attention_mask=attention_mask,\
token_type_ids=token_type_ids)
# pool the outputs into a mean vector
mean_last_hidden_state = self.pool_hidden_state(last_hidden_state)
logits = self.classifier(mean_last_hidden_state)
if labels is not None:
loss_fct = BCEWithLogitsLoss()
loss = loss_fct(logits.view(-1, self.num_labels),\
labels.view(-1, self.num_labels))
return loss
else:
return logits
def freeze_xlnet_decoder(self):
"""
Freeze XLNet weight parameters. They will not be updated during training.
"""
for param in self.xlnet.parameters():
param.requires_grad = False
def unfreeze_xlnet_decoder(self):
"""
Unfreeze XLNet weight parameters. They will be updated during training.
"""
for param in self.xlnet.parameters():
param.requires_grad = True
def pool_hidden_state(self, last_hidden_state):
"""
Pool the output vectors into a single mean vector
"""
last_hidden_state = last_hidden_state[0]
mean_last_hidden_state = torch.mean(last_hidden_state, 1)
return mean_last_hidden_state
# model = XLNetForMultiLabelSequenceClassification(num_labels=len(Y_train[0]))
# model = torch.nn.DataParallel(model)
# model.cuda()
cwd = os.getcwd()
model_save_path = output_model_file = os.path.join(cwd, "/content/drive/My Drive/xlnet_toxic.bin")
model, start_epoch, lowest_eval_loss, train_loss_hist, valid_loss_hist = load_model(model_save_path)
def generate_predictions(model, df, num_labels, device="cpu", batch_size=32):
num_iter = math.ceil(df.shape[0]/batch_size)
pred_probs = np.array([]).reshape(0, num_labels)
model.to(device)
model.eval()
for i in range(num_iter):
df_subset = df.iloc[i*batch_size:(i+1)*batch_size,:]
X = df_subset["features"].values.tolist()
masks = df_subset["masks"].values.tolist()
X = torch.tensor(X)
masks = torch.tensor(masks, dtype=torch.long)
X = X.to(device)
masks = masks.to(device)
with torch.no_grad():
logits = model(input_ids=X, attention_mask=masks)
logits = logits.sigmoid().detach().cpu().numpy()
pred_probs = np.vstack([pred_probs, logits])
return pred_probs
```
## Load Meeting Transcript
```
import pandas as pd
df=pd.read_csv('/content/sample_data/transcript.txt',names=["sent"],sep='/:')
df.to_csv('someFileName.csv')
df_list=df.values.tolist()
df.head()
import nltk
nltk.download('punkt')
from nltk.tokenize import sent_tokenize
new_list=[]
for i in df_list:
j=str(i).strip("[]'")
indx = j.find(":")#position of 'I'
intro = j[indx+1:]
new_list.append(intro)
# re.sub(r'[a-zA-Z]+[:]', 'I', stri)
# flat_list = [item for sublist in new_list for item in sublist]
new_list
import numpy as np
test_np=np.array(new_list)
test_np
# create input id tokens
test_np_input_ids = tokenize_inputs(test_np, tokenizer, num_embeddings=250)
test_np_input_ids
# create attention masks
test_np_attention_masks = create_attn_masks(test_np_input_ids)
test_np_attention_masks
import pandas as pd
dataset_unseen = pd.DataFrame()
dataset_unseen['Sent'] = test_np.tolist()
dataset_unseen.shape
dataset_unseen["features"] = test_np_input_ids.tolist()
dataset_unseen["masks"] = test_np_attention_masks
dataset_unseen1=dataset_unseen[["Sent","features","masks"]]
dataset_unseen1.head()
label_cols = ["label_ami_da_1","label_ami_da_11","label_ami_da_12","label_ami_da_13","label_ami_da_14","label_ami_da_15","label_ami_da_16","label_ami_da_2","label_ami_da_3","label_ami_da_4","label_ami_da_5","label_ami_da_6","label_ami_da_7","label_ami_da_8","label_ami_da_9"]
num_labels = len(label_cols)
pred_probs = generate_predictions(model, dataset_unseen1, num_labels, device="cuda", batch_size=32)
pred_probs
# label_cols = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
label_cols = ["label_ami_da_1","label_ami_da_11","label_ami_da_12","label_ami_da_13","label_ami_da_14","label_ami_da_15","label_ami_da_16","label_ami_da_2","label_ami_da_3","label_ami_da_4","label_ami_da_5","label_ami_da_6","label_ami_da_7","label_ami_da_8","label_ami_da_9"]
dataset_unseen1["label_ami_da_1"] = pred_probs[:,0]
dataset_unseen1["label_ami_da_11"] = pred_probs[:,1]
dataset_unseen1["label_ami_da_12"] = pred_probs[:,2]
dataset_unseen1["label_ami_da_13"] = pred_probs[:,3]
dataset_unseen1["label_ami_da_14"] = pred_probs[:,4]
dataset_unseen1["label_ami_da_15"] = pred_probs[:,5]
dataset_unseen1["label_ami_da_16"] = pred_probs[:,6]
dataset_unseen1["label_ami_da_2"] = pred_probs[:,7]
dataset_unseen1["label_ami_da_3"] = pred_probs[:,8]
dataset_unseen1["label_ami_da_4"] = pred_probs[:,9]
dataset_unseen1["label_ami_da_5"] = pred_probs[:,10]
dataset_unseen1["label_ami_da_6"] = pred_probs[:,11]
dataset_unseen1["label_ami_da_7"] = pred_probs[:,12]
dataset_unseen1["label_ami_da_8"] = pred_probs[:,13]
dataset_unseen1["label_ami_da_9"] = pred_probs[:,14]
dataset_unseen1['HighScore'] = dataset_unseen1.max(axis=1)
dataset_unseen1
Dict_label = {"label_ami_da_1":"Backchannel","label_ami_da_11":"Elicit-Assessment","label_ami_da_12":"Comment-About-Understanding","label_ami_da_13":"Elicit-Comment-Understanding","label_ami_da_14":"Be-Positive","label_ami_da_15":"Be-Negative","label_ami_da_16":"Other","label_ami_da_2":"Stall","label_ami_da_3":"Fragment","label_ami_da_4":"Inform","label_ami_da_5":"Elicit-Inform","label_ami_da_6":"Suggest","label_ami_da_7":"Offer","label_ami_da_8":"Elicit-Offer-Or-Suggestion","label_ami_da_9":"Assess"}
new_lst=[]
new_lst_label_name=[]
d=dataset_unseen1.columns[3:-1]
l=len(dataset_unseen1)
for i in range(l):
for col in d:
if dataset_unseen1[col][i] == dataset_unseen1["HighScore"][i]:
new_lst.append(col)
for k, v in Dict_label.items():
if k == col:
new_lst_label_name.append(v)
dataset_unseen1["Label"]=new_lst
dataset_unseen1["Label_name"]=new_lst_label_name
dataset_unseen1[["Sent","HighScore","Label","Label_name"]].to_csv("output_action_itmes")
```
## Trying to implement Abstractive Dialogue Summarization
```
from transformers import PegasusForConditionalGeneration, PegasusTokenizer
import torch
model_name = 'google/pegasus-xsum'
torch_device = 'cuda' if torch.cuda.is_available() else 'cpu'
tokenizer = PegasusTokenizer.from_pretrained(model_name)
model = PegasusForConditionalGeneration.from_pretrained(model_name).to(torch_device)
# batch = tokenizer.prepare_seq2seq_batch(src_text, truncation=True, padding='longest', return_tensors="pt").to(torch_device)
# translated = model.generate(**batch)
# tgt_text = tokenizer.batch_decode(translated, skip_special_tokens=True)
# assert tgt_text[0] == "California's largest electricity provider has turned off power to hundreds of thousands of customers."
src_text = [
""" we have completed four modules and started the fifth module and quality and testing also happening in prallel.
we have to give demo to the user on first week of next month, so be ready. please start the frontend work and complete it by end of the day. """
]
batch = tokenizer.prepare_seq2seq_batch(src_text, truncation=True, padding='longest', return_tensors="pt").to(torch_device)
translated = model.generate(**batch)
tgt_text = tokenizer.batch_decode(translated, skip_special_tokens=True)
tgt_text
```
| github_jupyter |
# Exercise 5.06: Visualising K-NN Classification
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.neighbors import KNeighborsClassifier as KNN
df = pd.read_csv('../Datasets/breast-cancer-data.csv')
df.head()
```
We need to replace the species strings with indices to allow plotting of the prediction boundaries
```
labelled_diagnoses = [
'benign',
'malignant',
]
for idx, label in enumerate(labelled_diagnoses):
df.diagnosis = df.diagnosis.replace(label, idx)
df.head()
model = KNN(n_neighbors=3)
model.fit(X=df[['mean radius', 'worst radius']], y=df.diagnosis)
```
We need to construct the prediction space to allow for the visualisation of class allocations, to do this we will create a matrix or mesh of *mean radius*, *worst radius* coordinates that is slightly greater and less than the corresponding maximum and minimum values for each. Creating a range of values for *mean radius* and *worst radius* with a spacing of 0.1.
```
spacing = 0.1
mean_radius_range = np.arange(df['mean radius'].min() - 1, df['mean radius'].max() + 1, spacing)
worst_radius_range = np.arange(df['worst radius'].min() - 1, df['worst radius'].max() + 1, spacing)
```
Mix the two ranges into a mesh
```
xx, yy = np.meshgrid(mean_radius_range, worst_radius_range) # Create the mesh
xx
yy
pred_x = np.c_[xx.ravel(), yy.ravel()] # Concatenate the results
pred_x
pred_y = model.predict(pred_x).reshape(xx.shape)
pred_y
# Create color maps
cmap_light = ListedColormap(['#6FF6A5', '#F6A56F',])
cmap_bold = ListedColormap(['#0EE664', '#E6640E',])
markers = {
'benign': {'marker': 'o', 'facecolor': 'g', 'edgecolor': 'g'},
'malignant': {'marker': 'x', 'facecolor': 'r', 'edgecolor': 'r'},
}
plt.figure(figsize=(10, 7))
for name, group in df.groupby('diagnosis'):
diagnoses = labelled_diagnoses[name]
plt.scatter(group['mean radius'], group['worst radius'],
c=cmap_bold.colors[name],
label=labelled_diagnoses[name],
marker=markers[diagnoses]['marker']
)
plt.title('Breast Cancer Diagnosis Classification Mean Radius vs Worst Radius');
plt.xlabel('Mean Radius');
plt.ylabel('Worst Radius');
plt.legend();
plt.figure(figsize=(10, 7))
plt.pcolormesh(xx, yy, pred_y, cmap=cmap_light);
plt.scatter(df['mean radius'], df['worst radius'], c=df.diagnosis, cmap=cmap_bold, edgecolor='k', s=20);
plt.title('Breast Cancer Diagnosis Decision Boundaries Mean Radius vs Worst Radius');
plt.xlabel('Mean Radius');
plt.ylabel('Worst Radius');
plt.text(15, 12, 'Benign', ha='center',va='center', size=20,color='k');
plt.text(15, 30, 'Malignant', ha='center',va='center', size=20,color='k');
```
| github_jupyter |
# MeanHamilMinimizer native scipy
The purpose of this notebook is to describe a naive but pedagogical,
first baby step, in the implementation
of what is called in Qubiter the Mean Hamiltonian Minimization problem.
The qc history of this problem started with quantum chemists planning to
use on a qc the phase estimation algo invented by Kitaev? (an algo that
is also implemented in Qubiter) to estimate the energy levels (
eigenvalues) of simple molecules, initially H2. Then a bunch of people
realized, heck, rather than trying to estimate the eigenvalues of a
Hamiltonian by estimating the phase changes it causes, we can estimate
those eigenvalues more efficiently by estimating the mean value of that
Hamiltonian as measured empirically on a qc. Basically, just the
Rayleigh-Ritz method, one of the oldest tricks in the book. One of the
first papers to propose this mean idea is
https://arxiv.org/abs/1304.3061 Their algo is commonly referred to by
the ungainly name VQE (Variational Quantum Eigensolver) VQE was
originally applied to do quantum chemistry with a qc. But now Rigetti
and others have renamed it hybrid quantum-classical quantum computing
and pointed out that it's an algo that has wide applicability, not just
to quantum chemistry.
The idea behind hybrid quantum-classical is very simple. One has a
classical box CBox and a quantum box QBox. The gates of QBox depend on N
gate parameters. QBox sends info to CBox. CBox sends back to QBox N new
gate parameters that will lower some cost function. This feedback
process between CBox and QBox continues until the cost is minimized. The
cost function is the mean value of a Hamiltonian which is estimated
empirically from data obtained from the qc which resides inside the QBox.
To minimize a function of N continuous parameters, one can use some
methods like simulated annealing and Powell that do not require
calculating derivatives, or one can use methods that do use derivatives.
Another possible separation is between methods that don't care which
local minimum they find, as long as they find one of them, and those
methods that try to find the best local minimum of them all, the so
called global minimum. Yet another separation is between methods that
allow constraints and those that don't.
Among the methods that do use derivatives, the so called gradient based
methods only use the 1st derivative, whereas other methods use both
first (Jacobian) and second (Hessian) derivatives. The performance of
those that use both 1st and 2nd derivatives degrades quickly as N grows.
Besides, calculating 2nd derivatives is very expensive. Hence, methods
that use the 2nd derivatives are practically useless in the neural
network field where N is usually very large. In that field, gradient
based methods rule.
A method that uses no derivatives is Powell. A gradient based method
that is designed to have a fast convergence rate is the Conjugate
Gradient (CG) method. Another gradient based method is back-propagation
(BP). BP can be implemented as distributed computing much more easily
than other gradient based methods so it is favored by the most popular
computer programs for doing distributed AI, such as PyTorch and
Tensorflow.
Qubiter can perform minimization using various minlibs (minimization
software libraries) such as 'scipy', 'autograd', 'pytorch', 'tflow'. It
can also use various devices (aka simulators or backends), either
virtual or real, to do the minimization.
Non-scipy minlibs implement backprop.
The 'scipy' minlib is a wrapper for the scipy function
`scipy.optimize.minimize`. This scipy umbrella method implements many
minimization methods, including Powell and CG.
https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html
By a native device, we mean one that uses Qubiter native simulators like
SEO_simulator.
So, without further ado, here is an example of the use of
class `MeanHamilMinimizer` with a scipy minlib and native device.
$\newcommand{\bra}[1]{\left\langle{#1}\right|}$
$\newcommand{\ket}[1]{\left|{#1}\right\rangle}$
test: $\bra{\psi}M\ket{\phi}$
First change the directory to the Qubiter directory and add it to the path environment variable
```
import os
import sys
print(os.getcwd())
os.chdir('../../')
print(os.getcwd())
sys.path.insert(0,os.getcwd())
```
Next we construct a simple 4 qubit circuit that depends on two placeholder variables
`#1` and `#2`. These are the continuous variables that we will vary
to minimize a cost function. The cost function will be specified later.
```
from qubiter.adv_applications.MeanHamil_native import *
from qubiter.adv_applications.MeanHamilMinimizer import *
num_bits = 4
file_prefix = 'mean_hamil_native_test'
emb = CktEmbedder(num_bits, num_bits)
wr = SEO_writer(file_prefix, emb)
wr.write_Rx(2, rads=np.pi/7)
wr.write_Rx(1, rads='#2*.5')
wr.write_Rn(3, rads_list=['#1', '-#1*3', '#2'])
wr.write_Rx(1, rads='-my_fun#2#1')
wr.write_cnot(2, 3)
wr.close_files()
```
The code above wrote inside Qubiter's `io_folder`, two files,
an English file and a Picture file. We next ask the writer object
wr to print those two files for us
```
wr.print_eng_file()
wr.print_pic_file()
```
The circuit depends on a placeholder function called `my_fun`.
This function will remain fixed during the
minimization process but it must be defined
and it must be passed in inside an input dictionary to
the constructor of class `MeanHamil`
```
def my_fun(x, y):
return x + .5*y
fun_name_to_fun = {'my_fun': my_fun}
```
One must also pass into the constructor of `MeanHamil`, the value of a Hamiltonian called `hamil`.
Qubiter stores `hamil` in an object of the class `QubitOperator`
from the open source Python library `OpenFermion`.
Note that the `QubitOperator`
constructor simplifies `hamil` automatically.
Hamiltonians are Hermitian, so after `QubitOperator`
finishes simplifying `hamil`, the coefficient of every "term"
of `hamil` must be real.
```
hamil = QubitOperator('X1 Y3 X1 Y1', .4) + QubitOperator('Y2 X1', .7)
print('hamil=\n', hamil)
```
So what is the purpose of this Hamiltonian `hamil`?
The cost function to be minimized is defined as the mean value of `hamil`.
More precisely, if $\ket{\psi}$ is the ouput of the circuit
specified above, then the cost function equals $\bra{\psi} | H | \ket{\psi}$.
The initial values for parameters `#1` and `#2` to be minimized
must also be passed in inside an input dictionary to
the constructor of class `MeanHamilMinimizer`. Variable `all_var_nums` contains the keys
of that dictionary.
> Note: Results are very heavily dependent on initial x_val, probably because,
due to the periodic nature of qubit rotations, there are many local minima
```
init_var_num_to_rads = {1: 1., 2: 3.}
all_var_nums = init_var_num_to_rads.keys()
```
For convenience, we define a function `case()` that creates an
object of `MeanHamilMinimizer`, asks that object to minimize the cost function,
and then `case()` returns the final result of that minimization process.
As advertised at the beginning of this notebook, we
use various sub-methods of `scipy.optimize.minimize`
```
num_samples = 0
print_hiatus = 25
verbose = False
np.random.seed(1234)
emp_mhamil = MeanHamil_native(file_prefix, num_bits, hamil,
all_var_nums, fun_name_to_fun, simulator_name='SEO_simulator', num_samples=num_samples)
targ_mhamil = MeanHamil_native(file_prefix, num_bits, hamil,
all_var_nums, fun_name_to_fun, simulator_name='SEO_simulator') # zero samples
def case(**kwargs):
return MeanHamilMinimizer(emp_mhamil, targ_mhamil,
all_var_nums, init_var_num_to_rads,
print_hiatus, verbose).find_min(minlib='scipy', **kwargs)
```
We focus on two cases, num_samples=0 and num_samples > 0.
When `MeanHamil` is told that
num_samples=0, it does no sampling. It just calculates
the mean value of `hamil` "exactly', using the exact
final state vector calculated by Qubiter's `SEO_simulator` class
```
min_method = 'Powell'
emp_mhamil.num_samples = 0
case(method=min_method)
```
When num_samples > 0, Qubiter calculates the final state vector
of the circuit, then it samples that num_samples times,
obtains an empirical probability distribution from that,
and then it calculates the mean value of `hamil`
using that empirical distribution
```
min_method = 'Powell'
emp_mhamil.num_samples = 1000
case(method=min_method)
# very sensitive to eps, seems to be hitting discontinuity
min_method='CG'
num_sample= 0
case(options={'disp': True, 'eps':1e-2})
```
| github_jupyter |
# Creating a loader with Kosh
In this example we will create a custom loader for some ASCII representation
The structure is
* Headers at the begining of the file starting with #
* *#* varname vs axis
* indicates new variable with name `varname`
* var_value axis_value
* repeated n times
* end
* marks end of the current variable
We will assume the user already has functions to read the data in. These can be found in the [some_user_io_functions.py](some_user_io_functions.py) file.
The function to read data in is called `load_variable_from_file`, the one to list the features in the file is called `get_variable_names`.
```
import os
import kosh
# Make sure local file is new sql file
kosh_example_sql_file = "my_store.sql"
# Create db on file
store = kosh.create_new_db(kosh_example_sql_file)
# Add dataset to the store
sample = store.create(name="example", metadata={'project':"example"})
# Associate file with datasets
sample.associate("example.ultra", mime_type="custom")
```
Let's create our `CustomLoader` inheriting from `kosh.KoshLoader`
For this we will need to:
* register the types we can read at the class level (not in `__init__`) and the format each type can be exported as.
* types : { "custom" : ["numpy",] }
* **IMPORTANT** the keys in this dictionary is what Kosh uses to tie the loader to a **mime_type**
* implement the `extract` function to read data in
* desired feature is in `self.feature`
* potential keywords are in: `self._user_passed_parameters[1]`
* kosh object describing the source is in `self.obj` (can query its attributes if desired)
* source uri is at: `self.obj.uri`
* The function to read a variable from a file is: `load_variable_from_file`
* implement the `list_features(self)` function, using the `get_variable_names` helper function.
* optionally implement the `describe_feature(self, feature)`
```
import sys, os
sys.path.append(".")
from some_user_io_functions import load_variable_from_file, get_variable_names
```
Let's query the function documentation
```
load_variable_from_file?
from kosh import KoshLoader
import numpy
class CustomLoader(KoshLoader):
types ={"custom": ["numpy", ]} # keys ties things back to mime_type in associate function
def extract(self, *args, **kargs):
return load_variable_from_file(self.obj.uri, self.feature)
def list_features(self):
return get_variable_names(self.obj.uri)
def describe_feature(self, feature):
var = load_variable_from_file(self.obj.uri, feature)
info = {"name": feature, "size": var.shape}
return info
```
At this point we need to register/[add](https://lc.llnl.gov/kosh/autoapi/kosh/core/index.html#kosh.core.KoshStoreClass.add_loader) our loader with the store (let's save it in the store as well).
```
store.add_loader(CustomLoader)
```
We can now [query](https://lc.llnl.gov/kosh/autoapi/kosh/core/index.html#kosh.core.KoshDataset.list_features) our dataset, as explained in the [previous](04_Associating_and_Reading_Data.ipynb) notebook.
```
print(sample.list_features())
sample.describe_feature("energy")
```
Or extract its features
```
print(sample.get("energy"))
```
| github_jupyter |
```
import cvxpy as cp
import numpy as np
import matplotlib.pyplot as plt
```
# <p style="font-family:arvo">1. Basic LP problem
<p style="font-family:arvo">
$A$ is $mxn$ matrix and $b$ is $mx1$ matrix.
Our objective function is to
<center>
Minimize: $(AX - b)^2$
</center>
subject to the constraints
<center>
$0 \leq X \leq 1$
</center>
Solve for $X$
```
# Problem data.
m = 30
n = 20
np.random.seed(1)
A = np.random.randn(m, n)
b = np.random.randn(m)
```
### <p style="font-family:arvo">We need to solve for $x$
```
x = cp.Variable(n)
```
### <p style="font-family:arvo">Minimizing the objective
```
objective = cp.Minimize(cp.sum_squares(A*x - b))
```
### <p style="font-family:arvo">Defining constraints
```
constraints = [x >= 0, x <= 1]
prob = cp.Problem(objective, constraints)
```
### <p style="font-family:arvo">The optimal objective is returned by prob.solve()
```
result = prob.solve()
print("Solution is:", prob.status)
print("Minimum Objective value is:", result)
```
### <p style="font-family:arvo">The optimal value for x is stored in x.value
```
print(x.value)
```
# <p style="font-family:arvo">2. Knapsack Problem
<p style="font-family:arvo">
<center>
$\max$ $\sum_{i}$ $x_{i} * c_{i}$
</center>
<br>
<b>subject to constraints-</b>
<br>
<center>
$\sum_{i}$ $x_{i} * w_{i}$ $\leq W$ where $W$ is maximum weight
<br>
$x_{i} \in$ {0,1}
</center>
</p>
### <p style="font-family:arvo">$W$ = 165 units
```
totalWeight = 165
```
### <p style="font-family:arvo">Specifying Constraints
### <p style="font-family:arvo">weights : weights$_{i}$ denotes weight of $i^{th}$ item
```
weights = np.array([23, 31, 29, 44, 53, 38, 63, 85, 89, 82])
len(weights)
```
### <p style="font-family:arvo">cost: cost$_{i}$ denotes cost of $i^{th}$ item
```
cost = np.array([92, 57, 49, 68, 60, 43, 67, 84, 87, 72])
len(cost)
```
### <p style="font-family:arvo"> take: take$_{i}$ denotes whether we are taking $i^{th}$ item or not, boolean variable
```
take = cp.Variable(len(weights),boolean=True)
take
```
### <p style="font-family:arvo">Weight Constraint: Total weight in bag <= totalWeight
```
weight_constraint = [weights * take <= totalWeight]
```
### <p style="font-family:arvo">Total cost for all the items present in the bag, need to maximize total_cost
```
total_cost = cost * take
```
### <p style="font-family:arvo">Defining problem in cvxpy
```
knapsack_problem = cp.Problem(cp.Maximize(total_cost), weight_constraint)
```
### <p style="font-family:arvo">Using cvxpy to solve the problem and also mentioning solver for Integer Programming Problems, return maximum value of the objective
```
knapsack_problem.solve(solver=cp.GLPK_MI)
```
### <p style="font-family:arvo">take$_{i}$ = 1, represents that we need to take $i^{th}$ item, otherwise don't take the item
```
print(take.value)
```
# <p style="font-family:arvo">3. Line Fitting Problem
<br>
Our objective function is -
<center>
$\min$ $\sum_{i} (y_{i} - (m * x_{i} + c))^2$
</center>
### <p style="font-family:arvo">This is an unconstrained optimization problem
### <p style="font-family:arvo">The number of data points are 50
```
np.random.seed(0)
num = 50
x = np.arange(num)
y = 2*x+4 + 5*np.random.randn(num,)
plt.figure(figsize=(10,10))
plt.scatter(x,y)
plt.show()
```
### <p style="font-family:arvo">The variables $m$ and $c$ are defined
```
m = cp.Variable()
c = cp.Variable()
```
### <p style="font-family:arvo">Objective function is posed as problem
```
objective = cp.Minimize(cp.sum((y - (m*x+c))**2))
prob = cp.Problem(objective)
result = prob.solve()
print("Optimal value is: ",result)
print("The values of m is {} and c is {}".format(m.value, c.value))
plt.figure(figsize=(10,10))
plt.scatter(x,y)
plt.plot(x, m.value * x + c.value, 'r')
plt.show()
```
# <p style="font-family:arvo">4. Min vertex cover
<p style="font-family:arvo">The objective function is defined as -
<center>
$\min \sum_{u} x_{u} , \forall u \in V$
</center>
<p style="font-family:arvo">subject to constraints -
<center>
$x_{u} + x_{v} \geq 1, \forall u, v \in E$
<br>
$x_{u} \in $ {0,1}
</center>

### <p style="font-family:arvo">The above graph has 6 vertices and 7 edges.
```
e = 7
v = 6
edges = [(1,2), (1,3), (2,3), (2,4), (3,5), (4,5), (4,6)]
```
### <p style="font-family:arvo">The variable $x$ , objective function and constraints are defined
```
x = cp.Variable(v, boolean=True)
objective = cp.Minimize(cp.sum(x))
constraints = [x[i-1]+x[j-1]>=1 for (i,j) in edges]
```
### <p style="font-family:arvo">Using cvxpy solving the problem
```
prob = cp.Problem(objective, constraints)
result = prob.solve(solver=cp.GLPK_MI)
print("The minimum value of the objective function is :", result)
print("The minimum vertex cover is :", np.sum(x.value == 1.0))
```
### <p style="font-family:arvo">The values in $x$ which correspond to 1 are included in minimum vertex cover
```
x.value
```
| github_jupyter |
# Burgers Optimization with a Physics-Informed NN
To illustrate how the physics-informed losses work, let's consider a reconstruction task
as an inverse problem example.
We'll use Burgers equation $\frac{\partial u}{\partial{t}} + u \nabla u = \nu \nabla \cdot \nabla u$ as a simple yet non-linear equation in 1D, for which we have a series of _observations_ at time $t=0.5$.
The solution should fulfill the residual formulation for Burgers equation and match the observations.
In addition, let's impose Dirichlet boundary conditions $u=0$
at the sides of our computational domain, and define the solution in
the time interval $t \in [0,1]$.
Note that similar to the previous forward simulation example,
we will still be sampling the solution with 128 points ($n=128$), but now we have a discretization via the NN. So we could also sample points in between without having to explicitly choose a basis function for interpolation. The discretization via the NN now internally determines how to use its degrees of freedom to arrange the activation functions as basis functions. So we have no direct control over the reconstruction.
[[run in colab]](https://colab.research.google.com/github/tum-pbs/pbdl-book/blob/main/physicalloss-code.ipynb)
## Formulation
In terms of the $x,y^*$ notation from {doc}`overview-equations` and the previous section, this reconstruction problem means we are solving
$$
\text{arg min}_{\theta} \sum_i ( f(x_i ; \theta)-y^*_i )^2 + R(x_i) ,
$$
where $x$ and $y^*$ are solutions of $u$ at different locations in space and time. As we're dealing with a 1D velocity, $x,y^* \in \mathbb{R}$.
They both represent two-dimensional solutions
$x(p_i,t_i)$ and $y^*(p_i,t_i)$ for a spatial coordinate $p_i$ and a time $t_i$, where the index $i$ sums over a set of chosen $p_i,t_i$ locations at which we evaluate the PDE and the approximated solutions. Thus $y^*$ denotes a reference $u$ for $\mathcal{P}$ being Burgers equation, which $x$ should approximate as closely as possible. Thus our neural network representation of $x$ will receive $p,t$ as input to produce a velocity solution at the specified position.
The residual function $R$ above collects additional evaluations of $f(;\theta)$ and its derivatives to formulate the residual for $\mathcal{P}$. This approach -- using derivatives of a neural network to compute a PDE residual -- is typically called a _physics-informed_ approach, yielding a _physics-informed neural network_ (PINN) to represent a solution for the inverse reconstruction problem.
Thus, in the formulation above, $R$ should simply converge to zero above. We've omitted scaling factors in the objective function for simplicity. Note that, effectively, we're only dealing with individual point samples of a single solution $u$ for $\mathcal{P}$ here.
## Preliminaries
Let's just load phiflow with the tensorflow backend for now, and initialize the random sampling. (_Note: this example uses an older version 1.5.1 of phiflow._)
```
!pip install --quiet phiflow==1.5.1
from phi.tf.flow import *
import numpy as np
#rnd = TF_BACKEND # for phiflow: sample different points in the domain each iteration
rnd = math.choose_backend(1) # use same random points for all iterations
```
We're importing phiflow here, but we won't use it to compute a solution to the PDE as in {doc}`overview-burgers-forw`. Instead, we'll use the derivatives of an NN (as explained in the previous section) to set up a loss formulation for training.
Next, we set up a simple NN with 8 fully connected layers and `tanh` activations with 20 units each.
We'll also define the `boundary_tx` function which gives an array of constraints for the solution (all for $=0.5$ in this example), and the `open_boundary` function which stores constraints for $x= \pm1$ being 0.
```
def network(x, t):
""" Dense neural network with 8 hidden layers and 3021 parameters in total.
Parameters will only be allocated once (auto reuse).
"""
y = math.stack([x, t], axis=-1)
for i in range(8):
y = tf.layers.dense(y, 20, activation=tf.math.tanh, name='layer%d' % i, reuse=tf.AUTO_REUSE)
return tf.layers.dense(y, 1, activation=None, name='layer_out', reuse=tf.AUTO_REUSE)
def boundary_tx(N):
x = np.linspace(-1,1,128)
# precomputed solution from forward simulation:
u = np.asarray( [0.008612174447657694, 0.02584669669548606, 0.043136357266407785, 0.060491074685516746, 0.07793926183951633, 0.0954779141740818, 0.11311894389663882, 0.1308497114054023, 0.14867023658641343, 0.1665634396808965, 0.18452263429574314, 0.20253084411376132, 0.22057828799835133, 0.23865132431365316, 0.25673879161339097, 0.27483167307082423, 0.2929182325574904, 0.3109944766354339, 0.3290477753208284, 0.34707880794585116, 0.36507311960102307, 0.38303584302507954, 0.40094962955534186, 0.4188235294008765, 0.4366357052408043, 0.45439856841363885, 0.4720845505219581, 0.4897081943759776, 0.5072391070000235, 0.5247011051514834, 0.542067187709797, 0.5593576751669057, 0.5765465453632126, 0.5936507311857876, 0.6106452944663003, 0.6275435911624945, 0.6443221318186165, 0.6609900633731869, 0.67752574922899, 0.6939334022562877, 0.7101938106059631, 0.7263049537163667, 0.7422506131457406, 0.7580207366534812, 0.7736033721649875, 0.7889776974379873, 0.8041371279965555, 0.8190465276590387, 0.8337064887158392, 0.8480617965162781, 0.8621229412131242, 0.8758057344502199, 0.8891341984763013, 0.9019806505391214, 0.9143881632159129, 0.9261597966464793, 0.9373647624856912, 0.9476871303793314, 0.9572273019669029, 0.9654367940878237, 0.9724097482283165, 0.9767381835635638, 0.9669484658390122, 0.659083299684951, -0.659083180712816, -0.9669485121167052, -0.9767382069792288, -0.9724097635533602, -0.9654367970450167, -0.9572273263645859, -0.9476871280825523, -0.9373647681120841, -0.9261598056102645, -0.9143881718456056, -0.9019807055316369, -0.8891341634240081, -0.8758057205293912, -0.8621229450911845, -0.8480618138204272, -0.833706571569058, -0.8190466131476127, -0.8041372124868691, -0.7889777195422356, -0.7736033858767385, -0.758020740007683, -0.7422507481169578, -0.7263049162371344, -0.7101938950789042, -0.6939334061553678, -0.677525822052029, -0.6609901538934517, -0.6443222327338847, -0.6275436932970322, -0.6106454472814152, -0.5936507836778451, -0.5765466491708988, -0.5593578078967361, -0.5420672759411125, -0.5247011730988912, -0.5072391580614087, -0.4897082914472909, -0.47208460952428394, -0.4543985995006753, -0.4366355580500639, -0.41882350871539187, -0.40094955631843376, -0.38303594105786365, -0.36507302109186685, -0.3470786936847069, -0.3290476440540586, -0.31099441589505206, -0.2929180880304103, -0.27483158663081614, -0.2567388003912687, -0.2386513127155433, -0.22057831776499126, -0.20253089403524566, -0.18452269630486776, -0.1665634500729787, -0.14867027528284874, -0.13084990929476334, -0.1131191325854089, -0.09547794429803691, -0.07793928430794522, -0.06049114408297565, -0.0431364527809777, -0.025846763281087953, -0.00861212501518312] );
t = np.asarray(rnd.ones_like(x)) * 0.5
perm = np.random.permutation(128)
return (x[perm])[0:N], (t[perm])[0:N], (u[perm])[0:N]
def _ALT_t0(N): # alternative, impose original initial state at t=0
x = rnd.random_uniform([N], -1, 1)
t = rnd.zeros_like(x)
u = - math.sin(np.pi * x)
return x, t, u
def open_boundary(N):
t = rnd.random_uniform([N], 0, 1)
x = math.concat([math.zeros([N//2]) + 1, math.zeros([N//2]) - 1], axis=0)
u = math.zeros([N])
return x, t, u
```
Most importantly, we can now also construct the residual loss function `f` that we'd like to minimize in order to guide the NN to retrieve a solution for our model equation. As can be seen in the equation at the top, we need derivatives w.r.t. $t$, $x$ and a second derivative for $x$. The first three lines of `f` below do just that.
Afterwards, we simply combine the derivates to form Burgers equation. Here we make use of phiflow's `gradient` function:
```
def f(u, x, t):
""" Physics-based loss function with Burgers equation """
u_t = gradients(u, t)
u_x = gradients(u, x)
u_xx = gradients(u_x, x)
return u_t + u*u_x - (0.01 / np.pi) * u_xx
```
Next, let's set up the sampling points in the inner domain, such that we can compare the solution with the previous forward simulation in phiflow.
The next cell allocates two tensors: `grid_x` will cover the size of our domain, i.e., the -1 to 1 range, with 128 cells, while `grid_t` will sample the time interval $[0,1]$ with 33 time stamps.
The last `math.expand_dims()` call simply adds another `batch` dimension, so that the resulting tensor is compatible with the following examples.
```
N=128
grids_xt = np.meshgrid(np.linspace(-1, 1, N), np.linspace(0, 1, 33), indexing='ij')
grid_x, grid_t = [tf.convert_to_tensor(t, tf.float32) for t in grids_xt]
# create 4D tensor with batch and channel dimensions in addition to space and time
# in this case gives shape=(1, N, 33, 1)
grid_u = math.expand_dims(network(grid_x, grid_t))
```
Now, `grid_u` contains a full graph to evaluate our NN at $128 \times 33$ positions, and returns the results in a $[1,128,33,1]$ array once we run it through `session.run`. Let's give this a try: we can initialize a TF session, evaluate `grid_u` and show it in an image, just like the phiflow solution we computed previously.
(Note, we'll use the `show_state` as in {doc}`overview-burgers-forw`. Hence, the x axis does not show actual simulation time, but is showing 32 steps "blown" up by a factor of 16 to make the changes over time easier to see in the image.)
```
import pylab as plt
print("Size of grid_u: "+format(grid_u.shape))
session = Session(None)
session.initialize_variables()
def show_state(a, title):
for i in range(4): a = np.concatenate( [a,a] , axis=3)
a = np.reshape( a, [a.shape[1],a.shape[2]*a.shape[3]] )
fig, axes = plt.subplots(1, 1, figsize=(16, 5))
im = axes.imshow(a, origin='upper', cmap='inferno')
plt.colorbar(im) ; plt.xlabel('time'); plt.ylabel('x'); plt.title(title)
print("Randomly initialized network state:")
show_state(session.run(grid_u),"Uninitialized NN")
```
This visualization already shows a smooth transition over space and time. So far, this is purely the random initialization of the NN that we're sampling here. So it has nothing to do with a solution of our PDE-based model up to now.
The next steps will actually evaluate the constraints in terms of data (from the `boundary` functions), and the model constraints from `f` to retrieve an actual solution to the PDE.
## Loss function and training
As objective for the learning process we can now combine the _direct_ constraints, i.e., the solution at $t=0.5$ and the Dirichlet $u=0$ boundary conditions with the loss from the PDE residuals. For both boundary constraints we'll use 100 points below, and then sample the solution in the inner region with an additional 1000 points.
The direct constraints are evaluated via `network(x, t)[:, 0] - u`, where `x` and `t` are the space-time location where we'd like to sample the solution, and `u` provides the corresponding ground truth value.
For the physical loss points, we have no ground truth solutions, but we'll only evaluate the PDE residual via the NN derivatives, to see whether the solution satisfies the PDE model. If not, this directly gives us an error to be reduced via an update step in the optimization. The corresponding expression is of the form `f(network(x, t)[:, 0], x, t)` below. Note that for both data and physics terms the `network()[:, 0]` expressions don't remove any data from the $L^2$ evaluation, they simply discard the last size-1 dimension of the $(n,1)$ tensor returned by the network.
```
# Boundary loss
N_SAMPLE_POINTS_BND = 100
x_bc, t_bc, u_bc = [math.concat([v_t0, v_x], axis=0) for v_t0, v_x in zip(boundary_tx(N_SAMPLE_POINTS_BND), open_boundary(N_SAMPLE_POINTS_BND))]
x_bc, t_bc, u_bc = np.asarray(x_bc,dtype=np.float32), np.asarray(t_bc,dtype=np.float32) ,np.asarray(u_bc,dtype=np.float32)
#with app.model_scope():
loss_u = math.l2_loss(network(x_bc, t_bc)[:, 0] - u_bc) # normalizes by first dimension, N_bc
# Physics loss inside of domain
N_SAMPLE_POINTS_INNER = 1000
x_ph, t_ph = tf.convert_to_tensor(rnd.random_uniform([N_SAMPLE_POINTS_INNER], -1, 1)), tf.convert_to_tensor(rnd.random_uniform([N_SAMPLE_POINTS_INNER], 0, 1))
loss_ph = math.l2_loss(f(network(x_ph, t_ph)[:, 0], x_ph, t_ph)) # normalizes by first dimension, N_ph
# Combine
ph_factor = 1.
loss = loss_u + ph_factor * loss_ph # allows us to control the relative influence of loss_ph
optim = tf.train.GradientDescentOptimizer(learning_rate=0.02).minimize(loss)
#optim = tf.train.AdamOptimizer(learning_rate=0.001).minimize(loss) # alternative, but not much benefit here
```
The code above just initializes the evaluation of the loss, we still didn't do any optimization steps, but we're finally in a good position to get started with this.
Despite the simple equation, the convergence is typically very slow. The iterations themselves are fast to compute, but this setup needs a _lot_ of iterations. To keep the runtime in a reasonable range, we only do 10k iterations by default below (`ITERS`). You can increase this value to get better results.
```
session.initialize_variables()
import time
start = time.time()
ITERS = 10000
for optim_step in range(ITERS+1):
_, loss_value = session.run([optim, loss])
if optim_step<3 or optim_step%1000==0:
print('Step %d, loss: %f' % (optim_step,loss_value))
#show_state(grid_u)
end = time.time()
print("Runtime {:.2f}s".format(end-start))
```
The training can take a significant amount of time, around 2 minutes on a typical notebook, but at least the error goes down significantly (roughly from around 0.2 to ca. 0.03), and the network seems to successfully converge to a solution.
Let's show the reconstruction of the network, by evaluating the network at the centers of a regular grid, so that we can show the solution as an image. Note that this is actually fairly expensive, we have to run through the whole network with a few thousand weights for all of the $128 \times 32$ sampling points in the grid.
It looks pretty good on first sight, though. There's been a very noticeable change compared to the random initialization shown above:
```
show_state(session.run(grid_u),"After Training")
```
---
## Evaluation
Let's compare solution in a bit more detail. Here are the actual sample points used for constraining the solution (at time step 16, $t=1/2$) shown in gray, versus the reconstructed solution in blue:
```
u = session.run(grid_u)
# solution is imposed at t=1/2 , which is 16 in the array
BC_TX = 16
uT = u[0,:,BC_TX,0]
fig = plt.figure().gca()
fig.plot(np.linspace(-1,1,len(uT)), uT, lw=2, color='blue', label="NN")
fig.scatter(x_bc[0:100], u_bc[0:100], color='gray', label="Reference")
plt.title("Comparison at t=1/2")
plt.xlabel('x'); plt.ylabel('u'); plt.legend()
```
Not too bad at the sides of the domain (the Dirichlet boundary conditions $u=0$ are fulfilled), but the shock in the center (at $x=0$) is not well represented.
Let's check how well the initial state at $t=0$ was reconstructed. That's the most interesting, and toughest part of the problem (the rest basically follows from the model equation and boundary conditions, given the first state).
It turns out that the accuracy of the initial state is actually not that good: the blue curve from the PINN is quite far away from the constraints via the reference data (shown in gray)... The solution will get better with larger number of iterations, but it requires a surprisingly large number of iterations for this fairly simple case.
```
# ground truth solution at t0
t0gt = np.asarray( [ [-math.sin(np.pi * x) * 1.] for x in np.linspace(-1,1,N)] )
velP0 = u[0,:,0,0]
fig = plt.figure().gca()
fig.plot(np.linspace(-1,1,len(velP0)), velP0, lw=2, color='blue', label="NN")
fig.plot(np.linspace(-1,1,len(t0gt)), t0gt, lw=2, color='gray', label="Reference")
plt.title("Comparison at t=0")
plt.xlabel('x'); plt.ylabel('u'); plt.legend()
```
Especially the maximum / minimum at $x=\pm 1/2$ are far off, and the boundaries at $x=\pm 1$ are not fulfilled: the solution is not at zero.
We have the forward simulator for this simulation, so we can use the $t=0$ solution of the network to
evaluate how well the temporal evaluation was reconstructed. This measures how well the temporal evolution of the model equation was captured via the soft constraints of the PINN loss.
The graph below shows the initial state in blue, and two evolved states at $t=8/32$ and $t=15/32$. Note that this is all from the simulated version, we'll show the learned version next.
(Note: The code segments below also have some optional code to show the states at `[STEPS//4]`. It's commented out by default, you can uncomment or add additional ones to visualize more of the time evolution if you like.)
```
# re-simulate with phiflow from solution at t=0
DT = 1./32.
STEPS = 32-BC_TX # depends on where BCs were imposed
INITIAL = u[...,BC_TX:(BC_TX+1),0] # np.reshape(u0, [1,len(u0),1])
print(INITIAL.shape)
DOMAIN = Domain([N], boundaries=PERIODIC, box=box[-1:1])
state = [BurgersVelocity(DOMAIN, velocity=INITIAL, viscosity=0.01/np.pi)]
physics = Burgers()
for i in range(STEPS):
state.append( physics.step(state[-1],dt=DT) )
# we only need "velocity.data" from each phiflow state
vel_resim = [x.velocity.data for x in state]
fig = plt.figure().gca()
pltx = np.linspace(-1,1,len(vel_resim[0].flatten()))
fig.plot(pltx, vel_resim[ 0].flatten(), lw=2, color='blue', label="t=0")
#fig.plot(pltx, vel_resim[STEPS//4].flatten(), lw=2, color='green', label="t=0.125")
fig.plot(pltx, vel_resim[STEPS//2].flatten(), lw=2, color='cyan', label="t=0.25")
fig.plot(pltx, vel_resim[STEPS-1].flatten(), lw=2, color='purple',label="t=0.5")
#fig.plot(pltx, t0gt, lw=2, color='gray', label="t=0 Reference") # optionally show GT, compare to blue
plt.title("Resimulated u from solution at t=0")
plt.xlabel('x'); plt.ylabel('u'); plt.legend()
```
And here is the PINN output from `u` at the same time steps:
```
velP = [u[0,:,x,0] for x in range(33)]
print(velP[0].shape)
fig = plt.figure().gca()
fig.plot(pltx, velP[BC_TX+ 0].flatten(), lw=2, color='blue', label="t=0")
#fig.plot(pltx, velP[BC_TX+STEPS//4].flatten(), lw=2, color='green', label="t=0.125")
fig.plot(pltx, velP[BC_TX+STEPS//2].flatten(), lw=2, color='cyan', label="t=0.25")
fig.plot(pltx, velP[BC_TX+STEPS-1].flatten(), lw=2, color='purple',label="t=0.5")
plt.title("NN Output")
plt.xlabel('x'); plt.ylabel('u'); plt.legend()
```
Judging via eyeball norm, these two versions of $u$ look quite similar, but not surprisingly the errors grow over time and there are significant differences. Especially the steepening of the solution near the shock at $x=0$ is not "captured" well. It's a bit difficult to see in these two graphs, though, let's quantify the error and show the actual difference:
```
error = np.sum( np.abs( np.asarray(vel_resim[0:16]).flatten() - np.asarray(velP[BC_TX:BC_TX+STEPS]).flatten() )) / (STEPS*N)
print("Mean absolute error for re-simulation across {} steps: {:7.5f}".format(STEPS,error))
fig = plt.figure().gca()
fig.plot(pltx, (vel_resim[0 ].flatten()-velP[BC_TX ].flatten()), lw=2, color='blue', label="t=5")
fig.plot(pltx, (vel_resim[STEPS//4].flatten()-velP[BC_TX+STEPS//4].flatten()), lw=2, color='green', label="t=0.625")
fig.plot(pltx, (vel_resim[STEPS//2].flatten()-velP[BC_TX+STEPS//2].flatten()), lw=2, color='cyan', label="t=0.75")
fig.plot(pltx, (vel_resim[STEPS-1 ].flatten()-velP[BC_TX+STEPS-1 ].flatten()), lw=2, color='purple',label="t=1")
plt.title("u Error")
plt.xlabel('x'); plt.ylabel('MAE'); plt.legend()
```
The code above will compute a mean absolute error of ca. $1.5 \cdot 10^{-2}$ between ground truth re-simulation and the PINN evolution, which is significant for the value range of the simulation.
And for comparison with the forward simulation and following cases, here are also all steps over time with a color map.
```
# show re-simulated solution again as full image over time
sn = np.concatenate(vel_resim, axis=-1)
sn = np.reshape(sn, list(sn.shape)+[1] ) # print(sn.shape)
show_state(sn,"Re-simulated u")
```
Next, we'll store the full solution over the course of the $t=0 \dots 1$ time interval, so that we can compare it later on to the full solution from a regular forward solve and compare it to the differential physics solution.
Thus, stay tuned for the full evaluation and the comparison. This will follow in {doc}`diffphys-code-burgers`, after we've discussed the details of how to run the differential physics optimization.
```
vels = session.run(grid_u) # special for showing NN results, run through TF
vels = np.reshape( vels, [vels.shape[1],vels.shape[2]] )
# save for comparison with other methods
import os; os.makedirs("./temp",exist_ok=True)
np.savez_compressed("./temp/burgers-pinn-solution.npz",vels) ; print("Vels array shape: "+format(vels.shape))
```
---
## Next steps
This setup is just a starting point for PINNs and physical soft-constraints, of course. The parameters of the setup were chosen to run relatively quickly. As we'll show in the next sections, the behavior of such an inverse solve can be improved substantially by a tighter integration of solver and learning.
The solution of the PINN setup above can also directly be improved, however. E.g., try to:
* Adjust parameters of the training to further decrease the error without making the solution diverge.
* Adapt the NN architecture for further improvements (keep track of the weight count, though).
* Activate a different optimizer, and observe the change in behavior (this typically requires adjusting the learning rate). Note that the more complex optimizers don't necessarily do better in this relatively simple example.
