text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Problem Statement
- BBC News Classification
# Data Description
- Dataset consists of 2225 news articles extracted from the BBC website between 2004 and 2005
- Published open source by Insight Resources and was collected by UC Davis for research
# Goal
- Create a text classifier that will streamline the process of categorizing news publications
- Classify BBC news articles into five categories using natural Language Processing and Machine Learning
- The five news topics are: Politics, Entertainment, Sports, Technology, and Business
# Business Constraint
- Misclassification is not a big issue
- latancy is could be an issue considering end user requirement
# Converting the data into CSV file.
```
# importing libraries
import os
import pandas as pd
from sklearn.model_selection import train_test_split
# asigning the location
data_folder = "/Users/abuzaid/Downloads/Machine Learning/Pianalystics/BBC_News"
folders = ["business","entertainment","politics","sport","tech"]
os.chdir(data_folder)
x = []
y = []
# loop to iterate through folder
for i in folders:
files = os.listdir(i)
for text_file in files:
file_path = i + "/" +text_file
print ("reading file:", file_path)
with open(file_path,encoding="utf8", errors='ignore') as f:
data = f.readlines()
data = ' '.join(data)
x.append(data)
y.append(i)
#Making dictionary
data = {'news': x, 'type': y}
# COnverting into Dataframe
df = pd.DataFrame(data)
print ('Converting csv flie ...')
data=df
data.head()
```
# Importing Libraries
```
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from prettytable import PrettyTable
```
# Performing simple EDA
```
data.columns #Checking columns
data.isnull().sum() #Checking for null values
data.isna().sum() #Checking for Nan values
```
# Text Preproccesing
```
from nltk.stem.porter import PorterStemmer
import re
import string
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from bs4 import BeautifulSoup
# DECONTRACTING
# eg: can't = can not
import re
def decontracted(phrase):
# specific
phrase = re.sub(r"won't", "will not", phrase)
phrase = re.sub(r"can\'t", "can not", phrase)
# general
phrase = re.sub(r"n\'t", " not", phrase)
phrase = re.sub(r"\'re", " are", phrase)
phrase = re.sub(r"\'s", " is", phrase)
phrase = re.sub(r"\'d", " would", phrase)
phrase = re.sub(r"\'ll", " will", phrase)
phrase = re.sub(r"\'t", " not", phrase)
phrase = re.sub(r"\'ve", " have", phrase)
phrase = re.sub(r"\'m", " am", phrase)
return phrase
# Defing Stopword Explictly
stopwords= set(['br', 'the', 'i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've",\
"you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', \
'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their',\
'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', \
'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', \
'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', \
'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after',\
'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further',\
'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more',\
'most', 'other', 'some', 'such', 'only', 'own', 'same', 'so', 'than', 'too', 'very', \
's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', \
've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn',\
"hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'mustn',\
"mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", \
'won', "won't", 'wouldn', "wouldn't"])
#Preprocessing the text
from tqdm import tqdm
preprocessed_text = []
# tqdm is for printing the status bar
for sentance in tqdm(data['news'].values):
sentance = re.sub(r"http\S+", "", sentance)
sentance = BeautifulSoup(sentance, 'lxml').get_text()
sentance = decontracted(sentance)
sentance = re.sub("\S*\d\S*", "", sentance).strip()
sentance = re.sub('[^A-Za-z]+', ' ', sentance)
sentance = ' '.join(e.lower() for e in sentance.split() if e.lower() not in stopwords)
preprocessed_text.append(sentance.strip())
#Appending processed text into DATA
data['news']=preprocessed_text
```
# Data After preprocessing
```
maping = { "business":0,
"entertainment":1,
"politics":2,
"sport":3,
"tech":4}
data.loc[:,"type"]=data.type.map(maping)
data
from wordcloud import WordCloud
import matplotlib.pyplot as plt
fig, (ax1) = plt.subplots(1, figsize=[9, 9])
wordcloud = WordCloud( background_color='white',
width=600,
height=600).generate(" ".join(data['news']))
ax1.imshow(wordcloud)
ax1.axis('off')
ax1.set_title('Frequent Words',fontsize=6);
import seaborn as sns
sns.distplot(data['type'])
```
# Business & Politics are rightly skwed , and Tech is rightly skewed , while Entertainment and Sport is following Gaussian Distribution
```
sns.distplot(data['type'], kde=False, rug=True,bins=20);
```
# This is unbalanced dataset so we need to keep in mind while creating model
```
sns.countplot(x="type", data=data)
```
# Spliting data
```
X = data.news
y = data.type
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size = 0.30, random_state = 100)
X_cv, X_test, y_cv, y_test = train_test_split(X_train, y_train, train_size = 0.30, random_state = 100)
```
# Using TF-IDF to convert text into vectors
```
tfidf_vectorizer = TfidfVectorizer(stop_words = 'english', max_df = 0.7) # Creating tfidf object
tfidf_train = tfidf_vectorizer.fit_transform(X_train) #Converting Text to tfidf vectors
tfidf_cv = tfidf_vectorizer.transform(X_cv)
tfidf_test = tfidf_vectorizer.transform(X_test)
```
# ********************** Creating Models *****************
# 1. PassiveAggressiveClassifier
# Training
```
pac = PassiveAggressiveClassifier(max_iter = 50)
pac.fit(tfidf_train, y_train)
cv_pred = pac.predict(tfidf_cv)
pac_score_cv = accuracy_score(y_cv, cv_pred)*100
print('Accuracy:', pac_score_cv)
cm= confusion_matrix(y_cv, cv_pred)
sns.heatmap(cm,annot=True)
```
# Testing
```
test_pred = pac.predict(tfidf_test)
pac_score_test = accuracy_score(y_test, test_pred)*100
print('~~~~~~~Accuracy on Test data~~~~~~~~~~~:', pac_score_test)
cm1= confusion_matrix(y_test, test_pred)
sns.heatmap(cm1,annot=True)
# NO Misclassification whatsoever
```
# 2. Logistic Reggression
# Training
```
lr = LogisticRegression()
lr.fit(tfidf_train, y_train)
cv_pred = lr.predict(tfidf_cv)
lr_score_cv = accuracy_score(y_cv, cv_pred)*100
print('Accuracy:',lr_score_cv)
print(confusion_matrix(y_cv, cv_pred))
```
# Testing
```
test_pred = lr.predict(tfidf_test)
lr_score_test = accuracy_score(y_test, test_pred)*100
print('Accuracy on test data: ',lr_score_test)
cm2= confusion_matrix(y_test, test_pred)
sns.heatmap(cm2,annot=True)
```
# 3. SVC
# Training
```
svc = SVC()
svc.fit(tfidf_train, y_train)
cv_pred = svc.predict(tfidf_cv)
svc_score_cv = accuracy_score(y_cv, cv_pred)*100
print('Accuracy: ', svc_score_cv)
print(confusion_matrix(y_cv, cv_pred))
```
# Testing
```
test_pred = svc.predict(tfidf_test)
svc_score_test = accuracy_score(y_test, test_pred)*100
print('Accuracy: ',svc_score_test)
cm3=confusion_matrix(y_test, test_pred)
sns.heatmap(cm3,annot=True)
```
# - Reason :
- the model fail as we used categorial handing diffrently which dont work very well with linear model
- hence SVC failed
# 4. Naive Bayes
- Base model for NLP task
```
## Training
nb = GaussianNB()
nb.fit(tfidf_train.toarray(), y_train)
cv_pred = nb.predict(tfidf_cv.toarray())
nb_score_cv = accuracy_score(y_cv, cv_pred)*100
print('Accuracy: ',nb_score_cv)
print(confusion_matrix(y_cv, cv_pred))
# Testing
test_pred = nb.predict(tfidf_test.toarray())
nb_score_test = accuracy_score(y_test, test_pred)*100
print('Accuracy:', nb_score_test)
cm4=confusion_matrix(y_test, test_pred)
sns.heatmap(cm4,annot=True)
```
# Summary of all trained and tested models
```
x = PrettyTable()
x.field_names = ['Model Name', 'CV_Accuracy', 'Testing_Accuracy']
x.add_row(['PassiveAggressiveClassifier', pac_score_cv , pac_score_test])
x.add_row([" -------------------------"," ----------- "," ----------- "])
x.add_row(['Logistic Regression', lr_score_cv , lr_score_test])
x.add_row([" -------------------------"," ----------- "," ----------- "])
x.add_row(['Support Vector Classifier', svc_score_cv , svc_score_test])
x.add_row([" -------------------------"," ----------- "," ----------- "])
x.add_row(['Naive Bayes', nb_score_cv , nb_score_test])
print(x)
```
# Conclusion : Passive Aggressive Classiffier having train and test best accuracy as well as Confusion matrix shows it classsify correctly
# TEST MODEL
```
text="The 2000 election wasn't settled until the Supreme Court decision Bush v. Gore on December 12, five weeks after the election. During that time the S&P 500 dropped 5% and the small-cap Russell 2000 tumbled 7%, according to DataTrek Research. Treasury yields dipped as demand rose and gold outperformed."
def preprocess(x):
x = str(x).lower()
x = x.replace(",000,000", "m").replace(",000", "k").replace("′", "'").replace("’", "'") .replace("won't", "will not").replace("cannot", "can not").replace("can't", "can not") .replace("n't", " not").replace("what's", "what is").replace("it's", "it is") .replace("'ve", " have").replace("i'm", "i am").replace("'re", " are") .replace("he's", "he is").replace("she's", "she is").replace("'s", " own") .replace("%", " percent ").replace("₹", " rupee ").replace("$", " dollar ") .replace("€", " euro ").replace("'ll", " will")
x = re.sub(r"([0-9]+)000000", r"\1m", x)
x = re.sub(r"([0-9]+)000", r"\1k", x)
x = re.sub(r"http\S+", "", x)
porter = PorterStemmer()
pattern = re.compile('\W')
if type(x) == type(''):
x = re.sub(pattern, ' ', x)
if type(x) == type(''):
x = porter.stem(x)
example1 = BeautifulSoup(x)
x = example1.get_text()
return x
text = preprocess(text)
# Passing text trough the preprocessing function
text
text_vect = tfidf_vectorizer.transform(([text])) #transforming test text using TFIDF
pred = pac.predict(text_vect)
if pred==0:
print('business')
elif pred == 1:
print('Entertainment')
elif pred== 2:
print("politics")
elif pred == 3:
print("Sport")
elif pred== 4 :
print("Tech")
```
| github_jupyter |
```
%logstop
%logstart -rtq ~/.logs/vc.py append
%matplotlib inline
import matplotlib
import seaborn as sns
sns.set()
matplotlib.rcParams['figure.dpi'] = 144
from static_grader import grader
```
# Object-oriented exercises
## Introduction
The objective of these exercises is to develop your familiarity with Python's `class` syntax and object-oriented programming. By deepening our understanding of Python objects, we will be better prepared to work with complex data structures and machine learning models. We will develop a `Point` class capable of handling some simple linear algebra operations in 2D.
## Exercise 1: `point_repr`
The first step in defining most classes is to define their `__init__` and `__repr__` methods so that we can construct and represent distinct objects of that class. Our `Point` class should accept two arguments, `x` and `y`, and be represented by a string `'Point(x, y)'` with appropriate values for `x` and `y`.
When you've written a `Point` class capable of this, execute the cell with `grader.score` for this question (do not edit that cell; you only need to modify the `Point` class).
```
class Point(object):
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return "Point(%d, %d)" % (self.x, self.y)
grader.score.vc__point_repr(lambda points: [str(Point(*point)) for point in points])
```
## Exercise 2: add_subtract
The most basic vector operations we want our `Point` object to handle are addition and subtraction. For two points $(x_1, y_1) + (x_2, y_2) = (x_1 + x_2, y_1 + y_2)$ and similarly for subtraction. Implement a method within `Point` that allows two `Point` objects to be added together using the `+` operator, and likewise for subtraction. Once this is done, execute the `grader.score` cell for this question (do not edit that cell; you only need to modify the `Point` class.)
(Remember that `__add__` and `__sub__` methods will allow us to use the `+` and `-` operators.)
```
class Point(object):
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return "Point(%d, %d)" % (self.x, self.y)
def __add__(self, number):
return Point(self.x + number.x, self.y + number.y)
def __sub__(self, number):
return Point(self.x - number.x, self.y - number.y)
from functools import reduce
def add_sub_results(points):
points = [Point(*point) for point in points]
return [str(reduce(lambda x, y: x + y, points)),
str(reduce(lambda x, y: x - y, points))]
grader.score.vc__add_subtract(add_sub_results)
```
## Exercise 3: multiplication
Within linear algebra there's many different kinds of multiplication: scalar multiplication, inner product, cross product, and matrix product. We're going to implement scalar multiplication and the inner product.
We can define scalar multiplication given a point $P$ and a scalar $a$ as
$$aP=a(x,y)=(ax,ay)$$
and we can define the inner product for points $P,Q$ as
$$P\cdot Q=(x_1,y_1)\cdot (x_2, y_2) = x_1x_2 + y_1y_2$$
To test that you've implemented this correctly, compute $2(x, y) \cdot (x, y)$ for a `Point` object. Once this is done, execute the `grader.score` cell for this question (do not edit that cell; you only need to modify the `Point` class.)
(Remember that `__mul__` method will allow us to use the `*` operator. Also don't forget that the ordering of operands matters when implementing these operators.)
```
class Point(object):
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return "Point(%d, %d)" % (self.x, self.y)
def __add__(self, number):
return Point(self.x + number.x, self.y + number.y)
def __sub__(self, number):
return Point(self.x - number.x, self.y - number.y)
def __mul__(self, number):
return self.x * number.x + self.y * number.y
# **if the point is multiplied by just a number**
if isinstance(number, int):
return Point(self.x * number, self.y * number)
def mult_result(points):
points = [Point(*point) for point in points]
return [point*point*2 for point in points]
grader.score.vc__multiplication(mult_result)
```
## Exercise 4: Distance
Another quantity we might want to compute is the distance between two points. This is generally given for points $P_1=(x_1,y_1)$ and $P_2=(x_2,y_2)$ as
$$D = |P_2 - P_1| = \sqrt{(x_1-x_2)^2 + (y_1-y_2)^2}.$$
Implement a method called `distance` which finds the distance from a point to another point.
Once this is done, execute the `grader.score` cell for this question (do not edit that cell; you only need to modify the `Point` class.)
### Hint
* *You can use the `sqrt` function from the math package*.
```
from math import sqrt
class Point(object):
def __init__(self, x, y):
self.x=x
self.y=y
def distance(self, other):
if isinstance(other, Point):
return sqrt((self.x-other.x)**2+(self.y-other.y)**2)
def __repr__(self):
return f"Point({self.x}, {self.y})"
def dist_result(points):
points = [Point(*point) for point in points]
return [points[0].distance(point) for point in points]
grader.score.vc__distance(dist_result)
```
## Exercise 5: Algorithm
Now we will use these points to solve a real world problem! We can use our Point objects to represent measurements of two different quantities (e.g. a company's stock price and volume). One thing we might want to do with a data set is to separate the points into groups of similar points. Here we will implement an iterative algorithm to do this which will be a specific case of the very general $k$-means clustering algorithm. The algorithm will require us to keep track of two clusters, each of which have a list of points and a center (which is another point, not necessarily one of the points we are clustering). After making an initial guess at the center of the two clusters, $C_1$ and $C_2$, the steps proceed as follows
1. Assign each point to $C_1$ or $C_2$ based on whether the point is closer to the center of $C_1$ or $C_2$.
2. Recalculate the center of $C_1$ and $C_2$ based on the contained points.
See [reference](https://en.wikipedia.org/wiki/K-means_clustering#Standard_algorithm) for more information.
This algorithm will terminate in general when the assignments no longer change. For this question, we would like you to initialize one cluster at `(1, 0)` and the other at `(-1, 0)`.
The returned values should be the two centers of the clusters ordered by greatest `x` value. Please return these as a list of numeric tuples $[(x_1, y_1), (x_2, y_2)]$
In order to accomplish this we will create a class called cluster which has two methods besides `__init__` which you will need to write. The first method `update` will update the center of the Cluster given the points contained in the attribute `points`. Remember, you after updating the center of the cluster, you will want to reassign the points and thus remove previous assignments. The other method `add_point` will add a point to the `points` attribute.
Once this is done, execute the `grader.score` cell for this question (do not edit that cell; you only need to modify the `Cluster` class and `compute_result` function.)
```
class Cluster(object):
def __init__(self, x, y):
self.center = Point(x, y)
self.points = []
def update(self):
len_ = len(self.points)
# Check if there are points (to aovid zero division). 0 means False any other number grater than zero True
if len_ :
xs = [p.x for p in self.points]
ys = [p.y for p in self.points]
self.center.x = sum(xs) / len_
self.center.y = sum(ys) / len_
# Empty points list
self.points = []
def add_point(self, point):
self.points.append(point)
def compute_result(points):
points = [Point(*point) for point in points]
a = Cluster(1,0)
b = Cluster(-1,0)
a_old = []
for _ in range(10000): # max iterations
for point in points:
if point.distance(a.center) < point.distance(b.center):
a.add_point(point)
else:
b.add_point(point)
if a_old == a.points:
break
a_old = list(a.points)
# Update centers
a.update()
b.update()
return [(a.center.x, a.center.y),(b.center.x,b.center.y)]
grader.score.vc__k_means(compute_result)
```
*Copyright © 2020 The Data Incubator. All rights reserved.*
| github_jupyter |

# _*Qiskit Aqua: Generating Random Variates*_
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-iqx-tutorials.
***
### Contributors
Albert Akhriev<sup>[1]</sup>, Jakub Marecek<sup>[1]</sup>
### Affiliation
- <sup>[1]</sup>IBMQ
## Introduction
While classical computers use only pseudo-random routines, quantum computers
can generate true random variates.
For example, the measurement of a quantum superposition is intrinsically random,
as suggested by Born's rule.
Consequently, some of the
best random-number generators are based on such quantum-mechanical effects.
Further, with a logarithmic amount of random bits, quantum computers can produce
linearly many more bits, which is known as
randomness expansion protocols.
In practical applications, one wishes to use random variates of well-known
distributions, rather than random bits.
In this notebook, we illustrate ways of generating random variates of several popular
distributions on IBM Q.
## Random Bits and the Bernoulli distribution
It is clear that there are many options for generating random bits (i.e., Bernoulli-distributed scalars, taking values either 0 or 1). Starting from a simple circuit such as a Hadamard gate followed by measurement, one can progress to vectors of Bernoulli-distributed elements. By addition of such random variates, we could get binomial distributions. By multiplication we could get geometric distributions, although perhaps leading to a circuit depth that may be impractical at the moment, though.
Let us start by importing the basic modules and creating a quantum circuit for generating random bits:
```
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import sys, math, time
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
from qiskit import Aer
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, execute
# In this example we use 'qasm_simulator' backend.
glo_backend = Aer.get_backend("qasm_simulator")
```
In the next step we create a quantum circuit, which will be used for generation:
```
# Number of qubits utilised simultaneously.
glo_num_qubits = 5
def create_circuit(num_target_qubits: int) -> QuantumCircuit:
"""
Creates and returns quantum circuit for random variate generation.
:param num_target_qubits: number of qubits to be used.
:return: quantum curcuit.
"""
assert isinstance(num_target_qubits, int) and num_target_qubits > 0
q = QuantumRegister(num_target_qubits)
c = ClassicalRegister(num_target_qubits)
circuit = QuantumCircuit(q, c)
circuit.h(q)
circuit.barrier()
circuit.measure(q, c)
return circuit
# Create and plot generating quantum circuit.
circuit = create_circuit(glo_num_qubits)
#print(circuit)
circuit.draw(output='mpl')
```
## Uniformly-distributed scalars and vectors
It is clear that there are many options for approximating uniformly-distributed scalars by the choice of an integer from a finite range uniformly at random, e.g., by a binary-code construction from the Bernoulli-distributed vectors. In the following snippet, we generate random bits, which we then convert using the binary-code construction, up to the machine precision of a classical computer.
```
def uniform_rand_float64(circuit: QuantumCircuit, num_target_qubits: int,
size: int, vmin: float, vmax: float) -> np.ndarray:
"""
Generates a vector of random float64 values in the range [vmin, vmax].
:param circuit: quantum circuit for random variate generation.
:param num_target_qubits: number of qubits to be used.
:param size: length of the vector.
:param vmin: lower bound.
:param vmax: upper bound.
:return: vector of random values.
"""
assert sys.maxsize == np.iinfo(np.int64).max # sizeof(int) == 64 bits
assert isinstance(size, int) and size > 0
assert isinstance(vmin, float) and isinstance(vmax, float) and vmin <= vmax
nbits = 7 * 8 # nbits > mantissa of float64
bit_str_len = (nbits * size + num_target_qubits - 1) // num_target_qubits
job = execute(circuit, glo_backend, shots=bit_str_len, memory=True)
bit_str = ''.join(job.result().get_memory())
scale = float(vmax - vmin) / float(2**nbits - 1)
return np.array([vmin + scale * float(int(bit_str[i:i+nbits], 2))
for i in range(0, nbits * size, nbits)], dtype=np.float64)
def uniform_rand_int64(circuit: QuantumCircuit, num_target_qubits: int,
size: int, vmin: int, vmax: int) -> np.ndarray:
"""
Generates a vector of random int64 values in the range [vmin, vmax].
:param circuit: quantum circuit for random variate generation.
:param num_target_qubits: number of qubits to be used.
:param size: length of the vector.
:param vmin: lower bound.
:param vmax: upper bound.
:return: vector of random values.
"""
assert sys.maxsize == np.iinfo(np.int64).max # sizeof(int) == 64 bits
assert isinstance(size, int) and size > 0
assert isinstance(vmin, int) and isinstance(vmax, int) and vmin <= vmax
assert abs(vmin) <= 2**52 and abs(vmax) <= 2**52 # 52 == mantissa of float64
return np.rint(uniform_rand_float64(circuit, num_target_qubits,
size, float(vmin), float(vmax))).astype(np.int64)
```
### Uniform distribution over floating point numbers.
In this example we draw a random vector of floating-point values uniformly distributed within some arbitrary selected interval:
```
# Draw a sample from uniform distribution.
start_time = time.time()
sample = uniform_rand_float64(circuit, glo_num_qubits, size=54321, vmin=-7.67, vmax=19.52)
sampling_time = time.time() - start_time
# Print out some details.
print("Uniform distribution over floating point numbers:")
print(" sample type:", type(sample), ", element type:", sample.dtype, ", shape:", sample.shape)
print(" sample min: {:.4f}, max: {:.4f}".format(np.amin(sample), np.amax(sample)))
print(" sampling time: {:.2f} secs".format(sampling_time))
# Plotting the distribution.
plt.hist(sample.ravel(),
bins=min(int(np.ceil(np.sqrt(sample.size))), 100),
density=True, facecolor='b', alpha=0.75)
plt.xlabel("value", size=12)
plt.ylabel("probability", size=12)
plt.title("Uniform distribution over float64 numbers in [{:.2f} ... {:.2f}]".format(
np.amin(sample), np.amax(sample)), size=12)
plt.grid(True)
# plt.savefig("uniform_distrib_float.png", bbox_inches="tight")
plt.show()
```
### Uniform distribution over integers.
Our next example is similar to the previous one, but here we generate a random vector of integers:
```
# Draw a sample from uniform distribution.
start_time = time.time()
sample = uniform_rand_int64(circuit, glo_num_qubits, size=54321, vmin=37, vmax=841)
sampling_time = time.time() - start_time
# Print out some details.
print("Uniform distribution over bounded integer numbers:")
print(" sample type:", type(sample), ", element type:", sample.dtype, ", shape:", sample.shape)
print(" sample min: {:d}, max: {:d}".format(np.amin(sample), np.amax(sample)))
print(" sampling time: {:.2f} secs".format(sampling_time))
# Plotting the distribution.
plt.hist(sample.ravel(),
bins=min(int(np.ceil(np.sqrt(sample.size))), 100),
density=True, facecolor='g', alpha=0.75)
plt.xlabel("value", size=12)
plt.ylabel("probability", size=12)
plt.title("Uniform distribution over int64 numbers in [{:d} ... {:d}]".format(
np.amin(sample), np.amax(sample)), size=12)
plt.grid(True)
# plt.savefig("uniform_distrib_int.png", bbox_inches="tight")
plt.show()
```
## Normal distribution
To generate random variates with a standard normal distribution using two independent
samples $u_1, u_2$ of the uniform distribution on the unit interval [0, 1], one can
consider the Box-Muller transform to obtain a 2-vector:
\begin{align}
\begin{bmatrix}
%R\cos(\Theta )=
{\sqrt {-2\ln u_{1}}}\cos(2\pi u_{2}) \\
% R\sin(\Theta )=
{\sqrt {-2\ln u_{1}}}\sin(2\pi u_{2})
\end{bmatrix},
\end{align}
wherein we have two independent samples of the standard normal distribution.
In IBM Q, this is implemented as follows:
```
def normal_rand_float64(circuit: QuantumCircuit, num_target_qubits: int,
size: int, mu: float, sigma: float) -> np.ndarray:
"""
Draws a sample vector from the normal distribution given the mean and standard
deviation, using the Box-Muller method.
"""
TINY = np.sqrt(np.finfo(np.float64).tiny)
assert isinstance(size, int) and size > 0
rand_vec = np.zeros((size,), dtype=np.float64)
# Generate array of uniformly distributed samples, factor 1.5 longer that
# actually needed.
n = (3 * size) // 2
x = np.reshape(uniform_rand_float64(circuit, num_target_qubits,
2*n, 0.0, 1.0), (-1, 2))
x1 = 0.0 # first sample in a pair
c = 0 # counter
for d in range(size):
r2 = 2.0
while r2 >= 1.0 or r2 < TINY:
# Regenerate array of uniformly distributed samples upon shortage.
if c >= n:
c = 0
n = max(size // 10, 1)
x = np.reshape(uniform_rand_float64(circuit, num_target_qubits,
2*n, 0.0, 1.0), (-1, 2))
x1 = 2.0 * x[c, 0] - 1.0 # first sample in a pair
x2 = 2.0 * x[c, 1] - 1.0 # second sample in a pair
r2 = x1 * x1 + x2 * x2
c += 1
f = np.sqrt(np.abs(-2.0 * np.log(r2) / r2))
rand_vec[d] = f * x1
return (rand_vec * sigma + mu)
```
The following example demonstrates how to draw a random vector of normally distributed variates:
```
# Mean and standard deviation.
mu = 2.4
sigma = 5.1
# Draw a sample from the normal distribution.
start_time = time.time()
sample = normal_rand_float64(circuit, glo_num_qubits, size=4321, mu=mu, sigma=sigma)
sampling_time = time.time() - start_time
# Print out some details.
print("Normal distribution (mu={:.3f}, sigma={:.3f}):".format(mu, sigma))
print(" sample type:", type(sample), ", element type:", sample.dtype, ", shape:", sample.shape)
print(" sample min: {:.4f}, max: {:.4f}".format(np.amin(sample), np.amax(sample)))
print(" sampling time: {:.2f} secs".format(sampling_time))
# Plotting the distribution.
x = np.linspace(mu - 4.0 * sigma, mu + 4.0 * sigma, 1000)
analyt = np.exp(-0.5 * ((x - mu) / sigma)**2) / (sigma * math.sqrt(2.0 * math.pi))
plt.hist(sample.ravel(),
bins=min(int(np.ceil(np.sqrt(sample.size))), 100),
density=True, facecolor='r', alpha=0.75)
plt.plot(x, analyt, '-b', lw=1)
plt.xlabel("value", size=12)
plt.ylabel("probability", size=12)
plt.title("Normal distribution: empirical vs analytic", size=12)
plt.grid(True)
# plt.savefig("normal_distrib.png", bbox_inches="tight")
plt.show()
```
There is a substantial amount of further work needed to either certify the quality of the source of random numbers (cf. NIST SP 800-90B, Recommendation for the Entropy Sources Used for Random Bit Generation) or to use random variates within quantum algorithms (cf. <a href="https://github.com/Qiskit/qiskit-aqua/tree/master/qiskit/aqua/components/uncertainty_models">uncertainty_models</a> within Qiskit Aqua).
```
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
# Figure. CNV eQTL Examples
```
import copy
import cPickle
import os
import subprocess
import cdpybio as cpb
import matplotlib as mpl
import matplotlib.gridspec as gridspec
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
pd.options.mode.chained_assignment = None # default='warn'
import pybedtools as pbt
import scipy.stats as stats
import seaborn as sns
import ciepy
import cardipspy as cpy
%matplotlib inline
%load_ext rpy2.ipython
dy_name = 'figure_cnv_eqtl_examples'
outdir = os.path.join(ciepy.root, 'output', dy_name)
cpy.makedir(outdir)
private_outdir = os.path.join(ciepy.root, 'private_output', dy_name)
cpy.makedir(private_outdir)
import socket
if socket.gethostname() == 'fl-hn1' or socket.gethostname() == 'fl-hn2':
dy = os.path.join(ciepy.root, 'sandbox', 'tmp', dy_name)
cpy.makedir(dy)
pbt.set_tempdir(dy)
fn = os.path.join(ciepy.root, 'output', 'cnv_analysis', 'cnv_gene_variants.pickle')
cnv_gv = pd.read_pickle(fn)
fn = os.path.join(ciepy.root, 'output', 'cnv_analysis', 'lead_variants.pickle')
lead_vars = pd.read_pickle(fn)
fn = os.path.join(ciepy.root, 'output', 'mcnv_analysis', 'reg_results.tsv')
mcnv_results = pd.read_table(fn, index_col=0)
mcnv_sig = mcnv_results[mcnv_results.bh_sig]
fn = os.path.join(ciepy.root, 'private_output', 'mcnv_analysis', 'filtered_mcnvs.tsv')
mcnv_genotypes = pd.read_table(fn, index_col=0)
cnv_lead_vars = cnv_gv[cnv_gv.cnv_is_lead]
cnv_lead_vars = cnv_lead_vars.sort_values(by='pvalue').drop_duplicates(subset=['gene_id'])
mcnv_genotypes.shape
mcnv_sig.cnv.value_counts().head(12)
n = 143953316 - 143951166
print('mCNV length is {:,}.'.format(n))
c = 'CNV_7_143951166_143953316'
fn = os.path.join(ciepy.root, 'output', 'mcnv_analysis', 'CNV_7_143951166_143953316_data.tsv')
data = pd.read_table(fn, index_col=0)
data.columns = list(data.columns[0:-1]) + ['Gene']
sns.set_style('whitegrid')
mcnv_sig[mcnv_sig.cnv == 'CNV_7_143951166_143953316'].gene_name
legend_colors = [
np.array((255,0,0)) / 255.,
np.array((255,105,105)) / 255.,
np.array((250,202,0)) / 255.,
np.array((255,252,4)) / 255.,
np.array((10,190,254)) / 255.,
np.array((0,176,80)) / 255.,
np.array((0,176,80)) / 255.,
np.array((153,255,102)) / 255.,
np.array((245,245,245)) / 255.,
]
ind = [
'Active promoter',
'Weak promoter',
'Strong enhancer',
'Weak/poised enhancer',
'Insulator',
'Transcriptional transition',
'Transcriptional elongation',
'Weak transcribed',
'Heterochromatin',
]
legend_colors = pd.Series(legend_colors, index=ind)
s,p = stats.mannwhitneyu(cnv_lead_vars.drop_duplicates('gene_id').beta.abs(),
lead_vars[lead_vars.cnv_sig == False].drop_duplicates('gene_id').beta.abs())
print('Effect sizes are different with p={:.3e}, Mann Whitney U.'.format(p))
cnv_abs_beta_pdf = pd.Series(index=np.arange(0, 3 + 0.1, 0.1))
se = cnv_lead_vars.drop_duplicates('gene_id').beta.abs()
density = stats.gaussian_kde(se)
cnv_abs_beta_pdf = pd.Series(density(cnv_abs_beta_pdf.index), index=cnv_abs_beta_pdf.index)
snv_abs_beta_pdf = pd.DataFrame(index=np.arange(0, 3 + 0.005, 0.005))
se = lead_vars[lead_vars.cnv_sig == False].drop_duplicates('gene_id').beta.abs()
density = stats.gaussian_kde(se)
snv_abs_beta_pdf = pd.Series(density(snv_abs_beta_pdf.index), index=snv_abs_beta_pdf.index)
fn = os.path.join(ciepy.root, 'output', 'cnv_analysis', 'sig_not_genic_vs_not_sig_roadmap_res.tsv')
intergenic_res = pd.read_table(fn, index_col=0)
repressive = ['H3K9me3', 'H3K27me3']
transcribed = ['H3K36me3']
intergenic_res['type'] = 'Active'
for i in repressive:
intergenic_res.ix[intergenic_res.mark == i, 'type'] = 'Repressive'
for i in transcribed:
intergenic_res.ix[intergenic_res.mark == i, 'type'] = 'Transcribed'
intergenic_res['mark_mean'] = np.nan
for m in set(intergenic_res.mark):
ind = intergenic_res[intergenic_res.mark == m].index
intergenic_res.ix[ind, 'mark_mean'] = intergenic_res.ix[ind, 'neg_log_pvalue'].mean()
intergenic_res.sort_values(by=['type', 'mark_mean'], inplace=True, ascending=[True, False])
data['CNV_7_143951166_143953316'] = data.CNV_7_143951166_143953316.astype(int)
sns.boxplot(x='CNV_7_143951166_143953316', y='exp', hue='Gene', data=data, width=0.5,
fliersize=0, linewidth=0.5);
```
For the purposes of plotting I'll group 7 and 8 and call it 7+. I have a feeling the
8 calls are really seven. I also think the one calls are really 2 but I'll leave it.
```
data_f = data.copy(deep=True)
data_f.ix[data.CNV_7_143951166_143953316 == 8, 'CNV_7_143951166_143953316'] = 7
fig = plt.figure(figsize=(4.5, 8), dpi=300)
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
ax.text(0, 1, 'Figure 4',
size=16, va='top')
ciepy.clean_axis(ax)
ax.set_xticks([])
ax.set_yticks([])
gs.tight_layout(fig, rect=[0, 0.92, 1, 1])
# SNV/indel lead vs. CNV lead effect size
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
cnv_abs_beta_pdf.plot(label='CNV', ax=ax, linestyle='--')
snv_abs_beta_pdf.plot(label='Not CNV', ax=ax)
for t in ax.get_xticklabels() + ax.get_yticklabels():
t.set_fontsize(8)
ax.set_xlabel('$\\left|\\beta\\right|$', fontsize=8)
ax.set_ylabel('Density', fontsize=8)
ax.legend(fontsize=7, frameon=True, fancybox=True)
gs.tight_layout(fig, rect=[0, 0.77, 0.52, 0.94])
# Lead variant CNV not overlapping gene
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
g = cnv_lead_vars.ix[cnv_lead_vars.cnv_overlaps_gene_cons, 'gene_id']
t = cnv_lead_vars[cnv_lead_vars.gene_id.apply(lambda x: x not in g.values)]
bins = np.arange(-3, 3.1, 0.1)
t.drop_duplicates('gene_id').beta.hist(bins=bins, histtype='stepfilled', lw=0)
p = stats.binom_test((t.drop_duplicates('gene_id').beta > 0).value_counts())
print('{:,} lead intergenic CNV eGenes.'.format(t.drop_duplicates('gene_id').shape[0]))
print('Effect sizes for intergenic lead CNVs are biased '
'(p={:.3e}, binomial test).'.format(p))
for t in ax.get_xticklabels() + ax.get_yticklabels():
t.set_fontsize(8)
ax.set_xlabel('$\\beta$', fontsize=8)
ax.set_ylabel('Number of\nlead variants', fontsize=8)
ax.set_xlim(-3, 3)
gs.tight_layout(fig, rect=[0.48, 0.77, 1, 0.94])
# Functional annotation enrichment
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
sns.stripplot(x='neg_log_pvalue', y='mark', data=intergenic_res, jitter=0.3,
ax=ax, orient='h', size=4.5)
for t in ax.get_xticklabels():
t.set_fontsize(8)
ax.set_xlabel('$-\\log_{10}$ enrichment $p$-value', fontsize=8)
xmin,xmax = ax.get_xlim()
ax.set_xlim(-0.1, xmax)
ymin,ymax = ax.get_ylim()
ax.vlines(-np.log10(0.05), ymin, ymax, linestyle='--', color='grey',
linewidth=1)
ax.text(-np.log10(0.05) + 0.1, ymax, '$p=0.05$', ha='left', va='top', fontsize=8)
ax.axhspan(-0.5, 4.5, facecolor='blue', alpha=0.2, label='Active', lw=0)
ax.axhspan(4.5, 6.5, facecolor='red', alpha=0.2, label='Repressed', lw=0)
ax.axhspan(6.5, 7.5, facecolor='grey', alpha=0.2, label='Transcribed', lw=0)
ax.legend(frameon=True, fancybox=True, loc=[0.74, 0.17], fontsize=7)
ax.set_ylabel('')
for t in ax.get_xticklabels() + ax.get_yticklabels():
t.set_fontsize(8)
gs.tight_layout(fig, rect=[0, 0.57, 1, 0.79])
# mCNV eQTL gene expression
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
sns.boxplot(x='CNV_7_143951166_143953316', y='exp', hue='Gene', data=data_f,
width=0.75, fliersize=0, linewidth=0.5)
ax.set_xticklabels([str(x) for x in range(1,7)] + ['7+'])
for t in ax.get_xticklabels() + ax.get_yticklabels():
t.set_fontsize(8)
ax.set_ylabel('$\log$ TPM $z$-score', fontsize=8)
ax.set_xlabel('Diploid copy number', fontsize=8)
ax.legend(fontsize=7, loc='upper left', bbox_to_anchor=(1, 1))
gs.tight_layout(fig, rect=[0, 0.4, 0.8, 0.6])
# mCNV legend
gs = gridspec.GridSpec(1, 1)
ax = fig.add_subplot(gs[0, 0])
ciepy.clean_axis(ax)
rects = []
labels = []
for k in legend_colors.index:
labels.append(k)
rects.append(plt.Rectangle((0, 0), 0, 0, fc=legend_colors[k]))
lgd = ax.legend(rects, labels, loc='center', prop={'size':7}, ncol=3)
for p in lgd.get_patches():
p.set_linewidth(0)
gs.tight_layout(fig, rect=[0, 0, 1, 0.05])
t = fig.text(0.005, 0.915, 'A', weight='bold',
size=12)
t = fig.text(0.5, 0.915, 'B', weight='bold',
size=12)
t = fig.text(0.005, 0.77, 'C', weight='bold',
size=12)
t = fig.text(0.005, 0.58, 'D', weight='bold',
size=12)
t = fig.text(0.005, 0.4, 'E', weight='bold',
size=12)
plt.savefig(os.path.join(outdir, 'cnv_examples_skeleton.pdf'))
%%R
suppressPackageStartupMessages(library(Gviz))
suppressPackageStartupMessages(library(GenomicFeatures))
chrom = 'chr7'
start = 143851293
end = 144061217
fontsize = 6
cnv_color = '#000000'
cepbp_color = "#20B2AA"
dnase_color = "#663399"
cnvs = os.path.join(ciepy.root, 'output', 'cnv_processing', 'gs_cnvs.bed')
%%R -i data,chrom,start,end,fontsize,cepbp_color,dnase_color,cnv_color,cnvs
cnvTrack <- AnnotationTrack(
range=cnvs,
genome="hg19",
chromosome=chrom,
start=start,
end=end,
collapse=FALSE,
stacking="dense",
fontsize=fontsize,
name="CNVs",
fontcolor.legend='black',
col.axis='black',
col.title='black',
background.title='transparent',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1,
lwd=0,
fontface=1,
fontface.title=1,
rotation.title=0
)
ideoTrack <- IdeogramTrack(
genome="hg19",
fontsize=fontsize,
fontsize.legend=fontsize,
fontcolor='black',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1
)
gtrack <- GenomeAxisTrack(
col="black",
cex=1,
fontsize=8,
col.id="black",
fontcolor="black",
fontface=1,
fontface.group=1,
lwd=1,
)
biomTrack <- BiomartGeneRegionTrack(
genome="hg19",
chromosome=chrom,
start=start,
end=end,
name="",
fontsize=fontsize,
collapseTranscripts='meta',
fontcolor.legend='black',
col.axis='black',
col.title='black',
fontcolor.legend="black",
background.title='transparent',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1,
geneSymbols=TRUE,
cex.group=1,
fontcolor.group="black",
fontface.group=1,
fontface.title=1,
alpha.title=1,
lwd=0.8,
)
hmmTrack <- UcscTrack(
track="Broad ChromHMM",
table="wgEncodeBroadHmmH1hescHMM",
genome="hg19",
chromosome=chrom,
from=start,
to=end,
trackType="AnnotationTrack",
shape="box",
start="chromStart",
end="chromEnd",
feature="itemRgb",
id="name",
collapse=FALSE,
stacking="dense",
fontsize=fontsize,
name="chromHMM",
fontcolor.legend='black',
col.axis='black',
col.title='black',
background.title='transparent',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1,
lwd=0,
fontface=1,
fontface.title=1,
rotation.title=0
)
feat <- unique(feature(hmmTrack))
featCol <- setNames(as.list(rgb(t(sapply(strsplit(feat, ","),
as.numeric)), maxColorValue=255)), feat)
displayPars(hmmTrack) <- featCol
cebpbTrack <- UcscTrack(
track="Uniform TFBS",
table="wgEncodeAwgTfbsSydhH1hescCebpbIggrabUniPk",
genome="hg19",
chromosome=chrom,
from=start,
to=end,
trackType="AnnotationTrack",
shape="box",
start="chromStart",
end="chromEnd",
feature="itemRgb",
id="name",
collapse=FALSE,
stacking="dense",
fontsize=fontsize,
name="CEBPB",
fontcolor.legend='black',
col.axis='black',
col.title='black',
background.title='transparent',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1,
lwd=0,
fontface=1,
fontface.title=1,
rotation.title=0
)
dnaseTrack <- UcscTrack(
track="Uniform DNaseI HS",
table="wgEncodeAwgDnaseUwdukeH1hescUniPk",
genome="hg19",
chromosome=chrom,
from=start,
to=end,
trackType="AnnotationTrack",
shape="box",
start="chromStart",
end="chromEnd",
feature="itemRgb",
id="name",
collapse=FALSE,
stacking="dense",
fontsize=fontsize,
name="DHS",
fontcolor.legend='black',
col.axis='black',
col.title='black',
background.title='transparent',
cex=1,
cex.id=1,
cex.axis=1,
cex.title=1,
fontface=1,
fontface.title=1,
lwd=0,
fontface=1,
fontface.title=1,
rotation.title=0
)
cnvTrack = setPar(cnvTrack, "fill", cnv_color)
cebpbTrack = setPar(cebpbTrack, "fill", cepbp_color)
dnaseTrack = setPar(dnaseTrack, "fill", dnase_color)
fn = os.path.join(outdir, 'CNV_7_143951166_143953316_region.pdf')
%%R -i fn,chrom,start,end
pdf(fn, 4.5, 3)
plotTracks(c(gtrack, biomTrack, cnvTrack, cebpbTrack, dnaseTrack, hmmTrack), chromosome=chrom,
from=start, to=end, col.title="black", sizes=c(0.22, 1, 0.12, 0.12, 0.12, 0.12))
dev.off()
gene_info = pd.read_table(cpy.gencode_gene_info, index_col=0)
gene_info[gene_info.gene_name == 'CTAGE15']
```
## Presentation
```
fig,ax = plt.subplots(1, 1)
fs = 14
for i,g in enumerate(sorted(list(set(data.Gene)))):
t = data[data.Gene == g]
sns.regplot(x='CNV_7_143951166_143953316', y='exp', data=t,
x_jitter=0.2, fit_reg=False, ax=ax, label=g,
color=sns.color_palette('husl', 7)[i], scatter_kws={'alpha':0.35, 's':40})
lgd = ax.legend(fontsize=fs, loc='upper left', bbox_to_anchor=(1, 1))
ax.set_ylabel('$\log$ TPM $z$-score', fontsize=fs)
ax.set_xlabel('Diploid copy number', fontsize=fs)
for t in ax.get_xticklabels() + ax.get_yticklabels():
t.set_fontsize(fs)
fig.tight_layout()
fig.savefig(os.path.join(outdir, 'CNV_7_143951166_143953316_reg.pdf'), bbox_extra_artists=(lgd,), bbox_inches='tight')
```
| github_jupyter |
# Curve Fitting
## Objective and Prerequisites
Try this Jupyter Notebook Modeling Example to learn how you can fit a function to a set of observations. We will formulate this regression problem as a linear programming problem using the Gurobi Python API and then solve it with the Gurobi Optimizer.
This model is example 11 from the fifth edition of Model Building in Mathematical Programming, by H. Paul Williams on pages 266 and 319-320.
This modeling example is at the beginner level, where we assume that you know Python and that you have some knowledge about building mathematical optimization models. The reader should also consult the
[documentation](https://www.gurobi.com/resources/?category-filter=documentation)
of the Gurobi Python API.
**Download the Repository** <br />
You can download the repository containing this and other examples by clicking [here](https://github.com/Gurobi/modeling-examples/archive/master.zip).
## Model Formulation
### Sets and Indices
$i \in \text{Observations}=\{1, .. ,n\}$.
### Parameters
$x_{i} \in \mathbb{R}$: Independent variable value at observation $i$.
$y_{i} \in \mathbb{R}$: Dependent variable value at observation $i$.
### Decision Variables
$a \in \mathbb{R}$: Value of the constant term in the function that explains the values of $y$ in terms of the values of $x$.
$b \in \mathbb{R}$: Coefficient of the linear term in the function that explains the values of $y$ in terms of the values of $x$.
$u_{i} \in \mathbb{R}^+$: Positive deviation of the proposed function of x with respect to the value of y at observation $i$.
$v_{i} \in \mathbb{R}^+$: Negative deviation of the proposed function of x with respect to the value of y at observation $i$.
$z$: Value of the maximum deviation.
We model the problem for the first goal:
* Fit a line $y=a+bx$ to the given data set in order to minimize the sum of absolute deviations of each observed value of $y$ from the value predicted by the linear relationship.
### Constraints Problem 1
**Deviation**: Each pair of corresponding data values $(x_{i},y_{i})$ gives rise to the following constraint.
\begin{equation}
bx_{i} + a + u_{i} - v_{i} = y_{i} \quad \forall i \in \text{Observations}
\end{equation}
Where $x_{i}$ and $y_{i}$ are the given values in the set of observations, $b$, $a$, $u_{i}$ and $v_{i}$ are variables.
The positive deviation $u_{i}$ and the negative deviation $v_{i}$ give the amounts by which the values of $y_{i}$ proposed by the linear expression differ from the observed values.
### Objective Function Problem 1
**Total deviation**: The objective is to minimize the total positive and negative deviations.
\begin{equation}
\text{Minimize} \quad \sum_{i \in \text{Observations}} (u_{i} + v_{i})
\end{equation}
We now provide a model formulation for the second goal:
* Fit a line $y=a+bx$ to the given data set in order to minimize the maximum deviation of all the observed values of $y$ from the value predicted by the linear relationship.
For this new formulation, in addition to the "Deviation constraints", we need to include the following constraints.
### Constraints Problem 2
**Maximum deviation**: The following constraints ensure that the decision variable $z$ takes the value of the maximum deviation.
\begin{equation}
z \geq u_{i} \quad \forall i \in \text{Observations}
\end{equation}
\begin{equation}
z \geq v_{i} \quad \forall i \in \text{Observations}
\end{equation}
### Objective Function Problem 2
**Minimum/Maximum deviation**: The objective is to minimize the maximum deviation.
\begin{equation}
\text{Minimize} \quad z
\end{equation}
## Python Implementation
We import the Gurobi Python Module.
```
%pip install gurobipy
import gurobipy as gp
from gurobipy import GRB
# tested with Python 3.7.0 & Gurobi 9.1.0
```
## Input data
We define the corresponding values for $x$ and $y$ in the set of observations.
```
# Sample data: values of independent variable x and dependent variable y
observations, x, y = gp.multidict({
('1'): [0,1],
('2'): [0.5,0.9],
('3'): [1,0.7],
('4'): [1.5,1.5],
('5'): [1.9,2],
('6'): [2.5,2.4],
('7'): [3,3.2],
('8'): [3.5,2],
('9'): [4,2.7],
('10'): [4.5,3.5],
('11'): [5,1],
('12'): [5.5,4],
('13'): [6,3.6],
('14'): [6.6,2.7],
('15'): [7,5.7],
('16'): [7.6,4.6],
('17'): [8.5,6],
('18'): [9,6.8],
('19'): [10,7.3]
})
```
## Model Deployment
We create a model and the variables. The variables of the model are the constant term and coefficient of the linear term of the function f(x), the positive and negative deviations, and the maximum deviation.
```
model = gp.Model('CurveFitting')
# Constant term of the function f(x). This is a free continuous variable that can take positive and negative values.
a = model.addVar(lb=-GRB.INFINITY, ub=GRB.INFINITY, vtype=GRB.CONTINUOUS, name="a")
# Coefficient of the linear term of the function f(x). This is a free continuous variable that can take positive
# and negative values.
b = model.addVar(lb=-GRB.INFINITY, ub=GRB.INFINITY, vtype=GRB.CONTINUOUS, name="b")
# Non-negative continuous variables that capture the positive deviations
u = model.addVars(observations, vtype=GRB.CONTINUOUS, name="u")
# Non-negative continuous variables that capture the negative deviations
v = model.addVars(observations, vtype=GRB.CONTINUOUS, name="v")
# Non-negative continuous variables that capture the value of the maximum deviation
z = model.addVar(vtype=GRB.CONTINUOUS, name="z")
```
Each pair of corresponding data values $x_{i}$ and $y_{i}$ gives rise to a constraint.
```
# Deviation constraints
deviations = model.addConstrs( (b*x[i] + a + u[i] - v[i] == y[i] for i in observations), name='deviations')
```
The objective function of problem 1 is to minimize the total positive and negative deviations.
```
# Objective function of problem 1
model.setObjective(u.sum('*') + v.sum('*'))
# Verify model formulation
model.write('CurveFitting.lp')
# Run optimization engine
model.optimize()
# Output report
print("\n\n_________________________________________________________________________________")
print(f"The best straight line that minimizes the absolute value of the deviations is:")
print("_________________________________________________________________________________")
print(f"y = {b.x:.4f}x + ({a.x:.4f})")
```
For Problem 2, it is necessary to introduce another variable $z$ to capture the value of the maximum deviations
```
# Maximum deviation constraints
maxPositive_deviation = model.addConstrs( (z >= u[i] for i in observations), name='maxPositive_deviation')
maxNegative_deviation = model.addConstrs( (z >= v[i] for i in observations), name='maxNegative_deviation')
```
The objective function for Problem 2 is to minimize the maximum deviation.
```
# Objective function for Problem 2
model.setObjective(z)
# Run optimization engine
model.optimize()
# Output report
print("\n\n_________________________________________________________________________________")
print(f"The best straight line that minimizes the maximum deviation is:")
print("_________________________________________________________________________________")
print(f"y = {b.x:.4f}x + ({a.x:.4f})")
```
---
## References
H. Paul Williams, Model Building in Mathematical Programming, fifth edition.
Copyright © 2020 Gurobi Optimization, LLC
| github_jupyter |
# 1-4.3 Intro Python Practice
## Conditionals
<font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
- **control code flow with `if`... `else` conditional logic**
- using Boolean string methods (`.isupper(), .isalpha(), startswith()...`)
- using comparision (`>, <, >=, <=, ==, !=`)
- using Strings in comparisons
## `if else`
```
# [ ] input avariable: age as digit and cast to int
# if age greater than or equal to 12 then print message on age in 10 years
# or else print message "It is good to be" age
age = int(input("enter age: "))
if age >= 12:
print("age in 10 years is", age + 10)
else:
print("It is good to be", age)
# [ ] input a number
# - if number is NOT a digit cast to int # WHY?
# - print number "greater than 100 is" True/False
number = input("input a number: ")
if number.isdigit() == False:
number = int(number)
print(str(number), "greater than 100 is", str(int(number) > 100))
```
### Guessing a letter A-Z
**check_guess()** takes 2 string arguments: **letter and guess** (both expect single alphabetical character)
- if guess is not an alpha character print invalid and return False
- test and print if guess is "high" or "low" and return False
- test and print if guess is "correct" and return True
```
# [ ] create check_guess()
# call with test
def check_guess(letter, guess):
if guess.isalpha() == False:
print('invalid')
return False
elif guess > letter:
print('your guess is high')
return False
elif guess < letter:
print('your guess is low')
return False
else:
print('correct')
return True
check_guess('c', 'b')
check_guess('c', 'd')
check_guess('c', 'c')
check_guess('c', '1')
# [ ] call check_guess with user input
letter = input("enter a letter: ")
guess = input("input a guess: ")
check_guess(letter, guess)
```
### Letter Guess
**create letter_guess() function that gives user 3 guesses**
- takes a letter character argument for the answer letter
- gets user input for letter guess
- calls check_guess() with answer and guess
- End letter_guess if
- check_guess() equals True, return True
- or after 3 failed attempts, return False
```
# [ ] create letter_guess() function, call the function to test
def letter_guess(letter):
guess = input("enter a letter guess: ")
if check_guess(letter, guess) == True:
return True
else:
print("You have 2 guess attempts. Try again!")
pass
guess = input("enter a letter guess: ")
if check_guess(letter, guess) == True:
return True
else:
print("You have 1 guess attempts. Try again!")
pass
guess = input("enter a letter guess: ")
if check_guess(letter, guess) == True:
return True
else:
print("You have 0 guess attempts. You fail.")
return False
letter = 'c'
letter_guess(letter)
```
### Pet Conversation
**ask the user for a sentence about a pet and then reply**
- get user input in variable: about_pet
- using a series of **if** statements respond with appropiet conversation
- check if "dog" is in the string about_pet (sample reply "Ah, a dog")
- check if "cat" is in the string about_pet
- check if 1 or more animal is in string about_pet
- no need for **else**'s
- finish with thanking for the story
```
# [ ] complete pet conversation
about_pet = input("Tell me about your pet: ")
if "dog" in about_pet:
print("Ah, a dog!")
if "cat" in about_pet:
print("Ah, a cat!")
if "parrot" in about_pet:
print("Ah, a parrot!")
```
[Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| github_jupyter |
# Multi-Qubit Noisy Simulator
*Copyright (c) 2021 Institute for Quantum Computing, Baidu Inc. All Rights Reserved.*
## Outline
This tutorial will introduce how to use multi-qubit simulator at the pulse level. The outline is as follows:
- Introduction
- Preparation
- Use multi-qubit noisy simulator at the gate level
- Use multi-qubit noisy simulator at the pulse level
- Modeling the system
- Rabi oscillation
- Cross-Resonance effect
- ZZ crosstalk characterization through Ramsey experiment
- Summary
## Introduction
Simulating time evolution of the qubits at the pulse level gives us more insight into the physics of quantum gates and the effects of noise. For superconducting quantum circuits, the transmon qubits are controlled by applying microwave pulses and magnetic flux. However, the performance of quantum gates is often suppressed by various factors: the decoherence of the qubit due to its interaction with the environment, the unwanted crosstalk effect, and leakage into the higher levels of the transmon.
The multi-qubit noisy simulator provided by Quanlse allows us to simulate quantum operations on a noisy quantum device consisting of multiple transmon qubits to understand the physics behind quantum computing better. Several main types of noise are included in our noisy simulator: decoherence noise, amplitude noise, and crosstalk noise. We will focus on several common applications in superconducting quantum computing based on this noisy simulator: Rabi oscillation, Cross-Resonance effect, and characterizing ZZ crosstalk through a Ramsey experiment.
## Preparation
After you have successfully installed Quanlse, you could run the Quanlse program below following this tutorial. To run this particular tutorial, you would need to import the following packages from Quanlse and other commonly-used Python libraries:
```
from Quanlse.remoteOptimizer import remoteOptimize1Qubit as optimize1q
from Quanlse.remoteSimulator import remoteSimulatorRunHamiltonian as runHamiltonian
from Quanlse.Simulator import PulseModel
from Quanlse.Simulator.PulseSim3Q import pulseSim3Q
from Quanlse.QWaveform import QJob, QJobList
from Quanlse.QOperator import driveZ
from Quanlse.QWaveform import square
from Quanlse.Utils.Functions import basis, tensor, expect, dagger, computationalBasisList
from Quanlse.Utils.Plot import plotBarGraph
from Quanlse.QOperation.FixedGate import H, X, CNOT
from Quanlse.Scheduler.Superconduct.PipelineCenterAligned import centerAligned
from math import pi
from scipy.optimize import curve_fit
import numpy as np
import matplotlib.pyplot as plt
```
To use Quanlse Cloud Service, we need to acquire a token to get access to the cloud.
```
# Import Define class and set the token for cloud service
# Please visit http://quantum-hub.baidu.com
from Quanlse import Define
Define.hubToken = ""
```
## Use multi-qubit noisy simulator at the gate level
We can also run the simulator at the gate level. In this section, we use a predefined `PulseModel()` instance with the default configuration. To create a 3-qubit physics model, we first instantiate the `PulseModel()` object by calling `pulseSim3Q()`.
```
model = pulseSim3Q(frameMode='lab', dt=0.01)
model.savePulse = False
model.pipeline.addPipelineJob(centerAligned)
```
To define a circuit of creating GHZ state by Quanlse scheduler, we add the gates to the model by `gate(model.Q[index])`.
```
# Hadamard gate
H(model.Q[0])
# CNOT
CNOT(model.Q[0], model.Q[1])
CNOT(model.Q[1], model.Q[2])
```
The pulse sequence of the quantum circuit defined above is generated by calling method `model.schedule`.
```
scheJob = model.schedule()
```
Define the initial state $|000\rangle$ and run the simulation. Then, plot the probability distribution of different outcome.
```
# Run the simulation
res = model.simulate(job=scheJob)
# Plot the result
popList = [abs(item ** 2) for item in res[0]['state'].T[0]]
basisList = computationalBasisList(3, 3)
plotBarGraph(basisList, popList, "Result", "Outcome", "Population")
```
As it shows, the measurement result included unexpected values due to the noise. The simulation under decoherence can be done by setting the parameter `isOpen=True` in module `runHamiltonian()`, which takes some time to run, and we will obtain the density matrix after the simulation. For more details about how decoherence noise affects superconducting quantum computing, please refer to [Single-Qubit Noisy Simulator](https://quanlse.baidu.com/#/doc/tutorial-single-qubit-noisy-simulator).
## Use multi-qubit noisy simulator at the pulse level
Multi-qubit noisy simulator supports the quantum control simulation at the pulse level - get the system's final state by defining the waveform of the pulse and other related parameters. It allows us to simulate the quantum control of the superconducting hardware at the lower level. Here, we simulate some of the common operations used in real experiments.
### Modeling the system
Usually, we use Duffing oscillator to describe the physics model of the superconducting circuits. In lab frame, the system Hamiltonian of a three-qubit system with coupling between qubits $q_0$ and $q_1$, $q_1$ and $q_2$ reads:
$$
\hat{H} = \sum_{i=0}^2 \omega_i \hat{a}^\dagger_i \hat{a}_i + \sum_{i=0}^2 \frac{\alpha_i}{2} \hat{a}^\dagger_i \hat{a}^\dagger_i \hat{a}_i \hat{a}_i + g_{01} (\hat{a}^\dagger_0 \hat{a}_1 + \hat{a}_0 \hat{a}^\dagger_1) + g_{12} (\hat{a}^\dagger_1 \hat{a}_2 + \hat{a}_1 \hat{a}^\dagger_2),
$$
where $\omega_i$ and $\alpha_i$ are the qubit frequency and anharmonicity of qubit $q_i$ respectively; $g_{i, j}$ is the coupling strength between qubit $q_i$ and qubit $q_j$; $a_i$, $a^\dagger_i$ denote the annihilation operator and creation operator of qubit $q_i$.
In this tutorial, we use a three-qubit system as an example. We first define the parameters of the hardware.
```
qubitNum = 3 # The number of qubits
level = 3 # The energy level for each qubit
anharm = -0.33 * 2 * pi # The anharmonicity of the qubit, in 2 pi GHz
wq0 = 4.914 * 2 * pi # The frequency for qubit 0, in 2 pi GHz
wq1 = 5.100 * 2 * pi # The frequency for qubit 1, in 2 pi GHz
wq2 = 5.200 * 2 * pi # The frequency for qubit 2, in 2 pi GHz
g01 = 0.0038 * 2 * pi # The coupling strength of the interaction between qubit 0 and qubit 1, in 2 pi GHz
g12 = 0.0020 * 2 * pi # The coupling strength of the interaction between qubit 1 and qubit 2, in 2 pi GHz
dt = 1. # The sampling time of AWG
# T1 relaxation time for qubit 0, qubit 1, and qubit 2, in nanoseconds
t01 = 1000
t11 = 1120
t21 = 1300
# T2 dephasing time for qubit 0, qubit 1, and qubit 2, in nanoseconds
t02 = 500
t12 = 450
t22 = 600
# The random amplitude distortion
ampNoise = 0.02
```
The physics model is created by instantiating an object of class `PulseModel`. The types of noise include $T_1$-relaxation noise, $T_2$-dephasing, and distortion of amplitudes.
```
qubitFreq = {0: wq0, 1: wq1, 2: wq2} # Qubit frequency for each qubit
qubitAnharm = {0: anharm, 1: anharm, 2: anharm} # Qubit anharmonicity for each qubit
qubitT1 = {0: t01, 1: t11, 2: t21} # Relaxation time
qubitT2 = {0: t02, 1: t12, 2: t22} # Dephasing time
couplingMap = {(0, 1): g01, (1, 2): g12} # Coupling map
# Create an instant of PulseModel
model = PulseModel(subSysNum=qubitNum,
sysLevel=level,
qubitFreq=qubitFreq,
qubitAnharm=qubitAnharm,
couplingMap=couplingMap,
T1=qubitT1,
T2=qubitT2,
dt=dt,
ampSigma=ampNoise)
```
We have constructed a noisy simulator including three superconducting qubits with three types of noises. The next step is to create a `QHamiltonian` object by calling method `createQHamiltonian()`.
```
ham = model.createQHamiltonian()
```
### Cross-Resonance effect
The all-microwave control is one of the strategies to realize quantum control on superconducting circuits. In this strategy, two-qubit operations harness the cross-resonance effect of two weakly-coupled qubits. This is done by driving the control qubit with the frequency of the weakly-coupled target qubit. Ideally, the desired $\hat{\sigma}_z \otimes \hat{\sigma}_x$ interaction between the control and target qubit is dominating interaction \[1\]. For more details about CR gate, please refer to [Cross-Resonance Gate](https://quanlse.baidu.com/#/doc/tutorial-cr).
In our simulation, we again drive qubit $q_0$ (control qubit) by various amplitudes (with the drive frequency of the qubit $q_1$). This can be done by `addWaveRot(index, waves, detuning)` where `index` is the index of qubit acted upon; `waves` is the waveform of the pulse; and `detuning` $\Delta$ is the frequency difference ($\Delta = \omega_q - \omega_d$, where $\omega_q$ is the qubit frequency and $\omega_d$ the drive frequency).
Here, we vary the amplitudes of the pulse and record the population of $|1\rangle$ for each qubit. In this example, $q_1$ is the control qubit driven by the pulse with the frequency of target qubit $q_0$.
```
dCoef = 0.03 * (2 * pi) # The drive strength of the pulse
ampCR = np.linspace(0, 0.5, 40) # The amplitudes in arbitrary unit
amps = ampCR * dCoef
detuning = wq1 - wq0 # The detuning of the pulse
# jobList = QJobList(subSysNum=qubitNum, sysLevel=level, dt=dt, title='cr')
jobList = ham.createJobList()
# Fix the gate time
tg = 950
# Append each job to the jobList
for amp in amps:
job = ham.createJob()
job.addWaveRot(1, waves=square(tg, amp), t0=0., detuning=detuning) # Apply pulse at qubit 1
job = model.getSimJob(job)
jobList.addJob(jobs=job)
```
Run the simulation with initial state $|\psi\rangle = |010\rangle$, and the control qubit $q_1$ starts at the excited state.
```
# Define the initial state of |010>
psi0 = tensor(basis(level, 0), basis(level, 1), basis(level, 0))
# Run the simulation
result = runHamiltonian(ham=ham, state0=psi0, jobList=jobList)
```
Define the projector of the first excited state for qubit $q_i$ and initialize the list of expected values.
```
prj01 = tensor(basis(3, 1) @ dagger(basis(3,1)), np.identity(level), np.identity(level)) # The projector of qubit 0
prj11 = tensor(np.identity(level), basis(3, 1) @ dagger(basis(3,1)), np.identity(level)) # The projector of qubit 1
prj21 = tensor(np.identity(level), np.identity(level), basis(3, 1) @ dagger(basis(3,1))) # The projector of qubit 1
```
Compute the expected values of the projector of each qubit, and plot them with respect to the different amplitudes.
```
# Initialize the list of expected values
num0List = []
num1List = []
num2List = []
for res in result.result:
state = res['state'] # The final state of each job
num0Expect = expect(prj01, state) # Compute the expected values of the projector |1><1|
num1Expect = expect(prj11, state)
num2Expect = expect(prj21, state)
num0List.append(num0Expect)
num1List.append(num1Expect)
num2List.append(num2Expect)
plt.figure(figsize=(8, 6))
# Plot the figure of CR effect
plt.plot(ampCR, num0List, label='qubit0')
plt.plot(ampCR, num1List, label='qubit1')
plt.plot(ampCR, num2List, label='qubit2')
plt.xlabel('Amplitudes (a.u.)')
plt.ylabel(r'Population of $|1\rangle$')
plt.title('Cross-Resonance effect')
plt.legend()
plt.show()
```
As it shows, the projector $|1\rangle \langle 1|$'s expected value of target qubit $q_0$ changes over the amplitude, while the control qubit $q_1$ is in the excited state when the amplitude is relatively small. Meanwhile, qubit $q_2$ is always in the ground state. It can also be seen that the increasing amplitude inevitably affects qubit $q_1$.
### ZZ crosstalk characterization by a Ramsey experiment
ZZ crosstalk is the major source of unwanted interaction between coupled qubits. It arises from the existence of states of higher energy levels. The effective Hamiltonian of two coupled qubits (directly or indirectly) in the two-qubit subspace is \[2\]:
$$
\hat{H}_{\rm eff} = \omega_{0}\frac{\hat{\sigma}_{z}^0 \otimes I_1}{2} + \omega_{1}\frac{I_0\otimes\hat{\sigma}_{z}^1}{2} + \xi \frac{\hat{\sigma}_{z}^0 \otimes \hat{\sigma}_{z}^1}{2},
$$
Where $\omega_0$, $\omega_1$ are the qubit frequencies and $\xi$ is the strength of ZZ crosstalk. $\xi$ is defined as the different of transition frequencies between $|11\rangle \leftrightarrow |10\rangle$ and $|01\rangle \leftrightarrow |00\rangle$:
$$
\xi = \left(E_{11} - E_{10}\right) - \left(E_{01} - E_{00}\right),
$$
where $E_{ij}$ is the energy level of state $|ij\rangle$. We can actually detect and measure this frequency shift-induced crosstalk by Ramsey experiment. This can be done by applying two Hadamard gates with an idle time apart \[3\].
To better illustrate the effect of ZZ crosstalk, we define a new 3-qubit model with stronger coupling strengths (6 ~ 40 MHz).
```
dt = 0.2 # The sampling time
level = 3 # The system level
qubitNum = 3 # The number of qubits
g01 = 0.0377 * (2 * pi)
g12 = 0.0060 * (2 * pi)
# Coupling map
couplingMap = {
(0, 1): g01,
(1, 2): g12
}
# Qubits frequency anharmonicity
anharm = - 0.33 * (2 * pi)
qubitAnharm = {0: anharm, 1: anharm, 2: anharm} # The anharmonicities for each qubit
# Qubit Frequency
qubitFreq = {
0: 5.5904 * (2 * pi),
1: 4.7354 * (2 * pi),
2: 4.8524 * (2 * pi)
}
```
Create the physics model by class `PulseModel()`, and create Hamiltonian `ham` by the model.
```
model = PulseModel(subSysNum=qubitNum, sysLevel=level, couplingMap=couplingMap,
qubitFreq=qubitFreq, dt=dt, qubitAnharm=qubitAnharm)
ham = model.createQHamiltonian()
```
Generate the pulses of the gates $H$ and $X$ on different qubits.
```
# Define function to generate the QJob for gate of specified qubit
def generateGate(gate, index):
job1q, _ = optimize1q(ham=ham.subSystem(index), uGoal=gate.getMatrix(), targetInfid=1e-5)
job3q = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
waves = job1q.waves
ops = job1q.ctrlOperators
for key, op in ops.items():
job3q.addWave(operators=op, onSubSys=index, waves=waves[key])
return job3q
# Generate the gates needed
h0 = generateGate(H, 0) # H gate on qubit 0
h1 = generateGate(H, 1) # H gate on qubit 1
x1 = generateGate(X, 1) # X gate on qubit 1
x2 = generateGate(X, 2) # X gate on qubit 2
maxTime = 500 # The delayed time in Ramsey experiment, in nanosecond.
freq = 3 / maxTime # Detuning.
# Generate job for delayed time
def generateIdle(tg, index):
jobIdle = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
jobIdle.appendWave(operators=driveZ, onSubSys=index, waves=square(tg, 2 * pi * freq))
return jobIdle
```
Define two different `jobList` objects - one begins with $|00\rangle$ and the other $|01\rangle$ by applying a $X$ gate on qubit $q_1$. Then perform Ramsey experiment on qubit $q_0$.
```
# jobList with initial state |00>
jobListGrd = QJobList(subSysNum=qubitNum, sysLevel=level, dt=dt)
# jobList with initial state |01> (by applying X gate)
jobListExd = QJobList(subSysNum=qubitNum, sysLevel=level, dt=dt)
# Define the delayed time
tgList = np.linspace(0, maxTime, 50)
# Define jobList with initial state |00>
for tg in tgList:
job = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
job += h0
job += generateIdle(tg, 0)
job += h0
jobListGrd.addJob(job)
# Define jobList with initial state |01>
for tg in tgList:
job = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
job += x1
job += h0
job += generateIdle(tg, 0)
job += h0
jobListExd.addJob(job)
# Run the simulation
stateInit = tensor(basis(level, 0), basis(level, 0), basis(level, 0))
resultGrd = runHamiltonian(ham, state0=stateInit, jobList=jobListGrd)
resultExd = runHamiltonian(ham, state0=stateInit, jobList=jobListExd)
```
Plot the population of excited state of qubit $q_0$ versus delayed time.
```
num0List = []
num1List = []
# projector |1><1| of qubit 0
prj1 = tensor(basis(level, 1) @ dagger(basis(level, 1)), np.identity(9))
# append the result to the list
for res0, res1 in zip(resultGrd, resultExd):
psi0, psi1 = res0['state'], res1['state']
rho0, rho1 = psi0 @ dagger(psi0), psi1 @ dagger(psi1)
num0List.append(expect(prj1, rho0))
num1List.append(expect(prj1, rho1))
# plot the result
plt.figure(figsize=(8, 6))
plt.plot(tgList, num0List, '.b', label=r'$|00\rangle$')
plt.plot(tgList, num1List, '.r', label=r'$|01\rangle$')
plt.xlabel('Delayed time (ns)')
plt.ylabel('Population of excited state of qubit 0')
plt.legend()
plt.show()
```
The strength of ZZ crosstalk $\xi$ can be estimated by computing the frequency difference in the Ramsey oscillation. Therefore, we use the cosine function to fit the result acquired by simulation to compute the frequencies $f_1$, $f_2$. The strength is given by $\xi / \left( 2\pi \right) = |f_1 - f_2|$.
```
# Define the fitting curve
def fit(x, omega, theta):
return - 0.5 * np.cos(omega * x + theta) + 0.5
# Fit the curve
para1Fit, _ = curve_fit(fit, tgList, num0List, [2.1 * pi * freq, 0])
para2Fit, _ = curve_fit(fit, tgList, num1List, [1. * pi * freq, 0])
step = 0.01
y1Fit = [fit(x, para1Fit[0], para1Fit[1]) for x in np.arange(tgList[0], tgList[-1], step)]
y2Fit = [fit(x, para2Fit[0], para2Fit[1]) for x in np.arange(tgList[0], tgList[-1], step)]
# Plot the curve
plt.figure(figsize=(8, 6))
plt.plot(np.arange(tgList[0], tgList[-1], step), y1Fit)
plt.plot(np.arange(tgList[0], tgList[-1], step), y2Fit)
plt.plot(tgList, num0List, '.b', label=r'$|00\rangle$')
plt.plot(tgList, num1List, '.r', label=r'$|01\rangle$')
plt.xlabel('Delayed time (ns)')
plt.ylabel('Population of excited state of qubit 0')
plt.title('Ramsey on Q0')
plt.legend()
plt.show()
# Calculate the crosstalk strength
xiEst = abs(para1Fit[0] - para2Fit[0])
print(f'Coupling strength: {g01 * 1e3 / (2 * pi)} MHz')
print(f'ZZ crosstalk strength: {xiEst * 1e3 / (2 * pi)} MHz')
```
Due to the strong coupling strength between qubits $𝑞_0$ and $𝑞_1$, it can be observed that the frequency difference is relatively large, that is, the 𝑍𝑍 crosstalk is relatively large.
We can repeat the same experiment to calculate ZZ crosstalk strength $\xi$ between qubit $q_1$ and qubit $q_2$ with smaller coupling strength.
```
# jobList with initial state |00>
jobListGrd = QJobList(subSysNum=qubitNum, sysLevel=level, dt=dt)
# jobList with initial state |01> (by applying X gate)
jobListExd = QJobList(subSysNum=qubitNum, sysLevel=level, dt=dt)
# Define the delayed time
tgList = np.linspace(0, maxTime, 50)
# Define jobList with initial state |00>
for tg in tgList:
job = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
job += h1
job += generateIdle(tg, 1)
job += h1
jobListGrd.addJob(job)
# Define jobList with initial state |01>
for tg in tgList:
job = QJob(subSysNum=qubitNum, sysLevel=level, dt=dt)
job += x2
job += h1
job += generateIdle(tg, 1)
job += h1
jobListExd.addJob(job)
# Run the simulation
stateInit = tensor(basis(level, 0), basis(level, 0), basis(level, 0))
resultGrd = runHamiltonian(ham, state0=stateInit, jobList=jobListGrd)
resultExd = runHamiltonian(ham, state0=stateInit, jobList=jobListExd)
num0List = []
num1List = []
# projector |1><1| of qubit 1
prj1 = tensor(np.identity(3), basis(level, 1) @ dagger(basis(level, 1)), np.identity(3))
# append the result to the list
for res0, res1 in zip(resultGrd, resultExd):
psi0, psi1 = res0['state'], res1['state']
rho0, rho1 = psi0 @ dagger(psi0), psi1 @ dagger(psi1)
num0List.append(expect(prj1, rho0))
num1List.append(expect(prj1, rho1))
# plot the result
plt.figure(figsize=(8, 6))
plt.plot(tgList, num0List, '.b', label=r'$|00\rangle$')
plt.plot(tgList, num1List, '.r', label=r'$|01\rangle$')
plt.xlabel('Delayed time (ns)')
plt.ylabel('Population of excited state of qubit 1')
plt.title('Ramsey on Q1')
plt.legend()
plt.show()
# Fit the curve
para1Fit, _ = curve_fit(fit, tgList, num0List, [2 * pi * freq / 1.2, 0.])
para2Fit, _ = curve_fit(fit, tgList, num1List, [2 * pi * freq / 1.1, 0.])
step = 0.01
y1Fit = [fit(x, para1Fit[0], para1Fit[1]) for x in np.arange(tgList[0], tgList[-1], step)]
y2Fit = [fit(x, para2Fit[0], para2Fit[1]) for x in np.arange(tgList[0], tgList[-1], step)]
# Plot the curve
plt.figure(figsize=(8, 6))
plt.plot(np.arange(tgList[0], tgList[-1], step), y1Fit)
plt.plot(np.arange(tgList[0], tgList[-1], step), y2Fit)
plt.plot(tgList, num0List, '.b', label=r'$|00\rangle$')
plt.plot(tgList, num1List, '.r', label=r'$|01\rangle$')
plt.xlabel('Delayed time (ns)')
plt.ylabel('Population of excited state of qubit 1')
plt.title('Ramsey on Q1')
plt.legend()
plt.show()
# Calculate the crosstalk strength
xiEst = abs(para1Fit[0] - para2Fit[0])
print(f'Coupling strength: {g12 * 1e3 / (2 * pi)} MHz')
print(f'ZZ crosstalk strength: {xiEst * 1e3 / (2 * pi)} MHz')
```
Due to the weaker coupling strength, the relatively small qubit frequency shift of $q_1$ indicates the weak ZZ crosstalk between qubit $q_1$ and $q_2$.
## Summary
After reading this tutorial on multi-qubit noisy simulator, the users could follow this link [tutorial-multi-qubit-noisy-simulator.ipynb](https://github.com/baidu/Quanlse/blob/main/Tutorial/EN/tutorial-multi-qubit-noisy-simulator.ipynb) to the GitHub page of this Jupyter Notebook document and run this program for themselves. The users are encouraged to explore other advanced research which is different from this tutorial.
## References
\[1\] [Malekakhlagh, Moein, Easwar Magesan, and David C. McKay. "First-principles analysis of cross-resonance gate operation." *Physical Review A* 102.4 (2020): 042605.](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.102.042605)
\[2\] [Magesan, Easwar, and Jay M. Gambetta. "Effective Hamiltonian models of the cross-resonance gate." *Physical Review A* 101.5 (2020): 052308.](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.101.052308)
\[3\] [Ku, Jaseung, et al. "Suppression of Unwanted ZZ Interactions in a Hybrid Two-Qubit System." *Physical review letters* 125.20 (2020): 200504.](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.125.200504)
| github_jupyter |
# 1. Introduction to Python
Python is a programming language.
It is not a mathematics-oriented language in and of itself.
It is a general-purpose language, meaning we can do pretty much what we want with it.
[](https://xkcd.com/353/)
Here is a (supershort) list of what humanity did with Python:
- Dropbox (Source: [Dropbox Blog](https://blogs.dropbox.com/tech/2018/09/how-we-rolled-out-one-of-the-largest-python-3-migrations-ever/))
- Image editing ([The GNU Image Manipulation Program](https://www.gimp.org/))
- Vector graphics ([Inkscape](https://inkscape.org/))
- 3D modeling ([Blender](https://www.blender.org/))
- Desktop publishing ([Scribus](https://www.scribus.net/))
- Web pages ([Reddit](https://www.reddit.com/), Source: [Reddit Blog](https://redditblog.com/2005/12/05/on-lisp/))
Also in economics we have a great sites like [QuantEcon](https://quantecon.org) by **Prof. Sargent** and his team and you can find many economics models on it.
# 2. How to work with the notebooks in colab/binder
Notebooks:
- notebook as a collection of cells
- two main cell types: text (markdown) and code
- click on a cell to edit the contents
- Shift+Enter to run the code within a code cell or render the text cell
- be aware of the execution order
- installing necessary modules with pip and conda
# 3. The Basics
Python alone cannot do much.
For this reason, we are almost always going to work with a package (see below).
However, it is fundamental to understand the basics.
This involves familiarizing with the _syntax_ and with the basic _data types_.
Syntax is the set of rules that govern writing code.
This includes how to write an _assignment_ (providing a variable with a value), how to call functions and how to access items in a iterable object (e.g., lists, arrays).
It also includes _code blocks,_ which execute conditionally on a given rule (e.g., `if`, `while`).
$$ \sqrt{2^x} \frac{x^3}{\sqrt{2^{3x}}}$$
```
import this
```
## 3.1 Arithmetic operations and variables
Let's start with basic calculations in Python. Entering some arithmetic operation in the code cell below (e.g. `2+2`):
<br><center> <b>Arithmetic Operators<b> <center>
<br>
| Symbol | Task Performed | Description| Example<br> (a = 10 & b = 20) |
|:----:|:---:|:---:|:---:|
| + | Addition | Adds values on either side of the operator. | a + b = 30
| - | Subtraction | Subtracts right hand operand from left hand operand. | a – b = -10
| / | division | Divides left hand operand by right hand operand | b / a = 2
| % | mod | Divides left hand operand by right hand operand and returns remainder | b % a = 0
| * | multiplication | Multiplies values on either side of the operator | a * b = 200
| // | floor division | he division of operands where the result is the quotient in<br> which the digits after the decimal point are removed. | b // a = 2
| ** | to the power of | Performs exponential (power) calculation on operators | a**b =$10^{20}$
```
# 2 + 10 * 2 sdkfhasdkjfsadf
# 2+10*2
```
Every code line in Python is interpreted as a `command`, unless it starts
with the hash/pound sign, in which case it is considered to be a `comment`:
```
# the line below will be executed
1+1
# the line below will not be executed
# 1+2
```
```
2 + 4
6 - 4
5 * 3
5 / 4
18 % 4
18 // 4
3 ** 5
5 + (4 - 3 * 2)**3 + 1
```
As with many programming languages, we are defining variables and changing their values all the time in Python.
We can create a variable simply by inventing a name and assigning a value to it, like the following.
```
a = 1
a
```
To store the results of a calculation in a `variable` we can use `=` sign.
```python
sample_number = 2 + 3
```
When working inside the notebooks, we can see the result of the last executed command
or a `variable`. For example, what do you see when you type the code below in a blank cell?
```
sample_number
```
```
# type here
sample_number = 2 + 3
sample_number
b = 5 + (4 - 3 * 2)**3 + 1
b
# note that only the last command/variable is displayed
# for example of the following two commands only the last one will be shown
2+2
2+4
2+3
b
b = 10
b
b = 10
b = 10 - 8
b = b + 10
b
```
To see the contents of the variable we can use the `print` function:
```python
print(sample_number)
```
Check that this works in the cell below:
```
# type here
print(sample_number)
print(b)
b
```
Later in the code we can re-use this variable, check what is the output of this command:
```python
sample_number + 1
```
```
b = b + 10
b
# uncomment the line below
sample_number + 1
sample_number = sample_number + 2
sample_number
```
## 3.2 Relational Operators
Relational operators are used for comparing the values. It either returns **True** or **False** according to the condition. These operators are also known as Comparison Operators.
| Symbol | Task Performed | Description|
|:----: |:---:|:---:|
|= |Assignment| Assigns values from right side operands to left side operand
|== |True, if it is equal| If the values of two operands are equal, then the condition becomes true.|
|!= |True, if not equal to| If values of two operands are not equal, then condition becomes true.|
|< |less than| If the value of left operand is less than the value of right operand,<br> then condition becomes true.
|> |greater than| If the value of left operand is greater than the value of right operand, <br>then condition becomes true.
|<= |less than or equal to| If the value of left operand is less than or equal to the value of right operand,<br> then condition becomes true.
|>= |greater than or equal to|If the value of left operand is greater than or equal to the value of right operand, <br>then condition becomes true.
```
x = 2
y = 6
print(x == y)
print(x != y)
print(x == 2)
print(x > y)
print(x > 2)
print(x >= 2)
x > 1 and y < 1
x >1 or y < 1
1 < x < 3
```
## 3.3 Type
Now we have a variable whose name is `a` and its value is `1`.
This variable has a _type_, which is the kind of value it contains.
In particular, `1` is an integer, whose type is represented in Python with the keyword `int`.
Understanding that variables can be of different types is important, because the type defines what we can do with that variable.
We can ask Python to tell us the type of a variable by using the `type()` function.
Data types are the types of variables we can create.
Python defines a few basic ones and packages (see below) provide new data types.
This programming language features _dynamic typing_, which means that we do not have to define what a variable can be.
Instead, Python infers the type of a variable when we create it.
Examples of basic data types are strings (`str`), lists (`list`), integer numbers (`int`).
<br><br>
| Types | Example |
| :--: | :--: |
| string | "Hello" , 'World' |
| integer | 1 , 2 , 3 |
| float | 1.2 , 4.6 , 112.6 |
| boolian | True , Flase |
|list | [1, 2, 3, 'python', 9, 7] |
| dictionary | {'Python': 18 , "Econ": 20}
### Text (`str`)
A basic object in Python is text.
This is formally represented with a `str` object.
Strings are denoted either with single or double quotes, and the choice between them is a matter of taste.
```
print("Hello world!")
print('python course')
```
There must always be consistency between opening a closing quotes.
This means that single quotes can only close single quotes and double quotes can only close double quotes.
```
"weird striing..?'
```
As mentioned, Python variables can also contain `string` data:
```python
sample_string = "abc"
```
```
# create a variable `sample_string` and assign it some string of your choice
sample_string = "abc"
print(sample_string)
```
Python allows performing some operations with string variables using notation similar to
arithmetic operators. For example, try this in the cell below. Is this what you would expect?
```python
sample_string*2
```
```
# type here
sample_string * 2
```
It does not matter if you put the strings in single or double quotes::
```python
sample_string = "abc" + 'def'
```
```
# uncomment the line below
sample_string + 'def'
```
### Numbers (`int` and a`float`)
We already encountered integer numbers, whose type is `int`.
Another numerical type we are going to use very often is `float`.
This essentially is a non-integer real number, although the inner workings of [floating-point numbers](https://en.wikipedia.org/wiki/Floating-point_arithmetic) are more complicated.
We can initialize a variable to be of type `float` by simply assigning a decimal number to a variable.
```
b = 1.5
type(b)
```
This works even when the digit on the right of the decimal point is non significant.
The simple fact that we typed a period in the number tells Python that we want to work with floating point numbers.
```
type(1.0)
```
Both `int` and `float` variables support conventional arithmetic operations, such as addition (`+`), subtraction (`-`), multiplication (`*`), division (`/`) and raise to power (`**`).
For example:
```
a + b
b = 1
b - 1
b ** 2
```
Other operations that might be handy are floor division (`//`) and the _mod_ operation (`%`).
The former returns the largest integer smaller than the quotient, while the latter returns the remainder of floor division.
```
16 % 4
16 // 4
```
### Iterable objects (`list`, `tuple` and `dict`)
Often we want to collect objects in arrays.
The most basic example is a vector, which is an organized array of numbers.
Python (mainly) provides three iterable objects.
#### **List**
We first look at **lists**.
These are arrays of heterogeneous objects.
They are created by collecting objects in square brackets.
A list of objects is an odered collection of items, for example `['a', 'b', 'c', 'a', 123]`. Since
lists are ordered, every item in the list has an associated index that starts with 0. This provides
access to specific elements of the list by specifying their index. For example, 'b' in the list above has index `1`.
The syntax for accessing a specific element is to provide its index in square brackets, e.g. `sample_list[1]`.
There are no restriction on the contents of the list, they can contain repeated values
(e.g. 'a' in the list above) or they can contain different data types (e.g. strings and integers in the list above).
```
z = ['text', 2, b]
z
type(z)
```
Lists can also nest.
```
['nesting!', z]
```
We can access the contents of any iterable object using square brackets.
```
z[1]
```
Note that Python starts counting from zero.
This means that `z[0]` will denote the first element of the list `z`, so that `z[1]` denotes the second element.
We can also refer to items at the end of a list, without knowing how many elements it contains.
```
z[-1]
```
Try defining the list below and accessing different elements inside the list:
```python
sample_list = ['a', 'b', 'c', 'a', 123, 0, 'X']
```
```
# type here
sample_list = ['a', 'b', 'c', 'a', 123, 0, 'X']
sample_list
sample_list[::2]
```
Sometimes we want to use sequences inside a list, e.g. all items from start until the third item.
For this we can use the slicing notation:
```
[start:stop:step_size]
```
If `start` is missing, the assumption is to start from 0, if `stop` is missing the assumption
is to run up to and including the last item. If `step_size` is missing then iteration
will go over all items between `start` and `stop`.
Specifying `step_size` will display every `step_size`'th item, e.g. `[::2]` will display every second
item starting from the first item. Note that specifying a negative number will reverse the order in which items in the list are
displayed. Try this:
```python
sample_list[:3:-1]
```
```
# type here
sample_list[ 2: 7: 2]
# type here
sample_list[::-1]
sample_list[: : -2]
```
#### **Tuple**
Another way to collect objects is using a **tuple**.
This is similar to a `list` and it is created by using round parentheses.
```
a = 2
b = 15
y = ('a', a, b)
y
type(y)
```
What distinguishes lists from tuples is the ability to replace items without creating the iterable from scratch.
We can replace items in a list, but we cannot do so with a tuple.
```
z = ['text', 2, b]
z
type(z)
z[0] = 'another text'
z
y
y[0] = 'text'
```
#### **Dictionary**
Finally, we have **dictionaries**.
These are essentially lists, with the difference that each element is assigned to a _key_.
We create dictionaries using curly braces.
`Dictionary` is a data type that contains a combination of key-value items, for example:
```python
sample_dictionary = dict(math=19, econ=20, physics=16, geology=14)
sample_dictionary_alt = {'math' =19, 'econ'= 20, 'physics'= 16, 'geology'= 14}
print(sample_dictionary, sample_dictionary_alt)
```
Once a dictionary is defined we can access specific values by using relevant key, specifying
it in square brackets, for example:
```python
print(sample_dictionary['econ'])
```
We can use a similar notation to add new key/value pairs to the dictionary:
```python
sample_dictionary['python'] = True
print(sample_dictionary)
```
```
# dict = { "keys" : values , 'math' : 18 , , , , }
# type here
sample_dictionary = {'math' : 19, 'econ' : 20, 'physics' : 16, 'geology': 14}
sample_dictionary
sample_dictionary['math']
sample_dictionary['econ']
sample_dictionary['physics'] = 18
sample_dictionary
sample_dictionary['history'] = 19
sample_dictionary
type(sample_dictionary)
```
The advantage of dictionaries is that we can access their elements by specifying the name of the key, rather than using a number to index the position.
```
sample_dictionary['history']
```
We can access to values and keys of dictionary with below commands:
```
sample_dictionary.values()
sample_dictionary.keys()
```
| github_jupyter |
# Inevitability Of The Time Domain
**version 0.2**
***
AA Miller (Northwestern/CIERA)
14 Sep 2021
The falling cost of silicon detectors (and data storage), has led to a consistent theme over the past ~decade.
The proliferation of wide field surveys.
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/large_detectors_J_Johansson.png" align="middle">
<div align="right"> <font size="-3">(credit: J. Johansson) </font></div>
The motivation is straightforward $\longrightarrow$ MORE
(more area, more sources, better statistics, more discoveries, etc.)
These efforts will in some ways culminate with the Vera C. Rubin Observatory.$^\dagger$
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/2015-SL_LSST_LSSTIllus.jpg" align="middle">
<div align="right"> <font size="-3">(credit: Kavli foundation) </font></div>
$^\dagger$ Though there are existing surveys that go nearly as deep as LSST, and other surveys with a wider field of view, none provide the same combination of wide and deep.
From Lucianne's Introduction to the Rubin Observatory, we know that the DOE camera can detect a source at $m_r \approx 24.5\,\mathrm{mag}$ in $\sim$${30}\,\mathrm{s}$.
We also know that Rubin weak lensing requires depths of $m_r \approx 27.5\,\mathrm{mag}$.
3 mag is a factor of $\sim$$16$ in flux, which corresponds to a factor of $\sim$$250$ in exposure time.
To reach a depth of $\sim$${27.5}\,\mathrm{mag}$ requires a $\sim$${7500}\,\mathrm{s}$ exposure with Rubin.
**Breakout Problem 1**
Should we take $\sim$${2\,\mathrm{hr}}$ long exposures with Rubin?
**Solution to Breakout 1**
*Write your answer here*
**Solution to Breakout 1**
There are many reasons hour-long exposures with Rubin are probably not a good idea (e.g., saturation, StarLink, telescope tracking accuracy), chief among them – cosmic rays:
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/lris-red-upgrade-dark-frame.png" align="middle">
<div align="right"> <font size="-3">(credit: Kassis, Kibrick, Rockosi, & Wirth; WMKO Technical Report 2009) </font></div>
Upshot – our detectors are bigger than ever but we are also limited to relatively short duration exposures.
We can move the telescope in between observations, but a $10\,\mathrm{deg}^2$ FOV means we will literally run out of sky.
As a result
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/thanos.png" align="middle">
<div align="right"> <font size="-3">(credit: Marvel; Rubin Observatory) </font></div>
The Rubin Observatory *requires* time-domain observations.
Time-domain observations beget new science goals.
Session 13 = how to analyze data obtained in the time-domain.
### Quick Aside
Ground-based, time-series photometry looks like this:
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/ASAS_lc.png" align="middle">
<div align="right"> <font size="-3">(credit: Richards et al. 2012) </font></div>
There are a few salient features to notice here:
$~~~~$There are large gaps in the observations
$~~~~$The observations are very noisy
$~~~~$There are non-gaussian processes present in the data
Relative to standard signal processing procedures in the statistical literature, these issues made the analysis of astronomical time-series difficult. "Standard" procedures assume regularly spaced observations or homoskedastic uncertainties, which is a regime that we did not have access to in astronomy.
... Until $\sim$10 years ago with the launch of *CoRoT* (and later *Kepler* and now *TESS*).$^\dagger$
$^\dagger$ X-ray observatories, and now gravitational wave observatories, also give us access to regularly sampled data.
Now we have this:
<img style="display: block; margin-left: auto; margin-right: auto" src="./images/13Cyg_fromAASposter1.jpg" align="middle">
<div align="right"> <font size="-3">(credit: NASA *Kepler* Science Center) </font></div>
These space-based missions provide precise (ppm accuracy) observations taken at regular intervals ($\sim$30 min) over long intervals ($\sim$4 yr for *Kepler*).
As a result, astronomical time-series analysis often falls into two different regimes: ground-based surveys and space-based planet hunting surveys. We will cover relevant problems for both throughout the week.
*Fin*
The sheer number of sources detected by LSST ($\sim$$37$ billion) means that spectroscopy will be out of reach for nearly everything observed by Rubin.
A *significant* challenge in the Rubin-era will be extracting information from photometric-only data sets.
[Note that the following is loaded with personal bias]
One way in which LSST is going to be exceptionally transformative is the time series of persistent objects. Everything brighter than $21\,\mathrm{mag}$ will have better than 1% light curves over a 10 year duration. This is completely unlike anything that has been previously done, and anything planned in the near future. For stars with $m < 21\,\mathrm{mag}$, *Gaia* will provide distance measurements.
What does this sub-1% photometry mean?
$~~~~$all $21\,\mathrm{mag}$ hot jupiters will be found
$~~~~$known rotation periods for all $21\,\mathrm{mag}$ asteroids
$~~~~$most distant stars in the MW halo will be identified
**Breakout Problem 2**
What information might we want to extract from light curves?
**Solution to Breakout 2**
Write you answer here.
Periodicity is the most fundamental signal in astronomical light curves. Periodic signals are always very closely related to fundamental physics (orbits, pulsations, rotation). If you can measure the period of a source, you can know something fundamental about its nature.
The previously mentioned challenges (observational gaps, heteroskedastic uncertainties, non-Gaussian outliers), make it very very difficult to measure periodic signals in ground-based astronomical light curves.
Measuring periodicity will be a significant theme for this session (as will dealing with gaps and heteroskedastic uncertainties).
And with that, let the analysis begin...
| github_jupyter |
# INFO 3402 – Class 13: Data cleaning exercise
[Brian C. Keegan, Ph.D.](http://brianckeegan.com/)
[Assistant Professor, Department of Information Science](https://www.colorado.edu/cmci/people/information-science/brian-c-keegan)
University of Colorado Boulder
Copyright and distributed under an [MIT License](https://opensource.org/licenses/MIT)
```
import pandas as pd
pd.options.display.max_columns = 100
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
```
I got this above-average gnarly data on [passenger traffic reports](https://www.flydenver.com/about/financials/passenger_traffic) from the Denver International Airport.
* Medium difficulty: `passengers_df` or `flights_df`
* Hard difficulty: `pass_type_df`
The goal is to turn this structured but very untidy and irregular data into something we can simply visualize in `seaborn`.
You'll almost certainly want these documentation resources available:
* pandas [`read_excel`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_excel.html)
* pandas [Reshaping and Pivot Tables](https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html)
* seaborn [`catplot`](https://seaborn.pydata.org/generated/seaborn.catplot.html)
```
# Read in the data
passengers_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passengers')
flights_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Flights')
pass_type_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passenger by Type')
```
Go through the EDA checklist:
1. **Formulate your question** → see “Characteristics of a good question”
2. **Read in your data** → Is it properly formatted? Perform cleanup activities
3. **Check the packaging** → Make sure there are the right number of rows & columns, formats, etc.
4. **Look at the top and bottom of data** → Confirm that all observations are there
5. **Check the “n”s** → Identify “landmark” values and to check expectations (number of states, etc.)
6. **Validate against an external data source** → Right order of magnitude, expected distribution, etc.
7. **Make a plot** → Checking and creating expectations about the shape of data and appropriate analyses
8. **Try an easy solution** → What is the simplest test for your question?
## Fix the columns and index
(Hint: The easiest way is to read in the data again, but use different parameters. Read the documentation!)
```
passengers_df = pd.read_excel('DIA_activity.xlsx',sheet_name='Passengers',header=(0,1),index_col=0)
passengers_df.head()
```
## Melt the data down
(Hint: pandas's `melt` and `stack` functions both turn columns into rows)
```
passengers_df2 = passengers_df.unstack()
passengers_df2.head(20)
```
## Make sure each variable has its own column... and rename the columns
(Hint: Mutate column names with wither [`rename`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.rename.html?highlight=rename#pandas.DataFrame.rename) function or [`columns`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.columns.html?highlight=columns#pandas.DataFrame.columns) attribute.
```
passengers_df3 = passengers_df2.reset_index()
passengers_df3.tail(20)
passengers_df3.head()
passengers_df3.columns = ['parent_carrier','carrier','date','passengers']
passengers_df3.head()
```
## Remove the miscellaneous rows
(Hint: Where did the "Unnamed: N_level_1" rows come from in the original spreadsheet?)
```
passengers_df4 = passengers_df3[~passengers_df3['carrier'].str.contains('Unnamed:')]
passengers_df4.tail()
passengers_df4.head()
```
## Separate the "date" column into year and months
(**Normie hint**: Access the month and year as attributes of a datetime/timestamp)
(**Elite hint**: Use `.str.extract` and pass a regular expression that matches a 4 digit number for year and 2 digit number of month)
```
passengers_df4['date'] = pd.to_datetime(passengers_df4['date'])
passengers_df4.tail()
passengers_df4.loc[0,'date'].day
passengers_df4['month'] = passengers_df4['date'].apply(lambda x:x.month)
passengers_df4['year'] = passengers_df4['date'].apply(lambda x:x.year)
passengers_df4.head()
```
## Pivot the data into total passengers by parent carrier per year
(Hint: Pivot table needs to know columns, indexes, values, and *probably* an aggfunc)
```
passengers_df5 = pd.pivot_table(columns='parent_carrier',
index='year',
values='passengers',
data=passengers_df4,aggfunc='sum')
passengers_df5
```
## Plot the pivot table
Is the drop-off in 2018 a "real" effect or caused by something else?
```
ax = passengers_df5.plot(legend=False)
ax.legend(loc='center left',bbox_to_anchor = (1,.5))
```
## Plot data
Make a seaborn `catplot` with "month" on the x-axis, "passengers" on the y-axis, and hues for the different parent carriers (probably excluding "Other"). What is the top month (on average) for passengers into DIA?
```
passengers_df4.head()
sb.catplot(x='year',y='passengers',hue='parent_carrier',row='month',data=passengers_df4,
aspect=3,kind='point')
```
## Re-do clean-up for `flights_df`
Make a nice clean dataset so we can use both of them.
## Compute average monthly passengers per flight
(Hint: Confirm cleaned `passengers_df` and `flights_df` are similar dimensions)
(Hint: You may also need to reference pandas's [`div`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.div.html) function.)
## Plot average monthly passengers per flight
Make a seaborn `catplot` with "month" on the x-axis, "passengers" on the y-axis, and hues for the different carriers. Is the top month still the same?
| github_jupyter |
# Week 2 Assignment: Feature Engineering
For this week's assignment, you will build a data pipeline using using [Tensorflow Extended (TFX)](https://www.tensorflow.org/tfx) to prepare features from the [Metro Interstate Traffic Volume dataset](https://archive.ics.uci.edu/ml/datasets/Metro+Interstate+Traffic+Volume). Try to only use the documentation and code hints to accomplish the tasks but feel free to review the 2nd ungraded lab this week in case you get stuck.
Upon completion, you will have:
* created an InteractiveContext to run TFX components interactively
* used TFX ExampleGen component to split your dataset into training and evaluation datasets
* generated the statistics and the schema of your dataset using TFX StatisticsGen and SchemaGen components
* validated the evaluation dataset statistics using TFX ExampleValidator
* performed feature engineering using the TFX Transform component
Let's begin!
## Table of Contents
- [1 - Setup](#1)
- [1.1 - Imports](#1-1)
- [1.2 - Define Paths](#1-2)
- [1.3 - Preview the Dataset](#1-3)
- [1.4 - Create the InteractiveContext](#1-4)
- [2 - Run TFX components interactively](#2)
- [2.1 - ExampleGen](#2-1)
- [Exercise 1 - ExampleGen](#ex-1)
- [Exercise 2 - get_records()](#ex-2)
- [2.2 - StatisticsGen](#2-2)
- [Exercise 3 - StatisticsGen](#ex-3)
- [2.3 - SchemaGen](#2-3)
- [Exercise 4 - SchemaGen](#ex-4)
- [2.4 - ExampleValidator](#2-4)
- [Exercise 5 - ExampleValidator](#ex-5)
- [2.5 - Transform](#2-5)
- [Exercise 6 - preprocessing_fn()](#ex-6)
- [Exercise 7 - Transform](#ex-7)
<a name='1'></a>
## 1 - Setup
As usual, you will first need to import the necessary packages. For reference, the lab environment uses *TensorFlow version: 2.3.1* and *TFX version: 0.24.0*.
<a name='1-1'></a>
### 1.1 Imports
```
import tensorflow as tf
from tfx.components import CsvExampleGen
from tfx.components import ExampleValidator
from tfx.components import SchemaGen
from tfx.components import StatisticsGen
from tfx.components import Transform
from tfx.orchestration.experimental.interactive.interactive_context import InteractiveContext
from google.protobuf.json_format import MessageToDict
import os
import pprint
pp = pprint.PrettyPrinter()
```
<a name='1-2'></a>
### 1.2 - Define paths
You will define a few global variables to indicate paths in the local workspace.
```
# location of the pipeline metadata store
_pipeline_root = 'metro_traffic_pipeline/'
# directory of the raw data files
_data_root = 'metro_traffic_pipeline/data'
# path to the raw training data
_data_filepath = os.path.join(_data_root, 'metro_traffic_volume.csv')
```
<a name='1-3'></a>
### 1.3 - Preview the dataset
The [Metro Interstate Traffic Volume dataset](https://archive.ics.uci.edu/ml/datasets/Metro+Interstate+Traffic+Volume) contains hourly traffic volume of a road in Minnesota from 2012-2018. With this data, you can develop a model for predicting the traffic volume given the date, time, and weather conditions. The attributes are:
* **holiday** - US National holidays plus regional holiday, Minnesota State Fair
* **temp** - Average temp in Kelvin
* **rain_1h** - Amount in mm of rain that occurred in the hour
* **snow_1h** - Amount in mm of snow that occurred in the hour
* **clouds_all** - Percentage of cloud cover
* **weather_main** - Short textual description of the current weather
* **weather_description** - Longer textual description of the current weather
* **date_time** - DateTime Hour of the data collected in local CST time
* **traffic_volume** - Numeric Hourly I-94 ATR 301 reported westbound traffic volume
* **month** - taken from date_time
* **day** - taken from date_time
* **day_of_week** - taken from date_time
* **hour** - taken from date_time
*Disclaimer: We added the last four attributes shown above (i.e. month, day, day_of_week, hour) to the original dataset to increase the features you can transform later.*
Take a quick look at the first few rows of the CSV file.
```
# Preview the dataset
!head {_data_filepath}
```
<a name='1-4'></a>
### 1.4 - Create the InteractiveContext
You will need to initialize the `InteractiveContext` to enable running the TFX components interactively. As before, you will let it create the metadata store in the `_pipeline_root` directory. You can safely ignore the warning about the missing metadata config file.
```
# Declare the InteractiveContext and use a local sqlite file as the metadata store.
# You can ignore the warning about the missing metadata config file
context = InteractiveContext(pipeline_root=_pipeline_root)
```
<a name='2'></a>
## 2 - Run TFX components interactively
In the following exercises, you will create the data pipeline components one-by-one, run each of them, and visualize their output artifacts. Recall that we refer to the outputs of pipeline components as *artifacts* and these can be inputs to the next stage of the pipeline.
<a name='2-1'></a>
### 2.1 - ExampleGen
The pipeline starts with the [ExampleGen](https://www.tensorflow.org/tfx/guide/examplegen) component. It will:
* split the data into training and evaluation sets (by default: 2/3 train, 1/3 eval).
* convert each data row into `tf.train.Example` format. This [protocol buffer](https://developers.google.com/protocol-buffers) is designed for Tensorflow operations and is used by the TFX components.
* compress and save the data collection under the `_pipeline_root` directory for other components to access. These examples are stored in `TFRecord` format. This optimizes read and write operations within Tensorflow especially if you have a large collection of data.
<a name='ex-1'></a>
#### Exercise 1: ExampleGen
Fill out the code below to ingest the data from the CSV file stored in the `_data_root` directory.
```
### START CODE HERE
# Instantiate ExampleGen with the input CSV dataset
example_gen = CsvExampleGen(input_base=_data_root)
# Run the component using the InteractiveContext instance
context.run(example_gen)
### END CODE HERE
```
You should see the output cell of the `InteractiveContext` above showing the metadata associated with the component execution. You can expand the items under `.component.outputs` and see that an `Examples` artifact for the train and eval split is created in `metro_traffic_pipeline/CsvExampleGen/examples/{execution_id}`.
You can also check that programmatically with the following snippet. You can focus on the `try` block. The `except` and `else` block is needed mainly for grading. `context.run()` yields no operation when executed in a non-interactive environment (such as the grader script that runs outside of this notebook). In such scenarios, the URI must be manually set to avoid errors.
```
try:
# get the artifact object
artifact = example_gen.outputs['examples'].get()[0]
# print split names and uri
print(f'split names: {artifact.split_names}')
print(f'artifact uri: {artifact.uri}')
# for grading since context.run() does not work outside the notebook
except IndexError:
print("context.run() was no-op")
examples_path = './metro_traffic_pipeline/CsvExampleGen/examples'
dir_id = os.listdir(examples_path)[0]
artifact_uri = f'{examples_path}/{dir_id}'
else:
artifact_uri = artifact.uri
```
The ingested data has been saved to the directory specified by the artifact Uniform Resource Identifier (URI). As a sanity check, you can take a look at some of the training examples. This requires working with Tensorflow data types, particularly `tf.train.Example` and `TFRecord` (you can read more about them [here](https://www.tensorflow.org/tutorials/load_data/tfrecord)). Let's first load the `TFRecord` into a variable:
```
# Get the URI of the output artifact representing the training examples, which is a directory
train_uri = os.path.join(artifact_uri, 'train')
# Get the list of files in this directory (all compressed TFRecord files)
tfrecord_filenames = [os.path.join(train_uri, name)
for name in os.listdir(train_uri)]
# Create a `TFRecordDataset` to read these files
dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
```
<a name='ex-2'></a>
#### Exercise 2: get_records()
Complete the helper function below to return a specified number of examples.
*Hints: You may find the [MessageToDict](https://googleapis.dev/python/protobuf/latest/google/protobuf/json_format.html#google.protobuf.json_format.MessageToDict) helper function and tf.train.Example's [ParseFromString()](https://googleapis.dev/python/protobuf/latest/google/protobuf/message.html#google.protobuf.message.Message.ParseFromString) method useful here. You can also refer [here](https://www.tensorflow.org/tutorials/load_data/tfrecord) for a refresher on TFRecord and tf.train.Example()*
```
def get_records(dataset, num_records):
'''Extracts records from the given dataset.
Args:
dataset (TFRecordDataset): dataset saved by ExampleGen
num_records (int): number of records to preview
'''
# initialize an empty list
records = []
### START CODE HERE
# Use the `take()` method to specify how many records to get
for tfrecord in dataset.take(num_records):
# Get the numpy property of the tensor
serialized_example = tfrecord.numpy()
# Initialize a `tf.train.Example()` to read the serialized data
example = tf.train.Example()
# Read the example data (output is a protocol buffer message)
example.ParseFromString(serialized_example)
# convert the protocol buffer message to a Python dictionary
example_dict = MessageToDict(example)
# append to the records list
records.append(example_dict)
### END CODE HERE
return records
# Get 3 records from the dataset
sample_records = get_records(dataset, 3)
# Print the output
pp.pprint(sample_records)
```
You should see three of the examples printed above. Now that `ExampleGen` has finished ingesting the data, the next step is data analysis.
<a name='2-2'></a>
### 2.2 - StatisticsGen
The [StatisticsGen](https://www.tensorflow.org/tfx/guide/statsgen) component computes statistics over your dataset for data analysis, as well as for use in downstream components. It uses the [TensorFlow Data Validation](https://www.tensorflow.org/tfx/data_validation/get_started) library.
`StatisticsGen` takes as input the dataset ingested using `CsvExampleGen`.
<a name='ex-3'></a>
#### Exercise 3: StatisticsGen
Fill the code below to generate statistics from the output examples of `CsvExampleGen`.
```
### START CODE HERE
# Instantiate StatisticsGen with the ExampleGen ingested dataset
statistics_gen = StatisticsGen(examples=example_gen.outputs['examples'])
# Run the component
context.run(statistics_gen)
### END CODE HERE
# Plot the statistics generated
context.show(statistics_gen.outputs['statistics'])
```
<a name='2-3'></a>
### 2.3 - SchemaGen
The [SchemaGen](https://www.tensorflow.org/tfx/guide/schemagen) component also uses TFDV to generate a schema based on your data statistics. As you've learned previously, a schema defines the expected bounds, types, and properties of the features in your dataset.
`SchemaGen` will take as input the statistics that we generated with `StatisticsGen`, looking at the training split by default.
<a name='ex-4'></a>
#### Exercise 4: SchemaGen
```
### START CODE HERE
# Instantiate SchemaGen with the output statistics from the StatisticsGen
schema_gen = SchemaGen(statistics=statistics_gen.outputs['statistics'])
# Run the component
context.run(schema_gen)
### END CODE HERE
```
If all went well, you can now visualize the generated schema as a table.
```
# Visualize the output
context.show(schema_gen.outputs['schema'])
```
Each attribute in your dataset shows up as a row in the schema table, alongside its properties. The schema also captures all the values that a categorical feature takes on, denoted as its domain.
This schema will be used to detect anomalies in the next step.
<a name='2-4'></a>
### 2.4 - ExampleValidator
The [ExampleValidator](https://www.tensorflow.org/tfx/guide/exampleval) component detects anomalies in your data based on the generated schema from the previous step. Like the previous two components, it also uses TFDV under the hood.
`ExampleValidator` will take as input the statistics from `StatisticsGen` and the schema from `SchemaGen`. By default, it compares the statistics from the evaluation split to the schema from the training split.
<a name='2-4'></a>
#### Exercise 5: ExampleValidator
Fill the code below to detect anomalies in your datasets.
```
### START CODE HERE
# Instantiate ExampleValidator with the statistics and schema from the previous steps
example_validator = ExampleValidator(statistics=statistics_gen.outputs['statistics'],schema=schema_gen.outputs['schema'])
# Run the component
context.run(example_validator)
### END CODE HERE
```
As with the previous steps, you can visualize the anomalies as a table.
```
# Visualize the output
context.show(example_validator.outputs['anomalies'])
```
If there are anomalies detected, you should examine how you should handle it. For example, you can relax distribution constraints or modify the domain of some features. You've had some practice with this last week when you used TFDV and you can also do that here.
For this particular case, there should be no anomalies detected and we can proceed to the next step.
<a name='2-5'></a>
### 2.5 - Transform
In this section, you will use the [Transform](https://www.tensorflow.org/tfx/guide/transform) component to perform feature engineering.
`Transform` will take as input the data from `ExampleGen`, the schema from `SchemaGen`, as well as a module containing the preprocessing function.
The component expects an external module for your Transform code so you need to use the magic command `%% writefile` to save the file to disk. We have defined a few constants that group the data's attributes according to the transforms you will perform later. This file will also be saved locally.
```
# Set the constants module filename
_traffic_constants_module_file = 'traffic_constants.py'
%%writefile {_traffic_constants_module_file}
# Features to be scaled to the z-score
DENSE_FLOAT_FEATURE_KEYS = ['temp', 'snow_1h']
# Features to bucketize
BUCKET_FEATURE_KEYS = ['rain_1h']
# Number of buckets used by tf.transform for encoding each feature.
FEATURE_BUCKET_COUNT = {'rain_1h': 3}
# Feature to scale from 0 to 1
RANGE_FEATURE_KEYS = ['clouds_all']
# Number of vocabulary terms used for encoding VOCAB_FEATURES by tf.transform
VOCAB_SIZE = 1000
# Count of out-of-vocab buckets in which unrecognized VOCAB_FEATURES are hashed.
OOV_SIZE = 10
# Features with string data types that will be converted to indices
VOCAB_FEATURE_KEYS = [
'holiday',
'weather_main',
'weather_description'
]
# Features with int data type that will be kept as is
CATEGORICAL_FEATURE_KEYS = [
'hour', 'day', 'day_of_week', 'month'
]
# Feature to predict
VOLUME_KEY = 'traffic_volume'
def transformed_name(key):
return key + '_xf'
```
<a name='ex-6'></a>
#### Exercise 6
Next, you will fill out the transform module. As mentioned, this will also be saved to disk. Specifically, you will complete the `preprocessing_fn` which defines the transformations. See the code comments for instructions and refer to the [tft module documentation](https://www.tensorflow.org/tfx/transform/api_docs/python/tft) to look up which function to use for a given group of keys.
For the label (i.e. `VOLUME_KEY`), you will transform it to indicate if it is greater than the mean of the entire dataset. For the transform to work, you will need to convert a [SparseTensor](https://www.tensorflow.org/api_docs/python/tf/sparse/SparseTensor) to a dense one. We've provided a `_fill_in_missing()` helper function for you to use.
```
# Set the transform module filename
_traffic_transform_module_file = 'traffic_transform.py'
%%writefile {_traffic_transform_module_file}
import tensorflow as tf
import tensorflow_transform as tft
import traffic_constants
# Unpack the contents of the constants module
_DENSE_FLOAT_FEATURE_KEYS = traffic_constants.DENSE_FLOAT_FEATURE_KEYS
_RANGE_FEATURE_KEYS = traffic_constants.RANGE_FEATURE_KEYS
_VOCAB_FEATURE_KEYS = traffic_constants.VOCAB_FEATURE_KEYS
_VOCAB_SIZE = traffic_constants.VOCAB_SIZE
_OOV_SIZE = traffic_constants.OOV_SIZE
_CATEGORICAL_FEATURE_KEYS = traffic_constants.CATEGORICAL_FEATURE_KEYS
_BUCKET_FEATURE_KEYS = traffic_constants.BUCKET_FEATURE_KEYS
_FEATURE_BUCKET_COUNT = traffic_constants.FEATURE_BUCKET_COUNT
_VOLUME_KEY = traffic_constants.VOLUME_KEY
_transformed_name = traffic_constants.transformed_name
def preprocessing_fn(inputs):
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
Returns:
Map from string feature key to transformed feature operations.
"""
outputs = {}
### START CODE HERE
# Scale these features to the z-score.
for key in _DENSE_FLOAT_FEATURE_KEYS:
# Scale these features to the z-score.
outputs[_transformed_name(key)] = tft.scale_to_z_score(inputs[key])
# Scale these feature/s from 0 to 1
for key in _RANGE_FEATURE_KEYS:
outputs[_transformed_name(key)] = tft.scale_to_0_1(inputs[key])
# Transform the strings into indices
# hint: use the VOCAB_SIZE and OOV_SIZE to define the top_k and num_oov parameters
for key in _VOCAB_FEATURE_KEYS:
outputs[_transformed_name(key)] = tft.compute_and_apply_vocabulary(
inputs[key],
top_k=_VOCAB_SIZE,
num_oov_buckets=_OOV_SIZE)
# Bucketize the feature
for key in _BUCKET_FEATURE_KEYS:
outputs[_transformed_name(key)] = tft.bucketize(
inputs[key],
_FEATURE_BUCKET_COUNT[key],
always_return_num_quantiles=False)
# Keep as is. No tft function needed.
for key in _CATEGORICAL_FEATURE_KEYS:
outputs[_transformed_name(key)] = inputs[key]
# Use `tf.cast` to cast the label key to float32 and fill in the missing values.
traffic_volume = tf.cast(_fill_in_missing(inputs[_VOLUME_KEY]), tf.float32)
# Create a feature that shows if the traffic volume is greater than the mean and cast to an int
outputs[_transformed_name(_VOLUME_KEY)] = tf.cast(
# Use `tf.greater` to check if the traffic volume in a row is greater than the mean of the entire traffic volumn column
tf.greater(traffic_volume,
tft.mean(tf.cast(traffic_volume,
tf.float32))),
tf.int64)
### END CODE HERE
return outputs
def _fill_in_missing(x):
"""Replace missing values in a SparseTensor and convert to a dense tensor.
Fills in missing values of `x` with '' or 0, and converts to a dense tensor.
Args:
x: A `SparseTensor` of rank 2. Its dense shape should have size at most 1
in the second dimension.
Returns:
A rank 1 tensor where missing values of `x` have been filled in.
"""
default_value = '' if x.dtype == tf.string else 0
return tf.squeeze(
tf.sparse.to_dense(
tf.SparseTensor(x.indices, x.values, [x.dense_shape[0], 1]),
default_value),
axis=1)
```
<a name='ex-7'></a>
#### Exercise 7
With the transform module defined, complete the code below to perform feature engineering on the raw data.
```
# ignore tf warning messages
tf.get_logger().setLevel('ERROR')
### START CODE HERE
# Instantiate the Transform component
transform = Transform(
examples=example_gen.outputs['examples'],
schema=schema_gen.outputs['schema'],
module_file=os.path.abspath(_traffic_transform_module_file))
# Run the component
context.run(transform)
### END CODE HERE
```
You should see the output cell by `InteractiveContext` above and see the three artifacts in `.component.outputs`:
* `transform_graph` is the graph that performs the preprocessing operations. This will be included during training and serving to ensure consistent transformations of incoming data.
* `transformed_examples` points to the preprocessed training and evaluation data.
* `updated_analyzer_cache` are stored calculations from previous runs.
The `transform_graph` artifact URI should point to a directory containing:
* The `metadata` subdirectory containing the schema of the original data.
* The `transformed_metadata` subdirectory containing the schema of the preprocessed data.
* The `transform_fn` subdirectory containing the actual preprocessing graph.
Again, for grading purposes, we inserted an `except` and `else` below to handle checking the output outside the notebook environment.
```
try:
# Get the uri of the transform graph
transform_graph_uri = transform.outputs['transform_graph'].get()[0].uri
except IndexError:
print("context.run() was no-op")
transform_path = './metro_traffic_pipeline/Transform/transformed_examples'
dir_id = os.listdir(transform_path)[0]
transform_graph_uri = f'{transform_path}/{dir_id}'
else:
# List the subdirectories under the uri
os.listdir(transform_graph_uri)
```
Lastly, you can also take a look at a few of the transformed examples.
```
try:
# Get the URI of the output artifact representing the transformed examples
train_uri = os.path.join(transform.outputs['transformed_examples'].get()[0].uri, 'train')
except IndexError:
print("context.run() was no-op")
train_uri = os.path.join(transform_graph_uri, 'train')
# Get the list of files in this directory (all compressed TFRecord files)
tfrecord_filenames = [os.path.join(train_uri, name)
for name in os.listdir(train_uri)]
# Create a `TFRecordDataset` to read these files
transformed_dataset = tf.data.TFRecordDataset(tfrecord_filenames, compression_type="GZIP")
# Get 3 records from the dataset
sample_records_xf = get_records(transformed_dataset, 3)
# Print the output
pp.pprint(sample_records_xf)
```
**Congratulations on completing this week's assignment!** You've just demonstrated how to build a data pipeline and do feature engineering. You will build upon these concepts in the next weeks where you will deal with more complex datasets and also access the metadata store. Keep up the good work!
| github_jupyter |
# Analyzing Movie Reviews
Notebook by [Wenyi Xu](https://github.com/xuwenyihust)
<br/>
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import linregress
from scipy.stats.stats import pearsonr
%matplotlib inline
```
## Import Data
**pd.read_csv()** can directly read url
```
url = "https://raw.githubusercontent.com/fivethirtyeight/data/master/fandango/fandango_score_comparison.csv"
reviews = pd.read_csv(url)
reviews.head(3)
```
## Data Summaries
Get some summaries about the raw data.
```
reviews.columns
```
## Rotten Tomatoes VS. IMDB
Want to compare the user scores of Rotten Tomatoes & IMDB.
First get some summaries of these 2 columns.
```
print('Rotton Tomato score min:\t' + str(reviews["RT_user_norm"].min()))
print('Rotton Tomato score max:\t' + str(reviews["RT_user_norm"].max()))
print('-'*50)
print('IMDB score min:\t\t\t' + str(reviews["IMDB_norm"].min()))
print('IMDB score max:\t\t\t' + str(reviews["IMDB_norm"].max()))
```
So we guess RT user scores have a **larger spread** compared to IMDB's.
<br/>
```
print(reviews["RT_user_norm"].mean())
print(reviews["IMDB_norm"].mean())
```
## Histograms Comparison
Compare the **distribution** of **Rotten Tomatoes** & **IMDB** user scores.
To have a fair comparison, choose col **"RT_user_norm"** & **"IMDB_norm"** to have them both in 0~5.
<br/>
```
fig = plt.figure(figsize=(10,4))
ax1 = fig.add_subplot(1,2,1)
ax1.hist(reviews["RT_user_norm"], bins=10, facecolor='blue', alpha=0.75)
ax1.set_title("Distribution of RT user scores")
ax1.set_xlabel("norm_score")
ax1.set_ylabel("num")
ax1.set_xlim(1,5.0)
ax2 = fig.add_subplot(1,2,2)
ax2.hist(reviews["IMDB_norm"], bins=8, facecolor='blue', alpha=0.75)
ax2.set_title("Distribution of IMDB user scores")
ax2.set_xlabel("norm_score")
ax2.set_ylabel("num")
ax2.set_xlim(1,5.0)
```
From the above charts, we can see that
the distribution of **RT's** rating is more **flat**.
While the **IMDB** ratings are much more **clustered** around some value between 3 & 3.5, and **skews left**.
<br/>
## Variance Comparison
To directly show how spread they are, can also calculate the **variance**.
```
print(np.var(reviews["RT_user_norm"]))
print(np.var(reviews["IMDB_norm"]))
```
The result conforms to the distribution visualization.
<br/>
## Correlations
Want to know the correlations between RT's & IMDB's scores.
First show the scatter plot.
```
plt.figure(figsize=(4,4))
plt.scatter(reviews["RT_user_norm"], reviews["IMDB_norm"])
plt.xlabel("IMDB")
plt.ylabel("Rotten Tomatoes")
plt.show()
```
Looks kind of linear.
<br/>
## Add the regression line
Use the line chart to plot the regression line.
```
slope, intercept, r_value, p_value, stderr_slope = linregress(reviews["RT_user_norm"], reviews["IMDB_norm"])
def predict(x):
return x * slope + intercept
plt.figure(figsize=(4,4))
plt.scatter(reviews["RT_user_norm"], reviews["IMDB_norm"])
plt.xlabel("IMDB")
plt.ylabel("Rotten Tomatoes")
x = [0.0, 1.0, 2.0, 3.0, 4.0, 5.0]
y = [predict(i) for i in x]
plt.plot(x,y)
```
## Correlation Coefficient
```
print(pearsonr(reviews["RT_user_norm"], reviews["IMDB_norm"])[0])
```
| github_jupyter |
```
!pip install dynet
!git clone https://github.com/neubig/nn4nlp-code.git
from __future__ import print_function
import time
from collections import defaultdict
import random
import math
import sys
import argparse
import dynet as dy
import numpy as np
# format of files: each line is "word1 word2 ..." aligned line-by-line
train_src_file = "nn4nlp-code/data/parallel/train.ja"
train_trg_file = "nn4nlp-code/data/parallel/train.en"
dev_src_file = "nn4nlp-code/data/parallel/dev.ja"
dev_trg_file = "nn4nlp-code/data/parallel/dev.en"
w2i_src = defaultdict(lambda: len(w2i_src))
w2i_trg = defaultdict(lambda: len(w2i_trg))
def read(fname_src, fname_trg):
"""
Read parallel files where each line lines up
"""
with open(fname_src, "r") as f_src, open(fname_trg, "r") as f_trg:
for line_src, line_trg in zip(f_src, f_trg):
sent_src = [w2i_src[x] for x in line_src.strip().split()]
sent_trg = [w2i_trg[x] for x in line_trg.strip().split()]
yield (sent_src, sent_trg)
# Read the data
train = list(read(train_src_file, train_trg_file))
unk_src = w2i_src["<unk>"]
w2i_src = defaultdict(lambda: unk_src, w2i_src)
unk_trg = w2i_trg["<unk>"]
w2i_trg = defaultdict(lambda: unk_trg, w2i_trg)
nwords_src = len(w2i_src)
nwords_trg = len(w2i_trg)
dev = list(read(dev_src_file, dev_trg_file))
# DyNet Starts
model = dy.Model()
trainer = dy.AdamTrainer(model)
# Model parameters
EMBED_SIZE = 64
HIDDEN_SIZE = 128
BATCH_SIZE = 16
# Lookup parameters for word embeddings
LOOKUP_SRC = model.add_lookup_parameters((nwords_src, EMBED_SIZE))
LOOKUP_TRG = model.add_lookup_parameters((nwords_trg, EMBED_SIZE))
# Word-level BiLSTMs
LSTM_SRC_FWD = dy.LSTMBuilder(1, EMBED_SIZE, HIDDEN_SIZE/2, model)
LSTM_SRC_BWD = dy.LSTMBuilder(1, EMBED_SIZE, HIDDEN_SIZE/2, model)
LSTM_TRG_FWD = dy.LSTMBuilder(1, EMBED_SIZE, HIDDEN_SIZE/2, model)
LSTM_TRG_BWD = dy.LSTMBuilder(1, EMBED_SIZE, HIDDEN_SIZE/2, model)
def encode_sents(look, fwd, bwd, sents):
embs = [[look[x] for x in sent] for sent in sents]
return [dy.concatenate([fwd.transduce(x)[-1], bwd.transduce(x)[-1]]) for x in embs]
# Calculate loss for one mini-batch
def calc_loss(sents):
dy.renew_cg()
src_fwd = LSTM_SRC_FWD.initial_state()
src_bwd = LSTM_SRC_BWD.initial_state()
trg_fwd = LSTM_TRG_FWD.initial_state()
trg_bwd = LSTM_TRG_BWD.initial_state()
# Encoding
src_reps = encode_sents(LOOKUP_SRC, src_fwd, src_bwd, [src for src, trg in sents])
trg_reps = encode_sents(LOOKUP_TRG, trg_fwd, trg_bwd, [trg for src, trg in sents])
# Concatenate the sentence representations to a single matrix
mtx_src = dy.concatenate_cols(src_reps)
mtx_trg = dy.concatenate_cols(trg_reps)
# Do matrix multiplication to get a matrix of dot product similarity scores
sim_mtx = dy.transpose(mtx_src) * mtx_trg
# Calculate the hinge loss over all dimensions
loss = dy.hinge_dim(sim_mtx, list(range(len(sents))), d=1)
return dy.sum_elems(loss)
# Calculate representations for one corpus
def index_corpus(sents):
# To take advantage of auto-batching, do several at a time
for sid in range(0, len(sents), BATCH_SIZE):
dy.renew_cg()
src_fwd = LSTM_SRC_FWD.initial_state()
src_bwd = LSTM_SRC_BWD.initial_state()
trg_fwd = LSTM_TRG_FWD.initial_state()
trg_bwd = LSTM_TRG_BWD.initial_state()
# Set up the computation graph
src_exprs = encode_sents(LOOKUP_SRC, src_fwd, src_bwd, [src for src, trg in sents[sid:min(sid+BATCH_SIZE,len(sents))]])
trg_exprs = encode_sents(LOOKUP_TRG, trg_fwd, trg_bwd, [trg for src, trg in sents[sid:min(sid+BATCH_SIZE,len(sents))]])
# Perform the forward pass to calculate everything at once
trg_exprs[-1][1].forward()
for src_expr, trg_expr in zip(src_exprs, trg_exprs):
yield (src_expr.npvalue(), trg_expr.npvalue())
# Perform retrieval, and return both scores and ranked order of candidates
def retrieve(src, db_mtx):
scores = np.dot(db_mtx,src)
ranks = np.argsort(-scores)
return ranks, scores
# Perform training
start = time.time()
train_mbs = all_time = dev_time = all_tagged = this_sents = this_loss = 0
for ITER in range(100):
random.shuffle(train)
for sid in range(0, len(train), BATCH_SIZE):
my_size = min(BATCH_SIZE, len(train)-sid)
train_mbs += 1
if train_mbs % int(1000/BATCH_SIZE) == 0:
trainer.status()
print("loss/sent=%.4f, sent/sec=%.4f" % (this_loss / this_sents, (train_mbs * BATCH_SIZE) / (time.time() - start - dev_time)), file=sys.stderr)
this_loss = this_sents = 0
# train on the minibatch
loss_exp = calc_loss(train[sid:sid+BATCH_SIZE])
this_loss += loss_exp.scalar_value()
this_sents += BATCH_SIZE
loss_exp.backward()
trainer.update()
# Perform evaluation
dev_start = time.time()
rec_at_1, rec_at_5, rec_at_10 = 0, 0, 0
reps = list(index_corpus(dev))
trg_mtx = np.stack([trg for src, trg in reps])
for i, (src, trg) in enumerate(reps):
ranks, scores = retrieve(src, trg_mtx)
if ranks[0] == i: rec_at_1 += 1
if i in ranks[:5]: rec_at_5 += 1
if i in ranks[:10]: rec_at_10 += 1
dev_time += time.time()-dev_start
print("epoch %r: dev recall@1=%.2f%% recall@5=%.2f%% recall@10=%.2f%%" % (ITER, rec_at_1/len(dev)*100, rec_at_5/len(dev)*100, rec_at_10/len(dev)*100))
```
| github_jupyter |
```
import string
from functools import partial
import matplotlib.pyplot as plt
import numpy as np
from tqdm import tqdm
from reservoirpy.nodes import Reservoir, FORCE
from reservoirpy.experimental import AsabukiNorm, RandomChoice
from reservoirpy import merge
from scipy.sparse import csr_matrix, coo_matrix
def asabuki_1alphachunks(n_timesteps,
chunk=('a', 'b', 'c', 'd'),
width=50,
input_gain=2,
min_length=5,
dt=1):
alphabet = tuple(string.ascii_lowercase)
random_elements = tuple(set(alphabet) - set(chunk))
input_shape = np.zeros(width * 2)
input_shape[0:width] = input_gain * (1 - np.exp(-(np.arange(0, width) / 10)))
input_shape[width:2 * width] = input_gain * np.exp(-(np.arange(0, width) / 10))
simtime = np.arange(0, n_timesteps * dt, dt)
simtime_len = len(simtime)
symbol_list = np.zeros(simtime_len)
target_list = np.zeros(simtime_len)
random_seq_len = np.random.randint(min_length, min_length + 4)
random_seq = [''] * random_seq_len
for i in range(random_seq_len):
random_seq[i] = random_elements[
np.random.randint(0, len(random_elements))]
input_type = random_seq
m = 0
I = np.zeros((len(alphabet), simtime_len))
for i in range(simtime_len):
if i % width == 0 and i > 0:
if input_type == chunk:
if m == len(chunk) - 1:
random_seq_len = np.random.randint(min_length,
min_length + 4)
random_seq = [''] * random_seq_len
for l in range(random_seq_len):
random_seq[l] = random_elements[
np.random.randint(0, len(random_elements))]
input_type = random_seq
m = 0
else:
input_type = chunk
m += 1
elif input_type == random_seq:
if m == len(random_seq) - 1:
input_type = chunk
m = 0
else:
input_type = random_seq
m += 1
I[alphabet.index(input_type[m]),
i:min(i + width * 2, simtime_len)] = input_shape[
0:min(i + width * 2,
simtime_len) - i]
return I.T
train_len = 500000
test_len = 2000
X = asabuki_1alphachunks(train_len + test_len)
samples = 2000
plt.figure(figsize=(15, 4))
plt.ylim(-0.01, 2.2)
for i in range(X.shape[1]):
letter = string.ascii_lowercase[i]
color = "red" if letter in {"a", "b", "c", "d"} else "black"
sig = X[:samples, i]
if sig.sum() > 0.0:
plt.plot(sig)
pos = np.where(np.isclose(sig, 2.0-1e-8))[0]
for p in pos:
plt.text(p, 2.1, letter, color=color)
dt = 1 # one timestep represents 1ms
tau = 10 # time contant of reservoirs
width = 50 # width in time of input stimuli
g_FG = 1 # feedback gain
N = 300 # number of reservoir units
n = 300 # number of synapses selected to be connected with readout
p = 1.0 # connectivity of reservoir
g = 1.5 # reservoir recurrent matrix gain
alpha = 100 # P = Id * 1/alpha (for FORCE learning)
sigma = 0.3 # noise gain
window = 300 # window for normalization
beta = 3 # coefficient for normalization
learn_every = 2 # readout is trained every 2 steps
scale = 1.0 / np.sqrt(p * N) # scale of reservoir matrix
M = np.zeros((N, N))
M2 = np.zeros((N, N))
for i in range(N):
for j in range(N):
if np.random.rand() < p:
M[i, j] = (np.random.randn()) * g * scale
if np.random.rand() < p:
M2[i, j] = (np.random.randn()) * g * scale
nRec2Out = N
nIn2Rec = X.shape[1]
wo1 = np.random.randn(n, 1) / np.sqrt(n)
wo2 = np.random.randn(n, 1) / np.sqrt(n)
wf1 = (np.random.rand(N, 1) - 0.5) * 2 * g_FG
wf2 = (np.random.rand(N, 1) - 0.5) * 2 * g_FG
win = np.zeros((N, nIn2Rec))
win2 = np.zeros((N, nIn2Rec))
for i in range(N):
win[i, np.random.randint(0, nIn2Rec)] = np.random.randn()
win2[i, np.random.randint(0, nIn2Rec)] = np.random.randn()
res1 = Reservoir(N, lr=dt/tau, input_bias=False, rc_connectivity=p,
W=csr_matrix(M), Win=csr_matrix(win),
Wfb=wf1,
noise_rc=sigma * np.sqrt(dt),
noise_type="normal",
equation="external",
name="h1")
res2 = Reservoir(N, lr=dt/tau, input_bias=False, rc_connectivity=p,
W=csr_matrix(M2), Win=csr_matrix(win2),
Wfb=wf2,
noise_rc=sigma * np.sqrt(dt),
noise_type="normal",
equation="external",
name="h2")
read1 = FORCE(1, alpha=alpha, name="r1", Wout=wo1, input_bias=False)
read2 = FORCE(1, alpha=alpha, name="r2", Wout=wo2, input_bias=False)
norm1 = AsabukiNorm(window=window * width, beta=beta)
norm2 = AsabukiNorm(window=window * width, beta=beta)
# feedack connections
teacher1 = (read2 >> norm2)
teacher2 = (read1 >> norm1)
res1 <<= read1
res2 <<= read2
# this is our training model
branch1 = res1 >> RandomChoice(n=n, name="rand1") >> read1
branch2 = res2 >> RandomChoice(n=n, name="rand2") >> read2
model = merge(branch1, branch2)
# to warmup the special normalization node, we also build a
# transient model
trans1 = branch1 >> norm1
trans2 = branch2 >> norm2
transient_model = merge(branch1, branch2)
init_states = {"h1": 0.5 * np.random.randn(1, N),
"h2": 0.5 * np.random.randn(1, N),
"r1": 0.5 * np.random.randn(1, 1),
"r2": 0.5 * np.random.randn(1, 1)}
# first, warmup the nodes a bit to be sure the rolling window of the normalization
# nodes is full of real data
transients = width * window
transient_model.run(X[:transients], from_state=init_states)
"""
for i, x in enumerate(tqdm(X[transients:train_len])):
s = model(x, return_states=["rand1", "rand2", "r1", "r2"])
y1 = norm1(s["r1"])
y2 = norm2(s["r2"])
if i % learn_every:
read1.train(s["rand1"], y2, call=False)
read2.train(s["rand2"], y1, call=False)
"""
# then, train the model
model.train(X[transients:train_len], Y={"r1": teacher1, "r2": teacher2}, learn_every=learn_every, force_teachers=False)
# test the model
res = model.run(X[train_len:])
fig, (ax0, ax1) = plt.subplots(2, 1, sharex=True)
ax0.imshow(X[train_len:].T, aspect="auto", interpolation="none")
ax1.plot(res["r1"], label="R1")
ax1.plot(res["r2"], label="R2")
plt.legend()
plt.show()
from reservoirpy import __version__
print(__version__)
```
| github_jupyter |
# A quick overview on preprocessing
These materials are mostly borrowed from Lane et al. (2019)
As usual, let us first import all the dependencies
```
import re
```
## Tokenisation
```
txt = "Thomas Jefferson started building Monticello at the age of 26."
```
A simple "tokeniser" which gets only alphabetical characters.
```
tokens = re.findall('[A-Za-z]+', txt)
print(tokens)
```
Python provides a ``similar'' tool to tokenise but, in general, it is not enough
```
tokens = txt.split()
print(tokens)
```
Obviously, we can design a better regular expression
```
tokens = re.split(r'([-\s.,;!?])+', txt)
print(tokens)
```
The community has created multiple libraries for pre-processing, which include options for tokenisation. One of the most popular ones is [NLTK](http://www.nltk.org).
Before using it, you should install it. If using pip, you should do:
\$ pip install --user -U nltk
\$ pip install --user -U numpy
An now we can import and use one of its tokenisers
```
from nltk.tokenize import TreebankWordTokenizer # import one of the many tokenizers available
tokenizer = TreebankWordTokenizer() # invoke it
tokens = tokenizer.tokenize(txt)
print(tokens)
```
Now, see the difference between tokenising with split() and with NLTK's treebank tokeniser on a different sentence.
```
sentence = "Monticello wasn't designated as UNESCO World Heritage Site until 1987."
tokens_split = sentence.split()
tokens_tree = tokenizer.tokenize(sentence)
print("OUTPUT USING split()\t\t", tokens_split)
print("OUTPUT USING TreebankWordTokenizer\t", tokens_tree)
```
## Normalisation
### Casefolding
```
sentence = sentence.lower()
print(sentence)
```
## Stemming
Once again, we can use a regular expression to do stemming
```
def stem(phrase):
return ' '.join([re.findall('^(.*ss|.*?)(s)?$',
word)[0][0].strip("'") for word in phrase.lower()
.split()])
print("'houses' \t\t->", stem('houses'))
print("'Doctor House's calls' \t->", stem("Doctor House's calls"))
print("'stress' \t\t->", stem("stress"))
```
But we would need to include many more expressions to deal with all cases and exceptions.
Instead, once again we can rely on a library. Let's consider the **Porter stemmer**, available in NLTK.
```
from nltk.stem.porter import PorterStemmer # Import the stemmer
stemmer = PorterStemmer() # invoke the stemmer
# Notice that we are "tokenising" and stemming in one line
x = ' '.join([stemmer.stem(w).strip("'") for w in "dish washer's washed dishes".split()])
print(x.split())
```
## Lemmatisation
This is a more complex process, compared to stemming. Let us go straight to use a library.
In this particular case we are going to use NLTK's WordNet lemmatiser. If it is the first time you use it (or you are in an ephemeral environment!), you should download it as follows:
```
import nltk
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer # importing the lemmatiser
lemmatizer = WordNetLemmatizer() # invoking it
print("'better' alone \t->",lemmatizer.lemmatize("better"))
print("'better' including it's part of speech (adj) \t->",lemmatizer.lemmatize("better", pos="a"))
```
## A quick overview on representations
### Bag of Words (BoW)
First, let us see a simple construction, using a dictionary
```
sentence = """Thomas Jefferson began building Monticello at the age of 26. Thomas"""
sentence_bow = {}
for token in sentence.split():
sentence_bow[token] = 1
sorted(sentence_bow.items())
```
Another option would be using **pandas**
```
import pandas as pd
# Loading the corpus
sentences = """Thomas Jefferson began building Monticello at the age of 26.\n"""
sentences += """Construction was done mostly by local masons and carpenters.\n"""
sentences += "He moved into the South Pavilion in 1770.\n"
sentences += """Turning Monticello into a neoclassical masterpiece was Jefferson's obsession."""
# Loading the tokens into a dictionary (notice that we asume that each line is a document)
corpus = {}
for i, sent in enumerate(sentences.split('\n')):
corpus['sent{}'.format(i)] = dict((tok, 1) for tok in
sent.split())
# Loading the dictionary contents into a pandas dataframe.
df = pd.DataFrame.from_records(corpus).fillna(0).astype(int).T
# SEE THE .T, which transposes the matrix for visualisation purposes.
df[df.columns[:10]]
```
### One-hot vectors
This is our input sentence (and its vocabulary)
```
import numpy as np
sentence = "Thomas Jefferson began building Monticello at the age of 26."
token_sequence = str.split(sentence)
vocab = sorted(set(token_sequence))
print(vocab)
```
And now, we produce the one-hot representation
```
num_tokens = len(token_sequence)
vocab_size = len(vocab)
onehot_vectors = np.zeros((num_tokens, vocab_size), int) # create the |tokens| x |vocabulary size| matrix of zeros
for i, word in enumerate(token_sequence):
onehot_vectors[i, vocab.index(word)] = 1 # set one to right dimension to 1
print("Vocabulary:\t", vocab)
print("Sentence:\t", token_sequence)
onehot_vectors
```
Let us bring pandas into the game
```
pd.DataFrame(onehot_vectors, columns=vocab)
```
| github_jupyter |
# Using AI Platform Pipelines (Hosted Kubeflow Pipelines) from a notebook
[Cloud AI Platform Pipelines](https://cloud.google.com/ai-platform/pipelines/docs) , currently in Beta, provides a way to deploy robust, repeatable machine learning pipelines along with monitoring, auditing, version tracking, and reproducibility, and gives you an easy-to-install, secure execution environment for your ML workflows. AI Platform Pipelines is based on [Kubeflow Pipelines](https://www.kubeflow.org/docs/pipelines/) (KFP) installed on a [Google Kubernetes Engine (GKE)](https://cloud.google.com/kubernetes-engine) cluster, and can run pipelines specified via both the KFP and TFX SDKs. See [this blog post](https://cloud.google.com/blog/products/ai-machine-learning/introducing-cloud-ai-platform-pipelines) for more detail on the Pipelines tech stack. You can create an AI Platform Pipelines installation with just a few clicks. After installing, you access AI Platform Pipelines by visiting the AI Platform Panel in the [Cloud Console](https://console.cloud.google.com/).
In this notebook:
- First we'll define a KFP pipeline based on predefined [_reusable components_](https://www.kubeflow.org/docs/pipelines/sdk/component-development/), and launch a pipeline run from the notebook.
- Then we'll show how to define a new component that's based on a python function-- a so-called 'lightweight component'.
- The notebook will also show how to grab the ID of a pipeline previously uploaded to the dashboard and use that to kick off a run.
- Finally, we'll show how to trigger a pipeline run when new data becomes available, via Cloud Functions.
## Set up to run this example
### Create an AI Platform Notebooks instance
If you're not doing so already, run this notebook on an AI Platform Notebook instance. See setup instructions [here](https://cloud.google.com/ai-platform/notebooks/docs). (It's possible to run the notebook using other Jupyter environments, but that requires some additional auth setup that we won't cover here). Once your notebook instance is set up, you should be able to use [this link](https://console.cloud.google.com/ai-platform/notebooks/deploy-notebook?name=KFP%20from%20a%20notebook&download_url=https%3A%2F%2Fraw.githubusercontent.com%2Famygdala%2Fcode-snippets%2Fmaster%2Fml%2Fnotebook_examples%2Fcaipp%2Fkfp_in_a_notebook.ipynb&url=https%3A%2F%2Fgithub.com%2Famygdala%2Fcode-snippets%2Fblob%2Fmaster%2Fml%2Fnotebook_examples%2Fcaipp%2Fkfp_in_a_notebook.ipynb) to upload the notebook.
### Install AI Platform Pipelines
The example assumes a Pipeline Installation set up as described [here](https://github.com/amygdala/code-snippets/tree/master/ml/kubeflow-pipelines/keras_tuner) (however, for this notebook you won't need the preemptible node pool, so you can skip that part if you like). The training step in the workflow does assume a GPU-enabled cluster node, though as indicated below, you can edit the pipeline definition to change that.
### Install the KFP SDK
Next, we'll install the KFP SDK, and then restart the kernel so it's available for import.
```
!pip install --user -U kfp kfp-server-api
# Restart kernel after the installs
import IPython
IPython.Application.instance().kernel.do_shutdown(True)
```
## Define and run a pipeline
Now we'll create a pipeline to train a model, then serve it using [TF-serving](xxx). We'll be training a [Keras](xxx) model to predict duration of London bike rentals given info about the start and end station as well as day of week, current weather, and other info. See [this README](xxx) for more detail.
First we'll do some imports:
```
import kfp # the Pipelines SDK.
from kfp import compiler
import kfp.dsl as dsl
import kfp.gcp as gcp
import kfp.components as comp
```
We'll use KFP [reusable components](https://www.kubeflow.org/docs/pipelines/sdk/component-development/) to construct the pipeline:
```
train_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/amygdala/code-snippets/master/ml/kubeflow-pipelines/keras_tuner/components/train_component.yaml'
)
serve_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/amygdala/code-snippets/master/ml/kubeflow-pipelines/keras_tuner/components/serve_component.yaml'
)
tb_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/master/components/tensorflow/tensorboard/prepare_tensorboard/component.yaml'
)
```
Now we'll define the pipeline itself, using the above component definitions.
This pipeline first sets up a TensorBoard visualization for monitoring the training run. Then it starts the training. Once training is finished, it uses TF-serving to set up a service on the Pipelines GKE cluster.
If you don't want to run the training step on GPUs, you can comment out the `train.set_gpu_limit(2)` line below.
```
@dsl.pipeline(
name='bikes_weather',
description='Model bike rental duration given weather'
)
def bikes_weather(
train_epochs: int = 5,
working_dir: str = 'gs://YOUR/GCS/PATH', # for the full training jobs
data_dir: str = 'gs://aju-dev-demos-codelabs/bikes_weather/',
steps_per_epoch: int = -1 , # if -1, don't override normal calcs based on dataset size
hptune_params: str = '[{"num_hidden_layers": %s, "learning_rate": %s, "hidden_size": %s}]' % (3, 1e-2, 64)
):
# create TensorBoard viz for the training run
tb_viz = tb_op(
log_dir_uri='%s/%s' % (working_dir, dsl.RUN_ID_PLACEHOLDER)
)
train = train_op(
data_dir=data_dir,
workdir='%s/%s' % (tb_viz.outputs['log_dir_uri'], 0),
tb_dir=tb_viz.outputs['log_dir_uri'],
epochs=train_epochs, steps_per_epoch=steps_per_epoch,
hp_idx=0,
hptune_results=hptune_params
)
serve = serve_op(
model_path=train.outputs['train_output_path'],
model_name='bikesw',
namespace='default'
)
train.set_gpu_limit(2)
```
You can see that data is being passed between the pipeline ops. [Here's a tutorial](https://gist.github.com/amygdala/bfa0f599a4814b3261367f558a852bfe) that goes into how that works in more detail.
Next we'll compile the pipeline (creating a local archive file):
```
compiler.Compiler().compile(bikes_weather, 'bikes_weather.tar.gz')
```
Then create a client object, and using that client, create (or get) an _Experiment_ (which lets you create semantic groupings of pipeline runs).
You'll need to set the correct host endpoint for your pipelines installation when you create the client. Visit the [Pipelines panel in the Cloud Console](https://console.cloud.google.com/ai-platform/pipelines/clusters) and click on the **SETTINGS** gear for the desired installation to get its endpoint.
```
# CHANGE THIS with the info for your KFP cluster installation
client = kfp.Client(host='xxxxxxxx-dot-us-centralx.pipelines.googleusercontent.com')
exp = client.create_experiment(name='bw_expers') # this is a 'get or create' call
```
Next, set some pipeline params, and run the pipeline, passing it the path to the compiled file:
```
WORKING_DIR = 'gs://YOUR_GCS/PATH'
TRAIN_EPOCHS = 2
run = client.run_pipeline(exp.id, 'bw_test', 'bikes_weather.tar.gz',
params={'working_dir': WORKING_DIR, 'train_epochs': TRAIN_EPOCHS})
```
Once you've kicked off the run, click the generated link to see the pipeline run in the Kubeflow Pipelines dashboard of your pipelines installation. (See the last section of this notebook for more info on how to use your trained and deployed model for prediction).
**Note**: It's also possible to start a pipeline run directly from the pipeline function definition, skipping the local compilation, like this:
```python
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(<pipeline_function_name>, arguments={})
```
One thing that might have occurred to you with the example thus far: what if the trained model's accuracy is not that great, and we don't necessarily want to deploy it? We'll address that in the next section.
## Defining a new 'lightweight component' based on a python function
'Lightweight' KFP python components allow you to create a component from a python function definition, and do not require you to build a new container image for every code change. They're helpful for fast iteration in a notebook environment. You can read more [here](https://github.com/kubeflow/pipelines/blob/master/samples/core/lightweight_component/lightweight_component.ipynb).
In this section, we'll create a lightweight component that uses training metrics info to decide whether to deploy a model.
We'll pass a "threshold" dict as a component arg, and compare those thresholds to the metrics values, and use that info to decide whether or not to deploy. Then we'll output a string indicating the decision.
(This is a pretty simple approach, that we're using for illustrative purposes; for production models you'd probably want to do more sophisticated analyses. The [TFMA library](https://www.tensorflow.org/tfx/model_analysis/get_started) might be of interest).
Then we'll update the pipeline to use the new component. In the new pipeline definition below, we'll make the 'serve' step conditional on that output.
First, we'll define the component function, `eval_metrics`:
```
from typing import NamedTuple
def eval_metrics(
metrics: str,
thresholds: str
) -> NamedTuple('Outputs', [('deploy', str)]):
import json
import logging
def regression_threshold_check(metrics_info):
# ...
for k, v in thresholds_dict.items():
logging.info('k {}, v {}'.format(k, v))
if k in ['root_mean_squared_error', 'mae']:
if metrics_info[k][-1] > v:
logging.info('{} > {}; returning False'.format(metrics_info[k][0], v))
return ('False', )
return ('deploy', )
logging.getLogger().setLevel(logging.INFO)
thresholds_dict = json.loads(thresholds)
logging.info('thresholds dict: {}'.format(thresholds_dict))
logging.info('metrics: %s', metrics)
metrics_dict = json.loads(metrics)
logging.info("got metrics info: %s", metrics_dict)
res = regression_threshold_check(metrics_dict)
logging.info('deploy decision: %s', res)
return res
```
Next we'll create a 'container op' from that function definition, via the `funct_to_container_op` method. As one of the method args, we specify the base container image that will run the function.
Here, we're using one of the [Deep Learning Container images](https://cloud.google.com/ai-platform/deep-learning-containers/docs/). (Admittedly, this is overkill for this simple function).
```
eval_metrics_op = comp.func_to_container_op(eval_metrics, base_image='gcr.io/deeplearning-platform-release/tf2-cpu.2-3:latest')
```
Now, we can define a new pipeline that uses the new op and makes the model serving conditional.
The new `eval_metrics_op` takes as an input one of the `train_op` outputs, which outputs a metrics dict. (We "cheated" a bit, as the training component was already designed to output this info; in other cases you might end up defining a new version of such an op that outputs the new info you need).
Then, we'll wrap the serving op in a conditional; we won't set up a TF-serving service unless the `eval_metrics` op has certified that it is okay.
Note that this new version of the pipeline also has a new input parameter— the `thresholds` dict.
```
@dsl.pipeline(
name='bikes_weather_metrics',
description='Model bike rental duration given weather'
)
def bikes_weather_metrics(
train_epochs: int = 2,
working_dir: str = 'gs://YOUR/GCS/PATH', # for the full training jobs
data_dir: str = 'gs://aju-dev-demos-codelabs/bikes_weather/',
steps_per_epoch: int = -1 , # if -1, don't override normal calcs based on dataset size
hptune_params: str = '[{"num_hidden_layers": %s, "learning_rate": %s, "hidden_size": %s}]' % (3, 1e-2, 64),
thresholds: str = '{"root_mean_squared_error": 2000}'
):
# create TensorBoard viz for the parent directory of all training runs, so that we can
# compare them.
tb_viz = tb_op(
log_dir_uri='%s/%s' % (working_dir, dsl.RUN_ID_PLACEHOLDER)
)
train = train_op(
data_dir=data_dir,
workdir='%s/%s' % (tb_viz.outputs['log_dir_uri'], 0),
tb_dir=tb_viz.outputs['log_dir_uri'],
epochs=train_epochs, steps_per_epoch=steps_per_epoch,
hp_idx=0,
hptune_results=hptune_params
)
eval_metrics = eval_metrics_op(
thresholds=thresholds,
metrics=train.outputs['metrics_output_path'],
)
with dsl.Condition(eval_metrics.outputs['deploy'] == 'deploy'):
serve = serve_op(
model_path=train.outputs['train_output_path'],
model_name='bikesw',
namespace='default'
)
train.set_gpu_limit(2)
```
Now, as before, we can compile and then run the pipeline.
```
compiler.Compiler().compile(bikes_weather_metrics, 'bikes_weather_metrics.tar.gz')
run2 = client.run_pipeline(exp.id, 'bw_metrics_test', 'bikes_weather_metrics.tar.gz',
params={'working_dir': WORKING_DIR, 'train_epochs': TRAIN_EPOCHS
# 'thresholds': THRESHOLDS
})
```
Again, once you've kicked off the run, click the link that will appear above to view the run in the Kubeflow Pipelines dashboard.
## Starting a pipeline run given the pipeline's ID
In the sections above, we compiled a pipeline definition to a local archive files, then passed the path to the file as part of the `run_pipeline` method.
It's also possible to start a pipeline run given the ID (not display name) of an already-uploaded pipeline. You can find these IDs via the Kubeflow Pipeline dashboard, but it's also possible to grab them programmatically.
As an example, we'll define a utility function to grab a pipeline's ID given its name, then use the given ID in the `run_pipeline` call.
> Note: Rather confusingly, with the the `run_pipeline` calls above, the created pipelines aren't actually given a display name (though they still have an ID). So the `get_pipeline_id` utility defined below is designed to be used for pipelines explicitly uploaded to the dashboard from the "Pipelines" tag, _not_ pipelines defined as we've done above in this notebook. You can visit the dashboard to get the IDs of the pipelines you've just created, or find them in the list of all pipelines.
```
# you can get a list of all the pipelines like this...
# pipelines_list = client.list_pipelines()
# pipelines_list
import json
_FILTER_OPERATIONS = {"UNKNOWN": 0,
"EQUALS" : 1,
"NOT_EQUALS" : 2,
"GREATER_THAN": 3,
"GREATER_THAN_EQUALS": 5,
"LESS_THAN": 6,
"LESS_THAN_EQUALS": 7}
def get_pipeline_id(client, name):
"""Find the id of a pipeline by name.
Args:
name: Pipeline name.
Returns:
Returns the pipeline id if a pipeline with the name exists.
"""
pipeline_filter = json.dumps({
"predicates": [
{
"op": _FILTER_OPERATIONS["EQUALS"],
"key": "name",
"stringValue": name,
}
]
})
result = client._pipelines_api.list_pipelines(filter=pipeline_filter)
if result.pipelines is None:
return None
if len(result.pipelines)==1:
return result.pipelines[0].id
elif len(result.pipelines)>1:
raise ValueError("Multiple pipelines with the name: {} found, the name needs to be unique".format(name))
return None
```
The following function call assumes the existence of a named pipeline that you've uploaded to the Kubeflow Pipelines dashboard:
```
# Replace with the name of your pipeline.
get_pipeline_id(client, 'YOUR_PIPELINE_NAME')
```
**Edit the following cell** to use the output pipeline ID. (And modify the params dict if you're using a different pipeline).
```
run3 = client.run_pipeline(exp.id, 'bw_metrics_test2', pipeline_id='YOUR_PIPELINE_ID',
params={'working_dir': WORKING_DIR, 'train_epochs': TRAIN_EPOCHS
})
```
## Triggering a pipeline run when new data comes in
Often, you may want to trigger a new run of the training pipeline when new data becomes available.
We can use [Cloud Functions](https://cloud.google.com/functions/docs/) (GCF) to do this. This section walks through how you could set this up.
> Note: Depending upon ML workflow context, in actuality you might want to fine-tune an existing model, or retrain on a different window of data than you did originally. For this example we're keeping things simple and just triggering another full model training job on the dataset after new data is added.
In this notebook, **we're skipping some of the prereqs, including some required auth setup. See [this notebook](https://github.com/amygdala/code-snippets/blob/master/ml/notebook_examples/functions/hosted_kfp_gcf.ipynb) for the details.**
The first step is to identify a GCS 'trigger bucket'. **It should be a new bucket or one that you don't use for anything else**. We'll set things up so that if any objects are added to this bucket, or modified, the GCF function will run, and it in turn will deploy a run of the pipeline. Edit and run the following to set your bucket:
```
%env TRIGGER_BUCKET=YOUR_TRIGGER_BUCKET
```
Next, copy the input directory used with the example model to your own trigger bucket, which you can do as follows:
```
%%bash
gsutil cp 'gs://aju-dev-demos-codelabs/bikes_weather/*' gs://${TRIGGER_BUCKET}/bikes_weather_temp
gsutil ls gs://${TRIGGER_BUCKET}/bikes_weather_temp
```
Then, delete one of the files from your temp data directory (we'll shortly add it back to trigger the GCF function):
```
%%bash
gsutil rm gs://${TRIGGER_BUCKET}/bikes_weather_temp/train-bw000000000007.csv
gsutil ls gs://${TRIGGER_BUCKET}/bikes_weather_temp
```
Next, we'll create a subdirectory for the GCF definition:
```
%%bash
mkdir -p functions
```
We'll first create a requirements.txt file, to indicate what packages the GCF code requires to be installed.
```
%%writefile functions/requirements.txt
kfp
```
Next we'll define the GCF function.
**Before you run the next cell, edit** the `HOST`, `WORKING_DIR`, `DATA_DIR`, and `PIPELINE_ID` values. Set `DATA_DIR` to the path under which your copied input files live, e.g. `gs://${TRIGGER_BUCKET}/bikes_weather_temp/`. Include the trailing slash. Set `PIPELINE_ID` to the ID of the pipeline that you want to run. Modify the params dict as necessary for the pipeline you're running.
```
%%writefile functions/main.py
import logging
import datetime
import logging
import time
import kfp
import kfp.compiler as compiler
import kfp.dsl as dsl
import requests
# TODO: replace with your Pipelines endpoint URL
HOST = 'xxxxxxxx-dot-us-centralx.pipelines.googleusercontent.com'
# TODO: replace with your working dir
WORKING_DIR = 'gs://YOUR_WORKING_DIR'
TRAIN_EPOCHS = 2
DATA_DIR = 'gs://PATH_TO/YOUR_COPIED_INPUT_DATA/' # include the trailing slash
PIPELINE_ID = 'YOUR_PIPELINE_ID'
def get_access_token():
url = 'http://metadata.google.internal/computeMetadata/v1/instance/service-accounts/default/token'
r = requests.get(url, headers={'Metadata-Flavor': 'Google'})
r.raise_for_status()
access_token = r.json()['access_token']
return access_token
def hosted_kfp_test(data, context):
logging.info('Event ID: {}'.format(context.event_id))
logging.info('Event type: {}'.format(context.event_type))
logging.info('Data: {}'.format(data))
logging.info('Bucket: {}'.format(data['bucket']))
logging.info('File: {}'.format(data['name']))
file_uri = 'gs://%s/%s' % (data['bucket'], data['name'])
logging.info('Using file uri: %s', file_uri)
logging.info('Metageneration: {}'.format(data['metageneration']))
logging.info('Created: {}'.format(data['timeCreated']))
logging.info('Updated: {}'.format(data['updated']))
token = get_access_token()
logging.info('attempting to launch pipeline run.')
ts = int(datetime.datetime.utcnow().timestamp() * 100000)
client = kfp.Client(host=HOST, existing_token=token)
exp = client.create_experiment(name='gcstriggered') # this is a 'get or create' op
res = client.run_pipeline(exp.id, 'bwmetrics_' + str(ts),
pipeline_id=PIPELINE_ID,
params={'working_dir': WORKING_DIR, 'train_epochs': TRAIN_EPOCHS,
'data_dir': DATA_DIR}
)
logging.info(res)
```
Deploy the GCF function as follows. (You'll need to wait a moment or two for output of the deployment to display in the notebook). You can also run this command from a notebook terminal window in the functions subdirectory.
```
%%bash
cd functions
gcloud functions deploy gcs_test --runtime python37 --trigger-resource ${TRIGGER_BUCKET} --trigger-event google.storage.object.finalize
```
Once the Cloud Function is successfully deployed, trigger the pipeline run by adding a new file to the `DATA_DIR`, e.g.:
```
%%bash
gsutil cp gs://aju-dev-demos-codelabs/bikes_weather/train-bw000000000007.csv gs://${TRIGGER_BUCKET}/bikes_weather_temp/train-bw000000000007.csv
```
You should see the newly launched pipeline running in the Kubeflow Pipelines dashboard.
> **Important note**: For simplicity, this example is a bit unrealistic in that for each file added to or updated in the bucket, a new separate Pipeline run is launched. So it would not suit for cases where multiple new files are added to a directory at once. A more general solution might keep the data directory separate from the trigger bucket, monitor it for new files, and periodically write a 'new file' notification to the trigger bucket.
(So, once you have your Cloud Function trigger set up, be careful of copying multiple new files to the trigger bucket at once).
## More detail on the code, and requesting predictions from your model
This notebook didn't focus on the details of the pipeline component (step) implementations. The training component uses a Keras model (TF 2.3). The serving component uses [TF-serving](https://www.tensorflow.org/tfx/guide/serving): once the serving service is up and running, you can send prediction requests to your trained model.
You can find more detail on these components, and an example of sending a prediction request, [here](https://github.com/amygdala/code-snippets/tree/master/ml/kubeflow-pipelines/keras_tuner).
----------------------------
Copyright 2020, Google, LLC. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
import numpy as np
import qiskit
import tqix
import sys, math
sys.path.insert(1, '../')
import qtm.constant, qtm.base
n = 5
def calculate_TML(trial_state):
###################
## Calculate Pr
###################
if (trial_state) is None:
return -100
trial_state = np.expand_dims(trial_state, 1)
model = tqix.qmeas(trial_state, 'MUB')
pr = model.probability()
for i in range(0, len(pr)):
if pr[i] == 0:
pr[i] = 10**(-10)
###################
## Calculate fr
###################
model = tqix.qmeas(state, 'MUB')
cdf_model = tqix.qsim(model, niter = int(qtm.constant.num_shots / len(pr)), backend='cdf')
fr = np.real(cdf_model.probability())
###################
## Calculate Tml
###################
tml = 0
for i in range(0, len(pr)):
tml += np.real(fr[i]*np.log(pr[i]))
return -tml
# state = tqix.random(4)
state = 2*np.random.rand(n)-1
state = state / np.linalg.norm(state)
state = np.expand_dims(state, axis = 0)
print(state)
from math import cos, atan
import numpy as np
from scipy.optimize import minimize
from scipy import optimize
from scipy.optimize import NonlinearConstraint, Bounds
def constraint(x):
"""Normalize condition
Args:
x (np.narray): state vector psi
Returns:
float: norm of psi
"""
norm = 0
for i in x:
norm += i**2
return norm
def create_bound(n):
bounds = []
for i in range(0, n):
bounds.append((-1, 1))
return bounds
progress = []
progress_val = []
def cb(x, convergence):
progress.append(x)
progress_val.append(calculate_TML(x))
nlc = NonlinearConstraint(constraint, -1.0, 1.0)
results = optimize.differential_evolution(calculate_TML, create_bound(n), constraints=(nlc), callback=cb)
progress = np.array(progress)
progress_val = np.array(progress_val)
state1 = np.squeeze(state, axis = 0)
trace, fidelity = qtm.base.get_metrics(state1, results.x)
print(trace)
print(fidelity)
print(state1)
print(results.x)
plt.ylabel("$-T_{ML}$")
plt.xlabel("$n_{iter}$")
plt.plot(progress_val)
```
### Randomness method
```
num_qubits = 2
from matplotlib import pyplot as plt
# psi = 2*np.random.rand(2**num_qubits)-1
# psi = psi / np.linalg.norm(psi)
# qc = qiskit.QuantumCircuit(num_qubits, num_qubits)
# qc.initialize(psi, range(0, num_qubits))
# state = qiskit.quantum_info.Statevector(qc).data
# state = np.real(state.reshape((2**num_qubits, 1)))
state = tqix.random(4)
max_tml = 0
tmls = []
iterations = 0
fidelities = []
for i in range(0, 10000000):
# trial_state = np.random.uniform(low = 0, high = 2*np.pi, size = (2**num_qubits, 1))
# trial_state = trial_state / np.linalg.norm(trial_state)
trial_state = tqix.random(4)
tml = calculate_TML(state, trial_state)
tmls.append(tml)
if i == 0 or max_tml < tml:
max_tml = tml
reconstructed_state = trial_state
iterations += 1
if iterations % 10 == 0:
print(iterations)
# if fidelity > 0.99:
# print('Founded')
# break
if tml >= 1 or iterations == 400:
break
state1 = np.squeeze(state, axis = 1)
reconstructed_state1 = np.squeeze(reconstructed_state, axis = 1)
trace, fidelity = qtm.base.get_metrics(state1, reconstructed_state1)
fidelities.append(fidelity)
print(tml)
print("State: ", state)
print("Reconstructed state: ", reconstructed_state)
plt.plot(fidelity)
```
first, we change ghz(1) by Haar random called from Qiskit. let fix N = 2.
then we calculate the cdf_model probabilites so, now we have f_r.
len(f_r) = (2^n + 1)*2^n
f_r(1) = (2^1 + 1)*2
Next, we calculate the trial probabilities, to calculate the trial probabilities, you can call the probability from model (not cdf_model)
1. We have $f_r$ based on measure random unknown state
2. For each $|\psi_{trial}\rangle$, we have different $P_r$.
3. And $T_{ML}(\psi_{trial})=\Sigma (f_r.\log(P_R)) \in C$?
We repeat 2-3 until $T_{ML}(\psi_{trial})$ max and $\psi_{trial}$ is $|\mu\rangle$
<img width = 500px src = '../../images/classical_tomography.jpg'/>
| github_jupyter |
<img src="images/numpy/numpy.png" style="width:50%;height:50%;">
### 1. Giới thiệu
Numpy là một thư viện của Python hỗ trợ việc tính toán trên mảng nhiều chiều. Một mảng Numpy là một tập hợp các giá trị cùng kiểu dữ liệu và được đánh số bằng các số nguyên dương. Numpy là Module quan trọng cho việc sử lý dữ liệu và có thể chuyển đổi qua kiểu dữ liệu Tensor trong Tensorflow và Pytorch.
Mọi chi tiết về Numpy đều được công bố trên trang chủ chính thức: https://numpy.org/doc/
Các bạn có thể cài đặt Numpy theo hướng dẫn dưới đây:
- [Dành cho máy Mac và Linux](https://machinelearningcoban.com/faqs/#-huong-dan-cai-dat-python-va-cac-thu-vien-tren-macos)
- [Dành cho Windows](https://machinelearningcoban.com/faqs/#-huong-dan-cai-dat-python-va-cac-thu-vien-tren-windows)
Sau khi cài đặt xong, chúng ta cần phải import thư viện vào để sử dụng:
```
import numpy as np
```
Từ khoá `as` trong Python giúp chúng ta viết gọn tên của thư viện giúp tiện lợi cho việc sử dụng về sau.
### 2. Nội dung
Nội dung chính trong bài ngày hôm nay chúng ta sẽ cùng tìm hiểu về:
- Tạo mảng Numpy (ndarray)
- Kiểu dữ liệu (Datatypes)
- Array Indexing
- Phép toán trên mảng
- Broadcasting
#### 2.1 Tạo mảng Numpy (ndarray)
Tạo ndarray từ List
```
# Tạo ndarray từ list
import numpy as np
# tạo list
l = list(range(1, 4))
# tạo ndarray
data = np.array(l)
print(data)
print(data[0], data[1])
```
<img src="images/numpy/chap5_np_1.png" style="width:20%;height:20%;">
Sử dụng thuộc tính `shape` và hàm `type()` cho mảng Numpy
```
# Tạo ndarray từ list
import numpy as np
# tạo list
l = list(range(1, 4))
# tạo ndarray
data = np.array(l)
print(data)
print(type(data))
print(type(data[0]))
print(data.shape)
```
Thay đổi `shape` của mảng sử dụng `reshape`
```
# thay đổi shape của một mảng
import numpy as np
arr1 = np.arange(12)
print(arr1.shape)
arr2 = arr1.reshape((3, 4))
print(arr2.shape)
```
Thay đổi giá trị của một phần tử
```
# thay đổi giá trị phần tử
import numpy as np
l = list(range(1, 4))
data = np.array(l)
print(data)
data[0] = 8
print(data)
```
<img src="images/numpy/chap5_np_2.png" style="width:20%;height:20%;">
Tạo ndarray với hàm `zeros()`
```
# tạo một numpy array với tất cả các phần tử là 0
import numpy as np
# shape: 2 dòng, 3 cột
arr = np.zeros((2, 3))
print(arr)
```
<img src="images/numpy/chap5_np_3.png" style="width:20%;height:20%;">
Tạo ndarray với hàm `ones()`
```
# tạo một numpy array với tất cả phần tử là 1
import numpy as np
# numpy.ones(shape, dtype=None, order='C')
# shape: 2 dòng, 3 cột
arr = np.ones((2, 3))
print(arr)
```
<img src="images/numpy/chap5_np_4.png" style="width:20%;height:20%;">
Tạo ndarray với hàm `full()`
```
# tạo một numpy array với tất cả phần tử là hằng số K
import numpy as np
# numpy.full(shape, fill_value, dtype=None, order='C')
# shape: 2 dòng, 3 cột - với K = 9
arr = np.full((2, 3), 9)
print(arr)
```
<img src="images/numpy/chap5_np_5.png" style="width:20%;height:20%;">
Tạo ma trận đường chéo
```
# tạo một numpy array với đường chéo là số 1
# số 0 được điền vào những ô phần tử còn lại
import numpy as np
# numpy.eye(N, M=None, k=0, dtype=<class 'float'>, order='C')
# shape: 2 dòng, 3 cột
arr = np.eye(3)
print(arr)
```
<img src="images/numpy/chap5_np_6.png" style="width:20%;height:20%;">
Tạo một numpy array với giá trị ngẫu nhiên
```
# tạo một numpy array với giá trị ngẫu nhiên
import numpy as np
# numpy.random.random(size=None)
# shape: 2 dòng, 3 cột; với phần tử có giá trị ngẫu nhiên
arr = np.random.random((2,3))
print(arr)
```
Điều kiện cho mảng numpy
```
import numpy as np
arr = np.arange(10)
print(arr)
out = np.where(arr < 5, arr, 10*arr)
print(out)
```
Chuyển mảng về một chiều
```
import numpy as np
arr = np.array([[1, 2], [3, 4]])
out = arr.flatten()
print(arr)
print(out)
```
#### 2.2 Kiểu dữ liệu (Datatypes)
Mảng numpy chứa các phần tử cùng kiểu dữ liệu. Numpy cung cấp một tập hợp các kiểu dữ liệu mà chúng có thể sử dụng để xây dựng các mảng.
```
import numpy as np
# int32
arr1 = np.array([1, 2])
print(arr1.dtype)
# float64
arr2 = np.array([1.0, 2.0])
print(arr2.dtype)
# int64
arr3 = np.array([1, 2], dtype = np.int64)
print(arr3.dtype)
```
#### 2.3 Array Indexing
Numpy cung cấp một số cách để truy xuất phần tử trong mảng. Truy xuất phần tử dùng kỹ thuật slicing tương tự như danh sách (list) trong Python.
**Ví dụ 1:** Lấy các phần tử từ mảng 2 chiều như sau:
```
import numpy as np
# Khởi tạo numpy array có shape = (2, 3) có giá trị như sau:
# [[ 1 2 3]
# [ 4 6 7]]
a_arr = np.array([[1,2,3],[5,6,7]])
print('a_arr: \n', a_arr)
# Sử dụng slicing để tạo mảng b bằng cách lấy 2 hàng đầu tiên
# và cột 1, 2. Như vậy b sẽ có shape = (2, 3):
# [[2 3]
# [6 7]]
b_arr = a_arr[:, 1:3]
print('b_arr: \n', b_arr)
```
<img src="images/numpy/chap5_np_9.png" style="width:30%;height:30%;">
Việc một mảng mới được tạo ra từ Slicing sẽ có **cùng địa chỉ** với mảng gốc. Nếu thay đổi một trong hai mảng này thì mảng còn lại cũng thay đổi theo.
```
import numpy as np
# Khởi tạo numpy array có shape = (2, 3) có giá trị như sau:
# [[ 1 2 3]
# [ 4 6 7]]
a_arr = np.array([[1,2,3],[5,6,7]])
print('a_arr: \n', a_arr)
# Sử dụng slicing để tạo mảng b bằng cách lấy 2 hàng đầu tiên
# và cột 1, 2. Như vậy b sẽ có shape = (2, 3):
# [[2 3]
# [6 7]]
b_arr = a_arr[:, 1:3]
print('b_arr: \n', b_arr)
print('Truoc khi thay doi: \n', a_arr[0, 1])
b_arr[0, 0] = 99
print('Sau khi thay doi: \n', a_arr[0, 1])
```
<img src="images/numpy/chap5_np_10.png" style="width:40%;height:40%;">
**Ví dụ 2:** Lấy một dòng dữ liệu
```
import numpy as np
# Tạo một numpy array có shape (3, 4) với giá trị như sau:
# [[ 1 2 3 ]
# [ 5 6 7 ]
# [ 9 10 11 ]]
arr = np.array([[1, 2, 3], [5, 6, 7], [9, 10, 11]])
# Hai cách truy cập dữ liệu ở hàng giữa của mảng
# Cách 1: Dùng kết hợp chỉ số và slice -> được array mới có số chiều thấp hơn
# Cách 2: Nếu chỉ dùng slice ta sẽ có array mới có cùng số chiều với array gốc
# Cách 1: số chiều giảm
row_r1 = arr[1, :]
# Cách 2: số chiều được giữ nguyên
row_r2 = arr[1:2, :]
print(row_r1, row_r1.shape)
print(row_r2, row_r2.shape)
```
<img src="images/numpy/chap5_np_11.png" style="width:40%;height:40%;">
**Ví dụ 3:** Lấy một cột dữ liệu
```
import numpy as np
# Tạo một numpy array có shape (3, 4) với giá trị như sau:
# [[ 1 2 3 ]
# [ 5 6 7 ]
# [ 9 10 11 ]]
arr = np.array([[1, 2, 3], [5, 6, 7], [9, 10, 11]])
# Hai cách truy cập dữ liệu ở cột giữa của mảng
# Cách 1: Dùng kết hợp chỉ số và slice -> được array mới có số chiều thấp hơn
# Cách 2: Nếu chỉ dùng slice ta sẽ có array mới có cùng số chiều với array gốc
# Cách 1: số chiều giảm
col_r1 = arr[:, 1]
# Cách 2: số chiều được giữ nguyên
col_r2 = arr[:, 1:2]
print(col_r1, col_r1.shape)
print(col_r2, col_r2.shape)
```
<img src="images/numpy/chap5_np_13.png" style="width:30%;height:30%;">
**Boolean indexing:** Cho phép chúng ta chọn ra các phần tử tùy ý của một mảng. Kiểu truy xuất này thường được sử dụng để chọn ra các phần tử thỏa mãn điều kiện nào đó.
**Ví dụ 4:** Tìm các vị trí thoả mãn điều kiện
```
import numpy as np
a_arr = np.array([[1, 2], [3, 4], [5, 6]])
print(a_arr)
# Tìm các phần tử lớn hơn 2
# Trả về 1 mảng Boolean có số chiều như mảng a_arr
# và giá trị tại mỗi phần từ là True nếu phần tử của a tại đó > 2,
# False cho trường hợp ngược lại
bool_idx = (a_arr > 2)
print(bool_idx)
```
<img src="images/numpy/chap5_np_15.png" style="width:30%;height:30%;">
**Ví dụ 5:** Tìm các vị trí thoả mãn điều kiện và lấy phần tử tương ứng
```
import numpy as np
a_arr = np.array([[1, 2], [3, 4], [5, 6]])
print(a_arr)
# Tìm các phần tử lớn hơn 2
# Trả về 1 mảng Boolean có số chiều như mảng a_arr
# và giá trị tại mỗi phần từ là True nếu phần tử của a tại đó > 2,
# False cho trường hợp ngược lại
bool_idx = (a_arr > 2)
print('bool_idx: \n', bool_idx)
# Chúng ta sẽ sử dụng boolean array indexing để xây dựng mảng 1 chiều
# Bao gồm các phần tử tương ứng với giá trị True của bool_idx
# Ví dụ ở đây in ra các giá trị của a_arr >2, sử dụng array bool_idx đã tạo
out = a_arr[bool_idx]
print('\nMethod 1:\n', out)
# một cách ngắn gọn hơn
print('\nMethod 2:\n', a_arr[a_arr > 2])
```
<img src="images/numpy/chap5_np_16.png" style="width:50%;height:50%;">
#### 2.4 Phép toán trên mảng
Phép cộng giữa hai mảng
```
import numpy as np
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Tổng của 2 mảng, cả 2 cách đều cho cùng một kết quả
# [[ 6.0 8.0]
# [10.0 12.0]]
print(x + y)
print(np.add(x, y))
```
<img src="images/numpy/chap5_np_18.png" style="width:30%;height:30%;">
Phép trừ giữa hai mảng
```
import numpy as np
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Phép trừ của 2 mảng, cả 2 cách đều cho cùng một kết quả
# [[-4.0 -4.0]
# [-4.0 -4.0]]
print(x - y)
print(np.subtract(x, y))
```
<img src="images/numpy/chap5_np_19.png" style="width:30%;height:30%;">
Phép nhân giữa hai mảng
```
import numpy as np
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Phép nhân
# [[ 5.0 12.0]
# [21.0 32.0]]
print(x * y)
print(np.multiply(x, y))
```
<img src="images/numpy/chap5_np_20.png" style="width:30%;height:30%;">
Phép chia giữa hai mảng
```
import numpy as np
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Phép chia
# [[ 0.2 0.33333333]
# [ 0.42857143 0.5 ]]
print(x / y)
print(np.divide(x, y))
```
<img src="images/numpy/chap5_np_21.png" style="width:30%;height:30%;">
Tính căn bậc 2 cho từng phần tử
```
import numpy as np
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Phép lấy căn
# [[ 1. 1.41421356]
# [ 1.73205081 2. ]]
print(np.sqrt(x))
```
Nhân giữa hai vector (inner product)
```
import numpy as np
v = np.array([9,10])
w = np.array([11, 12])
# Nhân giữa 2 vector, cả 2 đều cho kết quả 219
print(v.dot(w))
print(np.dot(v, w))
```
Nhân giữa vector và ma trận
```
import numpy as np
X = np.array([[1,2],[3,4]])
v = np.array([9,10])
# Nhân giữa Matrix và vector, cả 2 đều cho kết quả array [29 67]
print(X.dot(v))
print(np.dot(X, v))
```
Nhân giữa matrix và matrix
```
import numpy as np
X = np.array([[1,2],[3,4]])
Y = np.array([[5,6],[7,8]])
# Nhân giữa matrix và matrix; cả 2 cách đều cho cùng kết quả
# [[19 22]
# [43 50]]
print(X.dot(Y))
print(np.dot(X, Y))
```
Tính tổng cho một mảng numpy
```
import numpy as np
x = np.array([[1,2],[3,4]])
# Tổng các phần tử của mảng; prints "10"
print(np.sum(x))
# Tính tổng theo từng cột
print(np.sum(x, axis=0))
# Tính tổng theo từng hàng
print(np.sum(x, axis=1))
```
<img src="images/numpy/chap5_np_27.png" style="width:30%;height:30%;">
Chuyển vị cho một ma trận
```
import numpy as np
x = np.array([[1,2], [3,4]])
print(x) # Prints "[[1 2]
# [3 4]]"
print(x.T) # Prints "[[1 3]
# [2 4]]"
```
<img src="images/numpy/chap5_np_29.png" style="width:30%;height:30%;">
```
# Lưu ý rằng việc chuyển vị cho vector sẽ không làm gì cả.
v = np.array([1,2,3])
print(v) # Prints "[1 2 3]"
print(v.T) # Prints "[1 2 3]"
```
#### 2.5 Broadcasting
Broadcasting cho phép thực thi các phép toán trên các mảng có kích thước khác nhau.
Ví dụ: chúng ta muốn cộng vector cho mỗi hàng của ma trận. Chúng ta có thể làm như sau:
```
import numpy as np
X = np.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12]])
v = np.array([1,0,1])
Y = np.empty_like(X)
# Cách 1: Thêm vector v vào mỗi hàng của ma trận X bằng vòng lặp
for i in range(4):
Y[i, :] = X[i, :] + v
print('Matrix X: \n', X)
print('\nVector v: \n', v)
print('\nMatrix Y: \n', Y)
```
Cách này hoạt động bình thường với ma trận X nhỏ. Khi ma trận X lớn, việc sử dụng vòng lặp này sẽ rất chậm.
Chúng ta có thể thực hiện mục đích trên bằng cách xếp chồng nhiều bản sao của v theo chiều dọc, sau đó thực hiện phép tính tổng với X. Chúng ta có thể thực hiện phương pháp này như sau:
```
import numpy as np
X = np.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12]])
v = np.array([1,0,1])
# Xếp chồng 4 bản sao của v lên nhau:
V_t = np.tile(v, (4, 1))
# Thực hiện phép cộng
Y = X + V_t
print('Matrix X: \n', X)
print('\nVector v: \n', v)
print('\nMatrix v: \n', V_t)
print('\nMatrix Y: \n', Y)
```
<img src="images/numpy/chap5_np_31.png" style="width:50%;height:50%;">
Numpy broadcasting cho phép chúng ta thực thi tính toán này mà không cần phải làm thêm các bước thêm nào.
```
import numpy as np
X = np.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12]])
v = np.array([1,0,1])
Y = X + v
print(Y)
```
<img src="images/numpy/chap5_np_32.png" style="width:50%;height:50%;">
### Tài liệu tham khảo
[1] [CS231n Convolutional Neural Networks - Python Numpy Tutorial](http://cs231n.github.io/python-numpy-tutorial/#numpy-datatypes)
| github_jupyter |
```
import xarray as xr
import numpy as np
import matplotlib.ticker as ticker
from matplotlib import pyplot as plt
from matplotlib.dates import DateFormatter
import matplotlib.dates as mdates
import pandas as pd
np.set_printoptions(threshold=np.inf)
dirpath = '/local/data/soccom/SOCCOM_LoResQC_LIAR_05May2021_netcdf/'
f_4180 = xr.open_dataset(dirpath + '5904180QC.nc')
f_4860 = xr.open_dataset(dirpath + '5904860QC.nc')
f_4767 = xr.open_dataset(dirpath + '5904767QC.nc')
f_4180
#print(f_4180.Oxygen.values*100/f_4180.OxygenSat.values)
def get_mld_rho(fl, interp):
density = fl.Sigma_theta.transpose().values
if interp:
density = interp_var(density, fl)
density = np.flip(density, 0)
#density=density.reshape(1,(dim[0]*dim[1]), order='F')
depth = fl.Depth.transpose().values
if interp:
depth = interp_var(depth, fl)
depth = np.flip(depth, 0)
#depth=depth.reshape(1,(dim[0]*dim[1]), order='F')
rhocrit = 0.02
density[np.isnan(depth)] = float("nan")
depth[np.isnan(density)] = float("nan")
d10 = np.absolute(depth-10)
dref = np.empty((1, np.size(density, 1)))
dref[:] = np.NaN
mld = np.empty((1,np.size(density, 1)))
mld[:] = np.NaN
imld = np.empty((1,np.size(density, 1)))
imld[:] = np.NaN
for ii in range(np.size(density,1)):
idx = (d10[:, ii]==np.nanmin(d10[:,ii], axis=0)).argmax()
dref[0,ii]=depth[idx,ii]
if (idx.size != 0):
drho10 = density[idx, ii]
drho10 = np.absolute(density[:,ii] - drho10);
drho10[0:idx] = np.NaN;
idx = (drho10>=rhocrit).argmax()
if (idx.size == 0):
imld[0, ii] = np.NaN
mld[0, ii] = np.Nan
else:
imld[0,ii]=idx;
mld[0,ii]=depth[idx,ii]
else:
imld[0, ii] = np.NaN
mld[0, ii] = np.Nan
imld[mld>900]=np.NaN;
mld[mld>900]=np.NaN;
imld = imld.flatten()
mld = mld.flatten()
end = len(depth)
depth_cat = np.stack((depth[0:end-1,:],depth[1:end,:]),axis=2);
dpth_mid=np.nanmean(depth_cat, axis=2)
depth2 = np.vstack((depth[0,:],dpth_mid,depth[end-1,:]))
thickn=np.diff(depth2,axis=0);
return mld, imld, thickn
def set_oxygen_var(fl):
oxygen = fl.Oxygen.transpose().values
oxygen_qf = fl.Oxygen_QFA.transpose().values
oxygen_qf = oxygen_qf == 0
oxygen[~oxygen_qf] = np.nan
return oxygen
def set_oxygen_sat_var(fl):
oxygen = fl.OxygenSat.transpose().values
oxygen_qf = fl.OxygenSat_QFA.transpose().values
oxygen_qf = oxygen_qf == 0
oxygen[~oxygen_qf] = np.nan
return oxygen
def set_nitrate_var(fl):
nitrate = fl.Nitrate.transpose().values
nitr_qf = fl.Nitrate_QFA.transpose().values
nitr_qf = np.logical_or(nitr_qf ==0, nitr_qf == 4)
nitrate[~nitr_qf] = np.nan
return nitrate
```
# O2 Budget for One Float, One Year
```
np.set_printoptions(suppress=True)
def find_mld_avg(var):
variable = np.flip(var, 0)
variable[variable==0] = np.nan
thickn[np.isnan(variable)] = np.nan
avg_var = np.empty((1,np.size(variable, 1))).flatten()
for i in range(len(imld)):
avg_var[i] = np.nansum(variable[0:int(imld[i]),i]*thickn[0:int(imld[i]),i])/np.nansum(thickn[0:int(imld[i]),i])
return avg_var
floats = [f_4180, f_4860, f_4767]
fl_names = ['5904180', '5904860', '5904767']
fig = plt.figure(figsize=(10,15))
for fl in range(len(floats)):
time = floats[fl].JULD.values
dates = pd.to_datetime(time).year
dayofyear = floats[fl].JULD.dt.dayofyear
mld, imld, thickn = get_mld_rho(floats[fl], False)
#temporary (just for one yr)
mld = mld[dates==2017]
imld = imld[dates==2017]
thickn = thickn[:,dates==2017]
#set up o2, o2sat, no3
o2 = set_oxygen_var(floats[fl])
o2 = o2[:,dates==2017]
oxygenSat = set_oxygen_sat_var(floats[fl])
oxygenSat = oxygenSat[:,dates==2017]
#calculate O2sol
o2sol = (o2/oxygenSat*100)
#calculate AOU
aou = o2sol - o2
#calculate mld averages for O2, NO3, AOU
no3 = set_nitrate_var(floats[fl])
no3 = no3[:,dates==2017]
variables = [o2, no3, aou]
o2_mld = no3_mld = aou_mld = np.empty((1,np.size(o2, 1))).flatten()
o2_mld = find_mld_avg(o2)
no3_mld = find_mld_avg(no3)
aou_mld = find_mld_avg(aou)
#calculate ∆AOU
aou_am = np.nanmean(aou_mld, axis=0)
d_aou = aou_mld-aou_am
#calculate ∆O2
o2_am = np.nanmean(o2_mld, axis=0)
d_o2 = o2_mld-o2_am
#calculate ∆NO3
no3_am = np.nanmean(no3_mld, axis=0)
d_no3 = no3_mld-no3_am
#calculate ∆O2sol
d_o2sol = d_o2 + d_aou
#calculate ∆O2mix + ∆O2bio
r = -170/16
d_o2mixbio = d_o2 + r*d_no3
#calculate ∆O2gasex
d_o2gasex = np.negative(d_aou) - r*d_no3
#create plot
o2_vars = [d_o2, d_o2sol, d_o2mixbio, d_o2gasex]
varNames = ['∆O2', '∆O2sol', '∆O2mix + ∆O2bio', '∆O2gasex']
ax = fig.add_subplot(3,1, fl +1)
for var in range(len(o2_vars)):
ax.plot(dayofyear.values[dates==2017], o2_vars[var], linewidth=2, label=varNames[var])
#ax.plot(dayofyear.values[dates==2017], mld, color = 'black')
date_form = DateFormatter("%B") # %m for month number
ax.xaxis.set_major_formatter(date_form)
ax.xaxis.set_major_locator(mdates.MonthLocator(interval=1))
leg = ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
leg_lines = leg.get_lines()
leg_texts = leg.get_texts()
ax.set_xlabel('Month')
plt.xticks(rotation = 45)
plt.setp(leg_lines, linewidth=4)
plt.title('Oxygen Budget - Float ' + fl_names[fl] + ' in 2017', fontweight='bold', fontsize='15')
#plt.ylim([-90, 90])
fig.tight_layout()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/intel-analytics/analytics-zoo/blob/master/docs/docs/colab-notebook/orca/quickstart/pytorch_lenet_mnist_data_creator_func.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##### Copyright 2018 Analytics Zoo Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
```
## **Environment Preparation**
**Install Java 8**
Run the cell on the **Google Colab** to install jdk 1.8.
**Note:** if you run this notebook on your computer, root permission is required when running the cell to install Java 8. (You may ignore this cell if Java 8 has already been set up in your computer).
```
# Install jdk8
!apt-get install openjdk-8-jdk-headless -qq > /dev/null
import os
# Set environment variable JAVA_HOME.
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
!update-alternatives --set java /usr/lib/jvm/java-8-openjdk-amd64/jre/bin/java
!java -version
```
**Install Analytics Zoo**
[Conda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) is needed to prepare the Python environment for running this example.
**Note**: The following code cell is specific for setting up conda environment on Colab; for general conda installation, please refer to the [install guide](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) for more details.
```
import sys
# Get current python version
python_version = "3.7.10"
# Install Miniconda
!wget https://repo.continuum.io/miniconda/Miniconda3-4.5.4-Linux-x86_64.sh
!chmod +x Miniconda3-4.5.4-Linux-x86_64.sh
!./Miniconda3-4.5.4-Linux-x86_64.sh -b -f -p /usr/local
# Update Conda
!conda install --channel defaults conda python=$python_version --yes
!conda update --channel defaults --all --yes
# Append to the sys.path
_ = (sys.path
.append(f"/usr/local/lib/python3.7/site-packages"))
os.environ['PYTHONHOME']="/usr/local"
```
You can install the latest pre-release version using `pip install --pre analytics-zoo`.
```
# Install latest pre-release version of Analytics Zoo
# Installing Analytics Zoo from pip will automatically install pyspark, bigdl, and their dependencies.
!pip install --pre analytics-zoo
# Install python dependencies
!pip install torch==1.7.1 torchvision==0.8.2
!pip install six cloudpickle
!pip install jep==3.9.0
```
## **Distributed PyTorch using Orca APIs**
In this guide we will describe how to scale out PyTorch programs using Orca in 4 simple steps.
```
# import necesary libraries and modules
from __future__ import print_function
import os
import argparse
from zoo.orca import init_orca_context, stop_orca_context
from zoo.orca import OrcaContext
```
### **Step 1: Init Orca Context**
```
# recommended to set it to True when running Analytics Zoo in Jupyter notebook.
OrcaContext.log_output = True # (this will display terminal's stdout and stderr in the Jupyter notebook).
cluster_mode = "local"
if cluster_mode == "local":
init_orca_context(cores=1, memory="2g") # run in local mode
elif cluster_mode == "k8s":
init_orca_context(cluster_mode="k8s", num_nodes=2, cores=4) # run on K8s cluster
elif cluster_mode == "yarn":
init_orca_context(
cluster_mode="yarn-client", cores=4, num_nodes=2, memory="2g",
driver_memory="10g", driver_cores=1,
conf={"spark.rpc.message.maxSize": "1024",
"spark.task.maxFailures": "1",
"spark.driver.extraJavaOptions": "-Dbigdl.failure.retryTimes=1"}) # run on Hadoop YARN cluster
```
This is the only place where you need to specify local or distributed mode. View [Orca Context](https://analytics-zoo.readthedocs.io/en/latest/doc/Orca/Overview/orca-context.html) for more details.
**Note**: You should export HADOOP_CONF_DIR=/path/to/hadoop/conf/dir when you run on Hadoop YARN cluster.
### **Step 2: Define the Model**
You may define your model, loss and optimizer in the same way as in any standard (single node) PyTorch program.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
class LeNet(nn.Module):
def __init__(self):
super(LeNet, self).__init__()
self.conv1 = nn.Conv2d(1, 20, 5, 1)
self.conv2 = nn.Conv2d(20, 50, 5, 1)
self.fc1 = nn.Linear(4*4*50, 500)
self.fc2 = nn.Linear(500, 10)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2, 2)
x = x.view(-1, 4*4*50)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=1)
model = LeNet()
model.train()
criterion = nn.NLLLoss()
lr = 0.001
adam = torch.optim.Adam(model.parameters(), lr)
```
### **Step 3: Define Train Dataset**
You may use a Data Creator Function for your input data (as shown below), especially when the data size is very large.
```
import torch
from torchvision import datasets, transforms
torch.manual_seed(0)
dir='/tmp/dataset'
def train_loader_creator(config, batch_size):
train_loader = torch.utils.data.DataLoader(
datasets.MNIST(dir, train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=batch_size, shuffle=True)
return train_loader
def test_loader_creator(config, batch_size):
test_loader = torch.utils.data.DataLoader(
datasets.MNIST(dir, train=False,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=batch_size, shuffle=False)
return test_loader
```
### **Step 4: Fit with Orca Estimator**
First, Create an Estimator.
```
from zoo.orca.learn.pytorch import Estimator
from zoo.orca.learn.metrics import Accuracy
est = Estimator.from_torch(model=model, optimizer=adam, loss=criterion, metrics=[Accuracy()])
```
Next, fit and evaluate using the Estimator.
```
from zoo.orca.learn.trigger import EveryEpoch
est.fit(data=train_loader_creator, epochs=1, validation_data=test_loader_creator, batch_size=320,
checkpoint_trigger=EveryEpoch())
```
Finally, evaluate using the Estimator.
```
result = est.evaluate(data=test_loader_creator, batch_size=320)
for r in result:
print(r, ":", result[r])
```
The accuracy of this model has reached 98%.
```
# stop orca context when program finishes
stop_orca_context()
```
| github_jupyter |
# Do More with Twitter Data
Twitter is what's happening and what people are talking about right now, with hundreds of millions of Tweets sent each day. We're a group of data scientists on the Twitter Data team who are helping people do more with this vast amount of data in less time. In this spirit, we are starting a series of tutorials that aim to help people work with Twitter data effectively. Each of the posts in this series centers around a real-life example project and provides MIT-licensed code that you can use to bootstrap your projects with our enterprise and premium API products. We hope this series is fruitful for you and we are excited to see what you'll build.
## Data Collection, Filtering, and Parsing
- By Fiona Pigott, @[notFromShrek][fiona], Data Scientist at Twitter, Jan 2018
In this inaugural post, we'll introduce data collection, filtering, parsing, and summarizing. We want to be able to give readers an easy way to spin up on the Twitter Search APIs, tips around quickly examining data, and guidelines around one of the most difficult and most overlooked Twitter data challenges: making sure you get the right Tweets for the job.
### *Caveat*
This post is not meant to be a tutorial in Python or the PyData ecosystem and assumes that readers have a reasonable amount of technical sophistication, such that they could find the right Tweets for their use case in any language or framework. This tutorial uses Python because our group makes heavy use of the PyData stack (python, pandas, numpy, etc.), but a technical user could follow along in a programming language of their choice.
[fiona]: https://twitter.com/notFromShrek "Twitter Profile"
## Using Twitter data to answer a question
Typically you will start with a very high-level question that requires refinement to understand what data might be useful to you in answering that question. For this post, let's start with a broad question:
"*_I want to understand airline customers while they fly._*"
We'll begin our refinement process from here.
## *Why* do we want to know this?
Understanding the business need behind a question is paramount. Use it to understand where it is important to spend extra time to be exact, and where an estimation will suffice. Also use "why" to understand when the analysis needs to be complete--the best analysis is not useful if it is finished a month after a decision is made and presented.
## *What* are we interested in?
Use this question to help define where to begin with the analysis and which data to pull. To understand airline customers' behavior while they are flying and getting ready to fly, we are interested in people actually in an airport or on a plane, but we are not interested in people simply talking about airlines (sharing news stories about an airline, for instance).
## *When* is this information relevant?
Deciding on a relevant timeframe is all about the business use case: if you're tracking a trend over a long period of time, you may need years of data; if you're looking at the audience of a single TV premier, a few days of data may be enough.
In the case of our air travel question, we're only interested in any given Twitter user in the few hours around their plane flight, so we don't necessarily need to track Tweets over a long period of time. We do, however, want to get enough of a sample to make generalizations about how people Tweet before and after flights, so we should make sure to collect enough data over a long enough period of time to examine multiple flights.
## *Who* are we interested in?
Using simple rule filters like country and language can go a long way towards helping us analyze Tweets from the right people.
In this case, we're interested in people from all demographics, as long as they are *also* passengers on an airline. We can likely identify those people through their language ("About to take off, headed home to Boulder!"), their granular geo-tagged location, or Twitter place ("Denver International Airport").
## *Where* are they?
We can use Twitter's geo data and enrichments to make sure that our users are in relevant locations, if that is important to the question. Another way to approximate a user's location might be the language they speak, or even the time that they are Tweeting (if you're collecting Tweets at 2AM PST, don't expect to see a lot of content from California).
Remember, selecting only users for whom we have a very granular locations (like a geo-tag near an airport) means that we only get a sample of users. For studies where we want to know generally what people are talking about that might not be a problem, but it isn't as effective for studies where we want an exact count. Keep those trade-offs in mind when designing a data query.
---------------
Once we have a good understanding of our question, the next steps are to figure out how to get to the answer using Twitter data. We'll have to get the right data, understand it, and filter out any irrelevant data.
__Steps to get to an answer__
1. Consume Tweet data
- We need to get Tweets to analyze them. We'll walk through using the Python client for the Twitter Search API right inside this notebook. All you need is an account.
2. Parse Twitter data
- You can't analyze what you can't load. Understand the structure of the data, and get the pieces that are important to your analysis.
3. Describe Twitter data
- Descriptive statistics go a long way. What are the most popular hashtags? When are people Tweeting about these topics? Where is noise coming from? Are there specific URLs, hashtags, or @-mentions being shared that *aren't* relevant to your analysis?
4. Iterate on your search terms to filter the data
- We can't simply ask for every Tweet. A thoughtful analysis needs to work within the bounds of the Twitter data APIs to filter and retrieve the right data for the job--and for your data budget. Now that you know how to quickly edit rules to retrieve small quantities of data, as well as parse and describe a set of Tweets, it's time to iterate on filters to retrieve the data that is relevant to your question (and not pay for data that isn't).
---------------
# 1. Consume Tweet data: the Twitter Search APIs
## Twitter Search APIs query language
In order to pull Tweets from the Search APIs, you will first have to understand how to write a search query in Twitter's premium / enterprise search API's query language. The most important basics are these:
- Operator: Anything that can select or "match" a Tweet. One type of operator is a simple token match. A token match operator is simply the word, e.g. `cat`, and matches when that word occurs within the text of the Tweet. Token matches always disregard case.
- Logical "and": words (tokens) joined by a `space` are treated as queries and-ed together. For instance, the query "`cat dog`" would search for Tweets with both `cat` and `dog` somewhere in the text of the Tweet.
- Logical "or": the operator "`OR`" (capitalization is important here) between two tokens means that your rule will match on either term (without needing both to be present). The rule "`cat OR dog`" will match on a Tweet with _either_ "`cat`" or "`dog`" in the Tweet text.
- Grouping: use parentheses `()` to group operators together. In the query-language order of operations, "and" (`_`) is applied before "or" (`OR`), so use parentheses to make your groups explicit. "`cat dog OR bunny`" is different from "`cat (dog OR bunny)`" (can you see why?).
- Negation: use the operator "`-`" to negate operators (such as terms that you definitely _don't_ want in your dataset). You might search for Tweets about cats and _not_ about dogs with this type of query "`cat -dog`".
Detailed information on operators can be found [here](https://developer.twitter.com/en/docs/tweets/search/guides/premium-operators). We'll introduce and use some more advanced operators later.
__Caveat__
When using any rule operators, you need to be careful to understand which parts of a field a rule is matching on. On twitter.com a Tweet seems like a simple thing, but a [Twitter data payload](https://developer.twitter.com/en/docs/tweets/data-dictionary/overview/intro-to-tweet-json) can have more than 100 fields, and different operators match on different fields. For this reason, it's important to make sure that you read the documentation around each operator and test your rules to be sure that they are behaving as you expect.
One example: "`cat`" (the token match operator) will only match text that the user typed out for that Tweet (it won't for instance, match the word "`cat`" in the users's name or bio).
## Consuming Tweets
In order to Search for Tweets, we're going to use Twitter's [enterprise-grade Twitter search tool](https://developer.twitter.com/en/docs/tweets/search/overview/full-archive-search). This tools allows a user to make a request for Tweets by specifying a rule (more on that in a minute) that matches some Tweets and retrieve results. This tutorial is also compatible with the 30-day search API, though you will have to change some of the dates for searching due to the 30-day historic limitation.
__Requesting some Tweets__
In order to search for Tweets, we have to understand specifically how Twitter's search rules work. We'll outline a few simple rules, and we'll talk more about details later when we iterate on our rules.
**Time window for search**
Look for Tweets within a certain time window by specifying the time window in minute granularity.
**Search rules are simple, boolean, token matches**
Tweets are tokenized on spaces and punctuation, and those tokens are matched to a rule. Let's look at a simple example:
> **Tweet**: `"Boarding the plane to fly to Tokyo in a few hours! So excited!"`
> **Tokens** (capitalization is ignored): `"boarding", "the", "plane", "to", "fly", "to", "tokyo", "in", "a", "few", "hours", "so, "excited"`
A rule that collected this Tweet might have been `"plane"` (because the exact token `"plane"` is included in the token list). You should note that these matches rely on complete tokens: this Tweet would **not** match if our rule was `"airplane"`.
**You can search for more than one token in a rule**
A search for Tweets about flying on planes might include any Tweet with the word `"airplane"` *or* the word `"plane"` *or* the word `"flying"` to get all of the relevant Tweets. In a single rule, use the operator `OR` (capitalization is important here) to search for *any* of a set of keywords.
> `"airplane OR plane OR flying"`
**You can combine criteria using *and* logic, and combine tokens into phrases using quotation marks**
Maybe you want the to find Tweet that include the word `"flying"` and the word `"plane"` but only when they appear together. Here, you would want to use Boolean `AND` logic to combine tokens into a single rule. In the syntax of Twitter's Search API, `AND` is simply represented by a space.
> **rule**: `flying plane`
> **match**: `"I'm flying in a plane!"`
> **no match**: `"I'm flying!"` or `"I'm in a plane!"`
You can also look for specific phrases (combinations of tokens) using quotation marks. Use `"` to combine tokens and look for them in a specific phrase.
> **rule**: `"air travel"`
> **match**: `"Air travel is great"`
> **no match**: `"Travel by air is great"` or `"Traveling in an airplane"`
**You can exclude certain tokens that are irrelevant to your analysis**
Keep in mind that tokens do not exactly map to meaning (especially on a colloquial platform like Twitter). If you are looking for Tweets about flying (on an airplane), and you submit the rule `"flying"`, `"I don't give a flying f**k what you do."` would match your rule (true story: I've had to exclude the phrase `"flying f**k"` from this analysis 😳).
Use a "`-`" to exclude, and use parentheses to group together logical clauses:
> `"(airplane OR plane OR flying) -fuck"`
This isn't a comprehensive guide, and more clarification can be found in our [documentation](http://support.gnip.com/apis/search_api2.0/rules.html).
__Getting the results__
Twitter's Search APIs return results in *pages*, with up to 500 Tweets per page for paid tiers. For users in a sandbox environment, you can receive a maximum of 100 Tweets per call. For a very low-volume search, you might only need one page to retrieve all of the results. For a high-volume search, you can choose to make multiple API call to receive multiple pages of results.
__Python client__
Consuming data from Twitter APIs directly into an environment where we can analyze them is important for fast iteration on queries. The data science team has created some Python libraries that make it easy to consume data from Twitter's Search APIs directly into this notebook.
This package can be found at https://github.com/twitterdev/search-tweets-python and implements a Python wrapper to:
- Create appropriately formatted rule payloads (date and rule query parameters)
- Make requests of the API
- Gracefully handle pagination of Search APIs results (be aware--the tool can make multiple API calls, so but sure to set the `max_tweets` parameter, which we'll point out)
- Load the resultant Tweet text data as Python objects
## Running This Notebook
If you want to run this notebook, it is hosted [here](https://github.com/twitterdev/do_more_with_twitter_data). Clone this repo and you'll see this notebook in the `examples/finding_the_right_data` directory.
Please see the accompanying `README.md` file for full instructions. We've provided both a pip-ready `finding_the_right_data_requirements.txt` file and a conda environment file, `finding_the_right_data_conda_env.yml` that allows an easy virtual environment for this example. This example assumes Python 3.6.
__Credentials__
Please go ahead and make a YAML file named `.twitter_keys.yaml` in your home directory.
For premium customers, the simplest credential file should look like this:
```yaml
search_tweets_api:
account_type: premium
endpoint: <FULL_URL_OF_ENDPOINT>
consumer_key: <CONSUMER_KEY>
consumer_secret: <CONSUMER_SECRET>
```
For enterprise customers, the simplest credential file should look like this:
```yaml
search_tweets_api:
account_type: enterprise
endpoint: <FULL_URL_OF_ENDPOINT>
username: <USERNAME>
password: <PW>
```
The rest of the example will assume `~/.twitter_keys.yaml` exists, though you can specify your connection information directing in the notebook or using an environment variable if you want. For more information, please see the `searchtweets` [section on credential handling](https://twitterdev.github.io/search-tweets-python/#credential-handling).
The `load_credentials` function parses this file and we'll save the `search_args` variable for use throughout the session.
```
from tweet_parser.tweet import Tweet
from searchtweets import (ResultStream,
collect_results,
gen_rule_payload,
load_credentials)
search_args = load_credentials(filename="~/.twitter_keys.yaml",
account_type="enterprise")
```
In the following cells, we'll define our rule and rule payloads, then use the `collect_results` method to get our tweets.
First - it is often convenient to assign a rule to a variable. Rules are strings and can be simple strings, e.g., `"(flying OR plane)"` or be literal blocks, which can be easier to read for long rules:
```
"""
(flying
OR plane
OR landed)
"""
```
The remaining functions will parse newlines correctly.
Below, we'll start with a simple rule.
```
_rule_a = "(flying OR plane OR landed OR airport OR takeoff)"
```
The function, `gen_rule_payload`, will generate the JSON used to make requests to the search endpoint. Its full API doc lives [here](https://twitterdev.github.io/twitter_search_api/twittersearch.html#twittersearch.api_utils.gen_rule_payload), but it has several main parameters:
1. `pt_rule` (or the first positional arg) – The string version of a powertrack rule
2. `results_per_call` – number of tweets or counts returned per API. Defaults to 500 to reduce API call usage, but you can change it as needed. It can range from 100-500.
3. `from_date` – Date the starting time of your search.
4. `to_date` – Date for the end time of your search.
5. `count_bucket` – If using the counts api endpoint, this will define the count bucket for which tweets are aggregated. This must be set if you want to use the Counts API.
The time searches can be specified via the following dates or datetimes, down to minutes, e.g.:
- `2017-12-31`
- `2017-12-31 23:50`
- `2017-12-31T23:50`
- `201712312350`
Specifying a date will return a date starting at 00:00.
The `from_date` and `to_date` parameters are optional. The search API defaults to returning up to the last 30 day's worth of tweets.
```
rule_a = gen_rule_payload(_rule_a,
from_date="2017-07-01",
to_date="2017-08-01",
results_per_call=500)
rule_a
```
The `collect_results` function is the fastest entry point to getting Tweets. It has several parameters:
- `rule`: a valid payload made from `gen_rule_payload`
- `max_results`: the maximum number of Tweets or counts you want to receive
- `result_stream_args`: the search arguments and authentication info for this request.
And it returns a list of results, Tweets or counts.
```
results_list = collect_results(rule_a,
max_results=500,
result_stream_args=search_args)
# hark, a Tweet!
results_list[0]
```
## 2. Parse Twitter data
Let's take the 1st Tweet from our results list and discuss its various elements.
Tweet data is returned from the API as a single JSON payload with a `results` array, containing many JSON Tweet payloads. The `searchtweets` package parses that data for you (including abstracting away concerns about the specific format of the Tweet payloads) using our [`tweet_parser`](https://github.com/tw-ddis/tweet_parser) package and handles any errors caused by non-Tweet messages (like logging messages), returning `Tweet` objects (I'll explain about that in a second).
To better understand Tweets, you should check out [our developer website](https://developer.twitter.com/en/docs/tweets/data-dictionary/overview/intro-to-tweet-json ) for information on Tweet payload elements. For now, I'm going to explain a few key elements of a Tweet and how to access them.
## The nitty gritty: Tweet payloads
We'll grab a single example Tweet: just the first one from our list.
This is a Tweet object, which has all of the properties of a Python `dict` as well as some special, Tweet-specific attributes.
```
example_tweet = results_list[0]
example_tweet["id"]
```
**A note on Tweet formatting and payload element naming**
It just so happens that the format of this Tweet is "original format," so we can look up the keys
(names of payload elements) on our support website.
It's possible (but unlikely) that your Search API stream is configured slightly differently, and you're looking at "activity streams" formatted Tweets (if so, this is completely fine). We'll add the equivalent "activity streams" keys in the comments.
You can look up specific of Tweet payload elements [here](https://developer.twitter.com/en/docs/tweets/data-dictionary/overview/tweet-object).
Note that the `tweet_parser` package will abstract away the format of the Tweet, and either activity streams or original format will work fine in this notebook.
**Now, the text of a Tweet:**
The text of the Tweet (the thing you type, the characters that display in the Tweet) is stored as a top-level key called "text."
```
print(results_list[0]["text"])
# uncomment the following if you appear to have an activity-stream formatted Tweet
# (the names of the payload elements are different, the data is the same):
# results_list[0]["body"]
```
**Other Tweet elements**
Be sure to read the documentation, we cannot enumerate every Tweet element here. Note that certain fundamental and useful Tweet elements are extracted for you, such as a #hashtag or an @mention.
You can access, for instance, a Tweet's #hashtags like this:
```
results_list[0]["entities"]["hashtags"]
# uncomment the following if you appear to have an activity-stream formatted Tweet
# results_list[0]["twitter_entities"]["hashtags"]
# in case that first Tweet didn't actually have any hashtags:
[x["entities"]["hashtags"] for x in results_list[0:10]]
# uncomment the following if you appear to have an activity-stream formatted Tweet
# [x["twitter_entities"]["hashtags"] for x in results_list[0:10]]
```
## tweet_parser
You will always need to understand Tweet payloads to work with Tweets, but a Python package from this team (`pip install tweet_parser`) attempts to remove some of the hassle of accessing elements of a Tweet. This package works seamlessly with both possible Twitter data formats, and supplies the `Tweet` object we referred to earlier.
Before doing a lot of work with this package (but not right this minute!), we would encourage you to read the documentation for the `tweet_parser` package at https://twitterdev.github.io/tweet_parser/. The package is open-source and available on GitHub at https://github.com/twitterdev/tweet_parser. Feel free to submit issues or make pull requests.
The `Tweet` object has properties that allow you to access useful elements of a Tweet without dealing with its format. For instance, we can access some of the text fields of a Tweet with:
```
print("Literally the content of the 'text' field: \n{}\n" .format(results_list[0].text))
print("Other fields, like the content of a quoted tweet or the options of a poll: {}, {}".format(
results_list[0].quoted_tweet.text if results_list[0].quoted_tweet else None,
results_list[0].poll_options))
```
As the Twitter platform changes, so do the fields in the data payload. For instance, "extended" and "truncated" tweets have been introduced to stop 280 character from causing breaking changes. The `Tweet` object has a convenience property `.all_text` that gets "all" the text that a Tweet displays.
```
results_list[0].all_text
```
We cannot dive into detail on every property of `Tweet` that we'll use in this notebook, but hopefully you get the idea. When we call `.something` on a `Tweet`, that `.something` is provided in the `tweet_parser` package and documented there.
For now, we will describe the elements of the payload that we'll use in this analysis.
- **time**: The time that a Tweet was created. This is always reported in UTC in the Tweet payload, and you can look for the user's timezone in the "user" portion of a Tweet payload if you want to translate this time to the user's time-of-day. A string of time information can be found in the `created_at` element of a Tweet, but since we're using the `tweet_parser` package, we can access a Python datetime object with `tweet.created_at_datetime` or an integer representing seconds since Jan 1, 1970 with `tweet.created_at_seconds`.
- **user**: The user who created the Tweet. The "user" portion of a Tweet payload contains many valuable elements, including the user's id (`tweet["user"]["id"]`, or `tweet.user_id` with the `tweet_parser` package), the user's screen name (`tweet.screen_name`, keep in mind that the user name may change), the user's timezone, potentially their derived location, their bio (a user-entered description at, `tweet.bio`) and more.
- **text**: The text of the Tweet. As Tweets get more complex (Retweets, Quote Tweets, poll Tweets, Tweets with hidden @-mentions and links), the text that you read in a Tweet can appear in many different areas of a Tweet payload. Here using the `tweet_parser`-provided attribute `tweet.all_text`, which aggregates all of the possible text fields of a Tweet (the quoted text, the poll text, the hidden @-mentions, etc) into one string.
- **hashtags**: Tweet payloads contain a field that list hashtags present in the Tweet (you do not have to parse them out yourself). The `tweet_parser` Tweet attribute `tweet.hashtags` provides a list of hashtags.
- **mentions**: You don't have to parse out @-mentions yourself either. Use `tweet.user_mentions` for a list of users (names and ids) mentioned in a Tweet.
- **URLs**: Many Tweet contain links to outside sources, articles or media. You can pull the literal link text from a Tweet, or use Twitter's URL enrichments (if you've purchased them) to unroll URLs and get insight into the linked content.
- **geo**: For the small sample of Tweets where explicit lat/lon geo data is available, use `tweet.geo_coordinates` to get locations.
Now we can take all of those aforementioned elements, parse them out of my Tweets, and put them in a [Pandas](https://pandas.pydata.org/pandas-docs/stable/) [Dataframe](https://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe).
```
import pandas as pd
# make a pandas dataframe
data_df = pd.DataFrame([{"date": x.created_at_datetime,
"text": x.all_text,
"user": x.screen_name,
"bio": x.bio,
"at_mentions": [u["screen_name"] for u in x.user_mentions],
"hashtags": x.hashtags,
"urls": x.most_unrolled_urls,
"geo": x.geo_coordinates,
"type": x.tweet_type,
"id": x.id} for x in results_list]).set_index("date")
```
## Before you read these Tweets
You're going to look at this Tweet data and you (should) feel a little sad. We did all that work talking about rules, you've heard so much about Twitter data, and these Tweets mostly don't seem relevant to our usecase at all. They're not from our target audience (people flying on planes) and they're not even necessarily about plane trips. What gives?
There's a reason we started out with only one API call: we didn't want to waste calls pulling Tweets using an unrefined ruleset. This notebook is going to be about making better decisions about rules, iterating, and refining them until you get the Tweets that you want. And those Tweets are out there - there are, after all, *lots* of Tweets.
```
# data, parsed out and ready to analyze
data_df.head()
```
## 3. Describe your data
Descriptive statistics can tell you a lot about your dataset with very little special effort and customization. Having a good set of descriptive statistics coded up that you always run on a new dataset can be helpful. Our team maintains [Gnip-Tweet-Evaluation](https://github.com/tw-ddis/Gnip-Tweet-Evaluation) a repository on GitHub that contains some tools to do quick evaluation of a Tweet corpus (that package is useful as a command line tool, and as a template for Tweet evaluation).
I'm going to walk through a few different statistics that might help us understand the data better:
- **What is generating the Tweets?**: One statistic that is common and useful in understanding a group of Tweets is understanding how many of them are Retweets or Quote Tweets. Another interesting data point can be looking at the application that created the Tweet itself.
- **Common words**: What are the most common words in the corpus? Are they what we expected? Are they relevant to the topic of interest?
- **Who is Tweeting**: Who are the users who Tweet the most? Are there spammy users that should be removed? Are they important users that we need to pay attention to?
- **Where are they Tweeting from**: What's the distribution of Tweet locations? Is that surprising?
- **Time series**: Are there spikes in the Tweet volume? When do they occur? What drives the spikes? For the sake of filtering out noise (or news stories that we don't care about) it may be important to identify spikes and filter out the Tweets that drive those spikes.
```
# plotting library
import matplotlib.pyplot as plt
# pretty plots
plt.style.use("bmh")
# better sizing for the notebook
plt.rcParams['figure.figsize'] = (10, 5)
%matplotlib inline
```
## How are people Tweeting?
Briefly, let's note that there are several different Twitter platform actions that can create a Tweet.
- **original Tweet**: the user created a Tweet by typing it into the Tweet create box or otherwise hitting the `statuses/update` endpoint
- **Retweet**: the user reposted the Tweet to their own timeline, without adding and comment
- **Quote Tweet**: user added commentary to a reposted Tweet, essentially a Tweet with another Tweet embedded in it
This is sometimes an important distinction: how much do Retweets or Quote Tweets represent the experience of the user reposting? Do we actually want Retweets in our particular dataset? Are they likely to tell us anything about users who are actually traveling (put it this way: if a user Retweets a story about being at an airport, how likely does it seem that they are at an airport)?
```
data_df[["type","id"]].groupby("type").count()
```
## Common words
Definitions:
- **corpus**: A collection of all of the documents that we are analyzing.
- **document**: A single piece of writing, in our case, a single Tweet
- **token**: Some characters from a document. Ideally, tokens would be common enough to occur in many documents, and have some semantic meaning--they provide us with a way to infer semantic correlation across documents.
- **stop word**: A stop word (something like "and","but","the") is a very common word with little semantic meaning, and we often exclude these words from word counts or NLP models.
__Tokenization__
In order to count common terms, first we have to define our terms. The easiest thing for me to count is a token--a single unit of characters in a Tweet. Often we try to define tokens so that a token is basically a word.
I'm going to tokenize the Tweets (split each Tweet into a list of tokens) using the [nltk TweetTokenizer](http://www.nltk.org/api/nltk.tokenize.html#module-nltk.tokenize.casual), which splits on spaces and throws away some punctuation. We will also throw away any tokens that contain no letters, and throw away any tokens that are less than 3 characters long. These choices--how to define a token, and which tokens to count, are somewhat arbitrary, but depend greatly on how language is used in the dataset. For instance, many tokenizers would remove the symbols "@" and "#" as punctuation, but on Twitter, those symbols mean something ("@Jack" is potentially very different from "jack"), and should perhaps be preserved.
```
from nltk.tokenize import TweetTokenizer
from nltk.stem.porter import PorterStemmer
def tweet_tokenizer(verbatim):
try:
tokenizer = TweetTokenizer()
all_tokens = tokenizer.tokenize(verbatim.lower())
# this line filters out all tokens that are entirely non-alphabetic characters
filtered_tokens = [t for t in all_tokens if t.islower()]
# filter out all tokens that are <=2 chars
filtered_tokens = [x for x in filtered_tokens if len(x)>2]
except IndexError:
filtered_tokens = []
return(filtered_tokens)
```
__Counting__
Of course, we could simply create a dictionary of tokens in our corpus and iterate through the entire corpus of Tweets and adding "1" to a count every time we encounter a certain token, but it's more convenient to start using the scikit-learn API now and put our token counts into a format that's easy to use later. If you're unfamiliar with scikit-learn, it is an excellent Python machine learning framework that implements many common ML algorithms in a common building-block-able API.
I am going to use [scikit-learn's CountVectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) to count tokens and turn the corpus into a document-term matrix. This makes it easy to count token frequencies. Note that the CountVectorizer allows us to count not only terms ("tokens") but n-grams (collections of tokens). For instance, if "plane" and "trip" are both tokens split on spaces, then explicit phrase "plane trip" might be a 2-gram in our corpus.
```
# get the most common words in Tweets
from sklearn.feature_extraction.text import CountVectorizer
def get_frequent_terms(text_series, stop_words = None, ngram_range = (1,2)):
'''
Input:
text_series: a list or series of documents
stop_words: a list of stop_words to ignore, or the string 'english',
which uses a built-in stop word list for the english language.
By default, there are no stop words used
ngram_range: a single int, or a 2 tuple representing the range of ngrams to count.
the default (1,2) counts 1- and 2- grams.
Returns:
a dataframe of counts, indexed by n-gram
'''
count_vectorizer = CountVectorizer(analyzer = "word",
tokenizer = tweet_tokenizer,
stop_words = stop_words, # try changing the stopword sets that we use.
# notice that many top terms are words like
# "and" and "the"
ngram_range = ngram_range # you can change this to count frequencies of
# ngrams as well as single tokens
# a range of (1,2) counts 1-grams (single tokens) and
# 2-grams (2-token phrases)
)
term_freq_matrix = count_vectorizer.fit_transform(text_series)
terms = count_vectorizer.get_feature_names()
term_frequencies = term_freq_matrix.sum(axis = 0).tolist()[0]
term_freq_df = (pd.DataFrame(list(zip(terms, term_frequencies)), columns = ["token","count"])
.set_index("token")
.sort_values("count",ascending = False))
return term_freq_df
term_freq_df = get_frequent_terms(data_df["text"],
stop_words = "english") # stop_words = "english" removes words like 'and'
term_freq_df.head(20)
term_freq_df.tail(10)
```
__Hand labeling__
Never underestimate the value of reading through your data. Actually look at the terms in the Tweets. Identify the Tweets that look like ones you are actually interested in.
Consider doing something like searching for specific, relevant, phrases that you hope will show up in your data:
```
data_df[data_df['text'].str.contains('flying home')]
```
Or simply reading a few random Tweets in your data.
```
pd.set_option('display.max_colwidth', -1)
data_df[['text']].sample(5)
```
A subsequent post will cover unsupervised clustering, which can help you group together Tweets to more easily make sense of them (you might group Tweets into 10 groups, and read a few Tweets from each group).
## Who is Tweeting? Who is speaking?
I'm going to use some Pandas tricks (`.groupby` and `.agg`) to find the most commonly Tweeting users in the dataset.
```
(data_df[["user","bio","geo","id"]]
.groupby("user")
.agg({"id":"count","bio":"first","geo":"first"})
.sort_values("id",ascending = False)
.rename(columns={'id':'tweet_count'})
).head(15)
```
## Time series plot
Doing time-series analysis with Twitter data will be covered in depth in a subsequent post, but it's worth introducing here.
Another great way to get a broader picture of our data is to understand when people are Tweeting about these keywords. Now, recall that earlier we only grabbed a tiny sample of data, so those Tweets only cover a few minutes of activity--not a good way to build a time series. If you have access to the enterprise level API, you will be able to make an API call to the [counts endpoint](https://developer.twitter.com/en/docs/tweets/search/overview/enterprise) to retrieve a time series *without* needing to use many calls to retrieve all Tweets over a time period. Let's try it out.
__Counts endpoint__
The counts endpoint has exactly the same API as the search endpoint, but instead of returning Tweets, it returns counts of Tweets. This is especially useful when you want to quickly assess a large dataset without retrieving a lot of data.
```
# use the same "_rule" string
print("Recall, our rule is: {}".format(_rule_a))
count_rule_a = gen_rule_payload(_rule_a,
from_date="2017-07-01",
to_date="2017-08-01",
results_per_call=500,
count_bucket="hour")
counts_list = collect_results(count_rule_a, max_results=24*31, result_stream_args=search_args)
```
The resulting counts payload is a list of counts and UTC timestamps.
```
counts_list[0:5]
# plot a timeseries of the Tweet counts
tweet_counts_original = (pd.DataFrame(counts_list)
.assign(time_bucket = lambda x: pd.to_datetime(x["timePeriod"]))
.drop("timePeriod",axis = 1)
.set_index("time_bucket")
.sort_index()
)
fig, axes = plt.subplots(nrows=1, ncols=2, figsize = (15,5))
(tweet_counts_original
.plot(ax=axes[0],
title = "Count of Tweets per hour",
legend = False));
(tweet_counts_original
.resample("D")
.sum()
.plot(ax=axes[1],
title="Count of Tweets per day",
legend=False));
```
__How can we use this information to understand our search data?__
Consider the type of Tweets that we are looking for: we're looking for Tweets from people who are talking about air travel, hopefully people talking about their own experiences using air travel. It's pretty likely that air travel has a relatively regular pattern, with probably some big seasonal fluctuations (Thanksgiving?) and some daily fluctuations (people fly, for the most part during the day); it's unlikely that 2 or 3 times as many people fly on any given Monday vs the Mondays around it.
We can use an intuition about what spikes mean in our data to either filter out high volume noise, or, in the case where spikes are what we are seeking out (say we wanted the audience for a movie release that happens on a specific day), zooming in on important time periods.
Let's choose a few large spikes in this data and investigate further, then exclude that topic from our final Twitter dataset.
__Note:__
If you don't have access to the counts API, you should still take a few small, time-boxed samples of data across the entire period of interest and doing the same exercise. It's harder to specifically target spikes, but it will help you get a broader sample of data.
```
# you can look at the plots to get a sense of what the highest-volume time periods are, or just sort your dataframe
(tweet_counts_original
.resample("D")
.sum()
.sort_values(by = "count", ascending = False)
.head())
# let's look at the highest volume day
spike_rule_717 = gen_rule_payload(_rule_a,
from_date="2017-07-17",
to_date="2017-07-18",
results_per_call=500)
spike_results_list_717 = collect_results(spike_rule_717, max_results=500, result_stream_args=search_args)
# what are these Tweets about?
get_frequent_terms([x.all_text for x in spike_results_list_717],
stop_words = "english").head(20)
# It's sometimes important to get the context of a full Tweet
[x.all_text for x in spike_results_list_717 if "drowned" in x.all_text.lower()][0:10]
# let's look at the highest volume day
spike_rule_726 = gen_rule_payload(_rule_a,
from_date="2017-07-26",
to_date="2017-07-27",
results_per_call=500)
spike_results_list_726 = collect_results(spike_rule_726, max_results=500, result_stream_args=search_args)
get_frequent_terms([x.all_text for x in spike_results_list_726], stop_words = "english").head(20)
[(x.all_text, x.tweet_type) for x in spike_results_list_726 if "gimpo" in x.all_text.lower()][0:10]
[(x.all_text, x.tweet_type) for x in spike_results_list_726 if "utah" in x.all_text.lower()][0:10]
```
## 3. Iterate
## Now that we understand what we're matching on
- **Exclusions**: The most important part of refining rules is often excluding Tweets that are irrelevant. Earlier, we learned that the Search API provides a negation operator (the "`-`" operator), which you should use for exclusions. Use the "`-`" operator to exclude terms that show up in irrelevant spikes, to exclude certain kinds of Tweets (say, exclude Retweets), or anything else that might mark a Tweet as irrelevant (spammy hashtags, news articles that aren't interesting, etc).
- **Advanced operators**: So far we have only covered token matching operators. There are more advanced operators, which can be used to match different aspects of a Tweet (is is a Retweet? does it contain an image?) or words in different parts of the payload (does the string "cnn" appear in a link? does the phrase "soccer mom" appear in the bio?). We'll show a few examples of using those operators in this section--for a complete list, read our [operator documentation](https://developer.twitter.com/en/docs/tweets/rules-and-filtering/overview/premium-operators).
```
# expand based on what you see (these are top terms that didn't seem relevant, look at the frequent terms we saw)
_rule_b = """
(plane OR landed OR airport OR takeoff OR #wheelsup)
-is:retweet
-trump
-harry
-drowned
-g20
-lawyer
-gimpo
-fuck
"""
rule_b = gen_rule_payload(_rule_b,
from_date="2017-07-01",
to_date="2017-08-01",
results_per_call=500)
results_list = collect_results(rule_b, max_results=500, result_stream_args=search_args)
# look at frequent terms again
get_frequent_terms([x.all_text for x in results_list], stop_words = "english", ngram_range = (2,3)).head(20)
# use the same "_rule" string
print("Recall, our rule is: {}".format(_rule_b))
count_rule_b = gen_rule_payload(_rule_b,
from_date="2017-07-01",
to_date="2017-08-01",
count_bucket="hour")
counts_list = collect_results(count_rule_b, max_results=24*31, result_stream_args=search_args)
# plot a timeseries of the Tweet counts
tweet_counts = (pd.DataFrame(counts_list)
.assign(time_bucket = lambda x: pd.to_datetime(x["timePeriod"]))
.drop("timePeriod",axis = 1)
.set_index("time_bucket")
.sort_index())
fig, axes = plt.subplots(nrows=1, ncols=2, figsize = (15,5))
(tweet_counts
.plot(ax=axes[0],
title = "Count of Tweets per hour", legend = False));
(tweet_counts
.resample("D")
.sum()
.plot(ax=axes[1],
title="Count of Tweets per day",
legend=False));
```
## Look at that regular timeseries!
Remember when we noted that the Tweet timeseries around flying is unlikely to have big, irregular spikes? It seems like we got rid of most of them, save one or two. Let's look into those last spikes and exclude those too.
```
# let's look at the highest volume day
spike_rule_711 = gen_rule_payload(_rule_b,
from_date="2017-07-11",
to_date="2017-07-12",
)
# force results to evaluate (this step actually makes the API calls)
spike_results_list_711 = collect_results(spike_rule_711, max_results=500, result_stream_args=search_args)
get_frequent_terms([x.all_text for x in spike_results_list_711],
stop_words="english",
ngram_range = (1,1)).head(15)
```
## Frequent terms vs frequent "n-grams"
We've talked about using frequent terms to identify what a corpus of Tweets is about, but I want to mention frequent "n-grams" as a slightly more sophisticated alternative.
**n-gram**: a sequence of *n* tokens from a document
Using frequent n-grams, you might be able to identify when words show together, in the same sequence surprisingly often - potentially indicating spam, promotional material, memes, lyrics, or reposted content. Pay attention to words that appear together, and you can exclude an n-gram by excluding an "exact phrase" from your Search query.
```
# let's look at 2- and 3- grams
get_frequent_terms([x.all_text for x in spike_results_list_711],
stop_words = "english",
ngram_range = (2,3)).head(20)
```
In this data, we can see that `"mississippi plane crash"`, `"killed mississippi plane "`, "vermont killed mississippi" all show up frequently, and they all show up *the same number of times*. Let me find one of those Tweets to look at:
```
[x.all_text for x in spike_results_list_711 if "vermont" in x.all_text.lower()][0:5]
```
Doesn't have much to do with the experience of flying in a plane. Let's exclude it. In this case, we might exclude these Tweets pretty precisely by excluding "Marine" (it's good to pick specific, unambiguous terms for exclusions, so that you don't do anything too broad), but for this example we'll exclude "plane crash"--it's precise enough, will exclude other similar news-type content, and we can demonstrate exact phrase matching.
Add this to my rule: `-"plane crash"`
```
# expand based on what you see
_rule_c = """
(plane OR landed OR airport OR takeoff OR #wheelsup)
-is:retweet
-trump
-harry
-drowned
-g20
-lawyer
-gimpo
-fuck
-"plane crash"
"""
spike_rule_704 = gen_rule_payload(_rule_c,
from_date="2017-07-04",
to_date="2017-07-05",
)
spike_results_list_704 = collect_results(spike_rule_704, max_results=500, result_stream_args=search_args)
get_frequent_terms([x.all_text for x in spike_results_list_704], stop_words = "english", ngram_range = (2,3)).head(20)
```
Now, it's not the top term, but when a specific hashtag shows up with some frequency, it's time take a look. Hashtags are often used to group together topics, and by filtering the hashtag we could catch irrelevant topics. Let's look at "#gameinsight."
```
# try searching on the hashtag:
[x.all_text for x in spike_results_list_704 if "gameinsight" in x.hashtags][0:5]
```
Well, that looks like spam. Let's exclude a hashtag this time: it's specific, seems to appear in this content, and easy to exclude. Use the hashtag and negation operator in Search like this:
Add `-#gameinsight`
Also, it's not clear that "gameinsight" is all of this spike. Try searching for "hampton" too (as it is one of the most common terms):
```
[x.all_text for x in spike_results_list_704 if "hampton" in x.all_text.lower()][0:5]
```
Now, could add: `-"citizens of east hampton"` and exclude this one news story, but we might have identified a larger pattern.
__URL matching__
News stories about planes, crashes, etc are showing up a lot, and we do not want to see any of them in the final dataset (again, this is absolutely a choice, and it will eliminate a lot of data, for better or for worse). We're going to choose to exclude all Tweets with certain big news URLs included in them. We're not going to get this perfect here, but we will see how to use the URL matching rule, and give you more ideas about how to filter Tweets.
The `url:<token>` operator performs a tokenized match on words in the unrolled URL that was posted in the Tweet. We'll eliminate Tweets with: "nytimes","bbc","washingtonpost", "cbsnews", "reuters", "apnews", and "news" in the URL.
Add:
`-url:nytimes -url:bbc -url:washingtonpost -url:cbsnews -url:reuters -url:apnews -url:news`
```
# expand based on what you see
_rule_d = """
(plane OR landed OR airport OR takeoff OR #wheelsup)
-is:retweet
-trump
-harry
-drowned
-g20
-lawyer
-gimpo
-fuck
-"plane crash"
-"citizens of east hampton"
-url:nytimes -url:bbc -url:washingtonpost -url:cbsnews -url:reuters -url:apnews -url:news
"""
# finally, let's look at that high volume hour on July 3rd
tweet_counts.sort_values(by = "count", ascending = False).head(2)
spike_rule_703 = gen_rule_payload(_rule_d, #<-remember, same rule
from_date="2017-07-03T18:00",
to_date="2017-07-03T19:01",
)
# force results to evaluate (this step actually makes the API calls)
spike_results_list_703 = collect_results(spike_rule_703, max_results=500, result_stream_args=search_args)
get_frequent_terms([x.all_text for x in spike_results_list_703], stop_words = "english", ngram_range = (2,3)).head(20)
# read some Tweets
[x.all_text for x in spike_results_list_703 if "Boston" in x.all_text][0:10]
```
We don't want to eliminate all mentions of "Boston Airport" (because many of those mentions are likely relevant to my study). Instead, we'll eliminate a few common phrases like "pedestrians struck", "pedestrians injured", "car strikes", "taxi strikes".
Add: `-"pedestrians struck" -"pedestrians injured" -"car strikes" -"taxi strikes" -"car hits"`
```
# expand based on what you see
_rule_e = """
(plane OR landed OR airport OR takeoff OR #wheelsup)
-is:retweet
-trump
-harry
-drowned
-g20
-lawyer
-gimpo
-fuck
-"plane crash"
-#gameinsight
-"citizens of east hampton"
-url:nytimes -url:bbc -url:washingtonpost -url:cbsnews -url:reuters -url:apnews -url:news
-"pedestrians struck" -"pedestrians injured" -"car strikes" -"taxi strikes" -"car hits"
"""
rule_e = gen_rule_payload(_rule_e,
from_date="2017-07-01",
to_date="2017-08-01",
)
results_list = collect_results(rule_e, max_results=500, result_stream_args=search_args)
print("Recall, our new rule is: {}".format(_rule_e))
count_rule_e = gen_rule_payload(_rule_e,
from_date="2017-07-01",
to_date="2017-08-01",
count_bucket="hour")
counts_list = collect_results(count_rule_e, max_results=24*31, result_stream_args=search_args)
# plot a timeseries of the Tweet counts
tweet_counts = (pd.DataFrame(counts_list)
.assign(time_bucket = lambda x: pd.to_datetime(x["timePeriod"]))
.drop("timePeriod",axis = 1)
.set_index("time_bucket")
.sort_index()
)
fig, axes = plt.subplots(nrows=1, ncols=2, figsize = (15,5))
_ = tweet_counts.plot(ax=axes[0], title = "Count of Tweets per hour", legend = False)
_ = tweet_counts.resample("D").sum().plot(ax=axes[1], title = "Count of Tweets per day", legend = False)
```
## Look how much irrelevant data we've eliminated
Let's compare the volume of our first (unrefined) rule to our most recent one.
Spoiler: We've eliminated a huge amount of unhelpful data!
Now you can begin to think about using this data in an analysis.
```
fig, axes = plt.subplots(nrows=1, ncols=1, figsize = (15,5))
tweet_counts_original.resample("D").sum().plot(ax=axes, title = "Count of Tweets per day", legend = False)
tweet_counts.resample("D").sum().plot(ax=axes, title = "Count of Tweets per day", legend = False)
plt.legend(["original", "refined"]);
get_frequent_terms([x.all_text for x in results_list], stop_words = "english", ngram_range = (1,2)).head(20)
```
## Pulling our dataset
It's always possible to continue refining a dataset, and we should definitely continue that work after pulling the Tweets. The key in early data-cleaning steps is to eliminate a large bulk of irrelevant Tweets that you don't want to store or pay for (we did that).
Now (**warning!** this will use quite a few API calls and you might want to think twice before actually running it. The cell type is markdown so you can't run it on accident) let's pull data with our final rule.
Before you decide to pull this data, we'll check how many API calls we've used in this notebook (the `ResultStream` object provides a convenience variable for this).
```
# count API calls
print("You have used {} API calls in this session".format(ResultStream.session_request_counter))
```
__Store the data you pull!__
You don't want to pull this much data without saving it for later. We're going to use the `ResultStream` object to make API calls, stream data in an iterator, hold relevant data information in memory in my Python session, and (importantly!) stream raw Tweets to a file for later use.
Even if you don't want to run these exact API calls, pay attention to how this is done for your own work.
__Finalize your rule__
```
# this is our final rule
print(_rule_e)
```
__No surprises__
You should always have a guess at how many Tweets you're going to pull _before_ you pull them. Use the Counts API (if possible), or extrapolate based on a smaller time period of data.
```
# I can sum up the counts endpoint results to guess how may Tweets I'll get
# (or, if you don't have access to the counts endpoints, try extrapolating from a single day)
tweet_counts.sum()
```
__Narrowing your dataset further__
You might refine your rule and find that there are still millions of Tweets about a topic on Twitter (this happens, there are lots of Tweets).
If your rule is still going to pull an unrealistic number of Tweets, maybe narrow the scope of your investigation. You can:
- **Narrow the time period**: do you really need a month of data? or maybe you could sample just a few days?
- **Select more specific Tweets**: maybe it would be helpful to your analysis to have only Tweets that are geo tagged (this reduces data volume significantly)? Or only pull Tweets from users with profile locations? Putting stringent requirements on data can help speed up your analysis and make your data volume smaller.
- **Sampling**: Search API doesn't support random sampling the way that the Historical APIs do. You'll have to come up with balanced ways of selecting for less data, or you can sample based on geography, time, Tweet language--anything to narrow your scope.
I'm going to show one example of this by *only* pulling Tweets from users with a profile location (you can read up more on what a "profile location" means in our documentation) in the state of Colorado (this is for illustrative purposes, depending on your use case, we'd probably recommend narrowing the time scope first).
Add: `profile_country:US profile_region:Colorado` to my Twitter Search rule.
```
final_rule = """
(plane OR landed OR airport OR takeoff OR #wheelsup)
-is:retweet
-trump
-harry
-drowned
-g20
-lawyer
-gimpo
-fuck
-"plane crash"
-#gameinsight
-"citizens of east hampton"
-url:nytimes -url:bbc -url:washingtonpost -url:cbsnews -url:reuters -url:apnews -url:news
-"pedestrians struck" -"pedstrians injured" -"car strikes" -"taxi strikes" -"car hits"
profile_country:US profile_region:"Colorado"
"""
count_rule = gen_rule_payload(final_rule,
from_date="2017-07-01",
to_date="2017-08-01",
count_bucket="hour")
counts_list = collect_results(count_rule, max_results=24*31, result_stream_args=search_args)
# plot a timeseries of the Tweet counts
tweet_counts = (pd.DataFrame(counts_list)
.assign(time_bucket = lambda x: pd.to_datetime(x["timePeriod"]))
.drop("timePeriod",axis = 1)
.set_index("time_bucket")
.sort_index()
)
fig, axes = plt.subplots(nrows=1, ncols=2, figsize = (15,5))
_ = tweet_counts.plot(ax=axes[0], title = "Count of Tweets per hour", legend = False)
_ = tweet_counts.resample("D").sum().plot(ax=axes[1], title = "Count of Tweets per day", legend = False)
tweet_counts.sum()
```
## Save Tweets to a file, and stream them into memory
Seems like a reasonable number of Tweets. Let's go get them.
This time, we're going to use the ResultStream object and stream the full Tweet payloads to a file while simultaneously creating a Pandas DataFrame in memory of the limited Tweet fields that we care about.
If you actually want to run the cell below, you'll have to set the cell type to "code."
```python
final_rule_payload = gen_rule_payload(final_rule,
from_date="2017-07-01",
to_date="2017-08-01")
stream = ResultStream(**search_args,
rule_payload=final_rule_payload,
max_results=None) # should collect all of the results
# write_ndjson is a utility function that writes the results to a file and passes them through the iterator
from searchtweets.utils import write_ndjson
limited_fields = []
for x in write_ndjson("air_travel_data.json", stream.stream()):
limited_fields.append({"date": x.created_at_datetime,
"text": x.all_text,
"user": x.screen_name,
"bio": x.bio,
"at_mentions": [u["screen_name"] for u in x.user_mentions],
"hashtags": x.hashtags,
"urls": x.most_unrolled_urls,
"geo": x.geo_coordinates,
"type": x.tweet_type,
"id": x.id})
# create a dataframe
final_dataset_df = pd.DataFrame(limited_fields)
final_dataset_df.head()
```
# Conclusion
All Tweet data collection should be focused on a question, and our focused on finding out what people were talking about while they flew on airplanes. You can use the same basic steps that we used here to answer your own business questions using Twitter data.
1. Consume Tweet data
- We walked through how to access the Search API using the `searchtweets` [package](https://github.com/twitterdev/search-tweets-python) and how to write rules to get the Tweets that you are looking for.
2. Parse Twitter data
- We talked about the JSON Tweet payloads, `Tweet` objects (provided by the `tweet_parser` [package](https://github.com/twitterdev/tweet_parser)), and Tweet [payload elements](https://developer.twitter.com/en/docs/tweets/data-dictionary/overview/intro-to-tweet-json).
3. Describe Twitter data
- We talked about how to describe data, using frequent terms, top users, timeseries spikes, and Twitter elements like hashtags and links.
4. Iterate on your search terms to filter the data
- We talked about how to narrow down a search by negating irrelevant terms, narrowing the time or the area of your search, and iterating until you retrieve a reasonable number of Tweets.
| github_jupyter |
```
import numpy as np
import tensorflow as tf
from utils import read_data
from rdkit import Chem, DataStructs
from rdkit.Chem import AllChem
from scipy import stats
from sklearn.metrics import accuracy_score, roc_auc_score
import pandas as pd
import seaborn as sns
tox_types = ['nr-ahr', 'nr-ar-lbd', 'nr-ar',
'nr-aromatase', 'nr-er-lbd', 'nr-er',
'nr-ppar-gamma', 'sr-are', 'sr-atad5',
'sr-hse', 'sr-mmp', 'sr-p53']
num_layer = 2
hidden_dim = 256
init_lr = 0.001
drop_rate = 0
reg_scale = 0.1
tox_type = 'nr-ahr'
smiles, label = read_data('tox21/' + tox_type)
smiles.shape, label.shape
mols = np.array([Chem.MolFromSmiles(smile) for smile in smiles])
fps_total = []
labels_total = []
for i in range(len(mols)):
try:
fp = AllChem.GetMorganFingerprintAsBitVect(mols[i], 2)
arr = np.zeros((1,))
DataStructs.ConvertToNumpyArray(fp,arr)
fps_total.append(arr)
labels_total.append(label[i])
except:
pass
fps_total = np.asarray(fps_total)
labels_total = np.asarray(labels_total)
fps_total.shape, labels_total.shape
num_total = fps_total.shape[0]
num_train = int(num_total*0.7)
num_validation = int(num_total*0.15)
num_test = int(num_total*0.15)
labels = labels_total
labels_total = np.zeros((num_total, 2))
for i in range(num_total):
labels_total[i][labels[i]] = 1
fps_train = fps_total[:num_train]
labels_train = labels_total[:num_train]
fps_validation = fps_total[num_train:num_train+num_validation]
labels_validation = labels_total[num_train:num_train+num_validation]
fps_test = fps_total[num_total-num_test:]
labels_test = labels_total[num_total-num_test:]
X = tf.placeholder(tf.float64, shape=[None, 2048])
Y = tf.placeholder(tf.float64, shape=[None, 2])
is_training = tf.placeholder(tf.bool, shape=())
h = X
regularizer = tf.contrib.layers.l2_regularizer(scale=reg_scale)
for i in range(num_layer):
h = tf.layers.dense(h,
units=hidden_dim,
use_bias=True,
activation=tf.nn.relu,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
kernel_regularizer=regularizer,
bias_regularizer=regularizer)
h = tf.layers.dropout(h,
rate=drop_rate,
training=is_training)
'''
h = tf.layers.dense(h,
units=hidden_dim,
use_bias=True,
activation=tf.nn.sigmoid,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
kernel_regularizer=regularizer,
bias_regularizer=regularizer)
'''
Y_pred = tf.layers.dense(h,
units=2,
use_bias=True,
activation=tf.nn.sigmoid,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
kernel_regularizer=regularizer,
bias_regularizer=regularizer)
Y_pred.shape, tf.layers.flatten(Y_pred).shape
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits=Y_pred, labels=Y)
loss = cross_entropy
lr = tf.Variable(0.0, trainable = False)
opt = tf.train.AdamOptimizer(lr).minimize(loss)
sess = tf.Session()
init = tf.global_variables_initializer()
sess.run(init)
batch_size = 100
epoch_size = 100
decay_rate = 0.95
batch_train = int(num_train/batch_size)
batch_validation = int(num_validation/batch_size)
batch_test = int(num_test/batch_size)
for t in range(epoch_size):
pred_train = []
sess.run(tf.assign(lr, init_lr*(decay_rate**t)))
for i in range(batch_train):
X_batch = fps_train[i*batch_size:(i+1)*batch_size]
Y_batch = labels_train[i*batch_size:(i+1)*batch_size]
_opt, _Y, _loss = sess.run([opt, Y_pred, loss], feed_dict = {X: X_batch, Y: Y_batch, is_training: True})
pred_train.append(_Y)
print("Epoch: ", t, "\t batch: ", i, "\t Training")
pred_train = np.concatenate(pred_train, axis=0)
pred_validation = []
for i in range(batch_validation):
X_batch = fps_validation[i*batch_size:(i+1)*batch_size]
Y_batch = labels_validation[i*batch_size:(i+1)*batch_size]
_Y, _loss = sess.run([Y_pred, loss], feed_dict={X: X_batch, Y: Y_batch, is_training: False})
pred_validation.append(_Y)
print("Epoch: ", t, "\tbatch: ", i, "\tValidating")
pred_validation = np.concatenate(pred_validation, axis=0)
pred_validation
pred_test = []
for i in range(batch_test):
X_batch = fps_test[i*batch_size:(i+1)*batch_size]
Y_batch = labels_test[i*batch_size:(i+1)*batch_size]
_Y, _loss = sess.run([Y_pred, loss], feed_dict = {X: X_batch, Y: Y_batch, is_training: False})
print("Batch: ", i, "\t Testing")
pred_test.append(_Y)
pred_test = np.concatenate(pred_test, axis=0)
labels_test_result = labels[num_total-num_test:num_total-num_test+len(pred_test)]
labels_pred_result = np.zeros(len(pred_test))
for i in range(len(pred_test)):
labels_pred_result[i] = 0 if pred_test[i][0]>pred_test[i][1] else 1
print(labels_test_result.shape, labels_pred_result.shape)
accuracy = accuracy_score(labels_test_result, labels_pred_result)
auc_roc = roc_auc_score(labels_test_result, labels_pred_result)
accuracy, auc_roc
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
import torch
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.utils import shuffle
from tqdm import tqdm
np.random.seed(42)
torch.manual_seed(42)
```
# Study population
```
data_dir = '../datafiles/chexpert/'
input_csv = 'chexpert.sample.test.csv'
# data_dir = '../datafiles/mimic/'
# input_csv = 'mimic.sample.test.csv'
output_suffix = 'resample'
sample_size_factor = 3
```
#### Preprocessing
```
df = pd.read_csv(data_dir + input_csv).drop(columns='Unnamed: 0')
def bin_age(age):
if age <= 20:
return 0
if age <= 30:
return 1
if age <= 40:
return 2
if age <= 50:
return 3
if age <= 60:
return 4
if age <= 70:
return 5
if age <= 80:
return 6
return 7
df["age_bin"] = df["age"].apply(bin_age)
```
#### Resample df to have equal proportion of Asian, Black, White
```
n_samples = len(df) * sample_size_factor
w_race = (1 / df.value_counts("race_label", normalize=True)).to_dict()
ids = list(torch.utils.data.WeightedRandomSampler(df["race_label"].apply(lambda x: w_race[x]).values, n_samples, replacement=True))
df_balanced_race = df.iloc[ids]
df_balanced_race.value_counts("race_label", normalize=True)
```
#### Within each (race)-subgroup ensure equal prevalence. Within each (race, disease)-subsubgroup ensure equal age distribution.
```
subgroups = [df_balanced_race.loc[df_balanced_race["race_label"] == i] for i in df_balanced_race.race_label.unique()]
overall_proportion_disease = df.value_counts("disease_label", normalize=True)
disease_labels = df.disease_label.unique()
overall_proportion_age_per_disease_label = [df.loc[df.disease_label == d].value_counts("age_bin", normalize=True) for d in disease_labels]
for i, subgroup in enumerate(subgroups):
# 1. Ensure constant disease prevalence in each subgroup
n_samples = len(subgroup)
observed_proportion = subgroup.value_counts("disease_label", normalize=True)
w = (overall_proportion_disease / observed_proportion).to_dict()
ids = list(torch.utils.data.WeightedRandomSampler(subgroup["disease_label"].apply(lambda x: w[x]).values, n_samples, replacement=True))
subgroups[i] = subgroup.iloc[ids]
# 2. Further ensure constant age repartition across groups
subsubgroups = [subgroups[i].loc[subgroups[i].disease_label == d] for d in disease_labels]
for j, subsubgroup in enumerate(subsubgroups):
n_samples = len(subsubgroup)
observed_proportion = subsubgroup.value_counts("age_bin", normalize=True)
w = (overall_proportion_age_per_disease_label[j] / observed_proportion).to_dict()
ids = list(torch.utils.data.WeightedRandomSampler(subsubgroup["age_bin"].apply(lambda x: w[x]).values, n_samples, replacement=True))
subsubgroups[j] = subsubgroup.iloc[ids]
subgroups[i] = pd.concat(subsubgroups)
balanced_df = pd.concat(subgroups)
```
#### Check balanced dataset
```
balanced_df.groupby(["race"]).size()
balanced_df.groupby(["race", "disease_label"]).size()
balanced_df.groupby(["race", "disease_label", "age_bin"]).size()
```
#### Save balanced dataset
```
balanced_df.to_csv(data_dir + input_csv.replace('sample',output_suffix))
```
## Plots
```
df_cxr = pd.read_csv(data_dir + input_csv.replace('sample',output_suffix))
df_subgroup = df_cxr[df_cxr['race_label']==0]
prev_0 = len(df_subgroup[df_subgroup['disease_label']==0]) / len(df_subgroup)
prev_1 = len(df_subgroup[df_subgroup['disease_label']==1]) / len(df_subgroup)
prev_2 = len(df_subgroup[df_subgroup['disease_label']==2]) / len(df_subgroup)
print('')
print('Prevalence - White')
print('No-finding:\t\t{:.2f}'.format(prev_0))
print('Pleural effusion:\t{:.2f}'.format(prev_1))
print('Other:\t\t\t{:.2f}'.format(prev_2))
df_subgroup = df_cxr[df_cxr['race_label']==1]
prev_0 = len(df_subgroup[df_subgroup['disease_label']==0]) / len(df_subgroup)
prev_1 = len(df_subgroup[df_subgroup['disease_label']==1]) / len(df_subgroup)
prev_2 = len(df_subgroup[df_subgroup['disease_label']==2]) / len(df_subgroup)
print('')
print('Prevalence - Asian')
print('No-finding:\t\t{:.2f}'.format(prev_0))
print('Pleural effusion:\t{:.2f}'.format(prev_1))
print('Other:\t\t\t{:.2f}'.format(prev_2))
df_subgroup = df_cxr[df_cxr['race_label']==2]
prev_0 = len(df_subgroup[df_subgroup['disease_label']==0]) / len(df_subgroup)
prev_1 = len(df_subgroup[df_subgroup['disease_label']==1]) / len(df_subgroup)
prev_2 = len(df_subgroup[df_subgroup['disease_label']==2]) / len(df_subgroup)
print('')
print('Prevalence - Black')
print('No-finding:\t\t{:.2f}'.format(prev_0))
print('Pleural effusion:\t{:.2f}'.format(prev_1))
print('Other:\t\t\t{:.2f}'.format(prev_2))
fontsize = 14
plt.rc('xtick', labelsize=fontsize)
plt.rc('ytick', labelsize=fontsize)
fig = sns.catplot(x='sex', y='age', hue='race', order=['Male', 'Female'], hue_order=['White', 'Asian', 'Black'], kind="box", data=df_cxr)
fig._legend.remove()
plt.xlabel('Sex', fontsize=fontsize)
plt.ylabel('Age (years)', fontsize=fontsize)
plt.ylim([0, 100])
plt.legend(fontsize=14, loc='lower center', ncol=3)
plt.show()
# fig.savefig("CheXpert-resample-age-sex-race.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='disease', y='age', hue='race', order=['Other', 'Pleural Effusion', 'No Finding'], hue_order=['White', 'Asian', 'Black'], kind="box", data=df_cxr)
fig._legend.remove()
plt.xlabel('Disease', fontsize=fontsize)
plt.ylabel('Age (years)', fontsize=fontsize)
plt.ylim([0, 100])
plt.legend(fontsize=14, loc='lower center', ncol=3)
plt.show()
# fig.savefig("CheXpert-resample-age-pathology-race.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='disease', hue='sex', order=['Other', 'Pleural Effusion', 'No Finding'], hue_order=['Male', 'Female'], data=df_cxr, kind='count')
fig._legend.remove()
plt.xlabel('Disease', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.legend(fontsize=14)
plt.show()
# fig.savefig("CheXpert-resample-sex-pathology.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='disease', hue='race', order=['Other', 'Pleural Effusion', 'No Finding'], hue_order=['White', 'Asian', 'Black'], data=df_cxr, kind='count')
fig._legend.remove()
plt.xlabel('Disease', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.legend(fontsize=14)
plt.show()
# fig.savefig("CheXpert-resample-race-pathology.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='disease', y='age', order=['Other', 'Pleural Effusion', 'No Finding'], kind="box", data=df_cxr)
plt.xlabel('Disease', fontsize=fontsize)
plt.ylabel('Age (years)', fontsize=fontsize)
plt.ylim([0, 100])
plt.show()
# fig.savefig("CheXpert-resample-age-pathology.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='sex', y='age', order=['Male', 'Female'], kind="box", data=df_cxr)
sns.stripplot(x='sex', y='age', color='k', order=['Male', 'Female'], alpha=0.01, data=df_cxr, ax=fig.ax)
plt.xlabel('Sex', fontsize=fontsize)
plt.ylabel('Age (years)', fontsize=fontsize)
plt.ylim([0, 100])
plt.show()
# fig.savefig("CheXpert-resample-age-sex.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='race', y='age', order=['White', 'Asian', 'Black'], kind="box", data=df_cxr)
sns.stripplot(x='race', y='age', color='k', order=['White', 'Asian', 'Black'], alpha=0.01, data=df_cxr, ax=fig.ax)
plt.xlabel('Race', fontsize=fontsize)
plt.ylabel('Age (years)', fontsize=fontsize)
plt.ylim([0, 100])
plt.show()
# fig.savefig("CheXpert-resample-age-race.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='race', order=['White', 'Asian', 'Black'], data=df_cxr, kind='count')
plt.xlabel('Race', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.show()
# fig.savefig("CheXpert-resample-race.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='sex', order=['Male', 'Female'], data=df_cxr, kind='count')
plt.xlabel('Sex', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.show()
# fig.savefig("CheXpert-resample-sex.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='disease', order=['Other', 'Pleural Effusion', 'No Finding'], data=df_cxr, kind='count')
plt.xlabel('Disease', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.show()
# fig.savefig("CheXpert-resample-pathology.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='race', hue='sex', order=['White', 'Asian', 'Black'], hue_order=['Male', 'Female'], data=df_cxr, kind='count')
fig._legend.remove()
plt.xlabel('Race', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.legend(fontsize=14)
plt.show()
# fig.savefig("CheXpert-resample-sex-race.png", bbox_inches='tight', dpi=300)
fig = sns.catplot(x='sex', hue='race', order=['Male', 'Female'], hue_order=['White', 'Asian', 'Black'], data=df_cxr, kind='count')
fig._legend.remove()
plt.xlabel('Sex', fontsize=fontsize)
plt.ylabel('Count', fontsize=fontsize)
plt.legend(fontsize=14)
plt.show()
# fig.savefig("CheXpert-resample-race-sex.png", bbox_inches='tight', dpi=300)
```
| github_jupyter |

