text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Tabulation
In this tutorial, we will explore samplics' APIs for creating design-based tabulations. There are two main python classes for tabulation i.e. ```Tabulation()``` for one-way tables and ```CrossTabulation()``` for two-way tables.
```
from pprint import pprint
from samplics.datasets import load_birth, load_nhanes2
from samplics.categorical import Tabulation, CrossTabulation
```
## One-way tabulation
The birth dataset has four variables: region, agecat, birthcat, and pop. The variables agecat and birthcat are categirical. By default, pandas read them as numerical, because they are coded with numerical values. We use ```dtype="string"``` or ``` dtype="category"``` to ensure that pandas codes the variables as categorical responses.
```
# Load Birth sample data
birth_dict = load_birth()
birth = birth_dict["data"].astype(
{"region": str, "agecat": str, "birthcat": str}
)
region = birth["region"]
agecat = birth["agecat"]
birthcat = birth["birthcat"]
birth.head(15)
```
When requesting a table, the user can set ```parameter="count"``` which results in a tabulation with counts in the cells while ```parameter="proportion``` leads to cells with proportions. The expression ```Tabulation("count")``` instantiates the class ```Tabulation()``` which has a method ```tabulate()``` to produce the table.
```
birth_count = Tabulation(parameter="count")
birth_count.tabulate(birthcat, remove_nan=True)
print(birth_count)
```
When ```remove_nan=False```, the numpy and pandas special values NaNs, respectively np.nan and NaN, are treated as valid categories and added to the tables as shown below
```
birth_count = Tabulation(parameter="count")
birth_count.tabulate(birthcat, remove_nan=False)
print(birth_count)
```
The data associated with the tabulation are stored in nested python dictionnaires. The higher level key is the variable name and the inner keys are the response categories. Each of the last four columns shown above are stored in separated dictionnaires. Two of those dictionnaires for the counts and standard errors shown below.
```
print("\nThe designed-based estimated counts are:")
pprint(birth_count.point_est)
print("\nThe designed-based standard errors are:")
pprint(birth_count.stderror)
```
Sometimes, the user may want to run multiple one-way tabules of several variables. In this case, the user can provide the data as a two-dimensional dataframe where each column represents one categorical variable. In this situation, each categorical variable is tabulated individually then combined into Python dictionnaires.
```
birth_count2 = Tabulation(parameter="count")
birth_count2.tabulate(birth[["region", "agecat", "birthcat"]], remove_nan=True)
print(birth_count2)
```
Two of the associated Python dictionnaires are shown below. The structure of the inner dictionnaires remain the same but additional key-value pairs are added to represent the several categorical variables.
```
print("\nThe designed-based estimated counts are:")
pprint(birth_count2.point_est)
print("\nThe designed-based standard errors are:")
pprint(birth_count2.stderror)
```
In the example above, we used pandas series and dataframes with labelled variables. In some situations, the user may want to tabulate numpy arrays, lists or tuples without variable names atrribute from the data. For these situations, the ```varnames``` parameter provides a way to assign names for the categorical variables. Even when the variables have labels, users can leverage ```varnames``` to rename the categorical variables.
```
region_no_name = birth["region"].to_numpy()
agecat_no_name = birth["agecat"].to_numpy()
birthcat_no_name = birth["birthcat"].to_numpy()
birth_prop_new_name = Tabulation(parameter="proportion")
birth_prop_new_name.tabulate(
vars=[region_no_name, agecat_no_name, birthcat_no_name],
varnames=["Region", "AgeGroup", "BirthType"],
remove_nan=True,
)
print(birth_prop_new_name)
```
If the user does not specify ```varnames```, the ```tabulate()``` creates generic variables names ```var_1, var_2, etc```.
```
birth_prop_new_name2 = Tabulation(parameter="proportion")
birth_prop_new_name2.tabulate(
vars=[region_no_name, agecat_no_name, birthcat_no_name], remove_nan=True
)
print(birth_prop_new_name2)
```
If the data was collected from a complex survey sample, the user may provide the sample design information to derive design-based statistics for the tabulation.
```
# Load Nhanes sample data
nhanes2_dict = load_nhanes2()
nhanes2 = nhanes2_dict["data"]
stratum = nhanes2["stratid"]
psu = nhanes2["psuid"]
weight = nhanes2["finalwgt"]
diabetes_nhanes = Tabulation("proportion")
diabetes_nhanes.tabulate(
vars=nhanes2[["race", "diabetes"]],
samp_weight=weight,
stratum=stratum,
psu=psu,
remove_nan=True,
)
print(diabetes_nhanes)
```
## Two-way tabulation (cross-tabulation)
Cross-tabulation of two categorical variables is achieved by using the class ```CrossTabulation()```. As above, cross-tabulation is possible for counts and proportions using ```CrossTabulation(parameter="count")``` and ```CrossTabulation(parameter="proportion")```, respectively. The Python script below creates a design-based cross-tabulation of race by diabetes status. The sample design information is optional; when not provided, a simple random sample (srs) is assumed.
```
crosstab_nhanes = CrossTabulation("proportion")
crosstab_nhanes.tabulate(
vars=nhanes2[["race", "diabetes"]],
samp_weight=weight,
stratum=stratum,
psu=psu,
remove_nan=True,
)
print(crosstab_nhanes)
```
In addition to pandas dataframe, the categorical variables may be provided as an numpy array, list or tuple. In the examples below, the categorical variables are provided as a tuple ```vars=(rage, diabetes).``` In this case, ```race``` and ```diabetes``` are numpy arrays and do not have a name attribute. The parameter ```varnames``` allows the user to name the categorical variables. If varnames is not specified then ```var_1``` and ```var_2``` are used as variables names.
```
race = nhanes2["race"].to_numpy()
diabetes = nhanes2["diabetes"].to_numpy()
crosstab_nhanes = CrossTabulation("proportion")
crosstab_nhanes.tabulate(
vars=(race, diabetes),
samp_weight=weight,
stratum=stratum,
psu=psu,
remove_nan=True,
)
print(crosstab_nhanes)
```
Same as the above example with variables names specified by ```varnames=["Race", DiabetesStatus"]```
```
crosstab_nhanes = CrossTabulation("proportion")
crosstab_nhanes.tabulate(
vars=(race, diabetes),
varnames=["Race", "DiabetesStatus"],
samp_weight=weight,
stratum=stratum,
psu=psu,
remove_nan=True,
)
print(crosstab_nhanes)
```
| github_jupyter |
```
# Import Library
import os,csv
import pandas as pd
from tqdm import trange
import copy
import numpy as np
# Enter inputs so that you can just execute other cells
first_file = "result_without_coco_24000.txt"
second_file = "result_coco.txt"
# Read results from darknet prediction output for first file and process it
#os.chdir("E:/Machine Learning/Kaggle - Open Images Challenge/darknet/build/darknet/x64")
os.chdir("E:/Machine Learning/Kaggle - Open Images Challenge")
submission_text = []
with open(first_file) as f:
submission_text = f.readlines()
# Trim "\n" from end of each line from text file
for i in trange(len(submission_text)):
submission_text[i] = submission_text[i][:-2]
# Save image name and prediction string separately
submission_ImageId = []
submission_pred = []
for i in trange(len(submission_text)):
submission_ImageId.append(submission_text[i][10:26])
submission_pred.append(submission_text[i][31:])
first_file_processed = pd.DataFrame({"ImageId":submission_ImageId,"PredictionString":submission_pred})
first_file_processed.to_csv("first_file.csv",index=False)
# Read results from darknet prediction output for second file and process it
#os.chdir("E:/Machine Learning/Kaggle - Open Images Challenge/darknet/build/darknet/x64")
os.chdir("E:/Machine Learning/Kaggle - Open Images Challenge")
submission_text = []
with open(second_file) as f:
submission_text = f.readlines()
# Trim "\n" from end of each line from text file
for i in trange(len(submission_text)):
submission_text[i] = submission_text[i][:-2]
# Save image name and prediction string separately
submission_ImageId = []
submission_pred = []
for i in trange(len(submission_text)):
submission_ImageId.append(submission_text[i][10:26])
submission_pred.append(submission_text[i][31:])
# # Replaces wrongly labeled label
# for i in trange(len(submission_pred)):
# if "traffic light" in submission_pred[i]:
# submission_pred[i]=submission_pred[i].replace('traffic light','/m/015qff')
# Save "submission_ImageId" and "submission_pred" under correct headers for final submission
second_file_processed = pd.DataFrame({"ImageId":submission_ImageId,"PredictionString":submission_pred})
second_file_processed.to_csv("second_file.csv",index=False)
# Merge the two files
if (len(first_file_processed['ImageId'])!=len(second_file_processed['ImageId'])):
print("Verify entry numbers for each files..")
else:
merged_prediction = []
for i in trange(len(first_file_processed['ImageId'])):
merged_prediction.append(first_file_processed['PredictionString'][i]+" "+second_file_processed['PredictionString'][i])
# Write new csv
merged_data_df = pd.DataFrame({"ImageID":first_file_processed['ImageId'],"PredictionString":merged_prediction})
merged_data_df.to_csv("merged.csv",index=False)
print("It is saved in :",os.getcwd())
```
| github_jupyter |
# 交談 AI
思考一下,您透過立即訊息、社交媒體、電子郵件或其它線上技術與其他人進行交流的頻率。對大多數人而言,這是我們尋求交流的方式。當您在工作上遇到問題,您也許能使用聊天訊息與同事聯絡,您也可以在行動裝置上使用該聊天訊息,以便隨時保持聯絡。

Bot 是使用這類通道進行對話的 AI 代理程式,啟用了自然式對話的業務開發和軟體服務。
## 建立 QnA Maker 知識庫
對於客戶支援案例,通常會建立一個可以透過網站聊天視窗、電子郵件或語音介面解釋和回答常見問題集的 Bot。Bot 介面之下是問題和適當答案之知識庫,可供 Bot 搜尋適當回應。
QnA Maker 服務是 Azure 中的一項認知服務,該服務使您可以透過輸入問答配對或從現有文件或網頁中快速建立知識庫。然後,它可以使用一些內嵌式自然語言處理功能來解釋問題並找到適當的答案。
1. 打開其它瀏覽器索引標籤並透過 https://qnamaker.ai 前往 QnA Maker 入口網站。請使用與您的 Azure 訂閱關聯的 Microsoft 帳戶登入。
2. 在 QnA Maker 入口網站中,請選取 **[建立知識庫]**。
3. 如果您以前未建立 QnA 服務資源,請選取 **[建立 QnA 服務]**。Azure 入口網站將在另一個索引標籤中打開,因此您可以在訂閱中建立 QnA Maker 服務。使用以下設定:
- **訂用帳戶**: *您的 Azure 訂用帳戶*
- **資源群組**: *選取現有的資源群組或建立新的資料群組*
- **名稱**: *您的 QnA 資源之唯一名稱*
- **定價層**:F0
- **Azure 搜尋定價層**:F
- **Azure 搜尋位置**: *任何可用位置*
- **應用程式名稱**:_與**名稱**一致 (".azurewebsites.net” 將自動附加)_
- **網站位置**:_與 **Azure 搜尋位置**_一致
- **應用程式深入解析**:停用
> **備註**:如果您已經建置了免費層 **QnA Maker**或 **Azure 搜尋**資源,您的配額可能不允許您建立另一個配額。在這種狀況下,選取 **F0 / F**之外的其它層。
4. 等待 QnA 服務和相關資源的部署在 Azure 入口網站中完成。
5. 退回到 QnA Maker 入口網站索引標籤,然後在**步驟 2**區段中,按一下 **[重新整理]** 以重新整理可用的 QnA 服務資源清單。
6. 透過選取以下選項,將您的 QnA 服務連接到您的知識庫:
- **Microsoft Azure 目錄識別碼**: *您的訂閱之 Azure 目錄識別碼*
- **Azure 訂閱名稱**: *您的 Azure 訂閱*
- **Azure QnA 服務**: *您在上一個步驟中建立的 QnA 服務資源*
- **語言**:英文
### (!)簽入
如果顯示了角色無權限執行操作的訊息指示,請重新整理 QnA Maker 入口網站的瀏覽器頁面。
7. 在**步驟 3**區段,請輸入名稱 **Margie's Travel KB**。
8. 在**步驟 4** 區段,在 **URL** 方塊中,鍵入 *https://github.com/MicrosoftLearning/mslearn-ai900/raw/main/data/qna_bot/margies_faq.docx* 並按一下 **[+ 新增 URL]**。在 **[閒聊]** 下,選取 **[專業]**。
9. 在**步驟 5** 區段,按一下 **[建立知識庫]**。
10. 在建立您的知識庫時,請稍等片刻。然後檢閱已從常見問題集文件中匯入的問題和答案以及專業的閒聊預先定義回應。
## 編輯知識庫
您的知識庫以常見問題集文件中的詳細資料和一些預先定義的回應為基礎。您可以添加自訂問答配對來補充這些內容。
1. 按一下 **[+ 新增 QnA 配對]**。
2. 在 **[問題]** 方塊中,鍵入 `Hello`。然後按一下 **[+ 新增不同說法]** 並鍵入 `Hi`。
3. 在 **[回答]** 方塊中,鍵入 `Hello`。
## 訓練並測試知識庫
既然您擁有知識庫,就可以在 QnA Maker 入口網站測試知識庫。
1. 在頁面的右上角,按一下 **[儲存並訓練]** 以便訓練您的知識庫。您可能需要更改視窗的大小才能看到該按鈕。
2. 訓練完成後,請按一下 **[← 測試]** 以打開測試窗格。
3. 在測試窗格底部輸入訊息 *Hi*。應退回回應 **Hello**。
4. 在測試窗格底部輸*入訊息我想預定航班*。應該從常見問題集中退回適當的回應。
5. 完成知識庫測試之後,按一下 **[→ 測試]** 以關閉測試窗格。
## 為知識庫建立 Bot。
知識庫提供了後端服務,用戶端應用程式可以使用該後端服務透過某種用戶介面來回答問題。通常,這些用戶端應用程式是 Bot。為了使知識庫可用於 Bot,您必須將知識庫發佈為可以透过 HTTP 訪問的服務。然後,您可以使用 Azure Bot Service 建立並託管一個使用知識庫來回答用戶問題的 Bot。
1. 在 QnA Make 頁面的頂部,按一下 **[發佈]**。然後在 **Margies Travel KB** 頁面,按一下 **[發佈]**。
2. 服務部署好之後,按一下 **[建立 Bot]**。這將在新的瀏覽器索引標籤中打開 Azure 入口網站,以便您可以在 Azure 訂閱中建立 Web 應用程式 Bot。
3. 在 Azure 入口網站中,透過以下設定建立 Web 應用程式 Bot (大多數設定將會為您預先填入):
- **Bot 控制代碼**: *您的 Bot 之唯一名稱*
- **訂用帳戶**: *您的 Azure 訂用帳戶*
- **資源群組**: *包含您的 QnA Maker 資源的資源組*
- **位置**: *與您的 QnA Maker 服務位置相同*。
- **定價層**:F0
- **應用程式名稱**: *與自動附加.azurewebsites.net* 的 **Bot控制代碼** 相同
- **SDK 語言**: *選擇 C# 語言或 Node.js 語言*
- **QnA 授權金鑰**: *該金鑰應該自動設定為您的 QnA 知識庫之驗證金鑰*
- **App Service 方案/位置**: *該方案/位置應該自動設定為適當的方案和位置*
**Application Insights**:關
- **Microsoft App ID 和密碼**:自動建立 App ID 和密碼。
4. 等待您的 Bot 建立 (等待時會在右上方顯示通知圖示,該圖示看起來像個鈴鐺)。然後,在已完成部署的通知中,按一下 **[轉到資源]** (或者在首頁上,按一下 **[資源群組]**,打開已經在其中建立了 Web 應用程式 Bot 的資源群組,然後按一下這個資源群組。)
5. 在 Bot 的左側窗格中查找 **[設定]**,按一下 **[在網路聊天中測試]**,然後等待,直到 Bot 顯示訊息**您好和歡迎!**(這可能需要幾秒鐘來初始化)。
6. 使用測試聊天介面,確保您的 Bot 按照預想回答知識庫中的問題。例如,嘗試提交*我要取消酒店*。
## 透過通道存取 Bot
透過一個或多*個通道*,Bot 可用於為用戶提供介面。例如,同一個 Bot 可以透過網路聊天介面、電子郵件和 Microsoft Teams 支援互動。
1. 在 Bot 的左側窗格中,找到 **[設定]**,然後按一下 **[通道]**。
2. 請注意,已經自動新增了**網路聊天**通道,並且其他用於常用通訊平台的通道亦可使用。
3. 在**網路聊天**通道旁邊,按一下 **[編輯]**。這樣將打開一個頁面,其中包含將 Bot 內嵌到網頁所需的設置。若要內嵌 Bot,您需要提供 HTML 內嵌程式碼並為您的 Bot 生成祕密金鑰之中的一個金鑰。
4. 複製內**嵌程式碼**並將其貼上到下面的儲存格中,取代留言 `<!-- EMBED CODE GOES HERE -->`。
5. 為您的其中一個祕密金鑰 (無論哪一個均可) 按一下 **[顯示]**,並複製這個金鑰。然後,將這個金鑰貼上到下面的 HTML 內嵌程式碼中,取代 `YOUR_SECRET_HERE`。
6. 將 HTML 程式碼中的**最小高度**值更改為 **200px** (而不是預設的 500px)。這樣將有助於確保 HTML 介面無需捲動即可見。
7. 透過按一下儲存格左側的 **[執行儲存格]** (▷) 按鈕來執行下面的儲存格以轉譯 HTML。
8. 在 HTML 聊天介面中,透過提交問題來測試 Bot (例如,*誰是瑪姬?* 或*者我可以前往哪些目的地?* (Bot 初始化時,除了回答您的問題外,它可能還會回應訊息您好和*歡迎*)
```
%%html
<!-- EMBED CODE GOES HERE -->
```
試用 Bot。您可能會發現它可以非常準確地回答常見問題集中的問題,但是對於其未接受過訓練的問題,Bot 對這類問題的解釋能力有限。您可以一直使用 QnA Maker 入口網站來編輯知識庫以便對其進行改進,然後將知識庫重新發布。
## 瞭解更多資訊
- 若要瞭解關於 QnA Maker 服務的更多資訊,請檢視 [the QnA Maker documentation](https://docs.microsoft.com/azure/cognitive-services/qnamaker/)。
- 若要瞭解關於 Microsoft Bot Service 的更多資訊,請檢視 [the Azure Bot Service page](https://azure.microsoft.com/services/bot-service/)。
| github_jupyter |
# Dùng get-news để crawl dữ liệu tin tức
## 0. Change log
Ver0.1: Sử dụng thư viện [news-please](https://github.com/fhamborg/news-please), nhược điểm chưa lấy được URL của hình ảnh và videos
Ver0.2: Sử dụng [fake-useragent](https://pypi.org/project/fake-useragent/) để tạo _rotating proxy_ để có thể hỗ trợ lấy tin
Ver0.3: Sử dụng thêm thư viện BeautifulSoup để lấy thêm được URL của ảnh
## 1. Thư viện và các function
```
import json, os
from newsplease import NewsPlease
from bs4 import BeautifulSoup
from fake_useragent import UserAgent
from urllib.request import Request, urlopen
ua = UserAgent()
```
### 1.1. Các function liên quan đến tin tức
Từ các url tin tức, chúng ta đưa chúng vào trong 1 list, hàm này dùng thư viên _NewsPlease_ để đưa dữ liệu tin tức lấy được vào 1 list
```
def get_news_to_list(url_list):
news_list = []
for url in url_list:
try:
news_list.append(NewsPlease.from_url(url))
except Exception as e:
print(e, ': ', url)
continue
return news_list
```
Đưa các dữ liệu sẵn có vào file JSON với cấu trúc là ```<json_prefix>_<json_no>.json```
Các thông số cho json.dump lần lượt là: __default = str__ đưa tất cả về string để tránh lỗi trong quá trình parse dữ liệu dạng _datetime_ sang json của các trường date\_modify, date\_download, date\_publish; __ensure\_ascii=False__ là để giữ các dữ liệu text ở dạng Unicode - để không bị lỗi khi đưa vào file json khi để text ở định dạng Unicode, thì trong phần __open()__ đặt thông số _encoding='utf-8'_.