* Or modify the setup to make the test case more interesting: e.g., move the boundary conditions to a later point in simulation time, to give the reconstruction a larger time interval to reconstruct.
| github_jupyter |
```
import torch
from torch import nn, optim
from torch.utils.data import (Dataset, DataLoader, TensorDataset)
import tqdm
import re
import collections
import itertools
from statistics import mean
remove_marks_regex = re.compile("[\,\(\)\[\]\*:;¿¡]|<.*?>")
shift_marks_regex = re.compile("([?!\.])")
unk = 0
sos = 1
eos = 2
def normalize(text):
text = text.lower()
text = remove_marks_regex.sub("", text)
text = shift_marks_regex.sub(r" \1", text)
return text
def parse_line(line):
line = normalize(line.strip())
# src - target 각각의 토큰을 리스트화
src, trg = line.split("\t")
src_tokens = src.strip().split()
trg_tokens = trg.strip().split()
return src_tokens, trg_tokens
def build_vocab(tokens):
# 모든 무장에서 토큰 등장 횟수 확인
counts = collections.Counter(tokens)
sorted_counts = sorted(counts.items(), key=lambda c: c[1], reverse=True)
word_list = ["<UNK>", "<SOS>", "<EOS>"] + [x[0] for x in sorted_counts]
word_dict = dict((w, i) for i, w in enumerate(word_list))
return word_list, word_dict
def words2tensor(words, word_dict, max_len, padding=0):
# 종료 태그
words = words + ["<EOS>"]
words = [word_dict.get(w, 0) for w in words]
seq_len = len(words)
if seq_len < max_len + 1:
words = words + [padding] * (max_len + 1 - seq_len)
return torch.tensor(words, dtype=torch.int64), seq_len
class TranslationPairDataset(Dataset):
def __init__(self, path, max_len=15):
def filter_pair(p):
# 단어수가 많은 문장 제거
return not (len(p[0]) > max_len or len(p[1]) > max_len)
with open(path, encoding='utf8') as fp:
pairs = map(parse_line, fp)
pairs = filter(filter_pair, pairs)
pairs = list(pairs)
src = [p[0] for p in pairs]
trg = [p[1] for p in pairs]
self.src_word_list, self.src_word_dict = build_vocab(itertools.chain.from_iterable(src))
self.trg_word_list, self.trg_word_dict = build_vocab(itertools.chain.from_iterable(trg))
self.src_data = [words2tensor(words, self.src_word_dict, max_len) for words in src]
self.trg_data= [words2tensor(words, self.trg_word_dict, max_len, -100) for words in trg]
def __len__(self):
return len(self.src_data)
def __getitem__(self, idx):
src, lsrc = self.src_data[idx]
trg, ltrg = self.trg_data[idx]
return src, lsrc, trg, ltrg
batch_size = 64
max_len = 10
path = "d:/dataset/spa-eng/spa.txt"
ds = TranslationPairDataset(path, max_len=max_len)
loader = DataLoader(ds, batch_size=batch_size, shuffle=True, num_workers=0)
class Encoder(nn.Module):
def __init__(self, num_embeddings, embedding_dim=50, hidden_size=50, num_layers=1, dropout=0.2):
super().__init__()
self.emb = nn.Embedding(num_embeddings, embedding_dim=embedding_dim, padding_idx=0)
self.lstm = nn.LSTM(embedding_dim, hidden_size, num_layers, batch_first=True, dropout=dropout)
def forward(self, x, h0=None, l=None):
x = self.emb(x)
if l is not None:
x = nn.utils.rnn.pack_padded_sequence(x, l, batch_first=True)
_, h = self.lstm(x, h0)
return h
class Decoder(nn.Module):
def __init__(self, num_embeddings, embedding_dim=50, hidden_size=50, num_layers=1, dropout=0.2):
super().__init__()
self.emb = nn.Embedding(num_embeddings, embedding_dim, padding_idx=0)
self.lstm = nn.LSTM(embedding_dim, hidden_size, num_layers, batch_first=True, dropout=dropout)
self.linear = nn.Linear(hidden_size, num_embeddings)
def forward(self, x, h, l=None):
x = self.emb(x)
if l is not None:
x = nn.utils.rnn.pack_padded_sequence(x, l, batch_first=True)
x, h = self.lstm(x, h)
if l is not None:
x = nn.utils.rnn.pad_packed_sequence(x, batch_first=True, padding_value=0)[0]
x = self.linear(x)
return x, h
def translate(input_str, enc, dec, max_len=15, device="cpu"):
# 입력 문자열을 수치화해서 Tensor로 변환
words = normalize(input_str).split()
input_tensor, seq_len = words2tensor(words,
ds.src_word_dict, max_len=max_len)
input_tensor = input_tensor.unsqueeze(0)
# 엔코더에서 사용하므로 입력값의 길이도 리스트로 만들어둔다
seq_len = [seq_len]
# 시작 토큰 준비
sos_inputs = torch.tensor(sos, dtype=torch.int64)
input_tensor = input_tensor.to(device)
sos_inputs = sos_inputs.to(device)
# 입력 문자열을 엔코더에 넣어서 컨텍스트 얻기
ctx = enc(input_tensor, l=seq_len)
# 시작 토큰과 컨텍스트를 디코더의 초깃값으로 설정
z = sos_inputs
h = ctx
results = []
for i in range(max_len):
# Decoder로 다음 단어 예측
o, h = dec(z.view(1, 1), h)
# 선형 계층의 출력이 가장 큰 위치가 다음 단어의 ID
wi = o.detach().view(-1).max(0)[1]
if wi.item() == eos:
break
results.append(wi.item())
# 다음 입력값으로 현재 출력 ID를 사용
z = wi
# 기록해둔 출력 ID를 문자열로 변환
return " ".join(ds.trg_word_list[i] for i in results)
enc = Encoder(len(ds.src_word_list), 100, 100, 2)
dec = Decoder(len(ds.trg_word_list), 100, 100, 2)
translate("I am a student.", enc, dec)
enc.to("cuda:0")
dec.to("cuda:0")
opt_enc = optim.Adam(enc.parameters(), 0.002)
opt_dec = optim.Adam(dec.parameters(), 0.002)
loss_f = nn.CrossEntropyLoss()
def to2D(x):
shapes = x.shape
return x.reshape(shapes[0] * shapes[1], -1)
for epoch in range(30):
enc.train(), dec.train()
losses = []
for x, lx, y, ly in tqdm.tqdm(loader):
# x packed sequence를 위해 소스 길이로 내림차순 정렬
lx, sort_idx = lx.sort(descending=True)
x, y, ly = x[sort_idx], y[sort_idx], ly[sort_idx]
x, y = x.to("cuda:0"), y.to("cuda:0")
ctx = enc(x, l=lx)
ly, sort_idx = ly.sort(descending=True)
y = y[sort_idx]
h0 = (ctx[0][:, sort_idx, :], ctx[1][:, sort_idx, :])
z = y[:, :-1].detach()
z[z==-100] = 0
o, _ = dec(z, h0, l=ly-1)
loss = loss_f(to2D(o[:]), to2D(y[:, 1:max(ly)]).squeeze())
enc.zero_grad(), dec.zero_grad()
loss.backward()
opt_enc.step(), opt_dec.step()
losses.append(loss.item())
enc.eval(), dec.eval()
print(epoch, mean(losses))
with torch.no_grad():
print(translate("I am a student.", enc, dec, max_len=max_len, device="cuda:0"))
print(translate("He likes to eat pizza.", enc, dec, max_len=max_len, device="cuda:0"))
print(translate("She is my mother.", enc, dec, max_len=max_len, device="cuda:0"))
```
| github_jupyter |
```
import numpy as np
from scipy.spatial.distance import pdist, cdist,squareform
from scipy.stats import pearsonr
import pandas as pd
import matplotlib.pyplot as plt
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit.Chem import DataStructs
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
from sklearn.kernel_ridge import KernelRidge
from sklearn.ensemble import GradientBoostingRegressor, RandomForestRegressor
from sklearn.linear_model import *
# ECFP6
def FPBase64ToNumpy(fps):
X = []
for item in fps:
bv = DataStructs.ExplicitBitVect(4096)
DataStructs.ExplicitBitVect.FromBase64(bv, item)
arr = np.zeros( (1,) )
DataStructs.ConvertToNumpyArray( bv, arr )
X.append(arr)
return X
train_ecfp = pd.read_csv("../data/train.csv")
test_ecfp = pd.read_csv("../data/holdout.csv")
peptide_ecfp = pd.read_csv("../data/peptide.csv")
trainX_ecfp = FPBase64ToNumpy(train_ecfp.ECFP6)
trainY_ecfp = train_ecfp.ConfEntropy
testX_ecfp = FPBase64ToNumpy(test_ecfp.ECFP6)
testY_ecfp = test_ecfp.ConfEntropy
peptideX_ecfp = FPBase64ToNumpy(peptide_ecfp.ECFP6)
peptideY_ecfp = peptide_ecfp.ConfEntropy
Lasso_ecfp = LassoCV(cv=5,alphas=(0.01, 0.05, 0.1,0.5, 1.0, 5.0, 10.0)).fit(trainX_ecfp, trainY_ecfp)
Ridge_ecfp = RidgeCV(scoring='neg_mean_absolute_error',cv=5,alphas=(0.01, 0.05, 0.1,0.5, 1.0, 5.0, 10.0)).fit(trainX_ecfp, trainY_ecfp)
# Prediction
lassotest_ecfp = Lasso_ecfp.predict(testX_ecfp)
lassopeptide_ecfp = Lasso_ecfp.predict(peptideX_ecfp)
ridgetest_ecfp = Ridge_ecfp.predict(testX_ecfp)
ridgepeptide_ecfp = Ridge_ecfp.predict(peptideX_ecfp)
print("LASSO MAE (J/mol K)")
print("ZINC, Peptide")
print(mean_absolute_error(testY_ecfp,lassotest_ecfp),mean_absolute_error(peptideY_ecfp,lassopeptide_ecfp))
print(r"Pearson $R^{2}$")
print("ZINC, Peptide")
print(np.square(pearsonr(testY_ecfp,lassotest_ecfp)[0]), np.square(pearsonr(peptideY_ecfp,lassopeptide_ecfp)[0]))
print("Ridge MAE (J/mol K)")
print("ZINC, Peptide")
print(mean_absolute_error(testY_ecfp,ridgetest_ecfp),mean_absolute_error(peptideY_ecfp, ridgepeptide_ecfp))
print(r"Pearson $R^{2}$")
print("ZINC, Peptide")
print(np.square(pearsonr(testY_ecfp,ridgetest_ecfp)[0]), np.square(pearsonr(peptideY_ecfp, ridgepeptide_ecfp)[0]))
# CDDD
train_cddd = pd.read_csv("../data/train_cddd.zip",compression='zip')
test_cddd = pd.read_csv("../data/zinc_test_cddd.zip",compression='zip')
peptide_cddd = pd.read_csv("../data/peptide_cddd.zip",compression='zip')
trainX_cddd = train_cddd.iloc[:,2:]
trainY_cddd = train_cddd["ConfEntropy"]
testX_cddd = test_cddd.iloc[:,2:]
testY_cddd = test_cddd["ConfEntropy"]
pepX_cddd = peptide_cddd.iloc[:,2:]
pepY_cddd = peptide_cddd["ConfEntropy"]
# LASSO and Ridge
Lasso_cddd = LassoCV(cv=5,alphas=(0.01, 0.05, 0.1,0.5, 1.0, 5.0, 10.0)).fit(trainX_cddd, trainY_cddd)
Ridge_cddd = RidgeCV(scoring='neg_mean_absolute_error',cv=5,alphas=(0.01, 0.05, 0.1,0.5, 1.0, 5.0, 10.0)).fit(trainX_cddd, trainY_cddd)
# Prediction
lassotest_cddd = Lasso_cddd.predict(testX_cddd)
lassopeptide_cddd = Lasso_cddd.predict(pepX_cddd)
ridgetest_cddd = Ridge_cddd.predict(testX_cddd)
ridgepeptide_cddd = Ridge_cddd.predict(pepX_cddd)
print("LASSO MAE (J/mol K)")
print("ZINC, Peptide")
print(mean_absolute_error(testY_cddd,lassotest_cddd),mean_absolute_error(pepY_cddd,lassopeptide_cddd))
print("Pearson R^{2}")
print("ZINC, Peptide")
print(np.square(pearsonr(testY_cddd,lassotest_cddd)[0]), np.square(pearsonr(pepY_cddd,lassopeptide_cddd)[0]))
print("Ridge MAE (J/mol K)")
print("ZINC, Peptide")
print(mean_absolute_error(testY_cddd,ridgetest_cddd),mean_absolute_error(pepY_cddd, ridgepeptide_cddd))
print("Pearson R^{2}")
print("ZINC, Peptide")
print(np.square(pearsonr(testY_cddd,ridgetest_cddd)[0]), np.square(pearsonr(pepY_cddd, ridgepeptide_cddd)[0]))
```
| github_jupyter |
# Welcome to fastai
```
from fastai.vision import *
from fastai.gen_doc.nbdoc import *
from fastai.core import *
from fastai.basic_train import *
```
The fastai library simplifies training fast and accurate neural nets using modern best practices. It's based on research in to deep learning best practices undertaken at [fast.ai](http://www.fast.ai), including "out of the box" support for [`vision`](/vision.html#vision), [`text`](/text.html#text), [`tabular`](/tabular.html#tabular), and [`collab`](/collab.html#collab) (collaborative filtering) models. If you're looking for the source code, head over to the [fastai repo](https://github.com/fastai/fastai) on GitHub. For brief examples, see the [examples](https://github.com/fastai/fastai/tree/master/examples) folder; detailed examples are provided in the full documentation (see the sidebar). For example, here's how to train an MNIST model using [resnet18](https://arxiv.org/abs/1512.03385) (from the [vision example](https://github.com/fastai/fastai/blob/master/examples/vision.ipynb)):
```
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = create_cnn(data, models.resnet18, metrics=accuracy)
learn.fit(1)
jekyll_note("""This documentation is all built from notebooks;
that means that you can try any of the code you see in any notebook yourself!
You'll find the notebooks in the <a href="https://github.com/fastai/fastai/tree/master/docs_src">docs_src</a> folder of the
<a href="https://github.com/fastai/fastai">fastai</a> repo. For instance,
<a href="https://nbviewer.jupyter.org/github/fastai/fastai/blob/master/docs_src/index.ipynb">here</a>
is the notebook source of what you're reading now.""")
```
## Installation and updating
To install or update fastai, we recommend `conda`:
```
conda install -c pytorch -c fastai fastai pytorch
```
For troubleshooting, and alternative installations (including pip and CPU-only options) see the [fastai readme](https://github.com/fastai/fastai/blob/master/README.md).
## Reading the docs
To get started quickly, click *Applications* on the sidebar, and then choose the application you're interested in. That will take you to a walk-through of training a model of that type. You can then either explore the various links from there, or dive more deeply into the various fastai modules.
We've provided below a quick summary of the key modules in this library. For details on each one, use the sidebar to find the module you're interested in. Each module includes an overview and example of how to use it, along with documentation for every class, function, and method. API documentation looks, for example, like this:
### An example function
```
show_doc(rotate, full_name='rotate')
```
---
Types for each parameter, and the return type, are displayed following standard Python [type hint syntax](https://www.python.org/dev/peps/pep-0484/). Sometimes for compound types we use [type variables](/fastai_typing.html). Types that are defined by fastai or Pytorch link directly to more information about that type; try clicking *Image* in the function above for an example. The docstring for the symbol is shown immediately after the signature, along with a link to the source code for the symbol in GitHub. After the basic signature and docstring you'll find examples and additional details (not shown in this example). As you'll see at the top of the page, all symbols documented like this also appear in the table of contents.
For inherited classes and some types of decorated function, the base class or decorator type will also be shown at the end of the signature, delimited by `::`. For `vision.transforms`, the random number generator used for data augmentation is shown instead of the type, for randomly generated parameters.
## Module structure
### Imports
fastai is designed to support both interactive computing as well as traditional software development. For interactive computing, where convenience and speed of experimentation is a priority, data scientists often prefer to grab all the symbols they need, with `import *`. Therefore, fastai is designed to support this approach, without compromising on maintainability and understanding.
In order to do so, the module dependencies are carefully managed (see next section), with each exporting a carefully chosen set of symbols when using `import *`. In general, for interactive computing, you'll want to import from both `fastai`, and from one of the *applications*, such as:
```
from fastai.vision import *
```
That will give you all the standard external modules you'll need, in their customary namespaces (e.g. `pandas as pd`, `numpy as np`, `matplotlib.pyplot as plt`), plus the core fastai libraries. In addition, the main classes and functions for your application ([`fastai.vision`](/vision.html#vision), in this case), e.g. creating a [`DataBunch`](/basic_data.html#DataBunch) from an image folder and training a convolutional neural network (with [`create_cnn`](/vision.learner.html#create_cnn)), are also imported. If you don't wish to import any application, but want all the main functionality from fastai, use `from fastai.basics import *`. Of course, you can also just import the specific symbols that you require, without using `import *`.
If you wish to see where a symbol is imported from, either just type the symbol name (in a REPL such as Jupyter Notebook or IPython), or (in most editors) wave your mouse over the symbol to see the definition. For instance:
```
Learner
```
### Dependencies
At the base of everything are the two modules [`core`](/core.html#core) and [`torch_core`](/torch_core.html#torch_core) (we're not including the `fastai.` prefix when naming modules in these docs). They define the basic functions we use in the library; [`core`](/core.html#core) only relies on general modules, whereas [`torch_core`](/torch_core.html#torch_core) requires pytorch. Most type-hinting shortcuts are defined there too (at least the one that don't depend on fastai classes defined later). Nearly all modules below import [`torch_core`](/torch_core.html#torch_core).
Then, there are three modules directly on top of [`torch_core`](/torch_core.html#torch_core):
- [`data`](/vision.data.html#vision.data), which contains the class that will take a [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) or pytorch [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) to wrap it in a [`DeviceDataLoader`](/basic_data.html#DeviceDataLoader) (a class that sits on top of a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) and is in charge of putting the data on the right device as well as applying transforms such as normalization) and regroup then in a [`DataBunch`](/basic_data.html#DataBunch).
- [`layers`](/layers.html#layers), which contains basic functions to define custom layers or groups of layers
- [`metrics`](/metrics.html#metrics), which contains all the metrics
This takes care of the basics, then we regroup a model with some data in a [`Learner`](/basic_train.html#Learner) object to take care of training. More specifically:
- [`callback`](/callback.html#callback) (depends on [`data`](/vision.data.html#vision.data)) defines the basis of callbacks and the [`CallbackHandler`](/callback.html#CallbackHandler). Those are functions that will be called every step of the way of the training loop and can allow us to customize what is happening there;
- [`basic_train`](/basic_train.html#basic_train) (depends on [`callback`](/callback.html#callback)) defines [`Learner`](/basic_train.html#Learner) and [`Recorder`](/basic_train.html#Recorder) (which is a callback that records training stats) and has the training loop;
- [`callbacks`](/callbacks.html#callbacks) (depends on [`basic_train`](/basic_train.html#basic_train)) is a submodule defining various callbacks, such as for mixed precision training or 1cycle annealing;
- `learn` (depends on [`callbacks`](/callbacks.html#callbacks)) defines helper functions to invoke the callbacks more easily.
From [`data`](/vision.data.html#vision.data) we can split on one of the four main *applications*, which each has their own module: [`vision`](/vision.html#vision), [`text`](/text.html#text) [`collab`](/collab.html#collab), or [`tabular`](/tabular.html#tabular). Each of those submodules is built in the same way with:
- a submodule named <code>transform</code> that handles the transformations of our data (data augmentation for computer vision, numericalizing and tokenizing for text and preprocessing for tabular)
- a submodule named <code>data</code> that contains the class that will create datasets specific to this application and the helper functions to create [`DataBunch`](/basic_data.html#DataBunch) objects.
- a submodule named <code>models</code> that contains the models specific to this application.
- optionally, a submodule named <code>learn</code> that will contain [`Learner`](/basic_train.html#Learner) speficic to the application.
Here is a graph of the key module dependencies:

| github_jupyter |
# CardIO framework for deep research of ECG
#### CardIO is based on a very simple and natural approach:
* Split input dataset into batches to process data batch by batch (because input datasets can be indefinitely large)
* Describe workflow as a sequence of actions (e.g., load data — preprocess data — train model)
* Run the workflow for the whole dataset
#### In this notebook you will learn:
* Where to get ECG data
* How to start using CardIO
* How to apply actions to batch
* How different actions transform batch and its components
## Table of contents
* [Getting ECG data](#Getting-ECG-data)
* [Indexing of ECGs](#Indexing-of-ECGs)
* [Initialization of dataset](#Initialization-of-dataset)
* [Generating batches](#Generating-batches)
* [Apply actions](#Apply-actions)
* [Actions in CardIO](#Actions-in-CardIO)
* [convolve_signals](#convolve_signals)
* [band_pass_signals](#band_pass_signals)
* [random_split_signals](#random_split_signals)
* [unstack_signals](#unstack_signals)
* [resample_signals](#resample_signals)
* [rfft](#rfft)
* [spectrogram](#spectrogram)
* [cwt](#cwt)
* [apply_transform](#apply_transform)
* [drop_labels](#drop_labels)
* [rename_labels](#rename_labels)
* [rename_channels](#rename_channels)
* [other actions](#other-actions)
* [Summary](#Summary)
## Getting ECG data
In this and following tutorials you will work with ECG data through the CardIO framework.
We suggest loading ECG data from the 2017 PhysioNet/CinC Challenge database of short single lead ECG recording.
The PhysioNet archive contains 8.528 ECGs in [wfdb](https://www.physionet.org/physiotools/wpg/wpg_35.htm) format. Each ECG has a unique index ranging from "A00001" to "A08528". According to wbdf format, ECG record consists of several files with the same name but different extensions. We will work with files with ```.hea``` and ```.mat``` extensions. File ```.mat``` contains signal, while ```.hea``` contains meta information about the signal (e.g., sample rate).
***
### Important note on downloading
First way to get the data is to follow this [link](https://physionet.org/challenge/2017/) and download archive \"training2017.zip\" or use the [direct](https://physionet.org/challenge/2017/training2017.zip) link.
Since the release of the challenge wfdb library has changed and now complies with the data standard more strictly. Latest versions of the library can not read the data from this archive because of some discrepancies with the standard. To follow this course of tutorials you can use pn2017_data_to_wfdb_format.py script, which formats the data properly:
```
>>> python3 pn2017_data_to_wfdb.py -p path_to_data
```
Another way to get the data is to use `dl_database` function from wfdb library. This function downloads the data from this [database](https://physionet.org/physiobank/database/challenge/2017/training/). It may take some time, and the data will be located in several subfolders.
```python
import wfdb
wfdb.io.dl_database(db_dir='challenge/2017/training/',
dl_dir='/notebooks/data/ECG/training2017')
```
If you choose this way, you do not need to process the data with the script mentioned above because records in this database are already formatted properly.
***
Since the PhysioNet archive was prepared for arrhythmia classification challenge, it also contains REFERENCE.csv file, where each ECG index is labeled with one of four classes:
* Normal rhythm
* AF
* Other rhythms
* Noise
Read [here](https://physionet.org/challenge/2017/) more about PhysioNet database.
However, if you do not want to load large dataset now, you can find several ECGs from that database in folder cardio/tests/data (this folder is included in CardIO repository).
## Indexing of ECGs
Working with ECG begins with ```FilesIndex```. ```FilesIndex``` contains index and location of each ECG record we want to process.
Let all ECGs be stored in wfdb format in a folder with path '../cardio/tests/data/' (if you cloned the CardIO repository from [Git](https://github.com/analysiscenter/cardio), this path will contain several examples of ECGs indeed). Let's create a new ```FilesIndex``` with all ECGs from this folder.
```
import sys
import numpy as np
from matplotlib import pyplot as plt
sys.path.append('..')
import cardio.cardio.batchflow as bf
index = bf.FilesIndex(path='c:/cardio/cardio/tests/data/A*.hea', no_ext=True, sort=True)
```
Now each ECG is indexed with its filename without extension, as it is defined by ```no_ext``` argument of ```FilesIndex```. Indices are stored in ```index.indices```:
```
print(index.indices)
```
## Initialization of Dataset
We have ```index``` that knows which ECGs we want to process. CardIO processes data grouped in batches of class [EcgBatch](https://analysiscenter.github.io/cardio/modiles/core.html). To generate proper batches we create a dataset:
```
from cardio.cardio import EcgBatch
eds = bf.Dataset(index, batch_class=EcgBatch)
```
Note that the same result can be obtained in a shorter way:
```python
from cardio import EcgDataset
eds = EcgDataset(path='../cardio/tests/data/*.hea', no_ext=True, sort=True)
```
## Generating batches
Let's generate a first batch of some size, say 6:
```
batch = eds.next_batch(batch_size=6, unique_labels=['A', 'N', 'O'])
```
Here we pass argument ```unique_labels```, which contains all possible labels that we expect to be in the dataset. We have ECG signals with three different labels that indicate whether the ECG has come from a person with normal rhythm (label "N"), a person with atrial fibrillation ("A"), or a person with some other abnormal rhythm ("O").
However, ```batch``` still does not contain any data, it contains only indices and paths to ECGs. To fill it with data we need to apply load action. Next section shows how to apply actions.
## Apply actions
Any preprocess typically begins with the loading of data. Therefore, we start with an example how to apply action [```load```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.load).
Note that paths to ECGs are already stored in the batch, so simply indicate data format, which is wfdb, and components of the batch we want to load. We load components ```signal``` and ```meta```:
```
batch_with_data = batch.load(fmt='wfdb', components=['signal', 'meta'])
```
Now ```batch_with_data``` contatains loaded ECGs. Any ECG record can be accessed by its index, e.g., ```batch_with_data['A00001']```. ECG components, signal and meta, can be accessed as ```batch_with_data['A00001'].signal``` and ```batch_with_data['A00001'].meta``` correspondingly.
We can also load labels for the data using the same action, but with other arguments:
```
batch_with_data = batch_with_data.load(src='c:/cardio/cardio/tests/data/REFERENCE.csv', fmt='csv', components='target')
```
Labels are stored in the ```target``` component. To access e.g. label of signal 'A00001' call ```batch_with_data['A00001'].target```:
```
batch_with_data['A00004'].target
```
Let's plot signal from ECG with index```'A00001'```:
```
batch_with_data.show_ecg('A00004')
```
Any other action can be applied to ```batch_with_data``` in the same way as ```load```. For example, consider action [```flip_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.flip_signals). It flips signals whose R-peaks are directed downwards.
Note that ```flip_signals``` modifies batch inplace, so we create copy of the batch:
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
And then apply ```flip_signals```:
```
changed_batch.flip_signals()
```
Now we can compare results. R-peaks of the signal with index 'A00013' were originally directed downwards:
```
original_batch.show_ecg('A00013')
changed_batch.show_ecg('A00013')
```
For further analysis we apply ```flip_signals()``` to the ```batch_with_data```:
```
batch_with_data = batch_with_data.flip_signals()
```
## Actions in CardIO
CardIO contains various actions for ECG processsing. Here are some of them:
* convolve_signals()
* band_pass_signals()
* random_split_signals()
* unstack_signals()
* resample_signals()
* rfft()
* spectrogram()
* cwt()
* apply_transform()
* drop_labels()
* rename_labels()
* rename_channels()
* ...
In the following sections we will show how to use them and how these actions affect batch and its components.
### [```convolve_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.convolve_signals)
```convolve_signals()``` action convolves signal with the specified kernel.
Convolution is a mathematical operation on two functions (or signals) that produces a third function. This is a common operation to remove noise from signals.
$$(f*g)[n] = \sum_{m=-\infty}^{\infty} f[m]g[n-m] = \sum_{m=-\infty}^{\infty} f[n-m]g[m]$$
Let's convolve signals in batch with gaussian kernel, which look like this:
```
%matplotlib inline
import cardio
kernel = cardio.kernels.gaussian(size=11)
plt.plot(kernel)
plt.grid("on")
plt.show()
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
Now we will add some normally distributed noise to the signal and apply ```convolve_signals()```:
```
siglen = original_batch["A00001"].signal.shape[1]
noise = np.random.normal(scale=0.01, size=siglen)
original_batch["A00001"].signal += noise
changed_batch["A00001"].signal += noise
changed_batch.convolve_signals(kernel=kernel)
```
And now you can see how this transformation affected the signal:
```
original_batch.show_ecg('A00001', start=10, end=15)
changed_batch.show_ecg('A00001', start=10, end=15)
```
### [```band_pass_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.band_pass_signals)
A bandpass filter passes frequencies within a certain range and rejects frequencies outside that range. To demonstrate capabilities of ```band_pass_signal()``` action we are going to artificially modify some of the batch's signals.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
First, generate some low-frequency noise:
```
n_sin = 10
noise = np.zeros_like(original_batch['A00013'].signal)
siglen = original_batch["A00013"].signal.shape[1]
t = np.linspace(0, 30, siglen)
for i in range(n_sin):
a = np.random.uniform(0, 0.1)
omega = np.random.uniform(0.1, 0.8)
phi = np.random.uniform(0, 2 * np.pi)
noise += a * np.sin(omega * t + phi)
fig = plt.figure(figsize=(12, 4))
plt.plot(noise[0])
plt.grid("on")
plt.show()
```
And add this noise to the signal "A00013":
```
original_batch['A00013'].signal += noise
changed_batch['A00013'].signal += noise
```
Now apply band-pass filter with low frequency:
```
changed_batch.band_pass_signals(low=0.2)
```
And here you can see that the noise was removed:
```
original_batch.show_ecg('A00013')
changed_batch.show_ecg('A00013')
```
### [```random_split_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.random_split_signals)
```random_split_signals()``` does simple transformation: it splits 2-D signal along axis 1 (typically time axis) into ```n_segments``` with random starting point and defined ```length```.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
changed_batch.random_split_signals(length=2048, n_segments=4)
```
We can now compare shapes of the initial signal and after the random split. Notice, that resulting array is 3-D: ```[n_segments, n_channels, time]```.
```
print("Original shape of the signal: ", original_batch['A00013'].signal.shape)
print("Shape of the signal after random split: ", changed_batch['A00013'].signal.shape)
```
### [```unstack_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.unstack_signals)
```unstack_signals()``` is useful to maintain batch structure after transformations that change shape of the signal component items, like ```random_split_signals()```.
This action creates a new batch in which each signal's item along axis 0 is considered as a separate signal.
Here we apply ```random_split_signals()``` to both batches and then apply ```unstack_signals()``` to the changed batch:
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
original_batch.random_split_signals(length=3000, n_segments=4)
changed_batch.random_split_signals(length=3000, n_segments=4).unstack_signals()
```
And now you can see that nothing happened:
```
print('Original shape of the signal: ', original_batch['A00013'].signal.shape)
print('Shape of the signal after actions: ', changed_batch['A00013'].signal.shape)
```
This happened because ```unstack_signals()``` doesn't change existing batch, but returns a new one instead.
Also, ```unstack_signals()``` changes the ```index``` of the batch. New ```indices``` will be integers from ```0``` to ```N-1```, where N is the sum of the sizes of the first dimension over all signals (3-D) in the batch.
For example, you have batch with two 3-D signals with shapes ```[3, 12, 1000]``` and ```[2, 12, 1000]```:
``` python
>>> batch.indices
['A00001', 'A00002']
>>> batch['A00001'].signal.shape
[3, 12, 1000]
>>> batch['A00002'].signal.shape
[2, 12, 1000]
```
If you apply ```unstack_signals()``` to the batch you would get a new batch with five 2-D signals and new ```indices```:
``` python
>>> batch = batch.unstack_signals()
>>> batch.indices
[0, 1, 2, 3, 4]
>>> batch[0].signal.shape
[12, 1000]
>>> batch[3].signal.shape
[12, 1000]
```
Let's use ```unstack_signals()``` on our data and see what happens:
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
original_batch.random_split_signals(length=3000, n_segments=4)
changed_batch = changed_batch.random_split_signals(length=3000, n_segments=4).unstack_signals()
```
At first, take a glance at the ```signal``` component of the batches:
```
print('Original shape of `signal` component: ', original_batch.signal.shape)
print('Shape of `signal` component after unstack: ', changed_batch.signal.shape)
```
Now we compare ```indices``` of the original and changed batches:
```
print('Indices of the original batch: ', original_batch.indices)
print('Indices of the batch after unstack: ', changed_batch.indices)
```
And, finally, look at the shapes of the signals:
```
print('Original shape of the signal: ', original_batch['A00001'].signal.shape)
print('Shape of the signal after unstacking: ', changed_batch[0].signal.shape)
```
### [```resample_signals```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.resample_signals)
```resample_signals()``` resamples 2-D signals along time axis (axis 1) to given sampling rate. This action also changes ```signal``` and ```meta``` components.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
changed_batch.resample_signals(fs=120)
```
If you set new sampling rate to a very low value, you can notice changes by eye:
```
original_batch.show_ecg('A00001')
changed_batch.show_ecg('A00001')
```
And also you can see that signal length has changed:
```
print('Original shape of the signal: ', original_batch['A00001'].signal.shape)
print('Shape of the signal after resampling: ', changed_batch['A00001'].signal.shape)
```
New sampling rate is stored in ```meta``` component under key "fs":
```
print('Original sampling rate: {} Hz'.format(original_batch['A00001'].meta['fs']))
print('Sampling rate after resampling: {} Hz'.format(changed_batch['A00001'].meta['fs']))
```
### [```rfft```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.rfft)
This action computes the one-dimensional discrete Fourier Transform (DFT) of a real-valued array by means of an efficient algorithm called the Fast Fourier Transform (FFT). FFT divides a signal into its frequency components. These components are single sinusoidal oscillations at distinct frequencies each with their own amplitude and phase.
```
changed_batch = batch_with_data.deepcopy()
```
Now we apply ```rfft()``` to the ```signal``` component and store the result to the atribute ```fft_data```. Note that action has a ```dst``` argument, which defines batch component (or atribute) where the result of the action will be stored.
```
changed_batch.rfft(src='signal', dst='fft_data')
sig_index = changed_batch.get_pos(None, 'signal', index='A00013')
changed_batch.fft_data
```
And the next step: calculate power spectrum and plot it. To calculate power spectrum we will square the absolute value of the ```rfft()``` result, which is stored in batch attribute defined by ```dst```.
```
sig_index = changed_batch.get_pos(None, 'signal', index='A00013')
power_spectrum = np.abs(changed_batch.fft_data[sig_index][0])**2
fig= plt.figure(figsize=(15, 4))
plt.plot(power_spectrum)
plt.grid(True)
plt.xlabel('Frequency')
plt.ylabel('Amplitude')
plt.title('Power spectrum')
plt.show()
```
### [```spectrogram```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.spectrogram)
A spectrogram is a visual representation of the spectrum of frequencies of signal as they vary with time.
We'll use the ```spectrogram()``` action on ```changed_batch```.
```
changed_batch = batch_with_data.deepcopy()
WIN_SIZE = 16
changed_batch.spectrogram(nperseg=WIN_SIZE, window="hamming", dst='spectrogram_data')
```
Note that ```spectrogram()``` action calculates a spectrogram for each channel, so resulting array for one 2-D signal will have three dimensions: ```[n_channels, frequency, time]```.
```
channel = 0
siglen = changed_batch["A00013"].signal.shape[1]
sig_index = changed_batch.get_pos(None, 'signal', index='A00013')
freqs = np.fft.rfftfreq(WIN_SIZE, 1 / changed_batch.meta[sig_index]["fs"])
_, (ax1, ax2) = plt.subplots(nrows=2, figsize=(15, 8))
ax1.pcolormesh(changed_batch.spectrogram_data[sig_index][channel])
ax1.set_xticks([])
ax1.set_yticklabels(freqs)
ax1.set_ylabel("Frequency (Hz)")
ax1.set_title("Spectrogram")
ax2.plot(changed_batch["A00013"].signal[channel])
ax2.set_xlim(0, siglen)
ax2.set_xlabel("Time")
ax2.set_ylabel("Amplitude")
ax2.set_title("Signal A00013")
ax2.grid(True)
plt.show()
```
### [```cwt```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.cwt)
A continuous wavelet transform (CWT) is used to divide a continuous-time function into wavelets. Unlike Fourier transform, the continuous wavelet transform possesses the ability to construct a time-frequency representation of a signal that offers very good time and frequency localization.
```
changed_batch = batch_with_data.deepcopy()
```
We now apply ```cwt()``` with mexican hat wavelet (also known as mexh) and scales from 1 to 30 to the batch:
```
min_scale = 1
max_scale = 30
scales = np.arange(min_scale, max_scale+1)
changed_batch.cwt(scales=scales, wavelet='mexh', src='signal', dst='wavelet')
```
Note that result of ```cwt()``` action is a 3-D array: ```[n_channels, n_scales, time]```.
```
sig_index = changed_batch.get_pos(None, 'signal', index='A00013')
changed_batch.wavelet[sig_index].shape
```
And here is a visualisation of the result:
```
channel = 0
siglen = changed_batch["A00013"].signal.shape[1]
_, (ax1, ax2) = plt.subplots(nrows=2, figsize=(15, 8))
ax1.pcolormesh(changed_batch.wavelet[sig_index][channel])
ax1.set_xticks([])
ax1.set_ylabel("Scales")
ax1.set_title("Wavelets")
ax2.plot(changed_batch["A00013"].signal[channel])
ax2.set_ylabel("Amplitude")
ax2.set_title("Signal A00013")
ax2.set_xlim(0, siglen)
ax2.set_xlabel("Time")
ax2.grid(True)
plt.show()
```
### [```apply_transform```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.apply_transform)
This action applies function given in action's argument to each item in the batch
For a source ```src``` and a destination ```dst``` ```apply_transform()``` does the following:
```python
for item in range(len(batch)):
batch.dst[item] = func(batch.src[item], *args, **kwargs)
```
```src``` and ```dst``` should be a component or an attribute of a batch.
The main benefit of the ```apply_transform``` action is that it runs in parallel.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
Let's apply ```np.abs()``` function to the signals and take a look at the result:
```
changed_batch.apply_transform(np.abs)
original_batch.show_ecg('A00002')
changed_batch.show_ecg('A00002')
```
### [```drop_labels```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.drop_labels)
```drop_labels()``` removes those elements from batch, whose labels are listed in the argument ```drop_list```.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
Note that this action creates a new batch, so we will call it like this:
```
changed_batch = changed_batch.drop_labels(["O"])
```
Now we can inspect labels in the batches:
```
print('Labels in original batch: ', original_batch.target)
print('Labels in batch after drop: ', changed_batch.target)
```
As far as this action creates new batch, it changes ```index``` of the batch, removing indices that correspond to signals with undesired label:
```
print('Indices in original batch: ', original_batch.indices)
print('Indices in batch after drop: ', changed_batch.indices)
```
### [```rename_labels```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.rename_labels)
```rename_labels()``` simply replaces labels according to the argument ```rename_dict```.
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
Suppose, we want to build a classification model to find people with abnormal rhythms using ECG signal. But we have data with three labels: "N" (normal rhythm), "A" (atrial fibrillation), "O" (other abnormal rhythms). So, all we need to do to re-label the data is to combine signals with labels "A" and "O" under single label. Let's rename labels "A" and "O" to some other label, say "AO":
```
changed_batch = changed_batch.rename_labels(rename_dict={'A':'AO', 'O':'AO'})
```
Now we have only two unique labels in the target component:
```
print('Labels in original batch: ', original_batch.target)
print('Labels in batch after drop: ', changed_batch.target)
```
And you can see that the property ```unique_labels``` of the batch also has changed:
```
print('Labels in original batch: ', original_batch.unique_labels)
print('Labels in batch after drop: ', changed_batch.unique_labels)
```
### [```rename_channels```](https://analysiscenter.github.io/cardio/api/cardio.batch.html#cardio.batch.EcgBatch.rename_channels)
```rename_channels()``` changes names of the channels in ```meta``` component under key "signame".
```
original_batch = batch_with_data.deepcopy()
changed_batch = batch_with_data.deepcopy()
```
If you want to change channel name from "ECG" to "Channel №1" you need to write this:
```
changed_batch = changed_batch.rename_channels(rename_dict={'ECG':'Channel №1'})
print('Signal names in original batch: ', original_batch["A00001"].meta["signame"])
print('Signal names in batch after drop: ', changed_batch["A00001"].meta["signame"])
```
Note that method ```show_ecg()``` now will display new names:
```
original_batch.show_ecg('A00001')
changed_batch.show_ecg('A00001')
```
### other actions
You can find complete list of available actions in [documentation](https://analysiscenter.github.io/cardio/api/cardio.ecg_batch.html) on EcgBatch.
## Summary
Summarizing, in Notebook 1 we learned:
* How to get ECG data
* How to create datasets
* How to apply actions
* How actions change batch and its components
In the next [Notebook 2](https://github.com/analysiscenter/cardio/blob/master/tutorials/II.Pipelines.ipynb) we will combine actions in pipeline.
| github_jupyter |
# Collect Tweets into MongoDB
## Install Python libraries
You may need to restart your Jupyter Notebook instance after installed those libraries.
```
!pip install pymongo
!pip install pymongo[srv]
!pip install dnspython
!pip install tweepy
!pip install twitter
```
## Import Python libraries
```
import pymongo
from pymongo import MongoClient
import json
import tweepy
import twitter
from pprint import pprint
import configparser
import pandas as pd
```
## Load the Authorization Info
Save database connection info and API Keys in a config.ini file and use the configparse to load the authorization info.
```
config = configparser.ConfigParser()
config.read('config.ini')
CONSUMER_KEY = config['mytwitter']['api_key']
CONSUMER_SECRET = config['mytwitter']['api_secrete']
OAUTH_TOKEN = config['mytwitter']['access_token']
OATH_TOKEN_SECRET = config['mytwitter']['access_secrete']
mongod_connect = config['mymongo']['connection']
```
## Connect to the MongoDB Cluster
```
client = MongoClient(mongod_connect)
db = client.demo # use or create a database named demo
tweet_collection = db.tweet_collection #use or create a collection named tweet_collection
tweet_collection.create_index([("id", pymongo.ASCENDING)],unique = True) # make sure the collected tweets are unique
```
## Use the Streaming API to Collect Tweets
Authorize the Stream API
```
stream_auth = tweepy.OAuthHandler(CONSUMER_KEY, CONSUMER_SECRET)
stream_auth.set_access_token(OAUTH_TOKEN, OATH_TOKEN_SECRET)
strem_api = tweepy.API(stream_auth)
```
Define the query for the Stream API
```
track = ['election'] # define the keywords, tweets contain election
locations = [-78.9326449,38.4150904,-78.8816972,38.4450731] #defin the location, in Harrisonburg, VA
```
The collected tweets will contain 'election' <span style="color:red;font-weight:bold"> OR </span> are located in Harrisonburg, VA
```
class MyStreamListener(tweepy.StreamListener):
def on_status(self, status):
print (status.id_str)
try:
tweet_collection.insert_one(status._json)
except:
pass
def on_error(self, status_code):
if status_code == 420:
#returning False in on_data disconnects the stream
return False
myStreamListener = MyStreamListener()
myStream = tweepy.Stream(auth = strem_api.auth, listener=myStreamListener)
myStream.filter(track=track)# (locations = locations) #Use either track or locations
```
## Use the REST API to Collect Tweets
Authorize the REST API
```
rest_auth = twitter.oauth.OAuth(OAUTH_TOKEN,OATH_TOKEN_SECRET,CONSUMER_KEY,CONSUMER_SECRET)
rest_api = twitter.Twitter(auth=rest_auth)
```
Define the query for the REST API
```
count = 100 #number of returned tweets, default and max is 100
geocode = "38.4392897,-78.9412224,50mi" # defin the location, in Harrisonburg, VA
q = "election" #define the keywords, tweets contain election
```
The collected tweets will contain 'election' <span style="color:red;font-weight:bold"> AND </span> are located in Harrisonburg, VA
```
search_results = rest_api.search.tweets( count=count,q=q, geocode=geocode) #you can use both q and geocode
statuses = search_results["statuses"]
since_id_new = statuses[-1]['id']
for statuse in statuses:
try:
tweet_collection.insert_one(statuse)
pprint(statuse['created_at'])# print the date of the collected tweets
except:
pass
```
Continue fetching early tweets with the same query.
<p><span style="color:red;font-weight:bold">YOU WILL REACH YOUR RATE LIMIT VERY FAST</span></p>
```
since_id_old = 0
while(since_id_new != since_id_old):
since_id_old = since_id_new
search_results = rest_api.search.tweets( count=count,q=q,
geocode=geocode, max_id= since_id_new)
statuses = search_results["statuses"]
since_id_new = statuses[-1]['id']
for statuse in statuses:
try:
tweet_collection.insert_one(statuse)
pprint(statuse['created_at']) # print the date of the collected tweets
except:
pass
```
## View the Collected Tweets
Print the number of tweets and unique twitter users
```
print(tweet_collection.estimated_document_count())# number of tweets collected
user_cursor = tweet_collection.distinct("user.id")
print (len(user_cursor)) # number of unique Twitter users
```
Create a text index and print the Tweets containing specific keywords.
```
tweet_collection.create_index([("text", pymongo.TEXT)], name='text_index', default_language='english') # create a text index
```
Create a cursor to query tweets with the created index
```
tweet_cursor = tweet_collection.find({"$text": {"$search": "vote"}}) # return tweets contain vote
```
Use pprint to display tweets
```
for document in tweet_cursor[0:10]: # display the first 10 tweets from the query
try:
print ('----')
# pprint (document) # use pprint to print the entire tweet document
print ('name:', document["user"]["name"]) # user name
print ('text:', document["text"]) # tweets
except:
print ("***error in encoding")
pass
tweet_cursor = tweet_collection.find({"$text": {"$search": "vote"}}) # return tweets contain vote
```
Use pandas to display tweets
```
tweet_df = pd.DataFrame(list(tweet_cursor ))
tweet_df[:10] #display the first 10 tweets
tweet_df["favorite_count"].hist() # create a histogram show the favorite count
```
| github_jupyter |
# Machine Translation
- Understand the sequence of one language and then translate it into another language
## Study Links
- [A survey of formal grammars and algorithms for recognition and transformation in mechanical translation]
- Page 254-260
- Various Approaches to MT
- [Google Cloud Natural Language](https://cloud.google.com/natural-language/)
- Google Natural Language API
## Knowledge of NLP covered so far
- Trying to understand the meaning of the word without even looking at the dictionary
- Trying to generate the senetences and try to figutre-out the probability of that sentence formed using Machine Learning/ Statistical Learning (probability models)
## Next Step in NLP
- Machine Translation
- Identifying new model where we can feed a text in one language and we want to get the translated version through that model
- This will be interesting and a challenging task
## History of Machine Translation
- People are trying to do machine translation for over 50+ years
- When they started machine translation, the power of the machines are too small
- It is lesser than the current digital watch that we have today
- Eventhough we have capability to do parallel processing over two corpus, understand the words, texts, for machine translation, it is not 100% mature
- We still have errors, and the Machine Trnaslations should evolve to 1100%
## Machine Translation session details
- In order to understand how people have been doing machine translation over the last 50+ years
- We will first look at the basic element that they had looked at and how they progressed
- Later, we will look at some Statistical Model / Statistical Machine Translation aspect
- And later, we will try to apply Neurol Model into the Machine Translation and see how we can succcessfully translate a sentence in one language to the other langauge
1. Statistical Machine Translation
1. Varous Approaches to MT
2. Autoamtic Machine Translation
3. Statistical Machine Translation
4. Definitions
5. Parallel Corpora
6. ArgMax
7. The Noisy Channel Model
8. Bayes Rule
9 The Language Model - recap
10. Trnaslation Model
11. Alighment
- In next session, Neural Models for Machine Translation will be covered
## Warren Weaver's Note on Trnaslation
- Following quote from Watten Weaver, shows, hwo he looked at the translation from Russian to English
When I loot at an article in Russian, I say "This
is really written in English, but it has been coded
in some strange symbols. I will not proceed to
decode." (Warren Weaver, 1947)
## Vauquois Diagram - Various Approaches to MT
- Below diagram called "Vanquois Diagram" discusses various approaches to machine translation in one shot
- Shows various approaches that people have followed in MT
- Left Tree shows the Source Language
- RIght Tree shows the Target Language

- Assume we are translating a sentence from French to English
### Word2Word or Literal Translation
- __Steps__
- Every word from the source language is converted into the target language, one word at a time with out considering the whole sentence as context
- When we do not know the Source language
- We take the dictionary (say French to English Dictionary)
- Look at the source language sentence
- Take one word at a time
- Find out the actual meaning of that word in Englihs and then write that word
- Once we converted all the words in source language to target language, we will have a __*Bag of Words*__
- _Bag of Words_ because the structure of source language could be very different from target langauge
- We will be rearranging the words to make a sensible sentence in target language
- __Validation__
- In order to validate the translated sentence, we need to take help of 'Profressional Translating persons' to check our translation
- We may get feedback that this word should not be translated in this fashion
- Example: Polysemy words: the word 'bank' translation depends on the context.
- So, just translating using dictionary without context won't give proper translation
- __Limitation/ Issue__
- Without context information, we cannot translate a word in source language to the target language
- So it is not possible to use this method for translating form source to target language in this fashion without knowing the syntax structure of the language
### Syntactic Translation
- __Steps__
1. The source sentece is parsed to create a syntax tree
- Example [Syntax Tree](http://mshang.ca/syntree/) for the sentence 'This is a wug'
- 
- See syntax tree generated using Google Natural Language API Demo - <https://cloud.google.com/natural-language/>
- 
2. The nodes of the source tree is mapped to the nodes the similar syntax tree created for the target language
$(subject)_s \rightarrow (subject)_t$
$(noun)_s \rightarrow (noun)_t$
$(det)_s \rightarrow (det)_t$
$(adj)_s \rightarrow (adj)_t$
3. Generate the sentence in the target language sentece from the parse tree
- __Limitation/ Issue__
- Assumes that the target language also has similar syntax structure
- Sometimes words are swapped in target language when we compare with source language
- Because, every language is very different in its own
- So, we cannot expect that the words are exactly similar between languages
- Even if we construct a syntax tree and place words according to target language syntax structure, the translation may not be proper - because we do not understand the semantic of that langauge

### Semantic Based Translation
- The meaning of the source sentence is obtained
- Try to understand the meaning of the sentence
- Based on the meaning of the sentence, try to do the translation
- Using the semantics derived from the source sentence, the target sentence is generated
### Interlingua Translation
- A meta-language format for representing knowledge independent of any language
- Interlingua is something that represents the knowledge of a langugae
- A very ideal model for representing a language
- It is a neutral model, where you can represent a language
- Source language will be translated into Interlingua model, from where it is tranlsated into target langauge
- $\textit{French} \rightarrow \textit{Interlingua}$
- $\textit{Interlingua} \rightarrow \textit{English}$
- $\textit{Interlingua} \rightarrow \textit{German}$
- $\textit{Interlingua} \rightarrow \textit{Spanish}$
- to any language
- Creating a Interlingua is a critical aspect on this type of machine translation
- In scientific research papers, they have represented the Interlingua in terms of _XML_ or _JSON_
- Instead of Translation systems for all possible pairs of languages, one representation would be used to generate translations
- $O(n^2) \rightarrow O(n)$
- Difficult to design efficient and comprehensive knowledge representation formalisms and due to the large amount of ambiguity
- People are still not very successful in constructing Interlingua so far
| github_jupyter |
<table align="left" width="100%"> <tr>
<td style="background-color:#ffffff;">
<a href="http://qworld.lu.lv" target="_blank"><img src="../images/qworld.jpg" width="35%" align="left"> </a></td>
<td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
prepared by Maksim Dimitrijev (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
</td>
</tr></table>
<table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
$ \newcommand{\bra}[1]{\langle #1|} $
$ \newcommand{\ket}[1]{|#1\rangle} $
$ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
$ \newcommand{\dot}[2]{ #1 \cdot #2} $
$ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
$ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
$ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
$ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
$ \newcommand{\mypar}[1]{\left( #1 \right)} $
$ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
$ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
$ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
$ \newcommand{\onehalf}{\frac{1}{2}} $
$ \newcommand{\donehalf}{\dfrac{1}{2}} $
$ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
$ \newcommand{\vzero}{\myvector{1\\0}} $
$ \newcommand{\vone}{\myvector{0\\1}} $
$ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
$ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
$ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
$ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
$ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
$ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
$ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
$ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
$ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
<h1> Quantum states with complex numbers </h1>
The main properties of quantum states do not change whether we are using complex numbers or not. Let's recall the definition we had in Bronze:
**Recall: Quantum states with real numbers**
When a quantum system is measured, the probability of observing one state is the square of its value.
The summation of amplitude squares must be 1 for a valid quantum state.