# Working With File Streams
```
import azureml.dataprep as dprep
```
In addition to loading and parsing tabular data (see [here](./data-ingestion.ipynb) for more details), Data Prep also supports a variety of operations on raw file streams.
File streams are usually created by calling `Dataflow.get_files`.
```
dflow = dprep.Dataflow.get_files(path='../data/*.csv')
dflow.head(5)
```
The result of this operation is a Dataflow with a single column named "Path". This column contains values of type `StreamInfo`, each of which represents a different file matched by the search pattern specified when calling `get_files`. The string representation of a `StreamInfo` follows this pattern:
StreamInfo(_Location_://_ResourceIdentifier_\[_Arguments_\])
Location is the type of storage where the stream is located (e.g. Azure Blob, Local, or ADLS); ResouceIdentifier is the name of the file within that storage, such as a file path; and Arguments is a list of arguments required to load and read the file.
On their own, `StreamInfo` objects are not particularly useful; however, you can use them as input to other functions.
## Retrieving File Names
In the example above, we matched a set of CSV files by using a search pattern and got back a column with several `StreamInfo` objects, each representing a different file. Now, we will extract the file path and name for each of these values into a new string column.
```
dflow = dflow.add_column(expression=dprep.get_stream_name(dflow['Path']),
new_column_name='FilePath',
prior_column='Path')
dflow.head(5)
```
The `get_stream_name` function will return the full name of the file referenced by a `StreamInfo`. In the case of a local file, this will be an absolute path. From here, you can use the `derive_column_by_example` method to extract just the file name.
```
import os
first_file_path = dflow.head(1)['FilePath'][0]
first_file_name = os.path.basename(first_file_path)
dflow = dflow.derive_column_by_example(new_column_name='FileName',
source_columns=['FilePath'],
example_data=(first_file_path, first_file_name))
dflow = dflow.drop_columns(['FilePath'])
dflow.head(5)
```
## Writing Streams
Whenever you have a column containing `StreamInfo` objects, it's possible to write these out to any of the locations Data Prep supports. You can do this by calling `Dataflow.write_streams`:
```
dflow.write_streams(streams_column='Path', base_path=dprep.LocalFileOutput('./test_out/')).run_local()
```
The `base_path` parameter specifies the location the files will be written to. By default, the name of the file will be the resource identifier of the stream with any invalid characters replaced by `_`. In the case of streams referencing local files, this would be the full path of the original file. You can also specify the desired file names by referencing a column containing them:
```
dflow.write_streams(streams_column='Path',
base_path=dprep.LocalFileOutput('./test_out/'),
file_names_column='FileName').run_local()
```
Using this functionality, you can transfer files from any source to any destination supported by Data Prep. In addition, since the streams are just values in the Dataflow, you can use all of the functionality available.
Here, for example, we will write out only the files that start with the prefix "crime-". The resulting file names will have the prefix stripped and will be written to a folder named "crime".
```
prefix = 'crime-'
dflow = dflow.filter(dflow['FileName'].starts_with(prefix))
dflow = dflow.add_column(expression=dflow['FileName'].substring(len(prefix)),
new_column_name='CleanName',
prior_column='FileName')
dflow.write_streams(streams_column='Path',
base_path=dprep.LocalFileOutput('./test_out/crime/'),
file_names_column='CleanName').run_local()
```
## Converting Data Into Streams
Tabular data can be easily converted into a series of streams containing the data expressed in a binary or text format. These streams can then be written out using the capabilities outlined above. The number of resulting streams will depend on the number of partitions in the input data.
```
tabular_dflow = dprep.auto_read_file('../data/crime-full.csv')
streams_dflow = tabular_dflow.to_parquet_streams()
streams_dflow.head(1)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2_complexes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#AlphaFold2_complexes
---------
**UPDATE** (Aug. 13, 2021)
This notebook is being retired and no longer updated. The functionality for complex prediction (including going beyond dimers) has been integrated in our [new advanced notebook](https://github.com/sokrypton/ColabFold/blob/main/beta/AlphaFold2_advanced.ipynb).
---------
Credit to Minkyung Baek @minkbaek and Yoshitaka Moriwaki @Ag_smith for initially showing protein-complex prediction works in alphafold2.
- https://twitter.com/minkbaek/status/1417538291709071362
- https://twitter.com/Ag_smith/status/1417063635000598528
- [script](https://github.com/RosettaCommons/RoseTTAFold/blob/main/example/complex_modeling/make_joint_MSA_bacterial.py) from rosettafold for paired alignment generation
**Instructions**
- For *monomers* and *homo-oligomers*, see this [notebook](https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb).
- For prokaryotic protein complexes (found in operons), we recommend using the `pair_msa` option.
**Limitations**
- This notebook does NOT use templates or amber relax at the end for refinement.
- For a typical Google-Colab-GPU (16G) session, the max total length is **1400 residues**.
```
#@title Input protein sequences
import os
os.environ['TF_FORCE_UNIFIED_MEMORY'] = '1'
os.environ['XLA_PYTHON_CLIENT_MEM_FRACTION'] = '2.0'
from google.colab import files
import os.path
import re
import hashlib
def add_hash(x,y):
return x+"_"+hashlib.sha1(y.encode()).hexdigest()[:5]
query_sequence_a = 'AVLKIIQGALDTRELLKAYQEEACAKNFGAFCVFVGIVRKEDNIQGLSFDIYEALLKTWFEKWHHKAKDLGVVLKMAHSLGDVLIGQSSFLCVSMGKNRKNALELYENFIEDFKHNAPIWKYDLIHNKRIYAKERSHPLKGSGLLA' #@param {type:"string"}
query_sequence_a = "".join(query_sequence_a.split())
query_sequence_a = re.sub(r'[^A-Z]','', query_sequence_a.upper())
query_sequence_b = 'MMVEVRFFGPIKEENFFIKANDLKELRAILQEKEGLKEWLGVCAIALNDHLIDNLNTPLKDGDVISLLPPVCGG' #@param {type:"string"}
query_sequence_b = "".join(query_sequence_b.split())
query_sequence_b = re.sub(r'[^A-Z]','', query_sequence_b.upper())
# Using trick from @onoda_hiroki
# https://twitter.com/onoda_hiroki/status/1420068104239910915
# "U" indicates an "UNKNOWN" residue and it will not be modeled
# But we need linker of at least length 32
query_sequence_a = re.sub(r'U+',"U"*32,query_sequence_a)
query_sequence_b = re.sub(r'U+',"U"*32,query_sequence_b)
query_sequence = query_sequence_a + query_sequence_b
if len(query_sequence) > 1400:
print(f"WARNING: For a typical Google-Colab-GPU (16G) session, the max total length is 1400 residues. You are at {len(query_sequence)}!")
jobname = 'test' #@param {type:"string"}
jobname = "".join(jobname.split())
jobname = re.sub(r'\W+', '', jobname)
jobname = add_hash(jobname, query_sequence)
# number of models to use
#@markdown ---
#@markdown ### Advanced settings
num_models = 5 #@param [1,2,3,4,5] {type:"raw"}
msa_mode = "MMseqs2" #@param ["MMseqs2","single_sequence"]
use_msa = True if msa_mode == "MMseqs2" else False
pair_msa = False #@param {type:"boolean"}
disable_mmseqs2_filter = pair_msa
#@markdown ---
with open(f"{jobname}.log", "w") as text_file:
text_file.write("num_models=%s\n" % num_models)
text_file.write("use_msa=%s\n" % use_msa)
text_file.write("msa_mode=%s\n" % msa_mode)
text_file.write("pair_msa=%s\n" % pair_msa)
text_file.write("disable_mmseqs2_filter=%s\n" % disable_mmseqs2_filter)
#@title Install dependencies
%%bash -s $use_msa
USE_MSA=$1
if [ ! -f AF2_READY ]; then
# install dependencies
pip -q install biopython
pip -q install dm-haiku
pip -q install ml-collections
pip -q install py3Dmol
wget -qnc https://raw.githubusercontent.com/sokrypton/ColabFold/main/beta/colabfold.py
# download model
if [ ! -d "alphafold/" ]; then
git clone https://github.com/deepmind/alphafold.git --quiet
mv alphafold alphafold_
mv alphafold_/alphafold .
# remove "END" from PDBs, otherwise biopython complains
sed -i "s/pdb_lines.append('END')//" /content/alphafold/common/protein.py
sed -i "s/pdb_lines.append('ENDMDL')//" /content/alphafold/common/protein.py
fi
# download model params (~1 min)
if [ ! -d "params/" ]; then
wget -qnc https://storage.googleapis.com/alphafold/alphafold_params_2021-07-14.tar
mkdir params
tar -xf alphafold_params_2021-07-14.tar -C params/
rm alphafold_params_2021-07-14.tar
fi
touch AF2_READY
fi
#@title Import libraries
# setup the model
if "IMPORTED" not in dir():
import time
import requests
import tarfile
import sys
import numpy as np
import pickle
from string import ascii_uppercase
from alphafold.common import protein
from alphafold.data import pipeline
from alphafold.data import templates
from alphafold.model import data
from alphafold.model import config
from alphafold.model import model
from alphafold.data.tools import hhsearch
import colabfold as cf
# plotting libraries
import py3Dmol
import matplotlib.pyplot as plt
IMPORTED = True
def set_bfactor(pdb_filename, bfac, idx_res, chains):
I = open(pdb_filename,"r").readlines()
O = open(pdb_filename,"w")
for line in I:
if line[0:6] == "ATOM ":
seq_id = int(line[22:26].strip()) - 1
seq_id = np.where(idx_res == seq_id)[0][0]
O.write(f"{line[:21]}{chains[seq_id]}{line[22:60]}{bfac[seq_id]:6.2f}{line[66:]}")
O.close()
def predict_structure(prefix, feature_dict, Ls, random_seed=0, num_models=5):
"""Predicts structure using AlphaFold for the given sequence."""
# Minkyung's code
# add big enough number to residue index to indicate chain breaks
idx_res = feature_dict['residue_index']
L_prev = 0
# Ls: number of residues in each chain
for L_i in Ls[:-1]:
idx_res[L_prev+L_i:] += 200
L_prev += L_i
chains = list("".join([ascii_uppercase[n]*L for n,L in enumerate(Ls)]))
feature_dict['residue_index'] = idx_res
# Run the models.
plddts = []
paes = []
unrelaxed_pdb_lines = []
relaxed_pdb_lines = []
model_names = ["model_4","model_1","model_2","model_3","model_5"][:num_models]
for n,model_name in enumerate(model_names):
model_config = config.model_config(model_name+"_ptm")
model_config.data.eval.num_ensemble = 1
model_params = data.get_model_haiku_params(model_name+"_ptm", data_dir=".")
if model_name == "model_4":
model_runner = model.RunModel(model_config, model_params)
processed_feature_dict = model_runner.process_features(feature_dict,random_seed=0)
else:
# swap params
for k in model_runner.params.keys():
model_runner.params[k] = model_params[k]
print(f"running model_{n+1}")
prediction_result = model_runner.predict(processed_feature_dict)
# cleanup to save memory
if model_name == "model_5": del model_runner
del model_params
unrelaxed_protein = protein.from_prediction(processed_feature_dict,prediction_result)
unrelaxed_pdb_lines.append(protein.to_pdb(unrelaxed_protein))
plddts.append(prediction_result['plddt'])
paes.append(prediction_result['predicted_aligned_error'])
# Delete unused outputs to save memory.
del prediction_result
# rerank models based on predicted lddt
lddt_rank = np.mean(plddts,-1).argsort()[::-1]
plddts_ranked = {}
paes_ranked = {}
print("model\tplldt\tpae_ab")
L = Ls[0]
for n,r in enumerate(lddt_rank):
plddt = plddts[r].mean()
pae_ab = (paes[r][L:,:L].mean() + paes[r][:L,L:].mean()) / 2
print(f"model_{n+1}\t{plddt:.2f}\t{pae_ab:.2f}")
unrelaxed_pdb_path = f'{prefix}_unrelaxed_model_{n+1}.pdb'
with open(unrelaxed_pdb_path, 'w') as f:
f.write(unrelaxed_pdb_lines[r])
set_bfactor(unrelaxed_pdb_path, plddts[r], idx_res, chains)
plddts_ranked[f"model_{n+1}"] = plddts[r]
paes_ranked[f"model_{n+1}"] = paes[r]
return plddts_ranked, paes_ranked
# CODE FROM MINKYUNG/ROSETTAFOLD
def read_a3m(a3m_lines):
'''parse an a3m files as a dictionary {label->sequence}'''
seq = []
lab = []
is_first = True
for line in a3m_lines.splitlines():
if line[0] == '>':
label = line.rstrip().split()[0][1:]
is_incl = True
if is_first: # include first sequence (query)
is_first = False
lab.append(label)
continue
if "UniRef" in label:
code = label.split()[0].split('_')[-1]
if code.startswith("UPI"): # UniParc identifier -- exclude
is_incl = False
continue
elif label.startswith("tr|"):
code = label.split('|')[1]
else:
is_incl = False
continue
lab.append(code)
else:
if is_incl:
seq.append(line.rstrip())
else:
continue
return seq, lab
# https://www.uniprot.org/help/accession_numbers
def uni2idx(ids):
'''convert uniprot ids into integers according to the structure
of uniprot accession numbers'''
ids2 = [i.split("-")[0] for i in ids]
ids2 = [i+'AAA0' if len(i)==6 else i for i in ids2]
arr = np.array([list(s) for s in ids2], dtype='|S1').view(np.uint8)
for i in [1,5,9]:
arr[:,i] -= ord('0')
arr[arr>=ord('A')] -= ord('A')
arr[arr>=ord('0')] -= ord('0')-26
arr[:,0][arr[:,0]>ord('Q')-ord('A')] -= 3
arr = arr.astype(np.int64)
coef = np.array([23,10,26,36,36,10,26,36,36,1], dtype=np.int64)
coef = np.tile(coef[None,:],[len(ids),1])
c1 = [i for i,id_ in enumerate(ids) if id_[0] in 'OPQ' and len(id_)==6]
c2 = [i for i,id_ in enumerate(ids) if id_[0] not in 'OPQ' and len(id_)==6]
coef[c1] = np.array([3, 10,36,36,36,1,1,1,1,1])
coef[c2] = np.array([23,10,26,36,36,1,1,1,1,1])
for i in range(1,10):
coef[:,-i-1] *= coef[:,-i]
return np.sum(arr*coef,axis=-1)
def run_mmseqs2(query_sequence, prefix, use_env=True, filter=False):
def submit(query_sequence, mode):
res = requests.post('https://a3m.mmseqs.com/ticket/msa', data={'q':f">1\n{query_sequence}", 'mode': mode})
return res.json()
def status(ID):
res = requests.get(f'https://a3m.mmseqs.com/ticket/{ID}')
return res.json()
def download(ID, path):
res = requests.get(f'https://a3m.mmseqs.com/result/download/{ID}')
with open(path,"wb") as out: out.write(res.content)
if filter:
mode = "env" if use_env else "all"
else:
mode = "env-nofilter" if use_env else "nofilter"
path = f"{prefix}_{mode}"
if not os.path.isdir(path): os.mkdir(path)
# call mmseqs2 api
tar_gz_file = f'{path}/out.tar.gz'
if not os.path.isfile(tar_gz_file):
out = submit(query_sequence, mode)
while out["status"] in ["RUNNING","PENDING"]:
time.sleep(1)
out = status(out["id"])
download(out["id"], tar_gz_file)
# parse a3m files
a3m_lines = []
a3m = f"{prefix}_{mode}.a3m"
if not os.path.isfile(a3m):
with tarfile.open(tar_gz_file) as tar_gz: tar_gz.extractall(path)
a3m_files = [f"{path}/uniref.a3m"]
if use_env: a3m_files.append(f"{path}/bfd.mgnify30.metaeuk30.smag30.a3m")
a3m_out = open(a3m,"w")
for a3m_file in a3m_files:
for line in open(a3m_file,"r"):
line = line.replace("\x00","")
if len(line) > 0:
a3m_lines.append(line)
a3m_out.write(line)
else:
a3m_lines = open(a3m).readlines()
return "".join(a3m_lines), len(a3m_lines)
#@title Call MMseqs2 to get MSA for each gene
Ls = [len(query_sequence_a),len(query_sequence_b)]
msas = []
deletion_matrices = []
if use_msa:
os.makedirs('tmp', exist_ok=True)
prefix = hashlib.sha1(query_sequence.encode()).hexdigest()
prefix = os.path.join('tmp',prefix)
print(f"running mmseqs2 (use_env={True} filter={True})")
a3m_lines = cf.run_mmseqs2([query_sequence_a, query_sequence_b], prefix, use_env=True, filter=True)
if pair_msa:
a3m_lines.append([])
print(f"running mmseqs2 for pair_msa (use_env={False} filter={False})")
a3m_lines_pair = cf.run_mmseqs2([query_sequence_a, query_sequence_b], prefix, use_env=False, filter=False)
# CODE FROM MINKYUNG/ROSETTAFOLD
msa1, lab1 = read_a3m(a3m_lines_pair[0])
msa2, lab2 = read_a3m(a3m_lines_pair[1])
if len(lab1) > 1 and len(lab2) > 1:
# convert uniprot ids into integers
hash1 = uni2idx(lab1[1:])
hash2 = uni2idx(lab2[1:])
# find pairs of uniprot ids which are separated by at most 10
idx1, idx2 = np.where(np.abs(hash1[:,None]-hash2[None,:]) < 10)
if idx1.shape[0] > 0:
a3m_lines[2] = ['>query\n%s%s\n'%(msa1[0],msa2[0])]
for i,j in zip(idx1,idx2):
a3m_lines[2].append(">%s_%s\n%s%s\n"%(lab1[i+1],lab2[j+1],msa1[i+1],msa2[j+1]))
msa, deletion_matrix = pipeline.parsers.parse_a3m("".join(a3m_lines[2]))
msas.append(msa)
deletion_matrices.append(deletion_matrix)
print("pairs found:",len(msa))
msa, deletion_matrix = pipeline.parsers.parse_a3m(a3m_lines[0])
msas.append([seq+"-"*Ls[1] for seq in msa])
deletion_matrices.append([mtx+[0]*Ls[1] for mtx in deletion_matrix])
msa, deletion_matrix = pipeline.parsers.parse_a3m(a3m_lines[1])
msas.append(["-"*Ls[0]+seq for seq in msa])
deletion_matrices.append([[0]*Ls[0]+mtx for mtx in deletion_matrix])
else:
msas.append([query_sequence])
deletion_matrices.append([[0]*len(query_sequence)])
feature_dict = {
**pipeline.make_sequence_features(sequence=query_sequence,
description="none",
num_res=len(query_sequence)),
**pipeline.make_msa_features(msas=msas, deletion_matrices=deletion_matrices),
}
#@title Plot Number of Sequences per Position
dpi = 100#@param {type:"integer"}
# confidence per position
plt.figure(dpi=dpi)
plt.plot((feature_dict["msa"] != 21).sum(0))
plt.xlabel("positions")
plt.ylabel("number of sequences")
plt.savefig(jobname+"_msa_coverage.png")
plt.show()
#@title Predict structure
plddts, paes = predict_structure(jobname, feature_dict, Ls=Ls, num_models=num_models)
#@title Plot Predicted Alignment Error
dpi = 100#@param {type:"integer"}
# confidence per position
plt.figure(figsize=(3*num_models,2), dpi=dpi)
for n,(model_name,value) in enumerate(paes.items()):
plt.subplot(1,num_models,n+1)
plt.title(model_name)
plt.imshow(value,label=model_name,cmap="bwr",vmin=0,vmax=30)
plt.colorbar()
plt.savefig(jobname+"_PAE.png")
plt.show()
#@title Plot lDDT per residue
# confidence per position
dpi = 100#@param {type:"integer"}
plt.figure(dpi=dpi)
for model_name,value in plddts.items():
plt.plot(value,label=model_name)
plt.legend()
plt.ylim(0,100)
plt.ylabel("predicted lDDT")
plt.xlabel("positions")
plt.savefig(jobname+"_lDDT.png")
plt.show()
#@title Display 3D structure {run: "auto"}
model_num = 1 #@param ["1", "2", "3", "4", "5"] {type:"raw"}
color = "chain" #@param ["chain", "lDDT", "rainbow"]
show_sidechains = False #@param {type:"boolean"}
show_mainchains = False #@param {type:"boolean"}
def plot_plddt_legend():
thresh = ['plDDT:','Very low (<50)','Low (60)','OK (70)','Confident (80)','Very high (>90)']
plt.figure(figsize=(1,0.1),dpi=100)
########################################
for c in ["#FFFFFF","#FF0000","#FFFF00","#00FF00","#00FFFF","#0000FF"]:
plt.bar(0, 0, color=c)
plt.legend(thresh, frameon=False,
loc='center', ncol=6,
handletextpad=1,
columnspacing=1,
markerscale=0.5,)
plt.axis(False)
return plt
def plot_confidence(model_num=1):
model_name = f"model_{model_num}"
plt.figure(figsize=(10,3),dpi=100)
"""Plots the legend for plDDT."""
#########################################
plt.subplot(1,2,1); plt.title('Predicted lDDT')
plt.plot(plddts[model_name])
for x in [len(query_sequence_a)]:
plt.plot([x,x],[0,100],color="black")
plt.ylabel('plDDT')
plt.xlabel('position')
#########################################
plt.subplot(1,2,2);plt.title('Predicted Aligned Error')
plt.imshow(paes[model_name], cmap="bwr",vmin=0,vmax=30)
plt.colorbar()
plt.xlabel('Scored residue')
plt.ylabel('Aligned residue')
#########################################
return plt
def show_pdb(model_num=1, show_sidechains=False, show_mainchains=False, color="lDDT"):
model_name = f"model_{model_num}"
pdb_filename = f"{jobname}_unrelaxed_{model_name}.pdb"
view = py3Dmol.view(js='https://3dmol.org/build/3Dmol.js',)
view.addModel(open(pdb_filename,'r').read(),'pdb')
if color == "lDDT":
view.setStyle({'cartoon': {'colorscheme': {'prop':'b','gradient': 'roygb','min':50,'max':90}}})
elif color == "rainbow":
view.setStyle({'cartoon': {'color':'spectrum'}})
elif color == "chain":
for n,chain,color in zip(range(2),list("ABCDEFGH"),
["lime","cyan","magenta","yellow","salmon","white","blue","orange"]):
view.setStyle({'chain':chain},{'cartoon': {'color':color}})
if show_sidechains:
BB = ['C','O','N']
view.addStyle({'and':[{'resn':["GLY","PRO"],'invert':True},{'atom':BB,'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"GLY"},{'atom':'CA'}]},
{'sphere':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.addStyle({'and':[{'resn':"PRO"},{'atom':['C','O'],'invert':True}]},
{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
if show_mainchains:
BB = ['C','O','N','CA']
view.addStyle({'atom':BB},{'stick':{'colorscheme':f"WhiteCarbon",'radius':0.3}})
view.zoomTo()
return view
show_pdb(model_num,show_sidechains, show_mainchains, color).show()
if color == "lDDT": plot_plddt_legend().show()
plot_confidence(model_num).show()
#@title Package and download results
!zip -FSr $jobname".result.zip" $jobname".log" $jobname"_msa_coverage.png" $jobname"_"*"relaxed_model_"*".pdb" $jobname"_lDDT.png" $jobname"_PAE.png"
files.download(f"{jobname}.result.zip")
```
# Instructions
- If you having issues downloading results, try disable adblocker and run the last cell again. If that fails click on the little folder icon to the left, navigate to file:`jobname.result.zip`, right-click and select "download".
| github_jupyter |
```
# -*- coding: utf-8 -*-
"""
EVCで変換する.
詳細 : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580aabf534c4dbb8bc.pdf
Converting by EVC.
Check detail : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580abf534c4dbb8bc.pdf
"""
from __future__ import division, print_function
import os
from shutil import rmtree
import argparse
import glob
import pickle
import time
import numpy as np
from numpy.linalg import norm
from sklearn.decomposition import PCA
from sklearn.mixture import GMM # sklearn 0.20.0から使えない
from sklearn.preprocessing import StandardScaler
import scipy.signal
import scipy.sparse
%matplotlib inline
import matplotlib.pyplot as plt
import IPython
from IPython.display import Audio
import soundfile as sf
import wave
import pyworld as pw
import librosa.display
from dtw import dtw
import warnings
warnings.filterwarnings('ignore')
"""
Parameters
__Mixtured : GMM混合数
__versions : 実験セット
__convert_source : 変換元話者のパス
__convert_target : 変換先話者のパス
"""
# parameters
__Mixtured = 40
__versions = 'pre-stored0.1.3'
__convert_source = 'input/EJM10/V01/T01/TIMIT/000/*.wav'
__convert_target = 'adaptation/EJF01/V01/T01/ATR503/A/*.wav'
# settings
__same_path = './utterance/' + __versions + '/'
__output_path = __same_path + 'output/EJF01/' # EJF01, EJF07, EJM04, EJM05
Mixtured = __Mixtured
pre_stored_pickle = __same_path + __versions + '.pickle'
pre_stored_source_list = __same_path + 'pre-source/**/V01/T01/**/*.wav'
pre_stored_list = __same_path + "pre/**/V01/T01/**/*.wav"
#pre_stored_target_list = "" (not yet)
pre_stored_gmm_init_pickle = __same_path + __versions + '_init-gmm.pickle'
pre_stored_sv_npy = __same_path + __versions + '_sv.npy'
save_for_evgmm_covarXX = __output_path + __versions + '_covarXX.npy'
save_for_evgmm_covarYX = __output_path + __versions + '_covarYX.npy'
save_for_evgmm_fitted_source = __output_path + __versions + '_fitted_source.npy'
save_for_evgmm_fitted_target = __output_path + __versions + '_fitted_target.npy'
save_for_evgmm_weights = __output_path + __versions + '_weights.npy'
save_for_evgmm_source_means = __output_path + __versions + '_source_means.npy'
for_convert_source = __same_path + __convert_source
for_convert_target = __same_path + __convert_target
converted_voice_npy = __output_path + 'sp_converted_' + __versions
converted_voice_wav = __output_path + 'sp_converted_' + __versions
mfcc_save_fig_png = __output_path + 'mfcc3dim_' + __versions
f0_save_fig_png = __output_path + 'f0_converted' + __versions
converted_voice_with_f0_wav = __output_path + 'sp_f0_converted' + __versions
__measure_target = 'adaptation/EJF01/V01/T01/TIMIT/000/*.wav'
for_measure_target = __same_path + __measure_target
mcd_text = __output_path + __versions + '_MCD.txt'
EPSILON = 1e-8
class MFCC:
"""
MFCC() : メル周波数ケプストラム係数(MFCC)を求めたり、MFCCからスペクトルに変換したりするクラス.
動的特徴量(delta)が実装途中.
ref : http://aidiary.hatenablog.com/entry/20120225/1330179868
"""
def __init__(self, frequency, nfft=1026, dimension=24, channels=24):
"""
各種パラメータのセット
nfft : FFTのサンプル点数
frequency : サンプリング周波数
dimension : MFCC次元数
channles : メルフィルタバンクのチャンネル数(dimensionに依存)
fscale : 周波数スケール軸
filterbankl, fcenters : フィルタバンク行列, フィルタバンクの頂点(?)
"""
self.nfft = nfft
self.frequency = frequency
self.dimension = dimension
self.channels = channels
self.fscale = np.fft.fftfreq(self.nfft, d = 1.0 / self.frequency)[: int(self.nfft / 2)]
self.filterbank, self.fcenters = self.melFilterBank()
def hz2mel(self, f):
"""
周波数からメル周波数に変換
"""
return 1127.01048 * np.log(f / 700.0 + 1.0)
def mel2hz(self, m):
"""
メル周波数から周波数に変換
"""
return 700.0 * (np.exp(m / 1127.01048) - 1.0)
def melFilterBank(self):
"""
メルフィルタバンクを生成する
"""
fmax = self.frequency / 2
melmax = self.hz2mel(fmax)
nmax = int(self.nfft / 2)
df = self.frequency / self.nfft
dmel = melmax / (self.channels + 1)
melcenters = np.arange(1, self.channels + 1) * dmel
fcenters = self.mel2hz(melcenters)
indexcenter = np.round(fcenters / df)
indexstart = np.hstack(([0], indexcenter[0:self.channels - 1]))
indexstop = np.hstack((indexcenter[1:self.channels], [nmax]))
filterbank = np.zeros((self.channels, nmax))
for c in np.arange(0, self.channels):
increment = 1.0 / (indexcenter[c] - indexstart[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexstart[c], indexcenter[c])):
filterbank[c, i] = (i - indexstart[c]) * increment
decrement = 1.0 / (indexstop[c] - indexcenter[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexcenter[c], indexstop[c])):
filterbank[c, i] = 1.0 - ((i - indexcenter[c]) * decrement)
return filterbank, fcenters
def mfcc(self, spectrum):
"""
スペクトルからMFCCを求める.
"""
mspec = []
mspec = np.log10(np.dot(spectrum, self.filterbank.T))
mspec = np.array(mspec)
return scipy.fftpack.realtransforms.dct(mspec, type=2, norm="ortho", axis=-1)
def delta(self, mfcc):
"""
MFCCから動的特徴量を求める.
現在は,求める特徴量フレームtをt-1とt+1の平均としている.
"""
mfcc = np.concatenate([
[mfcc[0]],
mfcc,
[mfcc[-1]]
]) # 最初のフレームを最初に、最後のフレームを最後に付け足す
delta = None
for i in range(1, mfcc.shape[0] - 1):
slope = (mfcc[i+1] - mfcc[i-1]) / 2
if delta is None:
delta = slope
else:
delta = np.vstack([delta, slope])
return delta
def imfcc(self, mfcc, spectrogram):
"""
MFCCからスペクトルを求める.
"""
im_sp = np.array([])
for i in range(mfcc.shape[0]):
mfcc_s = np.hstack([mfcc[i], [0] * (self.channels - self.dimension)])
mspectrum = scipy.fftpack.idct(mfcc_s, norm='ortho')
# splrep はスプライン補間のための補間関数を求める
tck = scipy.interpolate.splrep(self.fcenters, np.power(10, mspectrum))
# splev は指定座標での補間値を求める
im_spectrogram = scipy.interpolate.splev(self.fscale, tck)
im_sp = np.concatenate((im_sp, im_spectrogram), axis=0)
return im_sp.reshape(spectrogram.shape)
def trim_zeros_frames(x, eps=1e-7):
"""
無音区間を取り除く.
"""
T, D = x.shape
s = np.sum(np.abs(x), axis=1)
s[s < 1e-7] = 0.
return x[s > eps]
def analyse_by_world_with_harverst(x, fs):
"""
WORLD音声分析合成器で基本周波数F0,スペクトル包絡,非周期成分を求める.
基本周波数F0についてはharvest法により,より精度良く求める.
"""
# 4 Harvest with F0 refinement (using Stonemask)
frame_period = 5
_f0_h, t_h = pw.harvest(x, fs, frame_period=frame_period)
f0_h = pw.stonemask(x, _f0_h, t_h, fs)
sp_h = pw.cheaptrick(x, f0_h, t_h, fs)
ap_h = pw.d4c(x, f0_h, t_h, fs)
return f0_h, sp_h, ap_h
def wavread(file):
"""
wavファイルから音声トラックとサンプリング周波数を抽出する.
"""
wf = wave.open(file, "r")
fs = wf.getframerate()
x = wf.readframes(wf.getnframes())
x = np.frombuffer(x, dtype= "int16") / 32768.0
wf.close()
return x, float(fs)
def preEmphasis(signal, p=0.97):
"""
MFCC抽出のための高域強調フィルタ.
波形を通すことで,高域成分が強調される.
"""
return scipy.signal.lfilter([1.0, -p], 1, signal)
def alignment(source, target, path):
"""
タイムアライメントを取る.
target音声をsource音声の長さに合うように調整する.
"""
# ここでは814に合わせよう(targetに合わせる)
# p_p = 0 if source.shape[0] > target.shape[0] else 1
#shapes = source.shape if source.shape[0] > target.shape[0] else target.shape
shapes = source.shape
align = np.array([])
for (i, p) in enumerate(path[0]):
if i != 0:
if j != p:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
else:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
j = p
return align.reshape(shapes)
covarXX = np.load(save_for_evgmm_covarXX)
covarYX = np.load(save_for_evgmm_covarYX)
fitted_source = np.load(save_for_evgmm_fitted_source)
fitted_target = np.load(save_for_evgmm_fitted_target)
weights = np.load(save_for_evgmm_weights)
source_means = np.load(save_for_evgmm_source_means)
"""
声質変換に用いる変換元音声と目標音声を読み込む.
"""
timer_start = time.time()
source_mfcc_for_convert = []
source_sp_for_convert = []
source_f0_for_convert = []
source_ap_for_convert = []
fs_source = None
for name in sorted(glob.iglob(for_convert_source, recursive=True)):
print("source = ", name)
x_source, fs_source = sf.read(name)
f0_source, sp_source, ap_source = analyse_by_world_with_harverst(x_source, fs_source)
mfcc_source = MFCC(fs_source)
#mfcc_s_tmp = mfcc_s.mfcc(sp)
#source_mfcc_for_convert = np.hstack([mfcc_s_tmp, mfcc_s.delta(mfcc_s_tmp)])
source_mfcc_for_convert.append(mfcc_source.mfcc(sp_source))
source_sp_for_convert.append(sp_source)
source_f0_for_convert.append(f0_source)
source_ap_for_convert.append(ap_source)
target_mfcc_for_fit = []
target_f0_for_fit = []
target_ap_for_fit = []
for name in sorted(glob.iglob(for_convert_target, recursive=True)):
print("target = ", name)
x_target, fs_target = sf.read(name)
f0_target, sp_target, ap_target = analyse_by_world_with_harverst(x_target, fs_target)
mfcc_target = MFCC(fs_target)
#mfcc_target_tmp = mfcc_target.mfcc(sp_target)
#target_mfcc_for_fit = np.hstack([mfcc_t_tmp, mfcc_t.delta(mfcc_t_tmp)])
target_mfcc_for_fit.append(mfcc_target.mfcc(sp_target))
target_f0_for_fit.append(f0_target)
target_ap_for_fit.append(ap_target)
# 全部numpy.arrrayにしておく
source_data_mfcc = np.array(source_mfcc_for_convert)
source_data_sp = np.array(source_sp_for_convert)
source_data_f0 = np.array(source_f0_for_convert)
source_data_ap = np.array(source_ap_for_convert)
target_mfcc = np.array(target_mfcc_for_fit)
target_f0 = np.array(target_f0_for_fit)
target_ap = np.array(target_ap_for_fit)
print("Load Input and Target Voice time = ", time.time() - timer_start , "[sec]")
def convert(source, covarXX, fitted_source, fitted_target, covarYX, weights, source_means):
"""
声質変換を行う.
"""
Mixtured = 40
D = source.shape[0]
E = np.zeros((Mixtured, D))
for m in range(Mixtured):
xx = np.linalg.solve(covarXX[m], source - fitted_source[m])
E[m] = fitted_target[m] + np.dot(covarYX[m], xx)
px = GMM(n_components = Mixtured, covariance_type = 'full')
px.weights_ = weights
px.means_ = source_means
px.covars_ = covarXX
posterior = px.predict_proba(np.atleast_2d(source))
return np.dot(posterior, E)
def calc_std_mean(input_f0):
"""
F0変換のために標準偏差と平均を求める.
"""
tempF0 = input_f0[ np.where(input_f0 > 0)]
fixed_logF0 = np.log(tempF0)
#logF0 = np.ma.log(input_f0) # 0要素にlogをするとinfになるのでmaskする
#fixed_logF0 = np.ma.fix_invalid(logF0).data # maskを取る
return np.std(fixed_logF0), np.mean(fixed_logF0) # 標準偏差と平均を返す
"""
距離を測るために,正しい目標音声を読み込む
"""
source_mfcc_for_measure_target = []
source_sp_for_measure_target = []
source_f0_for_measure_target = []
source_ap_for_measure_target = []
for name in sorted(glob.iglob(for_measure_target, recursive=True)):
print("measure_target = ", name)
x_measure_target, fs_measure_target = sf.read(name)
f0_measure_target, sp_measure_target, ap_measure_target = analyse_by_world_with_harverst(x_measure_target, fs_measure_target)
mfcc_measure_target = MFCC(fs_measure_target)
#mfcc_s_tmp = mfcc_s.mfcc(sp)
#source_mfcc_for_convert = np.hstack([mfcc_s_tmp, mfcc_s.delta(mfcc_s_tmp)])
source_mfcc_for_measure_target.append(mfcc_measure_target.mfcc(sp_measure_target))
source_sp_for_measure_target.append(sp_measure_target)
source_f0_for_measure_target.append(f0_measure_target)
source_ap_for_measure_target.append(ap_measure_target)
measure_target_data_mfcc = np.array(source_mfcc_for_measure_target)
measure_target_data_sp = np.array(source_sp_for_measure_target)
measure_target_data_f0 = np.array(source_f0_for_measure_target)
measure_target_data_ap = np.array(source_ap_for_measure_target)
def calc_mcd(source, convert, target):
"""
変換する前の音声と目標音声でDTWを行う.
その後,変換後の音声と目標音声とのMCDを計測する.
"""
dist, cost, acc, path = dtw(source, target, dist=lambda x, y: norm(x-y, ord=1))
aligned = alignment(source, target, path)
return 10.0 / np.log(10) * np.sqrt(2 * np.sum(np.square(aligned - convert))), aligned
"""
変換を行う.
"""
timer_start = time.time()
# 事前に目標話者の標準偏差と平均を求めておく
temp_f = None
for x in range(len(target_f0)):
temp = target_f0[x].flatten()
if temp_f is None:
temp_f = temp
else:
temp_f = np.hstack((temp_f, temp))
target_std, target_mean = calc_std_mean(temp_f)
# 変換
output_mfcc = []
filer = open(mcd_text, 'a')
for i in range(len(source_data_mfcc)):
print("voice no = ", i)
# convert
source_temp = source_data_mfcc[i]
output_mfcc = np.array([convert(source_temp[frame], covarXX, fitted_source, fitted_target, covarYX, weights, source_means)[0] for frame in range(source_temp.shape[0])])
# syntehsis
source_sp_temp = source_data_sp[i]
source_f0_temp = source_data_f0[i]
source_ap_temp = source_data_ap[i]
output_imfcc = mfcc_source.imfcc(output_mfcc, source_sp_temp)
y_source = pw.synthesize(source_f0_temp, output_imfcc, source_ap_temp, fs_source, 5)
np.save(converted_voice_npy + "s{0}.npy".format(i), output_imfcc)
sf.write(converted_voice_wav + "s{0}.wav".format(i), y_source, fs_source)
# calc MCD
measure_temp = measure_target_data_mfcc[i]
mcd, aligned_measure = calc_mcd(source_temp, output_mfcc, measure_temp)
filer.write("MCD No.{0} = {1} , shape = {2}\n".format(i, mcd, source_temp.shape))
# save figure spectram
range_s = output_imfcc.shape[0]
scale = [x for x in range(range_s)]
MFCC_sample_s = [source_temp[x][0] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][0] for x in range(range_s)]
MFCC_sample_t = [aligned_measure[x][0] for x in range(range_s)]
plt.subplot(311)
plt.plot(scale, MFCC_sample_s, label="source", linewidth = 1.0)
plt.plot(scale, MFCC_sample_c, label="convert", linewidth = 1.0)
plt.plot(scale, MFCC_sample_t, label="target", linewidth = 1.0, linestyle="dashed")
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=3, ncol=3, mode="expand", borderaxespad=0.)
#plt.xlabel("Flame")
#plt.ylabel("amplitude MFCC")
MFCC_sample_s = [source_temp[x][1] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][1] for x in range(range_s)]
MFCC_sample_t = [aligned_measure[x][1] for x in range(range_s)]
plt.subplot(312)
plt.plot(scale, MFCC_sample_s, label="source", linewidth = 1.0)
plt.plot(scale, MFCC_sample_c, label="convert", linewidth = 1.0)
plt.plot(scale, MFCC_sample_t, label="target", linewidth = 1.0, linestyle="dashed")
plt.ylabel("amplitude MFCC")
MFCC_sample_s = [source_temp[x][2] for x in range(range_s)]
MFCC_sample_c = [output_mfcc[x][2] for x in range(range_s)]
MFCC_sample_t = [aligned_measure[x][2] for x in range(range_s)]
plt.subplot(313)
plt.plot(scale, MFCC_sample_s, label="source", linewidth = 1.0)
plt.plot(scale, MFCC_sample_c, label="convert", linewidth = 1.0)
plt.plot(scale, MFCC_sample_t, label="target", linewidth = 1.0, linestyle="dashed")
plt.xlabel("Flame")
plt.savefig(mfcc_save_fig_png + "s{0}.png".format(i) , format='png', dpi=300)
plt.close()
# synthesis with conveted f0
source_std, source_mean = calc_std_mean(source_f0_temp)
std_ratio = target_std / source_std
log_conv_f0 = std_ratio * (source_f0_temp - source_mean) + target_mean
conv_f0 = np.maximum(log_conv_f0, 0)
np.save(converted_voice_npy + "f{0}.npy".format(i), conv_f0)
y_conv = pw.synthesize(conv_f0, output_imfcc, source_ap_temp, fs_source, 5)
sf.write(converted_voice_with_f0_wav + "sf{0}.wav".format(i) , y_conv, fs_source)
# save figure f0
F0_s = [source_f0_temp[x] for x in range(range_s)]
F0_c = [conv_f0[x] for x in range(range_s)]
plt.plot(scale, F0_s, label="source", linewidth = 1.0)
plt.plot(scale, F0_c, label="convert", linewidth = 1.0)
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=3, ncol=2, mode="expand", borderaxespad=0.)
plt.xlabel("Frame")
plt.ylabel("Amplitude")
plt.savefig(f0_save_fig_png + "f{0}.png".format(i), format='png', dpi=300)
plt.close()
filer.close()
print("Make Converted Spectram time = ", time.time() - timer_start , "[sec]")
```
| github_jupyter |
## Example of Textual Augmenter Usage<a class="anchor" id="home"></a>:
* [Character Augmenter](#chara_aug)
* [OCR](#ocr_aug)
* [Keyboard](#keyboard_aug)
* [Random](#random_aug)
* [Word Augmenter](#word_aug)
* [Spelling](#spelling_aug)
* [Word Embeddings](#word_embs_aug)
* [TF-IDF](#tfidf_aug)
* [Contextual Word Embeddings](#context_word_embs_aug)
* [Synonym](#synonym_aug)
* [Antonym](#antonym_aug)
* [Random Word](#random_word_aug)
* [Split](#split_aug)
* [Back Translatoin](#back_translation_aug)
* [Reserved Word](#reserved_aug)
* [Sentence Augmenter](#sent_aug)
* [Contextual Word Embeddings for Sentence](#context_word_embs_sentence_aug)
* [Abstractive Summarization](#abst_summ_aug)
```
import os
os.environ["MODEL_DIR"] = '../model'
```
# Config
```
import nlpaug.augmenter.char as nac
import nlpaug.augmenter.word as naw
import nlpaug.augmenter.sentence as nas
import nlpaug.flow as nafc
from nlpaug.util import Action
text = 'The quick brown fox jumps over the lazy dog .'
print(text)
```
# Character Augmenter<a class="anchor" id="chara_aug">
Augmenting data in character level. Possible scenarios include image to text and chatbot. During recognizing text from image, we need to optical character recognition (OCR) model to achieve it but OCR introduces some errors such as recognizing "o" and "0". `OCRAug` simulate these errors to perform the data augmentation. For chatbot, we still have typo even though most of application comes with word correction. Therefore, `KeyboardAug` is introduced to simulate this kind of errors.
### OCR Augmenter<a class="anchor" id="ocr_aug"></a>
##### Substitute character by pre-defined OCR error
```
aug = nac.OcrAug()
augmented_texts = aug.augment(text, n=3)
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
```
### Keyboard Augmenter<a class="anchor" id="keyboard_aug"></a>
##### Substitute character by keyboard distance
```
aug = nac.KeyboardAug()
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Random Augmenter<a class="anchor" id="random_aug"></a>
##### Insert character randomly
```
aug = nac.RandomCharAug(action="insert")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Substitute character randomly
```
aug = nac.RandomCharAug(action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Swap character randomly
```
aug = nac.RandomCharAug(action="swap")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Delete character randomly
```
aug = nac.RandomCharAug(action="delete")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
# Word Augmenter<a class="anchor" id="word_aug"></a>
Besides character augmentation, word level is important as well. We make use of word2vec (Mikolov et al., 2013), GloVe (Pennington et al., 2014), fasttext (Joulin et al., 2016), BERT(Devlin et al., 2018) and wordnet to insert and substitute similar word. `Word2vecAug`, `GloVeAug` and `FasttextAug` use word embeddings to find most similar group of words to replace original word. On the other hand, `BertAug` use language models to predict possible target word. `WordNetAug` use statistics way to find the similar group of words.
### Spelling Augmenter<a class="anchor" id="spelling_aug"></a>
##### Substitute word by spelling mistake words dictionary
```
aug = naw.SpellingAug()
augmented_texts = aug.augment(text, n=3)
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
aug = naw.SpellingAug()
augmented_texts = aug.augment(text, n=3)
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
```
### Word Embeddings Augmenter<a class="anchor" id="word_embs_aug"></a>
##### Insert word randomly by word embeddings similarity
```
# model_type: word2vec, glove or fasttext
aug = naw.WordEmbsAug(
model_type='word2vec', model_path=model_dir+'GoogleNews-vectors-negative300.bin',
action="insert")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Substitute word by word2vec similarity
```
# model_type: word2vec, glove or fasttext
aug = naw.WordEmbsAug(
model_type='word2vec', model_path=model_dir+'GoogleNews-vectors-negative300.bin',
action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### TF-IDF Augmenter<a class="anchor" id="tfidf_aug"></a>
##### Insert word by TF-IDF similarity
```
aug = naw.TfIdfAug(
model_path=os.environ.get("MODEL_DIR"),
action="insert")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Substitute word by TF-IDF similarity
```
aug = naw.TfIdfAug(
model_path=os.environ.get("MODEL_DIR"),
action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Contextual Word Embeddings Augmenter<a class="anchor" id="context_word_embs_aug"></a>
##### Insert word by contextual word embeddings (BERT, DistilBERT, RoBERTA or XLNet)
```
aug = naw.ContextualWordEmbsAug(
model_path='bert-base-uncased', action="insert")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Substitute word by contextual word embeddings (BERT, DistilBERT, RoBERTA or XLNet)
```
aug = naw.ContextualWordEmbsAug(
model_path='bert-base-uncased', action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
aug = naw.ContextualWordEmbsAug(
model_path='distilbert-base-uncased', action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
aug = naw.ContextualWordEmbsAug(
model_path='roberta-base', action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Synonym Augmenter<a class="anchor" id="synonym_aug"></a>
##### Substitute word by WordNet's synonym
```
aug = naw.SynonymAug(aug_src='wordnet')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Substitute word by PPDB's synonym
```
aug = naw.SynonymAug(aug_src='ppdb', model_path=os.environ.get("MODEL_DIR") + 'ppdb-2.0-s-all')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Antonym Augmenter<a class="anchor" id="antonym_aug"></a>
##### Substitute word by antonym
```
aug = naw.AntonymAug()
_text = 'Good boy'
augmented_text = aug.augment(_text)
print("Original:")
print(_text)
print("Augmented Text:")
print(augmented_text)
```
### Random Word Augmenter<a class="anchor" id="random_word_aug"></a>
##### Swap word randomly
```
aug = naw.RandomWordAug(action="swap")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Delete word randomly
```
aug = naw.RandomWordAug()
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
##### Delete a set of contunous word will be removed randomly
```
aug = naw.RandomWordAug(action='crop')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Split Augmenter<a class="anchor" id="split_aug"></a>
##### Split word to two tokens randomly
```
aug = naw.SplitAug()
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Back Translation Augmenter<a class="anchor" id="back_translation_aug"></a>
```
import nlpaug.augmenter.word as naw
text = 'The quick brown fox jumped over the lazy dog'
back_translation_aug = naw.BackTranslationAug(
from_model_name='transformer.wmt19.en-de',
to_model_name='transformer.wmt19.de-en'
)
back_translation_aug.augment(text)
# Load models from local path
import nlpaug.augmenter.word as naw
from_model_dir = os.path.join(os.environ["MODEL_DIR"], 'word', 'fairseq', 'wmt19.en-de')
to_model_dir = os.path.join(os.environ["MODEL_DIR"], 'word', 'fairseq', 'wmt19.de-en')
text = 'The quick brown fox jumped over the lazy dog'
back_translation_aug = naw.BackTranslationAug(
from_model_name=from_model_dir, from_model_checkpt='model1.pt',
to_model_name=to_model_dir, to_model_checkpt='model1.pt',
is_load_from_github=False)
back_translation_aug.augment(text)
```
### Reserved Word Augmenter<a class="anchor" id="reserved_aug"></a>
```
import nlpaug.augmenter.word as naw
text = 'Fwd: Mail for solution'
reserved_tokens = [
['FW', 'Fwd', 'F/W', 'Forward'],
]
reserved_aug = naw.ReservedAug(reserved_tokens=reserved_tokens)
augmented_text = reserved_aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
# Sentence Augmentation
### Contextual Word Embeddings for Sentence Augmenter<a class="anchor" id="context_word_embs_sentence_aug"></a>
##### Insert sentence by contextual word embeddings (GPT2 or XLNet)
```
# model_path: xlnet-base-cased or gpt2
aug = nas.ContextualWordEmbsForSentenceAug(model_path='xlnet-base-cased')
augmented_texts = aug.augment(text, n=3)
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
aug = nas.ContextualWordEmbsForSentenceAug(model_path='gpt2')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
aug = nas.ContextualWordEmbsForSentenceAug(model_path='gpt2')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
aug = nas.ContextualWordEmbsForSentenceAug(model_path='distilgpt2')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
```
### Abstractive Summarization Augmenter<a class="anchor" id="abst_summ_aug"></a>
```
article = """
The history of natural language processing (NLP) generally started in the 1950s, although work can be
found from earlier periods. In 1950, Alan Turing published an article titled "Computing Machinery and
Intelligence" which proposed what is now called the Turing test as a criterion of intelligence.
The Georgetown experiment in 1954 involved fully automatic translation of more than sixty Russian
sentences into English. The authors claimed that within three or five years, machine translation would
be a solved problem. However, real progress was much slower, and after the ALPAC report in 1966,
which found that ten-year-long research had failed to fulfill the expectations, funding for machine
translation was dramatically reduced. Little further research in machine translation was conducted
until the late 1980s when the first statistical machine translation systems were developed.
"""
aug = nas.AbstSummAug(model_path='t5-base', num_beam=3)
augmented_text = aug.augment(article)
print("Original:")
print(article)
print("Augmented Text:")
print(augmented_text)
```
| github_jupyter |
# Wedges in `bruges`
Let's make wedge models!
We're going to make **all sorts of models** using one magical function. Here's what it can do:
- Layer-cake models with no dip.
- Wedge models with arbitrary thickness on the left and right.
- Clinoform models.
- Models filled with arbitrary stratigraphy, from constant values to well logs.
All these models can have varying amounts of rock above and below them, and can be extended left and right beyond the wedgy part. You can also dip the wedge in either direction.
**This is a new feature introduced in v0.4.2; if you find bugs, please let us know by [raising an issue](https://github.com/agile-geoscience/bruges/issues).**
```
import numpy as np
import matplotlib.pyplot as plt
import bruges as bg
```
---
## The default model and a basic synthetic workflow
We can produce a simple wedge model just by calling the function.
```
w, top, base, ref = bg.models.wedge()
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
plt.show()
```
You can then use this integer model to index into an array of rock properties:
```
rocks = np.array([2.32 * 2.65, # Rock index 0
2.35 * 2.60, # Rock index 1
2.35 * 2.62, # Rock index 2
])
# Fancy indexing into the rocks with the model.
impedance = rocks[w]
# Make reflectivity.
rc = (impedance[1:] - impedance[:-1]) / (impedance[1:] + impedance[:-1])
# Get a wavelet.
ricker, _ = bg.filters.ricker(0.064, 0.001, 40, sym=True, return_t=True)
# Repeated 1D convolution for a synthetic.
syn = np.apply_along_axis(np.convolve, arr=rc, axis=0, v=ricker, mode='same')
fig, axs = plt.subplots(figsize=(17, 4), ncols=5, gridspec_kw={'width_ratios': (4, 4, 4, 1, 4)})
axs[0].imshow(w)
axs[0].set_title('Wedge model')
axs[1].imshow(impedance)
axs[1].set_title('Impedance')
axs[2].imshow(rc)
axs[2].set_title('Reflectivity')
axs[3].plot(ricker, np.arange(ricker.size))
axs[3].axis('off')
axs[3].set_title('Wavelet')
axs[4].imshow(syn)
axs[4].set_title('Synthetic')
axs[4].plot(top, 'w', alpha=0.5)
axs[4].plot(base, 'w', alpha=0.5)
plt.show()
```
Note that we could also have made the impedance model directly — it just depends how you want to make your models.
So we can use the `strat` argument, and pass in the rock properties there.
```
w, top, base, ref = bg.models.wedge(strat=rocks)
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
plt.show()
```
Now the wedge contains rock properties, not integer labels.
```
w[:, 50]
```
---
## A layered wedge model
We can modify the stratigraphy of any layer. E.g., let's pass `(1, 2, 1, 2, 1)` in as the wedge strat, instead of just `1`. We'll also change the bottom layer to a `3`, so now we have 4 rocks.
```
w, top, base, ref = bg.models.wedge(depth=(100, 600, 100),
width=(200, 1600, 200),
strat=(0, (1, 2, 1, 2, 1), 3),
mode='linear'
)
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
plt.show()
```
Notice that the `wedge` function returns 4 things:
- The wedge model as a 2D NumPy array of ints (or whatever numbers you gave as input).
- The top horizon, in pixel index units, for convenience.
- The base horizon, in pixel index units, for convenience.
- The horizontal (left-right) position, as an integer, of the 'reference' model. You may or may not be interested in this, depending on your application.
If you only want the wedge, you can call the function like so:
wedge, *_ = bg.models.wedge()
---
## Arbitrary thicknesses
You can provide the minimum and maximum thickness of the wedge.
**Note:** If the maximum thickness of the wedge if more than 1, then the total depth (i.e. number of rows) of the model will be more than the sum of the `depth` argument, so that the entire model can be accommodated. If you don't want the 'extra' depth, you can slice them off the model as with any NumPy array.
Here's a layer cake:
```
w, top, base, ref = bg.models.wedge(depth=(10., 80, 10),
width=(10, 80, 10),
strat=(0, (1, 2, 2, 1, 2, 1, 0, 1, 1,), 3), # Floats in the wedge
thickness=(1, 1),
mode='linear',
)
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
plt.show()
```
Here's another example. This time we'll also pass in floats — velocities perhaps.
```
w, top, base, ref = bg.models.wedge(depth=(10., 80, 10),
width=(10, 80, 10),
strat=(1.48, (2.10, 2.25, 2.35), 2.40), # Floats in the wedge.
thickness=(1, 0.5),
mode='linear',
)
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
cb = plt.colorbar()
cb.ax.invert_yaxis()
plt.show()
```
---
## Top and bottom conformance
The layers in the wedge can also be top or bottom conforming, rather than proportionally adjusted.
```
confs = ['both', 'top', 'bottom']
fig, axs = plt.subplots(ncols=len(confs), figsize=(12, 4))
for ax, conf in zip(axs, confs):
w, top, base, ref = bg.models.wedge(strat=((0, 1, 0), (2, 3, 2, 3, 2), (4, 5, 4)),
conformance=conf)
ax.imshow(w, interpolation='none')
ax.axvline(ref, color='k', ls='--')
ax.plot(top, 'r-', lw=4)
ax.plot(base, 'r-', lw=4)
ax.set_title(f"{conf} conformant")
plt.show()
```
---
## Different 'wedge' shapes
The linear wedge is familiar, but you can also have other shapes (`power` and `root` were new in v0.4.3):
```
modes = ['linear', 'root', 'power', 'sigmoid']
fig, axs = plt.subplots(ncols=len(modes), figsize=(15, 5))
for ax, mode in zip(axs, modes):
w, top, base, ref = bg.models.wedge(mode=mode)
ax.imshow(w, interpolation='none')
ax.axvline(ref, color='k', ls='--')
ax.plot(top, 'r-', lw=4)
ax.plot(base, 'r-', lw=4)
ax.set_title(mode)
plt.show()
```
If you're feeling creative, you can also give `wedge()` your own function (since version 0.4.3). Your function should have an API like `np.linspace()` (the function that produces the standard wedge shape). Here's an example:
```
def wavy(start, stop, num):
"""
Custom wedge shape.
"""
x = np.linspace(0, 10*np.pi, num)
y = np.sin(x) + x
# Normalize to 0-1.
y_ = (y - np.min(y)) / (np.max(y)-np.min(y))
# Scale to required output.
return min(start, stop) + abs(stop-start) * y_
# The wedge function will pass 'left' and 'right' thicknesses.
# You only need to worry about the case where left < right.
left, right = 1, 2
y = wavy(left, right, 100)
plt.plot(y)
plt.ylim(right, 0)
plt.show()
```
Let's use that function to make a model:
```
w, top, base, ref = bg.models.wedge(mode=wavy, thickness=(1, 0))
plt.imshow(w, interpolation='none')
plt.axvline(ref, color='k', ls='--')
plt.plot(top, 'r-', lw=4)
plt.plot(base, 'r-', lw=4)
plt.show()
```
---
## Varying net:gross across `breadth`
**This is a new feature introduced in v0.4.3; if you find bugs, please let us know by [raising an issue](https://github.com/agile-geoscience/bruges/issues).**
If you define a **binary wedge** — i.e. exactly 2 lithologies in the wedge layer — then you can pass a `breadth` argument to get a 3D model. The new dimension contains the 2 pure litholgies at each end, and pinches them out across the model's 'breadth'. Now the top and base are 2D arrays (surfaces through the wedge volume), while `ref` is still a scalar (the lateral position of the reference 'trace').
```
w, top, base, ref = bg.models.wedge(strat=(0, (1, 2, 1, 1, 2, 1), 3), # Binary wedge.
breadth=100)
w.shape, top.shape, base.shape, ref
```
Let's look at 3 slices: one from one end of the 'net:gross' axis (the last axis), one from the other end (right hand image), and one from halfway (middle image). These are the net:gross end-members.
```
slices = [0, 50, 99]
fig, axs = plt.subplots(ncols=len(slices), figsize=(16, 4))
for ax, slic in zip(axs, slices):
ax.imshow(w[..., slic], interpolation='none')
ax.plot(top[:, slic], 'r-', lw=4)
ax.plot(base[:, slic], 'r-', lw=4)
ax.set_title(f"Wedge slice: {slic}")
plt.show()
```
Slices in/out of the page, look like this:
```
slices = [30, 50, 90]
fig, axs = plt.subplots(ncols=len(slices), figsize=(16, 4))
for ax, slic in zip(axs, slices):
ax.imshow(w[:, slic], interpolation='none')
ax.plot(top[slic], 'r-', lw=4)
ax.plot(base[slic], 'r-', lw=4)
ax.set_title(f"Net:gross slice: {slic}")
plt.show()
```
Let's simulate the seismic:
```
rocks = np.array([2.32 * 2.65, # Rock index 0
2.35 * 2.60, # Rock index 1
2.35 * 2.62, # Rock index 2
2.37 * 2.61, # Rock index 3
])
impedance = rocks[w]
rc = (impedance[1:] - impedance[:-1]) / (impedance[1:] + impedance[:-1])
syn_ = [np.apply_along_axis(np.convolve, arr=line, axis=0, v=ricker, mode='same')
for line in np.moveaxis(rc, 1, 0)]
syn = np.moveaxis(syn_, 0, 1)
syn.shape
```
Let's look at the three orthognal profiles through this synthetic:
```
ma = np.percentile(syn, 99.9)
vols, cmaps = [w, syn], ['viridis', 'gray']
fig, axs = plt.subplots(ncols=3, nrows=2, figsize=(14, 8))
for row, vol, cm in zip(axs, vols, cmaps):
row[0].imshow(vol[:, :, 24], aspect='auto', interpolation='none', cmap=cm, vmin=-ma if vol is syn else None, vmax=ma if vol is syn else None)
row[0].axhline(40, c='w', lw=0.67)
row[0].axvline(50, c='w', lw=0.67)
row[0].set_title(f"Wedge axis")
row[1].imshow(vol[:, 50, :], aspect='auto', interpolation='none', cmap=cm, vmin=-ma if vol is syn else None, vmax=ma if vol is syn else None)
row[1].axhline(40, c='w', lw=0.67)
row[1].axvline(24, c='w', lw=0.67)
row[1].set_title(f"Net:gross axis")
row[2].imshow(vol[40, :, :], aspect='auto', interpolation='none', cmap=cm, vmin=0 if vol is w else -ma, vmax=ma if vol is syn else None)
row[2].axhline(50, c='w', lw=0.67)
row[2].axvline(24, c='w', lw=0.67)
row[2].set_title(f"Timeslice axis")
plt.show()
```
---
## Models from well logs
We can pass in arrays as `strat` and they will be used as the values in the model layers.
We'll start by loading a well and making the pieces we want to pass in as `strat`.
The middle piece will be fitted to the middle layer of the wedge (resulting in the number of pixels given in the `depth` argument. The upper and lower pieces will then be cropped to fit their layers, so you must provide enough data for this to happen. The safest thing to do is to provide the _entire_ log above and the same below. The tool `welly` makes this straightforward:
```
from welly import Well
w = Well.from_las('../data/R-39.las')
log, top, bot = 'GR', 2620, 2720 # The zone we want in the wedge
log_above = w.data[log].to_basis(stop=top)
log_wedge = w.data[log].to_basis(start=top, stop=bot)
log_below = w.data[log].to_basis(start=bot)
```
Now we can send these pieces to `wedge`; it will squeeze all of `log_wedge` into the 600 pixels allotted to the wedge layer in `depth`. But it will only use the bits of `log_above` and `log_below` that it needs for the 200 pixels above and the 400 below (adjusting for the scale implied by the wedge).
```
w, top, base, ref = bg.models.wedge(depth=(200, 600, 400),
width=(20, 260, 20),
strat=(log_above, log_wedge, log_below),
mode='sigmoid', conformance='bottom',
thickness=(0, 2)
)
log = w[:, ref]
depth = np.arange(len(log))
fig, axs = plt.subplots(figsize=(10, 6), ncols=2, gridspec_kw={'width_ratios': (1, 3)})
axs[0].plot(log, depth)
axs[0].set_ylim(depth[-1], depth[0])
axs[0].set_title('Well A: GR')
axs[1].imshow(w, aspect='auto', cmap='summer_r')
axs[1].axvline(ref, color='k', ls='--')
axs[1].plot(top, 'r-', lw=4)
axs[1].plot(base, 'r-', lw=4)
axs[1].fill_betweenx(depth, (10 - log/10)+ref, ref, color='k', alpha=0.4)
axs[1].set_title('Well A')
plt.show()
```
---
© 2021 Agile Scientific, licensed CC-BY / Apache 2.0
| github_jupyter |
```
import numpy as np
import pandas as pd
import statsrat as sr
from statsrat import rw
from statsrat import expr
import random
import nlopt
import seaborn as sns
import plotnine as p9
# IMPORTANT NOTE: the "data" in this example are synthetic, i.e. generated by simulation rather than from
# actual human participants. This is to avoid any worries privacy issues.
# DEFINE MODELS
# The derived attention model from Le Pelley, Mitchell, Beesley, George and Wills (2016).
drva = rw.model(name = 'drva',
pred = rw.pred.identity,
fbase = rw.fbase.elem,
fweight = rw.fweight.none,
lrate = rw.lrate.from_aux_feature,
drate = rw.drate.zero,
aux = rw.aux.drva)
# CompAct (with only elemental features); Model 4 from Paskewitz and Jones (2020).
CompAct = rw.model(name = 'CompAct',
pred = rw.pred.identity,
fbase = rw.fbase.elem,
fweight = rw.fweight.from_aux_norm,
lrate = rw.lrate.from_aux_norm,
drate = rw.drate.zero,
aux = rw.aux.gradcomp)
# DEFINE THE EXPERIMENT
# Loosely based on Le Pelley and McLaren 2003 (learned predictiveness)
# ADD COMMENTS TO GIVE MORE DETAIL
design = expr.schedule(resp_type = 'choice',
stages = {'training': expr.stage(
x_pn = [['a', 'v'], ['b', 'v'], ['a', 'w'], ['b', 'w'], ['c', 'x'], ['d', 'x'], ['c', 'y'], ['d', 'y']],
y = 4*[['cat1'], ['cat2']],
y_psb = ['cat1', 'cat2'],
n_rep = 14),
'transfer': expr.stage(
x_pn = [['a', 'x'], ['b', 'y'], ['c', 'v'], ['d', 'w'], ['e', 'f'], ['g', 'h'], ['i', 'j'], ['k', 'l']],
y = 4*[['cat3'], ['cat4']],
y_psb = ['cat3', 'cat4'],
n_rep = 4),
'test': expr.stage(
x_pn = [['a', 'c'], ['b', 'd'], ['v', 'x'], ['w', 'y'], ['e', 'h'], ['f', 'g'], ['i', 'j'], ['k', 'l']],
y_psb = ['cat3', 'cat4'],
lrn = False,
n_rep = 1)})
rel_irl = expr.oat(schedule_pos = ['design'],
behav_score_pos = expr.behav_score(stage = 'test',
trial_pos = ['a.c -> nothing', 'b.d -> nothing'],
trial_neg = ['v.x -> nothing', 'w.y -> nothing'],
resp_pos = ['cat3', 'cat4'],
resp_neg = ['cat3', 'cat4']))
lrn_pred = expr.experiment(schedules = {'design': design},
oats = {'rel_irl': rel_irl})
# Documentation on the read_csv method (used to import data)
help(lrn_pred.read_csv)
# IMPORT THE (SYNTHETIC) DATA
(ds, summary) = lrn_pred.read_csv(path = 'data/',
x_col = ['left_stim', 'right_stim'],
resp_col = ['key_press'],
resp_map = {'a': 'cat1', 's': 'cat2', 'd': 'cat3', 'f': 'cat4'},
ident_col = 'subject_id',
conf_col = 'confidence_rating')
# View trial by trial dataset
print(ds)
# View summary dataframe
print(summary)
# Take a random subset of the imported data to test optimization algorithms
# and figure out how long optimization takes
selection = random.sample(list(ds['ident'].values), k = 5)
subset = ds.loc[{'ident': selection}].copy()
# FIRST MODEL (DERIVED ATTENTION)
# Test different optimization algorithms (on a subset of data)
compare_drva = sr.fit_algorithm_plots(ds = subset,
model = drva,
time_interval_size = 5,
n_time_intervals = 1,
algorithm_list = [nlopt.GN_AGS, nlopt.GN_CRS2_LM, nlopt.GN_DIRECT, nlopt.GN_ESCH, nlopt.GN_MLSL])
compare_drva['plot']
print(compare_drva['df'])
# Determine how long the optimization algorithm needs to run (subset of data)
convg_drva = sr.fit_algorithm_plots(ds = subset,
model = drva,
time_interval_size = 2,
n_time_intervals = 4,
algorithm = nlopt.GN_DIRECT)
convg_drva['plot']
print(convg_drva['df'])
# Fit the model to the data (full dataset) by maximum likelihood
fit_drva = sr.fit_indv(model = drva,
ds = ds,
global_time = 5,
local_time = 5,
algorithm = nlopt.GN_DIRECT)
print(fit_drva)
# SECOND MODEL (COMPACT)
# Test different optimization algorithms (on a subset of data)
compare_CompAct = sr.fit_algorithm_plots(ds = subset,
model = CompAct,
time_interval_size = 5,
n_time_intervals = 1,
algorithm_list = [nlopt.GN_AGS, nlopt.GN_CRS2_LM, nlopt.GN_DIRECT, nlopt.GN_ESCH, nlopt.GN_MLSL])
compare_CompAct['plot']
print(compare_CompAct['df'])
# Determine how long the optimization algorithm needs to run (subset of data)
convg_CompAct = sr.fit_algorithm_plots(ds = subset,
model = drva,
time_interval_size = 2,
n_time_intervals = 4,
algorithm = nlopt.GN_DIRECT)
convg_CompAct['plot']
print(convg_CompAct['df'])
# Fit the model to the data (full dataset) by maximum likelihood
fit_CompAct = sr.fit_indv(model = CompAct,
ds = ds,
global_time = 5,
local_time = 5,
algorithm = nlopt.GN_DIRECT)
print(fit_CompAct)
# Create a data frame that combines fit data from the two models
fit_combined = pd.concat({'drva': fit_drva, 'CompAct': fit_CompAct})
print(fit_combined)
# Compare AIC (Akaike Information Criterion) values
# These are based on a log-likelihood but penalize the number of free parameters
# Higher is better
sns.stripplot(x = 'model', y = 'aic', data = fit_combined)
sns.distplot(fit_drva['aic'] - fit_CompAct['aic'])
help(sns.catplot)
p9.ggplot(p9.aes(x = 'model', y = 'aic'), data = fit_combined) + p9.geom_point()
```
| github_jupyter |
# Python Basics with Numpy (optional assignment)
Welcome to your first assignment. This exercise gives you a brief introduction to Python. Even if you've used Python before, this will help familiarize you with functions we'll need.
**Instructions:**
- You will be using Python 3.
- Avoid using for-loops and while-loops, unless you are explicitly told to do so.
- Do not modify the (# GRADED FUNCTION [function name]) comment in some cells. Your work would not be graded if you change this. Each cell containing that comment should only contain one function.
- After coding your function, run the cell right below it to check if your result is correct.
**After this assignment you will:**
- Be able to use iPython Notebooks
- Be able to use numpy functions and numpy matrix/vector operations
- Understand the concept of "broadcasting"
- Be able to vectorize code
Let's get started!
## About iPython Notebooks ##
iPython Notebooks are interactive coding environments embedded in a webpage. You will be using iPython notebooks in this class. You only need to write code between the ### START CODE HERE ### and ### END CODE HERE ### comments. After writing your code, you can run the cell by either pressing "SHIFT"+"ENTER" or by clicking on "Run Cell" (denoted by a play symbol) in the upper bar of the notebook.
We will often specify "(≈ X lines of code)" in the comments to tell you about how much code you need to write. It is just a rough estimate, so don't feel bad if your code is longer or shorter.
**Exercise**: Set test to `"Hello World"` in the cell below to print "Hello World" and run the two cells below.
```
### START CODE HERE ### (≈ 1 line of code)
test = "Hello World"
### END CODE HERE ###
print ("test: " + test)
```
**Expected output**:
test: Hello World
<font color='blue'>
**What you need to remember**:
- Run your cells using SHIFT+ENTER (or "Run cell")
- Write code in the designated areas using Python 3 only
- Do not modify the code outside of the designated areas
## 1 - Building basic functions with numpy ##
Numpy is the main package for scientific computing in Python. It is maintained by a large community (www.numpy.org). In this exercise you will learn several key numpy functions such as np.exp, np.log, and np.reshape. You will need to know how to use these functions for future assignments.
### 1.1 - sigmoid function, np.exp() ###
Before using np.exp(), you will use math.exp() to implement the sigmoid function. You will then see why np.exp() is preferable to math.exp().
**Exercise**: Build a function that returns the sigmoid of a real number x. Use math.exp(x) for the exponential function.
**Reminder**:
$sigmoid(x) = \frac{1}{1+e^{-x}}$ is sometimes also known as the logistic function. It is a non-linear function used not only in Machine Learning (Logistic Regression), but also in Deep Learning.
<img src="images/Sigmoid.png" style="width:500px;height:228px;">
To refer to a function belonging to a specific package you could call it using package_name.function(). Run the code below to see an example with math.exp().
```
# GRADED FUNCTION: basic_sigmoid
import math
def basic_sigmoid(x):
"""
Compute sigmoid of x.
Arguments:
x -- A scalar
Return:
s -- sigmoid(x)
"""
### START CODE HERE ### (≈ 1 line of code)
s = 1 / (1 + math.exp(-1 * x))
### END CODE HERE ###
return s
basic_sigmoid(3)
```
**Expected Output**:
<table style = "width:40%">
<tr>
<td>** basic_sigmoid(3) **</td>
<td>0.9525741268224334 </td>
</tr>
</table>
Actually, we rarely use the "math" library in deep learning because the inputs of the functions are real numbers. In deep learning we mostly use matrices and vectors. This is why numpy is more useful.
```
### One reason why we use "numpy" instead of "math" in Deep Learning ###
x = [1, 2, 3]
basic_sigmoid(x) # you will see this give an error when you run it, because x is a vector.
```
In fact, if $ x = (x_1, x_2, ..., x_n)$ is a row vector then $np.exp(x)$ will apply the exponential function to every element of x. The output will thus be: $np.exp(x) = (e^{x_1}, e^{x_2}, ..., e^{x_n})$
```
import numpy as np
# example of np.exp
x = np.array([1, 2, 3])
print(np.exp(x)) # result is (exp(1), exp(2), exp(3))
```
Furthermore, if x is a vector, then a Python operation such as $s = x + 3$ or $s = \frac{1}{x}$ will output s as a vector of the same size as x.
```
# example of vector operation
x = np.array([1, 2, 3])
print (x + 3)
```
Any time you need more info on a numpy function, we encourage you to look at [the official documentation](https://docs.scipy.org/doc/numpy-1.10.1/reference/generated/numpy.exp.html).
You can also create a new cell in the notebook and write `np.exp?` (for example) to get quick access to the documentation.
**Exercise**: Implement the sigmoid function using numpy.
**Instructions**: x could now be either a real number, a vector, or a matrix. The data structures we use in numpy to represent these shapes (vectors, matrices...) are called numpy arrays. You don't need to know more for now.
$$ \text{For } x \in \mathbb{R}^n \text{, } sigmoid(x) = sigmoid\begin{pmatrix}
x_1 \\
x_2 \\
... \\
x_n \\
\end{pmatrix} = \begin{pmatrix}
\frac{1}{1+e^{-x_1}} \\
\frac{1}{1+e^{-x_2}} \\
... \\
\frac{1}{1+e^{-x_n}} \\
\end{pmatrix}\tag{1} $$
```
# GRADED FUNCTION: sigmoid
import numpy as np # this means you can access numpy functions by writing np.function() instead of numpy.function()
def sigmoid(x):
"""
Compute the sigmoid of x
Arguments:
x -- A scalar or numpy array of any size
Return:
s -- sigmoid(x)
"""
### START CODE HERE ### (≈ 1 line of code)
s = 1 / (1 + np.exp(-1 * x))
### END CODE HERE ###
return s
x = np.array([1, 2, 3])
sigmoid(x)
```
**Expected Output**:
<table>
<tr>
<td> **sigmoid([1,2,3])**</td>
<td> array([ 0.73105858, 0.88079708, 0.95257413]) </td>
</tr>
</table>
### 1.2 - Sigmoid gradient
As you've seen in lecture, you will need to compute gradients to optimize loss functions using backpropagation. Let's code your first gradient function.
**Exercise**: Implement the function sigmoid_grad() to compute the gradient of the sigmoid function with respect to its input x. The formula is: $$sigmoid\_derivative(x) = \sigma'(x) = \sigma(x) (1 - \sigma(x))\tag{2}$$
You often code this function in two steps:
1. Set s to be the sigmoid of x. You might find your sigmoid(x) function useful.
2. Compute $\sigma'(x) = s(1-s)$
```
# GRADED FUNCTION: sigmoid_derivative
def sigmoid_derivative(x):
"""
Compute the gradient (also called the slope or derivative) of the sigmoid function with respect to its input x.
You can store the output of the sigmoid function into variables and then use it to calculate the gradient.
Arguments:
x -- A scalar or numpy array
Return:
ds -- Your computed gradient.
"""
### START CODE HERE ### (≈ 2 lines of code)
s = sigmoid(x)
ds = s * (1 - s)
### END CODE HERE ###
return ds
x = np.array([1, 2, 3])
print ("sigmoid_derivative(x) = " + str(sigmoid_derivative(x)))
```
**Expected Output**:
<table>
<tr>
<td> **sigmoid_derivative([1,2,3])**</td>
<td> [ 0.19661193 0.10499359 0.04517666] </td>
</tr>
</table>
### 1.3 - Reshaping arrays ###
Two common numpy functions used in deep learning are [np.shape](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.shape.html) and [np.reshape()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.reshape.html).
- X.shape is used to get the shape (dimension) of a matrix/vector X.
- X.reshape(...) is used to reshape X into some other dimension.
For example, in computer science, an image is represented by a 3D array of shape $(length, height, depth = 3)$. However, when you read an image as the input of an algorithm you convert it to a vector of shape $(length*height*3, 1)$. In other words, you "unroll", or reshape, the 3D array into a 1D vector.
<img src="images/image2vector_kiank.png" style="width:500px;height:300;">
**Exercise**: Implement `image2vector()` that takes an input of shape (length, height, 3) and returns a vector of shape (length\*height\*3, 1). For example, if you would like to reshape an array v of shape (a, b, c) into a vector of shape (a*b,c) you would do:
``` python
v = v.reshape((v.shape[0]*v.shape[1], v.shape[2])) # v.shape[0] = a ; v.shape[1] = b ; v.shape[2] = c
```
- Please don't hardcode the dimensions of image as a constant. Instead look up the quantities you need with `image.shape[0]`, etc.
```
# GRADED FUNCTION: image2vector
def image2vector(image):
"""
Argument:
image -- a numpy array of shape (length, height, depth)
Returns:
v -- a vector of shape (length*height*depth, 1)
"""
### START CODE HERE ### (≈ 1 line of code)
print(f"Actual shape of the image: {image.shape}")
v = image.reshape(image.shape[0]*image.shape[1]*image.shape[2], 1)
### END CODE HERE ###
return v
# This is a 3 by 3 by 2 array, typically images will be (num_px_x, num_px_y,3) where 3 represents the RGB values
image = np.array([[[ 0.67826139, 0.29380381],
[ 0.90714982, 0.52835647],
[ 0.4215251 , 0.45017551]],
[[ 0.92814219, 0.96677647],
[ 0.85304703, 0.52351845],
[ 0.19981397, 0.27417313]],
[[ 0.60659855, 0.00533165],
[ 0.10820313, 0.49978937],
[ 0.34144279, 0.94630077]]])
print ("image2vector(image) = " + str(image2vector(image)))
```
**Expected Output**:
<table style="width:100%">
<tr>
<td> **image2vector(image)** </td>
<td> [[ 0.67826139]
[ 0.29380381]
[ 0.90714982]
[ 0.52835647]
[ 0.4215251 ]
[ 0.45017551]
[ 0.92814219]
[ 0.96677647]
[ 0.85304703]
[ 0.52351845]
[ 0.19981397]
[ 0.27417313]
[ 0.60659855]
[ 0.00533165]
[ 0.10820313]
[ 0.49978937]
[ 0.34144279]
[ 0.94630077]]</td>
</tr>
</table>
### 1.4 - Normalizing rows
Another common technique we use in Machine Learning and Deep Learning is to normalize our data. It often leads to a better performance because gradient descent converges faster after normalization. Here, by normalization we mean changing x to $ \frac{x}{\| x\|} $ (dividing each row vector of x by its norm).
For example, if $$x =
\begin{bmatrix}
0 & 3 & 4 \\
2 & 6 & 4 \\
\end{bmatrix}\tag{3}$$ then $$\| x\| = np.linalg.norm(x, axis = 1, keepdims = True) = \begin{bmatrix}
5 \\
\sqrt{56} \\
\end{bmatrix}\tag{4} $$and $$ x\_normalized = \frac{x}{\| x\|} = \begin{bmatrix}
0 & \frac{3}{5} & \frac{4}{5} \\
\frac{2}{\sqrt{56}} & \frac{6}{\sqrt{56}} & \frac{4}{\sqrt{56}} \\
\end{bmatrix}\tag{5}$$ Note that you can divide matrices of different sizes and it works fine: this is called broadcasting and you're going to learn about it in part 5.
**Exercise**: Implement normalizeRows() to normalize the rows of a matrix. After applying this function to an input matrix x, each row of x should be a vector of unit length (meaning length 1).
```
# GRADED FUNCTION: normalizeRows
def normalizeRows(x):
"""
Implement a function that normalizes each row of the matrix x (to have unit length).
Argument:
x -- A numpy matrix of shape (n, m)
Returns:
x -- The normalized (by row) numpy matrix. You are allowed to modify x.
"""
### START CODE HERE ### (≈ 2 lines of code)
# Compute x_norm as the norm 2 of x. Use np.linalg.norm(..., ord = 2, axis = ..., keepdims = True)
x_norm = np.linalg.norm(x, axis=1, keepdims=True)
print(f'x_norm.shape = {x_norm.shape}')
print(f'x.shape = {x.shape}')
# Divide x by its norm.
x = x / x_norm
### END CODE HERE ###
return x
x = np.array([
[0, 3, 4],
[1, 6, 4]])
print("normalizeRows(x) = " + str(normalizeRows(x)))
```
**Expected Output**:
<table style="width:60%">
<tr>
<td> **normalizeRows(x)** </td>
<td> [[ 0. 0.6 0.8 ]
[ 0.13736056 0.82416338 0.54944226]]</td>
</tr>
</table>
**Note**:
In normalizeRows(), you can try to print the shapes of x_norm and x, and then rerun the assessment. You'll find out that they have different shapes. This is normal given that x_norm takes the norm of each row of x. So x_norm has the same number of rows but only 1 column. So how did it work when you divided x by x_norm? This is called broadcasting and we'll talk about it now!
### 1.5 - Broadcasting and the softmax function ####
A very important concept to understand in numpy is "broadcasting". It is very useful for performing mathematical operations between arrays of different shapes. For the full details on broadcasting, you can read the official [broadcasting documentation](http://docs.scipy.org/doc/numpy/user/basics.broadcasting.html).
**Exercise**: Implement a softmax function using numpy. You can think of softmax as a normalizing function used when your algorithm needs to classify two or more classes. You will learn more about softmax in the second course of this specialization.
**Instructions**:
- $ \text{for } x \in \mathbb{R}^{1\times n} \text{, } softmax(x) = softmax(\begin{bmatrix}
x_1 &&
x_2 &&
... &&
x_n
\end{bmatrix}) = \begin{bmatrix}
\frac{e^{x_1}}{\sum_{j}e^{x_j}} &&
\frac{e^{x_2}}{\sum_{j}e^{x_j}} &&
... &&
\frac{e^{x_n}}{\sum_{j}e^{x_j}}
\end{bmatrix} $
- $\text{for a matrix } x \in \mathbb{R}^{m \times n} \text{, $x_{ij}$ maps to the element in the $i^{th}$ row and $j^{th}$ column of $x$, thus we have: }$ $$softmax(x) = softmax\begin{bmatrix}
x_{11} & x_{12} & x_{13} & \dots & x_{1n} \\
x_{21} & x_{22} & x_{23} & \dots & x_{2n} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
x_{m1} & x_{m2} & x_{m3} & \dots & x_{mn}
\end{bmatrix} = \begin{bmatrix}
\frac{e^{x_{11}}}{\sum_{j}e^{x_{1j}}} & \frac{e^{x_{12}}}{\sum_{j}e^{x_{1j}}} & \frac{e^{x_{13}}}{\sum_{j}e^{x_{1j}}} & \dots & \frac{e^{x_{1n}}}{\sum_{j}e^{x_{1j}}} \\
\frac{e^{x_{21}}}{\sum_{j}e^{x_{2j}}} & \frac{e^{x_{22}}}{\sum_{j}e^{x_{2j}}} & \frac{e^{x_{23}}}{\sum_{j}e^{x_{2j}}} & \dots & \frac{e^{x_{2n}}}{\sum_{j}e^{x_{2j}}} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
\frac{e^{x_{m1}}}{\sum_{j}e^{x_{mj}}} & \frac{e^{x_{m2}}}{\sum_{j}e^{x_{mj}}} & \frac{e^{x_{m3}}}{\sum_{j}e^{x_{mj}}} & \dots & \frac{e^{x_{mn}}}{\sum_{j}e^{x_{mj}}}
\end{bmatrix} = \begin{pmatrix}
softmax\text{(first row of x)} \\
softmax\text{(second row of x)} \\
... \\
softmax\text{(last row of x)} \\
\end{pmatrix} $$
```
# GRADED FUNCTION: softmax
def softmax(x):
"""Calculates the softmax for each row of the input x.
Your code should work for a row vector and also for matrices of shape (n, m).
Argument:
x -- A numpy matrix of shape (n,m)
Returns:
s -- A numpy matrix equal to the softmax of x, of shape (n,m)
"""
### START CODE HERE ### (≈ 3 lines of code)
# Apply exp() element-wise to x. Use np.exp(...).
x_exp = np.exp(x)
# Create a vector x_sum that sums each row of x_exp. Use np.sum(..., axis = 1, keepdims = True).
x_sum = np.sum(x_exp, axis=1, keepdims=True)
# Compute softmax(x) by dividing x_exp by x_sum. It should automatically use numpy broadcasting.
s = x_exp / x_sum
print(f'x_exp.shape = {x_exp.shape}')
print(f'x_sum.shape = {x_sum.shape}')
print(f's.shape = {s.shape}')
### END CODE HERE ###
return s
x = np.array([
[9, 2, 5, 0, 0],
[7, 5, 0, 0 ,0]])
print("softmax(x) = " + str(softmax(x)))
```
**Expected Output**:
<table style="width:60%">
<tr>
<td> **softmax(x)** </td>
<td> [[ 9.80897665e-01 8.94462891e-04 1.79657674e-02 1.21052389e-04
1.21052389e-04]
[ 8.78679856e-01 1.18916387e-01 8.01252314e-04 8.01252314e-04
8.01252314e-04]]</td>
</tr>
</table>
**Note**:
- If you print the shapes of x_exp, x_sum and s above and rerun the assessment cell, you will see that x_sum is of shape (2,1) while x_exp and s are of shape (2,5). **x_exp/x_sum** works due to python broadcasting.
Congratulations! You now have a pretty good understanding of python numpy and have implemented a few useful functions that you will be using in deep learning.
<font color='blue'>
**What you need to remember:**
- np.exp(x) works for any np.array x and applies the exponential function to every coordinate
- the sigmoid function and its gradient
- image2vector is commonly used in deep learning
- np.reshape is widely used. In the future, you'll see that keeping your matrix/vector dimensions straight will go toward eliminating a lot of bugs.
- numpy has efficient built-in functions
- broadcasting is extremely useful
## 2) Vectorization
In deep learning, you deal with very large datasets. Hence, a non-computationally-optimal function can become a huge bottleneck in your algorithm and can result in a model that takes ages to run. To make sure that your code is computationally efficient, you will use vectorization. For example, try to tell the difference between the following implementations of the dot/outer/elementwise product.
### Matrix Multiplication in Numpy
1. **Dot product**
- Sum of corresponding element multiplication
- Results in a scalar
2. **Outer product**
- Each element of the first matrix gets multiplies to each element in the other matrix
- (m, n) * (p, q) ==> (m, p)
3. **Element-wise product**
- Each corresponding element of the matrices get multiplied
- Shape does not change
- np.multiply(x1, x2) or x1 \* x2 are equivalent. Both do element wise multiplication
4. **General Dot product**