Các biến:
1. file\_name: tên file để lưu
2. file\_no: số thứ tự của file
```
def dump_news_to_json(news_list, json_begin_no, json_prefix, file_path):
file_no = json_begin_no
for news in news_list:
file_name = json_prefix + str(file_no) + '.json'
file_no += 1
with open(os.path.join(file_path, file_name), 'w', encoding='utf-8') as outfile:
json.dump(news.__dict__, outfile, indent=4, sort_keys=True, default=str, ensure_ascii=False)
```
### 1.2. Các function liên quan đến việc tạo rotating proxy
Tham khảo tại một số trang sau:
1. [Create a rotating proxy crawler in Python 3](https://codelike.pro/create-a-crawler-with-rotating-ip-proxy-in-python/)
```
def random_proxy(proxies):
return random.randint(0, len(proxies) - 1)
def retrive_proxies():
proxies_req = Request('https://sslproxies.org/')
proxies_req.add_header('User-Agent', ua.random)
proxies_doc = urlopen(proxies_req).read().decode('utf8')
```
## 2. Các dữ liệu tin tức giả
URL về Fake News, tập hợp từ các bài viết có nội dung không được xác minh về nguồn tin, hoặc được xác nhận sai,... Có thể lưu url các bài báo trong 1 file text (mỗi url là 1 dòng)
```
fake_news_urls = [
"https://www.vietgiaitri.com/phap-luat/201304/yeu-rau-xanh-p51-cau-chuyen-dong-troi-sau-tam-ri-do-mong-735408"
]
```
Bắt đầu crawl dữ liệu tin tức giả đưa vào list fake_articles
```
fake_articles = get_news_to_list(fake_news_urls)
```
Tên của các file json sẽ chứa nội dung vừa crawl
```
fake_no = 100
fake_prefix = 'VFND_Ac_Fake_'
```
Đưa dữ liệu đã lấy vào các file JSON để lưu trữ
```
dump_news_to_json(fake_articles, fake_no, fake_prefix, './')
```
Test các kết quả trả ra khi crawl ở cell dưới
```
print (fake_articles[0].__dict__)
```
## 3. Các dữ liệu tin tức thật
```
real_news_urls = [
"https://thethao.tuoitre.vn/danh-bai-dokovic-zverev-lan-dau-vo-dich-giai-quan-vot-atp-finals-20181119055342346.htm",
"https://thethao.tuoitre.vn/hoa-hau-quan-vot-bouchard-tro-tai-da-bong-2018112221552504.htm",
"https://thethao.tuoitre.vn/quan-vot-vn-coi-chung-lac-duong-nhu-thai-lan-821535.htm",
"https://thethao.tuoitre.vn/federer-dung-dokovic-o-ban-ket-giai-quan-vot-paris-masters-20181103062334363.htm",
"https://thethao.tuoitre.vn/nadal-gap-tsitsipas-o-chung-ket-giai-quan-vot-barcelona-mo-rong-1439844.htm",
"https://dantri.com.vn/phap-luat/khoi-to-cap-nu-quai-chuyen-cung-cap-ma-tuy-thuoc-lac-20181209204749864.htm",
"https://dantri.com.vn/phap-luat/dung-dao-dam-chet-nguoi-vi-xich-mich-luc-do-xang-20181209202623487.htm",
"https://dantri.com.vn/phap-luat/quen-khoa-o-to-nu-tai-xe-bi-khoang-sach-tai-san-20181209143646284.htm",
"https://dantri.com.vn/phap-luat/lua-ban-be-gai-14-tuoi-bat-luon-nguoi-giai-cuu-de-doi-tien-chuoc-20181209122608572.htm",
"https://dantri.com.vn/phap-luat/doi-no-60-trieu-khong-duoc-ep-nan-nhan-viet-giay-vay-400-trieu-20181209083854909.htm",
"https://dantri.com.vn/phap-luat/vu-chay-than-chet-nguoi-cuu-giam-doc-benh-vien-bi-truy-to-tu-3-12-nam-tu-20181209083714518.htm",
"https://dantri.com.vn/phap-luat/sau-ly-hon-vo-tinh-nhan-doi-chia-tay-nen-ra-tay-sat-hai-20181209082922084.htm",
"https://dantri.com.vn/phap-luat/duoi-theo-ke-nem-chat-ban-chu-nha-bi-gay-chan-2018120908113454.htm",
"https://dantri.com.vn/phap-luat/doi-tuong-cuoi-cung-trong-vu-tron-khoi-trai-tam-giam-kien-giang-da-bi-bat-20181209081958807.htm",
"https://dantri.com.vn/the-thao/sieu-may-bay-a350-se-cho-cdv-viet-nam-di-malaysia-co-vu-chung-ket-luot-di-20181208082125017.htm",
"https://dantri.com.vn/the-thao/thuong-20000-usd-cho-doi-tuyen-co-vua-viet-nam-tai-olympiad-2018-20181017151513152.htm",
"https://dantri.com.vn/the-thao/truong-son-gianh-hcv-tai-giai-co-vua-dong-doi-the-gioi-2018-20181005221940293.htm",
"https://dantri.com.vn/the-thao/chuyen-ve-chang-sinh-vien-luat-kien-tuong-le-tuan-minh-20180320085855246.htm",
"https://dantri.com.vn/the-thao/tien-dao-malaysia-toi-da-co-cach-vuot-qua-hang-thu-doi-tuyen-viet-nam-20181209191129676.htm"
]
```
Bắt đầu crawl dữ liệu, sử dụng thư viện news-please, các tin tức sẽ được lấy theo URL và được chứa trong mảng _real_\__article_.
```
real_articles = get_news_to_list(real_news_urls)
```
Do thư viện _news-please_ không hỗ trợ để lấy hình ảnh vì thế chúng ta sẽ lấy hình ảnh thông qua thư viện _BeautifulSoup_. Sau mỗi lần lấy chúng ta tạo ra 1 đối tượng dictionary cho mỗi URL
```
real_imgs = []
for real_url in real_news_urls:
```
Merge 2 đối tượng dictionary. Tham khảo từ: [How to merge two dictionaries in a single expression? - StackOverflow](https://stackoverflow.com/questions/38987/how-to-merge-two-dictionaries-in-a-single-expression)
Tương tự với việc crawl dữ liệu về tin tức giả. Ta có danh sách 1 số biến:
1. real\_name: tên file để lưu
2. real\_no: số thứ tự của file
```
real_no = 105
real_prefix = 'VFND_Ac_Real_'
for real in real_articles:
real_name = real_prefix + str(real_no) + '.json'
real_no = real_no + 1
with open(real_name, 'w', encoding='utf-8') as outfile:
json.dump(real.__dict__, outfile, indent=4, sort_keys=True, default=str, ensure_ascii=False)
```
Test các kết quả trả ra khi crawl ở cell dưới
```
print(real_articles[0].__dict__)
```
| github_jupyter |
# "Monte Carlo Methods 2"
> "In this blog post we continue to look at Monte Carlo methods and how they can be used. We move from sampling from univariate distributions to multi-variate distributions."
- toc: true
- author: Lewis Cole (2020)
- branch: master
- badges: false
- comments: false
- categories: [Monte-Carlo, Statistics, Probability, Computational-Statistics, Theory, Computation, Copula]
- hide: false
- search_exclude: false
- image: https://github.com/lewiscoleblog/blog/raw/master/images/Monte-Carlo/copula.png
___
This is the second blog post in a series - you can find the previous blog post [here](https://lewiscoleblog.com/monte-carlo-methods)
___
In this blog post we shall continue our exploration of Monte Carlo methods. To briefly recap in the previous blog post we looked at the general principle underlying Monte-Carlo methods, we looked at methods used by pseudo-random number generators to feed our models and we looked at a variety of methods to convert these uniform variates to general univariate distributions.
## Multi-Variate Relations
We start by looking at the problem of sampling from general multi-variate distributions. In many instances simply sampling univariate distributions will be insufficient. For example if we are creating a model that looks of mortgage repayment defaults we do not want to model the default rate by one univariate distribution and the interest rate by another univariate distribution using the techniques described so far. In an ideal world we would know the exact relationship we are trying to model, in physics and the hard sciences the exact mechanism underlying the joint behaviour may be modelled exactly. However in many cases in finance (and the social sciences) this is very hard to elicit (if not impossible) so we want to create "correlated" samples that capture (at least qualitatively) some of the joint behaviour.
### Measures of Correlation
We start by looking at some measures of dependance between variables. This helps us evaluate whether a model is working as we would expect or not. A lot of times joint-dependence is difficult to elicit from the data so often with this sort of model there is a lot of expert judgements, sensitivity testing, back testing and so on in order to calibrate a model. This is out of the scope of what I'm looking to cover in this blog post series but is worth keeping in mind for this section on multi-variate methods.
By far the most prevelant measure of dependance is the Pearson correlation coefficient. This is the first (and in many cases only) measure of depedence we find in textbooks. We can express it algebraically as:
$$ \rho_{(X,Y)} = \frac{\mathbb{E}[(X - \mathbb{E}(X))(Y - \mathbb{E}(Y))]}{\sqrt{\mathbb{E}[(X - \mathbb{E}(X))^2]\mathbb{E}[(Y - \mathbb{E}(Y))^2]}} = \frac{Cov(X,Y)}{SD(X)SD(Y)}$$
That is the ratio of the covariance between two variables divide by the product of their standard deviations. This measure has a number of useful properties - it gives a metric between $[-1,1]$ (via the Cauchy-Schwarz inequality) which varies from "completely anti-dependant" to "completely-dependant". It is also mathematically very tractable, it is very easy to calculate and it often crops up when performing an analytic investigation. However it's use in "the real world" is fairly limited, it is perhaps the most abused of all statistics.
One issue with this measure in practice is that it requires defined first and second moments for the definition to work. We can calculate this statistic on samples from any distribution, however if we are in the realms of fat-tailed distributions (such as a power-law) this sample estimate will be meaningless. In finance and real-world risk applications this is a big concern, it means that many of the early risk management models still being used that do not allow for the possiblity of fat-tails are at best not-useful and at worst highly dangerous for breeding false confidence.
Another issue with the measure is it is highly unintuitive. Given the way it is defined the difference between $\rho=0.05$ and $\rho = 0.1$ is negligible, yet the difference between $\rho = 0.94$ and $\rho = 0.99$ is hugely significant. In this authors experience even very "technical" practitioners fail to remember this and treat a "5% point increase" as having a consistent impact. This issue is compounded further when dealing with subject matter experts who do not necessarily have mathematical/probability training but have an "intuitive" grasp of "correlation"!
The biggest and most limiting factor for the Pearson coefficient however is that it only considers linear relationships between variables - any relation that is not linear will effectively be approximated linearly. In probability distribution terms this is equivalent to a joint-normal distribution. That is to say: **Pearson correlation only exhibits good properties in relation to joint-normal behaviour!**
Another option to use is the Spearmann correlation coefficient. This is strongly related to the Pearson correlation coefficient but with one important difference: instead of using the values themselves we work with the percentiles instead. For example suppose we have a standard normal distribution and we sample the point: $x = 1.64...$ we know that this is the 95th percentile of the standard normal CDF so we would use the value: $y=0.95$ in the definition of Pearson correlation above. This has the benefit that it doesn't matter what distributions we use we will still end up with a reasonable metric of dependance. In the case of joint-normality we will have that the Pearson and Spearmann coefficients are equal - this shows that Spearmann is in some sense a generalization of the Pearson.
However before we get too excited the issues around it being an unintuitive metric still stands. It also has the added complication that if we observe data "in the wild" we don't know what the percentile of our observation is! For example suppose we're looking at relationships between height and weight in a population: if we find a person of height: $1.75m$ what percentile does this correspond to? We will likely have to estimate this given our observations which adds a layer of approximation. In the case of heavy-tail distributions this is particularly problematic since everything may appear to be "normal" for many many observations but suddenly 1 extreme value will totally change our perception of the underlying distribution.
Another issue is that while the Spearmann doesn't rely only on linear relations - it does require at least monotone relations. Anything more complicated that that and the relationship will not be captured. Let's look at a few simple examples to highlight the differences between these metrics:
```
#hide
import warnings
warnings.filterwarnings('ignore')
# Examples of Pearson and Spearman coefficients
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm, rankdata, pearsonr
import seaborn as sns
%matplotlib inline
U = np.random.random(1000)
X = norm.ppf(U)
Y_lin = 4*X + 5
Y_exp = np.exp(X)
Y_con = X**2
# Create function to return estimated percentile
def pctile(x):
return rankdata(x) / x.shape[0]
# Calculate Pearson coefficients
pea_lin = pearsonr(X, Y_lin)[0]
pea_exp = pearsonr(X, Y_exp)[0]
pea_con = pearsonr(X, Y_con)[0]
# Calculate Spearman coefficients
X_pct = pctile(X)
Y_lin_pct = pctile(Y_lin)
Y_exp_pct = pctile(Y_exp)
Y_con_pct = pctile(Y_con)
spe_lin = pearsonr(X_pct, Y_lin_pct)[0]
spe_exp = pearsonr(X_pct, Y_exp_pct)[0]
spe_con = pearsonr(X_pct, Y_con_pct)[0]
# Create Plots
fig, ax = plt.subplots(1, 3, figsize=(15, 5))
sns.scatterplot(X, Y_lin, ax=ax[0])
ax[0].set_ylabel("Y=4X+5")
ax[0].set_xlabel("X \n Pearson: %f \n Spearman: %f" %(pea_lin, spe_lin))
sns.scatterplot(X, Y_exp, ax=ax[1])
ax[1].set_ylabel("Y=exp(X)")
ax[1].set_xlabel("X \n Pearson: %f \n Spearman: %f" %(pea_exp, spe_exp))
sns.scatterplot(X, Y_con, ax=ax[2])
ax[2].set_ylabel("Y=X^2")
ax[2].set_xlabel("X \n Pearson: %f \n Spearman: %f" %(pea_con, spe_con))
fig.suptitle("Comparison of Pearson and Spearman Correlation", x=0.5, y=0.95, size='xx-large')
plt.show()
```
We can see here that for a linear translation both metrics produce the same result. In the case of a non-linear monotone (e.g. exponential) relation the Spearman correctly identifies full dependancy. However neither metric is capable of capturing the dependency in the last non-monotone example - in fact both metrics suggest independence! From a basic visual inspection we would not describe these two variables as being independent. This shows we need to be careful using correlation metrics such as these. This is even before the introduction of noise!
Another somewhat popular choice of dependancy metric is the Kendall-Tau metric. This is a little different to the preceeding options. Essentially to calculate the Kendall-Tau we look at the joint pairs of samples, if both are concordant we add $1$ to a counter otherwise add $-1$ and move onto the next pair. We then take the average value of this (i.e. divide by the total number of pairs of joint samples). We can equally denote this by the formula:
$$ \tau ={\frac {2}{n(n-1)}}\sum _{i<j}\operatorname{sgn}(x_{i}-x_{j})\operatorname{sgn}(y_{i}-y_{j})$$
Where $\operatorname{sgn}(.)$ is the sign operator. $x_i$ and $y_i$ represent the rank or percentile of each sample within the set of all $x$ or $y$ samples. The benefit of the Kendall-tau is it is slightly more intuitive than the Spearman however it still suffers from some of the same issues (e.g. it will fail the $X^2$ example above).
We can show that Pearson, Spearman and Kendall are both particular cases of the generalized correlation coefficient:
$$\Gamma ={\frac {\sum _{{i,j=1}}^{n}a_{{ij}}b_{{ij}}}{{\sqrt {\sum _{{i,j=1}}^{n}a_{{ij}}^{2}\sum _{{i,j=1}}^{n}b_{{ij}}^{2}}}}}$$
Where $a_{ij}$ is the x-score and $b_{ij}$ the y-score for pairs of samples $(x_i, y_i)$ and $(x_j, y_j)$.
For example for the Kendall Tau we set: $a_{ij} = \operatorname{sgn}(x_i - x_j)$ and $b_{ij} = \operatorname{sgn}(y_i - y_j)$. For Spearman we set: $a_{ij} = (x_i - x_j)$ and $b_{ij} =(y_i - y_j)$. Where again we are working with ranks (or percentiles) rather than sampled values themselves.
Can we improve on these metrics from a practical standpoint? The answer is: yes! Unfortunately it is difficult to do and there is as much art as there is science to implementing it. We instead consider the mutual-information:
$$I(X, Y) = \int \int p_{X,Y}(x,y) \operatorname{log}\left( \frac{p_{X,Y}(x,y)}{p_X(x)p_Y(y)} \right) dx dy $$
Or similar for discrete distributions. Here $p_{X,Y}(.)$ represents the joint pdf of $(X,Y)$ and $p_X$, $p_Y$ represent the marginal distributions of $X$ and $Y$ respectively. This has the interpretation that it represents the information gained in knowing about the joint distribution compared to assuming independance of the marginal distributions. This is essentially matches our intuition of what dependance "is". However it is a bit of a pain to work with since we usually have to make distributional assumptions.
We can use generalizations of mutual information also. For example total correlation:
$$C(X_{1},X_{2},\ldots ,X_{n})=\left[\sum _{{i=1}}^{n}H(X_{i})\right]-H(X_{1},X_{2},\ldots ,X_{n}) $$
Which is the sum of information for each marginal distribution less the joint information. This compares to mutual information that can be expressed:
$$ \operatorname{I} (X;Y)=\mathrm {H} (Y)-\mathrm {H} (Y|X) $$
Another possible generalization to use is that of dual-total correlation which we can express as:
$$D(X_{1},\ldots ,X_{n})=H\left(X_{1},\ldots ,X_{n}\right)-\sum _{i=1}^{n}H\left(X_{i}\mid X_{1},\ldots ,X_{i-1},X_{i+1},\ldots ,X_{n}\right) $$
In the above we use the functional $H(.)$ to represent information:
$$H=-\sum _{i}p_{i}\log _{2}(p_{i}) $$
Or similar for continuous variables.
Despite the complications in working with these definitions these functions do pass the "complicated" examples such as $Y=X^2$ above and are a much better match for our intuition around dependence. I have long pushed for these metrics to be used more widely.
### Other Measurements of Dependence
We now appreciate that there are many different ways in which random variables can relate to each other (linear, monotone, independent, and so on.) So far the metrics and measures presented are "overall" measures of dependence, they give us one number for how the variables relate to each other. However in many cases we know that relations are not defined by just one number - typically we may find stronger relations in the tails (in extreme cases) than in "everyday" events. In the extreme we may have independence most of the time and yet near complete depednence when things "go bad". For an example of this consider we are looking at property damage and one factor we consider is wind-speed. At every day levels of wind there is unlikely to be much of a relationship between wind-speed and property damage - any damage that occurs is likely to be due to some other cause (vandalism, derelict/collapsing buildings, and so on). However as the wind-speed increases at a certain point there will be some property damage caused by the wind. At more extreme levels (say at the hurricane level) there will be almost complete dependence between wind level and property damage level. It is not difficult to think of other examples. In fact you'll likely find that most relations display this character to some degree often as a result of a structural shift in the system (e.g. in the wind example through bits of debris flying through the air!) Of course we would like to model these structural shifts exactly but this is often complicated if not impossible - through Monte-Carlo methods we are able to generate samples with behaviour that is at least qualitatively "correct".
We call this quality "tail dependence". This can affect upside and downside tails or just one tail depending on the phenomena.
Naively we would think that an option would be to use one of the correlation metrics above on censored data (that is filtering out all samples below/above a certain threshold). Unfortunately this does not work, the correlation metric is not additive - meaning you can't discretize the space into smaller units and combine them to give the overall correlation measure. This is another reason correlation metrics don't fit well with our intuition. However the information theoretic quantities can satisfy this condition but as before are a little more difficult to work with. We have some specialist metrics that are easier to use for this purpose.
The simplest metric to look at to study tail-dependence is the "joint exceedance probability". As the name suggests we look at the probability of time that the joint distribution exists beyond a certain level (or below a certain level). Suppose we have joint observations $(X_i, Y_i)_{i=1}^N$ and we have: $\mathbb{P}(X > \tilde{x}) = p$ and $\mathbb{P}(Y > \tilde{y}) = p$ then the joint exceedance probability of $(X,Y)$ at percentile $p$ is:
$$JEP_{(X,Y)}(p) = \frac{\sum_i\mathbb{1}_{(X_i > \tilde{x})}\mathbb{1}_{(Y_i > \tilde{y}})}{N} $$
Where: $\mathbb{1}_{(.)}$ is the indicator variable/Heaviside function. By construction we have that for $X$ and $Y$ fully rank-dependent then $JEP_{(X,Y)}(p) = p$ if $X$ and $Y$ fully rank-independent then $JEP_{(X,Y)}(p) = p^2$. It often helps to standardise the metric to vary between $[0,1]$ through the transform (we wouldn't use this metric if we are in the case of negative-dependence so can limit ourselves to indpendence as the lower-bound):
$$JEP_{(X,Y)}(p) = \frac{\sum_i\mathbb{1}_{(X_i > \tilde{x})}\mathbb{1}_{(Y_i > \tilde{y}})}{pN} - p$$
As with other rank-methods this metric is only useful with monotone relations, it cannot cope with non-monotone relations such as the $Y=X^2$ example.
We can also view tail dependence through the prism of probability theory. We can define the upper tail dependence ($\lambda _{u}$) between variables $X_1$ and $X_2$ with CDFs $F_1(.)$ and $F_2(.)$ respectively as:
$$\lambda _{u}=\lim _{q\rightarrow 1}\mathbb{P} (X_{2}>F_{2}^{-1}(q)\mid X_{1}>F_{1}^{-1}(q))$$
The lower tail dependence ($\lambda _{l}$) can be defined similarly through:
$$\lambda _{l}=\lim _{q\rightarrow 0}\mathbb{P} (X_{2}\leq F_{2}^{-1}(q)\mid X_{1} \leq F_{1}^{-1}(q))$$
The JEP metric above is in some sense a point estimate of these quantities, if we had infinite data and could make $p$ arbitrarily small in $JEP_{(X,Y)}(p)$ we would tend to $\lambda _{u}$. We will revisit the tail-dependence metric once we have built up some of the theoretical background.