The second property also means that the overall probability must be 1 when we observe a quantum system. If we consider a quantum system as a vector, then the length of such vector should be 1.
<h2>How complex numbers affect probabilities</h2>
Suppose that we have a quantum state with the amplitude $a+bi$. What is the probability to observe such state when the quantum system is measured? We need a small update to our statement about <b>the probability of the measurement</b> - it is equal to <b>the square of the absolute value of the amplitude</b>. If amplitudes are restricted to real numbers, then this update makes no difference. With complex numbers we obtain the following:
$\mathopen|a+bi\mathclose| = \sqrt{a^2+b^2} \implies \mathopen|a+bi\mathclose|^2 = a^2+b^2$.
It is easy to see that this calculation works fine if we do not have imaginary part - we just obtain the real part $a^2$. Notice that for the probability $a^2 + b^2$ both real and imaginary part contribute in a similar way - with the square of its value.
<!--Let's check the square of the complex number:
$(a+bi)^2 = (a+bi)(a+bi) = a^2 + 2abi + b^2i^2 = (a^2-b^2) + 2abi$.
In such case we still obtain a complex number, but for a probability we need a real number. -->
Suppose that we have the following vector, representing a quantum system:
$$
\myvector{ \frac{1+i}{\sqrt{3}} \\ -\frac{1}{\sqrt{3}} }.
$$
This vector represents the state $\frac{1+i}{\sqrt{3}}\ket{0} - \frac{1}{\sqrt{3}}\ket{1}$. After doing measurement, we observe state $\ket{1}$ with probability $\mypar{-\frac{1}{\sqrt{3}}}^2 = \frac{1}{3}$. Let's decompose the amplitude of state $\ket{0}$ into form $a+bi$. Then we obtain $\frac{1}{\sqrt{3}} + \frac{1}{\sqrt{3}}i$, and so our probability is $\mypar{\frac{1}{\sqrt{3}}}^2 + \mypar{\frac{1}{\sqrt{3}}}^2 = \frac{2}{3}$.
<h3> Task 1 </h3>
Calculate on the paper the probabilities to observe state $\ket{0}$ and $\ket{1}$ for each quantum system:
$$
\myvector{ \frac{1-i\sqrt{2}}{2} \\ \frac{i}{2} }
\mbox{ , }
\myvector{ \frac{2i}{\sqrt{6}} \\ \frac{1-i}{\sqrt{6}} }
\mbox{ and }
\myvector{ \frac{1+i\sqrt{3}}{\sqrt{5}} \\ \frac{-i}{\sqrt{5}} }.
$$.
<a href="C02_Quantum_States_With_Complex_Numbers_Solutions.ipynb#task1">click for our solution</a>
<h3> Task 2 </h3>
If the following vectors are valid quantum states, then what can be the values of $a$ and $b$?
$$
\ket{v} = \myrvector{0.1 - ai \\ -0.7 \\ 0.4 + 0.3i }
~~~~~ \mbox{and} ~~~~~
\ket{u} = \myrvector{ \frac{1-i}{\sqrt{6}} \\ \frac{1+2i}{\sqrt{b}} \\ -\frac{1}{\sqrt{4}} }.
$$
```
#
# your code is here
# (you may find the values by hand (in mind) as well)
#
```
<a href="C02_Quantum_States_With_Complex_Numbers_Solutions.ipynb#task2">click for our solution</a>
<h3> Task 3</h3>
Randomly create a 2-dimensional quantum state, where both amplitudes are complex numbers.
<b>Write a function</b> that returns a randomly created 2-dimensional quantum state.
<i>Hint:
<ul>
<li> Pick four random values between -100 and 100 for the real and imaginary parts of the amplitudes of state 0 and state 1 </li>
<li> Find an appropriate normalization factor to divide each amplitude such that the length of quantum state should be 1 </li>
</ul>
</i>
<b>Repeat several times:</b>
<ul>
<li> Randomly pick a quantum state </li>
<li> Check whether the picked quantum state is valid </li>
_Note:_ Comment out the first line after writing your code to store the function you have written for later use.
```
#%%writefile random_complex_quantum_state.py
from random import randrange
def random_complex_quantum_state():
# quantum state
quantum_state=[0,0]
#
#
#
return quantum_state
#%%writefile is_quantum_state.py
# testing whether a given quantum state is valid
def is_quantum_state(quantum_state):
#
# your code is here
#
#Use the functions you have written to randomly generate and check quantum states
#
# your code is here
#
```
<a href="C02_Quantum_States_With_Complex_Numbers_Solutions.ipynb#task3">click for our solution</a>
| github_jupyter |
# Backprop Core Example: Text Generation
Generate text based on some provided input.
The default behavior here is that of a standard instance of GPT-2 -- it'll continue writing based on whatever context you've written.
Other generative models, such as T5, can be used as well. If you've trained a model, you can pass in the required tokenizer/model checkpoints and use generate for a variety of tasks.
## What's the deal with all the parameters?
Text generation has a *lot* of parameter options. Some tweaking may be needed for you to get optimal results for your use case. I'll cover what we make accessible, and how they can change generation.
- `min_length`: Forces the model to continue writing until at least the supplied `min_length` is reached.
---
- `temperature`: Alters the probability distribution of the model's softmax. Raising this above 1.0 will lead to an increase in 'out there' token choices, that the model would ordinarily be less confident to select. Lowering it below 1.0 makes the distribution sharper, leading to 'safer' choices.
---
- `top_k`: Method of sampling in which the *K* most likely next words are identified, and the probability is redistributed among those *K*.
---
- `top_p`: Method of sampling in which a probability threshold *p* is chosen. The smallest possible set of words with a combined probability exceeding *p* are selected, and the probability is redistributed among that set.
---
- `do_sample`: Determines whether or not sampling is performed.
---
- `repetition_penalty`: Adds a penalty to words that are present in the input context, and to words that are already included in the generated sequence.
---
- `length_penalty`: Penalty applied to the length of a generated sequence. Defaults to 1.0 (no penalty). Set to lower than 1.0 to get shorter sequences, or higher than 1.0 to get longer ones.
---
- `num_beams`: Number of beams to use in beam search.
**Beam search** maintains `num_beams` different branches of word generation sequences, and returns the one with the highest overall probability. In practice, this is a way to ensure that the generator doesn't miss probable word sequences that may be obscured by an early low-probability word choice.
Setting `num_beams` to 1 means no beam search will be used.
---
- `num_generations`: Number of times the generator will run on the given input. Will give you back a list of results from generation.
---
```
import backprop
# Set your API key to do inference on Backprop's platform
# Leave as None to run locally
api_key = None
```
Let's see what models we can use for this.
```
backprop.TextGeneration.list_models(display=True)
tg = backprop.TextGeneration("gpt2", api_key=api_key)
# The basic functionality, just picks up where you leave off.
tg("Geralt knew the signs: the monster was a", temperature=1.2, max_length=50)
```
### Supplying your own checkpoints
As mentioned, the default generator is GPT-2.
Let's try supplying another model -- one of Backprop's pretrained T5 models. I'll be using the same model that our Sentiment Detection and Summarisation modules use.
```
# Initialise Text Generation with our model checkpoint
tg_t5 = backprop.TextGeneration("t5-base-qa-summary-emotion", api_key=api_key)
# Our sentiment function automatically adds the 'emotion:' prefix.
# As we're accessing the generator directly, we need to do it.
input_text = """emotion: This food was just not good.
Sorry, but you need to do better.
Really gross and undercooked."""
tg_t5(input_text)
```
### Finetuning
As you just saw, text generation can be extremely powerful. The above model has been finetuned for conversational question answering, emotion detection and text summarisation.
As text generation models just take some text as input and produce some text as output, it makes them very versatile.
With further finetuning, it is possible to solve any text based task. Check out our finetuning notebook for an example!
| github_jupyter |
# FairWorkflows quickstart notebook
Demonstrates the use of the library to semantically annotate a workflow, execute it, and publish prospective and retrospective semantic annotations.
## Define the steps of your workflow
Each step should be its own function. Mark the function as such with the @fairstep decorator.
```
from fairworkflows import is_fairworkflow, is_fairstep, FairStep, FairWorkflow
@is_fairstep(label='Addition')
def add(a:float, b:float) -> float:
"""Adding up numbers!"""
return a + b
@is_fairstep(label='Subtraction')
def sub(a: float, b: float) -> float:
"""Subtracting numbers."""
return a - b
@is_fairstep(label='Multiplication')
def mul(a: float, b: float) -> float:
"""Multiplying numbers."""
return a * b
@is_fairstep(label='A strange step with little use')
def weird(a: float, b:float) -> float:
"""A weird function"""
return a * 2 + b * 4
```
## Define your workflow using @fairworkflow
Now write a function which describes your workflow. Mark this function with the @fairworkflow decorator.
```
@is_fairworkflow(label='My Workflow')
def my_workflow(in1, in2, in3):
"""
A simple addition, subtraction, multiplication workflow
"""
t1 = add(in1, in2)
t2 = sub(in1, in2)
t3 = mul(weird(t1, in3), t2)
return t3
```
## Create an instance of your workflow and display it
```
fw = FairWorkflow.from_function(my_workflow)
type(fw)
fw.display()
fw.display_rdf()
```
## Publish the (prospective) workflow
You may publish the workflow, and its steps, as nanopublications in the usual manner:
```
#fw.publish_as_nanopub()
```
Be warned though - the above will keep publishing to the 'real' nanopub server network. For testing you may prefer to publish to the test servers as follows (note that this will refuse to publish a workflow you have already published :
```
fw.publish_as_nanopub(use_test_server=True, publish_steps=True)
```
You can then find your nanopublications by replacing the base of the URI with http://test-server.nanopubs.lod.labs.vu.nl/
## Execute your workflow using .execute()
Set num_threads greater than 1 if you wish to exploit parallelisation in your workflow. The retrospective provenance is also returned as a (nano) Publication object, that can optionally be published.
```
result, prov = fw.execute(1, 4, 3)
result
```
### Retrospective prov
A WorkflowRetroProv object is returned along with the result of the execution.
```
type(prov)
print(prov)
```
### Retrospective prov for each step
You can iterate through a WorkflowRetroProv object to get the StepRetroProv objects for each step. Print these to see the RDF they contain (input/output variable values, start and end datetime of the step's execution etc.)
```
for sp in prov:
print(sp)
```
### Publish the retrospective provenance
You can use the .publish_as_nanopub() method as with FairStep and FairWorkflow objects. This publishes a nanopub per step and one for the whole workflow, mirroring the prospective RDF.
```
prov.publish_as_nanopub(use_test_server=True)
```
The last nanopub (whose URI ends in #fairworkflowprov) contains the links to all of the individual step retrospective provenances.
## Provide semantic annotations for input and output variables
If you wish to specify semantic types for the inputs/outputs to a step, you can do so in the arguments to the decorator.
For example, if you have an input parameter 'a', you can write a='http://www.example.org/distance' to assign that (semantic) type to a. As output of functions is not named in python, you can specify the same but with 'out1', 'out2' etc. See the following example:
```
@is_fairstep(label='Addition', a='http://www.example.org/distance', returns='http://www.example.org/mass')
def add(a:float, b:float) -> float:
return a + b
```
If we now look at the RDF generated for the step, we will see that input parameter 'a' and the step output ('out1') both have the (additional) semantic types specified.
```
# ACTIONS:
# Add language and version to nanopubs (i.e. what the description is written in)
print(add._fairstep)
```
### Specify more than one semantic type for a parameter
You can provide a list of URIs if you want to specify several semantic types for e.g. parameter 'a':
```
@is_fairstep(label='Addition', a=['http://www.example.org/distance', 'http://www.example.org/number'])
def another_step(a:float, b:float) -> float:
"""Add two numbers together"""
return a + b
print(another_step._fairstep)
```
You can check the programming language that was used for writing the step:
```
print(another_step._fairstep.language)
```
## Semantic types for function producing multiple outputs
Provide 'out' with a tuple of the same length as the number of function outputs. You can use None for any you do not wish to assign a particular semantic type to.
```
from typing import Tuple
@is_fairstep(label='Addition and subtraction', returns=('http://www.example.org/distance', 'http://www.example.org/number'))
def another_step(a:float, b:float) -> Tuple[float, float]:
return a + b, a - b
print(another_step._fairstep)
```
As before, you may provide a list of URIs for each output. If you do not want to provide semantic types for a particular output, simply pass None:
```
from typing import Tuple
@is_fairstep(label='Addition and subtraction', returns=(['http://www.example.org/distance', 'http://www.example.org/number'], None))
def another_step(a:float, b:float) -> Tuple[float, float]:
"""This step returns an addition and a subtraction of its inputs"""
return a + b, a - b
print(another_step._fairstep)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yukinaga/twitter_bot/blob/master/section_5/02_preprocessing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# データの前処理
対話文のデータセットに前処理を行い、保存します。
## ライブラリのインストール
分かち書きのためにjanomeを、テキストデータの前処理のためにtorchtextをインストールします。
```
!pip install janome==0.4.1
!pip install torchvision==0.7.0
!pip install torchtext==0.7.0
!pip install torch==1.6.0
```
## Google ドライブとの連携
以下のコードを実行し、認証コードを使用してGoogle ドライブをマウントします。
```
from google.colab import drive
drive.mount('/content/drive/')
```
## 対話文の取得
雑談対話コーパス「projectnextnlp-chat-dialogue-corpus.zip」をダウンロードします。
> Copyright (c) 2015 Project Next NLP 対話タスク 参加者一同
> https://sites.google.com/site/dialoguebreakdowndetection/chat-dialogue-corpus/LICENSE.txt
> Released under the MIT license
解凍したフォルダをGoogle ドライブにアップします。
フォルダからjsonファイルを読み込み、対話文として成り立っている文章を取り出してリストに格納します。
```
import glob # ファイルの取得に使用
import json # jsonファイルの読み込みに使用
import re
path = "/content/drive/My Drive/live_ai_data/projectnextnlp-chat-dialogue-corpus/json" # フォルダの場所を指定
files = glob.glob(path + "/*/*.json") # ファイルの一覧
dialogues = [] # 複数の対話文を格納するリスト
file_count= 0 # ファイル数のカウント
for file in files:
with open(file, "r") as f:
json_dic = json.load(f)
dialogue = [] # 単一の対話
for turn in json_dic["turns"]:
annotations = turn["annotations"] # 注釈
speaker = turn["speaker"] # 発言者
utterance = turn["utterance"] # 発言
# 空の文章や、特殊文字や数字が含まれる文章は除く
if (utterance=="") or ("\\u" in utterance) or (re.search("\d", utterance)!=None):
dialogue.clear() # 対話をリセット
continue
utterance = utterance.replace(".", "。").replace(",", "、")
utterance = utterance.split("。")[0]
if speaker=="U": # 発言者が人間であれば
dialogue.append(utterance)
else: # 発言者がシステムであれば
is_wrong = False
for annotation in annotations:
breakdown = annotation["breakdown"] # 分類
if breakdown=="X": # 1つでも不適切評価があれば
is_wrong = True
break
if is_wrong:
dialogue.clear() # 対話をリセット
else:
dialogue.append(utterance) # 不適切評価が無ければ対話に追加
if len(dialogue) >= 2: # 単一の会話が成立すれば
dialogues.append(dialogue.copy())
dialogue.pop(0) # 最初の要素を削除
file_count += 1
if file_count%100 == 0:
print("files:", file_count, "dialogues", len(dialogues))
print("files:", file_count, "dialogues", len(dialogues))
```
## データ拡張の準備
データ拡張の準備として、正規表現の設定および分かち書きを行います。
```
import re
from janome.tokenizer import Tokenizer
re_kanji = re.compile(r"^[\u4E00-\u9FD0]+$") # 漢字の検出用
re_katakana = re.compile(r"[\u30A1-\u30F4]+") # カタカナの検出用
j_tk = Tokenizer()
def wakati(text):
return [tok for tok in j_tk.tokenize(text, wakati=True)]
wakati_inp = [] # 単語に分割された入力文
wakati_rep = [] # 単語に分割された応答文
for dialogue in dialogues:
wakati_inp.append(wakati(dialogue[0])[:10])
wakati_rep.append(wakati(dialogue[1])[:10])
```
## データ拡張
対話データの数を水増しします。
ある入力文を、それに対応する応答文以外の複数の応答文と組み合わせます。
組み合わせる応答文は、入力文に含まれる漢字やカタカナの単語を含むものを選択します。
```
dialogues_plus = []
for i, w_inp in enumerate(wakati_inp): # 全ての入力文でループ
inp_count = 0 # ある入力から生成された対話文をカウント
for j, w_rep in enumerate(wakati_rep): # 全ての応答文でループ
if i==j:
dialogues_plus.append(["".join(w_inp), "".join(w_rep)])
continue
similarity = 0 # 類似度
for w in w_inp: # 入力文と同じ単語があり、それが漢字かカタカナであれば類似度を上げる
if (w in w_rep) and (re_kanji.fullmatch(w) or re_katakana.fullmatch(w)):
similarity += 1
if similarity >= 1:
dialogue_plus = ["".join(w_inp), "".join(w_rep)]
if dialogue_plus not in dialogues_plus:
dialogues_plus.append(dialogue_plus)
inp_count += 1
if inp_count >= 10: # ある入力から生成する対話文の上限
break
if i%1000 == 0:
print("i:", i, "dialogues_pus:", len(dialogues_plus))
print("i:", i, "dialogues_pus:", len(dialogues_plus))
```
拡張された対話データを、新たな対話データとします。
```
dialogues = dialogues_plus
```
## 対話データの保存
対話データをcsvファイルとしてGoogle Driveに保存します。
```
import csv
from sklearn.model_selection import train_test_split
dialogues_train, dialogues_test = train_test_split(dialogues, shuffle=True, test_size=0.05) # 5%がテストデータ
path = "/content/drive/My Drive/live_ai_data/" # 保存場所
with open(path+"dialogues_train.csv", "w") as f:
writer = csv.writer(f)
writer.writerows(dialogues_train)
with open(path+"dialogues_test.csv", "w") as f:
writer = csv.writer(f)
writer.writerows(dialogues_test)
```
## 対話文の取得
Googleドライブから、対話文のデータを取り出してデータセットに格納します。
```
import torch
import torchtext
from janome.tokenizer import Tokenizer
path = "/content/drive/My Drive/live_ai_data/" # 保存場所を指定
j_tk = Tokenizer()
def tokenizer(text):
return [tok for tok in j_tk.tokenize(text, wakati=True)] # 内包表記
# データセットの列を定義
input_field = torchtext.data.Field( # 入力文
sequential=True, # データ長さが可変かどうか
tokenize=tokenizer, # 前処理や単語分割などのための関数
batch_first=True, # バッチの次元を先頭に
lower=True # アルファベットを小文字に変換
)
reply_field = torchtext.data.Field( # 応答文
sequential=True, # データ長さが可変かどうか
tokenize=tokenizer, # 前処理や単語分割などのための関数
init_token = "<sos>", # 文章開始のトークン
eos_token = "<eos>", # 文章終了のトークン
batch_first=True, # バッチの次元を先頭に
lower=True # アルファベットを小文字に変換
)
# csvファイルからデータセットを作成
train_data, test_data = torchtext.data.TabularDataset.splits(
path=path,
train="dialogues_train.csv",
validation="dialogues_test.csv",
format="csv",
fields=[("inp_text", input_field), ("rep_text", reply_field)] # 列の設定
)
```
## 単語とインデックスの対応
単語にインデックスを割り振り、辞書として格納します。
```
input_field.build_vocab(
train_data,
min_freq=3,
)
reply_field.build_vocab(
train_data,
min_freq=3,
)
print(input_field.vocab.freqs) # 各単語の出現頻度
print(len(input_field.vocab.stoi))
print(len(input_field.vocab.itos))
print(len(reply_field.vocab.stoi))
print(len(reply_field.vocab.itos))
```
## データセットの保存
データセットの`examples`とFieldをそれぞれ保存します。
```
import dill
torch.save(train_data.examples, path+"train_examples.pkl", pickle_module=dill)
torch.save(test_data.examples, path+"test_examples.pkl", pickle_module=dill)
torch.save(input_field, path+"input_field.pkl", pickle_module=dill)
torch.save(reply_field, path+"reply_field.pkl", pickle_module=dill)
```
| github_jupyter |
# Parametrized model for onshore and offshore wind turbines
Authors: Romain Besseau [romain.besseau@mines-paristech.fr](mailto:romain.besseau@mines-paristech.fr) and Romain Sacchi [r_s@me.com](mailto:r_s@me.com)
The script below was used to generate life cycle inventories of wind turbines based on the [Danish wind turbines registry](https://ens.dk/sites/ens.dk/files/Statistik/anlaegprodtilnettet.xls).
Results were used in the following publications:
Romain Sacchi, Romain Besseau, Paula Pérez-López, Isabelle Blanc, Exploring technologically, temporally and geographically-sensitive life cycle inventories for wind turbines: A parameterized model for Denmark, Renewable Energy, Volume 132, 2019, Pages 1238-1250, ISSN 0960-1481, [doi.org/10.1016/j.renene.2018.09.020](https://doi.org/10.1016/j.renene.2018.09.020).
and
Romain Besseau, Romain Sacchi, Isabelle Blanc, Paula Pérez-López, Past, present and future environmental footprint of the Danish wind turbine fleet with LCA_WIND_DK, an online interactive platform, Renewable & Sustainable Energy Reviews, online, 2019, [doi.org/10.1016/j.rser.2019.03.030](https://doi.org/10.1016/j.rser.2019.03.030).
We need first to import a few librairies
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import brightway2 as bw
from scipy.interpolate import InterpolatedUnivariateSpline
plt.style.use('ggplot')
%matplotlib inline
```
And the following assumes you have an BW2 project within which ecoinvent 3.4 cutoff is installed. If not, you should create a project and install ecoinvent before going any further.
```
bw.projects.set_current('my_project')
eidb = bw.Database('ecoinvent 3.4 cut off')
```
## Creation of datasets for power transformers
Let's start by creating datasets for high voltage and medium voltage power transformer. These datasets are based on the EPD of ABB's power transformers.
This has to be executed once.
### creation of 500 MVA transformer dataset
```
if [act for act in eidb if 'Power transformer TrafoStar 500 MVA' in act['name']] == []:
act_transfo=[act for act in eidb if act["name"]=="transformer production, high voltage use"][0]
new_act=act_transfo.copy()
new_act["name"]="Power transformer TrafoStar 500 MVA"
new_act["unit"]="unit"
new_act.save()
for exc in new_act.exchanges():
exc.delete()
new_act.save()
#electric steel
steel=[act for act in eidb if "steel production, electric, low-alloyed" in act["name"] and "RER" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=99640,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#transformer oil
steel=[act for act in eidb if "market for lubricating oil" in act["name"] and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=63000,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#copper
steel=[act for act in eidb if act["name"]=="market for copper" and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=39960,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#insulation
steel=[act for act in eidb if act["name"]=="market for glass wool mat" and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=6500,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#wood
steel=[act for act in eidb if act["name"]== "planing, board, softwood, u=20%" and "CH" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=15000,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#porcelain
steel=[act for act in eidb if act["name"]=="market for ceramic tile" and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=2650,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#construction steel
steel=[act for act in eidb if act["name"]=="market for steel, unalloyed" and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=53618,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#paint
steel=[act for act in eidb if act["name"]== "market for electrostatic paint" and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=2200,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#electricity, medium
steel=[act for act in eidb if act["name"]=="market for electricity, medium voltage" and "SE" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=750000,unit="kilowatt hour",type='technosphere')
new_exc.save()
new_act.save()
#heat
steel=[act for act in eidb if act["name"]=="heat, from municipal waste incineration to generic market for heat district or industrial, other than natural gas" and "SE" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=1080000,unit="megajoule",type='technosphere')
new_exc.save()
new_act.save()
#output
steel=[act for act in eidb if act["name"]=="Power transformer TrafoStar 500 MVA"][0]
new_exc = new_act.new_exchange(input=new_act.key,amount=1,unit="unit",categories="",type='production')
new_exc.save()
new_act.save()
```
### creation of 10 MVA transformer dataset
```
if [act for act in eidb if 'Power transformer TrafoStar 10 MVA' in act['name']] == []:
act=[a for a in eidb if "Power transformer TrafoStar 500 MVA" in a["name"]][0]
new_act=act.copy()
new_act["name"]="Power transformer TrafoStar 10 MVA"
new_act.save()
for exc in new_act.exchanges():
print(exc.input['name'])
if exc.input['name']=="steel production, electric, low-alloyed":
exc["amount"]=6820
exc.save()
if exc.input['name']=="market for lubricating oil":
exc["amount"]=6780
exc.save()
if exc.input['name']=="market for copper":
exc["amount"]=3526
exc.save()
if exc.input['name']=="market for ceramic tile":
exc["amount"]=53
exc.save()
if exc.input['name']=="market for steel, unalloyed":
exc["amount"]=9066
exc.save()
if exc.input['name']=="market for electrostatic paint":
exc["amount"]=95
exc.save()
if exc.input['name']=="market for electricity, medium voltage":
exc["amount"]=105200
exc.save()
if exc.input['name']=="heat, from municipal waste incineration to generic market for heat district or industrial, other than natural gas":
exc["amount"]=68760
exc.save()
if exc.input['name']=="market for aluminium, cast alloy":
exc["amount"]=65
exc.save()
if exc.input['name']=="market for sheet rolling, steel":
exc.delete()
if exc.input['name']=="market for epoxy resin, liquid":
exc.delete()
if exc.input['name']=="market for glass fibre":
exc.delete()
if exc.input['name']=="market for kraft paper, bleached":
exc.delete()
if exc.input['name']=="market for paper, melamine impregnated":
exc.delete()
if exc.input['name']=="market for electrostatic paint":
exc.delete()
if exc.input['name']=="market for glass fibre":
exc.delete()
if exc.input['name']=="Power transformer TrafoStar 250 MVA":
exc.input['name']="Power transformer TrafoStar 10 MVA"
exc.input=new_act
exc.save()
#insulation
steel=[act for act in eidb if "market for glass wool mat" in act["name"] and "GLO" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=337,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
#wood
steel=[act for act in eidb if "planing, board, softwood, u=20%" in act["name"] and "CH" in act["location"]][0]
new_exc = new_act.new_exchange(input=steel.key,amount=366,unit="kilogram",type='technosphere')
new_exc.save()
new_act.save()
```
## Foundation functions
These functions are used to size foundation, electrical connexions, etc...
```
def pile_weight(p, pile_height):
#diameters, in meters
diameter=[5, 5.5, 5.75, 6.75 ,7.75 ]
#kW
power=[3000,3600,4000,8000,10000]
fit_diameter= np.polyfit(power, diameter, 1)
f_fit_diameter=np.poly1d(fit_diameter)
#diameter for given power, in m
outer_diameter=f_fit_diameter(p)
#Cross section area of pile
outer_area=(np.pi/4)*(outer_diameter**2)
#Pile volume, in m3
outer_volume=outer_area*pile_height
inner_diameter= outer_diameter
pile_thickness = np.interp(p, [2000, 3000, 3600, 4000, 8000, 10000], [0.07,0.10, 0.13, 0.16, 0.19, 0.22])
inner_diameter-= 2 * pile_thickness
inner_area=(np.pi/4)*(inner_diameter**2)
inner_volume=inner_area*pile_height
volume_steel=outer_volume-inner_volume
weight_steel=8000*volume_steel
return weight_steel
def transition_height():
pile_length=[35,55,35,60,40,65,50,70,50,80]
transition_length=[15,20,15,20,15,24,20,30,20,31]
fit_transition_length= np.polyfit(pile_length, transition_length, 1)
return np.poly1d(fit_transition_length)
fit_transition_height = transition_height()
def transition_weight():
transition_length=[15,20,15,20,15,24,20,30,20,31]
transition_weight=[150,250,150,250,160,260,200,370,250,420]
fit_transition_weight= np.polyfit(transition_length, transition_weight, 1)
return np.poly1d(fit_transition_weight)
fit_transition_weight = transition_weight()
def grout_volume():
transition_length=[15,20,15,20,15,24,20,30,20,31]
grout=[15,35,15,35,20,40,25,60,30,65]
fit_grout= np.polyfit(transition_length, grout, 1)
return np.poly1d(fit_grout)
fit_grout_volume = grout_volume()
def scour_volume():
scour=[2200, 2200, 2600, 3100, 3600]
turbine_power=[3000, 3600, 4000, 8000, 10000]
fit_scour=np.polyfit(turbine_power, scour, 1)
return np.poly1d(fit_scour)
fit_scour_volume = scour_volume()
def pile_height(P, fit_penetration_depth, sea_depth):
return 9 + fit_penetration_depth(P)+ sea_depth
def penetration_depth():
#meters
depth=[22.5,22.5,23.5,26,29.5]
#kW
P = [3000,3600,4000,8000,10000]
fit_penetration= np.polyfit(P, depth, 1)
f_fit_penetration=np.poly1d(fit_penetration)
return f_fit_penetration
fit_penetration_depth = penetration_depth()
def transport_requirements(M_nacelle, M_tower, M_rotor, M_foundation, M_all, LT):
#taken from Vestas 2012 LCA report
#https://www.vestas.com/~/media/vestas/about/sustainability/pdfs/lca_v903mw_version_1_1.pdf
trsp_truck_nacelle=1025*(M_nacelle/1000)
trsp_truck_rotor=600*(M_rotor/1000)
trsp_truck_tower=1100*(M_tower/1000)
trsp_ship_tower=8050*(M_tower/1000)
trsp_truck_foundation= 50*(M_foundation/1000)
#transport to local waste facilities
trsp_end_of_life=200*(M_all/1000)
#30 turbines per plant
trsp_maintenance_per_year = 2160/30/1000*LT
#600 km by ship assumed
trsp_ship_offshore = 600*(M_all/1000)
return trsp_truck_nacelle, trsp_truck_rotor, trsp_truck_tower, trsp_ship_tower,trsp_truck_foundation, trsp_end_of_life, trsp_maintenance_per_year, trsp_ship_offshore
def grout_and_monopile_requirements(P, sea_depth):
pile_length = pile_height(P, fit_penetration_depth, sea_depth)
transition_lenght = fit_transition_height(pile_length)
density = 2400 #kg/m**3
m_grout = fit_grout_volume(transition_lenght) * density
m_monopile = 1e3*fit_transition_weight(transition_lenght)
return m_grout, m_monopile
#Nexans cable at 150 kV, section, ampacity and power capacity in kW
df150 = pd.DataFrame(index = [400, 500, 630, 800,1000, 1200, 1600, 2000])
df150['I'] = [710, 815, 925, 1045, 1160, 1335, 1425, 1560]
df150['P'] = 150 * df150['I']
#Nexans cable at 150 kV, section, ampacity and power capacity in kW
df33 = pd.DataFrame(index = [95, 120, 150, 185, 240, 300, 400, 500, 630, 800])
df33['I'] = [352, 399, 446, 502, 581, 652, 726, 811, 904, 993]
df33['P'] = 33 * df33['I']
# defines cable requirements for offshore foundations
def cable_requirements(P, park_size, dist_transfo, dist_coast, affiche = False):
copper_density = 8960
#Cable of 300 mm² cross section allocated according to the wind turbine power
cross_section1 = 300 * P / df33.loc[300].P
if affiche:
print('cross_section1 : %s mm²' %cross_section1)
#
m_copper = (cross_section1*1e-6*(dist_transfo*1e3))*copper_density
energy_cable_laying_ship = 450 * 39 / 15 * dist_transfo # 450 l diesel/hour for the ship that lays the cable at sea bottom
#39 MJ/liter, 15 km/h as speed of laying the cable
#Cross section calculated based on the farm cumulated power,
#and the transport capacity of the Nexans cables @ 150kV if the cumulated power of the park cannot be
#transported @ 33kV
#Test if the cumulated power of teh wind farm is inferior to 30 MW,
# If so, we use 33 kV cables.
if P*park_size <= 30e3:
cross_section2 = np.interp(P*park_size, df33.P.values, df33.index.values)
#Otherwise, we use 150 kV cables
else:
cross_section2 = np.interp(P*park_size, df150.P.values, df150.index.values)
if affiche:
print('cross_section2 : %s mm²' %cross_section2)
m_copper += (cross_section2*1e-6*(dist_coast*1e3/park_size))*copper_density
energy_cable_laying_ship += 450 * 39 / 15 * dist_coast/park_size #450 l/h de conso, 39 MJ/l, 15 km/h de vitesse d'installation
m_cable = m_copper*617/220
return m_cable*0.5, energy_cable_laying_ship*0.5
# defines cable requirements for onshore foundations
def cable_requirements_Onshore(P, affiche = False):
copper_density = 8960
#Cable of 300 mm² cross section allocated according to the wind turbine power
cross_section1 = 300 * P / df33.loc[300].P
#Calculation of the cable length starting fromt eh road surface, with assumed with of 8 meters
cable_length = np.interp(P, [0, 2000],[0, 8000]) / 8
if affiche:
print('cross_section1 : %s mm²' %cross_section1)
print('cable length : %s m' %cable_length)
#
m_copper = (cross_section1*1e-6*(cable_length))*copper_density
m_cable = m_copper*617/220
return m_cable*0.5
def add_to_dict(dictionary, key, value):
if key not in dictionary:
dictionary[key] = value
else:
dictionary[key] = dictionary[key] + value
```
### Activities
Now, let's load inventories of a 30 kW, 150 kW, 600 kW, 800 kW, 2000 kW wind turbines. These files contains lifecycle activities that has to be linked with activities of the ecoinvent database. That will be done executing the function find_UUID(). As it takes quite long, the dataframe containing all the necessary information can be saved in a pikcle files and be loaded directly.
### To create a pickle
```
path = 'to define'
datasets=pd.read_excel(path + 'Wind turbines inventories.xlsx', sheetname="All", dtype=None, decimal=";", header=0)
del datasets['Unnamed: 9']
datasets.head()
#Use of cement for foundation and reinforcing steel for foundation
datasets = datasets.replace('market for concrete, normal','market for concrete, sole plate and foundation')
datasets.loc[datasets[(datasets['Component']=='Foundation')&(datasets['Market name']=='market for steel, low-alloyed')].index, 'Market name'] = 'market for reinforcing steel'
datasets.loc[datasets[(datasets['Component']=='Foundation')&(datasets['Market name']=='market for reinforcing steel')].index, 'Dataset'] = 'Reinforcing steel'
datasets.Dataset.unique()
def find_UUID():
list_unique_act=datasets["Market name"].unique()
order_preference=["DK", "DE", "CH", "RER", "RoW", "GLO"]
for act in list_unique_act:
print(act)
for pref in order_preference:
list_act=[a for a in eidb if act in a["name"] and pref in a["location"]]
if len(list_act)==1:
datasets.loc[:,"UUID"][datasets.loc[:,"Market name"]==act]=list_act[0]["code"]
break
if len(list_act)>1:
list_act=[a for a in eidb if act == a["name"] and pref in a["location"]]
datasets.loc[:,"UUID"][datasets.loc[:,"Market name"]==act]=list_act[0]["code"]
break
else:
list_act=[a for a in bw.Database("biosphere3") if act in a["name"]]
if len(list_act)>0:
datasets.loc[:,"UUID"][datasets.loc[:,"Market name"]==act]=list_act[0]["code"]
break
else:
datasets.loc[:,"UUID"][datasets.loc[:,"Market name"]==act]=[a for a in bw.Database("biosphere3") if 'Transformation, from pasture, man made, intensive' in a["name"]][0]['code']
def get_act_country():
list_unique_uuid=datasets.loc[:,"UUID"].unique()
for uuid in list_unique_uuid:
try:
datasets.loc[:,"Location"][datasets.loc[:,"UUID"]==uuid]=bw.get_activity((eidb.name, uuid))["location"]
except:
datasets.loc[:,"Location"][datasets.loc[:,"UUID"]==uuid]=None
datasets["UUID"]=0
find_UUID()
datasets["Location"]=0
get_act_country()
#We add recycling of aluminium and chromium steel
df_act = datasets
df_act2 = pd.DataFrame(index = df_act.columns)
new_act = [act for act in eidb if 'market for waste' in act['name'] and 'aluminium' in act['name'] ][0]
print(new_act)
df_act2[new_act['name']] = [None, 'Disposal', None, None,'Aluminium waste', 'kg', None, eidb.name, new_act['name'], new_act['code'], new_act['location']]
df_act = df_act.append(df_act2.T)
df_act = df_act.append(df_act[df_act['Dataset'] =='Steel, inert waste'].iloc[0].replace('Steel, inert waste','Chromium Steel waste'))
df_act
df_act = df_act.replace('ecoinvent 3.3 cutoff', eidb.name)
#Saving data into a pickle
df_act.to_pickle('activities and uuids.pkl')
```
### To load the pickle
```
#Loading data from the pickle
df_act = pd.read_pickle('activities and uuids.pkl')
```
## Inventories
Now let's use values of the inventories. Values expressed in kg are normalized to get the share of different material.
Values not expressed in kg are used directly. At that stage, 2 new dataframe are created and saved in a pickle. Once created, they can be directly loaded to save time.
### To create the pickle
```
#Loading wind turbine inventories dataframe and creating 'df_inv.pkl'
# All that has "kg" as unit
df_inv = pd.read_excel(path + 'Wind turbines inventories.xlsx')
df_inv = df_inv.replace('30kW', 30)
df_inv = df_inv.replace('150kW',150)
df_inv = df_inv.replace('600kW', 600)
df_inv = df_inv.replace('800kW', 800)
df_inv = df_inv.replace('2000kW', 2000)
df_inv = df_inv[ df_inv.Unit == 'kg']
#df_inv = df_inv.pivot_table(columns= 'Power', values= 'Quantity', index=['Phase', 'Component','Sub-component', 'Dataset'])
df_inv = df_inv.pivot_table(columns= 'Power', values=['Quantity','Unit'] , index=['Phase', 'Component','Sub-component', 'Dataset'])
df_inv = df_inv.T
#Filling nan values
df_inv = df_inv.loc['Quantity']
df_inv.loc[0] = 0
df_inv = df_inv.sort_index()
df_inv.interpolate(method = 'index', limit_direction = 'both', inplace = True)
df_inv = df_inv.drop(df_inv.index[0])
print('NaN filled')
df_inv_tot = df_inv.T.groupby(level=['Phase','Component']).sum(axis = 1).T
for phase in set(df_inv.columns.get_level_values(0)):
print(phase + ' is being normalised')
for component in set(df_inv[phase].columns.get_level_values(0)):
for sub_comp in set(df_inv[phase][component].columns.get_level_values(0)):
for dataset in set(df_inv[phase][component][sub_comp].columns.get_level_values(0)):
#print(phase, component, sub_comp, dataset)
df_inv[phase][component][sub_comp][dataset] = df_inv[phase][component][sub_comp][dataset] / df_inv_tot[phase][component]
df_inv.to_pickle('df_inv.pkl')
#Loading wind turbine inventories dataframe and creating 'df_inv_not_kg.pkl'
#All that has not "kg" as unit
df_inv_not_kg = pd.read_excel(path + 'Wind turbines inventories.xlsx')
df_inv_not_kg = df_inv_not_kg.replace('30kW', 30)
df_inv_not_kg = df_inv_not_kg.replace('150kW',150)
df_inv_not_kg = df_inv_not_kg.replace('600kW', 600)
df_inv_not_kg = df_inv_not_kg.replace('800kW', 800)
df_inv_not_kg = df_inv_not_kg.replace('2000kW', 2000)
df_inv_not_kg = df_inv_not_kg[ df_inv_not_kg.Unit != 'kg']
df_inv_not_kg = df_inv_not_kg.pivot_table(columns= 'Power', values=['Quantity','Unit'] , index=['Phase', 'Component','Sub-component', 'Dataset'])
df_inv_not_kg = df_inv_not_kg.T
for phase in set(df_inv_not_kg.columns.get_level_values(0)):
for component in set(df_inv_not_kg[phase].columns.get_level_values(0)):
df_inv_not_kg[phase][component].loc['Quantity'].plot()
plt.title('%s, %s'%(phase, component))
plt.ylim([0, df_inv_not_kg[phase][component].loc['Quantity'].max().max()])
plt.xlim([0, 8000])
#plt.show()
df_inv_not_kg['Assembly']['Tower']['Assembly']['Galvanizing [m]'] = df_inv_not_kg['Assembly']['Tower']['Assembly']['Steel arc welding [m]']
df_inv_not_kg = df_inv_not_kg.loc['Quantity']
df_inv_not_kg = df_inv_not_kg.drop('Electricity [kWh]', axis = 1, level = 3)
df_inv_not_kg.to_pickle('df_inv_not_kg.pkl')
```
Once the dataframe saved, they can be loaded and some new activities also considered :
### To load the pickle
```
df_inv = pd.read_pickle('df_inv.pkl')
df_inv_not_kg = pd.read_pickle('df_inv_not_kg.pkl')
diesel_burned_activity = [act for act in eidb if 'market for diesel, burned in building machine' in act['name']][0]
MV_transfo = [act for act in eidb if 'Power transformer TrafoStar 10 MVA' in act['name']][0]
HV_transfo = [act for act in eidb if 'Power transformer TrafoStar 500 MVA' in act['name']][0]
assembly_activities = list(set(df_inv['Assembly'].columns.get_level_values(2)))
#Assembly activites to add
Copper_wire_drawing = [act for act in eidb if 'market for wire drawing, copper' in act['name']][0]
print(Copper_wire_drawing)
Explosive = [act for act in eidb if 'market for explosive, tovex' in act['name']][0]
print(Explosive)
Steel_sheet_rolling = [act for act in eidb if 'market for sheet rolling, steel' in act['name']][0]
print(Steel_sheet_rolling)
Aluminium_sheet_rolling = [act for act in eidb if 'market for sheet rolling, aluminium' in act['name']][0]
print(Aluminium_sheet_rolling)
Chromium_sheet_rolling = [act for act in eidb if 'market for sheet rolling, chromium steel' in act['name']][0]
print(Chromium_sheet_rolling)
Road = [act for act in eidb if 'market for road' == act['name'] and 'CH' in act['location']][0]
print(Road)
Truck_transport = [act for act in eidb if 'market for transport, freight, lorry >32 metric ton, EURO6' in act['name']][0]
print(Truck_transport)
Ship_transport = [act for act in eidb if 'market for transport, freight, inland waterways, barge' in act['name']][0]
print(Ship_transport)
Digger = [act for act in eidb if 'market for excavation, hydraulic digger' in act['name']][0]
print(Digger)
Cement = [act for act in eidb if 'market for cement, Portland' in act['name']][0]
print(Cement)
disposal_activities = list(set(df_inv['Disposal'].columns.get_level_values(2)))
disposal_activities.append('Aluminium waste')
disposal_activities.append('Chromium Steel waste')
disposal_activities
#Dataset for electricity directly used and steel low-alloyed
Electricity_dataset = [act for act in eidb if 'market for electricity, high voltage' in act['name'] and 'DK' in act['location']][0]
print(Electricity_dataset)
Steel_dataset = [act for act in eidb if 'market for steel, low-alloyed' == act['name']][0]
print(Steel_dataset)
```
### Scaling models used in the publication are defined below
Scaling models are based on regression from TheWindPower database (see Supporting information of the article: "Exploring technologically, temporally and geographically-sensitive life cycle inventories for renewable energy systems: a parameterized model for wind turbines"
```
##Scaling model: Rotor diameter (m) - Rated power (kW)
def func_rotor_power(x, a, b, c, d):
y = a - b*np.exp(-(x-d)/c)
return y
p_rotor_power_ON = [152.66222073, 136.56772435, 2478.03511414, 16.44042379]
p_rotor_power_OFF = [191.83651588, 147.37205671, 5101.28555377, 376.62814798]
#Scaling model: Hub height (m) - Rated power (kW)
def func_height_power(x, a, b, c):
y = a - b*np.exp(-(x)/c)
return y
p_height_power_ON = [116.43035193, 91.64953366, 2391.88662558]
p_height_power_OFF = [120.75491612, 82.75390577, 4177.56520433]
#Scaling model: Nacelle weight (kg) - Rated power (kW)
def func_nacelle_weight_power(x, a, b):
y = a * x**2 + b*x
return 1e3 * y
p_nacelle_weight_power_ON = [ 1.66691134e-06, 3.20700974e-02]
p_nacelle_weight_power_OFF = [ 2.15668283e-06, 3.24712680e-02]
#Scaling model: Rotor weight (kg) - Rotor diameter (m)
def func_rotor_weight_rotor_diameter(x, a, b):
y = a * x**2 + b*x
return 1e3 * y
p_rotor_weight_rotor_diameter_ON = [ 0.00460956, 0.11199577]
p_rotor_weight_rotor_diameter_OFF = [ 0.0088365, -0.16435292]
#Scaling model: Tower (kg) - D²*h (m**3)
def func_tower_weight_d2h(d, h, a, b):
y = a * d**2*h + b
return 1e3 * y
p_tower_weight_d2h = [3.03584782e-04, 9.68652909e+00]
```
# Function to generate the wind turbine LCI
```
def create_dictionary(P, d = None, h = None, M_tower = None, M_foundation = None, M_reinfSteel_foundation = None,
V_conc_foundation = None, M_nacelle = None, M_power_supply = None, M_rotor = None,
M_electronics = None, offshore = False, park_size = 50, dist_transfo = 1, dist_coast = 5,
sea_depth = 5, print_details = False, lifetime = 20):
"""
This function generates the lifecycle inventory of a wind turbine
List of parameters : P (rated power) expressed in kW, d (rotor diameter) expressed in m, h (hub height) expressed in m,
offshore = True/False (False by default), park_size (number of wind turbines in a park) as integer,
dist_transfo (ditance to transformer) in meters, disct_coast (distance to coast for offshore) in meters,
sea_depth (for offshore) in meters, print_details (to print details) as boolean and lifetime (in years) as integer.