```
import time
x1 = [9, 2, 5, 0, 0, 7, 5, 0, 0, 0, 9, 2, 5, 0, 0]
x2 = [9, 2, 2, 9, 0, 9, 2, 5, 0, 0, 9, 2, 5, 0, 0]
### CLASSIC DOT PRODUCT OF VECTORS IMPLEMENTATION ###
tic = time.process_time()
dot = 0
for i in range(len(x1)):
dot+= x1[i]*x2[i]
toc = time.process_time()
print ("dot = " + str(dot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### CLASSIC OUTER PRODUCT IMPLEMENTATION ###
tic = time.process_time()
outer = np.zeros((len(x1),len(x2))) # we create a len(x1)*len(x2) matrix with only zeros
for i in range(len(x1)):
for j in range(len(x2)):
outer[i,j] = x1[i]*x2[j]
toc = time.process_time()
print ("outer = " + str(outer) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### CLASSIC ELEMENTWISE IMPLEMENTATION ###
tic = time.process_time()
mul = np.zeros(len(x1))
for i in range(len(x1)):
mul[i] = x1[i]*x2[i]
toc = time.process_time()
print ("elementwise multiplication = " + str(mul) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### CLASSIC GENERAL DOT PRODUCT IMPLEMENTATION ###
W = np.random.rand(3,len(x1)) # Random 3*len(x1) numpy array
tic = time.process_time()
gdot = np.zeros(W.shape[0])
for i in range(W.shape[0]):
for j in range(len(x1)):
gdot[i] += W[i,j]*x1[j]
toc = time.process_time()
print ("gdot = " + str(gdot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
x1 = [9, 2, 5, 0, 0, 7, 5, 0, 0, 0, 9, 2, 5, 0, 0]
x2 = [9, 2, 2, 9, 0, 9, 2, 5, 0, 0, 9, 2, 5, 0, 0]
### VECTORIZED DOT PRODUCT OF VECTORS ###
tic = time.process_time()
dot = np.dot(x1,x2)
toc = time.process_time()
print ("dot = " + str(dot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED OUTER PRODUCT ###
tic = time.process_time()
outer = np.outer(x1,x2)
toc = time.process_time()
print ("outer = " + str(outer) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED ELEMENTWISE MULTIPLICATION ###
tic = time.process_time()
mul = np.multiply(x1,x2)
toc = time.process_time()
print ("elementwise multiplication = " + str(mul) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED GENERAL DOT PRODUCT ###
tic = time.process_time()
dot = np.dot(W,x1)
toc = time.process_time()
print ("gdot = " + str(dot) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
```
As you may have noticed, the vectorized implementation is much cleaner and more efficient. For bigger vectors/matrices, the differences in running time become even bigger.
**Note** that `np.dot()` performs a matrix-matrix or matrix-vector multiplication. This is different from `np.multiply()` and the `*` operator (which is equivalent to `.*` in Matlab/Octave), which performs an element-wise multiplication.
### 2.1 Implement the L1 and L2 loss functions
**Exercise**: Implement the numpy vectorized version of the L1 loss. You may find the function abs(x) (absolute value of x) useful.
**Reminder**:
- The loss is used to evaluate the performance of your model. The bigger your loss is, the more different your predictions ($ \hat{y} $) are from the true values ($y$). In deep learning, you use optimization algorithms like Gradient Descent to train your model and to minimize the cost.
- L1 loss is defined as:
$$\begin{align*} & L_1(\hat{y}, y) = \sum_{i=0}^m|y^{(i)} - \hat{y}^{(i)}| \end{align*}\tag{6}$$
```
# GRADED FUNCTION: L1
def L1(yhat, y):
"""
Arguments:
yhat -- vector of size m (predicted labels)
y -- vector of size m (true labels)
Returns:
loss -- the value of the L1 loss function defined above
"""
### START CODE HERE ### (≈ 1 line of code)
loss = np.sum(abs(y - yhat))
### END CODE HERE ###
return loss
yhat = np.array([.9, 0.2, 0.1, .4, .9])
y = np.array([1, 0, 0, 1, 1])
print("L1 = " + str(L1(yhat,y)))
```
**Expected Output**:
<table style="width:20%">
<tr>
<td> **L1** </td>
<td> 1.1 </td>
</tr>
</table>
**Exercise**: Implement the numpy vectorized version of the L2 loss. There are several way of implementing the L2 loss but you may find the function np.dot() useful. As a reminder, if $x = [x_1, x_2, ..., x_n]$, then `np.dot(x,x)` = $\sum_{j=0}^n x_j^{2}$.
- L2 loss is defined as $$\begin{align*} & L_2(\hat{y},y) = \sum_{i=0}^m(y^{(i)} - \hat{y}^{(i)})^2 \end{align*}\tag{7}$$
```
# GRADED FUNCTION: L2
def L2(yhat, y):
"""
Arguments:
yhat -- vector of size m (predicted labels)
y -- vector of size m (true labels)
Returns:
loss -- the value of the L2 loss function defined above
"""
### START CODE HERE ### (≈ 1 line of code)
loss = np.sum(np.dot(y - yhat, y - yhat))
### END CODE HERE ###
return loss
yhat = np.array([.9, 0.2, 0.1, .4, .9])
y = np.array([1, 0, 0, 1, 1])
print("L2 = " + str(L2(yhat,y)))
```
**Expected Output**:
<table style="width:20%">
<tr>
<td> **L2** </td>
<td> 0.43 </td>
</tr>
</table>
Congratulations on completing this assignment. We hope that this little warm-up exercise helps you in the future assignments, which will be more exciting and interesting!
<font color='blue'>
**What to remember:**
- Vectorization is very important in deep learning. It provides computational efficiency and clarity.
- You have reviewed the L1 and L2 loss.
- You are familiar with many numpy functions such as np.sum, np.dot, np.multiply, np.maximum, etc...
| github_jupyter |
<h1 align="center">Segmentation Evaluation</h1>
Evaluating segmentation algorithms is most often done using reference data to which you compare your results.
In the medical domain reference data is commonly obtained via manual segmentation by an expert (don't forget to thank your clinical colleagues for their hard work). When you are resource limited, the reference data may be defined by a single expert. This is less than ideal. When multiple experts provide you with their input then you can potentially combine them to obtain reference data that is closer to the ever elusive "ground truth". In this notebook we show two approaches to combining input from multiple observers, majority vote and the Simultaneous Truth and Performance Level
Estimation [(STAPLE)](http://crl.med.harvard.edu/research/staple/).
Once we have a reference, we compare the algorithm's performance using multiple criteria, as usually there is no single evaluation measure that conveys all of the relevant information. In this notebook we illustrate the use of the following evaluation criteria:
* Overlap measures:
* Jaccard and Dice coefficients
* false negative and false positive errors
* Surface distance measures:
* mean, median, max and standard deviation between surfaces
* Volume measures:
* volume similarity $ \frac{2*(v1-v2)}{v1+v2}$
The relevant criteria are task dependent, so you need to ask yourself whether you are interested in detecting spurious errors or not (mean or max surface distance), whether over/under segmentation should be differentiated (volume similarity and Dice or just Dice), and what is the ratio between acceptable errors and the size of the segmented object (Dice coefficient may be too sensitive to small errors when the segmented object is small and not sensitive enough to large errors when the segmented object is large).
The data we use in the notebook is a set of manually segmented liver tumors from a single clinical CT scan. A larger dataset (four scans) is freely available from this [MIDAS repository](http://www.insight-journal.org/midas/collection/view/38). The relevant publication is: T. Popa et al., "Tumor Volume Measurement and Volume Measurement Comparison Plug-ins for VolView Using ITK", SPIE Medical Imaging: Visualization, Image-Guided Procedures, and Display, 2006.
<b>Note</b>: The approach described here can also be used to evaluate Registration, as illustrated in the [free form deformation notebook](65_Registration_FFD.ipynb).
```
library(SimpleITK)
source("downloaddata.R")
```
## Utility functions
Display related utility functions.
```
## save the default options in case you need to reset them
if (!exists("default.options"))
{
default.options <- options()
}
# display 2D images inside the notebook (colour and greyscale)
show_inline <- function(object, Dwidth=grid::unit(5, "cm"))
{
ncomp <- object$GetNumberOfComponents()
if (ncomp == 3) {
## colour
a <- as.array(object)
a <- aperm(a, c(2, 1, 3))
} else if (ncomp == 1) {
a <- t(as.array(object))
} else {
stop("Only deals with 1 or 3 component images")
}
rg <- range(a)
A <- (a - rg[1]) / (rg[2] - rg[1])
dd <- dim(a)
sp <- object$GetSpacing()
sz <- object$GetSize()
worlddim <- sp * sz
worlddim <- worlddim / worlddim[1]
W <- Dwidth
H <- Dwidth * worlddim[2]
WW <- grid::convertX(W*1.1, "inches", valueOnly=TRUE)
HH <- grid::convertY(H*1.1, "inches", valueOnly=TRUE)
## here we set the display size
## Jupyter only honours the last setting for a cell, so
## we can't reset the original options. That needs to
## be done manually, using the "default.options" stored above
## Obvious point to do this is before plotting graphs
options(repr.plot.width = WW, repr.plot.height = HH)
grid::grid.raster(A, default.units="mm", width=W, height=H)
}
# Tile images to create a single wider image.
color_tile <- function(images)
{
width <- images[[1]]$GetWidth()
height <- images[[1]]$GetHeight()
tiled_image <- Image(c(length(images) * width, height), images[[1]]$GetPixelID(), images[[1]]$GetNumberOfComponentsPerPixel())
for(i in 1:length(images))
{
tiled_image <- Paste(tiled_image, images[[i]], images[[i]]$GetSize(), c(0, 0), c((i - 1) * width, 0))
}
return( tiled_image )
}
```
## Fetch the data
Retrieve a single CT scan and three manual delineations of a liver tumor. Visual inspection of the data highlights the variability between experts.
All computations are done in 3D (the dimensionality of the images). For display purposes we selected a single slice_for_display. Change this variable's value to see other slices.
```
slice_for_display <- 77
image <- ReadImage(fetch_data("liverTumorSegmentations/Patient01Homo.mha"))
# For display we need to window-level the slice (map the high dynamic range to a reasonable display)
display_slice <- Cast(IntensityWindowing(image[,,slice_for_display],
windowMinimum=-1024,
windowMaximum=976),
"sitkUInt8")
segmentation_file_names <- list("liverTumorSegmentations/Patient01Homo_Rad01.mha",
"liverTumorSegmentations/Patient01Homo_Rad02.mha",
"liverTumorSegmentations/Patient01Homo_Rad03.mha")
segmentations <- lapply(segmentation_file_names, function(x) ReadImage(fetch_data(x),"sitkUInt8"))
# Overlay the segmentation contour from each of the segmentations onto the "slice_for_display"
display_overlays <- lapply(segmentations,
function(seg) LabelMapContourOverlay(Cast(seg[,,slice_for_display], "sitkLabelUInt8"),
display_slice,
opacity = 1))
show_inline(color_tile(display_overlays),grid::unit(15, "cm"))
```
## Derive a reference
There are a variety of ways to derive a reference segmentation from multiple expert inputs. Several options, there are more, are described in "A comparison of ground truth estimation methods", A. M. Biancardi, A. C. Jirapatnakul, A. P. Reeves.
Two methods that are available in SimpleITK are <b>majority vote</b> and the <b>STAPLE</b> algorithm.
```
# Use majority voting to obtain the reference segmentation. Note that this filter does not resolve ties. In case of
# ties, it will assign max_label_value+1 or a user specified label value (labelForUndecidedPixels) to the result.
# Before using the results of this filter you will have to check whether there were ties and modify the results to
# resolve the ties in a manner that makes sense for your task. The filter implicitly accommodates multiple labels.
labelForUndecidedPixels <- 10
reference_segmentation_majority_vote <- LabelVoting(segmentations, labelForUndecidedPixels)
show_inline(LabelMapContourOverlay(Cast(reference_segmentation_majority_vote[,,slice_for_display], "sitkLabelUInt8"), display_slice, opacity = 1),
grid::unit(5, "cm"))
# Use the STAPLE algorithm to obtain the reference segmentation. This implementation of the original algorithm
# combines a single label from multiple segmentations, the label is user specified. The result of the
# filter is the voxel's probability of belonging to the foreground. We then have to threshold the result to obtain
# a reference binary segmentation.
foregroundValue <- 1
threshold <- 0.95
reference_segmentation_STAPLE_probabilities <- STAPLE(segmentations, foregroundValue)
# We use the overloaded operator to perform thresholding, another option is to use the BinaryThreshold function.
reference_segmentation_STAPLE <- reference_segmentation_STAPLE_probabilities > threshold
show_inline(LabelMapContourOverlay(Cast(reference_segmentation_STAPLE[,,slice_for_display], "sitkLabelUInt8"), display_slice, opacity = 1),
grid::unit(5, "cm"))
```
## Evaluate segmentations using the reference
Once we derive a reference from our experts input we can compare segmentation results to it.
Note that in this notebook we compare the expert segmentations to the reference derived from them. This is not relevant for algorithm evaluation, but it can potentially be used to rank your experts.
### Utility functions
These functions compute standard overlap and surface distance measures used when comparing segmentations.
```
# Compare the two given segmentations using overlap measures (Jaccard, Dice, etc.)
compute_overlap_measures <- function(segmentation, reference_segmentation)
{
omf <- LabelOverlapMeasuresImageFilter()
omf$Execute(reference_segmentation, segmentation)
result <- c(omf$GetJaccardCoefficient(), omf$GetDiceCoefficient(),
omf$GetVolumeSimilarity(), omf$GetFalseNegativeError(), omf$GetFalsePositiveError())
names(result) <- c("JaccardCoefficient", "DiceCoefficient", "VolumeSimilarity",
"FalseNegativeError", "FalsePositiveError")
return (result)
}
# Compare a segmentation to the reference segmentation using distances between the two surfaces. To facilitate
# surface distance computations we use a distance map of the reference segmentation.
compute_surface_distance_measures <- function(segmentation, reference_distance_map)
{
segmented_label = 1
# Get the intensity statistics associated with each of the labels, combined
# with the distance map image this gives us the distances between surfaces.
lisf <- LabelIntensityStatisticsImageFilter()
# Get the pixels on the border of the segmented object
segmented_surface <- LabelContour(segmentation)
lisf$Execute(segmented_surface, reference_distance_map)
result <- c(lisf$GetMean(segmented_label), lisf$GetMedian(segmented_label),
lisf$GetStandardDeviation(segmented_label), lisf$GetMaximum(segmented_label))
names(result) <- c("Mean", "Median", "SD", "Max")
return (result)
}
```
Evaluate the three segmentations with respect to the STAPLE based reference.
```
overlap_measures <- t(sapply(segmentations, compute_overlap_measures,
reference_segmentation=reference_segmentation_STAPLE))
overlap_measures <- as.data.frame(overlap_measures)
overlap_measures$rater <- rownames(overlap_measures)
distance_map_filter <- SignedMaurerDistanceMapImageFilter()
distance_map_filter$SquaredDistanceOff()
STAPLE_reference_distance_map <-
abs(distance_map_filter$Execute(reference_segmentation_STAPLE))
surface_distance_measures <- t(sapply(segmentations,
compute_surface_distance_measures,
reference_distance_map=STAPLE_reference_distance_map))
surface_distance_measures <- as.data.frame(surface_distance_measures)
surface_distance_measures$rater <- rownames(surface_distance_measures)
# Look at the results using the notebook's default display format for data frames
overlap_measures
surface_distance_measures
```
## Improved output
If the [tidyr](https://cran.r-project.org/web/packages/tidyr/index.html) and [ggplot2](https://cran.r-project.org/web/packages/ggplot2/index.html) packages are installed in your R environment then you can easily produce high quality output.
```
library(tidyr)
library(ggplot2)
## reset the plot size
options(default.options)
overlap.gathered <- gather(overlap_measures, key=Measure, value=Score, -rater)
ggplot(overlap.gathered,
aes(x=rater, y=Score, group=Measure, fill=Measure)) +
geom_bar(stat="identity", position="dodge", colour='black', alpha=0.5)
surface_distance.gathered <- gather(surface_distance_measures, key=Measure, value=Score, -rater)
ggplot(surface_distance.gathered,
aes(x=rater, y=Score, group=Measure, fill=Measure)) +
geom_bar(stat="identity", position="dodge", colour='black', alpha=0.5)
```
You can also export the data as a table for your LaTeX manuscript using the [xtable](https://cran.r-project.org/web/packages/xtable/index.html) package, just copy paste the output of the following cell into your document.
```
library(xtable)
sd <- surface_distance_measures
sd$rater <- NULL
print(xtable(sd, caption="Segmentation surface distance measures per rater.",
label="tab:surfdist", digits=2))
```
| github_jupyter |
## Summarizing annotations to a term and descendants
This notebook demonstrates summarizing annotation counts for a term and its descendants.
An example use of this is a GO annotator exploring refactoring a subtree in GO
Of course, if this were a regular thing we would make a command line or even web interface,
but keeping as a notebook gives us some flexibility in logic, and anyway is intended largely
as a demonstration
### boilerplate
* importing relevant ontobiolibraries
* set up key objects
```
import pandas as pd
## Create an ontology factory in order to fetch GO
from ontobio.ontol_factory import OntologyFactory
ofactory = OntologyFactory()
## GOLR queries
from ontobio.golr.golr_query import GolrAssociationQuery
## rendering ontologies
from ontobio import GraphRenderer
## Load GO. Note the first time this runs Jupyter will show '*' - be patient
ont = ofactory.create("go")
```
### Finding descendants
Here we are using the in-memory ontology object, no external service calls are executed
Change the value of `term_id` to what you like
```
term_id = "GO:0009070" ## serine family amino acid biosynthetic process
descendants = ont.descendants(term_id, reflexive=True, relations=['subClassOf', 'BFO:0000050'])
descendants
```
### rendering subtrees
We use the good-old-fashioned Tree renderer
(this doesn't scale well for latticey-subontologies)
```
renderer = GraphRenderer.create('tree')
print(renderer.render_subgraph(ont, nodes=descendants))
```
### summarizing annotations
We write a short procedure to wrap calling Golr and returning a summary dict
The dict is keyed by taxon label. We also include an entry for `ALL`
```
DEFAULT_FACET_FIELDS = ['taxon_subset_closure_label', 'evidence_label', 'assigned_by']
def summarize(t: str,
evidence_closure='ECO:0000269', ## restrict to experimental
facet_fields=None) -> dict:
"""
Summarize a term
"""
if facet_fields == None:
facet_fields = DEFAULT_FACET_FIELDS
q = GolrAssociationQuery(object=t, rows=0, object_category='function',
fq={'evidence_closur'taxon_subset_closure_label'e_label':'experimental evidence'},
facet_fields=facet_fields)
#params = q.solr_params()
#print(params)
result = q.exec()
fc = result['facet_counts']
item = {'ALL': result['numFound']} ## make sure this is the first entry
for ff in facet_fields:
if ff in fc:
item.update(fc[ff])
return item
print(summarize(term_id))
def summarize_set(ids, facet_fields=None) -> pd.DataFrame:
"""
Summarize a set of annotations, return a dataframe
"""
items = []
for id in ids:
item = {'id': id, 'name:': ont.label(id)}
for k,v in summarize(id, facet_fields=facet_fields).items():
item[k] = v
items.append(item)
df = pd.DataFrame(items).fillna(0)
# sort using total number
df.sort_values('ALL', axis=0, ascending=False, inplace=True)
return df
```
## Summarize GO term and descendants
More advanced visualziations are easy with plotly etc. We leave as an exercise to the reader...
As an example, for the first query we bundle all facets (species, evidence, assigned by) together
```
pd.options.display.max_columns = None
df = summarize_set(descendants)
df
```
## Summary by assigned by
```
summarize_set(descendants, facet_fields=['assigned_by'])
```
### Summarize by species
use `taxon_subset_closure_label` facet
```
summarize_set(descendants, facet_fields=['taxon_subset_closure_label'])
```
| github_jupyter |
```
def load_pickle(fname):
with open(fname, 'rb') as f:
return pickle.load(f)
## time
def aexp2zred(aexp):
return [1.0/a - 1.0 for a in aexp]
def zred2aexp(zred):
return [1.0/(1.0 + z) for z in zred]
def lbt2aexp(lts):
import astropy.units as u
from astropy.cosmology import WMAP7, z_at_value
zreds = [z_at_value(WMAP7.lookback_time, ll * u.Gyr) for ll in lts]
return [1.0/(1+z) for z in zreds]
def density_map(x, y, sort=True):
from scipy.stats import gaussian_kde
xy = np.vstack([x,y])
z = gaussian_kde(xy)(xy)
z /= max(z)
idx = z.argsort()
xx, yy = x[idx], y[idx]
z = z[idx]
return xx, yy, z
def sigma_clip_ind(c, high, low):
"""
returns indices of sigma-clipping-safe elements.
"""
import numpy as np
ind = (np.mean(c) - np.std(c)*low < c) * (c < np.mean(c) + np.std(c)*high)
return ind
def mask_outlier(y, low=1.5, high=1.5):
"""
maks outlier assuming monotonic trend.
"""
x = np.arange(len(y))
# linear fitting .. more desirably, a very strong smoothing scheme that can reconstrcut mild curve.
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(x,y)
# extract linear fit
yy = y - (slope * x + intercept)
# sigma clipped value = mean of the rest
i_good = sigma_clip_ind(yy, low, high)
yy[~i_good] = np.mean(yy[i_good])
# add linear fit again
return yy + (slope * x + intercept)
def smooth(x, beta=5, window_len=20, monotonic=False, clip_tail_zeros=True):
"""
kaiser window smoothing.
If len(x) < window_len, window_len is overwritten to be len(x).
This ensures to return valid length fo array, but with modified window size.
beta = 5 : Similar to a Hamming
"""
if clip_tail_zeros:
x = x[:max(np.where(x > 0)[0])+1]
if monotonic:
"""
if there is an overall slope, smoothing may result in offset.
compensate for that.
"""
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(x, y=np.arange(len(x)))
xx = np.arange(len(x)) * slope + intercept
x = x - xx
# extending the data at beginning and at the end
# to apply the window at the borders
window_len = min([window_len, len(x)])
s = np.r_[x[window_len-1:0:-1], x, x[-1:-window_len:-1]] # concatenate along 0-th axis.
# periodic boundary.
w = np.kaiser(window_len,beta)
y = np.convolve(w/w.sum(), s, mode='valid')
if monotonic:
return y[int(window_len)/2:len(y)-int(window_len/2) + 1] + xx
else:
return y[int(window_len)/2:len(y)-int(window_len/2) + 1]
#return y[5:len(y)-5]
class MainPrg():
import tree.ctutils as ctu
import numpy as np
def __init__(self, treedata, final_gal, nout_ini=None, nout_fi=None):
temp_tree = ctu.extract_main_tree(treedata, final_gal)
if nout_ini == None:
nout_ini = min(temp_tree['nout'])
if nout_fi == None:
nout_fi = max(temp_tree['nout'])
self.nouts = np.arange(nout_fi, nout_ini -1, -1)
self.idxs = temp_tree['id'] # nout_ini, nout_fi consideration needed.
self.ids = temp_tree['Orig_halo_id']
self.data = None
def set_data(self, cat, nout):
"""
compile data from catalogs.
"""
if nout in self.nouts:
# Declare self.data first if there isn't.
if self.data == None:
self.data = np.zeros(len(self.nouts), dtype=cat.dtype)
inow = self.nouts == nout
a = np.where(cat['idx'] == self.idxs[inow])[0]
if len(a) > 0:
self.data[inow] = cat[a]
else:
pass
#print(self.ids[inow],cat['id'])
else:
pass
#print("No {} in the catalog".format(nout))
def clip_non_detection(self):
# end of galaxy tree = last non-zero position.
# Note that 'id' can be 0 if phantom. But phantom is a valid datapoint
i_first_nout = max(np.where(self.data['idx'] > 0)[0])
#print('i_first', i_first_nout)
# then, only [0-i_first_nout] are valid.
# earlier then 187 - 91-th are zero. so get rid of them.
self.data = self.data[:i_first_nout].copy()
self.nouts = self.nouts[:i_first_nout].copy()
self.ids = self.ids[:i_first_nout].copy()
self.idxs = self.idxs[:i_first_nout].copy()
def fill_missing_data(self):
assert (self.ids[-1] != 0)
# position angles cannot be linearly interpolated.
# skip.
#
# position and velocity are also not that linear..
# but let me just interpolate them.
#
#
# excluded=["lambda_arr2"]
filled_fields = ["eps", "epsh", "epsq", "lambda_12kpc",
"lambda_arr", "lambda_arrh",
"lambda_r","lambda_r12kpc",
"lambda_r2","lambda_rh","mgas","mrj","mstar",
"reff","reff2","rgal","rgal2","rhalo","rscale_lambda",
"sfr","sma","smah","smaq","smi","smih","smiq","ssfr",
"vxc", "vyc", "vzc", "xc", "yc", "zc"]
i_good_max = max(np.where(gal.data["reff"] > 0)[0])
i_bad = np.where(gal.data['idx'] == 0)[0]
i_bad = i_bad[i_bad < i_good_max]
if len(i_bad) > 0:
for field in filled_fields:
# do not modify index and id fields.
arr = gal.data[field] # it's a view.
for i_b in i_bad:
# neighbouring array might also be empty. Search for closest valid element.
# left point
i_l = i_b - 1
while(i_l in i_bad):
i_l = i_l - 1
# right point
i_r = i_b + 1
while(i_r in i_bad):
i_r = i_r + 1
arr[i_b] = (arr[i_b -1] + arr[i_b +1])/2.
def fixed_ind_Lr(gal):
nnouts = len(gal.nouts)
ind_reff_fix = np.zeros(nnouts, dtype='i4')
#print(gal.data['rgal'])
smooth_r = smooth(mask_outlier(gal.data['rgal'], 1.5, 1.5), 50, monotonic=False)
# fixed Reff array
for i in range(nnouts):
# 1Reff = 5 points
reff_real = smooth_r[i]
reff = gal.data['rgal'][i]
try:
ind_reff_fix[i] = np.round(reff_real/reff * 5) -1
except:
pass
return ind_reff_fix
def smoothed_reff(cat, nout_merger):
"""
returns "representative" lambda at each nout by assuming monotonic change in Reff.
During merger, Reff can fluctuate, and if has no physical meaning to infer Labda at Reff during merger stage.
So Reff' is derived by linear interpolating Reffs before and after the merger.
cat is one galaxy catalog over time.
"""
import utils.match as mtc
i_merger = np.where(cat['nout'] == nout_merger)[0]
ind_lower = 20
ind_upper = 20
reffs = cat['rgal']
# left and right values chosen by sigma-clipping
r_lefts, b, c = scipy.stats.sigmaclip(reffs[max([0,i_merger-ind_lower]):i_merger], sig_lower, sig_upper)
#print(r_lefts)
r_left = r_lefts[-1]
i_left = np.where(reffs == r_left)[0]
r_rights, b,c = scipy.stats.sigmaclip(reffs[i_merger:min([i_merger+ind_upper,len(reffs)])], sig_lower, sig_upper)
r_right = r_rights[0]
i_right = np.where(reffs == r_right)[0]
r_prime = reffs
#print("chekc")
#print(r_prime)
r_prime[i_left : i_right + 1] = np.linspace(r_left, r_right, i_right - i_left + 1)
return r_prime
def l_at_smoothed_r(gal, npix_per_reff=5):
npix_per_reff = 5
n_valid_points = sum(gal.data["reff"] > 0)
new_l_arr = np.zeros(n_valid_points)
new_reff=smooth(gal.data["reff"][gal.data["reff"] > 0])
for i in range(n_valid_points):
try:
lambdar = gal.data["lambda_arr"][i]
ind_org = npix_per_reff - 1
i_new = new_reff[i]/gal.data["reff"][i] * ind_org # 0-indexed.
il = np.fix(i_new).astype(int)
ir = il + 1
if ir >= len(lambdar):
new_l_arr[i] = lambdar[-1]
else:
new_l_arr[i] = lambdar[il]*(ir-ind_org) + lambdar[ir]*(ind_org-il)
except:
new_l_arr[i] = gal.data["lambda_arr"][i] # = 0 with bad measurements.
return new_l_arr
def get_mr_dl(cluster, mr, dl):
wdir = base + cluster + '/'
# Serialize catalogs. -> Only main galaxies
# main galaxy list
alltrees = ctu.load_tree(wdir, is_gal=True)
ad = alltrees.data
tn = ad[ad['nout'] == nout_fi]
cat = load_cat(wdir + cdir + 'catalog' + str(nout_fi) + '.pickle')
#idx_all = [tn['id'][tn['Orig_halo_id'] == id_final][0] for id_final in cat['id']]
idx_all = cat['idx'][cat["idx"] > 0].astype(int) # why idx are float???
mpg_tmp = []
for i, idx in enumerate(idx_all):
#print(i, idx)
mpg_tmp.append(MainPrg(ad, idx))
# mpg_tmp =[MainPrg(ad, idx) for idx in idx_all]
for nout in range(nout_ini, nout_fi + 1):
cat = load_cat(wdir + cdir + 'catalog' + str(nout) + '.pickle')
for gal in mpg_tmp:
gal.set_data(cat, nout)
# print(nout)
while len(mpg_tmp) > 0:
mpgs.append(mpg_tmp.pop())
with open('main_prgs_GM.pickle', 'wb') as f:
pickle.dump(mpgs, f)
#wdir = './' + cluster + '/' #'05427/'
# Serialize catalogs. -> Only main galaxies
# main galaxy list
#alltrees = ctu.load_tree(wdir, is_gal=True)
#ad = alltrees.data
#tn = ad[ad['nout'] == nout_fi] # tree now
#cat = load_cat(wdir + cdir + 'catalog' + str(nout_fi) + '.pickle')
#last_idx_all = cat['idx'] # idx of galaxies at the last snapshot.
# list of Main progenitor class.
# feed in alltree.data and idx to initialize MainPrg object.
#mpgs = [MainPrg(ad, idx) for idx in last_idx_all]
# Compile catalogs ##################
#for nout in range(nout_ini, nout_fi + 1):
# cat = pickle.load(open(wdir + cdir + 'catalog' + str(nout) + '.pickle', 'rb'))
# for mpg in mpgs:
# mpg.set_data(cat, nout) # fill up MainPrg data
# print(nout, end='\r')
# load merger galaxy list (geneated by scripts/notebooks/halo/Merter_no_cat.ipynb) ##################
with open(wdir + 'merger_list.txt', 'rb') as f:
mgl = np.genfromtxt(f, dtype=[('idx','i8'),('mr','f8'),('nout','i4')])
mids = mgl['idx'] # Merger-galaxy IDs
mrs = mgl['mr'] # Merger ratios
nout_mergers = mgl['nout'] # merger epoch
i_mergers = nout_fi - nout_mergers # merger epoch as nout index.
gal_idxs = [gal.idxs[0] for gal in mpgs] # idx of all galaxies from cluster catalog.
bad = 0
#1. merger list에 있는 은하들에 대해서 은하 catalog를 업데이트 한다.
#2. 은하의 catalog가 충분히 긴지 확인
#3. Rvir이 요동치는 경우 Reff를 보정. (요동치기 전, 후의 값을 linear interpolate)
#4. 전-후의 lambda를 측정해서 dl vs mr 측정.
for igal, mid in enumerate(mids):
#gal = mpgs[3]
if mid not in gal_idxs:
print("Merger gal {} is not in catalog, skipping".format(mid))
continue
else:
gal = mpgs[np.where(gal_idxs == mid)[0]]
if len(gal.nouts) < 20:
continue
gal.clip_non_detection()
try:
gal.fill_missing_data()
except:
bad = bad + 1
pass
if verbose: print("Galaxy ID at the final nout, idx = {}, id = {}".format(gal.idxs[0], gal.ids[0]))
i_merger = i_mergers[igal] #i_merger = 187 - mgl[igal][2]
merger_ratio = mrs[igal]
if verbose: print("nnouts: {}, i_merger {}".format(len(gal.nouts), i_merger))
if i_merger > len(gal.nouts):
print("Too short evolution history, aborting..")
continue
# fixed Lambda array based on Reff_fix.
if fix_ind:
ind_reff_fix = fixed_ind_Lr(gal)
lam = np.zeros(len(ind_reff_fix))
ind_max = len(gal.data['lambda_arr'][0]) - 1
for inout, ind in enumerate(ind_reff_fix):#[ind_reff_fix > 0]):
if ind == 0 : print(ind)
lam[inout] = gal.data['lambda_arr'][inout][min([ind_max,ind])] # fixed value
else:
lam = gal.data['lambda_r']
x_al = range(max([0,i_merger-ind_lower]), i_merger) # nout before merger
x_ar = range(i_merger,min([i_merger+ind_upper, len(lam)])) # nout after merger
# representative value of lambda BEFORE the merger
al, b1, c1 = scipy.stats.sigmaclip(lam[x_al], sig_lower, sig_upper)
# representative value of lambda AFTER the merger
ar, b2, c2 = scipy.stats.sigmaclip(lam[x_ar], sig_lower, sig_upper)
if (len(al) > 1) & (len(ar) > 0):
dl.append(np.median(ar) - np.median(al))
mr.append(merger_ratio)
else:
print("error in measuring lambda")
from analysis.misc import load_cat
import numpy as np
import scipy.stats
import tree.ctutils as ctu
import matplotlib.pyplot as plt
# Read a single galaxy evolution catalog.
import pickle
verbose=True
# In[4]:
base = './'
cdir = ['catalog/', 'easy/', 'catalog_GM/', "easy_final/"][3]
clusters = ['05427', '05420', '29172', \
'29176', '10002', '36415',
'06098', '39990', '36413', '04466','17891', '07206', '01605', '35663'][:-5]
# parameters used for lambda_arr clipping.
ind_upper = 20
ind_lower = 20
sig_upper = 2.0
sig_lower = 2.0
nout_ini = 70
nout_fi = 187
bad = 0
#ngals_tot = 0
#for cluster in clusters:
# wdir = base + cluster + '/'
# # main galaxy list
# cat = load_cat(wdir + cdir + 'catalog' + str(nout_fi) + '.pickle')
# ngals_tot = ngals_tot + len(cat['idx'] > 0)
# # some galaxies without complete tree are in the catalog.
# # exclude them for this analysis.
#nnouts = nout_fi - nout_ini + 1
mpgs = []
for cluster in clusters:
print(cluster)
wdir = base + cluster + '/'
# Serialize catalogs. -> Only main galaxies
# main galaxy list
alltrees = ctu.load_tree(wdir, is_gal=True)
ad = alltrees.data
tn = ad[ad['nout'] == nout_fi]
cat = load_cat(wdir + cdir + 'catalog' + str(nout_fi) + '.pickle')
#idx_all = [tn['id'][tn['Orig_halo_id'] == id_final][0] for id_final in cat['id']]
idx_all = cat['idx'][cat["idx"] > 0].astype(int) # why idx are float???
mpg_tmp = []
for i, idx in enumerate(idx_all):
#print(i, idx)
mpg_tmp.append(MainPrg(ad, idx))
# mpg_tmp =[MainPrg(ad, idx) for idx in idx_all]
for nout in range(nout_ini, nout_fi + 1):
cat = load_cat(wdir + cdir + 'catalog' + str(nout) + '.pickle')
for gal in mpg_tmp:
gal.set_data(cat, nout)
gal.cluster = int(cluster)
# print(nout)
while len(mpg_tmp) > 0:
mpgs.append(mpg_tmp.pop())
with open('main_prgs_GM.pickle', 'wb') as f:
pickle.dump(mpgs, f)
from matplotlib.backends.backend_pdf import PdfPages
with PdfPages("lambda_plots.pdf") as pdf:
for gal in mpgs:
#print(sum(gal.data["reff"] > 0),len(smooth(gal.data["reff"])))
#smoothed_reff = smooth(gal.data["reff"])
gal.fill_missing_data()
fig, ax = plt.subplots(2, sharex=True)
plt.title(str(gal.cluster) + " " + str(gal.idxs[0]) +" " + str(gal.ids[0]))
ax[0].plot(gal.data["reff"], label="org")
ax[0].plot(smooth(gal.data["reff"], window_len=10), "r--", label="smoothed")
ax[1].plot(gal.data["lambda_r"], label="org")
ax[1].plot(smooth(gal.data["lambda_r"], window_len=10), "r--", label="smoothed")
new_ls = l_at_smoothed_r(gal, npix_per_reff=5)
ax[1].plot(smooth(new_ls, window_len=10), "g--", label="smoothed_new")
#ax[2].plot(new_ls, label="org")
plt.legend()
pdf.savefig()
plt.close()
gal.idxs[0]
gal.cluster = 1234
```
| github_jupyter |
# description 6/03/19 clinically guided aggregation modeling w/ 2 elix variables
sklearn modeling using local methods of the median imputed training data using origional min/max clinically guided aggregation. note the preprocessing of data from 07.20-worst_case_model was performed in R (09.newagg2_preprocessing_med_impute.rmd). this eventually will be converted over to python, but for now works in r.
preprocessing includes variable formatting (categorical to factor variables in r, train/test split, and median imputation).
```
import pandas as pd
import matplotlib.pyplot as plt
import os, sys
from pathlib import Path
import seaborn as sns
import numpy as np
import glob
from sklearn.metrics import confusion_matrix, classification_report, roc_curve, roc_auc_score, accuracy_score, auc, precision_recall_fscore_support, pairwise, f1_score, log_loss, make_scorer
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.externals.joblib import Memory
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder, Imputer
from sklearn.model_selection import StratifiedKFold, GridSearchCV, RandomizedSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.utils import validation
from scipy.sparse import issparse
from scipy.spatial import distance
from sklearn import svm
#importin xg boost and all needed otherstuff
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from xgboost import XGBClassifier #conda install -c conda-forge xgboost to install
##adding these, lets see if it helps with xgboost crash
os.environ['KMP_DUPLICATE_LIB_OK']='True'
#reducing warnings that are super common in my model
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.simplefilter(action='ignore') #ignore all warnings
memory = Memory(cachedir='/tmp', verbose=0)
#@memory.cache above any def fxn.
RANDOM_STATE = 15485867
%matplotlib inline
plt.style.use('ggplot')
from notebook.services.config import ConfigManager
cm = ConfigManager()
cm.update('livereveal', {
'width': 1024,
'height': 768,
'scroll': True,
})
%load_ext autotime
```
# importing and formatting dataset
```
# # ##24 hr sensitivity
# # #importing in all clinical_variable files
# lower_window=0
# upper_window=1
# time_col="charttime"
# time_var="t_0"
# folder="{}_hr_window".format(timewindowdays)
# timewindowdays="24"
# date= '09062019'
# patient_df= final_pt_df2
# #48 hr sensitivity
# lower_window=0
# upper_window=2
# time_var="t_0"
# timewindowdays="48"
# folder="{}_hr_window".format(timewindowdays)
# date='16052019'
# time_col="charttime"
# time_var= 't_0'
# patient_df= final_pt_df2
#72 hr elixhauser-redo
date='11062019'
lower_window=0
upper_window=3
timewindowdays="72"
folder="{}_hr_window".format(timewindowdays)
time_col="charttime"
time_var= 't_0'
patient_df= final_pt_df2
#cohort import
os.chdir('/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling') #use to change working directory
wd= os.getcwd() #'/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling'
final_pt_df2 = pd.read_csv(Path(wd + '/data/raw/csv/04042019_final_pt_df2_v.csv') , index_col=0) #only for patients with minimum vitals, 14478 icustay_id
patients= list(final_pt_df2['subject_id'].unique())
hadm_id= list(final_pt_df2['hadm_id'].unique())
icustay_id= list(final_pt_df2['icustay_id'].unique())
icustay_id= [int(x) for x in icustay_id]
# #importing in all clinical_variable files
# os.chdir(r'/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling/data/processed/')
# train_data= pd.read_csv(Path(wd+'/data/processed/merged/{}_worst_df_train_preImp_{}.csv'.format(date,timewindowdays), index_col=0))
# test_data= pd.read_csv(Path(wd+'/data/processed/merged/{}_worst_df_test_preImp_{}.csv'.format(date,timewindowdays), index_col=0))
#importing in all clinical_variable files
os.chdir(r'/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling/data/processed/')
train_data= pd.read_csv(Path(wd+'/data/processed/merged/{}_worst_df_train_preImp_{}_2.csv'.format(date,timewindowdays), index_col=0))
test_data= pd.read_csv(Path(wd+'/data/processed/merged/{}_worst_df_test_preImp_{}_2.csv'.format(date,timewindowdays), index_col=0))
list(train_data)
vaso_active=['phenylephrine',
'norepinephrine',
'vasopressin',
'dobutamine',
'dopamine',
'epinephrine']
ordinal=[
'leukocyte',
'nitrite',
'vent_recieved',
'o2_flow',
'rrt',
'pao2fio2ratio',
'cancer_elix',
"any_vasoactives",
'bands',
'pco2'
]
categorical=[
"ethnicity",
'gender'
]
### 06/18/19 changed this to be consistent with the 07.20 model_aggregation notebook
categorical_variables= vaso_active+ordinal+categorical
def preprocessing(preimp_df):
"""
1) rename columns
2) standardize last 2 columns to be standardized
3) convert categorical columns to proper format
4) median impute
"""
from sklearn.impute import SimpleImputer
global categorical_variables
rename_dic={
"('max', 'sodium')": "maxSodium" ,
"('max', 'sodium')" : "maxSodium",
"('min', 'sodium')" : "minSodium",
"('max', 'calcium')" : "maxCalcium",
"('min', 'calcium')" : "minCalcium",
"('max', 'sodium')": "maxSodium",
"('min', 'sodium')": "minSodium",
"('max', 'wbc')": "maxWBC",
"('min', 'wbc')": "minWBC",
"any_vasoactive": "any_vasoactives"
}
data=preimp_df.copy()
data=data.rename(rename_dic, axis='columns').copy()
# ##changing the deitentified ages to 90.
# data.loc[data['first_admit_age']>90,"first_admit_age"]=90
# weight_median=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).median()
# weight_quant1=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).quantile(0.25)#.between(train_data['col'].quantile(.25), df['col'].quantile(.75), inclusive=True)]
# weight_quant3=np.log(data.loc[data['final_bin']=="C_neg/A_partial","weight"]+1).quantile(0.75)
# weight_iqr=weight_quant3-weight_quant1
# #print(weight_median,weight_quant3,weight_quant1, weight_iqr)
# age_median=np.log((data.loc[data['final_bin']=="C_neg/A_partial","first_admit_age"].median()+1))
# # age_quant1=np.log((data.loc[data['final_bin']=="C_neg/A_partial","first_admit_age"]+1).quantile(0.25))
# # age_quant3=np.log((data.loc[data['final_bin']=="C_neg/A_partial","first_admit_age"]+1).quantile(0.75))
# age_quant1=(data.loc[data['final_bin']=="C_neg/A_partial","first_admit_age"]).quantile(0.25)
# age_quant3=(data.loc[data['final_bin']=="C_neg/A_partial","first_admit_age"]).quantile(0.75)
# age_iqr=np.log((age_quant3-age_quant1)+1)
# print(age_median,age_quant1,age_quant3, age_iqr)
# #converting to log scaled standardized data for age/weight
# data['weight']=data['weight'].apply(lambda x: (np.log(x+1)-weight_median)/weight_iqr)
# data['first_admit_age']=data['first_admit_age'].apply(lambda x: (np.log(x+1)-age_median)/age_iqr)
### onehot encoding categorical var
cols_to_transform= categorical_variables
data = pd.get_dummies(data, columns = cols_to_transform, drop_first=True)
#binarizing and poping outcome for training data
data.loc[data['final_bin']=="C_pos/A_full","final_bin"]=1
data.loc[data['final_bin']=="C_neg/A_partial","final_bin"]=0
data['final_bin']=pd.to_numeric(data['final_bin'])
imp = SimpleImputer(missing_values=np.nan, strategy='median')
data=pd.DataFrame(imp.fit_transform(data), columns=list(data))
## establishing training data and labels
x_train= data.copy()
z_icustay_id=x_train.pop('icustay_id')
y_train= x_train.pop("final_bin").values
return(x_train, y_train, z_icustay_id)
# x_train, y_train, z_icustay_id= preprocessing(pd.merge(preimp_train_df, final_pt_df2[['icustay_id','final_bin']]))
def preprocessing(preimp_df):
"""
1) rename columns
2) standardize last 2 columns to be standardized
3) convert categorical columns to proper format
4) median impute
"""
from sklearn.impute import SimpleImputer
global categorical_variables
rename_dic={
"('max', 'sodium')": "maxSodium" ,
"('max', 'sodium')" : "maxSodium",
"('min', 'sodium')" : "minSodium",
"('max', 'calcium')" : "maxCalcium",
"('min', 'calcium')" : "minCalcium",
"('max', 'sodium')": "maxSodium",
"('min', 'sodium')": "minSodium",
"('max', 'wbc')": "maxWBC",
"('min', 'wbc')": "minWBC",
"any_vasoactive": "any_vasoactives"
}
data=preimp_df.copy()
data=data.rename(rename_dic, axis='columns').copy()
#binarizing and poping outcome for training data
data.loc[data['final_bin']=="C_pos/A_full","final_bin"]=1
data.loc[data['final_bin']=="C_neg/A_partial","final_bin"]=0
data['final_bin']=pd.to_numeric(data['final_bin'])
return(data)
# x_train, y_train, z_icustay_id= preprocessing(pd.merge(preimp_train_df, final_pt_df2[['icustay_id','final_bin']]))
train_data=preprocessing(pd.merge(train_data, final_pt_df2[['icustay_id','final_bin']]))
# ### onehot encoding categorical var
# cols_to_transform= categorical_variables
# data = pd.get_dummies(data, columns = cols_to_transform, drop_first=True)
# #binarizing and poping outcome for training data
# data.loc[data['final_bin']=="C_pos/A_full","final_bin"]=1
# data.loc[data['final_bin']=="C_neg/A_partial","final_bin"]=0
# data['final_bin']=pd.to_numeric(data['final_bin'])
# imp = SimpleImputer(missing_values=np.nan, strategy='median')
# data=pd.DataFrame(imp.fit_transform(data), columns=list(data))
# ## establishing training data and labels
# x_train= data.copy()
# z_icustay_id=x_train.pop('icustay_id')
# y_train= x_train.pop("final_bin").values
# return(x_train, y_train, z_icustay_id)
# x_train, y_train, z_icustay_id = preprocessing(pd.merge(train_data, final_pt_df2[['icustay_id','final_bin']]))
# x_test, y_test, z_icustay_id_test= preprocessing(pd.merge(test_data, final_pt_df2[['icustay_id','final_bin']]))
# x_train, y_train, z_icustay_id = preprocessing(pd.merge(train_data, final_pt_df2[['icustay_id','final_bin']]))
# x_test, y_test, z_icustay_id_test= preprocessing(pd.merge(test_data, final_pt_df2[['icustay_id','final_bin']]))
# #for local modeling
# # all_xy=xy_preprocessing(train_data)
# # all_xy_test=xy_preprocessing(test_data)
train_data
x_train['first_admit_age'].describe()
```
# experimenting with pipelines start
```
train_data['first_admit_age'].describe()
### in a sklearn pipeline each step just has a .transform applied to it.
### except the last step which has .fit applied
###here we are just making a factorextractor class that tells what to do when transform is applied to it.
###
from sklearn.base import TransformerMixin, BaseEstimator
class FactorExtractor(TransformerMixin, BaseEstimator):
def __init__(self, factor):
self.factor = factor
def transform(self, df):
return df[self.factor]
def fit(self, *_):
return self
#demonstrates this is the same thing:
(FactorExtractor("final_bin").transform(train_data) == train_data['final_bin']).all()
##estimator= anyclass in sklearn with a .fit and a .predict. this is what i will use. ie a rf classifier.
FactorExtractor("final_bin").transform(train_data)
class ConvertToDataFrame:
def transform(self, data):
df=pd.DataFrame(data)
self.columns= df.columns
return df
#return [{self.factor: self.normalize(tt)} for tt in data[self.factor]]
def fit(self, *_):
return self
# class DateImputer(TransformerMixin):
# def transform(self, df):
# # Observations don't always exist. Hence impute by lower/upper bound.
# df = df.fillna({'date_first_observation': df['date_creation'],
# 'date_last_observation': df['date_deletion']})
# return df
# def fit(self, df, labels=None):
# return self
class onehot(TransformerMixin):
def __init__(self, cols_to_transform):
self.cols_to_transform=cols_to_transform
def transform(self,df ):
data = pd.get_dummies(df, columns = self.cols_to_transform, drop_first=True)
return(data)
def fit(self, *_):
return self
pd.get_dummies(x_train, columns = cols_to_transform, drop_first=True)
##sklearn example https://scikit-learn.org/stable/auto_examples/compose/plot_column_transformer_mixed_types.html#sphx-glr-auto-examples-compose-plot-column-transformer-mixed-types-py
# We will train our classifier with the following features:
# Numeric Features:
# - age: float.
# - fare: float.
# Categorical Features:
# - embarked: categories encoded as strings {'C', 'S', 'Q'}.
# - sex: categories encoded as strings {'female', 'male'}.
# - pclass: ordinal integers {1, 2, 3}.
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_features = ['age', 'fare']
numeric_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler())])
categorical_features = ['embarked', 'sex', 'pclass']
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='constant', fill_value='missing')),
('onehot', OneHotEncoder(handle_unknown='ignore'))])
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
clf = Pipeline(steps=[('preprocessor', preprocessor),
('classifier', LogisticRegression(solver='lbfgs'))])
X = data.drop('survived', axis=1)
y = data['survived']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
clf.fit(X_train, y_train)
print("model score: %.3f" % clf.score(X_test, y_test))
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
pipeline=make_pipeline(onehot(categorical_variables),
SimpleImputer(missing_values=np.nan, strategy='median'),
#FactorExtractor('final_bin'),
ConvertToDataFrame(),
RandomForestClassifier(n_estimators=100, random_state=2065))
#binarizing and poping outcome for training data
train_data.loc[train_data['final_bin']=="C_pos/A_full","final_bin"]=1
train_data.loc[train_data['final_bin']=="C_neg/A_partial","final_bin"]=0
train_data['final_bin']=pd.to_numeric(train_data['final_bin'])
## establishing training data and labels
x_train= train_data.drop(columns=["final_bin",'icustay_id'])
z_icustay_id=train_data["icustay_id"].copy()
y_train= train_data["final_bin"].copy()
pipeline.fit(x_train, y_train)
pipeline.predict_proba(x_train)
x_train.columns.to_list()
# def var_imp(model,folder_name,model_name, n_var=4, save=True):
# model_name=type(model).__name__
# plot_title= "Top {} {} {} Variable Importance".format(n_var, folder_name,model_name)
# feat_importances = pd.Series(model.pipeline.predict_proba(x_train), index=x_train.columns)
# topn=feat_importances.nlargest(n_var).sort_values()
# ax=topn.plot(kind='barh', x='doop', title=plot_title)#.xlabel("xlab")
# ax.set_xlabel("Variable Importance")
# if save==True:
# saveplot(figure_name=plot_title, folder_name=folder_name)
# return(topn)
pd.Series(pipeline.predict_proba(x_train)[:,:], index=x_train.columns)
pipeline.named_steps['converttodataframe'].columns()#.shape
#x_train.columns
pipeline.steps#[-1][1].feature_importances_
pd.Series(pipeline.named_steps['randomforestclassifier'].feature_importances_, index=x_train.columns)
#pipeline.named_steps['randomforestclassifier'].feature_importances_
x_train
x_train
```
# experimenting with pipelines end
```
train_data.head()
```
#### optional qc
```
#x_train.iloc[1:5, 25:45]
#x_train.iloc[1:5, 35:65]
#x_train.iloc[1:5, 10:30]
```
## looking at correlation of all variables
```
corr = x_train.corr().abs()
plt.figure(figsize=(25, 20))
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
);
sol = (corr.where(np.triu(np.ones(corr.shape), k=1).astype(np.bool)).stack().sort_values(ascending=False))
cor_df=pd.DataFrame(sol)#.sort_values(kind="quicksort") #[-10:0])
cor_df=cor_df.reset_index()
cor_df=cor_df.rename(columns={'level_0': 'corx', 'level_1': 'cory', 0:'corr'})
cor_df2=cor_df[(cor_df['corx']!=cor_df['cory']) & (cor_df['corr']>0.7)].sort_values('corr', ascending=False)
cor_df2.head()
```
### DROPING one of the 2 columns with correlation >0.7
corx cory corr
0 ipco2_absent pao2fio2Ratio_(475, 3000] 0.872418
1 maxWBC minWBC 0.802373
2 bun creatinine 0.720861
3 maxSodium minSodium 0.704233
```
x_train.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
x_test.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
# all_xy.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
# all_xy_test.drop(columns=list(cor_df2['corx']), inplace=True, errors='raise')
```
### formatting x and y for modleing
```
#x=np.array(x_train.iloc[:,[1,2,3,4,5,6,7,8,9,38,39,40,41]]).copy() #copy of x_train
x=np.array(x_train.copy())
#x=np.array(train_data.iloc[:,[1,2,3,4]]).copy() #copy of x_train
#train_data.iloc[:,[1,2,3,4,5]] ###drastically reducing my dataframe size to test algorithm
y=y_train.copy() #copy of y_train
y=y.astype('int')
##all_xy: train data with finalbin:label and index=icustay_id
#all_xy=train_data.copy().set_index("icustay_id").rename(columns={'final_bin':"label"}) #
time_interval=4
print(len(x_train),len(x_test))
```
# Modelbuilding
* step1) hypertune xgb on 5fold cv.
* step2) test entire trainset and predict trainset.
* step3) run hypertuned model on 5fold cv with lr and get overall metrics.
* step4) local model testing
## step1) XGB hypertuning
```
def evaluate(model, test_features, test_labels):
from sklearn.metrics import log_loss
y_hat = model.predict(test_features)
errors = abs(y_hat - test_labels)
mape = 100 * np.mean(errors / test_labels)
accuracy = 100 - mape
auc=roc_auc_score(test_labels, y_hat)
loss= log_loss(test_labels, y_hat)
print ('the AUC is: {:0.2f}'.format(auc))
print ('the logloss is: {:0.2f}'.format(loss))
print(confusion_matrix(test_labels, y_hat))
return loss
def hypertuning_fxn(X, y, nfolds, model , param_grid, base_model, scoring="neg_log_loss", gridsearch=True, n_iter=20):
if gridsearch==True:
grid_search = GridSearchCV(estimator= model,
param_grid=param_grid,
cv=nfolds,
scoring=scoring,
return_train_score=True,
n_jobs = -1)
else:
grid_search = RandomizedSearchCV(estimator= model,
param_distributions= param_grid,
n_iter=n_iter,
cv=nfolds,
scoring=scoring,
return_train_score=True,
n_jobs = -1)
grid_search.fit(X, y)
print("Grid scores on development set:")
means = grid_search.cv_results_['mean_test_score']
stds = grid_search.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, grid_search.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
#grid_search.best_params_
print(grid_search.best_score_)
print("\n")
print(grid_search.best_params_)
print('\n base model:')
base_model = base_model#(random_state = 42)
base_model.fit(x, y)
base_auc = evaluate(base_model, x, y)
print('\n hypertuned model:')
best_random = grid_search.best_estimator_
random_auc = evaluate(best_random, x, y)
print('logloss change of {:0.2f}%. after hypertuning on training set (may be overfit)'.format( 100 * (random_auc - base_auc) / base_auc))
print(grid_search.best_estimator_)
return(grid_search)
###xgboost
model= XGBClassifier(n_estimators=100, min_child_weight=2, #changed: GridSearchCV ->RandomizedSearchCV
gamma=0, subsample=0.8, colsample_bytree=0.8,
objective='binary:logistic', n_jobs=-1, seed=27)
scale_pos_weight = [0.1, 1, 5, 10]
max_depth = [1, 2, 3, 4, 5]
learning_rate=[0.01, 0.1, 0.5, 1]
param_grid = {'scale_pos_weight': scale_pos_weight, 'max_depth' : max_depth, "learning_rate":learning_rate}
base_model=XGBClassifier(random_state = 42)
xgboost_hyper=hypertuning_fxn(x, y, nfolds=5, model=model , param_grid=param_grid, base_model= base_model, scoring="neg_log_loss", n_iter=20, gridsearch=True)
###rf
# Number of trees in random forest
#n_estimators = [100, 1000]#[int(x) for x in np.linspace(start = 10, stop = 1000, num = 5)]
# Number of features to consider at every split
max_features = [3,'auto', 10 ]
# Maximum number of levels in tree
max_depth = [5,10, 25]#[int(x) for x in np.linspace(5, 110, num = 5)]
#max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [2, 5, 10]
# Method of selecting samples for training each tree
#bootstrap = [True, False]
#class_weight is either a dictionary of each class to a uniform weight for that class (e.g., {1:.9, 2:.5, 3:.01}), or is a string telling sklearn how to automatically determine this dictionary.
class_weight= [None,{0:1, 1:4}, {0:(1/np.bincount(y))[0], 1:(1/np.bincount(y))[1]}]
param_grid = {#'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'class_weight': class_weight}
model= RandomForestClassifier(criterion='entropy')
base_model=RandomForestClassifier(random_state = 42, criterion='entropy')
rf_hyper=hypertuning_fxn(x, y, nfolds=10, model=model , param_grid=param_grid, base_model= base_model, scoring="neg_log_loss",n_iter = 30, gridsearch=True)
```
## Hypertune SVC
```
model= svm.SVC(probability=True)
kernel = ['linear', 'rbf', 'poly']
gamma = [0.1, 1, 'auto'] #Kernel coefficient for ‘rbf’, ‘poly’ and ‘sigmoid’. default=’auto’ uses 1 / n_features
C = [0.1, 1, 10, 100] #Penalty parameter C of the error term.
degree = [0, 1, 2]
class_weight=['balanced', None]
param_grid = {'kernel': kernel,
'gamma': gamma,
'C': C,
'degree': degree,
'class_weight':class_weight}
base_model=svm.SVC(probability=True)
svc_hyper=hypertuning_fxn(x, y, nfolds=4, model=model , param_grid=param_grid, base_model= base_model, scoring="neg_log_loss", n_iter=10, gridsearch=False)
###knn
from sklearn.neighbors import KNeighborsClassifier
model= KNeighborsClassifier()
n_neighbors = [3,4,5, 8, 10, 25]
weights=['uniform']
p=[1,2] #1= mmanhattan, 2= euclidian
param_grid = {'n_neighbors': n_neighbors,
'weights': weights,
'p': p}
base_model=KNeighborsClassifier()
knn_hyper=hypertuning_fxn(x, y, nfolds=10, model=model , param_grid=param_grid, base_model= base_model, scoring="neg_log_loss", n_iter=40, gridsearch=True)
knn_hyper.best_params_
```
# Hypertuned Model Initialization
```
def reset_model(model_name, hardcode=True):
if hardcode==True:
if model_name== 'xgboost':
model = XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bytree=0.8, gamma=0, learning_rate=0.1, max_delta_step=0,
max_depth=4, min_child_weight=2, missing=None, n_estimators=100,
n_jobs=-1, nthread=None, objective='binary:logistic',
random_state=0, reg_alpha=0, reg_lambda=1, scale_pos_weight=1,
seed=27, silent=True, subsample=0.8)
elif model_name== 'logreg':
model = LogisticRegression(penalty='l2', dual=False, tol=0.0001, C=10, fit_intercept=True,
intercept_scaling=1, class_weight='balanced', random_state=None)
elif model_name== 'rf':
model = RandomForestClassifier(bootstrap=False, class_weight={0: 1, 1: 4},
criterion='entropy', max_depth=10, max_features='auto',
max_leaf_nodes=None, min_impurity_decrease=0.0,
min_impurity_split=None, min_samples_leaf=2,
min_samples_split=2, min_weight_fraction_leaf=0.0,
n_estimators=600, n_jobs=None, oob_score=False,
random_state=None, verbose=0, warm_start=False)
elif model_name== 'svc':
model = svm.SVC(C=100, cache_size=200, class_weight='balanced', coef0=0.0,
decision_function_shape='ovr', degree=0, gamma=1, kernel='linear',
max_iter=-1, probability=True, random_state=None, shrinking=True,
tol=0.001, verbose=False)
elif model_name== 'knn':
model = KNeighborsClassifier(algorithm='auto', leaf_size=30, metric='minkowski',
metric_params=None, n_jobs=None, n_neighbors=25, p=1,
weights='uniform')
else:
if model_name== 'xgboost':
model = xgboost_hyper.best_estimator_
elif model_name== 'logreg':
model = LogisticRegression(penalty='l2', dual=False, tol=0.0001, C=10, fit_intercept=True,
intercept_scaling=1, class_weight='balanced', random_state=None)
elif model_name== 'rf':
model = rf_hyper.best_estimator_
elif model_name== 'svc':
model = svc_hyper.best_estimator_
elif model_name== 'knn':
model = knn_hyper.best_estimator_
return(model)
```
# test entire trainset and predict testset.
*<del> step1) hypertune xgb on 5fold cv.
* step2) test entire trainset and predict testset.
* step3) local model testing
```
def get_auc_score(model,train_index, x=x,y=y):
y_pred_proba = model.predict_proba(x[train_index])[:, 1]
roc_score=roc_auc_score(y[train_index], y_pred_proba)
return(roc_score)
```
## youden index and plotting functions
```
def optimal_youden_index(fpr, tpr, thresholds, tp90=True):
"""
inputs fpr, tpr, thresholds from metrics.roc(),
outputs the clasification threshold, roc dataframe, and the index of roc dataframe for optimal youden index
"""
#making dataframe out of the thresholds
roc_df= pd.DataFrame({"thresholds": thresholds,"fpr":fpr, "tpr": tpr})
roc_df.iloc[0,0] =1
roc_df['yuden']= roc_df['tpr']-roc_df['fpr']
if tp90==True:
idx= roc_df[roc_df['tpr']>=0.9]['yuden'].idxmax() #changed this so now finds optimial yuden threshold but tp>=90%
else:
idx=roc_df['yuden'].idxmax() #MAX INDEX
youden_threshold=roc_df.iloc[idx,0] #threshold for max youden
return(youden_threshold, roc_df, idx)
def plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=False,model_name=None, folder_name=None, file_name=None):
plt.title('ROC with optimal Youden Index')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
#finding the point on the line given threshold 0.5 (finding closest row in roc_df)
og_idx=roc_df.iloc[(roc_df['thresholds']-0.5).abs().argsort()[:1]].index[0]
plt.plot(roc_df.iloc[og_idx,1], roc_df.iloc[og_idx,2],marker='x', markersize=5, color="g")
plt.annotate(s="P(>=0.5)",xy=(roc_df.iloc[og_idx,1]+0.02, roc_df.iloc[og_idx,2]-0.04),color='g') #textcoords
plt.plot(roc_df.iloc[idx,1], roc_df.iloc[idx,2],marker='o', markersize=5, color="r") ##
plt.annotate(s="M_Youden",xy=(roc_df.iloc[idx,1]+0.02, roc_df.iloc[idx,2]-0.04),color='r' ) #textcoords
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
if save==True:
if folder_name != None:
address = 'figures/{}/'.format(folder_name)
else:
address = 'figures/'
if not os.path.exists(address):
os.makedirs(address)
plt.savefig(address+"/{}_{}.png".format(model_name,file_name),bbox_inches='tight')
else: pass
plt.show()
def plot_table_as_fig(table_in, col_labels, row_labels, save=False,model_name=None,folder_name=None, file_name=None,figsize=(6,1)):
fig = plt.figure(figsize=figsize)
table = plt.table(cellText = table_in,
colLabels = col_labels,
rowLabels = row_labels,
loc='best')
plt.axis("tight")
plt.axis('off')
if save==True:
if folder_name != None:
address = 'figures/{}/'.format(folder_name)
else:
address = 'figures/'
if not os.path.exists(address):
os.makedirs(address)
plt.savefig(address+"/{}_{}.png".format(model_name,file_name),bbox_inches='tight')
else: pass
plt.show()
def classifier_eval(model, x=x, y=y, proba_input=False,pos_label=1, print_default=True, save=False,model_name=None, folder_name=None):
import sklearn.metrics as metrics
from sklearn.metrics import precision_score, roc_auc_score, f1_score, recall_score
"""
catchall classification evaluation function. will print/save the following:
print/save the following:
ROC curve marked with threshold for optimal youden (maximizing tpr+fpr with constraint that tpr>0.9)
using 0.5 threshold:
confusion matrix
classification report
npv
accuracy
using optimal youden (maximizing tpr+fpr with constraint that tpr>0.9):
confusion matrix
classification report
npv
accuracy
output:
outputs modelname, auc, precision, recall, f1, and npv to a dictionary.
notes:
youden's J statistic:
J= sensitivity + specificity -1
(truepos/ truepos+falseneg) + (true neg/ trueneg + falsepos) -1
"""
if save==True: #making folder if one doesn't exist
if folder_name != None:
address = '../figures/{}/'.format(folder_name)
else:
address = 'train/'
if not os.path.exists(address):
os.makedirs(address)
if proba_input==True: #incorporating classifier_eval2() functionality into this (ie allowing user to input a y_proba instead of a model)
y_proba= model
y_pred=[1 if y >= 0.5 else 0 for y in y_proba]
else:
model_name=type(model).__name__
y_pred = model.predict(x)
y_proba = model.predict_proba(x)[:,1]
fpr, tpr, thresholds = metrics.roc_curve(y, y_proba, pos_label=pos_label)
roc_auc = metrics.auc(fpr, tpr)
#gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds,tp90=True)
#plotting roc
plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=save, model_name=model_name,folder_name=folder_name, file_name='roc')
plt.show(), plt.close()
#printing npv, recall, precision, accuracy
npv=confusion_matrix(y, y_pred)[0,0]/sum(np.array(y_pred)==0)*100
prec= precision_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
recall= recall_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
f1= f1_score(y_true=y, y_pred= y_pred, pos_label=pos_label)
if print_default==True: ###can opt to not print the 0.5 classification threshold classification report/conf matrix
#plotting confusion matrixs
print("\n******* Using 0.5 Classification Threshold *******\n")
print(confusion_matrix(y, y_pred))
print ('the Accuracy is: {:01.2f}'.format(accuracy_score(y, y_pred)))
print ("npv: {:01.2f}".format(npv))
print ('the classification_report:\n', classification_report(y,y_pred))
else:
pass
#### YOUDEN ADJUSTMENT #####
print("\n******* Using Optimal Youden Classification Threshold *******\n")
print("\nthe Youden optimal index is : {:01.2f}".format(youden_threshold))
y_pred_youden = [1 if y >= youden_threshold else 0 for y in y_proba]
npv_y=confusion_matrix(y, y_pred_youden)[0,0]/sum(np.array(y_pred_youden)==0)*100
prec_y= precision_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)*100
recall_y= recall_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)*100
f1_y= f1_score(y_true=y, y_pred= y_pred_youden, pos_label=pos_label)*100
auc_y=roc_auc_score(y_true=y, y_score= y_proba)*100
##plotting and saving confusion matrix
confusion_youden=confusion_matrix(y, y_pred_youden)
plot_table_as_fig(confusion_youden,
col_labels=['predicted_neg','predicted_pos'],
row_labels=['true_neg',"true_pos"],
save=save,
figsize=(6,1),
model_name=model_name,
folder_name=folder_name,
file_name='y_confusion')
plt.show(), plt.close()
#print(confusion_matrix(y, y_pred_youden))
print("the Accuracy is: {:01.2f}".format(accuracy_score(y, y_pred_youden)))
print("npv: {:01.2f}".format(npv_y))
###formatting classification report to be compatable with matplotlib table
report_youden=classification_report(y,y_pred_youden,output_dict=True)
report_youden = pd.DataFrame.from_dict(report_youden).transpose()[['precision','recall','f1-score','support']]
report_youden = np.round(report_youden,2)
##plotting and saving classification report
plot_table_as_fig(table_in=np.array(report_youden),#classification_report(y, xgboost.predict(x))),
col_labels=['precision','recall','f1-score','support'],
row_labels=['neg',"pos","micro_avg","macro_avg",'weighted_avg'],
figsize=(15,5),
save=save,
model_name=model_name,
folder_name=folder_name,
file_name='y_report')
plt.show(), plt.close()
youden_dic= {'model':model_name, 'auc':auc_y, 'precision':prec_y, 'recall':recall_y, 'f1':f1_y, 'npv':npv_y}
return(youden_dic)
### label youden index
```
# testing global model
## test entire trainset and predict trainset.
<del> * step1) hypertune xgb on 5fold cv.
* step2) test entire train set and predict testset.
* step3) local model testing
thresholds:
* Decreasing thresholds on the decision function used to compute
fpr and tpr. `thresholds[0]` represents no instances being predicted
and is arbitrarily set to `max(y_score) + 1`.
```
#setting up test table
test_summary_df= pd.DataFrame({'model':[],'auc':[], 'precision':[], 'recall':[], 'f1':[], 'npv':[]})
test_summary_df
```
### model fitting
```
xgboost = reset_model('xgboost', hardcode=False)
xgboost.fit(x, y)
logreg = reset_model('logreg', hardcode=False)
logreg.fit(x, y)
rf= reset_model('rf', hardcode=False)
rf.fit(x,y)
# from sklearn.naive_bayes import GaussianNB
# gnb =GaussianNB()
# nb_y_pred = gnb.fit(x, y)
from sklearn import svm
svc= reset_model('svc', hardcode=False)
svc.fit(x, y)
knn= reset_model('knn', hardcode=False)
knn.fit(x,y)
```
### global model test set evaluation
```
svc_eval= classifier_eval(svc, x=np.array(x_test), y=y_test, save=True, model_name='svc', folder_name='clinical_agg_elix72')
xgboost_eval= classifier_eval(xgboost, x=np.array(x_test), y=y_test, save=True, model_name='xgboost', folder_name='clinical_agg_elix72')
rf_eval= classifier_eval(rf, x=np.array(x_test), y=y_test, save=True, model_name='rf', folder_name='clinical_agg_elix72')
# logreg_eval= classifier_eval(logreg, x=np.array(x_test), y=y_test)
logreg_eval= classifier_eval(logreg, x=np.array(x_test), y=y_test, save=True, model_name='logreg', folder_name='clinical_agg_elix72')
knn_eval= classifier_eval(knn, x=np.array(x_test), y=y_test, save=True, model_name='knn', folder_name='clinical_agg_elix72')
```
## Ensemble of all models
```
from sklearn.ensemble import VotingClassifier
#create a dictionary of our models
estimators=[("xgboost", xgboost), ('rf', rf), ('log_reg', logreg), ('svc',svc)]
#create our voting classifier, inputting our models
ensemble = VotingClassifier(estimators, voting='soft', n_jobs=-1)
# If ‘hard’, uses predicted class labels for majority rule voting.
# Else if ‘soft’, predicts the class label based on the argmax of the sums of the predicted probabilities,
# which is recommended for an ensemble of well-calibrated classifiers.
#weights: array-like, shape (n_classifiers,), optional (default=`None`)
#Sequence of weights (float or int) to weight the occurrences of predicted class labels (hard voting) or class probabilities before averaging (soft voting).
#Uses uniform weights if None.
ensemble.fit(x, y)#, sample_weight=np.array([0.67289604, 1.94595562]))
ensemble2 = VotingClassifier(estimators, voting='hard', n_jobs=-1)
ensemble2.fit(x, y)
y_pred = ensemble2.predict(np.array(x_test))
#y_proba = ensemble2.predict_proba(np.array(x_test))[:,1]
from sklearn import metrics
from sklearn.metrics import precision_score, roc_auc_score, f1_score, recall_score
pos_label=1
fpr, tpr, thresholds = metrics.roc_curve(y_test, y_pred, pos_label=pos_label)
roc_auc = metrics.auc(fpr, tpr)
#gathering the optimal youden_index and df of tpr/fpr for auc and index of that optimal youden. idx is needed in the roc
youden_threshold, roc_df, idx= optimal_youden_index(fpr, tpr, thresholds,tp90=True)
#plotting roc
plot_roc(fpr, tpr, roc_auc, roc_df, idx, save=False, model_name=ensemble2,folder_name=None, file_name='roc')
plt.show(), plt.close()
#printing npv, recall, precision, accuracy
npv=confusion_matrix(y_test, y_pred)[0,0]/sum(np.array(y_pred)==0)*100
prec= precision_score(y_true=y_test, y_pred= y_pred, pos_label=pos_label)*100
recall= recall_score(y_true=y_test, y_pred= y_pred, pos_label=pos_label)*100
f1= f1_score(y_true=y_test, y_pred= y_pred, pos_label=pos_label)*100
acc=accuracy_score(y_test, y_pred)*100
print("npv:", npv,'\n')
print("prec:", prec,'\n')
print("recall:", recall,'\n')
print("f1:", f1,'\n')
print("acc:", acc,'\n')
hard_vote_summary={'model':"hard_voting_classifier",'auc':acc, 'precision':prec, 'recall':recall, 'f1':f1, 'npv':npv}
ensemble_eval= classifier_eval(ensemble, x=np.array(x_test), y=y_test, save=True, model_name='model_ensemble', folder_name='clinical_agg_elix72')
np.unique(y)
from sklearn.utils.class_weight import compute_class_weight
compute_class_weight('balanced', np.unique(y), y)
#classifier_eval(gnb, x=x_test, y=y_test)
test_summary_df= pd.DataFrame([rf_eval,
logreg_eval,
xgboost_eval,
svc_eval,
ensemble_eval])
test_summary_df.set_index('model').round(decimals=2)
```
# variable importance
```
def saveplot(figure_name,folder_name=None):
"""
simple function for saving plots
"""
if folder_name != None:
address = 'figures/{}/'.format(folder_name)
else:
address = 'figures/'
if not os.path.exists(address):
os.makedirs(address)
plt.savefig(address+"/{}.png".format(figure_name),bbox_inches='tight')
def var_imp(model,folder_name,model_name, n_var=4, save=True):
model_name=type(model).__name__
plot_title= "Top {} {} {} Variable Importance".format(n_var, folder_name,model_name)
feat_importances = pd.Series(model.pipeline.predict_proba(x_train), index=x_train.columns)
topn=feat_importances.nlargest(n_var).sort_values()
ax=topn.plot(kind='barh', x='doop', title=plot_title)#.xlabel("xlab")
ax.set_xlabel("Variable Importance")
if save==True:
saveplot(figure_name=plot_title, folder_name=folder_name)
return(topn)
# ###
# imp= model.feature_importances_
# var_index=[ x for x in range(0,len(rf.feature_importances_))]
# variables=list(x_train)
# return(pd.DataFrame({"imp":imp, 'index':var_index, 'variable': variables}).sort_values('imp', ascending=False))
#var_imp(rf,plot_title='RF_' n_var=4)
#var_imp(rf,"clinical_agg","RF", n_var=6, save=False)
var_imp(rf,"clinical_agg_elix","RF", n_var=20, save=True)
var_imp(xgboost2,"clinical_agg","xgboost", n_var=10, save=False)
var_imp(xgboost,"clinical_agg_elix_72","xgboost", n_var=10, save=True)
test_summary_df['model']
print(len(x_train),len(x_test))
len(x)
```
# model pickling
```
import pickle
from sklearn import model_selection
# save the model to disk
model_list=[rf,logreg, xgboost,svc ]
for element in model_list:#test_summary_df['model']:
filename = 'models/{}_{}_{}.sav'.format(date,dataset,type(element).__name__)
#os.makedirs(os.path.dirname(filename), exist_ok=True)
pickle.dump(element, open(filename, 'wb'))
# xgboost = reset_model('xgboost')
# xgboost.fit(x, y)
# logreg = reset_model('logreg')
# logreg.fit(x, y)
# rf= reset_model('rf')
# rf.fit(x,y)
# # from sklearn.naive_bayes import GaussianNB
# # gnb =GaussianNB()
# # nb_y_pred = gnb.fit(x, y)
# from sklearn import svm
# svc= reset_model('svc')
# svc.fit(x, y)
#$dataset= "clinagg_elix"
filename
os.path.exists(filename)
import pickle
from sklearn import model_selection
# save the model to disk
model_list=[rf,logreg, xgboost,svc ]
for element in model_list:#test_summary_df['model']:
filename = 'models/{}_{}_{}.sav'.format(date,dataset,type(element).__name__)
pickle.dump(element, open(filename, 'wb'))
# # load the model from disk
# loaded_model = pickle.load(open(filename, 'rb'))
# result = loaded_model.score(X_test, Y_test)
# print(result)
```
# tSNE visualization
* 5-29-19 changes: changed x_full to x_train, ie dimension reduction will be performed on train set since grower samples will only come from train set (since all_xy is made from trainset)
```
from __future__ import print_function
import time
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from mpl_toolkits.mplot3d import Axes3D
from sklearn.manifold import TSNE
def continuous_filter(df):
#global all_xy
categorical=['gender',
'ethnicity_black',
'ethnicity_hispanic',
'ethnicity_unknown/other',
'ethnicity_white/nonhispanic',
'ibands_absent',
'any_vasoactive_True',
'leukocyte_1',
'nitrite_1',
'pao2fio2ratio(200, 333]',
'pao2fio2ratio_(333, 475]',
'pao2fio2ratio_(475, 3000]',
'vent_recieved_1',
'vent_recieved_2',
'dobutamine_True',
'dopamine_True',
'epinephrine_True',
'norepinephrine_True',
'phenylephrine_True',
'rrt_True',
'vasopressin_True',
'ipco2_absent',
"cancer_elix_True"]
all_xy_label= list(all_xy)
in_both= list(set(categorical)& set(all_xy))
##restricting all_xy to only continuous variables
all_xy_cont = df.drop(in_both, axis=1)
return(all_xy_cont)
def saveplot(figure_name,folder_name=None):
"""
simple function for saving plots
"""
if folder_name != None:
address = 'figures/{}/'.format(folder_name)
else:
address = 'figures/'
if not os.path.exists(address):
os.makedirs(address)
plt.savefig(address+"/{}.png".format(figure_name),bbox_inches='tight')
#plt.close()
# dataprep
####### 5-29-19: changed this here for an easy fix and to realign the dimension reduction to be only on train data
x_full=x_train.copy()#pd.concat([x_train,x_test])
#x_full=pd.concat([x_train,x_test])
icu_full= z_icustay_id.copy()#icu_full= pd.concat([z_icustay_id,z_icustay_id_test])
y_full=pd.DataFrame(y_train)#pd.concat([pd.DataFrame(y_train), pd.DataFrame(y_test)])
##PCA
from sklearn.decomposition import PCA
time_start = time.time()
pca = PCA(n_components=4)
#continuous_filter(x_train)
pca_result = pca.fit_transform(continuous_filter(x_train)) #x_train
print('PCA done! Time elapsed: {} seconds'.format(time.time()-time_start))
pca_df = pd.DataFrame(columns = ['pca1','pca2','pca3','pca4'])
pca_df['pca1'] = pca_result[:,0]
pca_df['pca2'] = pca_result[:,1]
pca_df['pca3'] = pca_result[:,2]
pca_df['pca4'] = pca_result[:,3]
print ('Variance explained per principal component: {}'.format(pca.explained_variance_ratio_))
pca.explained_variance_ratio_[:2]
```
## PCA plot and grower labeling
* what is a good m value?
* do patients with different profiles: index 1 and 3457 differ in their grower samples on tsne?
```
# m=50
# loc_sample=z_icustay_id[1]
# grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
# pca_df['y']= y_full.values
# ##added to color grower
# pd.DataFrame(pca_df)['icustay_id']=icu_full.values
# pca_df['grower']=pca_df['icustay_id'].isin(list(grower_samples))
# pca_df['sample']=pca_df['icustay_id']==(testing_sample_icu)
# pal = sns.dark_palette("palegreen", as_cmap=True)
# plt.figure(figsize=(16,10))
# # sns.scatterplot(x="pca1",
# # y="pca2",
# # hue='grower',
# # size='grower',
# # sizes=[25, 25],
# # style='y',
# # palette=sns.color_palette(n_colors=2, desat=0.9),
# # data=pca_df,
# # legend="full",
# # alpha=[0.8,0.6]
# # )
# ax= sns.scatterplot(x="pca1",
# y="pca2",
# data= pca_df[pca_df.grower==True],
# #hue='y',
# # size='grower',
# # sizes=[15, 50],
# # style='y',
# palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
# alpha=1,
# legend="full",
# # alpha=[0.8,0.6]
# )
# sns.scatterplot(x="pca1",
# y="pca2",
# data= pca_df[pca_df.grower==False],
# alpha=0.2,
# #hue='y',
# #palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
# legend='full',
# ax=ax)
# # sns.scatterplot(x="pca1",
# # y="pca2",
# # data= pca_df[pca_df.grower==False],
# # alpha=0.1,
# # hue='y',
# # palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
# # #legend='full',
# # ax=ax)
pca_df['y']= y_full.values
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="pca1",
y="pca2",
hue='y',
data= pca_df,#[pca_df.grower==False],
alpha=0.7)
ax.set_title('global training PCA of pos vs neg patients')
saveplot(figure_name='global_training_PCA', folder_name='PCA')
# ## pcaplot and grower labeling [different style]
# m=100
# loc_sample=z_icustay_id[3457]
# grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
# pca_df['y']= y_full.values
# ##added to color grower
# pd.DataFrame(pca_df)['icustay_id']=icu_full.values
# pca_df['grower']=pca_df['icustay_id'].isin(list(grower_samples))
# pca_df['sample']=pca_df['icustay_id']==(testing_sample_icu)
# plt.figure(figsize=(16,10))
# sns.scatterplot(x="pca1",
# y="pca2",
# hue='grower',
# size='grower',
# sizes=[15, 80],
# #style='grower',
# #palette=sns.color_palette("hls", 2),
# data=pca_df,
# legend="full",
# alpha=0.6
# )
# ax.set_title('PCA of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
# ax.set(xlabel='PCA1 ({:.2f}% variance)'.format(pca.explained_variance_ratio_[:1][0]), ylabel='PCA2({:.2f}% variance)'.format(pca.explained_variance_ratio_[1:2][0]))
# saveplot(figure_name='local_training_PCA_m{}_icu{}'.format(m,index), folder_name='tsne')
## pcaplot and grower labeling
## pcaplot and grower labeling
m=100
index=1
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
pca_df['y']= y_full.values
##added to color grower
pd.DataFrame(pca_df)['icustay_id']=icu_full.values
pca_df['grower']=pca_df['icustay_id'].isin(list(grower_samples))
pca_df['sample']=pca_df['icustay_id']==(testing_sample_icu)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.grower==False],
alpha=0.2,
style="y",
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('PCA of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
ax.set(xlabel='PCA1 ({:.2f}% variance)'.format(pca.explained_variance_ratio_[:1][0]), ylabel='PCA2({:.2f}% variance)'.format(pca.explained_variance_ratio_[1:2][0]))
saveplot(figure_name='local_training_PCA_m{}_icu{}'.format(m,index), folder_name='PCA')
## pcaplot and grower labeling
m=100
index=3457
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
pca_df['y']= y_full.values
##added to color grower
pd.DataFrame(pca_df)['icustay_id']=icu_full.values
pca_df['grower']=pca_df['icustay_id'].isin(list(grower_samples))
#pca_df['sample']=pca_df['icustay_id']==(testing_sample_icu)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.grower==False],
alpha=0.2,
style="y",
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('PCA of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
ax.set(xlabel='PCA1 ({:.2f}% variance)'.format(pca.explained_variance_ratio_[:1][0]), ylabel='PCA2({:.2f}% variance)'.format(pca.explained_variance_ratio_[1:2][0]))
saveplot(figure_name='local_training_PCA_m{}_icu{}_allcont'.format(m,index), folder_name='PCA')
## pcaplot and grower labeling
m=100
index=3457
loc_sample=z_icustay_id[index]
euc_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
pca_df['y']= y_full.values
##added to color grower
pd.DataFrame(pca_df)['icustay_id']=icu_full.values
pca_df['euc']=pca_df['icustay_id'].isin(list(euc_samples))
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.euc==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="pca1",
y="pca2",
data=pca_df[pca_df.euc==False],
alpha=0.1,
style="y",
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax1.set_title('PCA of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
ax1.set(xlabel='PCA1 ({:.2f}% variance)'.format(pca.explained_variance_ratio_[:1][0]), ylabel='PCA2({:.2f}% variance)'.format(pca.explained_variance_ratio_[1:2][0]))
saveplot(figure_name='euc_training_PCA_m{}_icu{}_allcont'.format(m,index), folder_name='PCA')
```
## tSNE plot and grower labeling
* what is a good m value?
* do patients with different profiles: index 1 and 3457 differ in their grower samples on tsne?
```
##TSNE fit, run once
time_start = time.time()
tsne = TSNE(n_components=2, verbose=1, perplexity=40, n_iter=500)
tsne_results = tsne.fit_transform(continuous_filter(x_train))#x_full)
print('t-SNE done! Time elapsed: {} seconds'.format(time.time()-time_start))
##plotting all trainset labeling y==true, y==false.
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
plt.figure(figsize=(16,10))
ax= sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
hue='y',
data=tsne_df,
legend="full",
alpha=0.7
)
ax.set_title('global training t-SNE of pos vs neg patients')
#saveplot(figure_name='global_training_tsne', folder_name='tsne')
#m=100 tsne w patient at index 1: a y=0 patient and gender=1
m=100
index=1
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
#saveplot(figure_name='local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
#m=100 tsne w patient at index 3457: a y=1 patient and gender=0
m=100
index=3457
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
#saveplot(figure_name='local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
#####duplcicate plot with different style
#m=100 tsne w patient at index 3457: a y=1 patient and gender=0
# m=100
# loc_sample=z_icustay_id[3457]
# grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
# tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
# tsne_df['tsne-2d-one'] = tsne_results[:,0]
# tsne_df['tsne-2d-two'] = tsne_results[:,1]
# tsne_df['y']= y_full.values
# ##added to color grower
# pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
# tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
# tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
# plt.figure(figsize=(16,10))
# sns.scatterplot(x="tsne-2d-one",
# y="tsne-2d-two",
# hue='grower',
# size='grower',
# sizes=[30, 100],
# style='y',
# #palette=sns.color_palette("hls", 2),
# data=tsne_df,
# legend="full",
# alpha=0.7
# )
#m=250 tsne w patient at index 1
m=250
index=1
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='gower_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
#m=250 tsne w patient at index 3457
m=250
index=3457
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='gower_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
#m=50 tsne w patient at index 1
m=50
index=1
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='gower_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
#m=50 tsne w patient at index 3457
m=50
index=3457
loc_sample=z_icustay_id[index]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.grower==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='gower_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
## pcaplot and euclidean labeling
m=100
index=1
loc_sample=z_icustay_id[index]
euc_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
#grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['euc']=tsne_df['icustay_id'].isin(list(euc_samples))
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='euc_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
## pcaplot and euclidean labeling
m=100
index=3457
loc_sample=z_icustay_id[index]
euc_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
#grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['euc']=tsne_df['icustay_id'].isin(list(euc_samples))
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='euc_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
## pcaplot and euclidean labeling
m=250
index=1
loc_sample=z_icustay_id[index]
euc_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
#grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['euc']=tsne_df['icustay_id'].isin(list(euc_samples))
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='euc_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
## pcaplot and euclidean labeling
m=250
index=3457
loc_sample=z_icustay_id[index]
euc_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
#grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full.values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full.values
tsne_df['euc']=tsne_df['icustay_id'].isin(list(euc_samples))
plt.figure(figsize=(16,10))
ax=sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==True],
style="y",
size="y",
sizes=[80,80],
palette=sns.color_palette("Set2", n_colors=1, desat=.2),#sns.color_palette(sns.hls_palette(2, l=.3, s=.9)),#"hls",n_colors=2, desat=0.9),
alpha=1,
legend="full"
)
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
data=tsne_df[tsne_df.euc==False],
alpha=0.2,
#hue='y',
#palette=sns.color_palette(sns.color_palette("Set1", n_colors=2, desat=.9)),#sns.color_palette(sns.hls_palette(2, l=.5, s=.9)),#sns.color_palette("hls",n_colors=2, desat=0.8),
legend='full',
ax=ax)
ax.set_title('t-SNE of m={} gower closest ICU stay vs rest for icu_index={}'.format(m,index))
saveplot(figure_name='euc_local_training_tsne_m{}_icu{}'.format(m,index), folder_name='tsne')
```
## takeaway: the two patients do have similar tsne, however patient at index 1 is more dispersed across the reduced dimensions
```
len(all_xy)
# problem, currently select_train_samples are all producing the same m*2 cases regardless of the sample being input.
m=10
loc_sample=z_icustay_id[1]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
grower_samples
m=10
loc_sample=z_icustay_id[3457]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
grower_samples
```
## flip the script, perform a pca and tsne on only the grower samples
# 5/23/19
need to redo with only focusing on continuous variables for tsne and pca both (1) dimension reduction -> subset and (2) subset -> dimension reduction.
```
## pcaplot and gower labeling
m=50
loc_sample=z_icustay_id[3457]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
##PCA
from sklearn.decomposition import PCA
time_start = time.time()
pca = PCA(n_components=4)
##limiting xfull to just grower_samples
grower_index=icu_full.isin(grower_samples)
pca_result = pca.fit_transform(x_full[grower_index])
print('PCA done! Time elapsed: {} seconds'.format(time.time()-time_start))
pca_df = pd.DataFrame(columns = ['pca1','pca2','pca3','pca4'])
pca_df['pca1'] = pca_result[:,0]
pca_df['pca2'] = pca_result[:,1]
pca_df['pca3'] = pca_result[:,2]
pca_df['pca4'] = pca_result[:,3]
print ('Variance explained per principal component: {}'.format(pca.explained_variance_ratio_))
# ## pcaplot and grower labeling
pca_df['y']= y_full[grower_index].values
##added to color grower
pd.DataFrame(pca_df)['icustay_id']=icu_full[grower_index].values
pca_df['grower']=pca_df['icustay_id'].isin(list(grower_samples))
pca_df['sample']=pca_df['icustay_id']==(testing_sample_icu)
plt.figure(figsize=(16,10))
sns.scatterplot(x="pca1",
y="pca2",
hue='y',
#size='grower',
#sizes=[30, 70],
#style='grower',
#palette=sns.color_palette("hls", 2),
data=pca_df,
legend="full",
alpha=0.7
)
## pcaplot and gower labeling
m=50
loc_sample=z_icustay_id[3457]
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
##limiting xfull to just grower_samples
grower_index=icu_full.isin(grower_samples)
##TSNE fit
time_start = time.time()
tsne = TSNE(n_components=2, verbose=1, perplexity=40, n_iter=300)
tsne_results = tsne.fit_transform(x_full[grower_index])
print('t-SNE done! Time elapsed: {} seconds'.format(time.time()-time_start))
tsne_df = pd.DataFrame(columns = ['tsne-2d-one','tsne-2d-two'])
tsne_df['tsne-2d-one'] = tsne_results[:,0]
tsne_df['tsne-2d-two'] = tsne_results[:,1]
tsne_df['y']= y_full[grower_index].values
##added to color grower
pd.DataFrame(tsne_df)['icustay_id']=icu_full[grower_index].values
tsne_df['grower']=tsne_df['icustay_id'].isin(list(grower_samples))
tsne_df['sample']=tsne_df['icustay_id']==(loc_sample)
plt.figure(figsize=(16,10))
sns.scatterplot(x="tsne-2d-one",
y="tsne-2d-two",
hue='y',
data=tsne_df,
legend="full",
alpha=0.7
)
```
# 5-29-19:
### issue: tsne not in agreement with grower distance labeling. need to investigate
* Debug my t-sne visualization w/ grower labeling
* T-sne plot on only continuous variables
* Debug my grower distance:
* Grower can be an ensemble of distance:
* IE: Euclidian for continuous, count diff for categorical.
todo:
* How is it computing for continuous vs categorical?
* Maybe define own distance metric?
* Hamming distance? Can look into lots of different distance metrics.
* Can look into R package for grower distance. *probably fastest starting point for looking at other possibilities.
## local model testing
*<del> step1) hypertune xgb on 5fold cv.
*<del> step2) test entire trainset and predict trainset.
*<del> step3) run hypertuned model on 5fold cv with lr and get overall metrics.
* step4) local model testing
```
z_icustay_id
def single_split_training(m=250, n_sfk_split=5):
#######
skf = StratifiedKFold(n_splits=n_sfk_split) #Stratified K-Folds cross-validator
num_fold = 0
for train_index, test_index in skf.split(x, y):
X_train_0, X_test_0 = x[train_index], x[test_index] #assigning x_train and x_test sets within this cv fold
y_train_0, y_test_0 = y[train_index], y[test_index] #assigning y_train and y_test sets within this cv fold
#######
num_fold = num_fold + 1 ##silly to keep but it's from the loop
print('this is the results of the {} fold in 5 folds:'.format(num_fold))
print('the number of testing samples in this fold:', test_index.size)
train_z_icustay_id = z_icustay_id[train_index] # the icustay_id of samples in training set from 5 fold
test_z_icustay_id = z_icustay_id[test_index] # the icustay_id of samples in testing set from 5 fold
xg_one_fold_pred = [] # obtain the pred label of testing samples for one fold using xgboost
xg_one_fold_proba = [] # obtain the proba of testing samples for one fold using xgboost
lr_one_fold_pred = [] # obtain the pred label of testing samples for one fold using lr
lr_one_fold_proba = [] # obtain the proba of testing samples for one fold using lr
######
indicator_time = 0 # the indicator
for i, j in zip(test_z_icustay_id, test_index): #looping through the zipped indicies of the test indicies/test icustay_id
testing_sample_id = i #numerical index of first 1/2 of data ##??? this seems to be instead the
all_xy_0 = all_xy.loc[train_z_icustay_id] # select all TRAINING samples from the current fold using icustay_id index
all_xy_training = all_xy_0.append(all_xy.loc[i]) # append the current ith testing sample to the training set.
###important parameter. was at 400, i changed to X
m = m # m is the number of similar cases or similar controls
X_test_00 = x[j]
y_test = y[j]
X_test = X_test_00.reshape(1, -1)
# print 'start selecting......'
Id_train_set = select_train_samples(testing_sample_id, all_xy_training, m, time_interval) # individulization
ix = np.isin(z_icustay_id, Id_train_set)
Id_train_set_index = list(np.where(ix))
# Id_train_set_index = np.argwhere(z_icustay_id == Id_train_set)
X_train = x[Id_train_set_index]
y_train = y[Id_train_set_index]
#print('start training......')
# scoring = 'roc_auc'
# xgboost
#hyper parameter tuning F1 from gridsearchCV on 5cv:{'learning_rate': 0.1, 'max_depth': 5, 'scale_pos_weight': 5}
#hyper parameter tuning F1_macro from gridsearchCV on 5cv:{'learning_rate': 0.1, 'max_depth': 4, 'scale_pos_weight': 1}
#hyper parameter tuning recall_macro from gridsearchCV on 5cv:{'learning_rate': 0.1, 'max_depth': 5, 'scale_pos_weight': 5}
#hyper parameter tuning neg_log_loss from gridsearchCV on 5cv:{'learning_rate': 0.1, 'max_depth': 2, 'scale_pos_weight': 1}
# xgboost_mod = XGBClassifier(learning_rate=0.1, n_estimators=100, max_depth=2,
# min_child_weight=2, gamma=0, subsample=0.8, colsample_bytree=0.8,
# objective='binary:logistic', nthread=-1, scale_pos_weight=1, seed=27)
xgboost_mod= reset_model('xgboost')
xgboost_mod.fit(x[Id_train_set_index], y[Id_train_set_index])
xg_y_pred = xgboost_mod.predict(X_test)
xg_y_pred_proba = xgboost_mod.predict_proba(X_test)[:,1]
xg_one_fold_pred.append(xg_y_pred)
xg_one_fold_proba.append(xg_y_pred_proba)
# lr
logreg = LogisticRegression(penalty='l2', dual=False, tol=0.0001, C=10, fit_intercept=True,
intercept_scaling=1, class_weight='balanced', random_state=None)
logreg.fit(x[Id_train_set_index], y[Id_train_set_index])
lr_y_pred = logreg.predict(X_test)
lr_y_pred_proba = logreg.predict_proba(X_test)[:,1]
lr_one_fold_pred.append(lr_y_pred)
lr_one_fold_proba.append(lr_y_pred_proba)
indicator_time = indicator_time + 1
# print 'the next testing sample and total samples:', indicator_time, test_index.size
xg_y_individual_pred = np.array(xg_one_fold_pred)
xg_y_individual_proba = np.array(xg_one_fold_proba)
lr_y_individual_pred = np.array(lr_one_fold_pred)
lr_y_individual_proba = np.array(lr_one_fold_proba)
one_fold_y_test = y[test_index]
print ('---------new fold---------------')
print ('**** result of non-individual predictor using xgboost:')
print ('the Accuracy of one fold:', accuracy_score(y[test_index], xg_y_individual_pred))
print ('the AUC of one fold:', roc_auc_score(y[test_index], xg_y_individual_pred))
print ('the classification_report :', classification_report(y[test_index], xg_y_individual_pred))
print(confusion_matrix(y[test_index], xg_y_individual_pred))
print("\n")
print ('****this is the result of individual predictor using lr:')
print ('the Accuracy of one fold:', accuracy_score(y[test_index], lr_y_individual_pred))
print ('the AUC of one fold:', roc_auc_score(y[test_index], lr_y_individual_pred))
print ('the classification_report :', classification_report(y[test_index], lr_y_individual_pred))
print(confusion_matrix(y[test_index], lr_y_individual_pred))
print("\n")
#do we have multiple models for each patient? if so how do we aggregate?
single_split_training(m=100,n_sfk_split=2 )
#single_split_training(m=500,n_sfk_split=2 )
```
# M=50 Gower Local Modeling on entire training set
```
m=50
xg_fold_pred = [] # obtain the pred label of testing samples for one fold using xgboost
xg_fold_proba = [] # obtain the proba of testing samples for one fold using xgboost
lr_fold_pred = [] # obtain the pred label of testing samples for one fold using lr
lr_fold_proba = [] # obtain the proba of testing samples for one fold using lr
for i in range(0, len(z_icustay_id)):
if i%500==0:
print("iteration: ", i)
loc_sample=z_icustay_id[i]
test_index= z_icustay_id.index[z_icustay_id==loc_sample].tolist()[0]
test_sample= x[test_index].reshape((1,-1))
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
train_indices= z_icustay_id.index[z_icustay_id.isin(grower_samples)].tolist()
train_samples= x[train_indices].reshape((1,-1))
# xgboost
xgboost_mod= reset_model('xgboost')
xgboost_mod.fit(x[train_indices], y[train_indices])
xg_y_pred = xgboost_mod.predict(test_sample)
xg_y_proba = xgboost_mod.predict_proba(test_sample)[:,1]
xg_fold_pred.append(xg_y_pred)
xg_fold_proba.append(xg_y_proba)
# lr
lr_mod= reset_model('logreg')
lr_mod.fit(x[train_indices], y[train_indices])
lr_y_pred = lr_mod.predict(test_sample)
lr_y_proba = lr_mod.predict_proba(test_sample)[:,1]
lr_fold_pred.append(lr_y_pred)
lr_fold_proba.append(lr_y_proba)
classifier_eval(np.concatenate(xg_fold_proba), proba_input=True, x=x, y=y, save=False, model_name='local_xg', folder_name='clinical_agg')
classifier_eval(lr_fold_proba, proba_input=True, x=x, y=y, save=False, model_name='local_lr', folder_name='clinical_agg')
{'auc': 0.689467706613181,
'f1': 0.45111980092428017,
'model': 'local_xg',
'npv': 89.40397350993378,
'precision': 0.30007093875620716,
'recall': 0.9083750894774517}
{'auc': 0.6868647100930566,
'f1': 0.43407917383820993,
'model': 'local_lr',
'npv': 86.71875,
'precision': 0.285746657602538,
'recall': 0.9026485325697924}
```
# M=100 Gower Local Modeling on entire training set
```
m=100
xg_fold_pred_g100 = [] # obtain the pred label of testing samples for one fold using xgboost
xg_fold_proba_g100 = [] # obtain the proba of testing samples for one fold using xgboost
lr_fold_pred_g100 = [] # obtain the pred label of testing samples for one fold using lr
lr_fold_proba_g100 = [] # obtain the proba of testing samples for one fold using lr
for i in range(0, len(z_icustay_id)):
if i%500==0:
print("iteration: ", i)
loc_sample=z_icustay_id[i]
test_index= z_icustay_id.index[z_icustay_id==loc_sample].tolist()[0]
test_sample= x[test_index].reshape((1,-1))
grower_samples=select_train_samples(loc_sample, all_xy, m, time_interval)
train_indices= z_icustay_id.index[z_icustay_id.isin(grower_samples)].tolist()
train_samples= x[train_indices].reshape((1,-1))
# xgboost
xgboost_mod= reset_model('xgboost')
xgboost_mod.fit(x[train_indices], y[train_indices])
xg_y_pred = xgboost_mod.predict(test_sample)
xg_y_proba = xgboost_mod.predict_proba(test_sample)[:,1]
xg_fold_pred_g100.append(xg_y_pred)
xg_fold_proba_g100.append(xg_y_proba)
# lr
lr_mod= reset_model('logreg')
lr_mod.fit(x[train_indices], y[train_indices])
lr_y_pred = lr_mod.predict(test_sample)
lr_y_proba = lr_mod.predict_proba(test_sample)[:,1]
lr_fold_pred_g100.append(lr_y_pred)
lr_fold_proba_g100.append(lr_y_proba)
classifier_eval(np.concatenate(xg_fold_proba_g100), proba_input=True, x=x, y=y, save=True, model_name='local_xg_g100', folder_name='clinical_agg')
classifier_eval(np.concatenate(lr_fold_proba_g100), proba_input=True, x=x, y=y, save=True, model_name='local_lr_g100', folder_name='clinical_agg')
```
# euclidean local method
```
m=50
xg_fold_pred2 = [] # obtain the pred label of testing samples for one fold using xgboost
xg_fold_proba2 = [] # obtain the proba of testing samples for one fold using xgboost
lr_fold_pred2 = [] # obtain the pred label of testing samples for one fold using lr
lr_fold_proba2 = [] # obtain the proba of testing samples for one fold using lr
for i in range(0, len(z_icustay_id)):
if i%1000==0:
print("iteration: ", i)
loc_sample=z_icustay_id[i]
test_index= z_icustay_id.index[z_icustay_id==loc_sample].tolist()[0]
test_sample= x[test_index].reshape((1,-1))
grower_samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
train_indices= z_icustay_id.index[z_icustay_id.isin(grower_samples)].tolist()
train_samples= x[train_indices].reshape((1,-1))
# xgboost
xgboost_mod= reset_model('xgboost')
xgboost_mod.fit(x[train_indices], y[train_indices])
xg_y_pred = xgboost_mod.predict(test_sample)
xg_y_proba = xgboost_mod.predict_proba(test_sample)[:,1]
xg_fold_pred2.append(xg_y_pred)
xg_fold_proba2.append(xg_y_proba)
# lr
lr_mod= reset_model('logreg')
lr_mod.fit(x[train_indices], y[train_indices])
lr_y_pred = lr_mod.predict(test_sample)
lr_y_proba = lr_mod.predict_proba(test_sample)[:,1]
lr_fold_pred2.append(lr_y_pred)
lr_fold_proba2.append(lr_y_proba)
#x_test#.loc[z_icustay_id_test[0]]
#loading in data saved from past night
xg_fold_proba2=pd.read_csv("models/xg_fold_proba2.csv") #two class training data
lr_fold_proba2=pd.read_csv("models/lr_fold_proba2.csv") #two class training data
xg_fold_proba2['0'].describe()
classifier_eval(np.array(xg_fold_proba2['0']), proba_input=True, x=x, y=y, save=True, model_name='local_xg_euc', folder_name='clinical_agg')
classifier_eval(np.array(lr_fold_proba2['0']), proba_input=True, x=x, y=y, save=True, model_name='local_lr_euc', folder_name='clinical_agg')
pd.DataFrame(np.concatenate(xg_fold_proba2)).to_csv('models/xg_fold_proba2.csv')
pd.DataFrame(np.concatenate(lr_fold_proba2)).to_csv('models/lr_fold_proba2.csv')
xg_gow_50={'auc': 0.689467706613181,
'f1': 0.45111980092428017,
'model': 'local_xg_g50',
'npv': 89.40397350993378,
'precision': 0.30007093875620716,
'recall': 0.9083750894774517}
lr_gow_50={'auc': 0.6868647100930566,
'f1': 0.43407917383820993,
'model': 'local_lr_g50',
'npv': 86.71875,
'precision': 0.285746657602538,
'recall': 0.9026485325697924}
xg_gow_100={'auc': 0.7206768216191698,
'f1': 0.45500542103361047,
'model': 'local_xg_g100',
'npv': 89.38461538461539,
'precision': 0.30432680686487795,
'recall': 0.9012168933428776}
lr_gow_100={'auc': 0.6788229905667733,
'f1': 0.4141563786008231,
'model': 'local_lr_g100',
'npv': 81.68642951251647,
'precision': 0.2689183411714408,
'recall': 0.9005010737294202}
xg_euc_50={'auc': 0.7570882088208821,
'f1': 0.4844170834936514,
'model': 'local_xg_euc',
'npv': 91.56479217603912,
'precision': 0.3312286240463036,
'recall': 0.9012168933428776}
lr_euc_50={'auc': 0.6994438223349895,
'f1': 0.4324139112557821,
'model': 'local_lr_euc',
'npv': 86.45937813440321,
'precision': 0.2842342342342342,
'recall': 0.9033643521832498}
#classifier_eval(gnb, x=x_test, y=y_test)
local_summary_df= pd.DataFrame([xg_gow_50,
lr_gow_50,
xg_gow_100,
lr_gow_100,
xg_euc_50,
lr_euc_50])
local_summary_df.set_index('model').round(decimals=2)
```
# local modeling on test set
```
import time, sys
from IPython.display import clear_output
def update_progress(progress):
bar_length = 20
if isinstance(progress, int):
progress = float(progress)
if not isinstance(progress, float):
progress = 0
if progress < 0:
progress = 0
if progress >= 1:
progress = 1
block = int(round(bar_length * progress))
clear_output(wait = True)
text = "Progress: [{0}] {1:.1f}%".format( "#" * block + "-" * (bar_length - block), progress * 100)
print(text)
# number_of_elements = 1000
# for i in range(number_of_elements):
# time.sleep(0.1) #Replace this with a real computation
# update_progress(i / number_of_elements)
# update_progress(1)
def local_model(metric, m, test=True):
global all_xy, all_xy_test
xg_fold_proba = [] # obtain the proba of testing samples for one fold using xgboost
lr_fold_proba = [] # obtain the proba of testing samples for one fold using lr
## adding block to accomidate training or test sample specified.
if test==False:
z_icustay= z_icustay_id
x= np.array(x_train.copy())
else:
z_icustay= z_icustay_id_test
x= np.array(x_test.copy())
for i in range(0, len(z_icustay)):
# establishing index and icustay of the specified testing sample
loc_sample=z_icustay[i]
test_index= z_icustay.index[z_icustay==loc_sample].tolist()[0]
test_sample= x[test_index].reshape((1,-1)) #sample to be predicted.
#adding test set sample to all_xy for calculating 2m closest training samples
if loc_sample in all_xy_test.index: #if trying to find distance of a sample in test set
all_xy=all_xy.append(all_xy_test.loc[loc_sample])
###calc distance
if metric =='gower':
samples=select_train_samples(loc_sample, all_xy, m, time_interval)
elif metric =='euclidean':
samples=m_distance_samples(loc_sample, all_xy, m, time_interval, metric='euclidean')
#removing test set sample to all_xy for calculating 2m closest training samples
if loc_sample in all_xy_test.index & all_xy.index: #if trying to find distance of a sample in test set
all_xy.drop(index=loc_sample, axis=1,inplace=True)
##extracting out only the 2m closest samples to fit the model on.
train_indices= z_icustay_id.index[z_icustay_id.isin(samples)].tolist()
train_samples= np.array(x_train)[train_indices].reshape((1,-1))
# xgboost
xgboost_mod= reset_model('xgboost')
xgboost_mod.fit(np.array(x_train)[train_indices], y[train_indices])
xg_y_proba = xgboost_mod.predict_proba(test_sample)[:,1]
xg_fold_proba.append(xg_y_proba)
# lr
lr_mod= reset_model('logreg')
lr_mod.fit(np.array(x_train)[train_indices], y[train_indices])
lr_y_proba = lr_mod.predict_proba(test_sample)[:,1]
lr_fold_proba.append(lr_y_proba)
update_progress(i / len(z_icustay))
update_progress(1)
return(xg_fold_proba, lr_fold_proba)
xg_proba_euc_m50, lr_proba_euc_m50 = local_model(metric='euclidean', m=50, test=True) # works
xg_proba_euc_m100, lr_proba_euc_m100 = local_model(metric='euclidean', m=100, test=True)
xg_proba_gow_m50, lr_proba_gow_m50 = local_model(metric='gower', m=50, test=True)
xg_proba_gow_m100, lr_proba_gow_m100 = local_model(metric='gower', m=100, test=True)
#np.array(xg_proba_euc_m50)
y_test
local_xg_euc_50=classifier_eval(np.array(xg_proba_euc_m50), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_xg_euc_50', folder_name='clinical_agg')
local_lr_euc_50= classifier_eval(np.array(lr_proba_euc_m50), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_lr_euc_50', folder_name='clinical_agg')
local_xg_euc_100=classifier_eval(np.array(xg_proba_euc_m100), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_xg_euc_100', folder_name='clinical_agg')
local_lr_euc_100=classifier_eval(np.array(lr_proba_euc_m100), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_lr_euc_100', folder_name='clinical_agg')
local_xg_gow_50=classifier_eval(np.array(xg_proba_gow_m50), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_xg_gow_50', folder_name='clinical_agg')
local_lr_gow_50=classifier_eval(np.array(lr_proba_gow_m50), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_lr_gow_50', folder_name='clinical_agg')
local_xg_gow_100=classifier_eval(np.array(xg_proba_gow_m100), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_xg_gow_100', folder_name='clinical_agg')
local_lr_gow_100=classifier_eval(np.array(lr_proba_gow_m100), proba_input=True, x=x_test, y=y_test, save=True, model_name='local_lr_gow_100', folder_name='clinical_agg')
#classifier_eval(gnb, x=x_test, y=y_test)
local_summary_df= pd.DataFrame([local_xg_euc_50,
local_lr_euc_50,
local_xg_euc_100,
local_lr_euc_100,
local_xg_gow_50,
local_lr_gow_50,
local_xg_gow_100,
local_lr_gow_100])
local_summary_df.set_index('model').round(decimals=2)
pd.DataFrame(np.concatenate(xg_proba_euc_m50)).to_csv('models/xg_proba_euc_m50.csv')
pd.DataFrame(np.concatenate(lr_proba_euc_m50)).to_csv('models/lr_proba_euc_m50.csv')
pd.DataFrame(np.concatenate(xg_proba_euc_m100)).to_csv('models/xg_proba_euc_m100.csv')
pd.DataFrame(np.concatenate(lr_proba_euc_m100)).to_csv('models/lr_proba_euc_m100.csv')
pd.DataFrame(np.concatenate(xg_proba_gow_m50)).to_csv('models/xg_proba_gow_m50.csv')
pd.DataFrame(np.concatenate(lr_proba_gow_m50)).to_csv('models/lr_proba_gow_m50.csv')
pd.DataFrame(np.concatenate(xg_proba_gow_m100)).to_csv('models/xg_proba_gow_m100.csv')
pd.DataFrame(np.concatenate(lr_proba_gow_m100)).to_csv('models/lr_proba_gow_m100.csv')
```
| github_jupyter |
Kod bazowany na Roboflow Tensorflow detection: https://colab.research.google.com/drive/1Cx9cYmclvREcCQLf6iu4hjckNx_zNMap
Najpierw instalujemy TensorFlow
```
!pip install -U --pre tensorflow_gpu=="2.3.0"
import os
import pathlib
# Klonowanie modelów z githuba Tensorflow
if "models" in pathlib.Path.cwd().parts:
while "models" in pathlib.Path.cwd().parts:
os.chdir('..')
elif not pathlib.Path('models').exists():
!git clone --depth 1 https://github.com/tensorflow/models
# Instalacja Object Detection API
%%bash
cd models/research/
protoc object_detection/protos/*.proto --python_out=.
cp object_detection/packages/tf2/setup.py .
python -m pip install .
```
Importowanie wymaganych zależności
```
import matplotlib
import matplotlib.pyplot as plt
import os
import random
import io
import imageio
import glob
import scipy.misc
import numpy as np
from six import BytesIO
from PIL import Image, ImageDraw, ImageFont
from IPython.display import display, Javascript
from IPython.display import Image as IPyImage
import tensorflow as tf
from object_detection.utils import label_map_util
from object_detection.utils import config_util
from object_detection.utils import visualization_utils as viz_utils
from object_detection.utils import colab_utils
from object_detection.builders import model_builder
%matplotlib inline
#run model builder test
!python /content/models/research/object_detection/builders/model_builder_tf2_test.py
```
Definicja pomocnych funkcji:
```
def load_image_into_numpy_array(path): #Funkcja zwraca obraz jako macierz numpy
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
def plot_detections(image_np,
boxes,
classes,
scores,
category_index,
figsize=(12, 16),
image_name=None):
image_np_with_annotations = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_annotations,
boxes,
classes,
scores,
category_index,
use_normalized_coordinates=True,
min_score_thresh=0.8)
if image_name:
plt.imsave(image_name, image_np_with_annotations)
else:
plt.imshow(image_np_with_annotations)
```
Przygotowanie danych:
Wykorzystywany jest system roboflow tworzący dataseta w określonym formacie. Tutaj jako TFRecords.
```
#Downloading data from Roboflow
%cd /content
!curl -L "https://app.roboflow.com/ds/YMTofWuMb1?key=tHLKyQlgw0" > roboflow.zip; unzip roboflow.zip; rm roboflow.zip
test_record_fname = '/content/valid/potholes.tfrecord'
train_record_fname = '/content/train/potholes.tfrecord'
label_map_pbtxt_fname = '/content/train/potholes_label_map.pbtxt'
```
Konfiguracja modelu:
```
#W tym miejscu można dobrać różne modele dostępne w tensorflow.
MODELS_CONFIG = {
'efficientdet-d0': {
'model_name': 'efficientdet_d0_coco17_tpu-32',
'base_pipeline_file': 'ssd_efficientdet_d0_512x512_coco17_tpu-8.config',
'pretrained_checkpoint': 'efficientdet_d0_coco17_tpu-32.tar.gz',
'batch_size': 16
},
'efficientdet-d1': {
'model_name': 'efficientdet_d1_coco17_tpu-32',
'base_pipeline_file': 'ssd_efficientdet_d1_640x640_coco17_tpu-8.config',
'pretrained_checkpoint': 'efficientdet_d1_coco17_tpu-32.tar.gz',
'batch_size': 16
},
'efficientdet-d2': {
'model_name': 'efficientdet_d2_coco17_tpu-32',
'base_pipeline_file': 'ssd_efficientdet_d2_768x768_coco17_tpu-8.config',
'pretrained_checkpoint': 'efficientdet_d2_coco17_tpu-32.tar.gz',
'batch_size': 16
},
'efficientdet-d3': {
'model_name': 'efficientdet_d3_coco17_tpu-32',
'base_pipeline_file': 'ssd_efficientdet_d3_896x896_coco17_tpu-32.config',
'pretrained_checkpoint': 'efficientdet_d3_coco17_tpu-32.tar.gz',
'batch_size': 16
}
}
chosen_model = 'efficientdet-d0'
num_steps = 1000
num_eval_steps = 500
model_name = MODELS_CONFIG[chosen_model]['model_name']
pretrained_checkpoint = MODELS_CONFIG[chosen_model]['pretrained_checkpoint']
base_pipeline_file = MODELS_CONFIG[chosen_model]['base_pipeline_file']
batch_size = MODELS_CONFIG[chosen_model]['batch_size']
```
Do wybranego modelu należy pobrać wagi i konfiguracje:
```
%mkdir /content/models/research/deploy/
%cd /content/models/research/deploy/
import tarfile
download_tar = 'http://download.tensorflow.org/models/object_detection/tf2/20200711/' + pretrained_checkpoint
!wget {download_tar}
tar = tarfile.open(pretrained_checkpoint)
tar.extractall()
tar.close()
%cd /content/models/research/deploy
download_config = 'https://raw.githubusercontent.com/tensorflow/models/master/research/object_detection/configs/tf2/' + base_pipeline_file
!wget {download_config}
pipeline_fname = '/content/models/research/deploy/' + base_pipeline_file
fine_tune_checkpoint = '/content/models/research/deploy/' + model_name + '/checkpoint/ckpt-0'
def get_num_classes(pbtxt_fname):
from object_detection.utils import label_map_util
label_map = label_map_util.load_labelmap(pbtxt_fname)
categories = label_map_util.convert_label_map_to_categories(
label_map, max_num_classes=90, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
return len(category_index.keys())
num_classes = get_num_classes(label_map_pbtxt_fname)
```
Zmieniamy elemnty pliku konfiguracyjnego:
```
import re
%cd /content/models/research/deploy
print('writing custom configuration file')
with open(pipeline_fname) as f:
s = f.read()
with open('pipeline_file.config', 'w') as f:
# fine_tune_checkpoint
s = re.sub('fine_tune_checkpoint: ".*?"',
'fine_tune_checkpoint: "{}"'.format(fine_tune_checkpoint), s)
# tfrecord files train and test.
s = re.sub(
'(input_path: ".*?)(PATH_TO_BE_CONFIGURED/train)(.*?")', 'input_path: "{}"'.format(train_record_fname), s)
s = re.sub(
'(input_path: ".*?)(PATH_TO_BE_CONFIGURED/val)(.*?")', 'input_path: "{}"'.format(test_record_fname), s)
# label_map_path
s = re.sub(
'label_map_path: ".*?"', 'label_map_path: "{}"'.format(label_map_pbtxt_fname), s)
# Set training batch_size.
s = re.sub('batch_size: [0-9]+',
'batch_size: {}'.format(batch_size), s)
# Set training steps, num_steps
s = re.sub('num_steps: [0-9]+',
'num_steps: {}'.format(num_steps), s)
# Set number of classes num_classes.
s = re.sub('num_classes: [0-9]+',
'num_classes: {}'.format(num_classes), s)
#fine-tune checkpoint type
s = re.sub(
'fine_tune_checkpoint_type: "classification"', 'fine_tune_checkpoint_type: "{}"'.format('detection'), s)
f.write(s)
%cat /content/models/research/deploy/pipeline_file.config
pipeline_file = '/content/models/research/deploy/pipeline_file.config'
model_dir = '/content/training/'
```
Pętla treningowa:
```
!python /content/models/research/object_detection/model_main_tf2.py \
--pipeline_config_path={pipeline_file} \
--model_dir={model_dir} \
--alsologtostderr \
--num_train_steps={num_steps} \
--sample_1_of_n_eval_examples=1 \
--num_eval_steps={num_eval_steps}
```
Test modelu:
Pobranie danych testowych z Roboflow:
```
%mkdir /content/test/
%cd /content/test/
!curl -L "https://app.roboflow.com/ds/YMTofWuMb1?key=tHLKyQlgw0" > roboflow.zip; unzip roboflow.zip; rm roboflow.zipy
import matplotlib
import matplotlib.pyplot as plt
import io
import scipy.misc
import numpy as np
from six import BytesIO
from PIL import Image, ImageDraw, ImageFont
import tensorflow as tf
from object_detection.utils import label_map_util
from object_detection.utils import config_util
from object_detection.utils import visualization_utils as viz_utils
from object_detection.builders import model_builder
%matplotlib inline
def load_image_into_numpy_array(path):
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
%ls '/content/training/'
```
Załadowanie modelu:
```
pipeline_config = pipeline_file
model_dir = '/content/training/ckpt-1'
configs = config_util.get_configs_from_pipeline_file(pipeline_config)
model_config = configs['model']
detection_model = model_builder.build(
model_config=model_config, is_training=False)
ckpt = tf.compat.v2.train.Checkpoint(
model=detection_model)
ckpt.restore(os.path.join('/content/training/ckpt-2'))
def get_model_detection_function(model):
@tf.function
def detect_fn(image):
image, shapes = model.preprocess(image)
prediction_dict = model.predict(image, shapes)
detections = model.postprocess(prediction_dict, shapes)
return detections, prediction_dict, tf.reshape(shapes, [-1])
return detect_fn
detect_fn = get_model_detection_function(detection_model)
```
Załadowanie mapy z nazwami obiektu:
```
label_map_path = configs['eval_input_config'].label_map_path
label_map = label_map_util.load_labelmap(label_map_path)
categories = label_map_util.convert_label_map_to_categories(
label_map,
max_num_classes=label_map_util.get_max_label_map_index(label_map),
use_display_name=True)
category_index = label_map_util.create_category_index(categories)
label_map_dict = label_map_util.get_label_map_dict(label_map, use_display_name=True)
```
Test:
```
import random
TEST_IMAGE_PATHS = glob.glob('/content/test/test/*.jpg')
image_path = random.choice(TEST_IMAGE_PATHS)
image_np = load_image_into_numpy_array(image_path)
input_tensor = tf.convert_to_tensor(
np.expand_dims(image_np, 0), dtype=tf.float32)
detections, predictions_dict, shapes = detect_fn(input_tensor)
label_id_offset = 1
image_np_with_detections = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_detections,
detections['detection_boxes'][0].numpy(),
(detections['detection_classes'][0].numpy() + label_id_offset).astype(int),
detections['detection_scores'][0].numpy(),
category_index,
use_normalized_coordinates=True,
max_boxes_to_draw=200,
min_score_thresh=.5,
agnostic_mode=False,
)
plt.figure(figsize=(12,16))
plt.imshow(image_np_with_detections)
plt.show()
```
| github_jupyter |
# <center>Oil Properties Estimation</center>
### <center>Sept 2014<br>Author: Bill Lehr<br>Edited By: James L. Makela</center>
## <u>Minimum Requirements</u>
Adios 3 requires certain minimum amounts of data for any oil to be included in the library
Crude Oil:
<ul>
<li>API and/or density at a reference temperature</li>
<li>Viscosity (either kinematic or dynamic) at a reference temperature</li>
</ul>
Refined Product or 'Other' oil:
<ul>
<li>API and/or density at a reference temperature</li>
<li>Viscosity (either kinematic or dynamic) at a reference temperature</li>
<li>At least three distillation cuts giving mass or volume fraction at boiling(bubble) point</li>
</ul>
Unlike Adios 2, Adios 3 stores both structural and distillation fractional components <b><u>(see figure 1)</u></b>.
Certain properties are valid for the whole oil while others may be defined for each structural fraction and still others vary for both the distillation and chemical structural fraction.
The SQL oil database needs a complete set of oil data. If measured data exists, it takes priority over estimated values. Many of the more complex estimation formulas are based upon Adios 2 calculations and/or the following reference:
<b><i>Characterization and Properties of Petroleum Fractions</b></i><br>
<b>Author:</b> Dr. M. R. Riazi, Professor of Chemical Engineering, Kuwait University<br>
<b>Published:</b> 2005<br>
<b>Publisher:</b> American Society for Testing and Materials (ASTM), International<br>
<b>Stock No.:</b> MNL50<br>
This will be abbreviated further in this document as CPPF.
## <u>Setting Up for our Estimations</u>
This is just a bit of setup so that we may verify our estimations with code snippets.
```
%pylab inline
import numpy as np
from scipy.optimize import curve_fit
import oil_library
from oil_library.models import Oil, ImportedRecord, KVis, Density
from pprint import PrettyPrinter
pp = PrettyPrinter(indent=2, width=120)
session = oil_library._get_db_session()
# these are some reasonable samples of oil records in the oil library
ans_mp = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'ALASKA NORTH SLOPE (MIDDLE PIPELINE)').one()
ans_2002 = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'ALASKA NORTH SLOPE (2002)').one()
bahia = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'BAHIA').one()
arabian = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'ARABIAN MEDIUM, AMOCO').one()
# These samples are problematic in some way or another
canola = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'CANOLA OIL (REFERENCE)').one()
abu_safah = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'ABU SAFAH').one()
sajaa = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'SAJAA CONDENSATE, BP').one()
bontang = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'BONTANG MIX, BP').one()
geisum = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'NORTH GEISUM, GEISUM OIL').one()
cl_bitumen = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'COLD LAKE BITUMEN').one()
bonny_light = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'BONNY LIGHT').one()
petro_star = session.query(ImportedRecord).filter(ImportedRecord.oil_name ==
'FUEL OIL NO.1 (DIESEL/HEATING FUEL), PETRO STAR').one()
ifo_180 = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'IFO 180').one()
boscan = session.query(ImportedRecord).filter(ImportedRecord.oil_name == 'BOSCAN, OIL & GAS').one()
print ans_mp
print ans_2002
print bahia
print arabian
print canola
```
## <u>Procedural Order of Oil Estimation</u>
Our estimations of properties for an input oil record are based on a set of minimal data requirements.
We necessarily build upon this minimal set in an ordered fashion because some calculations are needed to provide input to later calculations.
If we could describe the general flow of calculations it might be something like this.
- Foundational Aggregate Oil Properties
- Oil Distillation Fractional Properties
- Oil SARA Component Properties
- Oil Miscellaneous Properties
## <u>Foundational Aggregate Oil Properties</u>
These are the baseline oil property estimations which take the properties supplied by the input oil record and produce the minimum set of whole oil properties necessary to estimate the more complex properties, such as the distillation fractions and SARA component properties. These are:
- Density
- API
- Viscosity
### 1. Density:
(A) If no density value exists, estimate it from the oil's API using the following equation:
$$
\begin{align}
\rho0_{oil} &= \text{density of the oil at a } T_{ref} \text{of } 288.15^\circ K \,\, (kg/m^3) \cr
&= {141.5 \over 131.5 + API} \cdot 1000 \qquad \qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 1)} \cr
\end{align}
$$
(B) If a density measurement $\rho_0$ at some temperature $T_0$ exists, but no API, then (eq. 1) can be inverted to give an API.
$$
\begin{align}
API &= \left( {141.5 \over \rho0_{oil}} \cdot 1000 \right) - 131.5
\qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 2)} \cr
\end{align}
$$
But this can only be done after the density value has been adjusted to be a density at $T_{ref} = 288.15^\circ K (15^\circ C)$. The density conversion formula for different temperatures is:
$$
\begin{align}
\rho_0 &= \text{measured density} \cr
T_0 &= \text{temperature at which density is measured} \cr
k_{\rho T} &= 0.0008 \, K^{-1} \cr
\cr
\rho_{ref} &= \rho_0 \cdot (1 - k_{\rho T} \cdot (T_{ref} - T_0 ))
\qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 3)} \cr
\end{align}
$$
<i>(Referenced Source: Adios2, File: InputData.cpp, Function: CAdiosData::CalcRhoOil() )</i>
```
def lowest_temperature(obj_list):
'''
General utility function.
From a list of objects containing a ref_temp_k attribute,
return the object that has the lowest temperature
'''
if len(obj_list) > 0:
return sorted(obj_list, key=lambda d: d.ref_temp_k)[0]
else:
return None
def closest_to_temperature(obj_list, temperature):
'''
General Utility Function
From a list of objects containing a ref_temp_k attribute,
return the object that is closest to the specified temperature
'''
temp_diffs = [(obj, abs(obj.ref_temp_k - temperature))
for obj in obj_list
if obj.ref_temp_k is not None]
if len(temp_diffs) > 0:
return sorted(temp_diffs, key=lambda d: d[1])[0][0]
else:
return None
def estimate_density_from_api(api):
kg_m_3 = 141.5 / (131.5 + api) * 1000.0
ref_temp_k = 273.15 + 15.0
return kg_m_3, ref_temp_k
def estimate_api_from_density(density):
return (141.5 / density * 1000.0) - 131.5
def estimate_density_at_temp(ref_density, ref_temp_k,
temperature):
k_pt = 0.0008
return ref_density / (1 - k_pt * (ref_temp_k - temperature))
def oil_density_at_temp(imported_rec, temperature, weathering=0.0):
density_list = [d for d in imported_rec.densities
if (d.kg_m_3 is not None and
d.ref_temp_k is not None and
d.weathering == weathering)]
closest_density = closest_to_temperature(density_list, temperature)
if closest_density is not None:
d_ref, t_ref = (closest_density.kg_m_3,
closest_density.ref_temp_k)
elif imported_rec.api is not None:
d_ref, t_ref = estimate_density_from_api(imported_rec.api)
else:
return None
return estimate_density_at_temp(d_ref, t_ref, temperature)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.api = ', obj.api
print 'oil.densities = ', obj.densities
if obj.api is not None:
print 'density(api) = ', estimate_density_from_api(obj.api)
else:
print 'density(api) = N/A'
if len(obj.densities) > 0:
print 'API(density) = ', estimate_api_from_density(oil_density_at_temp(obj, 288.15))
else:
print 'API(density) = N/A'
print 'density at 288.15K = ', (oil_density_at_temp(obj, 288.15), 288.15)
print
```
### 2. Kinematic Viscosity:
The input oil record could contain a number of viscosity properties, measured as either dynamic viscosity, kinematic viscosity, or a mix of both. Each viscosity property in the oil record is accompanied by a reference temperature $\,^\circ K$ which is the temperature at which the viscosity was measured at. We want to deal exclusively in kinematic viscosity.
Converting a dynamic viscosity value into a kinematic viscosity value is done as follows.
$$
\begin{align}
\eta 0_{oil} &= \text{measured dynamic viscosity } (kg/(m \cdot s)) \cr
\rho 0_{oil} &= \text{measured density } (kg/m^3) \cr
\cr
v0_{oil} &= \text{kinematic viscosity } (m^2/s) \cr
&= {\eta 0_{oil} \over \rho 0_{oil} }
\qquad \qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 4)} \cr
\end{align}
$$
Basically we want to take advantage of any and all measured viscosities that are available. We will of course prefer the kinematic measurements, but we will want to use any dynamic viscosity measurements that are not redundant. So the basic program flow will be as follows:
<ul>
<li>Collect all existing measured kinematic viscosities and their associated reference temperatures</li>
<li>Collect any dynamic viscosities measured at temperatures not already represented in the kinematic measurements</li>
<li>Convert the dynamic viscosities into kinematic viscosities using (eq. 3)</li>
<li>Combine our kinematic and converted dynamic viscosities into one aggregate list</li>
</ul>
Using the measured kinematic viscosities, we should be able to estimate the oil kinematic viscosity at an arbitrary temperature. I believe this is what Adios2 does:
$$
\begin{align}
T &= \text{temperature in } \,^\circ K \cr
k_{v2} &= 5000^\circ K \cr
T_{ref} &= \text{measured reference temperature} \cr
v_{ref} &= \text{measured reference viscosity} \cr
\cr
v_T &= \text{the viscosity at a specified temperature} \cr
&= v_{ref} \cdot exp \left( {k_{v2} \over T} - {k_{v2} \over T_{ref}} \right)
\qquad \qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 5)} \cr
\end{align}
$$
<i>(Referenced Source: Adios2, File: OilInitialize.cpp, Function: CAdiosData::InitialViscosity() )</i>
```
def dvis_to_kvis(dvis, density):
return dvis / density
def estimate_kvis_at_temp(temp_k, kvis_ref, ref_temp_k):
k_v2 = 5000.0
return kvis_ref * np.exp(k_v2 / temp_k - k_v2 / ref_temp_k)
def oil_aggregate_kvis(imported_rec):
kvis_list = [(k.ref_temp_k, k.m_2_s)
for k in imported_rec.kvis
if k.m_2_s is not None and
k.ref_temp_k is not None]
dvis_list = [(d.ref_temp_k,
dvis_to_kvis(d.kg_ms,
oil_density_at_temp(imported_rec, d.ref_temp_k)))
for d in imported_rec.dvis
if d.kg_ms is not None and
d.ref_temp_k is not None]
agg = dict(dvis_list)
agg.update(kvis_list)
return [KVis(ref_temp_k=i[0], m_2_s=i[1])
for i in agg.iteritems()]
def oil_kvis_at_temp(imported_rec, temp_k, weathering=0.0):
kvis_list = [kv for kv in oil_aggregate_kvis(imported_rec)
if (kv.weathering == weathering)]
closest_kvis = closest_to_temperature(kvis_list, temp_k)
if closest_kvis is not None:
kvis_ref, t_ref = (closest_kvis.m_2_s,
closest_kvis.ref_temp_k)
else:
return None
return estimate_kvis_at_temp(temp_k, kvis_ref, t_ref)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.kvis = ', obj.kvis
print 'oil.dvis = ', obj.dvis
print 'aggregate kvis = ', oil_aggregate_kvis(obj)
print 'kvis at 288k = ', oil_kvis_at_temp(obj, 288.0)
print
# TODO: Okay, the ANS (2002) oil record has a crazy list of dynamic viscosities
# This has got to be fixed.
```
## <u>Oil Distillation Fractional Properties</u>
A large number of our estimations depend upon a characterization of the fractional amounts of compounds that fit somewhere in a high level classification named SARA. SARA is an acronym meaning Saturates, Aromatics, Resins, and Asphaltenes. There is much literature on the characterization of these petroleum hydrocarbons, but generally:
- Saturates are volatile, and evaporate at an estimable rate with increasing temperature.
- Aromatics are similarly volatile, but at different rates.
- Resins are considered inert. That is to say they will not evaporate at any reasonable temperature.
- Asphaltenes are also considered inert.
Most oil records do not contain precise information regarding these fractional amounts. But they do often contain fractional distillation cut properties. The distillation cuts describe the rate at which the volatile portions of the oil evaporate. And from this we can attempt to estimate the SARA fractional amounts.
The steps involved in getting our distillation data are as follows:
- We will first get the fraction of the oil that is inert. That is the resins and asphaltenes.
- We will then get the data points that represent the fractions of oil that are evaporated at associated temperatures.
- We will then try to normalize the inert and volatile fractions so that they measure up to the total fraction of oil.
### 3. Initial Resin and Asphaltene Fractions and Densities:
<i>
Reference: Fingas empirical formulas that are based upon analysis of ESTC oil properties database.
</i>
If the imported oil record contains valid values for resins and asphaltenes, we will use those values when building our oil.<br>
Otherwise, we will need to estimate them.
First we define some values $A$ and $B$ that will be used for our component fraction formulas:
$$
\begin{align}
\rho 0_{oil} &= \text{oil aggregate density at } 288.15^\circ K \text{ (from eqs. 1,2)} \cr
v0_{oil} &= \text{oil kinematic viscosity at } 288.15^\circ K \text{ (from eq. 5)} \cr
\cr
A &= 10 \cdot exp(0.001 \cdot \rho 0_{oil}) \qquad \qquad \qquad \qquad \boldsymbol{(eq. 6)} \cr
B &= 10 \cdot ln(1000 \cdot \rho 0_{oil} \cdot v0_{oil}) \qquad \qquad \qquad \boldsymbol{(eq. 7)}\cr
\end{align}
$$
Then our component fraction formulas:
$$
\begin{align}
f_{res} &= (0.033 \cdot A + 0.00087 \cdot B - 0.74)_{\bot 0}^{\top 1}
\qquad \qquad \qquad \qquad \qquad \boldsymbol{(eq. 8)} \cr
f_{asph} &= (0.000014 \cdot A^3 + 0.000004 \cdot B^2 - 0.18)_{\bot 0}^{\top (1 - f_{res})}
\qquad \qquad \qquad \boldsymbol{(eq. 9)} \cr
\end{align}
$$
```
def estimate_A_coeff(density):
return 10.0 * np.exp(0.001 * density)
def estimate_B_coeff(density, viscosity):
return 10.0 * np.log(1000.0 * density * viscosity)
def estimate_resin_fraction(density, viscosity, f_other=0.0):
A = estimate_A_coeff(density)
B = estimate_B_coeff(density, viscosity)
f_res = 0.033 * A + 0.00087 * B - 0.74
f_res = np.clip(f_res, 0.0, 1.0 - f_other)
return f_res
def estimate_asphaltene_fraction(density, viscosity, f_other=0.0):
A = estimate_A_coeff(density)
B = estimate_B_coeff(density, viscosity)
f_asph = (0.000014 * A ** 3.0 +
0.000004 * B ** 2.0 -
0.18)
f_asph = np.clip(f_asph, 0.0, 1.0 - f_other)
return f_asph
def oil_inert_fractions(imported_rec):
f_res, f_asph = imported_rec.resins, imported_rec.asphaltenes
if f_res is not None and f_asph is not None:
return f_res, f_asph
else:
density = oil_density_at_temp(imported_rec, 288.15)
viscosity = oil_kvis_at_temp(imported_rec, 288.15)
if f_res is None:
f_res = estimate_resin_fraction(density, viscosity)
if f_asph is None:
f_asph = estimate_asphaltene_fraction(density, viscosity,
f_res)
return f_res, f_asph
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.api = ', obj.api
print 'oil.resins = ', obj.resins
print 'oil.asphaltenes = ', obj.asphaltenes
obj_density = oil_density_at_temp(obj, 288.15)
obj_viscosity = oil_kvis_at_temp(obj, 288.15)
print 'oil density at temp = ', obj_density
print 'oil viscosity at temp = ', obj_viscosity
print 'f_res, f_asph = ',
print estimate_resin_fraction(obj_density, obj_viscosity),
print estimate_asphaltene_fraction(obj_density, obj_viscosity)
print 'f_res, f_asph = ', oil_inert_fractions(obj)
print 'sum of inert fractions = ', np.sum(oil_inert_fractions(obj))
print
```
### 4. Distillation Cut Boiling Point:
<i>
<b>Reference:</b> Adios2 and Jones R. (1997), A Simplified Pseudo-component Oil Evaporation Model, Proceedings of the 20th Arctic and Marine Oil Spill Program (AMOP), Vancouver, CA, Vol. 1, pp. 43-62
</i>
If the oil record contains distillation data, then we can use the distillation cut temperatures.<br>
Otherwise, if no distillation data exists, then we can estimate the distillation data from the oil's API.
This is done as follows:
First we estimate our lower and upper temperature bounds:
$$
\begin{align}
T_0 &= \text{the lower temperature boundary} \cr
&= 457 - 3.34 \cdot API \qquad \qquad \qquad \qquad \boldsymbol{(eq. 10)} \cr
T_G &= \text{the upper temperature boundary} \cr
&= 1357 - 247.7 \cdot ln(API) \qquad \qquad \qquad \boldsymbol{(eq. 11)} \cr
\end{align}
$$
Next, we would like to generate a set of $N$ temperatures associated with our cuts $T_i$. We will use a default of 5 cuts for this.
$$
\begin{align}
N &= 5 \cr
T_i &= T_0 + T_G \cdot {i \over N} \quad \{ i \in \Bbb Z \,\,|\,\, 0 \leq i < N \}
\qquad \boldsymbol{(eq. 12)} \cr
\end{align}
$$
```
def estimate_cut_temps_from_api(api, N=5):
T_0 = 457.0 - 3.34 * api
T_G = 1357.0 - 247.7 * np.log(api)
return np.array([(T_0 + T_G * i / N)
for i in range(N)])
def oil_cut_temps(imported_rec):
culled_cuts = imported_rec.cuts
if len(culled_cuts) > 0:
return np.array([c.vapor_temp_k
for c in culled_cuts])
elif imported_rec.api is not None:
return estimate_cut_temps_from_api(imported_rec.api)
else:
est_api = estimate_api_from_density(oil_density_at_temp(imported_rec, 288.15))
return estimate_cut_temps_from_api(est_api)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.api = ', obj.api
print 'oil.densities = ', obj.densities
print 'oil.cut.vapor_temp_k = ', [c.vapor_temp_k for c in obj.cuts]
print 'oil.cut.fraction = ', [c.fraction for c in obj.cuts]
print 'cut temperatures = ', oil_cut_temps(obj)
print
```
### 5. Distillation Cut Mass Fraction:
The distillation cuts are a series of temperatures and mass fractions. And each temperature $T_i$ is associated with a fractional value $fevap_i$ representing the portion of substance that was evaporated at that temperature.
For any temperature below about $530^\circ K$, we can assume that the portion of substance that was evaporated contains a mix of saturates and aromatics, and that no resins or asphaltenes have been released.
The fractional quantity that was evaporated at a distillation cut temperature, is what we would like to consider to be our distillation cut mass fractions.
<b>(A)</b> If the oil record contains at least two distillation cuts exist for which $T_i < 530^\circ K$, then we will use the supplied distillation cut fractional masses to generate $fmass_i$.
The distillation cut mass fractions that we get in the imported oil record are cumulative in nature, so it is necessary to collect the differences in the distillation cut mass fractions when generating $fmass_i$
$$
\begin{align}
fmass_i &= {d \over di} \left( fevap_i \right)
\qquad \{ i \in \Bbb Z \,\,|\,\, 0 \leq i \leq N \}
\qquad \qquad \boldsymbol{(eq. 13)} \cr
\end{align}
$$
<b>(B)</b> If distillation cut temperatures were generated by approximation (eq. 18), then it can be assumed that there were no measured mass fractions. For this we will evenly distribute our fractional masses such that for all distillation cuts:
$$
\begin{align}
fmass_i &= {(1 - (f_{res} + f_{asph})) \over N}
\qquad \qquad \boldsymbol{(eq. 14)} \cr
\end{align}
$$
```
def estimate_fmasses_from_cuts(f_evap_i):
fmass_i = np.array(f_evap_i)
fmass_i[1:] = np.diff(fmass_i)
return fmass_i
def estimate_n_fmasses(f_res, f_asph, N=5):
return np.array([(1.0 - f_res - f_asph) / N] * N)
def oil_fmasses(imported_rec):
f_res, f_asph = oil_inert_fractions(imported_rec)
culled_cuts = imported_rec.cuts
if len(culled_cuts) > 0:
fractions = [c.fraction for c in culled_cuts]
return estimate_fmasses_from_cuts(fractions)
else:
return estimate_n_fmasses(f_res, f_asph)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil cut temps = ', oil_cut_temps(obj)
fractions = [c.fraction for c in obj.cuts]
print 'oil.cuts.fractions = ', fractions
f_res, f_asph = oil_inert_fractions(obj)
print 'f_res, f_asph = ', f_res, f_asph
fmasses = estimate_fmasses_from_cuts(fractions)
print 'sum(fmasses) = ', np.sum(fmasses) + f_res + f_asph
fmasses = estimate_n_fmasses(f_res, f_asph)
print 'sum(N fmasses) = ', np.sum(fmasses) + f_res + f_asph
fmasses = oil_fmasses(obj)
print 'sum(oil_fmasses()) = ', np.sum(fmasses) + f_res + f_asph
print
```
### 16. Normalizing our Distillation Cut Data:
Because there are so many calculations that propagate from the distillation cuts, it is important that we interpret them in such a way that results in a reasonable, or at least plausible, distillation curve. And since we are dealing with records that were manually entered into a database, data entry errors need to be anticipated.
We expect the evaporated fractions to be cumulative with rising temperature. And if someone incorrectly entered a value when generating the imported oil record, then our cut data could fail to follow this cumulative trend, and our evaporation curve is thrown off. Some specific errors and their implications are:
- <b>Smaller fraction</b> entered than what it should have been: This will likely cause the fraction to be smaller than the previous one, and the $fmass_i$ value will go negative.
- <b>Bigger fraction</b> entered than what it should have been: This will likely cause the next fraction to be smaller, and the next $fmass_i$ value will go negative.
- <b>Smaller temperature</b> entered than what it should have been: Optimistically the series of $(T_i, fevap_i)$ could be reordered by temperature. But if we reorder a mistaken temperature, then the fraction will be mistakenly reordered as well, and the error will manifest itself in a similar way as a <b>bigger fraction</b>.
I don't think we can treat this in a naive way.
- We need to treat the temperatures and their associated fractions as atomic units of data
- Temperatures are required to be in ascending order. If the temperature of a particular cut is lower than the previous, we throw out the cut.
- Cut fractions might seem to be less consequential. This will happen only once per error in the series, and we could simply clip the value to be $\geq 0$. But this will have implications in that the sum of the differences will add up to something bigger than the total evaporated amount, which we don't want. It's probably better to just throw out the cut.
```
def oil_culled_cuts(imported_rec):
prev_temp = prev_fraction = 0.0
for c in imported_rec.cuts:
if c.vapor_temp_k < prev_temp:
continue
if c.fraction < prev_fraction:
continue
prev_temp = c.vapor_temp_k
prev_fraction = c.fraction
yield c
# we need to redefine our cut temperatures function
# to utilize our culled cuts instead of raw cuts.
def oil_cut_temps(imported_rec):
culled_cuts = list(oil_culled_cuts(imported_rec))
if len(culled_cuts) > 0:
return np.array([c.vapor_temp_k
for c in culled_cuts])
elif imported_rec.api is not None:
return estimate_cut_temps_from_api(imported_rec.api)
else:
est_api = estimate_api_from_density(oil_density_at_temp(imported_rec, 288.15))
return estimate_cut_temps_from_api(est_api)
# we need to redefine our cut fractions function
# to utilize our culled cuts instead of raw cuts.
def oil_fmasses(imported_rec):
f_res, f_asph = oil_inert_fractions(imported_rec)
culled_cuts = list(oil_culled_cuts(imported_rec))
if len(culled_cuts) > 0:
fractions = [c.fraction for c in culled_cuts]
return estimate_fmasses_from_cuts(f_res, f_asph,
fractions
)
else:
return estimate_n_fmasses(f_res, f_asph)
obj = ans_2002
plt.figure(1, figsize=(12,6))
plt.subplot(111)
plt.plot(*zip(*[(c.vapor_temp_k, c.fraction) for c in obj.cuts]) + ['ro'],
label=ans_2002.oil_name)
plt.plot(*zip(*[(c.vapor_temp_k, c.fraction) for c in obj.cuts]) + ['r-'])
plt.plot(*zip(*[(c.vapor_temp_k, c.fraction) for c in oil_culled_cuts(obj)]) + ['bo'],
label=ans_2002.oil_name + ' culled')
plt.plot(*zip(*[(c.vapor_temp_k, c.fraction) for c in oil_culled_cuts(obj)]) + ['b-'])
plt.xlabel(r'$T_i$')
plt.ylabel('% Evap')
plt.grid()
plt.title(r'Distillation Cuts Comparison', fontsize=18)
plt.legend(loc='upper left')
```
Not only do we seem to have typographic errors in the distillation cut data, but we have incomplete representation of distillation. The cut data of some oil records have a maximum distillation amount that is considerably less than the total oil, and the inert fractions, calculated or read from the record, do not account for a complete fraction (1.0) of oil mass.
I will suggest this for our distillation curves. I think we need to calculate the maximum amount we expect to be evaporated with our distillation cuts. Call it $fevap_{max}$. We can obtain this from the following equation:
$$
\begin{align}
fevap_{max} &= 1 - f_{res} - f_{asph} \cr
\end{align}
$$
Then we take the culled distillation cuts that exist, and use them to approximate a linear curve of our fractional distillation. I believe Bill, Robert, and Chris would agree that the distillation data approximates a linear curve of the form $(ax + b)_{\bot 0}^{\top 1}$. But just for fun, we will also try a sigmoid.
```
plt.figure(1, figsize=(12,6))
plt.subplot(111)
obj = ans_2002
boiling_points = [c.vapor_temp_k for c in oil_culled_cuts(obj)]
cum_fractions = [c.fraction for c in oil_culled_cuts(obj)]
plt.plot(boiling_points, cum_fractions, '.b', label="Cut Fractions")
plt.xlabel(r'$T_i$', fontsize=16)
plt.ylabel(r'$f_{evap}$', fontsize=16)
plt.grid()
#def func(x, a, b, c, d):
# return a*x**3 + b*x**2 +c*x + d
def sigmoid(x, j, k, lamb):
# intuitively, our cut fractions could approximate
# a similar shape as a cumulative distribution function
# or sigmoid.
return (1.0 - j ** (-1.0 * (x / lamb) ** k))
def linear(x, a, b):
# intuitively, our cut fractions could approximate
# a similar shape as a linear function.
return (a * x + b)
new_bps = np.linspace(100., 1200., 100)
"""
curve_fit the sigmoid
"""
if len(boiling_points) >= 3:
popt, pcov = curve_fit(sigmoid, boiling_points, cum_fractions)
plt.title(r'$f(T_i)= '
'1 - {0}^{{-(T_i / {2})^{{{1}}} }} + '
'$'.format(*popt),
fontsize=18)
# now plot our sigmoid curve with a bunch of boiling points from 273K to 1015K
plt.plot(new_bps, sigmoid(new_bps, *popt), '-r', label="Fitted Sigmoid Curve")
else:
print 'not enough data for a sigmoid!'
"""
curve_fit the linear
"""
popt, pcov = curve_fit(linear, boiling_points, cum_fractions)
# now plot our linear curve with a bunch of boiling points from 273K to 1015K
plt.plot(new_bps, np.clip(linear(new_bps, *popt), 0.0, 1.0), '-g', label="Fitted Linear Curve")
plt.legend(loc='upper left')
```
Okay, the linear curve seems to fit somewhat better, although it would be interesting to test this with a lot of oils. We'll go with the linear fit for now.
What we would like to do now is inverse the linear curve function so that it will calculate the temperature at which a certain fraction has boiled off. Then we can take a set of N fractions that represent equidistant intervals between 0.0 and $fevap_{max}$.
Our inverse linear function looks like this:
$$
\begin{align}
x &= {y - b \over a} \cr
\end{align}
$$
```
plt.figure(1, figsize=(12,6))
ax = plt.subplot(111)
obj = ans_2002
#print list(oil_culled_cuts(ans_2002))
num_cuts = 10
boiling_points = [c.vapor_temp_k for c in oil_culled_cuts(obj)]
cum_fractions = [c.fraction for c in oil_culled_cuts(obj)]
f_res, f_asph = oil_inert_fractions(obj)
'''
plot the cut points
'''
plt.plot(cum_fractions, boiling_points, 'ro', label="Measured Cuts")
plt.xlabel(r'$f_{evap,i}$', fontsize=18)
plt.ylabel(r'$T_i$', fontsize=18)
plt.grid()
def linear(x, a, b):
return (a * x + b)
def inverse_linear(y, a, b):
return (y - b) / a
"""
curve_fit the linear
"""
popt, pcov = curve_fit(linear, boiling_points, cum_fractions)
plt.title(r'$T_i = '
'(f_{{evap,i}} - {1}) / {0}'
'$'.format(*popt),
fontsize=18)
# now plot our inverse linear curve with a bunch of fractions from 0.0001 to 0.9999
new_fracs = np.linspace(0.0001, 0.9999, 100)
plt.plot(new_fracs, inverse_linear(new_fracs, *popt), '-c', label="Fitted Curve")
# now plot our inverse linear curve with the N fractions we intend to use
new_fracs = np.linspace(0.0, 1.0 - f_res - f_asph, num_cuts + 1)[1:]
plt.plot(new_fracs, inverse_linear(new_fracs, *popt), 'go', label="Estimated Cuts")
# make the shaded region
low, high = 1.0 - f_res - f_asph, 1.0
ix = (low, high)
iy = (1000.0, 1000.0)
verts = [(low,0)] + list(zip(ix,iy)) + [(high,0)]
poly = Polygon(verts, facecolor='0.8', edgecolor='k')
ax.add_patch(poly)
# ticks for the shaded region
extraticks = [low,]
extralabels = [r'${f_{inert}}$',]
plt.xticks(list(plt.xticks()[0]) + extraticks,
list(plt.xticks()[0]) + extralabels)
for tick in ax.xaxis.get_major_ticks():
if tick.label.get_text().find('inert') >= 0:
tick.label.set_fontsize(14)
tick.label.set_rotation('vertical')
plt.legend(loc='upper left')
```
Alright, that doesn't look too bad, but we are still missing something.
First, it is possible for us to fit a curve that will give negative kelvin temperatures. Negative Kelvin temperatures, are of course impossible. We need to make sure our curve fit anticipates that.
What we have works ok for total evaporated amounts, and differencing them should give us reasonable values for $fmass_i$. But the temperatures we get from our curve represent only the maximum slice of distilled components within $fmass_i$. What we really need is the average temperature from $T_{i-1}$ to $T_i$
```
plt.figure(1, figsize=(12,6))
ax = plt.subplot(111)
obj = ans_2002
#print list(oil_culled_cuts(ans_2002))
num_cuts = 10
boiling_points = [c.vapor_temp_k for c in oil_culled_cuts(obj)]
cum_fractions = [c.fraction for c in oil_culled_cuts(obj)]
f_res, f_asph = oil_inert_fractions(obj)
print 'boiling_points:', boiling_points
print 'cum_fractions:', cum_fractions
print 'inert_fractions:', f_res, f_asph
plt.xlabel('% Evap')
plt.ylabel(r'$T_i$', fontsize=16)
plt.grid()
def linear(x, a, b):
return (a * x + b)
def inverse_linear(y, a, b):
return (y - b) / a
"""
curve_fit the linear
"""
popt, pcov = curve_fit(linear, boiling_points, cum_fractions)
# plot our distillation curve
new_fracs = np.linspace(0.0001, 0.9999, 100)
new_temps = np.clip(inverse_linear(new_fracs, *popt), 0.0, 5000.0)
plt.plot(new_fracs, new_temps, '-b', label="Distillation Curve")
# now plot the N fractions we intend to use
new_fracs = np.linspace(0.0, 1.0 - f_res - f_asph, (num_cuts * 2) + 1)[1:]
new_temps = inverse_linear(new_fracs, *popt)
new_fracs = new_fracs.reshape(-1,2)[:,1]
new_temps = new_temps.reshape(-1,2)[:,0]
above_zero = new_temps > 0.0
new_temps = new_temps[above_zero]
new_fracs = new_fracs[above_zero]
plt.plot(new_fracs, new_temps, 'go', label="Estimated Cuts")
# plot our inert fraction
plt.plot(1.0, 1015.0, 'go')
# now we would like to graph the quantized "areas"
# that our fractional masses represent
def make_shaded_region(f_low, f_high,
t_low, t_high,
facecolor='0.8'):
ix = (f_low, f_high)
iy = (t_low, t_high)
verts = [(x, y) for x in ix for y in iy]
verts[2:] = reversed(verts[2:])
poly = Polygon(verts, facecolor=facecolor, edgecolor='k')
ax.add_patch(poly)
for r in zip([0.0] + new_fracs.tolist(),
new_fracs,
[inverse_linear(0.0, *popt)] + new_temps.tolist(),
new_temps):
make_shaded_region(*r, facecolor='#ddeeff')
# make the shaded region for our inert fraction
make_shaded_region(new_fracs[-1], 1.0,
new_temps[-1], 1015.0)
# ticks for the shaded region
extraticks = [low,]
extralabels = [r'${f_{inert}}$',]
plt.xticks(list(plt.xticks()[0]) + extraticks,
list(plt.xticks()[0]) + extralabels)
for tick in ax.xaxis.get_major_ticks():
if tick.label.get_text().find('inert') >= 0:
tick.label.set_fontsize(16)
tick.label.set_rotation('vertical')
plt.legend(loc='upper left')
```
Alright, **now** this looks like a reasonable quantization of our distillation curve.
putting it all together in code...
```
def linear_curve(x, a, b):
return (a * x + b)
def inverse_linear_curve(y, a, b):
return (y - b) / a
def oil_normalized_cut_values(imported_rec, num_cuts=10):
f_res, f_asph = oil_inert_fractions(imported_rec)
culled_cuts = list(oil_culled_cuts(imported_rec))
if len(culled_cuts) == 0:
if imported_rec.api is not None:
oil_api = imported_rec.api
else:
oil_rho = oil_density_at_temp(imported_rec, 288.15)
oil_api = estimate_api_from_density(oil_rho)
BP_i = estimate_cut_temps_from_api(oil_api)
fevap_i = np.cumsum(estimate_n_fmasses(f_res, f_asph))
else:
BP_i, fevap_i = zip(*[(c.vapor_temp_k, c.fraction)
for c in culled_cuts])
popt, pcov = curve_fit(linear_curve, BP_i, fevap_i)
fevap_i = np.linspace(0.0, 1.0 - f_res - f_asph, (num_cuts * 2) + 1)[1:]
T_i = np.clip(inverse_linear(fevap_i, *popt), 0.0, 5000.0)
fevap_i = fevap_i.reshape(-1,2)[:,1]
T_i = T_i.reshape(-1,2)[:,0]
above_zero = T_i > 0.0
T_i = T_i[above_zero]
fevap_i = fevap_i[above_zero]
return T_i, fevap_i
def oil_normalized_cut_temps(imported_rec, num_cuts=10):
cut_temps, _f_evap_i = oil_normalized_cut_values(imported_rec)
return cut_temps
def oil_normalized_cut_fmasses(imported_rec, num_cuts=10):
_cut_temps, f_evap_i = oil_normalized_cut_values(imported_rec)
return estimate_fmasses_from_cuts(f_evap_i)
def oil_normalized_cut_temps_fmasses(imported_rec, num_cuts=10):
cut_temps, f_evap_i = oil_normalized_cut_values(imported_rec)
return cut_temps, estimate_fmasses_from_cuts(f_evap_i)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.api = ', obj.api
print 'oil cut temps = ', oil_cut_temps(obj)
fractions = [c.fraction for c in oil_culled_cuts(obj)]
print 'oil.cuts.fractions = ', fractions
f_res, f_asph = oil_inert_fractions(obj)
print 'f_res, f_asph = ', f_res, f_asph
if len(fractions) > 0:
print ('sum of all non-normalized fractions = {}'
.format(fractions[-1] + f_res + f_asph))
else:
print 'No measured fractions found.'
print ('normalized fmasses = {}'
.format(oil_normalized_cut_fmasses(obj) + f_res + f_asph))
print ('sum of all normalized fmasses = {}'
.format(np.sum(oil_normalized_cut_fmasses(obj)) + f_res + f_asph))
print
```
## <u>Oil Component Fractional Properties</u>
Once we have estimated the distilled and inert fractions of our oil, we can then begin to break it down even further, into SARA component fractional properties. This will include:
- Component Molecular Weight
- Component Density
- Component Specific Gravity <i>(basically a representation of density)</i>
- Saturate and Aromatic Mass Fractions <i>(we have already estimated our resins & asphaltenes, and they will not change)</i>
We will organize our components in arrays that are consistently ordered so that all properties of any particular component will share a common index. The ordering will be as follows:
$$
\begin{align}
[
&C_{sat,0} \, C_{arom,0} \cr
&C_{sat,1} \, C_{arom,1} \cr
&C_{sat,2} \, C_{arom,2} \cr
&... \cr
&C_{sat,N} \, C_{arom,N} \cr
&C_{res} \, C_{asph} ] \cr
\end{align}
$$
### 17. Component Temperatures:
Here we will define a temperature for each of our components. This is a fairly straight forward copy of the cut temperatures in regards to the saturate and aromatic components.
But our resins and asphaltenes need a temperature too. We will choose a default value of $1015^\circ K$ for them.
```
def oil_component_temps(imported_rec):
cut_temps = oil_normalized_cut_temps(imported_rec)
component_temps = np.append([1015.0, 1015.0],
zip(cut_temps, cut_temps))
return np.roll(component_temps, -2)
```
### 18. Initial Component Molecular Weight:
<i>
Reference: CPPF eq. 2.48 and table 2.6
</i>
The saturate and aromatic component molecular weights can be calculated using the distillation cut temperatures as follows:
$$
\begin{align}
M_{w,sat,i} &= \left( 49.677 \cdot \left[ 6.98291 - ln(1070 - T_i) \right] \right)^{3/2}
\qquad \qquad \qquad \boldsymbol{(eq. 15)} \cr
M_{w,arom,i} &= \left( 44.504 \cdot \left[ 6.911 - ln(1015 - T_i) \right] \right)^{3/2}
\qquad \qquad \qquad \boldsymbol{(eq. 16)} \cr
\end{align}
$$
So that we have a full complement of molecular weights for <u>all</u> components, We need some estimation method for our resins and asphaltenes. Bill recommends using the following average values:
$$
\begin{align}
M_{w,res} &= \text{molecular weight of our resin component} \cr
&= 800 \, g/mol \text{ at } 1015^\circ K \cr
M_{w,asph} &= \text{molecular weight of our asphaltene component} \cr
&= 1000 \, g/mol \text{ at } 1015^\circ K \cr
\end{align}
$$
```
def estimate_saturate_mol_wt(boiling_point):
T_i = np.array(boiling_point)
return (49.677 * (6.98291 - np.log(1070.0 - T_i))) ** (3.0 / 2.0)
def estimate_aromatic_mol_wt(boiling_point):
T_i = np.array(boiling_point)
return (44.504 * (6.911 - np.log(1015.0 - T_i))) ** (3.0 / 2.0)
def estimate_resin_mol_wt():
return 800.0
def estimate_asphaltene_mol_wt():
return 1000.0
def estimate_component_mol_wt(boiling_points):
rho_list = np.append([estimate_resin_mol_wt(),
estimate_asphaltene_mol_wt()],
zip(estimate_saturate_mol_wt(boiling_points),
estimate_aromatic_mol_wt(boiling_points)))
return np.roll(rho_list, -2)
def oil_component_mol_wt(imported_rec):
cut_temps = oil_normalized_cut_temps(imported_rec)
return estimate_component_mol_wt(cut_temps)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'component mol_wt = ', oil_component_mol_wt(obj).reshape(-1,2)
print
```
### 19. Component Density:
<i>
Reference: CPPF eq. 2.13 and table 9.6
</i>
Initial density estimate for both resin and asphaltene fractional componens is set to:
$$
\begin{align}
\rho_{res} &= \rho_{asph} = 1100 \, kg/m^3 \cr
\end{align}
$$
Then we define Watson characterization factors for saturates and aromatics:
$$
\begin{align}
K_{arom,w} &= \text{Watson characterization factor for aromatics} \cr
&= 10 \cr
K_{sat,w} &= \text{Watson characterization factor for saturates} \cr
&= 12 \cr
\end{align}
$$
Then we apply the appropriate Watson characterization to calculate an initial trial estimate for each density component:
$$
\begin{align}
T_i &= \text{component boiling point } (\,^\circ K) \cr
\cr
\rho try_{arom,i} &= 1000 \cdot {\root 3 \of { 1.8 \cdot T_i} \over K_{arom,w} }
\qquad \qquad \boldsymbol{(eq. 17)} \cr
\rho try_{sat,i} &= 1000 \cdot {\root 3 \of { 1.8 \cdot T_i} \over K_{sat,w} }
\qquad \qquad \boldsymbol{(eq. 18)} \cr
\end{align}
$$
This should be a reasonable estimate. However, the average density (fractionally weighted average of the component densities) must match the measured value or the value from approximation (eqs. 1, 2).<br>
To find the scaling factor between our densities and the aggregate density, we do the following:
$$
\begin{align}
j &= \text{index representing all oil components } \{ j \in \Bbb Z \} \cr
jMAX &= \text{index of the last oil component} \cr
&= 2N + 2 \cr
fmass0_j &= \text{the fractional mass of our jth component} \cr
\cr
Cf_{dens} &= { \rho 0_{oil} \over \sum_{j=1}^{jMAX} fmass0_j \cdot \rho try_j}
\qquad \qquad \boldsymbol{(eq. 19)} \cr
\end{align}
$$
Then we evenly apply this scaling factor to our trial densities to come up with densities that are consistent with the aggregate density.
$$
\begin{align}
\rho_j = Cf_{dens} \cdot \rho try_j \qquad \qquad \boldsymbol{(eq. 20)} \cr
\end{align}
$$
<center><b>--- Begin JamesM Comments ---</b></center>
Well, I don't know about the resin & asphaltene densities. It seems that at the higher temperatures, the Aromatic densities get slightly denser than the inert components. This doesn't seem right.
<center><b>--- End JamesM Comments ---</b></center>
```
def estimate_trial_densities(boiling_points, watson_factor):
return 1000.0 * (1.8 * boiling_points) ** (1.0 / 3.0) / watson_factor
def estimate_saturate_densities(boiling_points):
return estimate_trial_densities(boiling_points, 12)
def estimate_aromatic_densities(boiling_points):
return estimate_trial_densities(boiling_points, 10)
def estimate_resin_density():
return 1100.0
def estimate_asphaltene_density():
return 1100.0
def estimate_component_densities(boiling_points):
rho_list = np.append([estimate_resin_density(),
estimate_asphaltene_density()],
zip(estimate_saturate_densities(boiling_points),
estimate_aromatic_densities(boiling_points)))
return np.roll(rho_list, -2)
def oil_component_densities(imported_rec):
cut_temps = oil_normalized_cut_temps(imported_rec)
return estimate_component_densities(cut_temps)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'component densities from oil = ', oil_component_densities(obj).reshape(-1, 2)
print
```
### 20. Component Specific Gravity:
Specific Gravity of a substance is the ratio of density of the substance to the density of another substance (typically water) at a specified temperature.<br>
<i>
Reference: https://en.wikipedia.org/wiki/Specific_gravity
</i>
For simplicity, we will make the assumption that our water density is $1000 \, kg/m^3$. This is water's density at $4^\circ C$, Which is the maximum density it can achieve.<br>
So our estimation of the specific gravity of our oil components will be:
$$
\begin{align}
SG_j &= \rho_j / 1000 \qquad \qquad \boldsymbol{(eq. 21)} \cr
\end{align}
$$
```
def estimate_specific_gravity(rho_kg_m_3):
return rho_kg_m_3 / 1000.0
def oil_component_specific_gravity(imported_rec):
rho_list = oil_component_densities(imported_rec)
return estimate_specific_gravity(rho_list)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'specific gravity = ', oil_component_specific_gravity(obj).reshape(-1, 2)
print
```
### 21. Initial Saturate and Aromatic Mass Fractions:
<i>
References:
<ul>
<li>CPPF eqs. 2.114, 2.115, 3.77 and 3.78</li>
<li>Huang, E K., Characterization and Thermodynamic Correlations for Undefined Hydrocarbon Mixtures, Ph.D. Dissertation, Pennsylvania State University, University Park, PA, 1977</li>
</ul>
</i>
We start by computing our saturate mass fractions. This is done using the distillation cuts, and the specific gravity, and molecular weight of our saturate components.
Using Riazi, we perform a number of intermediate calculations to finally come up with the estimation for saturate component fractional mass.
First is a dimensionless parameter $I$, that was first used by Huang to correlate hydrocarbon properties. This is necessary to calculate the saturate refractive index $n$.<br>
Next is the calculation of refractive index $n$. This is necessary to calculate $m$.
The next intermediate calculation $m$ can be best described as a hydrocarbon grouping parameter, Riazi describes $m$ as a parameter that "not only separates paraffins and aromatics but also identifies various hydrocarbon types", and is (AFAIK) based on an observed correlation of refractive index and Molecular weight.
$$
\begin{align}
T_i &= \text{distillation cut boiling point} \cr
M_{w,sat,i} &= \text{molecular weight of our saturate component} \cr
SG_{sat, i} &= \text{specific gravity of saturate component} \cr
\cr
I &= \text{hydrocarbon characterization parameter} \cr
&= 0.3773 T_i^{-0.02269} SG_{sat,i}^{0.9182} \qquad \qquad \boldsymbol{(eq. 22)} \cr
\cr
n &= \text{refractive index of our saturate component at } 20^\circ C \cr
&= \left( 1 + 2 I \over 1 - I \right)^{1/2} \qquad \qquad \qquad \qquad \boldsymbol{(eq. 23)} \cr
\cr
m &= \text{hydrocarbon grouping parameter} \cr
&= M_{w,sat,i}(n - 1.475) \qquad \qquad \qquad \boldsymbol{(eq. 24)} \cr
\end{align}
$$
And finally, after these intermediate calculations are completed, we can estimate the saturate component fractional mass.
$$
\begin{align}
fmass_i &= \text{distillation cut fractional mass} \cr
\cr
f_{sat,i} &= (fmass_i \cdot (2.24 - 1.98 \cdot SG_{sat,i} - 0.009 \cdot m))_{\bot 0}^{\top fmass_i}
\qquad \qquad \boldsymbol{(eq. 25)} \cr
\end{align}
$$
After our saturate mass fraction has been computed, we compute our aromatic mass fraction simply as:
$$
\begin{align}
f_{arom,i} &= fmass_i - f_{sat,i} \qquad \qquad \qquad \boldsymbol{(eq. 26)} \cr
\end{align}
$$
```
def hydrocarbon_characterization_param(specific_gravity, temp_k):
T_i = temp_k
SG_i = specific_gravity
return 0.3773 * (T_i ** -0.02269) * (SG_i ** 0.9182)
def refractive_index(hc_char_param):
I = hc_char_param
return ((1 + 2 * I) / (1 - I)) ** (1.0 / 2.0)
def hydrocarbon_grouping_param(mol_wt, specific_gravity, temp_k):
I = hydrocarbon_characterization_param(specific_gravity, temp_k)
n = refractive_index(I)
return mol_wt * (n - 1.475)
def estimate_saturate_mass_fraction(fmass_i, mol_wt, specific_gravity, temp_k):
SG_sat_i = specific_gravity
m = hydrocarbon_grouping_param(mol_wt, SG_sat_i, temp_k)
X_P = 3.7387 - 4.0829 * SG_sat_i + 0.014772 * m
X_N = -1.5027 + 2.10152 * SG_sat_i - 0.02388 * m
f_sat_i = fmass_i * (X_P + X_N)
f_sat_i = np.clip(f_sat_i, 0.0, fmass_i)
return f_sat_i
def oil_component_mass_fractions(imported_rec):
f_res, f_asph = oil_inert_fractions(imported_rec)
cut_temps, fmass_i = oil_normalized_cut_temps_fmasses(imported_rec)
sat_mask = np.array(range(len(cut_temps))) * 2
sat_temps = oil_component_temps(imported_rec)[sat_mask]
sat_mol_wts = estimate_component_mol_wt(cut_temps)[sat_mask]
sat_rhos = estimate_component_densities(cut_temps)[sat_mask]
sat_SGs = estimate_specific_gravity(sat_rhos)
f_sat_i = estimate_saturate_mass_fraction(fmass_i,
sat_mol_wts,
sat_SGs,
sat_temps)
f_arom_i = fmass_i - f_sat_i
f_all = [n for l in zip(f_sat_i, f_arom_i) for n in l] + [f_res, f_asph]
return np.array(f_all)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.cuts.fractions = ', [c.fraction for c in obj.cuts]
print 'oil.cuts.vapor_temp_k = ', [c.vapor_temp_k for c in obj.cuts]
print 'component mass fractions = ', oil_component_mass_fractions(obj).reshape(-1, 2)
print 'sum(component mass fractions) = ', np.sum(oil_component_mass_fractions(obj))
print
```
<center><b>--- Begin JamesM Comments ---</b></center>
After staring at Riazi's equations for awhile and reading the surrounding descriptions, I am not convinced that we should be using $M_{w,sat,i}$ and $SG_{sat,i}$ for computing the saturate fractional mass. Instead, I think it is possible that we should use the average molecular weight and specific gravity of the distillation cut.<br>
That is, if we can determine it.
$$
\begin{align}
fmass_i &= \text{distillation cut fractional mass} \cr
T_i &= \text{distillation cut boiling point} \cr
M_{w,i} &= \text{distillation cut molecular weight} \cr
SG_i &= \text{distillation cut specific gravity} \cr
\cr
I &= 0.3773 T_i^{-0.02269} SG_i^{0.9182} \cr
\cr
n &= \left( 1 + 2 I \over 1 - I \right)^{1/2} \cr
\cr
m &= M_{w,i}(n - 1.475) \cr
\cr
f_{sat,i} &= (fmass_i \cdot (2.24 - 1.98 \cdot SG_i - 0.009 \cdot m))_{\bot 0}^{\top fmass_i} \cr
\end{align}
$$
I am not sure, but it might be possible to do the following:
- Start by assuming a 50/50 mix of saturates and aromatics for a particular cut
- Estimate the aggregate molecular weight and specific gravity of our mix
- Figure out the saturate and aromatic fractions using the Riazi correlation
- Repeat using the newly calculated mix until the mix converges (will it converge?)
```
def verify_cut_fractional_masses(fmass_i, T_i,
f_sat_i, f_arom_i):
assert np.all(fmass_i == f_sat_i + f_arom_i)
M_w_sat_i = estimate_saturate_mol_wt(T_i)
M_w_arom_i = estimate_aromatic_mol_wt(T_i)
M_w_avg_i = (M_w_sat_i * f_sat_i / fmass_i +
M_w_arom_i * f_arom_i / fmass_i)
# estimate specific gravity
rho_sat_i = estimate_saturate_densities(T_i)
SG_sat_i = estimate_specific_gravity(rho_sat_i)
rho_arom_i = estimate_aromatic_densities(T_i)
SG_arom_i = estimate_specific_gravity(rho_arom_i)
SG_avg_i = (SG_sat_i * f_sat_i / fmass_i +
SG_arom_i * f_arom_i / fmass_i)
f_sat_i = estimate_saturate_mass_fraction(fmass_i,
M_w_avg_i,
SG_avg_i,
T_i)
f_arom_i = fmass_i - f_sat_i
# Riazi states that CPPF eqs. 3.77 and 3.78 only work with
# molecular weights less than 200. So we will punt
# with Bill's recommendation of 50/50 in those cases.
# In the future we might be able to figure out how
# to implement CPPF eqs. 3.81 and 3.82, which work with
# the molecular weights above 200.
above_200 = M_w_avg_i > 200.0
try:
f_sat_i[above_200] = fmass_i[above_200] / 2.0
f_arom_i[above_200] = fmass_i[above_200] / 2.0
except TypeError:
# numpy array assignment failed, try a scalar assignment
if above_200:
f_sat_i = fmass_i / 2.0
f_arom_i = fmass_i / 2.0
return f_sat_i, f_arom_i
fmass_i, f_sat_i, f_arom_i = 1.0, 0.5, 0.5
print f_sat_i, f_arom_i
for i in range(20):
f_sat_i, f_arom_i = verify_cut_fractional_masses(fmass_i, 313.15,
f_sat_i, f_arom_i)
print f_sat_i, f_arom_i
```
Well, it does seem to converge, at least with the example above. Let's try plotting an oil record.
```
plt.figure(1, figsize=(12,6))
plt.subplot(111)
obj = ans_mp
plt.xlabel(r'$T_i$', fontsize=16)
plt.ylabel(r'$f_{sat,i}$', fontsize=16)
plt.grid()
plt.title(r'Convergence of $f_{{sat,i}}$ for oil {}'.format(obj.oil_name),
fontsize=18)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
cut_temps, fmass_i = oil_normalized_cut_temps_fmasses(obj)
f_sat_i = fmass_i / 2.0
f_arom_i = fmass_i / 2.0
label=r'Initial $f_{sat,i}$'
color, linestyle, marker = 'b', 'None', 'o'
plt.plot(cut_temps, f_sat_i,
color=color, linestyle=linestyle, marker=marker, label=label)
for i in range(20):
color, linestyle, marker = 'b', 'None', '.'
label=''
f_sat_i, f_arom_i = verify_cut_fractional_masses(fmass_i, cut_temps,
f_sat_i, f_arom_i)
plt.plot(cut_temps, f_sat_i,
color=color, linestyle=linestyle, marker=marker, label=label)
color, linestyle, marker = '#00ff00', 'None', 'o'
label=r'Final $f_{sat,i}$'
plt.plot(cut_temps, f_sat_i,
color=color, linestyle=linestyle, marker=marker, label=label)
color, linestyle, marker = '#00ff00', '-', 'o'
label=''
plt.plot(cut_temps, f_sat_i,
color=color, linestyle=linestyle, marker=marker, label=label)
plt.legend(loc='upper right')
```
Well, our saturate fractions do seem to converge into values that indicate the following:
- There is an inversely proportional relationship with boiling point. The trend looks more-or-less linear.
- The temperature at which we achieve a 50/50 mix of saturates and aromatics appears to be somewhere around $470^\circ K$.
- The temperature at which we have lost all saturates is somewhere between $600^\circ K$ and $630^\circ K$.
But I can't be completely sure if this is how our saturates are supposed to behave. For one thing, this would also indicate that the fraction of aromatics in our cuts would proportionally increase until about $600^\circ K$ or so, where the distilled fractions become 100% aromatics. Again, I can't be sure if this is how aromatics are supposed to behave.
I think I will submit this line of thinking to Bill & Co. before actually putting this into our estimations code.
<center><b>--- End JamesM Comments ---</b></center>
## <u>Oil Miscellaneous Properties</u>
These are oil properties that can be computed based on the previous computations, and in some cases are simply assigned a default value.
### 22. Oil-Water Surface Tension:
If the imported oil record does not contain a value for surface tension, then we will estimate it from the oil's API:
$$
\begin{align}
\sigma_{o-w} &= \text{oil/water surface tension at } 288.15^\circ K \,\, (N/m) \cr
&= 0.001 \cdot (39 - 0.2571 \cdot API) \qquad \qquad \qquad \qquad \boldsymbol{(eq. 27)} \cr
\end{align}
$$
<i>
Reference: Baker, O. and W. Swerdloff (1956), Calculation of Surface Tensions - Finding the Surface Tension of Hydrocarbon Liquids, Oil Gas J. (2 January 1956) pp. 125
</i>
```
def o_w_surface_tension_from_api(api):
if api is not None:
return 0.001 * (39.0 - 0.2571 * api)
else:
return None
def oil_o_w_surface_tension(imported_rec):
if (imported_rec.oil_water_interfacial_tension_n_m is not None and
imported_rec.oil_water_interfacial_tension_ref_temp_k is not None):
ow_st = imported_rec.oil_water_interfacial_tension_n_m
ref_temp_k = imported_rec.oil_water_interfacial_tension_ref_temp_k
return ow_st, ref_temp_k
elif imported_rec.api is not None:
ow_st = oil_water_surface_tension_from_api(imported_rec.api)
return ow_st, 273.15 + 15
else:
est_api = api_from_density(oil_density_at_temp(imported_rec, 288.15))
ow_st = oil_water_surface_tension_from_api(est_api)
return ow_st, 273.15 + 15
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil/water surface tension = ', o_w_surface_tension_from_api(obj.api)
print 'oil/water surface tension = ', oil_o_w_surface_tension(obj)
print
```
### 23. Pour Point:
If the imported oil record contains a pour point property then we will simply use it when building the final oil record.<br>
Otherwise, if we have measured molecular weights for the distillation fractions (unusual) then use method <b>(A)</b>.<br>
Otherwise, use method <b>(B)</b>
<b>(A)</b> If molecular weight and mass fractions are given for all the oil fractions $j = 1 \dots jMAX$, then an average molecular weight for the whole oil $\bar M_w$ can be estimated as:
$$
\begin{align}
N &= \text{number of distillation cuts} \cr
jMAX &= 2 (N + 1) \cr
M_{w,j} &= \text{molecular weight of component j} (kg/kmole)\cr
fmass_j &= \text{mass fraction of component j} \cr
\cr
\bar M_w &= \sum_1^{jMAX} M_{w,j} \cdot fmass_j \qquad \qquad \qquad \qquad \boldsymbol{(eq. 28)} \cr
\end{align}
$$
<i>(Note: The calculation for $jMAX$ may seem counterintuitive. It simply states that we sum over all the SARA fractions, each distillation cut represents 1 saturate and 1 aromatic fraction, and that resins and asphaltenes do not have distillation cut data. So for $N$ distillation cuts, we would calculate $2 N + 2 \rightarrow 2(N + 1)$)</i>
Define $SG = \rho_{oil} / 1000 \cdot kg$ as specific gravity
Then, using CPPF eq. 3.119, our pour point temperature is calculated as:
$$
\begin{align}
T_{API} &= \text{reference temperature for oil kinematic viscosity} \cr
&= 311^\circ K \cr
\cr
T_{pp} &= 130.47 SG^{2.97} \cdot \bar M_w^{0.61235 - 0.47357 SG} \cdot v_{oil}^{0.31 - 0.3283 SG} \cdot T_{API} \qquad \qquad \boldsymbol{(eq. 29)} \cr
\end{align}
$$
<b>(B)</b> Pour point is estimated by reversing the viscosity-temperature correction in Adios2 and assuming that, at the pour point, viscosity is equal to 1 million centistokes.
$$
\begin{align}
c_{v1} &= 5000 K \cr
\cr
T_{pp} &= { c_{v1} \cdot T_{ref} \over c_{v1} - T_{ref} ln(v_{ref}) }
\qquad \qquad \qquad \qquad \boldsymbol{(eq. 30)} \cr
\end{align}
$$
<center><b>--- Begin JamesM Comments ---</b></center>
In <b>(A)</b>, we reference CPPF eq. 3.119, which seems to not include $T_{API}$ as an input. Riazi describes this equation as:
$$
\begin{align}
SG &= \text{oil specific gravity} \cr
M &= \text{oil molecular weight} \cr
v_{38(100)} &= \text{oil kinematic viscosity at } 37.8^\circ C (100^\circ F) \cr
\cr
T_P &= \text{pour point (ASTM D 97) in } \,^\circ K \cr
&= 130.47 [SG^{2.970566}] \cdot [M^{(0.61235 - 0.47357 SG)}] \cdot [v_{38(100)}^{(0.310331 - 0.32834 SG)}] \cr
\end{align}
$$
So I think that eq. 6 is not correct.
Beyond that, I think that we would only use this formula if we had actual measured component densities in our imported oil record, which is very unlikely.
The component molecular weights and densities that we estimate already use viscosity as an input, at least in part. So I am not convinced that an estimation based on them would be any more accurate than <b>(B)</b>.
In <b>(B)</b>, exactly which $(T_{ref}, v_{ref})$ measurement should we use if multiple viscosity measurements exist? Since we are calculating pour point, which I understand to mean the lowest temperature for which a finite viscosity exists, I believe we should use the lowest measured temperature and its associated viscosity.
Does the viscosity-temperature correction formula in Adios2 define what $c_{v1}$ is?
<center><b>--- End JamesM Comments ---</b></center>
```
def estimate_pour_point_from_kvis(ref_kvis, ref_temp_k):
c_v1 = 5000.0
T_pp = (c_v1 * ref_temp_k) / (c_v1 - ref_temp_k * np.log(ref_kvis))
return T_pp
def oil_pour_point(imported_rec):
min_k = max_k = None
if (imported_rec.pour_point_min_k is not None or
imported_rec.pour_point_max_k is not None):
min_k = imported_rec.pour_point_min_k
max_k = imported_rec.pour_point_max_k
else:
lowest_kvis = lowest_temperature(oil_aggregate_kvis(imported_rec)[0])
max_k = pour_point_from_kvis(lowest_kvis.m_2_s, lowest_kvis.ref_temp_k)
return min_k, max_k
# pour point estimation based on (T_ref, v_ref)
# seems a bit inaccurate
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.pour_point_min = ', obj.pour_point_min_k
print 'oil.pour_point_max = ', obj.pour_point_max_k
lowest_kvis = lowest_temperature(oil_aggregate_kvis(obj))
ref_kvis, ref_temp_k = lowest_kvis.m_2_s, lowest_kvis.ref_temp_k
print 'pour point (T_ref, v_ref) = ', estimate_pour_point_from_kvis(ref_kvis, ref_temp_k)
print 'oil_pour_point() = ', oil_pour_point(obj)
print
```
### 24. Flash Point:
If a measured minimum flash point exists for the incoming record, we will use it.<br>
Otherwise, if a measured maximum flash point exists for the incoming record, we will use that instead.<br>
Otherwise, if measured distillation cut data exists, use method <b>(A)</b>.<br>
Otherwise, use method <b>(B)</b>.
<b>(A)</b> Flash point can be estimated from the first pseudo-component cut:
$$
\begin{align}
T_{cut1} &= \text{the boiling point of the first pseudo-component cut } (^\circ K) \cr
\cr
T_{flsh} &= 117 + 0.69 \cdot T_{cut1} \qquad \qquad \qquad \qquad \boldsymbol{(eq. 31)} \cr
\end{align}
$$
<i>
Reference: Chang A., K. Pashakanti, and Y. Liu (2012), Integrated Process Modeling and Optimization, Wiley Verlag.
</i>
<b>(B)</b> Flash point can be estimated from the API value:
$$
\begin{align}
T_{flsh} &= 457 - 3.34 \cdot API \qquad \qquad \qquad \qquad \boldsymbol{(eq. 32)} \cr
\end{align}
$$
<center><b>--- Begin JamesM Comments ---</b></center>
I have some reservations about simply using the first distillation cut boiling point.<br>
It seems that the results could vary wildly depending on the quality of the cut data.
I would be much more confident in a curve fit to the cuts, in which we take the temperature where a certain fraction is evaporated.
<center><b>--- End JamesM Comments ---</b></center>
```
def estimate_flash_point_from_bp(ref_temp_k):
return 117.0 + 0.69 * ref_temp_k
def estimate_flash_point_from_api(api):
return 457.0 - 3.34 * api
def oil_flash_point(imported_rec):
if imported_rec.flash_point_min_k is not None:
return imported_rec.flash_point_min_k
elif imported_rec.flash_point_max_k is not None:
return imported_rec.flash_point_max_k
elif len(list(oil_culled_cuts(imported_rec))) > 2:
cut_temps = oil_normalized_cut_temps(imported_rec)
return estimate_flash_point_from_bp(cut_temps[0])
elif imported_rec.api is not None:
return estimate_flash_point_from_api(imported_rec.api)
else:
est_api = estimate_api_from_density(oil_density_at_temp(imported_rec, 288.15))
return estimate_flash_point_from_api(est_api)
# ANS oil has both an api and distillation cuts.
# For this oil, methods (A) and (B) estimate different results.
# but the difference is only about 7%. Probably acceptable.
# we should try this on other oils to compare.
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.flash_point_min_k = ', obj.flash_point_min_k
print 'oil.flash_point_max_k = ', obj.flash_point_max_k
if len(list(oil_culled_cuts(obj))) > 0:
print 'oil.cuts[0] = ', oil_normalized_cut_values(obj)[0][0]
else:
print 'no cut data'
print 'oil.api = ', obj.api
print 'oil flash point = ', oil_flash_point(obj)
print
```
### 25. Maximum water fraction of emulsion:
This quantity will be set after the emulsification approach in Adios3 is finalized. It will vary depending upon the emulsion stability. For now the following rule will be applied:
If our substance is a crude oil, then $f_{w max} = 0.9$<br>
If our substance is a refined product, then $f_{w max} = 0$
### 26. Bullwinkle (time):
Adios3 needs to know when to initiate emulsification. The Adios2 development team called this term Bullwinkle. SINTEF has measured this parameter for some oils, and the new, not yet completed, analysis of emulsification may provide formulas for Bullwinkle. Bullwinkle may be either a time value (i.e. time delay after which the emulsification formulas are turned on) or a fraction of the oil that needs to evaporate or dissolve before emulsification can start.
Bullwinkle(time) is undefined unless the user explicitly sets a value. Then it overrides Bullwinkle(fraction) as the determining parameter for the onset of emulsification.
### 27. Bullwinkle (fraction):
<i>
Reference: Adios2
</i>
Bullwinkle is the mass fraction that must evaporate or dissolve before stable emulsification can begin. This formula will change when we complete the emulsification module.
If $f_{asph} > 0$, then we use method <b>(A)</b><br>
Otherwise, we use method <b>(B)</b><br>
<b>(A)</b> $f_{bull}$ can be calculated from $f_{asph}$:
$$
\begin{align}
f_{bull} &= 0.32 - 3.59 \cdot f_{asph} \qquad \qquad \qquad \boldsymbol{(eq. 33)} \cr
\end{align}
$$
<b>(B)</b> $f_{bull}$ can be calculated from the oil's API:
$$
\begin{align}
f_{bull} &= 0.5762 \cdot log10(API) - 0.6353 \qquad \qquad \boldsymbol{(eq. 34)} \cr
\end{align}
$$
A result of $f_{bull} < 0$ or $f_{bull} > 1$ indicates no emulsification.
<center><b>--- Begin JamesM Comments ---</b></center>
I don't think documentation exists for how Adios2 is __really__ calculating the bullwinkle fraction. But we can look at the C++ code for adios2 in the file OilInitialize.cpp. Here is what it basically does.
First we define the input terms that decide our bullwinkle fraction:
$$
\begin{align}
N_i &= \text{Nickel content of our oil} \cr
V_a &= \text{Vanadium content of our oil} \cr
f_{asph} &= \text{asphaltene fraction of our oil} \cr
API &= \text{The API density of our oil} \cr
\end{align}
$$
There are also some very briefly documented intermediate terms that are used for calculating the final result, which I will describe as best I can based on the code comments:
$$
\begin{align}
t_g &= \text{documented as '} dT/df \text{evaporation'} \cr
&\quad \, \text{I can only assume this is the rate of evaporation with temperature} \cr
t_{bp} &= \text{documented as the 'ADIOS 1 liquid boiling point (bubble point)'} \cr
f_{bull,adios1} &= \text{the ADIOS 1 calculated bullwinkle fraction possibly???} \cr
\end{align}
$$
And our decision tree goes a bit like this:
- if oil type is refined: then $f_{bull} = 1$
- else if oil has an emuls_constant_max property: then $f_{bull} = \text{emuls_constant_max}$
- else:
- if $N_i > 0$ and $V_a > 0$ and $N_i + V_a > 15$: then $f_{bull} = 0$
- else if $f_{asph} > 0$: then $f_{bull} = 0.20219 - 0.168 \cdot log_{10}(f_{asph})$
- else if $API < 26$: then $f_{bull} = 0.08$
- else if $API > 50$: then $f_{bull} = 0.303$
- else:
- $f_{bull} = -1.038 - 0.78935 \cdot log_{10}(1 / API)$
- $t_g = 1356.7 - 247.36 \cdot log(API)$
- $t_{bp} = 532.98 - 3.1295 \cdot API$
- $f_{bull,adios1} = \left( \frac{483 - t_{bp}}{t_g} \right)_{\bot 0}^{\top 0.4} $
- $f_{bull} = avg\left( (f_{bull}, f_{bull,adios1}) \right)$
<center><b>--- End JamesM Comments ---</b></center>
```
def estimate_bullwinkle_fraction_from_asph(f_asph):
return 0.32 - 3.59 * f_asph
def estimate_bullwinkle_fraction_from_api(api):
return 0.5762 * np.log10(api) - 0.6353
def oil_bullwinkle_fraction(imported_rec):
_f_res, f_asph = oil_inert_fractions(imported_rec)
if f_asph > 0.0:
return estimate_bullwinkle_fraction_from_asph(f_asph)
elif imported_rec.api is not None:
return estimate_bullwinkle_fraction_from_api(imported_rec.api)
else:
est_api = api_from_density(oil_density_at_temp(imported_rec, 288.15))
return estimate_bullwinkle_fraction_from_api(est_api)
#for obj in (ans_mp, ans_2002, bahia, arabian, canola):
for obj in (ans_mp,):
print 'oil = ', obj.oil_name
print 'oil.api = ', obj.api
print 'oil.asphaltenes = ', obj.asphaltenes
print 'oil.densities = ', obj.densities
print 'bullwinkle(fraction) = ', oil_bullwinkle_fraction(obj)
print
```
### 28. Solubility:
If the imported oil record contains a valid value for solubility, we will use that value when building our oil.<br>
Otherwise, $c_{solu} = 0 \, kg/m^3$
### 29. Adhesion <i>(not currently used by model)</i>:
If the imported oil record contains a valid value for adhesion, we will use that value when building our oil.<br>
Otherwise, $Adh_{oil} = 0.035 \, kg/m^2$
<i>
Reference: ESTC data
</i>
### 30. Sulphur Mass Fraction <i>(not currently used by model)</i>:
If the imported oil record contains a valid value for sulphur mass fraction, we will use that value when building our oil.<br>
Otherwise, $f_{sulf} = 0$
## <u>Addendum 1</u>
### <center>Addressing Heavy Volatile components</center>
In practice, we have seen that certain oil records in our database result in heavy estimated saturate and aromatic components. That is to say we have saturates and aromatic components that have a higher density than our resins and asphaltenes. Intuitively this does not seem right, and we can identify some things we are doing in our estimations that would cause this:
- We have set a hard-coded density of $1100 \, kg/m^3$ in the case of resins and asphaltenes, which is why we are getting the rejections. Maybe this value should be higher.
- We estimate our component densities based on a Watson characterization factor combined with a component boiling point. And as the boiling temperature rises, our component density calculation will eventually rise to a value higher than $1100 \, kg/m^3$. Maybe we should consider that the volatile component boiling points will not exceed a certain threshold.
And after discussing this at great length, we have decided that:
- $1100 \, kg/m^3$ is probably a reasonable value for our inert components.
- We should set a cap density for the volatile components so they do not exceed that of the involatile components.
So how might we do this? Well the most reasonable way to do this would probably be to alter our distillation temperature curve. Right now it is a linear function that extends from a fraction of $0$ to a fraction of $(1 - f_{res} - f_{asph})$. We could put a threshold limit that would level the curve at a certain upper temperature.
To do this, we will make use of a generalized logistic function or Richard's curve. This should allow us to make our distillation function flexible and not add to the software complexity of our estimations.
$$
\begin{align}
Y(x, M) &= \text{generalized logistic function} \cr
&= A + \frac{K - A}{(C + Qe^{-B(x - M)})^{1/\nu}} \cr
&= 0 + \frac{1 - 0}{(1 + e^{-20(x - M)})^{1/\nu}} \cr
&= \frac{1}{(1 + e^{-20(x - M)})^{1/\nu}} \cr
\cr
\zeta &= \text{smoothing factor for when we cross the M boundary} \cr
&\approx 0.03 \cr
\cr
x_s &= \text{clamped x value} \cr
&= x - x \cdot Y(x, M) + M \cdot Y(x, M) \cr
&= x - \frac{x}{(1 + e^{-20(x - M)})^{1/(1 + \zeta)}} + \frac{M}{(1 + e^{-20(x - M)})^{1/(1 - \zeta)}} \cr
\end{align}
$$
```
def clamp(x, M, zeta=0.03):
'''
The parameters nu, and B have been tweaked a bit to show a smooth
Transition as we cross the M boundary.
'''
return (x -
(x / (1.0 + np.e ** (-15 * (x - M))) ** (1.0 / (1 + zeta)) ) +
(M / (1.0 + np.e ** (-15 * (x - M))) ** (1.0 / (1 - zeta)) ))
def linear_curve(x, a, b, M, zeta=0.09):
x_c = clamp(x, M, zeta)
return (a * x_c + b)
plt.figure(figsize=(12, 6)) # make the drawing area a bit bigger
ax1 = plt.subplot(111)
x = np.linspace(-1.0, 5.0, 100)
# let's plot the function (x + 3), limited at 3
# this should give us a zero crossing of y=3.0,
# and an upper limit of y=6.0
y = linear_curve(x, 1.0, 3.0, 3.0, 0.03)
ax1.plot(x, x, linewidth=1)
ax1.plot(x, y, linewidth=1)
plt.axhline(0)
plt.axvline(0)
show()
```
Ok, this is a pretty reasonable, but general, example of what we would like to do. Let's see what it looks like using distillation cut data. First, let's estimate what our cutoff should be for $T_i$. From Section 19 (Component Density), our density formulas are:
$$
\begin{align}
\rho try_{arom,i} &= 1000 \cdot {\root 3 \of { 1.8 \cdot T_{arom,i}} \over K_{arom,w} } \cr
\frac{\rho try_{arom,i} \cdot K_{arom,w}}{1000} &= \root 3 \of { 1.8 \cdot T_{arom,i}} \cr
\left( \frac{\rho try_{arom,i} \cdot K_{arom,w}}{1000} \right)^3 &= 1.8 \cdot T_{arom,i} \cr
\left( \frac{\rho try_{arom,i} \cdot K_{arom,w}}{1000} \right)^3 \cdot \frac{1}{1.8} &= T_{arom,i} \cr
\left( \frac{1100 \cdot 10}{1000} \right)^3 \cdot \frac{1}{1.8} &= T_{arom,i} \cr
\frac{1331}{1.8} &= T_{arom,i} \cr
739.444 &= T_{arom,i} \cr
\cr
\rho try_{sat,i} &= 1000 \cdot {\root 3 \of { 1.8 \cdot T_{sat,i}} \over K_{sat,w} } \cr
\left( \frac{\rho try_{sat,i} \cdot K_{sat,w}}{1000} \right)^3 \cdot \frac{1}{1.8} &= T_{sat,i} \cr
\left( \frac{1100 \cdot 12}{1000} \right)^3 \cdot \frac{1}{1.8} &= T_{sat,i} \cr
\frac{2299.968}{1.8} &= T_{sat,i} \cr
1277.76 &= T_{sat,i} \cr
\end{align}
$$
Based on the above, it is likely that our aromatic components will become heavier than the inert components at around $739^\circ K$.<br>
Our saturate components will eventually appear too heavy, but at such a great temperature it is unlikely to happen.
So let's choose 739 as our temperature cutoff.
```
def linear_curve(x, a, b):
return (a * x + b)
def inverse_linear_old(y, a, b):
return (y - b) / a
def inverse_linear_curve(y, a, b, M, zeta=0.12):
y_c = clamp(y, M, zeta)
return (y_c - b) / a
def oil_normalized_cut_values(imported_rec, num_cuts=10):
f_res, f_asph = oil_inert_fractions(imported_rec)
culled_cuts = list(oil_culled_cuts(imported_rec))
if len(culled_cuts) == 0:
if imported_rec.api is not None:
oil_api = imported_rec.api
else:
oil_rho = oil_density_at_temp(imported_rec, 288.15)
oil_api = estimate_api_from_density(oil_rho)
BP_i = estimate_cut_temps_from_api(oil_api)
fevap_i = np.cumsum(estimate_n_fmasses(f_res, f_asph))
else:
BP_i, fevap_i = zip(*[(c.vapor_temp_k, c.fraction)
for c in culled_cuts])
popt, pcov = curve_fit(linear_curve, BP_i, fevap_i)
f_cutoff = linear_curve(732.0, *popt) # value of asymptote center (< 739)
popt = popt.tolist() + [f_cutoff]
fevap_i = np.linspace(0.0, 1.0 - f_res - f_asph, (num_cuts * 2) + 1)[1:]
T_i = np.clip(inverse_linear_curve(fevap_i, *popt), 0.0, 5000.0)
fevap_i = fevap_i.reshape(-1,2)[:,1]
T_i = T_i.reshape(-1,2)[:,0]
above_zero = T_i > 0.0
T_i = T_i[above_zero]
fevap_i = fevap_i[above_zero]
return T_i, fevap_i
plt.figure(1, figsize=(12,6))
ax = plt.subplot(111)
obj = boscan
plt.xlabel('% Evap')
plt.ylabel(r'$T_i$', fontsize=16)
plt.grid()
# plot our distillation curve
boiling_points = [c.vapor_temp_k for c in oil_culled_cuts(obj)]
cum_fractions = [c.fraction for c in oil_culled_cuts(obj)]
if len(boiling_points) == 0:
if obj.api is not None:
oil_api = obj.api
else:
oil_rho = oil_density_at_temp(obj, 288.15)
oil_api = estimate_api_from_density(oil_rho)
f_res, f_asph = oil_inert_fractions(obj)
boiling_points = estimate_cut_temps_from_api(oil_api)
cum_fractions = np.cumsum(estimate_n_fmasses(f_res, f_asph))
popt, pcov = curve_fit(linear_curve, boiling_points, cum_fractions)
new_fracs = np.linspace(0.0001, 0.9999, 100)
new_temps = np.clip(inverse_linear_old(new_fracs, *popt), 0.0, 5000.0)
plt.plot(new_fracs, new_temps, '-b', label="Distillation Curve")
# now plot the N fractions we intend to use
new_temps, new_fracs = oil_normalized_cut_values(obj)
plt.plot(new_fracs, new_temps, 'go', label="Estimated Cuts")
# plot our inert fraction
plt.plot(1.0, 1015.0, 'go')
# now we would like to graph the quantized "areas"
# that our fractional masses represent
def make_shaded_region(f_low, f_high,
t_low, t_high,
facecolor='0.8'):
ix = (f_low, f_high)
iy = (t_low, t_high)
verts = [(x, y) for x in ix for y in iy]
verts[2:] = reversed(verts[2:])
poly = Polygon(verts, facecolor=facecolor, edgecolor='k')
ax.add_patch(poly)
for r in zip([0.0] + new_fracs.tolist(),
new_fracs,
[inverse_linear(0.0, *popt)] + new_temps.tolist(),
new_temps):
make_shaded_region(*r, facecolor='#ddeeff')
# make the shaded region for our inert fraction
make_shaded_region(new_fracs[-1], 1.0,
new_temps[-1], 1015.0)
# ticks for the shaded region
extraticks = [low,]
extralabels = [r'${f_{inert}}$',]
plt.xticks(list(plt.xticks()[0]) + extraticks,
list(plt.xticks()[0]) + extralabels)
for tick in ax.xaxis.get_major_ticks():
if tick.label.get_text().find('inert') >= 0:
tick.label.set_fontsize(16)
tick.label.set_rotation('vertical')
plt.axhline(739, color='r')
plt.legend(loc='upper left')
plt.show()
```
Ok, that seems like a pretty reasonable response. This solution will require some peer review from Chris, Bill, and Robert, but it will certainly prevent our estimation methods from returning volatile oil fractions with higher densities than the inert fractions.
## <u>Addendum 2</u>
### <center>The Oil Density Temperature Expansion Coefficient</center>
Currently, we define the density conversion formula for different temperatures as:
$$
\begin{align}
\rho_0 &= \text{measured density} \cr
T_0 &= \text{temperature at which density is measured} \cr
k_{\rho T} &= \text{the volumetric expansion coefficient} \cr
&= 0.0008 \, K^{-1} \cr
\cr
\rho_{ref} &= \rho_0 \cdot (1 - k_{\rho T} \cdot (T_{ref} - T_0 ))
\end{align}
$$
This is what Adios2 did, and it seems to produce sane estimations of an oil's density at some arbitrary temperature. But the constant value that is used for the volumetric expansion coefficient is simply an average of a bunch of sampled oils. As such, it is a good middle-of-the-road value to use in lieu of a precise coefficient.
It would be better if we could determine a precise value for this coefficient, and if an oil has two or more density measurements, we should be able to do it by referring to the above equation and modifying it as follows:
$$
\begin{align}
\rho_{i} &= \text{a density value indexed by } i \cr
T_{i} &= \text{a temperature value indexed by } i \cr
k_{\rho T} &= \text{the volumetric expansion coefficient} \cr
\cr
\rho_{i+1} &= \rho_i \cdot (1 - k_{\rho T} \cdot (T_{i+1} - T_i)) \cr
\frac{\rho_{i+1}}{\rho_i} &= 1 - k_{\rho T} \cdot (T_{i+1} - T_i) \cr
\frac{\rho_{i+1}}{\rho_i} - 1 &= - k_{\rho T} \cdot (T_{i+1} - T_i) \cr
\frac{\rho_{i+1} - \rho_i}{\rho_i} &= - k_{\rho T} \cdot (T_{i+1} - T_i) \cr
\frac{\rho_{i+1} - \rho_i}{\rho_i \cdot (T_{i+1} - T_i)} &= - k_{\rho T} \cr
\frac{\rho_i - \rho_{i+1}}{\rho_i \cdot (T_{i+1} - T_i)} &= k_{\rho T} \cr
\cr
\end{align}
$$
Well, there it is. Let's see if this gives us anything sane.
```
def get_vol_expansion_coeff(rho_0, t_0, rho_1, t_1):
k_rho_t = (rho_0 - rho_1) / (rho_0 * (t_1 - t_0))
return k_rho_t
def oil_vol_expansion_coeff(imported_rec, weathering=0.0):
density_list = [(d.kg_m_3, d.ref_temp_k)
for d in imported_rec.densities
if (d.kg_m_3 is not None and
d.ref_temp_k is not None and
d.weathering == weathering)]
if len(density_list) >= 2:
d_args = [t for d in density_list[:2] for t in d]
k_rho_t = get_vol_expansion_coeff(*d_args)
else:
k_rho_t = 0.0008
return k_rho_t
print ans_mp.densities
print oil_vol_expansion_coeff(ans_mp)
```
Ok, so far so good. Let's see if we can try this out on all oils that have multiple density measurements.
```
mult_dens_oils = [ir for ir in session.query(ImportedRecord).all()
if (ir.product_type is not None and
ir.product_type.lower() == 'crude' and
len(ir.densities) >= 2)]
print len(mult_dens_oils)
for o in mult_dens_oils:
print '{} ({}):\t'.format(o.oil_name, o.product_type),
print '{}'.format(oil_vol_expansion_coeff(o))
mult_dens_oils = [ir for ir in session.query(ImportedRecord).all()
if (ir.product_type is not None and
ir.product_type.lower() == 'refined' and
len(ir.densities) >= 2)]
print '\n', len(mult_dens_oils)
for o in mult_dens_oils:
print '{} ({}):\t'.format(o.oil_name, o.product_type),
print '{}'.format(oil_vol_expansion_coeff(o))
```
Ok, that seems to be producing results that are in a reasonable range for crude as well as refined products.
Bill has given us a new document 'density_thermal_expansion.docx' (Jan 2017), in which he defines a couple of different expansion coefficients to be used if we don't have enough density data to perform a calculation.
- 0.0008: for crude oils and heavy refined products
- 0.0009: for refined products with an API > 30
This was amended, through discussion with Chris, to inclusively group both crude and refined products with API > 30. The document has not been updated at this time, but we will go with this amendment since it is truly what we want. So what we get is:
- 0.0008: if API <= 30
- 0.0009: if API > 30
So let's update our function:
```
def get_vol_expansion_coeff(rho_0, t_0, rho_1, t_1):
if t_0 == t_1:
k_rho_t = 0.0
else:
k_rho_t = (rho_0 - rho_1) / (rho_0 * (t_1 - t_0))
return k_rho_t
def oil_vol_expansion_coeff(imported_rec, weathering=0.0):
density_list = [(d.kg_m_3, d.ref_temp_k)
for d in imported_rec.densities
if (d.kg_m_3 is not None and
d.ref_temp_k is not None and
d.weathering == weathering)]
if len(density_list) >= 2:
d_args = [t for d in density_list[:2] for t in d]
k_rho_t = get_vol_expansion_coeff(*d_args)
else:
if imported_rec.api > 30:
k_rho_t = 0.0009
else:
k_rho_t = 0.0008
return k_rho_t
print 'Checking some low API oils:'
low_api_oils = (session.query(ImportedRecord)
.filter(ImportedRecord.api <= 30).all())
for o in low_api_oils[:10]:
print '{} ({}):\t'.format(o.oil_name, o.product_type),
print '{}'.format(oil_vol_expansion_coeff(o))
print '\nChecking some high API oils:'
high_api_oils = (session.query(ImportedRecord)
.filter(ImportedRecord.api > 30).all())
for o in high_api_oils[:10]:
print '{} ({}):\t'.format(o.oil_name, o.product_type),
print '{}'.format(oil_vol_expansion_coeff(o))
```
Alright, this seems to exhibit the right default behavior for oils that don't have enough density information to calculate the expansion coefficient.
```
imported_rec = session.query(ImportedRecord).filter(ImportedRecord.oil_name == "COLD LAKE BITUMEN").one()
print imported_rec.adios_oil_id
for d in imported_rec.densities:
print (d.kg_m_3, d.ref_temp_k, d.weathering)
fig, ax = plt.subplots()
op_obj = oil_library.get_oil_props("COLD LAKE BITUMEN")
for d in op_obj.get_densities():
print (d.kg_m_3, d.ref_temp_k, d.weathering)
densities = op_obj.get_densities()
ref_t = [d.ref_temp_k for d in densities]
ref_d = [d.kg_m_3 for d in densities]
ax.plot(ref_t, ref_d, 'o')
temps = np.linspace(273.0, 300.0, 100)
densities = op_obj.density_at_temp(temps)
ax.plot(temps, densities, '-')
ax.set_xlabel(r'temperature ($\degree K$)')
ax.set_ylabel(r'density ($kg/m^3$)')
```
Ok, I think we found an oil record that has some bad data. I see two issue that are slightly different:
- Bad density values. In this case it is a density value that is repeated at two different temperatures.
- Chris mentioned that he noticed 'glitches' midway between two density data points, and that he would like there to be a smooth interpolation between them.
### Smooth transition between Oil Density Data Values
We already have a method of determining the volumetric expansion coefficient. For oil records that have more than two density measurements, it is simply a problem of choosing the right two based on the temperature for which we are trying to estimate the density.
If we have two or more densities, then we should be able to determine the expansion coefficient, but there are some edge cases that we will list:
- temperature within bounds: we take the two closest measurements, specifically the closest one immediately lower than the specified temperature and the closest one immediately higher.
- low temperature out-of-bounds: Chris & Bill would like to take the lowest measurement and use the default expansion coefficient
- high temperature out-of-bounds: Chris & Bill would like to take the highest measurement and use the default expansion coefficient
Let's see if we can do this.
```
def closest_to_temperature(obj_list, temperature):
'''
General Utility Function
From a list of objects containing a ref_temp_k attribute,
return the object(s) that are closest to the specified temperature(s)
specifically:
- we want the ones that immediately bound our temperature.
- if our temperature is high and out of bounds of the temperatures
in our obj_list, then we return a range containing only the highest
temperature.
- if our temperature is low and out of bounds of the temperatures
in our obj_list, then we return a range containing only the lowest
temperature.
We accept only a scalar temperature or a sequence of temperatures
'''
temperature = np.array(temperature)
if len(obj_list) <= 1:
# range where the lowest and highest are basically the same.
return [(obj_list * 2) for t in temperature]
else:
geq_temps = temperature.reshape(-1, 1) >= [obj.ref_temp_k
for obj in obj_list]
high_and_oob = np.all(geq_temps, axis=1)
low_and_oob = np.all(geq_temps ^ True, axis=1)
rho_idxs0 = np.argmin(geq_temps, axis=1)
rho_idxs0[rho_idxs0 > 0] -= 1
rho_idxs0[high_and_oob] = len(obj_list) - 1
rho_idxs1 = (rho_idxs0 + 1).clip(0, len(obj_list) - 1)
rho_idxs1[low_and_oob] = 0
range_idxs = np.array((rho_idxs0, rho_idxs1)).T
return zip([obj_list[i] for i in rho_idxs0],
[obj_list[i] for i in rho_idxs1])
def oil_vol_expansion_coeff(densities, temperature):
closest_densities = closest_to_temperature(densities,
temperature)
temperature = np.array(temperature)
closest_values = np.array([[(d.kg_m_3, d.ref_temp_k)
for d in r]
for r in closest_densities])
args_list = [[t for d in v for t in d]
for v in closest_values]
k_rho_t = np.array([get_vol_expansion_coeff(*args)
for args in args_list])
greater_than = np.all((temperature > closest_values[:,:,1].T).T, axis=1)
less_than = np.all((temperature < closest_values[:,:,1].T).T, axis=1)
if imported_rec.api > 30:
k_rho_default = 0.0009
else:
k_rho_default = 0.0008
k_rho_t[greater_than | less_than] = k_rho_default
if k_rho_t.shape[0] == 1:
return k_rho_t[0]
else:
return k_rho_t
op_obj = oil_library.get_oil_props('COLD LAKE BITUMEN')
for d in op_obj.get_densities():
print d
temp_k = (272.0, 274.0, 279.0, 282.0, 284.0, 287.0, 288.1, 300.0)
print
pp.pprint(closest_to_temperature(op_obj.get_densities(), temp_k))
for t in temp_k:
print
print closest_to_temperature(op_obj.get_densities(), t),
print; print
pp.pprint(oil_vol_expansion_coeff(op_obj.get_densities(), temp_k))
for t in temp_k:
print
print oil_vol_expansion_coeff(op_obj.get_densities(), t),
```
Ok, now all we have to do is redefine our function for calculating density at a temperature so that it uses a temperature sensitive expansion coefficient.
```
def estimate_density_at_temp(ref_density, ref_temp_k, temp_k, k_rho_t=0.0008):
return ref_density / (1.0 - k_rho_t * (ref_temp_k - temp_k))
def get_reference_densities(densities, temperature):
closest_densities = closest_to_temperature(densities,
temperature)
try:
# sequence of ranges
density_values = np.array([[d.kg_m_3 for d in r]
for r in closest_densities])
ref_temp_values = np.array([[d.ref_temp_k for d in r]
for r in closest_densities])
greater_than = np.all((temperature > ref_temp_values.T).T, axis=1)
density_values[greater_than,0] = density_values[greater_than,1]
ref_temp_values[greater_than,0] = ref_temp_values[greater_than,1]
return density_values[:,0], ref_temp_values[:,0]
except TypeError:
# single range
density_values = np.array([d.kg_m_3 for d in closest_densities])
ref_temp_values = np.array([d.ref_temp_k for d in closest_densities])
if np.all(temperature > ref_temp_values):
return density_values[1], ref_temp_values[1]
else:
return density_values[0], ref_temp_values[0]
def oil_get_densities(imported_rec, weathering=0.0):
densities = [d for d in imported_rec.densities
if (d.kg_m_3 is not None and
d.ref_temp_k is not None and
d.weathering == weathering)]
if (weathering == 0.0 and
imported_rec.api is not None and
len([d for d in densities if d.ref_temp_k == 288.15]) == 0):
kg_m_3, ref_temp_k = estimate_density_from_api(imported_rec.api)
densities.append(Density(kg_m_3=kg_m_3,
ref_temp_k=ref_temp_k,
weathering=0.0))
return sorted(densities, key=lambda d: d.ref_temp_k)
def oil_density_at_temp(imported_rec, temperature=288.15, weathering=0.0):
densities = oil_get_densities(imported_rec, weathering=weathering)
# set the minimum temperature to be the oil's pour point
min_temp = np.min([t for t in oil_pour_point(imported_rec)] +
[d.ref_temp_k for d in densities])
temperature = np.clip(temperature, min_temp, 1000.0)
ref_density, ref_temp_k = get_reference_densities(densities, temperature)
k_rho_t = oil_vol_expansion_coeff(densities, temperature)
rho_t = estimate_density_at_temp(ref_density, ref_temp_k, temperature, k_rho_t)
return estimate_density_at_temp(ref_density, ref_temp_k, temperature, k_rho_t)
pp.pprint(oil_get_densities(imported_rec))
temp_k = (260.0, 270.0, 275.0, 280.0, 285.0, 300)
print
print oil_density_at_temp(imported_rec, temp_k)
for t in temp_k:
print
print oil_density_at_temp(imported_rec, t),
plt.figure(1, figsize=(12,8))
ax = plt.subplot(111)
oil_obj = session.query(ImportedRecord).filter(ImportedRecord.oil_name == "COLD LAKE BITUMEN").one()
#oil_obj = session.query(ImportedRecord).filter(ImportedRecord.adios_oil_id == 'AD00102').one()
print 'Oil: {}, Adios ID: {}'.format(oil_obj.oil_name, oil_obj.adios_oil_id)
densities = oil_get_densities(oil_obj)
for d in densities:
print (d.kg_m_3, d.ref_temp_k, d.weathering)
ref_t = [d.ref_temp_k for d in densities]
ref_d = [d.kg_m_3 for d in densities]
ax.plot(ref_t, ref_d, 'o')
temps = np.linspace(260.0, 320.0, 200)
rho_t = oil_density_at_temp(oil_obj, temps)
ax.plot(temps, rho_t, '-')
ax.set_xlabel(r'temperature ($\degree K$)')
ax.set_ylabel(r'density ($kg/m^3$)')
```
Ok that's not too bad. We have a linear interpolation between measured densities, the expansion coefficient above and below the range of our measured densities, at least for this oil, is 0.0008, and we cut off the lower temperature densities at the oil's pour point.
| github_jupyter |
# Optimization Sample: Multi-ship loading
In this example, we will take our learnings from the ship-loading sample and generalize to load-balancing between any number of ships. In addition, we'll see how we can make use of the parameter free-solvers to guide us on a selection of parameters which we can use with the parametrized solvers.
## Pre-requisites
1. [Create an Azure Quantum Workspace](https://github.com/MicrosoftDocs/quantum-docs-private/wiki/Create-quantum-workspaces-with-the-Azure-portal)
2. [Install the `azure-quantum` Python module](https://github.com/MicrosoftDocs/quantum-docs-private/wiki/Use-the-Python-SDK-for-Quantum-Inspired-Optimization).
3. [Complete the ship-loading sample](https://github.com/microsoft/qio-samples/tree/main/samples/ship-loading)
## The Problem
To tackle this problem we will use a PUBO format. As a reminder, these are [cost functions](https://docs.microsoft.com/en-us/azure/quantum/optimization-concepts-cost-functions) where the variables take the values of either 0 or 1 (rather than -1 or 1 for an Ising cost function).
In order to balance containers between multiple ships, one option is to define a cost function that:
1. Penalizes variance from a theoretical equal distribution (where an equal distribution is the total weight of the containers divided by the number of ships), and,
2. Penalizes the assignment of the same container on multiple ships
We will create two sub cost-functions $H1$ and $H2$ that we will then sum to evaluate the total cost of a solution. Let's begin with the first cost function, $H1$.
## Penalize variance from equal distribution between ships
Suppose we had 3 containers with respective weights $W0$, $W1$, $W2$, and we define an equal distribution of the container weights to be:
$$EqDistrib = (W0 + W1 + W2) / 3$$
A way to penalize a large variance from the equal distribution for a given ship is to express it in the following way:
$$(W0 + W1 + W2 - EqDistrib)^2$$
Let's take the following example:
| | |
|-----------------|---------------|
|Container weights| 1, 5, 9, 7, 3 |
|Total weight | 25 |
|Ships | A, B, C |
|EqualDistrib | 25 / 3 = 8.33 |
Suppose we were to assign those containers to the ships listed, we can calculate the variance from the equal distribution for each of the given ships, shown in the rightmost column:
| Ships\Containers | 1 | 5 | 9 | 7 | 3 | | |
|------------------|---|---|---|---|---|------------|--------|
| A | 0 | 0 | 9 | 0 | 0 | (9-8.33)^2 |= 0.4489|
| B | 0 | 5 | 0 | 0 | 3 |(5+3-8.33)^2|= 0.1089|
| C | 1 | 0 | 0 | 7 | 0 |(1+7-8.33)^2|= 0.1089|
As we need to represent our problem in a binary format we need to "encode" the presence ($x_i=1$) or absence ($x_i=0$) of a given container on a ship. To do this, we need to have a label for the weight of each container on each ship. The table below shows how we assign this continuous index by repeating the list of container weights for each ship and assigning a single list of weight labels across all three ships:
|| Ship A | | | | | Ship B | | | | | Ship C | | | | |
|---|--------|-----|-----|-----|-----|--------|-----|-----|-----|-----|----------|--------|--------|--------|--------|
|Container weight| 1 | 5 | 9 | 7 | 3 | 1 | 5 | 9 | 7 | 3 | 1 | 5 | 9 | 7 | 3 |
|Weight label|*w<sub>0</sub>*|*w<sub>1</sub>*|*w<sub>2</sub>*|*w<sub>3</sub>*|*w<sub>4</sub>*| *w<sub>5</sub>*|*w<sub>6</sub>*|*w<sub>7</sub>*|*w<sub>8</sub>*|*w<sub>9</sub>*|*w<sub>10</sub>*|*w<sub>11</sub>*|*w<sub>12</sub>*|*w<sub>13</sub>*|*w<sub>14</sub>*|
The cost function $H1$ becomes:
$$ H1 = H_{A} + H_{B} + H_{C} $$
where:
$$ H_{A} = (w_0 x_0 + w_1 x_1 + w_2 x_2 + w_3 x_3 + w_4 x_4 - EqDistrib)^2 $$
$$ H_{B} = (w_5 x_5 + w_6 x_6 + w_7 x_7 + w_8 x_8 + w_9 x_9 - EqDistrib)^2 $$
and
$$ H_{C} = (w_{10} x_{10} + w_{11} x_{11} + w_{12} x_{12} + w_{13} x_{13} + w_{14} x_{14} - EqDistrib)^2 $$
We can expand the above and group the common terms, for example if we expand $H_{A}$, we get:
$$
\begin{align}
H_{A} &= (\sum_i(w_i x_i) - EqDistrib)^2\\
&= (w_0 x_0 + w_1 x_1 + w_2 x_2 + w_3 x_3 + w_4 x_4 - EqDistrib)^2\\
\end{align}
$$
To simplify things for the expansion, let's rename the variables as follows:
$$
\begin{align}
w_0 x_0 &= a \\
w_1 x_1 &= b \\
w_2 x_2 &= c \\
w_3 x_3 &= d \\
w_4 x_4 &= e \\
EqDistrib &= f \\
\end{align}
$$
So now we have:
$$
\begin{align}
H_{A} &= (\sum_i(w_i x_i) - EqDistrib)^2\\
&= (w_0 x_0 + w_1 x_1 + w_2 x_2 + w_3 x_3 + w_4 x_4 - EqDistrib)^2\\
&= (a + b + c + d + e - f)^2\\
&= a^2 + b^2 + c^2 + d^2 + e^2 + f^2 + 2(ab + ac + ad + ae + bc + bd + be + cd + ce + de) - 2(af + bf + cf + df + ef)
\end{align}
$$
Substituting our original values back in, this gives us the following:
$$
\begin{align}
H_{A} &= (\sum_i(w_i x_i) - EqDistrib)^2\\
&= w_0^2 x_0^2 + w_1^2 x_1^2 + w_2^2 x_2^2 + w_3^2 x_3^2 + w_4^2 x_4^2 + EqDistrib ^2 + 2(w_0 x_0 \cdot w_1 x_1 + w_0 x_0 \cdot w_2 x_2 + w_0 x_0 \cdot w_3 x_3 + w_0 x_0 \cdot w_4 x_4 + w_1 x_1 \cdot w_2 x_2 + w_1 x_1 \cdot w_3 x_3 + w_1 x_1 \cdot w_4 x_4 + w_2 x_2 \cdot w_3 x_3 + w_2 x_2 \cdot w_4 x_4 + w_3 x_3 \cdot w_4 x_4) - 2(w_0 x_0 \cdot EqDistrib + w_1 x_1 \cdot EqDistrib + w_2 x_2 \cdot EqDistrib + w_3 x_3 \cdot EqDistrib + w_4 x_4 \cdot EqDistrib)
\end{align}
$$
We can do the same for $H_{B}$, and $H_{C}$.
## Penalize the assignment of the same container on multiple ships
Using the containers weight encoding above, we can devise a cost function such as this one for the first container:
$$ H_{D} = (w_0 x_0 + w_5 x_5 + w_{10} x_{10} - w_0)^2 $$
As $w_0$, $w_5$ and $w_{10}$ are actually the same value (it is the same container represented across multiple ships) we have:
$$ H_{D} = (w_0 x_0 + w_0 x_5 + w_0 x_{10} - w_0)^2 $$
If we expand and group the common terms, we get the following:
$$ H_{D} = {w_0}^2 {x_0}^2 + {w_0}^2 {x_5}^2 + {w_0}^2 {x_{10}}^2 + {w_0}^2 +
2 ({w_0}^2 x_0 x_5 + {w_0}^2 x_0 x_{10} + {w_0}^2 x_5 x_{10})
- 2({w_0}^2 x_0 + {w_0}^2 x_5 + {w_0}^2 x_{10}) $$
We can then repeat the above for each container across all ships:
So in addition to:
$$ H_{D} = (w_0 x_0 + w_0 x_5 + w_0 x_{10} - w_0)^2 $$
we also have:
$$ H_{E} = (w_1 x_1 + w_1 x_6 + w_1 x_{11} - w_1)^2 $$
$$ H_{F} = (w_2 x_2 + w_2 x_7 + w_2 x_{12} - w_2)^2 $$
$$ H_{G} = (w_3 x_3 + w_3 x_8 + w_3 x_{13} - w_3)^2 $$
$$ H_{H} = (w_4 x_4 + w_4 x_9 + w_4 x_{14} - w_4)^2 $$
Grouping these together into a single cost function, $H2$, we get:
$$ H2 = H_{D} + H_{E} + H_{F}+ H_{G}+ H_{H} $$
which we can expand and group the terms as we did with $H_D$ above.
## Combining our cost functions
The final part of the problem definition is to combine our cost functions $H1$ and $H2$:
$$H = H1 + H2 $$
You will notice that $H1$ and $H2$ have common indices $[i,i]/[m,m]$ and $[i]/[m]$. We will need to be careful to not duplicate them, but sum them, in our final list of terms describing the cost function.
## Solving the Problem in Python
First, we must instantiate a `Workspace` object which allows you to connect to the Workspace you've previously deployed in Azure. Be sure to fill in the settings below which can be retrieved by running `az quantum workspace show`.
```
# This allows you to connect to the Workspace you've previously deployed in Azure.
# Be sure to fill in the settings below which can be retrieved by running 'az quantum workspace show' in the terminal.
from azure.quantum import Workspace
# Copy the settings for your workspace below
workspace = Workspace (
subscription_id = "",
resource_group = "",
name = "",
location = ""
)
```
First, let's define a function to add terms according to our definition of the H1 cost function, where we penalized the variance from an equal distribution between the ships.
```
def AddTermsWeightVarianceCost(start, end, containers, EqDistrib):
terms: List[Term] = []
for i,w in enumerate(containers[start:end+1], start):
# -2*Wi*EqDistrib.xi -2Wi^2.xi (weight variance cost + duplicate container cost)
terms.append(Term(w=-2*w*EqDistrib - 2*w*w, indices=[i]))
# Wi^2.xi^2 + Wi^2.xi^2 (weight variance cost + duplicate container cost)
terms.append(Term(w=2*w*w, indices=[i,i]))
for c in combinations(range(start, end+1), 2):
w0 = containers[c[0]]
w1 = containers[c[1]]
# 2*Wi*Wj (weight variance cost)
terms.append(Term(w=2*w0*w1, indices=[c[0],c[1]]))
return terms
```
Next, let's incorporate our definition of the second part of our cost function, H2, where we wished to penalize the assignment of the same container on multiple ships.
```
def AddTermsDuplicateContainerCost(start, end, containers):
terms: List[Term] = []
# The following is integrated into AddTermsWeightVarianceCost to reduce the number of Terms and speed-up Terms generation
# for c in combinations(range(start, end+1), 1):
# w = containers[c[0]][0]
# i1 = containers[c[0]][1]
# terms.append(Term(w=w*w, indices=[i1,i1])) # Wi^2
# 2.w^2.x_i.x_j terms
for c in combinations(range(start, end+1), 2):
w = containers[c[0]][0]
i1 = containers[c[0]][1]
i2 = containers[c[1]][1]
terms.append(Term(w=2*w*w, indices=[i1,i2])) # Term(w=2*Wm^2, [m,n])
# The following is integrated into AddTermsWeightVarianceCost to reduce the number of Terms and speed-up Terms generation
# # for c in combinations(range(start, end+1), 1):
# w = containers[c[0]][0]
# i1 = containers[c[0]][1]
# terms.append(Term(w=-2*w*w, indices=[i1])) # -2*Wi^2
# w^2 term
terms.append(Term(w=containers[start][0]*containers[start][0], indices=[]))
return terms
```
Combining these together, we can create our cost function definition of the multi-ship problem.
```
from typing import List
from azure.quantum.optimization import Term
import numpy as np
from itertools import combinations
def createProblemForContainerWeights(containerWeights: List[int], Ships) -> List[Term]:
terms: List[Term] = []
containersWithinShip: List[int] = []
containersAcrossShips: List[int, int] = []
totalWeight = 0
EqDistrib = 0
for c in range (len(containerWeights)):
totalWeight = totalWeight + containerWeights[c]
EqDistrib = totalWeight / len(Ships)
print(Ships)
print(containerWeights)
print("Total Weight:", totalWeight)
print("Equal weight distribution:", EqDistrib)
# Create container weights in this format:
# 1 5 9 7 3 - 1 5 9 7 3 - 1 5 9 7 3
# W0 W1 W2 W3 W4 W5 W6 W7 W8 W9 W10 W11 W12 W13 W14
containersWithinShip = containerWeights*len(Ships)
# Create container weights in this format:
# 1 1 1 5 5 5 9 9 9 7 7 7 3 3 3
for i in range(len(containerWeights)):
for j in range(len(Ships)):
k = i + j*len(containerWeights)
containersAcrossShips.append([containersWithinShip[i], k])
for split in np.array_split(range(len(containersWithinShip)), len(Ships)):
terms = terms + AddTermsWeightVarianceCost(split[0], split[-1], containersWithinShip, EqDistrib)
for split in np.array_split(range(len(containersAcrossShips)), len(containerWeights)):
terms = terms + AddTermsDuplicateContainerCost(split[0], split[-1], containersAcrossShips)
return terms
```
Before we solve our problem, let's create a function to help us visualize the results of the solver:
```
def visualize_result(result, containers, ships, target):
print("\rResult received from: ", target)
nb_ships = len(ships)
try:
config = result['configuration']
config = list(config.values())
for ship, sub_config in enumerate(np.array_split(config, nb_ships)):
shipWeight = 0
for c,b in enumerate(sub_config):
shipWeight = shipWeight + b*containers[c]
print(f'Ship {ships[ship]}: \t' + ''.join(f'{b*containers[c]}' for c,b in enumerate(sub_config)) + ' - ' + str(shipWeight))
except:
print('No Configuration')
try:
print('Cost: {}'.format(result['cost']))
except:
print('No Cost')
try:
print('Parameters: {}'.format(result['parameters']))
except:
print('No Parameter')
```
Lastly, let's define a function that will submit our problem to Azure Quantum and visualize the results.
```
def SolveMyProblem(problem, s):
try:
# Optimize the problem
print("Optimizing with:", s.name)
Job = s.submit(problem)
Job.wait_until_completed()
duration = Job.details.end_execution_time - Job.details.begin_execution_time
if (Job.details.status == "Succeeded"):
visualize_result(Job.get_results(), containerWeights*len(Ships), Ships, s.name)
print("Execution duration: ", duration)
else:
print("\rJob ID", Job.id, "failed")
return Job.id
except BaseException as e:
print(e)
```
## Submitting our problem to Azure Quantum
Now, we can put it all together. In the following steps we'll:
1. Define the list of containers and their weights
2. Instantiate the problem, creating our list of terms
3. Submit these terms to a parameter-free solver
4. Use the parameters returned to submit the problem to a parametrized solver
```
# This array contains a list of the weights of the containers:
containerWeights = [3, 8, 3, 4, 1, 5, 2, 2, 7, 9, 5, 4, 8, 9, 4, 6, 8, 7, 6, 2, 2, 9, 4, 6, 3, 8, 5, 7, 2, 4, 9, 4]
Ships = ["A", "B", "C", "D", "E"]
# Create the Terms for this list of containers:
terms = createProblemForContainerWeights(containerWeights,Ships)
from azure.quantum.optimization import Problem, ProblemType
# Create the Problem to submit to the solver:
nbTerms = len(terms)
problemName = f'Balancing {str(len(containerWeights))} containers between {str(len(Ships))} Ships ({nbTerms:,} terms)'
print(problemName)
problem = Problem(name=problemName, problem_type=ProblemType.pubo, terms=terms)
from azure.quantum.optimization import SimulatedAnnealing
# Try to call a solver with different timeout value and see if it affects the results
jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace, timeout=10))
# jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace, timeout=20))
# jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace, timeout=30))
# jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace))
# Try using the parameters returned by the parameter free versions and observe the significant performance improvement
# First use the job id to view the parameters selected by the parameter free solver
job = workspace.get_job(jobid)
results = job.get_results()
print(results)
# From the results, let's extract the beta_start, beta_stop, restarts, and sweeps parameters selected
beta_start = results["parameters"]["beta_start"]
beta_stop = results["parameters"]["beta_stop"]
restarts = results["parameters"]["restarts"]
sweeps = results["parameters"]["sweeps"]
# Now let's call the solver again, this time the parametrized version, using these parameters
jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace, timeout=5, beta_start=beta_start, beta_stop=beta_stop, restarts=restarts, sweeps=sweeps))
from azure.quantum.optimization import ParallelTempering, Tabu, HardwarePlatform, QuantumMonteCarlo
# Here's how we could experiment with different solvers from Microsoft
jobid = SolveMyProblem(problem, SimulatedAnnealing(workspace, platform=HardwarePlatform.FPGA, timeout=5))
# jobid = SolveMyProblem(problem, Tabu(workspace, timeout=5))
# jobid = SolveMyProblem(problem, ParallelTempering(workspace, timeout=60))
# jobid = SolveMyProblem(problem, QuantumMonteCarlo(workspace))
from azure.quantum.target.oneqbit import PathRelinkingSolver
# And how we can submit the same jobs to solvers by third-party providers, such as 1QBit
# Note: PathRelinkingSolver is only available if the 1QBit provider is enabled in your quantum workspace
# jobid = SolveMyProblem(problem, PathRelinkingSolver(workspace))
```
| github_jupyter |
# 1. COLLECTIONS
A collections is a container that is used to store different objects


# A) List: "[ ]"
```
lst = ["one", 2, "three",2.2]
print(lst)
type(lst)
#How to insert new item in a lst? .append()
lst.append("abc")
lst
#How to remove item from a lst? .remove()
lst.remove("abc")
print(lst)
#How to print a specific item of a lst?
#We will use indexes for that…
#print(lst[0]) # print first item of a lst
print(lst[-1])
lst
#Check for a item in a lst…
if "one" in lst:
print("Found")
else:
print("Not found")
print(len(lst)) # print length of a list
#Join two lst
lst1 = ["three", "b" , "a"]
lst2 = [3, 2, 1]
lst3 = lst1 + lst2
print(lst3)
print(lst3[0])
lst2.sort()# this function will sort the lst
print(lst2)
lst2.reverse() # this function will print lst in reverse order
print(lst2)
lst
lst.clear() # will clear/remove all items in the lst
print(lst)
del lst # will delete lst
print(lst) # will give error
```
# B) Tuple: "( )"
```
cars = ('BMW', 'Suzuki', 'Nissan',2)
print(cars)
#How to insert new item in a tuple?
cars.append(4)
print(cars)
# This will give us an error because we cant insert in tuple once it defined
#How to remove item from a list?
cars.remove('BMW')
print(cars)
# This will give us an error because we cant insert in tuple once it defined
cars
#How to print a specific item of a tuple?
#We will use indexes for that…
print(cars[0]) # print first item of a tuple
print(cars[1]) # print second item of a tuple
print(cars[-1]) # print last item of a tuple
print(cars[-2]) # print second last item of a tuple
print(cars[1:3])
#Check for a item in a tuple…
if "asd" in cars:
print("Found")
else:
print("Not found")
print(len(cars)) # print length of a tuple
#Join two list
tup1 = ("b", "c" , "a")
tup2 = (1, 2, 3)
tup3 = tup1 + tup2
print(tup3)
type(cars)
cars = (1,2,3)
result = reversed(cars)# this function will print tuple in reverse order
print(tuple(result))
result = sorted(tup1)# this function will sort the tuple
print(tuple(result))
cars=list(cars) # convert type of tuple to list
cars.clear() # will clear/remove all items in the tuple
print(cars)
del cars # will delete tuple
print(cars) # will give error
```
# C) Sets: "{}"
```
cars_set = {"Nissan", "BMW", "Honda",2}
print(cars_set)
#How to insert new item in a set? .add()
cars_set.add('Eric')
print(cars_set)
#How to remove item from a set? .remove()
cars_set.remove('Eric')
print(cars_set)
#Check for a item in a set…
if "one" in cars_set:
print("Found")
else:
print("Not found")
print(len(cars_set)) # print length of a set
#Join two set
set1 = {"a", "b" , "c"}
set2 = {1, 2, 3}
#tuple = tup1 + tup2
set3 = set1.union(set2)
print(set3)
```
Advantages of Python Sets:
Because sets cannot have multiple occurrences of the same element, it makes sets highly useful to efficiently remove duplicate values from a list or tuple and to perform common math operations like unions and intersections.
# D) Dictionary: {key: value}
```
empty_dict = {} #empty dictionary
my_data = {'Name': 'Uzair', 'Age': 24, 'Education': 'Masters'}
print(my_data)
my_data['Education']
# Print dictionary element using key
print(my_data['Name'])
print(my_data['Age'])
# Print dictionary element using get() function
edu = my_data.get('Age')
print(edu)
# Change a key value
my_data['Age'] = 25
print(my_data)
#insert new item
my_data['Country'] = 'Pakistan'
print(my_data)
print(my_data.items())
print()
#print(my_data.keys()) #print all keys of dictionary
#print()
#print(my_data.values()) #print all values of dictionary
#print()
if "Uzair" in my_data:
print('Key found')
else:
print('Not found')
print(len(my_data))
del my_data['Name'] # remove entry with key 'Name'
print(my_data)
#del my_data # delete entire dictionary
```
# 2. LOOPS