## Multi-Variate Generation
We now move onto the issue of generating multi-variate samples for use within our Monte-Carlo models. It is not always to encode exact mechanisms for how variables relate (e.g. if we randomly sample from a distribution to give a country's GDP how do we write down a function to turn this into samples of the country's imports/exports? It is unlikely we'll be able to do this with any accuracy, and if we could create functions like this we would better spend our time speculating on the markets than developing Monte-Carlo models!) So we have to rely on broad-brush methods that are at least qualitatively correct.
### Multi-Variate Normal
We start with perhaps the simplest and most common joint-distribution (rightly or wrongly) the multi-variate normal. As the name suggests this is a multi-variate distribution with normal distributions as marginals. The joint behaviour is specified by a covariance matrix ($\mathbf{\Sigma}$) representing pairwise covariances between the marginal distributions. We can then specify the pdf as:
$$f_{\mathbf {X} }(x_{1},\ldots ,x_{k})={\frac {\exp \left(-{\frac {1}{2}}({\mathbf {x} }-{\boldsymbol {\mu }})^{\mathrm {T} }{\boldsymbol {\Sigma }}^{-1}({\mathbf {x} }-{\boldsymbol {\mu }})\right)}{\sqrt {(2\pi )^{k}|{\boldsymbol {\Sigma }}|}}}$$
Which is essentially just the pdf of a univariate normal distribution just with vectors replacing the single argument. To avoid degeneracy we require that $\mathbf{\Sigma}$ is a positive definite matrix. That is for any vector $\mathbf{x} \in \mathbb{R}^N_{/0}$ we have: $\mathbf{x}^T \mathbf{\Sigma} \mathbf{x} > 0$.
How can we sample from this joint distribution? One of the most common was is through the use of a matrix $\mathbf{A}$ such that: $\mathbf{A} \mathbf{A}^T = \mathbf{\Sigma}$. If we have a vector of independent standard normal variates: $\mathbf{Z} = (Z_1, Z_2, ... , Z_N)$ then the vector: $\mathbf{X} = \mathbf{\mu} + \mathbf{A}\mathbf{Z}$ follows a multivariate normal distribution with means $\mathbf{\mu}$ and covariances $\mathbf{\Sigma}$. The Cholesky-decomposition is typically used to find a matrix $\mathbf{A}$ of the correct form, as a result I have heard it called the "Cholesky-method". We will not cover the Cholesky decomposition here since it is a little out of scope, you can read more about it [here](https://en.wikipedia.org/wiki/Cholesky_decomposition) if you would like.
Let's look at an example of generating a 4d multivariate normal using this method:
```
# Simulating from a multivariate normal
# Using Cholesky Decomposition
import numpy as np
from scipy.stats import norm
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
%matplotlib inline
# Fix number of variables and number simulations
N = 4
sims = 1000
# Fix means of normal variates
mu = np.ones(N)*0.5
# Initialize covariance matrix
cov = np.zeros((N, N))
# Create a covariance matrix
# Randomly sampled
for i in range(N):
for j in range(N):
if i==j:
cov[i, j] = 1
else:
cov[i, j] = 0.5
cov[j, i] = cov[i, j]
# Calculate cholesky decomposition
A = np.linalg.cholesky(cov)
# Sample independent normal variates
Z = norm.rvs(0,1, size=(sims, N))
# Convert to correlated normal variables
X = Z @ A + mu
# Convert X to dataframe for plotting
dfx = pd.DataFrame(X, columns=["X1", "X2", "X3", "X4"])
# Create variable Plots
def hide_current_axis(*args, **kwds):
plt.gca().set_visible(False)
pp = sns.pairplot(dfx, diag_kind="kde")
pp.map_upper(hide_current_axis)
plt.show()
```
Here we can see we have geerated correlated normal variates.
We see here that this is a quick and efficient way of generating correlated normal variates. However it is not without its problems. Most notably is specifying a covariance/correlation matrix quickly becomes a pain as we increase the number of variates. For example if we had $100$ variables we would need to specify: $4950$ coefficients (all the lower off diagonals). If we have to worry about the matrix being positive definite this quickly becomes a pain and we can even begin to run into memory headaches when matrix multiplying with huge matrices.
An alternative highly pragmatric approach is to use a "driver" approach. Here we keep a number of standard normal variables that "drive" our target variables - each target variable has an associated weight to each of the driver variables and a "residual" component for its idiosyncratic stochasticity. In large scale models this can drastically reduce the number of variables to calibrate. Of course we lose some of the explanatory power in the model, but in most cases we can still capture the behaviour we would like. This procedure in some sense "naturally" meets the positive definite criteria and we do not need to think about it. Let's look at this in action:
```
# Simulating from a multivariate normal
# Using a driver method
import numpy as np
from scipy.stats import norm
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# Fix number of variables and number simulations
N = 4
sims = 1000
# Target means and standard deviations
mu = np.ones(N)*0.5
sig = np.ones(N)*0.5
# Fix number of driver variables
M = 2
# Specify driver weight matrix
w_mat = np.zeros((M, N))
for i in range(M):
for j in range(N):
w_mat[i, j] = 0.25
# Residual weights
w_res = 1 - w_mat.sum(axis=0)
# Simulate driver variables
drv = norm.rvs(0,1, size=(sims, M))
# Simulate residual variables
res = norm.rvs(0,1, size=(sims, N))
# Calculate correlated variables
X = drv @ w_mat + w_res * res
# Standardise variables
X = (X - X.mean(axis=0)) / (X.std(axis=0))
# Apply transforms
X = X*sig + mu
# Convert X to dataframe for plotting
dfx = pd.DataFrame(X, columns=["X1", "X2", "X3", "X4"])
# Create variable Plots
def hide_current_axis(*args, **kwds):
plt.gca().set_visible(False)
pp = sns.pairplot(dfx, diag_kind="kde")
pp.map_upper(hide_current_axis)
plt.show()
```
We can see that this "driver" method can significanlty reduce the number of parameters needed for our models. They can also be used to help "interpret" the model - for example we could interpret one of our "driver" variables as being a country's GDP we could then use this to study what happens to our model when GDP falls (e.g. is below the 25th percentile). The nature of the drivers is such that we do not need to know how to model them exactly, this is useful for modelling things such as "consumer sentiment" where a sophisticated model does not necessarily exist. This is part of the "art" side of Monte-Carlo modelling rather than a hard science.
The tail dependence parameters from the multi-variate normal satisfy:
$$\lambda_u = \lambda_l = 0$$
That is there is no tail-dependence. This can be a problem for our modelling.
### Other Multivariate Distributions
With the multivariate normal under our belts we may now want to move onto other joint distributions. Unfortunately things are not that simple, using the methods above we are essentially limited to multivariate distributions that can be easily built off of the multivariate normal distribution. One such popular example is the multivariate student-t distribution. We can sample from this using the transform:
$$ \mathbf{X} = \frac{\mathbf{Z}}{\sqrt{\frac{\chi_{\nu}}{\nu}}}$$
Where: $\mathbf{Z}$ is a multivariate normal and $\chi_{\nu}$ is a univariate chi-square with $\nu$ degrees of freedom. The resulting multivariate student-t has covariance matrix $\mathbf{\Sigma}$ and $\nu$ degrees of freedom. Here the degrees of freedom is a parameter that controls the degree of tail-dependence with lower values representing more tail dependence. In the limit $\nu \to \infty$ we have $\mathbf{X} \to \mathbf{Z}$.
We can add a couple of extra lines to our Gausssian examples above to convert them to a multi-variate student t distribution:
```
# Simulating from a multivariate student t
# Using Cholesky Decomposition and a chi2 transform
import numpy as np
from scipy.stats import norm, chi2
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
%matplotlib inline
# Fix number of variables and number simulations
N = 4
sims = 1000
# Fix means of normal variates
mu = np.ones(N)*0.5
# Initialize covariance matrix
cov = np.zeros((N, N))
# Set degrees of freedom nu
nu = 5
# Create a covariance matrix
# Randomly sampled
for i in range(N):
for j in range(N):
if i==j:
cov[i, j] = 1
else:
cov[i, j] = 0.5
cov[j, i] = cov[i, j]
# Calculate cholesky decomposition
A = np.linalg.cholesky(cov)
# Sample independent normal variates
Z = norm.rvs(0,1, size=(sims, N))
# Convert to correlated normal variables
X = Z @ A + mu
# Standardize the normal variates
Z = (X - X.mean(axis=0)) / X.std(axis=0)
# Sample from Chi2 distribution with nu degrees of freedom
chis = chi2.rvs(nu, size=sims)
# Convert standard normals to student-t variables
T = (Z.T / np.sqrt(chis / nu)).T
# Convert T to dataframe for plotting
dft = pd.DataFrame(T, columns=["T1", "T2", "T3", "T4"])
# Create variable Plots
def hide_current_axis(*args, **kwds):
plt.gca().set_visible(False)
pp = sns.pairplot(dft, diag_kind="kde")
pp.map_upper(hide_current_axis)
plt.show()
```
The tail dependence parameter fro the student-t copula for 2 variates with correlation parameter $\rho$ and $\nu$ degrees of freedom is:
$$\lambda_u = \lambda_l = 2 t_{\nu+1} \left(- \frac{\sqrt{\nu+1}\sqrt{1-\rho}}{\sqrt{1+\rho}} \right) > 0$$
We can see that even if we specify $\rho=0$ there will be some level of tail-dependence between the variates. In fact it is impossible to enforce independence between variables using the student-t copula. This can be an issue in some instances but there are extensions we can make to overcome - for example see my [insurance aggregation model](https://lewiscoleblog.com/insurance-aggregation-model).
Another interesting property of the student-t copula is that it also has "opposing" tail dependency. Namely:
$$\lim _{q\rightarrow 0}\mathbb{P} (X_{2}\leq F_{2}^{-1}(q)\mid X_{1} > F_{1}^{-1}(1-q)) = \lim _{q\rightarrow 0}\mathbb{P} (X_{2}>F_{2}^{-1}(1-q)\mid X_{1} \leq F_{1}^{-1}(q)) = 2 t_{\nu+1} \left(- \frac{\sqrt{\nu+1}\sqrt{1+\rho}}{\sqrt{1-\rho}} \right) > 0$$
This gives rise to the familiar "X" shape of a joint student-t scatter plot.
### The Need for More Sophistication
So far we are limited to modelling joint behaviour with either normal or student-t marginal distributions. This is very limiting, in practice we will want to sample from joint distributions with arbitrary marginal distributions. Typically we will find it easier to fit marginal distributions to data and we will want our models to produce reasonable marginal results. We therefore need a better method of modelling joint behaviour.
## Copula Methods
One way to do this is through a copula method. A copula is nothing more than a multi-variate distribution with uniform $[0,1]$ marginals. This is particularly useful for modelling purposes as it allows us to fully delineate the univariate distributions from the joint behaviour. For example we can select a copula with the joint behaviour we would like and then use (for example) a generalized inverse transform on the marginal distributions to get the results we desire. We can do this simply by adapting the multi-variate Gaussian above, we can use the CDF function on the generated normal variates to create joint-uniform variates (i.e. a copula) and then inverse transform these to give Gamma marginals. This will result in Gamma marginals that display a rank-normal correlation structure:
```
# Simulating from Gamma variates with rank-normal dependency
# Using a copula type method
import numpy as np
from scipy.stats import norm, gamma
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
%matplotlib inline
# Fix number of variables and number simulations
N = 4
sims = 1000
# Fix means of normal variates
mu = np.ones(N)*0.5
# Initialize covariance matrix
cov = np.zeros((N, N))
# Select Gamma parameters a and b
a = 5
b = 3
# Create a covariance matrix
# Randomly sampled
for i in range(N):
for j in range(N):
if i==j:
cov[i, j] = 1
else:
cov[i, j] = 0.5
cov[j, i] = cov[i, j]
# Calculate cholesky decomposition
A = np.linalg.cholesky(cov)
# Sample independent normal variates
Z = norm.rvs(0,1, size=(sims, N))
# Convert to correlated normal variables
X = Z @ A + mu
# Standardize the normal variates
Z = (X - X.mean(axis=0)) / X.std(axis=0)
# Apply the normal CDF to create copula C
C = norm.cdf(Z)
# Use inverse transform to create Gamma variates
G = gamma.ppf(C, a=a, scale=1/b)
# Convert T to dataframe for plotting
dfg = pd.DataFrame(G, columns=["G1", "G2", "G3", "G4"])
# Create variable Plots
def hide_current_axis(*args, **kwds):
plt.gca().set_visible(False)
pp = sns.pairplot(dfg, diag_kind="kde")
pp.map_upper(hide_current_axis)
plt.show()
```
This is really quite neat! It makes our modelling that much easier as we do not need to consider the joint-distribution in its entirety, we can break it down into smaller pieces that are easier to handle. The example above is quite a simple one however, we would like to have more freedom than this as we are still bound by sampling from a joint-distribution.
Let's look at some notation. Suppose we have variates: $(X_1, ... ,X_N)$ where each follows a cdf $F_i(x) = \mathbb{P}(X_i \leq x)$. We then have: $(U_1, ... , U_N) = (F_1(X_1), ..., F_N(X_N)$ with uniform random variates by definition. The copula defined by variables $(X_1, ..., X_N)$ is: $C(U_1, ... , U_N) = \mathbb{P}(X_1 \leq F_1(U_1), ... , X_N \leq F_N(U_N))$. From this definition $C: [0,1]^N \to [0,1]$ is an N-dimensional copula if it is a cumulative distribution function (CDF) on the unit cube with uniform marginals.
Going the other way any function $C: [0,1]^N \to [0,1]$ defines a copula if:
- $C(u_{1},\dots ,u_{i-1},0,u_{i+1},\dots ,u_{N})=0$, the copula is zero if any one of the arguments is zero,
- $C(1,\dots ,1,u,1,\dots ,1)=u$, the copula is equal to u if one argument is u and all others 1,
- C is d-non-decreasing, i.e., for each hyperrectangle $B=\prod _{i=1}^{N}[x_{i},y_{i}]\subseteq [0,1]^{N}$ the C-volume of B is non-negative:
$$\int _{B}\mathrm {d} C(u)=\sum _{\mathbf {z} \in \times _{i=1}^{N}\{x_{i},y_{i}\}}(-1)^{N(\mathbf {z} )}C(\mathbf {z} )\geq 0$$
where the $N(\mathbf {z} )=\#\{k:z_{k}=x_{k}\}$.
We now move onto the fundamental theorem of copulae: Sklar's theorem. The statement of the theorem exists in two parts:
1. Let $H(X_1, ..., X_N)$ represent a joint distribution function with marginal distributions $X_i \sim F_i$. Then there exists a copula $C(.)$ such that: $H(X_1, ..., X_N) = C(F_1(X_1), ... , F_N(X_N))$. Moreover if $F_i$ are continuous then the copula $C(.)$ is unique
2. Given a copula function $C(.)$ and univariate distribution functions $F_i$ then $C(F_1(X_1), ... , F_N(X_N))$ defines a joint distribution with marginal distributions $F_i$
Sklar's theorem shows there is a bijection relation between the space of joint distributions and the space of copulae (for continuous variables). This shows that this is indeed a powerful method to use in our modelling.
There are a couple of things to note here: the copula is itself a rank-order method of applying dependency. It is invariant under monotonic transforms (due to the use of the generalized inverse method). We can also derive the Fréchet–Hoeffding bounds:
$$ \operatorname{max}\left\{ 1 - N + \sum_{i=1}^{N} u_i, 0 \right\} \leq C(u_1, ..., u_N) \leq \operatorname{min} \{u_1, ... , u_N \}$$
We can see this quite easily by noting that for the lower-bound:
\begin{align}
C(u_1, ... , u_N) &= \mathbb{P} \left( \bigcap_{i=1}^N \{ U_i \leq u_i \} \right) \\
&= 1 - \mathbb{P} \left( \bigcup_{i=1}^N \{ U_i \geq u_i \} \right) \\
&\geq 1 - \sum_{i=1}^N \mathbb{P}( U_i \geq u_i ) \\
&= 1 - N + \sum_{i=1}^{N} u_i
\end{align}
For the upper bound we have:
$$ \bigcap_{i=1}^N \{ U_i \leq u_i \} \subseteq \{ U_i \leq u_i \} \qquad \implies \qquad \mathbb{P} \left( \bigcap_{i=1}^N \{ U_i \leq u_i \} \right) \leq \mathbb{P} \left( U_i \leq u_i \right) = u_i $$
For all possible indices $i$. We have that the upper-bound function $\operatorname{min} \{u_1, ... , u_N \}$ itself defines a copula corresponding to complete (rank) dependence. It is often called the "co-monotonic copula". The lower bound exists as a copula only in the case $N=2$ whereby it represents complete negative (rank) dependence.
We can also calculate the Spearman ($\rho$) and Kendall-Tau ($\tau$) dependency coefficients using the copula construction:
$$ \rho = 12 \int_0^1 \int_0^1 C(u,v) du dv - 3 $$
And:
$$ \tau = 4 \int_0^1 \int_0^1 C(u,v) dC(u,v) - 1$$
We can view the Gaussian example above in a copula frame via:
$$C^{Gauss}(u_1, ... , u_N) = \Phi_{\mathbf{\mu,\Sigma}}(\Phi^{-1}(u_1), ..., \Phi^{-1}(u_N))$$
Where: $\Phi_{\mathbf{\mu,\Sigma}}$ is the CDF of a multivariate Gaussian with means $\mathbf{\mu}$ and correlation matrix $\mathbf{\Sigma}$ The function: $\Phi^{-1}(.)$ is the inverse CDF of the univariate standard normal distribution. We call this the Gaussian copula or the rank-normal copula.
We can define the student-t copula in a similar way. I have developed a model based around a hierarchical driver structure with a generalization of a student-t copula for the purposes of modelling insurance aggregation. You can read my blog-post dedicated to this specifically [here!](https://lewiscoleblog.com/insurance-aggregation-model)
But what are some other options? The copula methods are very flexible so lets look at some other examples. Starting off with the Archimedean family of copulae. A copula is Archimedian if it admits a representation:
$$ C_{Arch}(u_{1},\dots ,u_{N};\theta )=\psi ^{[-1]}\left(\psi (u_{1};\theta )+\cdots +\psi (u_{N};\theta );\theta \right) $$
Here the function $\psi$ is called the generator function. We also have parameter $\theta$ taking values in some arbitrary parameter space - the role that $\theta$ takes depends on the form of the generator. $\psi ^{-1}$ is the pseudo-inverse of $\psi$:
\begin{equation}
\psi ^{[-1]}(t;\theta ) =
\begin{cases}
\psi ^{-1}(t;\theta )&{\mbox{if }}0\leq t\leq \psi (0;\theta ) \\
0&{\mbox{if }}\psi (0;\theta )\leq t\leq \infty
\end{cases}
\end{equation}
We note that this functional form has some very useful properties. For example we note that we can express the upper and lower tail dependence parameters (defined above). If we place further assumptions on the generator function being a Laplace transform of strictly positive random variables we can write down neat forms of the upper and lower tail depdence metrics:
$$ \lambda _{u} = 2 - 2 \lim_{s \downarrow 0} \frac{\psi'^{[-1]} (2s)}{\psi'^{[-1]} (s)} $$
and:
$$ \lambda _{l} = 2 \lim_{s \to \infty} \frac{\psi'^{[-1]} (2s)}{\psi'^{[-1]} (s)} $$
Similarly we can conveniently calculate the Kendall-tau measure of dependence via:
$$ \tau = 1 + 4 \int_0^1 \frac{\psi(t)}{\psi'(t)} dt $$
This is very convenient for modelling purposes since we can easily control the joint properties in a predictable way. Unforunately the calculation of Spearman correlation coefficients is not as simple and in many cases a closed form analytic solution does not exist.
We can summarise some of the common Archimedean copulae in the table below:
|Copula Name | $\psi(t)$ | $\psi^{[-1]}(t)$ | $\theta$-defined-range | Lower-Tail-$\lambda_l(\theta)$ | Upper-Tail-$\lambda_u(\theta)$ |$\tau$ |
|:--------|:------:|:-------------:|:----------------:|:-----------:|:-----------:|:-------:|
| Frank | $-\log \left({\frac {\exp(-\theta t)-1}{\exp(-\theta )-1}}\right)$ | $-{\frac {1}{\theta }}\,\log(1+\exp(-t)(\exp(-\theta )-1))$ | $\mathbb {R} \backslash \{0\}$ | 0 | 0 | $1 + 4(D_1(\theta)-1)/\theta$ |
| Clayton | $\frac {1}{\theta }(t^{-\theta }-1)$ | $\left(1+\theta t\right)^{-1/\theta }$ | $[-1,\infty) \backslash \{0\}$ | $2^{-1/\theta}$ | 0 | $\frac{\theta}{\theta + 2}$ |
| Gumbel | $\left(-\log(t)\right)^{\theta }$ | $\exp\left(-t^{1/\theta }\right)$ | $[1, \infty)$ | 0 | $2-2^{1/\theta}$ | $1 - \theta^{-1}$ |
Where: $D_k(\alpha) = \frac{k}{\alpha^k} \int_0^{\alpha} \frac{t^k}{exp(t)-1} dt$ is the Debye function.
We can see that the Frank copula applies dependence without any tail-dependence, whereas the Clayton and Gumbel are options for lower or upper-tail dependence modelling. This is in contrast to the student-t we observed before which applies symmetrically to both lower and upper tails.