"""
#Setting parameters for scaling model with onshore-offshore distinctions.
if not(offshore):
p_rotor_power = p_rotor_power_ON
p_height_power = p_height_power_ON
p_nacelle_weight_power = p_nacelle_weight_power_ON
p_rotor_weight_rotor_diameter = p_rotor_weight_rotor_diameter_ON
if offshore :
p_rotor_power = p_rotor_power_OFF
p_height_power = p_height_power_OFF
p_nacelle_weight_power = p_nacelle_weight_power_OFF
p_rotor_weight_rotor_diameter = p_rotor_weight_rotor_diameter_OFF
#Using scaling model for missing values
if d == None:
d = func_rotor_power(P, *p_rotor_power)
if h == None:
h = func_height_power(P, *p_height_power)
if M_tower == None:
M_tower = func_tower_weight_d2h(d, h, *p_tower_weight_d2h)
if M_foundation == None:
M_foundation = 1696e3 * h/80 * d**2/(100**2)
if M_reinfSteel_foundation == None:
M_reinfSteel_foundation = np.interp(P, [750, 2000, 4500], [10210, 27000, 51900])
if V_conc_foundation == None:
V_conc_foundation = (M_foundation - M_reinfSteel_foundation) / 2200
if M_nacelle == None:
M_nacelle = func_nacelle_weight_power(P, *p_nacelle_weight_power)
if M_power_supply == None:
M_power_supply = 620
if M_rotor == None:
M_rotor = func_rotor_weight_rotor_diameter(d, *p_rotor_weight_rotor_diameter)
if M_electronics == None:
M_electronics = np.interp(P, [30, 150, 600, 800, 2000], [150, 300, 862, 1112, 3946])
M_all = M_nacelle + M_tower + M_rotor + M_foundation + M_electronics
#A reprendre avec tous les paramètres possibles
if print_details:
print('Nominal power : %s kW' %P)
print('Rotor diameter : %s m'%d)
print('Hub height : %s m'%h)
print('Tower weight : %s kg'%M_tower)
print('Foundation weight : %s kg'%M_foundation)
print('Foundation reinforced steel: %s kg'%M_reinfSteel_foundation)
print('Foundation concrete volume: %s m3'%V_conc_foundation)
print('Nacelle weight: %s kg'%M_nacelle)
print('Rotor weight: %s'%M_rotor)
print('Electronics: %s'%M_electronics)
print('Total weight: %s kg'%M_all)
dict_activities={}
#Adding input element expressed in percentage of mass * M
phase = 'Input'
if print_details:
print(phase)
for component in set(df_inv[phase].columns.get_level_values(0)):
if print_details:
print('\t' + component)
if component == 'Electronics':
M = M_electronics
if component == 'Foundation':
continue
if component == 'Nacelle':
M = M_nacelle
if component == 'Power supply':
M = M_power_supply
if component == 'Rotor':
M = M_rotor
if component == 'Tower':
M = M_tower
if print_details:
print(M)
for sub_comp in set(df_inv[phase][component].columns.get_level_values(0)):
if print_details:
print('\t \t' + sub_comp)
for dataset in set(df_inv[phase][component][sub_comp].columns.get_level_values(0)):
df_inv_i = df_inv[phase][component][sub_comp][dataset]
if print_details:
print('\t \t \t' + dataset + '\t M = %s'%M + '\t pctg = %s' %(np.interp(P, df_inv_i.index.values, df_inv_i.values)) + '\t %s'%(np.interp(P, df_inv_i.index.values, df_inv_i.values)*M))
print('\t \t \t'+ df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_i.index.values, df_inv_i.values) * M )
#Adding onshore foundation:
if not(offshore):
if print_details:
print('\t Onshore foundation')
print('\t \t M_fond = %s'%M_foundation)
print('\t \t concrete = %s' %V_conc_foundation + 'm3')
print('\t \t reinforced steel = %s' %M_reinfSteel_foundation + 'kg')
#V_conc_foundation
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, df_act[(df_act['Component'] == 'Foundation')&(df_act['Market name'] == 'market for concrete, sole plate and foundation')]['UUID'].iloc[0])), value = V_conc_foundation)
#M_reinfSteel_foundation
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, df_act[(df_act['Component'] == 'Foundation')&(df_act['Market name'] == 'market for waste reinforced concrete')]['UUID'].iloc[0])), value = M_reinfSteel_foundation)
#Adding road
if not(offshore):
value = np.interp(P, [0, 2000],[0, 8000])
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Road['code'])), value = value)
#Adding land use
if False:
phase = 'Input'
if print_details:
print('Land use')
if not(offshore):
for dataset in df_inv_not_kg['Input']['Foundation']['Land use']:
df_inv_not_kg_i = df_inv_not_kg['Input']['Foundation']['Land use'][dataset].dropna()
if print_details:
print('\t' + dataset + '\t %s'%(np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values)))
print('\t' + df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values))
#Maintenance
if False:
if print_details:
print('Maintenance')
dataset = 'Car [km]'
df_inv_not_kg_i = df_inv_not_kg['Maintenance']['Nacelle']['Transport by car'][dataset].dropna()
if print_details:
print('\t' + dataset + '\t %s'%(np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values)))
print('\t' + df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values))
#Assembly activities not in kg
phase = 'Assembly'
if print_details:
print(phase + 'not in kg')
for component in set(df_inv_not_kg[phase].columns.get_level_values(0)):
if print_details:
print('\t' + component)
for sub_comp in set(df_inv_not_kg[phase][component].columns.get_level_values(0)):
if print_details:
print('\t \t' + sub_comp)
for dataset in set(df_inv_not_kg[phase][component][sub_comp].columns.get_level_values(0)):
df_inv_not_kg_i = df_inv_not_kg[phase][component][sub_comp][dataset].dropna()
if print_details:
print(dataset)
if component == 'Foundation':
df_inv_not_kg_i = df_inv_not_kg[phase][component][sub_comp][dataset].dropna()
if print_details:
print('\t' + dataset + '\t %s'%(np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values)))
print('\t' + df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_not_kg_i.index.values, df_inv_not_kg_i.values))
else:
s = InterpolatedUnivariateSpline(df_inv_not_kg_i.index.values, df_inv_not_kg_i.values, k=1)
if print_details:
print('\t' + dataset + '\t %s'%(s(P)))
print('\t' + df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[((df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = s(P))
#Connexion_requirements
if offshore:
if print_details:
print('Raccordement offshore')
M, E_CLS = cable_requirements(P = P, park_size = park_size, dist_transfo = dist_transfo, dist_coast = dist_coast)
phase = 'Input'
component = 'Power supply'
for sub_comp in set(df_inv[phase][component].columns.get_level_values(0)):
if print_details:
print('\t \t' + sub_comp)
for dataset in set(df_inv[phase][component][sub_comp].columns.get_level_values(0)):
df_inv_i = df_inv[phase][component][sub_comp][dataset].dropna()
if print_details:
print('\t \t \t' + dataset + '\t %s'%(np.interp(P, df_inv_i.index.values, df_inv_i.values)*M))
print('\t \t \t'+ df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[((df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_i.index.values, df_inv_i.values) * M )
#Energy for cable laying ship
add_to_dict(dict_activities, key = diesel_burned_activity, value = E_CLS)
#Transfo
add_to_dict(dict_activities, key = MV_transfo, value = P / 10e3 / 0.85 * lifetime/35) #calcul au proratat de la puissance, facteur 0.85 en puissance active et apparente et durée de vie
add_to_dict(dict_activities, key = HV_transfo, value = P / 500e3 / 0.85 * lifetime/35)
# Pour les onshore, Transfo moyenne tension au proratat de la puissance, et cable section et longueur suivant la puissance
if not(offshore):
add_to_dict(dict_activities, key = MV_transfo, value = P / 10e3 / 0.85 * 19/35)
M = cable_requirements_Onshore(P)
phase = 'Input'
component = 'Power supply'
for sub_comp in set(df_inv[phase][component].columns.get_level_values(0)):
if print_details:
print('\t \t' + sub_comp)
for dataset in set(df_inv[phase][component][sub_comp].columns.get_level_values(0)):
df_inv_i = df_inv[phase][component][sub_comp][dataset].dropna()
if print_details:
print('\t \t \t' + dataset + '\t %s'%(np.interp(P, df_inv_i.index.values, df_inv_i.values)*M))
print('\t \t \t'+ df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[((df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)) & (df_act['Phase'] == phase)].iloc[0]['UUID'])), value = np.interp(P, df_inv_i.index.values, df_inv_i.values) * M )
#Transport activities
trsp_truck_nacelle, trsp_truck_rotor, trsp_truck_tower, trsp_ship_tower,trsp_truck_foundation, trsp_end_of_life, trsp_maintenance_per_year, trsp_ship_offshore = transport_requirements(M_nacelle, M_tower, M_rotor, M_foundation, M_all, lifetime)
#Truck transportation
trsp_truck = trsp_truck_nacelle + trsp_truck_rotor + trsp_truck_tower + trsp_truck_foundation + trsp_end_of_life + trsp_maintenance_per_year
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Truck_transport['code'])), value = trsp_truck)
#Ship transportation
if offshore:
trsp_ship = trsp_ship_tower + trsp_ship_offshore
else:
trsp_ship = trsp_ship_tower
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Ship_transport['code'])), value = trsp_ship)
#Scour stuff
if offshore:
scour_volume = fit_scour_volume(P)
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Digger['code'])), value = scour_volume)
#Grout stuff and monopile
if offshore:
m_grout, m_monopile = grout_and_monopile_requirements(P, sea_depth)
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Cement['code'])), value = m_grout)
phase = 'Input'
component = 'Tower'
sub_comp = 'Material'
M = m_monopile
for dataset in set(df_inv[phase][component][sub_comp].columns.get_level_values(0)):
df_inv_i = df_inv[phase][component][sub_comp][dataset].dropna()
if print_details:
print('\t \t \t' + dataset + '\t %s'%(np.interp(P, df_inv_i.index.values, df_inv_i.values)*M))
print('\t \t \t'+ df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['UUID'])
add_to_dict(dict_activities, key = bw.get_activity((df_act[(df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)].iloc[0]['Database name'], df_act[((df_act['Dataset'] == dataset) & (df_act['Phase'] == phase)) & (df_act['Phase'] == 'Input')].iloc[0]['UUID'])), value = np.interp(P, df_inv_i.index.values, df_inv_i.values) * M )
#Assembly depending on kg
for AA in assembly_activities:
if print_details:
print(AA)
if AA == 'Explosives':
value = 10
add_to_dict(dict_activities, key = Explosive, value = value )
if AA == 'Copper wire drawing':
value = dict_activities[ bw.get_activity((eidb.name, df_act[ df_act['Dataset'] == 'Copper']['UUID'].iloc[0]))]
add_to_dict(dict_activities, key = Copper_wire_drawing, value = value )
if AA == 'Steel sheet rolling':
value = dict_activities[ bw.get_activity((eidb.name, df_act[ df_act['Dataset'] == 'Low-alloy steel']['UUID'].iloc[0]))] + dict_activities[bw.get_activity((eidb.name, df_act[ df_act['Dataset'] == 'Cast iron']['UUID'].iloc[0]))]
add_to_dict(dict_activities, key = Steel_sheet_rolling, value = value )
if AA == 'Aluminium sheet rolling':
value = dict_activities[ bw.get_activity((eidb.name, df_act[ df_act['Dataset'] == 'Aluminium 0% recycled']['UUID'].iloc[0]))]
add_to_dict(dict_activities, key = Aluminium_sheet_rolling, value = value )
if AA == 'Chromium sheet rolling':
value = dict_activities[ bw.get_activity((eidb.name, df_act[ df_act['Dataset'] == 'Chromium steel']['UUID'].iloc[0]))]
add_to_dict(dict_activities, key = Chromium_sheet_rolling, value = value )
#Disposal activities
for DA in disposal_activities:
if print_details:
print(DA)
ei_key_disposal = df_act[ (df_act['Dataset'] == DA) & (df_act['Phase'] == 'Disposal')]['UUID'].iloc[0]
DA = DA.replace(' -waste','').replace('Steel, inert waste', 'Low-alloy steel' ).replace('Concrete, inert waste', 'Concrete [m3]').replace('Aluminium waste', 'Aluminium 0% recycled').replace('Chromium Steel waste','Chromium steel')
ei_key_input = df_act[ (df_act['Dataset'] == DA) & (df_act['Phase'] == 'Input')]['UUID'].iloc[0]
try:
value = - dict_activities[ bw.get_activity((eidb.name, ei_key_input))]
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, ei_key_disposal)), value = value)
except:
pass
#Electicity dataset
value = 0.5 * (M_nacelle + M_rotor + M_tower)
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Electricity_dataset['code'])), value = value)
#Steel dataset
value = dict_activities[bw.get_activity((eidb.name, Steel_dataset['code']))]
del dict_activities[bw.get_activity((eidb.name, Steel_dataset['code']))]
add_to_dict(dict_activities, key = bw.get_activity((eidb.name, Steel_dataset['code'])), value = value)
return dict_activities
```
For example, here we create the inventory of a 2 MW onshore wind turbine
```
dict_activities = create_dictionary(P = 2000, print_details = True)
```
We print the dicitonnary that contains the inventory, in which quantities and corresponding datasets are listed.
```
dict_activities
```
# Calculation with Brightway2 !
### Methode d'évaluation d'impact
```
#Huge set of methods
#Acidification
Acid_method = [m for m in bw.methods if 'acidification' in str(m) and 'ILCD' in str(m) and not 'no LT' in str(m)][0]
#Climate change
CC_method = [m for m in bw.methods if 'IPCC 2013' in str(m) and 'climate change' in str(m) and 'GWP 100a' in str(m) and not 'no LT' in str(m) ][0]
#Eutrophication
Eutro_method = [m for m in bw.methods if 'freshwater eutrophication' in str(m) and 'ILCD' in str(m) and not 'no LT' in str(m)][0]
#Ecotoxicity
Ecotox_method = [m for m in bw.methods if 'freshwater ecotoxicity' in str(m) and 'ILCD' in str(m) and not 'no LT' in str(m)][0]
#CED
#non renewable
CEDfossil_method = [m for m in bw.methods if 'cumulative energy demand' in str(m) and 'non-renewable energy resources, fossil' in str(m)][0]
CEDnuclear_method = [ m for m in bw.methods if 'cumulative energy demand' in str(m) and 'nuclear' in str(m) ][0]
CEDbioNR_method = [ m for m in bw.methods if 'cumulative energy demand' in str(m) and 'non-renewable energy resources, primary forest' in str(m) ][0]
#renewable
CEDbiomR_method = [ m for m in bw.methods if 'cumulative energy demand' in str(m) and 'renewable energy resources, biomass' in str(m) ][0]
CEDgeoth_method = [ m for m in bw.methods if 'renewable energy resources, geothermal, converted' in str(m) and '' in str(m) ][0]
CEDsolar_method = [ m for m in bw.methods if 'renewable energy resources, solar, converted' in str(m) and '' in str(m) ][0]
CEDwater_method = [ m for m in bw.methods if 'renewable energy resources, potential (in barrage water), converted' in str(m) and '' in str(m) ][0]
CEDwind_method = [ m for m in bw.methods if 'renewable energy resources, kinetic (in wind), converted' in str(m) and '' in str(m) ][0]
#Land use
Land_method = [m for m in bw.methods if 'land' in str(m) and 'ILCD' in str(m) and not 'no LT' in str(m)][0]
#Depletion
Fossildeplet_method = [m for m in bw.methods if 'fossil depletion' in str(m) and 'resource' in str(m) and 'H' in str(m) and not 'w/o LT' in str(m)][0]
Metaldeplet_method = [m for m in bw.methods if 'metal depletion' in str(m) and 'resource' in str(m) and 'H' in str(m) and not 'w/o LT' in str(m)][0]
Deplet_method = [ m for m in bw.methods if 'ILCD' in str(m) and 'resource' in str(m) and 'mineral, fossils and renewables' in str(m) and not 'no LT' in str(m)][0]
#Ozone
Ozone_method = [m for m in bw.methods if 'ozone layer depletion' in str(m) and 'ILCD' in str(m) and not 'no LT' in str(m)][0]
#Human health
HH_carcinogenic_method = [m for m in bw.methods if 'carcinogenic effects' in str(m) and '' in str(m) and 'ILCD' in str(m) and not 'LT' in str(m) and not '-' in str(m)][0]
HH_noncarcinogenic_method = [m for m in bw.methods if '-carcinogenic effects' in str(m) and '' in str(m) and 'ILCD' in str(m) and not 'LT' in str(m)][0]
#PM emissions : ILCD
PM_method = [ m for m in bw.methods if 'ILCD' in str(m) and 'human health' in str(m) and 'respiratory effects, inorganics' in str(m)][0]
#Water
Water_method = [m for m in bw.methods if 'water' in str(m) and 'scarcity' in str(m) and 'water resources' in str(m) and not 'no LT' in str(m)][0]
list_methods = [Acid_method, CC_method, Eutro_method, Ecotox_method, Land_method, Fossildeplet_method, Metaldeplet_method, Deplet_method, Ozone_method, HH_carcinogenic_method, HH_noncarcinogenic_method, Water_method, PM_method, CEDfossil_method, CEDnuclear_method, CEDbioNR_method, CEDbiomR_method, CEDgeoth_method, CEDsolar_method, CEDwater_method, CEDwind_method]
print('set_of_methods: \n %s \n' %list_methods)
```
We start a LCA calculation that will calculate the impacts for all the methods contained in list_methods for as many wind turbines as contained in the dicitonnary dict_activities (in this case, one).
```
bw.calculation_setups['multiLCA'] = {'inv': [dict_activities], 'ia': list_methods}
bw.calculation_setups['multiLCA']
myMultiLCA = bw.MultiLCA('multiLCA')
myMultiLCA.results
df_impact = pd.DataFrame(columns = list_methods, data = myMultiLCA.results)
df_impact
```
We can formulate a calculation function that takes directly the attributes of the wind turbine as arguments and normalize the life cycle impacts by the electricity production, a given life time and load factor.
```
def calculate_impact(dict_activities = None, P = None, lifetime = 20, load_factor = 0.3, list_methods = list_methods):
bw.calculation_setups['multiLCA'] = {'inv': [dict_activities], 'ia': list_methods}
bw.calculation_setups['multiLCA']
myMultiLCA = bw.MultiLCA('multiLCA')
myMultiLCA.results
df_impact = pd.DataFrame(columns = list_methods, data = myMultiLCA.results)
df_impact
df_impact_kWh = df_impact / (P * lifetime * 8760 * load_factor)
return df_impact, df_impact_kWh
df_impact, df_impact_kWh = calculate_impact(dict_activities, P = 2000)
#Impacts per life cycle
df_impact
#Impacts per kWh
df_impact_kWh
```
### Visualization
There are many ways to visualize such results.
Here, we take the per kWh LCA results calculated for all currently operating wind turbines in Denmark.
Then we use Matplotlib's map library Basemap to plot them. Another very good library for static maps is Geopandas.
```
df_all=pd.read_csv("Complete_dataframe.csv")
df_all["Carbon footprint per kWh"]=df_all["('IPCC 2013', 'climate change', 'GWP 100a')"]/df_all.loc[:,"1977":"2050"].sum(axis=1)*1000
df_all["Carbon footprint per kWh"].head()
from mpl_toolkits.basemap import Basemap
from matplotlib import cm
plt.rcParams.update({'font.size': 16})
import matplotlib.colors as mcolors
cdict = {'red': ((0.0, 0.0, 0.0),
(0.5, 0.0, 0.0),
(1.0, 1.0, 1.0)),
'blue': ((0.0, 0.0, 0.0),
(1.0, 0.0, 0.0)),
'green': ((0.0, 0.0, 1.0),
(0.5, 0.0, 0.0),
(1.0, 0.0, 0.0))}
color_map = mcolors.LinearSegmentedColormap('my_colormap', cdict, 100)
plt.figure(figsize=(14,16))
my_map = Basemap(projection='merc', lat_0=56, lon_0=11,
resolution='h', area_thresh=0.1,
llcrnrlon = 7, llcrnrlat = 54,
urcrnrlon=13, urcrnrlat=58)
my_map.drawcoastlines()
my_map.drawcountries(color="grey", zorder=1)
my_map.fillcontinents(color='grey', zorder=1, alpha=.1)
my_map.drawmapboundary(fill_color="white")
#We filter out teh really bad wind tubrines with a too low load factor
df_data=df_all[(df_all["Estimated load factor"]>0.1)]
data=np.array(df_data["Carbon footprint per kWh"])
p_max=max(data)
x,y = my_map(df_data["lon"].tolist(), df_data["lat"].tolist())
my_map.scatter(x, y, marker='o', edgecolor=None, zorder=1, cmap=color_map, c=color_map(df_data['Carbon footprint per kWh']/1000))
l1 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(5), cmap=color_map)
l2 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(10), cmap=color_map)
l3 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(20), cmap=color_map)
l4 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(30), cmap=color_map)
l5 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(40), cmap=color_map)
l6 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(50), cmap=color_map)
l7 = plt.scatter([],[], s=100, edgecolor='black', linewidth='0.1', color=color_map(100), cmap=color_map)
labels = ["5 g", "10 g", "20 g", "30 g", "40 g", "50 g", "100 g"]
leg = plt.legend([l1, l2, l3, l4, l5, l6, l7], labels, ncol=7, frameon=True, fontsize=16,
handlelength=2, loc = 8, borderpad = 1, handletextpad=.2, title='$CO_2$ [g] per kWh produced', scatterpoints = 1)
plt.title('Carbon footprint per kWh', fontsize = 40)
plt.show()
```
Alternatively, you can serve interactive maps with Leaflet, which is what was done with [LCA_WIND_DK](http://viewer.webservice-energy.org/lca-wind-dk/) :-)
| github_jupyter |
# Deploying a MedNIST Classifier with Ray
This notebook demos the process of deploying a network with Ray Serve as a web service. Ray provides various ways of deploying models with existing platforms like AWS or Azure but we'll focus on local deployment here since researchers are more likely to do this. Ray also provides other libraries for tuning, reinforcement learning, and distributed training in addition to deployment. This tutorial will use MedNIST classifier from the BentoML tutorial so please run at least the training component of that notebook first. The documentation on Ray Serve [start here](https://docs.ray.io/en/master/serve/index.html#rayserve), this notebook will be using the Pytorch specific functionality [discussed here](https://docs.ray.io/en/master/serve/tutorials/pytorch.html).
To start install the Ray Serve component:
```
%pip install ray[serve]
!python -c "import monai" || pip install -q "monai-weekly[gdown, tqdm]"
```
The imports for MONAI are the same as for the BentoML tutorial (assuming it's already installed):
```
import os
import io
from PIL import Image
import torch
import numpy as np
import requests
import ray
from ray import serve
from monai.apps import download_url
from monai.config import print_config
from monai.transforms import (
AddChannel,
Compose,
ScaleIntensity,
EnsureType,
)
print_config()
resource = "https://drive.google.com/uc?id=1zKRi5FrwEES_J-AUkM7iBJwc__jy6ct6"
dst = os.path.join("..", "bentoml", "classifier.zip")
if not os.path.exists(dst):
download_url(resource, dst)
```
This class will represent the service for the model, which accepts an image sent as the body of a POST request and returns the class name in a JSON structure. Note that this class uses asyncio to define the `__call__` to be compatible with the server backend.
```
MEDNIST_CLASSES = ["AbdomenCT", "BreastMRI", "CXR", "ChestCT", "Hand", "HeadCT"]
@serve.deployment
class MedNISTClassifier:
def __init__(self):
# create the transform for normalizing the image data
self.transform = Compose([AddChannel(), ScaleIntensity(), EnsureType()])
# load the network on the CPU for simplicity and in eval mode
self.net = torch.jit.load("../bentoml/classifier.zip", map_location="cpu").eval()
async def __call__(self, request):
image_bytes = await request.body()
img = Image.open(io.BytesIO(image_bytes))
img = np.array(img)
image_tensor = self.transform(img)
with torch.no_grad():
outputs = self.net(image_tensor[None].float())
_, output_classes = outputs.max(dim=1)
return {"class_index": MEDNIST_CLASSES[output_classes[0]]}
```
Now the server is started and the classifier backend is associated with an endpoint, which is the route to the service relate to the server address.
```
serve.start()
MedNISTClassifier.deploy()
```
With the server running in another process we can send it a query with an image and get a response. By default the server will listen on port 8000.
```
image_bytes = open("./hand.jpg", "rb").read()
resp = requests.post("http://localhost:8000/MedNISTClassifier", data=image_bytes)
print(resp.json())
```
This can also be done on the command line with `curl`:
```
!curl -X POST "http://localhost:8000/MedNISTClassifier" --data-binary "@hand.jpg"
```
Finally shut down the server:
```
ray.shutdown()
```
### Command Line Usage
Ray can be started on the command line. Since it operates as a cluster of nodes the first thing to do is create the head node locally then start the serve component:
```
%%bash
ray start --head
serve start
```
A separate script with very similar code can then be used to add or replace the backend. This would be useful in an experimental setting where the server is running constantly in the background to which you can push updates quickly as you edit your script.
```
%%writefile mednist_classifier_start.py
import io
from PIL import Image
import torch
import numpy as np
import ray
from ray import serve
from monai.config import print_config
from monai.transforms import (
AddChannel,
Compose,
ScaleIntensity,
EnsureType,
)
MEDNIST_CLASSES = ["AbdomenCT", "BreastMRI", "CXR", "ChestCT", "Hand", "HeadCT"]
ray.init(address="auto", namespace="serve")
@serve.deployment
class MedNISTClassifier:
def __init__(self):
self.transform = Compose([AddChannel(), ScaleIntensity(), EnsureType()])
self.net = torch.jit.load("../bentoml/classifier.zip", map_location="cpu").eval()
async def __call__(self, request):
image_bytes = await request.body()
img = Image.open(io.BytesIO(image_bytes))
img = np.array(img)
image_tensor = self.transform(img)
with torch.no_grad():
outputs = self.net(image_tensor[None].float())
_, output_classes = outputs.max(dim=1)
return {"class_index": MEDNIST_CLASSES[output_classes[0]]}
MedNISTClassifier.deploy()
# ray.init(address="auto")
# client = serve.connect()
# # remove previous instance of this backend if present
# if "classifier" in client.list_backends():
# client.delete_endpoint("classifier")
# client.delete_backend("classifier")
# client.create_backend("classifier", MedNISTClassifier)
# client.create_endpoint("classifier", backend="classifier", route="/image_classify", methods=["POST"])
```
The endpoint is then added by running the script:
```
%%bash
python mednist_classifier_start.py
```
And checked once again for response:
```
!curl -X POST "http://localhost:8000/MedNISTClassifier" --data-binary "@hand.jpg"
```
Finally the service can be stopped:
```
%%bash
ray stop
```
| github_jupyter |
# Plotting the DC2 Run1.1p skyMap
<br>Owner: **Jim Chiang** ([@jchiang87](https://github.com/LSSTDESC/DC2-analysis/issues/new?body=@jchiang87))
<br>Last Verified to Run: **2018-10-26** (by @yymao)
In this notebook, we show how to use the data butler to obtain information on the skyMap used in the coadd analyses performed by the DRP pipeline. These skyMaps are composed of tracts and patches on the sky. Each tract is a rectangular region of the sky with a common map projection; each tract is further divided into rectangular patches, which use the same tract coordinate system and which are a convenient size for processing the coadd data. A more complete description of the skyMap geometry is given in the HSC Software Pipeline paper ([Bosch et al. 2017](https://arxiv.org/abs/1705.06766)).
Equipped with the info from the skyMap, we plot the tracts and patches that were used with the Run1.1p processing and overlay the WFD and uDDF simulation regions. We also use the butler to access the visit-level data and show how one can access the calexp (calibrated exposure) image data to obtain the PSF, zero-point, etc. as measured by the Stack for a given exposure. Finally, we show how to plot the sky region imaged on the focal plane for a given visit in two ways: the first using the CCD coordinates available from the calexps and the other using the lsst_sims code to compute those coordinates from the pointing information for the visit.
## Set Up
```
import os
import glob
import warnings
import sqlite3
import re
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.path import Path
import matplotlib.patches as patches
%matplotlib inline
import lsst.afw.geom as afw_geom
import lsst.afw.cameraGeom as cameraGeom
import lsst.daf.persistence as dp
# The lsst_sims code issues some ignorable warnings regarding ids used for querying the object
# databases.
with warnings.catch_warnings():
warnings.simplefilter("ignore")
import lsst.sims.coordUtils
from lsst.sims.catUtils.utils import ObservationMetaDataGenerator
from lsst.sims.utils import getRotSkyPos
from desc_dc2_dm_data import REPOS
```
## Plotting Functions
First, we define some functions to plot the tract, patch, and CCD regions on the sky. These are copied from [example code](https://github.com/yalsayyad/dm_notebooks/blob/master/desc-ssim/DESC-SSim%20Patch%20Geometry.ipynb) that Yusra AlSayyad presented at the [2017-06-29 SSim meeting](https://confluence.slac.stanford.edu/pages/viewpage.action?pageId=224461017).
```
def make_patch(vertexList, wcs=None):
"""
Return a Path in sky coords from vertex list in pixel coords.
Parameters
----------
vertexList: list of coordinates
These are the corners of the region to be plotted either in pixel coordinates or
sky coordinates.
wcs: lsst.afw.geom.skyWcs.skyWcs.SkyWcs [None]
The WCS object used to convert from pixel to sky coordinates.
Returns
-------
matplotlib.path.Path: The encapsulation of the vertex info that matplotlib uses to
plot a patch.
"""
if wcs is not None:
skyPatchList = [wcs.pixelToSky(pos).getPosition(afw_geom.degrees)
for pos in vertexList]
else:
skyPatchList = vertexList
verts = [(coord[0], coord[1]) for coord in skyPatchList]
verts.append((0,0))
codes = [Path.MOVETO,
Path.LINETO,
Path.LINETO,
Path.LINETO,
Path.CLOSEPOLY,
]
return Path(verts, codes)
def plot_skymap_tract(skyMap, tract=0, title=None, ax=None):
"""
Plot a tract from a skyMap.
Parameters
----------
skyMap: lsst.skyMap.SkyMap
The SkyMap object containing the tract and patch information.
tract: int [0]
The tract id of the desired tract to plot.
title: str [None]
Title of the tract plot. If None, the use `tract <id>`.
ax: matplotlib.axes._subplots.AxesSubplot [None]
The subplot object to contain the tract plot. If None, then make a new one.
Returns
-------
matplotlib.axes._subplots.AxesSubplot: The subplot containing the tract plot.
"""
if title is None:
title = 'tract {}'.format(tract)
tractInfo = skyMap[tract]
tractBox = afw_geom.Box2D(tractInfo.getBBox())
tractPosList = tractBox.getCorners()
wcs = tractInfo.getWcs()
xNum, yNum = tractInfo.getNumPatches()
if ax is None:
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
tract_center = wcs.pixelToSky(tractBox.getCenter())\
.getPosition(afw_geom.degrees)
ax.text(tract_center[0], tract_center[1], '%d' % tract, size=16,
ha="center", va="center", color='blue')
for x in range(xNum):
for y in range(yNum):
patchInfo = tractInfo.getPatchInfo([x, y])
patchBox = afw_geom.Box2D(patchInfo.getOuterBBox())
pixelPatchList = patchBox.getCorners()
path = make_patch(pixelPatchList, wcs)
patch = patches.PathPatch(path, alpha=0.1, lw=1)
ax.add_patch(patch)
center = wcs.pixelToSky(patchBox.getCenter())\
.getPosition(afw_geom.degrees)
ax.text(center[0], center[1], '%d,%d'%(x,y), size=6,
ha="center", va="center")
skyPosList = [wcs.pixelToSky(pos).getPosition(afw_geom.degrees)
for pos in tractPosList]
ax.set_xlim(max(coord[0] for coord in skyPosList) + 1,
min(coord[0] for coord in skyPosList) - 1)
ax.set_ylim(min(coord[1] for coord in skyPosList) - 1,
max(coord[1] for coord in skyPosList) + 1)
ax.grid(ls=':',color='gray')
ax.set_xlabel("RA (deg.)")
ax.set_ylabel("Dec (deg.)")
ax.set_title(title)
return ax
```
The following function loops over the available calexps as returned by the data butler to determine which CCDs to draw. Unfortunately, looping over those calexps using the butler is rather slow, but would be necessary if we wanted to access CCD-level information, like the PSF, from the calexps.
We include this function here for your edification, but then provide a faster function below.
```
def plot_focal_plane(butler, visit, ax, color='red'):
"""
Plot the CCDs in the LSST focal plane using the coordinate information in the calexps.
Notes
-----
By looping over the available calexps, we only plot the CCDs for which image data
are available.
Parameters
----------
butler: lsst.daf.persistence.Butler
The data butler serving up data from the desired repo.
visit: int
The visit or obsHistID number.
ax: matplotlib.axes._subplots.AxesSubplot
The matplotlib subplot object onto which to plot the focal plane.
color: str ['red']
Color to use for plotting the individual CCDs.
Returns
-------
matplotlib.axes._subplots.AxesSubplot: The subplot object used for plotting.
"""
# We use the `subset` method to obtain all of the `datarefs` (i.e., references to calexp
# data in this case) that satisfy an "incomplete" dataId. For visit-level calexp data,
# a unique dataset would specify visit, raft, and sensor. If we just give the visit, then
# references to the available data for all of the CCDs would be returned.
dataId = dict(visit=visit)
datarefs = list(butler.subset('calexp', dataId=dataid))
for i, dataref in enumerate(datarefs):
calexp = dataref.get('calexp')
# We're not going to do anything with it here, but we can get the PSF from the calexp
# like this:
# psf = calexp.getPsf()
# and we can get the zero-point (in ADU) like this
# zero_point = calexp.getCalib().getFluxMag0()
ccd_box = afw_geom.Box2D(calexp.getBBox())
wcs = calexp.getWcs()
path = make_patch(ccd_box.getCorners(), wcs)
ccd = patches.PathPatch(path, alpha=0.2, lw=1, color=color)
ax.add_patch(ccd)
center = wcs.pixelToSky(ccd_box.getCenter()).getPosition(afw_geom.degrees)
return ax
```
The following fast version of the focal plane plotting code uses the lsst_sims package to obtain the location and orientation of the CCDs based on the pointing information for the desired visit. That pointing information is extracted from the dithered minion_1016 OpSim db that has been prepared for DC2. Since this code does not access the individual calexps for each CCD, it runs much faster. However, it assumes that the obs_lsstSim package was used in the analysis of the data, and it needs to use the inferred locations of the calexp files to determine if calexp data for a given CCD is available.
```
def plot_focal_plane_fast(butler, visit, ax, color='red', opsimdb=None):
"""
Plot the CCDs in the LSST focal plane using CCD coordinates derived from the pointing
info using the lsst.sims code.
Notes
-----
This function assumes that the obs_lsstSims package was used to define the camera geometry
for the analysis of the simulated image data.
Parameters
----------
butler: lsst.daf.persistence.Butler
The data butler serving up data from the desired repo.
visit: int
The visit or obsHistID number.
ax: matplotlib.axes._subplots.AxesSubplot
The matplotlib subplot object onto which to plot the focal plane.
color: str ['red']
Color to use for plotting the individual CCDs.
opsimDb: str [None]
Filename of the OpSim sqlite database. If None, then the dithered opsim db for Run1.1p
is used.
Returns
-------
matplotlib.axes._subplots.AxesSubplot: The subplot object used for plotting.
"""
if opsimdb is None:
opsimdb = '/global/projecta/projectdirs/lsst/groups/SSim/DC2/minion_1016_desc_dithered_v4.db'
conn = sqlite3.connect(opsimdb)
obs_gen = ObservationMetaDataGenerator(database=opsimdb, driver='sqlite')
# The dithered pointing info was added to the baseline minion_1016 db. We query for the values
# used for the desired visit.
curs = conn.execute('''select descDitheredRA, descDitheredDec, descDitheredRotTelPos
from summary where obshistid={}'''.format(visit))
ra, dec, rottelpos = [np.degrees(x) for x in curs][0]
# An ObservationMetaData object used to pass the pointing info to the function in
# lsst.sims.coordUtils that provides the CCD coordinates.
obs_md = obs_gen.getObservationMetaData(obsHistID=visit, boundType='circle', boundLength=0.1)[0]
obs_md.pointingRA = ra
obs_md.pointingDec = dec
obs_md.OpsimMetaData['rotTelPos'] = rottelpos
# Convert the rotation angle of the sky relative to the telescope to the sky angle relative to
# the camera.
obs_md.rotSkyPos = getRotSkyPos(ra, dec, obs_md, rottelpos)
# Use the butler to get the camera appropriate for this observation. If the data were from a
# different camera, e.g., DECam or HSC, the corresponding camera objects with the associated
# CCD geometries would be returned.
camera = butler.get('camera')
# Grab one of the calexps via its dataref so that we can ask for its filename and thereby infer
# the location on disk of all of the calexps for this visit.
dataref = list(butler.subset('calexp', visit=visit))[0]
calexp_path = os.path.dirname(os.path.dirname(dataref.get('calexp_filename')[0]))
# The following code is specific to the obs_lsstSim package and how it names CCDs
# (e.g., "R:2,2 S:1,1") and formulates the path components for writing to disk. This
# code would not work for a different obs_ package/camera implementation.
# Re-order the CCD vertex list returned by the lsst_sims code so that a rectangle is plotted.
corner_index = (np.array([0, 1, 3, 2]),)
for det in camera:
# Skip the WAVEFRONT and GUIDER CCDs
if det.getType() != cameraGeom.SCIENCE:
continue
detname = det.getName()
raft, sensor = re.match(r'R:?(\d,?\d)[_ ]S:?(\d,?\d)', detname).groups()
raft = 'R' + raft.replace(',', '')
sensor = 'S{}.fits'.format(sensor.replace(',', ''))
if os.path.isfile(os.path.join(calexp_path, raft, sensor)):
corners = np.array(lsst.sims.coordUtils.getCornerRaDec(detname, camera, obs_md))
path = make_patch(corners[corner_index])
ccd = patches.PathPatch(path, alpha=0.2, lw=1, color=color)
ax.add_patch(ccd)
return ax
```
The following function just plots the boundaries of the Run1.1p regions as described in the [Run 1.1p Specifications document](https://docs.google.com/document/d/1aQOPL9smeDlhtlwDrp39Zuu2q8DKivDaHLQX3_omwOI/edit).
```
def plot_Run1_1p_region(ax):
"""Plot the WFD and uDDF regions for Run1.1p."""
uddf_ra = [53.764, 52.486, 52.479, 53.771, 53.764]
uddf_dec = [-27.533, -27.533, -28.667, -28.667, -27.533]
wfd_ra = [52.25, 52.11, 58.02, 57.87, 52.25]
wfd_dec = [-27.25, -32.25, -32.25, -27.25, -27.25]
ax.errorbar(wfd_ra, wfd_dec, fmt='-', color='green',
label='protoDC2 WFD boundary')
ax.errorbar(uddf_ra, uddf_dec, fmt='-', color='red',
label='uDDF boundary')
return ax
```
## Making the Plot
We now use the above functions to plot the tracts and patches on the sky.
```
repo = REPOS['1.2p']
butler = dp.Butler(repo)
# Single visit to overlay:
visit = 219976
# WARNING: this is a hack to search the coadd folder for the tracts that have data.
# Unfortunately, this information is not directly accessible from the data butler.
ref_path = butler.getUri('deepCoadd_forced_src', tract=4851, patch='0,0', filter='i')
ref_path, success, _ = ref_path.partition('/4851/0,0')
if not success:
ref_path = ref_path.partition('/i_t4851_p0,0')[0]
tracts = sorted(set((int(re.search(r'(\d{4})', t).groups()[0]) for t in os.listdir(ref_path) if os.path.isdir(os.path.join(ref_path, t)))))
# First, loop over all the tracts, plotting them as gray, numbered, rectangles:
ax = None
skyMap = butler.get('deepCoadd_skyMap')
for tract in tracts:
ax = plot_skymap_tract(skyMap, tract=tract, title='', ax=ax)
# Now overlay a single focal plane, for the chosen visit, in violet:
plot_focal_plane_fast(butler, visit, ax, color="violet")
# Finally, overlay the Run 1.1p main survey and uDDF regions, and then adjust the axes:
plot_Run1_1p_region(ax)
ax.set_xlim(60.5, 50.0)
ax.set_ylim(-33.5, -25.5)
plt.legend(loc=0);
```
| github_jupyter |
# Part 3 : Mitigate Bias, Train another unbiased Model and Put in the Model Registry
<a id='aup-overview'></a>
## [Overview](./0-AutoClaimFraudDetection.ipynb)
* [Notebook 0 : Overview, Architecture and Data Exploration](./0-AutoClaimFraudDetection.ipynb)
* [Notebook 1: Data Prep, Process, Store Features](./1-data-prep-e2e.ipynb)
* [Notebook 2: Train, Check Bias, Tune, Record Lineage, and Register a Model](./2-lineage-train-assess-bias-tune-registry-e2e.ipynb)
* **[Notebook 3: Mitigate Bias, Train New Model, Store in Registry](./3-mitigate-bias-train-model2-registry-e2e.ipynb)**
* **[Architecture](#train2)**
* **[Develop a second model](#second-model)**
* **[Analyze the Second Model for Bias](#analyze-second-model)**
* **[View Results of Clarify Bias Detection Job](#view-second-clarify-job)**
* **[Configure and Run Clarify Explainability Job](#explainability)**
* **[Create Model Package for the Second Trained Model](#model-package)**
* [Notebook 4: Deploy Model, Run Predictions](./4-deploy-run-inference-e2e.ipynb)
* [Notebook 5 : Create and Run an End-to-End Pipeline to Deploy the Model](./5-pipeline-e2e.ipynb)
In this notebook, we will describe how to detect bias using Clarify, Mitigate it with SMOTE, train another model, put it in the Model Registry along with all the Lineage of the Artifacts created along the way: data, code and model metadata.
### Install required and/or update third-party libraries
```
!python -m pip install -Uq pip
!python -m pip install -q awswrangler==2.2.0 imbalanced-learn==0.7.0 sagemaker==2.23.1 boto3==1.16.48
```
### Import libraries
```
import json
import time
import boto3
import sagemaker
import numpy as np
import pandas as pd
import awswrangler as wr
import matplotlib.pyplot as plt
from imblearn.over_sampling import SMOTE
from sagemaker.xgboost.estimator import XGBoost
from model_package_src.inference_specification import InferenceSpecification
%matplotlib inline
```
### Load stored variables
Run the cell below to load any prevously created variables. You should see a print-out of the existing variables. If you don't see anything you may need to create them again or it may be your first time running this notebook.
```
%store -r
%store
```
**<font color='red'>Important</font>: You must have run the previous sequential notebooks to retrieve variables using the StoreMagic command.**
### Set region, boto3 and SageMaker SDK variables
```
# You can change this to a region of your choice
import sagemaker
region = sagemaker.Session().boto_region_name
print("Using AWS Region: {}".format(region))
boto3.setup_default_session(region_name=region)
boto_session = boto3.Session(region_name=region)
s3_client = boto3.client("s3", region_name=region)
sagemaker_boto_client = boto_session.client("sagemaker")
sagemaker_session = sagemaker.session.Session(
boto_session=boto_session, sagemaker_client=sagemaker_boto_client
)
sagemaker_role = sagemaker.get_execution_role()
account_id = boto3.client("sts").get_caller_identity()["Account"]
# variables used for parameterizing the notebook run
model_2_name = f"{prefix}-xgboost-post-smote"
train_data_upsampled_s3_path = f"s3://{bucket}/{prefix}/data/train/upsampled/train.csv"
bias_report_2_output_path = f"s3://{bucket}/{prefix}/clarify-output/bias-2"
explainability_output_path = f"s3://{bucket}/{prefix}/clarify-output/explainability"
train_instance_count = 1
train_instance_type = "ml.m4.xlarge"
claify_instance_count = 1
clairfy_instance_type = "ml.c5.xlarge"
```
<a id ='train2'> </a>
## Architecture for this ML Lifecycle Stage : Train, Check Bias, Tune, Record Lineage, Register Model
[overview](#aup-overview)
----

<a id='second-model'></a>
## Develop a second model
[overview](#aup-overview)
----
In this second model, you will fix the gender imbalance in the dataset using SMOTE and train another model using XGBoost. This model will also be saved to our registry and eventually approved for deployment.
```
train = pd.read_csv("data/train.csv")
test = pd.read_csv("data/test.csv")
train
test
```
<a id='smote'></a>
### Resolve class imbalance using SMOTE
To handle the imbalance, we can over-sample (i.e. upsample) the minority class using [SMOTE (Synthetic Minority Over-sampling Technique)](https://arxiv.org/pdf/1106.1813.pdf). After installing the imbalanced-learn module, if you receive an ImportError when importing SMOTE, then try restarting the kernel.
#### Gender balance before SMOTE
```
gender = train["customer_gender_female"]
gender.value_counts()
```
#### Gender balance after SMOTE
```
sm = SMOTE(random_state=42)
train_data_upsampled, gender_res = sm.fit_resample(train, gender)
train_data_upsampled["customer_gender_female"].value_counts()
```
### Train new model
```
train_data_upsampled.to_csv("data/upsampled_train.csv", index=False)
s3_client.upload_file(
Filename="data/upsampled_train.csv",
Bucket=bucket,
Key=f"{prefix}/data/train/upsampled/train.csv",
)
xgb_estimator = XGBoost(
entry_point="xgboost_starter_script.py",
hyperparameters=hyperparameters,
role=sagemaker_role,
instance_count=train_instance_count,
instance_type=train_instance_type,
framework_version="1.0-1",
)
if 'training_job_2_name' not in locals():
xgb_estimator.fit(inputs = {'train': train_data_upsampled_s3_path})
training_job_2_name = xgb_estimator.latest_training_job.job_name
%store training_job_2_name
else:
print(f'Using previous training job: {training_job_2_name}')
```
### Register artifacts
```
training_job_2_info = sagemaker_boto_client.describe_training_job(
TrainingJobName=training_job_2_name
)
```
#### Code artifact
```
# return any existing artifact which match the our training job's code arn
code_s3_uri = training_job_2_info["HyperParameters"]["sagemaker_submit_directory"]
list_response = list(
sagemaker.lineage.artifact.Artifact.list(
source_uri=code_s3_uri, sagemaker_session=sagemaker_session
)
)
# use existing arifact if it's already been created, otherwise create a new artifact
if list_response:
code_artifact = list_response[0]
print(f"Using existing artifact: {code_artifact.artifact_arn}")
else:
code_artifact = sagemaker.lineage.artifact.Artifact.create(
artifact_name="TrainingScript",
source_uri=code_s3_uri,
artifact_type="Code",
sagemaker_session=sagemaker_session,
)
print(f"Create artifact {code_artifact.artifact_arn}: SUCCESSFUL")
```
#### Training data artifact
```
training_data_s3_uri = training_job_2_info["InputDataConfig"][0]["DataSource"]["S3DataSource"][
"S3Uri"
]
list_response = list(
sagemaker.lineage.artifact.Artifact.list(
source_uri=training_data_s3_uri, sagemaker_session=sagemaker_session
)
)
if list_response:
training_data_artifact = list_response[0]
print(f"Using existing artifact: {training_data_artifact.artifact_arn}")
else:
training_data_artifact = sagemaker.lineage.artifact.Artifact.create(
artifact_name="TrainingData",
source_uri=training_data_s3_uri,
artifact_type="Dataset",
sagemaker_session=sagemaker_session,
)
print(f"Create artifact {training_data_artifact.artifact_arn}: SUCCESSFUL")
```
#### Model artifact
```
trained_model_s3_uri = training_job_2_info["ModelArtifacts"]["S3ModelArtifacts"]
list_response = list(
sagemaker.lineage.artifact.Artifact.list(
source_uri=trained_model_s3_uri, sagemaker_session=sagemaker_session
)
)
if list_response:
model_artifact = list_response[0]
print(f"Using existing artifact: {model_artifact.artifact_arn}")
else:
model_artifact = sagemaker.lineage.artifact.Artifact.create(
artifact_name="TrainedModel",
source_uri=trained_model_s3_uri,
artifact_type="Model",
sagemaker_session=sagemaker_session,
)
print(f"Create artifact {model_artifact.artifact_arn}: SUCCESSFUL")
```
### Set artifact associations
```
trial_component = sagemaker_boto_client.describe_trial_component(
TrialComponentName=training_job_2_name + "-aws-training-job"
)
trial_component_arn = trial_component["TrialComponentArn"]
```
#### Input artifacts
```
input_artifacts = [code_artifact, training_data_artifact]
for a in input_artifacts:
try:
sagemaker.lineage.association.Association.create(
source_arn=a.artifact_arn,
destination_arn=trial_component_arn,
association_type="ContributedTo",
sagemaker_session=sagemaker_session,
)
print(f"Associate {trial_component_arn} and {a.artifact_arn}: SUCCEESFUL\n")
except:
print(f"Association already exists between {trial_component_arn} and {a.artifact_arn}.\n")
```
#### Output artifacts
```
output_artifacts = [model_artifact]
for artifact_arn in output_artifacts:
try:
sagemaker.lineage.association.Association.create(
source_arn=a.artifact_arn,
destination_arn=trial_component_arn,
association_type="Produced",
sagemaker_session=sagemaker_session,
)
print(f"Associate {trial_component_arn} and {a.artifact_arn}: SUCCEESFUL\n")
except:
print(f"Association already exists between {trial_component_arn} and {a.artifact_arn}.\n")
```
<pre>
</pre>
<a id ='analyze-second-model'></a>
## Analyze the second model for bias and explainability
[overview](#aup-overview)
----
Amazon SageMaker Clarify provides tools to help explain how machine learning (ML) models make predictions. These tools can help ML modelers and developers and other internal stakeholders understand model characteristics as a whole prior to deployment and to debug predictions provided by the model after it's deployed. Transparency about how ML models arrive at their predictions is also critical to consumers and regulators who need to trust the model predictions if they are going to accept the decisions based on them. SageMaker Clarify uses a model-agnostic feature attribution approach, which you can used to understand why a model made a prediction after training and to provide per-instance explanation during inference. The implementation includes a scalable and efficient implementation of SHAP ([see paper](https://papers.nips.cc/paper/2017/file/8a20a8621978632d76c43dfd28b67767-Paper.pdf)), based on the concept of a Shapley value from the field of cooperative game theory that assigns each feature an importance value for a particular prediction.
### Create model from estimator
```
model_matches = sagemaker_boto_client.list_models(NameContains=model_2_name)['Models']
if not model_matches:
model_2 = sagemaker_session.create_model_from_job(
name=model_2_name,
training_job_name=training_job_2_info['TrainingJobName'],
role=sagemaker_role,
image_uri=training_job_2_info['AlgorithmSpecification']['TrainingImage'])
%store model_2_name
else:
print(f"Model {model_2_name} already exists.")
```
<a id='bias-v1'></a>
### Check for data set bias and model bias
With SageMaker, we can check for pre-training and post-training bias. Pre-training metrics show pre-existing bias in that data, while post-training metrics show bias in the predictions from the model. Using the SageMaker SDK, we can specify which groups we want to check bias across and which metrics we'd like to show.
To run the full Clarify job, you must un-comment the code in the cell below. Running the job will take ~15 minutes. If you wish to save time, you can view the results in the next cell after which loads a pre-generated output if no bias job was run.
```
clarify_processor = sagemaker.clarify.SageMakerClarifyProcessor(
role=sagemaker_role,
instance_count=1,
instance_type="ml.c4.xlarge",
sagemaker_session=sagemaker_session,
)
bias_data_config = sagemaker.clarify.DataConfig(
s3_data_input_path=train_data_upsampled_s3_path,
s3_output_path=bias_report_2_output_path,
label="fraud",
headers=train.columns.to_list(),
dataset_type="text/csv",
)
model_config = sagemaker.clarify.ModelConfig(
model_name=model_2_name,
instance_type=train_instance_type,
instance_count=1,
accept_type="text/csv",
)
predictions_config = sagemaker.clarify.ModelPredictedLabelConfig(probability_threshold=0.5)
bias_config = sagemaker.clarify.BiasConfig(
label_values_or_threshold=[0],
facet_name="customer_gender_female",
facet_values_or_threshold=[1],
)
# # un-comment the code below to run the whole job
# if 'clarify_bias_job_2_name' not in locals():
# clarify_processor.run_bias(
# data_config=bias_data_config,
# bias_config=bias_config,
# model_config=model_config,
# model_predicted_label_config=predictions_config,
# pre_training_methods='all',
# post_training_methods='all')
# clarify_bias_job_2_name = clarify_processor.latest_job.name
# %store clarify_bias_job_2_name
# else:
# print(f'Clarify job {clarify_bias_job_2_name} has already run successfully.')
```
<a id ='view-second-clarify-job'></a>
## View results of Clarify job
[overview](#aup-overview)
----
Running Clarify on your dataset or model can take ~15 minutes. If you don't have time to run the job, you can view the pre-generated results included with this demo. Otherwise, you can run the job by un-commenting the code in the cell above.
```
if "clarify_bias_job_2_name" in locals():
s3_client.download_file(
Bucket=bucket,
Key=f"{prefix}/clarify-output/bias-2/analysis.json",
Filename="clarify_output/bias_2/analysis.json",
)
print(f"Downloaded analysis from previous Clarify job: {clarify_bias_job_2_name}\n")
else:
print(f"Loading pre-generated analysis file...\n")
with open("clarify_output/bias_1/analysis.json", "r") as f:
bias_analysis = json.load(f)
results = bias_analysis["pre_training_bias_metrics"]["facets"]["customer_gender_female"][0][
"metrics"
][1]
print(json.dumps(results, indent=4))
with open("clarify_output/bias_2/analysis.json", "r") as f:
bias_analysis = json.load(f)
results = bias_analysis["pre_training_bias_metrics"]["facets"]["customer_gender_female"][0][
"metrics"
][1]
print(json.dumps(results, indent=4))
```
<a id ='explainability' ></a>
## Configure and run explainability job
[overview](#aup-overview)
----
To run the full Clarify job, you must un-comment the code in the cell below. Running the job will take ~15 minutes. If you wish to save time, you can view the results in the next cell after which loads a pre-generated output if no explainability job was run.
```
model_config = sagemaker.clarify.ModelConfig(
model_name=model_2_name,
instance_type=train_instance_type,
instance_count=1,
accept_type="text/csv",
)
shap_config = sagemaker.clarify.SHAPConfig(
baseline=[train.median().values[1:].tolist()], num_samples=100, agg_method="mean_abs"
)
explainability_data_config = sagemaker.clarify.DataConfig(
s3_data_input_path=train_data_upsampled_s3_path,
s3_output_path=explainability_output_path,
label="fraud",
headers=train.columns.to_list(),
dataset_type="text/csv",
)
# un-comment the code below to run the whole job
# if 'clarify_expl_job_name' not in locals():
# clarify_processor.run_explainability(
# data_config=explainability_data_config,
# model_config=model_config,
# explainability_config=shap_config)
# clarify_expl_job_name = clarify_processor.latest_job.name
# %store clarify_expl_job_name
# else:
# print(f'Clarify job {clarify_expl_job_name} has already run successfully.')
```
### View Clarify explainability results (shortcut)
Running Clarify on your dataset or model can take ~15 minutes. If you don't have time to run the job, you can view the pre-generated results included with this demo. Otherwise, you can run the job by un-commenting the code in the cell above.
```
if "clarify_expl_job_name" in locals():
s3_client.download_file(
Bucket=bucket,
Key=f"{prefix}/clarify-output/explainability/analysis.json",
Filename="clarify_output/explainability/analysis.json",
)
print(f"Downloaded analysis from previous Clarify job: {clarify_expl_job_name}\n")
else:
print(f"Loading pre-generated analysis file...\n")
with open("clarify_output/explainability/analysis.json", "r") as f:
analysis_result = json.load(f)
shap_values = pd.DataFrame(analysis_result["explanations"]["kernel_shap"]["label0"])
importances = shap_values["global_shap_values"].sort_values(ascending=False)
fig, ax = plt.subplots()
n = 5
y_pos = np.arange(n)
importance_scores = importances.values[:n]
y_label = importances.index[:n]
ax.barh(y_pos, importance_scores, align="center")
ax.set_yticks(y_pos)
ax.set_yticklabels(y_label)
ax.invert_yaxis()
ax.set_xlabel("SHAP Value (impact on model output)");
```
To see the autogenerated SageMaker Clarify report, run the following code and use the output link to open the report.