```
while <condition>:
```
# A) While Loop
```
# Program to add natural
# numbers up to
# sum = 1+2+3+...+n
# To take input from the user,
# n = int(input("Enter n: "))
n = 10
# initialize sum and counter
sum = 0
i = 1
while i <= n:
sum = sum + i
i = i+1 # update counter
print(i)
# print the sum
print("The sum is", sum)
```
- In the above program, the test expression will be True as long as our counter variable i is less than or equal to n (10 in our program).
- We need to increase the value of the counter variable in the body of the loop. This is very important (and mostly forgotten). Failing to do so will result in an infinite loop (never-ending loop).
- Finally, the result is displayed.
```
i = 1
while i <= 6:
print(i)
#i += 1
i = i+ 1
n = 5
# initialize sum and counter
sum = 0
i = 1
while i <= n:
sum = sum + i
i = i+1 # update counter
# print the sum
print("The sum is", sum)
fruits = ["Mangoes","bananas","Oranges"]
for fruit in fruits:
print(fruit)
```
# B) For Loop
```
# Dictionary
my_data = {'Name': 'Uzair', 'Age': 24, 'Education': 'Masters'}
# my_data.items()
# for key in my_data.keys():
# print(key)
# # print()
# for value in my_data.values():
# print(value)
# # print()
for key, value in my_data.items():
print(key, value)
# List
lst = ["grapes", "apple", "banana"]
for item in lst:
print(item)
# Tuple
cars = ("UK", "Cricket", "Football","Paris")
for car in cars:
print(car)
```
# Nested Loops
```
Loop within a loop
adj = ["red", "big", "tasty"]
fruits = ["apple", "banana", "cherry"]
for x in adj:
for y in fruits:
pass
print(x, y)
1st Loop iteraion:
2nd loop 1st time
x = "red"
y ="apple"
2nd Loop 2nd time
x ="red"
y= "banana"
2nd Loop 3rd time
x ="red"
y= "cherry"
```
# BREAK
```
break
```
With the break statement we can stop the loop before it has looped through all the items
```
courses = ["ENG", "MATH", "BIO"]
for x in courses:
if x == "MATH":
break
print(x)
print('I am here')
```
# CONTINUE
```
continue
```
With the continue statement we can stop the current iteration of the loop, and continue with the next
```
courses = ["ENG", "MATH", "BIO","URDU"]
for x in courses:
if x == "BIO":
continue
print(x)
for x in range(10):
print(x)
```
# Assignment
```
# Solution
dict1 = {'a': 1, 'b': 2, 'c': 3, 'd': 4, 'e': 5}
even_list = []
odd_list = []
for val in dict1.values():
if val % 2 == 0:
even_list.append(val)
else:
odd_list.append(val)
print("Numbers in Even List are {}".format(even_list))
print("Numbers in Odd List are {}".format(odd_list))
```
```
text = '''
The video by digital artist Beeple, whose real name is Mike Winkelmann, was authenticated by blockchain, which serves as a digital signature to certify who owns it and that it is the original work.
It’s a new type of digital asset – known as a non-fungible token (NFT) – that has exploded in popularity during the pandemic as enthusiasts and investors scramble to spend enormous sums of money on items that only exist online.
Blockchain technology allows the items to be publicly authenticated as one-of-a-kind, unlike traditional online objects which can be endlessly reproduced.
'''
word_corpus = text.split()
unique_words = []
for word in word_corpus:
if word not in unique_words:
unique_words.append(word)
print(sorted(unique_words))
```
# ---- End of Session ----
| github_jupyter |
<p align="center">
<img src="https://github.com/GeostatsGuy/GeostatsPy/blob/master/TCG_color_logo.png?raw=true" width="220" height="240" />
</p>
## GeostatsPy: Spatial Bootstrap
### Michael Pyrcz, Associate Professor, University of Texas at Austin
##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
### A Demonstration of Spatial Boostrap
Here's a simple workflow to demonstrate spatial bootstrap and to compare with regular (non-spatial) bootstrap.
The workflow steps:
1. Build an initial sample set with $ndata$ samples. Specify locations. Sort the data in a data only array for
an equal weighted cumulative density function (CDF) for backtransform
2. Calculate the covariance matrix (between the data) for LU simulation. This provides a fast and easy method
to calculate spaitally correlated, unconditional, Gaussian realizations at the data locations.
3. Perform LU (lower upper) decomposition of the covariance matrix. Realizations only require L x random vector.
4. Draw from this initial sample set, with replacement, $ndata$ times to build a new realization of the sample.
Repeat this $nreal$ times to make realizations of the sample. Use LU simulation to ensure spaital correlation
between samples. Backtransform from Gaussian to original data values (use percentiles to sample).
5. Calculate the statistic of interest for each realization. This demonstration considers the mean only.
We could have considered any statistic including median, 13th percentile, skew etc.
6. (and 7.) Quantify and visualize uncertainty with histograms and summary statistics.
References:
Efron, 1982, The jackknife, the bootstrap, and other resampling plans, Society of Industrial and Applied Math,
CBMS-NSF Monographs, 38.
Journel, A. G. Resampling from stochastic simulations. Environmental and Ecological Statistics, 1:63–84, 1994.
#### Getting Started
Here's the steps to get setup in Python with the GeostatsPy package:
1. Install Anaconda 3 on your machine (https://www.anaconda.com/download/).
2. From Anaconda Navigator (within Anaconda3 group), go to the environment tab, click on base (root) green arrow and open a terminal.
3. In the terminal type: pip install geostatspy.
4. Open Jupyter and in the top block get started by copy and pasting the code block below from this Jupyter Notebook to start using the geostatspy functionality.
You will need to copy the data file to your working directory. They are available here:
* Tabular data - sample_data.csv at https://git.io/fh4gm.
There are exampled below with these functions. You can go here to see a list of the available functions, https://git.io/fh4eX, other example workflows and source code.
#### Load the required libraries
The following code loads the required libraries.
```
import os # to set current working directory
import sys # supress output to screen for interactive variogram modeling
import io
import numpy as np # arrays and matrix math
import pandas as pd # DataFrames
import matplotlib.pyplot as plt # plotting
from matplotlib.pyplot import cm # color maps
from matplotlib.patches import Ellipse # plot an ellipse
import math # sqrt operator
import scipy
import scipy.linalg # SciPy Linear Algebra Library
from scipy.stats import norm # Gaussian distribution
```
#### Set the working directory
I always like to do this so I don't lose files and to simplify subsequent read and writes (avoid including the full address each time).
```
os.chdir("c:/PGE383") # set the working directory
```
#### Assume Data Values and Locations
For a simple demonstration we assume 4 data values in 2D and make a CDF by sorting
* We will use the CDF for the back transform from Gaussian space.
```
ndata = 10 # number of data
vmin = 0.0; vmax = 10.0 # assume min and max values
data = np.zeros((ndata,3)) # x, y, value
data[0,0] = 25.0; data[0,1] = 50.0; data[0,2] = 2.3 # data 1
data[1,0] = 75.0; data[1,1] = 80.0; data[1,2] = 5.5 # data 2
data[2,0] = 10.0; data[2,1] = 25.0; data[2,2] = 1.0 # data 3
data[3,0] = 95.0; data[3,1] = 15.0; data[3,2] = 8.7 # data 4
data[4,0] = 25.0; data[4,1] = 53.0; data[4,2] = 3.7 # data 5
data[5,0] = 34.0; data[5,1] = 81.0; data[5,2] = 7.1 # data 6
data[6,0] = 72.0; data[6,1] = 68.0; data[6,2] = 8.2 # data 7
data[7,0] = 9.0; data[7,1] = 13.0; data[7,2] = 5.4 # data 8
data[8,0] = 95.0; data[8,1] = 92.0; data[8,2] = 6.2 # data 9
data[9,0] = 58.0; data[9,1] = 32.0; data[9,2] = 2.2 # data 9
print('The user specified data set:\n')
print(' X Y Value')
print(data)
cum_prob = np.zeros(ndata+2) # calculate the CDF (piece-wise linear)
var = np.zeros(ndata+2)
var[0] = vmin; var[ndata+1] = vmax
cum_prob[0] = 0.0; cum_prob[ndata+1] = 1.0
for i in range(1, ndata+1):
var[i] = data[i-1,2]
cum_prob[i] = i/(ndata+1)
var = np.sort(var)
print('\nThe data empirical CDF assuming known tails:')
plt.subplot(121)
im = plt.scatter(data[:,0],data[:,1],c=data[:,2],vmin=vmin,vmax=vmax,edgecolor='black',cmap=plt.cm.inferno)
plt.xlim([0,100]); plt.ylim([0,100])
cbar = plt.colorbar(im, orientation="vertical", ticks=np.linspace(vmin, vmax, 10))
cbar.set_label('Feature Value', rotation=270, labelpad=20)
plt.subplot(122)
plt.plot(var,cum_prob,c='red',zorder=1); plt.xlabel('Feature Value'); plt.xlim([vmin,vmax]); plt.ylim([0.0,1.0])
plt.scatter(var,cum_prob,edgecolor='black',c='red',zorder = 2)
plt.ylabel('Cumulative Probability'); plt.title('Feature CDF');
plt.grid();
plt.subplots_adjust(left=0.0, bottom=0.0, right=2.0, top=1.1, wspace=0.3, hspace=0.3)
plt.show()
```
#### Calculate the Covariance Matrix
The data-to-data covariance matrix is calculated from the distance matrix
* assuming a isotropic spherical variogram with range
```
cov = np.zeros((ndata,ndata)) # covarince matrix initialization
var_range = 100.0 # isotropic spherical variogram model range
for i in range(0, ndata):
for j in range(0, ndata):
distance = math.sqrt(math.pow((data[i,0]-data[j,0]),2) + math.pow((data[i,1]-data[j,1]),2))
cova = 0.0
if(distance < var_range):
hr = distance / var_range
cova = 1.0 - hr*(1.5 - 0.5*hr*hr) # spherical single, isotropic structure, no nugget
cov[i,j] = cova
print('The covariance matrix:\n')
print(np.round(cov,2))
```
#### Lower Upper (LU) Decomposition
Peform LU decomposition of the covariance matrix
* calculate the lower matrix $L$ from the covariance matrix.
```
P, L, U = scipy.linalg.lu(cov) # LU decomposition of the covariance matrix
print('The lower matrix:\n')
print(np.round(L,2))
```
#### Check the Lower Upper (LU) Decomposition of the Covariance Matrix
We can test the lower upper decomposition by multipling the lower and upper matrices and comparing them to the original covariance matrix.
```
test = cov - np.matmul(L,U) # check result should be close to zero
print('Covariance matrix minus the matrix multiplication of the lower and upper matrices:\n')
print(np.round(test,3))
```
#### Calculate Unconditional, Stationary Realizations at the Data Locations
The following steps:
1. Matrix multiplication of the nxn lower matrix with a 1D matrix of random standard normal values to calculate a realization with spatial correlation at data locations $i = 0,\ldots,n-1$.
2. Convert to p-values with the standard normal CDF
3. Transform to discrete values in the data CDF.
Realizations stored in output.
```
nreal = 10000 # number of bootstrap realizations
spatial_bootstrap = np.zeros((ndata,nreal)) # array to store spatial bootstrap results
for ireal in range(0, nreal): # loop over realizations
rand = np.random.normal(loc = 0.0, scale = 1.0, size = ndata) # random Gaussian values [ndata]
realization = np.matmul(L,rand) # impose spatial correlation between the values
pvalue = norm.cdf(realization) # calculate the p-values
spatial_bootstrap[:,ireal] = np.interp(pvalue,cum_prob,var,left=None, right=None, period=None) # backtransform ot feature values
```
#### Visualize Spatial Bootstrap Data Realizations
Let's make some histograms of several spatial bootstrap data realizations.
```
for ix in range(0,4):
plt.subplot(1,4,ix+1) # plot histograms of spatial bootstrap data realizations
plt.hist(spatial_bootstrap[:,ix],color='red',alpha=0.2,edgecolor='black',bins=np.linspace(vmin,vmax,20))
plt.xlim([vmin,vmax]); plt.ylim([0,4]); plt.xlabel('Average'); plt.ylabel('Frequency'); plt.title('Spatial Bootstrap Data Realization ' + str(ix+1) + '')
plt.subplots_adjust(left=0.0, bottom=0.0, right=4.0, top=1.1, wspace=0.3, hspace=0.3)
plt.show()
```
#### Calculate and Summarize the Statistic for Over All Spatial Bootstrap Realizations
Calculate the summary statistic over each spatial bootstrap data realization
* we show the L arithmetic averages calculated over the spatially correlated ndata realizations
```
summary_spatial_bootstrap = np.average(spatial_bootstrap,axis=0)
plt.hist(summary_spatial_bootstrap,color='red',alpha=0.2,edgecolor='black',bins=np.linspace(vmin,vmax,40))
plt.xlim([vmin,vmax]); plt.xlabel('Average'); plt.ylabel('Frequency'); plt.title('Spatial Bootstrap Uncertainty in the Average')
plt.subplots_adjust(left=0.0, bottom=0.0, right=1.0, top=1.1, wspace=0.3, hspace=0.3)
plt.show()
```
#### Calculate and Summarize the Statistic for Over All Regular Bootstrap Realizations and Compare with Spatial Bootstrap
Let's repeat the process with regular bootstrap and compare with our spatial bootstrap results.
* we will work in Gaussian space and backtransform with the data CDF for consistency
```
bootstrap = np.zeros((ndata,nreal)) # array to store spatial bootstrap results
for ireal in range(0, nreal): # loop over realizations
rand = np.random.normal(loc = 0.0, scale = 1.0, size = ndata) # random Gaussian values [ndata]
pvalue = norm.cdf(rand) # calculate the p-values
bootstrap[:,ireal] = np.interp(pvalue,cum_prob,var,left=None, right=None, period=None)
summary_bootstrap = np.average(bootstrap,axis=0)
plt.subplot(131)
im = plt.scatter(data[:,0],data[:,1],c=data[:,2],vmin=vmin,vmax=vmax,edgecolor='black',cmap=plt.cm.inferno)
plt.xlim([0,100]); plt.ylim([0,100]); plt.xlabel('X(m)'); plt.ylabel('Y(m)'); plt.title('Data Location Map')
cbar = plt.colorbar(im, orientation="vertical", ticks=np.linspace(vmin, vmax, 10))
cbar.set_label('Feature Value', rotation=270, labelpad=20)
plt.subplot(132)
plt.plot(var,cum_prob,c='red',zorder=1); plt.xlabel('Feature Value'); plt.xlim([vmin,vmax]); plt.ylim([0.0,1.0])
plt.scatter(var,cum_prob,edgecolor='black',c='red',zorder = 2)
plt.ylabel('Cumulative Probability'); plt.title('Feature CDF');
plt.grid();
plt.subplot(133)
plt.hist(summary_bootstrap,color='yellow',alpha=0.2,edgecolor='black',bins=np.linspace(vmin,vmax,40),label='Bootstrap')
plt.hist(summary_spatial_bootstrap,color='red',alpha=0.2,edgecolor='black',bins=np.linspace(vmin,vmax,40),label='Spatial Bootstrap')
plt.legend(loc = 'upper right')
plt.xlim([vmin,vmax]); plt.xlabel('Average'); plt.ylabel('Frequency'); plt.title('Bootstrap and Spatial Bootstrap Uncertainty in the Average')
plt.subplots_adjust(left=0.0, bottom=0.0, right=3.0, top=1.1, wspace=0.3, hspace=0.3)
plt.show()
```
#### Comments
This was a basic workflow for spatial and regular bootstrap for uncertainty in a samples statistic through sampling multiple data realizations.
I have other demonstrations on the basics of working with DataFrames, ndarrays, univariate statistics, plotting data, declustering, data transformations, trend modeling and many other workflows available at https://github.com/GeostatsGuy/PythonNumericalDemos and https://github.com/GeostatsGuy/GeostatsPy.
I hope this was helpful,
*Michael*
#### The Author:
### Michael Pyrcz, Associate Professor, University of Texas at Austin
*Novel Data Analytics, Geostatistics and Machine Learning Subsurface Solutions*
With over 17 years of experience in subsurface consulting, research and development, Michael has returned to academia driven by his passion for teaching and enthusiasm for enhancing engineers' and geoscientists' impact in subsurface resource development.
For more about Michael check out these links:
#### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#### Want to Work Together?
I hope this content is helpful to those that want to learn more about subsurface modeling, data analytics and machine learning. Students and working professionals are welcome to participate.
* Want to invite me to visit your company for training, mentoring, project review, workflow design and / or consulting? I'd be happy to drop by and work with you!
* Interested in partnering, supporting my graduate student research or my Subsurface Data Analytics and Machine Learning consortium (co-PIs including Profs. Foster, Torres-Verdin and van Oort)? My research combines data analytics, stochastic modeling and machine learning theory with practice to develop novel methods and workflows to add value. We are solving challenging subsurface problems!
* I can be reached at mpyrcz@austin.utexas.edu.
I'm always happy to discuss,
*Michael*
Michael Pyrcz, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
I have other demonstrations on the basics of working with DataFrames, ndarrays, univariate statistics, plotting data, declustering, data transformations, trend modeling and many other workflows available at https://github.com/GeostatsGuy/PythonNumericalDemos and https://github.com/GeostatsGuy/GeostatsPy. Also, I have posted a lot of other example workflows and source code to help you get learn about and get work done with **Spatial Data Analytics, Geostatistics and Machine Learning** in Python and R. Check out the [GeostatsGuy Inventory](https://github.com/GeostatsGuy/Resources) and I have a [YouTube](https://youtube.com/GeostatsGuyLectures) channel with all my university lectures recorded.
I hope this was helpful,
*Michael*
Michael Pyrcz, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
#### More Resources Available at: [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
| github_jupyter |
# GPU-Accelerated Numerical Computing with MatX
<img src="img/dli-matx-overview.png">
## Tutorial List
1. Introduction (this tutorial)
2. [Operators](02_operators.ipynb)
3. [Executors](03_executors.ipynb)
4. [Radar Pipeline Example](04_radar_pipeline.ipynb)
## Introduction
Welcome to NVIDIA's MatX training! In this course, you will learn how to use MatX, a modern C++ header-only library designed for generic matrix and n-dimensional array (tensor) operations. By borrowing heavily from both Python and MATLAB syntax, MatX is designed to provide native CUDA performance all behind a friendly and extensible API. Whereas other GPU-accelerated numerical computing libraries like [CuPy](https://cupy.dev/) and [Numba](http://numba.pydata.org/) allow the Python developer to build GPU-centric applications, MatX extends this "quick build, fast performance" thesis to the C++ developer. Our ultimate goal is to encourage developer productivity and provide a quick on-ramp from prototype code to statically-typed production code.
## Training Structure
This training contains a series of tutorials in increasing order of name that will guide you through basic and intermediate MatX features. Most tutorials will require you to open the source code, make a small change, and run it from the Jupyter notebook. As time permits, there is another notebook called `99_assignments.ipynb` that will give a series of problems to solve using MatX primitives. The assignments have verification code to make sure the answer is correct.
## Motivation
Writing high-performance GPU code usually requires writing CUDA code directly and having at least a basic knowledge of the underlying hardware. Users not wanting to invest the time to learn this will typically use higher-level languages with CUDA bindings, such as MATLAB or Python. Using these high-level languages may cause interoperability with existing libraries and software more difficult. High-level languages can also cause a performance penalty for applications that require the lowest latency or highest throughput.
MatX aims to provide syntax familiar to high-level languages with the performance and interoperability of C++. Most of the runtime costs are transferred into compile-time costs by using C++ templates, and kernels are auto-generated for most tensor and domain-specific functions. MatX does not require any CUDA or hardware knowledge; users can write algebraic expressions or use simple functions on tensor objects without every writing CUDA code.
## MatX Structure
MatX is structured into two separate, but compatible, APIs: tensor and frontend. The tensor API provides a suite of mathematical operations for operating on MatX tensor types. These operations are evaluated at compile-time and can be combined into chained expressions to improve performance. The frontend APIs are simple interfaces into numerous existing CUDA libraries. These include cuFFT, cuBLAS, cuSolver, and more. A common tensor type can be passed seamlessly between the tensor and frontend APIs, and MatX uses the information from these operations to call the appropriate kernels or libraries.
One of the key concepts in MatX's design and usage is the disassociation of memory allocation from processing. We will cover this concept in depth during the training, but MatX optimizes primitive performance by discouraging operations that negatively impact performance, such as deep copies. When creating an application, the user creates views into memory regions that provide zero-cost abstractions of looking at the data differently in memory.
## Terminology
The basic building block of all MatX operations is the tensor. Having the common tensor type for all operations provides a powerful tool for manipulating data, and more importantly, abstracts much of the complexity from the user. MatX does not use traditional tensor notation (Einstein), and instead treats a tensor simply as scalars (0-dimensional tensors), vectors (1-dimensional tensors), or multi-dimensonal matrices (n-dimensional tensors). Tensors have three main properties: rank, size, and type. The rank and type are specified at compile time, while the size can be specified at either runtime or compile-time. The rank describes how many dimensions the tensor has. The size of the tensor describes how many elements are in each dimension. Lastly, the type of the tensor is the type of data stored in the tensor. Different data types will change the size and capabilities of the tensor.
- **Tensors**
Tensors (`tensor_t`) provide views of the underlying tensor data and optionally allocate managed memory. Tensors describe properties such as stride, size of each dimension, and provide basic accessor functions for retrieving the data in a view. Tensors can be created from other tensors when convenient, which allows a view into another tensor without copying any data. For example, a sliced view of a tensor (which we'll get to soon) can be used to generate two more tensors of both real and imaginary parts of a complex number. The term `view` is sometimes interchangeably used with the term `tensor` in MatX since all tensors are simply a view into underlying memory. The memory backing tensors is reference counted; the last tensor to be destroyed either explicitly or implicitly will free any non-user-managed data.
<br>
- **Operators**
Operators are an abstract term used for types that can return a value at a given index. Currently there are three types of operators: tensors, operator expressions, and generators. Operators are covered in more detail in lesson 2.
## Our First Example
In this example, we give an introduction to the MatX library by performing basic operations on tensor objects. We will show the following operations on tensors:
0. Library Import
1. Creation
2. Initialization
3. Permuting (Rearrange dimensions of tensor)
4. Slicing
5. Cloning
All of these operations are on tensor object types only, and do not use any of the frontend API. If desired, open `example1.cu` in a separate tab to view the entire file. Let's walk through the example line-by-line.
### 0. Library Import
During this tutorial, we will be writing our MatX code in a CUDA file (`.cu`), compiled via the CUDA Compiler, `nvcc`. If you're curious, the specific command line to build and execute code can be found [here](exercises/compile_and_run.sh).
When using MatX, be sure to import the library via:
```c++
#include <matx.h>
using namespace matx;
```
### 1. Creation
First, we create a `tensor_t` object:
```c++
tensor_t<int, 2> t2({5,4});
```
tensor_t objects take two template parameters to instantiate:
1. The type of the underlying data
2. The rank of the tensor
On this line, we are creating a 2D tensor (matrix) of integer values. These template parameters are required for all tensor objects to allow compile-time optimizations. The constructor of `tensor_t` takes a `tensorShape_t` object to specify the size of each of the two dimensions. Creating a shape can be done with an initializer list as well, so we construct the shape object using this shortened format. In this case, we are asking for a 5x4 2D tensor, where 5 is the number of rows and 4 is the number of columns.
**NOTE** Unlike MATLAB, MatX follows the C-style for indexing, meaning we assume row-major formats rather than column-major, and 0-based indexing rather than 1-based.
By not passing extra arguments, we are asking the tensor object to allocate the memory using CUDA managed memory so that it is visible by both host and device code. Alternatively, users can opt to manage their own memory using other forms of the constructor (see documentation).
### 2. Initialization
After allocating the tensor, we initialize the underlying data:
```c++
t2.SetVals({ {1, 2, 3, 4},
{5, 6, 7, 8},
{9, 10, 11, 12},
{13, 14, 15, 16},
{17, 18, 19, 20}});
t2.PrefetchDevice(0);
```
The tensor is initialized using a nested initializer list inside of the `SetVals` member function, specifying the values of the matrix. The initializer list is a single-nested list to match a 2D tensor shape, but this can be extended up to 4D tensors. `operator()` is also available to set and get individual values of a tensor as an alternative:
```c++
t2(0,0) = 1;
t2(0,1) = 2;
...
```
The next call to `PrefetchDevice(0)` ensures that any data that may have been set on the host is now copied to the device. The `0` parameter is the CUDA stream, and for this example, we're operating only with the default stream (0). By default, prefetching will instruct the CUDA runtime to prefetch all data visible to the current view. After this step completes, the initialized data is visible on both the host and device. Note that if the user is not using managed memory, `operator()` to set values is not available on the host, and instead the values should be copied from a host-side tensor.
The prefetch line can be ommited and the program will still work correctly, but the first time `t2` is accesed on the device will trigger a page fault, and the data would be moved automatically later.
On the next line we print the size of the tensor dimensions and the current data inside the tensor:
```
t2.Print();
```
`Print` is a utility function of a tensor to print a tensor's contents to stdout. Printing tensors can be used with data that resides either on the host or device, and MatX will determine this at runtime. With no arguments it will print the entire contents of the tensor. However, the size of the printing can also be limited by passing a limit to each dimension. For example, `Print(3,2)` would print the first 2 columns and 3 rows of the 2D tensor. The contents of the tensor printed should appear as an increasing sequence of numbers from the top to bottom rows.
Open the file [exercises/example1_init.cu](exercises/example1_init.cu) and edit the contents where you see TODO markers.
```
!./exercises/compile_and_run.sh example1_init
```
Expected output:
```sh
000000: 1 2 3 4
000001: 5 6 7 8
000002: 9 10 11 12
000003: 13 14 15 16
000004: 17 18 19 20
```
### 3. Permute
The next section calls `Permute` on the returned view:
```c++
t2p = t2.Permute({1,0});
t2p.Print();
```
`Permute` returns a view of the data with the dimensions swapped to match the order of the initializer list argument. In this case there are only two dimensions being permuted on a 2D tensor, so it's equivalent to a matrix transpose. However, `Permute` can be used on higher-order tensors with the dimensions swapped in any particular order. Observe the data and size of the tensor is now transposed when using this view:

Open the file [exercises/example1_permute.cu](exercises/example1_permute.cu) and edit the contents where you see TODO markers.
```
!./exercises/compile_and_run.sh example1_permute
```
Expected output:
```sh
000000: 1 5 9 13 17
000001: 2 6 10 14 18
000002: 3 7 11 15 19
000003: 4 8 12 16 20
```
Note that none of the underlying data has been modified. A permuted view simply accesses the data as if it were transposed, but the actual order of data in memory has not changed. This can be confirmed by re-printing the previous non-transposed view. By not modifying the data in memory, a permuted view *may* be slower to access than a non-permuted view with contiguous entries. In general, permuted views are useful for infrequent accesses, but if the transposed data would be accessed repeatedly it may be faster to permute the data in memory using the `transpose` operator.
### 4. Slice
The next line takes a slice of the 2D tensor by selecting a subset of data in both dimensions:
```c++
t2s = t2.Slice({1,1}, {3, 3});
```
`t2s` is now a view of the same data, but starting at index 1 and ending at index 3 (exclusive) on both dimensions. This is equivalent to Python using `t2[1:3, 1:3]`. Since a new sliced view is returned, the new view will have dimensions `{2, 2}`.

Open the file [exercises/example1_simple_slice.cu](exercises/example1_simple_slice.cu) and edit the contents where you see TODO markers.
```
!./exercises/compile_and_run.sh example1_simple_slice
```
Expected output:
```sh
000000: 6 7
000001: 10 11
```
The next line shows a variant of `Slice` that can reduce the dimension of a tensor:
```c++
auto t1 = t2.Slice<1>({0, 1}, {matxEnd, matxDropDim});
```
Using this form of `Slice` requires a template argument with the rank of the new slice. The first parameter to `Slice` takes the starting index for each dimension, while the second takes the ending index. To include all values from the beginning on, a special sentinel of `matxEnd` can be used. Similarly, `matxDropDim` is used to indicate this dimension is the one being sliced (i.e. removed). In this case we are slicing the second column of the tensor and all rows, which produces a new 1D tensor containing only the second column of the original tensor. This is equivalent to `t2[:,1]` in Python.

Open the file [exercises/example1_adv_slice_col.cu](exercises/example1_adv_slice_col.cu) and edit the contents where you see TODO markers.
```
!./exercises/compile_and_run.sh example1_adv_slice_col
```
Expected output:
```sh
000000: 2
000001: 6
000002: 10
000003: 14
000004: 18
```
Instead of slicing a single column, we can also slice a single row:
```c++
auto t1 = t2.Slice<1>({1,0}, {matxDropDim, matxEnd});
```

Open the file [exercises/example1_adv_slice_row.cu](exercises/example1_adv_slice_row.cu) and edit the contents where you see TODO markers.
```
!./exercises/compile_and_run.sh example1_adv_slice_row
```
Expected output:
```sh
000000: 5
000001: 6
000002: 7
000003: 8
```
Note that since we reduced the dimension to a 1D tensor in both cases, printing a 1D tensor (vector) will appear the same in the direction the values are printed.
### 5. Clone
The last line shows `Clone`, which replicates a tensor's dimensions into a higher-rank tensor:
```c++
auto t2c = t1.Clone<2>({5, matxKeepDim});
```
`Clone` is used on a 1D tensor from the output of the previous example, and replicates the data of the `t1` vector into a 2D tensor with 5 rows where all rows match the data in `t1`. Cloning does not replicate the data in memory; instead, the same elements in `t1` are accessed repeatedly when different rows are accessed. This not only saves memory, but also benefits from the caches in the GPU by not hitting different addresses in memory for the same value.
In this case `Clone` was being used on a 1D view from a 2D tensor data set, but similar code works on taking any dimension tensor and increasing it to a higher dimension. The increase in dimensions is not restricted to one. For example, a scalar (0D tensor) can be cloned into a 4F tensor where a single value in memory would appear as a 4D tensor.

Open the file [exercises/example1_clone.cu](exercises/example1_clone.cu) and edit the first TODO.
```
!./exercises/compile_and_run.sh example1_clone
```
Expected output:
```sh
000000: 1 2 3 4
000001: 1 2 3 4
000002: 1 2 3 4
000003: 1 2 3 4
000004: 1 2 3 4
```
By changing which dimension is cloned, we can also take the same 1D tensor across columns. Edit the last file and clones across columns instead, and print the output of the cloned view.

Expected output:
```sh
000000: 1 1 1 1 1
000001: 2 2 2 2 2
000002: 3 3 3 3 3
000003: 4 4 4 4 4
```
As mentioned at the beginning, views do not modify the underlying data; they simply provide the metadata needed to access the elements. To show this, we will open the same [exercise](exercises/example1_clone.cu) again and modify the first value in the original 1D tensor to 10 and watch how multiple elements of the cloned view are modified.
```
!./exercises/compile_and_run.sh example1_clone
```
Expected output:
```sh
000000: 10 2 3 4
000001: 10 2 3 4
000002: 10 2 3 4
000003: 10 2 3 4
000004: 10 2 3 4
```
This concludes the first tutorial on MatX. In this tutorial, you learned the basics of accessing, slicing, permuting, and cloning a tensor. In the next example you will learn about operators and how to apply them to tensors.
[Start Next Tutorial](02_operators.ipynb)
| github_jupyter |
# Pandas Visualization
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
df1 = pd.read_csv('df1.csv', index_col = 0)
df1.head()
df2 = pd.read_csv('df2.csv')
df2.head()
```
## Historgram
```
df1['A'].hist(bins= 30)
df1['A'].plot(kind ='hist', bins = 30)
df1['A'].plot.hist(bins = 30)
df2
```
## Area Plot
```
df2.plot.area(alpha = 0.4)
```
## Bar Plot
```
# index will be the x-axis
df2.plot.bar()
df2.plot.bar(stacked = True)
df1['A'].plot.hist(bins = 50)
```
## Line Plot
```
df1.plot.line(x = df1.index, y = 'B', figsize=(12,3), lw = 0.5)
```
## Scatter Plot
```
df1.plot.scatter(x='A', y='B', c = 'C', cmap= 'coolwarm')
df1.plot.scatter(x='A', y='B', s = df1['C'] * 10) # s = size
```
## Box Plot
```
df2.plot.box()
df = pd.DataFrame(np.random.randn(1000,2), columns=['a', 'b'])
df.head(10)
```
## Hexbin Plot
```
df.plot.hexbin(x = 'a', y = 'b')
df.plot.hexbin(x = 'a', y = 'b', gridsize = 25, cmap = 'coolwarm')
```
## Kernel Density Estimation Plot
```
df2['a'].plot.kde()
df2['a'].plot.density()
df2.plot.kde()
```
# Pandas Visualization continued.... 2
```
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
mcdon = pd.read_csv('mcdonalds.csv', index_col = 'Date', parse_dates = True)
mcdon.head()
# Note the Adj. Volume is way scaled-up than Adj. Close, hence the plot is not so nice
mcdon.plot()
mcdon['Adj. Close'].plot()
mcdon['Adj. Volume'].plot(figsize=(12,4))
mcdon['Adj. Close'].plot(xlim = ['2007-01-01', '2009-01-01']) #xlimit
#xlimit and ylimit , [list] or (tuple) both accepted
mcdon['Adj. Close'].plot(xlim = ['2007-01-01', '2009-01-01'], ylim = (20,50))
mcdon['Adj. Close'].plot(xlim = ['2007-01-01', '2009-01-01'], ylim = (20,50), ls = '--', c='red')
import matplotlib.dates as pltdates
mcdon['Adj. Close'].plot(xlim = ['2007-01-01', '2009-01-01'], ylim = (20,50))
idx = mcdon.index
idx
idx = mcdon.loc['2007-01-01':'2007-05-01'].index
idx
stock = mcdon.loc['2007-01-01':'2007-05-01']['Adj. Close']
stock
fig, ax = plt.subplots()
ax.plot_date(idx, stock,'-')
ax.yaxis.grid(True)
ax.xaxis.grid(True)
fig.autofmt_xdate() # automatically format the x-date axis
plt.tight_layout()
```
## Date formating using major_locator, major_formatter, minor_locator, minor_locator
```
fig, ax = plt.subplots()
ax.plot_date(idx, stock,'-')
ax.xaxis.grid(True)
ax.yaxis.grid(True)
# http://strftime.org/
ax.xaxis.set_major_locator(pltdates.MonthLocator())
ax.xaxis.set_major_formatter(pltdates.DateFormatter('\n\n%b-%Y'))
ax.xaxis.set_minor_locator(pltdates.WeekdayLocator(byweekday=0))
ax.xaxis.set_minor_formatter(pltdates.DateFormatter('%d')) # try %a
fig.autofmt_xdate() # automatically format the x-date axis
plt.tight_layout()
```
| github_jupyter |
```
import torch
import torch.nn.functional as F
import torchsde
import math
import matplotlib.pyplot as plt
import numpy as np
from tqdm.notebook import tqdm
# from torch import datasets
from torch import _vmap_internals
from torchvision import datasets, transforms
# import torch.nn.functional as F
import pandas as pd
from cfollmer.objectives import log_g, relative_entropy_control_cost, stl_relative_entropy_control_cost_xu
from cfollmer.sampler_utils import FollmerSDE
from cfollmer.drifts import *
from cfollmer.trainers import basic_batched_trainer
```
# The Model
\begin{align}
\theta &\sim \mathcal{N}(\theta | 0, \sigma_w^2 \mathbb{I}) \\
y_i | x_i, \theta &\sim \mathrm{Bernouli}\left[\mathrm{NN}_{\theta}\left(x_i \right)\right]
\end{align}
We want samples from $p(\theta | \{(y_i, x_i)\})$. Note $f(x; \theta)$ is a neural net with params $\theta$
## Loading the iris dataset
```
images_train = datasets.MNIST("../data/mnist/", download=True, train=True)
images_test = datasets.MNIST("../data/mnist/", download=True, train=False)
transform = torch.nn.Sequential(transforms.Normalize((0.1307,), (0.3081)))
X_train, y_train = images_train.data, images_train.targets
X_test, y_test = images_test.data, images_test.targets
X_train = torch.flatten(transform(X_train.float()), 1)
X_test = torch.flatten(transform(X_test.float()), 1)
y_train = F.one_hot(y_train)
y_test = F.one_hot(y_test)
# X_train = np.concatenate((X_train, np.ones((X_train.shape[0],X_train.shape[1]))), axis=1)
# X_test = np.concatenate((X_test, np.ones((X_test.shape[0],X_train.shape[1]))), axis=1)
device = "cuda" if torch.cuda.is_available() else "cpu"
X_train, X_test, y_train, y_test = \
torch.tensor(X_train, dtype=torch.float32, device=device), \
torch.tensor(X_test, dtype=torch.float32, device=device), \
torch.tensor(y_train, dtype=torch.float32, device=device), \
torch.tensor(y_test, dtype=torch.float32, device=device)
X_train.shape
```
$$\DeclareMathOperator*{\argmin}{arg\,min}$$
$$\def\E{{\mathbb{E}}}$$
$$\def\rvu{{\mathbf{u}}}$$
$$\def\rvTheta{{\bm{\Theta}}}$$
$$\def\gU{{\mathcal{U}}}$$
$$\def\mX{{\mathbf{X}}}$$
## Controlled Schrodinger Follmer Sampler
The objevtive we are trying to implement is:
\begin{align}
\mathbf{u}_t^{*}= \argmin_{\rvu_t \in \mathcal{U}}\mathbb{E}\left[\frac{1}{2\gamma}\int_0^1||\rvu(t, \Theta_t)||^2 dt - \ln\left(\frac{ p(\mX | \Theta_1)p(\Theta_1)}{\mathcal{N}(\Theta_1|\mathbf{0}, \gamma \mathbb{I} )}\right)\right] \
\end{align}
Where:
\begin{align}
d\Theta_t = \rvu(t, \Theta_t)dt + \sqrt{\gamma} dB_t
\end{align}
To do so we use the EM discretisation.
```
import torch.nn.functional as F
class ClassificationNetwork(object):
def __init__(
self, input_dim=1, output_dim=1, depth=None,
width=20, width_seq=None, device="cpu", activation=F.relu
):
self.device = device
self.output_dim = output_dim
self.input_dim = input_dim
self.activation = activation
self.depth = depth
if not self.depth:
self.depth = 1
if not width_seq:
self.width = width
self.width_seq = [self.width] * (self.depth + 1)
self.shapes = [(self.width_seq[i-1], self.width_seq[i]) for i in range(1,self.depth)]
self.shapes += [(self.width_seq[-1], self.output_dim)]
self.shapes = [(self.input_dim, self.width_seq[0])] + self.shapes
self.dim = sum([wx * wy + wy for wx, wy in self.shapes])
def forward(self, x, Θ):
index = 0
n, d = x.shape
# dim_bl = sum([wx * wy + wy for wx, wy in self.shapes[:-1]])
# Θ[:dim_bl] = (Θ[:dim_bl] - Θ[:dim_bl].mean()) / Θ[:dim_bl].std()
# σ_Θ, μ_Θ = Θ.std(), Θ.mean()
# Θ = (Θ - μ_Θ) / σ_Θ
for wx, wy in self.shapes[:-1]:
x = F.linear(
x,
Θ[index: index + wx * wy].reshape(wy, wx),
Θ[index + wx * wy: index + wx * wy + wy].reshape(1,wy)
)
x = self.activation(x)
index += wx * wy + wy
wx, wy = self.shapes[-1]
x = F.linear(
x,
Θ[index: index + wx * wy].reshape(wy, wx), #* σ_Θ + μ_Θ,
Θ[index + wx * wy: index + wx * wy + wy].reshape(1,wy) # * σ_Θ + μ_Θ
)
return x.to(self.device)
def map_forward(self, x, Θ):
preds_func = lambda θ: self.forward(x, θ)
batched_preds = torch._vmap_internals.vmap(preds_func)
preds = torch.hstack(list(map(preds_func, Θ)))
return preds
dim = X_train.shape[1]
out_dim = y_train.shape[1]
net = ClassificationNetwork(
dim, out_dim, device=device, depth=1, width=50, activation=F.tanh
)
def gaussian_prior(Θ, σ_w=3.8):
"""
Logistic regresion bayesian prior
"""
return -0.5 * (Θ**2).sum(axis=1) / σ_w
def log_likelihood_vmap_nn(Θ, X, y, net=net):
"""
Hoping this implementation is less buggy / faster
still feels a bit slow.
"""
def loss(θ):
preds = net.forward(X, θ)
cel = torch.nn.CrossEntropyLoss(reduction="sum")
# import pdb; pdb.set_trace()
ll_cel = -1.0 * cel(preds, y.argmax(dim=1))
return ll_cel
batched_loss = torch._vmap_internals.vmap(loss)
return batched_loss(Θ)
net.dim
class SimpleForwardNetBN_larger(AbstractDrift):
def __init__(self, input_dim=1, width=300, activation=torch.nn.Softplus):
super(SimpleForwardNetBN_larger, self).__init__()
self.nn = torch.nn.Sequential(
torch.nn.Linear(input_dim + 1, width), torch.nn.BatchNorm1d(width, affine=False), activation(),
torch.nn.Linear(width, width), torch.nn.BatchNorm1d(width, affine=False), activation(),
torch.nn.Linear(width, width), torch.nn.BatchNorm1d(width, affine=False), activation(),
torch.nn.Linear(width, width), torch.nn.BatchNorm1d(width, affine=False), activation(),
torch.nn.Linear(width, input_dim )
)
self.nn[-1].weight.data.fill_(0.0)
γ = 0.1**2
Δt=0.01
dim= net.dim
prior = gaussian_prior
sde, losses = basic_batched_trainer(
γ, Δt, prior, log_likelihood_vmap_nn, dim, X_train, y_train,
method="euler", stl="stl_xu", adjoint=False, optimizer=None,
num_steps=79, batch_size_data=int(X_train.shape[0] // 5), batch_size_Θ=30,
batchnorm=True, device=device, lr=0.0001, drift=SimpleForwardNetBN_larger, schedule="uniform",
γ_min= 0.1**2, γ_max= 0.4**2
)
losses
plt.plot(losses[:])
X_train.shape
t_size = int(math.ceil(1.0/Δt))
ts = torch.linspace(0, 1, t_size).to(device)
no_posterior_samples = 100
Θ_0 = torch.zeros((no_posterior_samples, net.dim)).to(device)
Θ_1 = torchsde.sdeint(sde, Θ_0, ts, dt=Δt)[-1,...]
fig, (ax1,ax2,ax3) = plt.subplots(1,3)
ax1.hist(Θ_1[:,0].cpu().detach().numpy())
ax2.hist(Θ_1[:,1].cpu().detach().numpy())
ax3.hist(Θ_1[:,2].cpu().detach().numpy())
def predc(X, Θ):
return torch.vstack([(net.forward(X, θ)[None,...]).softmax(dim=-1) for θ in Θ]).mean(dim=0)
pred = predc(X_train, Θ_1)
pred.shape
((pred.argmax(dim=-1)).float().flatten()== y_train.argmax(dim=-1)).float().mean()
pred_test = predc(X_test.float(), Θ_1)
((pred_test.argmax(dim=-1)).float().flatten()== y_test.argmax(dim=-1)).float().mean()
```
## MAP Baseline
We run the point estimate approximation (Maximum a posteriori) to double check what the learned weights look like. We get the exact same training accuracy as with the controlled model and similarly large weights for the non bias weights.
```
Θ_map = torch.zeros((1, dim), requires_grad=True, device=device)
optimizer_map = torch.optim.Adam([Θ_map], lr=0.05)
# optimizer = torch.optim.LBFGS(gpr.parameters(), lr=0.01)
losses_map = []
num_steps = 1000
for i in tqdm(range(num_steps)):
optimizer_map.zero_grad()
if isinstance(optimizer_map, torch.optim.LBFGS):
def closure_map():
loss_map = log_likelihood_vmap()
optimizer_map.zero_grad()
loss_map.backward()
return loss
optimizer_map.step(closure_map)
losses_map.append(closure_map().item())
else:
loss_map = -(log_likelihood_vmap(Θ_map, X_train, y_train) + gaussian_prior(Θ_map))
optimizer_map.zero_grad()
loss_map.backward()
print(loss_map.item())
optimizer_map.step()
losses_map.append(loss_map.item())
Θ_map
pred_map = torch.sigmoid(X_train.mm(Θ_map.T)).mean(axis=1)
((pred_map < 0.5).float() == y_train).float().mean(), Θ_map
```
| github_jupyter |
# Tensorflow ImageNet Pixel Threshold demo
>⚠️ **Warning:** This demo assumes that you have access to an on-prem deployment of Dioptra that provides a copy of the ImageNet dataset and a CUDA-compatible GPU.
> This demo cannot be run on a typical personal computer.
This notebook contains a demonstration of how to use Dioptra to run experiments that investigate the effects of the pixel threshold attack when launched on a neural network model trained on the ImageNet dataset.
## Setup
**Note:** This demo is specifically for the NCCoE DGX Workstation with hostname `dgx-station-2`.
Port forwarding is required in order to run this demo.
The recommended port mapping is as follows:
- Map `*:20080` on laptop to `localhost:30080` on `dgx-station-2`
- Map `*:25000` on laptop to `localhost:35000` on `dgx-station-2`
A sample SSH config file that enables the above port forwarding is provided below,
> ⚠️ **Edits required**: replace `username` with your assigned username _on the NCCoE virtual machines_!
```conf
# vm hostname: jumphost001
Host nccoe-jumphost001
Hostname 10.33.53.98
User username # Change to your assigned username on the NCCoE virtual machines!
Port 54131
IdentityFile %d/.ssh/nccoe-vm
# vm hostname: dgx-station-2
Host nccoe-k8s-gpu002
Hostname 192.168.1.28
User username # Change to your assigned username on the NCCoE virtual machines!
Port 22
IdentityFile %d/.ssh/nccoe-vm
ProxyJump nccoe-jumphost001
LocalForward *:20080 localhost:30080
LocalForward *:25000 localhost:35000
```
Now, connect to the NCCoE VPN and SSH into the DGX Workstation,
```bash
ssh nccoe-k8s-gpu002
```
Next, we import the necessary Python modules and ensure the proper environment variables are set so that all the code blocks will work as expected.
> ⚠️ **Edits possibly required**: update the value of the `HOST_DOCKER_INTERNAL` variable
If this notebook is being served to you via Docker (i.e. you ran `make jupyter` to launch this notebook), **then you may need to change the value assigned to the variable `HOST_DOCKER_INTERNAL`** to make the port forwarding you configured in the previous step accessible within the container.
The value you need to assign to the variable to depends on your host device's operating system:
- **Case 1: Host operating system is Windows 10 or MacOS**
- Set `HOST_DOCKER_INTERNAL = "host.docker.internal"`. This is the default setting.
- **Case 2: Host operating system is Linux**
- Run either `ip address` or `ifconfig` to print a list of the available network interfaces on your host device
- Locate the `docker0` interface and take note of the associated IP address (this is commonly set to `172.17.0.1`)
- Set `HOST_DOCKER_INTERNAL` equal to the IP address for the `docker0` interface. So, if the IP address was `172.17.0.1`, then you would set `HOST_DOCKER_INTERNAL = "172.17.0.1"`
If you started your Jupyter Lab instance from a conda environment, then you do not need to change anything.
The code below uses an environment variable to check whether this notebook is being served via the `jupyter` service, and if that variable isn't found, then the connection address reverts to `localhost` and ignores the `HOST_DOCKER_INTERNAL` variable.
```
# Import packages from the Python standard library
import os
import pprint
import time
import warnings
from pathlib import Path
from typing import Tuple
# Filter out warning messages
warnings.filterwarnings("ignore")
# Address for connecting the docker container to exposed ports on the host device
HOST_DOCKER_INTERNAL = "host.docker.internal"
# HOST_DOCKER_INTERNAL = "172.17.0.1"
# Testbed API ports
RESTAPI_PORT = "30080"
MLFLOW_TRACKING_PORT = "35000"
# Default address for accessing the RESTful API service
RESTAPI_ADDRESS = (
f"http://{HOST_DOCKER_INTERNAL}:{RESTAPI_PORT}"
if os.getenv("IS_JUPYTER_SERVICE")
else f"http://localhost:{RESTAPI_PORT}"
)
# Override the AI_RESTAPI_URI variable, used to connect to RESTful API service
os.environ["AI_RESTAPI_URI"] = RESTAPI_ADDRESS
# Default address for accessing the MLFlow Tracking server
MLFLOW_TRACKING_URI = (
f"http://{HOST_DOCKER_INTERNAL}:{MLFLOW_TRACKING_PORT}"
if os.getenv("IS_JUPYTER_SERVICE")
else f"http://localhost:{MLFLOW_TRACKING_PORT}"
)
# Override the MLFLOW_TRACKING_URI variable, used to connect to MLFlow Tracking service
os.environ["MLFLOW_TRACKING_URI"] = MLFLOW_TRACKING_URI
# Base API address
RESTAPI_API_BASE = f"{RESTAPI_ADDRESS}/api"
# Path to workflows archive
WORKFLOWS_TAR_GZ = Path("workflows.tar.gz")
# Experiment name (note the username_ prefix convention)
EXPERIMENT_NAME = "jtsexton_imagenet_pixel_threshold"
# Path to dataset
data_path_imagenet = "/nfs/data/ImageNet-Kaggle-2017/images/ILSVRC/Data/CLS-LOC"
# Import third-party Python packages
import numpy as np
import requests
from mlflow.tracking import MlflowClient
# Import utils.py file
import utils
# Create random number generator
rng = np.random.default_rng(54399264723942495723666216079516778448)
```
## Submit and run jobs
The entrypoints that we will be running in this example are implemented in the Python source files under `src/` and the `MLproject` file.
To run these entrypoints within the testbed architecture, we need to package those files up into an archive and submit it to the Testbed RESTful API to create a new job.
For convenience, the `Makefile` provides a rule for creating the archive file for this example, just run `make workflows`,
```
%%bash
# Create the workflows.tar.gz file
make workflows
```
To connect with the endpoint, we will use a client class defined in the `utils.py` file that is able to connect with the Testbed RESTful API using the HTTP protocol.
We connect using the client below, which uses the environment variable `AI_RESTAPI_URI` to figure out how to connect to the Testbed RESTful API,
```
restapi_client = utils.SecuringAIClient()
```
We need to register an experiment under which to collect our job runs.
The code below checks if the relevant experiment exists.
If it does, then it just returns info about the experiment, if it doesn't, it then registers the new experiment.
```
response_experiment = restapi_client.get_experiment_by_name(name=experiment)
if response_experiment is None or "Not Found" in response_experiment.get("message", []):
response_experiment = restapi_client.register_experiment(name=experiment)
response_experiment
```
We should also check which queues are available for running our jobs to make sure that the resources that we need are available.
The code below queries the Lab API and returns a list of active queues.
```
restapi_client.list_queues()
```
Next, we need to create our model.
In this example, we will be using an existing ImageNet classifier, and not training a new one. So, we use the ```init_model``` entry point to simply initialize an existing model.
```
response_train = restapi_client.submit_job(
workflows_file=WORKFLOWS_TAR_GZ,
experiment_name=EXPERIMENT_NAME,
entry_point="init_model",
entry_point_kwargs=" ".join([
f"-P data_dir={data_path_imagenet}/val-sorted-5000",
]),
queue="tensorflow_gpu",
timeout="1h",
)
pprint.pprint(response_train)
```
Now that we have an ImageNet model, we can run the Pixel Threshold attack on it.
The Pixel Threshold attack attempts to change a limited number of pixels in a test image in an attempt to get it misclassified.
It has two main arguments:
| parameter | data type | description |
| --- | --- | --- |
| th | int | The maximum number of pixels it is allowed to change |
| `es` | int | If 0, then use the CMA-ES strategy, or if 1, use the DE strategy for evolution |
```
def mlflow_run_id_is_not_known(response_pt):
return response_pt["mlflowRunId"] is None and response_pt["status"] not in [
"failed",
"finished",
]
response_pt = restapi_client.submit_job(
workflows_file=WORKFLOWS_TAR_GZ,
experiment_name=EXPERIMENT_NAME,
entry_point="pt",
entry_point_kwargs=" ".join(
[
f"-P model=keras-model-imagenet-resnet50/6",
"-P model_architecture=alex_net",
f"-P data_dir={data_path_imagenet}",
"-P batch_size=32",
"-P th=1",
"-P es=1",
]
),
queue="tensorflow_gpu",
timeout="1h"
)
pprint.pprint(response_pt)
```
| github_jupyter |
# DenseNet Model Building Pipeline for 1D Signals with DEMO
DenseNet121, DenseNet161, DenseNet169, DenseNet201, DenseNet264
# Test GPU (Optional)
Before Starting, kindly check the available GPU from the Google Server, GPU model and other related information. It might help!
```
import torch
print("Is CUDA enabled GPU Available?", torch.cuda.is_available())
print("GPU Number:", torch.cuda.device_count())
print("Current GPU Index:", torch.cuda.current_device())
print("GPU Type:", torch.cuda.get_device_name(device=None))
print("GPU Capability:", torch.cuda.get_device_capability(device=None))
print("Is GPU Initialized yet?", torch.cuda.is_initialized())
```
# Connect to Google Drive (Optional for Google COLAB)
Copy-Paste the Authorization Code and Mount Google Drive to COLAB
```
from google.colab import drive
drive.mount('/content/GDrive')
```
Move to the Target Directory
```
%cd /content
```
List the Files and Folders Located in the Current Directory
```
!ls
```
Upload Files from Local Directory (if required)
```
from google.colab import files
uploaded = files.upload()
```
#Import Necessary Libraries
```
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from scipy import interp
from itertools import cycle
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.metrics import mean_absolute_error, accuracy_score, precision_score, recall_score, f1_score, roc_curve, plot_roc_curve
from sklearn.metrics import confusion_matrix, classification_report, auc, precision_recall_curve, average_precision_score
sns.set_theme(style="whitegrid")
# Import DenseNet Module
from DenseNet_1DCNN import DenseNet
```
# DEMO: Regression and Classification
## Regression
### Import and Prepare Dataset
Import Dataset from a CSV file using Pandas. This DEMO uses the Weather History dataset from Kaggle.
Source: https://www.kaggle.com/budincsevity/szeged-weather
```
dataset = pd.read_csv('WeatherHistory.csv')
print(dataset.shape)
dataset.head(5)
```
Have a look at the dataset! The CSV file includes a hourly/daily summary for Szeged, Hungary area, between 2006 and 2016.
Data available in the hourly response:
1. Time
2. Summary
3. PrecipType
4. Temperature
5. ApparentTemperature
6. Humidity
7. WindSpeed
8. WindBearing
9. Visibility
10. LoudCover
11. Pressure
Check for any existing 'NaN' value in the dataset which might create issues afterwards. We, here, check both overall and column by column.
```
print(f'Total NULL values in the Original DataFrame = {dataset.isnull().sum().sum()}')
print(dataset.isnull().sum())
```
There are total 517 NULL or NaN values and all of them are in the 'Precip_Type' Column. Let's print out the unique values in that column.
```
dataset['Precip Type'].unique()
```
So, it can be seen that there are mainly 2 classes in this column other than the NaN. So, we can replace the NaN cell with 'Unknown' instead of '0' (which is done more commonly).
```
dataset['Precip Type'] = dataset['Precip Type'].fillna('Unknown')
```
One important realization is that we could easily remove those rows in this case instead of filling up the NULLs, but that would remove some datapoints. For very large datasets, it might not matter. On the contrary, sometimes it might remove important observations. If the dataset is too sparse, then it is better to remove the rows instead of filling. But since in this case, value in only 1 column was missing, it was filled up with a suitable replacement.
```
dataset['Precip Type'].unique()
```
Now, at first slice the DataTime Frame into month and hour since they can be strong indicators of weather (fairly long term and short term). Years can also show gradual change in weather pattern (especially change in the climate) but that has been ignored in this study. Modify it according to your wish to improve performance.
```
dataset['Month'] = (dataset['Formatted Date']).str.slice(start=5, stop=7)
dataset['Hour'] = (dataset['Formatted Date']).str.slice(start=11, stop=13)
dataset.head(5)
```
The first DateTime Frame column and the column containing the real temperature for that hour have been dropped. The aim here is to predict "Apparent Temperature" from other weather indicators using Deep Learning based Regression.
```
dataset.drop(['Formatted Date', 'Temperature (C)'], axis=1, inplace=True)
dataset.head(5)
```
Rearrange the Columns, place the target column at the end for next step
```
dataset = dataset[['Month', 'Hour', 'Summary', 'Precip Type', 'Humidity',
'Wind Speed (km/h)', 'Wind Bearing (degrees)',
'Visibility (km)', 'Pressure (millibars)', 'Daily Summary',
'Apparent Temperature (C)']]
dataset.head(5)
```
At this stage, check the datatypes of the columns. You can see that 'Month' and 'Hour' columns are object types, convert them to numeric 'int64' (otherwise while creating dummy variables, each month or hour will get a separate column, which is unncessary).
```
dataset.info()
dataset['Month'] = dataset['Month'].astype('int64')
dataset['Hour'] = dataset['Hour'].astype('int64')
dataset.info()
```
Convert Text Data into Dummy Variables for Machine Learning.
It is important since Machine Learning models will not accept non-numerical data directly during training or testing.
```
dummy_dataset = pd.DataFrame()
for i in range(0,len(dataset.columns)):
X = dataset[dataset.columns[i]]
if type(X[0]) == str:
Y = pd.get_dummies(X)
dummy_dataset = pd.concat([dummy_dataset, Y], axis=1)
else:
dummy_dataset = pd.concat([dummy_dataset, X], axis=1)
#
dummy_dataset.shape
dummy_dataset.head(5)
```
At this stage, you can plot the dataset prior to check any pattern in the dataset or just for illustrative purposes.
Convert Pandas DataFrame into NumPy Arrays
```
X_Data = dummy_dataset.iloc[:,0:-1].values # All columns except the last are the predicting variables
Y_Data = dummy_dataset.iloc[:,-1].values # Last column (Weight Column in this case) is the label
print(X_Data.shape)
print(Y_Data.shape)
```
Train-Test Split
```
X_Train, X_Test, Y_Train, Y_Test = train_test_split(X_Data, Y_Data, test_size=0.20, random_state=42)
```
The third axis is to show the number of channels, which is 1 in this case. If same labels were appropriate for more than 1 dataset, there would be more than 1 channel.
```
X_Train = np.expand_dims(X_Train, axis=2)
X_Test = np.expand_dims(X_Test, axis=2)
print(X_Train.shape, X_Test.shape)
print(Y_Train.shape, Y_Test.shape)
```
### Build and Train Imported Data using the Regression Model
Configurations - Select the Configurations for the Regression Model. Vary them (while following the guidelines) to improve performance.
```
"Configurations for DenseNet in Regression Mode"
length = X_Train.shape[1] # Number of Features (or length of the signal)
model_width = 32 # Number of Filter or Kernel in the Input Layer (Power of 2 to avoid error)
num_channel = 1 # Number of Input Channels
problem_type = 'Regression' # Regression or Classification
output_number = 1 # Number of Outputs in the Regression Mode - 1 input is mapped to a single output
```
Build Model
```
Regression_Model = DenseNet(length, num_channel, model_width, problem_type=problem_type, output_nums=output_number).DenseNet169() # Build Model
Regression_Model.compile(loss='mae', optimizer='adam', metrics= ['mse']) # Compile Model
```
Model_Summary
```
Regression_Model.summary() # Summary of the Model
```
Upload Past Weights if available
```
Regression_Model.load_weights('Saved_Model.h5') # Load Previously Trained Weights for Transfer Learning
```
Train Model for 'n' number of Epochs with Batch size of 'm'
```
# Early Stopping and Model_Checkpoints are optional parameters
# Early Stopping is to stop the training based on certain condition set by the user
# Model Checkpoint is to save a model in a directory based on certain conditions so that it can be used later for Transfer Learning or avoiding retraining
callbacks = [tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=30, mode='min'),
tf.keras.callbacks.ModelCheckpoint('trained_models/DenseNet169_'+str(model_width)+'_'+str(num_channel)+'_'+str(output_number)+'_'+str(problem_type)+'.h5',
verbose=1, monitor='val_loss', save_best_only=True, mode='min')]
history = Regression_Model.fit(X_Train, Y_Train, epochs=500, batch_size=128, verbose=1, validation_split=0.2, shuffle=True, callbacks=callbacks)
# Save 'History' of the model for model performance analysis performed later
```
Test and Predict
```
# Preictions from the Test Set from the Trained Model
Predictions = Regression_Model.predict(X_Test, verbose=1)
print(Predictions.shape)
```
Error Performance (Mean Sqaured Error or MAE)
```
# Error of the prediction, one of many evaluation metrics
# Using Mean Absolute Error (MAE) in this case as a sample
Error = mean_absolute_error(Y_Test, Predictions)
print(f"MAE: {Error}")
```
Plot Train and Validation Error and Loss
```
def history_plot(history):
# list all dictionaries in history
print(history.history.keys())
# summarize history for error
plt.figure(figsize=(12,10))
plt.subplot(2,1,1)
plt.plot(history.history['mse'])
plt.plot(history.history['val_mse'])
plt.title('Model Error Performance')
plt.ylabel('Error')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
# summarize history for loss
plt.figure(figsize=(12,10))
plt.subplot(2,1,2)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
#
history_plot(history)
```
Plot Prediction Performance
```
def plot_prediction(test_labels, test_predictions):
plt.figure(figsize=(15,10))
plt.scatter(test_labels, test_predictions)
'''Add Trendline'''
z = np.polyfit(test_labels.ravel(), test_predictions.ravel(), 1)
p = np.poly1d(z)
plt.plot(test_labels, p(test_labels))
plt.text(np.max(test_labels)/3,np.max(test_predictions),f'y = {p[1]:.2f}x+{p[0]:.2f}', fontsize=15)
plt.title('Ground Truth vs. Prediction Scatter Plot', fontsize=20)
plt.xlabel('Ground Truth', fontsize=15)
plt.ylabel('Predictions', fontsize=15)
#
plot_prediction(Y_Test, Predictions)
```
Kernel Density Plot
```
df = pd.DataFrame(data = {'Ground Truth': Y_Test.ravel(), 'Predictions': Predictions.ravel()})
plt.figure(figsize=(15,10))
sns.set_style('whitegrid')
sns.kdeplot(data=df)
plt.title('Kernel Density Estimation (KDE) Plot for Ground Truth and Predictions', fontsize=20)
plt.xlabel('Magnitude', fontsize=15)
plt.ylabel('Density', fontsize=15)
plt.figure(figsize = (15,10))
df = pd.DataFrame(data = {'Ground Truth': Y_Test.ravel(), 'Predictions': Predictions.ravel()})
ax = sns.violinplot(data=df)
plt.title('Violin Plot for Ground Truth and Predictions', fontsize=20)
plt.ylabel('Magnitude', fontsize=15)
```
Both from the Kernel Density Plot or the Violin Plot, it can be seen that not only the MAE is good, but also the predicted values follow the same pattern as the ground truth, which proves the robustness of the model.
## Classification
### Data Preparation
Import the dataset from a CSV file and save it into a Pandas DataFrame
```
dataset = pd.read_excel('Drug_Persistency.xlsx', sheet_name = 'Dataset')
dataset.drop(columns=['Ptid'], inplace=True) # Drop Patient ID Column
print(dataset.shape)
dataset.head(10)
```
Create a Blank DataFrame
```
dummy_dataset = pd.DataFrame()
for i in range(0,len(dataset.columns)):
X = dataset[dataset.columns[i]]
if type(X[0]) == str:
Y = pd.get_dummies(X)
dummy_dataset = pd.concat([dummy_dataset, Y], axis=1)
else:
dummy_dataset = pd.concat([dummy_dataset, X], axis=1)
#
dummy_dataset.head(10)
```
Convert the DataFrame to a Numpy Array
```
X_Data = dummy_dataset.iloc[:,0:-1].values
Y_Data = dummy_dataset.iloc[:,-1].values
print(X_Data.shape)
print(Y_Data.shape)
```
Get Unique Labels
```
'''Print out the Classes to track the sequence, will be important in the evaluation section (e.g., Labelling Conf Mat)'''
L_E = LabelEncoder()
L_E.fit_transform(Y_Data)
labels = L_E.classes_
print(f'Classes: {labels}')
```
Train_Test Split
```
X_Train, X_Test, Y_Train_, Y_Test_ = train_test_split(X_Data, Y_Data, test_size=0.20, random_state=42)
```
One-Hot-Encoding for the Classification Labels
```
def one_hot_encoding(data):
L_E = LabelEncoder()
integer_encoded = L_E.fit_transform(data)
onehot_encoder = OneHotEncoder(sparse=False)
integer_encoded = integer_encoded.reshape(len(integer_encoded), 1)
one_hot_encoded_data = onehot_encoder.fit_transform(integer_encoded)
return one_hot_encoded_data
Y_Train = one_hot_encoding(Y_Train_.ravel())
Y_Test = one_hot_encoding(Y_Test_.ravel())
```
Train and Test Data Shapes
```
print(X_Train.shape, X_Test.shape)
print(Y_Train.shape, Y_Test.shape)
```
### Build and Train Imported Data using the Classification Model
Configurations
```
"Configurations for the 1D Network in Classification Mode"
length = X_Train.shape[1] # Number of Features (or length of the signal)
model_width = 32 # Number of Filter or Kernels in the Input Layer (Power of 2 to avoid error)
num_channel = 1 # Number of Input Channels
problem_type = 'Classification' # Regression or Classification
class_number = Y_Train.shape[1] # Number of Output Class in Classification Mode (>=2)
```
Build Model
```
Classification_Model = DenseNet(length, num_channel, model_width, problem_type=problem_type, output_nums=class_number).DenseNet169()
if class_number == 2:
Classification_Model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['mse','accuracy'])
elif class_number > 2:
Classification_Model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['mse','accuracy'])
```
Model Summary
```
Classification_Model.summary()
```
Upload Past Weights (Transfer Learning)
```
Classification_Model.load_weights('Saved_Model.h5') # Load Previously Trained Weights for Transfer Learning
```
Train Model
```
# Early Stopping and Model_Checkpoints are optional parameters
# Early Stopping is to stop the training based on certain condition set by the user
# Model Checkpoint is to save a model in a directory based on certain conditions so that it can be used later for Transfer Learning or avoiding retraining
callbacks = [tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=50, mode='min'),
tf.keras.callbacks.ModelCheckpoint('Saved_Model.h5', verbose=1, monitor='val_loss', save_best_only=True, mode='min')]
history = Classification_Model.fit(X_Train, Y_Train, epochs=300, batch_size=128, verbose=1, validation_split=0.2, shuffle=True, callbacks=callbacks)
```
Test
```
# Predictions from the Test Set from the Trained Model
Predictions = Classification_Model.predict(X_Test, verbose=1)
print(Predictions.shape)
```
Error Performance
```
# Error of the prediction, one of many evaluation metrics
# Using Mean Absolute Error (MAE) in this case as a sample
Error = mean_absolute_error(Y_Test, Predictions)
print(f"MAE: {Error}")
```
Plot Training History [Metrics] - Requires to Train the Model
```
def history_plot(history):
# list all dictionaries in history
print(history.history.keys())
# summarize history for error
plt.figure(figsize=(12,10))
plt.subplot(2,1,1)
plt.plot(history.history['mse'])
plt.plot(history.history['val_mse'])
plt.title('Model Error Performance')
plt.ylabel('Error')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
# summarize history for loss
plt.figure(figsize=(12,10))
plt.subplot(2,1,2)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper right')
plt.show()
#
history_plot(history)
```
Convert raw predictions into single-column, integer-based predictions
```
prediction_shape = Predictions.shape
prediction_length = prediction_shape[0]
Y_Preds = np.zeros((prediction_length, 1), dtype=int)
#
for i in range(0, prediction_length):
prediction = Predictions[i]
x = np.where(prediction == np.max(prediction))
x = int(x[0])
Y_Preds[i] = x
Y_Preds.shape
```
Print Confusion Matrix
```
print('-------------------------------------------')
print('Raw Confusion Matrix')
print(confusion_matrix(Y_Test_, Y_Preds, normalize=None))
print('-------------------------------------------')
print('Normalized Confusion Matrix')
print(confusion_matrix(Y_Test_, Y_Preds, normalize='true'))
print('-------------------------------------------')
```
Plot Confusion Matrix
```
# Use the numeric labels or replace with real-class names. But be careful about their respective sequence
labels = ['Class_0', 'Class_1', 'Class_2', 'Class_3', 'Class_4', 'Class_5', 'Class_6', 'Class_7']
def plot_conf_mat(Ground_Truth_Labels, Predictions):
confusion_matrix_raw = confusion_matrix(Ground_Truth_Labels, Predictions, normalize=None)
confusion_matrix_norm = confusion_matrix(Ground_Truth_Labels, Predictions, normalize='true')
shape = confusion_matrix_raw.shape
data = np.asarray(confusion_matrix_raw, dtype=int)
text = np.asarray(confusion_matrix_norm, dtype=float)
annots = (np.asarray(["{0:.2f} ({1:.0f})".format(text, data) for text, data in zip(text.flatten(), data.flatten())])).reshape(shape[0],shape[1])
fig = plt.figure(figsize=(20, 10))
sns.heatmap(confusion_matrix_norm, cmap='YlGnBu', annot=annots, fmt='', xticklabels=labels, yticklabels=labels)
plt.title('Confusion Matrix', fontsize=25)
plt.xlabel("Predicted", fontsize=15)
plt.ylabel("Actual", fontsize=15)
#
plot_conf_mat(Y_Test_, Y_Preds)
```
Here it can be seen that risk level 6 and 7 could not be predicted well due to less number of instances.
```
# Evaluating Overall Metrics - Accuracy, Precision, Recall, f1-Score
Accuracy = accuracy_score(Y_Test_, Y_Preds)
Precision = precision_score(Y_Test_, Y_Preds, average= 'weighted')
Recall = recall_score(Y_Test_, Y_Preds, average= 'weighted')
f1_Score = f1_score(Y_Test_, Y_Preds, average= 'weighted')
print(f'Accuracy = {Accuracy:.3f}')
print(f'Precision = {Precision:.3f}')
print(f'Recall = {Recall:.3f}')
print(f'f1-Score = {f1_Score:.3f}')
print(classification_report(Y_Test_, Y_Preds, target_names=labels, zero_division=0))
```
Multiclass Receiver Operating Characteristic(ROC) Curves Plot
```
def plot_multiclass_roc(Y_Test, Predictions):
# Compute ROC curve and Area Under Curve (AUC) for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(class_number):
fpr[i], tpr[i], _ = roc_curve(Y_Test[:, i], Predictions[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(Y_Test.ravel(), Predictions.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(class_number)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(class_number):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= class_number
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
plt.figure(figsize=(20, 10))
plt.plot(fpr["micro"], tpr["micro"],
label='micro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["micro"]),
color='deeppink', linestyle=':', linewidth=4)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-average ROC curve (area = {0:0.2f})'''.format(roc_auc["macro"]),
color='navy', linestyle=':', linewidth=4)
for i in range(class_number):
plt.plot(fpr[i], tpr[i], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'''.format(i, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate', fontsize=15)
plt.ylabel('True Positive Rate', fontsize=15)
plt.title('MultiClass ROC Plot with Respective AUC', fontsize=25)
plt.legend(loc="lower right")
plt.show()
#
plot_multiclass_roc(Y_Test, Predictions)
def plot_multiclass_precision_recall_curves(Y_Test, Predictions):
# For each class
precision = dict()
recall = dict()
average_precision = dict()
for i in range(class_number):
precision[i], recall[i], _ = precision_recall_curve(Y_Test[:, i], Predictions[:, i])
average_precision[i] = average_precision_score(Y_Test[:, i], Predictions[:, i])
# A "micro-average": quantifying score on all classes jointly
precision["micro"], recall["micro"], _ = precision_recall_curve(Y_Test.ravel(), Predictions.ravel())
average_precision["micro"] = average_precision_score(Y_Test, Predictions, average="micro")
print('Average precision score, micro-averaged over all classes: {0:0.2f}'.format(average_precision["micro"]))
from itertools import cycle
# setup plot details
colors = cycle(['navy', 'turquoise', 'darkorange', 'cornflowerblue', 'teal'])
plt.figure(figsize=(20, 10))
f_scores = np.linspace(0.2, 0.8, num=4)
lines = []
labels = []
for f_score in f_scores:
x = np.linspace(0.01, 1)
y = f_score * x / (2 * x - f_score)
l, = plt.plot(x[y >= 0], y[y >= 0], color='gray', alpha=0.2)
plt.annotate('f1={0:0.1f}'.format(f_score), xy=(0.9, y[45] + 0.02))
lines.append(l)
labels.append('iso-f1 curves')
l, = plt.plot(recall["micro"], precision["micro"], color='gold', lw=2)
lines.append(l)
labels.append('micro-average Precision-recall (area = {0:0.2f})'''.format(average_precision["micro"]))
for i, color in zip(range(class_number), colors):
l, = plt.plot(recall[i], precision[i], color=color, lw=2)
lines.append(l)
labels.append('Precision-recall for class {0} (area = {1:0.2f})'''.format(i, average_precision[i]))
fig = plt.gcf()
fig.subplots_adjust(bottom=0.25)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('Recall', fontsize=15)
plt.ylabel('Precision', fontsize=15)
plt.title('MultiClass Precision-Recall Curves', fontsize=25)
plt.legend(lines, labels, loc=(0, -.3), prop=dict(size=14))
plt.show()
#
plot_multiclass_precision_recall_curves(Y_Test, Predictions)
```
| github_jupyter |
# Loops
Loops are a way to repeatedly execute some code statement.
```
planets = ['Mercury', 'Venus', 'Earth', 'Mars', 'Jupiter', 'Saturn', 'Uranus', 'Neptune']
for planet in planets:
print(planet, end=' ') # print all on same line
```
Notice the simplicity of the ``for`` loop: we specify the variable we want to use, the sequence we want to loop over, and use the "``in``" keyword to link them together in an intuitive and readable way.
The object to the right of the "``in``" can be any object that supports iteration. Basically, if it can be thought of as a sequence or collection of things, you can probably loop over it. In addition to lists, we can iterate over the elements of a tuple:
```
multiplicands = (2, 2, 2, 3, 3, 5)
product = 1
for mult in multiplicands:
product = product * mult
product
```
And even iterate over each character in a string:
```
s = 'steganograpHy is the practicE of conceaLing a file, message, image, or video within another fiLe, message, image, Or video.'
msg = ''
# print all the uppercase letters in s, one at a time
for char in s:
if char.isupper():
print(char, end='')
```
### range()
`range()` is a function that returns a sequence of numbers. It turns out to be very useful for writing loops.
For example, if we want to repeat some action 5 times:
```
for i in range(5):
print("Doing important work. i =", i)
```
You might assume that `range(5)` returns the list `[0, 1, 2, 3, 4]`. The truth is a little bit more complicated:
```
r = range(5)
r
```
`range` returns a "range object". It acts a lot like a list (it's iterable), but doesn't have all the same capabilities. As we saw in the [previous tutorial](#$TUTORIAL_URL(4)$), we can call `help()` on an object like `r` to see Python's documentation on that object, including all of its methods. Click the 'output' button if you're curious about what the help page for a range object looks like.
```
help(range)
```
Just as we can use `int()`, `float()`, and `bool()` to convert objects to another type, we can use `list()` to convert a list-like thing into a list, which shows a more familiar (and useful) representation:
```
list(range(5))
```
Note that the range starts at zero, and that by convention the top of the range is not included in the output. `range(5)` gives the numbers from 0 up to *but not including* 5.
This may seem like a strange way to do things, but the documentation (accessed via `help(range)`) alludes to the reasoning when it says:
> `range(4)` produces 0, 1, 2, 3. These are exactly the valid indices for a list of 4 elements.
So for any list `L`, `for i in range(len(L)):` will iterate over all its valid indices.
```
nums = [1, 2, 4, 8, 16]
for i in range(len(nums)):
nums[i] = nums[i] * 2
nums
```
This is the classic way of iterating over the indices of a list or other sequence.
> **Aside**: `for i in range(len(L)):` is analogous to constructs like `for (int i = 0; i < L.length; i++)` in other languages.
### `enumerate`
`for foo in x` loops over the elements of a list and `for i in range(len(x))` loops over the indices of a list. What if you want to do both?
Enter the `enumerate` function, one of Python's hidden gems:
```
def double_odds(nums):
for i, num in enumerate(nums):
if num % 2 == 1:
nums[i] = num * 2
x = list(range(10))
double_odds(x)
x
```
Given a list, `enumerate` returns an object which iterates over the indices *and* the values of the list.
(Like the `range()` function, it returns an iterable object. To see its contents as a list, we can call `list()` on it.)
```
list(enumerate(['a', 'b']))
```
We can see that that the things we were iterating over are tuples. This helps explain that `for i, num` syntax. We're "unpacking" the tuple, just like in this example from the previous tutorial:
```
x = 0.125
numerator, denominator = x.as_integer_ratio()
```
We can use this unpacking syntax any time we iterate over a collection of tuples.
```
nums = [
('one', 1, 'I'),
('two', 2, 'II'),
('three', 3, 'III'),
('four', 4, 'IV'),
]
for word, integer, roman_numeral in nums:
print(integer, word, roman_numeral, sep=' = ', end='; ')
```
This is equivalent to the following (more tedious) code:
```
for tup in nums:
word = tup[0]
integer = tup[1]
roman_numeral = tup[2]
print(integer, word, roman_numeral, sep=' = ', end='; ')
```
## ``while`` loops
The other type of loop in Python is a ``while`` loop, which iterates until some condition is met:
```
i = 0
while i < 10:
print(i, end=' ')
i += 1
```
The argument of the ``while`` loop is evaluated as a boolean statement, and the loop is executed until the statement evaluates to False.
## List comprehensions
List comprehensions are one of Python's most beloved and unique features. The easiest way to understand them is probably to just look at a few examples:
```
squares = [n**2 for n in range(10)]
squares
```
Here's how we would do the same thing without a list comprehension:
```
squares = []
for n in range(10):
squares.append(n**2)
squares
```
We can also add an `if` condition:
```
short_planets = [planet for planet in planets if len(planet) < 6]
short_planets
```
(If you're familiar with SQL, you might think of this as being like a "WHERE" clause)
Here's an example of filtering with an `if` condition *and* applying some transformation to the loop variable:
```
# str.upper() returns an all-caps version of a string
loud_short_planets = [planet.upper() + '!' for planet in planets if len(planet) < 6]
loud_short_planets
```
People usually write these on a single line, but you might find the structure clearer when it's split up over 3 lines:
```
[
planet.upper() + '!'
for planet in planets
if len(planet) < 6
]
```
(Continuing the SQL analogy, you could think of these three lines as SELECT, FROM, and WHERE)
The expression on the left doesn't technically have to involve the loop variable (though it'd be pretty unusual for it not to). What do you think the expression below will evaluate to? Press the 'output' button to check.
```
[32 for planet in planets]
```
List comprehensions combined with some of the functions we've seen like `min`, `max`, `sum`, `len`, and `sorted`, can lead to some pretty impressive one-line solutions for problems that would otherwise require several lines of code.
For example, [the last exercise](#$EXERCISE_URL(4)$) included a brainteaser asking you to write a function to count the number of negative numbers in a list *without using loops* (or any other syntax we hadn't seen). Here's how we might solve the problem now that we have loops in our arsenal:
```
def count_negatives(nums):
"""Return the number of negative numbers in the given list.
>>> count_negatives([5, -1, -2, 0, 3])
2
"""
n_negative = 0
for num in nums:
if num < 0:
n_negative = n_negative + 1
return n_negative
```
Here's a solution using a list comprehension:
```
def count_negatives(nums):
return len([num for num in nums if num < 0])
```
Much better, right?
Well if all we care about is minimizing the length of our code, this third solution is better still!
```
def count_negatives(nums):
# Reminder: in the "booleans and conditionals" exercises, we learned about a quirk of
# Python where it calculates something like True + True + False + True to be equal to 3.
return sum([num < 0 for num in nums])
```
Which of these solutions is the "best" is entirely subjective. Solving a problem with less code is always nice, but it's worth keeping in mind the following lines from [The Zen of Python](https://en.wikipedia.org/wiki/Zen_of_Python):
> Readability counts.
> Explicit is better than implicit.
The last definition of `count_negatives` might be the shortest, but will other people reading your code understand how it works?
Writing Pythonic code doesn't mean never using for loops!
#$YOURTURN$
| github_jupyter |
<a href="https://colab.research.google.com/github/LeoVogiatzis/GNN_based_NILM/blob/main/notebooks/Embedinggs_AE.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install torch-scatter torch-sparse torch-cluster torch-spline-conv torch-geometric -f https://data.pyg.org/whl/torch-1.10.0+cu113.html
import networkx as nx
import pandas as pd
import torch
import torch_geometric
from torch_geometric.data import Dataset, Data
import numpy as np
import os
from tqdm import tqdm
import sklearn
import torch
from torch.utils.data import TensorDataset, DataLoader
import torch.nn.functional as F
from torch_geometric.utils import negative_sampling, remove_self_loops, add_self_loops
from torch_geometric.nn.inits import reset
from torch_geometric.nn import GCNConv, BatchNorm
from .utils import cluster_kl_loss
EPS = 1e-15
MAX_LOGSTD = 10
class GCNEncoder(torch.nn.Module):
def __init__(self, dims, dropout):
super(GCNEncoder, self).__init__()
self.dropout = dropout
self.layers = torch.nn.ModuleList()
for i in range(len(dims) - 1):
conv = GCNConv(dims[i], dims[i + 1])
self.layers.append(conv)
def forward(self, x, edge_index):
num_layers = len(self.layers)
for idx, layer in enumerate(self.layers):
if idx < num_layers - 1:
x = layer(x, edge_index)
x = F.relu(x)
x = F.dropout(x, p=self.dropout, training=self.training)
else:
x = layer(x, edge_index)
return x
class InnerProductDecoder(torch.nn.Module):
def forward(self, z, edge_index, sigmoid=True):
value = (z[edge_index[0]] * z[edge_index[1]]).sum(dim=1)
return torch.sigmoid(value) if sigmoid else value
class GAE(torch.nn.Module):
def __init__(self, dims, dropout):
super(GAE, self).__init__()
self.encoder = GCNEncoder(dims, dropout)
self.decoder = InnerProductDecoder()
GAE.reset_parameters(self)
def reset_parameters(self):
reset(self.encoder)
reset(self.decoder)
reset(self.cl_module)
def encode(self, *args, **kwargs):
return self.encoder(*args, **kwargs)
def decode(self, *args, **kwargs):
return self.decoder(*args, **kwargs)
def recon_loss(self, z, pos_edge_index, neg_edge_index=None):
"""Given latent variables, computes the binary cross entropy loss for positive edges and negative sampled edges.
Args:
z (Tensor): The latent space representations.
pos_edge_index (LongTensor): The positive edges to train against.
neg_edge_index (LongTensor, optional): The negative edges to train against.
If not given, uses negative sampling to calculate negative edges.
"""
pos_loss = -torch.log(self.decoder(z, pos_edge_index, sigmoid=True) + EPS).mean()
# Do not include self-loops in negative samples
pos_edge_index, _ = remove_self_loops(pos_edge_index)
pos_edge_index, _ = add_self_loops(pos_edge_index)
if neg_edge_index is None:
neg_edge_index = negative_sampling(pos_edge_index, z.size(0))
neg_loss = -torch.log(1 - self.decoder(z, neg_edge_index, sigmoid=True) + EPS).mean()
return pos_loss + neg_loss
model = GAE([2,2], dropout=0.2)
optimizer = optim.Adam(model.parameters(), lr=0.001)
criterion = torch.nn.MSELoss()
for i in range(epochs):
optimizer.zero_grad()
# z = model.encode(data.x, data.train_pos_edge_index)
z = model.encode(data.x, data.edge_index)
out = model.decode(z, data.edge_index)
loss = criterion(data.x, out, sigmoid=True)
# loss = model.recon_loss(z, data.train_pos_edge_index)
loss.backward()
optimizer.step()
model.encode(data.x, data.edge_index)
```
| github_jupyter |
```
%%writefile mul2triplet.c
//Sparse Matrix Multiplication(Triplet Original)
#include<stdio.h>
int c[10][3];
void print_triplet(int compactMatrix[][3], int size){
for (int i=0; i<3; i++)
{
for (int j=0; j<=size; j++)
printf("%2d ", compactMatrix[j][i]);
printf("\n");
}
}
void triplet_multiply(int a[][3], int b[][3]){
int r1,c1,d1,r2,c2,d2,i,j,k,l,flag1=0,flag2=0;
int t1,t2,i1,j1,t,x,y,m;
r1 = a[0][0];
c1 = a[0][1];
d1 = a[0][2];
r2 = b[0][0];
c2 = b[0][1];
d2 = b[0][2];
c[0][0] = r1;
c[0][1] = c2;
l=1;
t = 0;
x = 0;
y = 0;
for(i=0;i<r1;i++){
for(t1=1;t1<=d1;t1++){
if(a[t1][0]==i){
//printf("A:%d\t",a[t1][2]);
for(j=0;j<c2;j++){
for(t2=1;t2<=d2;t2++){
if(b[t2][1]==j){
//printf("B:%d\n",b[t2][2]);
for(k=0;k<c1;k++){
if(a[t1][1]==k && b[t2][0]==k && flag1 == 0) {
x = a[t1][0];
y = b[t2][1];
//printf("\n%d-%d\n",a[t1][0],b[t2][1]);
//printf("\n%d * %d\n",a[t1][2],b[t2][2]);
t = (a[t1][2]*b[t2][2]);
flag1 = 1;
}
else
flag1 = 0;
if (flag1==1){
for(m=1;m<=l;m++){
if(c[m][0] == x && c[m][1]==y){
c[m][2] = c[m][2] + t;
flag2 = 1;
}
}
if(flag2==0){
c[l][0] = x;
c[l][1] = y;
c[l][2] = t;
l = l + 1;
flag1 = 0;
flag2 = 0;
t = 0;
}
else{
if(l==1){
c[l][2] = t;
l = l + 1;
}
flag1 = 0;
flag2 = 0;
t = 0;
}
}
}
}
}
}
}
}
}
c[0][2] = l-1;
}
int main()
{
int r1,c1,d1,r2,c2,d2,i,j,temp;
//Enter the two Sparse Matrices
printf("\nSparse Matrix 1:\n");
int sparse1[5][3]={{2,2,4},
{0,0,1},
{0,1,2},
{1,0,3},
{1,1,4}};
print_triplet(sparse1,4);
printf("\nSparse Matrix 2:\n");
int sparse2[5][3]={{2,2,4},
{0,0,1},
{0,1,2},
{1,0,3},
{1,1,4}};
print_triplet(sparse2,4);
if(2 == 2){
triplet_multiply(sparse1, sparse2);
}
else
printf("\nMultiplication is not possible...!!\n");
printf("\nThe Result is:\n");
for(i=0;i<=c[0][2];i++)
printf("(i):%d--(j):%d--(value)%d\n",c[i][0],c[i][1],c[i][2]);
print_triplet(c,c[0][2]);
return 0;
}
%%script bash
gcc mul2triplet.c -std=c99 -o mul2triplet
!./mul2triplet
%%writefile mat_multiply.c
#include<stdio.h>
int m=1;
void print_mat(int mat[2][2]){
int i,j;
printf("\n Matrix:%d\n",m);
for(i=0;i<2;i++){
for(j=0;j<2;j++){
printf("%2d ",mat[i][j]);
}
printf("\n");
}
}
void sparsemat2tripletf(int sparseMatrix[][2], int row, int col){
int size = 0;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMatrix[i][j] != 0)
size++;
// number of columns in compactMatrix (size) must be
// equal to number of non - zero elements in
// sparseMatrix
int compactMatrix[3][size+1];
compactMatrix[0][0] = row;
compactMatrix[1][0] = col;
compactMatrix[2][0] = size;
// Making of new matrix
int k = 1;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMatrix[i][j] != 0)
{
compactMatrix[0][k] = i;
compactMatrix[1][k] = j;
compactMatrix[2][k] = sparseMatrix[i][j];
k++;
}
printf("\n Triplet Matrix:%d\n",m);
for (int i=0; i<3; i++)
{
for (int j=0; j<=size; j++)
printf("%2d ", compactMatrix[i][j]);
printf("\n");
}
}
int main(){
int a[2][2]={{1,2},
{3,4}};
int b[2][2]={{1,2},
{3,4}};
int c[2][2]={0}, i, j, k;
for(i=0;i<2;i++){
for(j=0;j<2;j++){
c[i][j]=0;
for(k=0;k<2;k++)
c[i][j] = c[i][j] + (a[i][k]*b[k][j]);
}
}
printf("\nThe Result:\n");
print_mat(a);
sparsemat2tripletf(a,2,2);
m = m + 1;
print_mat(b);
sparsemat2tripletf(b,2,2);
m = m + 1;
print_mat(c);
sparsemat2tripletf(c,2,2);
return 0;
}
%%script bash
gcc mat_multiply.c -std=c99 -o mat_multiply
!./mat_multiply
%%writefile mul2triplet33.c
//Sparse Matrix Multiplication(Triplet Original)
#include<stdio.h>
int c[10][3];
void print_triplet(int compactMatrix[][3], int size){
printf("\n");
for (int i=0; i<3; i++)
{
for (int j=0; j<=size; j++)
printf("%4d ", compactMatrix[j][i]);
printf("\n");
}
}
void triplet_multiply(int a[][3], int b[][3]){
int r1,c1,d1,r2,c2,d2,i,j,k,l,flag1=0,flag2=0;
int t1,t2,i1,j1,t,x,y,m;
r1 = a[0][0];
c1 = a[0][1];
d1 = a[0][2];
r2 = b[0][0];
c2 = b[0][1];
d2 = b[0][2];
c[0][0] = r1;
c[0][1] = c2;
l=1;
t = 0;
x = 0;
y = 0;
for(i=0;i<r1;i++){
for(t1=1;t1<=d1;t1++){
if(a[t1][0]==i){
//printf("A:%d\t",a[t1][2]);
for(j=0;j<c2;j++){
for(t2=1;t2<=d2;t2++){
if(b[t2][1]==j){
//printf("B:%d\n",b[t2][2]);
for(k=0;k<c1;k++){
if(a[t1][1]==k && b[t2][0]==k && flag1 == 0) {
x = a[t1][0];
y = b[t2][1];
//printf("\n%d-%d\n",a[t1][0],b[t2][1]);
//printf("\n%d * %d\n",a[t1][2],b[t2][2]);
t = (a[t1][2]*b[t2][2]);
flag1 = 1;
}
else
flag1 = 0;
if (flag1==1){
for(m=1;m<=l;m++){
if(c[m][0] == x && c[m][1]==y){
c[m][2] = c[m][2] + t;
flag2 = 1;
}
}
if(flag2==0){
c[l][0] = x;
c[l][1] = y;
c[l][2] = t;
l = l + 1;
flag1 = 0;
flag2 = 0;
t = 0;
}
else{
if(l==1){
c[l][2] = t;
l = l + 1;
}
flag1 = 0;
flag2 = 0;
t = 0;
}
}
}
}
}
}
}
}
}
c[0][2] = l-1;
}
int main()
{
int r1,c1,d1,r2,c2,d2,i,j,temp;
//Enter the two Sparse Matrices
printf("\nSparse Matrix 1:\n");
int sparse1[10][3]={{3,3,9},
{0,0,1},
{0,1,2},
{0,2,3},
{1,0,4},
{1,1,5},
{1,2,6},
{2,0,7},
{2,1,8},
{2,2,9},};
print_triplet(sparse1,9);
printf("\nSparse Matrix 2:\n");
int sparse2[10][3]={{3,3,9},
{0,0,1},
{0,1,2},
{0,2,3},
{1,0,4},
{1,1,5},
{1,2,6},
{2,0,7},
{2,1,8},
{2,2,9},};
print_triplet(sparse2,9);
if(3 == 3){
triplet_multiply(sparse1, sparse2);
}
else
printf("\nMultiplication is not possible...!!\n");
printf("\nThe Result is:\n");
for(i=0;i<=c[0][2];i++)
printf("(i):%d--(j):%d--(value)%d\n",c[i][0],c[i][1],c[i][2]);
print_triplet(c,c[0][2]);
return 0;
}
%%script bash
gcc mul2triplet33.c -std=c99 -o mul2triplet33
!./mul2triplet33
%%writefile mat_multiply33.c
#include<stdio.h>
int m=1;
void print_mat(int mat[][3]){
int i,j;
printf("\n Matrix:%d\n",m);
for(i=0;i<3;i++){
for(j=0;j<3;j++){
printf("%4d ",mat[i][j]);
}
printf("\n");
}
}
void sparsemat2tripletf(int sparseMatrix[][3], int row, int col){
int size = 0;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMatrix[i][j] != 0)
size++;
// number of columns in compactMatrix (size) must be
// equal to number of non - zero elements in
// sparseMatrix
int compactMatrix[3][size+1];
compactMatrix[0][0] = row;
compactMatrix[1][0] = col;
compactMatrix[2][0] = size;
// Making of new matrix
int k = 1;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMatrix[i][j] != 0)
{
compactMatrix[0][k] = i;
compactMatrix[1][k] = j;
compactMatrix[2][k] = sparseMatrix[i][j];
k++;
}
printf("\n Triplet Matrix:%d\n",m);
for (int i=0; i<3; i++)
{
for (int j=0; j<=size; j++)
printf("%4d", compactMatrix[i][j]);
printf("\n");
}
}
int main(){
int a[3][3]={{1,2,3},
{4,5,6},
{7,8,9}};
int b[3][3]={{1,2,3},
{4,5,6},
{7,8,9}};
int c[3][3]={0}, i, j, k;
for(i=0;i<3;i++){
for(j=0;j<3;j++){
c[i][j]=0;
for(k=0;k<3;k++)
c[i][j] = c[i][j] + (a[i][k]*b[k][j]);
}
}
printf("\nThe Result:\n");
print_mat(a);
sparsemat2tripletf(a,3,3);
m = m + 1;
print_mat(b);
sparsemat2tripletf(b,3,3);
m = m + 1;
print_mat(c);
sparsemat2tripletf(c,3,3);
return 0;
}
%%script bash
gcc mat_multiply33.c -std=c99 -o mat_multiply33
!./mat_multiply33
%%writefile allin1triplet.c
#include<stdio.h>
#define ROW 10
#define COL 10
#define MAX 100
int a[ROW][COL];
int b[ROW][COL];
int c[ROW][COL];
int atrip[3][MAX];
int btrip[3][MAX];
int ctrip[3][MAX];
int dtrip[3][MAX];
void read_sparse(int mat[][COL], int row, int col){
int i,j;
printf("\nEnter %d rows X %d cols elements:\n",row,col);
for(i=0;i<row;i++){
for(j=0;j<col;j++){
scanf("%d",&mat[i][j]);
}
}
}
void print_sparse(int mat[][COL], int row, int col){
int i,j;
printf("\nMatrix:\n");
for(i=0;i<row;i++){
for(j=0;j<col;j++){
printf("%4d ",mat[i][j]);
}
printf("\n");
}
}
void read_triplet(int mat[][3])
{
int i,t,m,n;
printf("\n Enter no. of rows and columns:\n");
scanf("%d%d",&m,&n);
printf("\n No. of non-zero triples:\n");
scanf("%d",&t);
mat[0][0]=m;
mat[0][1]=n;
mat[0][2]=t;
printf("\n Enter the triples(row, column, value):\n");
for(i=1;i<=t;i++){
scanf("%d%d%d",&mat[i][0],&mat[i][1],&mat[i][2]);
}
}
void print_triplet(int mat[3][MAX], int nzero){
int t;
printf("\nOnly Print (Triplet):\n");
for(t=0;t<=nzero;t++){
printf("\n%d--%d: %d",mat[0][t],mat[1][t],mat[2][t]);
}
}
void print_sparse_triplet(int sparseMat[][COL], int trip[3][MAX], int row, int col){
int size = 0;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMat[i][j] != 0)
size++;
// number of columns in compactMatrix (size) must be
// equal to number of non - zero elements in
// sparseMatrix
int tripletMatrix[3][size+1];
tripletMatrix[0][0] = row;
tripletMatrix[1][0] = col;
tripletMatrix[2][0] = size;
// Making of new matrix
int k = 1;
for (int i = 0; i < row; i++)
for (int j = 0; j < col; j++)
if (sparseMat[i][j] != 0)
{
tripletMatrix[0][k] = i;
tripletMatrix[1][k] = j;
tripletMatrix[2][k] = sparseMat[i][j];
k++;
}
printf("\n Triplet Matrix:\n");
for (int i=0; i<3; i++)
{
for (int j=0; j<=size; j++){
printf("%4d", tripletMatrix[i][j]);
trip[i][j] = tripletMatrix[i][j];
}
printf("\n");
}
}
void mul2sparse_mat(int mat1[][COL], int mat2[][COL], int row1, int col1, int row2, int col2){
int i, j, k;
for(i=0;i<row1;i++){
for(j=0;j<col2;j++){
c[i][j]=0;
for(k=0;k<col1;k++)
c[i][j] = c[i][j] + (mat1[i][k]*mat2[k][j]);
}
}
printf("\nThe multiplied matrix in traditional way:\n");
print_sparse(c, row1, col2);
}
void multiply_triplet(int a[3][MAX], int b[3][MAX]){
int r1,c1,d1,r2,c2,d2,i,j,k,l,flag1=0,flag2=0;
int t1,t2,i1,j1,t,x,y,m;
r1 = a[0][0];
c1 = a[1][0];
d1 = a[2][0];
r2 = b[0][0];
c2 = b[1][0];
d2 = b[2][0];
dtrip[0][0] = r1;
dtrip[1][0] = c2;
l=1;
t = 0;
x = 0;
y = 0;
for(i=0;i<r1;i++){
for(t1=1;t1<=d1;t1++){
if(a[0][t1]==i){
//printf("A:%d\t",a[2][t1]);
for(j=0;j<c2;j++){
for(t2=1;t2<=d2;t2++){
if(b[1][t2]==j){
//printf("B:%d\n",b[2][t2]);
for(k=0;k<c1;k++){
if(a[1][t1]==k && b[0][t2]==k && flag1 == 0) {
x = a[0][t1];
y = b[1][t2];
//printf("\n%d-%d\n",x,y);
//printf("\n%d * %d\n",a[2][t1],b[2][t2]);
t = (a[2][t1]*b[2][t2]);
flag1 = 1;
}
else
flag1 = 0;
if (flag1==1){
for(m=1;m<=l;m++){
if(dtrip[0][m] == x && dtrip[1][m]==y){
dtrip[2][m] = dtrip[2][m] + t;
flag2 = 1;
}
}
if(flag2==0){
dtrip[0][l] = x;
dtrip[1][l] = y;
dtrip[2][l] = t;
l = l + 1;
flag1 = 0;
flag2 = 0;
t = 0;
}
else{
if(l==1){
dtrip[2][l] = t;
l = l + 1;
}
flag1 = 0;
flag2 = 0;
t = 0;
}
}
}
}
}
}
}
}
}
dtrip[2][0] = l-1;
}
int main(){
int row1, col1,row2, col2;
printf("\nEnter the Row1 and Col1 values:\n");
scanf("%d%d",&row1,&col1);
printf("\nEnter the Row2 and Col2 values:\n");
scanf("%d%d",&row2,&col2);
printf("\nRead Sparse Matrix1:\n");
read_sparse(a,row1, col1);
printf("\nRead Sparse Matrix2:\n");
read_sparse(b,row2, col2);
printf("\nPrint Sparse Matrix1:\n");
print_sparse(a,row1, col1);
print_sparse_triplet(a, atrip, row1, col1);
printf("\nPrint Sparse Matrix2:\n");
print_sparse(b,row2, col2);
print_sparse_triplet(b, btrip, row2, col2);
//Multiplication in traditional way
mul2sparse_mat(a, b, row1, col1, row2, col2);
print_sparse_triplet(c, ctrip, row1, col2);
//Print the respective triplets only
print_triplet(atrip, row1*col1);
print_triplet(btrip, row1*col1);
print_triplet(ctrip, row1*col1);
printf("\n***Triplet Multiplication***");
multiply_triplet(atrip, btrip);
print_triplet(dtrip, row1*col1);
return 0;
}
%%script bash
gcc allin1triplet.c -std=c99 -o allin1triplet
!./allin1triplet
```
| github_jupyter |
# Using MXNet DALI plugin: using various readers
### Overview
This example shows how different readers could be used to interact with MXNet. It shows how flexible DALI is.
The following readers are used in this example:
- MXNetReader
- CaffeReader
- FileReader
- TFRecordReader
For details on how to use them please see other [examples](../../index.rst).
Let us start from defining some global constants
`DALI_EXTRA_PATH` environment variable should point to the place where data from [DALI extra repository](https://github.com/NVIDIA/DALI_extra) is downloaded. Please make sure that the proper release tag is checked out.
```
import os.path
test_data_root = os.environ['DALI_EXTRA_PATH']
# MXNet RecordIO
db_folder = os.path.join(test_data_root, 'db', 'recordio/')
# Caffe LMDB
lmdb_folder = os.path.join(test_data_root, 'db', 'lmdb')
# image dir with plain jpeg files
image_dir = "../../data/images"
# TFRecord
tfrecord = os.path.join(test_data_root, 'db', 'tfrecord', 'train')
tfrecord_idx = "idx_files/train.idx"
tfrecord2idx_script = "tfrecord2idx"
N = 8 # number of GPUs
BATCH_SIZE = 128 # batch size per GPU
ITERATIONS = 32
IMAGE_SIZE = 3
```
Create idx file by calling `tfrecord2idx` script
```
from subprocess import call
import os.path
if not os.path.exists("idx_files"):
os.mkdir("idx_files")
if not os.path.isfile(tfrecord_idx):
call([tfrecord2idx_script, tfrecord, tfrecord_idx])
```
Let us define:
- common part of pipeline, other pipelines will inherit it
```
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
class CommonPipeline(Pipeline):
def __init__(self, batch_size, num_threads, device_id):
super(CommonPipeline, self).__init__(batch_size, num_threads, device_id)
self.decode = ops.ImageDecoder(device = "mixed", output_type = types.RGB)
self.resize = ops.Resize(device = "gpu",
interp_type = types.INTERP_LINEAR)
self.cmn = ops.CropMirrorNormalize(device = "gpu",
dtype = types.FLOAT,
crop = (227, 227),
mean = [128., 128., 128.],
std = [1., 1., 1.])
self.uniform = ops.Uniform(range = (0.0, 1.0))
self.resize_rng = ops.Uniform(range = (256, 480))
def base_define_graph(self, inputs, labels):
images = self.decode(inputs)
images = self.resize(images, resize_shorter = self.resize_rng())
output = self.cmn(images, crop_pos_x = self.uniform(),
crop_pos_y = self.uniform())
return (output, labels)
```
- MXNetReaderPipeline
```
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
class MXNetReaderPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(MXNetReaderPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.MXNetReader(path = [db_folder+"train.rec"], index_path=[db_folder+"train.idx"],
random_shuffle = True, shard_id = device_id, num_shards = num_gpus)
def define_graph(self):
images, labels = self.input(name="Reader")
return self.base_define_graph(images, labels)
```
- CaffeReadPipeline
```
class CaffeReadPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(CaffeReadPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.CaffeReader(path = lmdb_folder,
random_shuffle = True, shard_id = device_id, num_shards = num_gpus)
def define_graph(self):
images, labels = self.input(name="Reader")
return self.base_define_graph(images, labels)
```
- FileReadPipeline
```
class FileReadPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(FileReadPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.FileReader(file_root = image_dir)
def define_graph(self):
images, labels = self.input(name="Reader")
return self.base_define_graph(images, labels)
```
- TFRecordPipeline
```
import nvidia.dali.tfrecord as tfrec
class TFRecordPipeline(CommonPipeline):
def __init__(self, batch_size, num_threads, device_id, num_gpus):
super(TFRecordPipeline, self).__init__(batch_size, num_threads, device_id)
self.input = ops.TFRecordReader(path = tfrecord,
index_path = tfrecord_idx,
features = {"image/encoded" : tfrec.FixedLenFeature((), tfrec.string, ""),
"image/class/label": tfrec.FixedLenFeature([1], tfrec.int64, -1)
})
def define_graph(self):
inputs = self.input(name="Reader")
images = inputs["image/encoded"]
labels = inputs["image/class/label"]
return self.base_define_graph(images, labels)
```
Let us create pipelines and pass them to MXNet generic iterator
```
import numpy as np
from nvidia.dali.plugin.mxnet import DALIGenericIterator
pipe_types = [[MXNetReaderPipeline, (0, 999)],
[CaffeReadPipeline, (0, 999)],
[FileReadPipeline, (0, 1)],
[TFRecordPipeline, (1, 1000)]]
for pipe_t in pipe_types:
pipe_name, label_range = pipe_t
print ("RUN: " + pipe_name.__name__)
pipes = [pipe_name(batch_size=BATCH_SIZE, num_threads=2, device_id = device_id, num_gpus = N) for device_id in range(N)]
pipes[0].build()
dali_iter = DALIGenericIterator(pipes, [('data', DALIGenericIterator.DATA_TAG),
('label', DALIGenericIterator.LABEL_TAG)],
pipes[0].epoch_size("Reader"))
for i, data in enumerate(dali_iter):
if i >= ITERATIONS:
break
# Testing correctness of labels
for d in data:
label = d.label[0].asnumpy()
image = d.data[0]
## labels need to be integers
assert(np.equal(np.mod(label, 1), 0).all())
## labels need to be in range pipe_name[2]
assert((label >= label_range[0]).all())
assert((label <= label_range[1]).all())
print("OK : " + pipe_name.__name__)
```
| github_jupyter |
# 2A.ML101.1: Introduction to data manipulation with scientific Python
In this section we'll go through the basics of the scientific Python stack for data manipulation: using numpy and matplotlib.
*Source:* [Course on machine learning with scikit-learn](https://github.com/GaelVaroquaux/sklearn_ensae_course) by Gaël Varoquaux
You can skip this section if you already know the scipy stack.
**To learn the scientific Python ecosystem**: http://scipy-lectures.org
```
# Start pylab inline mode, so figures will appear in the notebook
%matplotlib inline
```
## Numpy Arrays
Manipulating `numpy` arrays is an important part of doing machine learning
(or, really, any type of scientific computation) in Python. This will likely
be review for most: we'll quickly go through some of the most important features.
```
import numpy as np
# Generating a random array
X = np.random.random((3, 5)) # a 3 x 5 array
print(X)
# Accessing elements
# get a single element
print(X[0, 0])
# get a row
print(X[1])
# get a column
print(X[:, 1])
# Transposing an array
print(X.T)
# Turning a row vector into a column vector
y = np.linspace(0, 12, 5)
print(y)
# make into a column vector
print(y[:, np.newaxis])
```
There is much, much more to know, but these few operations are fundamental to what we'll
do during this tutorial.
## Scipy Sparse Matrices
We won't make very much use of these in this tutorial, but sparse matrices are very nice
in some situations. For example, in some machine learning tasks, especially those associated
with textual analysis, the data may be mostly zeros. Storing all these zeros is very
inefficient. We can create and manipulate sparse matrices as follows:
```
from scipy import sparse
# Create a random array with a lot of zeros
X = np.random.random((10, 5))
print(X)
# set the majority of elements to zero
X[X < 0.7] = 0
print(X)
# turn X into a csr (Compressed-Sparse-Row) matrix
X_csr = sparse.csr_matrix(X)
print(X_csr)
# convert the sparse matrix to a dense array
print(X_csr.toarray())
```
## Matplotlib
Another important part of machine learning is visualization of data. The most common
tool for this in Python is `matplotlib`. It is an extremely flexible package, but
we will go over some basics here.
First, something special to IPython notebook. We can turn on the "IPython inline" mode,
which will make plots show up inline in the notebook.
```
%matplotlib inline
# Here we import the plotting functions
import matplotlib.pyplot as plt
# plotting a line
x = np.linspace(0, 10, 100)
plt.plot(x, np.sin(x));
# scatter-plot points
x = np.random.normal(size=500)
y = np.random.normal(size=500)
plt.scatter(x, y);
# showing images
x = np.linspace(1, 12, 100)
y = x[:, np.newaxis]
im = y * np.sin(x) * np.cos(y)
print(im.shape)
# imshow - note that origin is at the top-left by default!
plt.imshow(im);
# Contour plot - note that origin here is at the bottom-left by default!
plt.contour(im);
```
There are many, many more plot types available. One useful way to explore these is by
looking at the matplotlib gallery: http://matplotlib.org/gallery.html
You can test these examples out easily in the notebook: simply copy the ``Source Code``
link on each page, and put it in a notebook using the ``%load`` magic.
For example:
```
# %load http://matplotlib.org/mpl_examples/pylab_examples/ellipse_collection.py
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.collections import EllipseCollection
x = np.arange(10)
y = np.arange(15)
X, Y = np.meshgrid(x, y)
XY = np.hstack((X.ravel()[:, np.newaxis], Y.ravel()[:, np.newaxis]))
ww = X/10.0
hh = Y/15.0
aa = X*9
fig, ax = plt.subplots()
ec = EllipseCollection(ww, hh, aa, units='x', offsets=XY,
transOffset=ax.transData)
ec.set_array((X + Y).ravel())
ax.add_collection(ec)
ax.autoscale_view()
ax.set_xlabel('X')
ax.set_ylabel('y')
cbar = plt.colorbar(ec)
cbar.set_label('X+Y');
```
| github_jupyter |
___
<a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
___
# K Nearest Neighbors with Python
You've been given a classified data set from a company! They've hidden the feature column names but have given you the data and the target classes.
We'll try to use KNN to create a model that directly predicts a class for a new data point based off of the features.
Let's grab it and use it!
## Import Libraries
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
```
## Get the Data
Set index_col=0 to use the first column as the index.
```
df = pd.read_csv("Classified Data",index_col=0)
df.head()
```
## Standardize the Variables
Because the KNN classifier predicts the class of a given test observation by identifying the observations that are nearest to it, the scale of the variables matters. Any variables that are on a large scale will have a much larger effect on the distance between the observations, and hence on the KNN classifier, than variables that are on a small scale.
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
dropped = df.drop('TARGET CLASS', axis=1)
scaler.fit(dropped)
scaled_features = scaler.transform(dropped)
scaled_features
df.columns
df_feat = pd.DataFrame(scaled_features, columns=df.columns[:-1])
df_feat.head()
```
## Train Test Split
```
from sklearn.model_selection import train_test_split
X = scaled_features
y = df['TARGET CLASS']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=101)
```
## Using KNN
Remember that we are trying to come up with a model to predict whether someone will TARGET CLASS or not. We'll start with k=1.
```
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train, y_train)
predictions = knn.predict(X_test)
```
## Predictions and Evaluations
Let's evaluate our KNN model!
```
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,predictions))
print(classification_report(y_test,predictions))
```
## Choosing a K Value
Let's go ahead and use the **elbow method** to pick a good K Value:
```
error_rate = []
# Will take some time
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
```
Here we can see that that after arouns K>23 the error rate just tends to hover around 0.06-0.05 Let's retrain the model with that and check the classification report!
```
# FIRST A QUICK COMPARISON TO OUR ORIGINAL K=1
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=1')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
# NOW WITH K=23
knn = KNeighborsClassifier(n_neighbors=23)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=23')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
```
# Great job!
We were able to squeeze some more performance out of our model by tuning to a better K value!
| github_jupyter |
```
""" Basic imports and configurations for efficient numerical calculations and beautiful plots.
"""
# Imports from __future__ in case we're running Python 2
from __future__ import division, print_function
from __future__ import absolute_import, unicode_literals
# Our numerical workhorses
import numpy as np
# import scipy.integrate
# Import pyplot for plotting
import matplotlib.pyplot as plt
from ipywidgets import widgets
# Seaborn, useful for graphics
import seaborn as sns
# Magic function to make matplotlib inline; other style specs must come AFTER
%matplotlib inline
# This enables SVG graphics inline. There is a bug, so uncomment if it works.
%config InlineBackend.figure_formats = {'svg',}
# This enables high resolution PNGs. SVG is preferred, but has problems
# rendering vertical and horizontal lines
%config InlineBackend.figure_formats = {'png', 'retina'}
rc = {'lines.linewidth': 2,
'axes.labelsize': 18,
'axes.titlesize': 18,
'axes.facecolor': 'DFDFE5',
'axes.autolimit_mode': 'round_numbers'}
sns.set_context('notebook', rc=rc)
sns.set_style('darkgrid', rc=rc)
class Speaker(object):
def __init__(self, params):
self.name = params.get('name') # Name for speaker parameter set
self.sensitivity = params.get('sensitivity') # sensitivity of the speaker, SPL at 1 meter for an input of 1 watt
# fundamental parameters
self.sd = params.get('sd') # driver diaphragm area in square meters
self.mms = params.get('mms') # driver mass
self.cms = params.get('cms') # driver compliance
self.rms = params.get('rms') # mechanical resistance
self.re = params.get('re') # driver impedance in ohm
self.le = params.get('le') # voice coil inductance
self.bl = params.get('bl') # driver magnetic field strength
self.vas = params.get('vas') # volume of air necessary for equal compliance as CMS
# small parameters
self.fs = params.get('fs') # resonance frequency
self.qts = params.get('qts') # overall quality factor
self.qms = params.get('qms') # mechanical quality factor
self.qes = params.get('qes') # electrical quality factor
# large parameters
self.vd = params.get('vd') # peak displacement volume
self.xmax = params.get('xmax') # max driver excursion in millimeters
def __repr__(self):
return '\n'.join(item+': '+str(self.__dict__[item]) for item in self.__dict__)
def fs(mms, cms):
""" Resonance frequency is inversely proportional to diaphragm's mass and suspension's compliance.
mms = driver mass in gramm
cms = driver compliance in mm/N
"""
return 1. / (2. * np.pi * np.sqrt(mms * cms))
def qts(qms, qes):
""" Quality factors are calculated from their mechanical and electrical inverse.
"""
return 1. / (1. / qms + 1. / qes)
def qms(qts, qes):
return 1. / (1. / qts - 1. / qes)
def qes(qts, qms):
return 1. / (1. / qts - 1. / qms)
def vas(cms, sd):
""" Calculate VAS from driver compliance and diaphragm diameter.
Return volume in cubic meters, multiply by 1000 to get litres
"""
air_density = 1.184 # k/m^3 at 25°C
speed_sound = 346.1 # m/s at 25°C
return air_density * speed_sound**2 * cms * sd**2
def calc_params(speaker_params):
"""Calculates system parameters from physical parameters.
"""
p = speaker_params
p.qes = qes(p.qts, p.qms)
p.qms = qms(p.qts, p.qes)
p.qts = qts(p.qms, p.qes)
p.rms = p.bl**2 / (p.qms / p.qes * p.re)
p.cms = 1. / 2. / np.pi / p.fs / p.qms / p.rms * 1000 # mm / N
p.mms = 1. / (p.cms * (2. * np.pi * p.fs)**2) * 1000 # grams
p.fs = fs(p.mms/1000, p.cms/1000)
p.vas = vas(p.cms/1000, p.sd/10000)
def calc_sensitivity(speaker_params):
p = speaker_params
p.sensitivity = (p.sd / 10000.)**2 * 1.184 / 346.1 / 2. / np.pi / p.re / (p.mms / 1000 / p.bl ** 2) ** 2 / p.bl ** 2
def calculate_impedance(speaker_params, v_box=np.inf, l_over_a=np.inf):
"""Calculate impedance magnitude, impedance phase, SPL magnitude,
acoustic output phase, and acoustic output group delay.
Code for calculations of transferfunction and pre-calculations from
Scott Howard's https://github.com/maqifrnswa/scimpy
l_over_a = Port/Vent Length to Area Ratio
v_box = "Box Volume"
"""
p = speaker_params
rms = p.bl**2 / (p.qms / p.qes * p.re)
res = p.bl**2 / rms
les = p.bl**2 * p.cms / 1000.
ces = p.mms / 1000. / p.bl**2
leb = p.bl**2 / (p.sd / 10000.)**2 * v_box / (1.184 * 346.1**2)
cev = (p.sd / 10000.)**2 / p.bl**2 * 1.184 * l_over_a
omega = np.logspace(1.3, 4.3, 1000) * 2. * np.pi
zvc = p.le / 1000 * (omega * 1j) # ** n_ = 1 anyway
re_ = p.re + zvc.real # freq. dependent resistance
le_ = zvc.imag / omega # freq. dependent inductance
y_acoustic = -1j / (leb * omega - 1. / (omega * cev))
z_mech = (1. / res + 1 / (omega * les * 1j) + omega * ces * 1j + y_acoustic)**(-1)
z_total = z_mech + re_ + 1j * omega * le_
transferfunc = 1j * (omega * z_mech / z_total) * re_ * ces
if l_over_a != np.inf:
transferfunc = transferfunc * (1j * omega * leb) / (1j * omega * leb + 1. / (1j * omega * cev))
freqs = omega / 2. / np.pi
imp_magnitude = np.abs(z_total)
imp_phase = np.angle(z_total) * 180. / np.pi
efficiency = ((p.sd / 10000.)**2 * 1.184 / 346.1 / 2. / np.pi / p.re / ces**2 / p.bl**2) * np.abs(transferfunc)**2
power_spl = 112.1 + 10. * np.log10(efficiency)
return {'omega': omega, 'power_spl': power_spl}
def spl_response(speaker_params):
""" Return the simulated pressure response of a speaker,
given its parameters, the frequency space and the resolution of simulation steps.
Pressure is proportional to driver velocity and frequency.
"""
calculate_impedance()
def f_c(vas, v_box):
""" Calculate the resonance frequency in a sealed enclosure volume.
"""
return np.sqrt(vas / (v_box-1))
# test speaker setup
fe103en = {
'name': 'Fostex FE103En',
'sensitivity': 89.,
'sd': 50.,
'mms': 25.5,
'cms': 1.65,
'rms': None,
're': 7.5,
'le': 0.0398,
'bl': 4.99,
'vas': 5.95,
'fs': 83.,
'qts': 0.33,
'qes': 0.377,
'qms': 2.747,
'vd': None,
'xmax': 0.6
}
alpha15 = {
'name': 'Eminence Alpha 15',
'sensitivity': 97.,
'sd': 856.3,
'mms': 59.,
'cms': 0.25,
're': 5.88,
'le': 0.84,
'bl': 7.7,
'vas': 260.,
'fs': 41.,
'qts': 1.26,
'qes': 1.53,
'qms': 7.23,
'rms': None
}
# alpha15, fe103en
parameters = alpha15
box_volume = 60. # liters
port_area_by_length = 30. * 3. / 20. # area to length ratio of the port
speaker = Speaker(parameters)
# fill up missing parameters if possible
# calc_params(speaker)
print(speaker)
r_ = calculate_impedance(speaker, v_box=box_volume, l_over_a=port_area_by_length)
fig = plt.figure()
ax_power = fig.add_subplot(111, title="Speaker SPL")
ax_power.plot(r_['omega'] / 2. / np.pi, r_['power_spl'])
ax_power.set_ylabel('SPL (dB 1W1m)')
ax_power.set_xlabel('Frequency (Hz)')
ax_power.set_xscale('log')
ax_power.set_xlim([20, 20000])
```
| github_jupyter |
# Identificando y modelando relaciones entre más de dos variables

> Como vimos anteriormente, el coeficiente de correlación se calcula por pares de variables, y es bastante útil para identificar cuando dos variables están relacionadas.
> Una vez identificamos la relación entre dos variables, digamos $x$ (independiente / explicativa) y $y$ (dependiente / objetivo), vimos un método para modelar esta relación, con el cual podemos predecir el valor de la variable objetivo $y$ ante un nuevo valor de la variable explicativa $x$.
> Pero, ¿Qué pasa si la variable objetivo depende de más de una variable explicativa $x$?
# 1. Visualizando la correlación entre múltiples variables
Incluso con dos variables, vimos que al calcular la correlación obteníamos una matriz:
```
# Importar pandas
# Cargamos datos de las casas
# Corrleación entre precio y tamaño
```
La interpretación de esta matriz es sencilla. La entrada $i$, $j$ corresponde la correlación de la variable $i$ con la variable $j$.
En general, podemos calcular esta matriz para un número arbitrario de variables:
```
# Matriz de correlación entre las variables de número de cuartos, tamaño y precio
```
Para este ejemplo particular, dado que tenemos pocos datos, es fácil identificar las relaciones observando directamente los números de la matriz. Sin embargo, cuando tenemos muchas variables, resulta complejo visualizar todas las correlaciones numéricas entre pares de variables.
Una manera práctica de identificar la relación entre más de dos variables es por medio de una visualización por mapa de calor:
```
# Importamos seaborn
# Mapa de calor de correlaciones
sns.heatmap(data.corr())
```
Veamos un ejemplo con más variables [Boston house prices dataset](https://scikit-learn.org/stable/datasets/toy_dataset.html):
```
# Dataset de precios de casas en Boston
# Encapsulamos en un data frame
# Mapa de calor de correlaciones
```
# 2. Regresión lineal con múltiples variables
El modelo de regresión lineal que vimos con una variable es fácilmente generalizable para considerar múltiples variables.
En el caso de dos variables, el objetivo de la regresión lineal es encontrar un modelo de un plano:
$$
y = \alpha_0 + \alpha_1 x^1 + \alpha_2 x^2
$$
que **"mejor" (en el sentido de mínimos cuadrados) se ajuste a los puntos**.
Matemáticamente, si los puntos son
$$
\{(x^1_1, x^2_1, y_1), (x^1_2, x^2_2, y_2), \dots, (x^1_n, x^2_n, y_n)\},
$$
lo que queremos es estimar los valores de los parámetros $m$ y $b$ que minimizan la siguiente función:
$$
J(m, b) = \sum_{i=1}^{n}(y_i - \alpha_0 - \alpha_1 x^1_i - \alpha_2 x^2_i)^2
$$
La buena noticia es que el código no se modifica en lo absoluto:
```
# Importar sklearn.linear_model.LinearRegression
# Ajustar el mejor plano a los datos de precios vs. tamaños, número de recámaras
# Obtener parámetros ajustados
```
Podemos comparar los scores ($R^2$: % de variación que explica el modelo) de ambos modelos:
```
# Coeficiente de determinación de modelo con una variable
# Coeficiente de determinación de modelo con dos variables
```
### ¡De esta manera, podemos incluir en nuestro sistema de avalúos automáticos, la variable de número de cuartos!
| github_jupyter |
Reference:https://blog.csdn.net/u013733326/article/details/79702148
## 开始之前
* 构建具有单隐藏层的2类分类神经网络。
* 使用具有非线性激活功能激活函数,例如tanh。
* 计算交叉熵损失(损失函数)。
* 实现向前和向后传播。
## 准备软件包
* numpy:是用Python进行科学计算的基本软件包。
* sklearn:为数据挖掘和数据分析提供的简单高效的工具。
* matplotlib :是一个用于在Python中绘制图表的库。
* testCases:提供了一些测试示例来评估函数的正确性,参见下载的资料或者在底部查看它的代码。
* planar_utils :提供了在这个任务中使用的各种有用的功能,参见下载的资料或者在底部查看它的代码。
```
import numpy as np
import matplotlib.pyplot as plt
from testCases import *
import sklearn
import sklearn.datasets
import sklearn.linear_model
from planar_utils import plot_decision_boundary, sigmoid, load_planar_dataset, load_extra_datasets
# %matplotlib inline #如果你使用用的是Jupyter Notebook的话请取消注释。把每一次执行的结果显示出来
np.random.seed(1) #设置一个固定的随机种子,以保证接下来的步骤中我们的结果是一致的。
```
## 加载和查看数据集
* X:一个numpy的矩阵,包含了这些数据点的数值
* Y:一个numpy的向量,对应着的是X的标签【0 | 1】(红色:0 , 蓝色 :1)
```
X, Y = load_planar_dataset()
# X:一个numpy的矩阵,包含了这些数据点的数值
# Y:一个numpy的向量,对应着的是X的标签【0 | 1】(红色:0 , 蓝色 :1)
print("X的维度:" + str(X.shape))
# print("X =:" + str(X))
# print( "Y :" + str(Y))
# plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral) #绘制散点图
# 上一语句如出现问题,请使用下面的语句:
# print("np.sequence : " + str(np.squeeze(Y))) # 一个[[]] or []
plt.scatter(X[0, :], X[1, :], c=np.squeeze(Y), s=40, cmap=plt.cm.Spectral) #绘制散点图
# 数据看起来像一朵红色(y = 0)和一些蓝色(y = 1)的数据点的花朵的图案
shape_X = X.shape
shape_Y = Y.shape
m = Y.shape[1] # 训练集数量
print("X的维度为:" + str(shape_X))
print("Y的维度为:" + str(shape_Y))
print("数据集里面的数据有:" + str(m) + "个")
```
## 查看简单的Logistic回归的分类效果
在构建完整的神经网络之前,先让我们看看逻辑回归在这个问题上的表现如何,我们可以使用sklearn的内置函数来做到这一点, 运行下面的代码来训练数据集上的逻辑回归分类器。
```
clf = sklearn.linear_model.LogisticRegressionCV()
clf.fit(X.T,Y.T)
plot_decision_boundary(lambda x: clf.predict(x), X, Y) #绘制决策边界
plt.title("Logistic Regression") #图标题
LR_predictions = clf.predict(X.T) #预测结果
print ("逻辑回归的准确性: %d " % float((np.dot(Y, LR_predictions) +
np.dot(1 - Y,1 - LR_predictions)) / float(Y.size) * 100) +
"% " + "(正确标记的数据点所占的百分比)")
# 准确性只有47%的原因是数据集不是线性可分的,所以逻辑回归表现不佳,现在我们正式开始构建神经网络。
```
准确性只有47%的原因是数据集不是线性可分的,所以逻辑回归表现不佳,现在我们正式开始构建神经网络。
## 搭建神经网络
## 4 - Neural Network model
*模型如图**:
<img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%203/Planar%20data%20classification%20with%20one%20hidden%20layer/images/classification_kiank.png" style="width:600px;height:300px;">
**Mathematically**:
For one example $x^{(i)}$:
$$z^{[1] (i)} = W^{[1]} x^{(i)} + b^{[1]}\tag{1}$$
$$a^{[1] (i)} = \tanh(z^{[1] (i)})\tag{2}$$
$$z^{[2] (i)} = W^{[2]} a^{[1] (i)} + b^{[2]}\tag{3}$$
$$\hat{y}^{(i)} = a^{[2] (i)} = \sigma(z^{ [2] (i)})\tag{4}$$
$$y^{(i)}_{prediction} = \begin{cases} 1 & \mbox{if } a^{[2](i)} > 0.5 \\ 0 & \mbox{otherwise } \end{cases}\tag{5}$$
给出所有示例的预测结果,可以按如下方式计算成本$J$:
$$J = - \frac{1}{m} \sum\limits_{i = 0}^{m} \large\left(\small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large \right) \small \tag{6}$$
构建神经网络的一般方法是:
1. 定义神经网络结构(输入单元的数量,隐藏单元的数量等)。
2. 初始化模型的参数
3. 循环:
* 实施前向传播
* 计算损失
* 实现向后传播
* 更新参数(梯度下降)
我们要它们合并到一个nn_model() 函数中,当我们构建好了nn_model()并学习了正确的参数,我们就可以预测新的数据。
## 定义神经网络
在构建之前,我们要先把神经网络的结构给定义好:
* n_x: 输入层的数量
* n_h: 隐藏层的数量(这里设置为4)
* n_y: 输出层的数量
```
def layer_sizes(X , Y):
"""
参数:
X - 输入数据集,维度为(输入的数量,训练/测试的数量)
Y - 标签,维度为(输出的数量,训练/测试数量)
返回:
n_x - 输入层的数量
n_h - 隐藏层的数量
n_y - 输出层的数量
"""
n_x = X.shape[0] #输入层
n_h = 4 #,隐藏层,硬编码为4
n_y = Y.shape[0] #输出层
return (n_x, n_h, n_y)
#测试layer_sizes
print("=========================测试layer_sizes=========================")
X_asses , Y_asses = layer_sizes_test_case()
print(X_asses.shape)
(n_x,n_h,n_y) = layer_sizes(X_asses,Y_asses)
print("输入层的节点数量为: n_x = " + str(n_x))
print("隐藏层的节点数量为: n_h = " + str(n_h))
print("输出层的节点数量为: n_y = " + str(n_y))
```
## 初始化模型的参数
在这里,我们要实现函数initialize_parameters()。我们要确保我们的参数大小合适,如果需要的话,请参考上面的神经网络图。
我们将会用随机值初始化权重矩阵。
* `np.random.randn(a,b)* 0.01`来随机初始化一个维度为(a,b)的矩阵。
将偏向量初始化为零。
* `np.zeros((a,b))`用零初始化矩阵`(a,b)`。
```
def initialize_parameters( n_x , n_h ,n_y):
"""
参数:
n_x - 输入层节点的数量
n_h - 隐藏层节点的数量
n_y - 输出层节点的数量
返回:
parameters - 包含参数的字典:
W1 - 权重矩阵,维度为(n_h,n_x)
b1 - 偏向量,维度为(n_h,1)
W2 - 权重矩阵,维度为(n_y,n_h)
b2 - 偏向量,维度为(n_y,1)
"""
np.random.seed(2) #指定一个随机种子,以便你的输出与我们的一样。
W1 = np.random.randn(n_h, n_x)* 0.01
b1 = np.zeros(shape = (n_h, 1))
W2 = np.random.randn(n_y,n_h)*0.01
b2 = np.zeros(shape=(n_y, 1))
#使用断言确保我的数据格式是正确的
assert(W1.shape == ( n_h , n_x ))
assert(b1.shape == ( n_h , 1 ))
assert(W2.shape == ( n_y , n_h ))
assert(b2.shape == ( n_y , 1 ))
parameters = {"W1" : W1,
"b1" : b1,
"W2" : W2,
"b2" : b2 }
return parameters
#测试initialize_parameters
print("=========================测试initialize_parameters=========================")
n_x , n_h , n_y = initialize_parameters_test_case()
parameters = initialize_parameters(n_x , n_h , n_y)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
循环
前向传播
我们现在要实现前向传播函数forward_propagation()。
我们可以使用sigmoid()函数,也可以使用np.tanh()函数。
步骤如下:
* 使用字典类型的parameters(它是initialize_parameters() 的输出)检索每个参数。
* 实现向前传播, 计算$Z^{[1]},A^{[1]},Z^{[2]},Z^{[1]}$和 $[2]A^{[2]}A [2]$
( 训练集里面所有例子的预测向量)。
* 反向传播所需的值存储在“cache”中,cache将作为反向传播函数的输入。
```
def forward_propagation( X , parameters ):
"""
参数:
X - 维度为(n_x,m)的输入数据。
parameters - 初始化函数(initialize_parameters)的输出
返回:
A2 - 使用sigmoid()函数计算的第二次激活后的数值
cache - 包含“Z1”,“A1”,“Z2”和“A2”的字典类型变量
"""
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
#前向传播计算A2
Z1 = np.dot(W1, X) + b1
A1 = np.tanh(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = sigmoid(Z2)
#使用断言确保我的数据格式是正确的
assert(A2.shape == (1,X.shape[1]))
cache = {"Z1": Z1,
"A1": A1,
"Z2": Z2,
"A2": A2}
return (A2, cache)
#测试forward_propagation
print("=========================测试forward_propagation=========================")
X_assess, parameters = forward_propagation_test_case()
A2, cache = forward_propagation(X_assess, parameters)
print(np.mean(cache["Z1"]), np.mean(cache["A1"]), np.mean(cache["Z2"]), np.mean(cache["A2"]))
```
现在我们已经计算了$A^{[2]},a^{[2](i)}$
包含了训练集里每个数值,现在我们就可以构建成本函数了。
## 计算损失
计算成本的公式如下:
$$J = - \frac{1}{m} \sum\limits_{i = 1}^{m} \large{(} \small y^{(i)}\log\left(a^{[2] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[2] (i)}\right) \large{)} \small\tag{13}$$
有很多的方法都可以计算交叉熵损失,比如下面的这个公式,我们在python中可以这么实现:
KaTeX parse error: \tag works only in display equations:
```
logprobs = np.multiply(np.log(A2),Y)
cost = - np.sum(logprobs) # 不需要使用循环就可以直接算出来。
```
当然,你也可以使用np.multiply()然后使用np.sum()或者直接使用np.dot()
```
def compute_cost(A2,Y,parameters):
"""
计算方程(6)中给出的交叉熵成本,
参数:
A2 - 使用sigmoid()函数计算的第二次激活后的数值
Y - "True"标签向量,维度为(1,数量)
parameters - 一个包含W1,B1,W2和B2的字典类型的变量
返回:
成本 - 交叉熵成本给出方程(13)
"""
m = Y.shape[1]
W1 = parameters["W1"]
W2 = parameters["W2"]
#计算成本
logprobs = logprobs = np.multiply(np.log(A2), Y) + np.multiply((1 - Y), np.log(1 - A2))
cost = - np.sum(logprobs) / m
cost = float(np.squeeze(cost))
assert(isinstance(cost,float))
return cost
#测试compute_cost
print("=========================测试compute_cost=========================")
A2 , Y_assess , parameters = compute_cost_test_case()
print("cost = " + str(compute_cost(A2,Y_assess,parameters)))
```
使用正向传播期间计算的cache,现在可以利用它实现反向传播。
现在我们要开始实现函数backward_propagation()。
向后传播
说明:反向传播通常是深度学习中最难(数学意义)部分,为了帮助你,这里有反向传播讲座的幻灯片, 由于我们正在构建向量化实现,因此我们将需要使用这下面的六个方程:
<img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%203/Planar%20data%20classification%20with%20one%20hidden%20layer/images/grad_summary.png" style="width:600px;height:300px;">
<!--
$\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } = \frac{1}{m} (a^{[2](i)} - y^{(i)})$
$\frac{\partial \mathcal{J} }{ \partial W_2 } = \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } a^{[1] (i) T} $
$\frac{\partial \mathcal{J} }{ \partial b_2 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)}}}$
$\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } = W_2^T \frac{\partial \mathcal{J} }{ \partial z_{2}^{(i)} } * ( 1 - a^{[1] (i) 2}) $
$\frac{\partial \mathcal{J} }{ \partial W_1 } = \frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)} } X^T $
$\frac{\partial \mathcal{J} _i }{ \partial b_1 } = \sum_i{\frac{\partial \mathcal{J} }{ \partial z_{1}^{(i)}}}$
- Note that $*$ denotes elementwise multiplication.
- The notation you will use is common in deep learning coding:
- dW1 = $\frac{\partial \mathcal{J} }{ \partial W_1 }$
- db1 = $\frac{\partial \mathcal{J} }{ \partial b_1 }$
- dW2 = $\frac{\partial \mathcal{J} }{ \partial W_2 }$
- db2 = $\frac{\partial \mathcal{J} }{ \partial b_2 }$
!-->
- Tips:
- 为了计算 dZ1需要计算 $g^{[1]'}(Z^{[1]})$. 其中 $g^{[1]}(.)$ 是tanh激活函数, 如果 $a = g^{[1]}(z)$ 则 $g^{[1]'}(z) = 1-a^2$. 所以,我们需要使用
`(1 - np.power(A1, 2))`来计算$g^{[1]'}(Z^{[1]})$。
```
def backward_propagation(parameters,cache,X,Y):
"""
使用上述说明搭建反向传播函数。
参数:
parameters - 包含我们的参数的一个字典类型的变量。
cache - 包含“Z1”,“A1”,“Z2”和“A2”的字典类型的变量。
X - 输入数据,维度为(2,数量)
Y - “True”标签,维度为(1,数量)
返回:
grads - 包含W和b的导数一个字典类型的变量。
"""
m = X.shape[1]
W1 = parameters["W1"]
W2 = parameters["W2"]
A1 = cache["A1"]
A2 = cache["A2"]
dZ2= A2 - Y
dW2 = (1 / m) * np.dot(dZ2, A1.T)
db2 = (1 / m) * np.sum(dZ2, axis=1, keepdims=True)
dZ1 = np.multiply(np.dot(W2.T, dZ2), 1 - np.power(A1, 2))
dW1 = (1 / m) * np.dot(dZ1, X.T)
db1 = (1 / m) * np.sum(dZ1, axis=1, keepdims=True)
grads = {"dW1": dW1,
"db1": db1,
"dW2": dW2,
"db2": db2 }
return grads
#测试backward_propagation
print("=========================测试backward_propagation=========================")
parameters, cache, X_assess, Y_assess = backward_propagation_test_case()
grads = backward_propagation(parameters, cache, X_assess, Y_assess)
print ("dW1 = "+ str(grads["dW1"]))
print ("db1 = "+ str(grads["db1"]))
print ("dW2 = "+ str(grads["dW2"]))
print ("db2 = "+ str(grads["db2"]))
```
## 更新参数
我们需要使用(dW1, db1, dW2, db2)来更新(W1, b1, W2, b2)。
更新算法如下:
$ \theta = \theta - \alpha \frac{\partial J }{ \partial \theta }$
* $\alpha$:学习速率
* $\theta$ :参数
我们需要选择一个良好的学习速率,我们可以看一下下面这两个图(由Adam Harley提供):
<img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%203/Planar%20data%20classification%20with%20one%20hidden%20layer/images/sgd.gif" style="width:400;height:400;">
<img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%203/Planar%20data%20classification%20with%20one%20hidden%20layer/images/sgd_bad.gif" style="width:400;height:400;">
上面两个图分别代表了具有良好学习速率(收敛)和不良学习速率(发散)的梯度下降算法。
```
def update_parameters(parameters,grads,learning_rate=1.2):
"""
使用上面给出的梯度下降更新规则更新参数
参数:
parameters - 包含参数的字典类型的变量。
grads - 包含导数值的字典类型的变量。
learning_rate - 学习速率
返回:
parameters - 包含更新参数的字典类型的变量。
"""
W1,W2 = parameters["W1"],parameters["W2"]
b1,b2 = parameters["b1"],parameters["b2"]
dW1,dW2 = grads["dW1"],grads["dW2"]
db1,db2 = grads["db1"],grads["db2"]
W1 = W1 - learning_rate * dW1
b1 = b1 - learning_rate * db1
W2 = W2 - learning_rate * dW2
b2 = b2 - learning_rate * db2
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
#测试update_parameters
print("=========================测试update_parameters=========================")
parameters, grads = update_parameters_test_case()
parameters = update_parameters(parameters, grads)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
## 整合
我们现在把上面的东西整合到nn_model()中,神经网络模型必须以正确的顺序使用先前的功能
```
def nn_model(X,Y,n_h,num_iterations,print_cost=False):
"""
参数:
X - 数据集,维度为(2,示例数)
Y - 标签,维度为(1,示例数)
n_h - 隐藏层的数量
num_iterations - 梯度下降循环中的迭代次数
print_cost - 如果为True,则每1000次迭代打印一次成本数值
返回:
parameters - 模型学习的参数,它们可以用来进行预测。
"""
np.random.seed(3) #指定随机种子
n_x = layer_sizes(X, Y)[0]
n_y = layer_sizes(X, Y)[2]
parameters = initialize_parameters(n_x,n_h,n_y)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
for i in range(num_iterations):
A2 , cache = forward_propagation(X,parameters)
cost = compute_cost(A2,Y,parameters)
grads = backward_propagation(parameters,cache,X,Y)
parameters = update_parameters(parameters,grads,learning_rate = 0.5)
if print_cost:
if i%1000 == 0:
print("第 ",i," 次循环,成本为:"+str(cost))
return parameters
#测试nn_model
print("=========================测试nn_model=========================")
X_assess, Y_assess = nn_model_test_case()
parameters = nn_model(X_assess, Y_assess, 4, num_iterations=10000, print_cost=False)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
## 预测
构建predict()来使用模型进行预测, 使用向前传播来预测结果。
**Reminder**: predictions = $y_{prediction} = \mathbb 1 \text{{activation > 0.5}} = \begin{cases}
1 & \text{if}\ activation > 0.5 \\
0 & \text{otherwise}
\end{cases}$
As an example, if you would like to set the entries of a matrix X to 0 and 1 based on a threshold you would do: ```X_new = (X > threshold)```
```
def predict(parameters,X):
"""
使用学习的参数,为X中的每个示例预测一个类
参数:
parameters - 包含参数的字典类型的变量。
X - 输入数据(n_x,m)
返回
predictions - 我们模型预测的向量(红色:0 /蓝色:1)
"""
A2 , cache = forward_propagation(X,parameters)
predictions = np.round(A2)
return predictions
#测试predict
print("=========================测试predict=========================")
parameters, X_assess = predict_test_case()
predictions = predict(parameters, X_assess)
print("预测的平均值 = " + str(np.mean(predictions)))
```
现在我们把所有的东西基本都做完了,我们开始正式运行。
## 正式运行
```
parameters = nn_model(X, Y, n_h = 4, num_iterations=10000, print_cost=True)
#绘制边界
# plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y) 修改
plot_decision_boundary(lambda x: predict(parameters, x.T), X, np.squeeze(Y))
plt.title("Decision Boundary for hidden layer size " + str(4))
predictions = predict(parameters, X)
print ('准确率: %d' % float((np.dot(Y, predictions.T) + np.dot(1 - Y, 1 - predictions.T)) / float(Y.size) * 100) + '%')
```
## 更改隐藏层节点数量
我们上面的实验把隐藏层定为4个节点,现在我们更改隐藏层里面的节点数量,看一看节点数量是否会对结果造成影响。
```
plt.figure(figsize=(16, 32))
hidden_layer_sizes = [1, 2, 3, 4, 5, 20, 50] #隐藏层数量
for i, n_h in enumerate(hidden_layer_sizes):
plt.subplot(5, 2, i + 1)
plt.title('Hidden Layer of size %d' % n_h)
parameters = nn_model(X, Y, n_h, num_iterations=5000)
plot_decision_boundary(lambda x: predict(parameters, x.T), X, np.squeeze(Y))
predictions = predict(parameters, X)
accuracy = float((np.dot(Y, predictions.T) + np.dot(1 - Y, 1 - predictions.T)) / float(Y.size) * 100)
print ("隐藏层的节点数量: {} ,准确率: {} %".format(n_h, accuracy))
```
较大的模型(具有更多隐藏单元)能够更好地适应训练集,直到最终的最大模型过度拟合数据。
最好的隐藏层大小似乎在n_h = 5附近。实际上,这里的值似乎很适合数据,而且不会引起过度拟合。
我们还将在后面学习有关正则化的知识,它允许我们使用非常大的模型(如n_h = 50),而不会出现太多过度拟合。
## 【可选】探索
* 当改变sigmoid激活或ReLU激活的tanh激活时会发生什么?
* 改变learning_rate的数值会发生什么
* 如果我们改变数据集呢?
```
# 数据集
noisy_circles, noisy_moons, blobs, gaussian_quantiles, no_structure = load_extra_datasets()
datasets = {"noisy_circles": noisy_circles,
"noisy_moons": noisy_moons,
"blobs": blobs,
"gaussian_quantiles": gaussian_quantiles}
dataset = "noisy_moons"
X, Y = datasets[dataset]
X, Y = X.T, Y.reshape(1, Y.shape[0])
if dataset == "blobs":
Y = Y % 2
# plt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral)
#上一语句如出现问题请使用下面的语句:
plt.scatter(X[0, :], X[1, :], c=np.squeeze(Y), s=40, cmap=plt.cm.Spectral)
```
| github_jupyter |
Cognizant Data Science Summit 2020 : July 1, 2020
Yogesh Deshpande [157456]
# Week 1 challenge - Python
Description
The eight queens puzzle is the problem of placing eight chess queens on an 8×8 chessboard so that no two queens threaten each other; thus, a solution requires that no two queens share the same row, column, or diagonal. The eight queens puzzle is an example of the more general n queens problem of placing n non-attacking queens on an n×n chessboard. (Source : https://en.wikipedia.org/wiki/Eight_queens_puzzle )
Challenge
The challenge is to generate one right sequence through Genetic Programming. The sequence has to be 8 numbers between 0 to 7. Each number represents the positions the Queens can be placed. Each number refers to the row number in the specific column
0 3 4 5 6 1 2 4
• 0 is the row number in the column 0 where the Queen can be placed
• 3 is the row number in the column 1 where the Queen can be placed
# Initiaze variables, functions' definitions
```
import random
# Set the variables as per the problem statement
NumberofQueens = 8
InitialPopulation = 1000000 # Initial population has number of chromozones out of which one or more are possible solutions
NumberofIterations = 1000 # Number of generations to check for possible solution
def create_chromozone(NumberofQueens):
chromozone = []
for gene in range(NumberofQueens):
chromozone.append(random.randint(0, NumberofQueens-1))
return chromozone
#print(chromozone)
# Unit testing
# create_chromozone(NumberofQueens)
def create_population(NumberofQueens, InitialPopulation):
Population = []
for chromozone in range(InitialPopulation):
Population.append(create_chromozone(NumberofQueens))
#print(Population)
return Population
# Unit testing
#create_population(NumberofQueens, InitialPopulation)
def fitness_calculation(chromosome, maxFitness):
horizontal_collisions = sum([chromosome.count(i) - 1 for i in chromosome])/2
diagonal_collisions = 0
for record in range(1,len(chromosome)+1):
column1 = record-1
row1 = chromosome[column1]
for i in range (column1+1, len(chromosome)):
column2 = i
row2 = chromosome[i]
deltaRow = abs(row1 - row2)
deltaCol = abs(column1 - column2)
if (deltaRow == deltaCol):
#print("######## Collision detected ##############")
diagonal_collisions = diagonal_collisions + 1
#print("Horizontal Collisions are {} and Diagonal are {} ".format(horizontal_collisions, diagonal_collisions))
fitness_score = maxFitness - (horizontal_collisions + diagonal_collisions)
#print("The fitness score is {}".format(fitness_score))
return fitness_score
#Unit Test
#itness_calculation([4, 1, 5, 8, 2, 7, 3, 6], 28)
def strength_of_chromosome(chromosome, maxFitness):
return fitness_calculation(chromosome, maxFitness) / maxFitness
#Unit Test
#strength_of_chromosome([1, 1, 1, 1, 1, 1, 1, 1], 28)
#strength_of_chromosome([4, 1, 5, 8, 2, 7, 3, 6], 28)
```
# Main Program for solution to get a 8-Queen sequence
```
# Main Program
if __name__ == "__main__":
# Calulate the target Fitness
TargetFitness = (NumberofQueens * (NumberofQueens - 1)) /2
print("Maximum score to achive is = {}".format(TargetFitness))
# Inital population
Population = create_population(NumberofQueens, InitialPopulation)
generation_counter = 0
for iteration in range(NumberofIterations):
MaxPopulationScore = max([fitness_calculation(chromozone, TargetFitness) for chromozone in Population])
print("generation counter = {}, MaxPopulationScore = {}".format(generation_counter, MaxPopulationScore))
if (MaxPopulationScore != TargetFitness):
# If the current population has no score matching target score, continue with next generation
generation_counter = generation_counter + 1
else:
# Target score is achieved at this stage
break
print("Solved in generation {}".format(generation_counter+1))
for chromosome in Population:
if (fitness_calculation(chromosome, TargetFitness) == TargetFitness):
print("Solution =======> {}".format(chromosome))
```
| github_jupyter |
# Enabling App Insights for Services in Production
With this notebook, you can learn how to enable App Insights for standard service monitoring, plus, we provide examples for doing custom logging within a scoring files in a model.
## What does Application Insights monitor?
It monitors request rates, response times, failure rates, etc. For more information visit [App Insights docs.](https://docs.microsoft.com/en-us/azure/application-insights/app-insights-overview)
## What is different compared to standard production deployment process?
If you want to enable generic App Insights for a service run:
```python
aks_service= Webservice(ws, "aks-w-dc2")
aks_service.update(enable_app_insights=True)```
Where "aks-w-dc2" is your service name. You can also do this from the Azure Portal under your Workspace--> deployments--> Select deployment--> Edit--> Advanced Settings--> Select "Enable AppInsights diagnostics"
If you want to log custom traces, you will follow the standard deplyment process for AKS and you will:
1. Update scoring file.
2. Update aks configuration.
3. Build new image and deploy it.
## 1. Import your dependencies
```
from azureml.core import Workspace, Run
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.webservice import Webservice, AksWebservice
from azureml.core.image import Image
from azureml.core.model import Model
import azureml.core
print(azureml.core.VERSION)
```
## 2. Set up your configuration and create a workspace
```
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
```
## 3. Register Model
Register an existing trained model, add descirption and tags.
```
#Register the model
from azureml.core.model import Model
model = Model.register(model_path = "sklearn_regression_model.pkl", # this points to a local file
model_name = "sklearn_regression_model.pkl", # this is the name the model is registered as
tags = {'area': "diabetes", 'type': "regression"},
description = "Ridge regression model to predict diabetes",
workspace = ws)
print(model.name, model.description, model.version)
```
## 4. *Update your scoring file with custom print statements*
Here is an example:
### a. In your init function add:
```python
print ("model initialized" + time.strftime("%H:%M:%S"))```
### b. In your run function add:
```python
print ("Prediction created" + time.strftime("%H:%M:%S"))```
```
%%writefile score.py
import pickle
import json
import numpy
from sklearn.externals import joblib
from sklearn.linear_model import Ridge
from azureml.core.model import Model
import time
def init():
global model
#Print statement for appinsights custom traces:
print ("model initialized" + time.strftime("%H:%M:%S"))
# note here "sklearn_regression_model.pkl" is the name of the model registered under the workspace
# this call should return the path to the model.pkl file on the local disk.
model_path = Model.get_model_path(model_name = 'sklearn_regression_model.pkl')
# deserialize the model file back into a sklearn model
model = joblib.load(model_path)
# note you can pass in multiple rows for scoring
def run(raw_data):
try:
data = json.loads(raw_data)['data']
data = numpy.array(data)
result = model.predict(data)
print ("Prediction created" + time.strftime("%H:%M:%S"))
# you can return any datatype as long as it is JSON-serializable
return result.tolist()
except Exception as e:
error = str(e)
print (error + time.strftime("%H:%M:%S"))
return error
```
## 5. *Create myenv.yml file*
```
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
```
## 6. Create your new Image
```
from azureml.core.image import ContainerImage
image_config = ContainerImage.image_configuration(execution_script = "score.py",
runtime = "python",
conda_file = "myenv.yml",
description = "Image with ridge regression model",
tags = {'area': "diabetes", 'type': "regression"}
)
image = ContainerImage.create(name = "myimage1",
# this is the model object
models = [model],
image_config = image_config,
workspace = ws)
image.wait_for_creation(show_output = True)
```
## Deploy to ACI (Optional)
```
from azureml.core.webservice import AciWebservice
aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1,
memory_gb = 1,
tags = {'area': "diabetes", 'type': "regression"},
description = 'Predict diabetes using regression model',
enable_app_insights = True)
from azureml.core.webservice import Webservice
aci_service_name = 'my-aci-service-4'
print(aci_service_name)
aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,
image = image,
name = aci_service_name,
workspace = ws)
aci_service.wait_for_deployment(True)
print(aci_service.state)
%%time
import json
test_sample = json.dumps({'data': [
[1,28,13,45,54,6,57,8,8,10],
[101,9,8,37,6,45,4,3,2,41]
]})
test_sample = bytes(test_sample,encoding='utf8')
if aci_service.state == "Healthy":
prediction = aci_service.run(input_data=test_sample)
print(prediction)
else:
raise ValueError("Service deployment isn't healthy, can't call the service")
```
## 7. Deploy to AKS service
### Create AKS compute if you haven't done so.
```
# Use the default configuration (can also provide parameters to customize)
prov_config = AksCompute.provisioning_configuration()
aks_name = 'my-aks-test3'
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
%%time
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
```
If you already have a cluster you can attach the service to it:
```python
%%time
resource_id = '/subscriptions/<subscriptionid>/resourcegroups/<resourcegroupname>/providers/Microsoft.ContainerService/managedClusters/<aksservername>'
create_name= 'myaks4'
attach_config = AksCompute.attach_configuration(resource_id=resource_id)
aks_target = ComputeTarget.attach(workspace = ws,
name = create_name,
attach_configuration=attach_config)
## Wait for the operation to complete
aks_target.wait_for_provisioning(True)```
### a. *Activate App Insights through updating AKS Webservice configuration*
In order to enable App Insights in your service you will need to update your AKS configuration file:
```
#Set the web service configuration
aks_config = AksWebservice.deploy_configuration(enable_app_insights=True)
```
### b. Deploy your service
```
if aks_target.provisioning_state== "Succeeded":
aks_service_name ='aks-w-dc5'
aks_service = Webservice.deploy_from_image(workspace = ws,
name = aks_service_name,
image = image,
deployment_config = aks_config,
deployment_target = aks_target
)
aks_service.wait_for_deployment(show_output = True)
print(aks_service.state)
else:
raise ValueError("AKS provisioning failed.")
```
## 8. Test your service
```
%%time
import json
test_sample = json.dumps({'data': [
[1,28,13,45,54,6,57,8,8,10],
[101,9,8,37,6,45,4,3,2,41]
]})
test_sample = bytes(test_sample,encoding='utf8')
if aks_service.state == "Healthy":
prediction = aks_service.run(input_data=test_sample)
print(prediction)
else:
raise ValueError("Service deployment isn't healthy, can't call the service")
```
## 9. See your service telemetry in App Insights
1. Go to the [Azure Portal](https://portal.azure.com/)
2. All resources--> Select the subscription/resource group where you created your Workspace--> Select the App Insights type
3. Click on the AppInsights resource. You'll see a highlevel dashboard with information on Requests, Server response time and availability.
4. Click on the top banner "Analytics"
5. In the "Schema" section select "traces" and run your query.
6. Voila! All your custom traces should be there.
# Disable App Insights
```
aks_service.update(enable_app_insights=False)
```
## Clean up
```
%%time
aks_service.delete()
aci_service.delete()
image.delete()
model.delete()
```
| github_jupyter |
## 1. Regression discontinuity: banking recovery
<p>After a debt has been legally declared "uncollectable" by a bank, the account is considered "charged-off." But that doesn't mean the bank <strong><em>walks away</em></strong> from the debt. They still want to collect some of the money they are owed. The bank will score the account to assess the expected recovery amount, that is, the expected amount that the bank may be able to receive from the customer in the future. This amount is a function of the probability of the customer paying, the total debt, and other factors that impact the ability and willingness to pay.</p>
<p>The bank has implemented different recovery strategies at different thresholds (\$1000, \$2000, etc.) where the greater the expected recovery amount, the more effort the bank puts into contacting the customer. For low recovery amounts (Level 0), the bank just adds the customer's contact information to their automatic dialer and emailing system. For higher recovery strategies, the bank incurs more costs as they leverage human resources in more efforts to obtain payments. Each additional level of recovery strategy requires an additional \$50 per customer so that customers in the Recovery Strategy Level 1 cost the company \$50 more than those in Level 0. Customers in Level 2 cost \$50 more than those in Level 1, etc. </p>
<p><strong>The big question</strong>: does the extra amount that is recovered at the higher strategy level exceed the extra \$50 in costs? In other words, was there a jump (also called a "discontinuity") of more than \$50 in the amount recovered at the higher strategy level? We'll find out in this notebook.</p>
<p>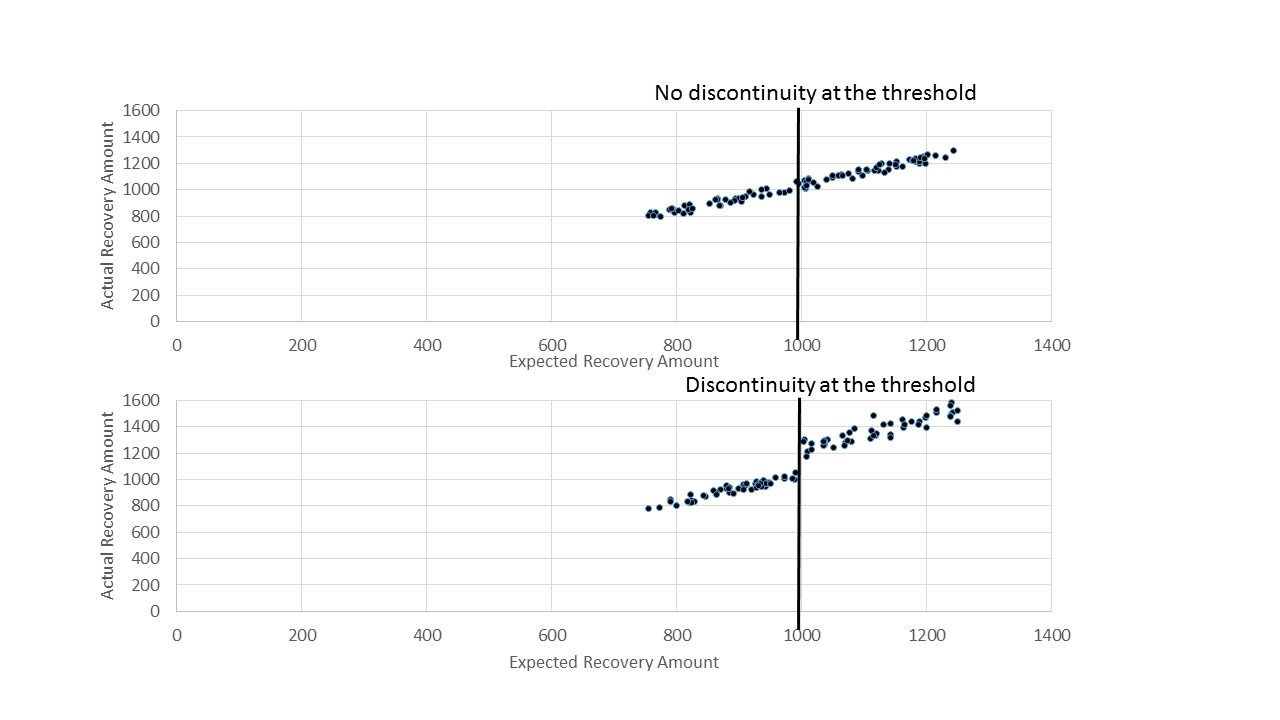</p>
<p>First, we'll load the banking dataset and look at the first few rows of data. This lets us understand the dataset itself and begin thinking about how to analyze the data.</p>
```
# Import modules
# ... YOUR CODE FOR TASK 1 ...
# ... YOUR CODE FOR TASK 1 ...
import numpy as np
import pandas as pd
# Read in dataset
df = pd.read_csv('datasets/bank_data.csv')
# Print the first few rows of the DataFrame
# ... YOUR CODE FOR TASK 1 ...
df.head()
```
## 2. Graphical exploratory data analysis
<p>The bank has implemented different recovery strategies at different thresholds (\$1000, \$2000, \$3000 and \$5000) where the greater the Expected Recovery Amount, the more effort the bank puts into contacting the customer. Zeroing in on the first transition (between Level 0 and Level 1) means we are focused on the population with Expected Recovery Amounts between \$0 and \$2000 where the transition between Levels occurred at \$1000. We know that the customers in Level 1 (expected recovery amounts between \$1001 and \$2000) received more attention from the bank and, by definition, they had higher Expected Recovery Amounts than the customers in Level 0 (between \$1 and \$1000).</p>
<p>Here's a quick summary of the Levels and thresholds again:</p>
<ul>
<li>Level 0: Expected recovery amounts >\$0 and <=\$1000</li>
<li>Level 1: Expected recovery amounts >\$1000 and <=\$2000</li>
<li>The threshold of \$1000 separates Level 0 from Level 1</li>
</ul>
<p>A key question is whether there are other factors besides Expected Recovery Amount that also varied systematically across the \$1000 threshold. For example, does the customer age show a jump (discontinuity) at the \$1000 threshold or does that age vary smoothly? We can examine this by first making a scatter plot of the age as a function of Expected Recovery Amount for a small window of Expected Recovery Amount, \$0 to \$2000. This range covers Levels 0 and 1.</p>
```
# Scatter plot of Age vs. Expected Recovery Amount
from matplotlib import pyplot as plt
%matplotlib inline
plt.scatter(x=df['expected_recovery_amount'], y=df['age'], c="g", s=2)
plt.xlim(0, 2000)
plt.ylim(0, 60)
plt.xlabel('Expected Recovery Amount')
plt.ylabel('Age')
plt.legend(loc=2)
# ... YOUR CODE FOR TASK 2 ...
```
## 3. Statistical test: age vs. expected recovery amount
<p>We want to convince ourselves that variables such as age and sex are similar above and below the \$1000 Expected Recovery Amount threshold. This is important because we want to be able to conclude that differences in the actual recovery amount are due to the higher Recovery Strategy and not due to some other difference like age or sex.</p>
<p>The scatter plot of age versus Expected Recovery Amount did not show an obvious jump around \$1000. We will now do statistical analysis examining the average age of the customers just above and just below the threshold. We can start by exploring the range from \$900 to \$1100.</p>
<p>For determining if there is a difference in the ages just above and just below the threshold, we will use the Kruskal-Wallis test, a statistical test that makes no distributional assumptions.</p>
```
# Import stats module
from scipy import stats
# Compute average age just below and above the threshold
era_900_1100 = df.loc[(df['expected_recovery_amount'] < 1100) &
(df['expected_recovery_amount'] >= 900)]
by_recovery_strategy = era_900_1100.groupby(['recovery_strategy'])
by_recovery_strategy['age'].describe().unstack()
# Perform Kruskal-Wallis test
Level_0_age = era_900_1100.loc[df['recovery_strategy']=="Level 0 Recovery"]['age']
Level_1_age = era_900_1100.loc[df['recovery_strategy']=="Level 1 Recovery"]['age']
stats.kruskal(Level_0_age, Level_1_age)
```
## 4. Statistical test: sex vs. expected recovery amount
<p>We have seen that there is no major jump in the average customer age just above and just
below the \$1000 threshold by doing a statistical test as well as exploring it graphically with a scatter plot. </p>
<p>We want to also test that the percentage of customers that are male does not jump across the \$1000 threshold. We can start by exploring the range of \$900 to \$1100 and later adjust this range.</p>
<p>We can examine this question statistically by developing cross-tabs as well as doing chi-square tests of the percentage of customers that are male vs. female.</p>
```
# Number of customers in each category
crosstab = pd.crosstab(df.loc[(df['expected_recovery_amount'] < 1100) &
(df['expected_recovery_amount'] >= 900)]['recovery_strategy'],
df['sex'])
# ... YOUR CODE FOR TASK 4 ...
print(crosstab)
# Chi-square test
chi2_stat, p_val, dof, ex = stats.chi2_contingency(crosstab)
# ... YOUR CODE FOR TASK 4 ...
p_val
```
## 5. Exploratory graphical analysis: recovery amount
<p>We are now reasonably confident that customers just above and just below the \$1000 threshold are, on average, similar in their average age and the percentage that are male. </p>
<p>It is now time to focus on the key outcome of interest, the actual recovery amount.</p>
<p>A first step in examining the relationship between the actual recovery amount and the expected recovery amount is to develop a scatter plot where we want to focus our attention at the range just below and just above the threshold. Specifically, we will develop a scatter plot of Expected Recovery Amount (Y) vs. Actual Recovery Amount (X) for Expected Recovery Amounts between \$900 to \$1100. This range covers Levels 0 and 1. A key question is whether or not we see a discontinuity (jump) around the \$1000 threshold.</p>
```
# Scatter plot of Actual Recovery Amount vs. Expected Recovery Amount
plt.scatter(x=df['expected_recovery_amount'], y=df['actual_recovery_amount'], c="g", s=2)
plt.xlim(900, 1100)
plt.ylim(0, 2000)
plt.xlabel("Expected Recovery Amount")
plt.ylabel("Actual Recovery Amount")
plt.legend(loc=2)
# ... YOUR CODE FOR TASK 5 ...
```
## 6. Statistical analysis: recovery amount
<p>As we did with age, we can perform statistical tests to see if the actual recovery amount has a discontinuity above the \$1000 threshold. We are going to do this for two different windows of the expected recovery amount \$900 to \$1100 and for a narrow range of \$950 to \$1050 to see if our results are consistent.</p>
<p>Again, we will use the Kruskal-Wallis test.</p>
<p>We will first compute the average actual recovery amount for those customers just below and just above the threshold using a range from \$900 to \$1100. Then we will perform a Kruskal-Wallis test to see if the actual recovery amounts are different just above and just below the threshold. Once we do that, we will repeat these steps for a smaller window of \$950 to \$1050.</p>
```
# Compute average actual recovery amount just below and above the threshold
by_recovery_strategy['actual_recovery_amount'].describe().unstack()
# Perform Kruskal-Wallis test
Level_0_actual = era_900_1100.loc[df['recovery_strategy']=='Level 0 Recovery']['actual_recovery_amount']
Level_1_actual = era_900_1100.loc[df['recovery_strategy']=='Level 1 Recovery']['actual_recovery_amount']
print(stats.kruskal(Level_0_actual, Level_1_actual))
# Repeat for a smaller range of $950 to $1050
era_950_1050 = df.loc[(df['expected_recovery_amount'] < 1050) &
(df['expected_recovery_amount'] >= 950)]
Level_0_actual = era_950_1050[era_950_1050['recovery_strategy'] == 'Level 0 Recovery']['actual_recovery_amount']
Level_1_actual = era_950_1050[era_950_1050['recovery_strategy'] == 'Level 1 Recovery']['actual_recovery_amount']
# ... YOUR CODE FOR TASK 6 ...
print(stats.kruskal(Level_0_actual, Level_1_actual))
```
## 7. Regression modeling: no threshold
<p>We now want to take a regression-based approach to estimate the program impact at the \$1000 threshold using data that is just above and below the threshold. </p>
<p>We will build two models. The first model does not have a threshold while the second will include a threshold.</p>
<p>The first model predicts the actual recovery amount (dependent variable) as a function of the expected recovery amount (independent variable). We expect that there will be a strong positive relationship between these two variables. </p>
<p>We will examine the adjusted R-squared to see the percent of variance explained by the model. In this model, we are not representing the threshold but simply seeing how the variable used for assigning the customers (expected recovery amount) relates to the outcome variable (actual recovery amount).</p>
```
# Import statsmodels
import statsmodels.api as sm
# Define X and y
X = era_900_1100['expected_recovery_amount']
y = era_900_1100['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y, X).fit()
predictions = model.predict(X)
# Print out the model summary statistics
# ... YOUR CODE FOR TASK 7 ...
model.summary()
```
## 8. Regression modeling: adding true threshold
<p>From the first model, we see that the expected recovery amount's regression coefficient is statistically significant. </p>
<p>The second model adds an indicator of the true threshold to the model (in this case at \$1000). </p>
<p>We will create an indicator variable (either a 0 or a 1) that represents whether or not the expected recovery amount was greater than \$1000. When we add the true threshold to the model, the regression coefficient for the true threshold represents the additional amount recovered due to the higher recovery strategy. That is to say, the regression coefficient for the true threshold measures the size of the discontinuity for customers just above and just below the threshold.</p>
<p>If the higher recovery strategy helped recovery more money, then the regression coefficient of the true threshold will be greater than zero. If the higher recovery strategy did not help recovery more money, then the regression coefficient will not be statistically significant.</p>
```
#np.where(df['expected_recovery_amount']<1000)
np.where(df['expected_recovery_amount'] < 1000, 0, 1)
# Create indicator (0 or 1) for expected recovery amount >= $1000
df['indicator_1000'] = np.where(df['expected_recovery_amount']<1000, 0, 1)
era_900_1100 = df.loc[(df['expected_recovery_amount']<1100) &
(df['expected_recovery_amount']>=900)]
# Define X and y
X = era_900_1100[['expected_recovery_amount', 'indicator_1000']]
y = era_900_1100['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y,X).fit()
# Print the model summary
# ... YOUR CODE FOR TASK 8 ...
model.summary()
```
## 9. Regression modeling: adjusting the window
<p>The regression coefficient for the true threshold was statistically significant with an estimated impact of around \$278. This is much larger than the \$50 per customer needed to run this higher recovery strategy. </p>
<p>Before showing this to our manager, we want to convince ourselves that this result wasn't due to choosing an expected recovery amount window of \$900 to \$1100. Let's repeat this analysis for the window from \$950 to \$1050 to see if we get similar results.</p>
<p>The answer? Whether we use a wide (\$900 to \$1100) or narrower window (\$950 to \$1050), the incremental recovery amount at the higher recovery strategy is much greater than the \$50 per customer it costs for the higher recovery strategy. So we conclude that the higher recovery strategy is worth the extra cost of \$50 per customer.</p>
```
# Redefine era_950_1050 so the indicator variable is included
era_950_1050 = df.loc[(df['expected_recovery_amount'] < 1050) &
(df['expected_recovery_amount'] >= 950)]
# Define X and y
X = era_950_1050[['expected_recovery_amount','indicator_1000']]
y = era_950_1050['actual_recovery_amount']
X = sm.add_constant(X)
# Build linear regression model
model = sm.OLS(y,X).fit()
# Print the model summary
model.summary()
```
| github_jupyter |
# 5. Support Vector Machines
Support Vector Machines (SVM) are a powerful and flexible ML model, capable of performing linear or nonlinear classification, regression, and even
outlier detection and particularly suited for classification of complex but small- or medium-sized datasets.
### Linear SVM Classification
We can think of a SVM classifier as fitting for the widest possible distance (_large margin classification_) between classes, relying on specific data points (_support vectors_) who lie at the edge of our boundary.
If we only rely on the most extreme data point, we are looking at **hard** margin classification. In addition to being very sensitive to outlier, sometimes it is not even possible to perform (because the data is not linearly separable).
To avoid this issues, we generally rely on **soft** margin classification, balancing margin wideness and margin violations.
In Scikit-learn, we do this by tuning the `C` hyperparameter. Higher C > Narrower margin.
Here is an implementation example based on the iris flowers dataset:
```
import numpy as np
from sklearn import datasets
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import LinearSVC
iris = datasets.load_iris()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.float64) # Iris-Virginica
svm_clf = Pipeline([
("scaler", StandardScaler()),
("linear_svc", LinearSVC(C=1, loss="hinge")),
])
svm_clf.fit(X, y)
svm_clf.predict([[5.5, 1.7]])
```
### Nonlinear SVM Classification
Sometimes, data is not linearly separable but it can be if we apply a transformation (e.g. polynomial features).
```
from sklearn.datasets import make_moons
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import PolynomialFeatures
from matplotlib import pyplot as plt
polynomial_svm_clf = Pipeline([
("poly_features", PolynomialFeatures(degree=3)),
("scaler", StandardScaler()),
("svm_clf", LinearSVC(C=10, loss="hinge"))
])
polynomial_svm_clf.fit(X, y)
```
In terms of kernels, we do actually have several options to choose from. The linear kernel should be our default choice, but we may need others in specific circumstances (e.g. string kernels for text docs / DNA sequences).
Another very good option is the Gaussian **Radial Basic Function** (RBF) kernel:
```
from sklearn.svm import SVC
rbf_kernel_svm_clf = Pipeline([
("scaler", StandardScaler()),
("svm_clf", SVC(kernel="rbf", gamma=5, C=0.001))
])
rbf_kernel_svm_clf.fit(X, y)
```
To summarize our three approaches for SVM classification:

### SVM Regression
On an intuitive level, regression using SVM works by reversing our logic: instead of fitting the largest possible street _between_ values, we try to fit the as many instances as possible _inside_ the street, with the width controlled by an hyperparameter $\epsilon$.
### Exercises
#### Ex8
Train a `LinearSVC` on a linearly separable dataset. Then train an `SVC` and a `SGDClassifier` on the same dataset.
See if you can get them to produce roughly the same model.
For this exercise, we will use the Iris dataset used in this chapter. First, we want to check which types are linearly separable, so we will plot them in 2D:
```
import matplotlib.pyplot as plt
iris = datasets.load_iris()
X = iris.data
y = iris.target
target_names = iris.target_names
plt.figure()
colors = ['navy', 'darkgreen', 'crimson']
lw = 2
for color, i, target_name in zip(colors, [0, 1, 2], target_names):
plt.scatter(X[y == i, 0], X[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
```
Ok, it is clear that we can build a linearly separable dataset out of setosa + virginica / versicolor. I will go for versicolor.
```
from sklearn.linear_model import SGDClassifier
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 1).astype(np.float64) # Iris-Versicolor
max_iter = 100000
scaler = StandardScaler()
X_scal = scaler.fit_transform(X)
lin = LinearSVC(loss="hinge", max_iter=max_iter)
svc = SVC(max_iter=max_iter)
sgd = SGDClassifier(loss="hinge", learning_rate="constant", eta0 = 0.01, max_iter=max_iter)
for mod in [lin, svc, sgd]:
mod.fit(X_scal, y)
print(mod.intercept_)
```
#### Ex9
Train an SVM classifier on the MNIST dataset. Since SVM classifiers are binary classifiers, you will need to use one-versus-the-rest to classify all 10 digits. You may want to tune the hyperparameters using small validation sets to speed up the process. What accuracy can you reach?
```
from sklearn.datasets import fetch_openml
# load MNIST dataset
X,y = fetch_openml('mnist_784', version=1, return_X_y=True)
from sklearn.model_selection import train_test_split
# split dataset
X_train, X_test, y_train, y_test = train_test_split(
X, y, train_size=1000, test_size=10000)
# scaling inputs
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# trying different SVM models
max_iter = 2000
lin = LinearSVC(loss="hinge", max_iter=max_iter)
svc = SVC(max_iter=max_iter)
sgd = SGDClassifier(loss="hinge", learning_rate="constant", eta0 = 0.05, max_iter=max_iter)
mods = [lin, svc, sgd]
for mod in mods:
mod.fit(X_train, y_train)
from sklearn.metrics import accuracy_score
# continuing with linear model
y_pred = lin.predict(X_train_scaled)
# cross validation score for accuracy
acc_score = accuracy_score(y_train, y_pred)
acc_score
```
Following a very raw approach, we got slightly less than 96.5% accuracy. This is not a bad result at all, especially for a linear model.
#### Ex10
Train an SVM regressor on the California housing dataset.
```
import os
import pandas as pd
def load_housing_data(housing_path):
csv_path = os.path.join(housing_path, datasets, housing, "housing.csv")
return pd.read_csv(csv_path)
cwd = os.getcwd()
load_housing_data(cwd)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Train and explain models locally and deploy model and scoring explainer
_**This notebook illustrates how to use the Azure Machine Learning Interpretability SDK to deploy a locally-trained model and its corresponding scoring explainer to Azure Container Instances (ACI) as a web service.**_
Problem: IBM employee attrition classification with scikit-learn (train and explain a model locally and use Azure Container Instances (ACI) for deploying your model and its corresponding scoring explainer as a web service.)
---
## Table of Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Run model explainer locally at training time](#Explain)
1. Apply feature transformations
1. Train a binary classification model
1. Explain the model on raw features
1. Generate global explanations
1. Generate local explanations
1. [Visualize explanations](#Visualize)
1. [Deploy model and scoring explainer](#Deploy)
1. [Next steps](#Next)
## Introduction
This notebook showcases how to train and explain a classification model locally, and deploy the trained model and its corresponding explainer to Azure Container Instances (ACI).
It demonstrates the API calls that you need to make to submit a run for training and explaining a model to AMLCompute, download the compute explanations remotely, and visualizing the global and local explanations via a visualization dashboard that provides an interactive way of discovering patterns in model predictions and downloaded explanations. It also demonstrates how to use Azure Machine Learning MLOps capabilities to deploy your model and its corresponding explainer.
We will showcase one of the tabular data explainers: TabularExplainer (SHAP) and follow these steps:
1. Develop a machine learning script in Python which involves the training script and the explanation script.
2. Run the script locally.
3. Use the interpretability toolkit’s visualization dashboard to visualize predictions and their explanation. If the metrics and explanations don't indicate a desired outcome, loop back to step 1 and iterate on your scripts.
5. After a satisfactory run is found, create a scoring explainer and register the persisted model and its corresponding explainer in the model registry.
6. Develop a scoring script.
7. Create an image and register it in the image registry.
8. Deploy the image as a web service in Azure.
## Setup
Make sure you go through the [configuration notebook](../../../../configuration.ipynb) first if you haven't.
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Initialize a Workspace
Initialize a workspace object from persisted configuration
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
```
## Explain
Create An Experiment: **Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments.
```
from azureml.core import Experiment
experiment_name = 'explain_model_at_scoring_time'
experiment = Experiment(workspace=ws, name=experiment_name)
run = experiment.start_logging()
# get IBM attrition data
import os
import pandas as pd
outdirname = 'dataset.6.21.19'
try:
from urllib import urlretrieve
except ImportError:
from urllib.request import urlretrieve
import zipfile
zipfilename = outdirname + '.zip'
urlretrieve('https://publictestdatasets.blob.core.windows.net/data/' + zipfilename, zipfilename)
with zipfile.ZipFile(zipfilename, 'r') as unzip:
unzip.extractall('.')
attritionData = pd.read_csv('./WA_Fn-UseC_-HR-Employee-Attrition.csv')
from sklearn.model_selection import train_test_split
from sklearn.externals import joblib
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn_pandas import DataFrameMapper
from interpret.ext.blackbox import TabularExplainer
os.makedirs('./outputs', exist_ok=True)
# Dropping Employee count as all values are 1 and hence attrition is independent of this feature
attritionData = attritionData.drop(['EmployeeCount'], axis=1)
# Dropping Employee Number since it is merely an identifier
attritionData = attritionData.drop(['EmployeeNumber'], axis=1)
attritionData = attritionData.drop(['Over18'], axis=1)
# Since all values are 80
attritionData = attritionData.drop(['StandardHours'], axis=1)
# Converting target variables from string to numerical values
target_map = {'Yes': 1, 'No': 0}
attritionData["Attrition_numerical"] = attritionData["Attrition"].apply(lambda x: target_map[x])
target = attritionData["Attrition_numerical"]
attritionXData = attritionData.drop(['Attrition_numerical', 'Attrition'], axis=1)
# Creating dummy columns for each categorical feature
categorical = []
for col, value in attritionXData.iteritems():
if value.dtype == 'object':
categorical.append(col)
# Store the numerical columns in a list numerical
numerical = attritionXData.columns.difference(categorical)
numeric_transformations = [([f], Pipeline(steps=[
('imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler())])) for f in numerical]
categorical_transformations = [([f], OneHotEncoder(handle_unknown='ignore', sparse=False)) for f in categorical]
transformations = numeric_transformations + categorical_transformations
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
clf = Pipeline(steps=[('preprocessor', DataFrameMapper(transformations)),
('classifier', RandomForestClassifier())])
# Split data into train and test
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(attritionXData,
target,
test_size = 0.2,
random_state=0,
stratify=target)
# preprocess the data and fit the classification model
clf.fit(x_train, y_train)
model = clf.steps[-1][1]
model_file_name = 'log_reg.pkl'
# save model in the outputs folder so it automatically get uploaded
with open(model_file_name, 'wb') as file:
joblib.dump(value=clf, filename=os.path.join('./outputs/',
model_file_name))
# Explain predictions on your local machine
tabular_explainer = TabularExplainer(model,
initialization_examples=x_train,
features=attritionXData.columns,
classes=["Not leaving", "leaving"],
transformations=transformations)
# Explain overall model predictions (global explanation)
# Passing in test dataset for evaluation examples - note it must be a representative sample of the original data
# x_train can be passed as well, but with more examples explanations it will
# take longer although they may be more accurate
global_explanation = tabular_explainer.explain_global(x_test)
from azureml.interpret.scoring.scoring_explainer import TreeScoringExplainer, save
# ScoringExplainer
scoring_explainer = TreeScoringExplainer(tabular_explainer)
# Pickle scoring explainer locally
save(scoring_explainer, exist_ok=True)
# Register original model
run.upload_file('original_model.pkl', os.path.join('./outputs/', model_file_name))
original_model = run.register_model(model_name='local_deploy_model',
model_path='original_model.pkl')
# Register scoring explainer
run.upload_file('IBM_attrition_explainer.pkl', 'scoring_explainer.pkl')
scoring_explainer_model = run.register_model(model_name='IBM_attrition_explainer', model_path='IBM_attrition_explainer.pkl')
```
## Visualize
Visualize the explanations
```
from interpret_community.widget import ExplanationDashboard
ExplanationDashboard(global_explanation, clf, datasetX=x_test)
```
## Deploy
Deploy Model and ScoringExplainer.
Please note that you must indicate azureml-defaults with verion >= 1.0.45 as a pip dependency, because it contains the functionality needed to host the model as a web service.
```
from azureml.core.conda_dependencies import CondaDependencies
# azureml-defaults is required to host the model as a web service.
azureml_pip_packages = [
'azureml-defaults', 'azureml-contrib-interpret', 'azureml-core', 'azureml-telemetry',
'azureml-interpret'
]
# Note: this is to pin the scikit-learn and pandas versions to be same as notebook.
# In production scenario user would choose their dependencies
import pkg_resources
available_packages = pkg_resources.working_set
sklearn_ver = None
pandas_ver = None
for dist in available_packages:
if dist.key == 'scikit-learn':
sklearn_ver = dist.version
elif dist.key == 'pandas':
pandas_ver = dist.version
sklearn_dep = 'scikit-learn'
pandas_dep = 'pandas'
if sklearn_ver:
sklearn_dep = 'scikit-learn=={}'.format(sklearn_ver)
if pandas_ver:
pandas_dep = 'pandas=={}'.format(pandas_ver)
# specify CondaDependencies obj
myenv = CondaDependencies.create(conda_packages=[sklearn_dep, pandas_dep],
pip_packages=['sklearn-pandas', 'pyyaml'] + azureml_pip_packages,
pin_sdk_version=False)
with open("myenv.yml","w") as f:
f.write(myenv.serialize_to_string())
with open("myenv.yml","r") as f:
print(f.read())
from azureml.core.model import Model
# retrieve scoring explainer for deployment
scoring_explainer_model = Model(ws, 'IBM_attrition_explainer')
from azureml.core.webservice import Webservice
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice
from azureml.core.model import Model
from azureml.core.environment import Environment
aciconfig = AciWebservice.deploy_configuration(cpu_cores=1,
memory_gb=1,
tags={"data": "IBM_Attrition",
"method" : "local_explanation"},
description='Get local explanations for IBM Employee Attrition data')
myenv = Environment.from_conda_specification(name="myenv", file_path="myenv.yml")
inference_config = InferenceConfig(entry_script="score_local_explain.py", environment=myenv)
# Use configs and models generated above
service = Model.deploy(ws, 'model-scoring-deploy-local', [scoring_explainer_model, original_model], inference_config, aciconfig)
service.wait_for_deployment(show_output=True)
import requests
import json
# Create data to test service with
sample_data = '{"Age":{"899":49},"BusinessTravel":{"899":"Travel_Rarely"},"DailyRate":{"899":1098},"Department":{"899":"Research & Development"},"DistanceFromHome":{"899":4},"Education":{"899":2},"EducationField":{"899":"Medical"},"EnvironmentSatisfaction":{"899":1},"Gender":{"899":"Male"},"HourlyRate":{"899":85},"JobInvolvement":{"899":2},"JobLevel":{"899":5},"JobRole":{"899":"Manager"},"JobSatisfaction":{"899":3},"MaritalStatus":{"899":"Married"},"MonthlyIncome":{"899":18711},"MonthlyRate":{"899":12124},"NumCompaniesWorked":{"899":2},"OverTime":{"899":"No"},"PercentSalaryHike":{"899":13},"PerformanceRating":{"899":3},"RelationshipSatisfaction":{"899":3},"StockOptionLevel":{"899":1},"TotalWorkingYears":{"899":23},"TrainingTimesLastYear":{"899":2},"WorkLifeBalance":{"899":4},"YearsAtCompany":{"899":1},"YearsInCurrentRole":{"899":0},"YearsSinceLastPromotion":{"899":0},"YearsWithCurrManager":{"899":0}}'
headers = {'Content-Type':'application/json'}
# send request to service
resp = requests.post(service.scoring_uri, sample_data, headers=headers)
print("POST to url", service.scoring_uri)
# can covert back to Python objects from json string if desired
print("prediction:", resp.text)
result = json.loads(resp.text)
#plot the feature importance for the prediction
import numpy as np
import matplotlib.pyplot as plt; plt.rcdefaults()
labels = json.loads(sample_data)
labels = labels.keys()
objects = labels
y_pos = np.arange(len(objects))
performance = result["local_importance_values"][0][0]
plt.bar(y_pos, performance, align='center', alpha=0.5)
plt.xticks(y_pos, objects)
locs, labels = plt.xticks()
plt.setp(labels, rotation=90)
plt.ylabel('Feature impact - leaving vs not leaving')
plt.title('Local feature importance for prediction')
plt.show()
service.delete()
```
## Next
Learn about other use cases of the explain package on a:
1. [Training time: regression problem](https://github.com/interpretml/interpret-community/blob/master/notebooks/explain-regression-local.ipynb)
1. [Training time: binary classification problem](https://github.com/interpretml/interpret-community/blob/master/notebooks/explain-binary-classification-local.ipynb)
1. [Training time: multiclass classification problem](https://github.com/interpretml/interpret-community/blob/master/notebooks/explain-multiclass-classification-local.ipynb)
1. Explain models with engineered features:
1. [Simple feature transformations](https://github.com/interpretml/interpret-community/blob/master/notebooks/simple-feature-transformations-explain-local.ipynb)
1. [Advanced feature transformations](https://github.com/interpretml/interpret-community/blob/master/notebooks/advanced-feature-transformations-explain-local.ipynb)
1. [Save model explanations via Azure Machine Learning Run History](../run-history/save-retrieve-explanations-run-history.ipynb)
1. [Run explainers remotely on Azure Machine Learning Compute (AMLCompute)](../remote-explanation/explain-model-on-amlcompute.ipynb)
1. [Inferencing time: deploy a remotely-trained model and explainer](./train-explain-model-on-amlcompute-and-deploy.ipynb)
1. [Inferencing time: deploy a locally-trained keras model and explainer](./train-explain-model-keras-locally-and-deploy.ipynb)
| github_jupyter |
### This notebook is optionally accelerated with a GPU runtime.
### If you would like to use this acceleration, please select the menu option "Runtime" -> "Change runtime type", select "Hardware Accelerator" -> "GPU" and click "SAVE"
----------------------------------------------------------------------
# Transformer (NMT)
*Author: Facebook AI (fairseq Team)*
**Transformer models for English-French and English-German translation.**
### Model Description
The Transformer, introduced in the paper [Attention Is All You Need][1], is a
powerful sequence-to-sequence modeling architecture capable of producing
state-of-the-art neural machine translation (NMT) systems.
Recently, the fairseq team has explored large-scale semi-supervised training of
Transformers using back-translated data, further improving translation quality
over the original model. More details can be found in [this blog post][2].
### Requirements
We require a few additional Python dependencies for preprocessing:
```
%%bash
pip install fastBPE regex requests sacremoses subword_nmt
```
### English-to-French Translation
To translate from English to French using the model from the paper [Scaling
Neural Machine Translation][3]:
```
import torch
# Load an En-Fr Transformer model trained on WMT'14 data :
en2fr = torch.hub.load('pytorch/fairseq', 'transformer.wmt14.en-fr', tokenizer='moses', bpe='subword_nmt')
# Use the GPU (optional):
en2fr.cuda()
# Translate with beam search:
fr = en2fr.translate('Hello world!', beam=5)
assert fr == 'Bonjour à tous !'
# Manually tokenize:
en_toks = en2fr.tokenize('Hello world!')
assert en_toks == 'Hello world !'
# Manually apply BPE:
en_bpe = en2fr.apply_bpe(en_toks)
assert en_bpe == 'H@@ ello world !'
# Manually binarize:
en_bin = en2fr.binarize(en_bpe)
assert en_bin.tolist() == [329, 14044, 682, 812, 2]
# Generate five translations with top-k sampling:
fr_bin = en2fr.generate(en_bin, beam=5, sampling=True, sampling_topk=20)
assert len(fr_bin) == 5
# Convert one of the samples to a string and detokenize
fr_sample = fr_bin[0]['tokens']
fr_bpe = en2fr.string(fr_sample)
fr_toks = en2fr.remove_bpe(fr_bpe)
fr = en2fr.detokenize(fr_toks)
assert fr == en2fr.decode(fr_sample)
```
### English-to-German Translation
Semi-supervised training with back-translation is an effective way of improving
translation systems. In the paper [Understanding Back-Translation at Scale][4],
we back-translate over 200 million German sentences to use as additional
training data. An ensemble of five of these models was the winning submission to
the [WMT'18 English-German news translation competition][5].
We can further improved this approach through [noisy-channel reranking][6]. More
details can be found in [this blog post][7]. An ensemble of models trained with
this technique was the winning submission to the [WMT'19 English-German news
translation competition][8].
To translate from English to German using one of the models from the winning submission:
```
import torch
# Load an En-De Transformer model trained on WMT'19 data:
en2de = torch.hub.load('pytorch/fairseq', 'transformer.wmt19.en-de.single_model', tokenizer='moses', bpe='fastbpe')
# Access the underlying TransformerModel
assert isinstance(en2de.models[0], torch.nn.Module)
# Translate from En-De
de = en2de.translate('PyTorch Hub is a pre-trained model repository designed to facilitate research reproducibility.')
assert de == 'PyTorch Hub ist ein vorgefertigtes Modell-Repository, das die Reproduzierbarkeit der Forschung erleichtern soll.'
```
We can also do a round-trip translation to create a paraphrase:
```
# Round-trip translations between English and German:
en2de = torch.hub.load('pytorch/fairseq', 'transformer.wmt19.en-de.single_model', tokenizer='moses', bpe='fastbpe')
de2en = torch.hub.load('pytorch/fairseq', 'transformer.wmt19.de-en.single_model', tokenizer='moses', bpe='fastbpe')
paraphrase = de2en.translate(en2de.translate('PyTorch Hub is an awesome interface!'))
assert paraphrase == 'PyTorch Hub is a fantastic interface!'
# Compare the results with English-Russian round-trip translation:
en2ru = torch.hub.load('pytorch/fairseq', 'transformer.wmt19.en-ru.single_model', tokenizer='moses', bpe='fastbpe')
ru2en = torch.hub.load('pytorch/fairseq', 'transformer.wmt19.ru-en.single_model', tokenizer='moses', bpe='fastbpe')
paraphrase = ru2en.translate(en2ru.translate('PyTorch Hub is an awesome interface!'))
assert paraphrase == 'PyTorch is a great interface!'
```
### References
- [Attention Is All You Need][1]
- [Scaling Neural Machine Translation][3]
- [Understanding Back-Translation at Scale][4]
- [Facebook FAIR's WMT19 News Translation Task Submission][6]
[1]: https://arxiv.org/abs/1706.03762
[2]: https://code.fb.com/ai-research/scaling-neural-machine-translation-to-bigger-data-sets-with-faster-training-and-inference/
[3]: https://arxiv.org/abs/1806.00187
[4]: https://arxiv.org/abs/1808.09381
[5]: http://www.statmt.org/wmt18/translation-task.html
[6]: https://arxiv.org/abs/1907.06616
[7]: https://ai.facebook.com/blog/facebook-leads-wmt-translation-competition/
[8]: http://www.statmt.org/wmt19/translation-task.html
| github_jupyter |
# Práctica 4: Entrenamiento de redes neuronales:
Mario Quiñones Pérez y Guillermo García Patiño Lenza
```
from scipy.io import loadmat
from displayData import displayData
import matplotlib.pyplot as plt
import numpy as np
import scipy.optimize as opt
```
## Función de coste:
```
def y_to_one_hot(num_etiquetas):
l = []
for i in range(num_etiquetas):
p = []
for j in range(num_etiquetas):
if( i == j):
p.append(1)
else:
p.append(0)
l.append(p)
return l
def cargaDatos(file = 'ex4data1.mat'):
data = loadmat(file)
X = data['X']
y = data['y']
return (X,y)
(X,y) = cargaDatos('ex4data1.mat')
l = np.array([X[i] for i in np.random.randint(5000, size = 20)])
displayData(l)
def cargaPesos(file = 'ex4weights.mat'):
weights = loadmat(file)
Theta1,Theta2 = weights['Theta1'] , weights['Theta2']
print(Theta1.shape)
print(Theta2.shape)
return Theta1,Theta2
T1,T2 = cargaPesos()
def pasa_a_onehot(y, y_onehot):
res = [y_onehot[i[0]-1] for i in y]
return res
(X,y) = cargaDatos('ex4data1.mat')
print(y)
r = y_to_one_hot(10)
s = np.array(pasa_a_onehot(y, r))
print(s.shape)
for i in np.random.randint(5000,size = 10):
print("{} = {}".format(s[i],y[i]))
def sigmoide(Z):
sigmoide = 1 / (1 + np.exp(-Z))
return sigmoide
def prop_delante(X,Theta1,Theta2):
m = X.shape[0]
A1 = np.hstack([np.ones([m,1]),X])
Z2 = np.dot(A1,Theta1.T)
A2 = np.hstack([np.ones([m,1]), sigmoide(Z2)])
Z3 = np.dot(A2, Theta2.T)
H = sigmoide(Z3)
return A1,A2,H
def prop_delante2(X,Theta):
m = X.shape[0]
A = []
Z = []
a = np.hstack([np.ones([m,1]), X])
z = np.dot(a,Theta[0].T)
A.append(a)
Z.append(z)
for i in range(1,Theta.shape[0],1):
a = np.hstack([np.ones([m,1]), sigmoide(Z[-1])])
A.append(a)
z = np.dot(A[-1],Theta[i].T)
Z.append(z)
H = sigmoide(Z[-1])
return A,Z,H
def prueba_prop_delante():
T1, T2 = cargaPesos()
T = np.array([T1,T2])
X, y = cargaDatos()
A,Z,H = prop_delante2(X,T)
print("Dimensiones de H = {}".format(H.shape))
print(H)
prueba_prop_delante()
def coste_redNeuronal(X,y,Theta1,Theta2):
m_inv = (1/X.shape[0])
m = X.shape[0]
c = 0
Theta = np.array([Theta1,Theta2])
A, Z, H = prop_delante2(X,Theta)
maux = -1 * y
m1 = maux * np.log(H)
m2 = ((1 - y) * np.log(1 - H))
m3 = m1 - m2
c = m_inv * sum(sum(m3))
return c
def prueba_coste():
(X,y) = cargaDatos()
T1, T2 = cargaPesos()
y = np.array(pasa_a_onehot(y, y_to_one_hot(10)))
c = coste_redNeuronal(X,y,T1,T2)
print(c)
prueba_coste()
def coste_redNeuronalReg(X,y,Theta1,Theta2,reg):
c = coste_redNeuronal(X,y,Theta1,Theta2)
r = reg / (2 * X.shape[0])
m1 = Theta1[1:][1:]
m2 = Theta2[1:][1:]
m1 = m1 ** 2
m2 = m2 ** 2
print("M1 es {}".format(m1.shape))
t = 0
t += sum(sum(m1)) + sum(sum(m2))
c += r*t
return c
def prueba_coste2():
(X,y) = cargaDatos()
T1, T2 = cargaPesos()
y = np.array(pasa_a_onehot(y, y_to_one_hot(10)))
c = coste_redNeuronalReg(X,y,T1,T2,1)
print(c)
prueba_coste2()
```
## Cálculo del Gradiente
```
# params_rn = array unidimensional de parámetros de la red neuronal
def backprop(params_rn, num_entradas, num_ocultas, num_etiquetas, X, y):
Theta1 = np.reshape(params_rn[:num_ocultas * (num_entradas +1)], (num_ocultas, (num_entradas + 1)))
Theta2 = np.reshape(params_rn[num_ocultas * (num_entradas +1):], (num_etiquetas, (num_ocultas + 1)))
Theta = np.array([Theta1, Theta2])
A2, Z, H = prop_delante2(X,Theta)
A = []
A.append(X)
for i in range(len(A2)):
A.append(A2[i])
Delta1 = np.zeros(np.shape(Theta1))
Delta2 = np.zeros(np.shape(Theta2))
m = X.shape[0]
d3 = H - y
print(len(A2))
print(len(A))
print(A[2])
g = A[2] * (1 - A[2])
print(g)
d2 = np.dot(Theta2.T, d3)
d2 = d2 * g
Delta1 = Delta1 + np.dot(d2 , A[1])
Delta2 = Delta2 + np.dot(d3 , A[2])
coste = coste_redNeuronal(X, y, Theta1, Theta2)
return coste, Delta1, Delta2
def backprop_reg(params_rn, num_entradas, num_ocultas, num_etiquetas, X, y, reg):
c, D1, D2 = backprop(params_rn, num_entradas, num_ocultas, num_etiquetas, X, y)
m = X.shape[0]
Theta1 = np.reshape(params_rn[:num_ocultas * (num_entradas +1)], (num_ocultas, (num_entradas + 1)))
Theta2 = np.reshape(params_rn[num_ocultas * (num_entradas +1):], (num_etiquetas, (num_ocultas + 1)))
c = coste_redNeuronalReg(X,y,Theta1,Theta2,reg)
D1 = D1 + (reg/m)*Theta1
D2 = D2 + (reg/m)*Theta2
return c,D1,D2
```
## Comprobación del Gradiente
```
from checkNNGradients import checkNNGradients
checkNNGradients(backprop_reg, 0.1)
```
| github_jupyter |
```
from IPython.display import Image
```
# Policy Based Methods
These notes are from all over the web. Lots of it is from [Udacity Deep Reinforcement Learning](https://classroom.udacity.com/nanodegrees/nd893) course.
Policy-based methods can learn either stochastic or deterministic policies, and they can be used to solve environments with either finite or continuous action spaces.
There are three reasons why we consider policy-based methods:
- Simplicity: Policy-based methods directly get to the problem at hand (estimating the optimal policy), without having to store a bunch of additional data (i.e., the action values / Q table) that may not be useful.
- Stochastic policies: Unlike value-based methods, policy-based methods can learn true stochastic policies.
- Continuous action spaces: Policy-based methods are well-suited for continuous action spaces.
-------
Below some attempts to solve environments using these methods:
- Black Box Methods
- [Hill-Climbing](./hill-climbing/Hill_Climbing.ipynb)
- [Mountain Car Continuous](./cross-entropy/CEM.ipynb) (run this with GPU)
- REINFORCE implementation
- [Cartpole](./reinforce/REINFORCE.ipynb)
- [Pong (from pixels)](./pong/pong-REINFORCE.ipynb) (run this with GPU)
- trained this for a long time with several different network architectures and hyperparameters, still didn't manage to beat the CPU opponent
- Proximal Policy Optimisation implementation (from pixels)
- [Pong (from pixels)](./pong/pong-PPO.ipynb) (run this with GPU)
The above exercises are from [Udacity Deep Reinforcement Learning Git](https://github.com/udacity/deep-reinforcement-learning).
Lessons learntthe hard way - `learning rate` is a very important hyperparameter and can have a huge impact on training process. (spent heaps of time tweaking with)
# 1. Black Box Optimisation Methods
Black box refers to method where we only need a function and change parameters to evaluate the function. Then we check the outcome and reiterate until convergence.
## 1.2. Hill Climbing Algorithm
.. is a relatively simple algorithm to find a (local) minima/maxima of a function.
When training an agent using this algorithm, we can use any function, for example a neural net with for example `tanh` (continuous action spaces) or `softmax` (discrete action spaces) output function to represent the policy function. We want to find a $\theta$ which maximises:

(formula copied from Udacity Lecture notes)
where *J* is expected return, $\theta$ are weights, $\tau$ trajectory (a state-action sequence ${s_0, a_0, ... s_t, a_t, s_{t+1}}$).
We can use `hill climbing` to find the optimal values for a policy function as follows:

(pseudocode copied from Udacity Lecture notes)
Replacing expected return *J* with sampled return *G* doesn't yield perfect result, but is often good enough in practice.
## 1.3. Improvements to Hill Climbing Algorithm
We cam add the following improvements to the algorithm to make training process smarter.
- `Steepest ascent hill climbing` is a variation of hill climbing that chooses a small number of neighboring policies at each iteration and chooses the best among them.
- `Simulated annealing` uses a pre-defined schedule to control how the policy space is explored, and gradually reduces the search radius as we get closer to the optimal solution.
- `Adaptive noise scaling` decreases the search radius with each iteration when a new best policy is found, and otherwise increases the search radius.
-----
# 2. Other Black Box Methods
## 2.1. Cross Entropy
The cross-entropy method iteratively suggests a small number of neighboring policies, and uses a small percentage of the best performing policies to calculate a new estimate.
## 2.2. Evolution Strategies
Another interesting improvement to blackbox methods is [Evolution Strategies](https://arxiv.org/abs/1703.03864). The evolution strategies technique considers the return corresponding to each candidate policy. The policy estimate at the next iteration is a weighted sum of all of the candidate policies, where policies that got higher return are given higher weight.
Look into [this](https://gist.github.com/karpathy/77fbb6a8dac5395f1b73e7a89300318d) piece of code to have a go.
----
# 3 Policy Gradient Methods
At high level, policy gradient method look into all state action pairs in an episode and increases probability of successful actions while it decreases probability of unsuccessful actions.
This is very similar to supervised learning where we put data through the network, get prediction and then adjust weights accordingly. The main difference is that in reinforcement learning setting the dataset changes (state, action) after each episode.
In reinforcement learning there can be conflicting opinions on which action to take given at a state. This would be equivalent to having a same picture twice with different labels in an image classification exercise.
Policy gradients learn stochastic policies so we can apply these to learn deterministic policies as well. `Exploration` is also embedded in the learned function and we don't have to force exploration like we have to when using value-based methods.
`
LOOP:
Collect an episode.
Change the weights of the policy network
if WON, increase the probability of each (state, action) combination
if LOST, decrease the probability of each (state, action) combination
`

## 3.1. Problem setup
The simplest policy gradient algorithm is called `Reinforce`. When training an agent with policy gradients, our goal is find weights ${\theta}$ in the `neural network` that maximises the `expected return` (on average the agent experiecnes trajectories with high return):

where ${\tau = s_0, a_0, ... s_H, a_H, s_{H+1}}$ represents `trajectory` and `return` ${R(\tau) = r_1, r_2, ... r_H,r_{H+1}}$ is a function of the `trajectory`. We calculate `weighted average` of the ${R(\tau)}$ where weights are given by ${P(\tau; \theta)}$, which is a probability of all possible values the ${R(\tau)}$ can take. ${H}$ denotes Horizon (often the length of the episode).
We use `trajectories` instead of `episodes`, since there we want to be able to apply this method to `continuous tasks` as well (the length of episode can be infinite).
## 3.2 Finding the ${\theta}$
To optimise this, one way is to use gradient ascent, where we use `estimated gradient` of ${m}$ trajectories, which is then multiplied by reward given the trajectory (calculating the true gradient is not feasible since that would require calculating every possible trajectory).

Very detailed way on deriving gradients and implementing reinforce algorithm can be found [here](https://medium.com/@thechrisyoon/deriving-policy-gradients-and-implementing-reinforce-f887949bd630)
In theory REINFORCE can be used in `continuous` spaces as well, just use an output layer that parametrizes a continuous probability distribution. However "vanilla" REINFORCE doesn't perform well with continuous action spaces.
## 3.3 Noise reduction
There are typically millions of gradients even in simple discrete problem, and infinite for continuous. Most of the time the sampled gradient doesn't contain much information about optimal policy since it's just randomly picked. We can reduce this noise by sampling more trajectories and then estimate the policy gradient by just averaging across all the different trajectories. It's a small tweak to vanilla version but stabilises the training process quite a bit.

When running multiple trajectories like this, we also obtain data to calculate mean and standard deviations, so we can normalise the rewards like so:

Best practice is to set rewards to 0 in case all rewards are the same and thus ${\sigma = 0}$, to avoid numerical problems.
Intuition behind this is picking half the actions to encourage / discourage while making sure the gradient ascent steps are not too large / small.
## 3.4 Credit Assignment
Since we model these problems as markov decision processes, we don't need to care about the past but only the current moment and the future. So we tweak the gradient calculation like so:

We assume the current reward includes rewards from the past as well so basically we just ignore all rewards which happened in the past. Future rewards:

While ignoring past rewards can change gradient for each `trajectory`, it doesn't change average gradient. It also reduces gradient noise and thus should speed up the training process.
# 3.5 Proximal Policy Optimisation
We can also reuse the sampled trajectories. However, we don't want to reuse same samples too many times.