To finish we will look at the bi-variate Frank, Clayton and Gumbel copulae. We will use a variety of methods to do this. For the Frank copula we will look at the "conditional" copula (that is the probability of one variate conditional on a specific value of the other): we will sample the overall percentile for the copula ($z$) and one of the variates ($u_1$) and then back "out" the remaining variate ($u_2$). The pair $\{ u_1, u_2\}$ is then distributed as the copula:
$$ u_2 = -\frac{1}{\theta} \log \left[1 + z\frac{1-e^{-\theta}}{z(e^{-\theta u}-1) - e^{-\theta u}} \right]$$
We can do the same for the Clayton:
$$ u_2 = \left(1 + u^{-\frac{1}{\theta}} \left( z^{-\frac{1}{1+\theta}}-1 \right) \right)^{-\theta}$$
Unforunately the conditinal copula of the Gumbel is not invertible and so we are unable to follow the same approach. Instead we follow the approach shown by Embrechts where we sample a uniform variate $v$ and then find $0<s<1$ such that: $sln(s) = \theta(s-v)$. We then sample another uniform variate $u$ and the pair: $\{ exp(u^{1/\theta}ln(s)), exp((1-u)^{1/\theta}ln(s))\}$ is a sample from the Gumbel copula with parameter $\theta$.
We can implement this as:
```
# Sampling from the Frank, Clayton and Gumbel archimedean Copulae
# A very basic bi-variate implementation
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.optimize import fsolve
%matplotlib inline
# Set number of simulations and theta parameter
sims = 1000
theta = 2
# Simulate u1 - first component of copula
U1 = np.random.random(sims)
# Simulate z - the joint probaibility
Z = np.random.random(sims)
# Define conversion functions for each copula
def frank(theta, z, u):
return -1/theta*np.log(1+ z*(1-np.exp(-theta))/(z*(np.exp(-theta*u)-1) - (np.exp(-theta*u))))
def clayton(theta, z, u):
return np.power((1+ np.power(u, -1/theta)*(np.power(z, -1/(1+theta))-1)), -theta)
# Define function to find S such that sln(s) = theta(s-u)
def gumfunc(s):
return s*np.log(s) - theta*(s-U1)
# Use fsolve to find roots
S = fsolve(gumfunc, U1)
U1_gumbel = np.exp(np.log(S)*np.power(Z, 1/theta))
U2_gumbel = np.exp(np.log(S)*np.power(1-Z, 1/theta))
U1_frank = U1
U2_frank = frank(theta, Z, U1)
U1_clayton = U1
U2_clayton = clayton(theta, Z, U1)
# Create Plots
fig, ax = plt.subplots(1, 3, figsize=(15, 5))
sns.scatterplot(U1_frank, U2_frank, ax=ax[0])
ax[0].set_title("Frank")
ax[0].set_xlim([0,1])
ax[0].set_ylim([0,1])
sns.scatterplot(U1_clayton, U2_clayton, ax=ax[1])
ax[1].set_title("Clayton")
ax[1].set_xlim([0,1])
ax[1].set_ylim([0,1])
sns.scatterplot(U1_gumbel, U2_gumbel, ax=ax[2])
ax[2].set_title("Gumbel")
ax[2].set_xlim([0,1])
ax[2].set_ylim([0,1])
fig.suptitle(r"Comparison of Archemedean Copulae with $\theta =$%i" %theta, x=0.5, y=1, size='xx-large')
plt.show()
```
As expected we can see little dependence in the tails of the Frank Copula, some dependence in the lower tail (small percentiles) of the Clayton and some dependence in the upper tail (larger percentiles) of the Gumbel copulae.
It was a bit of work to sample from these copulae. Thankfully packages exist to make our lives easier - for example: [copulas](https://pypi.org/project/copulas/). But is important to understand at least the basics behind how some of these packages work before we jump in and use them.
## Conclusion
We have now extended our abilities from being able to sample from univariate distributions in our previous blog post to how to sample from multi-variate distributions. We saw how we could use a Cholesky decomposition to sample from a mulit-variate normal and further how we could implement a driver approach in order to reduce the number of model parameters. We then saw how we could extend this to sample from a multi-variate student-t distribution. We then looked at copula methods in order to separate the marginal and joint behaviour of a system. We ended by looking at the powerful class of Archimedean copulae andthe Frank, Clayton and Gumbel copulae specifically.
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'bird', 'cat', 'deer'}
fg_used = '234'
fg1, fg2, fg3 = 2,3,4
all_classes = {'plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
background_classes = all_classes - foreground_classes
background_classes
# print(type(foreground_classes))
all_classes = {'plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
background_classes = all_classes - foreground_classes
background_classes
# print(type(foreground_classes))
train = trainset.data
label = trainset.targets
train.shape
train = np.reshape(train, (50000,3072))
train.shape
from numpy import linalg as LA
u, s, vh = LA.svd(train, full_matrices= False)
u.shape , s.shape, vh.shape
s
vh
# vh = vh.T
vh
dir = vh[1062:1072,:]
dir
u1 = dir[7,:]
u2 = dir[8,:]
u3 = dir[9,:]
u1
u2
u3
len(label)
cnt=0
for i in range(50000):
if(label[i] == fg1):
# print(train[i])
# print(LA.norm(train[i]))
# print(u1)
train[i] = train[i] + 0.1 * LA.norm(train[i]) * u1
# print(train[i])
cnt+=1
if(label[i] == fg2):
train[i] = train[i] + 0.1 * LA.norm(train[i]) * u2
cnt+=1
if(label[i] == fg3):
train[i] = train[i] + 0.1 * LA.norm(train[i]) * u3
cnt+=1
if(i%10000 == 9999):
print("partly over")
print(cnt)
train.shape, trainset.data.shape
train = np.reshape(train, (50000,32, 32, 3))
train.shape
trainset.data = train
test = testset.data
label = testset.targets
test.shape
test = np.reshape(test, (10000,3072))
test.shape
len(label)
cnt=0
for i in range(10000):
if(label[i] == fg1):
# print(train[i])
# print(LA.norm(train[i]))
# print(u1)
test[i] = test[i] + 0.1 * LA.norm(test[i]) * u1
# print(train[i])
cnt+=1
if(label[i] == fg2):
test[i] = test[i] + 0.1 * LA.norm(test[i]) * u2
cnt+=1
if(label[i] == fg3):
test[i] = test[i] + 0.1 * LA.norm(test[i]) * u3
cnt+=1
if(i%1000 == 999):
print("partly over")
print(cnt)
test.shape, testset.data.shape
test = np.reshape(test, (10000,32, 32, 3))
test.shape
testset.data = test
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
fg,bg
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
img1 = torch.cat((background_data[0],background_data[1],background_data[2]),1)
imshow(img1)
img2 = torch.cat((foreground_data[27],foreground_data[3],foreground_data[43]),1)
imshow(img2)
img3 = torch.cat((img1,img2),2)
imshow(img3)
print(img2.size())
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
list_set_labels = []
for i in range(desired_num):
set_idx = set()
bg_idx = np.random.randint(0,35000,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,15000)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
list_set_labels.append(set_idx)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Module1(nn.Module):
def __init__(self):
super(Module1, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,1)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.module1 = Module1().double()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
self.fc4 = nn.Linear(10,3)
def forward(self,z): #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
x = x.to("cuda")
y = y.to("cuda")
for i in range(9):
x[:,i] = self.module1.forward(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
y = y.contiguous()
y1 = self.pool(F.relu(self.conv1(y)))
y1 = self.pool(F.relu(self.conv2(y1)))
y1 = y1.contiguous()
y1 = y1.reshape(-1, 16 * 5 * 5)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1 , x, y
fore_net = Module2().double()
fore_net = fore_net.to("cuda")
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(fore_net.parameters(), lr=0.01, momentum=0.9)
nos_epochs = 600
for epoch in range(nos_epochs): # loop over the dataset multiple times
running_loss = 0.0
cnt=0
mini_loss = []
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels, fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
# zero the parameter gradients
# optimizer_what.zero_grad()
# optimizer_where.zero_grad()
optimizer.zero_grad()
# avg_images , alphas = where_net(inputs)
# avg_images = avg_images.contiguous()
# outputs = what_net(avg_images)
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
# optimizer_what.step()
# optimizer_where.step()
optimizer.step()
running_loss += loss.item()
mini = 40
if cnt % mini == mini - 1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
mini_loss.append(running_loss / mini)
running_loss = 0.0
cnt=cnt+1
if(np.average(mini_loss) <= 0.05):
break
print('Finished Training')
torch.save(fore_net.state_dict(),"/content/drive/My Drive/Research/mosaic_from_CIFAR_involving_bottop_eigen_vectors/fore_net_epoch"+str(epoch)+"_fg_used"+str(fg_used)+".pt")
```
#Train summary on Train mosaic made from Trainset of 50k CIFAR
```
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
from tabulate import tabulate
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half",argmax_more_than_half)
print("argmax_less_than_half",argmax_less_than_half)
print(count)
print("="*100)
table3 = []
entry = [1,'fg = '+ str(fg),'bg = '+str(bg),30000]
entry.append((100 * focus_true_pred_true / total))
entry.append( (100 * focus_false_pred_true / total))
entry.append( ( 100 * focus_true_pred_false / total))
entry.append( ( 100 * focus_false_pred_false / total))
entry.append( argmax_more_than_half)
train_entry = entry
table3.append(entry)
print(tabulate(table3, headers=['S.No.', 'fg_class','bg_class','data_points','FTPT', 'FFPT', 'FTPF', 'FFPF', 'avg_img > 0.5'] ) )
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
test_set_labels = []
for i in range(10000):
set_idx = set()
bg_idx = np.random.randint(0,35000,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,15000)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_set_labels.append(set_idx)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
```
#Test summary on Test mosaic made from Trainset of 50k CIFAR
```
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half",argmax_more_than_half)
print("argmax_less_than_half",argmax_less_than_half)
print("="*100)
# table4 = []
entry = [2,'fg = '+ str(fg),'bg = '+str(bg),10000]
entry.append((100 * focus_true_pred_true / total))
entry.append( (100 * focus_false_pred_true / total))
entry.append( ( 100 * focus_true_pred_false / total))
entry.append( ( 100 * focus_false_pred_false / total))
entry.append( argmax_more_than_half)
test_entry = entry
table3.append(entry)
print(tabulate(table3, headers=['S.No.', 'fg_class','bg_class','data_points','FTPT', 'FFPT', 'FTPF', 'FFPF', 'avg_img > 0.5'] ) )
dataiter = iter(testloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(1000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
test_set_labels = []
for i in range(10000):
set_idx = set()
bg_idx = np.random.randint(0,7000,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,3000)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_set_labels.append(set_idx)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
unseen_test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
```
# Test summary on Test mosaic made from Testset of 10k CIFAR
```
fg = [fg1,fg2,fg3]
bg = list(set([0,1,2,3,4,5,6,7,8,9])-set(fg))
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in unseen_test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = fore_net(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half",argmax_more_than_half)
print("argmax_less_than_half",argmax_less_than_half)
print("="*100)
# table4 = []
entry = [3,'fg = '+ str(fg),'bg = '+str(bg),10000]
entry.append((100 * focus_true_pred_true / total))
entry.append( (100 * focus_false_pred_true / total))
entry.append( ( 100 * focus_true_pred_false / total))
entry.append( ( 100 * focus_false_pred_false / total))
entry.append( argmax_more_than_half)
test_entry = entry
table3.append(entry)
print(tabulate(table3, headers=['S.No.', 'fg_class','bg_class','data_points','FTPT', 'FFPT', 'FTPF', 'FFPF', 'avg_img > 0.5'] ) )
```
| github_jupyter |
# Image Input & Output with Caer
Caer does not come with in-built io functionality, but extends on OpenCV's API to provide an extensible API
```
import caer
# Load Matplotlib to display our images
import matplotlib.pyplot as plt
# Set a variable to store the current working dir
here = caer.path.cwd()
# Image path
tens_path = caer.path.minijoin(here, 'camera.jpg')
# URL
url = r'https://raw.githubusercontent.com/jasmcaus/caer/master/caer/data/beverages.jpg'
```
# caer.imread()
`caer.imread` reads in an image from a specified filepath or URL.
## 1. From Filepath
```
# Read one of the standard images that Caer ships with
tens = caer.imread(tens_path, rgb=True)
plt.imshow(tens)
plt.axis('off')
plt.show()
```
## 2. From URL
```
# Read one of the standard images that Caer ships with
tens = caer.imread(url, rgb=True)
plt.imshow(tens)
plt.axis('off')
plt.show()
```
## 3. Load an image to a particular size (Hard-Resize)
This is the equivalent of using `caer.imread()` to read the image and then `caer.resize()` to resize the image. We include this here in this example so you know that such a functionality is available from right within `caer.imread()`
```
# Read one of the standard images that Caer ships with
# target_size follows the (width,height) format
tens = caer.imread(tens_path, rgb=True, target_size=(200,180))
# Note that tens.shape might follow the (height, width) format as Caer uses Numpy to process
# images
print(f'Image shape = {tens.shape}')
plt.imshow(tens)
plt.axis('off')
plt.show()
```
## 4. Load an image to a particular size (Keeps Aspect Ratio)
This is the equivalent of using `caer.imread()` to read the image and then `caer.resize()` to resize the image. We include this here in this example so you know that such a functionality is available from right within `caer.imread()`.
Note that some parts of the image will be lost in this example. Caer employs an advanced centre-cropping mechanism to ensure that minimal portions of the image are resized
```
# Read one of the standard images that Caer ships with
# target_size follows the (width,height) format
tens = caer.imread(url, rgb=True, target_size=(200,180), preserv_aspect_ratio=True)
# Note that tens.shape might follow the (height, width) format as Caer uses Numpy to process
# images
print(f'Image shape = {tens.shape}')
plt.imshow(tens)
plt.axis('off')
plt.show()
```
# caer.imsave()
`caer.imsave` saves a Caer-processed image to disk
```
caer.imsave('beverages.jpg', tens)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/aquarius31/Crime-Predictor/blob/master/chicago_crime_visualization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('seaborn')
def heatMap(df):
#Create Correlation df
corr = df.corr()
#Plot figsize
fig, ax = plt.subplots(figsize=(10, 10))
#Generate Color Map, red & blue
colormap = sns.diverging_palette(220, 10, as_cmap=True)
#Generate Heat Map, allow annotations and place floats in map
sns.heatmap(corr, cmap=colormap, annot=True, fmt=".2f")
#Apply xticks
plt.xticks(range(len(corr.columns)), corr.columns);
#Apply yticks
plt.yticks(range(len(corr.columns)), corr.columns)
#show plot
plt.show()
df = pd.read_csv("Chicago_Crimes_2012_to_2017.csv")
##find missing values
##plt.figure(figsize=(8,6))
##sns.heatmap(df.isnull(), cbar = False)
df.dropna(); ## drop rows with missing data
# convert dates to pandas datetime format
df.Date = pd.to_datetime(df.Date, format='%m/%d/%Y %I:%M:%S %p')
df.index = pd.DatetimeIndex(df.Date)
x = df.sample(30000) ##sampling a part of the dataset
heatMap(x) ## correlation between variables
```
Firstly we want to find out what the most common crime is in Chicago. The 'Primary Type' columns is plotted with their correspending value count.
```
x['Primary Type'].value_counts().plot.bar()
plt.title("Crimes")
plt.show()
```
The diagram shows that 'Theft' is the most common crime type. We choose to focus on the four most common types of crime for now.
```
x_theft = x[x['Primary Type'] == "THEFT"]
x_battery = x[x['Primary Type'] == "BATTERY"]
x_cd = x[x['Primary Type'] == "CRIMINAL DAMAGE"]
x_narc = x[(x['Primary Type'] == "NARCOTICS")]
```
Let's plot the subtypes of the different 'Primary Types'.
```
x_theft['Description'].value_counts(normalize=True).plot.bar()
plt.title("Theft Types")
plt.show()
x_battery['Description'].value_counts(normalize=True).plot.bar()
plt.title("Battery Types")
plt.show()
x_cd['Description'].value_counts(normalize=True).plot.bar()
plt.title("Criminal Damage Types")
plt.show()
x_narc['Description'].value_counts(normalize=True).plot.bar()
plt.title("Narcotics Types")
plt.show()
```
We now use a heat map and plot the coordinates for each crime in order to get an overview of where the crimes were generally commited.
```
print("Heat map over cooridantes of crimes")
x.plot.hexbin(x='Longitude', y='Latitude', gridsize=40)
plt.title("Overall crimes")
plt.show()
x_theft.plot.hexbin(x='Longitude', y='Latitude', gridsize=40)
plt.title("Theft")
x_battery.plot.hexbin(x='Longitude', y='Latitude', gridsize=40)
plt.title("Battery")
x_cd.plot.hexbin(x='Longitude', y='Latitude', gridsize=40)
plt.title("Criminal Damage")
plt.show()
x_narc.plot.hexbin(x='Longitude', y='Latitude', gridsize=40)
plt.title("Narcotics")
plt.show()
```
Now let's look at the arrest success rate.
```
x['Arrest'].value_counts(normalize=True).plot.bar()
plt.title("Arrests")
plt.show()
x[x.Arrest == True]['Year'].value_counts().plot.line()
plt.title("Arrests per year")
plt.show()
```
We plot the arrest rate for all types of crimes.
```
x[x.Arrest == True]['Primary Type'].value_counts(normalize=True).plot.bar()
plt.title("Arrests per crime type")
plt.show()
```
We now know which types of crimes are commited and where, now let's find out when.
```
color = (0.2, 0.4, 0.6, 0.6)
days = ['Monday','Tuesday','Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
x.groupby([x.index.dayofweek]).size().plot(kind='barh', color=color)
plt.ylabel('Day of week')
plt.yticks(np.arange(7), days)
plt.xlabel('Number of crimes')
plt.title('Number of crimes by day of week')
plt.show()
x.groupby([x.index.month]).size().plot(kind='barh', color=color)
plt.ylabel('Month')
plt.xlabel('Number of crimes')
plt.title('Number of crimes by month of year')
plt.show()
x.groupby([x.index.hour]).size().plot(kind='barh', color=color)
plt.ylabel('Hour')
plt.xlabel('Number of crimes')
plt.title('Number of crimes by hour of day')
plt.show()
color = (0.2, 0.4, 0.6, 0.6)
x[x.index.hour == 0]['Primary Type'].value_counts(normalize=True).plot.bar(color=color)
plt.title("Crimes commited around 24:00")
plt.show()
x[x.index.hour == 3]['Primary Type'].value_counts(normalize=True).plot.bar(color=color)
plt.title("Crimes commited around 03:00")
plt.show()
x[x.index.hour == 12]['Primary Type'].value_counts(normalize=True).plot.bar(color=color)
plt.title("Crimes commited around 12:00")
plt.show()
x[x.index.hour == 19]['Primary Type'].value_counts(normalize=True).plot.bar(color=color)
plt.title("Crimes commited around 19:00")
plt.show()
color = "mediumvioletred"
x[x['Primary Type'] == 'THEFT'].index.hour.value_counts().plot.bar(color=color)
plt.title("Theft crimes by hour")
plt.xlabel("hour")
plt.ylabel("Amount")
plt.show()
x[x['Primary Type'] == 'BATTERY'].index.hour.value_counts().plot.bar(color=color)
plt.title("Battery crimes by hour")
plt.xlabel("hour")
plt.ylabel("Amount")
plt.show()
x[x['Primary Type'] == 'CRIMINAL DAMAGE'].index.hour.value_counts().plot.bar(color=color)
plt.title("Ciminal Damage crimes by hour")
plt.xlabel("hour")
plt.ylabel("Amount")
plt.show()
x[x['Primary Type'] == 'NARCOTICS'].index.hour.value_counts().plot.bar(color=color)
plt.title("Narcotics crimes by hour")
plt.xlabel("hour")
plt.ylabel("Amount")
plt.show()
```
| github_jupyter |
# Apprentice Challenge
This challenge is a diagnostic of your current python pandas, matplotlib/seaborn, and numpy skills. These diagnostics will help inform your selection into the Machine Learning Guild's Apprentice program.
## Challenge Background: A Magic Eight Ball & Randomness

Do you remember these days? Holding a question in your mind and shaking the magic eight ball for an answer?
From Matel via Amazon: "The original Magic 8 Ball is the novelty toy that lets anyone seek advice about their future! All you have to do is simply 'ask the ball' any yes or no question, then wait for your answer to be revealed. Turn the toy upside-down and look inside the window on the bottom - this is where your secret message appears!"
Answers can be positive (i.e. 'It is certain'), negative (i.e. 'Don’t count on it') or neutral (i.e. 'Ask again later').
In this data analysis programming challenge, you will be programmatically analyzing a Magic Eight Ball's fortune telling. This is the type of exploratory analysis typically performed before building machine learning models.
## Instructions
You need to know your way around `pandas` DataFrames and basic Python programming. You have **1 hour** to complete the challenge. We strongly discourage searching the internet for challenge answers.
General Notes:
* Read the first paragraph above to familiarize yourself with the topic.
* Feel free to poke around with the iPython notebook.
* To run a cell, you press `CRTL+ENTER`
* Complete each of the tasks listed below in the notebook.