```
from IPython.display import FileLink, FileLinks
display(
"Click link below to view the SageMaker Clarify report", FileLink("clarify_output/report.pdf")
)
```
### What is SHAP?
SHAP is the method used for calculating explanations in this solution.
Unlike other feature attribution methods, such as single feature
permutation, SHAP tries to disentangle the effect of a single feature by
looking at all possible combinations of features.
[SHAP](https://github.com/slundberg/shap) (Lundberg et al. 2017) stands
for SHapley Additive exPlanations. 'Shapley' relates to a game theoretic
concept called [Shapley
values](https://en.wikipedia.org/wiki/Shapley_value) that is used to
create the explanations. A Shapley value describes the marginal
contribution of each 'player' when considering all possible 'coalitions'.
Using this in a machine learning context, a Shapley value describes the
marginal contribution of each feature when considering all possible sets
of features. 'Additive' relates to the fact that these Shapley values can
be summed together to give the final model prediction.
As an example, we might start off with a baseline credit default risk of
10%. Given a set of features, we can calculate the Shapley value for each
feature. Summing together all the Shapley values, we might obtain a
cumulative value of +30%. Given the same set of features, we therefore
expect our model to return a credit default risk of 40% (i.e. 10% + 30%).
<a id='model-package' ></a>
## Create Model Package for the Second Trained Model
[overview](#aup-overview)
----
#### Create and upload second model metrics report
```
model_metrics_report = {"classification_metrics": {}}
for metric in training_job_2_info["FinalMetricDataList"]:
stat = {metric["MetricName"]: {"value": metric["Value"]}}
model_metrics_report["classification_metrics"].update(stat)
with open("training_metrics.json", "w") as f:
json.dump(model_metrics_report, f)
metrics_s3_key = (
f"{prefix}/training_jobs/{training_job_2_info['TrainingJobName']}/training_metrics.json"
)
s3_client.upload_file(Filename="training_metrics.json", Bucket=bucket, Key=metrics_s3_key)
```
#### Define inference specification
```
mp_inference_spec = InferenceSpecification().get_inference_specification_dict(
ecr_image=training_job_2_info["AlgorithmSpecification"]["TrainingImage"],
supports_gpu=False,
supported_content_types=["text/csv"],
supported_mime_types=["text/csv"],
)
mp_inference_spec["InferenceSpecification"]["Containers"][0]["ModelDataUrl"] = training_job_2_info[
"ModelArtifacts"
]["S3ModelArtifacts"]
```
#### Define model metrics
```
model_metrics = {
"ModelQuality": {
"Statistics": {
"ContentType": "application/json",
"S3Uri": f"s3://{bucket}/{prefix}/{metrics_s3_key}",
}
},
"Bias": {
"Report": {
"ContentType": "application/json",
"S3Uri": f"{explainability_output_path}/analysis.json",
}
},
}
```
#### Register second model package to Model Package Group
```
mp_input_dict = {
"ModelPackageGroupName": mpg_name,
"ModelPackageDescription": "XGBoost classifier to detect insurance fraud with SMOTE.",
"ModelApprovalStatus": "PendingManualApproval",
"ModelMetrics": model_metrics,
}
mp_input_dict.update(mp_inference_spec)
mp2_response = sagemaker_boto_client.create_model_package(**mp_input_dict)
mp2_arn = mp2_response["ModelPackageArn"]
%store mp2_arn
```
#### Check status of model package creation
```
mp_info = sagemaker_boto_client.describe_model_package(
ModelPackageName=mp2_response["ModelPackageArn"]
)
mp_status = mp_info["ModelPackageStatus"]
while mp_status not in ["Completed", "Failed"]:
time.sleep(5)
mp_info = sagemaker_boto_client.describe_model_package(
ModelPackageName=mp2_response["ModelPackageArn"]
)
mp_status = mp_info["ModelPackageStatus"]
print(f"model package status: {mp_status}")
print(f"model package status: {mp_status}")
```
### View both models in the registry
```
sagemaker_boto_client.list_model_packages(ModelPackageGroupName=mpg_name)["ModelPackageSummaryList"]
```
----
### Next Notebook: [Deploy Model, Run Predictions](./4-deploy-run-inference-e2e.ipynb)
| github_jupyter |
<a href="https://colab.research.google.com/github/mohammadeunus/MachineLearningAlgorithms-MLA/blob/main/generic_algorithm_for_n_queen.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import random
chess=chessBoardSize= 8
initialPopulation=10000
MaxFitnessValue=0
while chess!=1:
chess-=1
MaxFitnessValue=MaxFitnessValue+chess
print(MaxFitnessValue)
mainList = []
#populating the samples for 1st generation
for _ in range(initialPopulation):
mainList.append([random.randint(0, chessBoardSize-1) for _ in range(chessBoardSize)])
print(mainList)
#measuring the fitnessValue of all sample data
listOffitnessValue= []
#print('number of attacks by every queen: on the far right side the attack count increaments for every individual queen ')
for sampleNumberInMainList in range(initialPopulation):
fitnessValue=0
for individualSampleNumberInMainList in range(chessBoardSize):
for k in range(individualSampleNumberInMainList+1,chessBoardSize):
col = individualSampleNumberInMainList
row = mainList[sampleNumberInMainList][individualSampleNumberInMainList]
colN= k
rowN= mainList[sampleNumberInMainList][k]
#print(col,row,sep=",",end=' ')
#print('vs',end=' ')
#condition: checking whether a queen attacks another or not
if col == colN or row == rowN or abs(col - colN) == abs(row - rowN):
fitnessValue+=1
#print(colN,rowN,sep=",",end=' -> ')
#print(fitnessValue)
#else:
#print(colN,rowN,sep=",")
#print()
#print('............................................')
listOffitnessValue.append(fitnessValue) #appending the total attack count(fitnessValue) for every individual queen
#end of forLoop
print('list of fitnessValues: ',end=" ")
print(listOffitnessValue)
print()
print('list of sample data: ',end=" ")
print(mainList)
#merged the fitnessValues as key with sample Data in a dictionary dictOfSamplesWithFitnessValue
dictOfSamplesWithFitnessValue=dict(zip(listOffitnessValue,mainList))
print('the dictionary before sorting : ',dictOfSamplesWithFitnessValue)
#sorted the data by key values
dictOfSamplesWithFitnessValue = dict(sorted(dictOfSamplesWithFitnessValue.items()))
print()
print('the dictionary after sorting : ',dictOfSamplesWithFitnessValue)
print(len(dictOfSamplesWithFitnessValue))
print()
refinedDic={}
if len(dictOfSamplesWithFitnessValue) >=9:
for i in dictOfSamplesWithFitnessValue:
if i <= int(MaxFitnessValue*.50): #deleting the sample values having more than 40percent of the max fitness value
refinedDic[i]= dictOfSamplesWithFitnessValue[i]
else:
refinedDic=dictOfSamplesWithFitnessValue
print()
print(refinedDic)
print()
print(len(refinedDic))
#crossover
afterCrossOverList=[]
def Crossover(p1,p2,point):
print(point)
for i in range(point,len(p1)):
p1[i],p2[i] = p2[i],p1[i] #swap the genetic information
afterCrossOverList.append(p1)
afterCrossOverList.append(p2)
count=-1
listOfSampleValues=list(refinedDic.values())
print(listOfSampleValues)
for i in range((len(refinedDic.values()))-2):
print(i)
count+=1
if(i==0):
point= random.randint(0,chessBoardSize)
Crossover(listOfSampleValues[i],listOfSampleValues[i+1],point)
if(count%2==0):
point= random.randint(0,chessBoardSize)
Crossover(listOfSampleValues[i],listOfSampleValues[i+1],point)
print(afterCrossOverList)
ggg=[[1,[212,3,2,5]],[2,[2]],[3232,[3]],[342,[44]]]
print(ggg)
ddd=list(zip(list(listOffitnessValue),list(mainList)))
print(ddd)
print(type(ddd[0]))
for i in refinedDic:
a_dictionary = {22: 1, "b": 2, "c":3}
keys_list = list(a_dictionary)
print(keys_list)
key = keys_list[0]
print(key)
```
| github_jupyter |
# Data pre-processing for Climate Spirals Visualisation
```
%matplotlib inline
import pandas as pd
from pandas_datapackage_reader import read_datapackage
import matplotlib.pyplot as plt
from scipy import stats
plt.style.use("ggplot")
```
## CO<sub>2</sub> emissions from the Global Carbon Budget
Historical CO<sub>2</sub> emissions are taken from the [Global Carbon Budget](https://www.globalcarbonproject.org/carbonbudget/20/data.htm). After downloading the Excel file we can read it in:
```
gcb = pd.read_excel(
'Global_Carbon_Budget_2020v1.0.xlsx',
sheet_name="Historical Budget",
skiprows=15,
index_col="Year",
usecols="A:H"
)
gcb = gcb.rename(columns={
"fossil emissions excluding carbonation": "Fossil-Fuel-Industry",
"land-use change emissions": "Land-Use-Change"
})
gcb.head()
gcb.tail()
gcb = gcb.dropna(how="all")
gcb = gcb.rename(index={"2020*": 2020})
gcb.loc[2020] = gcb.loc[2020].str.replace("~", "")
gcb = gcb.astype(float)
gcb
```
Converting to GtCO2 from GtC
```
co2 = (gcb[["Fossil-Fuel-Industry", "Land-Use-Change"]].loc[1850:] * 3.66 )
co2.tail()
co2["Total"] = co2[["Fossil-Fuel-Industry", "Land-Use-Change"]].sum(axis=1)
co2.tail()
co2.plot(subplots=True, sharey=True, figsize=(12,8))
```
Export for visualisation
```
export = pd.DataFrame({"value": co2["Total"]})
export.index.name = "year"
export.reset_index().to_csv("../public/emissions.csv", index=False, float_format='%g')
export.tail()
co2.loc[:2019].Total.sum()
```
## CO<sub>2</sub> concentrations
CO<sub>2</sub> concentrations are taken from the [CMIP6 concentration dataset](http://www.climate-energy-college.net/cmip6), version from 1 July 2016, combined with data from [ESRL/NOAA](https://www.esrl.noaa.gov/gmd/ccgg/trends/gl_data.html).
```
noaa = pd.read_csv("co2_mm_gl.txt", comment="#", parse_dates=[[0, 1]], index_col=[0], delim_whitespace=True, header=None,
names=["year", "month", "decimal", "average", "trend"])
noaa.head()
cmip6 = pd.read_csv(
"mole_fraction_of_carbon_dioxide_in_air_input4MIPs_GHGConcentrations_CMIP_UoM-CMIP-1-1-0_gr3-GMNHSH_000001-201412.csv",
)
cmip6.index = (cmip6.year.astype(str).apply(lambda x: x.zfill(4)) +
"-" +
cmip6.month.astype(str).apply(lambda x: x.zfill(2)) +
"-01"
)
cmip6 = cmip6.iloc[21000:]
cmip6.index = pd.to_datetime(cmip6.index)
cmip6 = cmip6.drop(['datenum', 'datetime', 'day'], axis=1)
cmip6.head()
fig, ax = plt.subplots(1,1, figsize=(16,10))
cmip6.loc["1850":]["data_mean_global"].plot(ax=ax)
noaa["average"].plot(ax=ax)
cmip6.loc["1850":].rename(columns={"data_mean_nh": "value"}).to_csv("../public/concentrations_nh.csv", index=False)
cmip6.loc["1850":].rename(columns={"data_mean_sh": "value"}).to_csv("../public/concentrations_sh.csv", index=False)
combined = pd.concat([cmip6["data_mean_global"].loc["1850":"1979-12"], noaa["average"]])
combined = pd.DataFrame({"value": combined})
combined["year"] = combined.index.year
combined["month"] = combined.index.month
combined.index.name = "date"
combined["day"] = 15
combined = combined[["year", "month", "value"]]
combined.to_csv("../public/concentrations.csv", index=False)
combined.tail()
```
## Global Temperatures
Global temperature data is taken from the HadCRUT4 near surface temperature dataset.
http://www.metoffice.gov.uk/hadobs/hadcrut4/data/current/download.html
```
hadcrut = pd.read_csv(
"HadCRUT.4.6.0.0.monthly_ns_avg.txt",
delim_whitespace=True,
usecols=[0, 1],
header=None
)
hadcrut['year'] = hadcrut.iloc[:, 0].apply(lambda x: x.split("/")[0]).astype(int)
hadcrut['month'] = hadcrut.iloc[:, 0].apply(lambda x: x.split("/")[1]).astype(int)
hadcrut = hadcrut.rename(columns={1: "value"})
hadcrut = hadcrut.iloc[:, 1:]
hadcrut = hadcrut.set_index(['year', 'month'])
hadcrut -= hadcrut.loc[1850:1900].mean()
hadcrut.plot(figsize=(16,10))
hadcrut = hadcrut.reset_index()
plt.xlabel("Time")
plt.ylabel(u"Temperature anomalies (°C) (1850-1990 mean)")
plt.legend("")
hadcrut.tail()
hadcrut.to_csv("../public/temperatures.csv", index=False)
```
| github_jupyter |
**This code is for CSoNet-2021 paper "Detecting Hate Speech Contents Using Embedding Models".**
**The resources of this paper are available at [here](https://github.com/duonghuuphuc/hate-speech-detection)**
# Import Library & Package
```
# Install Spacy and its en_core_web_lg
!pip install spacy
!python -m spacy download en_core_web_lg
from sklearn.utils import resample
from sklearn.model_selection import KFold, train_test_split, StratifiedKFold
from sklearn import metrics
from torch.utils.data import Dataset, DataLoader
from torch.autograd import Variable
from torch.nn.utils.rnn import pack_padded_sequence
from string import punctuation, ascii_lowercase
from imblearn.over_sampling import SMOTE
from gensim.models import KeyedVectors
from tqdm import tqdm
import os
import re
import random
import time
import copy
import logging
import spacy
import json
import gdown
import pandas as pd
pd.options.mode.chained_assignment = None
import numpy as np
import seaborn as sns
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import matplotlib.pyplot as plt
logging.basicConfig(format='%(message)s')
logger = logging.getLogger()
logger.setLevel(logging.INFO)
```
# Read & Prepare Dataset
* HASOC2019: Remove unused columns, change the column name to class
* HSOF-3: Remove unused columns, change the label 0 to 1 which mean label Hatespeech become label Offensive and 2 to 0 which mean label of Neither become 0.
* HS2-2021: Concatenate 2 dataset HASOC2019 and Davidson et Al.
**NOTE**: If you run all the blocks then the dataset's option will be our dataset, HS2-2021. Otherhands, make your decision on which dataset you want to perform.
## **Download Dataset**
```
!mkdir datasets
```
### HASOC-2019
```
!mkdir ./datasets/HASOC2019
!wget -O ./datasets/HASOC2019/english_dataset.tsv https://raw.githubusercontent.com/socialmediaie/HASOC2019/master/data/raw/training_data/english_dataset.tsv
!wget -O ./datasets/HASOC2019/test_english_dataset.tsv https://raw.githubusercontent.com/socialmediaie/HASOC2019/master/data/raw/test_data_gold/english_data.tsv
```
### Davidson-2017
```
!mkdir ./datasets/HSOF-3
!wget -O ./datasets/HSOF-3/labeled_data.csv https://raw.githubusercontent.com/t-davidson/hate-speech-and-offensive-language/master/data/labeled_data.csv
```
### HS2-2021
```
"""
In this case, you need to upload the file which download from our Github.
"""
!mkdir ./datasets/HS2-2021
```
## **Prepare Dataset**
```
class PreprocessData:
#Regular Expression
nlp = spacy.load("en_core_web_lg", disable=['parser', 'ner'])
space_pattern = '\s+'
giant_url_regex = ('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|''[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+')
mention_regex1 = '([@#][\w\s]+:)'
mention_regex2 = '([@#][\w]+)'
mention_regex3 = '(RT\s)'
special_character = '[^a-zA-Z\s]'
consecutive_space1 = '(?:\s\s)'
consecutive_space2 = '(?:\s{3})'
consecutive_space3 = '(?:\b\s{2})'
html_entities = '&([a-z0-9]+|#[0-9]{1,6}|#x[0-9a-fA-F]{1,6});'
def __init__(self):
self.data_df = None
def spacyLemmatize(self, sentence):
sentence = sentence.lower()
doc = self.nlp(sentence)
return " ".join([token.lemma_ for token in doc])
def run(self):
"""
Remove URL + Space + Emoji + Newline
"""
for t in tqdm(range(len(self.data_df['text'])), position=0, leave=True):
if("\n" in self.data_df['text'][t]):
self.data_df['text'][t] = self.data_df['text'][t].replace("\n", " ")
parsed_text = re.sub(self.space_pattern, ' ', self.data_df['text'][t])
parsed_text = re.sub(self.giant_url_regex, '', parsed_text)
if ("RT" in parsed_text):
parsed_text = re.sub(self.mention_regex3, '', parsed_text)
parsed_text = re.sub(self.mention_regex1, '', parsed_text)
parsed_text = re.sub(self.mention_regex2, '', parsed_text)
parsed_text = re.sub(self.html_entities, ' ', parsed_text)
parsed_text = re.sub(self.special_character, '', parsed_text)
parsed_text = re.sub(self.consecutive_space2, ' ', parsed_text)
parsed_text = re.sub(self.consecutive_space1, ' ', parsed_text)
parsed_text = re.sub(self.consecutive_space3, ' ', parsed_text)
parsed_text = self.spacyLemmatize(parsed_text)
parsed_text = parsed_text.replace('-PRON-', '')
parsed_text = re.sub(self.consecutive_space2, ' ', parsed_text)
parsed_text = re.sub(self.consecutive_space1, ' ', parsed_text)
parsed_text = re.sub(self.consecutive_space3, ' ', parsed_text)
self.data_df['text'][t] = parsed_text.strip()
def cvToCsv(self, path):
self.data_df.to_csv(path, index=False, columns=["text", "class"])
def pipeline(self, data_df, name):
self.data_df = data_df
self.run()
self.cvToCsv(name)
return self.data_df
```
### HASOC-2019
```
data_name = "HASOC-2019"
train_file_path = './datasets/HASOC2019/english_dataset.tsv'
test_file_path = './datasets/HASOC2019/test_english_dataset.tsv'
preprocessor = PreprocessData()
logger.info(f"Preprocess train dataset from {train_file_path}")
train_df = pd.read_csv(train_file_path, sep='\t')
train_df = train_df.drop(columns=['task_2', 'task_3', 'text_id'])
train_df = train_df.rename(columns={'task_1': 'class'})
train_df["class"] = train_df["class"].replace({'NOT': 0, 'HOF': 1})
train_df = preprocessor.pipeline(train_df, './datasets/HASOC2019/preprocessed_english_dataset.tsv')
logger.info(f"Preprocess test dataset from {test_file_path}")
test_df = pd.read_csv(test_file_path, sep='\t')
test_df = test_df.drop(columns=['task_2', 'task_3', 'text_id'])
test_df = test_df.rename(columns={'task_1': 'class'})
test_df["class"] = test_df["class"].replace({'NOT': 0, 'HOF': 1})
test_df = preprocessor.pipeline(test_df, './datasets/HASOC2019/preprocessed_hasoc2019_en_test-2919.tsv')
dataset_max_length = train_df['seq_len'] = train_df["text"].apply(lambda words: len(words.split(" ")))
MAX_SEQUENCE_LENGTH = max(dataset_max_length)
is_balance= False
```
### HSOF-3
```
data_name = "HSOF-3"
full_file_path = './datasets/HSOF-3/labeled_data.csv'
full_df = pd.read_csv(full_file_path)
train_df, test_df = train_test_split(full_df, random_state=42, test_size=0.1)
preprocessor = PreprocessData()
logger.info(f"Preprocess train dataset from {full_file_path}")
train_df = train_df.drop(columns=['count', 'hate_speech', 'offensive_language', 'neither', 'Unnamed: 0'])
train_df['class'] = train_df['class'].replace({0: 1, 2: 0})
train_df = train_df.reset_index().drop(columns=['index'])
train_df = train_df.rename(columns={'tweet': 'text'})
train_df = preprocessor.pipeline(train_df, './datasets/HSOF-3/preprocessed_labeled_data.csv')
logger.info(f"Preprocess train dataset from {full_file_path}")
test_df = test_df.drop(columns=['count', 'hate_speech', 'offensive_language', 'neither', 'Unnamed: 0'])
test_df['class'] = test_df['class'].replace({0: 1, 2: 0})
test_df = test_df.reset_index().drop(columns=['index'])
test_df = test_df.rename(columns={'tweet': 'text'})
test_df = preprocessor.pipeline(test_df, './datasets/HSOF-3/preprocessed_test_labeled_data.csv')
dataset_max_length = train_df['seq_len'] = train_df["text"].apply(lambda words: len(words.split(" ")))
MAX_SEQUENCE_LENGTH = max(dataset_max_length)
is_balance= True
```
### HS2-2021
```
data_name = "HS2-2021"
full_file_path = './datasets/HS2-2021/hs2_2021.csv'
full_df = pd.read_csv(full_file_path)
preprocessor = PreprocessData()
logger.info(f"Preprocess full dataset from {full_file_path}")
train_df = preprocessor.pipeline(full_df, './datasets/HS2-2021/preprocessed_hs2_2021.csv')
logger.info("Spliting Process ...")
train_df, test_df = train_test_split(train_df, random_state=42, test_size=0.1)
logger.info("Done")
dataset_max_length = train_df['seq_len'] = train_df["text"].apply(lambda words: len(words.split(" ")))
MAX_SEQUENCE_LENGTH = max(dataset_max_length)
is_balance= True
```
# Word2Vec
```
url = 'https://drive.google.com/u/0/uc?id=0B7XkCwpI5KDYNlNUTTlSS21pQmM&export=download'
output = 'GoogleNews-vectors-negative300.bin.gz'
if not os.path.exists(output):
gdown.download(url, output, quiet=False)
!gzip -d GoogleNews-vectors-negative300.bin.gz
"""
Upload the hate speech dictionary to the root folder before you run this block.
"""
hate_array = []
if os.path.exists('./Hatespeech.csv'):
logger.info("Load Hate Speech Dictionary ...")
hate_file = pd.read_csv('./Hatespeech.csv', header=None)
hate_array = hate_file[0].values
logger.info(f"Total unique hatespeech word: {len(hate_array)}")
def get_emb_matrix_by_gensim(word_index, max_features, embedding_file):
w2v_model = KeyedVectors.load_word2vec_format(embedding_file, binary=True)
embedding_dim = w2v_model.vectors.shape[1]
embedding_matrix = np.zeros((max_features, embedding_dim))
for word, i in word_index.items():
if i >= max_features:
continue
try:
embedding_vector = w2v_model.get_vector(word)
except:
embedding_vector = w2v_model.get_vector('unk')
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
return embedding_matrix
def build_vocab(texts):
unique_words = set()
for text in texts:
unique_words.update([i.strip() for i in text.split(" ") if i.strip()])
vocab = dict(zip(unique_words, list(range(1, len(unique_words) + 1))))
return vocab
merged_dataset = pd.concat([train_df, test_df], ignore_index=True)
logger.info("Build vocabulary ...")
dataset_vocab = build_vocab(list(merged_dataset['text'].values))
logger.info("Load word2vec model ...")
embedded_matrix = get_emb_matrix_by_gensim(dataset_vocab, len(dataset_vocab) + 1, './GoogleNews-vectors-negative300.bin')
embedded_matrix = torch.FloatTensor(embedded_matrix)
```
# Custom Dataloader
Note: Change self.balance value when you run the code on Davidson et Al. and HS2-2021, its default value is False on HASOC2020 which means that we perform SMOTE on these 2 datasets.
```
class MyDataset(Dataset):
def __init__(self, dataframe, max_seq_len, vocab, hate_array, is_balance = False, dataset_type="Train"):
self.dataset_type = dataset_type
self.max_seq_len = MAX_SEQUENCE_LENGTH
self.vocab = vocab
self.is_balance = False
self.examples = self.create_examples(dataframe, hate_array)
def __len__(self):
return len(self.examples)
def __getitem__(self, index):
example = self.examples[index]
ex_token_tensor = torch.tensor(example[0], dtype=torch.long)
ex_label_tensor = torch.tensor(example[1], dtype=torch.long)
ex_seq_length_tensor = torch.tensor(example[2], dtype=torch.long)
ex_hate_tensor = torch.tensor(example[3], dtype=torch.long)
return ex_token_tensor, ex_label_tensor, ex_seq_length_tensor, ex_hate_tensor
@staticmethod
def visualize(labels):
g = sns.countplot(labels)
g.set_xticklabels(['NOT','HOF'])
plt.show()
@staticmethod
def collate_fn(batch):
all_input_ids, all_labels, all_lens, all_hates = map(torch.stack, zip(*batch))
all_lens, indices = torch.sort(all_lens, descending=True)
all_input_ids = all_input_ids[indices]
all_labels = all_labels[indices]
return all_input_ids, all_labels, all_lens, all_hates
def create_examples(self, data_df, hate_array):
data_df = data_df.reset_index()
examples = []
sentences = []
hates = []
labels = []
self.visualize(data_df['class'])
tqdm_bar = tqdm(data_df.iterrows(), desc=f'Create {self.dataset_type} examples: ', total=len(data_df), position=0, leave=True)
for _, row in tqdm_bar:
hate_ids = []
token_ids = [self.vocab[token] for token in str(row['text']).lower().strip().split()[:self.max_seq_len]]
for token in str(row['text']).lower().strip().split()[:self.max_seq_len]:
if token in hate_array:
hate_ids.append(1)
else:
hate_ids.append(0)
seq_len = len(token_ids)
hate_len = len(hate_ids)
if seq_len == 0:
continue
if seq_len < self.max_seq_len:
num_pad = self.max_seq_len - seq_len
token_ids += [0] * num_pad
if hate_len < self.max_seq_len:
num_pad = self.max_seq_len - seq_len
hate_ids += [0] * num_pad
if not len(token_ids) == self.max_seq_len:
logger.info(f"{seq_len} => {len(token_ids)}")
if self.is_balance and self.dataset_type == 'Train':
token_ids = torch.LongTensor(token_ids)
hate_ids = torch.LongTensor(hate_ids)
sentences.append(token_ids)
hates.append(hate_ids)
labels.append(row['class'])
else:
examples.append((token_ids, row['class'], seq_len, hate_ids))
if self.is_balance and self.dataset_type == 'Train':
new_sentences = torch.stack(sentences)
new_labels = torch.LongTensor(labels)
new_hates = torch.stack(hates)
smote = SMOTE()
x_smote, y_smote = smote.fit_resample(new_sentences, labels)
hate_smote, y_hate_smote = smote.fit_resample(new_hates, labels)
for _,ins in enumerate(x_smote):
indices = np.where(ins==11380)
new_list = np.delete(ins, indices)
examples.append((ins, y_smote[_], len(new_list), hate_smote[_]))
self.visualize(y_smote)
for index in range(2):
logger.info(f"{self.dataset_type} example {index}: ")
logger.info(f"\tRaw text : {data_df['text'][index]}")
logger.info(f"\tToken IDS : {examples[index][0]}")
logger.info(f"\tHate IDS : {examples[index][3]}")
logger.info(f"\tLabel ID : {examples[index][1]}")
logger.info(f"\tSeq length : {examples[index][2]}")
return examples
if not (os.path.exists(f"train_dataset_{data_name}_cached.pt" ) and os.path.exists( f"test_dataset_{data_name}_cached.pt")):
train_dataset = MyDataset(train_df, MAX_SEQUENCE_LENGTH, dataset_vocab, hate_array, is_balance, dataset_type='Train')
test_dataset = MyDataset(test_df, MAX_SEQUENCE_LENGTH, dataset_vocab, hate_array, is_balance, dataset_type='Test')
torch.save(train_dataset, f"train_dataset_{data_name}_cached.pt" )
torch.save(test_dataset, f"test_dataset_{data_name}_cached.pt")
else:
train_dataset = torch.load(f"train_dataset_{data_name}_cached.pt")
test_dataset = torch.load( f"test_dataset_{data_name}_cached.pt")
```
# Models
## Hate Embedding Layer
```
class HateEmbedding(nn.Module):
def __init__(self, max_length):
super(HateEmbedding, self).__init__()
self.max_length = max_length
def forward(self, x):
result = []
for ins in x:
hate_embedding = [[1]] * self.max_length if 1 in ins else [[0]] * self.max_length
result.append(hate_embedding)
return torch.tensor(result, dtype=torch.long, device=x.device)
```
## Word2vec + MLP
```
class MLP(nn.Module):
def __init__(self, hidden_size, num_classes, vocab_size, embed_dim, pad_idx, dropout_prob, vectors, use_hatedict=False, requires_grad=True):
super(MLP, self).__init__()
if use_hatedict:
self.embedding_layer = nn.Embedding.from_pretrained(vectors, padding_idx=pad_idx)
self.hate_layer = HateEmbedding(MAX_SEQUENCE_LENGTH)
embed_dim += 1
else:
self.embedding_layer = nn.EmbeddingBag.from_pretrained(vectors, mode="sum")
if vectors is not None:
self.embedding_layer.weight.data = nn.Parameter(vectors, requires_grad=requires_grad)
self.dropout = nn.Dropout(dropout_prob)
self.layer1 = nn.Linear(embed_dim, hidden_size)
self.Relu = nn.ReLU()
self.layer2 = nn.Linear(hidden_size, num_classes)
self.softmax = nn.Softmax(dim=-1)
def forward(self, data_input, seq_length, hate_ids=None):
embedded_data_input = self.embedding_layer(data_input)
if hate_ids is not None:
hate_embedded = self.hate_layer(hate_ids)
embedded_data_input = torch.cat((embedded_data_input, hate_embedded), -1)
embedded_data_input = torch.sum(embedded_data_input, 1)
output_layer1 = self.layer1(embedded_data_input)
relu = self.Relu(output_layer1)
relu_drop = self.dropout(relu)
output_layer2 = self.layer2(relu_drop)
probs = self.softmax(output_layer2)
return probs
```
## Word2vec + BiLSTM
```
class TextLSTM(nn.Module):
def __init__(self, hidden_dim, bidirectional, num_lstm_layers, vocab_size, embed_dim, pad_idx, num_labels, dropout_prob, vectors, use_hatedict=False, requires_grad=True):
super(TextLSTM, self).__init__()
self.bidirectional = bidirectional
self.rnn_hidden_dim = hidden_dim // 2 if bidirectional else hidden_dim
self.embed_layer = nn.Embedding.from_pretrained(vectors, padding_idx=pad_idx)
if vectors is not None:
self.embed_layer.weight.data = nn.Parameter(vectors, requires_grad=requires_grad)
if use_hatedict:
self.hate_layer = HateEmbedding(MAX_SEQUENCE_LENGTH)
embed_dim += 1
self.lstm = nn.LSTM(embed_dim, self.rnn_hidden_dim, bidirectional=bidirectional,
num_layers=num_lstm_layers, batch_first=True,
dropout=dropout_prob if num_lstm_layers > 1 else 0)
self.dropout = nn.Dropout(dropout_prob)
self.fc_layer = nn.Linear(hidden_dim * num_lstm_layers, num_labels)
def forward(self, input_ids, seq_len, hate_ids=None):
batch_size = input_ids.shape[0]
token_reps = self.embed_layer(input_ids)
if hate_ids is not None:
hate_embedded = self.hate_layer(hate_ids)
token_reps = torch.cat((token_reps, hate_embedded), -1)
# token_reps = torch.sum(embedded_data_input, 1)
packed_reps = pack_padded_sequence(token_reps,
batch_first=True,
lengths=seq_len.tolist())
h_0 = Variable(torch.zeros(2, batch_size, self.rnn_hidden_dim).cuda())
c_0 = Variable(torch.zeros(2, batch_size, self.rnn_hidden_dim).cuda())
packed_output, (hn, cn) = self.lstm(packed_reps, (h_0, c_0))
hidden = self.dropout(hn)
hidden = torch.cat([hidden[i, :, :] for i in range(hidden.shape[0])], dim=-1)
logits = self.fc_layer(hidden)
return F.softmax(logits)
```
## Word2vec + CNN
```
class TextCNN(nn.Module):
def __init__(self, num_filters, num_labels, max_len, vocab_size, embed_dim, pad_idx, dropout_prob, use_hatedict=False, vectors=None):
super(TextCNN, self).__init__()
self.max_len = max_len
self.embed_layer = nn.Embedding.from_pretrained(vectors, padding_idx=pad_idx)
if vectors is not None:
self.embed_layer.weight.data = nn.Parameter(vectors, requires_grad=True)
if use_hatedict:
self.hate_layer = HateEmbedding(MAX_SEQUENCE_LENGTH)
embed_dim += 1
self.embed_dim = embed_dim
self.conv1 = nn.Conv1d(1, num_filters, 1 * embed_dim, stride= embed_dim)
self.conv2 = nn.Conv1d(1, num_filters, 2 * embed_dim, stride= embed_dim)
self.conv3 = nn.Conv1d(1, num_filters, 3 * embed_dim, stride= embed_dim)
self.dropout = nn.Dropout(dropout_prob)
self.fc_layer = nn.Linear(3 * num_filters, num_labels)
self.softmax = nn.Softmax(dim=-1)
def forward(self, input_ids, seq_len, hate_ids=None):
if hate_ids is not None:
hate_embedded = self.hate_layer(hate_ids)
word_embedded = self.embed_layer(input_ids)
input_representation = torch.cat((word_embedded, hate_embedded), -1)
x = input_representation.view(-1, 1, (self.embed_dim) * self.max_len)
else:
x = self.embed_layer(input_ids).view(-1, 1, (self.embed_dim * self.max_len))
# Convolution
conv1 = F.relu(self.conv1(x))
conv2 = F.relu(self.conv2(x))
conv3 = F.relu(self.conv3(x))
# Pooling
pooled_conv1 = F.max_pool1d(conv1, self.max_len - 1 + 1).squeeze(-1)
pooled_conv2 = F.max_pool1d(conv2, self.max_len - 2 + 1).squeeze(-1)
pooled_conv3 = F.max_pool1d(conv3, self.max_len - 3 + 1).squeeze(-1)
x = torch.cat((pooled_conv1, pooled_conv2, pooled_conv3), 1)
x = self.dropout(x)
logits = self.fc_layer(x)
probs = self.softmax(logits)
return probs
```
# **Experiments**
## Utils Function
```
def get_total_time(start_time):
end = time.time()
hours, rem = divmod(end - start_time, 3600)
minutes, seconds = divmod(rem, 60)
return sum([int(hours*3600), int(minutes*60), int(seconds)])
def caculate_score(actuals, predicts):
acc_score = metrics.accuracy_score(actuals, predicts)
f1_macro_score = metrics.f1_score(actuals, predicts, average="macro")
f1_weighted_score = metrics.f1_score(actuals, predicts, average="weighted")
report = metrics.classification_report(actuals, predicts, output_dict=True)
tn, fp, fn, tp = metrics.confusion_matrix(actuals, predicts).ravel()
f1_0 = round(report['0']['f1-score'], ndigits=2)
f1_1 = round(report['1']['f1-score'], ndigits=2)
acc_0 = round(tn/(tn+fp), ndigits=2)
acc_1 = round(tp/(tp+fn), ndigits=2)
return acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score
def plot_loss(history):
plt.plot(history['train_loss'])
plt.plot(history['eval_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Evaluate'], loc='upper left')
plt.show()
def eval(model, criterion, eval_iter):
sum_loss = 0
predicts, actuals = [], []
model.eval()
eval_bar = tqdm(eval_iter, desc=f'\tEval',total=len(eval_iter), position=0, leave=0)
start_time = time.time()
with torch.no_grad():
for batch in eval_bar:
if len(batch) == 3:
input_ids, label_ids, seq_lens = batch
else:
input_ids, label_ids, seq_lens, _ = batch
if device == 'cuda':
input_ids = input_ids.cuda()
label_ids = label_ids.cuda()
seq_lens = seq_lens.cuda()
outs = model(input_ids, seq_lens)
loss = criterion(outs, label_ids)
sum_loss += loss.item()
predicts += [y.argmax().item() for y in outs]
actuals += label_ids.tolist()
acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score = caculate_score(actuals, predicts)
avg_loss = sum_loss / len(eval_iter)
reports = metrics.classification_report(actuals, predicts, target_names=list(label_maps.keys()), output_dict=True)
return avg_loss, (acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score), reports
def train(model, criterion, optimizer, train_iter, eval_iter, scheduler=None):
history = {'train_loss':[], 'eval_loss':[]}
best_model = copy.deepcopy(model)
best_epoch = 0
min_loss = float('inf')
counter = 0
epoch_time = {}
for epoch in range(num_epochs):
start = time.time()
sum_loss = 0
predicts, actuals = [], []
model.train()
train_bar = tqdm(train_iter, desc=f'\tEpoch {epoch}', total=len(train_iter), position=0, leave=0)
start_time = time.time()
for batch in train_bar:
if len(batch) == 3:
input_ids, label_ids, seq_lens = batch
else:
input_ids, label_ids, seq_lens, _ = batch
if device == 'cuda':
input_ids = input_ids.cuda()
label_ids = label_ids.cuda()
seq_lens = seq_lens.cuda()
outs = model(input_ids, seq_lens)
loss = criterion(outs, label_ids)
loss.backward()
optimizer.step()
sum_loss += loss.item()
predicts += [y.argmax().item() for y in outs]
actuals += label_ids.tolist()
train_loss = sum_loss / len(train_iter)
acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score = caculate_score(actuals, predicts)
epoch_time[epoch] = get_total_time(start)
eval_loss, eval_scores, _ = eval(model, criterion, eval_iter)
if scheduler is not None:
scheduler.step(eval_loss)
history['train_loss'].append(train_loss)
history['eval_loss'].append(eval_loss)
if eval_loss <= min_loss:
best_model = copy.deepcopy(model)
min_loss = eval_loss
best_epoch = epoch
counter = 0
else:
counter += 1
if counter >= early_stop:
break
plot_loss(history)
return best_model, best_epoch, epoch_time
def eval_hate(model, criterion, eval_iter):
sum_loss = 0
predicts, actuals = [], []
model.eval()
eval_bar = tqdm(eval_iter, desc=f'\tEval',total=len(eval_iter), position=0, leave=0)
start_time = time.time()
with torch.no_grad():
for batch in eval_bar:
input_ids, label_ids, seq_lens, hate_ids = batch
if device == 'cuda':
input_ids = input_ids.cuda()
label_ids = label_ids.cuda()
seq_lens = seq_lens.cuda()
hate_ids = hate_ids.cuda()
outs = model(input_ids, seq_lens, hate_ids)
loss = criterion(outs, label_ids)
sum_loss += loss.item()
predicts += [y.argmax().item() for y in outs]
actuals += label_ids.tolist()
acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score = caculate_score(actuals, predicts)
avg_loss = sum_loss / len(eval_iter)
reports = metrics.classification_report(actuals, predicts, target_names=list(label_maps.keys()), output_dict=True)
return avg_loss, (acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score), reports
def train_hate(model, criterion, optimizer, train_iter, eval_iter, scheduler=None):
history = {'train_loss':[], 'eval_loss':[]}
best_model = copy.deepcopy(model)
best_epoch = 0
min_loss = float('inf')
counter = 0
epoch_time = {}
for epoch in range(num_epochs):
start = time.time()
sum_loss = 0
predicts, actuals = [], []
model.train()
train_bar = tqdm(train_iter, desc=f'\tEpoch {epoch}', total=len(train_iter), position=0, leave=0)
start_time = time.time()
for batch in train_bar:
input_ids, label_ids, seq_lens, hate_ids = batch
if device == 'cuda':
input_ids = input_ids.cuda()
label_ids = label_ids.cuda()
seq_lens = seq_lens.cuda()
hate_ids = hate_ids.cuda()
outs = model(input_ids, seq_lens, hate_ids)
loss = criterion(outs, label_ids)
loss.backward()
optimizer.step()
sum_loss += loss.item()
predicts += [y.argmax().item() for y in outs]
actuals += label_ids.tolist()
train_loss = sum_loss / len(train_iter)
acc_score, f1_0, f1_1, acc_0, acc_1, f1_macro_score = caculate_score(actuals, predicts)
epoch_time[epoch] = get_total_time(start)
eval_loss, eval_scores, _ = eval_hate(model, criterion, eval_iter)
if scheduler is not None:
scheduler.step(eval_loss)
history['train_loss'].append(train_loss)
history['eval_loss'].append(eval_loss)
if eval_loss <= min_loss:
best_model = copy.deepcopy(model)
min_loss = eval_loss
best_epoch = epoch
counter = 0
else:
counter += 1
if counter >= early_stop:
break
plot_loss(history)
return best_model, best_epoch, epoch_time
```
## Word2vec + MLP
### Parameters
```
# Model Options
device = 'cuda' if torch.cuda.is_available() else 'cpu' # device
embed_dim = 300 # embed dim
hidden_dim = 100 # hidden dim
dropout_prob = 0.6 # Dropout prob
pad_idx = 0 # Padding
num_labels = 2 # num labels
# Training Options
num_epochs = 100 # Epoch
num_fold = 5 # Num fold
batch_size = 32
early_stop = 10 # Early Stop
learning_rate = 2e-5 # learning rate
loss_function = nn.CrossEntropyLoss() # Loss Function
kfold = KFold(n_splits=num_fold, shuffle=True)
kfold = StratifiedKFold(n_splits=num_fold, random_state=1, shuffle=True)
```
### Normal Case
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = MLP2(hidden_dim, num_labels, embedded_matrix.shape[0], embedded_matrix.shape[1], pad_idx, dropout_prob, vectors=embedded_matrix, requires_grad=True)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate, weight_decay=1e-5)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train(model, criterion, optimizer, train_iter, eval_iter)
print(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True)
loss, scores, reports = eval(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
### With HateDict
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = MLP(hidden_dim, num_labels, embedded_matrix.shape[0], embedded_matrix.shape[1], pad_idx, dropout_prob, vectors=embedded_matrix, use_hatedict=True, requires_grad=True)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train_hate(model, criterion, optimizer, train_iter, eval_iter)
print(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True)
loss, scores, reports = eval_hate(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
## Word2vec + BiLSTM
### Parameters
```
# Model Options
device = 'cuda' if torch.cuda.is_available() else 'cpu' # device
embed_dim = 300 # embed dim
lstm_hidden_dim = 128 # lstm hidden dim
dropout_prob = 0.6 # Dropout prob
bidirectional = True # bidirectional
num_lstm_layers = 1 # Num Lstm Layers
pad_idx = 0 # Padding
num_labels = 2 # num labels
# Training Options
num_epochs = 100 # Epoch
num_fold = 5 # Num fold
batch_size = 32
early_stop = 10 # Early Stop
learning_rate = 2e-5 # learning rate
loss_function = nn.CrossEntropyLoss() # Loss Function
kfold = StratifiedKFold(n_splits=num_fold, random_state=1, shuffle=True)
```
### Normal Case
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = TextLSTM(lstm_hidden_dim, bidirectional, num_lstm_layers, len(dataset_vocab), embedded_matrix.shape[1], pad_idx, num_labels, dropout_prob, vectors=embedded_matrix, requires_grad=True)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate, weight_decay=1e-5)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler, collate_fn=MyDataset.collate_fn)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler, collate_fn=MyDataset.collate_fn)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train(model, criterion, optimizer, train_iter, eval_iter)
logger.info(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn=MyDataset.collate_fn)
loss, scores, reports = eval(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
### With HateDict
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
data_idxs = list(range(train_dataset.__len__()))
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = TextLSTM(lstm_hidden_dim, bidirectional, num_lstm_layers, len(dataset_vocab), embedded_matrix.shape[1], pad_idx, num_labels, dropout_prob, vectors=embedded_matrix, use_hatedict=True, requires_grad=True)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate, weight_decay=1e-5)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler, collate_fn=MyDataset.collate_fn)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler, collate_fn=MyDataset.collate_fn)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train_hate(model, criterion, optimizer, train_iter, eval_iter)
print(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn=MyDataset.collate_fn)
loss, scores, reports = eval_hate(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
## Word2Vec + CNN
### Parameters
```
# Model Options
device = 'cuda' if torch.cuda.is_available() else 'cpu' # device
embed_dim = 300 # embed dim
dropout_prob = 0.6 # Dropout prob
num_filters = 300 # num filters
pad_idx = 0 # Padding
num_labels = 2 # num labels
# Training Options
num_epochs = 100 # Epoch
num_fold = 5 # Num fold
batch_size = 32
early_stop = 30 # Early Stop
learning_rate = 2e-5 # learning rate
loss_function = nn.CrossEntropyLoss() # Loss Function
kfold = StratifiedKFold(n_splits=num_fold, random_state=1, shuffle=True)
```
### Normal Case
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = TextCNN(num_filters, num_labels, MAX_SEQUENCE_LENGTH,embedded_matrix.shape[0], embed_dim, pad_idx, dropout_prob, vectors=embedded_matrix)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate, weight_decay=1e-5)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler, collate_fn=MyDataset.collate_fn)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler, collate_fn=MyDataset.collate_fn)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train(model, criterion, optimizer, train_iter, eval_iter)
logger.info(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn=MyDataset.collate_fn)
loss, scores, reports = eval(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
### With HateDict
```
data_idxs = list(range(train_dataset.__len__()))
data_labels = [train_dataset.examples[i][1] for i in data_idxs]
data_idxs = list(range(train_dataset.__len__()))
label_maps = {"NOT": 0, "HOF":1}
avg_scores = {'f1_0': [],'f1_1':[],'acc_0': [],'acc_1': [], 'macro_f1': [], 'macro_acc': [], 'acc': [],'labels': {k:[] for k in label_maps.keys()}}
criterion = nn.CrossEntropyLoss()
total_time = []
total_epoch = []
epoches = []
for idx, (train_index, test_index) in enumerate(kfold.split(data_idxs, data_labels)):
start = time.time()
model = TextCNN(num_filters, num_labels, MAX_SEQUENCE_LENGTH,embedded_matrix.shape[0], embed_dim, pad_idx, dropout_prob, use_hatedict=True, vectors=embedded_matrix)
model.to(device)
optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=learning_rate, weight_decay=1e-5)
train_subsampler = torch.utils.data.SubsetRandomSampler(train_index)
eval_subsampler = torch.utils.data.SubsetRandomSampler(test_index)
train_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=train_subsampler, collate_fn=MyDataset.collate_fn)
eval_iter = DataLoader(train_dataset, batch_size=batch_size, sampler=eval_subsampler, collate_fn=MyDataset.collate_fn)
logger.info(f"Fold-{idx}")
best_model, best_epoch, epoch_time = train_hate(model, criterion, optimizer, train_iter, eval_iter)
logger.info(f"\nBest Epoch: {best_epoch}")
epoches.append(len(list(epoch_time.values())))
total_epoch.append(round(np.mean(list(epoch_time.values()))))
test_iter = DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn=MyDataset.collate_fn)
loss, scores, reports = eval_hate(best_model, criterion, test_iter)
logger.info(f"\tAVG Loss: {loss:.6f}; Accurancy: {scores[0]:.4f}; F1_macro: {scores[5]:.4f}")
logger.info(f"\tClass scores:")
avg_scores['acc'].append(scores[0])
avg_scores['f1_0'].append(scores[1])
avg_scores['f1_1'].append(scores[2])
avg_scores['acc_0'].append(scores[3])
avg_scores['acc_1'].append(scores[4])
avg_scores['macro_f1'].append(scores[5])
for label in label_maps.keys():
avg_scores['labels'][label].append(reports[label]['f1-score'])
logger.info(f"\t\t{label}: {reports[label]['f1-score']}")
total_time.append(get_total_time(start))
logger.info(f"{'='*35}SUMMARY{'='*35}")
logger.info(f"\tAVG Accurancy: {(sum(avg_scores['acc'])/num_fold):.4f}; AVG F1_macro: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info("\tAVG Class scores: ")
for label in label_maps.keys():
logger.info(f"\t\t{label}: {(sum(avg_scores['labels'][label])/num_fold):.4f}")
logger.info(f"Accuracy for label 0: {(sum(avg_scores['acc_0'])/num_fold):.4f}")
logger.info(f"F1 for label 0: {(sum(avg_scores['f1_0'])/num_fold):.4f}")
logger.info(f"Accuracy for label 1: {(sum(avg_scores['acc_1'])/num_fold):.4f}")
logger.info(f"F1 for label 1: {(sum(avg_scores['f1_1'])/num_fold):.4f}")
logger.info(f"Macro F1 Score: {(sum(avg_scores['macro_f1'])/num_fold):.4f}")
logger.info(f"Acc Score: {(sum(avg_scores['acc'])/num_fold):.4f}")
logger.info("-"*30)
logger.info(f'Total 5-Fold Time: {total_time}')
logger.info(f'Total 5-Fold Time: {np.sum(total_time)}')
logger.info(f'Avg Fold Time: {np.mean(total_time)}')
logger.info(f'Avg Epoch Time {np.mean(total_epoch)}')
logger.info(f'Num of Epoches: {epoches}')
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Custom layers
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/tutorials/eager/custom_layers"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/custom_layers.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/custom_layers.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/eager/custom_layers.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
We recommend using `tf.keras` as a high-level API for building neural networks. That said, most TensorFlow APIs are usable with eager execution.
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
```
## Layers: common sets of useful operations
Most of the time when writing code for machine learning models you want to operate at a higher level of abstraction than individual operations and manipulation of individual variables.
Many machine learning models are expressible as the composition and stacking of relatively simple layers, and TensorFlow provides both a set of many common layers as a well as easy ways for you to write your own application-specific layers either from scratch or as the composition of existing layers.
TensorFlow includes the full [Keras](https://keras.io) API in the tf.keras package, and the Keras layers are very useful when building your own models.
```
# In the tf.keras.layers package, layers are objects. To construct a layer,
# simply construct the object. Most layers take as a first argument the number
# of output dimensions / channels.
layer = tf.keras.layers.Dense(100)
# The number of input dimensions is often unnecessary, as it can be inferred
# the first time the layer is used, but it can be provided if you want to
# specify it manually, which is useful in some complex models.
layer = tf.keras.layers.Dense(10, input_shape=(None, 5))
```
The full list of pre-existing layers can be seen in [the documentation](https://www.tensorflow.org/api_docs/python/tf/keras/layers). It includes Dense (a fully-connected layer),
Conv2D, LSTM, BatchNormalization, Dropout, and many others.