PPO algorithm is implemented as follows
1. Collect some trajectories based on some policy ${\pi_\theta}$, and initialize theta prime ${\theta'=\theta}$
2. Next, compute the gradient of the clipped surrogate function using the trajectories
3. Update ${\theta'}$ using gradient ascent ${\theta'\leftarrow\theta+\alpha\nabla_{\theta'}L_{\rm sur}^{\rm clip}(\theta', \theta)}$
4. Then we repeat step 2-3 without generating new trajectories. Typically, step 2-3 are only repeated a few times
5. Set ${\theta=\theta'}$, go back to step 1, repeat.
Source: DRLND notes and [here](https://arxiv.org/pdf/1707.06347.pdf).
-----
# 3. Summary
- In deep reinforcement learning, it is common to represent the policy with a neural network.
- This network takes the environment state as input.
- If the environment has discrete actions, the output layer has a node for each possible action and contains the probability that the agent should select each possible action.
- The weights in this neural network are initially set to random values. Then, the agent updates the weights as it interacts with (and learns more about) the environment.
- Policy-based methods can learn either stochastic or deterministic policies, and they can be used to solve environments with either finite or continuous action spaces.
${\nabla_\theta log \pi(a_t|s_t) = (1-\theta)' 1/(1-\theta) = -1/(1-\theta) = -1/0.5}$
| github_jupyter |
# Evaluation of a Pipeline and its Components
[](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial5_Evaluation.ipynb)
To be able to make a statement about the quality of results a question-answering pipeline or any other pipeline in haystack produces, it is important to evaluate it. Furthermore, evaluation allows determining which components of the pipeline can be improved.
The results of the evaluation can be saved as CSV files, which contain all the information to calculate additional metrics later on or inspect individual predictions.
### Prepare environment
#### Colab: Enable the GPU runtime
Make sure you enable the GPU runtime to experience decent speed in this tutorial.
**Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
<img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/_src/img/colab_gpu_runtime.jpg">
```
# Make sure you have a GPU running
!nvidia-smi
# Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack
!pip install grpcio-tools==1.34.1
!pip install git+https://github.com/deepset-ai/haystack.git
# If you run this notebook on Google Colab, you might need to
# restart the runtime after installing haystack.
from haystack.modeling.utils import initialize_device_settings
devices, n_gpu = initialize_device_settings(use_cuda=True)
```
## Start an Elasticsearch server
You can start Elasticsearch on your local machine instance using Docker. If Docker is not readily available in your environment (eg., in Colab notebooks), then you can manually download and execute Elasticsearch from source.
```
# If Docker is available: Start Elasticsearch as docker container
# from haystack.utils import launch_es
# launch_es()
# Alternative in Colab / No Docker environments: Start Elasticsearch from source
! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(['elasticsearch-7.9.2/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT,
preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
! sleep 30
```
## Fetch, Store And Preprocess the Evaluation Dataset
```
from haystack.utils import fetch_archive_from_http
# Download evaluation data, which is a subset of Natural Questions development set containing 50 documents with one question per document and multiple annotated answers
doc_dir = "../data/nq"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/nq_dev_subset_v2.json.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# make sure these indices do not collide with existing ones, the indices will be wiped clean before data is inserted
doc_index = "tutorial5_docs"
label_index = "tutorial5_labels"
# Connect to Elasticsearch
from haystack.document_stores import ElasticsearchDocumentStore
# Connect to Elasticsearch
document_store = ElasticsearchDocumentStore(host="localhost", username="", password="", index=doc_index,
label_index=label_index, embedding_field="emb",
embedding_dim=768, excluded_meta_data=["emb"])
from haystack.nodes import PreProcessor
# Add evaluation data to Elasticsearch Document Store
# We first delete the custom tutorial indices to not have duplicate elements
# and also split our documents into shorter passages using the PreProcessor
preprocessor = PreProcessor(
split_length=200,
split_overlap=0,
split_respect_sentence_boundary=False,
clean_empty_lines=False,
clean_whitespace=False
)
document_store.delete_documents(index=doc_index)
document_store.delete_documents(index=label_index)
# The add_eval_data() method converts the given dataset in json format into Haystack document and label objects. Those objects are then indexed in their respective document and label index in the document store. The method can be used with any dataset in SQuAD format.
document_store.add_eval_data(
filename="../data/nq/nq_dev_subset_v2.json",
doc_index=doc_index,
label_index=label_index,
preprocessor=preprocessor
)
```
## Initialize the Two Components of an ExtractiveQAPipeline: Retriever and Reader
```
# Initialize Retriever
from haystack.nodes import ElasticsearchRetriever
retriever = ElasticsearchRetriever(document_store=document_store)
# Alternative: Evaluate dense retrievers (DensePassageRetriever or EmbeddingRetriever)
# DensePassageRetriever uses two separate transformer based encoders for query and document.
# In contrast, EmbeddingRetriever uses a single encoder for both.
# Please make sure the "embedding_dim" parameter in the DocumentStore above matches the output dimension of your models!
# Please also take care that the PreProcessor splits your files into chunks that can be completely converted with
# the max_seq_len limitations of Transformers
# The SentenceTransformer model "all-mpnet-base-v2" generally works well with the EmbeddingRetriever on any kind of English text.
# For more information check out the documentation at: https://www.sbert.net/docs/pretrained_models.html
# from haystack.retriever import DensePassageRetriever, EmbeddingRetriever
# retriever = DensePassageRetriever(document_store=document_store,
# query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
# passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
# use_gpu=True,
# max_seq_len_passage=256,
# embed_title=True)
# retriever = EmbeddingRetriever(document_store=document_store, model_format="sentence_transformers",
# embedding_model="all-mpnet-base-v2")
# document_store.update_embeddings(retriever, index=doc_index)
# Initialize Reader
from haystack.nodes import FARMReader
reader = FARMReader("deepset/roberta-base-squad2", top_k=4, return_no_answer=True)
# Define a pipeline consisting of the initialized retriever and reader
from haystack.pipelines import ExtractiveQAPipeline
pipeline = ExtractiveQAPipeline(reader=reader, retriever=retriever)
# The evaluation also works with any other pipeline.
# For example you could use a DocumentSearchPipeline as an alternative:
# from haystack.pipelines import DocumentSearchPipeline
# pipeline = DocumentSearchPipeline(retriever=retriever)
```
## Evaluation of an ExtractiveQAPipeline
Here we evaluate retriever and reader in open domain fashion on the full corpus of documents i.e. a document is considered
correctly retrieved if it contains the gold answer string within it. The reader is evaluated based purely on the
predicted answer string, regardless of which document this came from and the position of the extracted span.
The generation of predictions is seperated from the calculation of metrics. This allows you to run the computation-heavy model predictions only once and then iterate flexibly on the metrics or reports you want to generate.
```
from haystack.schema import EvaluationResult, MultiLabel
# We can load evaluation labels from the document store
eval_labels = document_store.get_all_labels_aggregated(drop_negative_labels=True, drop_no_answers=False)
# Alternative: Define queries and labels directly
# from haystack.schema import Answer, Document, Label, Span
# eval_labels = [
# MultiLabel(labels=[Label(query="who is written in the book of life",
# answer=Answer(answer="every person who is destined for Heaven or the World to Come",
# offsets_in_context=[Span(374, 434)]),
# document=Document(id='1b090aec7dbd1af6739c4c80f8995877-0',
# content_type="text",
# content='Book of Life - wikipedia Book of Life Jump to: navigation, search This article is about the book mentioned in Christian and Jewish religious teachings. For other uses, see The Book of Life. In Christianity and Judaism, the Book of Life (Hebrew: ספר החיים, transliterated Sefer HaChaim; Greek: βιβλίον τῆς ζωῆς Biblíon tēs Zōēs) is the book in which God records the names of every person who is destined for Heaven or the World to Come. According to the Talmud it is open on Rosh Hashanah, as is its analog for the wicked, the Book of the Dead. For this reason extra mention is made for the Book of Life during Amidah recitations during the Days of Awe, the ten days between Rosh Hashanah, the Jewish new year, and Yom Kippur, the day of atonement (the two High Holidays, particularly in the prayer Unetaneh Tokef). Contents (hide) 1 In the Hebrew Bible 2 Book of Jubilees 3 References in the New Testament 4 The eschatological or annual roll-call 5 Fundraising 6 See also 7 Notes 8 References In the Hebrew Bible(edit) In the Hebrew Bible the Book of Life - the book or muster-roll of God - records forever all people considered righteous before God'),
# is_correct_answer=True,
# is_correct_document=True,
# origin="gold-label")])
# ]
# Similar to pipeline.run() we can execute pipeline.eval()
eval_result = pipeline.eval(
labels=eval_labels,
params={"Retriever": {"top_k": 5}}
)
# The EvaluationResult contains a pandas dataframe for each pipeline node.
# That's why there are two dataframes in the EvaluationResult of an ExtractiveQAPipeline.
retriever_result = eval_result["Retriever"]
retriever_result.head()
reader_result = eval_result["Reader"]
reader_result.head()
# We can filter for all documents retrieved for a given query
retriever_book_of_life = retriever_result[retriever_result['query'] == "who is written in the book of life"]
# We can also filter for all answers predicted for a given query
reader_book_of_life = reader_result[reader_result['query'] == "who is written in the book of life"]
# Save the evaluation result so that we can reload it later and calculate evaluation metrics without running the pipeline again.
eval_result.save("../")
```
## Calculating Evaluation Metrics
Load an EvaluationResult to quickly calculate standard evaluation metrics for all predictions, such as F1-score of each individual prediction of the Reader node or recall of the retriever.
```
saved_eval_result = EvaluationResult.load("../")
metrics = saved_eval_result.calculate_metrics()
print(f'Retriever - Recall (single relevant document): {metrics["Retriever"]["recall_single_hit"]}')
print(f'Retriever - Recall (multiple relevant documents): {metrics["Retriever"]["recall_multi_hit"]}')
print(f'Retriever - Mean Reciprocal Rank: {metrics["Retriever"]["mrr"]}')
print(f'Retriever - Precision: {metrics["Retriever"]["precision"]}')
print(f'Retriever - Mean Average Precision: {metrics["Retriever"]["map"]}')
print(f'Reader - F1-Score: {metrics["Reader"]["f1"]}')
print(f'Reader - Exact Match: {metrics["Reader"]["exact_match"]}')
```
## Generating an Evaluation Report
A summary of the evaluation results can be printed to get a quick overview. It includes some aggregated metrics and also shows a few wrongly predicted examples.
```
pipeline.print_eval_report(saved_eval_result)
```
## Advanced Evaluation Metrics
As an advanced evaluation metric, semantic answer similarity (SAS) can be calculated. This metric takes into account whether the meaning of a predicted answer is similar to the annotated gold answer rather than just doing string comparison.
To this end SAS relies on pre-trained models. For English, we recommend "cross-encoder/stsb-roberta-large", whereas for German we recommend "deepset/gbert-large-sts". A good multilingual model is "sentence-transformers/paraphrase-multilingual-mpnet-base-v2".
More info on this metric can be found in our [paper](https://arxiv.org/abs/2108.06130) or in our [blog post](https://www.deepset.ai/blog/semantic-answer-similarity-to-evaluate-qa).
```
advanced_eval_result = pipeline.eval(
labels=eval_labels,
params={"Retriever": {"top_k": 1}},
sas_model_name_or_path="cross-encoder/stsb-roberta-large"
)
metrics = advanced_eval_result.calculate_metrics()
print(metrics["Reader"]["sas"])
```
## Evaluation of Individual Components: Retriever
Here we evaluate only the retriever, based on whether the gold_label document is retrieved.
```
## Evaluate Retriever on its own
retriever_eval_results = retriever.eval(top_k=5, label_index=label_index, doc_index=doc_index)
# Retriever Recall is the proportion of questions for which the correct document containing the answer is
# among the correct documents
print("Retriever Recall:", retriever_eval_results["recall"])
# Retriever Mean Avg Precision rewards retrievers that give relevant documents a higher rank
print("Retriever Mean Avg Precision:", retriever_eval_results["map"])
```
## Evaluation of Individual Components: Reader
Here we evaluate only the reader in a closed domain fashion i.e. the reader is given one query
and its corresponding relevant document and metrics are calculated on whether the right position in this text is selected by
the model as the answer span (i.e. SQuAD style)
```
# Evaluate Reader on its own
reader_eval_results = reader.eval(document_store=document_store, device=devices[0], label_index=label_index, doc_index=doc_index)
# Evaluation of Reader can also be done directly on a SQuAD-formatted file without passing the data to Elasticsearch
#reader_eval_results = reader.eval_on_file("../data/nq", "nq_dev_subset_v2.json", device=device)
# Reader Top-N-Accuracy is the proportion of predicted answers that match with their corresponding correct answer
print("Reader Top-N-Accuracy:", reader_eval_results["top_n_accuracy"])
# Reader Exact Match is the proportion of questions where the predicted answer is exactly the same as the correct answer
print("Reader Exact Match:", reader_eval_results["EM"])
# Reader F1-Score is the average overlap between the predicted answers and the correct answers
print("Reader F1-Score:", reader_eval_results["f1"])
```
## About us
This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
We bring NLP to the industry via open source!
Our focus: Industry specific language models & large scale QA systems.
Some of our other work:
- [German BERT](https://deepset.ai/german-bert)
- [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
- [FARM](https://github.com/deepset-ai/FARM)
Get in touch:
[Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
By the way: [we're hiring!](https://www.deepset.ai/jobs)
| github_jupyter |
# Chap 12: Implementing a Multilayer Artificial Neural Network from Scratch
Deep learning có thể hiểu là một trường con của Machine learning tập trung vào việc huấn luyện mạng nhân tạo thông qua nhiều lớp một cách hiệu quả. Trong chương này, ta sẽ học về concept cơ bản của NNs.
Những chủ đề trong phần này bao gồm:
* Hiểu được concept chung về Multilayer NNs.
* Cài đặt hàm backpropagation cơ bản cho quá trình training NN.
* Training NNs cơ bản để phân loại hình ảnh.
## Modeling complex functions with artificial neural networks
Lịch sử của quá trình hình thành NNs, đặc biệt là một giai đoạn gọi là [**AI_Winter**](https://en.wikipedia.org/wiki/AI_winter).
Một mô hình NNs bao gồm thằng kiến trúc cũng như là thuật toán để học nó nữa.
Nhiều ứng dụng của DNNs được phát triển tại nhiều công ty như:
* Facebook's DeepFace
* Baidu's DeepSpeech
* Google's new language translation service
* Novel techniques for drug discovery and toxicity prediction (Toxicity prediction using Deep Learning, T. Unterthiner, A. Mayr, G. Klambauer, and S. Hochreiter, arXiv preprint arXiv:1503.01445, 2015)
## Single-layer neural network recap
Giới thiệu lại những mô hình Single-Layer NNs được giới thiệu trong chương 2
Trong chương 2, ta cài đặt ADALINE để thực hiện bài toán phân loại nhị phân, ta sử dụng Gradient Descent Optimization đẻ học các hệ số trọng lượng của model. Trong mỗi epoch (mỗi lần duyệt hết qua các training trong dữ liệu), ta cập nhật trọng số cho vectorr **w** thông qua luật:
$w := w + \Delta w$ với giá trị $\Delta w = -\eta \triangledown J(w)$
Nói cách khác, ta tính toán giá trị Gradient dựa trên toàn bộ dữ liệu train và cập nhật trọng số weitghts bằng cách đi theo hướng ngược lại hướng của vector gradient $\triangledown J(w)$. Để có thể tìm được trọng số tối ưu cho model, ta tối thiểu hóa hàm mục tiêu của mình đó là hàm được định nghĩa mang tên **Sum of Square Errors (SSE)** ký hiệu cho cost function J(w). Hơn nữam ta nhân Gradient với một hệ số, gọi là **learning rate** $\eta$, hệ số này phải được lựa chọn kỹ để cân bằng giữa tốc độ học và nguy cơ dẫn đến overshooting global minimum của hàm cost function.
Trong Gradient Descent Optimization, ta cập nhật trọng số weights đồng thời sau mỗi Epoch, như ta định nghĩa đó là đạo hàm riêng phần của các $w_{j}$ trong vector **w** như sau:
$\frac{\partial }{\partial w_{j}} J(w) = - \sum_{}^{i}(y^{(i)} - a^{(i)})x_{j}^{(i)}$
Với giá trị các $y^{(i)}$ là nhãn đúng của mẫu $x^{(i)}$, và $a^{(i)}$ là giá trị sau khi ra khỏi hàm kích hoạt của neuron, là một hàm linear function trong trường hợp của Adaline.
Hơn nữa, ta định nghĩa hàm activation function như sau:
$\phi(z) = z = a$
Với z : netinput, là tích vector của mảng các weights kết nối giữa đầu vô và đầu ra
$z = \sum^{}_{j} w_{j}x_{j}=w^{T}x$
Trong khi sử dụng hàm Activation $\phi(z)$ để tính toán giá trị Update của gradient, ta cài đặt thêm hàm threshold để chuyển các giá trị liên tục thành các giá trị rời rạc nhị phân phục vụ việc dự đoán:
$\hat{y} = 1 (nếu g(z) \geq 0)hoặc -1$
#### Single-layer naming convention
Chú ý rằng mặc dù Adaline bao gồm 2 layers, một cho input và một cho output, nó vẫn được gọi là single-layer network bởi vì nó có 1 liên kết đơn giữa lớp input và output của nó thôi.
Ta cũng đã tìm hiểu về trick để tăng tốc độ học lên thông qua phương pháp mang tên **stochastic gradient descent (SGD)**. SGD xấp xỉ hàm cost function từ một mẫu training hoặc là một tập training nhỏ thôi (mini-batch training). Ta tận dụng concept này ở chương sau khi tiếp cận và train Multilayer perceptron (MLP). Ngoài việc học nhanh hơn do có nhiều trọng số được cập nhật thường xuyên hơn so với Gradient Descent - thuộc tính Noisy của nó cũng là một trong những lợi thế khi train multilayer NNs với hàm kích hoạt không tuyến tính, hàm đó không có hàm lồi. Ở đây các giá trị noise thêm vào có thể giúp ta thoát khỏi cực trị địa phương, ta sẽ thảo luận nhiều hơn trong chương này.
## Introducing the multilayer neural network architecture
Trong phần này, ta sẽ học cách kết nối các neuron đơn lại thành một mô hình đa lớp multilayer feedforward NN; trường hợp đặc biệt của nó là fully connected network còn được gọi là **MLP**.
Hình bên dưới minh hoạ cho concept của MLP trong đó chứa 3 layers:

**MLP** như hình trên bao gồm một input layer, một hidden layer và một output layer. Mỗi phần tử đơn vị trong hidden layer được kết nối fully connect với lớp input layer, và output layer cũng kết nối fully connect với lại thằng hidden layer. Nếu mô hình có nhiều hơn một hidden layer, ta gọi nó là **deep artificial NN**.
#### Adding additional hidden layers
Ta có thể thêm bất kỳ số lượng lớp ẩn nào vào trong MLP để có thể tạo ra một kiến trúc sâu hơn. Ta có thể tưởng tượng số lớp và số đơn vị trong mỗi lớp như là các tham số mà ta cần phải chọn sao cho tối ưu cho bài toán cụ thể sử dụng **cross-validation technique** (**Mình sẽ đọc về phần này ở trong chương 6 sau khi xong cái mục ni**)
Tuy nhiên rằng, càng nhiều lớp thì cái error gradients được tính toán thông qua hàm backpropagation nó sẽ hội tụ chậm hơn. Vấn đề này đặt ra một thách thức lớn cho các model hiện nay. Đo đó tạo ra các thuật toán đặc biệt được phát triển để có thể train được kiến trúc DNN này; những thuật toán đó được gọi là **deep learning**.
Trong hình ở trên, ký hiệu đơn vị **activation** thứ i của hớp thứ l là $a_{i}^{(l)}$. Ta ký hiệu **in** là input layer, **out** là output layer, h là **hidden layer**. Ví dụ $a_{i}^{(in)}$ là giá trị thứ i trong lớp input layer,$a_{i}^{(h)}$ là đơn vị thứ i trong lớp hidden layer, và $a_{i}^{(out)}$ là đơn vị thứ i trong lớp output. Các giá đơn vị kích hoạt $a_{0}^{(in)}$ và $a_{0}^{(h)}$ được gọi là các bias units, và có giá trị bằng 1. Hàm kích hoạt cho các units trong lớp input layer đơn giản chỉ là đầu vào và cộng thêm với thằng bias unit:

#### Notational convention for the bias unit
Trong phần sau của chương, ta sẽ cài đặt một MLP sử dụng vectơ riêng để chứa các giá trị bias, làm cho quá trình cài đặt hiệu quả và dễ đọc hơn. Concept này cũng đc sử dụng trong TensorFlow. Tuy nhiên công thức toán sẽ phức tạp hơn. Đây chỉ là một cách trình bày khác hơn thôi, chứ bản chất không có gì thay đổi cả.
## Mô tả cấu trúc
Mỗi đơn vị trong lớp l được kết nối với toàn bộ các unit khác trong lớp thứ l + 1 thông qua trọng số weight giữa chúng. Ví dụ, sự kết nối giữa thằng đơn vị thứ k trong lớp l với lại đơn vị thứ j trong lớp l + 1 được ký hiệu là $w_k,j^{l}$. Tham khảo lại cái hình phía trước, ta ký hiệu ma trận trọng số kết nối cái input với thằng lớp ẩn là $W^{(h)}$, và ký hiệu thằng ma trận kết nối lớp ẩn với lại thằng ouput là $W^{(out)}$.
Trong khi đó mỗi thằng đơn vị trong lớp output layer đảm nhiệm việc phân loại nhị phân, ta sẽ thấy loại tổng quan hơn của NN trong hình bên trên, cho phép chúng ta biểu diễn bài toán phân loại nhiều lớp (multiclass classification) thông qua kỹ thuật tổng quan hóa **one-versus-all(OvA)** technique. Để có thể hiểu hơn nó làm những gì, ta tưởng tượng lại kiểu **one-hot encoding** cho những thằng categorical variable trong chương 4.
Ví dụ, ta có thể encode 3 nhãn class labels trong dữ liệu IRIS (0=Setosa, 1=Versicolor, 2=Virginica) như sau:

Kiểu biểu diễn one-hot này cho phép chúng ta xử lý với những bài toán phân loại những lớp kiểu này.
Trong phần sau ta sẽ vector hóa những ký hiệu subscript và superscript. Ta tổng hợp lại thằng ma trận trọng số kết nối giữa thăng input và thằng hidden là ma trận $W^{(h)} \in \mathbb{R}^{m*d}$, với d là số lượng của thằng node con phía sau (không kể thằng bias) và thằng m là số lượng của thằng input units bao gồm cả bias unit. Bởi vì việc nắm được kiến trúc của thằng này là quan trong, nên ta tổng hợp lại những gì đã trình bày bằng hình vẽ cho MLP 3-4-3 sau:

## Activation a neural network via forward propagation
Trong phần này, ta sẽ mô tả tiến trình mang tên **forward propagation**(lan truyền thẳng) để tính toán giá trị output cho MLP model. Để có thể hiểu hơn concept của việc học một MLP Model, ta tổng hợp quá trình học thông qua 3 bước:
1. Bắt đầu tại input layer, ta sử dụng lan truyền thẳng dữ liệu training data qua mạng network để tạo ra output.
2. Dựa vào network output, ta tính toán giá trị lỗi mà chúng ta muốn tối thiểu bằng cách sử dụng hàm cost function mà ta sẽ định nghĩa trong phần sau.
3. Ta backpropagate cái lỗi đó, tìm cái đạo hàm đối với từng thằng trọng số trong network, và cập nhật lại model.
Cuối cùng, khi ta lặp lại các tiến trình này trải qua nhiều Epochs và học các trọng số có trong MLP, ta sẽ sử dụng cái forward propagation để tính toán giá trị output và áp dụng hàm ngưỡng thresh hold function để đạt được cái nhãn dự báo theo kiểu one-hot encoding.
Giờ ta mô tả chi tiết các bước của forward propagation để tạo ra output từ những thằng mẫu trong dữ liệu train. Do mỗi đơn vị trong lớp hidden layer được kết nối với tất cả những thằng trong lớp input layer, đầu tiên ta tính gía trị activation unit cho thằng hidden layer $a_{1}^{(h)}$ như sau:


(tức là giá trị a tại node sau bằng hàm activation của net input của toàn bộ thằng input trước đổ về thằng phía sau của nó, cũng dễ hiểu mà :v)
Bổ sung thêm, ở đây $z_1^{(h)}$ là giá trị của net input và $\phi(*)$ được gị là hàm kích hoạt activation function, hàm này nên phải khác biệt để có thể học đc các trọng số kết nối giữa những thằng neuron sử dụng phương thức Gradient Descent. Để có thể giải được vấn đề phức tạp này như là phân loại hình ảnh, ta sử dụng hàm phi tuyến **nonl-linear** cho model MLP của mình, ví dụ như hàm sigmoid (logistic) activation function mà ta đã được học trong chương 3:

Như ta đã gọi, thì hàm sigmoid có dạng hình chữ S và chiếu một giá trị bất kỳ của net input z sang dạng phân phối logistic trong khoảng từ 0 đến 1, và cắt trục y tại điểm z = 0, được biểu diễn theo hình vẽ:

MLP là một trong những dạng NN feedforward. Thuật ngữ **feedforward** liên quan đến sự thật là mỗi lớp đóng vai trò làm đầu vào cho những lớp tiếp theo mà không có vòng lặp giữa chúng, ở phía ngược lại trong **recurrent** NNs - một kiểu kiến trúc ta sẽ thảo luận trong phần sau và thảo luận chi tiết trong chương 16. Thuật ngữ Multilayer perceptron có thể làm chúng ta dễ nhầm lẫn. Do rằng những đơn vị trong từng lớp thực chất là những đơn vị kích họat sigmoid units, không phải là các perceptrons. Ta có thể tưởng tượng là mỗi một neuron trong MLP như là một đơn vị logistic regression units và return lại các giá trị trong khoảng từ 0 đến 1.
Để làm cho code dễ đọc và thực thi hiêụ quả hơn, ta sẽ viết hàm activation function dưới dạng phức tạp hơn sử dụng concept của đại số tuyến tính, cho phép chúng ta vectơ hóa cài đặt thuật toán của mình thông qua NumPy thay vì viết những hàm for loop tù:


(Tức là thay vì làm những vòng for dài ngoằng để tính toán các phần tử trong matrận thì ta sử dụng thằng có sẵn của NumPy để rút gọn công thức tính lại)
Với $a^{(in)}$ là vector 1 * m chiều (một dòng và m cột) và chính là mẫu $x^{(in)}$ có thêm thằng bias ở đầu tiên của mảng
$W^{(h)}$ là ma trận m * d chiều với d là số lượng đơn vị units trong lớp ẩn của chúng ta. Sau khi thực hiện phép nhân ma trận, ta được một ma trận mới kích thước 1 * d chiều gọi là net input $z^{(h)}$ để dùng nó áp vào tính giá trị activation $a^{(h)}$ (với $a^{(h)} \in \mathbb{R}^{1xd}$).
Hơn nữa, ta có thể tổng quát hóa quá trình tính toán này đối với n mẫu trong quá trình train dữ liệu của chúng ta:

với thằng $A^{(in)}$ bây giờ là ma trận n * m chiều, và phép nhân 2 ma trận sẽ cho ra kết quả ma trận $Z^{(h)}$ là n * d chiều. Cuối cùng, ta áp dụng hàm activation funtion $\phi(.)$ cho mỗi giá trị của thằng net input và thu được ma trận kích hoạt n * d chiều và là lớp tiếp theo của ta (trong phần này chính là cái lớp output):

Tương tự, ta có thể viết hàm activation function cho giá trị output layer:

Ở đây, ta nhân một ma trận với kích thước d * t chiều $W^{(out)}$ (t là số lượng output units) với lại ma trận có kích thước n * d chiều $A^{(h)}$ để tại ra ma trận input net n * t chiều $Z^{(out)}$ (các cột trong ma trận được tạo thành là đại diện cho các output của từng mẫu).
Cuối cúng, ta áp dụng hàm sigmoid function để có được các giá trị đầu ra liên tục trong khoảng từ 0 đến 1

Với $A^{(out)} \in \mathbb{R}^{n x t}$
## Classifying handwritten digits
Trong phần trước, ta đã trình bày lý thuyết về NNs. Trước khi tiếp tục thảo luận về thuật toán học được giá trị trọng số của MLP model (đó là giai đoạn backpropagation), giờ ta bắt đầu những bước cài đặt đầu tiên
#### Additional resource on backpropagation
* Lý thuyết về NN khá phức tạp, do đó có thêm những nguồn giải thích cụ thể về thằng NNs này trong chương 6:(http://www.deeplearningbook.org)
* Pattern Recognition and Machine Learning, C. M. Bishop and others, Volume 1. Springer New York, 2006
* Lecture slides from the deep learning course at the University of Wisconsin–Madison:
1. https://sebastianraschka.com/pdf/lecture-notes/stat479ss19/L08_logistic_slides.pdf
2. https://sebastianraschka.com/pdf/lecture-notes/stat479ss19/L09_mlp_slides.pdf
Trong phần này, ta sẽ cài đặt mô hình NN đầu tiên để phân loại chữ viết tay từ nguồn dữ liệu **Mixed National Institute of Standards and Technology (MNIST)** được xây dựng bởi Yann LeCun và các cộng sự bắt đầu từ năm 1998.
## Obtaining and preparing the MNIST dataset
Dữ liệu có thể tải về từ (http://yann.lecun.com/exdb/mnist/) và bao gồm 4 files:
* Training dataset images
* Training dataset labels
* Test dataset images
* Test dataset labels
Dữ liệu training bao hàm chữ viết tay của 250 người, 50% là của sinh viên, và 50% là của nhân viên trong công ty. Chú ý rằng dữ liệu test data chứa dữ liệu viết tay với tỉ lệ tương tự dữ liệu train.
Sau khi tải về, sử dụng Linux gzip tool để mà giải nén. Sử dụng command sau:
gzip *ubyte.gz -d
Ta cũng có thể sử dụng tool bất kỳ để giải nén khi làm việc với môi trường Windows.
Hình ảnh được lưu trữ với định dạng byte format, ta sẽ đọc chúng vào trong NumPy và sử dụng để train và Test MLP model. Để thực hiện được việc đó, ta định nghĩa một số hàm helper:
(Đoạn code ta có thể tìm thấy trong project đi kèm trong chương này)
Phương thức load_mnist trả lại 2 mảng, mảng đầu tiên có kích thước n * m chiều NumPy array, bới n là số mẫu và n là số lượng feature (ở đây là pixels). Dữ liệu train bao gồm 60000 training digits và test data chưa 10000 examples.
Hình ảnh trong MNIST dataset là định dạng 28 * 28 pixel, và mỗi pixel là giá trị của grayscale intesity values. Ở đây ta duỗi 28 * 28 pixels thành một mảng vector một chiều, đại diện cho các hàng trong mảng image array (784 features với mỗi một tấm hình). Mảng thứ 2 trả lại mảng các nhãn tương ứng cho các thành phần mình đã đọc được trong dữ liệu train hoặc là test.
cách mà chúng ta đọc ảnh cũng khá là lạ:
```python
magic, n = struct.unpack('>II', lpath.read(8))
labels = np.fromfile(lbpath, dtypnp.unit8)
```
Để hiểu cách những dòng code này thực hiện, giờ ta xem xét đến mô tả dữ liệu của MNIST

Sử dụng 2 dòng code trên, đầu tiên ta đọc vào magic number, được mô tả trong bảng trên, cũng như là số lượng mẫu n từ file buffer. Trươc khi ta load chúng vào trong mảng NumPy sử dụng phương thức fromfile. Tham số fmt, '>II' mà ta pass vào trong struct.unpack bao gồm 2 thành phần:
* > : Đây là ký hiệu của big-endian- định nghiã thứ tự của các byte được lưu trữ.
* I: Đây là ký hiệu của Unsigned Integer
Cuối cùng, ta normalized những giá trị pixels trong MNIST về lại khoảng -1 tới 1 (thông thường là 0 đến 255) thông qua bước xử lý sau:
```python
images = ((images / 255.) - .5) * 2
```
Lý do ở đây là Gradient-Descent Optimization ổn định hơn dưới những điều kiện mà ta đã tạo ra được,cái này đã được đề cập trong chương 2. Chú ý rằng image mà ta đã scale được là nhờ các phép toán cơ bản, điều này khác với những kỹ thuật scale mà ta đã nói trong chương 2 đó.
Ta đã có được các giá trị scaled từ training dataset và sử dụng nó để scale các cột trong training dataset và test dataset. Tuy nhiên, khi làm việc với image pixels, center chúng ở zero và chuyển trạng thái của chúng về khoảng [-1, 1] là thông dụng và được áp dụng rộng rãi trong thực tế.
#### Batch normalization
Một trong số những trick để cải thiện độ hội tụ của Gradient-based Optimization đó là batch normalization, kỹ thuật này được đề cập trong chường 17. Ta có thể đọc thêm về batch normalization tại nguồn tài liệu này (https://arxiv.org/abs/1502.03167)
Bằng cách thực thi đoạn code bên dưới, ta có thể load được 60000 mẫu trains cũng như là 10000 mẫu test.
Để quan sát được các mẫu test, ta có thể vẽ một vài tấm hình minh họa cho các số trong khoảng từ 0 đến 9 cũng như là các bản khác nhau của cùng một chữ số.
Sau khi qua các bước trên, một ý tưởng tốt hơn là ta save lại các giá trị scaled của các images trong một định dạng mới hơn mà ta có thể load ra một cách nhanh chóng hơn trong Python session để tránh tình trạng quá tải khi mà xử lý dữ liệu lại. Khi chúng ta làm việc với NumPy arrays, một biện pháp hiệu quả và thuận tiện đó là lưu giá trị của các array nhiều chiều đó vào trong đĩa thông qua hàm NumPy savez.
Tài liệu chi tiết của nó: (https://docs.scipy.org/doc/numpy/reference/generated/numpy.savez.html)
Hàm savez nó cũng tương tự như cái module Pickle của Python vậy, ta đã sử dụng nó trong chapter 9, nhưng nó tối ưu cho việc lưu trữ các dữ liệu NumPy Arrays. Hàm savez tạo ra một file nén định dạng .npz; ta đọc thêm về nó tại đây (https://docs.scipy.org/doc/numpy/neps/npy-format.html). Hơn nữa, thay vì sử dụng hàm savez, ta có thể sử dụng hàm savez_compressed, nó cũng có cú pháp tương tự như hàm savez, nhưng nó nén dữ liệu lại tốt hơn (chỉ còn lại 22Mb thay vì 400Mb khi dùng với savez). Đoạn code bên dưới mô tả cách lưu dữ liệu train và test data vào trong file mnist_scaled.npz:
Ta có thể load toàn bộ dữ liệu ra lại thông qua list comprehesion, và cái np.load() này nó trả lại những gì mình đã nén trước đó, thuộc tính files cho phép xem các trường có trong dữ liệu đã được load ra lại.
Chú ý rằng những thứ savez_compressed và np.load là không cần thiết cho ví dụ ở đây, tại vì ta đang thực hiện trên dữ liệu không lớn lắm. Nhưng mà trong thực tế thì ta vẫn hay dùng những thằng như vậy lắm.
#### Loading MNIST using sklearn
Sử dụng sklearn method fetch_openml, ta có thể load được dữ liệu của MNIST một cách thuận tiện. Ví dụ ta có thể sử dụng đoạn code bên dưới để có thể tạo ra 50000 training data set và 10000 test dataset.
(Code mình để trong thư mục hiện hành thôi)
Chú ý rằng cái phân bổ của MNIST trong training và testing dataset sẽ khác với cách tiếp cận thông thường. Do đó, ta có thể thấy được sự khác biệt nhẹ giữa 2 bộ dữ liệu được tạo bởi 2 cách khác nhau (Ta thấy thằng Sklearn nó chậm quá nên thôi, tải data về rồi chiến là được rồi)
## Implementing a multilayer perceptron
Trong phần này, ta sẽ cài đặt thằng MLP từ đầu để có thể phân biệt được dữ liệu trong Dataset của chúng ta. Để làm cho mọi thứ đơm giản, ta sẽ cài đặt 1 lớp ẩn thôi.
## Training an artificial neural network
Phần này trình bày kỹ hơn về cách áp dụng logistic regression mà đã trình bày trong chương 3
### Computing the logistic cost function
Hàm logistic cost function được cài đặt trong phương thức _compute_cost tương tự với hàm cost function mà ta đã cài đặt trong logistic regression trong chương 3:

Ở đây, giá trị $a^{[i]}$ là giá trị sigmoid activation của mẫu thứ i trong dữ liệu, và ta tính toán nó như sau: $a^{[i]} = \phi(z^{[i]})$
Chú ý, ký hiệu [i] là chỉ số của mẫu train, không phải là chỉ số của lớp. Giờ ta cho thêm thằng regularization, cho phép ta giảm thiểu overfitting. Hàm L2 được định nghĩa như sau (chú ý là ta không cần phải regularize các giá trị bias units):

Bằng việc thêm giá trị L2 regularization vào trong hàm logistic cost function, ta có được công thức sau:

Tưởng tượng mô hình của ta khi dự đoán kiểu dữ liệu cho thằng dữ liệu có nhãn là 2, thì hàm activation của lớp thứ 3 và target (one hot) tương ứng phải là:

Do đó, khi ta muốn tổng quá hàm logistic cost function cho tất cả t các activation units trong network của minh. Thì hàm cost function (khi chưa có regularization) trở thành:

Với giá trị [i] ký hiệu cho mẫu thứ i trong dữ liệu training dataset.
Hàm cost có thêm regularization có chút phức tạp
## Training neural networks via backpropagation
Trong phần này, ta nghiên các hàm back propagation hoạt động để update weights trong model MLP của chúng ta, trong code được trình bày đoạn # Backpropagation trong phương thức fit. Trong phần trướcm ta đầu tiên phải apply forward propagation để có thể đạt được activation của output layer, trình bày bằng công thức sau:
$Z^{(h)} = A^{(in)}W^{(h)}$ (net input of the hidden layer)
$A^{(h)} = \phi(Z^{(h)})$ (activation of the hidden layer)
$Z^{(out)} = A^{(h)}W^{(out)}$ (net input of the output layer)
$A^{(out)} = \phi(Z^{(out)})$ (activation of the hidden layer)
Quá trình này được miêu tả bằng hình vẽ sau:

Trong pha bacpropagation, ta tính toán giá trị error từ phải sang trái.Ta bắt đầu tính toán giá trị error vector của output layer:

Với y là vector của true class labels (giá trị trong công thức tương đương với thằng delta_out trong code của mình)
Tiếp theo, ta tính giá trị delta cho thằng hidden layer:

Với giá trị $\frac{\partial \phi (z^{(h)})}{\partial z^{(h)}$ là đạo hàm của hàm sigmoid activation function, giá trị này được tính bằng công thức sigmoid_derivative_h = a_h * (1. - a_h) trong method fit của NeuralNetMLP:

Chú ý rằng ký hiệu $\odot$ là element-wise multiplication. (Hay còn được gọi là Hadamard product).
### Note: chứng minh công thức
Chứng minh đạo hàm của thằng activation function với biến vào là z (net input):

Theo công thức này ta thấy được $\phi'(z) = a(1-a)$ với a là giá trị activation tại thằng z.
Tiếp theo, ta tính giá trị $\delta^{(h)}$ tại lớp hidden tính toán như sau:

Giải thích cho công thức này, ta dùng phép transpose ma trận h * t $W^{(out)}$. Với t là số lượng output class labels (trong bài toán cụ thể là 10) và h là số lượng hidden units. Phép nhân ma trận giữa ma trận n * t $\delta^{(out)}$ và ma trận t * h (W^{(out)})^{T} cho ra ma trận n * h và thông qua phép nhân hadamard với lại đạo hàm của sigmoid với cùng số chiều là n * h để có được ma trận $\delta^{(h)}$.
Sau đó, khi ta có được giá trị $\delta$ của các lớp, giờ ta có thể biểu thị gía trị của thằng cost function như sau:


Các công thức trên dễ hơn khi ta vectơ hóa chúng lại như sau:


Và ta thêm thằng regularization vào:

Chú ý rằng những thằng bias unit thông thường không cần phải regularized. Những thằng delta_w_h, delta_b_h, delta_w_out, delta_b_out được trình bày trong code của mình.
Cuối cùng, sau khi ta tính toán được giá trị của các gradients, ta có thể cập nhật trọng số bằng cách đi theo hướng ngược lại đối với mỗi layer l:

Công thức này được cài đặt trong thực tế sẽ như sau :
``` python
self.w_h -= self.eta * delta_w_h
self.b_h -= self.eta * delta_b_h
self.w_out -= self.eta * delta_w_out
self.b_out -= self.eta * delta_b_out
```
Hình ảnh tổng hợp những gì mà thăng backpropagation làm được:

# APPENDIX
## Phụ lục thêm về phần backpropagation này
Source:http://neuralnetworksanddeeplearning.com/chap2.html
4 công thức cần nhớ về hàm backpropagation:
 (BP1)
Trong trường hợp của quadratic cost tức là , ta có được điều này: , và do đó công thức tại (BL1) trở thành:

## Công thức cho error trong lớp trước thông qua lớp sau của nó:
 (BP2)
## Công thức tính lượng thay đổi cost tương ứng với bất kỳ bias nào trong network:
 (BP3)
Ta có thể viết lại dạng gọn hơn như sau:

## Công thức tính lượng thay đổi của cost tương ứng với weight bất kỳ trong network:
 (BP4)
## Chứng minh cho 4 công thức cơ bản nói trên
* Chứng minh thăng BP1:
Theo định nghĩa ta có công thức:

Sau khi áp dụng quy tắc chuối, ta có công thức sau:

Chú ý rằng thằng output activation $a_k^L$ chỉ phụ thuộc vào thằng net input $z_j^L$ khi mà thằng k = j. Còn lại các giá trị khác chúng ko phụ thuộc vào nhau, tại những điểm đó thì gía trị của thằng $\frac{\partial a_k^L}{\partial z_j^L}$ là bằng 0. Do đó biểu thức có thể rút gọn lại thành:

Mà ta lại có rằng $a_j^L=\sigma(z_j^L)$ nên là thành phần thứ 2 của biểu thức có thể viết lại thành:

Đây chính là công thức của thằng BP1.
* Chứng minh thằng BP2:
Ta cũng sử dụng chain rule:
Phần ni mai tìm hiểu tiếp, mệt cái ni rồi. Chuyển sang cái khác học.
| github_jupyter |
# Load cleaned data
```
import pandas as pd
df = pd.read_csv('data/sample_5000_clean.csv', index_col="Unnamed: 0")
print(df.shape)
df.head()
```
# Latent Dirichlet Allocation
We use [scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.LatentDirichletAllocation.html).
Make sure to also check out [gensim](https://radimrehurek.com/gensim/) and [spacy](https://spacy.io/) and the [transformers library](https://github.com/huggingface/transformers).
```
from IPython.display import IFrame
display(IFrame("https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.LatentDirichletAllocation.html", width=1300, height=500))
# Author: Olivier Grisel <olivier.grisel@ensta.org>
# Lars Buitinck
# Chyi-Kwei Yau <chyikwei.yau@gmail.com>
# License: BSD 3 clause
# https://scikit-learn.org/stable/auto_examples/applications/plot_topics_extraction_with_nmf_lda.html#sphx-glr-auto-examples-applications-plot-topics-extraction-with-nmf-lda-py
from time import time
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.decomposition import LatentDirichletAllocation
n_samples = 3000
min_df=0.005
max_df=0.6
n_components = 15
n_top_words = 20
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
message = "Topic #%d: " % topic_idx
message += " ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]])
print(message)
print()
t0 = time()
data = df.clean_text.values
data_samples = data[:n_samples]
# Use tf (raw term count) features for LDA.
print("Extracting tf features for LDA...")
tf_vectorizer = CountVectorizer(max_df=max_df, min_df=min_df, stop_words='english')
%time tf = tf_vectorizer.fit_transform(data_samples)
lda = LatentDirichletAllocation(n_components=n_components, max_iter=5,learning_method='online',random_state=0)
%time lda.fit(tf)
print("\nTopics in LDA model:")
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, n_top_words)
```
# Wordclouds
```
from wordcloud import WordCloud, STOPWORDS
import matplotlib.pyplot as plt
def get_top_words(model, feature_names, n_top_words, topics):
""" Get top words per topic from lda model """
for topic_idx, topic in enumerate(model.components_):
if not int(topics) == topic_idx:
continue
words = [feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]]
probs = [topic[i] for i in topic.argsort()[:-n_top_words - 1:-1]]
return dict(zip(words, probs))
def sklearn_lda_wordcloud(model, feature_names, n_top_words, topic):
""" Create wordcloud from lda model """
wc = WordCloud(background_color="white").fit_words(get_top_words(model, feature_names, n_top_words, topic))
plt.imshow(wc)
plt.axis("off")
plt.title("Topic #" + str(topic))
return wc
sklearn_lda_wordcloud(lda, tf_feature_names, 100, 1)
# for i in range(15):
# wc = sklearn_lda_wordcloud(lda, tf_feature_names, 100, i)
# plt.imshow(wc)
# plt.show()
lda.wordclouds = {i:sklearn_lda_wordcloud(lda, tf_feature_names, 100, i) for i in range(n_components)}
```
# Model evaluation
- We use Perplexity for simplicity, however it is better to use topic coherence.
- Still one should not bother too much about but evaluation metrics like perplexity or topic coherence but rather inform the choice of model by the area of application. Learning more about this isa direction of current research.
```
lda.perplexity(tf)
# train second model with same hyperparameters but different random seed
lda2 = LatentDirichletAllocation(n_components=n_components, max_iter=5, learning_method='online', random_state=1)
lda2.fit(tf)
print(lda.perplexity(tf_vectorizer.transform(data[n_samples:])))
lda2.perplexity(tf_vectorizer.transform(data[n_samples:]))
```
# Hyperparameter Tuning
https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html
```
display(IFrame("https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html", width=1300, height=200))
import numpy as np
np.random.seed(42)
from sklearn.externals import joblib
from sklearn.model_selection import RandomizedSearchCV
import scipy.stats
import os
randomState = 444
n_runs = 5 # number of tries for Random Search
min_df = 0.005 # tf min df; old version: min_df = 5
max_df = 0.60 # tf max df
lda_max_iter = 5
min_topics, max_topics = 10, 20
verbose = 1 # how much feedback during optimization. 0 = no feedback, 1 = a bit more, 2 = even more
# Random Search parameter grid
param_dist = {'n_components': list(range(min_topics, max_topics )),
'learning_decay': scipy.stats.uniform(0, 1),
'doc_topic_prior': scipy.stats.uniform(0, 1), # alpha
'topic_word_prior': scipy.stats.uniform(0, 1), # beta
}
# define models
lda_random = LatentDirichletAllocation(max_iter=lda_max_iter, random_state=randomState)
model_random = RandomizedSearchCV(lda_random,
param_distributions=param_dist,
random_state=randomState,
n_iter=n_runs,
verbose=verbose ,
cv=3
)
# Do the random Search
%time model_random.fit(tf)
# Best Model
best_lda_model_random = model_random.best_estimator_
# Model Parameters
print("Best Model's Params: ", model_random.best_params_)
# Log Likelihood Score
print("Best Log Likelihood Score: ", model_random.best_score_)
random_search_results = pd.DataFrame(model_random.cv_results_).sort_values('rank_test_score')
random_search_results.head()
for i in range(5):
wc = sklearn_lda_wordcloud(best_lda_model_random, tf_feature_names, 100, i)
plt.imshow(wc)
plt.show()
```
# Save Models for later
```
import os, joblib
# make dirs
os.makedirs("results/", exist_ok=True)
os.makedirs("models/", exist_ok=True)
# save objects
joblib.dump(lda, 'models/lda')
joblib.dump(tf_vectorizer, 'models/tfvec')
# results
random_search_results.to_csv('results/random_search_results.csv', sep="\t")
tf_vectorizer
```
| github_jupyter |
# Session 5: Source Detection and Measurement
<br>Owner(s): **Imran Hasan** ([@ih64](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@ih64))
<br>Last Verified to Run: **2020-06-03**
<br>Verified Stack Release: **Weekly 2020 22**
### Learning Objectives:
After working through this tutorial you should be able to:
1. Understand how the PSF is constructed
2. Run detection on an image, along with deblending and measurement
3. Understand Footprints and HeavyFootprints, and their relationship to Detection, Deblending, and Measurement;
4. Understand some of the different measurement algorithms
5. Understand Forced Photometry, and how we combine across bands
### Logistics
This notebook is intended to be runnable on `lsst-lsp-stable.ncsa.illinois.edu` from a local git clone of https://github.com/LSSTScienceCollaborations/StackClubCourse
You can find the Stack version that this notebook is running by using eups list -s on the terminal command line:
```
# What version of the Stack am I using?
! echo $HOSTNAME
! eups list lsst_distrib -s
```
For this tutorial we'll need the following modules:
```
%matplotlib inline
import os
import numpy as np
import matplotlib as mpl
mpl.style.use('seaborn-poster')
import matplotlib.pyplot as plt
import lsst.daf.persistence as dafPersist
import lsst.afw.table as afwTable
import lsst.afw.display as afwDisplay
from lsst.pipe.tasks.characterizeImage import CharacterizeImageTask, CharacterizeImageConfig
from lsst.meas.algorithms import SourceDetectionTask, SourceDetectionConfig
from lsst.meas.deblender import SourceDeblendTask, SourceDeblendConfig
from lsst.meas.base import SingleFrameMeasurementTask, SingleFrameMeasurementConfig
from lsst.meas.base import NoiseReplacer, NoiseReplacerConfig
import matplotlib.animation as animation
from matplotlib.colors import LogNorm
from IPython.display import IFrame, display, Markdown, HTML
import lsst.meas.extensions.photometryKron
import lsst.geom as lsstGeom
mpl.style.use('seaborn-poster')
```
## Prelude : Measuring the Point Spread Function (PSF)
The detection algorithm used in the DM stack uses the transpose of the PSF as a detection kernel. So before we dive into detection, we will discuss how the PSF is constructed in the stack. This is handled in a high level task called `CharacterizeImage`.
We will need an image to run on for this section. Let's grab a `calexp`, (a single processed CCD image) to use for this section. `calexps` already have had a lot of work done on it, including instrument signiture removal, and calibrations like astrometry and estimating the point spread function. They are not expected to be distributed as a high-level science product. Nonetheless, it will give us a great opportunity to understand PSF estimation.
```
band = 'HSC-I'
depth = 'WIDE' # WIDE, DEEP, UDEEP
field = 'SSP_WIDE' # SSP_WIDE, SSP_DEEP, SSP_UDEEP
butler = dafPersist.Butler('/datasets/hsc/repo/rerun/DM-13666/%s/'%(depth))
subset = butler.subset('calexp', dataId={'filter':band, 'field':field})
dataId = subset.cache[5]
calexp = butler.get('calexp', **dataId)
```
Roughly speaking, `CharacterizeImage` PSF estimation will first use a simple Gaussian PSF as a place holder to detect very bright sources, identify point sources that can be used to model the PSF, and finally model the PSF. Let's setup and run it now
```
#Set up the task, it requires a config when we instantiate it
charConfig = CharacterizeImageConfig()
charImageTask = CharacterizeImageTask(None, config=charConfig)
#run the task on our exposure and save the output. This takes a minute to run
charResult = charImageTask.run(calexp)
```
Let's take a quick look at the output catalog. It is convinient to print it as an astropy table so we can see all the columns. There are *a lot* of columns, often several for each measurement algorithm being run. We will revisit measurement later in this notebook, but for now take note of the `calib_psf_candidate` and `calib_psf_used` columns. This tells us if the source was a candidate to be used for modeling the PSF, and if it was actually used.
```
charResult.sourceCat.asAstropy()
```
The Science Pipelines identify sources on the stellar locus in a magnitude vs size diagram to pick out candidates for determining the PSF. Let us try to visualize this now by plotting all the sources in magnitude-size space, and color code points that were PSF candidates as orange. If you browsed the output table above, you would have noticed that the Science Pipelines report instrumental fluxes, and not magnitudes. To get magnitudes, we need to do a bit of leg work. We will call the `photoCalib` associated with our `calexp`. This will tell us how to get magnitudes out of our instrumental fluxes. We will pass in the catalog along with the flux column we want to use to the `instFluxToMagnitude` method. For now lets keep things simple and use 12 pixel aperture photometry to get our magnitudes. For sizes, we will get a rough estimate by adding the second moments together from the Sdss Shape algorithm.
```
# calculate the magnitude and magnitude error for sources
# we grab the photoCalib from the calexp, and use its instFluxToMagnitude method
# instFluxToMagnitude can return a vector of magnitude and magnitude errors if you pass in
# your source catalog and the name of the flux measurement algorithm you want to use
apMag = calexp.getPhotoCalib().instFluxToMagnitude(charResult.sourceCat,'base_CircularApertureFlux_12_0')
# use the sloan algorithm to calculate something like a size
# this is just one definition of size, and the size reported here in pix^2 is not the area of the object
size = charResult.sourceCat['base_SdssShape_xx'] + charResult.sourceCat['base_SdssShape_yy']
# mask if it was not a PSF candidate
calib_mask = charResult.sourceCat['calib_psf_candidate']
# color the points by their extendendness, it will be red for resolved sources and blue for point sources
plt.scatter(apMag[:,0], size, c=charResult.sourceCat['base_ClassificationExtendedness_value'],
cmap='bwr')
# over plot PSF candidates in orange
# to get the magnitues we just need the first column from apMag
# the second column would give you magnitude errors
plt.scatter(apMag[:,0][calib_mask], size[calib_mask], color='C1')
plt.ylim([0,50])
plt.ylabel('Size $px^2$')
plt.xlabel('Magnitude')
```
We can see all the point sources (colored in blue) live on the 'stellar locus' in this diagram, and the candidate PSF stars (orange) in fact live on the stellar locus. The extended sources shown in red are likely bright galaxies that were detected.
```
# now lets verify that the calexp has a psf object 'attached' to it
calexp.hasPsf()
```
We can exercise the PSF to evaluate it at different locations on the detector. However, it will require we pass it an `lsst.geom.Point2D` object that defines that location. We demonstrate a simple example below before moving onto more complicated tricks
```
myPoint = lsstGeom.Point2D(400,500)
myPsf = calexp.getPsf().computeImage(myPoint)
plt.imshow(myPsf.getArray())
```
We can visualize how good of a job the stack did in giving us a PSF. We can use the stars that were PSF candidates but ultimately not used, to validate the PSF model. Let's look at the residuals of the PSF model evaluated at the location of these reserve stars, and the reserve stars themselves. This exercise is borrowed from Rober Lupton's [PSF.ipynb](https://github.com/RobertLuptonTheGood/notebooks/blob/master/Demos/PSF.ipynb)
```
# setup a mask to give us stars that were PSF candidates but ultimately not used for PSF estimation
calib_not_used = np.logical_and.reduce([charResult.sourceCat['calib_psf_candidate'],
~charResult.sourceCat['calib_psf_used'],
charResult.sourceCat['base_PixelFlags_flag_saturatedCenter'] == 0])
# grab the cadidate-but-not-used stars
reserve_sources = charResult.sourceCat[calib_not_used]
```
Now we are prepared to visualize some PSF model residuals. We will evaluate the PSF at the location of some candidate stars which were not used to make the PSF. Then we will take the difference between the model and the actual star
```
residuals = calexp.maskedImage.getImage()
psf = calexp.getPsf()
f, ax = plt.subplots(ncols=3, nrows=3, sharex=True, sharey=True)
for i in (0,1,2):
# we can evaluate the PSF at different locations on the CCD
# lets evaluate it at the location of our reserve sources
psfImage = psf.computeImage(reserve_sources[i].getCentroid()).convertF()
# we should scale up the modle PSF so its flux matches that of the star we will compare it too
psfImage *= reserve_sources[i].getPsfInstFlux()/np.sum(psfImage.getArray())
ax[0,i].imshow(psfImage.getArray(), origin='lower') #plot the model PSF
ax[1,i].imshow(residuals[psfImage.getBBox()].getArray(), origin='lower') #plot the star
ax[2,i].imshow(psfImage.getArray() - residuals[psfImage.getBBox()].getArray(), origin='lower') #plot residuals
ax[0,0].set_ylabel('PSF model')
ax[1,0].set_ylabel('Image')
ax[2,0].set_ylabel('Residual')
plt.tight_layout()
```
### EXERCISE
Make a plot showing the difference between the model PSF evaluatad at two different arbitrary points
The PSF is spatially varying, so the PSF at two different points should be different. Remember you need to pass in lsstGeom.Point2D to the PSF model
```
# put your code here
```
## Introduction to SourceDetectionTask
We will now examine the source detection task. Typically, source detection is a subtask of some other higher level tasks. For example, detection is used multiple times in ProcessCcd.py, which you learned about in the previous lesson with Andrew Bradshaw. And it was used as a subtask to CharacterizeImage above. While that is the norm, it will be useful to start of working with the SourceDetectionTask on its own so we can get to know it a little better.
As we learned in previous lessons, `calexp` data contain a mask plane, where pixels are marked as being DETECTED, among other properties. This calexp we are working with has already been processed, and so it's mask plane already has marked pixels. Let's clear out the mask plane so that we can have a clean slate to work with.
```
calexp.mask.clearAllMaskPlanes()
```
Now that we have an image to work with, we will set up a minimal source detection task which we will run on our calexp. Detection requires that we pass a schema for a catalog to it to begin with. That schema will be used to populate a catalog with data from detection. Let's set up a minimal schema to start with.
```
schema = afwTable.SourceTable.makeMinimalSchema()
```
We also need to pass a `config` to our detection task when we instantiate it. Tasks have configs, which let you explicitly set certain parameters of the task to adjust it to your specific needs. You can usually find a task's config in the same file as the task. To see all the adjustable parameters, you can take a look [the detection source code on github](https://github.com/lsst/meas_algorithms/blob/master/python/lsst/meas/algorithms/detection.py). It will also allow you to see what types are allowed per each parameter, e.g. thresholdType expects a string, and specifically one of the following "stdev", "variance", "value", or "pixel_stdev", while thresholdValue expects a float larger than 0.0. While there are many to choose from, for now, we will set parameters that you are most likely to be interested in.
```
#create an instance of the source detection config
config = SourceDetectionConfig()
#drill down on the atributes of the config and set them
config.thresholdType = 'stdev'
config.thresholdValue = 5.0 #we want 5 sigma detection threshold
config.includeThresholdMultiplier = 1.0 # this scales the threshold value, so that the threshold is thresholdValue*thresholdMultiplyer
config.minPixels = 5 # sources less with fewer than 5 pixels will not be counted as detected sources
```
We are now ready to create an instance of the source detection class
```
srcDetection = SourceDetectionTask(config=config, schema=schema)
```
While we are here and have our schema defined, we will also define our deblending task and source measurement task as well. We will use them later in this notebook. In fact this all *must* be done before running any of these tasks. This is because each of the tasks need to add their own columns to the schema, which hapens durint the task init. A consequence of this is that the same schema object needs to be passed to all of them.
```
sourceDeblendTask = SourceDeblendTask(schema=schema)
measConfig = SingleFrameMeasurementConfig()
measConfig.plugins.names.add('ext_photometryKron_KronFlux')
sourceMeasurementTask = SingleFrameMeasurementTask(schema=schema, config=measConfig)
```
Tasks have a run method, a high level method that does everything the task is meant to do. We saw this previously when we ran `characterizeImage`. Lets finally run our calexp through detection. This will require a table to populate with results, as well as the calexp we want to run detection on.
```
tab = afwTable.SourceTable.make(schema)
detRes = srcDetection.run(tab, calexp)
```
We get several data products back from detection. `detRes` is a type `struct`. We can view the items by using print
```
print(detRes)
```
We will delve into footprints, and peaks later in this notebook. We have previously seen a bit of the source catalog earlier in this notebook, but we will take a closer look at it later when we discuss source measurement.
Our calexp itself has also been changed by running the detection task. It's mask plane has now been updated by the detection algorithm. Lets look at the mask plane bit values
```
calexp.getMask().getMaskPlaneDict()
```
The mask bit values are the exponent of the bitmask. For example, pixels only marked detected will have a value of 2^5 = 32, while pixels that are both on the edge and detected will be 2^5 + 2^4 = 48.
Finally we can visualize a small corner of our image. WHen you use AFW display and the matplotlib backend, you will get some colored overlays in the image. Detected objects will have a blue overlay, while interoplated pixels will have a green overlay, pixels close to the edge will be yellow, and bad pixels will have red overlay. For more information on how colors map to mask bits, you can view the documentation [here](https://pipelines.lsst.io/v/DM-11392/getting-started/display.html#interpreting-displayed-mask-colors)
```
afwDisplay.setDefaultBackend('matplotlib')
plt.figure()
afw_display = afwDisplay.Display()
afw_display.scale('asinh', 'zscale')
afw_display.mtv(calexp.maskedImage[:1000,:1000])
plt.gca().axis('off')
```
We also recieve a source catalog. Let's take a peek at it
```
sources = detRes.sources
sources
```
The resulting table doesn't say very much right now . This is because we only ran detection, and the table merely gives us information on detected sources--i.e., their ID, position, and 'parent'. The parent value for a source is 0 if it is its own parent, otherwise it will be the id of its parent source from which it was deblended. All of the sources have 0 as their parent right now, because we have not run deblending. Later on we will run deblending and source measurement as well, the resulting table will be more substantive. Before we go onto deblending and measurement, we will examine Footprints, another data product produced by detection.
## Altering the PSF for detection
The transpose of the PSF is formally the optimal filter for isolated point sources in the limit that the background is known perfectly and noise is dominated by the sky. But what if you are interested in detecting something else, and therefore want to tweak the detection kernel? We will take a moment to examine the effects of convolution with different PSFs to see the effect it has on convolution. This section is borrowed from Alex Drlica-Wagner's [Low Surface Brightness](https://nbviewer.jupyter.org/github/LSSTScienceCollaborations/StackClub/blob/rendered/SourceDetection/LowSurfaceBrightness.nbconvert.ipynb) Stack Club Notebook.
Something we can tinker with is the width of a gaussian PSF. We are going to delve into the guts of the [Source Detection Task](https://github.com/lsst/meas_algorithms/blob/master/python/lsst/meas/algorithms/detection.py) so we can gain some intuition for how convolution happens, and how the PSF factors into it. If we look at the [getPsf](https://github.com/lsst/meas_algorithms/blob/master/python/lsst/meas/algorithms/detection.py#L486) method, we can see that if we specify the parameter sigma, a Gaussian PSF with width sigma will be used to perform convolution in lieu of the PSF that is attached to the `calexp`. Going a little further down, we can see the
[convolveImage](https://github.com/lsst/meas_algorithms/blob/master/python/lsst/meas/algorithms/detection.py#L513) method is where the actual convolution happens. Let's do an exercise where we pass in increasingly wider PSFs to use in convolution, and visualize the effect they have on the smoothed images.
```
doSmooth = True # actually do the smoothing
# Loop over PSF convolution size in pixels
for i,sigma in enumerate([5, 7, 10]):
# Make a deep copy of the calexp
exp = calexp.clone()
# this will give us a gaussian PSF with width sigma
psf = srcDetection.getPsf(exp, sigma=sigma)
convolveResults = srcDetection.convolveImage(exp.maskedImage, psf, doSmooth=doSmooth)
middle = convolveResults.middle
sigma = convolveResults.sigma
plt.figure(i, figsize=(5,5))
afw_display = afwDisplay.Display(frame=i)
afw_display.scale('asinh', 'zscale')
afw_display.mtv(middle.image[:1000,:1000])
plt.gca().axis('off')
plt.title("Convolution Kernel: %s pix"%sigma)
```
We can see the detection kernel's width affects the degree to which different features 'pop out' in the smoothed images. In particular, broadening the filter has the effect of giving more contrast to larger more diffuse sources.
As Alex shows in his notebook, once you have settled on a sigma that suits your best detection needs, you can perform detection using your user defined sigma. Looking at the [run method](https://github.com/lsst/meas_algorithms/blob/master/python/lsst/meas/algorithms/detection.py#L298), we can see if you use the call pattern `srcDetection.run(tab, calexp, sigma=mySigma)` detection will carry on using a kernel with the specified width instead of the default width defined by the PSF. In Alex's notebook, we see that by broadening the detection filter, he is able to detect a low surface brightness galaxy that was not originally detected in the initial pass with a smaller PSF. So which is the right PSF to use? It really depends on the science you are doing and the goals you have.
## Footprints
Along with an updated mask plane to the calexp, and a minimal source catalog, detection also produces objects called `Footprints`. To start our discussion to Footprints we can draw an analogy between them and segmentation maps from Source Extractor, as they both identify above threshold pixels. As we first introduce footprints, we will concentrate on this similarity as it gives us a place to start understanding the location and gemoteric properties of footprints. We will soon see, however, that they are much more powerful, and play a role in deblending and source measurement.
Lets grab the positive footprints from our detection result, and introduce some basic methods footprints have that inform us of their location and geometry, and build some inutition for them.
```
fpset = detRes.fpSets.positive
#fps will be a list of the footprints
fps = detRes.fpSets.positive.getFootprints()
#This will return a rectangle in pixel coordinates of the CCD that the first footprint lies on
fps[0].getRegion()
#center of the footprint in detector coordinates
fps[0].getCentroid()
#the smallest box you could enscribe around the footprint
#not all the pixels in the bounding box strictly belong to the footprint,
#since the footprint isnt necessarily rectangular
fps[0].getBBox()
#area of the Footprint, not to be confused with the area of the bounding box
fps[0].getArea()
```
Right now the footprints do not know about the data contained in the pixels that they identify. We can fix this by defining `HeavyFootprints`. HeavyFootprints have all the qualities of Footprints, but additionally 'know' about pixel level data from their calexp's image, variance, and mask planes. Right now we can see the footprints are not 'heavy'
```
fps[0].isHeavy()
```
We can make the parent footprint set heavy by passing in the calexp, then confirm our HeavyFootprints are indeed heavy.
```
fpset.makeHeavy(calexp.maskedImage)
# we have to redefine fps
hfps = fpset.getFootprints()
hfps[0].isHeavy()
```
Now we can dump the pixel values for our heavy footprints. However, we will get back a flatted 1D array. To restructure this 1D array into the 2D array as it appears on the CCD, we will need to use the heavy footprints' `span`.
NB I went ahead and shearched for a footprint that would correspond do a nice image :) feel free to choose a different HeavyFootprint
```
hfps[21].getImageArray()
```
We can also grab the indicies on the chip that this footprint lives on by using the SpanSet's indices method. Once you have those, you can grab the pixel level data that corresponds to the footprint
```
yind, xind = hfps[21].spans.indices()
calexp.maskedImage.image.array[yind, xind]
```
We get information about which pixels in the footprint's bounding box actually belong to the footprint by using its `SpanSet`. The SpanSet is an arithmetic set of individual spans. We can print it out and get some informative informaation. Printed we will find several rows containing ones and zeros. If an element in the printed text is zero, this means the corresponding pixel in the SpanSet is inside the footprint's bounding box but does not belong to the footprint. If an element in the printed text is one, the pixel is both in the bounding box and associated with the footprint.
```
hfps[21].spans
```
We can use the footprint SpanSet and the pixel level information from the heavy footprints to rebuild the footprint image. In particular, you want to use the `unflatten` method from the spans to reshuffle the 1D image data into 2 dimensions as it appears on the CCD.
```
plt.imshow(hfps[21].spans.unflatten(hfps[21].getImageArray()),
norm=LogNorm(vmin=0.01, vmax=500))
```
## Erosion and Dilation on Footprints
We can grow or shrink footprints by using the `dilate()` and `erode()` methods. You may imagine using dilate if you want to ensure you are getting all the flux associated with the detected source, and you may imagine using erode if you want to chip away at the edges of a footprint, making it smaller. Using these methods requires that we provide a scale by which to grow or shrink the footprint. This scale is often related to the PSF scale, e.g., the RMS or σ of a gaussian PSF multiplied by a user defined factor. In the dilation process, a circular stencil-whose size is set by the scale the user passes in-is run along the edge of the spanset of the footprint, growing it. During erosion, the stencil is passed along the edge of the span set, this time shrinking it.
Let's demonstrate this below
```
# first get the length scale of the PSF
psf = calexp.getPsf()
sigma = psf.computeShape().getDeterminantRadius()
# print how many pixels belong to the footprint before we dilate it
print('before dilating the footprint has an area of {}'.format(hfps[0].getArea()))
# now we will grow the footprint by 2 x sigma. We need to pass an int
hfps[0].dilate(int(2*sigma))
# print how many pixels belong to the footprint ater dilation
print('after dilating the footprint has an area of {}'.format(hfps[0].getArea()))
# lets chip away at the footprint by using the erode method now
hfps[0].erode(int(2*sigma))
# print how many pixels belong to the footprint ater dilation
print('after eroding the footprint has an area of {}'.format(hfps[0].getArea()))
```
It is possible to erode such that your footprint is disconnected. You can use the `isContiguous()` method to check continuity of a footprint. If a footprint is no longer contiguous, you can use `split()` to split a multi-component Footprint into a vector of contiguous Footprints
### EXERCISE
Count the number of peaks in a footprint. Then Erode it and report the number of peaks after erosion.
You can get a table of the peaks by using the .getPeaks(deep=True) method on a a footprint
Be sure to choose a footprint that has a fair number of peaks so you can see the effect of erosion
```
# Your code here
```
### EXERCISE
Make two plots of a footprint by uising its span. First, plot an image of a footprint and overlay the peaks associated with it as red crosses. In the second plot, erode the footprint and overlay the remaining peaks still in the footprint again with red crosses. This will give you a better visualization of how erosion works and how it leads to culling peaks
```
# Your code here
```
## Deblending and Source Measurement
Like `CharacterizeImage`, and `SourceDetection`, deblending and measurement are also tasks that have their own run methods. Both of these tasks operate on Footprints, so our discussion in the previous section will be highly relevant.
A blend is considered to be a footprint that contains more than one peak. A footprint's peaks are assumed to belong to distinct astrophysical sources. The deblender's job is to allocate flux from the parent footprint to the different peaks, and assign them their own deblended child footprints. That is to say, in addition to identifying the location and real estate on the CCD that blended sources occupy, the deblender also allocates flux from the parent object to its deblended children. As a result, the flux in a deblended heavy footprint will not necessarily equal the flux at the same pixels in its image, since the pixels are shared between the parent and child footprints. This is a departure from something like `Source Extractor`, where you merely get segmentation maps.
After deblending is run, we run source measurement. During this process footprints are all exchanged with noise, and footprints are individually inserted, measured, and replaced with noise. We will visualize this process later in this notebook.
Now that detection has been run, we can pass our calexp and source catalog to the deblender, which will separate sources out into their own footprints. Subseqently, we will run source measurement so that each footprint will be measured by several algorithms. Lets run them now
```
#Source deblending
sourceDeblendTask.run(calexp, sources)
#Source measurement
sourceMeasurementTask.run(measCat=sources, exposure=calexp)
```
Notice that the new catalog contains new `deblended_` columns that the deblender imparted to it. The deblended_nChild column tells you how many child sources this source created. The parent column will list zero if this source has no parent (it was not deblended) or the object ID of its parent source if it was deblended.
There are also several detection algorithms that have been run, each imparting several columns to the catalog as well.
```
# make sources contiguous in memory by making a deep copy.
# this will ensure we can get an astropy version of it
sources = sources.copy(deep=True)
sources.asAstropy()
```
### EXERCISE
Make a plot showing how well aperture magnitudes agree with kron magnitudes
The x axis should be the aperture magnitude, and the y axis will be the difference between aperture magnitude and kron magnitude
You can search the schema for the column names you will need by doing sources.getSchema().extract('\*Kron\*'), or sources.getSchema().extract(\*CircularApertureFlux_12_0\*). This exercise is borrowed from Robert Lupton's [Kron.ipynb](https://github.com/RobertLuptonTheGood/notebooks/blob/master/Demos/Kron.ipynb)
```
# Your code here
```
we will do an exercise together that will help us gain intuition for how delbending works and how source measurement works. First lets grab an object that has many deblended children. Note, that the next few cells will involve some tricks with afwTables. We will explain step by step what is happening with comments, but going into detail on afwTables is outside the scope of our notebook. If you would like to know more about afwTables, you can checkout [the afw table guided tour](https://nbviewer.jupyter.org/github/LSSTScienceCollaborations/StackClub/blob/rendered/Basics/afw_table_guided_tour.nbconvert.ipynb)
```
# First get the key for the deblend_nChild field, and id. We need these to sort on them
nChild_key = sources.getSchema().find('deblend_nChild').key
idKey = sources.getIdKey()
# Sort the catalog on number of children
sources.sort(nChild_key)
# The catalog is sorted low to high, so grab an element towards the end
# Use the nChild key to get the number of children, and the id key to get the parent's ID
# The first few sources with many children are very bright stars that have been shreded
# I took the liberty of finding a nice source that has blended galaxies for you
num_children = sources[-14].get(nChild_key)
parent_id = sources[-14].getId()
# Footprints are stored in the source catalog, we can access them easily
parent_fp = sources[-14].getFootprint()
print('Parent source {} has {} deblended children'.format(parent_id, num_children))
```
The footprint of the parent object contains catalog of the peaks it contains. This catalog gives the peaks and their locations and peak pixel value in the smoothed image. During the detection process, the image in question is smoothed with a spatial filter. A nice demonstration of this is avaliable in the [LowSurfaceBrightness notebook](https://nbviewer.jupyter.org/github/LSSTScienceCollaborations/StackClub/blob/rendered/SourceDetection/LowSurfaceBrightness.nbconvert.ipynb). The peaks' centers and counts are in this smoothed image, and may differ slightly from their subsequent children footprints after deblending.
```
# deep copy to make it contiguous in memory
peaks = parent_fp.getPeaks().copy(deep=True)
peaks
```
Now we will set up a visualization to see our parent source, the locations of its peaks (as red crosses), and the centers of the deblended children footprints (as orange circles).
```
# Get the centroid of the parent
# So we can zoom in on it when we display it
parent_x, parent_y = sources[-14].getCentroid()
parent_x = int(parent_x)
parent_y = int(parent_y)
# Sort the catalog on ID
# It needs to be sorted on ID so we can use the getChildren method
sources.sort(idKey)
# child_cat is a smaller afwTable that only has the deblended children
child_cat = sources.getChildren(parent_id)
```
Now we will use afwDisplay to zoom in on the parent source's location, and use the child_cat to overplot the peaks in orange circles and centers of the deblended children in red croses. You will notice they are not completely coincident, and that is because the peak locations are in the smoothed image, as we mentioned above.
You will also notice that there are some circle's and crosses where-by eye-it's not clear that there is a astropysical source. Although we can't say for sure since we do not know the ground truth, there's a strong chance these are just noise peaks that were found in the extended tails of the diffuse galaxies' tails in the frame. None the less, these are regarded as true astrophysical sources in the stack. As a result, the deblender will consider them and appropriate flux to them. Subsequently the Source Measurement Task will perform measurements on these deblended children as well.
```
# credit to ADW for this afwDisplay snippet
plt.figure()
afw_display = afwDisplay.Display()
afw_display.setMaskTransparency(100)
afw_display.scale('asinh', 'zscale')
afw_display.mtv(calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100])
# We use display buffering to avoid re-drawing the image after each source is plotted
with afw_display.Buffering():
# loop over child footprints, placing a red plus on their centers
for s in child_cat:
afw_display.dot('+', s.getX(), s.getY(), ctype=afwDisplay.RED)
# loop over peaks, placing an orange circle on their centers
for p in parent_fp.getPeaks():
afw_display.dot('o', p.getIx(), p.getIy(), ctype=afwDisplay.ORANGE, size=4)
```
We will concentrate on this example to help us visualize how measurement works on deblended sources. Drawing from the HSC Pipeline paper, the high level steps are outlined below
> 1. We replace all Footprints in the image with random Gaussian noise with the same variance as the original noise in those pixels.
2. We insert the deblended pixels for a particular source to be measured back into the image (replacing the noise pixels).
3. We run all measurement plug-ins on the current source.
4. We re-replace the Footprint of the current source with noise.
5. Repeat this process for the next Footprint
To exchange footprints with noise, and insert them back into the image, we will use an object called the `NoiseReplacer`. To set it up we need to provide it with a dictionary that contains source IDs as keys, and a tuple pair of parent ID and footprint.
```
fp_dict = {measRecord.getId(): (measRecord.getParent(), measRecord.getFootprint())
for measRecord in sources}
nr_config = NoiseReplacerConfig()
noiseReplacer = NoiseReplacer(nr_config, calexp, fp_dict)
plt.figure()
afw_display = afwDisplay.Display()
afw_display.setMaskTransparency(100)
afw_display.scale('asinh', 'zscale')
afw_display.mtv(calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100])
```
As advertised above, the image now only contains noise, as all the detected footprints have been replaced. Now we will insert the parent source back into the frame. A series of measurements would then be applied to the source, while all other detected sources are still replaced with noise.
```
noiseReplacer.insertSource(parent_id)
plt.figure()
afw_display = afwDisplay.Display()
afw_display.setMaskTransparency(100)
afw_display.scale('asinh', 'zscale')
afw_display.mtv(calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100])
# we will need the ids of the deblended children so we can insert them
# back into the image one at a time
child_ids = child_cat['id'].tolist()
fig = plt.figure()
ims = []
src_ids = [parent_id] + child_ids #tack on the parent ID to the list
for i in src_ids:
# replace the ith source using noiseReplacer
noiseReplacer.insertSource(i)
# grab a postage stamp centered on the parent object and plot it
data = calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100].getImage().array
im = plt.imshow(data, origin='lower', animated=True, vmin=-100, vmax=100, visible=False)
# keep a running list of our plotted images so we can make a movie out of them later
ims.append([im])
noiseReplacer.removeSource(i)
# the plt.close catches a spurious extra image
plt.close()
# organize our plotted images into an animation
HTML(animation.ArtistAnimation(fig, ims, interval=1000, blit=True,
repeat_delay=1000).to_jshtml())
# takes a minute to get going,
```
Let's make a similar visualization, now showing mug shots of all of the deblended sources.
### EXERCISE
Familiarize yourself with the noise replacer.
Finish the missing code in following for loop so that we make mugshots of the parent and 9 deblended children objects. You can iterate over the `src_ids` list
```
f, ax = plt.subplots(nrows=3, ncols=3, sharey=True, sharex=True, figsize=(12,8))
for i, a in zip(range(0, 9), ax.flatten()):
noiseReplacer.insertSource() # fill in the argument here
data = calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100].getImage().array
a.imshow(data, origin='lower', vmin=-100, vmax=100)
noiseReplacer.removeSource() # fill in this argument too
plt.tight_layout()
```
We can see the parent source in the top left pannel, and two of its deblended children in the top middle and top right. The remaining plots show the remaining deblended children. Each of these scenarios would be passed to the measurement algorithms to be measured individually, as distinct astrophysical sources.
Looking at the bottom six pannels, we can see the deblender allocated them nearly no flux and gave almost all the flux to the two deblended children in the top middle and top right plots. You may notice that some of the deblended children look 'flat' in that they are surrounded by noiseless signal. This is because nearly all the pixels in those deblended children were given zero or near zero flux, giving a uniform color to the deblended children footprints.
Now that we are ready to move on, lets tidy up and end the `noiseReplacer`. This will restore the image to its state before we started removing and inserting sources
```
noiseReplacer.end()
# replot the coadd at the location of the deblended source to prove
# everything is back to the way it was before we were inserting and removing footprints
plt.figure()
afw_display = afwDisplay.Display()
afw_display.setMaskTransparency(100)
afw_display.scale('asinh', 'zscale')
afw_display.mtv(calexp.maskedImage[parent_x-100:parent_x+100, parent_y-100:parent_y+100])
```
## Detection, deblending, measurement on Deep Coadds
Up until now we have been using `calexp`s and discussing detection, deblending, and measurement that happens at the individual CCD level. However, as we discussed above, most of these operaons are to characterize or calibrate properties of these chips, like setting the PSF, or setting the WCS for example. Other high level science will come from detection, deblending, and measurement on coadds.
With that in mind, we will take a look at how these play out on the Coadd Level. Borrowing from the HSC pipelines paper
1. Run detection on coadds each in different bands
2. Merge footprints and peak positions across bands using a priority list irzyg. This is roughly in order of decreasing S/N
3. Deal with peaks in merged footprints. If peaks in a newly considered band are more than 1 arcsecond away from any other peaks detected in former bands, this peak is added and considered a distinct source. If the peak is .3 arcseconds away from a nearest source, it is considered marked as appearing in both bands. If the peak is between .3 and 1 arcsecond away from a nearest source, it is discarded as it is not clear if it belongs to a peak in the new band belongs to its nearest neighboor in the previous bands.
4. Deblend the merged catalog independently in each band and run measurement on individual frames
5. Define a reference band for each object
6. Perform forced photometry in all bands. Hold the position and shape properties as measured in the reference band constant in all bands while running measurement algorithms.
For now let's begin by using the butler to grab a coadd in the I band, and demonstrate how detection is performed. We will run a high level task called `DetectCoaddSourcesTask`. Inside the task is a detection subtask, however it differs from the detection task we have encountered before. This new detection task is called `DynamicDetection`. In the previous detection task, the threshold was set across the whole image. However, convolution with the transpose of the PSF creates a smoothed image with correlated noise. Because of this the previous detection task will underperform and fail to detect fainter-but above threshold-sources. As a fix, the `DynamicDetection` task adjusts the threshold dynamically across the smoothed image.
```
from lsst.pipe.tasks.multiBand import DetectCoaddSourcesTask, DetectCoaddSourcesConfig
deepCoadd = butler.get("deepCoadd_calexp", dataId={'filter':"HSC-I", 'field':field, 'tract':15830, 'patch':'0,3'})
schema = afwTable.SourceTable.makeMinimalSchema()
dcconfig = DetectCoaddSourcesConfig()
# subaru configs and in detectcoaddsources defaults
# recall we drill down in the config to the detection subtask
# and its parameters to set them
dcconfig.detection.isotropicGrow = True
dcconfig.detection.doTempWideBackground = True
dcconfig.detection.tempWideBackground.binSize = 128
dcconfig.detection.tempWideBackground.useApprox = False
dcconfig.detection.reEstimateBackground = True
dcconfig.detection.background.binSize = 128
dcconfig.detection.background.useApprox = False
dcconfig.detection.thresholdType = "pixel_stdev"
sourceCoaddDetectionTask = DetectCoaddSourcesTask(schema=schema, config=dcconfig)
tab = afwTable.SourceTable.make(schema)
# detection needs this to be happy
# you can pass none for exposure id
idFactory = afwTable.IdFactory.makeSimple()
result = sourceCoaddDetectionTask.run(deepCoadd, idFactory, None)
result
plt.figure()
afw_display = afwDisplay.Display()
afw_display.scale('asinh', 'zscale')
afw_display.mtv(deepCoadd.maskedImage[2000:2500, 12000:12500])
plt.gca().axis('off')
```
## Forced Photometry: Combining Across Bands
Let's do one last example together, showing how you might exercise the forced photometry. As listed above in the steps that outline multi-band processing, the forced photometry catalogs unify objects across bands. To take advantage of this, we will make a color-color diagram. It's important to note that this would not be possible with the other detection products, as they have not had their peaks merged. Because Forced photometry happens after peaks have been merged, the jth row in the g band table is the same source as the jth row in the r band table, and so on. This is not the case in the other catalogs.
We need to first grab the forced photometry catalogs using the butler. Remember, these catalogs report instrumental fluxes, not magnitudes
```
objects = []
for filter in ("HSC-I", "HSC-R", "HSC-G", "HSC-Z"):
objects.append(butler.get("deepCoadd_forced_src", dataId={'filter':filter, 'field':field, 'tract':15830, 'patch':'0,3'}))
iSources, rSources, gSources, zSources = objects
```
To get magnitudes we will need photoCalibs from each filter. Fetch those with the butler too
```
calibs = []
for filter in ("HSC-I", "HSC-R", "HSC-G", "HSC-Z"):
calibs.append(butler.get("deepCoadd_calexp_photoCalib", dataId={'filter':filter, 'field':field, 'tract':15830, 'patch':'0,3'}))
iCalib, rCalib, gCalib, zCalib = calibs
```
Set up some quality control flags so we wont be dealing with too many junky sources. Partition the stars and galaxies into different samples
```
noChildren = iSources['deblend_nChild'] == 0
isGoodFlux = ~iSources['modelfit_CModel_flag']
isStellar = iSources['base_ClassificationExtendedness_value'] < 1.
snr = iSources['modelfit_CModel_instFlux']/iSources['modelfit_CModel_instFluxErr'] > 10
gal_flag = noChildren & isGoodFlux & ~isStellar & snr
star_flag = noChildren & isGoodFlux & isStellar & snr
```
Calculate magnitudes and magnitude errors
```
iMags = iCalib.instFluxToMagnitude(iSources, 'modelfit_CModel')
rMags = rCalib.instFluxToMagnitude(rSources, 'modelfit_CModel')
gMags = gCalib.instFluxToMagnitude(gSources, 'modelfit_CModel')
zMags = zCalib.instFluxToMagnitude(zSources, 'modelfit_CModel')
```
Make a color-color diagram. The stellar locus in this diagram will appear as red points. These are mostly Milky Way zero age main sequence stars, which overlap with galaxies that have simmilar stellar populations
```
# plot r-i vs g-r for galaxies
plt.scatter(rMags[:,0][gal_flag] - iMags[:,0][gal_flag],
gMags[:,0][gal_flag] - rMags[:,0][gal_flag],
s=10, color='k', label='galaxies')
# plot r-i vs g-r fo stars on the same axis
plt.scatter(rMags[star_flag]-iMags[star_flag], gMags[star_flag]-rMags[star_flag],
s=10, color='red', label='stars')
# zoom in on the interesting features
plt.xlim([-.5,2])
plt.ylim([-.5,2])
plt.xlabel('r - i')
plt.ylabel('g - r')
plt.legend()
```
### Exercise
Make a color *magnitude* diagram of stars only, and use psf fluxes to calculate magnitudes instead of CModel. This is borrowed from the [lsst science pipelines demo](https://pipelines.lsst.io/getting-started/multiband-analysis.html)
## Summary
Now you know how to detect, measure, and deblend images. You also know how to play around with detection configurations and measurement configurations to suit specific needs you may have if you are doing your own re-analysis. You should also be familiar with footprints, and their interplay between detection, deblending, and measurement. Finally, you learned how to navigate schemas and make plots using catalogs produced my the Science Pipelines.
| github_jupyter |
# Setuptools
En Python todo el tema de empaquetar puede ser un poco lioso, ya que encontramos varios módulos desintados a ello. Nosotros vamos a centrarnos en **setuptools**, ya que es la forma más utilizada, nos proporciona todo lo necesario para distribuir nuestros propios módulos e incluso nos permite publicar paquetes en el respositorio público PyPI (Python Package Index) de forma directa desde la propia terminal.
Si lo recordáis, en la lección de módulos ya os enseñé como crear un distribuible con setuptools, a lo largo de esta lección vamos a repasar y aprender varios conceptos nuevos.
## Paquete básico
Antes de comenzar es importante repasar la estructura de un paquete en Python,ya que para distribuir nuestro código es indispensable estructurarlo dentro de un paquete:
```
| setup.py # Fichero que contiene toda la información de instalación
+ prueba/ # Directorio del paquete al mismo nivel que setup.py
| __init__.py # Fichero que indica que el directorio es un paquete
| modulo.py # Módulo o script que contiene definiciones
```
Por lo tanto vamos a empaquetar el paquete de nombre **prueba**, que contiene código en el fichero *modulo.py*.
Vamos a aprender un poco más sobre el fichero de instalación.
## setup.py
El fichero de configuración incluye toda la información necesaria para realizar la instalación de nuestro paquete. Algunos campos incluyen sólo metadatos como el nombre, la versión, la descripción o el autor. Pero otros sirven para extender la instalación.
Como sería un caos que cada desarrollador pusiera los campos que quisiera, hay una serie de parámetros comunes y avanzados, pero como son muchos lo más común es utilizar una plantilla base como la siguiente que pasa la configuración a la función **setup**:
```python
from setuptools import setup
setup(name="Prueba", # Nombre
version="0.1", # Versión de desarrollo
description="Paquete de prueba", # Descripción del funcionamiento
author="Hector Costa", # Nombre del autor
author_email='me@hcosta.info', # Email del autor
license="GPL", # Licencia: MIT, GPL, GPL 2.0...
url="http://ejemplo.com", # Página oficial (si la hay)
packages=['prueba'],
)
```
¿Hasta aquí fácil no? Son simples metadatos para definir el paquete, con la excepción de **packages**, en el que tenemos que indicar todos los paquetes que formarán parte del paquete distribuido en forma de lista.
Aunque en este caso únicamente tendríamos al paquete **prueba**, imaginaros que tenemos docenas de subpaquetes y tubiéramos que añadirlos uno a uno... Pues para estos casos podemos importar una función que se encargará de buscar automáticamente los subpaquetes, se trata de **find_packages** y la podemos encontrar dentro de **setuptools**:
```python
from setuptools import setup, find_packages
setup(...
packages=find_packages()
)
```
## Dependencias
Ahora imaginaros que en vuestro paquete algún código utiliza funciones de un módulo externo o paquete que hay que instalar manualmente. Esto se conoce como dependencias del paquete, y por suerte podemos indicar a un parámetro que descargue todos los paquetes en la versión que nosotros indiquemos, se trata de **install_requires**.
Por ejemplo imaginad que dentro de nuestro paquete necesitamos utilizar el módulo **Pillow** para manejar imágenes. Por regla general podemos instalarlo desde la terminal con el comando:
```
pip install pillow
```
Pero si queremos que el paquete lo instale automáticamente sólo tenemos que indicarlo de esta forma:
```python
setup(...,
install_requires=["pillow"],
)
```
Y así iríamos poniendo todas las dependencias en la lista.
Lo bueno que tiene es que podemos indicar la versión exacta que queremos instalar, por ejemplo. Si mi programa utilizase la versión 1.1.0 de Pillow tendría que poner:
```python
setup(...,
install_requires=["pillow==1.1.0"],
)
```
En cambio si fuera compatible con cualquier versión a partir de la 1.1.5 podría poner:
```python
setup(...,
install_requires=["pillow>=1.1.5"],
)
```
Si no indicamos una versión, se instalará automáticamente la más actual.
### Utilizando un fichero de dependencias
De forma similar a antes, quizá llega el momento donde tenemos muchísimas dependencias y es un engorro tener que cambiar directamente el fichero **setup.py**. Para solucionarlo podemos utilizar una técnica que se basa en crear un fichero de texto y escribir las dependencias, una por línea.
Luego podemos abrir el fichero y añadir las dependencias automáticamente en forma de lista. Generalmente a este fichero se le llama **requirements.txt** y debe estar en el mismo directorio que **setup.py**:
#### requirements.txt
```
pillow==1.1.0
django>=1.10.0,<=1.10.3
pygame
```
Luego en las dependencias indicaríamos lo siguiente:
```python
setup(...,
install_requires=[i.strip() for i in open("requirements.txt").readlines()],
)
```
## Suite Test
Otra cosa interesante que podemos hacer es adjuntar una suite de tests unitarios para nuestro paquete, ya sabéis, los que aprendimos en la unidad anterior.
Para incluirlos tendremos indicar un parámetro en el instalador llamado **test_suite**, al que le pasaremos el nombre del directorio que los contiene, por lo general llamado **tests**:
```
| setup.py
| requeriments.txt
+ prueba/
| __init__.py
| modulo.py
+ tests/
| test_pillow.py
| test_django.py
| test_pygame.py
```
En el **setup.py**:
```python
setup(...,
test_suite="tests"
)
```
Luego para ejecutarlos podemos utilizar el comando:
```python
python setup.py test
```
# PyPI y PIP
Por último hablemos un poco más del **Python Package Index**.
Como ya sabéis se trata de un repositorio público con miles y miles de paquetes creados por la enorme comunidad de Python. De hecho yo mismo creé hace años un pequeño módulo para el framework django, os dejo el enlace por si os pica la curiosidad: https://pypi.python.org/pypi/django-easyregistration
Sea como sea, la forma de instalar cómodamente los paquetes de PyPI es con la herramienta PIP (un acrónimo recursivo de Pip Installs Packages), utilizando el comando **pip install nombre_paquete**.
Además podemos listar los paquetes instalados con **pip list**, borrar alguno con **pip uninstall nombre_paquete** o incluso instalar todas las dependencias de un fichero **requisites.txt** utilizando **pip install requisites.txt**.
Si queréis saber más sobre pip, simplemente escribid **pip** en la terminal.
## Clasificadores
Por lo tanto tenemos un repositorio inmenso, así que ¿cómo podemos añadir información para categorizar nuestro paquete en PyPI? Pues utilizando un parámetro llamado **classifiers** de la siguiente forma:
```python
setup(...,
classifiers=[
"Development Status :: 3 - Alpha",
"Topic :: Utilities",
"License :: OSI Approved :: GNU General Public License (GPL)",
],
)
```
Hay un montón de clasificadores, desde el estado del proyecto, el tema, las licencias, etc. Una lista completa de los clasificadores disponibles podemos encontrarla en la propia web de PyPI:
https://pypi.python.org/pypi?%3Aaction=list_classifiers
## Probando el paquete
Una vez tenemos toda la información configurada, podemos probar nuestro paquete fácilmente realizando una instalación en modo desarrollo. Para ello utilizaríamos el siguiente comando:
```
python setup.py develop
```
Este modo es muy práctico, ya que nos permite utilizar nuestro módulo en cualquier lugar y hacer modificacione sin necesidad de reinstalarlo constamente. Eso es posible porque se utiliza desde el propio directorio.
Una vez hayamos hecho las probaturas y estemos satisfechos, podemos desinstalar el paquete de desarrollo:
```
python setup.py develop --uninstall
```
Para instalar el paquete definitivo utilizaríamos:
```
python setup.py install
```
Pero tenemos que tener en cuenta que una vez hecho esto, el paquete se instala en una copia interna y ya no podremos modificarlo sin antes desinstalarlo, algo que tendremos que hacer con PIP, buscando el nombre del paquete con **pip list** y haciendo un **pip uninstall nombre_paquete**.
# Distribuyendo el paquete
Ya tenemos el paquete, hemos creado el instalador, lo hemos probado y estamos preparados para distribuirlo. Hay dos formas:
* **Localmente**: Generando un fichero comprimido que podemos compartir con nuestros conocidos.
* **Públicamente**: En el repositorio PyPI para que todo el mundo pueda utilizarlo.
Evidentemente si distribuimos localmente no tenemos que tener mucho cuidado, y además podemos hacer pruebas. Pero si decidimos hacerlo públicamente tendremos que intentar que el paquete tenga un mínimo de calidad.
### Localmente
Distribuir el paquete localmente es muy fáci. Simplemente tenemos que utilizar el comando:
```
python setup.py sdist
```
Esto generará un directorio **dist/** en la carpeta del paquete. Dentro encontraremos un fichero zip o tar.gz dependiendo de nuestro sistema operativo.
Este fichero ya podremos compartirlo con quien queramos, y para instalarlo sólo tendremos que utilizar la herramienta **pip**:
```
pip install nombre_del_fichero.zip # La extensión depende del sistema operativo
```
Luego para desinstalarlo de la misma forma pero utilizando el nombre del paquete:
```
pip uninstall nombre_paquete
```
### Públicamente
Aunque no voy a hacer la demostración porque ahora mismo no dispongo de un paquete para publicar en el repositorio de PyPI, sí que os voy a enseñar los pasos a seguir para hacerlo. Lo bueno de registrar un paquete en PyPI es que podemos instalarlo desde cualquier lugar a través de internet utilizando la herramienta PIP.
Dicho ésto, si algún día creáis un paquete de calidad y queréis compartirlo con la comunidad, lo primero es registrar una cuenta en PyPI: https://pypi.python.org/pypi?%3Aaction=register_form
A continuación desde el directorio de nuestro paquete tenemos que ejecutar el comando:
```
python setup.py register
```
Así iniciaremos una petición para registrar nuestro paquete en el repositorio. Luego tendremos que seguir los pasos e identificarnos cuando lo pida con nuestro usuario y contraseña (que hemos creado antes).
Una vez hecho esto ya hemos creado nuestro paquete, pero todavía no hemos publicado una versión, así que vamos a hacerlo utilizando el comando:
```
python setup.py sdist upload
```
¡Y ya está! Ahora podremos instalar nuestro paquete desde en cualquier lugar con PIP:
```
pip install nombre_paquete
```
| github_jupyter |
# Hello
```
%matplotlib inline
import torchvision
import torchvision.datasets as dset
import torchvision.transforms as transforms
from torch.utils.data import DataLoader, Dataset, TensorDataset
import matplotlib.pyplot as plt
import torchvision.utils
import numpy as np
import random
from PIL import Image
import torch
from torch.autograd import Variable
import PIL.ImageOps
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
import torch.optim as optim
import time
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
```
## Load Data
```
#load .npz-file from folder
data_folder = "data/"
loader = np.load(data_folder+"gray_image_data_xs.npz")
train_data = loader["train_features"]
train_labels = loader["train_labels"].reshape((-1,1))
val_data = loader["val_features"]
val_labels = loader["val_labels"].reshape((-1,1))
labels_length = train_labels.shape[0]
all_labels = np.append(train_labels, val_labels)
# numeric encoding
en = LabelEncoder()
all_labels = en.fit_transform(all_labels)
train_labels = all_labels[:labels_length]
val_labels = all_labels[labels_length:]
num_classes = max(train_labels)+1
train_data = np.expand_dims(train_data, axis=1)
val_data = np.expand_dims(val_data, axis=1)
train_data.shape
train_labels.shape
```
## Load data for pytorch
```
"""
# doesnt work yet:
class MyDataset(Dataset):
def __init__(self, data, labels):
#super(MyDataset, self).__init__()
self.data = data
self.labels = labels
def __len__(self):
return len(self.data)
def __getitem__(self, index):
return self.data[index], self.labels[index]
train = MyDataset(train_data, train_labels)
train_loader = DataLoader(train, batch_size=64, shuffle=True)
val = MyDataset(val_data, train_labels)
val_loader = DataLoader(val, batch_size=64, shuffle=True)
"""
train_data = torch.FloatTensor(train_data)
train_labels = torch.LongTensor(train_labels)
train = TensorDataset(train_data, train_labels)
train_loader = DataLoader(train, batch_size=64, shuffle=True)
val_data = torch.FloatTensor(val_data)
val_labels = torch.LongTensor(val_labels)
val = TensorDataset(val_data, val_labels)
val_loader = DataLoader(val, batch_size=64, shuffle=True)
```
## Build Model
```
# the Classifier stolen from Pytorch Tutorial
class CNN(nn.Module):
def __init__(self, num_classes, dropout=0.5):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(1, 16, 5)
self.conv2 = nn.Conv2d(16, 32, 5)
self.fc1 = nn.Linear(32*22*22 , 64)
self.fc2 = nn.Linear(64, num_classes)
self.dropout = nn.Dropout(p=dropout)
self.optimizer = optim.Adam(self.parameters(), lr=learning_rate)
self.criterion = nn.CrossEntropyLoss()
def forward(self, x):
x = F.max_pool2d(F.relu(self.conv1(x)), 2)
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = F.relu(self.dropout(self.fc1(x)))
x = self.fc2(x)
return x
def num_flat_features(self, x):
size = x.size()[1:] # all dimensions except the batch dimension
num_features = 1
for s in size:
num_features *= s
return num_features
def make_plot(self):
train_acc_dropoutreg = self.hist['train_accuracy'][-1]
val_acc_dropoutreg = self.hist['val_accuracy'][-1]
print("dr:",f"{train_acc_dropoutreg:.3f}",f"{val_acc_dropoutreg:.3f}")
print(epochs, batch_size, learning_rate)
print("maxacc,minloss at epoch: ", np.argmax(self.hist['val_accuracy']),np.argmin(self.hist['val_loss']),\
"acc there: ", self.hist['val_accuracy'][np.argmax(self.hist['val_accuracy'])],\
self.hist['val_accuracy'][np.argmin(self.hist['val_loss'])])
plt.figure(figsize=(10,5))
plt.plot(self.hist['train_loss'][0::],
label="Training (dropout regularization)")
plt.plot(self.hist['val_loss'][0::],
label="Validation (dropout regularization)", linestyle="--")
plt.xlabel("Epoch", fontsize=20)
plt.ylabel("Loss", fontsize=20)
plt.legend()
plt.show()
def train(self, train_data, train_labels, val_data, val_labels, epochs=20, dropout=0.0, batch_size=512):
train_losses, train_accs = [], []
val_losses, val_accs = [], []
tr_loss, tr_acc = self.runForward(train_loader)
val_loss, val_acc = self.runForward(val_loader)
train_losses.append(tr_loss)
train_accs.append(tr_acc)
val_losses.append(val_loss)
val_accs.append(val_acc)
for epoch in range(epochs): # loop over the dataset multiple times
self.running_loss = 0.0
self.run(train_loader)
tr_loss, tr_acc = self.run(train_loader, backward=False)
val_loss, val_acc = self.run(val_loader, backward=False)
train_losses.append(tr_loss)
train_accs.append(tr_acc)
val_losses.append(val_loss)
val_accs.append(val_acc)
self.hist={'train_loss': np.array(train_losses),
'train_accuracy': np.array(train_accs),
'val_loss': np.array(val_losses),
'val_accuracy': np.array(val_accs)}
if (epoch + 1) % 2 == 0:
print(f"Epoch {epoch+1}/{epochs}")
self.make_plot()
# hyperparams
epochs = 10
batch_size = 64
learning_rate = 0.001
dropout = 0.5
width, height = 100,100
# train model
t = time.time()
cnn = CNN(num_classes = num_classes)
#cnn.train(train_data, train_labels, val_data, val_labels, epochs=epochs, dropout=dropout, batch_size=batch_size)
optimizer = optim.Adam(cnn.parameters(), lr=learning_rate)
criterion = nn.CrossEntropyLoss()
for epoch in range(epochs): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(train_loader):
inputs, labels = data
outputs = cnn(inputs)
loss = criterion(outputs, labels)
ti = time.time()
loss.backward()
optimizer.step()
optimizer.zero_grad()
print("minibatch step",time.time() - ti)
running_loss += loss.item()
###end batch
if (epoch + 1) % 2 == 0:
print(f"Epoch {epoch+1}/{epochs}")
#cnn.make_plot()
print('Finished Training', time.time()-t)
# evaluate model
plt.figure(figsize=(10,5))
plt.plot(cnn.hist['train_loss'][5::],
label="Training (dropout regularization)")
plt.plot(cnn.hist['val_loss'][5::],
label="Validation (dropout regularization)", linestyle="--")
plt.xlabel("Epoch", fontsize=20)
plt.ylabel("Loss", fontsize=20)
plt.legend()
plt.show()
# Accuracy Plot
plt.figure(figsize=(10,5))
plt.plot(cnn.hist['train_accuracy'],
label="Training (dropout regularization)", color="purple")
plt.plot(cnn.hist['val_accuracy'],
label="Validation (dropout regularization)", color="purple", linestyle="--")
plt.xlabel("Epoch", fontsize=20)
plt.ylabel("Accuracy", fontsize=20)
plt.legend()
plt.show()
```
| github_jupyter |
# ACA-Py & ACC-Py Prover Template
## Copy this template into the root folder of your notebook workspace to get started.
## NOTE: A prover can only respond to proof requests if they have credentials previously stored in their wallet (unless using self-attested attributes). The prover_template works well with the templates in the issue-credential folder.
### Imports
```
from aries_cloudcontroller import AriesAgentController
import os
from termcolor import colored
```
### Initialise the Agent Controller
```
api_key = os.getenv("ACAPY_ADMIN_API_KEY")
admin_url = os.getenv("ADMIN_URL")
print(f"Initialising a controller with admin api at {admin_url} and an api key of {api_key}")
agent_controller = AriesAgentController(admin_url,api_key)
```
### Start a Webhook Server
```
webhook_port = int(os.getenv("WEBHOOK_PORT"))
webhook_host = "0.0.0.0"
await agent_controller.init_webhook_server(webhook_host, webhook_port)
print(f"Listening for webhooks from agent at http://{webhook_host}:{webhook_port}")
```
## Register Agent Event Listeners
You can see some examples within the webhook_listeners recipe. Copy any relevant cells across and customise as needed.
```
listeners = []
# Receive connection messages
def connections_handler(payload):
state = payload['state']
connection_id = payload["connection_id"]
their_role = payload["their_role"]
routing_state = payload["routing_state"]
print("----------------------------------------------------------")
print("Connection Webhook Event Received")
print("Connection ID : ", connection_id)
print("State : ", state)
print("Routing State : ", routing_state)
print("Their Role : ", their_role)
print("----------------------------------------------------------")
if state == "invitation":
# Your business logic
print("invitation")
elif state == "request":
# Your business logic
print("request")
elif state == "response":
# Your business logic
print("response")
elif state == "active":
# Your business logic
print(colored("Connection ID: {0} is now active.".format(connection_id), "green", attrs=["bold"]))
connection_listener = {
"handler": connections_handler,
"topic": "connections"
}
listeners.append(connection_listener)
def prover_proof_handler(payload):
role = payload["role"]
connection_id = payload["connection_id"]
pres_ex_id = payload["presentation_exchange_id"]
state = payload["state"]
print("\n---------------------------------------------------------------------\n")
print("Handle present-proof")
print("Connection ID : ", connection_id)
print("Presentation Exchange ID : ", pres_ex_id)
print("Protocol State : ", state)
print("Agent Role : ", role)
print("Initiator : ", payload["initiator"])
print("\n---------------------------------------------------------------------\n")
if state == "request_received":
presentation_request = payload["presentation_request"]
print("Recieved Presentation Request\n")
print("\nRequested Attributes - Note the restrictions. These limit the credentials we could respond with\n")
print(presentation_request["requested_attributes"])
elif state == "presentation_sent":
print("Presentation sent\n")
elif state == "presentation_acked":
print("Presentation has been acknowledged by the Issuer")
prover_listener = {
"topic": "present_proof",
"handler": prover_proof_handler
}
listeners.append(prover_listener)
agent_controller.register_listeners(listeners)
```
## Accept Invitation
Copy an invitation object from another agent playing the role inviter (see the inviter_template recipe)
```
invitation = {<some agent invitation>}
auto_accept="false"
alias=None
invite_response = await agent_controller.connections.receive_invitation(invitation, alias, auto_accept)
connection_id = invite_response["connection_id"]
# Label for the connection
my_label = None
# Endpoint you expect to recieve messages at
my_endpoint = None
accept_response = await agent_controller.connections.accept_invitation(connection_id, my_label, my_endpoint)
```
## Send Trust Ping
Once connection moves to response state one agent, either inviter or invitee needs to send a trust ping.
Note: you may not need to run this cell. It depends one of the agents has the ACAPY_AUTO_PING_CONNECTION=true flag set.
```
comment = "Some Optional Comment"
message = await agent_controller.messaging.trust_ping(connection_id, comment)
```
## Optional: Send Proposal
Propose a presentation to a verifier
```
# TODO: Example proposal object below
# proposal_object = {
# "auto_present": true,
# "comment": "string",
# "connection_id": "3fa85f64-5717-4562-b3fc-2c963f66afa6",
# "presentation_proposal": {
# "@type": "did:sov:BzCbsNYhMrjHiqZDTUASHg;spec/present-proof/1.0/presentation-preview",
# "attributes": [
# {
# "cred_def_id": "WgWxqztrNooG92RXvxSTWv:3:CL:20:tag",
# "mime-type": "image/jpeg",
# "name": "favourite_drink",
# "referent": "0",
# "value": "martini"
# }
# ],
# "predicates": [
# {
# "cred_def_id": "WgWxqztrNooG92RXvxSTWv:3:CL:20:tag",
# "name": "high_score",
# "predicate": ">=",
# "threshold": 0
# }
# ]
# },
# "trace": false
# }
# proposal_response = await agent_controller.proofs.send_proposal(proposal_object)
```
## Fetch Presentation Records
Before you can present a presentation, you must identify the presentation record which you wish to respond to with a presentation. This could also be done through the present_proof listeners which have access to a presentation record in the payload.
```
# Optional Query parameters
verifier_connection_id = connection_id
thread_id=None
state = "request_received"
role = "prover"
proof_records_response = await agent_controller.proofs.get_records(verifier_connection_id, thread_id, state, role)
# We fetch the first record from the response. You may want to customise this further
presentation_record = proof_records_response["results"][0]
presentation_exchange_id = presentation_record["presentation_exchange_id"]
```
## Search For Available Credentials to Construct Presentation From
The presentation record can be used to query your agents wallet and return all credentials that could be used to construct valid presentation
```
# select credentials to provide for the proof
credentials = await agent_controller.proofs.get_presentation_credentials(presentation_exchange_id)
print("Credentials stored that could be used to satisfy the request. In some situations you applications may have a choice which credential to reveal\n")
attribute_by_reft = {}
revealed = {}
self_attested = {}
predicates = {}
# Note we are working on a friendlier api to abstract this away
if credentials:
for credential in credentials:
for attribute_reft in credential["presentation_referents"]:
if attribute_reft not in attribute_by_reft:
attribute_by_reft[attribute_reft] = credential
for (key, value) in attribute_by_reft.items():
print(f"Attribute {presentation_record['presentation_request']['requested_attributes'][key]} can be satisfied by Credential with Referent -- {value['cred_info']['referent']}")
for attribute_reft in presentation_record["presentation_request"]["requested_attributes"]:
if attribute_reft in attribute_by_reft:
revealed[attribute_reft] = {
"cred_id": attribute_by_reft[attribute_reft]["cred_info"][
"referent"
],
"revealed": True,
}
print("\nGenerate the proof")
presentation = {
"requested_predicates": predicates,
"requested_attributes": revealed,
"self_attested_attributes": self_attested,
}
print(presentation)
```
## Send Presentation
A presentation is sent in represent to a presentation record that has previously been created.
```
presentation_response = await agent_controller.proofs.send_presentation(presentation_exchange_id, presentation)
```
## Your Own Business Logic
Now you should have an established, active connection you can write any custom logic you want to engage with protocols with the connection
```
## Custom Logic
```
## Terminate Controller
Whenever you have finished with this notebook, be sure to terminate the controller. This is especially important if your business logic runs across multiple notebooks.
```
await agent_controller.terminate()
```
| github_jupyter |
# CCMP Winds in a cloud-optimized-format for Pangeo
The Cross-Calibrated Multi-Platform (CCMP) Ocean Surface Wind Vector Analyses is part of the NASA Making Earth System Data Records for Use in Research Environments (MEaSUREs) Program. MEaSUREs, develops consistent global- and continental-scale Earth System Data Records by supporting projects that produce data using proven algorithms and input. If you use this data, please give [credit](https://podaac.jpl.nasa.gov/MEaSUREs-CCMP?sections=about). For more information, please review the [documentation](https://podaac-tools.jpl.nasa.gov/drive/files/allData/ccmp/L2.5/docs/ccmp_users_guide.pdf). Please note that this data is not recommended for trend calculations.
# Accessing cloud satellite data
- CCMP zarr conversion funding: Interagency Implementation and Advanced Concepts Team [IMPACT](https://earthdata.nasa.gov/esds/impact) for the Earth Science Data Systems (ESDS) program and AWS Public Dataset Program
### Credits: Tutorial development
* [Dr. Chelle Gentemann](mailto:gentemann@faralloninstitute.org) - [Twitter](https://twitter.com/ChelleGentemann) - Farallon Institute
### Zarr data format
[Zarr](https://zarr.readthedocs.io/en/stable/)
### Data proximate computing
These are BIG datasets that you can analyze on the cloud without downloading the data. You can run this on your phone, a Raspberry Pi, laptop, or desktop.
By using public cloud data, your science is reproducible and easily shared!
### To run this notebook
Code is in the cells that have <span style="color: blue;">In [ ]:</span> to the left of the cell and have a colored background
To run the code:
- option 1) click anywhere in the cell, then hold `shift` down and press `Enter`
- option 2) click on the Run button at the top of the page in the dashboard
Remember:
- to insert a new cell below press `Esc` then `b`
- to delete a cell press `Esc` then `dd`
### First start by importing libraries
```
#libs for reading data
import xarray as xr
import gcsfs
import glob
import numpy as np
import matplotlib.pyplot as plt
from xhistogram.xarray import histogram
#lib for dask gateway
from dask_gateway import Gateway
from dask.distributed import Client
from dask import delayed
```
### Start a cluster, a group of computers that will work together.
(A cluster is the key to big data analysis on on Cloud.)
- This will set up a [dask kubernetes](https://docs.dask.org/en/latest/setup/kubernetes.html) cluster for your analysis and give you a path that you can paste into the top of the Dask dashboard to visualize parts of your cluster.
- You don't need to paste the link below into the Dask dashboard for this to work, but it will help you visualize progress.
- Try 20 workers to start (during the tutorial) but you can increase to speed things up later
```
gateway = Gateway()
cluster = gateway.new_cluster()
cluster.adapt(minimum=1, maximum=75)
client = Client(cluster)
cluster
```
** ☝️ Don’t forget to click the link above or copy it to the Dask dashboard  on the left to view the scheduler dashboard! **
### Initialize Dataset
Here we load the dataset from the zarr store. Note that this very large dataset (273 GB) initializes nearly instantly, and we can see the full list of variables and coordinates.
### Examine Metadata
For those unfamiliar with this dataset, the variable metadata is very helpful for understanding what the variables actually represent
Printing the dataset will show you the dimensions, coordinates, and data variables with clickable icons at the end that show more metadata and size.
```
from intake import open_catalog
cat = open_catalog("https://raw.githubusercontent.com/pangeo-data/pangeo-datastore/master/intake-catalogs/atmosphere.yaml")
ds = cat['nasa_ccmp_wind_vectors'].to_dask()
ds['wspd']=np.sqrt(ds.uwnd**2+ds.vwnd**2) #calculate wind speed
ds
```
# Plot a global image of the data on 7/28/2020
``xarray`` makes plotting the data very easy. A nice overview of plotting with xarray is [here](http://xarray.pydata.org/en/stable/plotting.html). Details on [.plot](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.plot.html#xarray.DataArray.plot)
```
day = ds.sel(time='2020-07-04T00')
day.nobs.plot()
```
## Make a land/ocean/ice mask to show where there is actually data
### Three different ways to mask the data
1. A daily mask that removes data with sea ice and land
- sum over time for nobs (number of observations) variable
- average over a month so that land and monthly sea ice are masked out
2. A mask that removes all data that over land or where there is 'permanent' sea ice
- find when nobs is > 0
3. A climatology mask that removes all data that over land or where there has ever been sea ice
- sum over time for nobs (number of observations) variable
- average over a month so that land and monthly sea ice are masked out
# Apply the mask
- over land, CCMP is ERA5 data
- for many ocean applications a land / sea ice mask is needed
- below are some different mask options that use the CCMP data to generate a mask
```
def mask_data(ds,type):
if type=='daily': #daily mask removes sea ice and land
mask_obs = ds.nobs.rolling(time=180,center=True).max('time') #4 per day 30 days = 180 rolling window
cutoff = 0
if type=='land': # land mask only (includes data over sea ice)
mask_obs = ds.nobs.sum({'time'},keep_attrs=True) #this will give you a LAND mask
cutoff = 0
if type=='climatology': #climatology mask removes max sea ice extent and land
mask_obs = ds.nobs.rolling(time=180,center=True).max('time') #4 per day 30 days = 180 rolling window
mask_obs = mask_obs.sum({'time'},keep_attrs=True)
cutoff = 125000
dy_mask = mask_obs>cutoff
dy_mask = dy_mask.compute() #computing the mask speeds up subsequent operations
masked = ds.where(dy_mask)
return masked,dy_mask
```
# Print what the different masks look like
- This next cell block will take a while as the masks are computed.
```
%%time
subset=ds.isel(time=slice(500,3500))
masked1,dy_mask = mask_data(subset,'daily')
masked2,land_mask = mask_data(subset,'land')
masked3,clim_mask = mask_data(ds,'climatology')
fig, ax = plt.subplots(1,3, figsize=(18,6))
masked1.wspd.isel(time=500).plot(ax=ax[0])
masked2.wspd.isel(time=500).plot(ax=ax[1])
masked3.wspd.isel(time=1000).plot(ax=ax[2])
masked1
```
# For this we will use the climatology mask
```
# decide which mask to use 1=land/ice, 2=land, 3=climatology
masked,mask_obs = mask_data(ds,'climatology')
mask_obs.plot()
fig, ax = plt.subplots(1,3, figsize=(18,6))
masked.wspd[100,:,:].plot(ax=ax[0])
masked.wspd[-100,:,:].plot(ax=ax[1])
masked.wspd[5000,:,:].plot(ax=ax[2])
```
# create a weighted global mean function
```
# from http://gallery.pangeo.io/repos/pangeo-gallery/cmip6/global_mean_surface_temp.html
def global_mean(ds):
lat = ds.latitude
weight = np.cos(np.deg2rad(lat))
weight /= weight.mean()
other_dims = set(ds.dims) - {'time'}
return (ds * weight).mean(other_dims)
```
# calculate the global mean
- I wish I didn't have to have these loops. Programatically, it would be much cleaner to just do:
```python
#glb_mn = global_mean(masked)
#glb_mn = glb_mn.compute()
#print(glb_mn)
```
- but this code doesn't run, it kills my kernel (memory?) every time I try
- for some reason if I run it year by year it runs fine.
```
m,x=[],[]
for lyr in range(1988,2020):
subset = masked.sel(time=str(lyr))
m1 = global_mean(subset)
m1 = m1.mean()
m1_computed = m1.compute()
m.append(m1_computed)
x.append(lyr)
print(lyr)
mn_yr = xr.concat(m, dim='time')
mn_yr['time']=np.arange(1988,2020)
glb_mn = np.mean(mn_yr)
print(glb_mn)
```
# Results
-glb_mean = 1988 - 2019 41 years
- nobs 1.296
- uwnd -0.4763
- vwnd 0.2749
- wspd 8.558
```
plt.rcParams['figure.figsize'] = (12,6)
mn_yr.wspd.plot()
#plt.legend(fontsize=8)
plt.xlim(1988,2020)
#plt.ylim()
plt.ylabel('CCMPv2 Wind Speed (m s$^{-1}$)',fontsize=18)
plt.xlabel('Year',fontsize=18)
#plt.text(10,0.011,'CCMPv2 1988-2019 ',fontsize=18)
plt.text(2005,8.5,'Global mean = 8.6 m s$^{-1}$',fontsize=16)
#plt.text(10,0.009,'67% of winds are > 6 m s$^{-1}$',fontsize=16)
plt.savefig('./../../figures/ccmp_ts_mean.png')
```
# global Histogram figure
```
bins = np.arange(0,30,.1)
h,x=[],[]
for lyr in range(1988,2020):
subset = masked.wspd.sel(time=str(lyr))
h1 = histogram(subset, bins=[bins])
h1 = h1.compute()
print('start',lyr)
h.append(h1)
x.append(lyr)
hh = xr.concat(h, dim='time')
hh.to_netcdf('./../../data/ccmp/ccmp_annual_hist_20210507a.nc')
print('end',lyr)
hh=xr.open_dataset('./../../data/ccmp/ccmp_annual_hist_20210507.nc')
hh1=xr.open_dataset('./../../data/ccmp/ccmp_annual_hist_20210507a.nc')
#hh1.assign_coords['time']=hh1.time+27
hh=xr.concat([hh,hh1],dim='time')
hh['time']=np.arange(1988,2020)
hh.to_netcdf('./../../data/ccmp/ccmp_annual_hist_20210507_final.nc')
hh = xr.open_dataset('./../../data/ccmp/ccmp_annual_hist_20210507_final.nc')
hhall = hh.histogram_wspd.sum('time')
hhall
yr = hh.histogram_wspd[0,:].load()
yr.plot()
print('percentage of winds =< 2 m/s',hhall[0:21].sum()/hhall.sum())
print('percentage of winds =< 6 m/s',hhall[0:60].sum()/hhall.sum())
print('percentage of winds > 6 m/s',hhall[60:].sum()/hhall.sum())
hh2=hh
x=hh.time
plt.rcParams['figure.figsize'] = (8,8)
for iyr in range(32):
plt.plot(hh.wspd_bin,hh2.histogram_wspd[iyr,:]/hh2.histogram_wspd[iyr,:].sum(),label=str(x[iyr].data))
plt.legend(fontsize=8)
plt.xlim(-0,32)
plt.ylim(0,.013)
plt.xlabel('CCMP Wind Speed (m s$^{-1}$)',fontsize=18)
plt.ylabel('PDF (s m$^{-1}$)',fontsize=18)
plt.text(11,0.011,'CCMPv2 1988-2019 ',fontsize=18)
plt.text(11,0.010,'Global mean = 8.6 m s$^{-1}$',fontsize=16)
plt.text(11,0.009,'68% of winds are > 6 m s$^{-1}$',fontsize=16)
plt.savefig('./../../figures/ccmp_annual_hist.png')
plt.rcParams['figure.figsize'] = (8,8)
hhall = hh2.sum('time')
plt.plot(hh.wspd_bin,hhall.histogram_wspd/hhall.histogram_wspd.sum(),linewidth=5)
plt.xlim(-0,30)
plt.ylim(0,.012)
plt.xlabel('CCMP Wind Speed (m s$^{-1}$)',fontsize=18)
plt.ylabel('PDF (s m$^{-1}$)',fontsize=18)
plt.text(10,0.011,'CCMPv2 1988-2019 ',fontsize=18)
plt.text(10,0.010,'Global mean = 8.6 m s$^{-1}$',fontsize=18)
plt.text(10,0.009,'68% of winds are > 6 m s$^{-1}$',fontsize=18)
plt.savefig('./../../figures/ccmp_all_hist2.png')
bins = np.arange(0,30,.1)
h,x=[],[]
for lyr in range(1988,2020):
subset = masked.wspd.sel(time=str(lyr))
h1 = histogram(subset, bins=[bins])
h1 = h1.compute()
print('start',lyr)
h.append(h1)
x.append(lyr)
hh = xr.concat(h, dim='time')
hh.to_netcdf('./../../data/ccmp/ccmp_annual_hist_20210507a.nc')
print('end',lyr)
```
# maps of wind speed distributions for c.donlon
```
%%time
# calc % winds
#a spatial map showing a climatology of roughness.
#Ideally in 3 panels - (a) at Hs=<2, (b) at Hs=mean wind speed (c) Hs> 10
wnd = ds.wspd.where(ds.wspd<=2)
f2 = (wnd/wnd).sum({'time'})/len(wnd.time)*100 # percent less than or equal to 2 m/s
wnd = ds.wspd.where((ds.wspd>=8)&(ds.wspd<=9))
f8 = (wnd/wnd).sum({'time'})/len(wnd.time)*100 # percent 8-9 m/s
wnd = ds.wspd.where(ds.wspd>10)
f10 = (wnd/wnd).sum({'time'})/len(wnd.time)*100 # percent >= 10 m/s
%%time
f2 = f2.compute()
f8 = f8.compute()
f10 = f10.compute()
ff = xr.concat([f2,f8,f10],dim='frac')
plt.rcParams['figure.figsize'] = (15.0,8.0)
plt.rcParams.update({'font.size': 16})
fg = ff.plot(aspect=1, size=10, vmin=0, vmax=100,
col="frac",
transform=ccrs.PlateCarree(), # remember to provide this!
subplot_kws={
"projection": ccrs.PlateCarree()
},
cbar_kwargs={"label":'Percent',"orientation": "horizontal", "shrink": 0.8, "aspect": 40},
robust=True,
)
tstr = ['< 2 m/s','8-9 m/s','> 10 m/s']
for i, ax in enumerate(fg.axes.flat):
ax.set_title(tstr[i])
fg.map(lambda: plt.gca().coastlines())
fig_fname = '../../figures/map_global_wind_distributions.png'
plt.savefig(fig_fname, transparent=False, format='png')
ff2 = ff.where(mask_obs>0)
fg = ff2.plot(aspect=1, size=10, vmin=0, vmax=100,
col="frac",
transform=ccrs.PlateCarree(), # remember to provide this!
subplot_kws={
"projection": ccrs.PlateCarree()
},
cbar_kwargs={"label":'Percent',"orientation": "horizontal", "shrink": 0.8, "aspect": 40},
robust=True,
)
tstr = ['< 2 m/s','8-9 m/s','> 10 m/s']
for i, ax in enumerate(fg.axes.flat):
ax.set_title(tstr[i])
fg.map(lambda: plt.gca().coastlines())
fig_fname = '../../figures/map_ocean_wind_distributions.png'
plt.savefig(fig_fname, transparent=False, format='png')
import cartopy.crs as ccrs
plt.rcParams['figure.figsize'] = (18.0,5.0)
plt.rcParams.update({'font.size': 16})
ax = plt.subplot(131,projection=ccrs.PlateCarree())
cs=f2.plot(ax=ax,vmin=0,vmax=100,cbar_kwargs={'shrink':.35,'label': 'Wind < 2 m/s'})
ax.coastlines()
ax = plt.subplot(132,projection=ccrs.PlateCarree())
cs=f8.plot(ax=ax,vmin=0,vmax=100,cbar_kwargs={'shrink':.35,'label': 'Wind 8-9 m/s'})
ax.coastlines()
ax = plt.subplot(133,projection=ccrs.PlateCarree())
cs=f10.plot(ax=ax,vmin=0,vmax=100,cbar_kwargs={'shrink':.35,'label': 'Wind > 10 m/s'})
ax.coastlines()
# calculate weibull distributions TESTING STILL
ds
#test out weibull at one point with data and without data
import scipy.stats as stats
data = ds.wspd[:,0,400].load()
params = stats.exponweib.fit(data, floc=0, f0=1)
shape = params[1]
scale = params[3]
values,bins,hist = plt.hist(data,bins=51,range=(0,25),density=True)
center = (bins[:-1] + bins[1:]) / 2.
# Using all params and the stats function
params = stats.exponweib.fit(data, floc=0, f0=1)
plt.plot(center,stats.exponweib.pdf(center,*params),lw=4,label='scipy exp')
params = stats.exponweib.fit(ds.wspd, floc=0, f0=1)
params.to_netcdf('./../../data/weib.nc')
params
#adapted from https://gist.github.com/luke-gregor/4bb5c483b2d111e52413b260311fbe43
def dataset_encoding(xds):
cols = ['source', 'original_shape', 'dtype', 'zlib', 'complevel', 'chunksizes']
info = pd.DataFrame(columns=cols, index=xds.data_vars)
for row in info.index:
var_encoding = xds[row].encoding
for col in info.keys():
info.ix[row, col] = var_encoding.pop(col, '')
return info
def xarray_trend(xarr):
from scipy import stats
import numpy as np
# getting shapes
m = np.prod(xarr.shape[1:]).squeeze()
n = xarr.shape[0]
# creating x and y variables for linear regression
#x = xarr.time.to_pandas().index.to_julian_date().values[:, None]
y = xarr.to_masked_array().reshape(n, -1)
# ############################ #
# LINEAR REGRESSION DONE BELOW #
params = stats.exponweib.fit(y, floc=0, f0=1)
shape = params[1]
scale = params[3]
# preparing outputs
out = xarr[:2].mean('time')
# first create variable for slope and adjust meta
xarr_slope = out.copy()
xarr_slope.name += '_shape'
xarr_slope.attrs['units'] = 'none'
xarr_slope.values = shape.reshape(xarr.shape[1:])
# do the same for the p value
xarr_p = out.copy()
xarr_p.name += '_scale'
xarr_p.attrs['info'] = "none"
xarr_p.values = p.reshape(xarr.shape[1:])
# join these variables
xarr_out = xarr_slope.to_dataset(name='shape')
xarr_out['scale'] = xarr_p
return xarr_out
sst_slope2=[]
for inc in range(0,1):
mlon=inc*5
mlon2 = (inc+1)*5-1
subset = ds.wspd.sel(longitude=slice(mlon,mlon2),latitude=slice(-78,-68)).load()
sst_slope = xarray_trend(subset)
sst_slope2.append(sst_slope)
from scipy import stats
import numpy as np
# getting shapes
xarr = subset
m = np.prod(xarr.shape[1:]).squeeze()
n = xarr.shape[0]
# creating x and y variables for linear regression
#x = xarr.time.to_pandas().index.to_julian_date().values[:, None]
y = xarr.to_masked_array().reshape(n, -1)
# ############################ #
# LINEAR REGRESSION DONE BELOW #
params = stats.exponweib.fit(y, floc=0, f0=1)
shape = params[1]
scale = params[3]
# preparing outputs
out = xarr[:2].mean('time')
# first create variable for slope and adjust meta
xarr_slope = out.copy()
xarr_slope.name += '_shape'
xarr_slope.attrs['units'] = 'none'
xarr_slope.values = shape.reshape(xarr.shape[1:])
# do the same for the p value
xarr_p = out.copy()
xarr_p.name += '_scale'
xarr_p.attrs['info'] = "none"
xarr_p.values = p.reshape(xarr.shape[1:])
# join these variables
xarr_out = xarr_slope.to_dataset(name='shape')
xarr_out['scale'] = xarr_p
return xarr_out
sst_slope2=[]
for inc in range(0,35):
mlon=inc*10
mlon2 = (inc+1)*10-1
subset = ds.wspd.sel(longitude=slice(mlon,mlon2))
sst_slope = xarray_trend(subset)
sst_slope2.append(sst_slope)
sst_slope
cluster.close()
```
| github_jupyter |
### Exploratory Data Analysis with Python
We will explore the NYC MTA turnstile dataset. These data files are from the New York Subway. It tracks the hourly entries and exits to turnstiles by day in the subway system.
The data files are available on MTA's [website](http://web.mta.info/developers/turnstile.html).
```
!pip install wget
import os, wget
url_template = "http://web.mta.info/developers/data/nyct/turnstile/turnstile_%s.txt"
for date in ['160206', '160213', '160220', '160227', '160305']:
url = url_template % date
if os.path.isfile('data/turnstile_{}.txt'.format(date)):
print(date, 'file already downloaded')
else:
wget.download(url, out='data/')
print(date, 'file downloaded')
```
Our first step will be to create a dictionary of which the key will be the columns representing a turnstile (C/A, UNIT, SCP, STATION) and the value will be a list of the entries for that turnstile. It should look like so:
{
('A002','R051','02-00-00','LEXINGTON AVE'):
[
['NQR456', 'BMT', '01/03/2015', '03:00:00', 'REGULAR', '0004945474', '0001675324'],
['NQR456', 'BMT', '01/03/2015', '07:00:00', 'REGULAR', '0004945478', '0001675333'],
['NQR456', 'BMT', '01/03/2015', '11:00:00', 'REGULAR', '0004945515', '0001675364'],
...
]
}
```
import csv, glob
from collections import defaultdict
def read_csv(csv_file_name):
turnstile_to_count_reading = defaultdict(list)
with open(csv_file_name, 'r') as csv_file:
mta_reader = csv.reader(csv_file)
for i, row in enumerate(mta_reader):
if i == 0:
continue
turnstile_info = tuple(row[:4])
count_reading = row[4:]
turnstile_to_count_reading[turnstile_info].append(count_reading)
return turnstile_to_count_reading
weekly_data_dicts = [read_csv(csvfile) for csvfile in glob.glob('data/turnstile_*.txt')]
sample_dict = list(weekly_data_dicts[0].items())[:1]
sample_dict
```
Now let's turn this into a time series. This time our data will be comprised of just the point in time and the cumulative count of entries.
It should look like something like:
{
('A002','R051','02-00-00','LEXINGTON AVE'):
[
[datetime.datetime(2013, 3, 2, 3, 0), 3788],
[datetime.datetime(2013, 3, 2, 7, 0), 2585],
[datetime.datetime(2013, 3, 2, 12, 0), 10653],
[datetime.datetime(2013, 3, 2, 17, 0), 11016],
[datetime.datetime(2013, 3, 2, 23, 0), 10666],
[datetime.datetime(2013, 3, 3, 3, 0), 10814],
[datetime.datetime(2013, 3, 3, 7, 0), 10229],
...
],
...
}
```
from datetime import datetime
from dateutil.parser import parse
def convert_week_data_to_time_series(week_data_dict):
turnstile_to_time_series = defaultdict(list)
for i, (turnstile, row_data) in enumerate(week_data_dict.items()):
if i % 200 == 0:
print('Processing turnstile', turnstile)
for lines, division, datestr, timestr, event, cum_entries, cum_exits in row_data:
timestamp = parse('%sT%s' % (datestr, timestr))
turnstile_to_time_series[turnstile].append([timestamp, int(cum_entries)])
return turnstile_to_time_series
weekly_time_series = list(map(convert_week_data_to_time_series, weekly_data_dicts))
sample_turnstile_to_time_series = list(weekly_time_series[0].items())[:2]
sample_turnstile_to_time_series
```
These counts are grouped by dataset file (e.g., by week). Let's make it a high-res timeseries by combining multiple weeks.
```
def combine_multiple_weeks_into_single_high_res_timeseries(weekly_time_series):
combined_time_series = defaultdict(list)
for turnstile_to_weeklong_time_series in weekly_time_series:
for turnstile, weeklong_time_series in turnstile_to_weeklong_time_series.items():
combined_time_series[turnstile] += weeklong_time_series
# It's already sorted due to the nature of the files
return combined_time_series
turnstile_to_full_time_series = combine_multiple_weeks_into_single_high_res_timeseries(
weekly_time_series)
sample_turnstile_to_full_time_series = list(turnstile_to_full_time_series.items())[:2]
sample_turnstile_to_full_time_series
```
This seems to be a good time to take a break and ignore January, March, weekends and 2016 NYC holidays (Feb 12th and 15th). The sooner we do it the faster our code will execute.
```
feb_nyc_holidays = [12, 15]
removed = 0
for turnstile, turnstile_data in turnstile_to_full_time_series.items():
# iterate over a copy of the list in order to be able to remove items from the original
for timestamp, cum_entries in list(turnstile_data):
if timestamp.month != 2 or timestamp.weekday() >= 5 or timestamp.day in feb_nyc_holidays:
if not (timestamp.month == 1 and timestamp.day == 31):
# leave the last of january in order to be able to make the cumulative count
turnstile_data.remove([timestamp, cum_entries])
removed = removed + 1
print(removed)
```
Let's also further analyze the timestamps to see if we can easily filter entries between 8am and 8pm. Again, the sooner we do it the faster our code will execute.
```
turnstiles_timestamps = dict()
for turnstile, turnstile_data in turnstile_to_full_time_series.items():
timestamps_set = set()
for timestamp, cum_entries in list(turnstile_data):
timestamps_set.add(timestamp.time())
turnstiles_timestamps[turnstile] = timestamps_set
turnstiles_timestamps_items_list = list(turnstiles_timestamps.items())
n_turnstiles = len(turnstiles_timestamps_items_list)
n_samples = 4
sample_turnstiles_timestamps = [turnstiles_timestamps_items_list[i]
for i in range(0, n_turnstiles - 1, n_turnstiles // n_samples)]
sample_turnstiles_timestamps
```
Unfortunately, with only a few turnstile samples we can see that the timestamps in which data was recorded is not regular, with some timesamples being seemingly random. Since each station has multiple turnstiles it makes it hard even to compile timestamp data on a station basis.
Let's ignore the timestamps going forward and work with daily entries. We will have a single value for a single day, which is not cumulative counts but the total number of passengers that entered through this turnstile on this day.
```
sample_turnstile_to_full_time_series = list(turnstile_to_full_time_series.items())[:2]
sample_turnstile_to_full_time_series
from itertools import groupby
def count_within_normal_bounds(count):
if count is None:
return True
else:
return 10000 > count >= 0
def convert_time_series_to_daily(high_res_time_series):
daily_time_series = []
def day_of_timestamp(time_series_entry):
timestamp, tot_entries = time_series_entry
return timestamp.date()
# groupby() requires data to be sorted. It is sorted already here.
count_on_previous_day = None
for day, entries_on_this_day in groupby(high_res_time_series, key=day_of_timestamp):
# get the maximum cumulative count among the entries on this day
cum_entry_count_on_day = max([count for time, count in entries_on_this_day])
# skip the first entry if we don't know the previous day
if count_on_previous_day is None:
daily_entries = None
else:
daily_entries = cum_entry_count_on_day - count_on_previous_day
# Save today's count for tomorrow's calculation
count_on_previous_day = cum_entry_count_on_day
# Only append if the cumulative increased.
# Otherwise there is something wrong in the data - skip with a warning.
if count_within_normal_bounds(daily_entries):
daily_time_series.append((day, daily_entries))
else:
print('WARNING. Abnormal entry count found on day %s: %s' % (day, daily_entries))
daily_time_series.append((day, None))
return daily_time_series
def convert_turnstile_to_high_res_time_series_to_daily(turnstile_to_time_series):
turnstile_to_daily_time_series = {}
for i, (turnstile, time_series) in enumerate(turnstile_to_time_series.items()):
print('Processing turnstile', turnstile)
turnstile_to_daily_time_series[turnstile] = convert_time_series_to_daily(time_series)
return turnstile_to_daily_time_series
turnstile_to_daily_time_series = convert_turnstile_to_high_res_time_series_to_daily(
turnstile_to_full_time_series)
turnstile_to_daily_time_series[('N300', 'R113', '01-00-04', '7 AV')]
```
So far we've been operating on a single turnstile level, let's combine turnstiles in the same ControlArea/Unit/Station combo.
```
from collections import Counter
def booth_of_a_time_series_item(item):
turnstile, time_series = item
control_area, unit, device_id, station = turnstile
return (control_area, unit, station)
def reduce_turnstile_time_series_to_booths(turnstile_to_daily_time_series):
turnstile_time_series_items = sorted(turnstile_to_daily_time_series.items())
booth_to_time_series = {}
for booth, item_list_of_booth in groupby(turnstile_time_series_items,
key=booth_of_a_time_series_item):
daily_counter = Counter()
for turnstile, time_series in item_list_of_booth:
for day, count in time_series:
if count is not None:
daily_counter[day] += count
booth_to_time_series[booth] = sorted(daily_counter.items())
return booth_to_time_series
booth_to_daily_time_series = reduce_turnstile_time_series_to_booths(turnstile_to_daily_time_series)
booth_to_daily_time_series[('N300', 'R113', '7 AV')]
```
Similarly, we will combine everything in each station, and come up with a time series for each station by adding up all the turnstiles in a station.
```
def station_of_a_booth(booth):
control_area, unit, station = booth
return station
def station_of_a_time_series_item(item):
booth, time_series = item
return station_of_a_booth(booth)
def reduce_booth_time_series_to_stations(booth_to_daily_time_series):
booth_time_series_items = sorted(booth_to_daily_time_series.items())
station_to_time_series = {}
for station, item_list_of_station in groupby(booth_time_series_items,
key=station_of_a_time_series_item):
daily_counter = Counter()
for turnstile, time_series in item_list_of_station:
for day, count in time_series:
daily_counter[day] += count
station_to_time_series[station] = sorted(daily_counter.items())
return station_to_time_series
station_to_daily_time_series = reduce_booth_time_series_to_stations(booth_to_daily_time_series)
station_to_daily_time_series['7 AV']
```
We'll now make a list of the average ridership values per station and plot it to get an idea about its distribution among different stations.
```
feb_business_days = len(station_to_daily_time_series['7 AV'])
def station_time_series_item_to_station_avg_traffic(item):
station, time_series = item
avg_traffic = sum([count for day, count in time_series]) // feb_business_days
return avg_traffic, station
traffic = list(map(station_time_series_item_to_station_avg_traffic,
station_to_daily_time_series.items()))
traffic_report = sorted(traffic, reverse=True)
for avg_traffic, station in traffic_report[:30]:
print('{:<18} {:.0f}'.format(station, avg_traffic))
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
avg_ridership_counts = [ridership for ridership, station in traffic_report]
fig, ax = plt.subplots(figsize=(20, 10))
sns.distplot(avg_ridership_counts, bins=range(0, 165000, 5000), ax=ax)
ax.set_xlim(0, 165000)
```
We can see that most stations have a small traffic and the histogram bins for large traffic volumes have small bars.
Let's plot a histogram with logarithmic scale instead.
```
import math
log_counts = []
for count in avg_ridership_counts:
try:
log_result = math.log10(count)
except:
pass
log_counts.append(log_result)
fig, ax = plt.subplots(figsize=(20, 10))
sns.distplot(log_counts, bins=15)
def log_count_to_label(log_count):
if log_count <= 3:
return '{0:.0f} Hundred'.format(10 ** (log_count))
else:
return '{0:.1f} Thousand'.format(10 ** (log_count-3))
tick_labels = map(log_count_to_label, bins)
ticks = plt.xticks(bins, tick_labels, rotation=70)
plt.xlabel('Average Ridership per Day (log 10)')
plt.ylabel('Number of Stations with this Total Count')
plt.title('Distribution of ridership among NYC Subway Stations')
plt.savefig('figures/log.png', bbox_inches='tight')
```
Since we are interested in filtering out at least 90% of stations let's select the top 30.
```
import pandas as pd
top_stations = traffic_report[:30]
avgs, stations = zip(*top_stations)
indices = range(len(avgs))
df = pd.DataFrame([stations, avgs]).T
df.columns = ['Station', 'Average Daily Entries']
df.head(20)
fig, ax = plt.subplots(figsize=(20, 10))
sns.barplot(x='Station', y='Average Daily Entries', data=df,
palette=sns.diverging_palette(255, 240, n=len(df)))
ticks = plt.xticks(indices, stations, rotation = 70)
plt.title('Top 30 NYC Subway Stations by Traffic in February Weekdays')
plt.savefig('figures/mvp.png', bbox_inches='tight')
```
This could be our MVP. Let's save the traffic report as a binary `pickle` file and take a break to figure out how to improve the recommendations.
```
import pandas as pd
reversed_traffic_report = [reversed(t) for t in traffic_report]
df_to_pickle = pd.DataFrame(reversed_traffic_report, columns=['station', 'avg_daily_traffic_feb'])
df_to_pickle.head()
df_to_pickle.to_pickle('pickle/stations_traffic.p')
```
| github_jupyter |
```
import os
os.chdir("../../")
import sqlite3
import pandas as pd
from api.utils.database import rows_to_dicts
con = sqlite3.connect("./pipeline/database.db")
```
Compare 2017 and 2018 rent-burden rates
tables: rent_burdened, belonging, community_areas
columns: years, area_number, value
joins: self-join with both rent_burden and belonging
```
cur = con.cursor()
rows = cur.execute("""
SELECT
rent_2018.area_number,
rent_2017.value AS rb_rate_2017,
rent_2018.value AS rb_rate_2018,
belonging_2017.value as belonging_rate_2017,
belonging_2018.value as belonging_rate_2018,
rent_2018.value - rent_2017.value as rb_change,
belonging_2018.value - belonging_2017.value as belonging_change
FROM (
SELECT *
FROM rent_burdened_households
WHERE period_end_year=2017
AND segment="all"
) rent_2017
LEFT JOIN (
SELECT *
FROM rent_burdened_households
WHERE period_end_year=2018
AND segment="all"
) rent_2018
ON rent_2017.area_number = rent_2018.area_number
LEFT JOIN (
SELECT *
FROM belonging
WHERE period_end_year=2017
AND segment="all"
) belonging_2017
ON rent_2017.area_number = belonging_2017.area_number
LEFT JOIN (
SELECT *
FROM belonging
WHERE period_end_year=2018
AND segment="all"
) belonging_2018
ON rent_2017.area_number = belonging_2018.area_number
""").fetchall()
df = pd.DataFrame(rows_to_dicts(cur, rows))
df
import seaborn as sns
sns.scatterplot(x=df["rb_change"],y=df["belonging_change"])
```
Compare the rate of belonging to the number of rideshare pickups divided by population.
That is, the number of rideshare pickups per day in each area divided by that area's population
Should show area_number,
population,
number of rideshare pickups per day,
belonging rate of that area,
and the rideshares per day divded by population
```
cur = con.cursor()
rows = cur.execute("""
SELECT
rideshare_2018.area_number AS area_number,
population_2018.value AS area_population,
rideshare_2018.value / 365 AS rs_per_day_2018,
belonging_2018.value AS belonging_rate_2018,
((rideshare_2018.value/365) / population_2018.value) AS rideshare_by_population
FROM (
SELECT
pickup_community_area as area_number,
sum(n_trips) as value,
strftime('%Y', week) as period_end_year
FROM rideshare
WHERE period_end_year = '2018'
GROUP BY area_number
) rideshare_2018
LEFT JOIN (
SELECT *
FROM belonging
WHERE period_end_year=2018
AND segment="all"
) belonging_2018
ON rideshare_2018.area_number = belonging_2018.area_number
LEFT JOIN (
SELECT *
FROM population
WHERE period_end_year=2018
AND segment="all"
) population_2018
ON rideshare_2018.area_number = population_2018.area_number
""").fetchall()
df = pd.DataFrame(rows_to_dicts(cur, rows))
df
sns.scatterplot(x=df["belonging_rate_2018"],y=df["rideshare_by_population"])
```
| github_jupyter |
```
import pandas as pd
import networkx as nx
import subprocess
import matplotlib.pyplot as plt
import gensim
import os
from networkx.drawing.nx_agraph import graphviz_layout
from chinese_whispers import chinese_whispers, aggregate_clusters
from gensim.models.poincare import PoincareModel
from nltk.corpus import wordnet as wn
```
### Construct the Networkx graph
From a csv file
```
def display_taxonomy(graph):
""" Display the taxonomy in a hierarchical layout """
pos = graphviz_layout(graph, prog='dot', args="-Grankdir=LR")
plt.figure(3,figsize=(48,144))
nx.draw(graph, pos, with_labels=True, arrows=True)
plt.show()
# Construct the networkx graph
def process_input(taxonomy):
""" Read the taxonomy and generate a networkx graph """
# Generated
df = pd.read_csv(
taxonomy,
sep='\t',
header=None,
names=['hyponym', 'hypernym'],
usecols=[1,2],
)
G = nx.DiGraph()
for rel in zip(list(df['hypernym']), list(df['hyponym'])):
rel_0 = rel[0]
rel_1 = rel[1]
# Simplify the compound words by replacing the whitespaces with underscores
if ' ' in rel[0]:
rel_0 = '_'.join(rel[0].split())
if ' ' in rel[1]:
rel_1 = '_'.join(rel[1].split())
G.add_edge(rel_0, rel_1)
return G
taxo_path = 'taxi_output/simple_full/science/science_en.csv-relations.csv-taxo-knn1.csv'
gs_path = 'eval/taxi_eval_archive/gold_standard/science.taxo'
G_taxo = process_input(taxo_path)
G_gold = process_input(gs_path)
print('Nodes in GS:', len(set(G_gold.nodes())))
print('Nodes in G Taxo:', len(set(G_taxo.nodes())))
new_nodes = set(G_gold.nodes()) - set(G_taxo.nodes())
len(new_nodes)
```
## Load Word Vectors
```
def load_vectors():
""" Load word vectors. """
embedding_dir = '/home/5aly/taxi/distributed_semantics/embeddings/'
poincare_model = model = PoincareModel.load(embedding_dir + 'embeddings_poincare_wordnet') # parent-cluster relationship
own_model = gensim.models.KeyedVectors.load(embedding_dir + 'own_embeddings_w2v') # family-cluster relationship
return poincare_model, own_model
poincare_w2v, own_w2v = load_vectors()
```
# Improving Taxonomy with Distributional Semantics
Create a networkx graph for each node containing only its children. Draw edges among the children based on the similarity with one another using word vectors.
```
def create_children_clusters(own_model, graph):
""" This function returns a dictionary where corresponding to each key(node) is a graph of its children """
clustered_graph = {}
for node in graph.nodes():
clustered_graph[node] = nx.Graph()
successors = [s.lower() for s in graph.successors(node)]
for successor in successors:
try:
for word, _ in own_model.most_similar(successor, topn=100):
if word.lower() in successors:
clustered_graph[node].add_edge(successor, word.lower())
except KeyError: # If the word in not in vocabulary, check using the substring based method
successor_terms = successor.split('_')
if node in successor_terms:
clustered_graph[node].add_node(successor)
return clustered_graph
GC = create_children_clusters(own_w2v, G_taxo)
posI = graphviz_layout(GC['engineering'])
# plt.figure(2, figsize=(20, 20))
nx.draw(GC['engineering'], posI, with_labels=True, arrows=True)
plt.show()
```
## Implementing Chinese Whispers Algorithm
### Adding new nodes
- Loop through all the new nodes.
- For each removed node, find out the family and parent in the graph that has the maximum similarity with it.
```
G_improved = G_taxo.copy()
def calculate_similarity(poincare_model, own_model, parent, family, node, exclude_parent, exclude_family):
# Similarity between the parent and a cluster
parent_similarity = 0
if not exclude_parent:
node_senses = [n_sense.name() for n_sense in wn.synsets(node) if node in n_sense.name()]
parent_senses = [p_sense.name() for p_sense in wn.synsets(parent) if parent in p_sense.name()]
for parent_sense in parent_senses:
for node_sense in node_senses:
try:
similarity = poincare_model.kv.similarity(parent_sense, node_sense)
if similarity > parent_similarity:
parent_similarity = similarity
except KeyError as e:
if parent_sense in str(e):
break
else:
continue
# Similarity between a family and a cluster
family_similarity = 0
if not exclude_family:
family_similarities = []
for f_item in family:
try:
family_similarities.append(own_model.similarity(f_item, node))
except KeyError as e: # skip the terms not in vocabulary
if node in str(e):
break
else:
continue
if len(family_similarities) > 0:
family_similarity = sum(family_similarities) / len(family_similarities)
# Final score is the average of both the similarities
return (parent_similarity + family_similarity) / 2
for node in new_nodes:
max_score = 0
max_score_node = ''
for p_node, graph in GC.items():
gc = chinese_whispers(graph, weighting='top', iterations=60)
for label, family in aggregate_clusters(gc).items():
score = calculate_similarity(poincare_w2v, own_w2v, p_node, family, node, False, False)
if score > max_score:
max_score = score
max_score_node = p_node
G_improved.add_edge(max_score_node, node)
```
### Tuning the nodes and the edges
```
def tune_result(g_improved):
""" Filter the results i.e. remove all the isolated nodes and nodes with blank labels """
print('\nTuning the result...')
if '' in g_improved.nodes():
g_improved.remove_node('')
hypernyms = {x[0] for x in g_improved.edges()}
isolated_nodes = list(nx.isolates(g_improved))
for isolated_node in isolated_nodes:
terms = isolated_node.split('_')
if terms[-1] in hypernyms:
g_improved.add_edge(terms[-1], isolated_node)
elif terms[0] in hypernyms:
g_improved.add_edge(terms[0], isolated_node)
else:
g_improved.remove_node(isolated_node)
return g_improved
tune_result(G_improved)
print('Tuned.')
```
## Save the result
```
def save_result(result, path):
print('\nSaving the result...')
df_improved = pd.DataFrame(list(result.edges()), columns=['hypernym', 'hyponym'])
df_improved = df_improved[df_improved.columns.tolist()[::-1]]
# Replace the underscores with blanks
df_improved['hyponym'] = df_improved['hyponym'].apply(lambda x: x.replace('_', ' '))
df_improved['hypernym'] = df_improved['hypernym'].apply(lambda x: x.replace('_', ' '))
# Store the result
output_path = os.path.join(
'taxi_output', 'distributional_semantics',
os.path.basename(path) + '-' + 'new_ds' + os.path.splitext(path)[-1]
)
df_improved.to_csv(output_path, sep='\t', header=False)
print('Output saved at:', output_path)
return output_path
output_path = save_result(G_improved, taxo_path)
```
## Results visualization
### Clusters
```
def visualize_clusters(graph):
""" Clusterize the nodes of a particular domain in a given graph """
graph_cluster = chinese_whispers(graph, weighting='top', iterations=60)
# Visualize the clustering of graph_cluster using NetworkX (requires matplotlib)
colors = [1. / graph_cluster.node[node]['label'] for node in graph_cluster.nodes()]
fig = plt.gcf()
fig.set_size_inches(20, 20)
nx.draw_networkx(graph_cluster, cmap=plt.get_cmap('jet'), node_color=colors, font_color='black')
plt.show()
GC_improved = create_children_clusters(own_w2v, G_improved)
domain = 'mechanical_engineering'
# Original clusters
visualize_clusters(GC[domain])
# Clusters after detaching
visualize_clusters(GC_detached[domain])
# Clusters after detaching and re-attaching the clusters
visualize_clusters(GC_improved[domain])
```
### Taxonomy
```
# View the original taxonomy
display_taxonomy(G)
# View the modified taxonomy
display_taxonomy(G_improved)
len(list(G.nodes()))
len(list(G_improved.nodes()))
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
from collections import defaultdict, Counter
import seaborn as sns
import numpy as np
import os
from datetime import datetime, timedelta
import matplotlib
from matplotlib.ticker import MaxNLocator
set1 = sns.color_palette('Set1')
sns.set_palette('Set1')
import requests
from bs4 import BeautifulSoup
%load_ext autoreload
%autoreload 2
%matplotlib inline
headers = requests.utils.default_headers()
headers.update({ 'User-Agent': 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:52.0) Gecko/20100101 Firefox/52.0'})
url = "https://www.worldometers.info/coronavirus/"
req = requests.get(url, headers)
soup = BeautifulSoup(req.content, 'html.parser')
t = soup.find(id="main_table_countries_today").text
a = np.array(t.strip().split("\n"))
b = a[153:]
columns = ['Country', 'TotalCases', 'NewCases', 'TotalDeaths',
'NewDeaths', 'TotalRecovered', 'ActiveCases', 'Serious,Critical',
'TotCases/1M pop', 'Deaths/1M pop', 'TotalTests', 'Tests/1M pop', '', '', 'Continent']
df = pd.DataFrame(np.split(b, range(15, len(b), 15)), columns=columns).iloc[:212]
def convert_float(field):
field = field.replace(',', '')
try:
return float(field)
except ValueError:
return None
for c in ['TotCases/1M pop', 'Deaths/1M pop', 'Tests/1M pop']:
df[c] = df[c].apply(convert_float)
for c in ['TotalCases', 'NewCases', 'TotalDeaths',
'NewDeaths', 'TotalRecovered', 'ActiveCases', 'Serious,Critical',
'TotalTests']:
df[c] = df[c].apply(convert_float)
df.sort_values('TotalCases', ascending=False).head(20)
hun_list = []
page_i = 0
while True:
try:
t = pd.read_html(f"https://koronavirus.gov.hu/elhunytak/?page={page_i}")[0]
page_i += 1
hun_list.append(t)
except ValueError:
break
hun = pd.concat(hun_list)
hun = hun.rename(columns={
'Sorszám': 'id',
'Nem': 'gender',
'Kor': 'age',
'Alapbetegségek': 'conditions'
})
hun = hun.drop('id', axis=1)
```
# Általános statisztikák
```
print("Összes elhunyt:", len(hun))
print("Átlagéletkor:", hun.age.mean().round(1))
print("Medián életkor:", hun.age.median())
print("Életkor szórása:", hun.age.std().round(1))
gc = hun.gender.value_counts()
print("Férfi: {}, Nő: {}".format(gc.loc['Férfi'], gc.loc['Nő']))
daily_stats = pd.read_table("daily_stats.tsv", index_col="day", parse_dates=['day'])
today = pd.Timestamp.today()
if today.date() not in daily_stats.index:
print("Adding today's stats")
daily_stats.loc[today, "cnt"] = len(hun)
daily_stats.loc[today, 'M'] = gc.loc['Férfi']# - daily_stats.loc[(datetime.now() - timedelta(1)).date(), 'M']
daily_stats.loc[today, 'F'] = gc.loc['Nő']# - daily_stats.loc[(datetime.now() - timedelta(1)).date(), 'F']
daily_stats.index = daily_stats.index.map(lambda i: i.date())
headers = requests.utils.default_headers()
headers.update({ 'User-Agent': 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:52.0) Gecko/20100101 Firefox/52.0'})
url = "https://koronavirus.gov.hu/"
req = requests.get(url, headers)
soup = BeautifulSoup(req.content, 'html.parser')
for d in soup.find_all("div", class_="diagram-a"):
label = d.find("span", class_="label").get_text()
number = d.find("span", class_="number")
if number:
number = number.get_text().replace(" ", "")
daily_stats.loc[today.date(), label] = int(number)
daily_stats.to_csv("daily_stats.tsv", sep="\t")
daily_stats = daily_stats.rename(columns={'Fertőzött': 'case', 'Gyógyult': 'recovered'})
lineplot_kws = dict(marker='X', linewidth=3, markersize=10)
sns.palplot(sns.color_palette('Set1'))
d = daily_stats.copy()
d = d.rename(columns={'M': 'Férfi', 'F': 'Nő', 'cnt': 'Összes'})
m = sns.color_palette('Set1')
cmap = [m[0], m[1], m[2]]
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.plot(y=['Nő', 'Férfi', 'Összes'], color=cmap, ax=ax, **lineplot_kws)
ax.set_ylim(0, len(hun)+5)
ax.grid(axis='y')
ax.set_xlabel("")
ax.set_ylabel("Elhunytak")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
sns.despine()
plt.tight_layout()
fig.savefig("fig/line_death_count.png", dpi=200)
d = daily_stats.copy()
d = d.rename(columns={'recovered': 'Gyógyult', 'case': 'Összes', 'cnt': 'Elhunyt'})
d['Nyitott'] = d['Összes'] - d['Gyógyult'] - d['Elhunyt']
m = sns.color_palette('Set1')
cmap = [m[2], m[4], m[1], m[0]]
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.plot(y=['Összes', 'Nyitott', 'Gyógyult', 'Elhunyt'], color=cmap, ax=ax, **lineplot_kws)
ax.grid(axis='y')
ax.set_xlabel("")
ax.set_ylabel("Esetek")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
sns.despine()
plt.tight_layout()
fig.savefig("fig/line_all.png", dpi=200)
d = daily_stats.cnt.diff().dropna()
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.plot(ax=ax, **lineplot_kws)
ax.set_ylim(0, d.max()+1)
sns.despine()
ax.grid(axis='y')
ax.set_ylabel("Elhunytak")
ax.set_xlabel("")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
plt.tight_layout()
fig.savefig("fig/line_new_deaths.png", dpi=200)
```
# Meggyógyult / elhunyt naponta
```
d = daily_stats[['cnt', 'recovered']].diff()
d['recovery/death'] = d['recovered'] / d['cnt']
d = d['recovery/death']
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.plot(ax=ax, **lineplot_kws)
ax.set_ylim(0, d.max()+1)
sns.despine()
ax.grid(axis='y')
ax.set_ylabel("Meggyógyult / elhunyt")
ax.set_xlabel("")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
plt.tight_layout()
fig.savefig("fig/line_recovered_per_death.png", dpi=200)
daily_stats
d = daily_stats['Mintavétel'].diff().dropna()
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.plot(ax=ax, **lineplot_kws)
sns.despine()
ax.grid(axis='y')
ax.set_ylabel("Napi új mintavétel")
ax.set_xlabel("")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
plt.tight_layout()
fig.savefig("fig/line_tests_per_day.png", dpi=200)
d = daily_stats[['case', 'Mintavétel']].diff()
d['Mintavétel/új eset'] = d['Mintavétel'] / d['case']
d = d['Mintavétel/új eset']
with sns.plotting_context('notebook', font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 5))
d.dropna().iloc[1:].plot(ax=ax, **lineplot_kws)
#ax.set_ylim(0, d.max()+1)
sns.despine()
ax.grid(axis='y')
ax.set_ylabel("Mintavétel / új eset")
ax.set_xlabel("")
plt.xticks(rotation=45, ha='right')
ax.xaxis.set_major_locator(matplotlib.dates.DayLocator())
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter("%b-%d"))
plt.tight_layout()
fig.savefig("fig/line_test_per_case.png", dpi=200)
```
# Korosztályonként
```
labels = ['40 alatt', '40-49', '50-59', '60-64', '65-69', '70-74', '75-79', '80-84', '85-90', '90 felett']
hun['age_group'] = pd.cut(hun.age, bins=[0, 39, 49, 59, 64, 69, 74, 79, 84, 89, 120], labels=labels)
barplot_kws = dict(alpha=0.8, color=sns.color_palette('Set1')[2], zorder=3)
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(14, 4))
sns.countplot(hun.age_group, ax=ax, **barplot_kws)
ax.set_ylabel("Elhunytak")
ax.set_xlabel("")
ax.grid(axis='y')
sns.despine()
plt.tight_layout()
fig.savefig("fig/bar_death_by_age_group.png", dpi=200)
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 4))
h = hun.groupby(['gender', 'age_group']).size().reset_index()
h = h.rename(columns={0: 'cnt'})
sns.barplot(x=h.age_group, y=h.cnt, hue=h.gender, ax=ax, hue_order=['Nő', 'Férfi'],
palette=[set1[0], set1[1]], **barplot_kws)
ax.legend(title="", loc='upper left', fancybox=True)
ax.yaxis.set_major_locator(MaxNLocator(integer=True))
ax.grid(axis='y')
ax.set_ylabel("Elhunytak")
ax.set_xlabel("")
sns.despine()
plt.tight_layout()
fig.savefig("fig/bar_death_by_age_group_and_gender.png", dpi=200)
```
# Alapbetegségek
```
name_mapping = {}
with open("name_mapping.tsv") as f:
for line in f:
fd = line.strip().split("\t")
name_mapping[fd[0]] = fd[1]
list_of_conditions = []
norm_map = {
"anyagcsere és daganatos betegség": "anyagcsere betegség, daganatos betegség",
"szív- és veseelégtelenség": "szívelégtelenség, veseelégtelenség",
}
def split_conditions(conditions):
conditions = conditions.strip()
cnt = 0
if conditions in ("adat feltöltés alatt", "adatok feltöltése folyamatban", "adat feltöltése folyamatban", "nincs adat", "adatok feltöltés alatt"):
return None
conditions = conditions.replace("\xa0", " ")
for src, tgt in norm_map.items():
conditions = conditions.replace(src, tgt)
for c in conditions.split(","):
c = c.strip()
if not c:
continue
c = name_mapping.get(c, c)
list_of_conditions.append(c)
cnt += 1
return cnt
hun['condition_count'] = hun.conditions.apply(split_conditions)
lc = Counter(list_of_conditions)
conditions = pd.Series(lc)
conditions = conditions.reset_index().rename(columns={'index': 'condition', 0: 'cnt'})
categories = pd.read_table("categories.tsv", names=['condition', 'category'], skiprows=1)
all_cond = set(categories.condition.values)
for c in categories.category.unique():
if c not in all_cond:
categories = categories.append(pd.Series({'condition': c, 'category': c}), ignore_index=True)
conditions = conditions.merge(categories, on='condition', how='left')
```
# Alapbetegségek száma
Hány olyan elhunyt van, akinek 0, 1 vagy több alapbetegsége van.
```
cond_cnt = hun.condition_count.value_counts(dropna=False).sort_index()
cond_cnt['NA'] = cond_cnt.loc[np.nan]
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(10, 5))
#sns.countplot(hun.condition_count, ax=ax, **barplot_kws)
sns.barplot(x=cond_cnt.index, y=cond_cnt, ax=ax, **barplot_kws)
ax.set_ylabel("Elhunytak")
ax.set_xlabel("Alapbetegségek száma")
sns.despine()
ax.grid(axis='y', zorder=0)
plt.tight_layout()
fig.savefig("fig/condition_histogram.png", dpi=200)
```
# 10 leggyakoribb betegség
```
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 7))
common = conditions.sort_values('cnt', ascending=False).head(10)
sns.barplot(y=common.condition, x=common.cnt, ax=ax, **barplot_kws)
ax.set_ylabel("")
ax.set_xlabel("Előfordulás")
ax.grid(axis='x')
sns.despine()
plt.tight_layout()
fig.savefig("fig/bar_most_common_diseases.png", dpi=200)
```
# Kategóriák
```
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(1, figsize=(12, 7))
common = conditions.groupby('category')['cnt'].sum().sort_values(ascending=False).head(10)
sns.barplot(y=common.index, x=common, ax=ax, **barplot_kws)
ax.set_ylabel("")
ax.grid(axis='x')
ax.set_xlabel("Előfordulás")
sns.despine()
plt.tight_layout()
fig.savefig("fig/bar_icd.png", dpi=200)
print("\n".join(conditions[conditions.category.isnull()].condition))
```
| github_jupyter |
```
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
"""
# If you're using Google Colab and not running locally, run this cell
# install NeMo
BRANCH = 'v1.0.0b3'
!python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[nlp]
from nemo.collections import nlp as nemo_nlp
from nemo.utils.exp_manager import exp_manager
from nemo.utils import logging
import os
import wget
import torch
import pytorch_lightning as pl
from omegaconf import OmegaConf
```
# Task Description
**Joint Intent and Slot classification** - is a task of classifying an Intent and detecting all relevant Slots (Entities)
for this Intent in a query.
For example, in the query: `What is the weather in Santa Clara tomorrow morning?`, we would like to classify the query
as a `weather` Intent, and detect `Santa Clara` as a `location` slot and `tomorrow morning` as a `date_time` slot.
Intents and Slots names are usually task specific and defined as labels in the training data.
This is a fundamental step that is executed in any task-driven Conversational Assistant.
Our Bert based model implementation enables to train and then detect both of these tasks together.
# Dataset and NeMo data format
In this tutorial we are going to use a virtual assistant interaction data set that can be downloaded from here: https://github.com/xliuhw/NLU-Evaluation-Data.
There are about 10K training and 1K testing queries which cover 64 various Intents and 55 Slots.
To work with NeMo NLP classification model, this dataset should be first converted to the NeMo format, which requires next files:
- **dict.intents.csv** - list of all intent names in the data. One line per an intent name.
- **dict.slots.csv** - list of all slot names in the data. One line per a slot name. It is possible to use both: B- I- notations, for separating between first and intermediate tokens for multi token slots. Or just use one slot type for each token of multi token slot. Our recommendation is to use later one, since it is simpler and there is no visible degradation in performance.
- **train.tsv/test.tsv** - contain original queries, one per line, and intent number separated by tab. For example: `what alarms do i have set right now 0`. Intent numbers are according to the intent line in the intent dictionary file (dict.intents.csv) starting from 0. First line of these files contains a header line: `sentence \tab label`.
- **train_slot.tvs/test_slot.tsv** - contain one line per a query, where instead each token there is a number of the token from the slots dictionary file (dict.slots.csv), starting from 0. Last 'out-of scope' token is usually located in the last line of the dictionary. Example: `54 0 0 54 54 12 12` (numbers separated by space). No header line in these files.
NeMo provides **import_dataset.py** converter for few reference datasets (Assistant / Atis / Snips) which converts them to the NeMo data format for the Intent and Slot classification model. If you have your own annotated dataset in a different format, you will need to write a data converter. Possible recommended format for your own annotation, is to have one text file per all examples of one intent. With one line per query in a form like: `did i set an alarm to [alarm_type : wake up] in the [timeofday : morning]`, using brackets to define slot names. This is very similar to the assistant format from this example and you can use its converter to NeMo format with small changes.
You can run this utility as follows:
**python examples/nlp/intent_slot_classification/data/import_datasets.py --dataset_name=assistant --source_data_dir=source_dir_name --target_data_dir=target_dir_name**
# Download, preprocess and explore the dataset
## Download the dataset and convert it to the NeMo format
```
# you can replace DATA_DIR and NEMO_DIR with your own locations
DATA_DIR = "."
NEMO_DIR = '.'
# download the converter files from github for the purpose of this tutorial
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/intent_slot_classification/data/import_datasets.py', NEMO_DIR)
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/intent_slot_classification/data/assistant_utils.py', NEMO_DIR)
# download and unzip the example dataset from github
print('Downloading dataset...')
wget.download('https://github.com/xliuhw/NLU-Evaluation-Data/archive/master.zip', DATA_DIR)
! unzip {DATA_DIR}/NLU-Evaluation-Data-master.zip -d {DATA_DIR}
# convert the dataset to the NeMo format
!python {NEMO_DIR}/import_datasets.py --dataset_name=assistant --source_data_dir={DATA_DIR}/NLU-Evaluation-Data-master --target_data_dir={DATA_DIR}/nemo_format
```
## Data exploration
You can see the dataset in both the original and NeMo's formats. We have here 65 different Intents and 55 Slots, which could be typical commands for virtual assistants. Out of scope slot has the name 'O' and is the last in the dictionary of Slots. And we can see examples of queries and also format of training intent and slot files.
```
# list of queries divided by intent files in the original training dataset
! ls -l {DATA_DIR}/NLU-Evaluation-Data-master/dataset/trainset
# print all intents from the NeMo format intent dictionary
!echo 'Intents: ' $(wc -l < {DATA_DIR}/nemo_format/dict.intents.csv)
! cat {DATA_DIR}/nemo_format/dict.intents.csv
# print all slots from the NeMo format slot dictionary
!echo 'Slots: ' $(wc -l < {DATA_DIR}/nemo_format/dict.slots.csv)
! cat {DATA_DIR}/nemo_format/dict.slots.csv
# examples from the intent training file
! head -n 10 {DATA_DIR}/nemo_format/train.tsv
# examples from the slot training file
! head -n 10 {DATA_DIR}/nemo_format/train_slots.tsv
```
# Training model
## Model configuration
Our Joint Intent and Slot classification model is comprised of the pretrained [BERT](https://arxiv.org/pdf/1810.04805.pdf) model with an Intent and Slot Classification layer on top of it.
All model and training parameters are defined in the **intent_slot_classification_config.yaml** config file. This file is located in the folder **examples/nlp/intent_slot_classification/conf/**. It contains 2 main sections:
- **model**: All arguments that are related to the Model - language model, token classifier, optimizer and schedulers, datasets and any other related information
- **trainer**: Any argument to be passed to PyTorch Lightning
We will download the config file from repository for the purpose of the tutorial. If you have a version of NeMo installed locally, you can use it from the above folder.
```
# download the model config file from repository for the purpose of this example
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/intent_slot_classification/conf/intent_slot_classification_config.yaml', NEMO_DIR)
# print content of the config file
config_file = "intent_slot_classification_config.yaml"
print(config_file)
config = OmegaConf.load(config_file)
print(OmegaConf.to_yaml(config))
```
## Setting up Data within the config
Among other things, the config file contains dictionaries called train_ds and validation_ds. These are configurations used to setup the Dataset and DataLoaders of the corresponding config.
The converter utility creates both training and evaluation files in the same directory, so we need to specify `model.data_dir` parameter to this directory. Also notice that some config lines, including `model.data_dir`, have `???` in place of paths, this means that values for these fields are required to be specified by the user.
`config.model.intent_loss_weight` parameter - is a balance of training loss between Intent and Slot losses, a number between 0 to 1. Its default value is 0.6 which gives slightly higher priority to the Intent loss and it empirically works quite well. You can experiment with this value if you like.
Also you can try to change `config.model.class_balancing` parameter to `weighted_loss` and see if you get better accuracy.
Let's now add the data directory path to the config.
```
config.model.data_dir = f'{DATA_DIR}/nemo_format'
```
## Building the PyTorch Lightning Trainer
NeMo models are primarily PyTorch Lightning modules - and therefore are entirely compatible with the PyTorch Lightning ecosystem. `config.trainer.max_epochs` - param defines number of training epochs. Usually 50-100 epochs or less should be enough to train on your data. Let's instantiate the Trainer object.
```
# lets modify some trainer configs
# checks if we have GPU available and uses it
cuda = 1 if torch.cuda.is_available() else 0
config.trainer.gpus = cuda
config.trainer.precision = 16 if torch.cuda.is_available() else 32
# for mixed precision training, uncomment the line below (precision should be set to 16 and amp_level to O1):
# config.trainer.amp_level = O1
# remove distributed training flags
config.trainer.accelerator = None
# setup a small number of epochs for demonstration purposes of this tutorial
config.trainer.max_epochs = 5
trainer = pl.Trainer(**config.trainer)
```
## Setting up a NeMo Experiment
NeMo has an experiment manager that handles logging and checkpointing for us, so let's use it. Model check points during training will be saved in this directory.
```
exp_dir = exp_manager(trainer, config.get("exp_manager", None))
# the exp_dir provides a path to the current experiment for easy access
print(str(exp_dir))
```
## Initializing the model and Training
Initial statistics of the dataset will be displayed at the beginning of the training and then Intent and Slot classification report will be displayed after each training epoch.
```
# initialize the model
model = nemo_nlp.models.IntentSlotClassificationModel(config.model, trainer=trainer)
# train
trainer.fit(model)
```
After training for 5 epochs, which should take no more than few minutes, you can expect training precision for this data set to be around these numbers (the accuracy will gradually continue to improve for this dataset up to about 50 epochs of training):
```
Intents:
label precision recall f1 support
alarm_query (label_id: 0) 94.74 94.74 94.74 19
alarm_remove (label_id: 1) 100.00 100.00 100.00 11
alarm_set (label_id: 2) 85.71 94.74 90.00 19
audio_volume_down (label_id: 3) 0.00 0.00 0.00 8
audio_volume_mute (label_id: 4) 100.00 86.67 92.86 15
audio_volume_up (label_id: 5) 56.52 100.00 72.22 13
calendar_query (label_id: 6) 55.00 57.89 56.41 19
calendar_remove (label_id: 7) 88.89 84.21 86.49 19
calendar_set (label_id: 8) 81.25 68.42 74.29 19
cooking_recipe (label_id: 9) 86.36 100.00 92.68 19
datetime_convert (label_id: 10) 0.00 0.00 0.00 8
datetime_query (label_id: 11) 65.52 100.00 79.17 19
email_addcontact (label_id: 12) 100.00 12.50 22.22 8
email_query (label_id: 13) 83.33 78.95 81.08 19
email_querycontact (label_id: 14) 62.50 78.95 69.77 19
email_sendemail (label_id: 15) 70.83 89.47 79.07 19
general_affirm (label_id: 16) 95.00 100.00 97.44 19
general_commandstop (label_id: 17) 100.00 100.00 100.00 19
general_confirm (label_id: 18) 100.00 100.00 100.00 19
general_dontcare (label_id: 19) 100.00 100.00 100.00 19
general_explain (label_id: 20) 100.00 94.74 97.30 19
general_joke (label_id: 21) 100.00 100.00 100.00 12
general_negate (label_id: 22) 95.00 100.00 97.44 19
general_praise (label_id: 23) 100.00 94.74 97.30 19
general_quirky (label_id: 24) 40.00 10.53 16.67 19
general_repeat (label_id: 25) 100.00 100.00 100.00 19
iot_cleaning (label_id: 26) 84.21 100.00 91.43 16
iot_coffee (label_id: 27) 94.74 94.74 94.74 19
iot_hue_lightchange (label_id: 28) 94.44 89.47 91.89 19
iot_hue_lightdim (label_id: 29) 100.00 83.33 90.91 12
iot_hue_lightoff (label_id: 30) 89.47 89.47 89.47 19
iot_hue_lighton (label_id: 31) 0.00 0.00 0.00 3
iot_hue_lightup (label_id: 32) 81.25 92.86 86.67 14
iot_wemo_off (label_id: 33) 60.00 100.00 75.00 9
iot_wemo_on (label_id: 34) 100.00 14.29 25.00 7
lists_createoradd (label_id: 35) 78.95 78.95 78.95 19
lists_query (label_id: 36) 78.95 78.95 78.95 19
lists_remove (label_id: 37) 90.00 94.74 92.31 19
music_likeness (label_id: 38) 70.59 66.67 68.57 18
music_query (label_id: 39) 77.78 73.68 75.68 19
music_settings (label_id: 40) 0.00 0.00 0.00 7
news_query (label_id: 41) 77.78 73.68 75.68 19
play_audiobook (label_id: 42) 90.00 94.74 92.31 19
play_game (label_id: 43) 80.00 84.21 82.05 19
play_music (label_id: 44) 53.85 73.68 62.22 19
play_podcasts (label_id: 45) 89.47 89.47 89.47 19
play_radio (label_id: 46) 93.75 78.95 85.71 19
qa_currency (label_id: 47) 95.00 100.00 97.44 19
qa_definition (label_id: 48) 85.00 89.47 87.18 19
qa_factoid (label_id: 49) 45.16 73.68 56.00 19
qa_maths (label_id: 50) 100.00 100.00 100.00 14
qa_stock (label_id: 51) 95.00 100.00 97.44 19
recommendation_events (label_id: 52) 94.44 89.47 91.89 19
recommendation_locations (label_id: 53) 94.74 94.74 94.74 19
recommendation_movies (label_id: 54) 100.00 100.00 100.00 10
social_post (label_id: 55) 90.00 94.74 92.31 19
social_query (label_id: 56) 94.74 100.00 97.30 18
takeaway_order (label_id: 57) 93.75 78.95 85.71 19
takeaway_query (label_id: 58) 85.71 94.74 90.00 19
transport_query (label_id: 59) 83.33 78.95 81.08 19
transport_taxi (label_id: 60) 100.00 100.00 100.00 18
transport_ticket (label_id: 61) 89.47 89.47 89.47 19
transport_traffic (label_id: 62) 100.00 100.00 100.00 19
weather_query (label_id: 63) 100.00 89.47 94.44 19
-------------------
micro avg 85.04 85.04 85.04 1076
macro avg 81.13 80.81 79.36 1076
weighted avg 84.10 85.04 83.54 1076
Slots:
label precision recall f1 support
alarm_type (label_id: 0) 0.00 0.00 0.00 0
app_name (label_id: 1) 0.00 0.00 0.00 6
artist_name (label_id: 2) 0.00 0.00 0.00 21
audiobook_author (label_id: 3) 0.00 0.00 0.00 1
audiobook_name (label_id: 4) 0.00 0.00 0.00 18
business_name (label_id: 5) 60.00 56.60 58.25 53
business_type (label_id: 6) 0.00 0.00 0.00 24
change_amount (label_id: 7) 0.00 0.00 0.00 25
coffee_type (label_id: 8) 0.00 0.00 0.00 4
color_type (label_id: 9) 0.00 0.00 0.00 12
cooking_type (label_id: 10) 0.00 0.00 0.00 0
currency_name (label_id: 11) 84.09 75.51 79.57 49
date (label_id: 12) 57.95 91.07 70.83 112
definition_word (label_id: 13) 0.00 0.00 0.00 20
device_type (label_id: 14) 74.55 51.25 60.74 80
drink_type (label_id: 15) 0.00 0.00 0.00 0
email_address (label_id: 16) 0.00 0.00 0.00 14
email_folder (label_id: 17) 0.00 0.00 0.00 1
event_name (label_id: 18) 100.00 13.24 23.38 68
food_type (label_id: 19) 51.72 69.77 59.41 43
game_name (label_id: 20) 60.00 14.29 23.08 21
game_type (label_id: 21) 0.00 0.00 0.00 0
general_frequency (label_id: 22) 0.00 0.00 0.00 9
house_place (label_id: 23) 93.33 42.42 58.33 33
ingredient (label_id: 24) 0.00 0.00 0.00 6
joke_type (label_id: 25) 0.00 0.00 0.00 4
list_name (label_id: 26) 0.00 0.00 0.00 21
meal_type (label_id: 27) 0.00 0.00 0.00 0
media_type (label_id: 28) 0.00 0.00 0.00 37
movie_name (label_id: 29) 0.00 0.00 0.00 0
movie_type (label_id: 30) 0.00 0.00 0.00 0
music_album (label_id: 31) 0.00 0.00 0.00 0
music_descriptor (label_id: 32) 0.00 0.00 0.00 3
music_genre (label_id: 33) 0.00 0.00 0.00 9
news_topic (label_id: 34) 0.00 0.00 0.00 17
order_type (label_id: 35) 0.00 0.00 0.00 17
person (label_id: 36) 44.86 92.31 60.38 52
personal_info (label_id: 37) 0.00 0.00 0.00 20
place_name (label_id: 38) 71.25 77.03 74.03 148
player_setting (label_id: 39) 0.00 0.00 0.00 1
playlist_name (label_id: 40) 0.00 0.00 0.00 1
podcast_descriptor (label_id: 41) 0.00 0.00 0.00 13
podcast_name (label_id: 42) 0.00 0.00 0.00 4
radio_name (label_id: 43) 66.67 10.53 18.18 38
relation (label_id: 44) 0.00 0.00 0.00 17
song_name (label_id: 45) 0.00 0.00 0.00 22
time (label_id: 46) 70.27 78.20 74.02 133
time_zone (label_id: 47) 0.00 0.00 0.00 9
timeofday (label_id: 48) 0.00 0.00 0.00 28
transport_agency (label_id: 49) 0.00 0.00 0.00 9
transport_descriptor (label_id: 50) 0.00 0.00 0.00 0
transport_name (label_id: 51) 0.00 0.00 0.00 4
transport_type (label_id: 52) 78.38 82.86 80.56 35
weather_descriptor (label_id: 53) 0.00 0.00 0.00 17
O (label_id: 54) 92.42 98.80 95.50 5920
-------------------
micro avg 89.10 89.10 89.10 7199
macro avg 21.86 18.56 18.18 7199
weighted avg 84.42 89.10 86.01 7199
```
## Evaluation
To see how the model performs, we can evaluate the performance of the trained model on a test data file. Here we would load the best checkpoint (the one with the lowest validation loss) and create a model (eval_model) from the checkpoint. We will use the same trainer for testing.
```
# extract the path of the best checkpoint from the training, you may update it to any other saved checkpoint file
checkpoint_path = trainer.checkpoint_callback.best_model_path
# load the model from this checkpoint
eval_model = nemo_nlp.models.IntentSlotClassificationModel.load_from_checkpoint(checkpoint_path=checkpoint_path)
# we will setup testing data reusing the same config (test section)
eval_model.setup_test_data(test_data_config=config.model.test_ds)
# run the evaluation on the test dataset
trainer.test(model=model, ckpt_path=None, verbose=False)
```
## Inference from Examples
Next step to see how the trained model will classify Intents and Slots for given queries from this domain. To improve the predictions you may need to train the model for more than 5 epochs.
```
queries = [
'set alarm for seven thirty am',
'lower volume by fifty percent',
'what is my schedule for tomorrow',
]
pred_intents, pred_slots = eval_model.predict_from_examples(queries, config.model.test_ds)
logging.info('The prediction results of some sample queries with the trained model:')
for query, intent, slots in zip(queries, pred_intents, pred_slots):
logging.info(f'Query : {query}')
logging.info(f'Predicted Intent: {intent}')
logging.info(f'Predicted Slots: {slots}')
```
## Training Script
If you have NeMo installed locally (eg. cloned from the Github), you can also train the model with the example script: `examples/nlp/intent_slot_classification/intent_slot_classification.py.`
This script contains an example on how to train, evaluate and perform inference with the IntentSlotClassificationModel.
To run a training script, use:
`cd examples/nlp/intent_slot_classification`
`python intent_slot_classification.py model.data_dir=PATH_TO_DATA_DIR`
By default, this script uses examples/nlp/intent_slot_classification/conf/intent_slot_classification_config.py config file, and you may update all the params inside of this config file or alternatively providing them in the command line.
| github_jupyter |
# In this note book the following steps are taken:
1. Remove highly correlated attributes
2. Find the best hyper parameters for estimator
3. Find the most important features by tunned random forest
4. Find f1 score of the tunned full model
5. Find best hyper parameter of model with selected features
6. Find f1 score of the tuned seleccted model
7. Compare the two f1 scores
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.feature_selection import RFECV,RFE
from sklearn.model_selection import train_test_split, GridSearchCV, KFold,RandomizedSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn import metrics
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score,f1_score
import numpy as np
from sklearn.metrics import make_scorer
f1_score = make_scorer(f1_score)
#import data
Data=pd.read_csv("Waterloo-Transfomed-Data-BS-NoBreak - Copy.csv")
X = Data.iloc[:,:-1]
y = Data.iloc[:,-1]
#split test and training set.
np.random.seed(60)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2,
random_state = 1000)
#Define estimator and model
classifiers = {}
classifiers.update({"Random Forest": RandomForestClassifier(random_state=1000)})
#Define range of hyperparameters for estimator
np.random.seed(60)
parameters = {}
parameters.update({"Random Forest": { "classifier__n_estimators": [100,105,110,115,120,125,130,135,140,145,150,155,160,170,180,190,200],
# "classifier__n_estimators": [2,4,5,6,7,8,9,10,20,30,40,50,60,70,80,90,100,110,120,130,140,150,160,170,180,190,200],
#"classifier__class_weight": [None, "balanced"],
"classifier__max_features": ["auto", "sqrt", "log2"],
"classifier__max_depth" : [4,6,8,10,11,12,13,14,15,16,17,18,19,20,22],
#"classifier__max_depth" : [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20],
"classifier__criterion" :["gini", "entropy"]
}})
# Make correlation matrix
corr_matrix = X_train.corr(method = "spearman").abs()
# Draw the heatmap
sns.set(font_scale = 1.0)
f, ax = plt.subplots(figsize=(11, 9))
sns.heatmap(corr_matrix, cmap= "YlGnBu", square=True, ax = ax)
f.tight_layout()
plt.savefig("correlation_matrix.png", dpi = 1080)
# Select upper triangle of matrix
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k = 1).astype(np.bool))
# Find index of feature columns with correlation greater than 0.8
to_drop = [column for column in upper.columns if any(upper[column] > 0.85)]
# Drop features
X_train = X_train.drop(to_drop, axis = 1)
X_test = X_test.drop(to_drop, axis = 1)
X_train
FEATURE_IMPORTANCE = {"Random Forest"}
selected_classifier = "Random Forest"
classifier = classifiers[selected_classifier]
scaler = StandardScaler()
steps = [("scaler", scaler), ("classifier", classifier)]
pipeline = Pipeline(steps = steps)
#Define parameters that we want to use in gridsearch cv
param_grid = parameters[selected_classifier]
# Initialize GridSearch object for estimator
gscv = RandomizedSearchCV(pipeline, param_grid, cv = 3, n_jobs= -1, verbose = 1, scoring = f1_score, n_iter=30)
# Fit gscv (Tunes estimator)
print(f"Now tuning {selected_classifier}. Go grab a beer or something.")
gscv.fit(X_train, np.ravel(y_train))
#Getting the best hyperparameters
best_params = gscv.best_params_
best_params
#Getting the best score of model
best_score = gscv.best_score_
best_score
#Check overfitting of the estimator
from sklearn.model_selection import cross_val_score
mod = RandomForestClassifier(#class_weight= None,
criterion= 'entropy',
max_depth= 10,
max_features= 'log2',
n_estimators= 155 ,random_state=10000)
scores_test = cross_val_score(mod, X_test, y_test, scoring='f1', cv=5)
scores_test
tuned_params = {item[12:]: best_params[item] for item in best_params}
classifier.set_params(**tuned_params)
#Find f1 score of the model with all features (Model is tuned for all features)
results={}
model=classifier.set_params(criterion= 'entropy',
max_depth= 10,
max_features= 'log2',
n_estimators= 155 ,random_state=10000)
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
F1 = metrics.f1_score(y_test, y_pred)
results = {"classifier": model,
"Best Parameters": best_params,
"Training f1": best_score*100,
"Test f1": F1*100}
results
# Select Features using RFECV
class PipelineRFE(Pipeline):
# Source: https://ramhiser.com/post/2018-03-25-feature-selection-with-scikit-learn-pipeline/
def fit(self, X, y=None, **fit_params):
super(PipelineRFE, self).fit(X, y, **fit_params)
self.feature_importances_ = self.steps[-1][-1].feature_importances_
return self
steps = [("scaler", scaler), ("classifier", classifier)]
pipe = PipelineRFE(steps = steps)
np.random.seed(60)
# Initialize RFECV object
feature_selector = RFECV(pipe, cv = 5, step = 1, verbose = 1)
# Fit RFECV
feature_selector.fit(X_train, np.ravel(y_train))
# Get selected features
feature_names = X_train.columns
selected_features = feature_names[feature_selector.support_].tolist()
performance_curve = {"Number of Features": list(range(1, len(feature_names) + 1)),
"F1": feature_selector.grid_scores_}
performance_curve = pd.DataFrame(performance_curve)
# Performance vs Number of Features
# Set graph style
sns.set(font_scale = 1.75)
sns.set_style({"axes.facecolor": "1.0", "axes.edgecolor": "0.85", "grid.color": "0.85",
"grid.linestyle": "-", 'axes.labelcolor': '0.4', "xtick.color": "0.4",
'ytick.color': '0.4'})
colors = sns.color_palette("RdYlGn", 20)
line_color = colors[3]
marker_colors = colors[-1]
# Plot
f, ax = plt.subplots(figsize=(13, 6.5))
sns.lineplot(x = "Number of Features", y = "F1", data = performance_curve,
color = line_color, lw = 4, ax = ax)
sns.regplot(x = performance_curve["Number of Features"], y = performance_curve["F1"],
color = marker_colors, fit_reg = False, scatter_kws = {"s": 200}, ax = ax)
# Axes limits
plt.xlim(0.5, len(feature_names)+0.5)
plt.ylim(0.60, 1)
# Generate a bolded horizontal line at y = 0
ax.axhline(y = 0.625, color = 'black', linewidth = 1.3, alpha = .7)
# Turn frame off
ax.set_frame_on(False)
# Tight layout
plt.tight_layout()
#Define new training and test set based based on selected features by RFECV
X_train_rfecv = X_train[selected_features]
X_test_rfecv= X_test[selected_features]
np.random.seed(60)
classifier.fit(X_train_rfecv, np.ravel(y_train))
#Finding important features
np.random.seed(60)
feature_importance = pd.DataFrame(selected_features, columns = ["Feature Label"])
feature_importance["Feature Importance"] = classifier.feature_importances_
feature_importance = feature_importance.sort_values(by="Feature Importance", ascending=False)
feature_importance
# Initialize GridSearch object for model with selected features
np.random.seed(60)
gscv = RandomizedSearchCV(pipeline, param_grid, cv = 3, n_jobs= -1, verbose = 1, scoring = f1_score, n_iter=30)
#Tuning random forest classifier with selected features
np.random.seed(60)
gscv.fit(X_train_rfecv,y_train)
#Getting the best parameters of model with selected features
best_params = gscv.best_params_
best_params
#Getting the score of model with selected features
best_score = gscv.best_score_
best_score
#Check overfitting of the tuned model with selected features
from sklearn.model_selection import cross_val_score
mod = RandomForestClassifier(#class_weight= None,
criterion= 'gini',
max_depth= 11,
max_features= 'log2',
n_estimators= 135 ,random_state=10000)
scores_test = cross_val_score(mod, X_test_rfecv, y_test, scoring='f1', cv=5)
scores_test
results={}
model=classifier.set_params(criterion= 'gini',
max_depth= 11,
max_features= 'log2',
n_estimators= 135 ,random_state=10000)
scores_test = cross_val_score(mod, X_test_rfecv, y_test, scoring='f1', cv=5)
model.fit(X_train_rfecv,y_train)
y_pred = model.predict(X_test_rfecv)
F1 = metrics.f1_score(y_test, y_pred)
results = {"classifier": model,
"Best Parameters": best_params,
"Training f1": best_score*100,
"Test f1": F1*100}
results
```
| github_jupyter |
<center><img src="../static/images/python.png" width=500></center>
# Python
This section is meant as a general introduction to Python and is by far not complete. It is based amongst others on the [IPython notebooks from J. R. Johansson](http://github.com/jrjohansson/scientific-python-lectures), on http://www.stavros.io/tutorials/python/ and on http://www.swaroopch.com/notes/python.
<font color='red'>Important:</font> a very good interactive tutorial for Python can also be found on https://www.codecademy.com/learn/python
The goal of this section is to give you a short introduction to Python and help beginners to get familiar with this programming language.
Following chapters are available:
- [Module](#Module)
- [Help and Descriptions](#Help-and-Descriptions)
- [Variables and types](#Variables-and-types)
- [Symbol names](#Symbol-names)
- [Assignment](#Assignment)
- [Fundamental types](#Fundamental-types)
- [Operators and comparisons](#Operators-and-comparisons)
- [Shortcut math operation and assignment](#Shortcut-math-operation-and-assignment)
- [Strings, List and dictionaries](#Strings,-List-and-dictionaries)
- [Strings](#Strings)
- [List](#List)
- [Tuples](#Tuples)
- [Dictionaries](#Dictionaries)
- [Indentation](#Indentation)
- [Control Flow](#Control-Flow)
- [Conditional statements: `if`, `elif`, `else`](#Conditional-statements:-if,-elif,-else)
- [Loops](#Loops)
- [`for` loops](#for-loops)
- [`break`, `continue` and `pass`](#break,-continue-and-pass)
- [Functions](#Functions)
- [Default argument and keyword arguments](#Default-argument-and-keyword-arguments)
- [`*args` and `*kwargs` parameters](#*args-and-*kwargs-parameters)
- [Unnamed functions: `lambda` function](#Unnamed-functions:-lambda-function)
- [Classes](#Classes)
- [Modules](#Modules)
- [Exceptions](#Exceptions)
- [File I/O](#File-I/O)
- [Reading CSV files](#Reading-CSV-files)
- [Writing CSV files](#Writing-CSV-files)
- [Reading TXT files](#Reading-TXT-files)
- [Writing TXT files](#Writing-TXT-files)
- [with open](#with-open)
## Module
Most of the functionality in Python is provided by *modules*. To use a module in a Python program it first has to be imported. A module can be imported using the `import` statement. For example, to import the module `math`, which contains many standard mathematical functions, we can do:
```
import math
```
This includes the whole module and makes it available for use later in the program. For example, we can do:
```
import math
x = math.cos(2 * math.pi)
print(x)
```
Importing the whole module us often times unnecessary and can lead to longer loading time or increase the memory consumption. An alternative to the previous method, we can also choose to import only a few selected functions from a module by explicitly listing which ones we want to import:
```
from math import cos, pi
x = cos(2 * pi)
print(x)
```
It is also possible to give an imported module or symbol your own access name with the `as` additional:
```
import numpy as np
from math import pi as number_pi
x = np.rad2deg(number_pi)
print(x)
```
## Help and Descriptions
Using the function `help` we can get a description of almost all functions.
```
help(math.log)
math.log(10)
math.log(10, 2)
```
## Variables and types
### Symbol names
Variable names in Python can contain alphanumerical characters `a-z`, `A-Z`, `0-9` and some special characters such as `_`. Normal variable names must start with a letter.
By convention, variable names start with a lower-case letter, and Class names start with a capital letter.
In addition, there are a number of Python keywords that cannot be used as variable names. These keywords are:
and, as, assert, break, class, continue, def, del, elif, else, except, exec, finally, for, from, global, if, import, in, is, lambda, not, or, pass, print, raise, return, try, while, with, yield
### Assignment
The assignment operator in Python is `=`. Python is a dynamically typed language, so we do not need to specify the type of a variable when we create one.
Assigning a value to a new variable creates the variable:
```
# variable assignments
x = 1.0
```
Although not explicitly specified, a variable does have a type associated with it. The type is derived from the value it was assigned.
```
type(x)
```
If we assign a new value to a variable, its type can change.
```
x = 1
type(x)
```
If we try to use a variable that has not yet been defined we get an `NameError` (Note, that we will use in the notebooks `try/except` blocks to handle the exception, so the notebook doesn't stop. The code below will try to execute `print` function and if the `NameError` occurs the error message will be printed. Otherwise, an error will be raised. Later in this notebook you will learn more about exception handling.):
```
try:
print(y)
except(NameError) as err:
print("NameError", err)
else:
raise
```
### Fundamental types
```
# integers
x = 1
type(x)
# float
x = 1.0
type(x)
# boolean
b1 = True
b2 = False
type(b1)
# string
s = "hello world"
type(s)
```
## Operators and comparisons
Most operators and comparisons in Python work as one would expect:
* Arithmetic operators `+`, `-`, `*`, `/`, `**` power, `%` modulo
```
[1 + 2,
1 - 2,
1 * 2,
1 % 2]
```
In Python 2.7, what kind of division (`/`) will be executed, depends on the type of the numbers involved. If all numbers are integers, the division will be an integer division, otherwise, it will be a float division. In Python 3 this has been changed and fractions aren't lost when dividing integers (for integer division you can use another operator, `//`).
```
# In Python 3 these two operations will give the same result
# (in Python 2 the first one will be treated as an integer division).
print(1 / 2)
print(1 / 2.0)
# Note! The power operator in python isn't ^, but **
2 ** 2
```
* The boolean operators are spelled out as words `and`, `not`, `or`.
```
True and False
not False
True or False
```
* Comparison operators `>`, `<`, `>=` (greater or equal), `<=` (less or equal), `==` (equal), `!=` (not equal) and `is` (identical).
```
2 > 1, 2 < 1
2 > 2, 2 < 2
2 >= 2, 2 <= 2
# equal to
[1,2] == [1,2]
# not equal to
2 != 3
```
- boolean operator
```
x = True
y = False
print(not x)
print(x and y)
print(x or y)
```
- String comparison
```
"lo W" in "Hello World"
"x" not in "Hello World"
```
### Shortcut math operation and assignment
```
a = 2
a = a * 2
print(a)
```
The command `a = a * 2`, can be shortcut to `a *= 2`. This also works with `+=`, `-=` and `/=`.
```
b = 3
b *= 3
print(b)
```
## Strings, List and dictionaries
### Strings
Strings are the variable type that is used for storing text messages.
```
s = "Hello world"
type(s)
# length of the string: number of characters in string
len(s)
# replace a substring in a string with something else
s2 = s.replace("world", "test")
print(s2)
```
We can index a character in a string using `[]`:
```
s[0]
```
**Heads up MATLAB users:** Indexing start at 0!
We can extract a part of a string using the syntax `[start:stop]`, which extracts characters between index `start` and `stop`:
```
s[0:5]
```
If we omit either (or both) of `start` or `stop` from `[start:stop]`, the default is the beginning and the end of the string, respectively:
```
s[:5]
s[6:]
s[:]
```
We can also define the step size using the syntax `[start:end:step]` (the default value for `step` is 1, as we saw above):
```
s[::1]
s[::2]
```
This technique is called *slicing*.
#### String formatting examples
```
print("str1" + "str2" + "str3") # strings added with + are concatenated without space
print("str1" "str2" "str3") # The print function concatenates strings differently
print("str1", "str2", "str3") # depending on how the inputs are specified
print(("str1", "str2", "str3")) # See the three different outputs below
print("str1", 1.0, False) # The print function converts all arguments to strings
print("value = %f" %1.0) # we can use C-style string formatting
```
Python has two string formatting styles. An example of the old style is below, specifier `%.2f` transforms the input number into a string, that corresponds to a floating point number with 2 decimal places and the specifier `%d` transforms the input number into a string, corresponding to a decimal number.
```
s2 = "value1 = %.2f. value2 = %d" % (3.1415, 1.5)
print(s2)
```
The same string can be written using the new style string formatting.
```
s3 = 'value1 = {:.2f}, value2 = {}'.format(3.1415, 1.5)
print(s3)
print("Newlines are indicated by \nAnd tabs by \t.")
print(r"Newlines are indicated by \nAnd tabs by \t. Printed as rawstring")
print("Name: {}\nNumber: {}\nString: {}".format("Nipype", 3, 3 * "-"))
strString = """This is
a multiline
string."""
print(strString)
print("This {verb} a {noun}.".format(noun = "test", verb = "is"))
```
#### Single Quote
You can specify strings using single quotes such as `'Quote me on this'`.
All white space i.e. spaces and tabs, within the quotes, are preserved as-is.
#### Double Quotes
Strings in double quotes work exactly the same way as strings in single quotes. An example is `"What's your name?"`.
#### Triple Quotes
You can specify multi-line strings using triple quotes - (`"""` or `'''`). You can use single quotes and double quotes freely within the triple quotes. An example is:
```
'''This is a multi-line string. This is the first line.
This is the second line.
"What's your name?," I asked.
He said "Bond, James Bond."
'''
```
### List
Lists are very similar to strings, except that each element can be of any type.
The syntax for creating lists in Python is `[...]`:
```
l = [1,2,3,4]
print(type(l))
print(l)
```
We can use the same slicing techniques to manipulate lists as we could use on strings:
```
print(l)
print(l[1:3])
print(l[::2])
```
**Heads up MATLAB users:** Indexing starts at 0!
```
l[0]
```
Elements in a list do not all have to be of the same type:
```
l = [1, 'a', 1.0]
print(l)
```
Python lists can be inhomogeneous and arbitrarily nested:
```
nested_list = [1, [2, [3, [4, [5]]]]]
nested_list
```
Lists play a very important role in Python and are for example used in loops and other flow control structures (discussed below). There are a number of convenient functions for generating lists of various types, for example, the `range` function (note that in Python 3 `range` creates a generator, so you have to use `list` function to get a list):
```
start = 10
stop = 30
step = 2
list(range(start, stop, step))
# convert a string to a list by type casting:
print(s)
s2 = list(s)
s2
# sorting lists
s2.sort()
print(s2)
```
#### Adding, inserting, modifying, and removing elements from lists
```
# create a new empty list
l = []
# add an elements using `append`
l.append("A")
l.append("d")
l.append("d")
print(l)
```
We can modify lists by assigning new values to elements in the list. In technical jargon, lists are *mutable*.
```
l[1] = "p"
l[2] = "t"
print(l)
l[1:3] = ["s", "m"]
print(l)
```
Insert an element at an specific index using `insert`
```
l.insert(0, "i")
l.insert(1, "n")
l.insert(2, "s")
l.insert(3, "e")
l.insert(4, "r")
l.insert(5, "t")
print(l)
```
Remove first element with specific value using 'remove'
```
l.remove("A")
print(l)
```
Remove an element at a specific location using `del`:
```
del l[7]
del l[6]
print(l)
```
### Tuples
Tuples are like lists, except that they cannot be modified once created, that is they are *immutable*.
In Python, tuples are created using the syntax `(..., ..., ...)`, or even `..., ...`:
```
point = (10, 20)
print(type(point))
print(point)
```
If we try to assign a new value to an element in a tuple we get an error:
```
try:
point[0] = 20
except(TypeError) as er:
print("TypeError:", er)
else:
raise
```
### Dictionaries
Dictionaries are also like lists, except that each element is a key-value pair. The syntax for dictionaries is `{key1 : value1, ...}`:
```
params = {"parameter1" : 1.0,
"parameter2" : 2.0,
"parameter3" : 3.0,}
print(type(params))
print(params)
```
Dictionary entries can only be accessed by their key name.
```
params["parameter2"]
print("parameter1 = " + str(params["parameter1"]))
print("parameter2 = " + str(params["parameter2"]))
print("parameter3 = " + str(params["parameter3"]))
params["parameter1"] = "A"
params["parameter2"] = "B"
# add a new entry
params["parameter4"] = "D"
print("parameter1 = " + str(params["parameter1"]))
print("parameter2 = " + str(params["parameter2"]))
print("parameter3 = " + str(params["parameter3"]))
print("parameter4 = " + str(params["parameter4"]))
```
## Indentation
Whitespace is important in Python. Actually, whitespace at the beginning of the line is important. This is called indentation. Leading whitespace (spaces and tabs) at the beginning of the logical line is used to determine the indentation level of the logical line, which in turn is used to determine the grouping of statements.
This means that statements which go together must have the same indentation, for example:
```
i = 5
print('Value is ', i)
print('I repeat, the value is ', i)
```
Each such set of statements is called a block. We will see examples of how blocks are important later on.
One thing you should remember is that wrong indentation rises `IndentationError`.
## Control Flow
### Conditional statements: if, elif, else
The Python syntax for conditional execution of code use the keywords `if`, `elif` (else if), `else`:
```
statement1 = False
statement2 = False
if statement1:
print("statement1 is True")
elif statement2:
print("statement2 is True")
else:
print("statement1 and statement2 are False")
```
For the first time, here we encountered a peculiar and unusual aspect of the Python programming language: Program blocks are defined by their indentation level. In Python, the extent of a code block is defined by the indentation level (usually a tab or say four white spaces). This means that we have to be careful to indent our code correctly, or else we will get syntax errors.
**Examples:**
```
# Good indentation
statement1 = statement2 = True
if statement1:
if statement2:
print("both statement1 and statement2 are True")
# Bad indentation! This would lead to error
#if statement1:
# if statement2:
# print("both statement1 and statement2 are True") # this line is not properly indented
statement1 = False
if statement1:
print("printed if statement1 is True")
print("still inside the if block")
if statement1:
print("printed if statement1 is True")
print("now outside the if block")
```
## Loops
In Python, loops can be programmed in a number of different ways. The most common is the `for` loop, which is used together with iterable objects, such as lists. The basic syntax is:
## `for` loops
```
for x in [1,2,3]:
print(x),
```
The `for` loop iterates over the elements of the supplied list and executes the containing block once for each element. Any kind of list can be used in the `for` loop. For example:
```
for x in range(4): # by default range start at 0
print(x),
```
Note: `range(4)` does not include 4 !
```
for x in range(-3,3):
print(x),
for word in ["scientific", "computing", "with", "python"]:
print(word)
```
To iterate over key-value pairs of a dictionary:
```
for key, value in params.items():
print(key + " = " + str(value))
```
Sometimes it is useful to have access to the indices of the values when iterating over a list. We can use the `enumerate` function for this:
```
for idx, x in enumerate(range(-3,3)):
print(idx, x)
```
### `break`, `continue` and `pass`
To control the flow of a certain loop you can also use `break`, `continue` and `pass`.
```
rangelist = list(range(10))
print(list(rangelist))
for number in rangelist:
# Check if number is one of
# the numbers in the tuple.
if number in [4, 5, 7, 9]:
# "Break" terminates a for without
# executing the "else" clause.
break
else:
# "Continue" starts the next iteration
# of the loop. It's rather useless here,
# as it's the last statement of the loop.
print(number)
continue
else:
# The "else" clause is optional and is
# executed only if the loop didn't "break".
pass # Do nothing
```
**List comprehensions: Creating lists using `for` loops**:
A convenient and compact way to initialize lists:
```
l1 = [x**2 for x in range(0,5)]
print(l1)
```
**`while` loops**:
```
i = 0
while i < 5:
print(i)
i = i + 1
print("done")
```
Note that the `print "done"` statement is not part of the `while` loop body because of the difference in the indentation.
## Functions
A function in Python is defined using the keyword `def`, followed by a function name, a signature within parentheses `()`, and a colon `:`. The following code, with one additional level of indentation, is the function body.
```
def say_hello():
# block belonging to the function
print('hello world')
say_hello() # call the function
```
Following an example where we also feed two arguments into the function.
```
def print_max(a, b):
if a > b:
print( a, 'is maximum')
elif a == b:
print(a, 'is equal to', b)
else:
print(b, 'is maximum')
# directly pass literal values
print_max(3, 4)
x = 7
y = 7
# pass variables as arguments
print_max(x, y)
```
**Very important**: Variables inside a function are treated as local variables and therefore don't interfere with variables outside the scope of the function.
```
x = 50
def func(x):
print('x is', x)
x = 2
print('Changed local x to', x)
func(x)
print('x is still', x)
```
The local scope of a variable inside a function can be extended with the keyword `global`.
```
x = 50
def func():
global x
print('x is', x)
x = 2
print('Changed global x to', x)
func()
print('Value of x is', x)
```
Optionally, but highly recommended, we can define a so called "docstring", which is a description of the functions purpose and behavior. The docstring should follow directly after the function definition, before the code in the function body.
```
def func1(s):
"""
Print a string 's' and tell how many characters it has
"""
print(s + " has " + str(len(s)) + " characters")
help(func1)
func1("test")
```
Functions that return a value use the `return` keyword:
```
def square(x):
"""
Return the square of x.
"""
return x ** 2
square(4)
```
We can return multiple values from a function using tuples (see above):
```
def powers(x):
"""
Return a few powers of x.
"""
return x ** 2, x ** 3, x ** 4
powers(3)
```
And if we know that a function returns multiple outputs, we can store them directly in multiple variables.
```
x2, x3, x4 = powers(3)
print(x3)
```
### Default argument and keyword arguments
In a definition of a function, we can give default values to the arguments the function takes:
```
def myfunc(x, p=2, debug=False):
if debug:
print("evaluating myfunc for x = " + str(x) + " using exponent p = " + str(p))
return x**p
```
If we don't provide a value of the `debug` argument when calling the the function `myfunc` it defaults to the value provided in the function definition:
```
myfunc(5)
myfunc(5, debug=True)
```
If we explicitly list the name of the arguments in the function calls, they do not need to come in the same order as in the function definition. This is called *keyword* arguments and is often very useful in functions that take a lot of optional arguments.
```
myfunc(p=3, debug=True, x=7)
```
### `*args` and `*kwargs` parameters
Sometimes you might want to define a function that can take any number of parameters, i.e. variable number of arguments, this can be achieved by using one (`*args`) or two (`**kwargs`) asterisks in the function declaration. `*args` is used to pass a non-keyworded, variable-length argument list and the `**kwargs` is used to pass a keyworded, variable-length argument list.
```
def args_func(arg1, *args):
print("Formal arg:", arg1)
for a in args:
print("additioanl arg:", a)
args_func(1, "two", 3, [1, 2, 3])
def kwargs_func(arg1, **kwargs):
print("kwargs is now a dictionary...\nType: %s\nContent: %s\n" % (type(kwargs), kwargs))
print("Formal arg:", arg1)
for key in kwargs:
print("another keyword arg: %s: %s" % (key, kwargs[key]))
kwargs_func(arg1=1, myarg2="two", myarg3=3)
```
### Unnamed functions: lambda function
In Python we can also create unnamed functions, using the `lambda` keyword:
```
f1 = lambda x: x**2
# is equivalent to
def f2(x):
return x**2
f1(2), f2(2)
```
This technique is useful for example when we want to pass a simple function as an argument to another function, like this:
```
# map is a built-in python function
list(map(lambda x: x**2, range(-3,4)))
```
## Classes
Classes are the key features of object-oriented programming. A class is a structure for representing an object and the operations that can be performed on the object.
In Python, a class can contain *attributes* (variables) and *methods* (functions).
A class is defined almost like a function, but using the `class` keyword, and the class definition usually contains a number of class method definitions (a function in a class).
* Each class method should have an argument `self` as it first argument. This object is a self-reference.
* Some class method names have special meaning, for example:
* `__init__`: The name of the method that is invoked when the object is first created.
* `__str__` : A method that is invoked when a simple string representation of the class is needed, as for example when printed.
* There are many more, see http://docs.python.org/3.6/reference/datamodel.html#special-method-names
```
class Point:
"""
Simple class for representing a point in a Cartesian coordinate system.
"""
def __init__(self, x, y):
"""
Create a new Point at x, y.
"""
self.x = x
self.y = y
def translate(self, dx, dy):
"""
Translate the point by dx and dy in the x and y direction.
"""
self.x += dx
self.y += dy
def __str__(self):
return("Point at [%f, %f]" % (self.x, self.y))
```
To create a new instance of a class:
```
p1 = Point(0, 0) # this will invoke the __init__ method in the Point class
print(p1) # this will invoke the __str__ method
```
To invoke a class method in the class instance `p`:
```
p2 = Point(1, 1)
print(p2)
p2.translate(0.25, 1.5)
print(p2)
```
You can access any value of a class object directly, for example:
```
print(p1.x)
p1.x = 10
print(p1)
```
## Modules
One of the most important concepts in good programming is to reuse code and avoid repetitions.
The idea is to write functions and classes with a well-defined purpose and scope, and reuse these instead of repeating similar code in different part of a program (modular programming). The result is usually that readability and maintainability of a program are greatly improved. What this means in practice is that our programs have fewer bugs, are easier to extend and debug/troubleshoot.
Python supports modular programming at different levels. Functions and classes are examples of tools for low-level modular programming. Python modules are a higher-level modular programming construct, where we can collect related variables, functions, and classes in a module. A python module is defined in a python file (with file-ending `.py`), and it can be made accessible to other Python modules and programs using the `import` statement.
Consider the following example: the file `mymodule.py` contains simple example implementations of a variable, function and a class:
```
%%file mymodule.py
"""
Example of a python module. Contains a variable called my_variable,
a function called my_function, and a class called MyClass.
"""
my_variable = 0
def my_function():
"""
Example function
"""
return my_variable
class MyClass:
"""
Example class.
"""
def __init__(self):
self.variable = my_variable
def set_variable(self, new_value):
"""
Set self.variable to a new value
"""
self.variable = new_value
def get_variable(self):
return self.variable
```
**Note:** `%%file` is called a cell-magic function and creates a file that has the following lines as content.
We can import the module `mymodule` into our Python program using `import`:
```
import mymodule
```
Use `help(module)` to get a summary of what the module provides:
```
help(mymodule)
mymodule.my_variable
mymodule.my_function()
my_class = mymodule.MyClass()
my_class.set_variable(10)
my_class.get_variable()
```
If we make changes to the code in `mymodule.py`, we need to reload it using `reload`:
```
from importlib import reload
reload(mymodule)
```
## Exceptions
In Python errors are managed with a special language construct called "Exceptions". When errors occur exceptions can be raised, which interrupts the normal program flow and fallback to somewhere else in the code where the closest try-except statement is defined.
To generate an exception we can use the `raise` statement, which takes an argument that must be an instance of the class `BaseExpection` or a class derived from it.
```
try:
raise Exception("description of the error")
except(Exception) as err:
print ("Exception:", err)
```
A typical use of exceptions is to abort functions when some error condition occurs, for example:
def my_function(arguments):
if not verify(arguments):
raise Exception("Invalid arguments")
# rest of the code goes here
To gracefully catch errors that are generated by functions and class methods, or by the Python interpreter itself, use the `try` and `except` statements:
try:
# normal code goes here
except:
# code for error handling goes here
# this code is not executed unless the code
# above generated an error
For example:
```
try:
print("test")
# generate an error: the variable test is not defined
print(test)
except:
print("Caught an exception")
```
To get information about the error, we can access the `Exception` class instance that describes the exception by using for example:
except Exception as e:
```
try:
print("test")
# generate an error: the variable test is not defined
print(test)
except Exception as e:
print("Caught an exception:" + str(e))
finally:
print("This block is executed after the try- and except-block.")
def some_function():
try:
# Division by zero raises an exception
10 / 0
except ZeroDivisionError:
print("Oops, invalid.")
else:
# Exception didn't occur, we're good.
pass
finally:
# This is executed after the code block is run
# and all exceptions have been handled, even
# if a new exception is raised while handling.
print("We're done with that.")
some_function()
```
You will see more exception handling examples in this and other notebooks.
## File I/O
This section should give you a basic knowledge about how to read and write CSV or TXT files. First, let us create a CSV and TXT file about demographic information of 10 subjects (experiment_id, subject_id, gender, age).
```
%%file demographics.csv
ds102,sub001,F,21.94
ds102,sub002,M,22.79
ds102,sub003,M,19.65
ds102,sub004,M,25.98
ds102,sub005,M,23.24
ds102,sub006,M,23.27
ds102,sub007,D,34.72
ds102,sub008,D,22.22
ds102,sub009,M,22.7
ds102,sub010,D,25.24
%%file demographics.txt
ds102 sub001 F 21.94
ds102 sub002 M 22.79
ds102 sub003 M 19.65
ds102 sub004 M 25.98
ds102 sub005 M 23.24
ds102 sub006 M 23.27
ds102 sub007 D 34.72
ds102 sub008 D 22.22
ds102 sub009 M 22.7
ds102 sub010 D 25.24
```
### Reading CSV files
Parsing comma-separated-values (CSV) files is a common task. There are many tools available in Python to deal with this. Let's start by using the built-in `csv` module.
```
import csv
```
Before you can read or write any kind of file, you first have to open the file and go through its content with a reader function or write the output line by line with a write function.
```
f = open('demographics.csv','r') # open the file with reading rights = 'r'
data = [i for i in csv.reader(f) ] # go through file and read each line
f.close() # close the file again
for line in data:
print(line)
```
### Writing CSV files
Now, we want to write the same data without the first experiment_id column in CSV format to a csv-file. First, let's delete the first column in the dataset.
```
data_new = [line[1:] for line in data]
for line in data_new:
print(line)
```
Now, we first have to open a file again, but this time with writing permissions = `'w'`. After it, we can go through the file and write each line to the new csv-file.
```
f = open('demographics_new.csv','w') # open a file with writing rights = 'w'
fw = csv.writer(f) # create csv writer
fw.writerows(data_new) # write content to file
f.close() # close file
```
Lets now check the content of `demographics_new.csv`.
```
!cat demographics_new.csv
```
### Reading TXT files
The reading of txt files is quite similar to the reading of csv-files. The only difference is in the name of the reading function and the formatting that has to be applied to the input or output.
```
f = open('demographics.txt','r') # open file with reading rights = 'r'
# go through file and trim the new line '\n' at the end
datatxt = [i.splitlines() for i in f.readlines()]
# go through data and split elements in line by tabulators '\t'
datatxt = [i[0].split('\t') for i in datatxt]
f.close() # close file again
for line in datatxt:
print(line)
```
### Writing TXT files
The writing of txt files is as follows:
```
f = open('demograhics_new.txt', 'w') # open file with writing rights = 'w'
datatxt_new = [line[1:] for line in datatxt] # delete first column of array
# Go through datatxt array and write each line with specific format to file
for line in datatxt_new:
f.write("%s\t%s\t%s\n"%(line[0],line[1],line[2]))
f.close() # close file
```
### `with open`
The previous methods to open or write a file always required that you also close the file again with the `close()` function. If you don't want to worry about this, you can also use the `with open` approach. For example:
```
with open('demographics.txt','r') as f:
datatxt = [i.splitlines() for i in f.readlines()]
datatxt = [i[0].split('\t') for i in datatxt]
for line in datatxt:
print(line)
```
## File modes
* Read-only: `r`
* Write-only: `w` (Create a new file or overwrite existing file)
* Append a file: `a`
* Read and Write: `r+`
* Binary mode: `b` (Use for binary files, especially on Windows)
| github_jupyter |
## BERT as a service
This notebook demonstrates how to build a complete machine learning pipeline using TensorFlow Extended ([TFX](https://www.tensorflow.org/tfx)) to serve a BERT model for text sentiment classification.
Notes:
- Data: IMDB Movie Reviews (5000 samples) [original source](https://ai.stanford.edu/~amaas/data/sentiment/)
- Model: BERT base uncased (english) from [HuggingFace](https://huggingface.co/bert-base-uncased)
- Processor: BERT uncased (english seq_length=128) from [TF HUB](https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3)
You can check references, additional information, and resources at the [GitHub repository](https://github.com/dimitreOliveira/bert-as-a-service_TFX).
## Setup
```
# Use the latest version of pip.
!pip install --upgrade pip
# Install tfx and kfp Python packages.
!pip install -q --upgrade tfx[kfp]==1.0.0rc1
```
### Import packages
```
import os
import sys
import urllib
import tfx
import tensorflow as tf
print(f'TensorFlow version: {tf.__version__}')
print(f'TFX version: {tfx.__version__}')
```
[Kubeflow Pipelines](https://www.kubeflow.org/docs/pipelines/overview/pipelines-overview/) environment variables
```
# Read GCP project id from env.
shell_output=!gcloud config list --format 'value(core.project)' 2>/dev/null
GOOGLE_CLOUD_PROJECT=shell_output[0]
%env GOOGLE_CLOUD_PROJECT={GOOGLE_CLOUD_PROJECT}
print("GCP project ID:" + GOOGLE_CLOUD_PROJECT)
```
KFP endpoint
"AI Platform > Pipeline > pipeline instance `settings`"
**ENDPOINT should contain only the hostname part of the URL.** For example, if the URL of the KFP dashboard is `https://1e9deb537390ca22-dot-asia-east1.pipelines.googleusercontent.com/#/start`, ENDPOINT value becomes `1e9deb537390ca22-dot-asia-east1.pipelines.googleusercontent.com`.
```
# This refers to the KFP cluster endpoint
ENDPOINT='4c4887d40ceb4e53-dot-us-central1.pipelines.googleusercontent.com'
if not ENDPOINT:
from absl import logging
logging.error('Set your ENDPOINT in this cell.')
```
Set the image name as `tfx-pipeline` under the current GCP project.
```
# Docker image name for the pipeline image.
CUSTOM_TFX_IMAGE='gcr.io/' + GOOGLE_CLOUD_PROJECT + '/tfx-pipeline'
print(CUSTOM_TFX_IMAGE)
PIPELINE_NAME="my_pipeline"
PROJECT_DIR=os.path.join(os.path.expanduser("~"), "imported", PIPELINE_NAME)
```
Change the working directory context in this notebook to the project directory.
```
%cd {PROJECT_DIR}
```
## Source files
Here is brief introduction to each of the Python files.
- `data` - This directory contains the datasets.
- `models` - This directory contains ML model definitions.
- `bert_aas_utils.py` — defines utility functions for the pipeline
- `pipeline` - This directory contains the definition of the pipeline
- `configs.py` — defines common constants for pipeline runners
- `pipeline.py` — defines TFX components and a pipeline
- `local_runner.py`, `kubeflow_runner.py` — define runners for each orchestration engine
### Download data
```
data_csv_url = 'https://raw.githubusercontent.com/dimitreOliveira/bert-as-a-service_TFX/main/Data/IMDB_5k_dataset.csv'
data_csv_filename = 'IMDB_dataset.csv'
_data_dir = 'data/'
if not os.path.exists(_data_dir):
os.makedirs(_data_dir)
# Download data
urllib.request.urlretrieve(data_csv_url, f'{_data_dir}{data_csv_filename}')
```
Upload our sample data to GCS bucket so that we can use it in our pipeline later.
```
!gsutil cp data/IMDB_dataset.csv gs://{GOOGLE_CLOUD_PROJECT}-kubeflowpipelines-default/bert-aas/data/IMDB_dataset.csv
```
## Create TFX pipeline
Components in the TFX pipeline will generate outputs for each run as [ML Metadata Artifacts](https://www.tensorflow.org/tfx/guide/mlmd), and they need to be stored somewhere. You can use any storage which the KFP cluster can access, and for this example we will use Google Cloud Storage (GCS). A default GCS bucket should have been created automatically. Its name will be `<your-project-id>-kubeflowpipelines-default`.
Let's create a TFX pipeline using the `tfx pipeline create` command.
>Note: When creating a pipeline for KFP, we need a container image which will be used to run our pipeline. And `skaffold` will build the image for us. Because skaffold pulls base images from the docker hub, it will take 5~10 minutes when we build the image for the first time, but it will take much less time from the second build.
```
!tfx pipeline create \
--pipeline-path=kubeflow_runner.py \
--endpoint={ENDPOINT} \
--build-image
```
While creating a pipeline, `Dockerfile` will be generated to build a Docker image. Don't forget to add it to the source control system (for example, git) along with other source files.
NOTE: `kubeflow` will be automatically selected as an orchestration engine if `airflow` is not installed and `--engine` is not specified.
```
!tfx run create \
--pipeline-name={PIPELINE_NAME} \
--endpoint={ENDPOINT}
```
Or, you can also run the pipeline in the KFP Dashboard. The new execution run will be listed under Experiments in the KFP Dashboard. Clicking into the experiment will allow you to monitor progress and visualize the artifacts created during the execution run.
However, we recommend visiting the KFP Dashboard. You can access the KFP Dashboard from the Cloud AI Platform Pipelines menu in Google Cloud Console. Once you visit the dashboard, you will be able to find the pipeline, and access a wealth of information about the pipeline.
For example, you can find your runs under the *Experiments* menu, and when you open your execution run under Experiments you can find all your artifacts from the pipeline under *Artifacts* menu.
>Note: If your pipeline run fails, you can see detailed logs for each TFX component in the Experiments tab in the KFP Dashboard.
One of the major sources of failure is permission related problems. Please make sure your KFP cluster has permissions to access Google Cloud APIs. This can be configured [when you create a KFP cluster in GCP](https://cloud.google.com/ai-platform/pipelines/docs/setting-up), or see [Troubleshooting document in GCP](https://cloud.google.com/ai-platform/pipelines/docs/troubleshooting).
### In case the TFX pipeline needs to be updated
```
# Update the pipeline
!tfx pipeline update \
--pipeline-path=kubeflow_runner.py \
--endpoint={ENDPOINT} \
# --build-image
# You can run the pipeline the same way.
!tfx run create \
--pipeline-name {PIPELINE_NAME} \
--endpoint={ENDPOINT}
```
### Check pipeline outputs
Visit the KFP dashboard to find pipeline outputs in the page for your pipeline run. Click the *Experiments* tab on the left, and *All runs* in the Experiments page. You should be able to find the latest run under the name of your pipeline.
**NOTE:** If we changed anything in the model code, we have to rebuild the
container image, too. We can trigger rebuild using `--build-image` flag in the
`pipeline update` command.
**NOTE:** You might have noticed that every time we create a pipeline run, every component runs again and again even though the input and the parameters were not changed.
It is waste of time and resources, and you can skip those executions with pipeline caching. You can enable caching by specifying `enable_cache=True` for the `Pipeline` object in `pipeline.py`.
| github_jupyter |
## Dependencies
```
import os
import cv2
import shutil
import random
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from tensorflow import set_random_seed
from sklearn.utils import class_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, cohen_kappa_score
from keras import backend as K
from keras.models import Model
from keras.utils import to_categorical
from keras import optimizers, applications
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, Callback, LearningRateScheduler
from keras.layers import Dense, Dropout, GlobalAveragePooling2D, Input
# Set seeds to make the experiment more reproducible.
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(0)
seed = 0
seed_everything(seed)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
```
## Load data
```
hold_out_set = pd.read_csv('../input/aptos-data-split/hold-out.csv')
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
test = pd.read_csv('../input/aptos2019-blindness-detection/test.csv')
print('Number of train samples: ', X_train.shape[0])
print('Number of validation samples: ', X_val.shape[0])
print('Number of test samples: ', test.shape[0])
# Preprocecss data
X_train["id_code"] = X_train["id_code"].apply(lambda x: x + ".png")
X_val["id_code"] = X_val["id_code"].apply(lambda x: x + ".png")
test["id_code"] = test["id_code"].apply(lambda x: x + ".png")
X_train['diagnosis'] = X_train['diagnosis'].astype('str')
X_val['diagnosis'] = X_val['diagnosis'].astype('str')
display(X_train.head())
```
# Model parameters
```
# Model parameters
N_CLASSES = X_train['diagnosis'].nunique()
BATCH_SIZE = 16
EPOCHS = 40
WARMUP_EPOCHS = 5
LEARNING_RATE = 1e-4
WARMUP_LEARNING_RATE = 1e-3
HEIGHT = 320
WIDTH = 320
CHANNELS = 3
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
def kappa(y_true, y_pred, n_classes=5):
y_trues = K.cast(K.argmax(y_true), K.floatx())
y_preds = K.cast(K.argmax(y_pred), K.floatx())
n_samples = K.cast(K.shape(y_true)[0], K.floatx())
distance = K.sum(K.abs(y_trues - y_preds))
max_distance = n_classes - 1
kappa_score = 1 - ((distance**2) / (n_samples * (max_distance**2)))
return kappa_score
def step_decay(epoch):
lrate = 30e-5
if epoch > 3:
lrate = 15e-5
if epoch > 7:
lrate = 7.5e-5
if epoch > 11:
lrate = 3e-5
if epoch > 15:
lrate = 1e-5
return lrate
def focal_loss(y_true, y_pred):
gamma = 2.0
epsilon = K.epsilon()
pt = y_pred * y_true + (1-y_pred) * (1-y_true)
pt = K.clip(pt, epsilon, 1-epsilon)
CE = -K.log(pt)
FL = K.pow(1-pt, gamma) * CE
loss = K.sum(FL, axis=1)
return loss
```
# Pre-procecess images
```
train_base_path = '../input/aptos2019-blindness-detection/train_images/'
test_base_path = '../input/aptos2019-blindness-detection/test_images/'
train_dest_path = 'base_dir/train_images/'
validation_dest_path = 'base_dir/validation_images/'
test_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
# Creating train, validation and test directories
os.makedirs(train_dest_path)
os.makedirs(validation_dest_path)
os.makedirs(test_dest_path)
def crop_image(img, tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
img = np.stack([img1,img2,img3],axis=-1)
return img
def circle_crop(img):
img = crop_image(img)
height, width, depth = img.shape
largest_side = np.max((height, width))
img = cv2.resize(img, (largest_side, largest_side))
height, width, depth = img.shape
x = width//2
y = height//2
r = np.amin((x, y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x, y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image(img)
return img
def preprocess_image(base_path, save_path, image_id, HEIGHT, WIDTH, sigmaX=10):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = circle_crop(image)
image = cv2.resize(image, (HEIGHT, WIDTH))
image = cv2.addWeighted(image, 4, cv2.GaussianBlur(image, (0,0), sigmaX), -4 , 128)
cv2.imwrite(save_path + image_id, image)
# Pre-procecss train set
for i, image_id in enumerate(X_train['id_code']):
preprocess_image(train_base_path, train_dest_path, image_id, HEIGHT, WIDTH)
# Pre-procecss validation set
for i, image_id in enumerate(X_val['id_code']):
preprocess_image(train_base_path, validation_dest_path, image_id, HEIGHT, WIDTH)
# Pre-procecss test set
for i, image_id in enumerate(test['id_code']):
preprocess_image(test_base_path, test_dest_path, image_id, HEIGHT, WIDTH)
```
# Data generator
```
datagen=ImageDataGenerator(rescale=1./255,
rotation_range=360,
horizontal_flip=True,
vertical_flip=True)
train_generator=datagen.flow_from_dataframe(
dataframe=X_train,
directory=train_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="categorical",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
valid_generator=datagen.flow_from_dataframe(
dataframe=X_val,
directory=validation_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="categorical",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
test_generator=datagen.flow_from_dataframe(
dataframe=test,
directory=test_dest_path,
x_col="id_code",
batch_size=1,
class_mode=None,
shuffle=False,
target_size=(HEIGHT, WIDTH),
seed=seed)
```
# Model
```
def create_model(input_shape, n_out):
input_tensor = Input(shape=input_shape)
base_model = applications.NASNetLarge(weights=None,
include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/keras-notop/NASNet-large-no-top.h5')
x = GlobalAveragePooling2D()(base_model.output)
x = Dropout(0.5)(x)
x = Dense(2048, activation='relu')(x)
x = Dropout(0.5)(x)
final_output = Dense(n_out, activation='softmax', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
```
# Train top layers
```
model = create_model(input_shape=(HEIGHT, WIDTH, CHANNELS), n_out=N_CLASSES)
for layer in model.layers:
layer.trainable = False
for i in range(-5, 0):
model.layers[i].trainable = True
class_weights = class_weight.compute_class_weight('balanced', np.unique(X_train['diagnosis'].astype('int').values), X_train['diagnosis'].astype('int').values)
metric_list = ["accuracy", kappa]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history_warmup = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
class_weight=class_weights,
verbose=1).history
```
# Fine-tune the complete model (1st step)
```
for layer in model.layers:
layer.trainable = True
# lrstep = LearningRateScheduler(step_decay)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
callback_list = [es, rlrop]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metric_list)
model.summary()
history_finetunning = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=int(EPOCHS*0.8),
callbacks=callback_list,
class_weight=class_weights,
verbose=1).history
```
# Fine-tune the complete model (2nd step)
```
optimizer = optimizers.SGD(lr=LEARNING_RATE, momentum=0.9, nesterov=True)
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=metric_list)
history_finetunning_2 = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=int(EPOCHS*0.2),
callbacks=callback_list,
class_weight=class_weights,
verbose=1).history
```
# Model loss graph
```
history = {'loss': history_finetunning['loss'] + history_finetunning_2['loss'],
'val_loss': history_finetunning['val_loss'] + history_finetunning_2['val_loss'],
'acc': history_finetunning['acc'] + history_finetunning_2['acc'],
'val_acc': history_finetunning['val_acc'] + history_finetunning_2['val_acc'],
'kappa': history_finetunning['kappa'] + history_finetunning_2['kappa'],
'val_kappa': history_finetunning['val_kappa'] + history_finetunning_2['val_kappa']}
sns.set_style("whitegrid")
fig, (ax1, ax2, ax3) = plt.subplots(3, 1, sharex='col', figsize=(20, 18))
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['acc'], label='Train accuracy')
ax2.plot(history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
ax3.plot(history['kappa'], label='Train kappa')
ax3.plot(history['val_kappa'], label='Validation kappa')
ax3.legend(loc='best')
ax3.set_title('Kappa')
plt.xlabel('Epochs')
sns.despine()
plt.show()
# Create empty arays to keep the predictions and labels
lastFullTrainPred = np.empty((0, N_CLASSES))
lastFullTrainLabels = np.empty((0, N_CLASSES))
lastFullValPred = np.empty((0, N_CLASSES))
lastFullValLabels = np.empty((0, N_CLASSES))
# Add train predictions and labels
for i in range(STEP_SIZE_TRAIN+1):
im, lbl = next(train_generator)
scores = model.predict(im, batch_size=train_generator.batch_size)
lastFullTrainPred = np.append(lastFullTrainPred, scores, axis=0)
lastFullTrainLabels = np.append(lastFullTrainLabels, lbl, axis=0)
# Add validation predictions and labels
for i in range(STEP_SIZE_VALID+1):
im, lbl = next(valid_generator)
scores = model.predict(im, batch_size=valid_generator.batch_size)
lastFullValPred = np.append(lastFullValPred, scores, axis=0)
lastFullValLabels = np.append(lastFullValLabels, lbl, axis=0)
lastFullComPred = np.concatenate((lastFullTrainPred, lastFullValPred))
lastFullComLabels = np.concatenate((lastFullTrainLabels, lastFullValLabels))
train_preds = [np.argmax(pred) for pred in lastFullTrainPred]
train_labels = [np.argmax(label) for label in lastFullTrainLabels]
validation_preds = [np.argmax(pred) for pred in lastFullValPred]
validation_labels = [np.argmax(label) for label in lastFullValLabels]
complete_labels = [np.argmax(label) for label in lastFullComLabels]
```
# Threshold optimization
```
def find_best_fixed_threshold(preds, targs, do_plot=True):
best_thr_list = [0 for i in range(preds.shape[1])]
for index in range(1, preds.shape[1]):
score = []
thrs = np.arange(0, 1, 0.01)
for thr in thrs:
preds_thr = [index if x[index] > thr else np.argmax(x) for x in preds]
score.append(cohen_kappa_score(targs, preds_thr))
score = np.array(score)
pm = score.argmax()
best_thr, best_score = thrs[pm], score[pm].item()
best_thr_list[index] = best_thr
print('Label %s: thr=%.3f, Kappa=%.3f' % (index, best_thr, best_score))
if do_plot:
plt.plot(thrs, score)
plt.vlines(x=best_thr, ymin=score.min(), ymax=score.max())
plt.text(best_thr+0.03, best_score-0.01, ('Kappa[%s]=%.3f' % (index, best_score)), fontsize=14);
plt.show()
return best_thr_list
threshold_list = find_best_fixed_threshold(lastFullComPred, complete_labels, do_plot=True)
threshold_list[0] = 0 # In last instance assign label 0
# Apply optimized thresholds to the train predictions
train_preds_opt = [0 for i in range(lastFullTrainPred.shape[0])]
for idx, thr in enumerate(threshold_list):
for idx2, pred in enumerate(lastFullTrainPred):
if pred[idx] > thr:
train_preds_opt[idx2] = idx
# Apply optimized thresholds to the validation predictions
validation_preds_opt = [0 for i in range(lastFullValPred.shape[0])]
for idx, thr in enumerate(threshold_list):
for idx2, pred in enumerate(lastFullValPred):
if pred[idx] > thr:
validation_preds_opt[idx2] = idx
index_order = [0, 2, 1, 4, 3]
# Apply optimized thresholds to the train predictions by class distribution
train_preds_opt2 = [0 for i in range(lastFullTrainPred.shape[0])]
for idx in index_order:
thr = threshold_list[idx]
for idx2, pred in enumerate(lastFullTrainPred):
if pred[idx] > thr:
train_preds_opt2[idx2] = idx
# Apply optimized thresholds to the validation predictions by class distribution
validation_preds_opt2 = [0 for i in range(lastFullValPred.shape[0])]
for idx in index_order:
thr = threshold_list[idx]
for idx2, pred in enumerate(lastFullValPred):
if pred[idx] > thr:
validation_preds_opt2[idx2] = idx
```
# Model Evaluation
## Confusion Matrix
### Original thresholds
```
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
def plot_confusion_matrix(train, validation, labels=labels):
train_labels, train_preds = train
validation_labels, validation_preds = validation
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 7))
train_cnf_matrix = confusion_matrix(train_labels, train_preds)
validation_cnf_matrix = confusion_matrix(validation_labels, validation_preds)
train_cnf_matrix_norm = train_cnf_matrix.astype('float') / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
validation_cnf_matrix_norm = validation_cnf_matrix.astype('float') / validation_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=labels, columns=labels)
validation_df_cm = pd.DataFrame(validation_cnf_matrix_norm, index=labels, columns=labels)
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues",ax=ax1).set_title('Train')
sns.heatmap(validation_df_cm, annot=True, fmt='.2f', cmap=sns.cubehelix_palette(8),ax=ax2).set_title('Validation')
plt.show()
plot_confusion_matrix((train_labels, train_preds), (validation_labels, validation_preds))
```
### Optimized thresholds
```
plot_confusion_matrix((train_labels, train_preds_opt), (validation_labels, validation_preds_opt))
```
### Optimized thresholds by class
```
plot_confusion_matrix((train_labels, train_preds_opt2), (validation_labels, validation_preds_opt2))
```
## Quadratic Weighted Kappa
```
def evaluate_model(train, validation):
train_labels, train_preds = train
validation_labels, validation_preds = validation
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds, train_labels, weights='quadratic'))
print("Validation Cohen Kappa score: %.3f" % cohen_kappa_score(validation_preds, validation_labels, weights='quadratic'))
print("Complete set Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds+validation_preds, train_labels+validation_labels, weights='quadratic'))
print(" Original thresholds")
evaluate_model((train_preds, train_labels), (validation_preds, validation_labels))
print(" Optimized thresholds")
evaluate_model((train_preds_opt, train_labels), (validation_preds_opt, validation_labels))
print(" Optimized thresholds by class")
evaluate_model((train_preds_opt2, train_labels), (validation_preds_opt2, validation_labels))
```
## Apply model to test set and output predictions
```
def apply_tta(model, generator, steps=10):
step_size = generator.n//generator.batch_size
preds_tta = []
for i in range(steps):
generator.reset()
preds = model.predict_generator(generator, steps=step_size)
preds_tta.append(preds)
return np.mean(preds_tta, axis=0)
preds = apply_tta(model, test_generator)
predictions = np.argmax(preds, axis=1)
predictions_opt = [0 for i in range(preds.shape[0])]
for idx, thr in enumerate(threshold_list):
for idx2, pred in enumerate(preds):
if pred[idx] > thr:
predictions_opt[idx2] = idx
predictions_opt2 = [0 for i in range(preds.shape[0])]
for idx in index_order:
thr = threshold_list[idx]
for idx2, pred in enumerate(preds):
if pred[idx] > thr:
predictions_opt2[idx2] = idx
results = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions})
results['id_code'] = results['id_code'].map(lambda x: str(x)[:-4])
results_opt = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions_opt})
results_opt['id_code'] = results_opt['id_code'].map(lambda x: str(x)[:-4])
results_opt2 = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions_opt2})
results_opt2['id_code'] = results_opt2['id_code'].map(lambda x: str(x)[:-4])
# Cleaning created directories
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
```
# Predictions class distribution
```
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, sharex='col', figsize=(24, 8.7))
sns.countplot(x="diagnosis", data=results, palette="GnBu_d", ax=ax1).set_title('Test')
sns.countplot(x="diagnosis", data=results_opt, palette="GnBu_d", ax=ax2).set_title('Test optimized')
sns.countplot(x="diagnosis", data=results_opt2, palette="GnBu_d", ax=ax3).set_title('Test optimized by class')
sns.despine()
plt.show()
val_kappa = cohen_kappa_score(validation_preds, validation_labels, weights='quadratic')
val_opt_kappa = cohen_kappa_score(validation_preds_opt, validation_labels, weights='quadratic')
val_opt_kappa2 = cohen_kappa_score(validation_preds_opt2, validation_labels, weights='quadratic')
results_name = 'submission.csv'
results_opt_name = 'submission_opt.csv'
results_opt2_name = 'submission_opt2.csv'
# if (val_kappa > val_opt_kappa) and (val_kappa > val_opt_kappa2):
# results_name = 'submission.csv'
# results_opt_name = 'submission_opt.csv'
# results_opt2_name = 'submission_opt2.csv'
# elif (val_opt_kappa > val_kappa) and (val_opt_kappa > val_opt_kappa2):
# results_name = 'submission_norm.csv'
# results_opt_name = 'submission.csv'
# results_opt2_name = 'submission_opt2.csv'
# else:
# results_name = 'submission_norm.csv'
# results_opt_name = 'submission_opt.csv'
# results_opt2_name = 'submission.csv'
results.to_csv(results_name, index=False)
display(results.head())
results_opt.to_csv(results_opt_name, index=False)
display(results_opt.head())
results_opt2.to_csv(results_opt2_name, index=False)
display(results_opt2.head())
```
| github_jupyter |
<a href="https://colab.research.google.com/github/AbhilashPal/BanglaOCR/blob/master/BanglaOCR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import files
files.upload()
!pip install bijoy2unicode
!ls
import random
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from bijoy2unicode import converter
test = converter.Unicode()
FSNew = np.load("FinalSolution.npy",allow_pickle = True).item()
x_offset=45
y_offset=5
FSFinal = []
for i in range(len(FSNew['x'])):
p = np.ones((40,300))
s_img = FSNew['x'][i]
p[y_offset:y_offset+s_img.shape[0], x_offset:x_offset+s_img.shape[1]] = s_img
FSFinal.append(p)
ra = []
for i in range(2):
ra.append(random.randint(0,300))
for i in ra:
plt.imshow(FSNew['x'][i])
# print(FSNew['x'][i])
plt.show()
print(test.convertBijoyToUnicode(FSNew['y'][i]+" "))
x_offset=45
y_offset=5
FSFinal = []
for i in range(len(FSNew['x'])):
p = np.ones((40,300))
s_img = FSNew['x'][i]
x_offset = int(150 - (s_img.shape[1]/2 ))
p[y_offset:y_offset+s_img.shape[0], x_offset:x_offset+s_img.shape[1]] = s_img
FSFinal.append(p)
FSNew['x'] = FSFinal
ra = []
for i in range(2):
ra.append(random.randint(0,3000))
for i in ra:
plt.imshow(FSNew['x'][i])
plt.show()
print(test.convertBijoyToUnicode(FSNew['y'][i]+" "))
print(FSNew['y'][i])
len(FSNew['x'])
tlist = FSNew['y']
vocab = list(set(" ".join(tlist)))
c2i = {vocab[i]:i for i in range(len(vocab))}
i2c = {i:vocab[i] for i in range(len(vocab))}
def char2id(s):
r = []
for c in s:
r.append(c2i[c])
return(r)
def id2char(i):
r = []
for c in i:
r.append(i2c[c])
return(r)
def conv2ctc(d):
res = {"the_input":d['x']}
res['the_labels'] = []
res['input_length'] = []
res['label_length'] = []
res['source_str'] = []
ylist = d['y']
for i in ylist:
r = char2id(i)
ll = [len(r)]
labels = [len(vocab)+1] * 30
labels[:len(r)] = r
inp_leng = [30]
res['the_labels'].append(labels)
res['input_length'].append(inp_leng)
res['label_length'].append(ll)
res['source_str'].append(i)
return res
ctcdata = conv2ctc(FSNew)
def ttsplit(d):
X_train, X_test, y_train, y_test = train_test_split(
d['x'], d['y'], test_size=0.2, random_state=42)
return(X_train,X_test,y_train,y_test)
X_train, X_test, y_train, y_test = ttsplit(FSNew)
FSTrain = {"x":X_train,"y":y_train}
FSTest = {"x":X_test,"y":y_test}
ra = []
for i in range(2):
ra.append(random.randint(0,len(FSTrain['x'])))
for i in ra:
plt.imshow(FSTrain['x'][i])
plt.show()
print(test.convertBijoyToUnicode(FSTrain['y'][i]+" "))
print(FSTrain['y'][i])
ctcTrain = conv2ctc(FSTrain)
ctcTest = conv2ctc(FSTest)
def createbatches(d,batchsize=32):
n = batchsize
batched = {}
for j in d:
final = [d[j][i * n:(i + 1) * n] for i in range((len(d[j]) + n - 1) // n )]
final.pop()
batched[j] = final
return batched
btTest = createbatches(ctcTest)
btTrain = createbatches(ctcTrain)
def ffinalbtc(btc):
finalbtc = []
for i in range(len(btc["the_input"])):
t = (
{
"the_input" : np.array(btc['the_input'][i]).reshape(32,300,40,1) ,
"the_labels" : np.array(btc["the_labels"][0]),
"input_length" : np.array(btc["input_length"][i]),
"label_length" : np.array(btc["label_length"][i]),
"source_str" : btc["source_str"][i]
},
{
'ctc':np.array([0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.,
0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.])
}
)
finalbtc.append(t)
return finalbtc
finalTrain = ffinalbtc(btTrain)
finalTest = ffinalbtc(btTest)
len(finalTrain)
finalTrain[0]
import os
import itertools
import codecs
import re
import datetime
from keras.models import load_model
import editdistance
import numpy as np
from scipy import ndimage
# import pylab
import matplotlib.pyplot as plt
from keras import backend as K
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.layers import Input, Dense, Activation
from keras.layers import Reshape, Lambda
from keras.layers.merge import add, concatenate
from keras.models import Model
from keras.layers.recurrent import GRU
from keras.optimizers import SGD
from keras.utils.data_utils import get_file
from keras.preprocessing import image
import keras.callbacks
m = FSNew['x']
n = FSNew['y']
```
Converting strings to one hot encoding.
```
def data_gen():
while True:
for i in finalTrain:
yield i
def val_gen():
while True:
for x in finalTest:
yield x
# the actual loss calc occurs here despite it not being
# an internal Keras loss function
def ctc_lambda_func(args):
y_pred, labels, input_length, label_length = args
# the 2 is critical here since the first couple outputs of the RNN
# tend to be garbage:
y_pred = y_pred[:, 2:, :]
return K.ctc_batch_cost(labels, y_pred, input_length, label_length)
# For a real OCR application, this should be beam search with a dictionary
# and language model. For this example, best path is sufficient.
def decode_batch(test_func, word_batch):
out = test_func([word_batch])[0]
ret = []
for j in range(out.shape[0]):
out_best = list(np.argmax(out[j, 2:], 1))
out_best = [k for k, g in itertools.groupby(out_best)]
outstr = labels_to_text(out_best)
ret.append(outstr)
return ret
# Input Parameters
img_w = 300
img_h = 40
words_per_epoch = 16000
val_split = 0.2
val_words = int(words_per_epoch * (val_split))
# Network parameters
conv_filters = 16
kernel_size = (3, 3)
pool_size = 2
time_dense_size = 32
rnn_size = 512
minibatch_size = 32
OutputSize = 84
absolute_max_string_len = 30
start_epoch = 0
stop_epoch = 10
if K.image_data_format() == 'channels_first':
input_shape = (1, img_w, img_h)
else:
input_shape = (img_w, img_h, 1)
# Model
act = 'relu'
input_data = Input(name='the_input', shape=input_shape, dtype='float32')
inner = Conv2D(conv_filters, kernel_size, padding='same',
activation=act, kernel_initializer='he_normal',
name='conv1')(input_data)
inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max1')(inner)
inner = Conv2D(conv_filters, kernel_size, padding='same',
activation=act, kernel_initializer='he_normal',
name='conv2')(inner)
inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max2')(inner)
conv_to_rnn_dims = (img_w // (pool_size ** 2), (img_h // (pool_size ** 2)) * conv_filters)
inner = Reshape(target_shape=conv_to_rnn_dims, name='reshape')(inner)
# cuts down input size going into RNN:
inner = Dense(time_dense_size, activation=act, name='dense1')(inner)
# Two layers of bidirectional GRUs
# GRU seems to work as well, if not better than LSTM:
gru_1 = GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal', name='gru1')(inner)
gru_1b = GRU(rnn_size, return_sequences=True, go_backwards=True, kernel_initializer='he_normal', name='gru1_b')(inner)
gru1_merged = add([gru_1, gru_1b])
gru_2 = GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal', name='gru2')(gru1_merged)
gru_2b = GRU(rnn_size, return_sequences=True, go_backwards=True, kernel_initializer='he_normal', name='gru2_b')(gru1_merged)
# transforms RNN output to character activations:
inner = Dense(OutputSize, kernel_initializer='he_normal',
name='dense2')(concatenate([gru_2, gru_2b]))
y_pred = Activation('softmax', name='softmax')(inner)
Model(inputs=input_data, outputs=y_pred).summary()
labels = Input(name='the_labels', shape=[absolute_max_string_len], dtype='float32')
input_length = Input(name='input_length', shape=[1], dtype='int64')
label_length = Input(name='label_length', shape=[1], dtype='int64')
# Keras doesn't currently support loss funcs with extra parameters
# so CTC loss is implemented in a lambda layer
loss_out = Lambda(ctc_lambda_func, output_shape=(1,), name='ctc')([y_pred, labels, input_length, label_length])
# clipnorm seems to speeds up convergence
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True, clipnorm=5)
model = Model(inputs=[input_data, labels, input_length, label_length], outputs=loss_out)
# the loss calc occurs elsewhere, so use a dummy lambda func for the loss
model.compile(loss={'ctc': lambda y_true, y_pred: y_pred}, optimizer=sgd)
if start_epoch > 0:
weight_file = os.path.join(OUTPUT_DIR, os.path.join(run_name, 'weights%02d.h5' % (start_epoch - 1)))
model.load_weights(weight_file)
# captures output of softmax so we can decode the output during visualization
test_func = K.function([input_data], [y_pred])
# viz_cb = VizCallback(run_name, test_func, img_gen.next_val())
model.fit_generator(generator=data_gen(),
steps_per_epoch=108,
epochs=100,
validation_data=val_gen(),
validation_steps=27,
initial_epoch=0)
from keras.utils import plot_model
plot_model(model, to_file='model.png', show_shapes=True)
from IPython.display import Image
Image(filename='model.png')
model.save('my_model.h5') # creates a HDF5 file 'my_model.h5'
files.download('my_model.h5')
files.upload()
!ls
weight_file = "my_model3.h5"
# Input Parameters
img_w = 300
img_h = 40
words_per_epoch = 16000
val_split = 0.2
val_words = int(words_per_epoch * (val_split))
# Network parameters
conv_filters = 16
kernel_size = (3, 3)
pool_size = 2
time_dense_size = 32
rnn_size = 512
minibatch_size = 32
OutputSize = 84
absolute_max_string_len = 30
start_epoch = 0
stop_epoch = 10
if K.image_data_format() == 'channels_first':
input_shape = (1, img_w, img_h)
else:
input_shape = (img_w, img_h, 1)
# Model
act = 'relu'
input_data = Input(name='the_input', shape=input_shape, dtype='float32')
inner = Conv2D(conv_filters, kernel_size, padding='same',
activation=act, kernel_initializer='he_normal',
name='conv1')(input_data)
inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max1')(inner)
inner = Conv2D(conv_filters, kernel_size, padding='same',
activation=act, kernel_initializer='he_normal',
name='conv2')(inner)
inner = MaxPooling2D(pool_size=(pool_size, pool_size), name='max2')(inner)
conv_to_rnn_dims = (img_w // (pool_size ** 2), (img_h // (pool_size ** 2)) * conv_filters)
inner = Reshape(target_shape=conv_to_rnn_dims, name='reshape')(inner)
# cuts down input size going into RNN:
inner = Dense(time_dense_size, activation=act, name='dense1')(inner)
# Two layers of bidirectional GRUs
# GRU seems to work as well, if not better than LSTM:
gru_1 = GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal', name='gru1')(inner)
gru_1b = GRU(rnn_size, return_sequences=True, go_backwards=True, kernel_initializer='he_normal', name='gru1_b')(inner)
gru1_merged = add([gru_1, gru_1b])
gru_2 = GRU(rnn_size, return_sequences=True, kernel_initializer='he_normal', name='gru2')(gru1_merged)
gru_2b = GRU(rnn_size, return_sequences=True, go_backwards=True, kernel_initializer='he_normal', name='gru2_b')(gru1_merged)
# transforms RNN output to character activations:
inner = Dense(OutputSize, kernel_initializer='he_normal',
name='dense2')(concatenate([gru_2, gru_2b]))
y_pred = Activation('softmax', name='softmax')(inner)
Model(inputs=input_data, outputs=y_pred).summary()
labels = Input(name='the_labels', shape=[absolute_max_string_len], dtype='float32')
input_length = Input(name='input_length', shape=[1], dtype='int64')
label_length = Input(name='label_length', shape=[1], dtype='int64')
# Keras doesn't currently support loss funcs with extra parameters
# so CTC loss is implemented in a lambda layer
loss_out = Lambda(ctc_lambda_func, output_shape=(1,), name='ctc')([y_pred, labels, input_length, label_length])
# clipnorm seems to speeds up convergence
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True, clipnorm=5)
model_ = Model(inputs=[input_data, labels, input_length, label_length], outputs=loss_out)
# the loss calc occurs elsewhere, so use a dummy lambda func for the loss
model_.compile(loss={'ctc': lambda y_true, y_pred: y_pred}, optimizer=sgd)
model_.load_weights(weight_file)
# captures output of softmax so we can decode the output during visualization
test_func = K.function([input_data], [y_pred])
model_.fit_generator(generator=data_gen(),
steps_per_epoch=108,
epochs=100,
validation_data=val_gen(),
validation_steps=27,
initial_epoch=0)
model_.save('my_model4.h5') # creates a HDF5 file 'my_model.h5'
files.download('my_model4.h5')
model_.fit_generator(generator=data_gen(),
steps_per_epoch=108,
epochs=100,
validation_data=val_gen(),
validation_steps=27,
initial_epoch=0)
model_.save('my_model5.h5') # creates a HDF5 file 'my_model.h5'
files.download('my_model5.h5')
model_p = Model(inputs=input_data, outputs=y_pred)
def decode_predict_ctc(out, top_paths = 1):
results = []
beam_width = 5
if beam_width < top_paths:
beam_width = top_paths
for i in range(top_paths):
lables = K.get_value(K.ctc_decode(out, input_length=np.ones(out.shape[0])*out.shape[1],
greedy=False, beam_width=beam_width, top_paths=top_paths)[0][i])[0]
# text = labels_to_text(lables)
results.append(lables)
return results
img = FSTest['x'][-1]
plt.imshow(img)
img = img.reshape(1,40,300)
c = np.expand_dims(img.T, axis=0)
c.shape
net_out_value = model_p.predict(c)
net_out_value
pred_texts = decode_predict_ctc(net_out_value)
pred_texts
q = id2char(pred_texts[0])
plt.imshow(net_out_value[0].T, cmap='binary', interpolation='nearest')
plt.show()
q
m = "".join(q)
test.convertBijoyToUnicode(m)
model.fit_generator(generator=data_gen(),
steps_per_epoch=108,
epochs=100,
validation_data=val_gen(),
validation_steps=27,
initial_epoch=0)
model.save('my_model2.h5')
files.download('my_model2.h5')
model_p = Model(inputs=input_data, outputs=y_pred)
net_out_value = model_p.predict(c)
pred_texts = decode_predict_ctc(net_out_value)
q = id2char(pred_texts[0])
plt.imshow(net_out_value[0].T, cmap='binary', interpolation='nearest')
plt.show()
q
m = "".join(q)
test.convertBijoyToUnicode(m)
```
| github_jupyter |
## Fashion MNIST Image Classification - Multi-GPU training
**Code tested on:**
- Tensorflow==2.1.0
- Tensorflow-datasets==2.1.0
**Key activities**
- Extract and process Fashion-MNIST data
- Build Tensorflow keras model
- Training on Multiple GPU using MirroredStrategy
- Evaluate model
```
!pip3 install tensorflow-datasets==2.1.0 --user
# restart kernel
from IPython.display import display_html
def restartkernel() :
display_html("<script>Jupyter.notebook.kernel.restart()</script>",raw=True)
restartkernel()
```
### Import libraries
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow_datasets as tfds
import tensorflow as tf
import numpy as np
tfds.disable_progress_bar()
import logging
from datetime import datetime
logger = tf.get_logger()
logging.basicConfig(
format="%(asctime)s %(levelname)-8s %(message)s",
datefmt="%Y-%m-%dT%H:%M:%SZ",
level=logging.INFO)
print('Tensorflow-version: {0}'.format(tf.__version__))
# clear the logs
!rm -rf logs/
```
### Data extraction & processing
```
# prepare data
def prepare_data(batch_size=64, shuffle_size=1000):
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
# Split the training set into 80% and 20% for training and validation
train_validation_split = tfds.Split.TRAIN.subsplit([8, 2])
((train_data, validation_data), test_data),info = tfds.load(name="fashion_mnist:1.0.0",
split=(train_validation_split, tfds.Split.TEST),
as_supervised=True, with_info=True)
print("Training data count : ", int(info.splits['train'].num_examples * 0.8))
print("Validation data count : ", int(info.splits['train'].num_examples * 0.2))
print("Test data count : ", int(info.splits['test'].num_examples))
# create dataset to be used for training process
train_dataset = train_data.map(scale).shuffle(shuffle_size).batch(batch_size).repeat().prefetch(tf.data.experimental.AUTOTUNE)
val_dataset = validation_data.map(scale).batch(batch_size).prefetch(tf.data.experimental.AUTOTUNE)
test_dataset = test_data.map(scale).batch(batch_size)
return train_dataset, val_dataset, test_dataset
```
### Build Model
```
def build_model(learning_rate=0.001):
# define model architecture
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(filters=32, kernel_size=(3,3), activation='relu', input_shape=(28, 28, 1), name='x'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
# compile model with loss, optimizer and accuracy
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate),
metrics=['accuracy'])
return model
```
### Model Callback
```
def get_callbacks():
# callbacks
# folder to store current training logs
logdir="logs/fit/" + datetime.now().strftime("%Y%m%d-%H%M%S")
class customLog(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
logging.info('epoch: {}'.format(epoch + 1))
logging.info('loss={}'.format(logs['loss']))
logging.info('accuracy={}'.format(logs['accuracy']))
logging.info('val_accuracy={}'.format(logs['val_accuracy']))
callbacks = [
tf.keras.callbacks.TensorBoard(logdir),
customLog()
]
return callbacks
```
### Multi-GPU Training
```
# list physical devices available
tf.config.list_physical_devices('GPU')
# using MirroredStrategy
NUM_GPUS = 2
strategy = tf.distribute.MirroredStrategy(cross_device_ops=tf.distribute.HierarchicalCopyAllReduce())
print('Number of devices: {}'.format(strategy.num_replicas_in_sync))
with strategy.scope():
# Data extraction and processing
# set variables
BUFFER_SIZE = 10000
BATCH_SIZE = 64 * strategy.num_replicas_in_sync
train_dataset, val_dataset, test_dataset = prepare_data(batch_size=BATCH_SIZE, shuffle_size=BUFFER_SIZE)
TF_LEARNING_RATE = 0.001
# build model
model = build_model(learning_rate=TF_LEARNING_RATE)
model.summary()
# train model
TF_EPOCHS=20
TF_STEPS_PER_EPOCHS = int(np.ceil(60000 / float(BATCH_SIZE)))
model.fit(train_dataset,
epochs=TF_EPOCHS,
steps_per_epoch=3,
validation_data=val_dataset,
callbacks=get_callbacks())
```
**Track GPU Usage**
If you want to track the GPU usage then, open a terminal and use `nvidia-smi` command. To get refreshed value you can use the `watch -n <NUM_SECONDS>` command.
`watch -n 1 nvidia-smi`
```
# evaluate model
result = model.evaluate(test_dataset, steps=1)
loss = result[0]
accuracy = result[1]
print("loss : {0} accuracy : {1}".format(loss, accuracy))
```
#### Tensorboard
Note : If you want to use Tensorboard : use tensorboard command
```
tensorboard --logdir=/home/jovyan/logs/ --bind_all
```
if you are running inside a **container** you can use **port-mapping**. if you are running inside **kubernetes pod**, then use the pod **port-forward feature** on the port 6006 (default for tensorboard, change it as per the tensorboard command output ). When a notebook is created, a pod with name <NOTEBOOK_NAME>-0 is created in the users namespace. So you can use the port-forward to access tensorboard.
```
kubectl port-forward -n <NAMESPACE> <NOTEBOOK_NAME>-0 6006:6006
```
| github_jupyter |
<h1 align="center">Segmentation: Region Growing <a href="https://mybinder.org/v2/gh/InsightSoftwareConsortium/SimpleITK-Notebooks/master?filepath=Python%2F30_Segmentation_Region_Growing.ipynb"><img style="float: right;" src="https://mybinder.org/badge_logo.svg"></a>
</h1>
In this notebook we use one of the simplest segmentation approaches, region growing. We illustrate
the use of three variants of this family of algorithms. The common theme for all algorithms is that a voxel's neighbor is considered to be in the same class if its intensities are similar to the current voxel. The definition of similar is what varies:
* <b>ConnectedThreshold</b>: The neighboring voxel's intensity is within explicitly specified thresholds.
* <b>ConfidenceConnected</b>: The neighboring voxel's intensity is within the implicitly specified bounds $\mu\pm c\sigma$, where $\mu$ is the mean intensity of the seed points, $\sigma$ their standard deviation and $c$ a user specified constant.
* <b>VectorConfidenceConnected</b>: A generalization of the previous approach to vector valued images, for instance multi-spectral images or multi-parametric MRI. The neighboring voxel's intensity vector is within the implicitly specified bounds using the Mahalanobis distance $\sqrt{(\mathbf{x}-\mathbf{\mu})^T\Sigma^{-1}(\mathbf{x}-\mathbf{\mu})}<c$, where $\mathbf{\mu}$ is the mean of the vectors at the seed points, $\Sigma$ is the covariance matrix and $c$ is a user specified constant.
We will illustrate the usage of these three filters using a cranial MRI scan (T1 and T2) and attempt to segment one of the ventricles.
```
# To use interactive plots (mouse clicks, zooming, panning) we use the notebook back end. We want our graphs
# to be embedded in the notebook, inline mode, this combination is defined by the magic "%matplotlib notebook".
%matplotlib notebook
import SimpleITK as sitk
%run update_path_to_download_script
from downloaddata import fetch_data as fdata
import gui
# Using an external viewer (ITK-SNAP or 3D Slicer) we identified a visually appealing window-level setting
T1_WINDOW_LEVEL = (1050,500)
```
## Read Data and Select Seed Point(s)
We first load a T1 MRI brain scan and select our seed point(s). If you are unfamiliar with the anatomy you can use the preselected seed point specified below, just uncomment the line.
```
img_T1 = sitk.ReadImage(fdata("nac-hncma-atlas2013-Slicer4Version/Data/A1_grayT1.nrrd"))
# Rescale the intensities and map them to [0,255], these are the default values for the output
# We will use this image to display the results of segmentation
img_T1_255 = sitk.Cast(sitk.IntensityWindowing(img_T1,
windowMinimum=T1_WINDOW_LEVEL[1]-T1_WINDOW_LEVEL[0]/2.0,
windowMaximum=T1_WINDOW_LEVEL[1]+T1_WINDOW_LEVEL[0]/2.0),
sitk.sitkUInt8)
point_acquisition_interface = gui.PointDataAquisition(img_T1, window_level=(1050,500))
#preselected seed point in the left ventricle
point_acquisition_interface.set_point_indexes([(132,142,96)])
initial_seed_point_indexes = point_acquisition_interface.get_point_indexes()
```
## ConnectedThreshold
We start by using explicitly specified thresholds, you should modify these (lower/upper) to see the effects on the
resulting segmentation.
```
seg_explicit_thresholds = sitk.ConnectedThreshold(img_T1, seedList=initial_seed_point_indexes, lower=100, upper=170)
# Overlay the segmentation onto the T1 image
gui.MultiImageDisplay(image_list = [sitk.LabelOverlay(img_T1_255, seg_explicit_thresholds)],
title_list = ['connected threshold result'])
```
## ConfidenceConnected
This region growing algorithm allows the user to implicitly specify the threshold bounds based on the statistics estimated from the seed points, $\mu\pm c\sigma$. This algorithm has some flexibility which you should familiarize yourself with:
* The "multiplier" parameter is the constant $c$ from the formula above.
* You can specify a region around each seed point "initialNeighborhoodRadius" from which the statistics are estimated, see what happens when you set it to zero.
* The "numberOfIterations" allows you to rerun the algorithm. In the first run the bounds are defined by the seed voxels you specified, in the following iterations $\mu$ and $\sigma$ are estimated from the segmented points and the region growing is updated accordingly.
```
seg_implicit_thresholds = sitk.ConfidenceConnected(img_T1, seedList=initial_seed_point_indexes,
numberOfIterations=0,
multiplier=2,
initialNeighborhoodRadius=1,
replaceValue=1)
gui.MultiImageDisplay(image_list = [sitk.LabelOverlay(img_T1_255, seg_implicit_thresholds)],
title_list = ['confidence connected result'])
```
## VectorConfidenceConnected
We first load a T2 image from the same person and combine it with the T1 image to create a vector image. This region growing algorithm is similar to the previous one, ConfidenceConnected, and allows the user to implicitly specify the threshold bounds based on the statistics estimated from the seed points. The main difference is that in this case we are using the Mahalanobis and not the intensity difference.
```
img_T2 = sitk.ReadImage(fdata("nac-hncma-atlas2013-Slicer4Version/Data/A1_grayT2.nrrd"))
img_multi = sitk.Compose(img_T1, img_T2)
seg_implicit_threshold_vector = sitk.VectorConfidenceConnected(img_multi,
initial_seed_point_indexes,
numberOfIterations=2,
multiplier=4)
gui.MultiImageDisplay(image_list = [sitk.LabelOverlay(img_T1_255, seg_implicit_threshold_vector)],
title_list = ['vector confidence connected result'])
```
## Clean up, Clean up...
Use of low level segmentation algorithms such as region growing is often followed by a clean up step. In this step we fill holes and remove small connected components. Both of these operations are achieved by using binary morphological operations, opening (BinaryMorphologicalOpening) to remove small connected components and closing (BinaryMorphologicalClosing) to fill holes.
SimpleITK supports several shapes for the structuring elements (kernels) including:
* sitkAnnulus
* sitkBall
* sitkBox
* sitkCross
The size of the kernel can be specified as a scalar (same for all dimensions) or as a vector of values, size per dimension.
The following code cell illustrates the results of such a clean up, using closing to remove holes in the original segmentation.
```
vectorRadius=(1,1,1)
kernel=sitk.sitkBall
seg_implicit_thresholds_clean = sitk.BinaryMorphologicalClosing(seg_implicit_thresholds,
vectorRadius,
kernel)
```
And now we compare the original segmentation to the segmentation after clean up (using the GUI you can zoom in on the region of interest for a closer look).
```
gui.MultiImageDisplay(image_list = [sitk.LabelOverlay(img_T1_255, seg_implicit_thresholds),
sitk.LabelOverlay(img_T1_255, seg_implicit_thresholds_clean)],
shared_slider=True,
title_list = ['before morphological closing', 'after morphological closing'])
```
| github_jupyter |
# MACD with real data
This notebook shows the basic steps to begin to use downloaded NASDAQ csv stock price data.
*Follow the links below to download the csv and save it in the same directory as this notebook.*
The steps are:
1. Reading the data from the csv file into a pandas dataframe.
2. Cleaning up the data.
3. Plotting the data.
Data Source: [NASDAQ](https://www.nasdaq.com/), [Historical Quotes](https://www.nasdaq.com/market-activity/stocks/pep/historical)
## Libraries
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# MACD and EMA Functions
from MACDRealDataHelperFunctions import *
```
## Data Import and Data Cleaning
```
# Import and Clean Data
def importAndCleanData(file_name):
# Ingest the csv file into a pandas dataframe.
df = pd.read_csv(str(file_name))
# Remove dollar signs using df.replace()
try:
columns_with_dollars = [' Close/Last', ' Open', ' High', ' Low']
df[columns_with_dollars] = df[columns_with_dollars].replace({'\$':''}, regex=True)
except:
print("No dollar signs ($) to remove.")
# Convert object datatype to float
df[columns_with_dollars] = df[columns_with_dollars].astype(float)
# Reverse the spreadsheet to begin with oldest date and end with newest date.
df = df[::-1] # This reverses the order, but doesn't change the index numbers.
df = df.reset_index(drop=True) # This overwrites the index numbers to start from zero.
#df # Prints the dataframe. Useful for debugging.
return df
file_name = 'HistoricalQuotes-PEP-6M.csv'
df = importAndCleanData(file_name)
```
## Data Plotting
```
# Plot raw data from dataframe
def plotDataframe(df, file_name):
# Plot the data.
ax = plt.gca() # gca stands for 'get current axis'
df.plot(kind='line',x='Date',y=' High', color='blue',ax=ax)
df.plot(kind='line',x='Date',y=' Low', color='red', ax=ax)
df.plot(kind='line',x='Date',y=' Close/Last', color='cyan',ax=ax)
df.plot(kind='line',x='Date',y=' Open', color='orange', ax=ax, title=str(file_name))
#df.plot(title="ya dangus")
plt.show()
plotDataframe(df, file_name)
# Wrapper Function for EMAs and MACD Calculation/Plotting
def wrapper(df, ema_short_period, ema_long_period, macd_signal_period, plot_x_start, plot_x_end, plot_after_long_period):
# We will use the closing price for our analysis.
value_data = df[' Close/Last']
# We can use the dates as labels for the x axis.
date_data = df['Date']
# Run MACD Calculations
x, short_data, long_data, MACD, MACD_signal, MACD_bars = getMACD(value_data, ema_short_period, ema_long_period, macd_signal_period)
# Plot Results
calculateAndPlotEMA(value_data, ema_short_period, ema_long_period, plot_after_long_period)
plotMACD(x, MACD, MACD_signal, macd_signal_period, MACD_bars, plot_x_start, plot_x_end)
# MACD Parameters
ema_short_period = 12
ema_long_period = 26
macd_signal_period = 9
plot_after_long_period = True
# MACD Plotting Parameters
plot_x_start = 80
plot_x_end = 130
# Run everything
wrapper(df, ema_short_period, ema_long_period, macd_signal_period, plot_x_start, plot_x_end, plot_after_long_period)
```
## Let's see what a different ticker looks like
```
# Prepare the data
file_name = 'HistoricalQuotes-TGT-6M.csv' # Target Corp. 6 months of data.
df = importAndCleanData(file_name)
plotDataframe(df, file_name)
# MACD Parameters
ema_short_period = 12
ema_long_period = 26
macd_signal_period = 9
plot_x_start = 80
plot_x_end = 130
plot_after_long_period = True
# Run everything
wrapper(df, ema_short_period, ema_long_period, macd_signal_period, plot_x_start, plot_x_end, plot_after_long_period)
# Prepare the data
file_name = 'HistoricalQuotes-VOO-6M.csv' # Vanguard 500 ETF, 6 months of data. Note, does NOT have $ signs.
df = importAndCleanData(file_name)
plotDataframe(df, file_name)
# MACD Parameters
ema_short_period = 12
ema_long_period = 26
macd_signal_period = 9
plot_x_start = 80
plot_x_end = 130
plot_after_long_period = True
# Run everything
wrapper(df, ema_short_period, ema_long_period, macd_signal_period, plot_x_start, plot_x_end, plot_after_long_period)
```
| github_jupyter |
# Regressão Linear Regularizada e Bias vs Variância
```
# usado para manipular caminhos de diretório
import os
# pacote usado para realizar operações com matrizes
import numpy as np
# pacote de visualização gráfica
from matplotlib import pyplot as plt
# pacote de otimização (escolha de alguns hiperparâmetros)
from scipy import optimize
# carrega datasets executáveis em matlab
from scipy.io import loadmat
# incorporando plotagem do matplotlib no arquivo
%matplotlib inline
```
## Regressão Linear Regularizada
Na primeira metade do exercício, você implementará a regressão linear regularizada para prever a quantidade de água que flui de uma barragem usando a mudança do nível de água em um reservatório. Na próxima metade, você fará alguns diagnósticos de algoritmos de aprendizagem de depuração e examinará os efeitos do viés vs. variância.
### Visualizando o dataset
Começaremos visualizando o conjunto de dados contendo registros históricos sobre a mudança no nível da água, $ x $, e a quantidade de água fluindo para fora da barragem, $ y $. Este conjunto de dados é dividido em três partes:
- Um conjunto de **treinamento** que seu modelo aprenderá em: **X**, **y**;
- Um conjunto de **validação cruzada** definida para determinar o parâmetro de regularização: **Xval**, **yval**;
- Um conjunto de **teste** definido para avaliar o desempenho. Estes são exemplos que seu modelo não viu durante o treinamento: **Xtest**, **ytest**;
```
# carregando os dados do arquivo ex5data1.mat, todas as variáveis serão armazenadas em um dicionário
dataset = loadmat(os.path.join('datasets', 'ex5data1.mat'))
# visulizando a organização do dicionário
for keys, values in dataset.items():
print(keys)
# separando os dados de treinamento, teste e validação do dicionário
# além disso, os dados foram convertidos para um formato de vetor numpy
X, y = dataset['X'], dataset['y'][:, 0]
Xtest, ytest = dataset['Xtest'], dataset['ytest'][:, 0]
Xval, yval = dataset['Xval'], dataset['yval'][:, 0]
# m = número de exemplos treináveis
m = y.size
print(m)
# Visualizando os dados com matplotlib
plt.figure(figsize = (10, 5))
plt.plot(X, y, 'ro', ms=10, mec='k', mew=1)
plt.xlabel('Mudança no nível da água (x)')
plt.ylabel('Água fluindo para fora da barragem (y)')
```
### Função de custo de regressão linear regularizada
Lembre-se de que a regressão linear regularizada tem a seguinte função de custo:
$$ J(\theta) = \frac{1}{2m} \left( \sum_{i=1}^m \left( h_\theta\left( x^{(i)} \right) - y^{(i)} \right)^2 \right) + \frac{\lambda}{2m} \left( \sum_{j=1}^n \theta_j^2 \right)$$
Onde $\lambda$ é um parâmetro de regularização que controla o grau de regularização (assim, ajuda a prevenir overfitting). O termo de regularização impõe uma penalidade ao custo geral J. À medida que as magnitudes dos parâmetros do modelo $\theta_j$ aumentam, a penalidade também aumenta. Observe que você não deve regularizar o termo $\theta_0$.
### Gradiente de regressão linear regularizado
Correspondentemente, a derivada parcial da função de custo para regressão linear regularizada é definida como:
$$
\begin{align}
& \frac{\partial J(\theta)}{\partial \theta_0} = \frac{1}{m} \sum_{i=1}^m \left( h_\theta \left(x^{(i)} \right) - y^{(i)} \right) x_j^{(i)} & \qquad \text{for } j = 0 \\
& \frac{\partial J(\theta)}{\partial \theta_j} = \left( \frac{1}{m} \sum_{i=1}^m \left( h_\theta \left( x^{(i)} \right) - y^{(i)} \right) x_j^{(i)} \right) + \frac{\lambda}{m} \theta_j & \qquad \text{for } j \ge 1
\end{align}
$$
```
def linearRegCostFunction(X, y, teta, lambda_= 0.0):
"""
Compute cost and gradient for regularized linear regression
with multiple variables. Computes the cost of using theta as
the parameter for linear regression to fit the data points in X and y.
Parameters
----------
X : array_like
The dataset. Matrix with shape (m x n + 1) where m is the
total number of examples, and n is the number of features
before adding the bias term.
y : array_like
The functions values at each datapoint. A vector of
shape (m, ).
theta : array_like
The parameters for linear regression. A vector of shape (n+1,).
lambda_ : float, optional
The regularization parameter.
Returns
-------
J : float
The computed cost function.
grad : array_like
The value of the cost function gradient w.r.t theta.
A vector of shape (n+1, ).
Instructions
------------
Compute the cost and gradient of regularized linear regression for
a particular choice of theta.
You should set J to the cost and grad to the gradient.
"""
m = y.size # número de exemplos treináveis
# You need to return the following variables correctly
J = 0
grad = np.zeros(teta.shape)
# ====================== YOUR CODE HERE ======================
# computando a equação da função de custo
h = X.dot(teta)
J = (1 / (2 * m)) * np.sum(np.square(h - y)) + (lambda_ / (2 * m)) * np.sum(np.square(teta[1:]))
# computando o valor dos parâmetros através do método de gradiente descendente (via derivadas parciais)
grad = (1 / m) * (h - y).dot(X)
grad[1:] = grad[1:] + (lambda_ / m) * teta[1:]
# ============================================================
return J, grad
# hstack = concatena por colunas, vstack = concatena por linhas
#np.hstack((np.ones((m, 1)), X))
# axis = 1 concatena por colunas, axis = 0 concatena por linhas
#np.concatenate([np.ones((m, 1)), X], axis=1)
# definindo uma hipótese inicial para os valores dos parâmetros
teta = np.array([1, 1])
# adicionando um bias aos atributos previsores de treinamento
X_bias = np.concatenate([np.ones((m, 1)), X], axis=1)
# computando o valor da função de custo para esses valores
J, grad = linearRegCostFunction(X_bias, y, teta, 1)
print('Custo dos valores de teta = {}: {}'.format(teta, J))
print('Valores dos parâmetros teta para esse custo: {}'.format(grad))
```
### Realizando o treinamento com a regressão linear
Uma vez que sua função de custo e gradiente estão funcionando corretamente, a próxima célula irá executar o código em `trainLinearReg` para calcular os valores ótimos de $\theta$. Esta função de treinamento usa o módulo de otimização `scipy` para minimizar a função de custo.
Nesta parte, definimos o parâmetro de regularização $\lambda$ como zero. Como nossa implementação atual de regressão linear está tentando ajustar um $\theta$ bidimensional, a regularização não será extremamente útil para um $\theta$ de dimensão tão baixa. Nas partes posteriores do exercício, você usará a regressão polinomial com regularização.
Finalmente, o código na próxima célula também deve traçar a linha de melhor ajuste, que deve ser semelhante à figura abaixo.