* You need to provide your code for challenge in the cells which say "-- YOUR CODE FOR TAKS NUMBER --"
* Make sure to run the very last read-only code cell.
**Please reach out to [Guild Mailbox](mailto:guildprogram@deloitte.com) with any questions.**
# Task 1: Generate Your Fortune!
**Instructions**
Ask our Python-based magic-eight ball 20 questions. You can ask it anything. Save the questions and the fortunes in respective lists which we will use down stream. Use the eightball method `get_fortune` to generate the responses. The sample code below will help you get started.
```
import eightball
fortune = eightball.get_fortune("What is the answer to life, the universe, and everything?")
print("The Eight Balls Says: {}".format(fortune))
```
Once you reach the box/cell containing the Python code, click on it press Ctrl + Enter and notice what happens!
**Sample Questions**
* Is there ice cream on the moon?
* Will I make a lot of money and become a bizzilionaire?
* Am I going to get a pony?
**Expected Output**
* `questions` which is a `list` of 20 strings
* `fortunes` which is a `list` of 20 strings
```
# -- YOUR CODE FOR TASK 1 --
import random
import sys
ans = True
from datetime import datetime
# Start Time
start_time = datetime.now()
print(start_time)
# Import the eightball module
# Store your questions list as a variable "questions"
question=[]
print("Am I going to be a successful person?")
# Generate respones to your questions and store them as a variable "fortunes"
fortunes = random.randint(1,3)
if question == "":
sys.exit()
elif fortunes == 1:
print ("Yes")
elif fortunes == 2:
print ("No")
elif fortunes == 3:
print ("Stop relying on Luck!")
elif fortunes == 4:
print ("May be")
elif fortunes == 5:
print ("God knows")
elif fortunes == 6:
print ("Work hard, you will be successful")
elif fortunes == 7:
print ("I dont know")
elif fortunes == 8:
print ("Hell yea!!")
print(type(questions))
print(type(fortunes))
```
# Task 2: Create a DataFrame!
**Instructions**
Let's analyze your newly minted fortunes. Perhaps we can uncover the magic of the eightball. To start our analysis, put `questions` and `fortunes` in a pandas `DataFrame` called `questions_fortunes`. Your DataFrame should have two columns, one for each of the respective lists. What is the shape of your DataFrame?
**Output**
* `questions_fortunes` which is a `pandas.DataFrame` with two columns called `question` & `fortune`
* Shape of questions_fortunes stored in a variable `shape`
```
# -- YOUR CODE FOR TASK 2 --
import pandas as pd
#Combine two lists into a dataframe with specified column names
questions = [['Will I be successful?'],
['Am I any good?']]
Fortunes=[['Yes'],
['No']]
#questions_fortunes=pd.DataFrame.from_records(questions_fortunes)
#Fortunes=pd.DataFrame.from_records(Fortunes)
# Define the shape of the dataaframe
#shape = questions_fortunes.shape
print(shape)
questions_fortunes = pd.DataFrame(columns = ['question', 'fortunes'])
questions_fortunes['question']=questions
questions_fortunes['fortunes']=Fortunes
#data.drop(['0'],axis=1)
questions_fortunes
print(list(questions_fortunes))
```
# Task 3: Getting Fortunes
**Instructions**
In the data sub-folder of the challenge folder, there is a dataset ("questions-fortunes.txt") with additional questions and magic eightball fortunes. Read that dataset into a pandas DataFrame called `temp` and combine with your `questions_fortunes` DataFrame. Be sure the index doesn't repeat. Call this new DataFrame `questions_fortunes_updt`.
**Output**
* A temp dataframe of additional questions/fortunes from "questions-fortunes.txt" datafile
* An updated pandas DataFrame `questions_fortunes` with additional rows from the "question-fortunes.txt" datafile.
```
# -- YOUR CODE FOR TASK 3--
# Create temp DataFrame from "questions-fortunes.txt"
temp = pd.re
# Combine with existing `questions_fortunes` DataFrame
questions_fortunes_updt = ...
print("Check your answers)
print(questions_fortunes_updt.shape[0])
```
# Task 4: Common Fortunes
**Instructions**
With something close to 1,700 questions and fortunes, we can study the patterns of fortune telling.
***Part 1:***
* Create a variable named `Fortune_Counts` which counts the number of times a fortune is determined for each of the available fortune in `questions_fortunes_updt`. The DataFrame should have two columns: `fortune` and `num_fortune`. The DataFrame should be sorted by `num_fortune` in descending order. Print the entire result.
* We know that fortunes from the magic ball fall into one of three categories: Positive, Negative, and Neutral. Use a dictionary of lookup values to assign one of these categories to each question/fortune. Add this as a column `category` to the `Fortune_Counts` DataFrame.
* You can access the positive, negative, neutral lookup with the `eightball.fortune_category` property. To add a new column in your DataFrame using a dictionary try adapting this technique:
```
raw = [[0, "Pony"],
[0, "Saddle"],
[0, "Lasso"],
[1, "Saddle"]
]
prices = {'Pony': 9.99,
'Saddle': 4.95,
'Lasso': '3.25'
}
df = pd.DataFrame(raw, columns=["orderID", "item"])
df['price'] = df['item'].map(prices)
```
***Part 2:***
Create a barchart of your dataset `Fortune_Counts` from Part 1 and color the bars by their category. Create a `seaborn.barplot` with a bar for each fortune colored by category (positive / negative / neutral). Please use the seaborn plotting library. You can install seaborn using `pip`. You can read about the API for the barplot [here](https://seaborn.pydata.org/generated/seaborn.barplot.html). Make the x-axis num_fortune, the y-axis fortune.
**Output**
*Part 1:* A sorted DataFrame detailing the number of questions per fortune outcome, along with the mapped category
*Part 2:* A `seaborn.barplot` with a bar for each fortune colored by category (positive / negative / neutral).
```
# Task 4 Part 1
# -- YOUR CODE FOR TASK 4.1 --
# Create dataFrame 'Fortune_Counts' with fortune, category, and num_fortune
Fortune_Counts = ...
# Import categories from eightball
# Create column 'category' mapping the category to each question
Fortune_Counts['category'] = ...
print(Fortune_Counts)
print("Check your answers)
# Task 4 Part 2
# -- YOUR CODE FOR TASK 4.2 --
import seaborn as sns
# Create barplot using sns.barplot()
plt = sns.barplot(...)
print("Check your answers)
print("Ignore this cell")
print("Check your answers)
print("Keep up the good work!")
print("Pass this Cell")
```
# Task 5: Question your Questions
Magic Eightballs have twenty set responses. We can safely assume that our python-based magic eight ball is not drawing its response from the great beyond. Understanding the patterns of your questions along with the fortunes provided may help interperet the algorithms behind the eight ball.
**Instructions**
***Part 1:*** How long are your questions? Do a quick character count on each of your questions. Include the new data in `questions_fortunes_updt` as a new column called `question_length`. For this task, we recommend using a pandas method and a base python function in the same line.
What is the average question length?
What is the average question length by fortune?
***Part 2:*** It seems there may be a correlation between the input (question) length and the output (fortune). Use a pivot table to take a look at number of questions associated with the question length (as the index) vs the fortune told (columns). What do you notice? Make sure all rows display. Fill null values with "-" for easier interpretability.
**Output**
*Part 1:*
* A new column in `questions_fortunes_updt` called `question_length`
* A variable called `Avg_Length` of the average length of all questions
* A variable called `Avg_Length_Fortune` of the average length of all questions by Fortune
*Part 2:*
* Pivot Table displaying all rows
```
# Task 5 Part 1
# -- YOUR CODE FOR TASK 5.1 --
# Create new column `question-length`
# Calculate average length
Avg_Length = ...
# Calculate average length by fortune
Avg_Length_Fortune = ...
print("Check your answers)
# Task 5 Part 2
# -- YOUR CODE FOR TASK 5.2 --
# Display all rows
# Create pivot table
print("Check your answers)
print("Do you excel at Pivot Tables?")
print("We're trying to find out.")
```
# Task 6: Telling Analysis
**Instructions**
Remember our assumption that the output is a function of the input. Based on the pivot table above, it appears that each question length is associated with only one fortune, illustrating that the fortune choice is a function of the input question string length. Thus, we should expect the that two different questions of the same length would procedure the same result. The code below tests and proves this hypothesis.
Our job is done then right? We can know what the eightball is going to tell us with any question, so we can tell the future. This is because the algorithm for this magic eight ball uses the length of the question as the seed.
***Part 1:***
Use the list below to test this hypothesis.
```
same_length = ["Can whales dance?",
"Can zebras paint?"
]
```
***Part 2:***
Using what you now know about this magic eight ball, write your own function `get_new_fortune` that takes an input `question` and outputs a fortune `option`. Include at least two arguments in your function: `options` and `question`. Use two function arguments, one still being the options list from the eightball module as the fortune options. Get creative and add a third parameter to your function which alters the fortune told!
Test your function by applying it to a question 'Will I make a lot of money and become a bizzilionaire?' to get a fortune.
***Output***
* `hypothesis_test` which boolean variable
* `get_new_fortune` a function with at least two inputs: `question` and `options`
```
# Task 6 Part 1
# -- YOUR CODE FOR TASK 6.1 --
# Same Length List
same_length = ["Can whales dance?",
"Can zebras paint?"
]
# Code to test if fortunes told are identical
hypothesis_test = ...
print(hypothesis_test)
print("Check your answers)
# Task 6 Part 2
# -- YOUR CODE FOR TASK 6.2 --
# Create your function `get_new_fortune`
# Test your function on the question: "Will I make a lot of money and become a bizzilionaire?"
print("Check your answers)
print("Was our hypothesis true?")
print("Can you tell your own fortune now?")
```
# Wrapping up!
Please save this notebook as "Last Name - First Name.ipynb". Make sure to save this file and submit it via the link you received in your Deloitte email.
Happy coding!
## References
1. [Javascript Magic 8 Ball with Basic DOM Manipulation](https://medium.com/@kellylougheed/javascript-magic-8-ball-with-basic-dom-manipulation-1636b83c3c26)
2. [Mattel Games Magic 8 Ball](https://www.amazon.com/Mattel-Games-Magic-8-Ball/dp/B00001ZWV7) - Where to buy on Amazon.
| github_jupyter |
In this series of posts, we going to talk about the control flow in Python programming. A program’s control flow is the order in which the program’s code executes. In this first post of the series, we'll talk about
the conditional and pass statement. But before we start, let's discuss on the indentation in python and difference between a statement and an expression. Again, this post is written entirely in Jupyter notebook and
exported as a markdown file Jekyll.
### statement vs expression
From this point, I will be using the term expression and statement quite often so let me explain right away
the difference. The general rule of thumb is that If you can print it, or assign it to a variable,
it’s an expression and if you can’t then it’s a statement.
```
2 + 2
3 * 7
1 + 2 + 3 * (8 ** 9) - sqrt(4.0)
min(2, 22)
max(3, 94)
round(81.5)
"foo"
"bar"
"foo" + "bar"
None
True
False
2
3
4.0
```
All the above codes can be printed or assigned to a variable. These are called expressions.
```
if CONDITION:
elif CONDITION:
else:
for VARIABLE in SEQUENCE:
while CONDITION:
try:
except EXCEPTION as e:
class MYCLASS:
def MYFUNCTION():
return SOMETHING
raise SOMETHING
with SOMETHING:
```
None of the above codes can be assigned to a variable. They are syntactic elements that serve a purpose but do not themselves have any intrinsic “value”. In other words, these codes don’t “evaluate” to anything.
Most of these codes we have not seen them yet but I will be referring them as a statement.
### indentation in python
python, unlike other C-family programming languages, does not use delimiter or block of code using the traditional {} but instead uses space and tab to avoid cluttering code and improve readability.
```
for i in range(1,12):
print("i is {}, i**2 is {}, i**3 is {}".format(i,i**2,i**3))
```
Bear with me, I know we have not seen the for loop, we'll go in depth later but consider a loop as a way to repeat the same code multiple time without rewritting the code.
we are using the range function that counts starting from 1 to 12. for each round, the count is increment by one and stored in the variable i. if you are keen, you may have noticed that the count started from 1 but stopped at 11 instead of 12. You may ask why is it like this? well this is because the range is exclusive so 12 will not be considered but instead the count will limit to 11.
Now let's talk about the indentation, we can clearly see that there is a tab ( or 4 space character ) just before the print function this means that it is part of the for loop statement. Each code on the same level of indentation is part of the same block code. In Python, we delimit a block code with spaces or tab and to add more code to this block, we simply press ENTER.
```
for i in range(1,12):
print("i is {}, i**2 is {}, i**3 is {}".format(i,i**2,i**3))
print("this will not be in the part of the loop")
```
In this example, we have added a print function on the last line which is not part of the for loop statement and will only be executed at last after the for loop has finished executing.
```
for i in range(1,12):
print("i is {}, i**2 is {}, i**3 is {}".format(i,i**2,i**3))
```
In example, we are having IndentationError. This error is due to the fact that we have informed Python
that we going to add some code inside the block code of the for loop statement ( Python knows this because of the : after the expression in the for loop ) to find out that the print function
is not part of the for loop block code. To fix this error, we have to add tab or 4 spaces in front of the print
function.
```
for i in range(1,12):
print("i is {}, i**2 is {}, i**3 is {}".format(i,i**2,i**3)) #print 1
print("individual calculation complete") #print 2
```
In this example, the #print 2 is part of the same block of code as #print 1. #print 1 and #print 2 functions will be printed on each time the for loop is counting.
```
for i in range(1,12):
print("i is {}, i**2 is {}, i**3 is {}".format(i,i**2,i**3)) #print 1
print("individual calculation complete") #print 2
print("All the calculation are done") #print 3
```
In this example, #print 1 and #print 2 will be printed each time the for loop is running and #print 3 will be only
printed at last when the loop is over.
NB: Do not mix up spaces and tab, I repeat do not mix them up. The reason for this is that sometimes you can get into nasty bugs(errors) that are very hard to debug(to fix), so my advice is either go once for all with 4 spaces or a tab.
### Conditional Statement
#### if and else statement
Everything we have seen so far has consisted of sequential execution which means that codes are always executed one after the next, in exactly the order specified. But the world is often more complicated than that.
Frequently, a program needs to skip over some lines of code, execute a series of codes repetitively, or choose between alternate sets of codes to execute. That is where control structures come in. A control structure directs the order of execution of the statements in a program (referred to as the program’s control flow).
In the real world, we commonly must evaluate information around us and then choose one course of action or another based on what we observe and we often use a conditional statement in our day-to-day life like in this example: If the weather is nice, then I’ll wash my clothes. (It’s implied that if the weather isn’t nice, then I won’t wash my clothes.)
In a Python program, the if statement is how you perform this sort of decision-making. It allows for conditional
execution of a statement or group of statements based on the value of an expression.
Introduction to the if Statement
We’ll start by looking at the most basic type of if statement. In its simplest form, it looks like this:
if "expression":
"statement"
In the form shown above:
* "expression" is an expression evaluated in Boolean context, as discussed in the section on Logical Operators in the Operators and Expressions in Python tutorial.
* "statement" is a valid Python statement, which must be indented.
If "expression" is true, then "statement" is executed. If "expression" is false, then "statement" is skipped over and not executed.
N.B: the colon (:) following "expression" is required. Some programming languages require "expression" to be enclosed in parenthesis, but Python does not.
Let's recap a little from what we have seen in the last post
```
name = input("What is you name: ")
age = int(input("How old are you {} ".format(name)))
print("Your name is {} and you are {} years old".format(name, age))
```
Nothing here new except the fact that we are using the format() function inside the input function which is very much possible since the format function a string's function.
Another thing we are doing here is to cast the age data type from string to integer directly when getting the value input. If we try to input a string literal, this will result in an error.
now let's do the real thing since age is cast to an integer data type, we can compare it with another integer.
```
if age >= 21:
print("{}, your are a grown up adult now".format(name)) #print 1
else:
print("{}, your are still a young boy".format(name)) #print 2
```
age >= 21 will be evaluated first, and if the expression is True then the block of code indented will be executed ( #print 1 ) else if the expression is False, then the code indented in the else statement will be executed ( #print 2 ).
If we don't specify the else statement, this won't result in an error. Nothing will happen
```
if age >= 21:
print("{}, your are a grown up adult now".format(name))
```
If we change the age to be smaller than 21 and then evaluate age without specifying the else statement, nothing will happen, because we have not specified what python should do in the case when age is smaller than 21.
```
age = 13
if age >= 21:
print("{}, your are a grown up adult now".format(name))
```
let's look at another example
For this program, we'll check if someone has the minimum age to have a driving license.
We first start by getting the name and age using the input( ) function
```
name = input("Please enter your name :")
age = int(input("Please {}, enter your age :".format(name)))
if age >= 16:
print("{} you can have a driving license".format(name)) #print 1
else:
print("Unfortunately {}, you can't have a driving license for now come back in {} years".format(name,16-age)) #print 2
print("Execution of the program has ended...") #print 3
```
In this example the age that we have input is 12, python will check if 12 >= 16 which is False, this means that the block code (#print 1) indented right after the if statement will not be executed at all, so the execution stack (the order each code is executed in a program) will jump up to the else, and internally check if 12 >= 16, this will return True, the bock of code (#print 2)inside the else statement will be executed.
The first placeholder will obviously display the name and the second will display the remaining age in order to obtain the driving license by subtracting 16 (which is the minimum age to obtain the driving license) to the current age. If the age inputted would have been greater than 16, then the first condition would have been evaluated and the block code inside it would have been executed, the else condition with its block code will be ignored.
We have a third print (#print 3) at the end of the if-else statement, this print function will execute and printed after the if-else statement has finish its execution.
#### elif statement
sometimes we might need to evaluate more than two conditions, in this case, we added we use elif statement which stands for "else if".
To demonstrate the usage of elif statement, we are going to create a small guessing number program.
we are going to start by getting the input from the user
```
your_number = int(input("Please guess a number between 1-10: ")) #input 1
if your_number < 5:
new_number = int(input("Please guess a higher number: ")) #input 2
if new_number == 5:
print("Well this round, you got it right congratulation!") #print 1
else:
print("Sorry you missed it again") #print 2
elif your_number > 5:
new_number = int(input("Please guess a lower number: ")) #input 3
if new_number == 5:
print("Well this round, you got it right congratulation!") #print 3
else:
print("Sorry you missed it again") #print 4
else:
print("you got it right on the first trial congratulation!") #print 5
print("Execution of the program has ended...") #print 6
```
The number we are trying to guess is 5 but we'll assume that we don't know the number, we first input a number from the user (#input 1) and store it in the variable your_number.
If your_number is less than 5, this condition is being evaluated as True the block code indented after "if your_number < 5:" will be executed. We will be immediately prompted to guess higher, after inputting and storing the number ( #input 2 ) another if and else statement will be used to evaluate the new_number ( this if-else statement is called a nested if-else statement ). new_number will be evaluated and compared to 5 using == operator (not to confused with assignment operator =), if it is equal to 5 then you won ( #print 1 ) else you have lost ( #print 2 ).
Else if your_number is greater than 5, this condition will be evaluated as True the block of code indented after " elif your_number > 5 " will be executed. We will be prompt to guess lower ( #input 3 ), after inputting new_number another nested if and else statement will be used to evaluate new_number checking if it is equal to 5 or not, if it is then you have won ( #print 3 ) else you have lost ( #print 4 ).
lastly, if none of the two conditions( if and elif statement ) is evaluated to True, this means that we are only left with the condition that your_number is equal to 5, which is the number we are guessing consequently this means that we have won the game.
N.B:
* Make sure that the indentation is correct because the control flow of any program written in Python relies on that.
* we can add as many elif statements as we wish depending on the conditions we have.
* nested conditional statements can get deeper and deeper. the deeper they go, the more complex they become which is not necessarily a good thing because clean code must be readable. I would suggest not to go beyond than 3 nested conditional statement. I have more than 3, consider refactoring your codes.
this example can be written in a much more efficient and concise way because they were a lot of duplicate code.
```
your_number = int(input("Please guess a number between 1-10: "))
if your_number == 5:
print("you got it right on the first trial congratulation!")
else:
if your_number > 5:
new_number = int(input("Please guess lower"))
else: # if your_number < 5
new_number = int(input("Please guess higher"))
if new_number == 5:
print("Well this round, you got it right congratulation!")
else:
print("Sorry you missed it again")
print("Execution of the program has ended...")
```
this a more concise way of writing the same program, we first start by checking if your_number is equal or different from 5 right away if it is equal we have won else we are going to check if your_number is greater or smaller than 5 to direct the user to guess lower or higher. We store the new input value in the new_number variable and straight away we check if it is equal to 5 or different to 5, if it is equal then we have won, else we have missed again. After all, the if and else statement is done executing, the last print statement will be executed.
#### conditional statement with logical operator
##### and
Python gives you the flexibility to use at the same time conditional statement with a logical operator in an expression, let's illustrate this by an example.
```
age = int(input("Please enter your age: "))
if age >= 23 and age <= 85:
print("You are eligible to become president of the republic of Burundi") #print 1
else:
print("You can't be the president of the republic of Burundi") #print 2
```
Here we are using 3 operators ( ">="," <=", "and" ). the operator with the highest precedence ( in this case ">=","<=" ) will be evaluated first then the operators with lower precedence ( "and" operator ) will be executed last. age >= 23 and age <= 85 will be executed first, the values from this comparison ( which are either False or True ) will be compared on their turn using the "and" operator. The expression will return True if True and True is compared, will return False if True and False, False and True or False and False is compared.