```
# To use a layer, simply call it.
layer(tf.zeros([10, 5]))
# Layers have many useful methods. For example, you can inspect all variables
# in a layer using `layer.variables` and trainable variables using
# `layer.trainable_variables`. In this case a fully-connected layer
# will have variables for weights and biases.
layer.variables
# The variables are also accessible through nice accessors
layer.kernel, layer.bias
```
## Implementing custom layers
The best way to implement your own layer is extending the tf.keras.Layer class and implementing:
* `__init__` , where you can do all input-independent initialization
* `build`, where you know the shapes of the input tensors and can do the rest of the initialization
* `call`, where you do the forward computation
Note that you don't have to wait until `build` is called to create your variables, you can also create them in `__init__`. However, the advantage of creating them in `build` is that it enables late variable creation based on the shape of the inputs the layer will operate on. On the other hand, creating variables in `__init__` would mean that shapes required to create the variables will need to be explicitly specified.
```
class MyDenseLayer(tf.keras.layers.Layer):
def __init__(self, num_outputs):
super(MyDenseLayer, self).__init__()
self.num_outputs = num_outputs
def build(self, input_shape):
self.kernel = self.add_variable("kernel",
shape=[int(input_shape[-1]),
self.num_outputs])
def call(self, input):
return tf.matmul(input, self.kernel)
layer = MyDenseLayer(10)
print(layer(tf.zeros([10, 5])))
print(layer.trainable_variables)
```
Overall code is easier to read and maintain if it uses standard layers whenever possible, as other readers will be familiar with the behavior of standard layers. If you want to use a layer which is not present in `tf.keras.layers`, consider filing a [github issue](http://github.com/tensorflow/tensorflow/issues/new) or, even better, sending us a pull request!
## Models: composing layers
Many interesting layer-like things in machine learning models are implemented by composing existing layers. For example, each residual block in a resnet is a composition of convolutions, batch normalizations, and a shortcut.
The main class used when creating a layer-like thing which contains other layers is tf.keras.Model. Implementing one is done by inheriting from tf.keras.Model.
```
class ResnetIdentityBlock(tf.keras.Model):
def __init__(self, kernel_size, filters):
super(ResnetIdentityBlock, self).__init__(name='')
filters1, filters2, filters3 = filters
self.conv2a = tf.keras.layers.Conv2D(filters1, (1, 1))
self.bn2a = tf.keras.layers.BatchNormalization()
self.conv2b = tf.keras.layers.Conv2D(filters2, kernel_size, padding='same')
self.bn2b = tf.keras.layers.BatchNormalization()
self.conv2c = tf.keras.layers.Conv2D(filters3, (1, 1))
self.bn2c = tf.keras.layers.BatchNormalization()
def call(self, input_tensor, training=False):
x = self.conv2a(input_tensor)
x = self.bn2a(x, training=training)
x = tf.nn.relu(x)
x = self.conv2b(x)
x = self.bn2b(x, training=training)
x = tf.nn.relu(x)
x = self.conv2c(x)
x = self.bn2c(x, training=training)
x += input_tensor
return tf.nn.relu(x)
block = ResnetIdentityBlock(1, [1, 2, 3])
print(block(tf.zeros([1, 2, 3, 3])))
print([x.name for x in block.trainable_variables])
```
Much of the time, however, models which compose many layers simply call one layer after the other. This can be done in very little code using tf.keras.Sequential
```
my_seq = tf.keras.Sequential([tf.keras.layers.Conv2D(1, (1, 1),
input_shape=(
None, None, 3)),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(2, 1,
padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(3, (1, 1)),
tf.keras.layers.BatchNormalization()])
my_seq(tf.zeros([1, 2, 3, 3]))
```
# Next steps
Now you can go back to the previous notebook and adapt the linear regression example to use layers and models to be better structured.
```
```
| github_jupyter |
```
import fitsio
import numpy as np
from matplotlib import pyplot as plt
params = {'legend.fontsize': 'x-large',
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large',
'figure.facecolor':'w'}
plt.rcParams.update(params)
fs = fitsio.read('/global/cscratch1/sd/adematti/legacysim/dr9/ebv1000shaper/south/file0_rs0_skip0/merged/matched_input.fits') #obiwan outputs in DECaLS
seld = fs['flux_g']*0 == 0 #select detected
seld &= fs['flux_g'] > 0.2 #also select within this relevant gflux range
seld &= fs['flux_g'] < 2
fsd = fs[seld]
a = plt.hist(fsd['fiberflux_g']/fsd['flux_g'],bins=30)
plt.xlabel('output fiberflux_g/flux_g')
plt.ylabel('# in bin')
plt.title('Obiwan outputs with 0.2 < flux_g < 2')
plt.show()
```
The spike is for type PSF, take a look at what the histograms look like by type:
```
for tp in np.unique(fsd['type']):
wt = fsd['type'] == tp
if tp != 'PSF':
plt.hist(fsd[wt]['fiberflux_g']/fsd[wt]['flux_g'],bins=a[1],density=True,label=tp,histtype='step',linewidth=3)
plt.legend(loc='upper left')
plt.xlabel('output fiberflux_g/flux_g')
plt.ylabel('relative fraction in bin')
plt.title('Obiwan outputs with 0.2 < flux_g < 2')
plt.show()
```
Generally, this makes sense. The more complex the type, the more extended it is and the smaller the fraction of within the fiber.
What happens if we now look at the recovered fiberflux vs input?
```
b = plt.hist(fsd['fiberflux_g']/fsd['input_flux_g'],bins=30,range=(0,1.5))
plt.xlabel('output fiberflux_g / input flux_g')
plt.ylabel('# in bin')
plt.title('Obiwan outputs with 0.2 < flux_g < 2')
plt.show()
for tp in np.unique(fsd['type']):
wt = fsd['type'] == tp
#if tp != 'PSF':
plt.hist(fsd[wt]['fiberflux_g']/fsd[wt]['input_flux_g'],bins=b[1],density=True,label=tp,histtype='step',linewidth=3)
plt.legend(loc='upper left')
plt.xlabel('output fiberflux_g / input flux_g')
plt.ylabel('relative fraction in bin')
plt.title('Obiwan outputs with 0.2 < flux_g < 2')
plt.show()
a = plt.hist(fsd['input_galdepth_g'],range=(200,3000))
b = plt.hist(fsd['input_galdepth_g'],weights=fsd['fiberflux_g']/fsd['input_flux_g'],bins=a[1])
c = plt.hist(fsd['input_galdepth_g'],weights=fsd['flux_g']/fsd['input_flux_g'],bins=a[1])
plt.clf()
plt.plot(a[1][:-1],b[0]/a[0],label='fiberflux_g / input_flux_g')
plt.plot(a[1][:-1],c[0]/a[0]*.56,label='0.56 x flux_g / input_flux_g')
plt.legend()
plt.xlabel('galdepth_g')
plt.ylabel('ratio to input flux')
plt.title('Obiwan outputs with 0.2 < flux_g < 2')
plt.grid(alpha=0.5)
plt.show()
```
| github_jupyter |
# <center> Policy Gradient Methods</center>
## Table of Content
1) Introduction: Policy Gradient vs the World, Advantages and Disadvantages
2) REINFORCE: Simplest Policy Gradient Method
3) Actor-Critic Methods
4) Additional Enhancements to Actor-Critic Methods
# <center>Introduction</center>
## <center> Introduction </center>
<center><img src="img/pg_1.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Value-Based Vs Policy-Based RL</center>
<center><img src="img/pg_2.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Why Policy-Based RL</center>
<center><img src="img/pg_3.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Can Learning Policy be easier than Learning Values of states?</center>
* The policy may be a simpler function to approximate.
* This is the simplest advantage that policy parameterization may have over action-value parameterization.
Why?
* Problems vary in the complexity of their policies and action-value functions.
* For some, the action-value function is simpler and thus easier to approximate.
* For others, the policy is simpler.
** In the latter case a policy-based method will typically be faster to learn and yield a superior asymptotic policy.**
Example: In Robotics Tasks with continuous Action space.
## <center> Example of Stochastic Optimal Policy</center>
<center><img src="img/pg_4.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Example of Stochastic Optimal Policy</center>
<center><img src="img/pg_5.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Example of Stochastic Optimal Policy</center>
<center><img src="img/pg_6.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Example of Stochastic Optimal Policy</center>
<center><img src="img/pg_7.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## Why not use softmax of Action-Values for stochastic Policies?
* This alone would not approach determinism if and when required.
* The action-value estimates would differ by a finite amount, translating to specific probabilities other than 0 and 1.
* If softmax + Temprature Paramenter T: T could be reduced over time to approach determinism.
* However, in practice it would be difficult to choose the reduction schedule, or even the initial temperature, without more knowledge of the true action values.
* Whereas, Policy gradient is driven to produce the optimal stochastic policy.
* If the optimal policy is deterministic, then the preferences of the optimal actions will be driven infinitely higher than all suboptimal actions
## <center>REINFORCE: Simplest Policy Gradient Method</center>
## <center>Quality Measure of Policy</center>
<center><img src="img/pg_8.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Policy Optimisation</center>
<center><img src="img/pg_9.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Gradient Ascent</center>
<center><img src="img/pg_10.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Gradient Ascent - FDM</center>
<center><img src="img/pg_11.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Analytic Gradient Ascent</center>
<center><img src="img/pg_12.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Example- Softmax Policy</center>
<center><img src="img/pg_13.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Example- Gaussian Policy</center>
<center><img src="img/pg_14.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>One-step MDP</center>
<center><img src="img/pg_15.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Policy Gradient Theorem</center>
<center><img src="img/pg_16.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Policy Gradient Theorem-Proof</center>
<center><img src="img/sutton_1.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Monte-Carlo Policy Gradient (REINFORCE)</center>
<center><img src="img/pg_17.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> PuckWorld Example</center>
<center><img src="img/pg_18.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
** DQN Demo [Reinforce.js](http://cs.stanford.edu/people/karpathy/reinforcejs/puckworld.html) **
## <center> REINFORCE with Baseline</center>
* REINFORCE has good theoretical convergence properties.
* The expected update over an episode is in the same direction as the performance gradient.
* This assures:
* An improvement in expected performance for sufficiently small $\alpha$, and
* Convergence to a local optimum under standard stochastic approximation conditions.
* **However**,
* Monte Carlo method REINFORCE may be of high variance, and thus
* slow to learn.
** Can we reduce the variance somehow? **
## <center> REINFORCE with Baseline</center>
* The derivative of the quality $\eta(\theta)$ of policy network can be written as
<center><img src="img/sutton_5.JPG" alt="Multi-armed Bandit" style="width: 400px;"/></center>
* Instead of using the Rewards/Action Vaules generated directly, we first compare it with a baseline:
<center><img src="img/sutton_2.JPG" alt="Multi-armed Bandit" style="width: 500px;"/></center>
* The baseline can be any function, even a random variable,
* **Only Condition**: Should not vary with action $a$;
* **Any guesses why?**
<center><img src="img/sutton_3.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
* Finally the Update equation becomes:
<center><img src="img/sutton_4.JPG" alt="Multi-armed Bandit" style="width: 500px;"/></center>
# <center> Actor Critic Methods</center>
## <center> Reducing Variance Using a Critic</center>
<center><img src="img/pg_19.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Estimating the Action-Value Function</center>
<center><img src="img/pg_20.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Action Value Actor Critic</center>
<center><img src="img/pg_21.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Bias In Actor-Critic Algorithm</center>
<center><img src="img/pg_22.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Compatible Function Approximation</center>
<center><img src="img/pg_23.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Compatible Function Approximation- Proof</center>
<center><img src="img/pg_24.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
# <center>Additional Enhancements to Actor Critic</center>
## <center> Actor Critic with Baseline</center>
<center><img src="img/pg_25.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Estimating the Advantage Function</center>
<center><img src="img/pg_26.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Estimating the Advantage Function</center>
<center><img src="img/pg_27.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center> Critics at different Time-Scales</center>
<center><img src="img/pg_27.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>`
## <center> Actors at different Time-Scale</center>
<center><img src="img/pg_28.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Policy Gradient with Eligibility Traces</center>
<center><img src="img/pg_29.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Policy Gradient with Eligibility Traces</center>
<center><img src="img/pg_30.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
## <center>Summary</center>
<center><img src="img/pg_34.JPG" alt="Multi-armed Bandit" style="width: 700px;"/></center>
| github_jupyter |
# Decision Tree Classification with Standard Scalar
### Required Packages
```
!pip install imblearn
import numpy as np
import pandas as pd
import seaborn as se
import warnings
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import RandomOverSampler
from sklearn.tree import DecisionTreeClassifier,plot_tree
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
file_path= ""
```
List of features which are required for model training .
```
features=[]
```
Target feature for prediction.
```
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path);
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
<h3>Data Scaling </h3><br>
Standardizing a dataset involves rescaling the distribution of values so that the mean of observed values is 0 and the standard deviation is 1.
Like normalization, standardization can be useful, when your data has input values with differing scales. Standardization assumes that your observations fit a Gaussian distribution (bell curve) with a well-behaved mean and standard deviation.
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
x_train = scaler.fit_transform(x_train)
x_test = scaler.fit_transform(x_test)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
Decision tree is the most powerful and popular tool for classification and prediction. A Decision tree is a flowchart like tree structure, where each internal node denotes a test on an attribute, each branch represents an outcome of the test, and each leaf node holds a outcome label.
As with other classifiers, DecisionTreeClassifier takes as input two arrays: an array X, sparse or dense, of shape (n_samples, n_features) holding the training samples, and an array Y of integer values, shape (n_samples,), holding the class labels for the training samples.
It is capable of both binary ([-1,1] or [0,1]) classification and multiclass ([0, …,K-1]) classification.
#### Model Tuning Parameter
> - criterion -> The function to measure the quality of a split. Supported criteria are “gini” for the Gini impurity and “entropy” for the information gain.
> - max_depth -> The maximum depth of the tree. If None, then nodes are expanded until all leaves are pure or until all leaves contain less than min_samples_split samples.
> - max_leaf_nodes -> Grow a tree with max_leaf_nodes in best-first fashion. Best nodes are defined as relative reduction in impurity. If None then unlimited number of leaf nodes.
> - max_features -> The number of features to consider when looking for the best split: **{auto , sqrt, log2}**
```
model = DecisionTreeClassifier(random_state=123)
model.fit(x_train,y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
Plotting confusion matrix for the predicted values versus actual values.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* where:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Feature Importances.
The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
```
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
```
#### Tree Plot
Plot a decision tree.The visualization is fit automatically to the size of the axis. Use the figsize or dpi arguments of plt.figure to control the size of the rendering.
```
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (3,3), dpi=400)
cls_target = [str(x) for x in pd.unique(y_train)]
cls_target.sort()
plot_tree(model,feature_names = X.columns, class_names=cls_target,filled = True)
fig.savefig('./tree.png')
```
#### Creator: Anu Rithiga B , Github: [Profile - Iamgrootsh7](https://github.com/iamgrootsh7)
| github_jupyter |
```
from selenium import webdriver
import pandas as pd
import numpy as np
import time
import datetime
import subprocess
import re
import os
import requests
keywords = ["wikipedia"]
journals = ['information systems research', 'mis quarterly', 'journal of management information systems',
'journal of the association for information systems', 'management science', 'operational research'] # source: "MISQ"
# journals = ['information systems research']
# authors = ["Kuznets"] # author:Kuznets
fpath = "/Users/Nico/test/test_googlecrawer"
alias = {'information systems research':"ISR", 'mis quarterly':'MISQ', 'journal of management information systems':"JMIS",
'journal of the association for information systems':"JAIS", 'management science':'MS', 'operational research':"OR"}
# options = webdriver.ChromeOptions()
# # options.add_argument('headless')
# options.binary_location = '/usr/local/bin/chromedriver'
# options.add_argument('headless')
# options.add_argument("--no-sandbox");
# options.add_argument("--disable-dev-shm-usage")
def getbibTeX(article, driver):
bib = article.find_element_by_css_selector("div[class=gs_fl]").find_element_by_css_selector("a[class=gs_or_cit\ gs_nph]")
bib.click()
time.sleep(2) ## sleep wait for the ajax to load
driver.find_element_by_css_selector("div[id=gs_citi]").find_element_by_css_selector("a[class=gs_citi]").click()
bib_text = driver.find_element_by_tag_name("body").text.replace("\n", "")
driver.back()
driver.find_element_by_css_selector("span[class=gs_ico]").click()
return bib_text
getbibTeX(article[2], driver)
def getInfo(article, driver):
default = {"title": "NA", "author": "NA", "journal": "NA", "year":"NA"}
default['title'] = article.find_element_by_class_name("gs_rt").text.lower()
infobox = article.find_element_by_class_name("gs_a").text
default['author'], default['journal'], default['year'] = parse(infobox)
return default
getInfo(article[0], driver)
def getPdf(article, driver):
try:
tmp = article.find_element_by_css_selector("div[class=gs_or_ggsm")
pdf_link = tmp.find_element_by_tag_name("a").get_attribute("href")
except:
pdf_link = "NA"
return pdf_link
getPdf(article[1], driver)
def downloadPdf(output, link):
response = requests.get(link)
with open(output, 'wb') as f:
f.write(response.content)
def parse(infobox):
infobox = infobox.lower().split("-")
infobox = [c.strip() for c in infobox]
author = infobox[0].split(",")[0]
journal = infobox[1].split(",")[0]
year = infobox[1].split(",")[1].strip()
return author, journal, year
# for test
search_keyword = '''online communities'''
driver = webdriver.Chrome()
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
driver.get('https://scholar.google.com/')
input_element = driver.find_element_by_name("q")
input_element.clear()
input_element.send_keys(search_keyword)
input_element.submit()
elements = driver.find_elements_by_css_selector("div[class=gs_r\ gs_or\ gs_scl]")
elements[0].find_elements_by_class_name("gs_fl")
tmp = elements[5].find_elements_by_class_name("gs_fl")
[c.find_element_by_css_selector("a[href^='/scholar?cites']").text for c in tmp]
class Article:
def __init__(self, keywords, target_journal, folder):
self.keywords = keywords
self.target_journal = target_journal
self.output_folder = folder
self.create_folder()
self.total_article = {}
def create_folder(self):
self.output_fpath = "/".join([self.output_folder, self.keywords, self.target_journal])
if not os.path.exists(self.output_fpath):
os.makedirs(self.output_fpath)
print('created folder {0}'.format(self.output_fpath))
def getInfo(self, article, driver):
default = {"title": "NA", "author": "NA", "journal": "NA", "year":"NA", "log": "NA"}
default['title'] = article.find_element_by_class_name("gs_rt").text.lower()
default['title'] = re.sub("[^a-z0-9 ]", "", default['title'])
infobox = article.find_element_by_class_name("gs_a").text
default['author'], default['journal'], default['year'] = parse(infobox)
return default
def getPdf(self, article, driver):
tmp = article.find_element_by_css_selector("div[class=gs_or_ggsm")
pdf_link = tmp.find_element_by_tag_name("a").get_attribute("href")
return pdf_link
def getFileName(self, alias=alias):
by = ["author", "year", "title", "journal"]
if alias:
if self.info['journal'] in alias.keys():
self.info['journal-short'] = alias[self.info['journal']]
by = ["author", "year", "title", "journal-short"]
if len(self.info['title'].split(" ")) > 10:
self.info['title-short'] = " ".join(self.info['title'].split(" ")[:10])
if "journal-short" in self.info.keys():
by = ["author", "year", "title-short", "journal-short"]
else:
by = ["author", "year", "title-short", "journal"]
filename = "-".join([self.info[c] for c in by]) + ".pdf"
return filename
def fit(self, article, driver, num):
try:
self.info = self.getInfo(article, driver)
except:
print("article info parse error!")
self.info = None
if self.info:
try:
self.pdf = self.getPdf(article, driver)
except:
self.info['log'] = "pdf missing"
self.pdf = None
if self.pdf:
self.filename = self.getFileName()
output = self.output_fpath + "/" + self.filename
try:
downloadPdf(output, self.pdf)
except:
self.log = self.info['log'] + "||| pdf download error"
self.total_article[num] = self.info
# if hasattr(self, 'log'):
# now = datetime.datetime.now()
# logfile_path = self.output_fpath + "/" + "log_{0}.txt".format(now.strftime("%m-%d-%Y"))
# if not os.path.exists(logfile_path):
# subprocess.call("touch {0}".format(logfile_path), shell=True)
# with open(logfile_path, "a") as f:
# f.writelines(self.log)
def run(keywords, journals, recursive = 6):
driver = webdriver.Chrome()
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
driver.get('https://scholar.google.com/')
for i in keywords:
for j in journals:
cnt = 1
articles = Article(i, j, fpath)
search_keyword = " ".join([i.lower(), '''source:"{}"'''.format(j.lower())])
print("current search key: {0}".format(search_keyword))
input_element = driver.find_element_by_name("q")
input_element.clear()
input_element.send_keys(search_keyword)
input_element.submit()
time.sleep(2)
for n in range(recursive):
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
elements = driver.find_elements_by_css_selector("div[class=gs_r\ gs_or\ gs_scl]")
for e in elements:
try:
articles.fit(e, driver, cnt)
except:
print("page {} number {} parse error!".format(n, cnt))
cnt += 1
try:
driver.find_element_by_css_selector("span[class=gs_ico\ gs_ico_nav_next]").click()
time.sleep(5)
except:
pass
log = pd.DataFrame(articles.total_article).T
now = datetime.datetime.now()
log.to_csv(articles.output_fpath+"/"+"logfile_{}.txt".format(now.strftime("%m-%d-%Y")), sep="\t")
driver.quit()
# article = driver.find_element_by_css_selector("div[class^=gs_ri")
journals = ['journal of management information systems',
'journal of the association for information systems', 'management science', 'operational research']
run(keywords, journals, recursive=6)
```
| github_jupyter |
```
import keras.models
import os
from scipy.io import wavfile
import pandas as pd
import numpy as np
from keras.models import load_model
from sklearn.metrics import accuracy_score
from tqdm import tqdm
from python_speech_features import mfcc
import pickle
class Config:
def __init__(self, mode='conv', nfilt=26, nfeat = 13, nfft = 512, rate = 16000): #filtered out
self.mode = mode
self.nfilt = nfilt
self.nfeat = nfeat
self.rate = rate
self.nfft = nfft
self.step = int(rate/10) #0.1 sec, how much data computing while creating window
self.model_path = os.path.join('/content/gdrive/My Drive/models', mode + '.model')
self.p_path = os.path.join('/content/gdrive/My Drive/pickles', mode + '.p')
def build_predictions(audio_dir):
y_true = []
y_pred = []
fn_prob = {}
print('Extracting features from audio')
for fn in tqdm(os.listdir(audio_dir)):
print(audio_dir)
print(fn)
rate, wav = wavfile.read(os.path.join(audio_dir, fn))
label = fn2class[fn]
c = classes.index(label)
y_prob = []
for i in range(0, wav.shape[0]-config.step, config.step):#cannot go further
sample = wav[i:i+config.step]
x = mfcc(sample, rate, numcep = config.nfeat, nfilt = config.nfilt, nfft = config.nfft)
x = (x - config.min)/(config.max - config.min)#range
if config.mode == 'conv':
x = x.reshape(1, x.shape[0], x.shape[1], 1)
elif config.mode=='time':
x = np.expand_dims(x, axis = 0) #expand to 1 sample
elif config.mode == 'convtime':
x = x.reshape(1, x.shape[0], x.shape[1], 1)
y_hat = model.predict(x)
y_prob.append(y_hat)
y_pred.append(np.argmax(y_hat))
y_true.append(c)
fn_prob[fn] = np.mean(y_prob, axis = 0).flatten() #or that would be crap
return y_true, y_pred, fn_prob
df = pd.read_csv('/content/gdrive/My Drive/valrandmus10.csv')#can remove if no classification
classes = list(np.unique(df.label))#all names of genres
print(classes)
fn2class = dict(zip(df.fname, df.label))
p_path = os.path.join('/content/gdrive/My Drive/pickles', 'conv.p')
with open(p_path, 'rb') as handle:
config = pickle.load(handle)
model = load_model("/content/gdrive/My Drive/models/conv.model")
y_true, y_pred, fn_prob = build_predictions('/content/gdrive/My Drive/valrandmus1/')
acc_score = accuracy_score(y_true = y_true, y_pred = y_pred)
y_probs = []
for i,row in df.iterrows():
print(row.fname)
y_prob = fn_prob[row.fname]
y_probs.append(y_prob)
for c ,p in zip(classes, y_prob):
df.at[i, c] = p
print(classes)
y_pred = [classes[np.argmax(y)] for y in y_probs]
df['y_pred'] = y_pred
df.to_csv('/content/gdrive/My Drive/convconvconv.csv', index = False)
!pip install python_speech_features
from google.colab import drive
drive.mount('/content/gdrive')
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/FeatureCollection/select_columns.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/FeatureCollection/select_columns.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/FeatureCollection/select_columns.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
fc = ee.FeatureCollection('TIGER/2018/States')
print(fc.first().getInfo())
new_fc = fc.select(['STUSPS', 'NAME', 'ALAND'], ['abbr', 'name', 'area'])
print(new_fc.first().getInfo())
propertyNames = new_fc.first().propertyNames()
print(propertyNames.getInfo())
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Project: Titanic Dataset
## Table of Contents
<ul>
<li><a href="#intro">Introduction</a></li>
<li><a href="#wrangling">Data Wrangling</a></li>
<li><a href="#eda">Exploratory Data Analysis</a></li>
</ul>
<a id='intro'></a>
## Introduction
We will explore Titanic Dataset from Kaggle and will implement Machine Learning model to predict the Survival chance.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
data = pd.read_csv("../Data/titanic_data.csv")
data.head()
data.info()
data.describe()
```
-----
<a id='eda'></a>
# Exploratory data analysis (EDA)
### Number of Survived Passengers
```
data["Survived"].value_counts()
```
It seem like 342 peple survived.
### Gender Distribution
```
data["Sex"].value_counts()
data["Sex"].value_counts().plot(kind = "bar");
```
### Fare Distribution
```
data["Fare"].hist();
```
---------
## Bar Charts plot for categorical features
- PClass
- Sex
- SibSp
- Parch
- Embarked
- Cabin
```
def bar_chart(feature):
survived = data[data["Survived"] == 1][feature].value_counts()
not_survived = data[data["Survived"] == 0][feature].value_counts()
data_new = pd.DataFrame([survived, not_survived])
data_new.index = ["Survived", "Dead"]
data_new.plot(kind = "bar", stacked = True, figsize = (10, 5))
def total_ratio(feature):
total = data[feature].count()
print("Total Passenger: ", total)
print()
survived = (data[data["Survived"] == 1][feature].value_counts() / total) * 100
not_survived = (data[data["Survived"] == 0][feature].value_counts() /total) * 100
print("Survival Ratio %")
print(survived)
print()
print("Not Survival Ratio %")
print(not_survived)
```
### Survival by Gender
```
bar_chart("Sex")
plt.title("Survial by Gender");
```
It seems like more Female survived than Male passengers.
```
total_ratio("Sex")
```
-------
### Survival by Pclass
```
bar_chart("Pclass")
plt.title("Survial by Pclass");
total_ratio("Pclass")
```
-------
### Survival by Sibsp
```
bar_chart("SibSp")
plt.title("Survival by Sibilings and Spouse Numbers");
total_ratio("SibSp")
```
---------
### Survival by Parch
```
bar_chart("Parch")
plt.title("Survival by Parent and Childern Numbers");
total_ratio("Parch")
```
### Survival by Title
it may not make sense. but we will explore this too.
```
temp = data
temp.head()
#extract title from Name column
title = temp["Name"].apply(lambda name: name.split(", ")[1].split(".")[0])
title.value_counts()
data["Title"] = data["Name"].apply(lambda name: name.split(", ")[1].split(".")[0])
# temp_list = [data]
# for passenger in temp_list:
# passenger["Title"] = passenger["Name"].str.extract(" ([A-Za-z]+)\.", expand = False)
data.head()
# type(temp_list)
# temp_list
bar_chart("Title")
plt.title("Survival by Title");
total_ratio("Title")
```
-------
<a id = "wrangling"></a>
# Data Cleaning
## Mapping the features to Numbers
### Change `Title` column values to Numbers
```
data.dtypes
title_mapping = {"Mr": 0, "Miss":1, "Mrs": 2, "Ms": 1,
"Sir": 3,"Master": 3, "Rev": 3, "Dr": 3, "Capt": 3, "Major": 3, "Col": 3, "Jonkheer": 3, "Don": 3, "Mlle": 3, "Mme": 3, "Lady": 3, "the Countess": 3, "Dona": 3}
temp_list = [data]
for passenger in temp_list:
passenger["Title"] = passenger["Title"].map(title_mapping)
data.head(2)
```
### Change `Sex` column values to Number
```
data.replace(["male", "female"], ["1", "0"], inplace = True)
data.head(2)
```
--------
# Handling Missing Values
```
data.info()
data.isna().sum()
```
It seems like Age, Cabin and Embarked columns have some missing values.
### Fill mean value for `Age` null values
```
#mean Age by "sex" and "pclass"
mean_age = data.groupby(["Sex", "Pclass"])["Age"].mean()
mean_age
# function to fill the null age values
def fill_nan_age(row):
if pd.isnull(row["Age"]):
return mean_age[row["Sex"], row["Pclass"]]
else:
return row["Age"]
data["Age"] = data.apply(fill_nan_age, axis = 1)
data["Age"].isna().sum()
```
-----
# Seperating/ Categorizing the uncategorized data in the dataset
As passenger had various ages, we will categorize the ages into different bins.
- child: 0
- young: 1
- adult: 2
- mid-age: 3
- senior: 4
```
def binning_age(age):
if age <= 16:
return 0
elif age > 16 and age <= 26:
return 1
elif age > 26 and age <= 36:
return 2
elif age > 36 and age <= 62:
return 3
else:
return 4
data["Age"] = data["Age"].apply(binning_age)
data.head(2)
data.info()
data["Age"].value_counts()
bar_chart("Age")
data.head()
data.info()
```
Cabin and Embarked still have missing values. but as we won't use them later, we will skip handling for them.
--------
### Save the cleaned data
```
data.to_csv("../Data/titanic_data_cleaned.csv", index=False)
cleaned_data = pd.read_csv("../Data/titanic_data_cleaned.csv")
cleaned_data.head()
```
------
| github_jupyter |
# Tile Coding
---
Tile coding is an innovative way of discretizing a continuous space that enables better generalization compared to a single grid-based approach. The fundamental idea is to create several overlapping grids or _tilings_; then for any given sample value, you need only check which tiles it lies in. You can then encode the original continuous value by a vector of integer indices or bits that identifies each activated tile.
### 1. Import the Necessary Packages
```
# Import common libraries
import sys
import gym
import numpy as np
import matplotlib.pyplot as plt
# Set plotting options
%matplotlib inline
plt.style.use('ggplot')
np.set_printoptions(precision=3, linewidth=120)
```
### 2. Specify the Environment, and Explore the State and Action Spaces
We'll use [OpenAI Gym](https://gym.openai.com/) environments to test and develop our algorithms. These simulate a variety of classic as well as contemporary reinforcement learning tasks. Let's begin with an environment that has a continuous state space, but a discrete action space.
```
# Create an environment
env = gym.make('Acrobot-v1')
env.seed(505);
# Explore state (observation) space
print("State space:", env.observation_space)
print("- low:", env.observation_space.low)
print("- high:", env.observation_space.high)
# Explore action space
print("Action space:", env.action_space)
```
Note that the state space is multi-dimensional, with most dimensions ranging from -1 to 1 (positions of the two joints), while the final two dimensions have a larger range. How do we discretize such a space using tiles?
### 3. Tiling
Let's first design a way to create a single tiling for a given state space. This is very similar to a uniform grid! The only difference is that you should include an offset for each dimension that shifts the split points.
For instance, if `low = [-1.0, -5.0]`, `high = [1.0, 5.0]`, `bins = (10, 10)`, and `offsets = (-0.1, 0.5)`, then return a list of 2 NumPy arrays (2 dimensions) each containing the following split points (9 split points per dimension):
```
[array([-0.9, -0.7, -0.5, -0.3, -0.1, 0.1, 0.3, 0.5, 0.7]),
array([-3.5, -2.5, -1.5, -0.5, 0.5, 1.5, 2.5, 3.5, 4.5])]
```
Notice how the split points for the first dimension are offset by `-0.1`, and for the second dimension are offset by `+0.5`. This might mean that some of our tiles, especially along the perimeter, are partially outside the valid state space, but that is unavoidable and harmless.
```
def create_tiling_grid(low, high, bins=(10, 10), offsets=(0.0, 0.0)):
"""Define a uniformly-spaced grid that can be used for tile-coding a space.
Parameters
----------
low : array_like
Lower bounds for each dimension of the continuous space.
high : array_like
Upper bounds for each dimension of the continuous space.
bins : tuple
Number of bins or tiles along each corresponding dimension.
offsets : tuple
Split points for each dimension should be offset by these values.
Returns
-------
grid : list of array_like
A list of arrays containing split points for each dimension.
"""
# TODO: Implement this
pass
low = [-1.0, -5.0]
high = [1.0, 5.0]
create_tiling_grid(low, high, bins=(10, 10), offsets=(-0.1, 0.5)) # [test]
```
You can now use this function to define a set of tilings that are a little offset from each other.
```
def create_tilings(low, high, tiling_specs):
"""Define multiple tilings using the provided specifications.
Parameters
----------
low : array_like
Lower bounds for each dimension of the continuous space.
high : array_like
Upper bounds for each dimension of the continuous space.
tiling_specs : list of tuples
A sequence of (bins, offsets) to be passed to create_tiling_grid().
Returns
-------
tilings : list
A list of tilings (grids), each produced by create_tiling_grid().
"""
# TODO: Implement this
pass
# Tiling specs: [(<bins>, <offsets>), ...]
tiling_specs = [((10, 10), (-0.066, -0.33)),
((10, 10), (0.0, 0.0)),
((10, 10), (0.066, 0.33))]
tilings = create_tilings(low, high, tiling_specs)
```
It may be hard to gauge whether you are getting desired results or not. So let's try to visualize these tilings.
```
from matplotlib.lines import Line2D
def visualize_tilings(tilings):
"""Plot each tiling as a grid."""
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
linestyles = ['-', '--', ':']
legend_lines = []
fig, ax = plt.subplots(figsize=(10, 10))
for i, grid in enumerate(tilings):
for x in grid[0]:
l = ax.axvline(x=x, color=colors[i % len(colors)], linestyle=linestyles[i % len(linestyles)], label=i)
for y in grid[1]:
l = ax.axhline(y=y, color=colors[i % len(colors)], linestyle=linestyles[i % len(linestyles)])
legend_lines.append(l)
ax.grid('off')
ax.legend(legend_lines, ["Tiling #{}".format(t) for t in range(len(legend_lines))], facecolor='white', framealpha=0.9)
ax.set_title("Tilings")
return ax # return Axis object to draw on later, if needed
visualize_tilings(tilings);
```
Great! Now that we have a way to generate these tilings, we can next write our encoding function that will convert any given continuous state value to a discrete vector.
### 4. Tile Encoding
Implement the following to produce a vector that contains the indices for each tile that the input state value belongs to. The shape of the vector can be the same as the arrangment of tiles you have, or it can be ultimately flattened for convenience.
You can use the same `discretize()` function here from grid-based discretization, and simply call it for each tiling.
```
def discretize(sample, grid):
"""Discretize a sample as per given grid.
Parameters
----------
sample : array_like
A single sample from the (original) continuous space.
grid : list of array_like
A list of arrays containing split points for each dimension.
Returns
-------
discretized_sample : array_like
A sequence of integers with the same number of dimensions as sample.
"""
# TODO: Implement this
pass
def tile_encode(sample, tilings, flatten=False):
"""Encode given sample using tile-coding.
Parameters
----------
sample : array_like
A single sample from the (original) continuous space.
tilings : list
A list of tilings (grids), each produced by create_tiling_grid().
flatten : bool
If true, flatten the resulting binary arrays into a single long vector.
Returns
-------
encoded_sample : list or array_like
A list of binary vectors, one for each tiling, or flattened into one.
"""
# TODO: Implement this
pass
# Test with some sample values
samples = [(-1.2 , -5.1 ),
(-0.75, 3.25),
(-0.5 , 0.0 ),
( 0.25, -1.9 ),
( 0.15, -1.75),
( 0.75, 2.5 ),
( 0.7 , -3.7 ),
( 1.0 , 5.0 )]
encoded_samples = [tile_encode(sample, tilings) for sample in samples]
print("\nSamples:", repr(samples), sep="\n")
print("\nEncoded samples:", repr(encoded_samples), sep="\n")
```
Note that we did not flatten the encoding above, which is why each sample's representation is a pair of indices for each tiling. This makes it easy to visualize it using the tilings.
```
from matplotlib.patches import Rectangle
def visualize_encoded_samples(samples, encoded_samples, tilings, low=None, high=None):
"""Visualize samples by activating the respective tiles."""
samples = np.array(samples) # for ease of indexing
# Show tiling grids
ax = visualize_tilings(tilings)
# If bounds (low, high) are specified, use them to set axis limits
if low is not None and high is not None:
ax.set_xlim(low[0], high[0])
ax.set_ylim(low[1], high[1])
else:
# Pre-render (invisible) samples to automatically set reasonable axis limits, and use them as (low, high)
ax.plot(samples[:, 0], samples[:, 1], 'o', alpha=0.0)
low = [ax.get_xlim()[0], ax.get_ylim()[0]]
high = [ax.get_xlim()[1], ax.get_ylim()[1]]
# Map each encoded sample (which is really a list of indices) to the corresponding tiles it belongs to
tilings_extended = [np.hstack((np.array([low]).T, grid, np.array([high]).T)) for grid in tilings] # add low and high ends
tile_centers = [(grid_extended[:, 1:] + grid_extended[:, :-1]) / 2 for grid_extended in tilings_extended] # compute center of each tile
tile_toplefts = [grid_extended[:, :-1] for grid_extended in tilings_extended] # compute topleft of each tile
tile_bottomrights = [grid_extended[:, 1:] for grid_extended in tilings_extended] # compute bottomright of each tile
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
for sample, encoded_sample in zip(samples, encoded_samples):
for i, tile in enumerate(encoded_sample):
# Shade the entire tile with a rectangle
topleft = tile_toplefts[i][0][tile[0]], tile_toplefts[i][1][tile[1]]
bottomright = tile_bottomrights[i][0][tile[0]], tile_bottomrights[i][1][tile[1]]
ax.add_patch(Rectangle(topleft, bottomright[0] - topleft[0], bottomright[1] - topleft[1],
color=colors[i], alpha=0.33))
# In case sample is outside tile bounds, it may not have been highlighted properly
if any(sample < topleft) or any(sample > bottomright):
# So plot a point in the center of the tile and draw a connecting line
cx, cy = tile_centers[i][0][tile[0]], tile_centers[i][1][tile[1]]
ax.add_line(Line2D([sample[0], cx], [sample[1], cy], color=colors[i]))
ax.plot(cx, cy, 's', color=colors[i])
# Finally, plot original samples
ax.plot(samples[:, 0], samples[:, 1], 'o', color='r')
ax.margins(x=0, y=0) # remove unnecessary margins
ax.set_title("Tile-encoded samples")
return ax
visualize_encoded_samples(samples, encoded_samples, tilings);
```
Inspect the results and make sure you understand how the corresponding tiles are being chosen. Note that some samples may have one or more tiles in common.
### 5. Q-Table with Tile Coding
The next step is to design a special Q-table that is able to utilize this tile coding scheme. It should have the same kind of interface as a regular table, i.e. given a `<state, action>` pair, it should return a `<value>`. Similarly, it should also allow you to update the `<value>` for a given `<state, action>` pair (note that this should update all the tiles that `<state>` belongs to).
The `<state>` supplied here is assumed to be from the original continuous state space, and `<action>` is discrete (and integer index). The Q-table should internally convert the `<state>` to its tile-coded representation when required.
```
class QTable:
"""Simple Q-table."""
def __init__(self, state_size, action_size):
"""Initialize Q-table.
Parameters
----------
state_size : tuple
Number of discrete values along each dimension of state space.
action_size : int
Number of discrete actions in action space.
"""
self.state_size = state_size
self.action_size = action_size
# TODO: Create Q-table, initialize all Q-values to zero
# Note: If state_size = (9, 9), action_size = 2, q_table.shape should be (9, 9, 2)
print("QTable(): size =", self.q_table.shape)
class TiledQTable:
"""Composite Q-table with an internal tile coding scheme."""
def __init__(self, low, high, tiling_specs, action_size):
"""Create tilings and initialize internal Q-table(s).
Parameters
----------
low : array_like
Lower bounds for each dimension of state space.
high : array_like
Upper bounds for each dimension of state space.
tiling_specs : list of tuples
A sequence of (bins, offsets) to be passed to create_tilings() along with low, high.
action_size : int
Number of discrete actions in action space.
"""
self.tilings = create_tilings(low, high, tiling_specs)
self.state_sizes = [tuple(len(splits)+1 for splits in tiling_grid) for tiling_grid in self.tilings]
self.action_size = action_size
self.q_tables = [QTable(state_size, self.action_size) for state_size in self.state_sizes]
print("TiledQTable(): no. of internal tables = ", len(self.q_tables))
def get(self, state, action):
"""Get Q-value for given <state, action> pair.
Parameters
----------
state : array_like
Vector representing the state in the original continuous space.
action : int
Index of desired action.
Returns
-------
value : float
Q-value of given <state, action> pair, averaged from all internal Q-tables.
"""
# TODO: Encode state to get tile indices
# TODO: Retrieve q-value for each tiling, and return their average
pass
def update(self, state, action, value, alpha=0.1):
"""Soft-update Q-value for given <state, action> pair to value.
Instead of overwriting Q(state, action) with value, perform soft-update:
Q(state, action) = alpha * value + (1.0 - alpha) * Q(state, action)
Parameters
----------
state : array_like
Vector representing the state in the original continuous space.
action : int
Index of desired action.
value : float
Desired Q-value for <state, action> pair.
alpha : float
Update factor to perform soft-update, in [0.0, 1.0] range.
"""
# TODO: Encode state to get tile indices
# TODO: Update q-value for each tiling by update factor alpha
pass
# Test with a sample Q-table
tq = TiledQTable(low, high, tiling_specs, 2)
s1 = 3; s2 = 4; a = 0; q = 1.0
print("[GET] Q({}, {}) = {}".format(samples[s1], a, tq.get(samples[s1], a))) # check value at sample = s1, action = a
print("[UPDATE] Q({}, {}) = {}".format(samples[s2], a, q)); tq.update(samples[s2], a, q) # update value for sample with some common tile(s)
print("[GET] Q({}, {}) = {}".format(samples[s1], a, tq.get(samples[s1], a))) # check value again, should be slightly updated
```
If you update the q-value for a particular state (say, `(0.25, -1.91)`) and action (say, `0`), then you should notice the q-value of a nearby state (e.g. `(0.15, -1.75)` and same action) has changed as well! This is how tile-coding is able to generalize values across the state space better than a single uniform grid.
### 6. Implement a Q-Learning Agent using Tile-Coding
Now it's your turn to apply this discretization technique to design and test a complete learning agent!
| github_jupyter |
```
%matplotlib notebook
import control as c
import ipywidgets as w
import numpy as np
from IPython.display import display, HTML
import matplotlib.pyplot as plt
import matplotlib.animation as animation
#display(HTML('<script> $(document).ready(function() { $(\"div.input\").hide(); }); </script>'))
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Promijeni vidljivost <a href="javascript:code_toggle()">ovdje</a>.''')
display(tag)
```
## Stvaranje PI-regulatora korištenjem operacijskih pojačala
U analognoj elektronici, operacijska pojačala uobičajeno se koriste za realizaciju proporcionalno-integracijski-derivacijskih (PID) regulatora. Dok matematički modeli linearnih vremenski-nepromjenjivih (LTI) sustava pretpostavljaju idealne uvjete, realni sklopovi možda im ne odgovaraju u potpunosti.
U većini slučajeva idealni model daje prihvatljive rezultate, ali frekvencijske karakteristike mogu se bolje aproksimirati proširivanjem modela s pojačanjem otvorene petlje:
<br><br>
$$G_{ideal}(s)=\frac{V_{out}}{V_{in}}=-\frac{Z_F}{Z_G}\qquad\qquad G_{approx}(s)=\frac{V_{out}}{V_{in}}=-\frac{\frac{-A\cdot Z_F}{Z_G+Z_F}}{1+\frac{A\cdot Z_G}{Z_G+Z_F}}$$
<br>
U ovom ćemo primjeru istražiti neke od konfiguracija PI regulatora zasnovanih na operacijskim pojačalima.<br>
<b>Prvo, odaberite vrijednost pojačanja otvorene petlje za prikazane izračune!</b>
```
# Model selector
opampGain = w.ToggleButtons(
options=[('10 000', 10000), ('50 000', 50000), ('200 000', 200000),],
description='Pojačanje operacijskog pojačala: ', style={'description_width':'30%'})
display(opampGain)
```
PI regulator može se implementirati pomoću otpornika u unaprijednoj vezi i kondenzatora u povratnoj vezi. Idealni model točno odgovara matematičkom obliku regulatora. Ali, nakon uključivanja pojačanja otvorene petlje, integrator se zamjenjuje sustavom prvog reda s ogromnom vremenskom konstantom, ograničavajući na taj način amplitudu na niskim frekvencijama.
<br><br>
<img src="Images/int1.png" width="30%" />
<br>
<b>Prilagodite pasivne komponente tako da neidealni model bude najbliži idealnom! Gdje karakteristike značajno odstupaju od idealnih? Što se može reći o grafu faze?</b>
```
# Figure definition
fig1, ((f1_ax1), (f1_ax2)) = plt.subplots(2, 1)
fig1.set_size_inches((9.8, 5))
fig1.set_tight_layout(True)
l1 = f1_ax1.plot([], [], color='red')
l2 = f1_ax2.plot([], [], color='red')
l3 = f1_ax1.plot([], [], color='blue')
l4 = f1_ax2.plot([], [], color='blue')
f1_line1 = l1[0]
f1_line2 = l2[0]
f1_line3 = l3[0]
f1_line4 = l4[0]
f1_ax1.legend(l1+l3, ['Ne-idealno', 'Idealno'], loc=1)
f1_ax2.legend(l2+l4, ['Ne-idealno', 'Idealno'], loc=1)
f1_ax1.grid(which='both', axis='both', color='lightgray')
f1_ax2.grid(which='both', axis='both', color='lightgray')
f1_ax1.autoscale(enable=True, axis='x', tight=True)
f1_ax2.autoscale(enable=True, axis='x', tight=True)
f1_ax1.autoscale(enable=True, axis='y', tight=False)
f1_ax2.autoscale(enable=True, axis='y', tight=False)
f1_ax1.set_title('Bodeov graf amplitude', fontsize=11)
f1_ax1.set_xscale('log')
f1_ax1.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f1_ax1.set_ylabel(r'$A\/$[dB]', labelpad=0, fontsize=10)
f1_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
f1_ax2.set_title('Bodeov graf faze', fontsize=11)
f1_ax2.set_xscale('log')
f1_ax2.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f1_ax2.set_ylabel(r'$\phi\/$[°]', labelpad=0, fontsize=10)
f1_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
# System model
def system_model(rg, cf, a):
Rg = rg / 1000 # Convert to Ohm
Cf = cf * 1000000 # Convert to Farad
W_ideal = c.tf([-1], [Rg*Cf, 0])
W_ac = c.tf([-a], [Cf*Rg*(1+a), 1])
global f1_line1, f1_line2, f1_line3, f1_line4
f1_ax1.lines.remove(f1_line1)
f1_ax2.lines.remove(f1_line2)
f1_ax1.lines.remove(f1_line3)
f1_ax2.lines.remove(f1_line4)
mag, phase, omega = c.bode_plot(W_ac, Plot=False) # Non-ideal Bode-plot
f1_line1, = f1_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='red')
f1_line2, = f1_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='red')
mag, phase, omega = c.bode_plot(W_ideal, omega=omega, Plot=False) # Ideal Bode-plot at the non-ideal points
f1_line3, = f1_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f1_line4, = f1_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
f1_ax1.relim()
f1_ax2.relim()
f1_ax1.autoscale_view()
f1_ax2.autoscale_view()
print('Prijenosna funkcija za idealni PI:')
print(W_ideal)
print('\nPrijenosna funkcija za ne-idealni PI:')
print(W_ac)
# GUI widgets
rg_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_g\ [k\Omega]\ :$', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
cf_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$C_f\ [\mu H]\ :$', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
input_data = w.interactive_output(system_model, {'rg':rg_slider, 'cf':cf_slider, 'a':opampGain})
display(w.HBox([rg_slider, cf_slider]), input_data)
```
Ovaj PI regulator može biti jednostavan, ali nije moguće kontrolirati DC pojačanje putem pasivnih komponenata. Zbog toga se obično u povratnu vezu paralelno spaja odgovarajući otpornik.
<br><br>
<img src="Images/int2.png" width="30%" />
<br>
<b>Prilagodite pasivne komponente tako da neidealni model bude najbliži idealnom! Koje su razlike s obzirom na prethodni model?</b>
```
# Filtered PI - parallel
fig2, ((f2_ax1), (f2_ax2)) = plt.subplots(2, 1)
fig2.set_size_inches((9.8, 5))
fig2.set_tight_layout(True)
l1 = f2_ax1.plot([], [], color='red')
l2 = f2_ax2.plot([], [], color='red')
l3 = f2_ax1.plot([], [], color='blue')
l4 = f2_ax2.plot([], [], color='blue')
f2_line1 = l1[0]
f2_line2 = l2[0]
f2_line3 = l3[0]
f2_line4 = l4[0]
f2_ax1.legend(l1+l3, ['Ne-idealno', 'Idealno'], loc=1)
f2_ax2.legend(l2+l4, ['Ne-idealno', 'Idealno'], loc=1)
f2_ax1.grid(which='both', axis='both', color='lightgray')
f2_ax2.grid(which='both', axis='both', color='lightgray')
f2_ax1.autoscale(enable=True, axis='x', tight=True)
f2_ax2.autoscale(enable=True, axis='x', tight=True)
f2_ax1.autoscale(enable=True, axis='y', tight=False)
f2_ax2.autoscale(enable=True, axis='y', tight=False)
f2_ax1.set_title('Bodeov graf amplitude', fontsize=11)
f2_ax1.set_xscale('log')
f2_ax1.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f2_ax1.set_ylabel(r'$A\/$[dB]', labelpad=0, fontsize=10)
f2_ax1.tick_params(axis='both', which='both', pad=0, labelsize=8)
f2_ax2.set_title('Bodeov graf faze', fontsize=11)
f2_ax2.set_xscale('log')
f2_ax2.set_xlabel(r'$f\/$[Hz]', labelpad=0, fontsize=10)
f2_ax2.set_ylabel(r'$\phi\/$[°]', labelpad=0, fontsize=10)
f2_ax2.tick_params(axis='both', which='both', pad=0, labelsize=8)
# System model
def system2_model(rg, rf, cf, a):
Rg = rg / 1000 # Convert to Ohm
Rf = rf / 1000
Cf = cf * 1000000 # Convert to Farad
W_ideal = c.tf([-1], [Rg*Cf, Rg/Rf])
W_ac = c.tf([-a], [Cf*Rg*(a+1), Rg*(a+1)/Rf+1])
global f2_line1, f2_line2, f2_line3, f2_line4
f2_ax1.lines.remove(f2_line1)
f2_ax2.lines.remove(f2_line2)
f2_ax1.lines.remove(f2_line3)
f2_ax2.lines.remove(f2_line4)
mag, phase, omega = c.bode_plot(W_ac, Plot=False) # Non-ideal Bode-plot
f2_line1, = f2_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='red')
f2_line2, = f2_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='red')
mag, phase, omega = c.bode_plot(W_ideal, omega=omega, Plot=False) # Ideal Bode-plot at the non-ideal points
f2_line3, = f2_ax1.plot(omega/2/np.pi, 20*np.log10(mag), lw=1, color='blue')
f2_line4, = f2_ax2.plot(omega/2/np.pi, phase*180/np.pi, lw=1, color='blue')
f2_ax1.relim()
f2_ax2.relim()
f2_ax1.autoscale_view()
f2_ax2.autoscale_view()
print('Prijenosna funckija za idealni filtrirani PI:')
print(W_ideal)
print('\nPrijenosna funckija za ne-idealni filtrirani PI::')
print(W_ac)
# GUI widgets
rg2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_g$ [k$\Omega$]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
rf2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$R_f$ [k$\Omega$]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
cf2_slider = w.FloatLogSlider(value=1, base=10, min=-3, max=3, description=r'$C_f$ [$\mu$H]', continuous_update=False,
layout=w.Layout(width='75%'), style={'description_width':'30%'})
input_data = w.interactive_output(system2_model, {'rg':rg2_slider, 'rf':rf2_slider, 'cf':cf2_slider, 'a':opampGain})
display(w.HBox([rg2_slider, rf2_slider, cf2_slider]), input_data)
```
| github_jupyter |
# Using KERMIT - Building dataset -
This first notebook explains how to construct the syntactic input for KERMIT_system via KERMIT encoder.