A linha de melhor ajuste nos diz que o modelo não é um bom ajuste para os dados porque os dados têm um padrão não linear. Embora visualizar o melhor ajuste conforme mostrado seja uma maneira possível de depurar seu algoritmo de aprendizado, nem sempre é fácil visualizar os dados e o modelo. Na próxima seção, você implementará uma função para gerar curvas de aprendizado que podem ajudá-lo a depurar seu algoritmo de aprendizado, mesmo que não seja fácil visualizar os dados.
```
def trainLinearReg(linearRegCostFunction, X, y, lambda_ = 0.0, maxiter = 200):
"""
Trains linear regression using scipy's optimize.minimize.
Parameters
----------
X : array_like
The dataset with shape (m x n+1). The bias term is assumed to be concatenated.
y : array_like
Function values at each datapoint. A vector of shape (m,).
lambda_ : float, optional
The regularization parameter.
maxiter : int, optional
Maximum number of iteration for the optimization algorithm.
Returns
-------
theta : array_like
The parameters for linear regression. This is a vector of shape (n+1,).
"""
# definindo valores iniciais para teta
teta_inicial = np.zeros(X.shape[1])
# criando uma função lambda relativa a função de custo
costFunction = lambda t: linearRegCostFunction(X, y, t, lambda_)
# a função de custo recebe apenas um argumento
options = {'maxiter': maxiter}
# minimização da função de custo através do scipy (por meio da modificação dos parâmetros)
res = optimize.minimize(costFunction, teta_inicial, jac = True, method = 'TNC', options = options)
return res
# entendendo como uma função lambda pode ser invocada (observe que ela funciona só com a chamada de um argumento ao invés de 4)
lambda_ = 0
costFunction = lambda teta: linearRegCostFunction(X_bias, y, teta, lambda_)
costFunction(np.array([1, 1]))
# obtendo a função de custo e os parâmetros após a otimização
valores_otimizados = trainLinearReg(linearRegCostFunction, X_bias, y, lambda_ = 0)
print('Função de custo após a otimização: {}'.format(valores_otimizados.fun))
print('Valores de teta após a otimização: {}'.format(valores_otimizados.x))
# Visualizando a reta obtida com o algoritmo de regressão linear
plt.plot(X, y, 'ro', ms=10, mec='k', mew=1.5)
plt.xlabel('Mudança no nível da água (x)')
plt.ylabel('Água fluindo para fora da barragem (y)')
plt.plot(X, np.dot(X_bias, valores_otimizados.x), '--', lw=2)
```
## Bias - Variância
Um conceito importante no aprendizado de máquina é a compensação de bias - variância. Os modelos com um viés (bias) alto não são complexos o suficiente para os dados e tendem a se ajustar mal, enquanto os modelos com alta variância se ajustam excessivamente aos dados de treinamento.
### Curvas de Aprendizagem
Agora, você implementará o código para gerar as curvas de aprendizado que serão úteis na depuração de algoritmos de aprendizado. Lembre-se de que uma curva de aprendizado traça o treinamento e o erro de validação cruzada como uma função do tamanho do conjunto de treinamento. Seu trabalho é preencher a função **learningCurve** na próxima célula, de modo que ela retorne um vetor de erros para o conjunto de treinamento e conjunto de validação cruzada.
Para traçar a curva de aprendizado, precisamos de um erro de conjunto de treinamento e validação cruzada para diferentes tamanhos de conjunto de treinamento. Para obter tamanhos de conjunto de treinamento diferentes, você deve usar subconjuntos diferentes do conjunto de treinamento original `X`. Especificamente, para um tamanho de conjunto de treinamento de $i$, você deve usar os primeiros $i$ exemplos (i.e., `X[:i, :]`
and `y[:i]`).
Depois de aprender os parâmetros $\theta$, você deve calcular o erro nos conjuntos de treinamento e validação cruzada. Lembre-se de que o erro de treinamento para um conjunto de dados é definido como
$$ J_{\text{train}} = \frac{1}{2m} \left[ \sum_{i=1}^m \left(h_\theta \left( x^{(i)} \right) - y^{(i)} \right)^2 \right] $$
Em particular, observe que o erro de treinamento não inclui o termo de regularização. Uma maneira de calcular o erro de treinamento é usar sua função de custo existente e definir $\lambda$ como 0 apenas ao usá-la para calcular o erro de treinamento e o erro de validação cruzada. Ao calcular o erro do conjunto de treinamento, certifique-se de calculá-lo no subconjunto de treinamento (ou seja, `X [: n,:]` e `y [: n]`) em vez de no conjunto de treinamento inteiro. No entanto, para o erro de validação cruzada, você deve computá-lo em todo o conjunto de validação cruzada.
```
def learningCurve(X, y, Xval, yval, lambda_=0):
"""
Generates the train and cross validation set errors needed to plot a learning curve
returns the train and cross validation set errors for a learning curve.
In this function, you will compute the train and test errors for
dataset sizes from 1 up to m. In practice, when working with larger
datasets, you might want to do this in larger intervals.
Parameters
----------
X : array_like
The training dataset. Matrix with shape (m x n + 1) where m is the
total number of examples, and n is the number of features
before adding the bias term.
y : array_like
The functions values at each training datapoint. A vector of
shape (m, ).
Xval : array_like
The validation dataset. Matrix with shape (m_val x n + 1) where m is the
total number of examples, and n is the number of features
before adding the bias term.
yval : array_like
The functions values at each validation datapoint. A vector of
shape (m_val, ).
lambda_ : float, optional
The regularization parameter.
Returns
-------
error_train : array_like
A vector of shape m. error_train[i] contains the training error for
i examples.
error_val : array_like
A vecotr of shape m. error_val[i] contains the validation error for
i training examples.
Instructions
------------
Fill in this function to return training errors in error_train and the
cross validation errors in error_val. i.e., error_train[i] and
error_val[i] should give you the errors obtained after training on i examples.
Notes
-----
- You should evaluate the training error on the first i training
examples (i.e., X[:i, :] and y[:i]).
For the cross-validation error, you should instead evaluate on
the _entire_ cross validation set (Xval and yval).
- If you are using your cost function (linearRegCostFunction) to compute
the training and cross validation error, you should call the function with
the lambda argument set to 0. Do note that you will still need to use
lambda when running the training to obtain the theta parameters.
Hint
----
You can loop over the examples with the following:
for i in range(1, m+1):
# Compute train/cross validation errors using training examples
# X[:i, :] and y[:i], storing the result in
# error_train[i-1] and error_val[i-1]
....
"""
# número de exemplos treináveis
m = y.size
# criando um array numpy para armazenar os erros de predição associados aos atributos de treinamento e de validação
erro_treinamento = np.zeros(m)
erro_validacao = np.zeros(m)
# ====================== YOUR CODE HERE ======================
for i in range(1, m + 1):
teta_t = trainLinearReg(linearRegCostFunction, X[:i], y[:i], lambda_ = 0)
erro_treinamento[i - 1], _ = linearRegCostFunction(X[:i], y[:i], teta_t.x, lambda_ = 0)
erro_validacao[i - 1], _ = linearRegCostFunction(Xval, yval, teta_t.x, lambda_ = 0)
# =============================================================
return erro_treinamento, erro_validacao
```
Quando você terminar de implementar a função `learningCurve`, executar a próxima célula imprimirá as curvas de aprendizado e produzirá um gráfico semelhante à figura abaixo.

Na figura da curva de aprendizado, você pode observar que tanto o erro do treinamento quanto o erro de validação cruzada são altos quando o número de exemplos de treinamento é aumentado. Isso reflete um problema de viés (bias) alto no modelo - o modelo de regressão linear é muito simples e não consegue se ajustar bem ao nosso conjunto de dados.
```
# adicionando um parâmetro bias nos atributos previsores de treinamento e de validação
X_bias = np.concatenate([np.ones((m, 1)), X], axis=1)
Xval_bias = np.concatenate([np.ones((yval.size, 1)), Xval], axis=1)
# obtendo os valores de erro associados aos dados de treinamento e aos dados de validação
erro_treinamento, erro_validacao = learningCurve(X_bias, y, Xval_bias, yval, lambda_=0)
# visulizando o gráfico de erro nas predições com os dados de treinamento e os dados de validação
plt.figure(figsize = (10, 5))
plt.plot(np.arange(1, m+1), erro_treinamento, np.arange(1, m+1), erro_validacao, lw=2)
plt.title('Curva de aprendizado para regressão linear')
plt.legend(['Treinamento', 'Validação Cruzada'])
plt.xlabel('Número de exemplos treináveis')
plt.ylabel('Erro')
print('# Exemplos de Treinamento\tErro de Treinamento\tErro de Validação Cruzada')
for i in range(m):
print('{}\t\t\t\t{}\t{}'.format(i+1, erro_treinamento[i], erro_validacao[i]))
```
## Regressão Polinomial Regularizada
O problema com nosso modelo linear é que ele é muito simples para os dados e resultava em subajuste (viés alto). Nesta parte do exercício, você tratará desse problema adicionando mais recursos. Para regressão polinomial, nossa hipótese tem a forma:
$$
\begin{align}
h_\theta(x) &= \theta_0 + \theta_1 \times (\text{nivelAgua}) + \theta_2 \times (\text{nivelAgua})^2 + \cdots + \theta_p \times (\text{nivelAgua})^p \\
& = \theta_0 + \theta_1 x_1 + \theta_2 x_2 + \cdots + \theta_p x_p
\end{align}
$$
Observe que ao definir $x_1 = (\text{nivelAgua})$, $x_2 = (\text{nivelAgua})^2$ , $\cdots$, $x_p =
(\text{nivelAgua})^p$, obtemos um modelo de regressão linear onde os recursos são as várias potências do valor original (nivelAgua).
Agora, você adicionará mais recursos usando as potências mais altas do recurso existente $x$ no conjunto de dados. Sua tarefa nesta parte é completar o código na função `polyFeatures` na próxima célula. A função deve mapear o conjunto de treinamento original $X$ de tamanho $m \times 1$ em suas potências superiores. Especificamente, quando um conjunto de treinamento $X$ de tamanho $m \times 1$ é passado para a função, a função deve retornar uma matriz $m \times p$ `X_poli`, onde a coluna 1 contém os valores originais de X, coluna 2 contém os valores de $X^2$, a coluna 3 contém os valores de $ X^3$ e assim sucessivamente.
```
def polyFeatures(X, p):
"""
Maps X (1D vector) into the p-th power.
Parameters
----------
X : array_like
A data vector of size m, where m is the number of examples.
p : int
The polynomial power to map the features.
Returns
-------
X_poly : array_like
A matrix of shape (m x p) where p is the polynomial
power and m is the number of examples. That is:
X_poly[i, :] = [X[i], X[i]**2, X[i]**3 ... X[i]**p]
Instructions
------------
Given a vector X, return a matrix X_poly where the p-th column of
X contains the values of X to the p-th power.
"""
# iniciar um array numpy para armazenar os valores das potências obtidas
X_polinomios = np.zeros((X.shape[0], p))
# ====================== YOUR CODE HERE ======================
# iteração para obter as potências relativas em cada execução
for i in range(p):
X_polinomios[:, i] = X[:, 0] ** (i + 1)
# ============================================================
return X_polinomios
```
Agora você tem uma função que mapeará os recursos para uma dimensão superior. A próxima célula o aplicará ao conjunto de treinamento, ao conjunto de teste e ao conjunto de validação cruzada.
```
def featureNormalize(X):
"""
Normalizes the features in X returns a normalized version of X where the mean value of each
feature is 0 and the standard deviation is 1. This is often a good preprocessing step to do when
working with learning algorithms.
Parameters
----------
X : array_like
An dataset which is a (m x n) matrix, where m is the number of examples,
and n is the number of dimensions for each example.
Returns
-------
X_norm : array_like
The normalized input dataset.
mu : array_like
A vector of size n corresponding to the mean for each dimension across all examples.
sigma : array_like
A vector of size n corresponding to the standard deviations for each dimension across
all examples.
"""
# obtendo a médio dos dados
mu = np.mean(X, axis = 0)
X_norm = (X - mu)
# obtendo o desvio padrão dos dados
sigma = np.std(X_norm, axis = 0, ddof = 1)
# aplicando a distribuição normal (ou distribuição gaussiana)
X_norm = (X - mu) / sigma
return X_norm, mu, sigma
# definindo polinômios até grau 8
p = 8
# realizar o mapeamento nos atributos previsores de treinamento
X_polinomial = polyFeatures(X, p)
X_polinomial, mu, sigma = featureNormalize(X_polinomial)
X_polinomial = np.concatenate([np.ones((m, 1)), X_polinomial], axis=1)
# Realizar o mapeamento e aplicar a normalização (usando mu e sigma)
X_polinomial_teste = polyFeatures(Xtest, p)
X_polinomial_teste -= mu
X_polinomial_teste /= sigma
# adicionando um parâmetro de viés
X_polinomial_teste = np.concatenate([np.ones((ytest.size, 1)), X_polinomial_teste], axis=1)
# Realizar o mapeamento e aplicar a normalização (usando mu e sigma)
X_polinomial_validacao = polyFeatures(Xval, p)
X_polinomial_validacao -= mu
X_polinomial_validacao /= sigma
# adicionando um parâmetro de viés
X_polinomial_validacao = np.concatenate([np.ones((yval.size, 1)), X_polinomial_validacao], axis=1)
print('Exemplos de treinamento normalizados: ')
X_polinomial[0, :]
```
Depois de concluir a função `polyFeatures`, continuaremos a treinar a regressão polinomial usando sua função de custo de regressão linear.
Lembre-se de que, embora tenhamos termos polinomiais em nosso vetor de recursos, ainda estamos resolvendo um problema de otimização de regressão linear. Os termos polinomiais simplesmente se transformaram em recursos que podemos usar para regressão linear. Estamos usando a mesma função de custo e gradiente que você escreveu para a parte anterior deste exercício.
Para esta parte do exercício, você usará um polinômio de grau 8. Acontece que se executarmos o treinamento diretamente nos dados projetados, não funcionará bem, pois os recursos seriam mal dimensionados (por exemplo, um exemplo com $ x = 40 $ agora terá um recurso de $x_8 = 40^8 = 6.5 \times 10^{12}$). Portanto, você vai precisa usar a normalização.
Antes de aprender os parâmetros $\theta$ para a regressão polinomial, primeiro chamamos `featureNormalize` e normalizamos os recursos do conjunto de treinamento, armazenando os parâmetros mu, sigma separadamente.
Depois de aprender os parâmetros $\theta$, você deve ver dois gráficos gerados para regressão polinomial com $\lambda = 0 $, que devem ser semelhantes aos aqui:
<table>
<tr>
<td><img src="imagens/polynomial_regression.png"></td>
<td><img src="imagens/polynomial_learning_curve.png"></td>
</tr>
</table>
Você deve ver que o ajuste polinomial é capaz de acompanhar muito bem os pontos de dados, obtendo assim um erro de treinamento baixo. A figura à direita mostra que o erro de treinamento permanece essencialmente zero para todos os números de amostras de treinamento. No entanto, o ajuste polinomial é muito complexo e até mesmo cai nos extremos. Este é um indicador de que o modelo de regressão polinomial está super ajustando os dados de treinamento e não generalizará bem.
Para entender melhor os problemas com o modelo não regularizado ($\lambda = 0$), você pode ver que a curva de aprendizado mostra o mesmo efeito onde o erro de treinamento é baixo, mas o erro de validação cruzada é alto. Há uma lacuna entre os erros de treinamento e de validação cruzada, indicando um problema de alta variância.
```
def plotFit(polyFeatures, min_x, max_x, mu, sigma, teta, p):
"""
Plots a learned polynomial regression fit over an existing figure.
Also works with linear regression.
Plots the learned polynomial fit with power p and feature normalization (mu, sigma).
Parameters
----------
polyFeatures : func
A function which generators polynomial features from a single feature.
min_x : float
The minimum value for the feature.
max_x : float
The maximum value for the feature.
mu : float
The mean feature value over the training dataset.
sigma : float
The feature standard deviation of the training dataset.
theta : array_like
The parameters for the trained polynomial linear regression.
p : int
The polynomial order.
"""
# traçamos um intervalo ligeiramente maior do que os valores mínimo e máximo para obter
# uma ideia de como o ajuste irá variar fora do intervalo dos pontos de dados
x = np.arange(min_x - 15, max_x + 25, 0.05).reshape(-1, 1)
# realizando um mapeamento nos valores de X_polinomio
X_polinomio = polyFeatures(x, p)
X_polinomio -= mu
X_polinomio /= sigma
# adicionando o parâmetro de viés
X_polinomio = np.concatenate([np.ones((x.shape[0], 1)), X_polinomio], axis=1)
# plotando o gráfico da curva
plt.plot(x, np.dot(X_polinomio, teta), '--', lw=2)
return None
lambda_ = 1
# obtendo os valores de teta otimizador
teta = trainLinearReg(linearRegCostFunction, X_polinomial, y, lambda_=lambda_, maxiter = 55)
# plotandos os dados e realizando treinamento para obter a curva polinomial
plt.figure(figsize = (10, 5))
plt.plot(X, y, 'ro', ms=10, mew=1.5, mec='k')
plotFit(polyFeatures, np.min(X), np.max(X), mu, sigma, teta.x, p)
plt.xlabel('Mudança no nível da água (x)')
plt.ylabel('Água fluindo para fora da barragem (y)')
plt.title('Ajuste de regressão polinomial (lambda =% f)' % lambda_)
plt.ylim([-20, 50])
plt.figure(figsize = (10, 5))
# obtendo os erros de predição para os dados de treinamento e dados de validação
erro_treinamento, erro_validacao = learningCurve(X_polinomial, y, X_polinomial_validacao, yval, lambda_)
plt.plot(np.arange(1, 1+m), erro_treinamento, np.arange(1, 1+m), erro_validacao)
plt.title('Curva de aprendizado de regressão polinomial (lambda =% f)' % lambda_)
plt.xlabel('Número de exemplos treináveis')
plt.ylabel('Erro')
plt.axis([0, 13, 0, 100])
plt.legend(['Treinamento', 'Validação Cruzada'])
# visualizando os erros associados aos dados de treinamento e aos dados de validação
print('Regressão polinomial (lambda =% f) \n' % lambda_)
print('# Exemplos de Treinamento\tErro de Treinamento\tErro de Validação Cruzada')
for i in range(m):
print('{}\t\t\t\t{}\t{}'.format(i+1, erro_treinamento[i], erro_validacao[i]))
```
Uma maneira de combater o problema de overfitting (alta variância) é adicionar regularização ao modelo. Na próxima seção, você experimentará diferentes parâmetros $\lambda$ para ver como a regularização pode levar a um modelo melhor.
### Ajustando o hiperparâmetro de regularização
Nesta seção, você verá como o parâmetro de regularização afeta a variação de polarização da regressão polinomial regularizada. Agora você deve modificar o parâmetro lambda e tentar $\lambda = 1, 100$. Para cada um desses valores, o script deve gerar um ajuste polinomial aos dados e também uma curva de aprendizado.
Para $\lambda = 1$, os gráficos gerados devem ser semelhantes à figura abaixo. Você deve ver um ajuste polinomial que segue bem a tendência dos dados (à esquerda) e uma curva de aprendizado (à direita) mostrando que a validação cruzada e o erro de treinamento convergem para um valor relativamente baixo. Isso mostra que o modelo de regressão polinomial regularizado $\lambda = 1$ não tem problemas de viés alto ou alta variância. Na verdade, ele consegue um bom equilíbrio entre o viés e a variância.
<table>
<tr>
<td><img src="imagens/polynomial_regression_reg_1.png"></td>
<td><img src="imagens/polynomial_learning_curve_reg_1.png"></td>
</tr>
</table>
Para $\lambda = 100$, você deve ver um ajuste polinomial (figura abaixo) que não segue bem os dados. Nesse caso, há muita regularização e o modelo não consegue ajustar os dados de treinamento.

### Selecionando $\lambda$ usando validação cruzada
Nas partes anteriores do exercício, você observou que o valor de $\lambda$ pode afetar significativamente os resultados da regressão polinomial regularizada no conjunto de treinamento e validação cruzada. Em particular, um modelo sem regularização ($\lambda = 0$) se ajusta bem ao conjunto de treinamento, mas não generaliza. Por outro lado, um modelo com muita regularização ($\lambda = 100$) não se ajusta bem ao conjunto de treinamento e teste. Uma boa escolha de $\lambda$ (por exemplo, $\lambda = 1$) pode fornecer um bom ajuste aos dados.
Nesta seção, você implementará um método automatizado para selecionar o parâmetro $\lambda$. Concretamente, você usará um conjunto de validação cruzada para avaliar a qualidade de cada valor de $\lambda$. Depois de selecionar o melhor valor $\lambda$ usando o conjunto de validação cruzada, podemos avaliar o modelo no conjunto de teste para estimar o quão bem o modelo terá um desempenho em dados reais não vistos.
Sua tarefa é completar o código na função `validationCurve`. Especificamente, você deve usar a função `trainLinearReg` para treinar o modelo usando diferentes valores de $\lambda$ e calcular o erro de treinamento e o erro de validação cruzada. Você deve tentar $\lambda$ no seguinte intervalo: {0, 0,001, 0,003, 0,01, 0,03, 0,1, 0,3, 1, 3, 10}.
```
def validationCurve(X, y, Xval, yval):
"""
Generate the train and validation errors needed to plot a validation
curve that we can use to select lambda_.
Parameters
----------
X : array_like
The training dataset. Matrix with shape (m x n) where m is the
total number of training examples, and n is the number of features
including any polynomial features.
y : array_like
The functions values at each training datapoint. A vector of
shape (m, ).
Xval : array_like
The validation dataset. Matrix with shape (m_val x n) where m is the
total number of validation examples, and n is the number of features
including any polynomial features.
yval : array_like
The functions values at each validation datapoint. A vector of
shape (m_val, ).
Returns
-------
lambda_vec : list
The values of the regularization parameters which were used in
cross validation.
error_train : list
The training error computed at each value for the regularization
parameter.
error_val : list
The validation error computed at each value for the regularization
parameter.
Instructions
------------
Fill in this function to return training errors in `error_train` and
the validation errors in `error_val`. The vector `lambda_vec` contains
the different lambda parameters to use for each calculation of the
errors, i.e, `error_train[i]`, and `error_val[i]` should give you the
errors obtained after training with `lambda_ = lambda_vec[i]`.
Note
----
You can loop over lambda_vec with the following:
for i in range(len(lambda_vec))
lambda = lambda_vec[i]
# Compute train / val errors when training linear
# regression with regularization parameter lambda_
# You should store the result in error_train[i]
# and error_val[i]
....
"""
# selecionando valores de análise para o hiperparâmetro lambda
lambda_vec = [0, 0.001, 0.003, 0.01, 0.03, 0.1, 0.3, 1, 3, 10]
# computando o erro na predição dos dados de treinamento e dados de validação
erro_treinamento = np.zeros(len(lambda_vec))
erro_validacao = np.zeros(len(lambda_vec))
# ====================== YOUR CODE HERE ======================
for i in range(len(lambda_vec)):
lambda_try = lambda_vec[i]
teta_t = trainLinearReg(linearRegCostFunction, X, y, lambda_ = lambda_try)
erro_treinamento[i], _ = linearRegCostFunction(X, y, teta_t.x, lambda_ = 0)
erro_validacao[i], _ = linearRegCostFunction(Xval, yval, teta_t.x, lambda_ = 0)
# ============================================================
return lambda_vec, erro_treinamento, erro_validacao
```
Depois de concluir o código, a próxima célula executará sua função e traçará uma curva de validação cruzada de erro vs. $\lambda$ que permite que você selecione qual parâmetro $\lambda$ usar. Você deve ver um gráfico semelhante à figura abaixo.