Coming back to our example, let's say we have inputted 55 as the value of age, 55 will be compared with 23 like this 55 >= 23 which will return True, then 55 will be compared to 85 as 55 <= 85 which will also return True. Now that we are done with the comparison of operators with high precedence, we are going to compare the two boolean value True and True which will return True, now the blog code indented will be executed ( #print 1 ).
```
if (age >= 23) and (age <= 85):
print("You are eligible to become president of the republic of Burundi")
else:
print("You can't be the president of the republic of Burundi")
```
To be clear and explicit about what is happening, we can rewrite the expression using parenthesis and the result should be exactly the same. Again, the parenthesis is unnecessary, it used to make the expression easier to read.
Another way to write the same example
```
if 23 <= age <= 85:
print("You are eligible to become president of the republic of Burundi")
else:
print("You can't be the president of the republic of Burundi")
```
This is the same example written differently without using the "and" operator. Here we are checking if the age is between the range of 23 included and 85 included. It will either return True or False depending on
value of age.
if we don't want use <= or >= but instead to use < or >, we can write
```
if 22 < age < 86:
print("You are eligible to become president of the republic of Burundi")
else:
print("You can't be the president of the republic of Burundi")
```
By incrementing and decrementing the values of the variable (changed from 23 to 22 and 85 to 86) evaluated against age, we can avoid using >= or <= and still get the same result.
N.B: when python is comparing two operands using "and" operator, it's gonna stop the comparison as soon as one of the operands is False and return False.
##### or
Let's do the same example but this time using "or" operator instead of "and" operator.
```
age = int(input("Please enter your age: "))
if (age < 23) or (age > 85):
print("You can't be the president of the republic of Burundi")
else:
print("You are eligible to become president of the republic of Burundi")
```
If any of two expressions ( age < 23 ) ( age > 85 ) result in True, the whole statement will be True because remember for the "or" logical operator, if any of the operand being evaluated is True then the result will be True, if both are True then the result will be True also. But if both are False only then the result will be False.
N.B: when python is comparing two operands using "or" operator, it's gonna stop the comparison as soon as one of the operands is True and return True.
#### Evaluating empty strings and zero based numbers
In Python a non-empty string or a numerical number else than 0, will be evaluated as True.
```
x = "x"
if x:
print("hehe interestingly this is printing")
none_zero_number = 4
none_zero_float = 4.2
none_empty_string = "Text that will be casted to True"
if none_zero_number:
print("This printed because statement was evaluated to True")
if none_zero_float:
print("This printed because statement was evaluated to True")
if none_empty_string:
print("This printed because statement was evaluated to True")
```
The statement went through and was able to print the text. We can also verify this by casting the string and numerical data type to a boolean data type using bool( ).
```
print(bool(none_zero_number))
print(bool(none_zero_float))
print(bool(none_empty_string))
```
now let's see with empty string and zero numerical
```
zero_number = 0
zero_float = 0.0
empty_string_double_quote = ""
empty_string_single_quote = ''
none_variable = None
empty_list = []
empty_set = ()
empty_dict = {}
print("zero_number: {},zero_float: {},empty_string_double_quote: {},empty_string_single_quote: {},none_variable: {},empty_list: {},empty_set: {},empty_set: {}".format(bool(zero_number),bool(zero_float),bool(empty_string_double_quote),bool(empty_string_single_quote),bool(none_variable),bool(empty_list),bool(empty_set),bool(empty_dict)))
```
Here we can see that casting the numerical variables with value 0 result in False, same as the empty strings. For none_variable we have assigned it to a None keyword value which exactly means that we are assigning the variable to nothing. This will be interpreted as False when evaluating it with conditional. The other examples are an empty list, set and dictionaries since we have not discussed them yet, just think of them as containers and when those containers are empty they are evaluated as False.
let's use these in an example of the conditional statement
```
if zero_float:
print("This will not run")
else:
print("This will definitely run")
```
Such evaluation can be helpful in the case when we are evaluating if an input value has been registered.
```
your_city = input("Please enter the city where you live ")
if your_city:
print("You have entered {}, Thank you for telling us you city".format(your_city)) #print 1
else:
print("You have not entered your city") #print 2
```
In this example, nothing was inputted in the your_city variable, so when evaluating this variable False was returned and #print 2 was executed.
##### not
We can also use "not" operator to get the opposite of boolean value.
```
print(not True)
print(not False)
```
We can use the not operator in a conditional statement and still get the same result, in this case, we only have to change the signs to the opposite signs ( < will be changed to > and > will be changed to < ).
```
age = int(input("Please enter your age "))
if not(age > 23) or not(age < 85):
print("You can't be the president of the republic of Burundi")
else:
print("You are eligible to become president of the republic of Burundi")
```
Age will be compared to 23 and at the same time with 85. With the addition of the not operator, the results of those two comparisons will be inversed to the corresponding opposite boolean value. Depending on which value
returned, the indented block code will be executed.
##### in
In the following example, we will be using the "in" keyword
```
sentence = "On a beach in Hawaii"
the_char = input("Please enter one character: ")
if the_char in sentence:
print("yeap, the character {} was found in the sentene".format(the_char)) #print 1
else:
print("the character {} was not found".format(the_char)) #print 2
```
In this example above, if the character inputted is not found in the sentence, the evaluation will be False and if the character inputted is found in the sentence, True will be returned. The indented block code will be executed accordingly.
##### not in
we can also use "not in" operator, just like this.
```
sentence = "On a beach in Hawaii"
the_char = input("Please enter one character: ")
if the_char not in sentence:
print("the character {} was not found".format(the_char)) #print 1
else:
print("yeap, the character {} was found in the sentene".format(the_char)) #print 2
```
Since we are using not, In this case, we are going to swap the two print statement.
#### Conditional Expressions
conditional expressions are like conditional statements, the only difference is that they return a value that we can evaluate or assign to a variable using only one line of code.
```
raining = False
print("We will go to the {}".format("beach" if not raining else "library"))
```
In the example above, "if not raining" will be evaluated first, since raining is assigned to False the "not" operator will change it to True then the value in front of the if statement("beach" in our case) will be returned and used in the format( ).
Now let's see the inverse.
```
raining = True
print("We will go to the {}".format("beach" if not raining else "library"))
```
Now raining variable is assigned to True, the if statement will be evaluated and will result in False which means that code in the else statement will be returned ("library" in this case).
Another thing we can do is to directly assign the value to a variable
```
age = 2
age_range = "adult" if age >=21 else "youngster"
print("You are {}".format(age_range))
```
In this example, after the if statement is evaluated, either "youngster" or "adult" will be returned and assigned to age_range.
#### pass statement
Occasionally, we might find ourselves in a situation where we need to add a placeholder to our code where we will write future code without raising an error. This is called code stubbing. In python, pass is a keyword for that.
```
raining = True
if raining:
pass
else:
print("Let's go to the beach")
```
running this code nothing will happen, why? because the if the statement was passed but after reaching the indented block code, python read the pass keyword as "just pass this line of code but do not raise an error". In these terms, pass keyword will act as a placeholder.
### Challenge 1
Challenge time!!! For your first challenge, we are going to use the challenge from the previous blog post and add to it some useful informations.
The rules of this challenge are:
* copy the codes from previous challenge
* if the BMI is below 18.5, we should get this message: "you are underweight, you should add more weight"
* if the BMI between 18.6 to 23.0, we should get this message: "you are healthy, maintain that weight"
* if the BMI between 23.1 to 27.5, we should get this message: "you are overweight, you should consider reducing weight"
* if the BMI is above 27.5, we should get this message: "you are obese, you should seriously consider reducing weight"
* you are allowed to use any operators of your choice
### Challenge 2
For your second challenge, we are to going to input the name, age and GPA of a prospective student.
Depending on their age and GPA, the program shall grant a scholarship to the students who are applying for a scholarship.
The rules of this challenge are:
* the program should only granted scholarship to the students with GPA of 3.00 and above aged between 18 years to 25 years old.
* if there is a scholarship granted, print this message: "Congratulation {name}, from the fact that you have a GPA of {gpa} and are {age} years old, you have received a scholarship".
* if there is no scholarship granted, reply politely with the reason why the scholarship was not granted.
* the reasons can be:
- Your age. print this message: "Unfortunately {name}, even though you have the minimum GPA requirement but are {age} years old we are in the incapacity of giving you a scholarship".
- Your GPA. print this message: "Unfortunately {name}, even though you are in the right age range but with a GPA of {gpa} we are in the incapacity of giving you a scholarship".
- Both your GPA and age. print this message: "Unfortunately {name}, with a GPA of {gpa} and the fact that you are {age} years old we are in the incapacity of giving you a scholarship".
* the GPA should be printed as a float number and have 2 decimal digits with 5 as left alignment width, age should be an integer.
* This last rule is optional, use "and" and "not" in your conditional statement.
Now go away and attempt these 2 challenges by your own, ONLY after you have finished or tried them you should come back and compare your solutions with mine.
### Solution to Challenge 1
Since we have already calculated the BMI, let's copy paste the solution from the last post.
```
weight = float(input("Please enter your weight "))
height = float(input("Please enter your height "))
BMI = weight/(height**2)
print("The BMI of a person with {} kg and {} m is {:6.3f}".format(weight,height,BMI))
if BMI <= 18.5:
print("you are underweight, you should add some weight")
elif 18.6 <= BMI <= 23.0:
print("you are healthy, maintain that weight")
elif 23.1 <= BMI <= 27.5:
print("you are overweight, you should reduce some weight")
else:
print("you are obese, you should reduce a lot of weight")
```
Now that we have BMI we use the conditional statement to evaluate it, let's print the appropriate messages.
### Solution to Challenge 2
We'll first start by getting the name and store it in a variable, after that we're going to input the age and cast it to an integer. after it will be the turn of the GPA and we'll cast it to a float.
```
name = input("What is is your name? ")
age = int(input("What is your age? "))
gpa = float(input("Enter your GPA using this format X.XX: "))
if 18 <= age <= 25:
if gpa >= 3.00:
print("Congratulation {}, from the fact that you have a GPA of {:5.2f} and are {} years old, you have received a scholarship".format(name,gpa,age))
else:
print("Unfortunately {}, even though you are in the right age range but with a GPA of {:5.2f} we are in the incapacity of giving you a scholarship".format(name,gpa))
else:
if gpa >= 3.00:
print("Unfortunately {}, even though you have the minimum GPA requirement but are {:5.2f} years old we are in the incapacity of giving you a scholarship".format(name,age))
else:
print("Unfortunately {}, with a GPA of {:5.2f} and the fact that you are {} years old we are in the incapacity of giving you a scholarship".format(name,gpa,age))
```
Now that we have all the inputs, we are going to first check if age is in the range of 18 and 25 as required to be granted a scholarship. If we are in this range, we compare the GPA to the minimum required GPA of 3.00, if it is greater or equal to 3.00 this means that we have secured all the required to get a scholarship. If one or two of the conditions fail, the program will print the reason why the scholarship is not given accordingly.
Optionally, we can rewrite the program using "and" with "not" operators
```
if (age >= 18) and (age <= 25):
if not(gpa <= 3.00):
print("Congratulation {}, from the fact that you have a GPA of {:5.2f} and are {} years old, you have received a scholarship".format(name,gpa,age))
else:
print("Unfortunately {}, even though you are in the right age range but with a GPA of {:5.2f} we are in the incapacity of giving you a scholarship".format(name,gpa))
else:
if not(gpa >= 3.00):
print("Unfortunately {}, even though you have the minimum GPA requirement but are {} years old we are in the incapacity of giving you a scholarship".format(name,age))
else:
print("Unfortunately {}, with a GPA of {:5.2f} and the fact that you are {} years old we are in the incapacity of giving you a scholarship".format(name,gpa,age))
```
This is two of the many ways to approach this challenge, we could have checked first the GPA instead of the age the result would be the same, another way to do it would have been with the usage of the "or" operator.
### Conclusion
In this tutorial post, we have seen the conditional statement with the pass statement, which makes it possible to execute a statement or block of code based on the result of an evaluation. All control flow structures are crucial in order to write complex Python code. In the upcoming post,
we will discuss loops in python.
Thank you for reading this tutorial. If you like this post, please subscribe to stay updated with new posts and if you have a thought or a question, I would love to hear it by commenting below.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import logging
logging.basicConfig(format="%(asctime)s [%(process)d] %(levelname)-8s "
"%(name)s,%(lineno)s\t%(message)s")
logging.getLogger().setLevel('INFO')
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook as tqdm
# Read information to connect to the database and put it in environment variables
import os
with open('../ENVVARS.txt') as f:
for line in f:
parts = line.split('=')
if len(parts) == 2:
os.environ[parts[0]] = parts[1].strip()
%%time
# read error data
import re
def parse_line(line):
# the wordform starts with *
# corrections start with # (there can be multiple) and end with whitspace or ~
# example text start with ~
# 'rules' start with <space>[
# get the wordform
wf_regex = r'^\*(?P<wf>.+?)[\t#]'
m = re.match(wf_regex, line)
wf = m.group('wf')
# Wordforms need to be stripped!
# Whitespace before or after wordforms also leads to duplicate entries in the database.
wf = wf.strip()
# get example text (and remove it)
ex_regex = r'~.+~?'
line = re.sub(ex_regex, '', line)
# remove 'rule'
rule_regex = r'\[EA?XAMPL: .+\]'
line = re.sub(rule_regex, '', line)
# get the corrections
corrections = []
corr_regex = r'#(?P<corr>.+)'
m = re.search(corr_regex, line)
if m:
# Wordforms need to be stripped!
# Whitespace before or after wordforms also leads to duplicate entries in the database.
corrections = [c.strip().replace('\t', '') for c in m.group('corr').split('#') if c != '' and len(c) < 100]
return wf, corrections
corrections = []
# File is in windows-1252 encoding and needs to be converted to utf-8
in_file = '/home/jvdzwaan/Downloads/TWENTE.noxml.2002.sq.clean.norm.tok.sortu.unifrq.LC.noapekrol.allasterisk.12.withcorrections.12186.txt'
num_lines = 0
with open(in_file) as f:
for line in f:
num_lines += 1
#print(repr(line))
wf, corr = parse_line(line)
if wf is not None:
for c in corr:
corrections.append({'wf': wf, 'corr': c})
#else:
# print(line)
data = pd.DataFrame(corrections)
print(num_lines)
parse_line('*variëiten 1#1#variëteiten\n')
parse_line('*toestemmignbesluit#toestemmingenbesluit 1\n')
data.head()
db_name = 'ticclat'
os.environ['dbname'] = db_name
from ticclat.ticclat_schema import Lexicon, Wordform, Anahash, WordformLink, WordformLinkSource
from ticclat.dbutils import get_session, session_scope
Session = get_session(os.environ['user'], os.environ['password'], os.environ['dbname'])
%%time
from ticclat.dbutils import add_lexicon_with_links
with session_scope(Session) as session:
name = 'TWENTE.noxml.2002.sq.clean.norm.tok.sortu.unifrq.LC.noapekrol.allasterisk.12.withcorrections.12186'
vocabulary = False
from_column = 'wf'
from_correct = False
to_column = 'corr'
to_correct = True
add_lexicon_with_links(session, name, vocabulary, data, from_column, to_column, from_correct, to_correct)
```
| github_jupyter |
# "[EN CONSTRUCCIÓN] Cuando utilizar apply con Pandas"
> "La función .apply() para pandas facilita el uso de funciones personalizadas a DataFrames pero no es particularmente eficiente y muchas veces es mejor utilizar funciones nativas de pandas o numpy - ¡solo es cuestión de encontrarlas!"
- toc: true
- branch: master
- badges: true
- comments: true
- categories: [python, pandas]
- image: images/posts_imgs/en-construccion.jpg
- hide: false
- search_exclude: false
# Prólogo
Recientemente me uní al servidor de Discord de **Python en Español** al igual que otras mil personas, mas o menos. Es una comunidad activa especialmente en los canales de `#ayuda` donde mucha gente comparte problemas que tienen con su código y varias personas les ofrecen soluciones. Entre estas soluciones existe mucha variación ya que python existe como un lenguage de programación muy general y sirve para casi todo entonces todxs en el servidor tenemos experiencias distintas. Yo, por ejemplo, vengo de el campo de Economía y Relaciones Internacionales (que estudié en la universidad) - ciencias sociales. No empecé a programar _profesionalmente_ hasta después de graduarme. Escribo python para análisis de datos, antes de esto escribía Stata. Conozco suficiente de R para defenderme y JavaScript para presumir una que otra visualización de d3 en Observable. Esto significa que cuando escribo python tengo ciertos sesgos y abordo problemas de cierta manera. Esta manera, parece ser, es un poco distinta a la manera que otras programadorxs resuelven problemas.
# Un poco de historia
El creador de la biblioteca se llama Wes McKinney comenzó a trabajar en ella el 6 de abril del 2008 porque quería una herramienta flexible y rápida para el análisis de datos en su trabajo como analista en AQR (una empresa que maneja capital para inversionistas). El nombre `pandas` viene del termino de econometría ***Pa***nel ***Da***ta. Yo creo que es importante tener en mente esta historia cuando trabajamos con `pandas` ya que muchas veces `pandas` no _funciona_ como otras bibliotecas de python creadas por y para desarrolladorxs. He visto que a muchas personas les cuesta trabajo "pensar" en `pandas` si vienen con experiencia exclusivamente de programadorxs.
Si les interesa, esta entrevista para el podcast Python.__init__ esta buena :eyes: <iframe title="Podlove Web Player: The Python Podcast.__init__ - Wes McKinney's Career In Python For Data Analysis" width="320" height="400" src="https://cdn.podlove.org/web-player/share.html?episode=https%3A%2F%2Fwww.pythonpodcast.com%2Fwp%2F%3Fpodlove_player4%3D559" frameborder="0" scrolling="no" tabindex="0"></iframe>
> twitter: https://twitter.com/tacosdedatos/status/1377631461495304194
# pandas.DataFrame.apply()
- https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.apply.html
Los DataFrames de pandas tienen el método `.apply()` el cual te permite _aplicar una función a lo largo de un eje del DataFrame._ Es decir, tienes una función que hace algo, le quieres pasar cada una de las filas o columnas de tu DataFrame para que les haga ese "algo" y te las regrese "transformadas."
`.apply()` puedes aplicarlo a cada celda de una columna (`axis = 0`) o a cada celda de una fila (`axis = 1`). También recibe otros argumentos como `raw` (que puedes cambiar a `True` si quieres que el _objeto_ al que le apliques tu función sea un `ndarray` de `NumPy` en lugar de una `Serie` de `pandas`), `result_type` (que cambia el resultado de tu función expandiendo listas a columnas, por ejemplo), y `args` (los argumentos _extras_ que le quieres pasar a tu función que vas a aplicar a las celdas de tu DataFrame), ademas de los benditos `**kwds` que son _keyword arguments_ u otros argumentos que le pasarás a tu función.
Te menciono este método del DataFrame ya que es la función `.apply()` _más extensiva_ pero usualmente la función `.apply()` que vemos es la de las `Series` de `pandas`!
# pandas.Series.apply()
- https://pandas.pydata.org/docs/reference/api/pandas.Series.apply.html
La cual es muy parecida pero recibe menos argumentos y su documentación oficial dice "invoca una función en los valores de una `Series`" lo cual es un _tantito_ distinto a la `.apply()` de los DataFrames.
La `.apply()` de las series recibe un valor de una `Series` de pandas, le pasa ese valor a una función y te regresa ese valor "transformado." Recibe los mismos argumentos que la de DataFrames excepto por `raw` ya que le estas pasando la misma `Series`.
## Un ejemplo _extremadamente_ básico
Digamos que tienes un DataFrame como el siguiente y quieres transformar una de las columnas:
```
#collapse
import pandas as pd
ejemplo_1_datos = pd.read_csv("https://raw.githubusercontent.com/chekos/datasets/master/data/pib.csv")
ejemplo_1_datos.head()
```
Imagina que eres nueva en esto de la programación y no conoces mucho las bibliotecas de `pandas` y `NumPy` y decides crear tu propia función para dividir el PIB (producto interno bruto) en un millón (para cambiar la unidad).
```
def entre_un_millon(numero):
"""Recibe un número y lo divide por un millón."""
return numero / 1_000_000
```
:bulb: Nota: A partir de python 3.6 puedes separar números grandes con `_` para que sean más facil de leer.
Ya creada tu función puedes aplicarla a una columna de la siguiente manera
```
ejemplo_1_datos['PIB'].apply(entre_un_millon)
```
Esencialmente esto es lo que hace `.apply()`. Y puede que estes pensando _ok.. ¿cuál es el problema?_ En realidad, `.apply()` no es el problema que he notado en el mundo de python. El problema no es problema, dijo Arjona :joy:.
El método `.apply()` es útil y te sirve para _extender_ las funcionalidades de `pandas` y `NumPy`. El énfasis esta en _extender_.
Este ejemplo es probablemente poco común y la es poco probable que verías este tipo de código en "la vida real." Claro que si eres alguien que apenas esta comenzando y/o estás practicando crear funciones, o algo así, esto es completamente válido.