There are also links from where to download used datasets or you can use a dataset of your choice.
## Install Packages
Before starting, it is essential to have the following requirements:
- stanford-corenlp-full-2018-10-05 : which will be used to build the trees in parenthetical form.
- KERMIT : that it is obvious to have it but we specify it anyway.
```
#Install stanfordcorenlp
#!pip install stanfordcorenlp
#!wget http://nlp.stanford.edu/software/stanford-corenlp-full-2018-10-05.zip
#!unzip stanford-corenlp-full-2018-10-05.zip
#Install KERMIT
#!git clone https://github.com/ART-Group-it/kerMIT
```
# Import Statments
```
import time, pickle, ast, os
from tqdm import tqdm
import pandas as pd
import numpy as np
#script for reading/writing trees
from scripts.script import readP, writeTree
#script for build DTK
from scripts.script import createDTK
#script for parse sentences
from scripts.script import parse
```
## Download Dataset
The datasets used in our work have been randomly sampled as follows:
* *Note: in this tutorial we use ag_news (train and test set) but remember that the user can choose others as he prefers.*
```
#download dataset ag_news
#!wget wget https://data.deepai.org/agnews.zip
#!unzip agnews.zip
#! wget "https://multi-classification.s3.eu-central-1.amazonaws.com/dbpedia_csv.tar.gz"
#! wget "https://multi-classification.s3.eu-central-1.amazonaws.com/yelp_review_polarity_csv.tar.gz"
#! wget "https://multi-classification.s3.eu-central-1.amazonaws.com/yelp_review_full_csv.tar.gz"
data_train = pd.read_csv('train.csv')
data_test = pd.read_csv('test.csv')
#if you want to sample the dataset in this way you exchange the examples and then take the first n lines
#data = data_original.iloc[np.random.permutation(len(data_original))]
#data = data[:70000]
data_train.head()
```
## Building parenthetical trees and encode in Universal Syntactic Embeddings
Here the loaded dataset is processed, transformed into tree form and encoded via kerMIT encoder.
In realtime the trees are saved on file, a log file is made showing the number of processed rows of the dataset and the encoded trees are saved in pickle format.
## Building parenthetical trees for Training Set
```
#insert here your dataset name
dataset_name = 'ag_news_train'
name = 'dtk_trees_'+dataset_name+'.pkl'
name2 = 'log_'+dataset_name+'.txt'
name3 = 'dt_'+dataset_name+'.txt'
i = 0
cont = 0
listTree = []
newList = []
oldList = []
tree = "(S)"
treeException = createDTK(tree)
for line in tqdm(data_train['Description']):
cont += 1
try:
tree = (parse(line))
treeDTK = createDTK(tree)
except Exception:
tree, treeDTK = "(S)", treeException
listTree.append(treeDTK)
#write parenthetical tree
writeTree(name3,tree)
#every 5000 shafts saves the corresponding DTKs
if i>5000:
time.sleep(1)
if os.path.isfile(name):
#append new encoded tree in pickle file
oldList = readP(name)
newList = oldList+listTree
else:
newList = listTree
f=open(name, 'wb')
for x in newList:
pickle.dump(x, f)
f.close()
f=open(name2, "a+")
f.write(str(cont)+'..')
f.close()
i = 0
listTree = []
newList = []
oldList = []
else:
i +=1
#checking consistency
if os.path.isfile(name):
oldList = readP(name)
newList = oldList+listTree
else:
newList = listTree
f=open(name, 'wb')
for x in newList:
pickle.dump(x, f)
f.close()
```
## Building parenthetical trees for Test Set
```
#insert here your dataset name
dataset_name = 'ag_news_test'
name = 'dtk_trees_'+dataset_name+'.pkl'
name2 = 'log_'+dataset_name+'.txt'
name3 = 'dt_'+dataset_name+'.txt'
i = 0
cont = 0
listTree = []
newList = []
oldList = []
tree = "(S)"
treeException = createDTK(tree)
for line in tqdm(data_test['Description']):
cont += 1
try:
tree = (parse(line))
treeDTK = createDTK(tree)
except Exception:
tree, treeDTK = "(S)", treeException
listTree.append(treeDTK)
#write parenthetical tree
writeTree(name3,tree)
#every 5000 shafts saves the corresponding DTKs
if i>5000:
time.sleep(1)
if os.path.isfile(name):
#append new encoded tree in pickle file
oldList = readP(name)
newList = oldList+listTree
else:
newList = listTree
f=open(name, 'wb')
for x in newList:
pickle.dump(x, f)
f.close()
f=open(name2, "a+")
f.write(str(cont)+'..')
f.close()
i = 0
listTree = []
newList = []
oldList = []
else:
i +=1
#checking consistency
if os.path.isfile(name):
oldList = readP(name)
newList = oldList+listTree
else:
newList = listTree
f=open(name, 'wb')
for x in newList:
pickle.dump(x, f)
f.close()
```
| github_jupyter |
# PRMT-2131 - TPP-EMIS transfers with error code 30s all contain attachments over a certain size
## Hypothesis
We believe that TPP-->EMIS transfers that failed due to error code 30 all contain at least one attachment over a certain size
We will know this to be true when we see that these transfers all contain an attachment over a certain size, where successful TPP-->EMIS transfers do not contain attachments of this size
## Scope
Take a months worth of TPP-->EMIS transfers
User attachments dataset to identify, for transfers failed with error code 30,
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Import transfer files to extract whether message creator is sender or requester
transfer_file_location = "s3://prm-gp2gp-data-sandbox-dev/transfers-sample-5/"
transfer_files = [
"2021-1-transfers.parquet",
"2021-2-transfers.parquet",
"2021-3-transfers.parquet"
]
transfer_input_files = [transfer_file_location + f for f in transfer_files]
transfers_raw = pd.concat((
pd.read_parquet(f)
for f in transfer_input_files
))
# In the data from the PRMT-1742-duplicates-analysis branch, these columns have been added , but contain only empty values.
transfers_raw = transfers_raw.drop(["sending_supplier", "requesting_supplier"], axis=1)
transfers = transfers_raw.copy()
# Correctly interpret certain sender errors as failed.
# This is explained in PRMT-1974. Eventually this will be fixed upstream in the pipeline.
# Step Two: reclassifying the relevant transfers with pending sender error codes to FAILED DUE TO SENDER ERROR CODE status for comparison
pending_sender_error_codes=[6,7,10,24,30,23,14,99]
transfers_with_pending_sender_code_bool=transfers['sender_error_code'].isin(pending_sender_error_codes)
transfers_with_pending_with_error_bool=transfers['status']=='PENDING_WITH_ERROR'
transfers_which_need_pending_to_failure_change_bool=transfers_with_pending_sender_code_bool & transfers_with_pending_with_error_bool
transfers.loc[transfers_which_need_pending_to_failure_change_bool,'status']='FAILED DUE TO SENDER ERROR CODE'
# Add integrated Late status
eight_days_in_seconds=8*24*60*60
transfers_after_sla_bool=transfers['sla_duration']>eight_days_in_seconds
transfers_with_integrated_bool=transfers['status']=='INTEGRATED'
transfers_integrated_late_bool=transfers_after_sla_bool & transfers_with_integrated_bool
transfers.loc[transfers_integrated_late_bool,'status']='INTEGRATED LATE'
# If the record integrated after 28 days, change the status back to pending.
# This is to handle each month consistently and to always reflect a transfers status 28 days after it was made.
# TBD how this is handled upstream in the pipeline
twenty_eight_days_in_seconds=28*24*60*60
transfers_after_month_bool=transfers['sla_duration']>twenty_eight_days_in_seconds
transfers_pending_at_month_bool=transfers_after_month_bool & transfers_integrated_late_bool
transfers.loc[transfers_pending_at_month_bool,'status']='PENDING'
transfers_with_early_error_bool=(~transfers.loc[:,'sender_error_code'].isna()) |(~transfers.loc[:,'intermediate_error_codes'].apply(len)>0)
transfers.loc[transfers_with_early_error_bool & transfers_pending_at_month_bool,'status']='PENDING_WITH_ERROR'
# Supplier name mapping
supplier_renaming = {
"EGTON MEDICAL INFORMATION SYSTEMS LTD (EMIS)":"EMIS",
"IN PRACTICE SYSTEMS LTD":"Vision",
"MICROTEST LTD":"Microtest",
"THE PHOENIX PARTNERSHIP":"TPP",
None: "Unknown"
}
# Generate ASID lookup that contains all the most recent entry for all ASIDs encountered
asid_file_location = "s3://prm-gp2gp-data-sandbox-dev/asid-lookup/"
asid_files = [
"asidLookup-Nov-2020.csv.gz",
"asidLookup-Dec-2020.csv.gz",
"asidLookup-Jan-2021.csv.gz",
"asidLookup-Feb-2021.csv.gz",
"asidLookup-Mar-2021.csv.gz",
"asidLookup-Apr-2021.csv.gz"
]
asid_lookup_files = [asid_file_location + f for f in asid_files]
asid_lookup = pd.concat((
pd.read_csv(f)
for f in asid_lookup_files
))
asid_lookup = asid_lookup.drop_duplicates().groupby("ASID").last().reset_index()
lookup = asid_lookup[["ASID", "MName", "NACS","OrgName"]]
transfers = transfers.merge(lookup, left_on='requesting_practice_asid',right_on='ASID',how='left')
transfers = transfers.rename({'MName': 'requesting_supplier', 'ASID': 'requesting_supplier_asid', 'NACS': 'requesting_ods_code','OrgName':'requesting_practice_name'}, axis=1)
transfers = transfers.merge(lookup, left_on='sending_practice_asid',right_on='ASID',how='left')
transfers = transfers.rename({'MName': 'sending_supplier', 'ASID': 'sending_supplier_asid', 'NACS': 'sending_ods_code','OrgName':'sending_practice_name'}, axis=1)
transfers["sending_supplier"] = transfers["sending_supplier"].replace(supplier_renaming.keys(), supplier_renaming.values())
transfers["requesting_supplier"] = transfers["requesting_supplier"].replace(supplier_renaming.keys(), supplier_renaming.values())
# Making the status to be more human readable here
transfers["status"] = transfers["status"].str.replace("_", " ").str.title()
# selecting TPP -> EMIS
is_tpp_to_emis = (transfers["requesting_supplier"]=="EMIS") & (transfers["sending_supplier"]=="TPP")
tpp_to_emis_transfers = transfers.loc[is_tpp_to_emis]
# import attachment data for tpp -> emis
tpp_to_emis_attachments = pd.read_csv("s3://prm-gp2gp-data-sandbox-dev/PRMT-2131-tpp-to-emis-attachments/tpp-to-emis-jan-march-2021-attachment-data.csv")
```
Athena query to get the above data:
```sql
select * from gp2gp_attachment_metadata
where from_iso8601_timestamp(time) >= from_iso8601_timestamp('2021-01-01T00:00:00')
and from_iso8601_timestamp(time) < from_iso8601_timestamp('2021-04-01T00:00:00')
and from_system='SystmOne'
and to_system='EMIS Web'
limit 10
```
```
# convert length into int (was being read as string)
tpp_to_emis_attachments['length'] = pd.to_numeric(tpp_to_emis_attachments['length'], errors='coerce')
tpp_to_emis_attachments.head()
# Add column with max attachment size and total size of attachments per conversation
attachments_by_conversation = tpp_to_emis_attachments.groupby(by="conversation_id").agg({'length': ['max', 'sum']})['length']
attachments_by_conversation = attachments_by_conversation.rename({'max': 'Largest Attachment Size', 'sum': 'Total Size of Attachments'}, axis=1)
# combine datasets and convert length to megabytes
tpp_to_emis_transfers_and_attachments = tpp_to_emis_transfers.merge(attachments_by_conversation, left_on='conversation_id', right_index=True, how='left')
tpp_to_emis_transfers_and_attachments[['Largest Attachment Size', 'Total Size of Attachments']] = (tpp_to_emis_transfers_and_attachments[['Largest Attachment Size', 'Total Size of Attachments']].fillna(0))/(1024**2)
tpp_to_emis_transfers_and_attachments.head()
# checking which columns actually contain error code 30
tpp_to_emis_transfers_and_attachments['sender_error_code'].value_counts()
tpp_to_emis_transfers_and_attachments['final_error_codes'].apply(lambda x: 30 in x).value_counts()
tpp_to_emis_transfers_and_attachments['intermediate_error_codes'].apply(lambda x: 30 in x).value_counts()
# anything containing error 30
# Add contains error code 30 column
error_code_30 = (tpp_to_emis_transfers_and_attachments['sender_error_code']== 30).fillna(False)
tpp_to_emis_transfers_and_attachments.loc[error_code_30, 'Histogram categories'] = 'Contains error code 30'
successfully_integrated = (tpp_to_emis_transfers_and_attachments['status'].isin(['Integrated', 'Integrated Late'])).fillna(False)
tpp_to_emis_transfers_and_attachments.loc[successfully_integrated, 'Histogram categories'] = 'Successfully Integrated'
# anything successfully integrated (including integrated late and does not have err 30)
tpp_to_emis_transfers_and_attachments.groupby(by='Histogram categories').agg({'Largest Attachment Size': 'describe'})
tpp_to_emis_transfers_and_attachments.groupby(by='Histogram categories').agg({'Total Size of Attachments': 'describe'})
has_error_30_cat = tpp_to_emis_transfers_and_attachments['Histogram categories']== 'Contains error code 30'
tpp_to_emis_transfers_and_attachments[has_error_30_cat].sample(5)
# Generate histogram
successfully_integrated_bool = tpp_to_emis_transfers_and_attachments['Histogram categories']=='Successfully Integrated'
tpp_to_emis_attachment_size_histogram = tpp_to_emis_transfers_and_attachments.loc[successfully_integrated_bool, 'Largest Attachment Size'].plot.hist(xlim=[0, 65], bins=10)
tpp_to_emis_attachment_size_histogram.set_xlabel('File size (MB)')
tpp_to_emis_attachment_size_histogram.set_ylabel('Number of transfers')
tpp_to_emis_attachment_size_histogram.set_title('Distribution of TPP to EMIS file sizes')
tpp_to_emis_attachment_size_histogram
# plot data
successfully_integrated_bool = tpp_to_emis_transfers_and_attachments['Histogram categories']=='Successfully Integrated'
tpp_to_emis_transfers_and_attachments[successfully_integrated_bool].hist(column='Total Size of Attachments', grid=False)
#Extract some sample conversation IDs to send to TPP
april_transfers = pd.read_parquet("s3://prm-gp2gp-data-sandbox-dev/transfers-sample-5/2021-4-transfers.parquet")
tpp_to_emis_error30_failed_bool = (april_transfers['sender_error_code']==30) & (april_transfers['status'] == 'PENDING_WITH_ERROR') & (april_transfers['sending_supplier'] == 'SystmOne') & (april_transfers['requesting_supplier'] == 'EMIS')
tpp_to_emis_error30_failed = april_transfers.loc[tpp_to_emis_error30_failed_bool, 'conversation_id']
tpp_to_emis_error30_failed.sample(20).values
april_transfers['status'].value_counts().index
```
## Check EMIS to EMIS transfer metadata for comparison.
This is to see if the file size distribution for successful transfers is similar to EMIS -> EMIS
```
# import attachment data for emis -> emis
emis_to_emis_attachments = pd.read_csv("s3://prm-gp2gp-data-sandbox-dev/PRMT-2131-tpp-to-emis-attachments/emis-to-emis-jan-march-2021-attachment-data.csv")
# convert length into int (was being read as string)
emis_to_emis_attachments['length'] = pd.to_numeric(emis_to_emis_attachments['length'], errors='coerce')
emis_to_emis_attachments.head()
# Add column with max attachment size and total size of attachments per conversation
emis_attachments_by_conversation = emis_to_emis_attachments.groupby(by="conversation_id").agg({'length': ['max', 'sum']})['length']
emis_attachments_by_conversation = emis_attachments_by_conversation.rename({'max': 'Largest Attachment Size', 'sum': 'Total Size of Attachments'}, axis=1)
# selecting EMIS -> EMIS
is_emis_to_emis = (transfers["requesting_supplier"]=="EMIS") & (transfers["sending_supplier"]=="EMIS")
emis_to_emis_transfers = transfers.loc[is_emis_to_emis]
# combine datasets and convert length to megabytes
emis_to_emis_transfers_and_attachments = emis_to_emis_transfers.merge(emis_attachments_by_conversation, left_on='conversation_id', right_index=True, how='left')
emis_to_emis_transfers_and_attachments[['Largest Attachment Size', 'Total Size of Attachments']] = (emis_to_emis_transfers_and_attachments[['Largest Attachment Size', 'Total Size of Attachments']].fillna(0))/(1024**2)
emis_to_emis_transfers_and_attachments.head()
# checking which columns actually contain error code 30
emis_to_emis_transfers_and_attachments['sender_error_code'].value_counts()
emis_to_emis_transfers_and_attachments['final_error_codes'].apply(lambda x: 30 in x).value_counts()
emis_to_emis_transfers_and_attachments['intermediate_error_codes'].apply(lambda x: 30 in x).value_counts()
# mark all that are successfully integrated
emis_successfully_integrated = (emis_to_emis_transfers_and_attachments['status'].isin(['Integrated', 'Integrated Late'])).fillna(False)
emis_to_emis_transfers_and_attachments.loc[emis_successfully_integrated, 'Histogram categories'] = 'Successfully Integrated'
emis_to_emis_transfers_and_attachments.groupby(by='Histogram categories').agg({'Largest Attachment Size': 'describe'})
# plot data
emis_successfully_integrated_bool = emis_to_emis_transfers_and_attachments['Histogram categories']=='Successfully Integrated'
emis_to_emis_transfers_and_attachments[emis_successfully_integrated_bool].hist(column='Largest Attachment Size', grid=False, bins=60, figsize=(10,5))
# Generate histogram
emis_successfully_integrated_bool = emis_to_emis_transfers_and_attachments['Histogram categories']=='Successfully Integrated'
emis_to_emis_attachment_size_histogram = emis_to_emis_transfers_and_attachments.loc[emis_successfully_integrated_bool, 'Largest Attachment Size'].plot.hist(xlim=[0, 65], bins=100)
emis_to_emis_attachment_size_histogram.set_xlabel('File size (MB)')
emis_to_emis_attachment_size_histogram.set_ylabel('Number of transfers')
emis_to_emis_attachment_size_histogram.set_title('Distribution of EMIS to EMIS file sizes')
emis_to_emis_attachment_size_histogram
```
| github_jupyter |
# Example of optimizing a convex function
# Goal is to test the objective values found by Random
# Benchmarking test with different iterations for serial executions
```
from scipy.stats import uniform
from mango.domain.distribution import loguniform
from mango.tuner import Tuner
def get_param_dict():
param_dict = {"gamma": uniform(0.1, 4),
"C": loguniform(-7, 7)}
return param_dict
from sklearn import svm, datasets
from sklearn.model_selection import cross_val_score
iris = datasets.load_iris()
# Take only the first two features.
X = iris.data[:,:2]
Y = iris.target
def objectiveSVM(args_list):
global X,Y
results = []
for hyper_par in args_list:
clf = svm.SVC(**hyper_par)
result = cross_val_score(clf, X, Y, scoring='accuracy').mean()
results.append(result)
return results
def get_conf():
conf = dict()
conf['batch_size'] = 1
conf['num_iteration'] = 100
conf['optimizer'] = "Random"
return conf
def get_optimal_x():
param_dict = get_param_dict()
conf = get_conf()
tuner = Tuner(param_dict, objectiveSVM,conf)
results = tuner.maximize()
return results
Store_Optimal_X = []
Store_Results = []
num_of_tries = 20
for i in range(num_of_tries):
results = get_optimal_x()
Store_Results.append(results)
#print(results)
print(i,":",results['best_objective'])
```
# Extract from the results returned the true optimal values for each iteration
```
import numpy as np
total_experiments = 20
initial_random = 5
plotting_itr =[10, 20,30,40,50,60,70,80,90,100]
plotting_list = []
for exp in range(total_experiments): #for all exp
local_list = []
for itr in plotting_itr: # for all points to plot
# find the value of optimal parameters in itr+ initial_random
max_value = np.array(Store_Results[exp]['objective_values'][:itr+initial_random]).max()
local_list.append(max_value)
plotting_list.append(local_list)
plotting_array = np.array(plotting_list)
Y = []
range_min = -100
range_max = 100
#count range between -1 and 1 and show it
for i in range(len(plotting_itr)):
y_value = plotting_array[:,i].mean()
Y.append(y_value)
# Printing Y Values
Y
plotting_itr
import numpy as np
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,10))
plt.plot(plotting_itr,Y,label = 'Random',linewidth=3.0) #x, y
plt.xlabel('Number of Iterations',fontsize=25)
plt.ylabel('Mean best accuracy achieved',fontsize=25)
plt.title('Variation of SVM Accuracy',fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.grid(True)
#plt.yticks(np.arange(0.915, 0.925, step=0.005))
plt.legend(fontsize=20)
plt.show()
```
| github_jupyter |
# Theoretical Uncertainties in the SPS model
In this notebook I will quantify the theoretical uncertainties in SPS models by comparing the SEDs constructed using different spectral and isochrone libraries
```
import os,sys
import h5py
import numpy as np
# -- provabgs --
from provabgs import infer as Infer
from provabgs import models as Models
# --- plotting ---
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
```
## read in ($\theta$, SED $f_\lambda$) constructed using the fiducial setup
The fiducial setup uses a **MILES spectral and MIST isochrones libraries**. This sample has SPS parameters that generate SEDs that fall within the rough color cuts of BGS
```
dat_dir = '/Users/chahah/data/arcoiris/provabgs_cnf/'
ftheta = os.path.join(dat_dir, 'train.set0.thetas.npy')
thetas = np.load(ftheta)
flssps = os.path.join(dat_dir, 'train.set0.miles_mist.lssp.lssps.npy')
lssps_fid = np.load(flssps)
fwaves = os.path.join(dat_dir, 'train.set0.miles_mist.lssp.waves.npy')
waves_fid = np.load(fwaves)
waves = waves_fid[0]
print(thetas.shape)
```
# compare fiducial SEDs to SEDs using different isochrone libraries
```
lssps_m2 = np.load(os.path.join(dat_dir, 'train.set0.miles_pdva.lssp.lssps.npy'))
lssps_m3 = np.load(os.path.join(dat_dir, 'train.set0.miles_prsc.lssp.lssps.npy'))
lssps_m4 = np.load(os.path.join(dat_dir, 'train.set0.miles_bsti.lssp.lssps.npy'))
fig = plt.figure(figsize=(10,10))
sub = fig.add_subplot(211)
sub.plot(waves, lssps_fid[0], c='k', lw=2, label='MIST (fiducial)')
sub.plot(waves, lssps_m2[0], c='C0', lw=1, label='PADOVA')
sub.plot(waves, lssps_m3[0], c='C1', lw=1, label='PARSEC')
sub.plot(waves, lssps_m4[0], c='C2', lw=1, label='BASTI')
sub.legend(loc='lower right', fontsize=20)
#sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1.5e3, 1e4)
sub.set_xticklabels([])
sub.set_ylabel(r'${\rm SED}_{\rm isochrone}$ [$L_\odot/A$]', fontsize=25)
sub = fig.add_subplot(212)
sub.plot(waves, lssps_m2[0]/lssps_fid[0], c='C0', lw=1, label='MILES, Padova')
sub.plot(waves, lssps_m3[0]/lssps_fid[0], c='C1', lw=1, label='MILES, PARSEC')
sub.plot(waves, lssps_m4[0]/lssps_fid[0], c='C2', lw=1, label='MILES, BaSTI')
sub.plot(waves, np.ones(len(waves)), c='k', ls='--')
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1.5e3, 1e4)
sub.set_ylabel(r'${\rm SED} / {\rm SED}_{\rm MIST}$', fontsize=25)
sub.set_ylim(0.75, 1.25)
fig.savefig('../doc/paper/figs/sed_isochrone.pdf', bbox_inches='tight')
```
There are clearly some significant wavelength-dependent differents for the different isochrone libraries...
## SED ratios
```
lssp_ratios_m2 = lssps_m2/lssps_fid
lssp_ratios_m3 = lssps_m3/lssps_fid
lssp_ratios_m4 = lssps_m4/lssps_fid
lssp_ratios_m2_q = np.quantile(lssp_ratios_m2, [0.025, 0.16, 0.5, 0.84, 0.975], axis=0)
lssp_ratios_m3_q = np.quantile(lssp_ratios_m3, [0.025, 0.16, 0.5, 0.84, 0.975], axis=0)
lssp_ratios_m4_q = np.quantile(lssp_ratios_m4, [0.025, 0.16, 0.5, 0.84, 0.975], axis=0)
fig = plt.figure(figsize=(10,5))
sub = fig.add_subplot(111)
sub.fill_between(waves, lssp_ratios_m2_q[0], lssp_ratios_m2_q[-1], facecolor='C0', edgecolor='none', alpha=0.25)
sub.fill_between(waves, lssp_ratios_m2_q[1], lssp_ratios_m2_q[-2], facecolor='C0', edgecolor='none', alpha=0.5)
sub.plot(waves, lssp_ratios_m2_q[2], c='C0')
sub.plot([1e3, 1e4], [1., 1.], c='k', ls='--')
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1e3, 1e4)
sub.set_ylabel(r'$f_\lambda^{\rm MILES, PADOVA}/f_\lambda^{\rm MILES, MIST}$', fontsize=25)
sub.set_ylim(0., 2.)
fig = plt.figure(figsize=(10,5))
sub = fig.add_subplot(111)
sub.fill_between(waves, lssp_ratios_m3_q[0], lssp_ratios_m3_q[-1], facecolor='C0', edgecolor='none', alpha=0.25)
sub.fill_between(waves, lssp_ratios_m3_q[1], lssp_ratios_m3_q[-2], facecolor='C0', edgecolor='none', alpha=0.5)
sub.plot(waves, lssp_ratios_m3_q[2], c='C0')
sub.plot([1e3, 1e4], [1., 1.], c='k', ls='--')
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1e3, 1e4)
sub.set_ylabel(r'$f_\lambda^{\rm MILES, PARSEC}/f_\lambda^{\rm MILES, MIST}$', fontsize=25)
sub.set_ylim(0., 2.)
fig = plt.figure(figsize=(10,5))
sub = fig.add_subplot(111)
sub.fill_between(waves, lssp_ratios_m3_q[0], lssp_ratios_m3_q[-1], facecolor='C0', edgecolor='none', alpha=0.25)
sub.fill_between(waves, lssp_ratios_m3_q[1], lssp_ratios_m3_q[-2], facecolor='C0', edgecolor='none', alpha=0.5)
sub.plot(waves, lssp_ratios_m3_q[2], c='C0')
sub.plot([1e3, 1e4], [1., 1.], c='k', ls='--')
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1e3, 1e4)
sub.set_ylabel(r'$f_\lambda^{\rm MILES, BASTI}/f_\lambda^{\rm MILES, MIST}$', fontsize=25)
sub.set_ylim(0., 2.)
```
The differences are quite dramatic for UV wavelengths
# compare fiducial SEDs to SEDs using different spectral libraries
```
waves_mm2 = np.load(os.path.join(dat_dir, 'train.set0.basel_mist.lssp.waves.npy'))
lssps_mm2 = np.load(os.path.join(dat_dir, 'train.set0.basel_mist.lssp.lssps.npy'))
waves2 = waves_mm2[0]
fig = plt.figure(figsize=(10,5))
sub = fig.add_subplot(111)
for i in range(5):
sub.plot(waves, lssps_fid[i], c='C%i' % i)
sub.plot(waves2, lssps_mm2[i], c='k', ls='--', lw=1)
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1e3, 1e4)
sub.set_ylabel('SED', fontsize=25)
lssp_ratios_mm2 = np.array([lssps_mm2[i]/np.interp(waves2, waves, lssps_fid[i]) for i in range(lssps_fid.shape[0])])
lssp_ratios_mm2_q = np.quantile(lssp_ratios_mm2, [0.025, 0.16, 0.5, 0.84, 0.975], axis=0)
fig = plt.figure(figsize=(10,5))
sub = fig.add_subplot(111)
sub.fill_between(waves2, lssp_ratios_mm2_q[0], lssp_ratios_mm2_q[-1], facecolor='C0', edgecolor='none', alpha=0.25)
sub.fill_between(waves2, lssp_ratios_mm2_q[1], lssp_ratios_mm2_q[-2], facecolor='C0', edgecolor='none', alpha=0.5)
sub.plot(waves2, lssp_ratios_mm2_q[2], c='C0')
sub.plot([1e3, 1e4], [1., 1.], c='k', ls='--')
sub.set_xlabel('wavelength', fontsize=25)
sub.set_xlim(1e3, 1e4)
sub.set_ylabel(r'$f_\lambda^{\rm BASEL, MIST}/f_\lambda^{\rm MILES, MIST}$', fontsize=25)
sub.set_ylim(0., 2.)
```
There isn't a significant bias from using a different spectral library but there seems to be a significant scatter.
| github_jupyter |
```
from transformers import BertTokenizer
from pathlib import Path
import torch
from box import Box
import pandas as pd
import collections
import os
from tqdm import tqdm, trange
import sys
import random
import numpy as np
import apex
from sklearn.model_selection import train_test_split
import datetime
from fast_bert.modeling import BertForMultiLabelSequenceClassification
from fast_bert.data_cls import BertDataBunch, InputExample, InputFeatures, MultiLabelTextProcessor, convert_examples_to_features
from fast_bert.learner_cls import BertLearner
from fast_bert.metrics import accuracy_multilabel, accuracy_thresh, fbeta, roc_auc
torch.cuda.empty_cache()
pd.set_option('display.max_colwidth', -1)
run_start_time = datetime.datetime.today().strftime('%Y-%m-%d_%H-%M-%S')
DATA_PATH = Path('../data/')
LABEL_PATH = Path('../labels/')
AUG_DATA_PATH = Path('../data/data_augmentation/')
MODEL_PATH=Path('../models/')
LOG_PATH=Path('../logs/')
MODEL_PATH.mkdir(exist_ok=True)
model_state_dict = None
# BERT_PRETRAINED_PATH = Path('../../bert_models/pretrained-weights/cased_L-12_H-768_A-12/')
BERT_PRETRAINED_PATH = Path('../../bert_models/pretrained-weights/uncased_L-12_H-768_A-12/')
# BERT_PRETRAINED_PATH = Path('../../bert_fastai/pretrained-weights/uncased_L-24_H-1024_A-16/')
# FINETUNED_PATH = Path('../models/finetuned_model.bin')
FINETUNED_PATH = None
# model_state_dict = torch.load(FINETUNED_PATH)
LOG_PATH.mkdir(exist_ok=True)
OUTPUT_PATH = MODEL_PATH/'output'
OUTPUT_PATH.mkdir(exist_ok=True)
args = Box({
"run_text": "multilabel toxic comments with freezable layers",
"train_size": -1,
"val_size": -1,
"log_path": LOG_PATH,
"full_data_dir": DATA_PATH,
"data_dir": DATA_PATH,
"task_name": "toxic_classification_lib",
"no_cuda": False,
"bert_model": BERT_PRETRAINED_PATH,
"output_dir": OUTPUT_PATH,
"max_seq_length": 512,
"do_train": True,
"do_eval": True,
"do_lower_case": True,
"train_batch_size": 8,
"eval_batch_size": 16,
"learning_rate": 5e-5,
"num_train_epochs": 6,
"warmup_proportion": 0.0,
"no_cuda": False,
"local_rank": -1,
"seed": 42,
"gradient_accumulation_steps": 1,
"optimize_on_cpu": False,
"fp16": True,
"fp16_opt_level": "O1",
"weight_decay": 0.0,
"adam_epsilon": 1e-8,
"max_grad_norm": 1.0,
"max_steps": -1,
"warmup_steps": 500,
"logging_steps": 50,
"eval_all_checkpoints": True,
"overwrite_output_dir": True,
"overwrite_cache": False,
"seed": 42,
"loss_scale": 128,
"task_name": 'intent',
"model_name": 'xlnet-base-cased',
"model_type": 'xlnet'
})
import logging
logfile = str(LOG_PATH/'log-{}-{}.txt'.format(run_start_time, args["run_text"]))
logging.basicConfig(
level=logging.INFO,
format='%(asctime)s - %(levelname)s - %(name)s - %(message)s',
datefmt='%m/%d/%Y %H:%M:%S',
handlers=[
logging.FileHandler(logfile),
logging.StreamHandler(sys.stdout)
])
logger = logging.getLogger()
logger.info(args)
# tokenizer = BertTokenizer.from_pretrained(BERT_PRETRAINED_PATH, do_lower_case=args['do_lower_case'])
device = torch.device('cuda')
if torch.cuda.device_count() > 1:
args.multi_gpu = True
else:
args.multi_gpu = False
label_cols = ["toxic", "severe_toxic", "obscene", "threat", "insult", "identity_hate"]
databunch = BertDataBunch(args['data_dir'], LABEL_PATH, args.model_name, train_file='train.csv', val_file='val.csv',
test_data='test.csv',
text_col="comment_text", label_col=label_cols,
batch_size_per_gpu=args['train_batch_size'], max_seq_length=args['max_seq_length'],
multi_gpu=args.multi_gpu, multi_label=True, model_type=args.model_type)
databunch.train_dl.dataset[0][3]
# train_df.head(20)
# databunch = BertDataBunch.load(args['data_dir'])
num_labels = len(databunch.labels)
num_labels
# databunch.train_dl.dataset[10]
# torch.distributed.init_process_group(backend="nccl",
# init_method = "tcp://localhost:23459",
# rank=0, world_size=1)
metrics = []
metrics.append({'name': 'accuracy_thresh', 'function': accuracy_thresh})
metrics.append({'name': 'roc_auc', 'function': roc_auc})
metrics.append({'name': 'fbeta', 'function': fbeta})
learner = BertLearner.from_pretrained_model(databunch, args.model_name, metrics=metrics,
device=device, logger=logger, output_dir=args.output_dir,
finetuned_wgts_path=FINETUNED_PATH, warmup_steps=args.warmup_steps,
multi_gpu=args.multi_gpu, is_fp16=args.fp16,
multi_label=True, logging_steps=0)
learner.fit(args.num_train_epochs, args.learning_rate, validate=True)
learner.validate()
learner.save_model()
learner.predict_batch(list(pd.read_csv('../data/test.csv')['comment_text'].values))
```
| github_jupyter |
```
import re, sys, math, json, os, urllib.request
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image
from IPython.display import display
from time import gmtime, strftime
#Loading data from DataTurks
try:
model_data = pd.read_json('data.json',
lines=True,
orient='columns')
except Exception as e:
print('Data load error: ',e)
import pandas as pd
import numpy as np
import re
import nltk
from nltk.corpus import stopwords
import keras
from numpy import array
from keras.preprocessing.text import one_hot
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers.core import Activation, Dropout, Dense
from keras.layers import Flatten, LSTM
from keras.layers import GlobalMaxPooling1D
from keras.layers.embeddings import Embedding
from sklearn.model_selection import train_test_split
from keras.preprocessing.text import Tokenizer
raw_data = model_data
raw_data.head()
import seaborn as sns
import re
from sklearn.model_selection import train_test_split
from string import punctuation
from collections import Counter
import numpy as np
#Deleting the metadata, getting only the title
titles = [re.split("^(.+?),", title)[-1] for title in raw_data.content]
#Deleting whitespace
titles = [re.sub(r'\s+', ' ', title) for title in titles]
#Checking for titles without labels
labels = [md['labels'] for md in raw_data.annotation]
labels2 = []
for i in range(0, len(labels)):
if i:
labels2.append(labels[i])
else:
titles.pop(i)
labels = labels2
#One-hot encoding
distressed_labels = []
not_distressed_labels = []
na_labels = []
for label in labels:
if "Distress" in label:
distressed_labels.append(1)
else:
distressed_labels.append(0)
if"Not Distress" in label:
not_distressed_labels.append(1)
else:
not_distressed_labels.append(0)
if "N/A" in label:
na_labels.append(1)
else:
na_labels.append(0)
encoded_labels = list(zip(distressed_labels, not_distressed_labels, na_labels))
#Dividing the data into testing and training sets
X_train, X_test, y_train, y_test = train_test_split(titles, encoded_labels, test_size=0.20, random_state=42)
dist_train = [item[0] for item in y_train]
not_dist_train = [item[1] for item in y_train]
na_train = [item[2] for item in y_train]
dist_test = [item[0] for item in y_test]
not_dist_test = [item[1] for item in y_test]
na_test = [item[2] for item in y_test]
#Preparing testing and training dataframes
train_df = pd.DataFrame({
'text': X_train,
'distressed': dist_train,
'not_distressed': not_dist_train,
'na': na_train,
'labels':y_train
})
print(train_df.head())
eval_df = pd.DataFrame({
'text': X_test,
'distressed': dist_test,
'not_distressed': not_dist_test,
'na': na_test,
'labels': y_test
})
print([eval_df['labels']])
from simpletransformers.classification import MultiLabelClassificationModel
#Preparing the simpletransformers model
model = MultiLabelClassificationModel('roberta', 'roberta-base', num_labels=3, use_cuda=False, args={'output_dir': 'third_outputs/'})
#Training the model
model.train_model(train_df, multi_label=True)
#Evaluating the model
result, model_outputs, wrong_predictions = model.eval_model(eval_df)
#Trying out a prediction
model.predict(['High income flux expected.'])
#Viewing evaluation
result
import numpy as np
from sklearn.metrics import label_ranking_average_precision_score
#A quick implementation of the zero rule algorithm, to compare metrics
def zero_rule_algorithm(train_df, eval_df):
outputs = [t for t in train_df['labels']]
most_common = max(set(outputs), key = outputs.count)
return [most_common for i in range(len(eval_df))]
predictions = zero_rule_algorithm(train_df, eval_df)
values = [i for i in eval_df['labels']]
label_ranking_average_precision_score(values, predictions)
```
| github_jupyter |
# Problem Set 4, Spring 2021, Villas-Boas
Due <u>As posted on bCourses Assignments page</u>
This problem set is to be done using R. To receive full credit, answers must include a correct answer, demonstrate all steps (codes and corresponding output) used to obtain the answer, and be uploaded to [Gradescope](https://www.gradescope.com/courses/226571), correctly indicating on which pages your answers to each question can be found.
## Data
The data set `Pset4_2021.dta` contains state-level data on the percent of children born with low birth-weight, infant mortality, and the percent of population on AFDC (Aid to Families with Dependent Children). AFDC was an entitlement program that guaranteed benefits to all recipients whose income and resources were below state-determined eligibility levels. The data set also contains variables describing additional state characteristics, including per-capita income, per-capita number of doctors, etc. for the year 1987.
|Variable Name | Description |
| :----------- | :----------------------- |
|lowbrth | percent low weight births |
|infmort | infant mortality rate (unit: deaths per 1,000 live births) |
|afdcprt | # participants in AFDC (unit: 1000 persons) |
|popul | population (unit: 1000 persons) |
|pcinc | per capita income (unit: \$ per capita) |
|afdcprc | percent of population on AFDC |
|afdcpay | average monthly AFDC payment (unit: \$) |
|afdcinc | afdcpay as % per capita income |
|stateabb | state postal code |
|state | name of state |
|bedspc | hospital beds per capita |
|povrate | % population below poverty line |
|physicpc | physicians per capita |
|GovDem | = 1 if the state had a democratic governor in 1986, = 0 otherwise (i.e. Republican) \*|
\* Source: https://en.wikipedia.org/wiki/1986 United States gubernatorial elections
## Preamble
Use the following code cell to load the dataset and any packages you plan to use (at least **tidyverse** and **haven**).
```
# Add your preamble code here
```
## Exercise 1
**1.** Please provide a table of summary statistics (mean, median, std, min and max) for percent low birth weight, infant
mortality rate, physicians per capita, average AFDC monthly payment, hospital beds per capita, percent population below the poverty line, and the indicator for Democratic Governor.
*(Hint: See [Coding Bootcamp Part 5](https://r.datahub.berkeley.edu/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fds-modules%2FENVECON-118&urlpath=tree%2FENVECON-118%2FSpring2021-J%2FSections%2FCoding+Bootcamps) for how to do this using the `stargazer` package.)*
➡️ Type your written work for and answer to part Exercise 1 - Q1 here.
```
# insert code here
```
**2.** Please graph the histogram of percent low birthweight for all the states. Then, plot another histogram for states with average monthly AFDC payment (afdcpay) larger than the median. Finally, plot the same graph for states with average monthly AFDC payment (afdcpay) less than or equal to the median. What is the average low birthweight percentage for each group?
➡️ Type your written work for and answer to part Exercise 1 - Q2 here.
```
# insert code here
```
**3.** Produce a new plot that overlaps the histograms for the below and above median groups. What do you conclude in terms of similarities and differences in the two histograms? (Keep answers brief: max 2-4 sentences)
(Hint: see the Histograms section of [Coding Bootcamp Part 4](https://r.datahub.berkeley.edu/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fds-modules%2FENVECON-118&urlpath=tree%2FENVECON-118%2FSpring2021-J%2FSections%2FCoding+Bootcamps) for help overlapping histograms)
➡️ Type your written work for and answer to part Exercise 1 - Q3 here.
```
# insert code here
```
**4.** Estimate the model of low percent birth weight as a linear function of a constant, average monthly AFDC payment, and physicians per capita. Comment on the estimated intercept and each of the right hand side variables' estimated parameters in terms of Sign, Size, and Significance (SSS).
➡️ Type your written work for and answer to part Exercise 1 - Q4 here.
```
# insert code here
```
**5.** In absolute terms, does the average monthly AFDC payment or the per capita number of physicians matter more in predicting the percent low birth weight in a state? (Use standardized approach)
➡️ Type your written work for and answer to part Exercise 1 - Q5 here.
```
# insert code here
```
**6.** Estimate the model of low percent birth weight as a linear function of a constant, average monthly AFDC payment, hospital beds per capita, physicians per capita, and the percent below poverty rate. Test the joint significance of the hospital beds per capita and physicians per capita variables at the 10% significance level.
➡️ Type your written work for and answer to part Exercise 1 - Q6 here.
```
# insert code here
```
**7.** Specify a model to predict the **average low percent birthweight** of a state with a monthly average AFDC payment equal to \$550, 0.008 hospital beds per capita, and 0.092 physicians per capita.
➡️ Type your written work for and answer to part Exercise 1 - Q7 here.
```
# insert code here
```
**8.** Run that same regression model specified in 7 that allows you to easily obtain "inputs" for a confidence interval for that average prediction and report the 99% confidence interval.
(Hint for 7 and 8: Generate the new x variables such that when you regress y on newx1 newx2 newx3 the constant is what you need, <u>as we did in lecture 14</u>.)
➡️ Type your written work for and answer to part Exercise 1 - Q8 here.
```
# insert code here
```
**9.** Estimate the model that allows you to easy obtain the 99% confidence interval <u>for the average predicted</u> **infant mortality rate** for a state with a monthly average AFDC payment equal to \$550, 0.008 hospital beds per capita, and 0.092 physicians per capita.
➡️ Type your written work for and answer to part Exercise 1 - Q9 here.
```
# insert code here
```
**10.** Suppose I told you I want you to construct the 99% confidence interval for a <u>specific state's predicted</u> **infant mortality rate** according to the model you estimated in 9 above. <u>**True or False?**</u> "The 99% confidence interval for a specific state's predicted infant mortality rate is wider than the CI you obtained in 9 for the predicted average infant mortality rate?" <u>Explain briefly.</u>
(Hint: See lectures 14 / 15)
➡️ Type your written work for and answer to part Exercise 1 - Q10 here.
```
# insert code here
```
**11.** First, estimate the model (A) of infant mortality rate as a linear function of a constant, monthly average AFDC payment, per capita hospital beds, per capita hospital beds squared, per capita physicians, and the poverty rate. Next, consider and estimate an alternative model (B) of infant mortality rate as a linear function of a constant, the monthly average AFDC payment, the log of per capita hospital beds, per capita physicians, and the poverty rate. Which one do you prefer,
model (A) or (B)?
(Hint: See lecture 13)
➡️ Type your written work for and answer to part Exercise 1 - Q11 here.
```
# insert code here
```
## Exercise 2
Consider two models below:
\begin{align*}
lowbrth &= \beta_0 + \beta_1 AFDC + \beta_2 bedspc + \beta_3 physicpc + u & (\text{model 2.A}) \\
log(lowbrth) &= \alpha_0 + \alpha_1 AFDC + \alpha_2 bedspc + \alpha_3 physicpc + v & (\text{model 2.B})
\end{align*}
**(a)** Predict the average percent low birth-weight for a state with afdcpay = 400, bedspc = 0.001 and physicpc = 0.1 using model 2.B.
(Hint: See lectures 14/15)
➡️ Type your written work for and answer to part Exercise 2 - Part (a) here.
```
# insert code here
```
**(b)** Which of the models do you prefer? Show all the calculations and the required values obtained via R to answer this question.
(Hint: See lecture 15)
➡️ Type your written work for and answer to part Exercise 2 - Part (b) here.
```
# insert code here
```
## Exercise 3
**(a)** Estimate model 2.B separately for states with a democratic governor and for those with a republican governor in 1986. Formally test at the 10% significance level whether the estimation regression should be done separately or whether we can pool the data.
(Hint: See lecture 17 slide 37 onward)
➡️ Type your written work for and answer to part Exercise 3 - Part (a) here.
```
# insert code here
```
**(b)** I would like to know whether the effect of the monthly AFDC payments on the log of percent low birthweight (in model 2.B) differs depending on whether the governor is democrat or not (republican in that case). Estimate a model that enables you to test this and please interpret your findings. Compare the p value for the estimated coefficient of interest to the 10 percent significance level to conclude whether you reject the null of no heterogeneity in the effect of monthly AFDC payments on the percent low birth weight due to the governor's party affliation, against a two sided alternative, holding all else equal.
(Hint: Generate the interaction you need and add it to the regression.)
➡️ Type your written work for and answer to part Exercise 3 - Part (b) here.
```
# insert code here
```
| github_jupyter |
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
```
# Lecture 3A - Apply & Map, Misc
# Table of Contents
* [Lecture 3A - Apply & Map, Misc](#Lecture-3A---Apply-&-Map,-Misc)
*
* [Content](#Content)
* [Learning Outcomes](#Learning-Outcomes)
* [1. Functions and Dataframes - Using *apply()* and *applymap()*](#1.-Functions-and-Dataframes---Using-*apply%28%29*-and-*applymap%28%29*)
* [Functions along an axis](#Functions-along-an-axis)
* [Functions applied element-wise](#Functions-applied-element-wise)
* [Dummy Variables](#Dummy-Variables)
* [2. Removing Duplicates](#2.-Removing-Duplicates)
* [3. Transpose](#3.-Transpose)
---
### Content
1. Applying functions to dataframes
2. Removing duplicates
3. Re-shaping dataframes with transpose
4. Shift operations for time series
### Learning Outcomes
At the end of this lecture, you should be able to:
* apply functions to dataframes
* remove duplicate rows in dataframes
* transpose dataframes
* apply shift operations to dataframes for time series data
---
```
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
from pylab import rcParams
%matplotlib inline
# Set some Pandas options as you like
pd.set_option('max_columns', 30)
pd.set_option('max_rows', 30)
rcParams['figure.figsize'] = 15, 10
rcParams['font.size'] = 20
```
## 1. Functions and Dataframes - Using *apply()* and *applymap()*
Built-in or user-defines functions can be applied along the entire axes of a dataframe.
To apply a function to an entire axis (or multiple axes) of a dataframe, we resort to the apply() method, which can take an optional axis argument to determine if the axis is vertical/column-wise (0) or horizontal/row-wise (1).
### Functions along an axis
```
df = pd.DataFrame({'one' : pd.Series(np.random.randn(3), index=['a', 'b', 'c']),
'two' : pd.Series(np.random.randn(4), index=['a', 'b', 'c', 'd']),
'three' : pd.Series(np.random.randn(3), index=['b', 'c', 'd'])})
df = df[['one','two','three']]
df
```
Below is an example of applying a built in sum function
```
df.apply(np.sum, axis=0)
```
**Exercise**: Apply the mean function to the above dataframe in a row-wise manner.
```
df.apply(np.mean, axis=1)
```
**Exercise**: Apply the sum function to columns 'one' and 'two' only in a row-wise manner, and assign the result to a new column in the dataframe called 'four'.
```
df['four'] = df[['one', 'two']].apply(np.sum, axis=1)
df
```
**Exercise**: Replace the missing value in both columns with the row-wise mean value.
**Exercise**: Calculate the column-wise product for the first and third columns only.
**Exercise**: Write a function which calculates the sum of a vector and then returns the square of the sum. Once you have done this, apply your function to the dataframe in a row-wise manner, whilst creating a new column 'five', to which you will add insert the result.