Nesta figura, podemos ver que o melhor valor de $\lambda$ está em torno de 3. Devido à aleatoriedade nas divisões de treinamento e validação do conjunto de dados, o erro de validação cruzada pode às vezes ser menor do que o erro de treinamento.
```
lambda_vec, erro_treinamento, erro_validacao = validationCurve(X_polinomial, y, X_polinomial_validacao, yval)
plt.figure(figsize = (10, 5))
plt.plot(lambda_vec, erro_treinamento, '-o', lambda_vec, erro_validacao, '-o', lw = 2)
plt.legend(['Treinamento', 'Validação Cruzada'])
plt.xlabel('lambda')
plt.ylabel('Erro')
print('lambda\t\tErro de Treinamento\tErro de Validação')
for i in range(len(lambda_vec)):
print('{}\t\t{}\t{}'.format(lambda_vec[i], erro_treinamento[i], erro_validacao[i]))
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import torch
from gluonts.dataset.multivariate_grouper import MultivariateGrouper
from gluonts.dataset.repository.datasets import dataset_recipes, get_dataset
from gluonts.evaluation.backtest import make_evaluation_predictions
from gluonts.evaluation import MultivariateEvaluator
from pts.model.tempflow import TempFlowEstimator
from pts.model.time_grad import TimeGradEstimator
from pts.model.transformer_tempflow import TransformerTempFlowEstimator
from pts import Trainer
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
def plot(target, forecast, prediction_length, prediction_intervals=(50.0, 90.0), color='g', fname=None):
label_prefix = ""
rows = 4
cols = 4
fig, axs = plt.subplots(rows, cols, figsize=(24, 24))
axx = axs.ravel()
seq_len, target_dim = target.shape
ps = [50.0] + [
50.0 + f * c / 2.0 for c in prediction_intervals for f in [-1.0, +1.0]
]
percentiles_sorted = sorted(set(ps))
def alpha_for_percentile(p):
return (p / 100.0) ** 0.3
for dim in range(0, min(rows * cols, target_dim)):
ax = axx[dim]
target[-2 * prediction_length :][dim].plot(ax=ax)
ps_data = [forecast.quantile(p / 100.0)[:,dim] for p in percentiles_sorted]
i_p50 = len(percentiles_sorted) // 2
p50_data = ps_data[i_p50]
p50_series = pd.Series(data=p50_data, index=forecast.index)
p50_series.plot(color=color, ls="-", label=f"{label_prefix}median", ax=ax)
for i in range(len(percentiles_sorted) // 2):
ptile = percentiles_sorted[i]
alpha = alpha_for_percentile(ptile)
ax.fill_between(
forecast.index,
ps_data[i],
ps_data[-i - 1],
facecolor=color,
alpha=alpha,
interpolate=True,
)
# Hack to create labels for the error intervals.
# Doesn't actually plot anything, because we only pass a single data point
pd.Series(data=p50_data[:1], index=forecast.index[:1]).plot(
color=color,
alpha=alpha,
linewidth=10,
label=f"{label_prefix}{100 - ptile * 2}%",
ax=ax,
)
legend = ["observations", "median prediction"] + [f"{k}% prediction interval" for k in prediction_intervals][::-1]
axx[0].legend(legend, loc="upper left")
if fname is not None:
plt.savefig(fname, bbox_inches='tight', pad_inches=0.05)
print(f"Available datasets: {list(dataset_recipes.keys())}")
# exchange_rate_nips, electricity_nips, traffic_nips, solar_nips, wiki-rolling_nips, ## taxi_30min is buggy still
dataset = get_dataset("electricity_nips", regenerate=False)
dataset.metadata
min(2000, int(dataset.metadata.feat_static_cat[0].cardinality))
train_grouper = MultivariateGrouper(max_target_dim=min(2000, int(dataset.metadata.feat_static_cat[0].cardinality)))
test_grouper = MultivariateGrouper(num_test_dates=int(len(dataset.test)/len(dataset.train)),
max_target_dim=min(2000, int(dataset.metadata.feat_static_cat[0].cardinality)))
dataset_train = train_grouper(dataset.train)
dataset_test = test_grouper(dataset.test)
estimator = TimeGradEstimator(
target_dim=int(dataset.metadata.feat_static_cat[0].cardinality),
prediction_length=dataset.metadata.prediction_length,
context_length=dataset.metadata.prediction_length,
cell_type='GRU',
input_size=1484,
freq=dataset.metadata.freq,
loss_type='l2',
scaling=True,
diff_steps=100,
beta_end=0.1,
beta_schedule="linear",
trainer=Trainer(device=device,
epochs=1,
learning_rate=1e-3,
num_batches_per_epoch=100,
batch_size=64,)
)
predictor = estimator.train(dataset_train, num_workers=0)
forecast_it, ts_it = make_evaluation_predictions(dataset=dataset_test,
predictor=predictor,
num_samples=100)
forecasts = list(forecast_it)
targets = list(ts_it)
plot(
target=targets[0],
forecast=forecasts[0],
prediction_length=dataset.metadata.prediction_length,
)
plt.show()
evaluator = MultivariateEvaluator(quantiles=(np.arange(20)/20.0)[1:],
target_agg_funcs={'sum': np.sum})
agg_metric, item_metrics = evaluator(targets, forecasts, num_series=len(dataset_test))
print("CRPS:", agg_metric["mean_wQuantileLoss"])
print("ND:", agg_metric["ND"])
print("NRMSE:", agg_metric["NRMSE"])
print("")
print("CRPS-Sum:", agg_metric["m_sum_mean_wQuantileLoss"])
print("ND-Sum:", agg_metric["m_sum_ND"])
print("NRMSE-Sum:", agg_metric["m_sum_NRMSE"])
```
| github_jupyter |
## Get Suspicious DNS
```
import urllib2
import json
import os
import csv
# getting date from the parent path.
path = os.getcwd().split("/")
date = path[len(path)-1]
dsource = path[len(path)-2]
dpath = '/'.join(['data' if var == 'ipynb' else var for var in path]) + '/'
sconnect = dpath + 'dns_scores.csv'
sconnectbu = dpath + 'dns_scores_bu.csv'
score_tmp = dpath + 'score_tmp.csv'
score_fbk = dpath + 'dns_scores_fb.csv'
def apply_css_to_select(select):
select._css = (
(None, 'height', '90%'),
(None, 'width', '90%'),
('select', 'overflow-x', 'auto'),
('select', 'width', '100%'),
('select', 'margin', 0)
)
try:
import ipywidgets as widgets # For jupyter/ipython >= 1.4
except ImportError:
from IPython.html import widgets
from IPython.display import display, HTML, clear_output, Javascript
def fill_list(list_control,source):
options_list = ['--Select--']
options_list.extend([s for s in source])
list_control.options = options_list
# client panel
client_header = widgets.HTML(value="Client IP")
client_select = widgets.Select(height='90%')
apply_css_to_select(client_select)
client_box = widgets.Box(width='20%', height='100%')
client_box.children = [client_header, client_select]
# query panel
query_header = widgets.HTML(value="Query")
query_select = widgets.Select(height='90%')
apply_css_to_select(query_select)
query_box = widgets.Box(width='60%', height='100%')
query_box.children = [query_header, query_select]
# Actions Panel
actions_header = widgets.HTML(value=" ")
quick_text = widgets.Text(value='', width='100%', placeholder='Quick scoring')
quick_text._css = (
(None, 'width', '100%'),
)
rating_btn = widgets.RadioButtons(description='Rating:', options=['1', '2', '3'], width='100%')
assign_btn = widgets.Button(description='Score', width='45%')
assign_btn.button_style = 'primary'
save_btn = widgets.Button(description='Save', width='45%')
save_btn.button_style = 'primary'
save_btn._css = (
(None, 'margin-left', '10%'),
)
actions_box = widgets.Box(width='20%', height='100%')
actions_box.children = [actions_header,quick_text,rating_btn, assign_btn,save_btn]
scoring_form = widgets.HBox(width='90%', height=250)
scoring_form.children = [client_box,query_box,actions_box]
def data_loader():
us_ips = []
us_dns = []
with open(sconnect, 'r') as f:
reader = csv.DictReader(f, delimiter=',')
for row in reader:
if row['ip_dst'] not in us_ips and row['ip_sev'] == '0':
us_ips.append(row['ip_dst'])
if row['dns_qry_name'] not in us_dns and row['dns_sev'] == '0':
us_dns.append(row['dns_qry_name'])
fill_list(client_select,us_ips)
fill_list(query_select,us_dns)
client_select.value = "--Select--"
query_select.value = "--Select--"
display(Javascript("$('.widget-area > .widget-subarea > *').remove();"))
data_loader()
display(scoring_form)
```
# Update Suspicious DNS
```
import csv
import datetime
import subprocess
def assign_score(b):
score_values = []
scored_threats = []
ip_sev = int(rating_btn.selected_label) if not "--Select--" in client_select.value else ""
dns_sev = int(rating_btn.selected_label) if not "--Select--" in query_select.value else ""
if quick_text.value:
ip = ""
dns = quick_text.value
dns_sev = int(rating_btn.selected_label)
# Loop over every element in query_select widget
score_values = []
for query in query_select.options:
if query.endswith(dns):
# Matching element, create one row
score_values.append((ip,query,ip_sev,dns_sev))
else:
ip = client_select.value if not "--Select--" in client_select.value else ""
dns = query_select.value if not "--Select--" in query_select.value else ""
score_values.append((ip,dns,ip_sev,dns_sev))
with open(sconnect, 'r') as f:
reader = csv.DictReader(f, delimiter=',')
reader.next()
rowct = 0
with open(score_tmp, 'w') as score:
wr = csv.writer(score, delimiter=',',quotechar='|', quoting=csv.QUOTE_MINIMAL)
wr = csv.DictWriter(score, delimiter=',', quoting=csv.QUOTE_NONE, fieldnames=reader.fieldnames)
wr.writeheader()
for row in reader:
for value in score_values:
if row['ip_dst'] == value[0]:
row['ip_sev'] = value[2]
scored_threats.append(row)
rowct += 1
break
if row['dns_qry_name'] == value[1]:
row['dns_sev'] = value[3]
scored_threats.append(row)
rowct += 1
break
wr.writerow(row)
if not os.path.exists(score_fbk):
with open(score_fbk, 'w') as feedback:
wr = csv.DictWriter(feedback, delimiter='\t', quoting=csv.QUOTE_NONE, fieldnames=reader.fieldnames)
wr.writeheader()
with open(score_fbk, 'a') as feedback:
for row in scored_threats:
wr = csv.DictWriter(feedback, delimiter='\t', quoting=csv.QUOTE_NONE, fieldnames=reader.fieldnames)
wr.writerow(row)
clear_output()
print "{0} matching connections scored".format(rowct)
!mv $score_tmp $sconnect
if ip != "--Select--":
display(Javascript("$(\"option[data-value='" + ip +"']\").remove();"))
if quick_text.value:
display(Javascript("$(\"option[data-value$='" + quick_text.value +"']\").remove();"))
elif dns != "--Select--":
display(Javascript("$(\"option[data-value='" + dns +"']\").remove();"))
client_select.value = "--Select--"
query_select.value = "--Select--"
quick_text.value = ""
def save(b):
clear_output()
display(Javascript("$('.widget-area > .widget-subarea > *').remove();"))
data_loader()
display(scoring_form)
display(Javascript('reloadParentData();'))
ml_feedback()
print "Suspicious connects successfully updated"
assign_btn.on_click(assign_score)
save_btn.on_click(save)
def ml_feedback():
dst_name = os.path.basename(sconnect)
str_fb="DSOURCE={0} &&\
FDATE={1} &&\
source /etc/duxbay.conf &&\
usr=$(echo $LUSER | cut -f3 -d'/') &&\
mlnode=$MLNODE &&\
lpath=$LPATH &&\
scp {2} $usr@$mlnode:$lpath/{3}".format(dsource,date,score_fbk,dst_name)
subprocess.call(str_fb, shell=True)
#!cp $sconnectbu $sconnect
```
| github_jupyter |
**Comparison of Batch, Mini-Batch and Stochastic Gradient Descent**
*This notebook displays an animation comparing Batch, Mini-Batch and Stochastic Gradient Descent (introduced in Chapter 4). Thanks to [Daniel Ingram](https://github.com/daniel-s-ingram) who contributed this notebook.*
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/ageron/handson-ml/blob/master/extra_gradient_descent_comparison.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
</table>
**Warning**: this notebook accompanies the 1st edition of the book. Please visit https://github.com/ageron/handson-ml2 for the 2nd edition code, with up-to-date notebooks using the latest library versions.
```
from __future__ import print_function, division, unicode_literals
import numpy as np
%matplotlib nbagg
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
m = 100
X = 2*np.random.rand(m, 1)
X_b = np.c_[np.ones((m, 1)), X]
y = 4 + 3*X + np.random.rand(m, 1)
def batch_gradient_descent():
n_iterations = 1000
learning_rate = 0.05
thetas = np.random.randn(2, 1)
thetas_path = [thetas]
for i in range(n_iterations):
gradients = 2*X_b.T.dot(X_b.dot(thetas) - y)/m
thetas = thetas - learning_rate*gradients
thetas_path.append(thetas)
return thetas_path
def stochastic_gradient_descent():
n_epochs = 50
t0, t1 = 5, 50
thetas = np.random.randn(2, 1)
thetas_path = [thetas]
for epoch in range(n_epochs):
for i in range(m):
random_index = np.random.randint(m)
xi = X_b[random_index:random_index+1]
yi = y[random_index:random_index+1]
gradients = 2*xi.T.dot(xi.dot(thetas) - yi)
eta = learning_schedule(epoch*m + i, t0, t1)
thetas = thetas - eta*gradients
thetas_path.append(thetas)
return thetas_path
def mini_batch_gradient_descent():
n_iterations = 50
minibatch_size = 20
t0, t1 = 200, 1000
thetas = np.random.randn(2, 1)
thetas_path = [thetas]
t = 0
for epoch in range(n_iterations):
shuffled_indices = np.random.permutation(m)
X_b_shuffled = X_b[shuffled_indices]
y_shuffled = y[shuffled_indices]
for i in range(0, m, minibatch_size):
t += 1
xi = X_b_shuffled[i:i+minibatch_size]
yi = y_shuffled[i:i+minibatch_size]
gradients = 2*xi.T.dot(xi.dot(thetas) - yi)/minibatch_size
eta = learning_schedule(t, t0, t1)
thetas = thetas - eta*gradients
thetas_path.append(thetas)
return thetas_path
def compute_mse(theta):
return np.sum((np.dot(X_b, theta) - y)**2)/m
def learning_schedule(t, t0, t1):
return t0/(t+t1)
theta0, theta1 = np.meshgrid(np.arange(0, 5, 0.1), np.arange(0, 5, 0.1))
r, c = theta0.shape
cost_map = np.array([[0 for _ in range(c)] for _ in range(r)])
for i in range(r):
for j in range(c):
theta = np.array([theta0[i,j], theta1[i,j]])
cost_map[i,j] = compute_mse(theta)
exact_solution = np.linalg.inv(X_b.T.dot(X_b)).dot(X_b.T).dot(y)
bgd_thetas = np.array(batch_gradient_descent())
sgd_thetas = np.array(stochastic_gradient_descent())
mbgd_thetas = np.array(mini_batch_gradient_descent())
bgd_len = len(bgd_thetas)
sgd_len = len(sgd_thetas)
mbgd_len = len(mbgd_thetas)
n_iter = min(bgd_len, sgd_len, mbgd_len)
fig = plt.figure(figsize=(10, 5))
data_ax = fig.add_subplot(121)
cost_ax = fig.add_subplot(122)
cost_ax.plot(exact_solution[0,0], exact_solution[1,0], 'y*')
cost_img = cost_ax.pcolor(theta0, theta1, cost_map)
fig.colorbar(cost_img)
def animate(i):
data_ax.cla()
cost_ax.cla()
data_ax.plot(X, y, 'k.')
cost_ax.plot(exact_solution[0,0], exact_solution[1,0], 'y*')
cost_ax.pcolor(theta0, theta1, cost_map)
data_ax.plot(X, X_b.dot(bgd_thetas[i,:]), 'r-')
cost_ax.plot(bgd_thetas[:i,0], bgd_thetas[:i,1], 'r--')
data_ax.plot(X, X_b.dot(sgd_thetas[i,:]), 'g-')
cost_ax.plot(sgd_thetas[:i,0], sgd_thetas[:i,1], 'g--')
data_ax.plot(X, X_b.dot(mbgd_thetas[i,:]), 'b-')
cost_ax.plot(mbgd_thetas[:i,0], mbgd_thetas[:i,1], 'b--')
data_ax.set_xlim([0, 2])
data_ax.set_ylim([0, 15])
cost_ax.set_xlim([0, 5])
cost_ax.set_ylim([0, 5])
data_ax.set_xlabel(r'$x_1$')
data_ax.set_ylabel(r'$y$', rotation=0)
cost_ax.set_xlabel(r'$\theta_0$')
cost_ax.set_ylabel(r'$\theta_1$')
data_ax.legend(('Data', 'BGD', 'SGD', 'MBGD'), loc="upper left")
cost_ax.legend(('Normal Equation', 'BGD', 'SGD', 'MBGD'), loc="upper left")
animation = FuncAnimation(fig, animate, frames=n_iter)
plt.show()
```
| github_jupyter |
# t-SNE Animations
*openTSNE* includes a callback system, with can be triggered every *n* iterations and can also be used to control optimization and when to stop.
In this notebook, we'll look at an example and use callbacks to generate an animation of the optimization. In practice, this serves no real purpose other than being fun to look at.
```
import openTSNE
from examples import utils
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import gzip
import pickle
with gzip.open("data/macosko_2015.pkl.gz", "rb") as f:
data = pickle.load(f)
x = data["pca_50"]
y = data["CellType1"].astype(str)
print("Data set contains %d samples with %d features" % x.shape)
```
We pass a callback that will take the current embedding, make a copy (this is important because the embedding is changed inplace during optimization) and add it to a list. We can also specify how often the callbacks should be called. In this instance, we'll call it at every iteration.
```
embeddings = []
tsne = openTSNE.TSNE(
perplexity=50, metric="cosine", n_jobs=32, verbose=True,
# The embedding will be appended to the list we defined above, make sure we copy the
# embedding, otherwise the same object reference will be stored for every iteration
callbacks=lambda it, err, emb: embeddings.append(np.array(emb)),
# This should be done on every iteration
callbacks_every_iters=1,
)
%time tsne_embedding = tsne.fit(x)
```
Now that we have all the iterations in our list, we need to create the animation. We do this here using matplotlib, which is relatively straightforward. Generating the animation can take a long time, so we will save it as a gif so we can come back to it whenever we want, without having to wait again.
```
%%time
fig, ax = plt.subplots(figsize=(7, 7))
ax.set_xticks([]), ax.set_yticks([])
colors = list(map(utils.MACOSKO_COLORS.get, y))
pathcol = ax.scatter(embeddings[0][:, 0], embeddings[0][:, 1], c=colors, s=1, rasterized=True)
def update(embedding, ax, pathcol):
# Update point positions
pathcol.set_offsets(embedding)
# Adjust x/y limits so all the points are visible
ax.set_xlim(np.min(embedding[:, 0]), np.max(embedding[:, 0]))
ax.set_ylim(np.min(embedding[:, 1]), np.max(embedding[:, 1]))
return [pathcol]
anim = animation.FuncAnimation(
fig, update, fargs=(ax, pathcol), interval=20,
frames=embeddings, blit=True,
)
anim.save("macosko.mp4", dpi=150, writer="ffmpeg")
plt.close()
```
| github_jupyter |
## Download data from Google Drive to colab environment
First we need to mount the Google Drive folder into colab. <br>
Then we copy the data for this exercise to the colab VM and untar it "locally".
```
from google.colab import drive
drive.mount('/content/drive')
!echo "Copying Data Locally (Male/Female Radiograph)"
!tar xf "/content/drive/My Drive/ML4MI_BOOTCAMP_DATA/MaleFemaleRadiograph.tar" --directory /home/
```
## Setup packages and data
First import the packages you'll need. From Keras, we'll need an data generator package, layers package, a package containing optimizres, and a package that builds/configures models.
```
import tensorflow as tf
from tensorflow.keras import optimizers
from tensorflow.keras.models import Model
from tensorflow.keras import layers
from tensorflow.keras.preprocessing.image import ImageDataGenerator
```
Define data location and image dimensions. Data is split into train (50%), validate (25%), and test (25%).
We'll use Kera's ImageDataGenerator method to read in the data. Data (.png files) is sorted into folders with the following structure <br>
>train/<br>
 Class1/<br>
  xx1.png<br>
  xx2.png<br>
  ...<br>
 Class2/<br>
  yy1.png<br>
  yy2.png<br>
test/<br>
 Class1/ ...<br>
 Class2/ ...<br>
validation/<br>
 Class1/ ...<br>
 Class2/ ...<br>
We tell Keras where the directories are. It counts the number of subfolders and makes each one a class.
```
data_home_dir = '/home/MaleFemaleRadiograph/data/'
train_dir = data_home_dir + 'train'
validation_dir = data_home_dir + 'validation'
dims = 256
```
When we define the ImageDataGenerator object, we tell it to normalize the .png images by the max (255)
```
train_datagen = ImageDataGenerator(rescale=1./255)
valid_datagen = ImageDataGenerator(rescale=1./255)
```
Keras will read the files continuously from disk. We tell it where to read, how many to read at a time, what dimensions to resample the images to, and how many image channels there are. These generators will then generate batches of images.
```
train_generator = train_datagen.flow_from_directory(train_dir, batch_size=20, target_size=(dims,dims), class_mode='binary', color_mode='grayscale')
validation_generator = valid_datagen.flow_from_directory(validation_dir,batch_size=20, target_size=(dims,dims), class_mode='binary',color_mode='grayscale')
```
## Build network
First part of the graph is the input, which, at this point, we only need to tell it its shape (we'll define where the inputs come from when we build the model later)
```
img_input = layers.Input(shape=(dims,dims,1), dtype='float32')
```
Now we build our layers of the network. The format is layer_name(_config_info_)(_input_to_layer_).
Try a simple layer with 1 convolution, max pooling, and a fully-connected layer (these are _not_ the best parameters).
```
x = layers.Conv2D(15, (3, 3), strides=(4,4), padding='same')(img_input)
x = layers.Activation('relu')(x)
x = layers.MaxPooling2D((2, 2), strides=None)(x)
x = layers.Flatten()(x) #reshape to 1xN
x = layers.Dense(20, activation='relu')(x)
x = layers.Dense(1, activation='sigmoid')(x) #sigmoid for binary
```
## Configure and train model
We define our model, define the input(s) and output(s).
```
model = Model(inputs=img_input, outputs=x)
```
We then compile it and determine our loss function, our optimizer, and the metrics we want to calculate. This builds the "graph" of our model and computes the functions needed to train it.
```
model.compile(loss = "binary_crossentropy", optimizer = optimizers.RMSprop(learning_rate=1e-5), metrics=["accuracy"])
```
This next steps kicks off the network training. This is where we actually feed the compiled model the data (in batches).
```
history = model.fit(train_generator, steps_per_epoch=130, epochs=15,
validation_data=validation_generator, validation_steps=30)
```
## Evaluate performance
First, let's calculate the performance on our testing dataset
```
test_dir = data_home_dir + 'test'
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_directory(test_dir,batch_size=20, target_size=(dims,dims), class_mode='binary',color_mode='grayscale')
#now evaluate the model using the generator
[test_loss, test_acc] = model.evaluate(test_generator, steps=600/20)
print("Test_acc: "+str(test_acc))
```
Plot the results using matplotlib
```
from matplotlib import pyplot as plt
import numpy as np
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
epochs = range(1,len(acc)+1)
plt.plot(epochs,acc,'bo', label='Training acc')
plt.plot(epochs,val_acc,'b', label='Validation acc')
plt.legend()
plt.show()
```
| github_jupyter |
```
# reload packages
%load_ext autoreload
%autoreload 2
```
### Choose GPU
```
%env CUDA_DEVICE_ORDER=PCI_BUS_ID
%env CUDA_VISIBLE_DEVICES=1
import tensorflow as tf
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
if len(gpu_devices)>0:
tf.config.experimental.set_memory_growth(gpu_devices[0], True)
print(gpu_devices)
tf.keras.backend.clear_session()
```
### dataset information
```
from datetime import datetime
dataset = "mnist"
dims = (28, 28, 1)
num_classes = 10
labels_per_class = 16 # full
batch_size = 128
datestring = datetime.now().strftime("%Y_%m_%d_%H_%M_%S_%f")
datestring = (
str(dataset)
+ "_"
+ str(labels_per_class)
+ "____"
+ datestring
+ '_baseline_augmented'
)
print(datestring)
```
### Load packages
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
from IPython import display
import pandas as pd
import umap
import copy
import os, tempfile
```
### Load dataset
```
from tfumap.load_datasets import load_MNIST, mask_labels
X_train, X_test, X_valid, Y_train, Y_test, Y_valid = load_MNIST(flatten=False)
X_train.shape
if labels_per_class == "full":
X_labeled = X_train
Y_masked = Y_labeled = Y_train
else:
X_labeled, Y_labeled, Y_masked = mask_labels(
X_train, Y_train, labels_per_class=labels_per_class
)
```
### Build network
```
from tensorflow.keras import datasets, layers, models
from tensorflow_addons.layers import WeightNormalization
def conv_block(filts, name, kernel_size = (3, 3), padding = "same", **kwargs):
return WeightNormalization(
layers.Conv2D(
filts, kernel_size, activation=None, padding=padding, **kwargs
),
name="conv"+name,
)
#CNN13
#See:
#https://github.com/vikasverma1077/ICT/blob/master/networks/lenet.py
#https://github.com/brain-research/realistic-ssl-evaluation
lr_alpha = 0.1
dropout_rate = 0.5
num_classes = 10
input_shape = dims
model = models.Sequential()
model.add(tf.keras.Input(shape=input_shape))
### conv1a
name = '1a'
model.add(conv_block(name = name, filts = 128, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
### conv1b
name = '1b'
model.add(conv_block(name = name, filts = 128, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
### conv1c
name = '1c'
model.add(conv_block(name = name, filts = 128, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
# max pooling
model.add(layers.MaxPooling2D(pool_size=(2, 2), strides=2, padding='valid', name="mp1"))
# dropout
model.add(layers.Dropout(dropout_rate, name="drop1"))
### conv2a
name = '2a'
model.add(conv_block(name = name, filts = 256, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha))
### conv2b
name = '2b'
model.add(conv_block(name = name, filts = 256, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
### conv2c
name = '2c'
model.add(conv_block(name = name, filts = 256, kernel_size = (3,3), padding="same"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
# max pooling
model.add(layers.MaxPooling2D(pool_size=(2, 2), strides=2, padding='valid', name="mp2"))
# dropout
model.add(layers.Dropout(dropout_rate, name="drop2"))
### conv3a
name = '3a'
model.add(conv_block(name = name, filts = 512, kernel_size = (3,3), padding="valid"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
### conv3b
name = '3b'
model.add(conv_block(name = name, filts = 256, kernel_size = (1,1), padding="valid"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
### conv3c
name = '3c'
model.add(conv_block(name = name, filts = 128, kernel_size = (1,1), padding="valid"))
model.add(layers.BatchNormalization(name="bn"+name))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelu'+name))
# max pooling
model.add(layers.AveragePooling2D(pool_size=(3, 3), strides=2, padding='valid'))
model.add(layers.Flatten())
model.add(layers.Dense(256, activation=None, name='z'))
model.add(WeightNormalization(layers.Dense(256, activation=None)))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelufc1'))
model.add(WeightNormalization(layers.Dense(256, activation=None)))
model.add(layers.LeakyReLU(alpha=lr_alpha, name = 'lrelufc2'))
model.add(WeightNormalization(layers.Dense(num_classes, activation=None)))
model.summary()
```
### Augmentation
```
#https://github.com/tanzhenyu/image_augmentation/blob/master/image_augmentation/image/image_ops.py
IMAGE_DTYPES = [tf.uint8, tf.float32, tf.float16, tf.float64]
def _check_image_dtype(image):
assert image.dtype in IMAGE_DTYPES, "image with " + str(image.dtype) + " is not supported for this operation"
@tf.function
def invert(image, name=None):
"""Inverts the pixels of an `image`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "invert"):
if image.dtype == tf.uint8:
inv_image = 255 - image
else:
inv_image = 1. - image
return inv_image
@tf.function
def cutout(image, size=16, color=None, name=None):
"""This is an implementation of Cutout as described in "Improved
Regularization of Convolutional Neural Networks with Cutout" by
DeVries & Taylor (https://arxiv.org/abs/1708.04552).
It applies a random square patch of specified `size` over an `image`
and by replacing those pixels with value of `color`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
size: A 0-D int tensor or single int value that is divisible by 2.
color: A single pixel value (grayscale) or tuple of 3 values (RGB),
in case a single value is used for RGB image the value is tiled.
Gray color (128) is used by default.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "cutout"):
image_shape = tf.shape(image)
height, width, channels = image_shape[0], image_shape[1], image_shape[2]
loc_x = tf.random.uniform((), 0, width, tf.int32)
loc_y = tf.random.uniform((), 0, height, tf.int32)
ly, lx = tf.maximum(0, loc_y - size // 2), tf.maximum(0, loc_x - size // 2)
uy, ux = tf.minimum(height, loc_y + size // 2), tf.minimum(width, loc_x + size // 2)
gray = tf.constant(128)
if color is None:
if image.dtype == tf.uint8:
color = tf.repeat(gray, channels)
else:
color = tf.repeat(tf.cast(gray, tf.float32) / 255., channels)
else:
color = tf.convert_to_tensor(color)
color = tf.cast(color, image.dtype)
cut = tf.ones((uy - ly, ux - lx, channels), image.dtype)
top = image[0: ly, 0: width]
between = tf.concat([
image[ly: uy, 0: lx],
cut * color,
image[ly: uy, ux: width]
], axis=1)
bottom = image[uy: height, 0: width]
cutout_image = tf.concat([top, between, bottom], axis=0)
return cutout_image
@tf.function
def solarize(image, threshold, name=None):
"""Inverts the pixels of an `image` above a certain `threshold`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
threshold: A 0-D int / float tensor or int / float value for setting
inversion threshold.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "solarize"):
threshold = tf.cast(threshold, image.dtype)
inverted_image = invert(image)
solarized_image = tf.where(image < threshold, image, inverted_image)
return solarized_image
@tf.function
def solarize_add(image, addition, threshold=None, name=None):
"""Adds `addition` intensity to each pixel and inverts the pixels
of an `image` above a certain `threshold`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
addition: A 0-D int / float tensor or int / float value that is to be
added to each pixel.
threshold: A 0-D int / float tensor or int / float value for setting
inversion threshold. 128 (int) / 0.5 (float) is used by default.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "solarize_add"):
if threshold is None:
threshold = tf.image.convert_image_dtype(tf.constant(128, tf.uint8), image.dtype)
addition = tf.cast(addition, image.dtype)
added_image = image + addition
dark, bright = tf.constant(0, tf.uint8), tf.constant(255, tf.uint8)
added_image = tf.clip_by_value(added_image, tf.image.convert_image_dtype(dark, image.dtype),
tf.image.convert_image_dtype(bright, image.dtype))
return solarize(added_image, threshold)
@tf.function
def posterize(image, num_bits, name=None):
"""Reduces the number of bits used to represent an `image`
for each color channel.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
num_bits: A 0-D int tensor or integer value representing number of bits.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "posterize"):
orig_dtype = image.dtype
image = tf.image.convert_image_dtype(image, tf.uint8)
num_bits = tf.cast(num_bits, tf.int32)
mask = tf.cast(2 ** (8 - num_bits) - 1, tf.uint8)
mask = tf.bitwise.invert(mask)
posterized_image = tf.bitwise.bitwise_and(image, mask)
posterized_image = tf.image.convert_image_dtype(posterized_image, orig_dtype, saturate=True)
return posterized_image
@tf.function
def equalize(image, name=None):
"""Equalizes the `image` histogram. In case of an RGB image, equalization
individually for each channel.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "equalize"):
orig_dtype = image.dtype
image = tf.image.convert_image_dtype(image, tf.uint8, saturate=True)
image = tf.cast(image, tf.int32)
def equalize_grayscale(image_channel):
"""Equalizes the histogram of a grayscale (2D) image."""
bins = tf.constant(256, tf.int32)
histogram = tf.math.bincount(image_channel, minlength=bins)
nonzero = tf.where(tf.math.not_equal(histogram, 0))
nonzero_histogram = tf.reshape(tf.gather(histogram, nonzero), [-1])
step = (tf.reduce_sum(nonzero_histogram) - nonzero_histogram[-1]) // (bins - 1)
# use a lut similar to PIL
def normalize(histogram, step):
norm_histogram = (tf.math.cumsum(histogram) + (step // 2)) // step
norm_histogram = tf.concat([[0], norm_histogram], axis=0)
norm_histogram = tf.clip_by_value(norm_histogram, 0, bins - 1)
return norm_histogram
return tf.cond(tf.math.equal(step, 0),
lambda: image_channel,
lambda: tf.gather(normalize(histogram, step), image_channel))
channels_first_image = tf.transpose(image, [2, 0, 1])
channels_first_equalized_image = tf.map_fn(equalize_grayscale, channels_first_image)
equalized_image = tf.transpose(channels_first_equalized_image, [1, 2, 0])
equalized_image = tf.cast(equalized_image, tf.uint8)
equalized_image = tf.image.convert_image_dtype(equalized_image, orig_dtype)
return equalized_image
@tf.function
def auto_contrast(image, name=None):
"""Normalizes `image` contrast by remapping the `image` histogram such
that the brightest pixel becomes 1.0 (float) / 255 (unsigned int) and
darkest pixel becomes 0.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "auto_contrast"):
orig_dtype = image.dtype
image = tf.image.convert_image_dtype(image, tf.float32)
min_val, max_val = tf.reduce_min(image, axis=[0, 1]), tf.reduce_max(image, axis=[0, 1])
norm_image = (image - min_val) / (max_val - min_val)
norm_image = tf.image.convert_image_dtype(norm_image, orig_dtype, saturate=True)
return norm_image
@tf.function
def blend(image1, image2, factor, name=None):
"""Blends an image with another using `factor`.
Args:
image1: An int or float tensor of shape `[height, width, num_channels]`.
image2: An int or float tensor of shape `[height, width, num_channels]`.
factor: A 0-D float tensor or single floating point value depicting
a weight above 0.0 for combining the example_images.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image1` and `image2`.
"""
_check_image_dtype(image1)
_check_image_dtype(image2)
assert image1.dtype == image2.dtype, "image1 type should exactly match type of image2"
if factor == 0.0:
return image1
elif factor == 1.0:
return image2
else:
with tf.name_scope(name or "blend"):
orig_dtype = image2.dtype
image1, image2 = tf.image.convert_image_dtype(image1, tf.float32), tf.image.convert_image_dtype(image2, tf.float32)
scaled_diff = (image2 - image1) * factor
blended_image = image1 + scaled_diff
blended_image = tf.image.convert_image_dtype(blended_image, orig_dtype, saturate=True)
return blended_image
@tf.function
def sample_pairing(image1, image2, weight, name=None):
"""Alias of `blend`. This is an implementation of SamplePairing
as described in "Data Augmentation by Pairing Samples for Images Classification"
by Inoue (https://arxiv.org/abs/1801.02929).
Args:
image1: An int or float tensor of shape `[height, width, num_channels]`.
image2: An int or float tensor of shape `[height, width, num_channels]`.
weight: A 0-D float tensor or single floating point value depicting
a weight factor above 0.0 for combining the example_images.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image1`.
"""
with tf.name_scope(name or "sample_pairing"):
paired_image = blend(image1, image2, weight)
return paired_image
@tf.function
def color(image, magnitude, name=None):
"""Adjusts the `magnitude` of color of an `image`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
magnitude: A 0-D float tensor or single floating point value above 0.0.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "color"):
tiled_gray_image = tf.image.grayscale_to_rgb(tf.image.rgb_to_grayscale(image))
colored_image = blend(tiled_gray_image, image, magnitude)
return colored_image
@tf.function
def sharpness(image, magnitude, name=None):
"""Adjusts the `magnitude` of sharpness of an `image`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
magnitude: A 0-D float tensor or single floating point value above 0.0.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "sharpness"):
orig_dtype = image.dtype
image = tf.image.convert_image_dtype(image, tf.uint8, saturate=True)
image = tf.cast(image, tf.float32)
blur_kernel = tf.constant([[1, 1, 1],
[1, 5, 1],
[1, 1, 1]], tf.float32, shape=[3, 3, 1, 1]) / 13
blur_kernel = tf.tile(blur_kernel, [1, 1, 3, 1])
strides = [1, 1, 1, 1]
# add extra dimension to image before conv
blurred_image = tf.nn.depthwise_conv2d(image[None, ...], blur_kernel,
strides, padding="VALID")
blurred_image = tf.clip_by_value(blurred_image, 0., 255.)
# remove extra dimension
blurred_image = blurred_image[0]
mask = tf.ones_like(blurred_image)
extra_padding = tf.constant([[1, 1],
[1, 1],
[0, 0]], tf.int32)
padded_mask = tf.pad(mask, extra_padding)
padded_blurred_image = tf.pad(blurred_image, extra_padding)
blurred_image = tf.where(padded_mask == 1, padded_blurred_image, image)
sharpened_image = blend(blurred_image, image, magnitude)
sharpened_image = tf.cast(sharpened_image, tf.uint8)
sharpened_image = tf.image.convert_image_dtype(sharpened_image, orig_dtype)
return sharpened_image
@tf.function
def brightness(image, magnitude, name=None):
"""Adjusts the `magnitude` of brightness of an `image`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
magnitude: A 0-D float tensor or single floating point value above 0.0.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "brightness"):
dark = tf.zeros_like(image)
bright_image = blend(dark, image, magnitude)
return bright_image
@tf.function
def contrast(image, magnitude, name=None):
"""Adjusts the `magnitude` of contrast of an `image`.
Args:
image: An int or float tensor of shape `[height, width, num_channels]`.
magnitude: A 0-D float tensor or single floating point value above 0.0.
name: An optional string for name of the operation.
Returns:
A tensor with same shape and type as that of `image`.
"""
_check_image_dtype(image)
with tf.name_scope(name or "contrast"):
orig_dtype = image.dtype
image = tf.image.convert_image_dtype(image, tf.uint8, saturate=True)
grayed_image = tf.image.rgb_to_grayscale(image)
grayed_image = tf.cast(grayed_image, tf.int32)
bins = tf.constant(256, tf.int32)
histogram = tf.math.bincount(grayed_image, minlength=bins)
histogram = tf.cast(histogram, tf.float32)
mean = tf.reduce_sum(tf.cast(grayed_image, tf.float32)) / tf.reduce_sum(histogram)
mean = tf.clip_by_value(mean, 0.0, 255.0)
mean = tf.cast(mean, tf.uint8)
mean_image = tf.ones_like(grayed_image, tf.uint8) * mean
mean_image = tf.image.grayscale_to_rgb(mean_image)
contrast_image = blend(mean_image, image, magnitude)
contrast_image = tf.image.convert_image_dtype(contrast_image, orig_dtype, saturate=True)
return contrast_image
import tensorflow_addons as tfa
def get_augment(
augment_probability=0.25,
brightness_range=[1e-5, 1.5],
contrast_range=[1e-5, 1],
cutout_range=[0, 0.5],
rescale_range=[0.5, 1],
rescale_range_x_range=0.5,
rescale_range_y_range=0.5,
rotate_range=[-3.14, 3.14],
shear_x_range=[-0.3, 0.3],
shear_y_range=[-0.3, 0.3],
translate_x_range=0.3,
translate_y_range=0.3,
dims=(28, 28, 1),
):
def augment(image, label):
#image = tf.image.random_flip_left_right(image)
random_switch = tf.cast(
tf.random.uniform(
(1,), minval=0, maxval=1 + int(1 / augment_probability), dtype=tf.int32
)[0]
== 1,
tf.bool,
)
if random_switch:
return image, label
# Brightness 0-1
brightness_factor = tf.random.uniform(
(1,),
minval=brightness_range[0],
maxval=brightness_range[1],
dtype=tf.float32,
)[0]
image = brightness(image, brightness_factor)
# rescale 0.5-1
rescale_factor = tf.random.uniform(
(1,), minval=rescale_range[0], maxval=rescale_range[1], dtype=tf.float32
)[0]
image = tf.image.random_crop(image, [dims[0]*rescale_factor, dims[1]*rescale_factor, dims[2]])
image = tf.image.resize(image, [dims[0], dims[1]])
# sqeeze x or y
randint_hor = tf.random.uniform(
(2,),
minval=0,
maxval=tf.cast(rescale_range_x_range * dims[0], tf.int32),
dtype=tf.int32,
)[0]
randint_vert = tf.random.uniform(
(2,),
minval=0,
maxval=tf.cast(rescale_range_y_range * dims[1], tf.int32),
dtype=tf.int32,
)[0]
image = tf.image.resize(
image, (dims[0] + randint_vert * 2, dims[1] + randint_hor * 2)
)
image = tf.image.resize_with_pad(image, dims[0], dims[1])
image = tf.image.resize_with_crop_or_pad(
image, dims[0] + 3, dims[1] + 3
) # crop 6 pixels
image = tf.image.random_crop(image, size=dims)
# rotate -45 45
rotate_factor = tf.random.uniform(
(1,),
minval=rotate_range[0],
maxval=rotate_range[1],
dtype=tf.float32,
)[0]
image = tfa.image.rotate(image, rotate_factor, interpolation="BILINEAR",)
# shear_x -0.3, 3
shear_x_factor = tf.random.uniform(
(1,), minval=shear_x_range[0], maxval=shear_x_range[1], dtype=tf.float32
)[0]
img = tf.repeat(tf.cast(image * 255, tf.uint8), 3, axis=2)
image = tf.cast(tfa.image.shear_x(
img, shear_x_factor, replace=0
)[:,:,:1], tf.float32) / 255
# shear_y -0.3, 3
shear_y_factor = tf.random.uniform(
(1,), minval=shear_x_range[0], maxval=shear_y_range[1], dtype=tf.float32
)[0]
img = tf.repeat(tf.cast(image * 255, tf.uint8), 3, axis=2)
image = tf.cast(tfa.image.shear_y(
img, shear_y_factor, replace=0
)[:,:,:1], tf.float32) / 255.
#print(image.shape)
# translate x -0.3, 0.3
translate_x_factor = tf.random.uniform(
(1,), minval=0, maxval=translate_x_range * 2, dtype=tf.float32
)[0]
# translate y -0.3, 0.3
translate_y_factor = tf.random.uniform(
(1,), minval=0, maxval=translate_y_range * 2, dtype=tf.float32
)[0]
image = tf.image.resize_with_crop_or_pad(
image,
dims[0] + tf.cast(translate_x_factor * dims[0], tf.int32),
dims[1] + tf.cast(translate_x_factor * dims[1], tf.int32),
) # crop 6 pixels
image = tf.image.random_crop(image, size=dims)
# contrast 0-1
contrast_factor = tf.random.uniform(
(1,), minval=contrast_range[0], maxval=contrast_range[1], dtype=tf.float32
)[0]
image = tf.image.adjust_contrast(image, contrast_factor)
image = image - tf.reduce_min(image)
# cutout 0-0.5
cutout_factor = tf.random.uniform(
(1,), minval=cutout_range[0], maxval=cutout_range[1], dtype=tf.float32
)[0]
image = cutout(image, tf.cast(cutout_factor * dims[0], tf.int32))
image = tf.clip_by_value(image, 0.0,1.0)
return image, label
return augment
augment = get_augment(
augment_probability=0.1,
brightness_range=[0.5, 1],
contrast_range=[0.5, 2],
cutout_range=[0, 0.75],
rescale_range=[0.75, 1],
rescale_range_x_range=0.9,
rescale_range_y_range=0.9,
rotate_range=[-0.5, 0.5],
shear_x_range=[-0.3, 0.3],
shear_y_range=[-0.3, 0.3],
translate_x_range=0.2,
translate_y_range=0.2,
dims=(28, 28, 1),
)
nex = 10
for i in range(5):
fig, axs = plt.subplots(ncols=nex +1, figsize=((nex+1)*2, 2))
axs[0].imshow(np.squeeze(X_train[i]), cmap = plt.cm.Greys)
axs[0].axis('off')
for ax in axs.flatten()[1:]:
aug_img = np.squeeze(augment(X_train[i], Y_train[i])[0])
ax.matshow(aug_img, cmap = plt.cm.Greys, vmin=0, vmax=1)
ax.axis('off')
```
### train
```
early_stopping = tf.keras.callbacks.EarlyStopping(
monitor='val_accuracy', min_delta=0, patience=100, verbose=1, mode='auto',
baseline=None, restore_best_weights=True
)
import tensorflow_addons as tfa
opt = tf.keras.optimizers.Adam(1e-4)
opt = tfa.optimizers.MovingAverage(opt)
loss = tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.2, from_logits=True)
model.compile(opt, loss = loss, metrics=['accuracy'])
Y_valid_one_hot = tf.keras.backend.one_hot(
Y_valid, num_classes
)
Y_labeled_one_hot = tf.keras.backend.one_hot(
Y_labeled, num_classes
)
from livelossplot import PlotLossesKerasTF
# plot losses callback
plotlosses = PlotLossesKerasTF()
train_ds = (
tf.data.Dataset.from_tensor_slices((X_labeled, Y_labeled_one_hot))
.repeat()
.shuffle(len(X_labeled))
.map(augment, num_parallel_calls=tf.data.experimental.AUTOTUNE)
.batch(batch_size)
.prefetch(tf.data.experimental.AUTOTUNE)
)
steps_per_epoch = int(len(X_train)/ batch_size)
history = model.fit(
train_ds,
epochs=500,
validation_data=(X_valid, Y_valid_one_hot),
callbacks = [early_stopping, plotlosses],
steps_per_epoch = steps_per_epoch,
)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
submodel = tf.keras.models.Model(
[model.inputs[0]], [model.get_layer('z').output]
)
z = submodel.predict(X_train)
np.shape(z)
reducer = umap.UMAP(verbose=True)
embedding = reducer.fit_transform(z.reshape(len(z), np.product(np.shape(z)[1:])))
plt.scatter(embedding[:, 0], embedding[:, 1], c=Y_train.flatten(), s= 1, alpha = 0.1, cmap = plt.cm.tab10)
z_valid = submodel.predict(X_valid)
np.shape(z_valid)
reducer = umap.UMAP(verbose=True)
embedding = reducer.fit_transform(z_valid.reshape(len(z_valid), np.product(np.shape(z_valid)[1:])))
plt.scatter(embedding[:, 0], embedding[:, 1], c=Y_valid.flatten(), s= 1, alpha = 0.1, cmap = plt.cm.tab10)
fig, ax = plt.subplots(figsize=(10,10))
ax.scatter(embedding[:, 0], embedding[:, 1], c=Y_valid.flatten(), s= 1, alpha = 1, cmap = plt.cm.tab10)
predictions = model.predict(X_valid)
fig, ax = plt.subplots(figsize=(10,10))
ax.scatter(embedding[:, 0], embedding[:, 1], c=np.argmax(predictions, axis=1), s= 1, alpha = 1, cmap = plt.cm.tab10)
Y_test_one_hot = tf.keras.backend.one_hot(
Y_test, num_classes
)
result = model.evaluate(X_test, Y_test_one_hot)
```
### save results
```
# save score, valid embedding, weights, results
from tfumap.paths import MODEL_DIR, ensure_dir
save_folder = MODEL_DIR / 'semisupervised-keras' / dataset / str(labels_per_class) / datestring
ensure_dir(save_folder)
```
#### save weights
```
encoder = tf.keras.models.Model(
[model.inputs[0]], [model.get_layer('z').output]
)
encoder.save_weights((save_folder / "encoder").as_posix())
classifier = tf.keras.models.Model(
[tf.keras.Input(tensor=model.get_layer('weight_normalization').input)], [model.outputs[0]]
)
print([i.name for i in classifier.layers])
classifier.save_weights((save_folder / "classifier").as_posix())
```
#### save score
```
Y_test_one_hot = tf.keras.backend.one_hot(
Y_test, num_classes
)
result = model.evaluate(X_test, Y_test_one_hot)
np.save(save_folder / 'test_loss.npy', result)
```
#### save embedding
```
z = encoder.predict(X_train)
reducer = umap.UMAP(verbose=True)
embedding = reducer.fit_transform(z.reshape(len(z), np.product(np.shape(z)[1:])))
plt.scatter(embedding[:, 0], embedding[:, 1], c=Y_train.flatten(), s= 1, alpha = 0.1, cmap = plt.cm.tab10)
np.save(save_folder / 'train_embedding.npy', embedding)
```
#### save results
```
import pickle
with open(save_folder / 'history.pickle', 'wb') as file_pi:
pickle.dump(history.history, file_pi)
```
| github_jupyter |
## Preparation: start the websocket server workers
Each worker is represented by two parts, a local handle (websocket client worker) and the remote instance that holds the data and performs the computations. The remote part is called a websocket server worker.
So first, we need to create the remote workers. For this, you need to run in a terminal (not possible from the notebook):
```bash
python start_websocket_servers.py
```
## Setting up the websocket client workers
We first need to perform the imports and setup some arguments and variables.
```
%load_ext autoreload
%autoreload 2
import sys
import syft as sy
from syft.workers import WebsocketClientWorker
import torch
from torchvision import datasets, transforms
from syft.frameworks.torch.federated import utils
import run_websocket_client as rwc
args = rwc.define_and_get_arguments(args=[])
use_cuda = args.cuda and torch.cuda.is_available()
torch.manual_seed(args.seed)
device = torch.device("cuda" if use_cuda else "cpu")
print(args)
```
Now let's instantiate the websocket client workers, our local access point to the remote workers.
Note that **this step will fail, if the websocket server workers are not running**.
```
hook = sy.TorchHook(torch)
kwargs_websocket = {"host": "http://ec2-13-233-99-209.ap-south-1.compute.amazonaws.com", "hook": hook, "verbose": args.verbose}
#kwargs_websocket = {"host":"172.20.10.2", "hook": hook, "verbose": args.verbose}
alice = WebsocketClientWorker(id="alice", port=8777, **kwargs_websocket)
bob = WebsocketClientWorker(id="bob", port=8778, **kwargs_websocket)
charlie = WebsocketClientWorker(id="charlie", port=8779, **kwargs_websocket)
workers = [alice, bob, charlie]
print(workers)
federated_train_loader = sy.FederatedDataLoader(
datasets.MNIST(
"../data",
train=True,
download=True,
transform=transforms.Compose(
[transforms.ToTensor(), transforms.Normalize((0.1307,), (0.3081,))]
),
).federate(tuple(workers)),
batch_size=args.batch_size,
shuffle=True,
iter_per_worker=True
)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST(
"../data",
train=False,
transform=transforms.Compose(
[transforms.ToTensor(), transforms.Normalize((0.1307,), (0.3081,))]
),
),
batch_size=args.test_batch_size,
shuffle=True
)
model = rwc.Net().to(device)
print(model)
def train(model, device, federated_train_loader, lr, federate_after_n_batches):
model.train()
nr_batches = federate_after_n_batches
models = {}
loss_values = {}
iter(federated_train_loader) # initialize iterators
batches = rwc.get_next_batches(federated_train_loader, nr_batches)
counter = 0
while True:
print("Starting training round, batches [{}, {}]".format(counter, counter + nr_batches))
data_for_all_workers = True
for worker in batches:
curr_batches = batches[worker]
if curr_batches:
models[worker], loss_values[worker] = rwc.train_on_batches(
worker, curr_batches, model, device, lr
)
else:
data_for_all_workers = False
counter += nr_batches
if not data_for_all_workers:
logger.debug("At least one worker ran out of data, stopping.")
break
model = utils.federated_avg(models)
batches = rwc.get_next_batches(federated_train_loader, nr_batches)
return model
import logging
FORMAT = "%(asctime)s %(levelname)s %(filename)s(l:%(lineno)d) - %(message)s"
LOG_LEVEL = logging.DEBUG
logging.basicConfig(format=FORMAT, level=LOG_LEVEL)
logger = logging.getLogger("main")
for epoch in range(1, args.epochs + 1):
print("Starting epoch {}/{}".format(epoch, args.epochs))
model = train(model, device, federated_train_loader, args.lr, args.federate_after_n_batches)
rwc.test(model, device, test_loader)
```
| github_jupyter |
```
import numpy as np
import sunpy.map
from sunpy.coordinates import frames
from sunpy.net import Fido, attrs as a
from astropy import units as u
from astropy.coordinates import SkyCoord
from sunpy import timeseries as ts
from sunpy.coordinates import get_body_heliographic_stonyhurst
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
```
# 1. Fido Unified Downloader
Fido (Federated Internet Data Obtainer) is a unified interface for the search and retrival of solar physics data regardless of the underlying client or webservice through which the data is obtained, e.g. VSO, JSOC, etc.
In SunPy 1.0 Fido now makes use of an asynchronous download stream through the module `parfive`
## Lets look at how we can search and download some solar physics data
```
#lets search for some SDO/AIA data over some time
results = Fido.search(a.Time('2014/05/15 08:00', '2014/05/15 08:10'), a.Instrument('AIA'))
results
results.file_num
```
Queries to Fido can be make more specific by using other attributes. Lets narrow the search to only look at one wavelength, with a cadence of 1 minute. This makes use of Astropy units
```
results = Fido.search(a.Time('2014/05/15 08:00', '2014/05/15 08:10'), a.Instrument('AIA'),
a.Wavelength(171*u.angstrom), a.vso.Sample(1*u.minute))
results
```
## The search results can then be downloaded via the fetch function
```
aia_files = Fido.fetch(results)
aia_files[0:10]
```
## More detailed searchs with Fido
Fido searches can include searchs from multiple instruments, wavelengths, times etc. This is achieved through the | opertaior which acts as a logical OR operator.
```
results = Fido.search(a.Time('2014/05/15 08:00', '2014/05/15 08:10'), a.Instrument('AIA')|a.Instrument('SWAP'))
# the results can be indexed to access a subset of the search results
results[:,0]
files = Fido.fetch(results[:,0])
```
# 2. Handeling DataTypes (Map and Timeseries)
SunPy provides core data type classes that are designed to provide a consistent interface acress data types (lightcurves and images) as well as data sources from numerous instruments and observations. They handle all of the manipulation necessary to read data in from mission-specific files. The two main datatypes in SunPy are the TimeSeries and Map classes.
## 2.1 TimeSeries
The structure of a TimeSeries consists of times and measurements and the underlying structure is that of a `pandas.DataFrame`. SunPy TimeSeries supports time-series data from a wide range of solar-focussed instruments.
Lets look at an example of using TimeSeries to deal with GOES/XRS observations
```
search_results = Fido.search(a.Time('2013-10-28 00:00','2013-10-28 12:00'),a.Instrument('XRS'))
goes_files = Fido.fetch(search_results[0])
goes_lc = ts.TimeSeries(goes_files)
goes_lc.peek()
```
We can inspect the meta information in the TimeSeries object.
```
goes_lc.meta
```
And inspect the associated units
```
goes_lc.units
```
We can also inspect and manipulate the data
```
goes_lc.data[0:10]
goes_lc.data['xrsb'][0:5] = 10
goes_lc.data['xrsb'][0:10]
```
We can also truncate the data to specific times of interest
```
#new_goes_lc = goes_lc.truncate('2013-10-28 01:00', '2013-10-28 02:30')
#new_goes_lc.peek()
```
## 2.2 Map
The SunPy Map class provides the data type structure to store 2-dimensional data associated with a coordinate system. This allows users to store and manipulate images of the Sun and the heliosphere. Like TimeSeries, Map explicitly supports observations from mulitple instruments, as well as 2D data with associated WCS coordinate information.
Lets look at an example of image analysis with Map
```
aia_map = sunpy.map.Map(aia_files[0])
aia_map.peek()
```
We can inspect the map, the meta data etc
```
aia_map
aia_map.meta
```
One of the key features is the coordinate frame underlying Map through the defined WCS and utilizes the SunPy coordinate subpackage
```
aia_map.coordinate_frame
```
# 3. Coordinates
SunPy coordinates allows us to deal with points in physical space, and the SunPy coordinates subpackage provides definitions of and transformations between several reference frames commonly used in solar physics. This allows us to manipulate Maps and take advantage of WCS Axes for plotting
These reference frames and their associated transformations are implemented using the `astropy.coordinates` subpackage and extend Astropy’s coordinate frame transformation graph to include solar coordinates
```
# # Helioviewer Client
# from sunpy.net.helioviewer import HelioviewerClient
# hv = HelioviewerClient()
# file_stereo = hv.download_jp2('2014/05/15 08:00', observatory="STEREO_A", instrument="SECCHI",detector='COR2' )
# file_stereo
```
## 3.1 Using WCS Axes
```
file_stereo = '2014_05_15__07_54_00_005__STEREO-A_SECCHI_COR2_white-light.jp2'
map_stereo = sunpy.map.Map(file_stereo)
map_stereo
#print the coordinate frame
map_stereo.coordinate_frame
```
We can now take advantage of WCS Axes to plot the STEREO map in its coordinate frame
```
fig = plt.figure(figsize=(15, 15))
ax1 = fig.add_subplot(1, 2, 1, projection=map_stereo)
map_stereo.plot(axes=ax1, vmax=800)
map_stereo.draw_limb()
```
This plot is in the coordinate frame with respect to the observer location of the STEREO spacecraft. A number of bright object are also seen from this field of view. We can search for local astronomical bodies, get their coordinates at the time of the observation and in the coordinate frame of the observer (i.e. as seen from STEREO)
```
# get the location of mars
mars = get_body_heliographic_stonyhurst('mars', map_stereo.date, observer=map_stereo.observer_coordinate)
mars_hpc = mars.transform_to(frames.Helioprojective(observer=map_stereo.observer_coordinate))
```
We can now again make use of WCS axes to plot this coordinate of mars on the same plot as STEREO map
```
fig = plt.figure(figsize=(15, 15))
ax1 = fig.add_subplot(1, 2, 1, projection=map_stereo)
map_stereo.plot(axes=ax1, vmax=800)
ax1.plot_coord(mars_hpc, 's', color='white', fillstyle='none', markersize=12, label='Mars')
plt.legend()
```
## 3.1.1 Positions of planets with respect to observer position
Building upon this, we can search for the other planets with respect to the STEREO spacecraft (i.e. with respect to the observer of this Map)
```
#print the coordinate of the observer location
map_stereo.observer_coordinate
```
We can now get coordinates of other planets in Heliographic stoneyhurst to compare positions to the observer location
```
planet_list = ['earth', 'venus', 'mars', 'mercury']
planet_coord = [get_body_heliographic_stonyhurst(this_planet, time=map_stereo.date) for this_planet in planet_list]
#Plotting the locations of the STEREO instrument and planets all with respect to the Sun
fig = plt.figure(figsize=(10, 10))
ax1 = plt.subplot(2, 1, 1, projection='polar')
plt.polar(np.deg2rad(map_stereo.observer_coordinate.lon), map_stereo.observer_coordinate.radius.to('au'), 'x',label='STEREO')
for this_planet, this_coord in zip(planet_list, planet_coord):
plt.polar(np.deg2rad(this_coord.lon), this_coord.radius, 'o', label=this_planet)
plt.legend()
ax2 = plt.subplot(2, 1, 2)
ax2.plot(map_stereo.observer_coordinate.radius.to('au'), map_stereo.observer_coordinate.lat, 'x', label='STEREO')
for this_planet, this_coord in zip(planet_list, planet_coord):
ax2.plot(this_coord.radius, this_coord.lat, 'o', label=this_planet)
ax2.set_ylabel('Heliographic Latitude [deg]')
plt.legend()
```
## 3.1.2 Venus Transit
Lets look at another example to making use of SunPy coordinates and WCS axes to look at the Venus transit observation from SDO/AIA
```
file_venus = '20120606_040731_UTC.0041.fits'
map_venus = sunpy.map.Map(file_venus)
map_venus.peek()
from astropy.coordinates import solar_system_ephemeris
solar_system_ephemeris.set('de432s')
```
In this plot we can clearly see an object transiting the Sun. We can now get the position of Venus in the coordinate frame of the observer (SDO) and plot the location of Venus on this same plot
```
# now get venus position
venus = get_body_heliographic_stonyhurst('venus', map_venus.date, observer=map_venus.observer_coordinate)
venus_hpc = venus.transform_to(frames.Helioprojective(observer=map_venus.observer_coordinate))
fov = 100 * u.arcsec
top_right = SkyCoord(venus_hpc.Tx + fov, venus_hpc.Ty + fov, frame=map_venus.coordinate_frame)
bottom_left = SkyCoord(venus_hpc.Tx - fov, venus_hpc.Ty - fov, frame=map_venus.coordinate_frame)
smap = map_venus.submap(top_right, bottom_left)
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1, projection=smap)
# now get venus position
smap.plot(axes=ax)
smap.draw_limb()
ax.plot_coord(venus_hpc, 'x', color='white')
log.info('Test')
```
| github_jupyter |
<font size="+5">#03 | Model Selection. Decision Tree vs Support Vector Machines vs Logistic Regression</font>
<div class="alert alert-warning">
<ul>
<li>
<b>Python</b> + <b>Data Science</b> Tutorials in ↓
<ul>
<li>
<a href="https://www.youtube.com/c/PythonResolver?sub_confirmation=1"
>YouTube</a
>
</li>
<li>
<a href="https://blog.pythonresolver.com/">Blog</a>
</li>
<li>
<a href="https://github.com/jsulopz/00-python-resolver-discipline">GitHub</a>
</li>
</ul>
</li>
<li>
Author: <a href="https://twitter.com/jsulopz"><b>@jsulopz</b></a>
</li>
</ul>
</div>
<a href="https://colab.research.google.com/github/jsulopz/machine-learning/blob/main/03_Model%20Selection.%20Decision%20Tree%20vs%20Support%20Vector%20Machines%20vs%20Logistic%20Regression/03_model-selection_session_solution.ipynb">
<img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/>
</a>
# Discipline to Search Solutions in Google
> Apply the following steps when **looking for solutions in Google**:
>
> 1. **Necesity**: How to load an Excel in Python?
> 2. **Search in Google**: by keywords
> - `load excel python`
> - ~~how to load excel in python~~
> 3. **Solution**: What's the `function()` that loads an Excel in Python?
> - A Function to Programming is what the Atom to Phisics.
> - Every time you want to do something in programming
> - **You will need a `function()`** to make it
> - Theferore, you must **detect parenthesis `()`**
> - Out of all the words that you see in a website
> - Because they indicate the presence of a `function()`.
# Load the Data
Load the dataset from [CIS](https://www.cis.es/cis/opencms/ES/index.html) executing the lines of code below:
> - The goal of this dataset is
> - To predict `internet_usage` of **people** (rows)
> - Based on their **socio-demographical characteristics** (columns)
# Build & Compare Models
## `DecisionTreeClassifier()` Model in Python
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/7VeUPuFGJHk" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
```
> - Build the model `model.fit()`
> - And see how good it is `model.score()`
## `SVC()` Model in Python
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/efR1C6CvhmE" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
```
> - Build the model `model.fit()`
> - And see how good it is `model.score()`
## `LogisticRegression()` Model in Python
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/yIYKR4sgzI8" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
```
> - Build the model `model.fit()`
> - And see how good it is `model.score()`
# Function to Automate Lines of Code
> - We repeated all the time the same code:
```python
model.fit()
model.score()
```
> - Why not turning the lines into a `function()`
> - To automate the process?
> - In a way that you would just need
```python
calculate_accuracy(model=dt)
calculate_accuracy(model=svm)
calculate_accuracy(model = lr)
```
> - To calculate the `accuracy`
## Make a Procedure Sample for `DecisionTreeClassifier()`
## Automate the Procedure into a `function()`
**Code Thinking**
> 1. Think of the functions `result`
> 2. Store that `object` to a variable
> 3. `return` the `result` at the end
> 4. **Indent the body** of the function to the right
> 5. `def`ine the `function():`
> 6. Think of what's gonna change when you execute the function with `different models`
> 7. Locate the **`variable` that you will change**
> 8. Turn it into the `parameter` of the `function()`
## `DecisionTreeClassifier()` Accuracy
## `SVC()` Accuracy
## `LogisticRegression()` Accuracy
# Which is the Best Model?
> Which model has the **highest accuracy**?
## University Access Exams Analogy
> Let's **imagine**:
>
> 1. You have a `math exam` on Saturday
> 2. Today is Monday
> 3. You want to **calculate if you need to study more** for the math exam
> 4. How do you calibrate your `math level`?
> 5. Well, you've got **100 questions `X` with 100 solutions `y`** from past years exams
> 6. You may study the 100 questions with 100 solutions `fit(questions, solutions)`
> 7. Then, you may do a `mock exam` with the 100 questions `predict(questions)`
> 8. And compare `your_solutions` with the `real_solutions`
> 9. You've got **90/100 correct answers** `accuracy` in the mock exam
> 10. You think you are **prepared for the maths exam**
> 11. And when you do **the real exam on Saturday, the mark is 40/100**
> 12. Why? How could have we prevented this?
> 13. **Solution**: separate the 100 questions in
> - `70 train` to study & `30 test` for the mock exam.
# `train_test_split()` the Data
> 1. **`fit()` the model with `Train Data`**
>
> - `model.fit(70%questions, 70%solutions)`
> 2. **`.predict()` answers with `Test Data` (mock exam)**
>
> - `your_solutions = model.predict(30%questions)`
> **3. Compare `your_solutions` with `correct answers` from mock exam**
>
> - `your_solutions == real_solutions`?
# Optimize All Models & Compare Again
## Make a Procedure Sample for `DecisionTreeClassifier()`
## Automate the Procedure into a `function()`
**Code Thinking**
> 1. Think of the functions `result`
> 2. Store that `object` to a variable
> 3. `return` the `result` at the end
> 4. **Indent the body** of the function to the right
> 5. `def`ine the `function():`
> 6. Think of what's gonna change when you execute the function with `different models`
> 7. Locate the **`variable` that you will change**
> 8. Turn it into the `parameter` of the `function()`
## `DecisionTreeClassifier()` Accuracy
## `SVC()` Accuracy
## `LogisticRegression()` Accuracy
# Which is the Best Model with `train_test_split()`?
> Which model has the **highest accuracy**?
# Reflect
> - Banks deploy models to predict the **probability for a customer to pay the loan**
> - If the Bank used the `DecisionTreeClassifier()` instead of the `LogisticRegression()`
> - What would have happened?
> - Is `train_test_split()` always required to compare models?
| github_jupyter |
Univariate analysis of block design, one condition versus rest, single subject
==============================================================================
Authors: Bertrand Thirion, Elvis Dohmatob , Christophe Pallier, 2015--2017
Modified: Ralf Schmaelzle, 2019
In this tutorial, we compare the fMRI signal during periods of auditory stimulation
versus periods of rest, using a General Linear Model (GLM). We will
use a univariate approach in which independent tests are performed at
each single-voxel.
The dataset comes from experiment conducted at the FIL by Geriant Rees
under the direction of Karl Friston. It is provided by FIL methods
group which develops the SPM software.
According to SPM documentation, 96 acquisitions were made (RT=7s), in
blocks of 6, giving 16 42s blocks. The condition for successive blocks
alternated between rest and auditory stimulation, starting with rest.
Auditory stimulation was bi-syllabic words presented binaurally at a
rate of 60 per minute. The functional data starts at acquisiton 4,
image fM00223_004.
The whole brain BOLD/EPI images were acquired on a modified 2T Siemens
MAGNETOM Vision system. Each acquisition consisted of 64 contiguous
slices (64x64x64 3mm x 3mm x 3mm voxels). Acquisition took 6.05s, with
the scan to scan repeat time (RT) set arbitrarily to 7s.
This analyse described here is performed in the native space, on the
original EPI scans without any spatial or temporal preprocessing.
(More sensitive results would likely be obtained on the corrected,
spatially normalized and smoothed images).
How does this "jupyter-thing" work?
It is very easy. A notebook basically consists of cells that can be of two main kinds: Code cells and comment cells. The one you are reading is a comment cell, the one-line-cell below ("Import modules") as well, then comes a code cell. The idea behind this is called literate programming, that is the comment-cells explain stuff in natural language, and the code cells exectute some computer code (and can also display output).
How do I get ahead?
To click through cells, you can either click the 'Run' button above, or you can click in to a cell and press (SHIFT + ENTER). Try it now, and then go....
## Import modules
```
import os, sys, nibabel
!pip install nistats
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from os.path import join
import seaborn as sns
from nilearn import plotting, datasets, image
from nilearn.image import concat_imgs
from nilearn.input_data import NiftiSpheresMasker
from nistats.first_level_model import FirstLevelModel
from nistats.datasets import fetch_spm_auditory
from nistats.reporting import plot_design_matrix
from nilearn.plotting import plot_stat_map, plot_anat, plot_img
from nibabel.affines import apply_affine
```
If you have made it until here, you will see some confusing code about requirements or so that you can safely ignore. Just go ahead by shift-enter-clicking or pressing the run-button
Retrieving the data
-------------------
The next line of code will fetch a whole fMRI dataset from a server into our litte "data-oven" (the Binder platform runs a virtual machine that runs this jupyter notebook).
```
subject_data = fetch_spm_auditory()
```
We can list the filenames of the functional images by just entering the name of the data-structure that holds the fetched data.
```
subject_data
```
As you can see, there are a ton of files. Specifically, we downloaded functional files (from the fMRI experiment), one structural file (a high-res image of the person's brain), and some events (with info what was presented when).
Display the first functional image:
There will be some ugly red error code. Just re-run the cell again and it should be gone!
```
%matplotlib inline
plot_img(subject_data.func[0]);
```
***Exercise:***
in the next cell, please change the code so you display another than the first (in python this is counted as 0-th) image:
```
plot_img(subject_data.func[0]);
```
You can also display the subject's anatomical image:
```
plot_anat(subject_data.anat);
```
Next, we concatenate all the "3D EPI images" (i.e. the functionals) into a single 4D image.
```
fmri_img = concat_imgs(subject_data.func)
print(fmri_img.shape)
```
***Exercise:***
What does this output of the above cell actually mean? Explain to yourself by drawing and to your neighbor.
Let's plot the data from one voxel, that is retrieve the over-time activity series from one spot of the brain
```
data_from_one_voxel = fmri_img.get_data()[22,30,26,:] #22,30,26
plt.figure(figsize = (10,2))
plt.plot(data_from_one_voxel);
plt.xlabel('Time (volumes)');
plt.ylabel('fMRI Signal');
```
***Exercise:***
Next, please try to insert a new empty cell below (so you also learn how to edit this notebook)
Once you have successfully done that, copy the code for displaying one voxel's data into the new cell and play around by changing the voxel-coordinates. Try this multiple times. Can you find a particular wild voxel?
To show you how we can do math with these data, we can, for instance, average all the EPI images into one in order to create a background image that will be used to display the activations:
```
mean_img = image.mean_img(fmri_img)
plot_anat(mean_img);
```
Specifying the experimental paradigm
------------------------------------
Our goal is to actually analyze this single-person experiment (auditory stimuluation).
***Exercise:*** What do you expect to find?
For the actual analysis, we must provide a description of the experiment, that is, define the timing of the auditory stimulation and rest periods. According to the documentation of the dataset, there were 16 42s blocks --- in
which 6 scans were acquired --- alternating between rest and auditory stimulation, starting with rest. We use standard python functions to create a pandas.DataFrame object that specifies the timings:
(you don't need to understand the details of this yet - just execute the code and try to get the gist)
```
tr = 7.
slice_time_ref = 0.
n_scans = 96
epoch_duration = 6 * tr # duration in seconds
conditions = ['rest', 'active'] * 8
n_blocks = len(conditions)
duration = epoch_duration * np.ones(n_blocks)
onset = np.linspace(0, (n_blocks - 1) * epoch_duration, n_blocks)
events = pd.DataFrame({'onset': onset, 'duration': duration, 'trial_type': conditions})
```
The ``events`` object contains the information for the design:
```
print(events)
```
***Exercise:*** What kind of design are we dealing with?
Performing the GLM analysis
---------------------------
We need to construct a *design matrix* using the timing information provided by the ``events`` object. The design matrix contains regressors of interest as well as regressors of non-interest modeling temporal drifts.
With all these ingredients, we then create a ``FirstLevelModel`` object.
```
frame_times = np.linspace(0, (n_scans - 1) * tr, n_scans)
drift_model = 'Cosine'
period_cut = 4. * epoch_duration
hrf_model = 'glover + derivative'
fmri_glm = FirstLevelModel(tr, slice_time_ref, noise_model='ar1',
standardize=False, hrf_model=hrf_model,
drift_model=drift_model)# period_cut=period_cut) #not nice, but didn't work otherwise
```
Drumroll: Now, we fit the FirstLevelModel 'fmri_glm' to the data (4D dataset). Note that this is also called a mass-univariate analysis, that is we fit this model for each voxel individually, getting a beta-coefficient for each voxel. This is also why it takes a bit.
```
fmri_glm = fmri_glm.fit(fmri_img, events)
```
One can inspect the design matrix (rows represent time, and
columns contain the predictors):
```
design_matrix = fmri_glm.design_matrices_[0]
fig, ax1 = plt.subplots(figsize=(6, 8), nrows=1, ncols=1)
plot_design_matrix(design_matrix, ax= ax1, rescale= True);
```
***Exercise:*** What does this matrix exactly mean? What are the critical columns that we care about in order to test our hypothesis? Hey, speaking of this, what's our hypthesis, actually? Explain to your neighbor or create new cell (change it so it is a comment cell) and write the answer in (it's better to be explicit about the hypothesis)!
The first column contains the expected reponse profile of regions which are
sensitive to the auditory stimulation. Let's grab that from the design matrix and plot it:
```
plt.plot(design_matrix['active'])
plt.xlabel('scan')
plt.title('Expected Auditory Response')
plt.show()
```
***Exercise:*** Bonus question: Why is this expected timecourse so oddly shaped and not just a on-off-pattern?
Detecting voxels with significant effects
-----------------------------------------
To access the estimated coefficients (Betas of the GLM model), we created constrasts with a single '1' in each of the columns.
(Again, you don't have to understand the details of the code, just try to follow the overall gist for now)
```
contrast_matrix = np.eye(design_matrix.shape[1])
contrasts = dict([(column, contrast_matrix[i])
for i, column in enumerate(design_matrix.columns)])
"""
contrasts::
{
'active': array([ 1., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.]),
'active_derivative': array([ 0., 1., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0.]),
'constant': array([ 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 1.]),
'drift_1': array([ 0., 0., 0., 0., 1., 0., 0., 0., 0., 0., 0., 0.]),
'drift_2': array([ 0., 0., 0., 0., 0., 1., 0., 0., 0., 0., 0., 0.]),
'drift_3': array([ 0., 0., 0., 0., 0., 0., 1., 0., 0., 0., 0., 0.]),
'drift_4': array([ 0., 0., 0., 0., 0., 0., 0., 1., 0., 0., 0., 0.]),
'drift_5': array([ 0., 0., 0., 0., 0., 0., 0., 0., 1., 0., 0., 0.]),
'drift_6': array([ 0., 0., 0., 0., 0., 0., 0., 0., 0., 1., 0., 0.]),
'drift_7': array([ 0., 0., 0., 0., 0., 0., 0., 0., 0., 0., 1., 0.]),
'rest': array([ 0., 0., 1., 0., 0., 0., 0., 0., 0., 0., 0., 0.]),
'rest_derivative': array([ 0., 0., 0., 1., 0., 0., 0., 0., 0., 0., 0., 0.])}
"""
```
We can then compare the two conditions 'active' and 'rest' by
generating the relevant contrast that we can use to ~~ isolate ~~ the psychological process
(there are some tricky problems with this notion of process isolation and we're glossing over a lot of detail here, but for our purpose to localize brain regions associated with sound-stimulation this isn't so critical)
```
active_minus_rest = contrasts['active'] - contrasts['rest']
eff_map = fmri_glm.compute_contrast(active_minus_rest,
output_type='effect_size')
z_map = fmri_glm.compute_contrast(active_minus_rest,
output_type='z_score')
```
Plot thresholded z scores map
```
plot_stat_map(z_map, bg_img=mean_img, threshold=3.0,
display_mode='z', cut_coords=3, black_bg=True,
title='Active minus Rest (Z>3)');
```
***Exercise:*** Tada, what do you see? Where in the brain are we? What do you know about this territory?
***Exercise:*** Below, you can also get a cooler view that you can surf in 3D. Try to do that and find whether there are other hotspots.
```
plotting.view_img(z_map, bg_img=mean_img, threshold=3., title="Active vs. Rest contrast")
```
We can use ``nibabel.save`` to save the effect and zscore maps to the disk (Unfortunately, since we're running this in the cloud, you cannot get the data, but you could download the notebook, run the same code on your computer, and then you have it forever).
```
outdir = 'results'
if not os.path.exists(outdir):
os.mkdir(outdir)
nibabel.save(z_map, join('results', 'active_vs_rest_z_map.nii'))
nibabel.save(eff_map, join('results', 'active_vs_rest_eff_map.nii'))
```
Extract the signal from a voxels
--------------------------------
We search for the voxel with the larger z-score and plot the signal
(warning: double dipping!)
```
# Find the coordinates of the peak
values = z_map.get_fdata()
coord_peaks = np.dstack(np.unravel_index(np.argsort(values.ravel()),
values.shape))[0, 0, :]
coord_mm = apply_affine(z_map.affine, coord_peaks)
```
We create a masker for the voxel (allowing us to detrend the signal) and extract the time course.
(Again, no details needed here - we're just using a tool to extract from the large 4d-dataset the activity in a region around where we found something - this will then be put into the variable 'sig')
```
mask = NiftiSpheresMasker([coord_mm], radius=3,
detrend=True, standardize=True,
high_pass=None, low_pass=None, t_r=7.)
sig = mask.fit_transform(fmri_img)
```
***Exercise:*** Create a new cell below, enter 'sig' and press enter. What does that mean?
Let's plot the signal and the theoretical response
```
plt.plot(frame_times, sig, label='voxel %d %d %d' % tuple(coord_mm))
plt.plot(design_matrix['active'], color='red', label='model')
plt.xlabel('scan')
plt.legend()
plt.show()
```
***Exercise:*** What do you see? What does it mean? Why isn't the result perfect?
***Exercise:*** Congratulations, you have just completed a first analysis. Try to lean back and think about the steps you did!
What would be the next steps?
| github_jupyter |
Ben and I moved to New Hampshire in August. We moved into a small cottage on his parents' new house property, and figured that because we weren't going to be paying rent we could splurge on house expenses. But we went kinda wild, so let's see if the gamble was right...
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
df = pd.read_csv('Claire and Ben moving costs - Sheet1.csv')
df.columns = ['_'.join(c.lower().split()) for c in df.columns]
df.head()
```
First question, and the goal of this whole thing: are we settled up, or did one of us pay way more than the other?
```
df['how_much'] = (
df['how_much'].str.strip('$').str.replace(',', '').astype(float)
)
df.groupby('who_paid').sum()
```
Wow, at the beginning we thought that Ben would have paid way more because he got all the Amazon stuff. But it looks like I've now outpaced him -- probably due to the couple of large expenses I paid for (couch, our part of the hut tub).
Let's break it down by category and see how things stack up.
```
(df.groupby(['category', 'who_paid']).sum()
.reset_index() # get category, who paid, and total amount in columns
.pivot(index='category', columns='who_paid', values='how_much') # reshape to get categories per row, person per columns; for plotting with pandas
.plot(kind='bar') #, x='category', y='how_much', color='who_paid')
)
```
Sure enough, I paid the \\$3000 for the hot tub/Ben's mom's bike and ~\\$2000 for the couch. If we remove these, how do we stack up?
## Moving expenses only
```
# Keep just appliances, house, kitchen, and moving expenses
keep_cats = ['appliances', 'house', 'kitchen', 'moving']
print(df.query('category == @keep_cats').groupby('who_paid').sum())
(df.query('category == @keep_cats')
.groupby(['category', 'who_paid']).sum()
.reset_index() # get category, who paid, and total amount in columns
.pivot(index='category', columns='who_paid', values='how_much') # reshape to get categories per row, person per columns; for plotting with pandas
.plot(kind='bar') #, x='category', y='how_much', color='who_paid')
)
```
Ok, so if we just look at the things that would be more directly considered as moving expenses, then Ben spent about \\$1000 more than I did.
It's interesting that we did end up just about equaling out on moving expenses. We tried to go back and forth for the two weekends that we spent moving, but we weren't sure if it had evened out. Also wow moving is expensive -- a little over a thousand dollars to buy all the boxes and moving supplies, and to rent a truck for two weekends. I wonder how much hiring a moving crew would have cost, especially considering that spending two full weekends moving was _quite_ miserable and would be worth a lot of money to not have to repeat. But good to know how much it all added up to for next time we move, when we've both agreed to hire someone to help!
Out of curiosity, how much of these expenses were the truck vs. other things?
```
df.query('category == "moving"')
```
Looks like about half of it was the moving truck: ~\\$300 each weekend, looks like.
## An oxo tangent
Lol we spent almost as much on kitchen knick knacks than we did on our appliances! For appliances, we bought a fridge, toaster oven, microwave, and induction stove. But I guess that has no chance against the millions of Oxo and other kitchen gadgets we got!
```
keep_cats = ['appliances', 'kitchen']
df.query('category == @keep_cats').groupby('category').sum()
```
Speaking of Oxo... I joked to Ben that he was buying out every single Oxo item and that when we did this analysis we should see what percent of our expenses were Oxo. Unfortunately, getting that info would take way too long becuase we used his Amazon statements to get these expenses, and they don't show everything itemized. So instead, let's see how many of the kitchen-related orders included something Oxo (I made sure to write that down).
```
df[df['for_what'].str.lower().str.contains('oxo')]
```
Ahahah okay but actually a lot of these were just straight-up Oxo. Let's remove the one that contains the toaster oven (because the majority of that expense was the toaster oven itself), and see how much the rest adds up to.
```
# Oxo-only expenses
print(f"Oxo-only expenses = ${round(df[df['for_what'].str.lower() == 'oxo'].sum()['how_much'])}")
print(f"Expenses including Oxo = $ {round(df[df['for_what'].str.lower().str.contains('oxo')].sum()['how_much'] - 349.54)}")
```
So our Oxo expenses are somewhere between \\$100 and \\$400. Not bad, actually -- way less than I would have guessed!
Ok last Oxo-focused question: what proportion of our kitchen-related expenses included Oxo products?
```
# Get total number of kitchen expenses
n_kitchen = df.query('category == "kitchen"').shape[0]
# Get number of expenses containing oxo
n_oxo = df['for_what'].str.lower().str.contains('oxo').sum()
print(f"Fraction of kitchen expenses containing oxo = {round(n_oxo/n_kitchen, 2)} ({n_oxo}/{n_kitchen})")
```
Nice. Over a third of our kitchen expenses included something Oxo. We really just couldn't get enough!
## Back to expenses
Ok, back to the overall expenses. One thing sticks out -- what on earth did we spend over \\$500 on "house" things for?
```
df.query('category == "house"')['for_what'].unique()
```
Oh god that's right we spent \\$50 on deodorant and then a bunch on masks to keep us safe while we cleaned out a box that used to serve as some sort of pump (the house we live in used to be a jeweller's workshop) but had since become inhabited by many birds and wasps who filled the entire box with hay and dust.
# Conclusion
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import glob
import nibabel as nib
import os
import time
import pandas as pd
import numpy as np
import cv2
from skimage.transform import resize
from mricode.utils import log_textfile
from mricode.utils import copy_colab
from mricode.utils import return_iter
from mricode.utils import return_csv
from mricode.models.SimpleCNN import SimpleCNN
from mricode.models.DenseNet import MyDenseNet
import tensorflow as tf
from tensorflow.keras.layers import Conv3D
from tensorflow import nn
from tensorflow.python.ops import nn_ops
from tensorflow.python.framework import tensor_shape
from tensorflow.python.keras.engine.base_layer import InputSpec
from tensorflow.python.keras.utils import conv_utils
tf.__version__
tf.test.is_gpu_available()
path_output = './output/'
path_tfrecords = '/data2/res64/down/'
path_csv = '/data2/csv/'
filename_res = {'train': 'intell_residual_train.csv', 'val': 'intell_residual_valid.csv', 'test': 'intell_residual_test.csv'}
filename_final = filename_res
sample_size = 'allimages'
batch_size = 8
onlyt1 = False
modelname = 'runAllImages64_DenseNet_T1T2_site16_norm_20e_tfexplain_all_simple'
Model = SimpleCNN
t1_mean=1.3779395849814497
t1_std=3.4895845243139503
t2_mean=2.22435586968901
t2_std=5.07708743178319
ad_mean=1.3008901218593748e-05
ad_std=0.009966655860940228
fa_mean=0.0037552628409334037
fa_std=0.012922319568740915
md_mean=9.827903909139596e-06
md_std=0.009956973204022659
rd_mean=8.237404999587111e-06
rd_std=0.009954672598675338
train_iter, val_iter, test_iter = return_iter(path_tfrecords, sample_size, batch_size, onlyt1=onlyt1)
if False:
t1_mean = 0.
t1_std = 0.
t2_mean = 0.
t2_std = 0.
ad_mean = 0.
ad_std = 0.
fa_mean = 0.
fa_std = 0.
md_mean = 0.
md_std = 0.
rd_mean = 0.
rd_std = 0.
n = 0.
for b in train_iter:
t1_mean += np.mean(b['t1'])
t1_std += np.std(b['t1'])
t2_mean += np.mean(b['t2'])
t2_std += np.std(b['t2'])
a = np.asarray(b['ad'])
a = a.copy()
a[np.isnan(a)] = 0
ad_mean += np.mean(a)
ad_std += np.std(a)
a = np.asarray(b['fa'])
a = a.copy()
a[np.isnan(a)] = 0
fa_mean += np.mean(a)
fa_std += np.std(a)
a = np.asarray(b['md'])
a = a.copy()
a[np.isnan(a)] = 0
md_mean += np.mean(a)
md_std += np.std(a)
a = np.asarray(b['rd'])
a = a.copy()
a[np.isnan(a)] = 0
rd_mean += np.mean(a)
rd_std += np.std(a)
n += np.asarray(b['t1']).shape[0]
t1_mean /= n
t1_std /= n
t2_mean /= n
t2_std /= n
ad_mean /= n
ad_std /= n
fa_mean /= n
fa_std /= n
md_mean /= n
md_std /= n
rd_mean /= n
rd_std /= n
t1_mean, t1_std, t2_mean, t2_std, ad_mean, ad_std, fa_mean, fa_std, md_mean, md_std, rd_mean, rd_std
train_df, val_df, test_df, norm_dict = return_csv(path_csv, filename_final, False)
norm_dict
cat_cols = {'female': 2, 'race.ethnicity': 5, 'high.educ_group': 4, 'income_group': 8, 'married': 6}
num_cols = [x for x in list(val_df.columns) if '_norm' in x]
def calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict):
for col in num_cols:
tmp_col = col
tmp_std = norm_dict[tmp_col.replace('_norm','')]['std']
tmp_y_true = tf.cast(y_true[col], tf.float32).numpy()
tmp_y_pred = np.squeeze(y_pred[col].numpy())
if not(tmp_col in out_loss):
out_loss[tmp_col] = np.sum(np.square(tmp_y_true-tmp_y_pred))
else:
out_loss[tmp_col] += np.sum(np.square(tmp_y_true-tmp_y_pred))
if not(tmp_col in out_acc):
out_acc[tmp_col] = np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
else:
out_acc[tmp_col] += np.sum(np.square((tmp_y_true-tmp_y_pred)*tmp_std))
for col in list(cat_cols.keys()):
tmp_col = col
if not(tmp_col in out_loss):
out_loss[tmp_col] = tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
else:
out_loss[tmp_col] += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y_true[col]), tf.squeeze(y_pred[col])).numpy()
if not(tmp_col in out_acc):
out_acc[tmp_col] = tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
else:
out_acc[tmp_col] += tf.reduce_sum(tf.dtypes.cast((y_true[col] == tf.argmax(y_pred[col], axis=-1)), tf.float32)).numpy()
return(out_loss, out_acc)
def format_output(out_loss, out_acc, n, cols, print_bl=False):
loss = 0
acc = 0
output = []
for col in cols:
output.append([col, out_loss[col]/n, out_acc[col]/n])
loss += out_loss[col]/n
acc += out_acc[col]/n
df = pd.DataFrame(output)
df.columns = ['name', 'loss', 'acc']
if print_bl:
print(df)
return(loss, acc, df)
@tf.function
def train_step(X, y, model, optimizer, cat_cols, num_cols):
with tf.GradientTape() as tape:
predictions = model(X)
i = 0
loss = tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for i in range(1,len(num_cols)):
loss += tf.keras.losses.MSE(tf.cast(y[num_cols[i]], tf.float32), tf.squeeze(predictions[num_cols[i]]))
for col in list(cat_cols.keys()):
loss += tf.keras.losses.SparseCategoricalCrossentropy()(tf.squeeze(y[col]), tf.squeeze(predictions[col]))
gradients = tape.gradient(loss, model.trainable_variables)
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
with tf.control_dependencies(mean_std):
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
return(y, predictions, loss)
@tf.function
def test_step(X, y, model):
predictions = model(X)
return(y, predictions)
def epoch(data_iter, df, model, optimizer, cat_cols, num_cols, norm_dict):
out_loss = {}
out_acc = {}
n = 0.
n_batch = 0.
total_time_dataload = 0.
total_time_model = 0.
start_time = time.time()
for batch in data_iter:
total_time_dataload += time.time() - start_time
start_time = time.time()
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
subjectid = decoder(batch['subjectid'])
y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
X = tf.concat([t1, t2], axis=4)
#X = tf.concat([t1, t2], axis=4)
if optimizer != None:
y_true, y_pred, loss = train_step(X, y, model, optimizer, cat_cols, num_cols)
else:
y_true, y_pred = test_step(X, y, model)
out_loss, out_acc = calc_loss_acc(out_loss, out_acc, y_true, y_pred, cat_cols, num_cols, norm_dict)
n += X.shape[0]
n_batch += 1
if (n_batch % 10) == 0:
print(n_batch)
total_time_model += time.time() - start_time
start_time = time.time()
return (out_loss, out_acc, n, total_time_model, total_time_dataload)
def get_labels(df, subjectid, cols = ['nihtbx_fluidcomp_uncorrected_norm']):
subjects_df = pd.DataFrame(subjectid)
result_df = pd.merge(subjects_df, df, left_on=0, right_on='subjectkey', how='left')
output = {}
for col in cols:
output[col] = np.asarray(result_df[col].values)
return output
def best_val(df_best, df_val, df_test):
df_best = pd.merge(df_best, df_val, how='left', left_on='name', right_on='name')
df_best = pd.merge(df_best, df_test, how='left', left_on='name', right_on='name')
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_test'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_test']
df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'best_loss_val'] = df_best.loc[df_best['best_loss_val']>=df_best['cur_loss_val'], 'cur_loss_val']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']<=df_best['cur_acc_val'])&(df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_test'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_test']
df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'best_acc_val'] = df_best.loc[(df_best['best_acc_val']>=df_best['cur_acc_val'])&(~df_best['name'].isin(['female', 'race.ethnicity', 'high.educ_group', 'income_group', 'married'])), 'cur_acc_val']
df_best = df_best.drop(['cur_loss_val', 'cur_acc_val', 'cur_loss_test', 'cur_acc_test'], axis=1)
return(df_best)
decoder = np.vectorize(lambda x: x.decode('UTF-8'))
template = 'Epoch {0}, Loss: {1:.3f}, Accuracy: {2:.3f}, Val Loss: {3:.3f}, Val Accuracy: {4:.3f}, Time Model: {5:.3f}, Time Data: {6:.3f}'
for col in [0]:
log_textfile(path_output + modelname + 'multitask_test' + '.log', cat_cols),
log_textfile(path_output + modelname + 'multitask_test' + '.log', num_cols)
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam(lr = 0.001)
model = Model(cat_cols, num_cols)
df_best = None
for e in range(20):
log_textfile(path_output + modelname + 'multitask_test' + '.log', 'Epochs: ' + str(e))
loss = tf.Variable(0.)
acc = tf.Variable(0.)
val_loss = tf.Variable(0.)
val_acc = tf.Variable(0.)
test_loss = tf.Variable(0.)
test_acc = tf.Variable(0.)
tf.keras.backend.set_learning_phase(True)
train_out_loss, train_out_acc, n, time_model, time_data = epoch(train_iter, train_df, model, optimizer, cat_cols, num_cols, norm_dict)
tf.keras.backend.set_learning_phase(False)
val_out_loss, val_out_acc, n, _, _ = epoch(val_iter, val_df, model, None, cat_cols, num_cols, norm_dict)
test_out_loss, test_out_acc, n, _, _ = epoch(test_iter, test_df, model, None, cat_cols, num_cols, norm_dict)
loss, acc, _ = format_output(train_out_loss, train_out_acc, n, list(cat_cols.keys())+num_cols)
val_loss, val_acc, df_val = format_output(val_out_loss, val_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_val.columns = ['name', 'cur_loss_val', 'cur_acc_val']
df_test.columns = ['name', 'cur_loss_test', 'cur_acc_test']
if e == 0:
df_best = pd.merge(df_test, df_val, how='left', left_on='name', right_on='name')
df_best.columns = ['name', 'best_loss_test', 'best_acc_test', 'best_loss_val', 'best_acc_val']
df_best = best_val(df_best, df_val, df_test)
print(df_best[['name', 'best_loss_test', 'best_acc_test']])
print(df_best[['name', 'best_loss_val', 'best_acc_val']])
log_textfile(path_output + modelname + 'multitask_test' + '.log', template.format(e, loss, acc, val_loss, val_acc, time_model, time_data))
if list(np.asarray(df_test.loc[df_test['name']=='female','cur_acc_test'].values)>0.70)[0]:
break
if e in [10, 15]:
optimizer.lr = optimizer.lr/3
log_textfile(path_output + modelname + 'multitask_test' + '.log', 'Learning rate: ' + str(optimizer.lr))
df_best.to_csv(path_output + modelname + 'multitask_test' + '.csv')
test_loss, test_acc, df_test = format_output(test_out_loss, test_out_acc, n, list(cat_cols.keys())+num_cols, print_bl=False)
df_test.to_csv('final_output_all_simple.csv')
inputs = tf.keras.Input(shape=(64,64,64,2), name='inputlayer123')
a = model(inputs)['female']
mm = tf.keras.models.Model(inputs=inputs, outputs=a)
from tf_explain.core.smoothgrad import SmoothGrad
import pickle
explainer = SmoothGrad()
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
X_i = X[i]
X_i = tf.expand_dims(X_i, axis=0)
y_i = y_list[i]
grid = explainer.explain((X_i, _), mm, y_i, 20, 1.)
output_grid[y_i] += grid
output_n[y_i] += 1
pickle.dump([output_grid, output_n], open( "smoothgrad_female_all_simple.p", "wb" ) )
#output_grid, output_n = pickle.load(open( "smoothgrad_female.p", "rb" ))
def apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size):
"""
Replace a part of the image with a grey patch.
Args:
image (numpy.ndarray): Input image
top_left_x (int): Top Left X position of the applied box
top_left_y (int): Top Left Y position of the applied box
patch_size (int): Size of patch to apply
Returns:
numpy.ndarray: Patched image
"""
patched_image = np.array(image, copy=True)
patched_image[
top_left_x : top_left_x + patch_size, top_left_y : top_left_y + patch_size, top_left_z : top_left_z + patch_size, :
] = 0
return patched_image
import math
def get_sensgrid(image, mm, class_index, patch_size):
sensitivity_map = np.zeros((
math.ceil(image.shape[0] / patch_size),
math.ceil(image.shape[1] / patch_size),
math.ceil(image.shape[2] / patch_size)
))
for index_z, top_left_z in enumerate(range(0, image.shape[2], patch_size)):
patches = [
apply_grey_patch(image, top_left_x, top_left_y, top_left_z, patch_size)
for index_x, top_left_x in enumerate(range(0, image.shape[0], patch_size))
for index_y, top_left_y in enumerate(range(0, image.shape[1], patch_size))
]
coordinates = [
(index_y, index_x)
for index_x, _ in enumerate(range(0, image.shape[0], patch_size))
for index_y, _ in enumerate(range(0, image.shape[1], patch_size))
]
predictions = mm.predict(np.array(patches), batch_size=1)
target_class_predictions = [prediction[class_index] for prediction in predictions]
for (index_y, index_x), confidence in zip(coordinates, target_class_predictions):
sensitivity_map[index_y, index_x, index_z] = 1 - confidence
sm = resize(sensitivity_map, (64,64,64))
heatmap = (sm - np.min(sm)) / (sm.max() - sm.min())
return(heatmap)
output_grid = {}
output_n = {}
for i in range(2):
output_grid[i] = np.zeros((64,64,64))
output_n[i] = 0
counter = 0
for batch in test_iter:
counter+=1
print(counter)
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
X = tf.concat([t1, t2], axis=4)
subjectid = decoder(batch['subjectid'])
y = get_labels(test_df, subjectid, list(cat_cols.keys())+num_cols)
y_list = list(y['female'])
for i in range(X.shape[0]):
print(i)
X_i = X[i]
y_i = y_list[i]
grid = get_sensgrid(X_i, mm, y_i, 4)
output_grid[y_i] += grid
output_n[y_i] += 1
if counter==6:
break
pickle.dump([output_grid, output_n], open( "heatmap_female_all_simple.p", "wb" ) )
error
batch = next(iter(train_iter))
t1 = (tf.cast(batch['t1'], tf.float32)-t1_mean)/t1_std
t2 = (batch['t2']-t2_mean)/t2_std
ad = batch['ad']
ad = tf.where(tf.math.is_nan(ad), tf.zeros_like(ad), ad)
ad = (ad-ad_mean)/ad_std
fa = batch['fa']
fa = tf.where(tf.math.is_nan(fa), tf.zeros_like(fa), fa)
fa = (fa-fa_mean)/fa_std
md = batch['md']
md = tf.where(tf.math.is_nan(md), tf.zeros_like(md), md)
md = (md-md_mean)/md_std
rd = batch['rd']
rd = tf.where(tf.math.is_nan(rd), tf.zeros_like(rd), rd)
rd = (rd-rd_mean)/rd_std
#subjectid = decoder(batch['subjectid'])
#y = get_labels(df, subjectid, list(cat_cols.keys())+num_cols)
#X = tf.concat([t1, t2, ad, fa, md, rd], axis=4)
X = tf.concat([t1, t2], axis=4)
tf.keras.backend.set_learning_phase(True)
model(X)['female']
tf.keras.backend.set_learning_phase(False)
model(X)['female']
mean_std = [x.name for x in model.non_trainable_variables if ('batch_norm') in x.name and ('mean' in x.name or 'variance' in x.name)]
model = Model(cat_cols, num_cols)
model.non_trainable_variables
```
| github_jupyter |
```
from keras.datasets import cifar10
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.layers.advanced_activations import LeakyReLU, PReLU, ParametricSoftplus
from keras.utils import np_utils, generic_utils
from keras.callbacks import ModelCheckpoint, EarlyStopping
import numpy as np
batch_size = 64
nb_classes = 10
nb_epoch = 100
img_channels = 3
img_rows = 32
img_cols = 32
# the data, shuffled and split between tran and test sets
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
datagen = ImageDataGenerator(
featurewise_center=False,
samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,
rotation_range=None,
width_shift_range=None,
height_shift_range=None,
horizontal_flip=True,
vertical_flip=False)
batch = 0
for X_batch, y_batch in datagen.flow(X_train, y_train, batch_size=2048):
print(batch, end='...', flush=True)
X_train = np.vstack((X_train, X_batch))
y_train = np.vstack((y_train, y_batch))
batch += 1
print('X_train shape:', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
Y_train = np_utils.to_categorical(y_train, nb_classes)
Y_test = np_utils.to_categorical(y_test, nb_classes)
print('Y_train shape:', Y_train.shape)
print('Y_test shape:', Y_test.shape)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
model = Sequential()
model.add(Convolution2D(32, 3, 3, border_mode='full',
input_shape=(img_channels, img_rows, img_cols)))
model.add(LeakyReLU(alpha=0.2))
model.add(Convolution2D(32, 3, 3))
model.add(LeakyReLU(alpha=0.2))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.2))
model.add(Convolution2D(64, 3, 3, border_mode='full'))
model.add(LeakyReLU(alpha=0.2))
model.add(Convolution2D(64, 3, 3))
model.add(LeakyReLU(alpha=0.2))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.3))
model.add(Convolution2D(128, 3, 3, border_mode='full'))
model.add(LeakyReLU(alpha=0.2))
model.add(Convolution2D(128, 3, 3))
model.add(LeakyReLU(alpha=0.2))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(nb_classes))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', class_mode='categorical')
checkpointer = ModelCheckpoint(filepath='cifar10_cnn_keras_weights.hdf5', verbose=1, save_best_only=True)
earlystopping = EarlyStopping(monitor='val_loss', patience=10, verbose=1)
model.fit(X_train, Y_train,
batch_size=batch_size,
nb_epoch=nb_epoch,
show_accuracy=True,
validation_data=(X_test, Y_test),
callbacks=[checkpointer, earlystopping])
import h5py
import json
import gzip
layer_name_dict = {
'Dense': 'denseLayer',
'Dropout': 'dropoutLayer',
'Flatten': 'flattenLayer',
'Embedding': 'embeddingLayer',
'BatchNormalization': 'batchNormalizationLayer',
'LeakyReLU': 'leakyReLULayer',
'PReLU': 'parametricReLULayer',
'ParametricSoftplus': 'parametricSoftplusLayer',
'ThresholdedLinear': 'thresholdedLinearLayer',
'ThresholdedReLu': 'thresholdedReLuLayer',
'LSTM': 'rLSTMLayer',
'GRU': 'rGRULayer',
'JZS1': 'rJZS1Layer',
'JZS2': 'rJZS2Layer',
'JZS3': 'rJZS3Layer',
'Convolution2D': 'convolution2DLayer',
'MaxPooling2D': 'maxPooling2DLayer'
}
layer_params_dict = {
'Dense': ['weights', 'activation'],
'Dropout': ['p'],
'Flatten': [],
'Embedding': ['weights'],
'BatchNormalization': ['weights', 'epsilon'],
'LeakyReLU': ['alpha'],
'PReLU': ['weights'],
'ParametricSoftplus': ['weights'],
'ThresholdedLinear': ['theta'],
'ThresholdedReLu': ['theta'],
'LSTM': ['weights', 'activation', 'inner_activation', 'return_sequences'],
'GRU': ['weights', 'activation', 'inner_activation', 'return_sequences'],
'JZS1': ['weights', 'activation', 'inner_activation', 'return_sequences'],
'JZS2': ['weights', 'activation', 'inner_activation', 'return_sequences'],
'JZS3': ['weights', 'activation', 'inner_activation', 'return_sequences'],
'Convolution2D': ['weights', 'nb_filter', 'nb_row', 'nb_col', 'border_mode', 'subsample', 'activation'],
'MaxPooling2D': ['pool_size', 'stride', 'ignore_border']
}
layer_weights_dict = {
'Dense': ['W', 'b'],
'Embedding': ['E'],
'BatchNormalization': ['gamma', 'beta', 'mean', 'std'],
'PReLU': ['alphas'],
'ParametricSoftplus': ['alphas', 'betas'],
'LSTM': ['W_xi', 'W_hi', 'b_i', 'W_xc', 'W_hc', 'b_c', 'W_xf', 'W_hf', 'b_f', 'W_xo', 'W_ho', 'b_o'],
'GRU': ['W_xz', 'W_hz', 'b_z', 'W_xr', 'W_hr', 'b_r', 'W_xh', 'W_hh', 'b_h'],
'JZS1': ['W_xz', 'b_z', 'W_xr', 'W_hr', 'b_r', 'W_hh', 'b_h', 'Pmat'],
'JZS2': ['W_xz', 'W_hz', 'b_z', 'W_hr', 'b_r', 'W_xh', 'W_hh', 'b_h', 'Pmat'],
'JZS3': ['W_xz', 'W_hz', 'b_z', 'W_xr', 'W_hr', 'b_r', 'W_xh', 'W_hh', 'b_h'],
'Convolution2D': ['W', 'b']
}
def serialize(model_json_file, weights_hdf5_file, save_filepath, compress):
with open(model_json_file, 'r') as f:
model_metadata = json.load(f)
weights_file = h5py.File(weights_hdf5_file, 'r')
layers = []
num_activation_layers = 0
for k, layer in enumerate(model_metadata['layers']):
if layer['name'] == 'Activation':
num_activation_layers += 1
prev_layer_name = model_metadata['layers'][k-1]['name']
idx_activation = layer_params_dict[prev_layer_name].index('activation')
layers[k-num_activation_layers]['parameters'][idx_activation] = layer['activation']
continue
layer_params = []
for param in layer_params_dict[layer['name']]:
if param == 'weights':
weights = {}
weight_names = layer_weights_dict[layer['name']]
for p, name in enumerate(weight_names):
weights[name] = weights_file.get('layer_{}/param_{}'.format(k, p)).value.tolist()
layer_params.append(weights)
else:
layer_params.append(layer[param])
layers.append({
'layerName': layer_name_dict[layer['name']],
'parameters': layer_params
})
if compress:
with gzip.open(save_filepath, 'wb') as f:
f.write(json.dumps(layers).encode('utf8'))
else:
with open(save_filepath, 'w') as f:
json.dump(layers, f)
import json
model_metadata = json.loads(model.to_json())
with open('cifar10_cnn_keras_model.json', 'w') as f:
json.dump(model_metadata, f)
model_metadata
serialize('cifar10_cnn_keras_model.json',
'cifar10_cnn_keras_weights.hdf5',
'cifar10_cnn_model_params.json.gz',
True)
serialize('cifar10_cnn_keras_model.json',
'cifar10_cnn_keras_weights.hdf5',
'cifar10_cnn_model_params.json',
False)
import numpy as np
import gzip
randidx = np.random.randint(0, X_test.shape[0], size=500)
X_rand = X_test[randidx, :]
y_rand = y_test[randidx]
with gzip.open('sample_data.json.gz', 'wb') as f:
f.write(json.dumps({'data': X_rand.tolist(), 'labels': y_rand.tolist()}).encode('utf8'))
with open('sample_data.json', 'w') as f:
json.dump({'data': X_rand.tolist(), 'labels': y_rand.tolist()}, f)
%%time
model.predict(X_rand[0:1,:])
```
| github_jupyter |
```
import numpy as np
import matplotlib.cm as cm
import matplotlib.pyplot as plt
from math import cos, sin, pi
%matplotlib inline
def distance(p1,p2):
dx = p1[0]-p2[0]
dy = p1[1]-p2[1]
distance = np.sqrt(dx*dx+dy*dy)
return distance
def circle_intersection(circle1, circle2):
'''
Find 2 intersection points from 2 circles.
if circles are non overlap, return middle point between 2 perimeters
'''
x1,y1,r1 = circle1
x2,y2,r2 = circle2
dx,dy = x2-x1,y2-y1
d = np.sqrt(dx*dx+dy*dy)
#d = distance([x1,y1],[x2,y2])
# non-overlapping circles
if d >= r1+r2:
return (( (d+r1-r2)/(2*d)*(x2-x1)+x1, (d+r1-r2)/(2*d)*(y2-y1)+y1 ),)
# one circle inside another
elif r1 > (d+r2) or r2 > (d+r1):
if dx == 0: angle = pi/2
else: angle = np.arctan(dy/dx)
p11 = [ x1+r1*cos(angle), y1+r1*sin(angle) ]
p12 = [ x1-r1*cos(angle), y1-r1*sin(angle) ]
p21 = [ x2+r2*cos(angle), y2+r2*sin(angle) ]
p22 = [ x2-r2*cos(angle), y2-r2*sin(angle) ]
point_set = [[p11,p21],[p11,p22],[p12,p21],[p12,p22]]
dist_set = [distance(item[0],item[1]) for item in point_set]
points = point_set[np.argmin(dist_set)]
points = np.array(points)
points = sum(points)/len(points)
return (points,)
# overlapping circles
else:
a = (r1*r1-r2*r2+d*d)/(2*d)
h = np.sqrt(r1*r1-a*a)
xm = x1 + a*dx/d
ym = y1 + a*dy/d
xs1 = xm + h*dy/d
xs2 = xm - h*dy/d
ys1 = ym - h*dx/d
ys2 = ym + h*dx/d
return (xs1,ys1),(xs2,ys2)
def plot(cs):
cmap = cm.plasma
fig, ax = plt.subplots()
ax.set(xlim=[-10, 10], ylim=[-10, 10], aspect=1)
for i,c in enumerate(cs):
ax.add_artist(plt.Circle(c[:2], c[2], color=cmap(i/len(cs)), alpha=0.5))
ax.plot(c[0], c[1], 'or')
return fig, ax
def choose_point(points, circle, order):
'''
1. choose point nearest to perimeter of given circle
2. between choosen point and perimeter of circle, calculate location from weight average
Args:
points: [[x1,y1],[x2,x2],...]
circle: [center_x, center_y, radius]
order = number of circle in this calculation
Returns:
result: [x,y]
'''
distance = ()
for point in points:
dx = point[0]-circle[0]
dy = point[1]-circle[1]
dist = np.sqrt(dx*dx+dy*dy) - circle[2]
distance = distance + (dist,)
arg = np.argmin([abs(i) for i in distance])
choosen_point = points[arg]
# ref_point = point on c3 perimeter closest to choosen_point
ref_point = [circle[0]+(choosen_point[0]-circle[0])*circle[2]/(circle[2]+distance[arg]),
circle[1]+(choosen_point[1]-circle[1])*circle[2]/(circle[2]+distance[arg])]
result = [((order-1)*choosen_point[0]+ref_point[0])/order, ((order-1)*choosen_point[1]+ref_point[1])/order]
return result
def trilateration(c1, c2, c3):
points = circle_intersection(c1, c2)
result = choose_point(points, c3, 3)
fig, ax = plot([c1, c2, c3])
ax.plot(result[0], result[1], "*")
return result
c1 = (2, 0, 7)
c2 = (1, 4, 3)
c3 = (5, 3, 3)
c4 = (2, 2, 4)
trilateration(c1, c2, c3)
def trilateration2(c1, c2, c3):
circles = [c1,c2,c3]
results = []
for i in range(3):
first = circles[0]
circles.pop(0)
circles.append(first)
points = circle_intersection(circles[0], circles[1])
result = choose_point(points, circles[2], 3)
results.append(result)
results = np.array(results)
location = sum(results)/len(results)
fig, ax = plot([c1, c2, c3])
ax.plot(location[0],location[1], "*")
return location
c1 = (2, 0, 7)
c2 = (1, 4, 3)
c3 = (5, 3, 3)
trilateration2(c1, c2, c3)
def trilateration3(cs):
results = []
for i in range(len(cs)):
if i!=0:
first = cs[0]
cs.pop(0)
cs.append(first)
points = circle_intersection(cs[0], cs[1])
for j, circle in enumerate(cs[2:]):
points = choose_point(points, circle, order = j+3)
points = np.expand_dims(points, axis=0)
results.append(points[0])
results = np.array(results)
location = sum(results)/len(results)
print('error: ')
for result in results:
err = distance(result, location)
print(err)
fig, ax = plot(cs)
ax.plot(location[0], location[1], "*")
return location
c1 = (2, 0, 6)
c2 = (1, 4, 3)
c3 = (5, 3, 3)
c4 = (2, 2, 4)
trilateration3([c1, c2, c3, c4])
c1 = (2, 3, 3)
c2 = (-4, -4, 5.5)
c3 = (-2, 2, 3)
c4 = (6, -2, 5.5)
trilateration3([c1, c2, c3, c4])
```
| github_jupyter |
# Pandas 4: Combining data
Sometimes we need to combine data from two or more dataframes. That's colloquially known as a **merge** or a **join**. There are lots of ways to do this. We do a couple but supply references to more at the end.
Along the way we take an extended detour to review methods for **downloading** and **unzipping** compressed files. The tools we use here have a broad range of other applications, including web scraping.
Outline:
* [MovieLens data](#movielens). A collection of movies and individual ratings.
* [Automate file download](#requests). Use the requests package to get a zipped file, then other tools to unzip it and read in the contents.
* [Merge movie names and ratings](#merge-movies). Merge information from two dataframes with Pandas' `merge` function.
* [UN population data](#example). We merge ("concatenate") estimates from the past with projections of the future.
**Note: requires internet access to run.**
This IPython notebook was created by Dave Backus, Chase Coleman, Brian LeBlanc, and Spencer Lyon for the NYU Stern course [Data Bootcamp](http://databootcamp.nyuecon.com/).
<a id=prelims></a>
## Preliminaries
Import packages, etc.
```
import pandas as pd # data package
import matplotlib.pyplot as plt # graphics
import sys # system module, used to get Python version
import os # operating system tools (check files)
import datetime as dt # date tools, used to note current date
# these are new
import requests, io # internet and input tools
import zipfile as zf # zip file tools
import shutil # file management tools
%matplotlib inline
print('\nPython version: ', sys.version)
print('Pandas version: ', pd.__version__)
print('Requests version: ', requests.__version__)
print("Today's date:", dt.date.today())
```
<a id=movielens></a>
## MovieLens data
The data comes as a zip file that contains several csv's. We get the details from the README inside. (It's written in Markdown, so it's easier to read if we use a browser to format it. Or we could cut and paste into a Markdown cell in an IPython notebook.)
The file descriptions are:
* `ratings.csv`: each line is an individual film rating with the rater and movie id's and the rating. Order: `userId, movieId, rating, timestamp`.
* `tags.csv`: each line is a tag on a specific film. Order: `userId, movieId, tag, timestamp`.
* `movies.csv`: each line is a movie name, its id, and its genre. Order: `movieId, title, genres`. Multiple genres are separated by "pipes" `|`.
* `links.csv`: each line contains the movie id and corresponding id's at [IMBd](http://www.imdb.com/) and [TMDb](https://www.themoviedb.org/).
The easy way to input this data is to download the zip file onto our computer, unzip it, and read the individual csv files using `read.csv()`. But **anyone can do it the easy way**. We want to automate this, so we can redo it without any manual steps. This takes some effort, but once we have it down we can apply it to lots of other data sources.
<a id=requests></a>
## Automate file download
We're looking for an automated way, so that if we do this again, possibly with updated data, the whole process is in our code. Automated data entry involves these steps:
* Get the file. We use the [requests](http://docs.python-requests.org/) package, which handles internet files and comes pre-installed with Anaconda. This kind of thing was hidden behind the scenes in the Pandas `read_csv` function, but here we need to do it for ourselves. The package authors add:
>Recreational use of other HTTP libraries may result in dangerous side-effects, including: security vulnerabilities, verbose code, reinventing the wheel, constantly reading documentation, depression, headaches, or even death.
* Convert to zip. Requests simply loads whatever's at the given url. The [io](https://docs.python.org/3.5/library/io.html) module's `io.Bytes` reconstructs it as a file, here a zip file.
* Unzip the file. We use the [zipfile](https://docs.python.org/3.5/library/zipfile.html) module, which is part of core Python, to extract the files inside.
* Read in the csv's. Now that we've extracted the csv files, we use `read_csv` as usual.
We found this [Stack Overflow exchange](http://stackoverflow.com/questions/23419322/download-a-zip-file-and-extract-it-in-memory-using-python3) helpful.
**Digression.** This is probably more than you want to know, but it's a reminder of what goes on behind the scenes when we apply `read_csv` to a url. Here we grab whatever is at the url. Then we get its contents, convert it to bytes, identify it as a zip file, and read its components using `read_csv`. It's a lot easier when this happens automatically, but a reminder what's involved if we ever have to look into the details.
```
# get "response" from url
url = 'http://files.grouplens.org/datasets/movielens/ml-latest-small.zip'
r = requests.get(url)
# describe response
print('Response status code:', r.status_code)
print('Response type:', type(r))
print('Response .content:', type(r.content))
print('Response headers:\n', r.headers, sep='')
# convert bytes to zip file
mlz = zf.ZipFile(io.BytesIO(r.content))
print('Type of zipfile object:', type(mlz))
# what's in the zip file?
mlz.namelist()
# extract and read csv's
movies = pd.read_csv(mlz.open(mlz.namelist()[2]))
ratings = pd.read_csv(mlz.open(mlz.namelist()[3]))
# what do we have?
for df in [movies, ratings]:
print('Type:', type(df))
print('Dimensions:', df.shape)
print('Variables:', list(df))
print('First few rows', df.head(3), '\n')
```
**Exercise.** Something to do together. suppose we wanted to save the files on our computer. How would we do it? Would we prefer individual csv's or a single zip?
```
# experiment via http://stackoverflow.com/a/18043472/804513
with open('test.zip', 'wb') as out_file:
shutil.copyfileobj(io.BytesIO(r.content), out_file)
```
<a id=merge-movies></a>
## Merging ratings and movie titles
The movie ratings in the dataframe `ratings` give us individual opinions about movies, but they don't include the name of the movie. Why not? Rather than include the name every time a movie is rated, the MovieLens data associates each rating with a movie code, than stores the names of movies associatd with each movie code in the dataframe `movies`. We run across this a lot: some information is in one data table, other information is in another.
Our **want** is therefore to add the movie name to the `ratings` dataframe. We say we **merge** the two dataferames. There are lots of ways to merge. Here we do one as an illustration.
Let's start by reminding ourselves what we have.
```
ratings.head(3)
movies.head(3)
```
### Merging
Here's roughly what's involved in what we're doing. We take the `movieId` variable from `ratings` and look it up in `movies`. When we find it, we look up the `title` and add it as a column in `ratings`. The variable `movieId` is common, so we can use it to link the two dataframes.
```
combo = pd.merge(ratings, movies, # left and right df's
how='left', # add to left
on='movieId' # link with this variable/column
)
print('Dimensions of ratings:', ratings.shape)
print('Dimensions of movies:', movies.shape)
print('Dimensions of new df:', combo.shape)
combo.head(20)
# save as csv file for future use
combo.to_csv('mlcombined.csv')
print('Current directory:\n', os.getcwd(), sep='')
print('List of files:', os.listdir(), sep='\n')
```
**Exercise.** Some of these we know how to do, the others we don't. For the ones we know, what is the answer? For the others, what (in loose terms) do we need to be able to do to come up with an answer?
* What is the overall average rating?
* What is the overall distribution of ratings?
* What is the average rating of each movie?
* How many ratings does each movie get?
```
"""
fig, ax = plt.subplots()
bins = [bin/100 for bin in list(range(25, 575, 50))]
print(bins)
combo['rating'].plot(kind='hist', ax=ax, bins=bins, color='blue', alpha=0.5)
ax.set_xlim(0,5.5)
ax.set_ylabel('Number')
ax.set_xlabel('Rating')
"""
```
<a id=population></a>
## Population "estimates" and "projections"
We look (again) at the UN's [population data](http://esa.un.org/unpd/wpp/Download/Standard/Population/), specifically the age distribution of the population. The data comes in two sheets: *estimates* that cover the period 1950-2015 and *projections* that cover 2016-2100. Our mission is to combine them.
### Load data
We start, as usual, by loading the data. This takes a minute or so.
```
url1 = 'http://esa.un.org/unpd/wpp/DVD/Files/'
url2 = '1_Indicators%20(Standard)/EXCEL_FILES/1_Population/'
url3 = 'WPP2017_POP_F07_1_POPULATION_BY_AGE_BOTH_SEXES.XLSX'
url = url1 + url2 + url3
cols = [2, 5] + list(range(6,28))
est = pd.read_excel(url, sheetname=0, skiprows=16, parse_cols=cols, na_values=['…'])
prj = pd.read_excel(url, sheetname=1, skiprows=16, parse_cols=cols, na_values=['…'])
print('Dimensions and dtypes of estimates: ', est.shape, '\n', est.dtypes.head(), sep='')
print('\nDimensions and dtypes of projections: ', prj.shape, '\n', prj.dtypes.head(), sep='')
```
**Comment.** Note that they have different numbers of columns. Let's see where that comes from.
```
list(est)[15:]
list(prj)[15:]
```
### Clean data
Pick a useable subset and fix extra column so that we can combine them. The problem here is that until 1990, the highest age category was `'80+`. From 1990 on, we have a finer breakdown.
We fix this by reassigning `'80+'` to `'80-84'` and not worrying that some of these people are 85 or older. Note that `df.fillna(0.0)` replaces missing values with zeros.
```
def cleanpop(df, countries, years):
"""
take df as input and select countries and years
"""
# rename first two columns
names = list(df)
df = df.rename(columns={names[0]: 'Country', names[1]: 'Year'})
# select countries and years
newdf = df[df['Country'].isin(countries) & df['Year'].isin(years)]
return newdf
countries = ['Japan']
past = [1950, 2000]
future = [2050, 2100]
e = cleanpop(est, countries, past)
p = cleanpop(prj, countries, future)
# make copie sfor later use
ealt = e.copy()
palt = p.copy()
# fix top-coding in estimates
e['80-84'] = e['80-84'].fillna(0.0) + e['80+'].fillna(0.0)
e = e.drop(['80+'], axis=1)
# check dimensions again
print('Dimensions of cleaned estimates: ', e.shape)
print('Dimensions of cleaned projections: ', p.shape)
# check to see if we have the same variables
list(e) == list(p)
```
### Merge estimates and projections
If we have two blocks of data, and want just want to put them on top of each other, we use the Pandas' `concatenate` function. Ditto two blocks next to each other.
But first we need to fix the difference in the columns of the two dataframes.
```
pop = pd.concat([e, p], axis=0).fillna(0.0)
pop
```
**Exercise.** What happens if we try to merge the original dataframes, including the one with the extra `80+` column? Run the code below and comment on what you get.
```
"""
popalt = pd.concat([ealt, palt], axis=0)
popalt
"""
```
### Shape data
We **want** age categories in the index (the default x axis in a plot) and the years in the columns. The country we don't care about because there's only one.
```
pop = pop.drop('Country', axis=1)
popi = pop.set_index('Year')
popi
popi.columns.name = 'Age'
popt = popi.T
popt.head()
ax = popt.plot(kind='bar', color='blue', alpha=0.5,
subplots=True,
sharey=True,
figsize=(8,12))
```
**Exercise.** Use `set_index`, `stack`, and `unstack` to shape the dataframe `popi` into `popt`.
## Resources
The [Pandas docs](http://pandas.pydata.org/pandas-docs/stable/merging.html) are ok, but we prefer the Data Carpentry [guide](http://www.datacarpentry.org/python-ecology-lesson/04-merging-data)
| github_jupyter |
## 2.3 Modeling quality: 2. [small | medium] size tubers
### The data set
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import gridspec
import scipy.stats as stats
import time # Afficher la durée à l'optimisation
np.random.seed(seed = 505730)
```
Custom function to generate a grid with all possible combinations of vectors in a dictionnary (source: [pandas documentation](http://pandas.pydata.org/pandas-docs/stable/user_guide/cookbook.html?highlight=expand_grid#creating-example-data)).
```
import itertools
def expand_grid(data_dict):
rows = itertools.product(*data_dict.values())
return pd.DataFrame.from_records(rows, columns = data_dict.keys())
```
Load data.
```
df_ml = pd.read_csv('output/df_tuberSize.csv')
df_ml.columns
```
I create vectors categorizing response and predictors.
```
num_vars = ['DensitePlants', 'growing.season',
'temp_moy_5years', 'prec_tot_5years', 'sdi_5years', 'gdd_5years',
'NtotDose', 'PtotDose', 'KtotDose',
'soilTextIlr1', 'soilTextIlr2', 'soilTextIlr3',
'soilTypeIlr1_3', 'soilTypeIlr2_3',
'soil_pH',
'soil_P1_Fv.AlP', 'soil_P1_Al.P', 'soil_K2_FvMgCa.K', 'soil_K2_Fv.MgCa', 'soil_K2_Mg.Ca']
cat_vars = ['PrecCropFiveClasses']
resp_vars = ['tsizeS_M']
dose_vars = ['NtotDose', 'PtotDose', 'KtotDose']
```
In __N__ tests, __P__ and __K__ doses are in excess, which drags recommandations upwards. In all N tests, we set the __P__ and __K__ doses according to the CRAAQ fertilisation guide. The next cells are used to correct abnormally high non-limiting __P__ and __K__ dosage for some trials.
```
def craaq_P(x): # x, ISP
if x <= 2.5:
recommandation = 200
elif x <= 5:
recommandation = 150
elif x <= 10:
recommandation = 150
elif x <= 15:
recommandation = 120
elif x <= 25:
recommandation = 75
else:
recommandation = 50
return recommandation
def craaq_K(x): # x, K in kg K_M3 / ha
if x <= 75:
recommandation = 240
elif x <= 150:
recommandation = 215
elif x <= 225:
recommandation = 160
elif x <= 300:
recommandation = 120
elif x <= 375:
recommandation = 80
elif x <= 450:
recommandation = 50
else:
recommandation = 20
return recommandation
df_ml.loc[df_ml.test_type == 'N', "PtotDose"] = df_ml.ISP1.apply(lambda x: craaq_P(x))
df_ml.loc[df_ml.test_type == 'N', "KtotDose"] = df_ml.soil_K.apply(lambda x: craaq_K(x))
df_ml.loc[df_ml.test_type.isna(), "PtotDose"] = df_ml.ISP1.apply(lambda x: craaq_P(x))
df_ml.loc[df_ml.test_type.isna(), "KtotDose"] = df_ml.soil_K.apply(lambda x: craaq_K(x))
df_ml.loc[df_ml.test_type == 'P', "KtotDose"] = df_ml.soil_K.apply(lambda x: craaq_K(x))
df_ml.loc[df_ml.test_type == 'K', "PtotDose"] = df_ml.ISP1.apply(lambda x: craaq_P(x))
```
Verify max doses per element: N-P-K respectively, and drop other useless columns (related to soil elements and trial identifiers).
```
np.max(df_ml.NtotDose), np.max(df_ml.PtotDose), np.max(df_ml.KtotDose)
id_table = df_ml[['ID', 'NoEssai', 'NoBloc','ID_bl', 'NoTraitement', 'test_type']]
id_table.index = df_ml.index
soil_table = df_ml[['soil_P', 'soil_Al', 'soil_K', 'ISP1']]
soil_table.index = df_ml.index
df_ml = df_ml.drop(['ID', 'NoEssai', 'NoBloc', 'ID_bl', 'NoTraitement', 'soil_P', 'soil_Al', 'soil_K', 'ISP1', 'tsizeMS_L'], axis = 1)
```
I use the `panda.get_dummies()` function to transform each categorical feature with **m** possible values into **m** binary features, *with only one active*. The output will be a sparse matrix where each column corresponds to one possible value of one feature: one (__1__) indicates category membership; otherwise zero (__0__) is indicated.
```
df_ml["PrecCropFiveClasses"] = df_ml["PrecCropFiveClasses"].astype('category')
df_ml["test_type"] = df_ml["test_type"].astype('category')
df_ml["Cultivar"] = df_ml["Cultivar"].astype('category')
df_ml["Maturity5"] = df_ml["Maturity5"].astype('category')
df_ml = pd.get_dummies(df_ml, columns = ["PrecCropFiveClasses"])
df_ml = pd.get_dummies(df_ml, columns = ["test_type"])
df_ml = pd.get_dummies(df_ml, columns = ["Cultivar"])
df_ml = pd.get_dummies(df_ml, columns = ["Maturity5"])
```
If `test_type`, `Cultivar` or `Maturity5` variables should be removed before modeling, the next cell is activated or not:
```
df_ml = df_ml.loc[:, ~df_ml.columns.str.startswith('test_type')]
df_ml = df_ml.loc[:, ~df_ml.columns.str.startswith('Cultivar')]
df_ml = df_ml.loc[:, ~df_ml.columns.str.startswith('Maturity5')]
```
### Data set __train__ and __test__ split
If we need to predict a whole dose-response curve, we need to split by trials.
The split proportions are set to obtain __~70% of data__ in the training set. This kind of split is the most robust since the training and test sets are independant data. I split at _bloc level_. This split allows to mix data from the same experimental plots, but on different blocks. Data in both sets are independant, but because blocks are similar units the model might behave wrongly on new sites with less usual data.
```
np.random.seed(seed = 48327)
blocks = id_table['ID_bl']
N_blocks = blocks[id_table.test_type == 'N'].unique()
N_blocks_inTrain = np.random.choice(a = N_blocks,
size = np.around(0.7 * len(N_blocks)).astype(int),
replace = False)
P_blocks = blocks[id_table.test_type == 'P'].unique()
P_blocks_inTrain = np.random.choice(a = P_blocks,
size = np.around(0.7 * len(P_blocks)).astype(int),
replace = False)
K_blocks = blocks[id_table.test_type == 'K'].unique()
K_blocks_inTrain = np.random.choice(a = K_blocks,
size = np.around(0.7 * len(K_blocks)).astype(int),
replace = False)
npk_blocks = blocks[id_table.test_type == 'NPK'].unique()
npk_blocks_inTrain = np.random.choice(a = npk_blocks,
size = np.around(len(npk_blocks) * 0.7).astype(int),
replace = False)
na_blocks = blocks[id_table.test_type.isna()].unique()
na_blocks_inTrain = np.random.choice(a = na_blocks,
size = np.around(len(na_blocks) * 0.7).astype(int),
replace = False)
blocks_inTrain = np.concatenate([N_blocks_inTrain,
P_blocks_inTrain,
K_blocks_inTrain,
npk_blocks_inTrain,
na_blocks_inTrain])
blocks_inTest = blocks[~blocks.isin(blocks_inTrain)].unique()
```
Numerical variables are scaled before modeling. Both predictive and response variables are set to null (__0__) mean and unit (__1__) variance using the training set mean and standard error. This chunk computes the mean and standard error of predictive variables.
```
mean_numvars = df_ml.loc[blocks.isin(blocks_inTrain), num_vars].apply(axis = 0, func = np.mean)
std_numvars = df_ml.loc[blocks.isin(blocks_inTrain), num_vars].apply(axis = 0, func = np.std)
df_ml_sc = df_ml.copy()
for var in num_vars:
df_ml_sc[var] = (df_ml[var] - df_ml.loc[blocks.isin(blocks_inTrain), var].mean()) / df_ml.loc[blocks.isin(blocks_inTrain), var].std()
```
The response is also scaled:
```
mean_respvars = df_ml.loc[blocks.isin(blocks_inTrain), resp_vars].mean()
std_respvars = df_ml.loc[blocks.isin(blocks_inTrain), resp_vars].std()
for var in resp_vars:
df_ml_sc[var] = (df_ml[var] - mean_respvars[var]) / std_respvars[var]
```
Then, separate train and test arrays are created for predictive and response variables.
```
X_train = df_ml_sc.loc[blocks.isin(blocks_inTrain), df_ml.columns.isin(num_vars) | df_ml.columns.str.startswith(cat_vars[0])].values
X_test = df_ml_sc.loc[~blocks.isin(blocks_inTrain), df_ml.columns.isin(num_vars) | df_ml.columns.str.startswith(cat_vars[0])].values
Y_train = df_ml_sc[resp_vars][blocks.isin(blocks_inTrain)].values
Y_test = df_ml_sc[resp_vars][~blocks.isin(blocks_inTrain)].values
X_train.shape, X_test.shape, Y_train.shape, Y_test.shape
```
Let's check the proportion of data in the traning set:
```
round(df_ml.loc[blocks.isin(blocks_inTrain)].shape[0]/df_ml.shape[0], 3)
```
### Feature importance
I use the python scikit-learn _ExtraTreesRegressor_ algorithm to select features that contributed most as predictive variables. The process ranks features importance.
```
from sklearn.ensemble import ExtraTreesRegressor
import altair as alt
fs_reg = ExtraTreesRegressor(n_estimators = 50)
fs_reg.fit(X_train, Y_train.ravel())
```
There is a need to show column names in English rather than their names in the data frame. We created a `csv` as translation table (`translate_col2.csv`). This table is merged to the variable importance table, which is then ranked and plotted.
```
rf_varimp = pd.DataFrame({'columns': df_ml_sc.drop('tsizeS_M', axis = 1).columns,
'importance': fs_reg.feature_importances_})
translate_columns = pd.read_csv("csv/translate_col2.csv")
rf_varimp_en = pd.merge(rf_varimp,
translate_columns.drop("fr", axis = 1),
on = "columns", how = 'left').\
sort_values(by = ["importance"])
rf_varimp_en.to_csv('output/varImp_ilrSM.csv', index_label = "ID") # for common plot (1.4.1)
plt.figure(figsize = (4, 10))
plt.barh(rf_varimp_en.en, rf_varimp_en.importance, color = "black");
plt.grid(axis = 'x')
```
The preceeding crop categories could be removed.
```
remove_var_names = ['PrecCropFiveClasses_Cereal',
'PrecCropFiveClasses_Grassland',
'PrecCropFiveClasses_HRC',
'PrecCropFiveClasses_LRC',
'PrecCropFiveClasses_Legume']
keep_vars = ~ df_ml_sc.drop('tsizeS_M', axis = 1).columns.isin(remove_var_names)
X_train = X_train[:, keep_vars]
X_test = X_test[:, keep_vars]
```
### Modeling
#### _General packages_
```
#from sklearn.model_selection import GridSearchCV # Grid Search
from sklearn.model_selection import RandomizedSearchCV # Random Search
from sklearn.metrics import r2_score
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
from pprint import pprint # print default parameters of models
```
### k-nearest neighbors (knn) algorithm
#### _The default parameters_
```
from sklearn.neighbors import KNeighborsRegressor
knn_reg = KNeighborsRegressor()
print('Parameters currently in use:\n')
pprint(knn_reg.get_params())
```
#### _Optimisation (paraneters tuning)_
```
knn_param = {'n_neighbors': stats.randint(3, 20),
'weights': ['uniform', 'distance']}
n_iter_search = 50
print("\nRunning random search to tune up KNN hyperparameters ...")
start_time = time.time()
knn_random_search = RandomizedSearchCV(estimator = knn_reg, param_distributions = knn_param, n_iter = n_iter_search, cv = 5, n_jobs = -1)
knn_random_search.fit(X_train, Y_train)
print("Duration: %s secondes ---" % (time.time() - start_time))
knn_random_search.best_estimator_
knn_random_search.best_params_
```
#### _The __knn__ model_
```
knn_reg = KNeighborsRegressor(algorithm='auto', leaf_size=30, metric='minkowski',
metric_params=None, n_jobs=None, n_neighbors=18, p=2,
weights='distance')
knn = knn_reg.fit(X_train, Y_train)
```
Performance in train and test
```
Y_train_obs = df_ml[resp_vars][blocks.isin(blocks_inTrain)].values
Y_test_obs = df_ml[resp_vars][~blocks.isin(blocks_inTrain)].values
Y_train_predknn = knn.predict(X_train) * std_respvars.values + mean_respvars.values
Y_test_predknn = knn.predict(X_test) * std_respvars.values + mean_respvars.values
index = ['k-nearest neighbors']
knn_scores = pd.DataFrame({"Train_R2": r2_score(Y_train, knn.predict(X_train)).round(3),
"Train_MAE": mean_absolute_error(Y_train_obs, Y_train_predknn).round(4),
"Train_RMSE": np.sqrt(mean_squared_error(Y_train_obs, Y_train_predknn)).round(4),
"Test_R2": r2_score(Y_test, knn.predict(X_test)).round(3),
"Test_MAE": mean_absolute_error(Y_test_obs, Y_test_predknn).round(4),
"Test_RMSE": np.sqrt(mean_squared_error(Y_test_obs, Y_test_predknn)).round(4)},
index = index)
knn_scores
```
The __knn__ ressults plot
```
plt.subplot(1, 2, 1)
plt.plot(Y_train, knn.predict(X_train), marker = 'x', color = 'black', linestyle = 'None', alpha = 0.1)
plt.plot([-3, 3], [-3, 3], color = 'black', linewidth = 2)
plt.title('Train')
plt.subplot(1, 2, 2)
plt.plot(Y_test, knn.predict(X_test), marker = 'x', color = 'black', linestyle = 'None', alpha = 0.2)
plt.plot([-3, 3], [-3, 3], color = 'black', linewidth = 2)
plt.title('Test')
```
### Random forest (RF) algorithm
#### _The default parameters_
```
from sklearn.ensemble import RandomForestRegressor
rf_reg = RandomForestRegressor(random_state = 1)
print('Parameters currently in use:\n')
pprint(rf_reg.get_params())
```
#### _Optimisation_
The most important settings are the number of trees in the forest (`n_estimators`) and the number of features considered for splitting at each leaf node (`max_features`). The `max_features` is set to its default value `'auto'`, while I try out a wide range of values to see what works for `n_estimators`.
#### _The best parameters_
The best parameters from fitting the random search could be viewed with `rf_random_search.best_params_` :
#### _The optimal model_
```
rf_reg = RandomForestRegressor(bootstrap=True, criterion='mse', max_depth=None,
max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=17,
n_jobs=None, oob_score=False, random_state=1, verbose=0,
warm_start=False)
rfm = rf_reg.fit(X_train, Y_train.ravel())
```
Performance in train and test (rfm)
```
Y_train_predrf = rfm.predict(X_train) * std_respvars.values + mean_respvars.values
Y_test_predrf = rfm.predict(X_test) * std_respvars.values + mean_respvars.values
model = ['Random forest']
rf_scores = pd.DataFrame({"Train_R2": r2_score(Y_train, rfm.predict(X_train)).round(3),
"Train_MAE": mean_absolute_error(Y_train_obs, Y_train_predrf).round(4),
"Train_RMSE": np.sqrt(mean_squared_error(Y_train_obs, Y_train_predrf)).round(4),
"Test_R2": r2_score(Y_test, rfm.predict(X_test)).round(3),
"Test_MAE": mean_absolute_error(Y_test_obs, Y_test_predrf).round(4),
"Test_RMSE": np.sqrt(mean_squared_error(Y_test_obs, Y_test_predrf)).round(4)},
index = model)
rf_scores
```
The __rf__ ressults plot
```
#fig = plt.figure(figsize=(10, 5))
plt.subplot(1, 2, 1)
plt.plot(Y_train, rfm.predict(X_train), marker = 'o', color = 'grey', linestyle = 'None', alpha = 0.2)
plt.plot([-4, 3], [-4, 3], color = 'black', linewidth = 2)
plt.title('Training', fontsize = 12)
plt.xlabel(r'Mesured yield ($Mg~ha^{-1})$', fontsize = 12)
plt.ylabel(r'Predicted yield ($Mg~ha^{-1})$', fontsize = 12)
plt.subplot(1, 2, 2)
plt.plot(Y_test, rfm.predict(X_test), marker = 'o', color = 'grey', linestyle = 'None', alpha = 0.2)
plt.plot([-3, 3], [-3, 3], color = 'black', linewidth = 2)
plt.title('Testing', fontsize = 12)
plt.xlabel(r'Mesured yield ($Mg~ha^{-1})$', fontsize = 12)
```
### Neural Networks __NN__
#### _The default parameters_
```
from sklearn.neural_network import MLPRegressor
nn_reg = MLPRegressor(random_state = 1)
print('Parameters currently in use:\n')
pprint(nn_reg.get_params())
```
#### _Neural net parameters tuning_
nn_random_search.best_estimator_ =
MLPRegressor(activation='tanh', alpha=0.0001, batch_size='auto', beta_1=0.9,
beta_2=0.999, early_stopping=False, epsilon=1e-08,
hidden_layer_sizes=100, learning_rate='constant',
learning_rate_init=0.001, max_iter=500, momentum=0.9,
n_iter_no_change=10, nesterovs_momentum=True, power_t=0.5,
random_state=None, shuffle=True, solver='adam', tol=0.0001,
validation_fraction=0.1, verbose=False, warm_start=False)
#### _The neural network_
```
nn_reg = MLPRegressor(activation='tanh', alpha=0.0001, batch_size='auto', beta_1=0.9,
beta_2=0.999, early_stopping=False, epsilon=1e-08,
hidden_layer_sizes=100, learning_rate='constant',
learning_rate_init=0.001, max_iter=500, momentum=0.9,
n_iter_no_change=10, nesterovs_momentum=True, power_t=0.5,
random_state=None, shuffle=True, solver='adam', tol=0.0001,
validation_fraction=0.1, verbose=False, warm_start=False)
nn = nn_reg.fit(X_train, Y_train.ravel())
```
The __nn__ performance in train and test
```
Y_train_prednn = nn.predict(X_train) * std_respvars.values + mean_respvars.values
Y_test_prednn = nn.predict(X_test) * std_respvars.values + mean_respvars.values
index = ['Neural networks']
nn_scores = pd.DataFrame({"Train_R2": r2_score(Y_train, nn.predict(X_train)).round(3),
"Train_MAE": mean_absolute_error(Y_train_obs, Y_train_prednn).round(4),
"Train_RMSE": np.sqrt(mean_squared_error(Y_train_obs, Y_train_prednn)).round(4),
"Test_R2": r2_score(Y_test, nn.predict(X_test)).round(3),
"Test_MAE": mean_absolute_error(Y_test_obs, Y_test_prednn).round(4),
"Test_RMSE": np.sqrt(mean_squared_error(Y_test_obs, Y_test_prednn)).round(4)},
index = index)
nn_scores
```
The network ressults plot
```
plt.subplot(1, 2, 1)
plt.plot(Y_train, nn.predict(X_train), marker = 'x', color = 'blue', linestyle = 'None', alpha = 0.2)
plt.plot([-4, 3], [-4, 3], color = 'black', linewidth = 2)
plt.title('Train')
plt.subplot(1, 2, 2)
plt.plot(Y_test, nn.predict(X_test), marker = 'x', color = 'red', linestyle = 'None', alpha = 0.2)
plt.plot([-3, 3], [-3, 3], color='black', linewidth=2)
plt.title('Test')
```
### Gaussian process model __GP__
#### _The default parameters_
```
from sklearn.gaussian_process import GaussianProcessRegressor
gp_reg = GaussianProcessRegressor()
print('Parameters currently in use:\n')
pprint(gp_reg.get_params())
```
#### _Optimisation_
```
from sklearn.gaussian_process.kernels import RBF, Matern, WhiteKernel
```
The optimal model
Once the model has been optimized, I ran it again with more restarts. The __gp__ object (the model) was run then exported as a joblib file. To avoid fitting the model on each kernel restart, the fitting has been greyed out in favor of an import of the joblib file.
```
gp_reg = GaussianProcessRegressor(alpha=0.03060552017732754, copy_X_train=True,
kernel=RBF(length_scale=1), n_restarts_optimizer=0,
normalize_y=False, optimizer='fmin_l_bfgs_b',
random_state=None)
gpms = gp_reg.fit(X_train, Y_train)
```
The __gp__ performance in train and test
```
Y_train_predgp = gpms.predict(X_train) * std_respvars.values + mean_respvars.values
Y_test_predgp = gpms.predict(X_test) * std_respvars.values + mean_respvars.values
index = ['Gaussian processes']
gp_scores = pd.DataFrame({"Train_R2": r2_score(Y_train, gpms.predict(X_train)).round(3),
"Train_MAE": mean_absolute_error(Y_train_obs, Y_train_predgp).round(4),
"Train_RMSE": np.sqrt(mean_squared_error(Y_train_obs, Y_train_predgp)).round(4),
"Test_R2": r2_score(Y_test, gpms.predict(X_test)).round(3),
"Test_MAE": mean_absolute_error(Y_test_obs, Y_test_predgp).round(4),
"Test_RMSE": np.sqrt(mean_squared_error(Y_test_obs, Y_test_predgp)).round(4)},
index = index)
gp_scores
```
The __gp__ ressults plot
```
counter = 1
for i in range(len(resp_vars)):
plt.subplot(len(resp_vars), 2, counter)
plt.plot(Y_train[:, 0], gpms.predict(X_train), marker = 'x', color = 'blue', linestyle = 'None', alpha = 0.2)
plt.plot([-3, 3], [-3, 3], color = 'black', linewidth = 2)
plt.title('Train')
plt.subplot(1, 2, counter + 1)
plt.plot(Y_test[:, 0], gpms.predict(X_test), marker = 'x', color = 'red', linestyle = 'None', alpha = 0.2)
plt.plot([-3, 3], [-3, 3], color = 'black', linewidth = 2)
plt.title('Test')
counter = counter + 1
```
This chain concatenates scores of the models.
```
scoresTsms = pd.concat([knn_scores, rf_scores, nn_scores, gp_scores], axis=0, ignore_index=False)
scoresTsms['Target'] = '[S | M]'
scoresTsms.to_csv('output/scoresTsms.csv')
scoresTsms
```
### Point estimation: prediction of agronomic optimal __NPK__ dosage
#### Create a table to model
The same random row used in the preceding notebook `1.4.1_ml-marketable-yield-model.ipynb` is repoted here as the example where the model is used for diagnosis for comparison. The row is referenced by its index.
```
r_sample = pd.read_csv('output/r_sample.csv')
r_sample = id_table.loc[id_table.ID == r_sample.ID.values[0], ["NoEssai", "test_type", "ID"]]
r_sample
# Verify the row index
index = r_sample.index[0]
index
```
We grab this row with only the columns we need for modeling from the `df_ml_sc` table, which is already on the right scale for modeling.
```
reference_row = df_ml_sc.loc[df_ml_sc.index == index, df_ml.columns.isin(num_vars)]# | df_ml.columns.str.startswith(cat_vars[0])]
NoEssai = id_table.loc[df_ml_sc.index == index].NoEssai.values[0]
```
Here, we need to maximise the proportion of medium size tuber aiming to yield maximum tubers for table (market). There are no economic considerations. The optimal doses are the NPK combination where the computed ilr values reach the maximum.
Instead of generating a grid of values (which is numerically inefficient), we generate random NPK doses from uniform distributions on plausible doses. The `random.uniform()` function gives a random floating point number in a given range.
```
n_grid_samples = 1000
N_range = [0, 250]
P_range = [0, 250]
K_range = [0, 250]
np.random.seed(936492)
dose_grid = pd.DataFrame({'NtotDose': np.random.uniform(N_range[0], N_range[1], n_grid_samples),
'PtotDose': np.random.uniform(P_range[0], P_range[1], n_grid_samples),
'KtotDose': np.random.uniform(K_range[0], K_range[1], n_grid_samples)})
```
Doses are variables among others, and also need to be scaled on the scale on which they were put in to create the model.
```
dose_grid_sc = dose_grid.copy()
#dose_vars = dose_grid.columns
for var in dose_vars:
dose_grid_sc[var] = (dose_grid[var] - mean_numvars[var]) / std_numvars[var]
```
The last part to create the table is to stack the observation (randomly slected previously) a number of times equal to the `dose_grid` table, so that it conveys the same information at each row. Then we replace only NPK doses with the one sampled randomly.
```
reference_stack = pd.DataFrame(index = range(dose_grid.shape[0]), columns = reference_row.columns)
for i in range(reference_stack.shape[1]):
reference_stack.iloc[:, i] = np.repeat(reference_row.values[0, i], reference_stack.shape[0])
reference_stack.NtotDose = dose_grid_sc.NtotDose
reference_stack.PtotDose = dose_grid_sc.PtotDose
reference_stack.KtotDose = dose_grid_sc.KtotDose
```
The yield proportion is predicted based on the table, then put back to its original scale and inserted in the `dose_grid` table.
### Prediction with __knn__ model
```
ilr_sc = knn.predict(reference_stack)
dose_grid["ilr_"] = ilr_sc[:, 0] * std_respvars.values + mean_respvars.values
```
We isolate the row where `ilr_` is maximum.
```
opt_doses_knn = dose_grid.loc[dose_grid["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]].round(2)
opt_doses_knn
```
### Prediction with __rf__ model
```
ilr_sc = rfm.predict(reference_stack)
dose_grid["ilr_"] = ilr_sc * std_respvars.values + mean_respvars.values # yield_sc specifically not indexed needed with rf
opt_doses_rf = dose_grid.loc[dose_grid["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]].round(2)
opt_doses_rf
```
### Prediction with __nn__ model
```
ilr_sc = nn.predict(reference_stack)
dose_grid["ilr_"] = ilr_sc * std_respvars.values + mean_respvars.values # yield_sc specifically not indexed needed with rf
opt_doses_nn = dose_grid.loc[dose_grid["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]].round(2)
opt_doses_nn
```
### Prediction with __gp__ model
```
ilr_sc = gpms.predict(reference_stack)
dose_grid["ilr_"] = ilr_sc[:, 0] * std_respvars.values + mean_respvars.values
opt_doses_gp = dose_grid.loc[dose_grid["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]].round(2)
opt_doses_gp
```
This chain concatenates the points estimates of the models, and converts P2O5 and K2O in P and K respectively.
```
keys = ["NtotDose", "PtotDose", "KtotDose", "ilr_"]
predMs = pd.DataFrame({"k-nearest neighbors": opt_doses_knn,
"Random forest": opt_doses_rf,
"Neural networks": opt_doses_nn,
"Gaussian processes": opt_doses_gp},
index = keys).T
predMs['PtotDose'] = predMs.PtotDose*0.436
predMs['KtotDose'] = predMs.KtotDose*0.830
predMs['Target'] = '[S | M]'
predMs = predMs.rename(columns = {"ilr_":"Output"})
predMs.to_csv('output/predMs.csv')
predMs
```
### Prediction specific to trial type
I randomly select different test type (N, P, K) trials and plot their responses to compare the goodness of fit of the four models. The same codes are repeated for each element to generate new tables, one for each dose type. The treatment element varied while the others are kept constant at their experimental dosage.
Reference trial where only __N__ dose varied: `(test_type = N)`
```
test_types = pd.read_csv('output/test_types.csv')
id_sample_n = id_table.loc[id_table.ID == test_types.ID.values[0], ["NoEssai", "test_type", "ID"]]
id_sample_n
reference_row_N = df_ml_sc.loc[df_ml_sc.index == id_sample_n.index[0],
df_ml.columns.isin(num_vars)]# | df_ml.columns.str.startswith(cat_vars[0])]
dose_grid_N = pd.DataFrame({'NtotDose': np.linspace(0, 250, 251),
'PtotDose': reference_row_N.PtotDose.values[0],
'KtotDose': reference_row_N.KtotDose.values[0]})
dose_grid_N_sc = dose_grid_N.copy()
for var in dose_vars:
dose_grid_N_sc[var] = (dose_grid_N[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_N = pd.DataFrame(index = range(dose_grid_N.shape[0]), columns = reference_row_N.columns)
for i in range(reference_stack_N.shape[1]):
reference_stack_N.iloc[:, i] = np.repeat(reference_row_N.values[0, i], reference_stack_N.shape[0])
reference_stack_N.NtotDose = dose_grid_N_sc.NtotDose
reference_stack_N.PtotDose = dose_grid_N_sc.PtotDose
reference_stack_N.KtotDose = dose_grid_N_sc.KtotDose
# Point estimate
knn_N_sc = knn.predict(reference_stack_N)
rf_N_sc = rfm.predict(reference_stack_N)
nn_N_sc = nn.predict(reference_stack_N)
gp_N_sc = gpms.predict(reference_stack_N)
dose_grid_N["ilr_knn"] = knn_N_sc[:, 0] * std_respvars.values + mean_respvars.values
dose_grid_N["ilr_rf"] = rf_N_sc * std_respvars.values + mean_respvars.values
dose_grid_N["ilr_nn"] = nn_N_sc * std_respvars.values + mean_respvars.values
dose_grid_N["ilr_gp"] = gp_N_sc[:, 0] * std_respvars.values + mean_respvars.values
doses_opt_N_point = dose_grid_N.loc[dose_grid_N["ilr_gp"].idxmax(), ["NtotDose", "ilr_gp"]]#.round(2)
Ndata = df_ml.loc[id_table.NoEssai == id_sample_n.NoEssai.values[0]][["tsizeS_M", "NtotDose"]]
```
Reference trial where only __P__ dose varied: `(test_type = P)`
```
id_sample_p = id_table.loc[id_table.ID == test_types.ID.values[1], ["NoEssai", "test_type", "ID"]]
id_sample_p
```
```
reference_row_P = df_ml_sc.loc[df_ml_sc.index == id_sample_p.index[0],
df_ml.columns.isin(num_vars)]# | df_ml.columns.str.startswith(cat_vars[0])]
dose_grid_P = pd.DataFrame({'NtotDose': reference_row_P.NtotDose.values[0],
'PtotDose': np.linspace(0, 250, 251),
'KtotDose': reference_row_P.KtotDose.values[0]})
dose_grid_P['P'] = 0.436*dose_grid_P.PtotDose
dose_grid_P_sc = dose_grid_P.copy()
for var in dose_vars:
dose_grid_P_sc[var] = (dose_grid_P[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_P = pd.DataFrame(index = range(dose_grid_P.shape[0]), columns = reference_row_P.columns)
for i in range(reference_stack_P.shape[1]):
reference_stack_P.iloc[:, i] = np.repeat(reference_row_P.values[0, i], reference_stack_P.shape[0])
reference_stack_P.NtotDose = dose_grid_P_sc.NtotDose
reference_stack_P.PtotDose = dose_grid_P_sc.PtotDose
reference_stack_P.KtotDose = dose_grid_P_sc.KtotDose
# Point estimate
knn_P_sc = knn.predict(reference_stack_P)
rf_P_sc = rfm.predict(reference_stack_P)
nn_P_sc = nn.predict(reference_stack_P)
gp_P_sc = gpms.predict(reference_stack_P)
dose_grid_P["ilr_knn"] = knn_P_sc[:, 0] * std_respvars.values + mean_respvars.values
dose_grid_P["ilr_rf"] = rf_P_sc * std_respvars.values + mean_respvars.values
dose_grid_P["ilr_nn"] = nn_P_sc * std_respvars.values + mean_respvars.values
dose_grid_P["ilr_gp"] = gp_P_sc[:, 0] * std_respvars.values + mean_respvars.values
doses_opt_P_point = dose_grid_P.loc[dose_grid_P["ilr_gp"].idxmax(), ["PtotDose", "ilr_gp"]]#.round(2)
Pdata = df_ml.loc[id_table.NoEssai == id_sample_p.NoEssai.values[0]][["tsizeS_M", "PtotDose"]]
Pdata['P'] = 0.436*Pdata.PtotDose
```
Reference trial where only __K__ dose varied: `(test_type = K)`
```
id_sample_k = id_table.loc[id_table.ID == test_types.ID.values[2], ["NoEssai", "test_type", "ID"]]
id_sample_k
```
```
reference_row_K = df_ml_sc.loc[df_ml_sc.index == id_sample_k.index[0],
df_ml.columns.isin(num_vars)]# | df_ml.columns.str.startswith(cat_vars[0])]
dose_grid_K = pd.DataFrame({'NtotDose': reference_row_K.NtotDose.values[0],
'PtotDose': reference_row_K.PtotDose.values[0],
'KtotDose': np.linspace(0, 250, 251)})
dose_grid_K['K'] = 0.830*dose_grid_K.KtotDose
dose_grid_K_sc = dose_grid_K.copy()
for var in dose_vars:
dose_grid_K_sc[var] = (dose_grid_K[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_K = pd.DataFrame(index = range(dose_grid_K.shape[0]), columns = reference_row_K.columns)
for i in range(reference_stack_K.shape[1]):
reference_stack_K.iloc[:, i] = np.repeat(reference_row_K.values[0, i], reference_stack_K.shape[0])
reference_stack_K.NtotDose = dose_grid_K_sc.NtotDose
reference_stack_K.PtotDose = dose_grid_K_sc.PtotDose
reference_stack_K.KtotDose = dose_grid_K_sc.KtotDose
# Point estimate
knn_K_sc = knn.predict(reference_stack_K)
rf_K_sc = rfm.predict(reference_stack_K)
nn_K_sc = nn.predict(reference_stack_K)
gp_K_sc = gpms.predict(reference_stack_K)
dose_grid_K["ilr_knn"] = knn_K_sc[:, 0] * std_respvars.values + mean_respvars.values
dose_grid_K["ilr_rf"] = rf_K_sc * std_respvars.values + mean_respvars.values
dose_grid_K["ilr_nn"] = nn_K_sc * std_respvars.values + mean_respvars.values
dose_grid_K["ilr_gp"] = gp_K_sc[:, 0] * std_respvars.values + mean_respvars.values
doses_opt_K_point = dose_grid_K.loc[dose_grid_K["ilr_gp"].idxmax(), ["KtotDose", "ilr_gp"]]#.round(2)
Kdata = df_ml.loc[id_table.NoEssai == id_sample_k.NoEssai.values[0]][["tsizeS_M", "KtotDose"]]
Kdata['K'] = 0.830*Kdata.KtotDose
```
### Output data for future visualisation
The results are saved in the `output` folder and are loaded in the vizualisation notebook.
```
Ndata["Sample"] = id_sample_n.NoEssai.values[0]
Pdata["Sample"] = id_sample_p.NoEssai.values[0]
Kdata["Sample"] = id_sample_k.NoEssai.values[0]
Ndata.to_csv("output/tsizeM_S_model_Ndata.csv")
Pdata.to_csv("output/tsizeM_S_model_Pdata.csv")
Kdata.to_csv("output/tsizeM_S_model_Kdata.csv")
dose_grid_N.to_csv("output/tsizeM_S_model_N.csv")
dose_grid_P.to_csv("output/tsizeM_S_model_P.csv")
dose_grid_K.to_csv("output/tsizeM_S_model_K.csv")
```
### Probabilistic prediction with gaussian process
One advantage of gaussian processes is that it's probabilistic: it doesn't only returns point estimates, but can returns posterior samples. For the demonstration of this section, I use the __reference_stack__ data frame. I generate `500` gaussian process samples for each `NPK` triplet from the random doses __dose_grid__ table used for initial point estimation.
```
n_gp_samples = 500
gp_samples_sc = gpms.sample_y(reference_stack, n_samples = n_gp_samples)[:, 0, :] # [:, 0, :] to remove dummy middle dimension
```
... and put predicted `ilr` values back to scale.
```
gp_samples = gp_samples_sc.copy()
for i in range(n_gp_samples):
gp_samples[:, i] = gp_samples_sc[:, i] * std_respvars.values + mean_respvars.values
```
GP samples are put in a pandas data frame with pretty column names.
```
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_sm = pd.DataFrame(gp_samples, columns = gp_names)
```
Then, for each gaussian process, I find the agronomic (not economic) dosage, corresponding to maximum ratio (ilr). The dosages are gathered in a pandas data frame. The corresponding ilr is also reported.
```
doses_opt = pd.DataFrame({"NtotDose":0, "PtotDose":0, "KtotDose":0}, index = gp_names)
for i in gp_names:
doses_opt.loc[i] = dose_grid.loc[gp_sm.loc[:, i].idxmax(), ["NtotDose", "PtotDose", "KtotDose"]]
doses_opt['P'] = 0.436*doses_opt.PtotDose
doses_opt['K'] = 0.830*doses_opt.KtotDose
```
This chain plots the distribution of each fertilizer doses of the same trial (_the randomly selected one `r_sample`_). The red dotted vertical lines show the agronomic dosage from the initial gaussian process point estimate.
```
plt.figure(figsize=(12, 3), dpi= 80, facecolor='w', edgecolor='k')
gs = gridspec.GridSpec(1, 3,
width_ratios=[1,1,1],
height_ratios=[1]
)
ax0 = plt.subplot(gs[0])
ax0.hist(doses_opt.loc[:, "NtotDose"], color = "lightgrey", edgecolor = 'grey')
ax0.axvline(x = opt_doses_gp[0], color = 'r', linestyle = '--')
ax0.set_title("Trial N° " + str(NoEssai))
ax0.set_xlabel(r"Optimal dose N ($kg~ha^{-1})$")
ax0.set_ylabel("Count")
ax1 = plt.subplot(gs[1])
ax1.hist(doses_opt.loc[:, "P"], color = "lightgrey", edgecolor = 'grey')
ax1.axvline(x = 0.436*opt_doses_gp[1], color = 'r', linestyle = '--')
ax1.set_title("Trial N° " + str(NoEssai))
ax1.set_xlabel(r"Optimal dose P ($kg~ha^{-1})$")
ax2 = plt.subplot(gs[2])
ax2.hist(doses_opt.loc[:, "K"], color = "lightgrey", edgecolor = 'grey')
ax2.axvline(x = 0.830*opt_doses_gp[2], color = 'r', linestyle = '--')
ax2.set_title("Trial N° " + str(NoEssai))
ax2.set_xlabel(r"Optimal dose K ($kg~ha^{-1})$")
plt.tight_layout()
```
These distributions show optimal doses (_most of the time_) on the edge of ranges. (Let's explore why !!!). How can we use this model for decision making while it's difficult to obtain credible optimal dosage? Using the same trial, I generate new tables, one for each fertilizer type, and where one dose is varying while the others are kept constant _at their __gp__ model computed optimum_.
```
opt_doses_gp
```
#### Prediction where only __N__ dosage varied, trial `r_sample`
```
dose_grid_N = pd.DataFrame({'NtotDose': np.linspace(0, 250, 251),
'PtotDose': opt_doses_gp.PtotDose,
'KtotDose': opt_doses_gp.KtotDose})
dose_grid_N_sc = dose_grid_N.copy()
for var in dose_vars:
dose_grid_N_sc[var] = (dose_grid_N[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_N = pd.DataFrame(index = range(dose_grid_N.shape[0]), columns = reference_row.columns)
for i in range(reference_stack_N.shape[1]):
reference_stack_N.iloc[:, i] = np.repeat(reference_row.values[0, i], reference_stack_N.shape[0])
reference_stack_N.NtotDose = dose_grid_N_sc.NtotDose
reference_stack_N.PtotDose = dose_grid_N_sc.PtotDose
reference_stack_N.KtotDose = dose_grid_N_sc.KtotDose
```
Point estimate
```
ilr_N_sc = gpms.predict(reference_stack_N)
dose_grid_N["ilr_"] = ilr_N_sc[:, 0] * std_respvars.values + mean_respvars.values
doses_opt_N_point = dose_grid_N.loc[dose_grid_N["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]]#.round(2)
doses_opt_N_point
```
GP samples
```
n_gp_samples = 5
gp_samples_N_sc = gpms.sample_y(reference_stack_N, n_samples = n_gp_samples, random_state = 615108)[:, 0, :] # [:, 0, :] to remove dummy middle dimension
# Scale back to original
gp_samples_N = gp_samples_N_sc.copy()
for i in range(n_gp_samples):
gp_samples_N[:, i] = gp_samples_N_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_N = pd.DataFrame(gp_samples_N, columns = gp_names)
doses_opt_N_samples = pd.DataFrame({"NtotDose":0, "PtotDose":0, "KtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_N_samples.loc[i, ["NtotDose", "PtotDose", "KtotDose"]] = dose_grid_N.loc[gp_ilr_N.loc[:, i].idxmax(), ["NtotDose", "PtotDose", "KtotDose"]]
doses_opt_N_samples.loc[i, "ilr_"] = gp_ilr_N.loc[gp_ilr_N.loc[:, i].idxmax(), i]
```
#### Prediction where only __P__ dosage varied, trial `r_sample`
```
dose_grid_P = pd.DataFrame({'NtotDose': opt_doses_gp.NtotDose,
'PtotDose': np.linspace(0, 250, 251),
'KtotDose': opt_doses_gp.KtotDose})
dose_grid_P_sc = dose_grid_P.copy()
for var in dose_vars:
dose_grid_P_sc[var] = (dose_grid_P[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_P = pd.DataFrame(index = range(dose_grid_P.shape[0]), columns = reference_row.columns)
for i in range(reference_stack_P.shape[1]):
reference_stack_P.iloc[:, i] = np.repeat(reference_row.values[0, i], reference_stack_P.shape[0])
reference_stack_P.NtotDose = dose_grid_P_sc.NtotDose
reference_stack_P.PtotDose = dose_grid_P_sc.PtotDose
reference_stack_P.KtotDose = dose_grid_P_sc.KtotDose
```
Point estimate
```
ilr_P_sc = gpms.predict(reference_stack_P)
dose_grid_P["ilr_"] = ilr_P_sc[:, 0] * std_respvars.values + mean_respvars.values
dose_grid_P['P'] = 0.436*dose_grid_P.PtotDose
doses_opt_P_point = dose_grid_P.loc[dose_grid_P["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]]#.round(2)
doses_opt_P_point['P'] = 0.436*doses_opt_P_point.PtotDose
doses_opt_P_point
```
GP samples
```
n_gp_samples = 5
gp_samples_P_sc = gpms.sample_y(reference_stack_P, n_samples = n_gp_samples, random_state = 615108)[:, 0, :] # [:, 0, :] to remove dummy middle dimension
# Scale back to original
gp_samples_P = gp_samples_P_sc.copy()
for i in range(n_gp_samples):
gp_samples_P[:, i] = gp_samples_P_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_P = pd.DataFrame(gp_samples_P, columns = gp_names)
doses_opt_P_samples = pd.DataFrame({"NtotDose":0, "PtotDose":0, "KtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_P_samples.loc[i, ["NtotDose", "PtotDose", "KtotDose"]] = dose_grid_P.loc[gp_ilr_P.loc[:, i].idxmax(), ["NtotDose", "PtotDose", "KtotDose"]]
doses_opt_P_samples.loc[i, "ilr_"] = gp_ilr_P.loc[gp_ilr_P.loc[:, i].idxmax(), i]
doses_opt_P_samples["P"] = 0.436*doses_opt_P_samples.PtotDose
```
#### Prediction where only __K__ dosage varied, trial `r_sample`
```
dose_grid_K = pd.DataFrame({'NtotDose': opt_doses_gp.NtotDose,
'PtotDose': opt_doses_gp.PtotDose,
'KtotDose': np.linspace(0, 250, 251)})
dose_grid_K_sc = dose_grid_K.copy()
for var in dose_vars:
dose_grid_K_sc[var] = (dose_grid_K[var] - mean_numvars[var]) / std_numvars[var]
reference_stack_K = pd.DataFrame(index = range(dose_grid_K.shape[0]), columns = reference_row.columns)
for i in range(reference_stack_K.shape[1]):
reference_stack_K.iloc[:, i] = np.repeat(reference_row.values[0, i], reference_stack_K.shape[0])
reference_stack_K.NtotDose = dose_grid_K_sc.NtotDose
reference_stack_K.PtotDose = dose_grid_K_sc.PtotDose
reference_stack_K.KtotDose = dose_grid_K_sc.KtotDose
```
Point estimate
```
ilr_K_sc = gpms.predict(reference_stack_K)
dose_grid_K["ilr_"] = ilr_K_sc[:, 0] * std_respvars.values + mean_respvars.values
dose_grid_K['K'] = 0.830*dose_grid_K.KtotDose
doses_opt_K_point = dose_grid_K.loc[dose_grid_K["ilr_"].idxmax(), ["NtotDose", "PtotDose", "KtotDose", "ilr_"]]#.round(2)
doses_opt_K_point['K'] = 0.830*doses_opt_K_point.KtotDose
doses_opt_K_point
```
GP samples
```
n_gp_samples = 5
gp_samples_K_sc = gpms.sample_y(reference_stack_K, n_samples = n_gp_samples, random_state = 615108)[:, 0, :] # [:, 0, :] to remove dummy middle dimension
# Scale back to original
gp_samples_K = gp_samples_K_sc.copy()
for i in range(n_gp_samples):
gp_samples_K[:, i] = gp_samples_K_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_K = pd.DataFrame(gp_samples_K, columns = gp_names)
doses_opt_K_samples = pd.DataFrame({"NtotDose":0, "PtotDose":0, "KtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_K_samples.loc[i, ["NtotDose", "PtotDose", "KtotDose"]] = dose_grid_K.loc[gp_ilr_K.loc[:, i].idxmax(), ["NtotDose", "PtotDose", "KtotDose"]]
doses_opt_K_samples.loc[i, "ilr_"] = gp_ilr_K.loc[gp_ilr_K.loc[:, i].idxmax(), i]
doses_opt_K_samples["K"] = 0.830*doses_opt_K_samples.KtotDose
```
#### Plot all cases
Results are presented as the average GP and its point optimal dose in blue, with `10` gaussian processes and their optimum. Histograms of probabilistic dosages of maximum ilr values are shown under the response curves.
```
fig = plt.figure(figsize=(12, 5))
gs = gridspec.GridSpec(2, 3, height_ratios=[3, 1.5])
# N
ax0 = plt.subplot(gs[0])
ax0.plot(dose_grid_N.NtotDose, dose_grid_N.ilr_, linewidth = 2, color = "black")
ax0.plot(doses_opt_N_point.NtotDose, doses_opt_N_point.ilr_, marker = "o", color = "blue")
for i in gp_ilr_N.columns:
ax0.plot(dose_grid_N.NtotDose, gp_ilr_N[i], color = "black", alpha=0.2)
ax0.plot(doses_opt_N_samples.loc[i, "NtotDose"], doses_opt_N_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.2)
if r_sample.test_type.values[0] == "N":
plot_Ndata = df_ml.loc[id_table.NoEssai == r_sample.NoEssai.values[0]][["tsizeS_M", "NtotDose"]]
plt.plot(plot_Ndata.NtotDose, plot_Ndata.tsizeS_M, "o", color = "black", alpha = 0.2)
ax0.set_ylim([-2, 3])
ax0.set_xlabel(r"N dose ($kg~ha^{-1})$")
ax0.set_ylabel("Balance [S|M], GP model")
ax0.set_title("Trial N° " + str(NoEssai))
ax1 = plt.subplot(gs[3])
ax1.hist(doses_opt.loc[:, "NtotDose"], color = "lightgrey", edgecolor = 'grey')
ax1.set_xlabel("Optimal N dose ($kg~ha^{-1})$")
ax1.set_ylabel("Counts")
# P
ax2 = plt.subplot(gs[1])
ax2.plot(dose_grid_P.P, dose_grid_P.ilr_, linewidth = 2, color = "black")
ax2.plot(doses_opt_P_point.P, doses_opt_P_point.ilr_, marker = "o", color = "blue")
for i in gp_ilr_P.columns:
ax2.plot(dose_grid_P.P, gp_ilr_P[i], color = "black", alpha=0.2)
ax2.plot(doses_opt_P_samples.loc[i, "P"], doses_opt_P_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.2)
if r_sample.test_type.values[0] == "P":
plot_Pdata = df_ml.loc[id_table.NoEssai == r_sample.NoEssai.values[0]][["tsizeS_M", "PtotDose"]]
plot_Pdata['P'] = 0.436*plot_Pdata.PtotDose
plt.plot(plot_Pdata.P, plot_Pdata.tsizeS_M, "o", color = "black", alpha = 0.2)
ax2.set_ylim([-2, 3])
ax2.set_title("Trial N° " + str(NoEssai))
ax3 = plt.subplot(gs[4])
ax3.hist(doses_opt.loc[:, "P"], color = "lightgrey", edgecolor = 'grey')
ax3.set_xlabel(r"Optimal dose P ($kg~ha^{-1})$")
# K
ax4 = plt.subplot(gs[2])
ax4.plot(dose_grid_K.K, dose_grid_K.ilr_, linewidth = 2, color = "black")
ax4.plot(doses_opt_K_point.K, doses_opt_K_point.ilr_, marker = "o", color = "blue")
for i in gp_ilr_K.columns:
ax4.plot(dose_grid_K.K, gp_ilr_K[i], color = "black", alpha=0.2)
ax4.plot(doses_opt_K_samples.loc[i, "K"], doses_opt_K_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.2)
if r_sample.test_type.values[0] == "K":
plot_Kdata = df_ml.loc[id_table.NoEssai == r_sample.NoEssai.values[0]][["tsizeS_M", "KtotDose"]]
plot_Kdata['K'] = 0.830*plot_Kdata.KtotDose
plt.plot(plot_Kdata.K, plot_Kdata.tsizeS_M, "o", color = "black", alpha = 0.2)
ax4.set_ylim([-2, 3])
ax4.set_title("Trial N° " + str(NoEssai))
ax5 = plt.subplot(gs[5])
ax5.hist(doses_opt.loc[:, "K"], color = "lightgrey", edgecolor = 'grey')
ax5.set_xlabel(r"Optimal dose K ($kg~ha^{-1})$")
plt.tight_layout()
```
The distributions are almost similar to those from the uniform random grid doses table. Probabilistic optimum are on the edges of ranges corresponding or not to the prior optimum (_blue point_).
### Probabilistic predictions based on __trial type__
These chains make probabilistic predictions for the three N, P and K trials already modeled using `id_sample_n`, `id_sample_p` and `id_sample_k` respectively.
```
# GP samples
n_gp_samples = 1000 #500
gp_samples_N_sc = gpms.sample_y(reference_stack_N, n_samples = n_gp_samples, random_state = 615108)[:, 0, :]
# Scale back to original
gp_samples_N = gp_samples_N_sc.copy()
for i in range(n_gp_samples):
gp_samples_N[:, i] = gp_samples_N_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_N = pd.DataFrame(gp_samples_N, columns = gp_names)
doses_opt_N_samples = pd.DataFrame({"NtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_N_samples.loc[i, ["NtotDose"]] = dose_grid_N.loc[gp_ilr_N.loc[:, i].idxmax(), ["NtotDose"]]
doses_opt_N_samples.loc[i, "ilr_"] = gp_ilr_N.loc[gp_ilr_N.loc[:, i].idxmax(), i]
# GP samples
n_gp_samples = 1000 #500
gp_samples_P_sc = gpms.sample_y(reference_stack_P, n_samples = n_gp_samples, random_state = 615108)[:, 0, :]
# Scale back to original
gp_samples_P = gp_samples_P_sc.copy()
for i in range(n_gp_samples):
gp_samples_P[:, i] = gp_samples_P_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_P = pd.DataFrame(gp_samples_P, columns = gp_names)
doses_opt_P_samples = pd.DataFrame({"PtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_P_samples.loc[i, ["PtotDose"]] = dose_grid_P.loc[gp_ilr_P.loc[:, i].idxmax(), ["PtotDose"]]
doses_opt_P_samples.loc[i, "ilr_"] = gp_ilr_P.loc[gp_ilr_P.loc[:, i].idxmax(), i]
doses_opt_P_samples["P"] = 0.436*doses_opt_P_samples.PtotDose
# GP samples
n_gp_samples = 1000 #500
gp_samples_K_sc = gpms.sample_y(reference_stack_K, n_samples = n_gp_samples, random_state = 615108)[:, 0, :]
# Scale back to original
gp_samples_K = gp_samples_K_sc.copy()
for i in range(n_gp_samples):
gp_samples_K[:, i] = gp_samples_K_sc[:, i] * std_respvars.values + mean_respvars.values
# Pretty table
gp_names = list(range(n_gp_samples))
for i in range(n_gp_samples):
gp_names[i] = "gp" + str(gp_names[i])
gp_ilr_K = pd.DataFrame(gp_samples_K, columns = gp_names)
doses_opt_K_samples = pd.DataFrame({"KtotDose":0, "ilr_": 0}, index = gp_names)
for i in gp_names:
doses_opt_K_samples.loc[i, ["KtotDose"]] = dose_grid_K.loc[gp_ilr_K.loc[:, i].idxmax(), ["KtotDose"]]
doses_opt_K_samples.loc[i, "ilr_"] = gp_ilr_K.loc[gp_ilr_K.loc[:, i].idxmax(), i]
doses_opt_K_samples["K"] = 0.830*doses_opt_K_samples.KtotDose
fig = plt.figure(figsize=(10, 5))
gs = gridspec.GridSpec(2, 3,
width_ratios=[1,1,1],
height_ratios=[3, 1]
)
# N
ax0 = plt.subplot(gs[0])
ax0.plot(dose_grid_N.NtotDose, dose_grid_N.ilr_, "-", color = "black")
ax0.plot(Ndata.NtotDose, Ndata.tsizeS_M, "x", color = "black", alpha = 0.15)
ax0.plot(doses_opt_N_point.NtotDose, doses_opt_N_point.ilr_, marker = "o", color = "black")
for i in gp_ilr_N.columns[0:5]:
ax0.plot(dose_grid_N.NtotDose, gp_ilr_N[i], color = "black", alpha=0.15)
ax0.plot(doses_opt_N_samples.loc[i, "NtotDose"], doses_opt_N_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.15)
#ax0.set_ylim([-1, 2])
ax0.set_xlabel(r"N dose ($kg~ha^{-1})$")
ax0.set_ylabel("Balance [S|M], GP model")
ax0.set_title("N, sample no " + str(id_sample_n.NoEssai.values[0]))
ax1 = plt.subplot(gs[3])
ax1.hist(doses_opt_N_samples.loc[:, "NtotDose"], color = "lightgrey", edgecolor = 'grey')
ax1.axvline(x = doses_opt_N_point.NtotDose, color = 'black', linestyle = '--')
ax1.set_xlabel(r"Optimal dose N ($kg~ha^{-1})$")
ax1.set_ylabel("Count")
# P
ax2 = plt.subplot(gs[1])
ax2.plot(dose_grid_P.P, dose_grid_P.ilr_, "-", color = "black")
ax2.plot(Pdata.P, Pdata.tsizeS_M, "x", color = "black", alpha = 0.15)
ax2.plot(doses_opt_P_point.P, doses_opt_P_point.ilr_, marker = "o", color = "black")
for i in gp_ilr_P.columns[0:5]:
ax2.plot(dose_grid_P.P, gp_ilr_P[i], color = "black", alpha=0.15)
ax2.plot(doses_opt_P_samples.loc[i, "P"], doses_opt_P_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.15)
#ax2.set_ylim([-1, 2])
ax2.set_title("P, sample no " + str(id_sample_p.NoEssai.values[0]))
ax2.set_xlabel(r"P dose ($kg~ha^{-1})$")
ax3 = plt.subplot(gs[4])
ax3.hist(doses_opt_P_samples.loc[:, "P"], color = "lightgrey", edgecolor = 'grey')
ax3.axvline(x = doses_opt_P_point.P, color = 'black', linestyle = '--')
ax3.set_xlabel(r"Optimal dose P ($kg~ha^{-1})$")
# K
ax4 = plt.subplot(gs[2])
ax4.plot(dose_grid_K.K, dose_grid_K.ilr_, "-", color = "black")
ax4.plot(Kdata.K, Kdata.tsizeS_M, "x", color = "black", alpha = 0.15)
ax4.plot(doses_opt_K_point.K, doses_opt_K_point.ilr_, marker = "o", color = "black")
for i in gp_ilr_K.columns[0:5]:
ax4.plot(dose_grid_K.K, gp_ilr_K[i], color = "black", alpha=0.15)
ax4.plot(doses_opt_K_samples.loc[i, "K"], doses_opt_K_samples.loc[i, "ilr_"],
marker = "o", color = "black", alpha = 0.15)
#ax4.set_ylim([-1, 2])
ax4.set_title("K, sample no " + str(id_sample_k.NoEssai.values[0]))
ax4.set_xlabel(r"K dose ($kg~ha^{-1})$")
ax5 = plt.subplot(gs[5])
ax5.hist(doses_opt_K_samples.loc[:, "K"], color = "lightgrey", edgecolor = 'grey')
ax5.axvline(x = doses_opt_K_point.K, color = 'black', linestyle = '--')
ax5.set_xlabel(r"Optimal dose K ($kg~ha^{-1})$")
plt.tight_layout();
plt.savefig("images/MS_probab_doses.png", bbox_inches = "tight", dpi = 300)
index = ["NtotDose", "PtotDose", "KtotDose", "ilr_"]
opt_points = pd.DataFrame({"N trial": doses_opt_N_point,
"P trial": doses_opt_P_point,
"K trial": doses_opt_K_point},
index = index)
opt_points
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Random number generation
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/random_numbers"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/random_numbers.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/random_numbers.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/random_numbers.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
TensorFlow provides a set of pseudo-random number generators (RNG), in the `tf.random` module. This document describes how you can control the random number generators, and how these generators interact with other tensorflow sub-systems.
TensorFlow provides two approaches for controlling the random number generation process:
1. Through the explicit use of `tf.random.Generator` objects. Each such object maintains a state (in `tf.Variable`) that will be changed after each number generation.
2. Through the purely-functional stateless random functions like `tf.random.stateless_uniform`. Calling these functions with the same arguments (which include the seed) and on the same device will always produce the same results.
Warning: The old RNGs from TF 1.x such as `tf.random.uniform` and `tf.random.normal` are not yet deprecated but strongly discouraged.
Warning: The random numbers are not guaranteed to be consistent across TensorFlow versions, see: [Version Compatibility](https://www.tensorflow.org/guide/versions#what_is_not_covered)
## Setup
```
import tensorflow as tf
# Creates 2 virtual devices cpu:0 and cpu:1 for using distribution strategy
physical_devices = tf.config.list_physical_devices("CPU")
tf.config.experimental.set_virtual_device_configuration(
physical_devices[0], [
tf.config.experimental.VirtualDeviceConfiguration(),
tf.config.experimental.VirtualDeviceConfiguration()
])
```
## The `tf.random.Generator` class
The `tf.random.Generator` class is used in cases where you want each RNG call to produce different results. It maintains an internal state (managed by a `tf.Variable` object) which will be updated every time random numbers are generated. Because the state is managed by `tf.Variable`, it enjoys all facilities provided by `tf.Variable` such as easy checkpointing, automatic control-dependency and thread safety.
You can get a `tf.random.Generator` by manually creating an object of the class or call `tf.random.get_global_generator()` to get the default global generator:
```
g1 = tf.random.Generator.from_seed(1)
print(g1.normal(shape=[2, 3]))
g2 = tf.random.get_global_generator()
print(g2.normal(shape=[2, 3]))
```
There are multiple ways to create a generator object. The easiest is `Generator.from_seed`, as shown above, that creates a generator from a seed. A seed is any non-negative integer. `from_seed` also takes an optional argument `alg` which is the RNG algorithm that will be used by this generator:
```
g1 = tf.random.Generator.from_seed(1, alg='philox')
print(g1.normal(shape=[2, 3]))
```
See the *Algorithms* section below for more information about it.
Another way to create a generator is with `Generator.from_non_deterministic_state`. A generator created this way will start from a non-deterministic state, depending on e.g. time and OS.
```
g = tf.random.Generator.from_non_deterministic_state()
print(g.normal(shape=[2, 3]))
```
There are yet other ways to create generators, such as from explicit states, which are not covered by this guide.
When using `tf.random.get_global_generator` to get the global generator, you need to be careful about device placement. The global generator is created (from a non-deterministic state) at the first time `tf.random.get_global_generator` is called, and placed on the default device at that call. So, for example, if the first site you call `tf.random.get_global_generator` is within a `tf.device("gpu")` scope, the global generator will be placed on the GPU, and using the global generator later on from the CPU will incur a GPU-to-CPU copy.
There is also a function `tf.random.set_global_generator` for replacing the global generator with another generator object. This function should be used with caution though, because the old global generator may have been captured by a `tf.function` (as a weak reference), and replacing it will cause it to be garbage collected, breaking the `tf.function`. A better way to reset the global generator is to use one of the "reset" functions such as `Generator.reset_from_seed`, which won't create new generator objects.
```
g = tf.random.Generator.from_seed(1)
print(g.normal([]))
print(g.normal([]))
g.reset_from_seed(1)
print(g.normal([]))
```
### Creating independent random-number streams
In many applications one needs multiple independent random-number streams, independent in the sense that they won't overlap and won't have any statistically detectable correlations. This is achieved by using `Generator.split` to create multiple generators that are guaranteed to be independent of each other (i.e. generating independent streams).
```
g = tf.random.Generator.from_seed(1)
print(g.normal([]))
new_gs = g.split(3)
for new_g in new_gs:
print(new_g.normal([]))
print(g.normal([]))
```
`split` will change the state of the generator on which it is called (`g` in the above example), similar to an RNG method such as `normal`. In addition to being independent of each other, the new generators (`new_gs`) are also guaranteed to be independent of the old one (`g`).
Spawning new generators is also useful when you want to make sure the generator you use is on the same device as other computations, to avoid the overhead of cross-device copy. For example:
```
with tf.device("cpu"): # change "cpu" to the device you want
g = tf.random.get_global_generator().split(1)[0]
print(g.normal([])) # use of g won't cause cross-device copy, unlike the global generator
```
Note: In theory, you can use constructors such as `from_seed` instead of `split` here to obtain a new generator, but by doing so you lose the guarantee that the new generator is independent of the global generator. You will also run the risk that you may accidentally create two generators with the same seed or with seeds that lead to overlapping random-number streams.
You can do splitting recursively, calling `split` on splitted generators. There are no limits (barring integer overflow) on the depth of recursions.
### Interaction with `tf.function`
`tf.random.Generator` obeys the same rules as `tf.Variable` when used with `tf.function`. This includes three aspects.
#### Creating generators outside `tf.function`
`tf.function` can use a generator created outside of it.
```
g = tf.random.Generator.from_seed(1)
@tf.function
def foo():
return g.normal([])
print(foo())
```
The user needs to make sure that the generator object is still alive (not garbage-collected) when the function is called.
#### Creating generators inside `tf.function`
Creation of generators inside a `tf.function` can only happend during the first run of the function.
```
g = None
@tf.function
def foo():
global g
if g is None:
g = tf.random.Generator.from_seed(1)
return g.normal([])
print(foo())
print(foo())
```
#### Passing generators as arguments to `tf.function`
When used as an argument to a `tf.function`, different generator objects with the same state size (state size is determined by the RNG algorithm) won't cause retracing of the `tf.function`, while those with different state sizes will.
```
num_traces = 0
@tf.function
def foo(g):
global num_traces
num_traces += 1
return g.normal([])
foo(tf.random.Generator.from_seed(1))
foo(tf.random.Generator.from_seed(2))
print(num_traces)
```
### Interaction with distribution strategies
There are three ways in which `Generator` interacts with distribution strategies.
#### Creating generators outside distribution strategies
If a generator is created outside strategy scopes, all replicas’ access to the generator will be serialized, and hence the replicas will get different random numbers.
```
g = tf.random.Generator.from_seed(1)
strat = tf.distribute.MirroredStrategy(devices=["cpu:0", "cpu:1"])
with strat.scope():
def f():
print(g.normal([]))
results = strat.run(f)
```
Note that this usage may have performance issues because the generator's device is different from the replicas.
#### Creating generators inside distribution strategies
Creating generators inside strategy scopes is disallowed, because there is ambiguity on how to replicate a generator (e.g. should it be copied so that each replica gets the same random numbers, or 'split' so that each replica gets different random numbers).
```
strat = tf.distribute.MirroredStrategy(devices=["cpu:0", "cpu:1"])
with strat.scope():
try:
tf.random.Generator.from_seed(1)
except ValueError as e:
print("ValueError:", e)
```
Note that `Strategy.run` will run its argument function in a strategy scope implicitly:
```
strat = tf.distribute.MirroredStrategy(devices=["cpu:0", "cpu:1"])
def f():
tf.random.Generator.from_seed(1)
try:
strat.run(f)
except ValueError as e:
print("ValueError:", e)
```
#### Passing generators as arguments to `Strategy.run`
If you want each replica to use its own generator, you need to make `n` generators (either by copying or splitting), where `n` is the number of replicas, and then pass them as arguments to `Strategy.run`.
```
strat = tf.distribute.MirroredStrategy(devices=["cpu:0", "cpu:1"])
gs = tf.random.get_global_generator().split(2)
# to_args is a workaround for the absence of APIs to create arguments for
# run. It will be replaced when such APIs are available.
def to_args(gs):
with strat.scope():
def f():
return [gs[tf.distribute.get_replica_context().replica_id_in_sync_group]]
return strat.run(f)
args = to_args(gs)
def f(g):
print(g.normal([]))
results = strat.run(f, args=args)
```
## Stateless RNGs
Usage of stateless RNGs is simple. Since they are just pure functions, there is no state or side effect involved.
```
print(tf.random.stateless_normal(shape=[2, 3], seed=[1, 2]))
print(tf.random.stateless_normal(shape=[2, 3], seed=[1, 2]))
```
Every stateless RNG requires a `seed` argument, which needs to be an integer Tensor of shape `[2]`. The results of the op are fully determined by this seed.
## Algorithms
### General
Both the `tf.random.Generator` class and the `stateless` functions support the Philox algorithm (written as `"philox"` or `tf.random.Algorithm.PHILOX`) on all devices.
Different devices will generate the same integer numbers, if using the same algorithm and starting from the same state. They will also generate "almost the same" float-point numbers, though there may be small numerical discrepancies caused by the different ways the devices carry out the float-point computation (e.g. reduction order).
### XLA devices
On XLA-driven devices (such as TPU, and also CPU/GPU when XLA is enabled) the ThreeFry algorithm (written as `"threefry"` or `tf.random.Algorithm.THREEFRY`) is also supported. This algorithm is fast on TPU but slow on CPU/GPU compared to Philox.
See paper ['Parallel Random Numbers: As Easy as 1, 2, 3'](https://www.thesalmons.org/john/random123/papers/random123sc11.pdf) for more details about these algorithms.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.