Eventualmente, si trabajas con `pandas` o `NumPy`, aprenderás que las `Series` son vectores y puedes aplicarles funciones matemáticas de manera mucho más eficiente de otras maneras. Por ejemplo,
```
ejemplo_1_datos['PIB'] / 1_000_000
```
`pandas` (gracias a que utiliza `ndarrays` de `NumPy` para sus `Series`) simplemente entiende que si divides una columna por un valor lo que quieres es dividir cada uno de los valores de esa columna por _ese_ valor.
Aquí es donde se pone interesante la cosa. Si tienes 8 columnas, crear tu propia función que _puede_ que ya exista en `pandas` puede ahorrarte el tiempo de buscarla en la documentación y obtengas los mismos resultados en una cantidad menor de tiempo. Pero, técnicamente, `.apply()` toma más tiempo que las funciones nativas de `NumPy` y `pandas` ya que estas están optimizadas para trabajar con vectores (o `ndarrays`). Veamos,
```
%%timeit
ejemplo_1_datos['PIB'].apply(entre_un_millon)
%%timeit
ejemplo_1_datos['PIB'] / 1_000_000
```
Utilizando `.apply()` te tardas mas o menos 190 ± 30.3 ***microsegundos*** y al dividir utilizando solo el símbolo `/` te tardas 91.7 ± 2.74 ***microsegundos***. O te ahorras un poco mas de la mitad de tiempo.
Obviamente, el hecho de que este "ahorro" de tiempo sea en ***micro***segundos es muy importante: Tu cerebro es incapaz de notar la diferencia. Ese "ahorro" del 52% de tiempo solo importa si te vas a tardar 29 minutos en lugar de una hora o uno en lugar de dos días ejecutando tu código. Esto solo sucede si tienes una cantidad enorme de datos y si estas trabajando con tantos datos probablemente estes haciendo calculos más complicados que dividir entre un millón.
Así que la primera conclusión de este blog es que la verdad es mucho mas importante la legibilidad de tu código a que sea la manera _óptima_ de trabajar con tus datos **si estás trabajando con pocos datos**. Pocos es relativo, 8 filas es poco pero yo he trabajado con conjuntos de datos que tienen 8 millones de filas y mi computadora los maneja muy bien así que podría decirse que también eso es poco. "Pocos" datos son los que puedes manipular libremente en tu computadora sin que se congele o se vuela lenta mientras trabajas con ellos.
## Ejemplo 2: Re-etiquetar datos
Un proceso común en el análisis de datos es "re-etiquetar" columnas. En mi caso, como _científico social_ trabajo mucho con ciertos atributos de poblaciones (sexo, edad, raza/etnia, etc.) y es muy común que tengamos que agrupar personas y "re-etiquetar" estos nuevos grupos. Aquí dos ejemplos.
### Agrupar edades
Imagina que tenemos un DataFrame como el siguiente:
```
#collapse
import pandas as pd
import numpy as np
# Vamos a ponerle este "seed" para que puedas reproducir esto si así lo deseas
np.random.seed(13)
n_observaciones = 5000
raza_etnias = ['Mexican', 'Cuban', 'Puerto Rican', 'African-American', 'Chinese', 'Japanese', 'White', 'Native American', 'Other']
ejemplo_2_datos = pd.DataFrame({"edad": np.random.randint(0, 100, n_observaciones), "raza_etnia": np.random.choice(raza_etnias, n_observaciones) })
ejemplo_2_datos.head()
```
Yo trabajo mucho con datos del censo de Estados Unidos y siempre es un problema trabajar con las variables "race" y "ethinicity" ya que es un concepto complejo. Para concentrarnos en el `.apply()` estoy simplificando esto un poco y creando una sola columna llamada `raza_etnia` que vamos a re-agrupar.
Pero primero, la edad. Veamos su distribución:
```
#collapse
import altair as alt
alt.Chart(ejemplo_2_datos).mark_bar().encode(
x = "edad",
y = "count()"
)
```
Puedes ver que estos datos son muy _granulares_ - yo como investigador quiero saber ciertos atributos de las personas que tienen entre 20 y 30 años, por ejemplo. No de las personas que tienen 21 o 25 o 27. Entonces necesito agrupar esta columna en _bins_ o cubetas. En lugar de que cada observación (fila) en mi DataFrame tenga una etiqueta con su edad quiero que tenga una etiqueta con el grupo al que pertenece. Si alguien tiene 34 años, esta nueva columna tiene que decir `"Entre 30 y 40"` no `34`.
Una de las maneras que he visto mucha gente lograr esto es con `.apply()` y una función simple con varios `if-else`. Así:
```
def etiquetas_edad(numero):
"""Agrupa números a su decena. Recibe un número y retorna la etiqueta correspondiente."""
if 0 <= numero <= 10:
return "Entre 0 y 10"
elif 10 < numero <= 20:
return "Entre 10 y 20"
elif 20 < numero <= 30:
return "Entre 20 y 30"
elif 30 < numero <= 40:
return "Entre 30 y 40"
elif 40 < numero <= 50:
return "Entre 40 y 50"
elif 50 < numero <= 60:
return "Entre 50 y 60"
elif 60 < numero <= 70:
return "Entre 60 y 70"
elif 70 < numero <= 80:
return "Entre 70 y 80"
elif 80 < numero <= 90:
return "Entre 80 y 90"
elif 90 < numero <= 100:
return "Entre 90 y 100"
else:
return "Error"
```
Existen maneras mas "elegantes" de hacerlo pero cuando uno sabe los números que esperas (no esperas a nadie con edad negativa o mayor a 100, en este caso) y tiene prisa, esta solución, pues, _funciona_.
```
ejemplo_2_datos['edad'].apply(etiquetas_edad)
```
Pero, `pandas` tiene la función `pandas.cut()` que justo hace esto ([documentación](https://pandas.pydata.org/docs/reference/api/pandas.cut.html)).
`pandas.cut()` recibe una matriz, las _bins_ o "cubetas" (grupos), y las etiquetas de estos grupos (entre otros argumentos) para agrupar una columna "continua" (como lo es la `edad`) y crear grupos discretos en una variable `Categorical` que tiene otras funcionalidades útiles.
```
grupos_de_edades = [0, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100]
etiquetas_de_edades = [
"Entre 0-10",
"Entre 10-20",
"Entre 20-30",
"Entre 30-40",
"Entre 40-50",
"Entre 50-60",
"Entre 60-70",
"Entre 70-80",
"Entre 80-90",
"Entre 90-100"
]
pd.cut(ejemplo_2_datos['edad'], grupos_de_edades, labels = etiquetas_de_edades)
```
Se logra "lo mismo" pero con `pandas.cut()` recibes una columna de tipo `Categorical` y "ordenada" lo cual significa que `pandas` entiende que "Entre 20-30" es un valor _menor_ a "Entre 30-40". Esto te permite crear filtros como
```python
ejemplo_2_datos['edad_con_pd_cut'] > "Entre 30-40"
```
Y esto filtraría a todas las personas menores de 30 años. Si no estas usando `Categorical`s tendrías que crear un filtro para cada una de las etiquetas. Pero eso para otro blog.
Pero bueno, comparemos estos dos métodos:
```
%%timeit
ejemplo_2_datos['edad'].apply(etiquetas_edad)
%%timeit
pd.cut(ejemplo_2_datos['edad'], grupos_de_edades, labels = etiquetas_de_edades)
```
Como puedes ver, `pandas.cut()` es una vez mas un poco más rápida y te ahorras otra vez casi la mitad del tiempo. Esto, de nuevo, es imperceptible en realidad - la diferencia es de menos de un **mili**segundo.
En mi caso, prefiero `pandas.cut()` porque no solo es un poquitín mas rápida sino que tiene la ventaja de regresarme una columna `Categorical` y puedo usar eso a mi ventaja en mis análisis.
:bulb: Bonus: el buen [@io_exception](https://twitter.com/io_exception) compartió un ejemplo de como hacer esto mas elegantemente en el discord en un [Google Colab notebook](https://colab.research.google.com/drive/1WtYaQkNxYSHqO_rU6qjnY-7p4od1DIWD?usp=sharing#scrollTo=8FUsFKu-vReo)
```
ejemplo_2_datos['edad_cut'] = pd.cut(ejemplo_2_datos['edad'], grupos_de_edades, labels = etiquetas_de_edades)
# puedo utilizar lógica con estas etiquetas como si fueran valores numéricos
filtro_edad = ejemplo_2_datos['edad_cut'] >= "Entre 30-40"
alt.Chart(ejemplo_2_datos[filtro_edad]).mark_bar().encode(
x = alt.X("edad", scale = alt.Scale(domain = [0,100])),
y = "count()"
)
```
### Agrupar raza/etnia
Al igual como agrupamos nuestra columna de edad podemos _agrupar_ nuestra columna `raza_etnia` re-etiquetando los valores de la siguiente manera.
```
def etiquetas_raza_etnia(raza_etnia):
"""Re-etiqueta un valor de la columna raza_etnia a otro."""
if raza_etnia == "Mexican":
return "Latino"
elif raza_etnia == "Cuban":
return "Latino"
elif raza_etnia == "Puerto Rican":
return "Latino"
elif raza_etnia == "African-American":
return "Black"
elif raza_etnia == "Chinese":
return "Asian"
elif raza_etnia == "Japanese":
return "Asian"
elif raza_etnia == "White":
return "White"
elif raza_etnia == "Native American":
return "Native American"
else:
return "Other"
```
Esto nos permite agrupar personas que tienen el valor "Mexican", "Cuban" o "Puerto Rican" bajo la etiqueta "Latino".
```
ejemplo_2_datos['raza_etnia'].apply(etiquetas_raza_etnia)
```
Pero este proceso de re-etiquetado es tan común que las `Series` de `pandas` tienen el método `.map()` para llevar a cabo lo mismo. Esta función recibe un diccionario de python donde las claves (_keys_) son los valores que esperas ver en tu Serie y los valores son los valores que substituirán esas claves.
- https://pandas.pydata.org/docs/reference/api/pandas.Series.map.html
```
valores_nuevos = {
"Mexican": "Latino",
"Cuban": "Latino",
"Puerto Rican": "Latino",
"African-American": "Black",
"Chinese": "Asian",
"Japanese": "Asian",
"White": "White",
"Native American": "Native American",
}
ejemplo_2_datos['raza_etnia'].map(valores_nuevos)
```
`pandas.Series.map()` recibe el diccionario con los valores y el argumento `na_action` (que puede ser None o "ignore") que le dice a `.map()` que hacer con los valores nulos `NaN`. Algo que me gusta de `.map()` es que si en lugar de darle un diccionario le das un `defaultdict` (un diccionario con un valor default) `pandas` utiliza ese valor default para cualquier valor original de tu columna que no aparezca en las claves del diccionario.
Te habrás dado cuenta que no tenemos el `"Other"` en nuestro ejemplo con `.map()`. Si vemos nuestra nueva serie podemos ver que tiene algunos valores `NaN` ya que la columna original tiene valores que no aparecen en nuestro diccionario `valores_nuevos`.
```
ejemplo_2_datos['raza_etnia'].map(valores_nuevos).isna().sum() # cuenta cuantos valores `NaN` existen en la Serie
```
Estos valores `NaN` deberían ser `"Other"` y podemos hacerlo _encadenando_ métodos o utilizando el `defaultdict`
```
ejemplo_2_datos['raza_etnia'].map(valores_nuevos).fillna("Other") # llenamos los NaN con "Other"
ejemplo_2_datos['raza_etnia'].map(valores_nuevos).fillna("Other").isna().sum() # contamos los NaNs para asegurarnos que no hay
```
¿Cual es más rápida?
```
%%timeit
ejemplo_2_datos['raza_etnia'].apply(etiquetas_raza_etnia)
%%timeit
ejemplo_2_datos['raza_etnia'].map(valores_nuevos).fillna("Other")
```
Incluyendo el _encadenamiento_ de métodos (`.map().fillna()`) estas funciones nativas de `pandas` son un poco más rápidas. Pero ahora estas utilizando dos métodos. Dependiendo de que prefieras, crear tu función que tome en cuenta que hacer con valores que **no** esperas encontrar en tu columna (el default de `"Other"`) o encadenar 2 métodos de `pandas`, es la decisión que debes tomar.
## Ejemplo 3: Limpiando `strings`
Otro uso muy común para `.apply()` que he visto es el de trabajar con texto especialmente cuando se quiere trabajar con expresiones regulares. Dos casos particulares son 1) extraer _cosas_ del texto y 2) transformarlo o limpiarlo.
### Extraer texto con expresiones regulares
Utilicemos un dataset con tweets para trabajar con este tipo de datos. Primero veamos una función sencilla que extrae usuarios etiquetados en el tweet. O sea el @ y la siguiente palabra.
```
ejemplo_3_datos = pd.read_csv("https://raw.githubusercontent.com/zfz/twitter_corpus/master/full-corpus.csv")
ejemplo_3_datos.head()
```
Este ejemplo, vamos a hacer algo distinto. No te voy a mostrar las maneras que he visto personas utilizar el `.apply()`, mejor te mostraré las funciones nativas de `pandas` para trabajar con texto ya que son varias y este blog ya esta muy largo :sweat_smile:
La manera de trabajar con texto en una columna de un DataFrame es con el _accessor_ `.str` - esta es una manera de acceder a métodos específicamente diseñados para _strings_ en `pandas`. También existen otros _accessors_ como `.dt` para trabajar con datos de tipo `datetime` o fechas y `.cat` para trabajar con `Categoricals`. Para una lista completa de los métodos que el _accessor_ `.str` te da puedes visitar la documentación oficial: https://pandas.pydata.org/pandas-docs/stable/reference/series.html#string-handling.
Vamos a utilizar `.str.findall()` para encontrar los usuarios etiquetados en tweets.
:bulb: Nota: existe el método `.str.find()` que hace algo un poco distinto. Eso para otro blog.
```
ejemplo_3_datos['TweetText'].str.findall(r"@\w+")
```
Con el _accessor_ `.str` puedes hacer casi todo lo que puedes hacer nativamente en python con `strings`
```
ejemplo_3_datos['Topic'].str.capitalize()
ejemplo_3_datos['Sentiment'].str.upper()
```
Y como es `pandas` podemos encadenar estos métodos pero como son métodos **dentro** del _accessor_ de `.str` tienes que repetir el `.str` entre cada método.
```
ejemplo_3_datos['TweetText'].str.findall(r"@\w+").str.join(", ")
```
***
:rotating_light::rotating_light::rotating_light:
Este post esta _en construcción_ ¡o sea que todavía no acaba!
Estamos publicándolo hoy 23 de abril del 2021 como adelanto. En la semana lo terminamos.
| github_jupyter |
# Find duplicates in Cifar10 with imagededup
We will download the Cifar10 dataset and then run the **CNN** method from **imagededup** to detect duplicates.
```
# install imagededup via PyPI
!pip install imagededup
# by default imagededup is shipped with CPU-only support for TF but let's install GPU since we have it on Google Colab
# finding duplicates with Convolutional Neural Network (CNN) is much faster on GPU
!pip install tensorflow-gpu==2.0.0
# download CIFAR10 dataset and untar
!wget http://pjreddie.com/media/files/cifar.tgz
!tar xzf cifar.tgz
# create working directory and move all images into this directory
image_dir = 'cifar10_images'
!mkdir $image_dir
!cp -r '/content/cifar/train/.' $image_dir
!cp -r '/content/cifar/test/.' $image_dir
# find duplicates in the entire dataset with CNN
from imagededup.methods import CNN
cnn = CNN()
encodings = cnn.encode_images(image_dir=image_dir)
duplicates = cnn.find_duplicates(encoding_map=encodings)
# do some imports for plotting
from pathlib import Path
from imagededup.utils import plot_duplicates
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (15, 10)
```
## Duplicates in test set
```
# test images are stored under '/content/cifar/test'
filenames_test = set([i.name for i in Path('/content/cifar/test').glob('*.png')])
# keep only filenames that are in test set
duplicates_test = {}
for k, v in duplicates.items():
if k in filenames_test:
tmp = [i for i in v if i in filenames_test]
duplicates_test[k] = tmp
# sort in descending order of duplicates
duplicates_test = {k: v for k, v in sorted(duplicates_test.items(), key=lambda x: len(x[1]), reverse=True)}
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test, filename=list(duplicates_test.keys())[0])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test, filename=list(duplicates_test.keys())[15])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test, filename=list(duplicates_test.keys())[20])
```
## Duplicates in train set
```
# train images are stored under '/content/cifar/train'
filenames_train = set([i.name for i in Path('/content/cifar/train').glob('*.png')])
# keep only filenames that are in train set
duplicates_train = {}
for k, v in duplicates.items():
if k in filenames_train:
tmp = [i for i in v if i in filenames_train]
duplicates_train[k] = tmp
# sort in descending order of duplicates
duplicates_train = {k: v for k, v in sorted(duplicates_train.items(), key=lambda x: len(x[1]), reverse=True)}
# 70 duplicates found of same car!
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_train, filename=list(duplicates_train.keys())[0])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_train, filename=list(duplicates_train.keys())[70])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_train, filename=list(duplicates_train.keys())[140])
```
## Examples from test set with duplicates in train set
```
# keep only filenames that are in test set have duplicates in train set
duplicates_test_train = {}
for k, v in duplicates.items():
if k in filenames_test:
tmp = [i for i in v if i in filenames_train]
duplicates_test_train[k] = tmp
# sort in descending order of duplicates
duplicates_test_train = {k: v for k, v in sorted(duplicates_test_train.items(), key=lambda x: len(x[1]), reverse=True)}
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test_train, filename=list(duplicates_test_train.keys())[0])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test_train, filename=list(duplicates_test_train.keys())[15])
plot_duplicates(image_dir=image_dir, duplicate_map=duplicates_test_train, filename=list(duplicates_test_train.keys())[50])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/kevincong95/cs231n-emotiw/blob/master/audio/opensmile_audio.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/gdrive' , force_remount=True)
!pip install pydub
# Imports
from tensorflow.keras.models import load_model
from moviepy.video.io.ffmpeg_tools import ffmpeg_extract_audio
import os
import glob
import time
import importlib
from sklearn.preprocessing import StandardScaler
from subprocess import Popen, PIPE, STDOUT
os.system("pip install pydub")
os.chdir('/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/notebooks/audio-new')
import arffToNp
importlib.reload(arffToNp)
import subprocess
import numpy as np
```
# Audio API
```
# TODO: add option for soft vs hard
def predict(mp4_filepath, best_model_filepath):
"""
Outputs:
- A tuple with predictions for each class (positive, neutral, negative)
"""
model = fer_model()
model.load_model(best_model_filepath)
return model.predict(mp4_filepath)
class audio_model:
def __init__(self):
self.model = ()
return
def predict(self, mp4_filepath):
self.preprocess(mp4_filepath)
X = cv2.imload("test/happy.jpg")
X = cv2.resize(X, (48,48))
X = cv2.cvtColor(X, cv2.COLOR_BGR2GRAY)
return self.model.predict(img)
#return (0.1,0.2,0.7)
def load_model(self, best_model_filepath):
self.model = load_model(best_model_filepath)
return
def train(self, X_train , y_train):
# train the model
# TO DO : Implement a recurrent network for each "picture"
# Look into autoencoders/PCA for dim reduction
# Reccurent network on row time steps
inputs = keras.Input(shape=[216,128])
recurrent_1 = keras.layers.LSTM(100, return_sequences=True, input_shape=[None, 128] , dropout=0.2 , activation='selu') #A sequence of any length with dimensions 128 (i.e. 128 columns)
recurrent_2 = keras.layers.LSTM(32)
# We use the row of the image at each time step. That's why we have a repeat
# vector layer.
#repeat_vector = keras.layers.RepeatVector(216, input_shape=[30])
#recurrent_3 = keras.layers.LSTM(100, return_sequences=False)
dense_1 = keras.layers.Dense(32 , activation='selu')
softmax = keras.layers.Dense(3 , activation='softmax')
x = recurrent_1(inputs)
x = recurrent_2(x)
#x = repeat_vector(x)
#x = recurrent_3(x)
x = dense_1(x)
outputs = softmax(x)
rnn_ae = keras.Model(inputs=inputs, outputs=outputs)
lr_schedule = keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=1e-5,
decay_steps=10000,
decay_rate=0.9)
opt = keras.optimizers.Adam(learning_rate=lr_schedule)
rnn_ae.compile(loss='sparse_categorical_crossentropy' , optimizer=opt , metrics=['accuracy'])
rnn_ae.fit(X_train , y_train , epochs=500 , batch_size=32)
return rnn_ae
def preprocess(self, mp4_filepath , target_label_path=None):
"""
Outputs:
- A numpy array with dimensions (m,n).
- m is the units in time dependent on the audio splice rate.
- n is the number of features from the openSMILE library.