```
def square_of_summed_vector(x):
return result
```
### Functions applied element-wise
The apply() method produces some form of aggregate calculations on the axes of a dataframe. applymap() on the other hand extends us the flexibility of applying functions which manipulate single elements in a dataframe.
Say we would like to define a function which returns 'pos' for a positive number and alternatively 'neg'
```
def pos_neg_to_string(x):
if x >= 0:
return 'pos'
else: return 'neg'
```
We can apply this to our dataframe as follows:
```
df.applymap(pos_neg_to_string)
```
Having the ability to apply element-wise operations on dataframes is extremely useful when it comes to dataset cleaning and transformations.
Let's take a look at a sample from a real-world dataset used for gathering results from a survey:
```
assig = pd.read_csv("../datasets/surveySample.csv")
assig.head()
assig.OCCUPATION_M.head(20)
```
Clearly the values in this column need to be cleaned up.
Let's first find out what all the unique values are in this dataset.
```
assig.OCCUPATION_M.unique()
```
We can now write a function that removes the first 3 characters in each entry in order to tidy the values.
```
def remove_first_three_chars(x):
return x.replace(x[:3], '')
assig[['OCCUPATION_M']].applymap(remove_first_three_chars)
```
In order to make the change permanent, we need to assign the result to the dataframe:
```
assig['OCCUPATION_M'] = assig[['OCCUPATION_M']].applymap(remove_first_three_chars)
```
## Dummy Variables
A dummy variable is a numerical variable used in data analysis to represent subgroups of the sample in under study.
In research design, a dummy variable is often used to distinguish different treatment groups. This is accomplished by taking distinct values from a column and creating new columns out of them which are populated with 0 or 1 in order to indicate whether or not the particular data point belongs to this.
This is a frequent operation that can be easily in Python.
```
assig
assig['OCCUPATION_M'].str.get_dummies()
```
We can also specify if there are multiple values within some cells that should be treated as separate columns. In this example we will say that the forward slash indicates a distinct value for which we would like to generate a column for.
```
assig['OCCUPATION_M'].str.get_dummies('/')
```
**Exercise:** From the assignment dataset, consider the column 'supermarket spend in a week'. The '\$' character can cause issues in some applications. We want to clean up this column in such a way that the first 3 characters are replaced as well as the '\$' character, and we also want to change entries with 'No Answer' to reflect that they are actually missing values so replace them with np.NaN. Write a function to do this and apply this function to this column.
Verify that your code works.
## 2. Removing Duplicates
Duplicate rows may be naturally occurring in some datasets or they might arise from input errors. In many instances, like machine learning, these duplicate entries need to be removed from the datasets.
Dataframes provide straightforward functionality to remove such records.
Here is an example:
```
df = pd.DataFrame({'c1': ['one'] * 3 + ['two'] * 4,
'c2': [1, 1, 2, 3, 3, 4, 4]})
df
```
`drop_duplicates` returns a DataFrame where the duplicated rows **across all columns** are dropped:
```
df.drop_duplicates()
```
We can also pass a particular column we would like the duplicates removed from. Let's first make a change to the dataframe:
```
df.loc[1, 'c1'] = 'five'
df
df.drop_duplicates(['c2'])
```
Notice that `drop_duplicates` by default keep the first observed value combination.
## 3. Transpose
Transposing is a special form of reshaping tabular data in such a way that the rows become columns and likewise the columns become rows.
```
df = pd.DataFrame({'one' : pd.Series(np.random.randn(3), index=['a', 'b', 'c']),
'two' : pd.Series(np.random.randn(4), index=['a', 'b', 'c', 'd']),
'three' : pd.Series(np.random.randn(3), index=['b', 'c', 'd'])})
df = df[['one','two','three']]
df
```
Transpose of a dataframe can be accomplished using either the transpose() method call or simple .T
```
df.T
```
Transpose operations are not permanent unless you re-assign the result back tothe original dataframe.
```
df
```
**Exercise:** Slice and select out a dataframe with rows 'c' and 'd' and columns 'one' and 'two', then execute a transpose.
| github_jupyter |
<a href="https://colab.research.google.com/github/Siddharthgolecha/HighQ/blob/main/QOSF_Mentorship_Task_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#Basic Setup
##Installing required packages
For this task, I am using Qiskit along with PyTorch to complete it.
Installing packages:
* Qiskit v 0.18.3
* Pytorch v 1.9.0+cu102
```
!pip install qiskit
!pip install torch
```
##Importing the required libraries
The libraries to be imported are:
* Qiskit
* Pytorch
* Numpy
* Matplotlib
```
import qiskit
import numpy as np
import torch
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, Aer
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
print(qiskit.__version__)
print(qiskit.__qiskit_version__)
print(torch.__version__)
```
#Generating Random States
##Creating TrainingData and Label classes to faciliate training
* TrainingData - Creates a dataset of random values of required bit length and size
* Labels - Wrapper class for the predefined output labels
```
class TrainingData:
"""
Creates a Training Dataset of random quantum states.
Args:
bit_len (int): The number of bits required to represent a state.
size (int): The size of dataset required for training. Minimum can be 1
and maximum (n^2)
backend (AerBackend): The backend required to run the quantum simulation
for random state generation. If not set, the
QASM Simulator will be chosen as default.
shots (int): The number of shots provided to the backend. Default value
is 1024.
Raises:
TypeError: Raises the error, if the type determined is not valid.
ValueError: Raises the error, if the size is either less than 1 or
greater than n^2.
"""
def __init__(self, bit_len, size, backend=Aer.get_backend('qasm_simulator'),
shots=1024):
if not isinstance(bit_len, int):
raise TypeError("The length of bits must be an integer instead of {}.".
format(type(bit_len).__name__))
if not isinstance(size, int):
raise TypeError("The size of dataset must be an integer instead of {}.".
format(type(size).__name__))
if size < 1:
raise ValueError("The minimum size of dataset must be 1.")
if size > bit_len**2:
raise ValueError("The maximum size of dataset can be generated from {} \
qubits is {}".format(bit_len, bit_len**2))
self.bit_len = bit_len
self.backend = backend
self.shots = shots
self.size = size
counts = self.generate_samples(self.bit_len, self.backend, self.shots)
while len(counts) < self.size:
#Generates samples again if they less the required size
counts = self.generate_samples(self.bit_len, self.backend, self.shots)
#Sort the dictionary based on the count of states
self.counts = dict(sorted(counts.items(),
key=lambda item: item[1]))
self._samples = list(self.counts.keys())[:self.size]
self.params = None
def generate_samples(self, qubits, backend=Aer.get_backend('qasm_simulator'),
shots=1024):
"""
Returns the samples and its counts generated by a random circuit.
Args:
qubits (int): The number of quantum bits and classical bits to generate
samples.
backend (AerBackend): The backend required to run the quantum simulation
for random state generation. If not set, the
QASM Simulator will be chosen as default.
shots (int): The number of shots provided to the backend. Default value
is 1024.
"""
circuit = QuantumCircuit(qubits,qubits)
for i in range(qubits):
args = np.random.rand(3)*2*np.pi
circuit.u(args[0], args[1], args[2], i)
circuit.measure(i,i)
result = backend.run(circuit, shots=shots).result()
counts = result.get_counts()
return counts
def to_params(self):
"""
Returns the dataset converted from states into angles(radians) for quantum
circuit.
Returns:
Ndarray: A 2D array of shape (self.size, self.bit_len) with paramaters
to be passed to QNNCircuit for training.
"""
self.params = np.zeros((self.size,self.bit_len))
for index in range(self.size):
sample = self.samples[index]
for i in range(self.bit_len):
state = sample[i]
if state=="1":
#RY(π/2)|0> = X|0> = |1>
self.params[index][self.bit_len-i-1] = np.pi/2
return self.params
@property
def samples(self):
"""
Returns the samples generated by the random circuit
Returns:
List: A list of length 'size' containing different samples generated by
the quantum circuits
"""
return self._samples
@samples.setter
def samples(self, samples):
self._samples = samples
def __str__(self):
return str(self.samples)
class Labels:
"""
Creates a wrapper for labels for training the model
Args:
data (list(str)): A list of strings containing different states to be
generated by the model.
Raises:
TypeError: Raises an error, if any of the element is either not string or
a binary string
"""
def __init__(self, *data):
if isinstance(data[0], list):
data = data[0]
for label in data:
if not isinstance(label, str):
raise TypeError("Expected a string of strings but recieved a {}".
format(type(label).__name__))
try:
check = int(label,2)
except TypeError:
raise TypeError("The label {} is not a binary string".format(label))
self._data = list(data)
self.size = len(self.data)
self.bit_len = len(self.data[0])
@property
def data(self):
"""
Returns the labels
Returns:
List: List of binary labels for validation
"""
return self._data
@data.setter
def data(self, data):
self._data = data
def __str__(self):
return str(self._data)
random_states = TrainingData(4, size=4)
labels = Labels('0011', '0101', '1010', '1100')
print(random_states)
```
**Aim**: To map these newly random states to the predefined label states
For example -
* 0010 -> 0011
* 0011 -> 0101
* 0111 -> 1010
* 1000 -> 1100
**Proposed Solution**: Use Hybrid QGANs with Quantum Generators and Classical Discriminator to generate a required states for the random states.
#Design Quantum Circuit for QNN
The quantum circuit consists of alternating group of RY and RZ gates followed by full entanglement of CNOT gates. Taking the 32 theta and 4 label values, the quantum circuit can look like -

```
class QNNCircuit:
"""
Creates a Quantum Neural Network Circuit for Quantum Generator
Args:
kernel_size (int): The number of qubits in the Quantum Neural Network
Circuit.
backend (AerBackend): The backend required to run the quantum simulation
for random state generation. If not set, the
QASM Simulator will be chosen as default.
shots (int): The number of shots provided to the backend. Default value
is 1024.
Raises:
TypeError: Raises the error, if the type determined is not valid.
"""
def __init__(self, kernel_size,
backend=Aer.get_backend('qasm_simulator'),
shots = 1024):
if not isinstance(kernel_size, int):
raise TypeError("The number of qubits must be an integer instead of {}.".
format(type(kernel_size).__name__))
# --- Circuit definition ---
self.kernel_size = kernel_size
self.training_qubits = QuantumRegister(kernel_size, name="training")
self.measuring_register = ClassicalRegister(kernel_size, "meas")
self.circuit = QuantumCircuit(self.training_qubits,
self.measuring_register)
# ---------------------------
self.backend = backend
self.shots = shots
def create_circuit(self, thetas, label):
"""
Add the gates and assign parameters to Quantum Circuit.
Args:
thetas (list(float), ndarray(float)): Training parameters to assign to
the quantum circuit.
label (list(float), ndarray(float)): Parameters to initialize the
state to the circuit.
Raises:
TypeError: Raises an error if the arguments are neither list nor
ndarray.
ValueError: Raises an erorr if label size is not equal to kernel_size
"""
if not isinstance(thetas, (list, np.ndarray)):
raise TypeError("Expected a list of floating or integer values but \
recieved a {} for thetas".format(type(thetas).__name__))
if not isinstance(label, (list, np.ndarray)):
raise TypeError("Expected a list of floating or integer values but \
recieved a {} for label".format(type(label).__name__))
if len(label) != self.kernel_size:
raise ValueError("Expected a size of {} for label but recieved {}".
format(self.kernel_size, len(label)))
if isinstance(thetas, np.ndarray):
thetas = thetas.tolist()
if isinstance(label, np.ndarray):
label = label.tolist()
#Initialising the Quantum state on the Quantum Circuit
#RY(π/2)|0> = |1>
#RY(0)|0> = |0>
for i in range(len(label)):
self.circuit.ry(label[i], self.training_qubits[i])
for i in range(len(thetas)):
#Fully entangled CNOT gates after each RY and RZ gates pair
if (i%(2*self.kernel_size))==0 and i!=0:
for j in range(self.kernel_size):
for k in range(j+1,self.kernel_size):
self.circuit.cnot(self.training_qubits[j],
self.training_qubits[k])
#Alternating RY and RZ gates
if (i//self.kernel_size)%2 == 0:
self.circuit.ry(thetas[i],
self.training_qubits[i%self.kernel_size])
else:
self.circuit.rz(thetas[i],
self.training_qubits[i%self.kernel_size])
def expval(self):
"""
Calculates the expectation value of the created circuit.
Expectation Value = Sum(state(i)*probability(i))
Returns:
Float: Expectation value of the quantum circuit.
"""
expval = 0
result = self.get_states()
for key,val in result.items():
expval += key*val
return expval
def get_states(self):
"""
Measures the quantum circuit and outputs the probability of each state
Returns:
Dict(string: float): Probaility of different states measured by the
quantum circuit
"""
self.circuit.measure(self.training_qubits, self.measuring_register)
job = self.backend.run(self.circuit, shots=self.shots)
counts = job.result().get_counts()
output = {key: val/self.shots for key, val in counts.items()}
return output
```
#Defining Hybrid Quantum GAN
The Hybrid Quantum GAN consists of Quantum Generator and Classical Discriminator
* **Quantum Generator** - It takes the random states and uses QNNCircuit to generate the probability of all the possible states.
* **Classical Discriminator** - It takes the probabilties generated by the generator to check if it is producing real or fake outputs.
```
def get_accuracy(pred_list, actual):
"""
Determines the average accuracy of the generated data.
Args:
pred_list (list(list(float))): Probabilty of different states of different
random input states
actual (list(str)): Correct labels which is used to determine the accuracy
Returns:
float: Returns the accuracy of the output generated by the generator
"""
total = 0
for i in range(len(pred_list)):
total += pred_list[i][int(actual[i],2)]
return total/len(pred_list)
def hotshot(counts, bit_len):
"""
Converts the different states and its probability into a list of all state
relative probabilities wrt highest probable state.
Args:
counts (dict(str:float)): Dictionary with different states as the keys and
probailities as its values
bit_len (int): The length of bits provided
Returns:
list(float): List of probabilities of state converted into the index of
the list
"""
new_count = []
max_val = max(list(counts.values()))
for i in range(bit_len*bit_len):
key = "{0:b}".format(i).zfill(bit_len)
if key in counts:
new_count.append(counts[key]/max_val)
else:
new_count.append(0.0)
return new_count
class QNNModel(nn.Module):
"""
Quantum Nueral Network model to generate different states
Args:
kernel_size (int): The number of qubits in the Quantum Neural Network
Circuit.
backend (AerBackend): The backend required to run the quantum simulation
for random state generation. If not set, the
QASM Simulator will be chosen as default.
shots (int): The number of shots provided to the backend. Default value
is 1024.
n_params (int): Number of parameters to be trained. If not set, 8 will be
chosen as deafult.
Raises:
ValueError: Raises the error, if the size is less than 1
"""
def __init__(self,kernel_size,
backend = Aer.get_backend('qasm_simulator'),
shots = 1024, n_params = 8):
super(QNNModel, self).__init__()
if n_params < 1:
raise ValueError("The minimum number of parameters must be 1.")
self.backend = backend
self.kernel_size = kernel_size
self.shots = shots
self.n_params = n_params
self.param = nn.Parameter(torch.rand(self.n_params,
requires_grad=True)*np.pi)
def forward(self, data):
"""
Performs forward propogation
Args:
data (TrainingData): Random training data to initialize the state
Returns:
tensor: Tensor of different probabilities of different input states
"""
size = len(data.samples)
self.circuits = [QNNCircuit(self.kernel_size) for i in range(size)]
param = self.param.data.numpy()
data_params = data.to_params()
for i in range(size):
self.circuits[i].create_circuit(param, data_params[i])
output = [hotshot(self.circuits[i].get_states(),
self.kernel_size) for i in range(size)]
return torch.tensor(output)
class Discriminator(nn.Module):
"""
Creates a discriminator for the GANs. Contains a single Linear layer
followed by the sigmoid activation function.
Args:
n (int): The number of neurons in the hidden layer.
Returns:
float: The probailility of given output state
"""
def __init__(self, n):
super(Discriminator, self).__init__()
self.dense = nn.Linear(int(n), 1);
self.activation = nn.Sigmoid()
def forward(self, x):
return self.activation(self.dense(x))
def GAN_training(generator, discriminator, g_optimizer, d_optimizer,
loss_fn, g_loss_list = [], d_loss_list = [],
g_accuracy_list = [], d_accuracy_list = [], epochs = 15000):
"""
Trains the GAN with given generator and discriminator
Args:
generator (torch.nn.Module): Generator for GAN
discriminator (torch.nn.Module): Discriminator for GAN
g_optimizer (torch.optim): Generator's Optimizer
d_optimizer (torch.optim): Discriminator's Optimizer
loss_fn (torch.nn.Loss): Loss function for Discriminator
g_loss_list (list): List to store Generator's Loss
d_loss_list (list): List to store Discriminator's Loss
g_accuracy_list (list): List to store Generator's Accuracy
d_accuracy_list (list): List to store Discriminator's Accuracy
epochs (int): Number of iterations for GAN. Deafult value is 15000
"""
true_labels = torch.full((labels.size,), 1, dtype=torch.float)
true_data = torch.tensor([hotshot({state:1},labels.bit_len)
for state in labels.data])
fake_labels = torch.zeros(labels.size)
generator.train()
discriminator.train()
for epoch in range(1,epochs+1):
# Forward pass
g_optimizer.zero_grad()
# Generate the data on the random states
g_out = generator(random_states)
g_accuracy = get_accuracy(g_out, labels.data)
# Calculate the Discriminator's loss on output of random states and true
# labels
g_d_out = discriminator(g_out).view(-1)
g_loss = loss_fn(g_d_out, true_labels)
# Train the discriminator on the true data and true labels
d_optimizer.zero_grad()
true_d_out = discriminator(true_data).view(-1)
true_d_loss = loss_fn(true_d_out, true_labels)
# Train the discriminator on the output of random states and fake labels
g_d_f_out = discriminator(g_out.detach()).view(-1)
g_d_loss = loss_fn(g_d_f_out, fake_labels)
d_loss = true_d_loss + g_d_loss
d_loss.backward()
d_optimizer.step()
# Backpropogation of Generator
g_loss.backward()
g_optimizer.step()
g_loss_list.append(g_loss.item())
d_loss_list.append(d_loss.item())
g_accuracy_list.append(g_accuracy)
d_accuracy_list.append(true_d_out.mean(0).detach().numpy())
if epoch%1000==0:
print('Epoch {:03d}\tGenerator Loss: {:.4f}\tGenerator Accuracy: {:.4f}\t'.
format(epoch, g_loss.item(), g_accuracy), end='')
print('Discriminator Loss: {:.4f}\tDiscriminator Accuracy: {:.4f}'.
format(d_loss.item(), d_accuracy_list[-1]))
```
#Train the Hybrid Quantum GAN
```
qgenerator = QNNModel(kernel_size=4, n_params=32)
qdiscriminator = Discriminator(qgenerator.kernel_size**2)
qg_optimizer = optim.Adam(qgenerator.parameters())
qd_optimizer = optim.Adam(qdiscriminator.parameters(), lr=0.001)
qloss_fn = nn.BCELoss()
epochs = 15000
qg_loss_list = []
qd_loss_list = []
qg_accuracy_list = []
qd_accuracy_list = []
GAN_training(qgenerator, qdiscriminator, qg_optimizer, qd_optimizer,
qloss_fn, qg_loss_list, qd_loss_list, qg_accuracy_list,
qd_accuracy_list, epochs = 15000)
```
#Results and Conclusions
```
plt.figure(figsize=(10,5))
plt.title("Generator and Discriminator Loss During Hybrid Quantum \
GAN Training")
plt.plot(qg_loss_list, label='Generator')
plt.plot(qd_loss_list, label='Discriminator')
plt.xlabel('Training Iterations')
plt.ylabel('Training Loss')
plt.legend()
plt.show()
plt.figure(figsize=(10,5))
plt.title("Generator and Discriminator Accuracy During Hybrid Quantum \
GAN Training")
plt.plot(qg_accuracy_list, label='Generator')
plt.plot(qd_accuracy_list, label='Discriminator')
plt.xlabel('Training Iterations')
plt.ylabel('Training Accuracy')
plt.legend()
plt.show()
```
##Conclusions
By analyzing the Generator's and Discriminator's Loss Graph of Hybrid Quantum GAN, we can see that the loss of the Generator is continously increasing while Discriminator's loss is decreasing. This indicates that the Hybrid QGAN model is reaching failure mode. The discriminator can easily identify that the random quantum states cannot produce the predefined output states. It can also be said that the generator is not able to produce the states with probable states.
This can also be verified by analysing the accuracy graph. It shows that the disciminator is becoming good at finding that the given outputs don't correspond to the actual labels. For the ideal GAN model,
* The accuracy of generator should reach 100%.
* The Discriminator's loss should be around 0.5 indicating it is facing hard time deciding if the generator is producing the correct results.
###Possible Solutions
* **Increasing the number of iterations**: Increasing the number of iterations in GANs usually help the generator to produce good data. However, in this case it seem unlikely it will help because with increasing iterations, the loss does not seem to rise to 0.5 or become constant.
* **Decreasing the learning rate of Discriminator's optimizer**: This ensures that the discriminator learns slowly so that generator could get enough time to generate good data. However, during my trails, it also does not seem to work much.
* **Change the Quantum Circuit**: The reason why the generator is not producing the convincing results is that the circuit is not good enough. However, I tried different quantum circuit but none of them seemed to work much good.
#Testing the Quantum Generator
```
def GAN_test(generator, states, labels):
"""
Tests the GAN generator.
Args:
generator (torch.nn.Module): GAN Generator
states (TrainingData): Training Data to generate result
labels (Labels): Output Labels
"""
generator.eval()
with torch.no_grad():
output = generator(states)
for i in range(len(labels.data)):
output_dict = {"{0:b}".format(key).zfill(labels.bit_len):output[i][key]
for key in range(len(output[i]))}
out_state = max(output_dict, key = lambda x: output_dict[x])
print("Random State = {}\tActual Label = {}\t".
format(states.samples[i], labels.data[i]), end=' ')
print("Output Label = {}\tActual Label Accuracy = {:.2f}%".
format(out_state, output_dict[labels.data[i]]*100))
GAN_test(qgenerator, random_states, labels)
new_states = TrainingData(4, size=4)
GAN_test(qgenerator, new_states, labels)
```
| github_jupyter |
```
from datetime import datetime
import pandas as pd
from covid_xprize.scoring.predictor_scoring import load_dataset
from covid_xprize.validation.scenario_generator import generate_scenario
from covid_xprize.validation.scenario_generator import phase1_update
```
# Scenario generator
## Latest data
```
LATEST_DATA_URL = 'https://raw.githubusercontent.com/OxCGRT/covid-policy-tracker/master/data/OxCGRT_latest.csv'
GEO_FILE = "../../countries_regions.csv"
latest_df = load_dataset(LATEST_DATA_URL, GEO_FILE)
len(latest_df.CountryName.unique())
len(latest_df.RegionName.unique())
```
# Scenario: historical IP until 2020-09-30
Latest historical data, truncated to the specified end date
```
start_date_str = None
end_date_str = "2020-09-30"
countries = None
output_file = "data/2020-09-30_historical_ip.csv"
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Historical")
scenario_df[scenario_df.CountryName == "Italy"].Date.max()
truncation_date = pd.to_datetime(end_date_str, format='%Y-%m-%d')
scenario_df = scenario_df[scenario_df.Date <= truncation_date]
scenario_df.tail()
# Write to file
# scenario_df.to_csv(output_file, index=False)
```
# Scenario: frozen NPIs
Latest historical data + frozen NPIS between last known date and end of Januaray 2021 for India and Mexico
## Generate
```
start_date_str = "2021-01-01"
end_date_str = "2021-01-31"
countries = ["India", "Mexico"]
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Freeze")
len(scenario_df)
scenario_df.CountryName.unique()
scenario_df.tail()
```
## Save
```
# Write to a file
# hist_file_name = "data/future_ip.csv"
# scenario_df.to_csv(hist_file_name, index=False)
```
# Robojudge test: December
IP file to test robojudge for the month of December
## Generate
```
today = datetime.utcnow().strftime('%Y%m%d_%H%M%S')
start_date_str = "2020-12-01"
end_date_str = "2020-12-31"
latest_df = load_dataset(LATEST_DATA_URL, GEO_FILE)
countries = None
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Freeze")
# Check: should contain all 366 days of 2020
nb_countries = len(scenario_df.CountryName.unique())
nb_regions = len(scenario_df.RegionName.unique()) - 1 # Ignore the '' region
len(scenario_df) / (nb_countries + nb_regions)
```
## Save
```
from datetime import datetime
sd = 20200101 # IP file always contains data since inception
ed = end_date_str.replace('-', "")
december_file_name = f"../../../covid-xprize-robotasks/ips/tests/{today}_{sd}_{ed}_ips.csv"
scenario_df.to_csv(december_file_name, index=False)
print(f"Saved to {december_file_name}")
```
# Robojudge: Official
IP file robojudge uses for its daily submissions evaluation
## Generate
```
# Handle US Virgin Islands
LATEST_DATA_URL = 'https://raw.githubusercontent.com/OxCGRT/covid-policy-tracker/master/data/OxCGRT_latest.csv'
PHASE1_FILE = "../../countries_regions_phase1_fix.csv"
latest_df = load_dataset(LATEST_DATA_URL, PHASE1_FILE)
latest_df = phase1_update(latest_df)
today = datetime.utcnow().strftime('%Y%m%d_%H%M%S')
start_date_str = "2020-12-22"
end_date_str = "2021-06-19"
countries = None
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Freeze")
# Check: should contain 536 days:
# 366 days of 2020 + 170 days of 2021 (10 days in 2020 + 170 days in 2021 = 180 days of eval)
nb_countries = len(scenario_df.CountryName.unique())
nb_regions = len(scenario_df.RegionName.unique()) - 1 # Ignore the 'nan' region
len(scenario_df) / (nb_countries + nb_regions)
len(scenario_df.CountryName.unique())
len(scenario_df.RegionName.unique())
```
## Save
```
from datetime import datetime
sd = start_date_str.replace('-', "")
ed = end_date_str.replace('-', "")
december_file_name = f"../../../covid-xprize-robotasks/ips/live/{today}_{sd}_{ed}_ips.csv"
scenario_df.to_csv(december_file_name, index=False)
print(f"Saved to {december_file_name}")
```
# Prescriptions
## UK future test
```
start_date_str = None
end_date_str = "2021-02-14"
latest_df = load_dataset(LATEST_DATA_URL, GEO_FILE)
countries = ["United Kingdom"]
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Freeze")
scenario_df.to_csv("~/workspace/covid-xprize-robotasks/ips/prescriptions/uk_future_test_ips.csv", index=False)
```
## All, past and future, test
```
start_date_str = None
end_date_str = "2020-12-31"
latest_df = load_dataset(LATEST_DATA_URL, GEO_FILE)
countries = None
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Historical")
end_date = pd.to_datetime(end_date_str, format='%Y-%m-%d')
scenario_df = scenario_df[scenario_df.Date <= end_date]
scenario_df.Date.max()
scenario_df.to_csv("~/workspace/covid-xprize-robotasks/ips/prescriptions/all_2020_ips.csv", index=False)
```
## China early 2020 test
```
start_date_str = None
end_date_str = "2020-02-14"
latest_df = load_dataset(LATEST_DATA_URL, GEO_FILE)
countries = ["China"]
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries, scenario="Historical")
end_date = pd.to_datetime(end_date_str, format='%Y-%m-%d')
scenario_df = scenario_df[scenario_df.Date <= end_date]
scenario_df.tail()
len(scenario_df)
scenario_df.to_csv("~/workspace/covid-xprize-robotasks/ips/prescriptions/china_early_2020_ips.csv", index=False)
```
| github_jupyter |
# Linear Models with SparkR: uses and present limitations
[**Introduction to Apache Spark with R by J. A. Dianes**](https://github.com/jadianes/spark-r-notebooks)
In this notebook we will use SparkR machine learning capabilities in order to predict property value in relation to other variables in the [2013 American Community Survey](http://www.census.gov/programs-surveys/acs/data/summary-file.html) dataset. The whole point of R on Spark is to introduce Spark scalability into R data analysis pipelines. With this idea in mind, we have seen how [SparkR](http://spark.apache.org/docs/latest/sparkr.html) introduces data types and functions that are very similar to what we are used to when using regular R libraries. The next step in our series of notebooks will deal with its machine learning capabilities. While building a linear model we also want to check the significance of each of the variables involved in building such a predictor for property value.
## Creating a SparkSQL context and loading data
In order to explore our data, we first need to load it into a SparkSQL data frame. But first we need to init a SparkSQL context. The first thing we need to do is to set up some environment variables and library paths as follows. Remember to replace the value assigned to `SPARK_HOME` with your Spark home folder.
```
# Set Spark home and R libs
Sys.setenv(SPARK_HOME='/home/cluster/spark-1.5.0-bin-hadoop2.6')
.libPaths(c(file.path(Sys.getenv('SPARK_HOME'), 'R', 'lib'), .libPaths()))
```
Now we can load the `SparkR` library as follows.
```
library(SparkR)
```
And now we can initialise the Spark context as [in the official documentation](http://spark.apache.org/docs/latest/sparkr.html#starting-up-sparkcontext-sqlcontext). In our case we are use a standalone Spark cluster with one master and seven workers. If you are running Spark in local node, use just `master='local'`. Additionally, we require a Spark package from Databricks to read CSV files (more on this in the [previous notebook](https://github.com/jadianes/spark-r-notebooks/blob/master/notebooks/nb1-spark-sql-basics/nb1-spark-sql-basics.ipynb)).
```
sc <- sparkR.init(master='spark://169.254.206.2:7077', sparkPackages="com.databricks:spark-csv_2.11:1.2.0")
```
And finally we can start the SparkSQL context as follows.
```
sqlContext <- sparkRSQL.init(sc)
```
Now that we have our SparkSQL context ready, we can use it to load our CSV data into data frames. We have downloaded our [2013 American Community Survey dataset](http://www.census.gov/programs-surveys/acs/data/summary-file.html) files in [notebook 0](https://github.com/jadianes/spark-r-notebooks/tree/master/notebooks/nb0-starting-up/nb0-starting-up.ipynb), so they should be stored locally. Remember to set the right path for your data files in the first line, ours is `/nfs/data/2013-acs/ss13husa.csv`.
```
housing_a_file_path <- file.path('', 'nfs','data','2013-acs','ss13husa.csv')
housing_b_file_path <- file.path('', 'nfs','data','2013-acs','ss13husb.csv')
```
Now let's read into a SparkSQL dataframe. We need to pass four parameters in addition to the `sqlContext`:
- The file path.
- `header='true'` since our `csv` files have a header with the column names.
- Indicate that we want the library to infer the schema.
- And the source type (the Databricks package in this case).
And we have two separate files for both, housing and population data. We need to join them.
```
housing_a_df <- read.df(sqlContext,
housing_a_file_path,
header='true',
source = "com.databricks.spark.csv",
inferSchema='true')
housing_b_df <- read.df(sqlContext,
housing_b_file_path,
header='true',
source = "com.databricks.spark.csv",
inferSchema='true')
housing_df <- rbind(housing_a_df, housing_b_df)
```
Let's check that we have everything there by counting the files and listing a few of them.
```
nrows <- nrow(housing_df)
nrows
head(housing_df)
```
## Preparing our data
We need to convert `ST` (or any other categorical variable) from a numeric variable into a factor.
```
housing_df$ST <- cast(housing_df$ST, "string")
housing_df$REGION <- cast(housing_df$REGION, "string")
```
Additionally, we need either to impute values or to remove samples with null values in any of our predictors or desponse. For the response (`VALP`) we will use just those samples with actual values.
```
housing_with_valp_df <- filter(
housing_df,
isNotNull(housing_df$VALP)
& isNotNull(housing_df$TAXP)
& isNotNull(housing_df$INSP)
& isNotNull(housing_df$ACR)
)
```
Let's count the remaining samples.
```
nrows <- nrow(housing_with_valp_df)
nrows
```
## Preparing a train / test data split
We don't have a split function in SparkR, but we can use `sample` in combination with the `SERIALNO` column in order to prepare two sets of IDs for training and testing.
```
housing_df_test <- sample(housing_with_valp_df,FALSE,0.1)
nrow(housing_df_test)
test_ids <- collect(select(housing_df_test, "SERIALNO"))$SERIALNO
```
Unfortunately SparkR doesn't support negative %in% expressions, so we need to do this in two steps. First we add a flag to the whole dataset indicating that a sample belongs to the test set.
```
housing_with_valp_df$IS_TEST <- housing_with_valp_df$SERIALNO %in% test_ids
```
And then we use that flag to subset out the train dataset as follows.
```
housing_df_train <- subset(housing_with_valp_df, housing_with_valp_df$IS_TEST==FALSE)
nrow(housing_df_train)
```
However this approach is not very scalable since we are collecting all the test IDs and passing them over to build the new flag column. What if we have a much larger test set? Hopefully futre versions of SparkR will come up with a proper `split` functionality.
## Training a linear model
In order to train a linear model, we call `glm` with the following parameters:
- A formula: sadly, `SparkR::glm()` gives us an error when we pass more than eight variables using `+` in the formula.
- The dataset we want to use to train the model.
- The type of model (gaussian or binomial).
This doesn't differ much from the usual R `glm` command, although right now is more limited.
The list of variables we have used includes:
- `RMSP` or number of rooms.
- `ACR` the lot size.
- `INSP` or insurance cost.
- `TAXP` or taxes cost.
- `ELEP` or electricity cost.
- `GASP` or gas cost.
- `ST` that is the state code.
- `REGION` that identifies the region.
```
model <- glm(
VALP ~ RMSP + ACR + INSP + TAXP + ELEP + GASP + ST + REGION,
data = housing_df_train,
family = "gaussian")
summary(model, signif.stars=TRUE)
```
Sadly, the current version of `SparkR::summary()` doesn't provide significance starts. That makes model interpretation and selection very difficult. But at least we know how each variables influences a property value. For example, the Midwest region decreases property value, while the West increases it, etc. In order to interpret that we need to have a look at our [data dictionary](http://www2.census.gov/programs-surveys/acs/tech_docs/pums/data_dict/PUMSDataDict13.txt).
In any case, since we don't have significance starts, we can iterate through adding/removing variables and calculating the R2 value. In our case we ended up with the previous model.
## Evaluating our model using the test data
First of all let's obtain the average value for `VALP` that we will use as a reference of a base predictor model.
```
VALP_mean <- collect(agg(
housing_df_train,
AVG_VALP=mean(housing_df_train$VALP)
))$AVG_VALP
VALP_mean
```
Let's now predict on our test dataset as follows.
```
predictions <- predict(model, newData = housing_df_test)
```
Let's add the squared residuals and squared totals so later on we can calculate [R2](https://en.wikipedia.org/wiki/Coefficient_of_determination).
```
predictions <- transform(
predictions,
S_res=(predictions$VALP - predictions$prediction)**2,
S_tot=(predictions$VALP - VALP_mean)**2)
head(select(predictions, "VALP", "prediction", "S_res", "S_tot"))
nrows_test <- nrow(housing_df_test)
residuals <- collect(agg(
predictions,
SS_res=sum(predictions$S_res),
SS_tot=sum(predictions$S_tot)
))
residuals
R2 <- 1.0 - (residuals$SS_res/residuals$SS_tot)
R2
```
In regression, the R2 coefficient of determination is a statistical measure of how well the regression line approximates the real data points. An R2 of 1 indicates that the regression line perfectly fits the data.
A value of 0.41 doesn't speak very well about our model.
## Conclusions
We still need to improve our model if we really want to be able to predict property values. However there are some limitations in the current SparkR implementation that stop us from doing so. Hopefully these limitations won't be there in further versions. Moreover, we are using a linear model, and the relationships between our predictors and the target variable might not be linear at all.
But right now, in Spark v1.5, the R machine learning capabilities are still very limited. We are missing a few things, such as:
- Accepting more than 8 variables in formulas using `+`.
- Having significance stars that help model interpretation and selection.
- Having other indicators (e.g. R2) in summary objects so we don't have to calculate them ourselves.
- Being able to create more complex formulas (e.g. removing intercepts using 100 + ...) so we don't get negative values, etc.
- Although we have a `sample` method, we are missing a `split` one that we can use to easier have train/test splits.
- Being able to use more powerful models (or at least models that deal better with non linearities), and not just linear ones.
| github_jupyter |
<a href="https://colab.research.google.com/github/RecursiveOuroboros/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/High_Elipitcal_Orbit_Sat_Data_IoT_Telementary_Predictive_Maitenance_Jason_Meil_DS3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# Importing Appropriate Libraries
import random
import seaborn as sns
import numpy as np
import sklearn as skl
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import RobustScaler
from sklearn.cluster import KMeans
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
import pandas as pd
import matplotlib.pyplot as plt
# Importing the DataSet
data = pd.read_csv('http://aws-proserve-data-science.s3.amazonaws.com/device_failure.csv', encoding='latin1')
data.head(10)
#exporing data types and looking for null values
data.dtypes
data.isnull()
data['failure'].value_counts()
# Dropping uncessecary data
data_2 = data.drop(columns=['device', 'date'])
data_2.columns
data_2.groupby('failure').count()
sns.set()
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute1']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute2']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute3']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute4']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute5']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute6']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['failure']
y = data_2['attribute7']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['attribute8']
y = data_2['failure']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# same plotting code as above!
x = data_2['attribute9']
y = data_2['failure']
plt.plot(x, y)
plt.legend('ABCDEF', ncol=2, loc='upper left');
# Synthetic Minority Oversampling Algorithm
X = data_2.loc[:, data_2.columns != 'failure']
y = data_2.loc[:, data_2.columns == 'failure']
from imblearn.over_sampling import SMOTE
os = SMOTE(random_state=0)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=30)
columns = X_train.columns
os_data_X,os_data_y=os.fit_sample(X_train, y_train)
os_data_X = pd.DataFrame(data=os_data_X,columns=columns )
os_data_y= pd.DataFrame(data=os_data_y,columns=['y'])
# we can Check the numbers of our data
print("length of oversampled data is ",len(os_data_X))
print("Number of no subscription in oversampled data",len(os_data_y[os_data_y['y']==0]))
print("Number of subscription",len(os_data_y[os_data_y['y']==1]))
print("Proportion of no subscription data in oversampled data is ",len(os_data_y[os_data_y['y']==0])/len(os_data_X))
print("Proportion of subscription data in oversampled data is ",len(os_data_y[os_data_y['y']==1])/len(os_data_X))
# Logit Variable Assignment
y = data_2['failure']
X = data_2[['attribute1', 'attribute2', 'attribute3', 'attribute4', 'attribute5', 'attribute6', 'attribute7', 'attribute8', 'attribute9']]
#logisticRegr = LogisticRegression()
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(X,y, train_size=.85, test_size=.15)
X_train.shape, X_val.shape, y_train.shape, y_val.shape
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
y_pred = logreg.predict(X_test)
print('Accuracy of logistic regression classifier on test set: {:.2f}'.format(logreg.score(X_test, y_test)))
# Run Confusion matrix shows 37315 correct and 34 incorrect
from sklearn.metrics import confusion_matrix
confusion_matrix = confusion_matrix(y_test, y_pred)
print(confusion_matrix)
import statsmodels.api as sm
logit_model=sm.Logit(y,X)
result=logit_model.fit()
print(result.summary2())
cols= [ 'attribute2', 'attribute4', 'attribute5', 'attribute7', 'attribute8', 'attribute9']
X=data_2 [cols]
y=data_2 ['failure']
logit_model=sm.Logit(y,X)
result=logit_model.fit()
print(result.summary2())
# Comptute Precision, recall, and F-Measure Support
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
#Plot ROC Curve
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
logit_roc_auc = roc_auc_score(y_test, logreg.predict(X_test))
fpr, tpr, thresholds = roc_curve(y_test, logreg.predict_proba(X_test)[:,1])
plt.figure()
plt.plot(fpr, tpr, label='Logistic Regression (area = %0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
from sklearn.feature_extraction.text import CountVectorizer
clf = RandomForestClassifier()
clf.fit(X, y)
rfc = RandomForestClassifier(n_estimators=1000,
min_samples_split=6,
criterion='gini',
max_features='auto',
oob_score=True,
random_state=1,
n_jobs=-1)
rfc.fit(X_train, y_train)
# Running a Decision Tree Algorithm
# Confirming that we still have failure target and
# 9 attributes across axis 1
print ("Dataset Length:: ", len(data_2))
print ("Dataset Shape:: ", data_2.shape)
data_2.head()
#Slicing to split data appropriatley for TTS
Y = data_2.values[:,1]
X = data_2.values[:, 2:9]
# 30% of whole data set and 70% Training Set
X_train, X_test, y_train, y_test = train_test_split( X, Y, test_size = 0.3, random_state = 100)
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn import tree
clf_gini = DecisionTreeClassifier(criterion = "gini", random_state = 100,
max_depth=3, min_samples_leaf=5)
clf_gini.fit(X_train, y_train)
plt.show()
clf_entropy = DecisionTreeClassifier(criterion = "entropy", random_state = 100,
max_depth=3, min_samples_leaf=5)
clf_entropy.fit(X_train, y_train)
```
| github_jupyter |
## View original data
View original data provided by telegram
```
import pandas as pd
from tqdm import tqdm_notebook
import json
file = 'dc0202'
with open(f'telegram_data/{file}-input.txt', 'r', encoding='utf-8') as f:
input_lines = f.read().split('\n')
with open(f'telegram_data/{file}-output.txt', 'r', encoding='utf-8') as f:
output_lines = f.read().split('\n')
data = {
"title": [],
"description": [],
"recent_posts": [],
"language": []
}
for line in tqdm_notebook(input_lines):
try:
line = json.loads(line)
except:
pass
if line != '':
data['title'].append(line['title'])
data['description'].append(line['description'])
data['recent_posts'].append('\n'.join(line['recent_posts']))
for line in tqdm_notebook(output_lines):
try:
line = json.loads(line)
except:
pass
if line != '':
data['language'].append(line['lang_code'])
data = pd.DataFrame(data)
```
## tgstat2octoparse
Parse tgstat links to octoparse links format
```
import json
import pandas as pd
CATEGORY = 'Hobbies & Activities'
print(CATEGORY)
with open("data/category.txt", 'r', encoding='utf-8-sig') as f:
data = json.loads(f.read())
with open("data/channelsEN.json", 'r', encoding='utf-8') as f:
channels = json.loads(f.read())
# channels[CATEGORY] = []
# with open("data/channelsEN.json", 'w+', encoding='utf-8') as f:
# json.dump(channels, f)
channels_data = []
for i, item in enumerate(data['items']['list']):
channels_data.append(item['channelIdCode'])
channels[CATEGORY] += channels_data
with open("data/channelsEN.json", 'w+', encoding='utf-8') as f:
json.dump(channels, f)
for i, item in enumerate(channels.keys()):
print(i, item, len(channels[item]))
# write to links for octoparse
links = []
for cat in list(channels.keys())[30:]:
for l in channels[cat]:
links.append(f'https://in.tgstat.com/channel/{l}?category={cat}'.replace(' ', '%20'))
with open('data/EN_TGSTAT_LINKS/links4.txt', 'w') as f:
f.write('\n'.join(links))
```
## vk2octoparse
Parse vk links to octoparse format
```
import json
import pandas as pd
CATEGORY = 'Other'
with open("tgparser/categoryVK.txt", 'r', encoding='utf-8-sig') as f:
data = json.loads(f.read())
with open("tgparser/vkgroupsRU.json", 'r', encoding='utf-8') as f:
groups = json.loads(f.read())
import urllib
import time
links = []
for query in ['архитектура', 'дизайн', 'design', 'искусство', 'architecture']:
for i in range(10):
OFFSET = f'&offset={25 * i}'
QUERY = urllib.parse.quote(query)
URL = f'https://allsocial.ru/entity?direction=1&is_closed=0&is_verified=-1&list_type=1&order_by=diff_abs&period=day&platform=1&range=1000:12000000&str={QUERY}&type_id=-1'
response = urllib.request.urlopen(URL + OFFSET).read().decode('utf-8')
response = json.loads(response)
for item in response['response']['entity']:
links.append(f'''https://vk.com/public{item['vk_id']}''')
time.sleep(0.5)
```
## octoparse2json
Parse octoparse data to final data
```
LANG = 'EN'
import json
import pandas as pd
import os
data = pd.DataFrame(columns = ['Text', 'Text1', 'Field1', 'Page_URL'])
for file in os.listdir(f'data/{LANG}_TGSTAT_LINKS'):
data = pd.concat([data, pd.read_excel(os.path.join(f'data/{LANG}_TGSTAT_LINKS', file))])
data.columns = ['title', 'description', 'recent_posts', 'category']
print(data.isna().sum())
data = data[data['category'].notna()]
data['category'] = data['category'].apply(lambda x: x.split('?category=')[-1].replace('%20', ' '))
data = data[~data['category'].apply(lambda x: 'http' in x)].reset_index(drop = True)
import re
regex = re.compile(r'\\(?![/u"])')
for cat in data['category'].unique():
tmp = data.query('category == @cat')
for title in tmp['title'].unique():
json_data = {}
try:
channel_data = tmp.query('title == @title')
json_data["title"] = str(title).replace("'", '').replace('"', '')
json_data["description"] = str(channel_data['description'].iloc[0]).replace("'", '').replace('"', '')
json_data["recent_posts"] = []
for i in range(channel_data.shape[0]):
post = str(channel_data.iloc[i, 2]).replace("'", '').replace('"', '')
json_data["recent_posts"].append(post)
except:
print('parse error')
json_data = regex.sub(r"\\\\", str(json_data).replace("'", '"'))
json_data = regex.sub(r"\\", json_data)
with open(f'data/{LANG}_TGSTAT_DATA/{cat}.txt', 'a+', encoding='utf-8') as f:
#json.dump(json_data, f, ensure_ascii=False)
f.write(json_data+'\n')
from tqdm import tqdm_notebook
with open(f'data/{LANG}_TGSTAT_DATA/Bets & Gambling.txt', 'r', encoding='utf-8') as f:
input_lines = f.read().split('\n')
data = {
"title": [],
"description": [],
"recent_posts": [],
}
for line in input_lines:
try:
line = json.loads(line.replace('\\', ''))
except:
pass
if line != '':
data['title'].append(line['title'])
data['description'].append(line['description'])
data['recent_posts'].append('\n'.join(line['recent_posts']))
data = pd.DataFrame(data)
```
## json2translate
Parse final data into tsv for translation
```
SEP = '<-------------новоесообщение------------->'
from tqdm import tqdm_notebook
LANG = 'EN'
for cat in os.listdir(f'data/{LANG}_TGSTAT_DATA'):
data = {
"title": [],
"description": [],
"recent_posts": [],
}
with open(f'data/{LANG}_TGSTAT_DATA/{cat}', 'r', encoding='utf-8') as f:
input_lines = f.read().split('\n')
for line in input_lines:
try:
line = json.loads(line.replace('\\', ''))
if line != '':
title, desc, posts = line['title'], line['description'], SEP.join(line['recent_posts'])
data['title'].append(title)
data['description'].append(desc)
data['recent_posts'].append(posts)
except:
print(line)
data = pd.DataFrame(data)
cat = cat.split('.')[0]
data.to_excel(f'Translate/{cat}.xlsx', index = None)
with open(f'Translate/EN-RU/{cat}.tsv', 'w+') as f:
f.write('')
import os
import pandas as pd
cats = [cat for cat in os.listdir(f'data/RU_TGSTAT_DATA')]
pd.read_csv(f'''Translate/RU-EN/{cats[26].split('.')[0]}.tsv''', sep='\t')
```
## translate2json
Parse translated data back to json
```
SEP_EN = '<-------------newpost------------->'
SEP_RU = '<-------------новоесообщение------------->'
import os
import re
import pandas as pd
FOLDER = 'EN_TGSTAT_DATA_TRANSLATE'
regex = re.compile(r'\\(?![/u"])')
for cat in os.listdir(f'data/EN_TGSTAT_DATA'):
cat = cat.split(".")[0]
try:
data = pd.read_csv(f'''Translate/RU-EN/{cat}.tsv''', sep='\t')
except:
continue
data.columns = ['title', 'description', 'recent_posts']
for i in range(data.shape[0]):
channel_data = data.iloc[i]
json_data = {}
json_data["title"] = str(channel_data['title']).replace("'", '').replace('"', '')
json_data["description"] = str(channel_data['description']).replace("'", '').replace('"', '')
json_data["recent_posts"] = str(channel_data['recent_posts']).replace("'", '').replace('"', '').split(SEP_RU)
json_data = regex.sub(r"\\\\", str(json_data).replace("'", '"'))
json_data = regex.sub(r"\\", json_data)
with open(f'data/{FOLDER}/{cat}.txt', 'a+', encoding='utf-8') as f:
f.write(json_data+'\n')
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Visualization/nwi_wetlands_symbology.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/nwi_wetlands_symbology.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Visualization/nwi_wetlands_symbology.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# NWI legend: https://www.fws.gov/wetlands/Data/Mapper-Wetlands-Legend.html
def nwi_add_color(fc):
emergent = ee.FeatureCollection(
fc.filter(ee.Filter.eq('WETLAND_TY', 'Freshwater Emergent Wetland')))
emergent = emergent.map(lambda f: f.set(
'R', 127).set('G', 195).set('B', 28))
# print(emergent.first())
forested = fc.filter(ee.Filter.eq(
'WETLAND_TY', 'Freshwater Forested/Shrub Wetland'))
forested = forested.map(lambda f: f.set('R', 0).set('G', 136).set('B', 55))
pond = fc.filter(ee.Filter.eq('WETLAND_TY', 'Freshwater Pond'))
pond = pond.map(lambda f: f.set('R', 104).set('G', 140).set('B', 192))
lake = fc.filter(ee.Filter.eq('WETLAND_TY', 'Lake'))
lake = lake.map(lambda f: f.set('R', 19).set('G', 0).set('B', 124))
riverine = fc.filter(ee.Filter.eq('WETLAND_TY', 'Riverine'))
riverine = riverine.map(lambda f: f.set(
'R', 1).set('G', 144).set('B', 191))
fc = ee.FeatureCollection(emergent.merge(
forested).merge(pond).merge(lake).merge(riverine))
base = ee.Image(0).mask(0).toInt8()
img = base.paint(fc, 'R') \
.addBands(base.paint(fc, 'G')
.addBands(base.paint(fc, 'B')))
return img
fromFT = ee.FeatureCollection("users/wqs/Pipestem/Pipestem_HUC10")
Map.addLayer(ee.Image().paint(fromFT, 0, 2), {}, 'Watershed')
huc8_id = '10160002'
nwi_asset_path = 'users/wqs/NWI-HU8/HU8_' + huc8_id + '_Wetlands' # NWI wetlands for the clicked watershed
clicked_nwi_huc = ee.FeatureCollection(nwi_asset_path)
nwi_color = nwi_add_color(clicked_nwi_huc)
Map.centerObject(clicked_nwi_huc, 10)
Map.addLayer(nwi_color, {'gamma': 0.3, 'opacity': 0.7}, 'NWI Wetlands Color')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.