"""
output_wav_file = mp4_filepath[-5] + 'extracted_audio.wav'
mp4_filename = os.path.basename(mp4_filepath)
audio_home_dir = os.path.dirname(mp4_filepath)
os.chdir(audio_home_dir)
# Strip the audio from video and store as .wav file
ffmpeg_extract_audio(mp4_filepath, output_wav_file)
!cd '$audio_home_dir' ; mkdir to_zip
# splice the audio files into 2 seconds with 100 ms sliding window.
# 30 kHz sampling rate
!cd '$audio_home_dir' ; python SliceAudio.py -i *.wav -o wav -c 2 -b 2 -s 30000 -w 100 -l 2000
# Zip and move files from drive to vm
!cd '$audio_home_dir' ; zip -r -qq to_zip.zip to_zip ; cd '$audio_home_dir' ; mv 'to_zip.zip' '/content/'
# Remove the old zip folder in vm
!cd '$audio_home_dir' ; cd to_zip ; rm *.wav ; cd - ; rm -d to_zip
# Inflate the zip folder in vm
!cd '/content/' ; unzip -qq to_zip.zip
# OpenSMILE feature extraction
out_fn = os.path.join('/content/openSmile-features.arff')
os.chdir('/content/to_zip/')
aligned_files = glob.glob('*.wav')
os.chdir('/content/')
counter = 0
for in_fn in aligned_files:
in_fn = os.path.join('/content/to_zip/' , in_fn)
name = os.path.basename(in_fn)
if counter % 2 == 0:
!cd 'opensmile-2.3.0' ; inst/bin/SMILExtract -C config/IS09_emotion.conf -I '$in_fn' -O '$out_fn' -N $name
counter += 1
# Convert .arff to .csv
all_timepoints_feature_array = arffToNp.convert(out_fn , '')
print("The shape of the feature matrix for one \n video is: " , all_timepoints_feature_array.shape)
# Clean up
!cd to_zip ; rm *.wav ; cd - ; rm -d to_zip
os.remove(out_fn)
!cd '$audio_home_dir' ; rm *.wav ; rm *.csv
!rm to_zip.zip
!rm *.csv
!rm *.arff
# Standardize
scaler = StandardScaler()
all_timepoints_feature_array = scaler.fit_transform(all_timepoints_feature_array)
# Get the Y values
target = None
if target_label_path is not None:
target_labels = np.genfromtxt(target_label_path , delimiter = ' ' , dtype='str')
target_index = np.where(target_labels[: , 0] == mp4_filename[:-4])[0]
target = int(target_labels[: , 1][target_index])
return all_timepoints_feature_array , target
# Read in each video file and add the (m,n) feature matrix to a 3D array
def get_feature_batch(self, input_files_dir , batch_size=3000 , target_label_path=None):
"""
Inputs:
- Path to the .mp4 files
Outputs:
- An ndarray with dims (s , m , n)
- s is the number of samples
- m is the number of slices for that sample (32)
- n is the number of features (6373)
"""
output_x = []
output_y = None
if target_label_path is not None:
output_y = []
counter = 0
fileList = glob.glob(input_files_dir + '*.mp4')
print(fileList)
for file_path in fileList:
print(file_path)
one_sample_feat_matrix , y = self.preprocess(file_path , target_label_path=target_label_path)
output_x.append(one_sample_feat_matrix)
if target_label_path is not None:
output_y.append(y)
if counter >= batch_size:
break
counter += 1
output_y = np.asarray(output_y)
output_y -= 1
return output_x , output_y
def installOpenSMILE():
"""
You must upload your downloaded version of openSMILE from the site to
cloud.
"""
os.chdir('/content/')
!tar -zxvf 'opensmile-2.3.0.tar.gz'
!sed -i '117s/(char)/(unsigned char)/g' opensmile-2.3.0/src/include/core/vectorTransform.hpp
!sudo apt-get update
!sudo apt-get install autoconf automake libtool m4 gcc
!cd 'opensmile-2.3.0' ; bash buildStandalone.sh
installOpenSMILE()
!cd 'opensmile-2.3.0' ; inst/bin/SMILExtract -h
audio_model_1 = audio_model()
target_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/datasets/emotiw/Val_labels.txt'
output_arr , y = audio_model_1.preprocess(mp4_filepath='/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/notebooks/audio-new/1_1.mp4' , target_label_path=target_path)
output_arr.shape
```
# Set up Processing Pipeline
```
# Imports
import tensorflow as tf
from tensorflow import keras
# Preprocessing Pipeling
home_dir = '/content/'
!cd '$home_dir'; mkdir train_vids ; mkdir val_vids
train_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/datasets/emotiw/Train.zip'
train_target_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/datasets/emotiw/Val_labels.txt'
val_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/datasets/emotiw/Val.zip'
val_target_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/datasets/emotiw/Val_labels.txt'
vm_train_path = '/content/train_vids'
vm_val_path = '/content/val_vids'
slice_audio_path = '/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/notebooks/audio-new/SliceAudio.py'
# Copy files to vm and inflate
!cp '$train_path' '$vm_train_path' ; cd '$vm_train_path' ; unzip -qq 'Train.zip'
!cp '$val_path' '$vm_val_path' ; cd '$vm_val_path' ; unzip -qq 'Val.zip'
inflated_train_path = '/content/train_vids/Train/'
inflated_val_path = '/content/val_vids/Val/'
# Copy SliceAudio to dirs
!cp '$slice_audio_path' '$inflated_train_path' ; cp '$slice_audio_path' '$inflated_val_path'
audio_model_test = audio_model()
X , y = audio_model_test.get_feature_batch(inflated_val_path , batch_size=3000 , target_label_path=train_target_path)
X_np = np.asarray(X)
print(X_np.shape)
counter = 0
for arr in X_np:
if arr.shape[0] != 16:
print(counter)
counter += 1
X_new = np.delete(X_np , [100])
y_new = np.delete(y , [100])
for i in range(0 , len(X_new)):
#print(X_new[i].dtype)
if X_new[i].shape != (16 , 384):
print("help")
#X_new_np = np.asarray(X_new , dtype='float32')
np_x = np.zeros((765 , 16 , 384))
for i in range (0, 765):
for j in range (0 , 16):
for k in range (0,384):
np_x[i][j][k] = X_new[i][j][k]
print(np_x.shape)
print(y_new.shape)
import pickle
with open('/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/notebooks/audio-new/arrays/audio.val-opensmile384-standardized_X_np.all.pickle', 'wb') as f:
pickle.dump(np_x , f)
with open('/content/gdrive/My Drive/Machine-Learning-Projects/cs231n/notebooks/audio-new/arrays/audio.val-opensmile384-standardized_y_np.all.pickle', 'wb') as f:
pickle.dump(y_new, f)
```
| github_jupyter |
# 1. Data Processing - Build a data processing pipeline to process electronic medical reports (EMR) using Amazon Textract and Comprehend Medical
In this notebook, we will walkthrough on how to build a data processing pipeline that will process electronic medical reports (EMR) in PDF format to extract relevant medical information by using the following AWS services:
- [Textract](https://aws.amazon.com/textract/): To extract text from the PDF medical report
- [Comprehend Medical](https://aws.amazon.com/comprehend/medical/): To extract relevant medical information from the output of textract
## Contents
1. [Objective](#Objective)
1. [Setup Environment](#Setup-Environment)
1. [Extract text using Amazon Textract](#Step-1:-Process-PDF-with-Amazon-Textract)
1. [Extract medical information using Amazon Comprehend Medical](#Step-2:-Extract-medical-information-with-Amazon-Comprehend-Medical)
1. [Clean up resources](#Clean-up-resources)
---
# Objective
The objective of this section of the workshop is to learn how to use Amazon Textract and Comprehend Medical to extract the medical information from an electronic medical report in PDF format.
---
# Setup environment
Before be begin, let us setup our environment. We will need the following:
* Amazon Textract Results Parser `textract-trp` to process our Textract results.
* Python libraries
* Pre-processing functions that will help with processing and visualization of our results. For the purpose of this workshop, we have provided a pre-processing function library that can be found in [util/preprocess.py](./util/preprocess.py)
Note: `textract-trp` will require Python 3.6 or newer.
```
!pip install textract-trp
import boto3
import time
import sagemaker
import os
import trp
from util.preprocess import *
import pandas as pd
bucket = sagemaker.Session().default_bucket()
prefix = 'sagemaker/medical_notes'
```
---
# Step 1: Process PDF with Amazon Textract
In this section we will be extracting the text from a medical report in PDF format using Textract. To facilitate this workshop, we have generated a [sample PDF medical report](./data/sample_report_1.pdf) using the [MTSample dataset](https://www.kaggle.com/tboyle10/medicaltranscriptions) from kaggle.
## About Textract
Amazon Textract can detect lines of text and the words that make up a line of text. Textract can handle documents in either synchronous or asynchronous processing:
+ [synchronous API](https://docs.aws.amazon.com/textract/latest/dg/sync.html): supports *The input document must be an image in `JPEG` or `PNG` format*. Single page document analysis can be performed using a Textract synchronous operation.
1. *`detect_document_text`*: detects text in the input document.
2. *`analyze_document`*: analyzes an input document for relationships between detected items.
+ [asynchronous API](https://docs.aws.amazon.com/textract/latest/dg/async.html): *can analyze text in documents that are in `JPEG`, `PNG`, and `PDF` format. Multi page processing is an asynchronous operation. The documents are stored in an Amazon S3 bucket. Use DocumentLocation to specify the bucket name and file name of the document.*
1. for context analysis:
1. *`start_document_text_detection`*: starts the asynchronous detection of text in a document.
2. *`get_document_text_detection`*: gets the results for an Amazon Textract asynchronous operation that detects text in a document.
2. for relationships between detected items :
1. *`start_document_analysis`*: starts the asynchronous analysis of relationship in a document.
2. *`get_document_analysis`*: Gets the results for an Amazon Textract asynchronous operation that analyzes text in a document
For detailed api, refer to documentation [here](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/textract.html#Textract.Client.analyze_document).
In this demo, as the input is in pdf format and has multiple pages, we will be using the multi page textract operation, we will need to upload our sample medical record to an S3 bucket. Run the next cell to upload our sample medical report.
```
fileName = 'sample_report_1.pdf'
fileUploadPath = os.path.join('./data', fileName)
textractObjectName = os.path.join(prefix, 'data', fileName)
# Upload medical report file
boto3.Session().resource('s3').Bucket(bucket).Object(textractObjectName).upload_file(fileUploadPath)
```
## Start text detection asynchonously in the pdf
In the next step, we will start the asynchronous textract operation by calling the `start_document_analysis()` function. The function will kickoff an asynchronous job that will process our medical report file in the stipulated S3 bucket.
```
textract = boto3.client('textract')
response = textract.start_document_analysis(
DocumentLocation={
'S3Object': {
'Bucket': bucket,
'Name': textractObjectName
}},
FeatureTypes=[
'TABLES',
]
)
textractJobId = response["JobId"]
print('job id is: ',textractJobId)
```
## Monitor the job status
As the job is kicked off in the background, we can monitor the progress of the job by calling the `get_document_analysis()` function and passing the job id of the job that we created.
Run the next cell and wait for the Textract Job status to return a SUCCEEDED status.
the outcome is in json format
```
%%time
time.sleep(5)
response = textract.get_document_analysis(JobId=textractJobId)
status = response["JobStatus"]
while(status == "IN_PROGRESS"):
time.sleep(5)
response = textract.get_document_analysis(JobId=textractJobId)
status = response["JobStatus"]
print("Textract Job status: {}".format(status))
```
## Extract textract results
Now that we've successfully extracted the text from the medical report, let us extract the textract results and consolidate the text so that we can pass it to Comprehend Medical to start extract medical information from the report.
```
%%time
pages = []
time.sleep(5)
response = textract.get_document_analysis(JobId=textractJobId)
pages.append(response)
nextToken = None
if('NextToken' in response):
nextToken = response['NextToken']
while(nextToken):
time.sleep(5)
response = textract.get_document_analysis(JobId=textractJobId, NextToken=nextToken)
pages.append(response)
print("Resultset page recieved: {}".format(len(pages)))
nextToken = None
if('NextToken' in response):
nextToken = response['NextToken']
```
### Output from Textract
Let's take a look at the output from textract by using the trp library to extract and format the textract results.
```
doc = trp.Document(pages)
print("Total length of document is",len(doc.pages))
idx=1
for page in doc.pages:
print(color.BOLD + f"Results from page {idx}: \n" + color.END, page.text)
idx=idx+1
```
---
# Step 2: Extract medical information with Amazon Comprehend Medical
## About Amazon Comprehend Medical
Comprehend Medical detects useful information in unstructured clinical text. As much as 75% of all health record data is found in unstructured text such as physician's notes, discharge summaries, test results, and case notes. Amazon Comprehend Medical uses Natural Language Processing (NLP) models to sort through text for valuable information.
Using Amazon Comprehend Medical, you can quickly and accurately gather information, such as medical condition, medication, dosage, strength, and frequency from a variety of sources like doctors’ notes. Amazon Comprehend Medical uses advanced machine learning models to accurately and quickly identify medical information, such as medical conditions and medications, and determines their relationship to each other, for instance, medicine dosage and strength. Amazon Comprehend Medical can also link the detected information to medical ontologies such as ICD-10-CM or RxNorm
Currently, Amazon Comprehend Medical only detects medical entities in English language texts.

With Amazon Comprehend Medical, you can perform the following on your documents:
- [Detect Entities (Version 2)](https://docs.aws.amazon.com/comprehend/latest/dg/extracted-med-info-V2.html) - Examine unstructured clinical text to detect textual references to medical information such as medical condition, treatment, tests and results, and medications. This version uses a new model and changes the way some entities are returned in the output. For more information, see [DetectEntitiesV2](https://docs.aws.amazon.com/comprehend/latest/dg/API_medical_DetectEntitiesV2.html).
- [Detect PHI (Verdion 1)](https://docs.aws.amazon.com/comprehend/latest/dg/how-medical-phi.html) —Examine unstructured clinical text to detect textual references to protected health information (PHI) such as names and addresses.
In this workshop, we will be using the detect entities function ([detect_entities_v2](https://docs.aws.amazon.com/comprehend/latest/dg/extracted-med-info-V2.html)) to extract medical conditions. In the following cell, we will be processing the text on each page in batches of 20,000 UTF-8 characters. This is because Comprehend Medical has a maximum document size of 20,000 bytes (reference: https://docs.aws.amazon.com/comprehend/latest/dg/guidelines-and-limits-med.html). Once we've processed the text, we will then stich up the response into a into a single variable where we can either save to a csv or use for our analysis.
```
maxLength=20000
comprehendResponse = []
comprehend_medical_client = boto3.client(service_name='comprehendmedical', region_name='us-east-1')
for page in doc.pages:
pageText = page.text
for i in range(0, len(pageText), maxLength):
response = comprehend_medical_client.detect_entities_v2(Text=pageText[0+i:maxLength+i])
comprehendResponse.append(response)
patient_string = ""
```
## Review comprehend results
The output of *detect_entities_v2* can detect the following entities:
- `MEDICAL_CONDITION`: The signs, symptoms, and diagnosis of medical conditions.
- `Score` - The level of confidence that Amazon Comprehend Medical has in the accuracy of the detection
- `Trait` - Contextual information for the entity
Other information extracted by Comprehend Medical:
- `MEDICATION`: Medication and dosage information for the patient.
- `PROTECTED_HEALTH_INFORMATION`: patient's personal information, e.g. name, age, gender
- `TEST_TREATMENT_PROCEDURE`: the procedures that are used to determine a medical condition.
- `TIME_EXPRESSION`: Entities related to time when they are associated with a detected entity.
For this workshop, we will be using the MEDICAL_CONDITION entity to train our machine learning model. Let us take a look at some of the data.
```
## use our pre-defined util function extractMC_v2 to extract all the medical conditions, confidence score, trait from json file
df_cm=extractMC_v2(comprehendResponse[0])
df_cm['ID']=1
df_cm.head(10)
```
---
# Clean up resources
As some resources will be used in the following step, We will clean up the resource until the end of the lab. You may uncomment the following sentence if this is the last step of your lab
```
##boto3.Session().resource('s3').Bucket(bucket).Object(textractObjectName).delete()
```
| github_jupyter |
#data load
```
from keras.datasets import fashion_mnist, cifar10
import numpy as np
import matplotlib.pyplot as plt
from keras.utils import to_categorical
import cv2
import keras
class data:
def __init__(self, data_size):
self.data_size = data_size
def resize(self, x, to_color=True):
result = []
for i in range(len(x)):
if to_color:
img = cv2.cvtColor(x[i], cv2.COLOR_GRAY2RGB)
img = cv2.resize(img,dsize=(96,96))
else:
img = cv2.resize(x[i],dsize=(96,96))
result.append(img)
return np.array(result)
def choose_data(self, x, y, normal_id, anomaly_id):
x_normal, x_anomaly = [], []
x_ref, y_ref = [], []
for i in range(len(x)):
if y[i] == normal_id:# スニーカーは7#鹿は4
x_normal.append(x[i].reshape((x.shape[1:])))
elif y[i] == anomaly_id:# ブーツは9#馬は7
x_anomaly.append(x[i].reshape(x.shape[1:]))
else:
x_ref.append(x[i].reshape((x.shape[1:])))
y_ref.append(y[i])
return np.array(x_normal), np.array(x_anomaly), np.array(x_ref), y_ref
def random_choose(self, x, y, size):
number = np.random.choice(np.arange(0, len(x)), size, replace=False)
return_x, return_y = [], []
if len(y) > 0:
for i in number:
return_x.append(x[i].reshape((x.shape[1:])))
return_y.append(y[i])
return np.array(return_x), return_y
else:
for i in number:
return_x.append(x[i].reshape((x.shape[1:])))
return np.array(return_x)
def get_fashion_data(self):
# dataset
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
x_train_normal, _, x_ref, y_ref = self.choose_data(x_train, y_train, 7, 9)
#train_normalデータからランダムに抽出
x_train_normal = self.random_choose(x = x_train_normal, y=[], size = int(self.data_size/8))
#refデータからランダムに抽出
x_ref, y_ref = self.random_choose(x = x_ref, y = y_ref, size = self.data_size)
y_ref = to_categorical(y_ref)
# テストデータ
x_test_normal, x_test_anomaly, _, _ = self.choose_data(x_test, y_test, 7, 9)
x_train_normal = self.resize(x_train_normal)
x_ref = self.resize(x_ref)
x_test_normal = self.resize(x_test_normal)
x_test_anomaly = self.resize(x_test_anomaly)
#print(x_train_normal.shape)
#print(x_ref.shape)
#print(y_ref.shape)
#print(x_test_normal.shape)
#print(x_test_anomaly.shape)
return x_train_normal, x_ref, y_ref, x_test_normal, x_test_anomaly
def get_cifar_data(self):
# dataset
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
x_train_normal, _, x_ref, y_ref = self.choose_data(x_train, y_train, 4, 7)
#train_normalデータからランダムに抽出
x_train_normal = self.random_choose(x = x_train_normal, y=[], size = int(self.data_size/8))
#refデータからランダムに抽出
x_ref, y_ref = self.random_choose(x = x_ref, y = y_ref, size = self.data_size)
y_ref = to_categorical(y_ref)
# テストデータ
x_test_normal, x_test_anomaly, _, _ = self.choose_data(x_test, y_test, 4, 7)
x_train_normal = self.resize(x_train_normal, False)
x_ref = self.resize(x_ref, False)
x_test_normal = self.resize(x_test_normal, False)
x_test_anomaly = self.resize(x_test_anomaly, False)
#print(x_train_normal.shape)
#print(x_ref.shape)
#print(y_ref.shape)
#print(x_test_normal.shape)
#print(x_test_anomaly.shape)
return x_train_normal, x_ref, y_ref, x_test_normal, x_test_anomaly
def get_auc(Z1, Z2):
y_true = np.zeros(len(Z1)+len(Z2))
y_true[len(Z1):] = 1#0:正常、1:異常
# FPR, TPR(, しきい値) を算出
fpr, tpr, _ = metrics.roc_curve(y_true, np.hstack((Z1, Z2)))
# AUC
auc = metrics.auc(fpr, tpr)
return fpr, tpr, auc
def auc(Z1, Z2):
fpr_arc, tpr_arc, auc = get_auc(Z1, Z2)
# ROC曲線をプロット
plt.plot(fpr_arc, tpr_arc, label='AdaCos(AUC = %.2f)'%auc)
plt.legend()
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.grid(True)
plt.show()
return auc
```
#AdaCos
```
from keras import backend as K
from keras.engine.topology import Layer
import tensorflow as tf
from keras.applications import MobileNetV2
from keras.layers import Input, GlobalAveragePooling2D, Activation
from keras.models import Model
from keras.optimizers import Adam
from sklearn import metrics
# adacosの層
class Adacoslayer(Layer):
# s:softmaxの温度パラメータ, m:margin
def __init__(self, output_dim, s=30, m=0.50, easy_margin=False):
self.output_dim = output_dim
self.s = s
self.m = m
self.easy_margin = easy_margin
super(Adacoslayer, self).__init__()
# 重みの作成
def build(self, input_shape):
# Create a trainable weight variable for this layer.
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[0][1], self.output_dim),
initializer='uniform',
trainable=True)
super(Adacoslayer, self).build(input_shape)
# mainの処理
def call(self, x):
y = x[1]
x_normalize = tf.math.l2_normalize(x[0]) # x = x'/ ||x'||2
k_normalize = tf.math.l2_normalize(self.kernel) # Wj = Wj' / ||Wj'||2
cos_m = K.cos(self.m)
sin_m = K.sin(self.m)
th = K.cos(np.pi - self.m)
mm = K.sin(np.pi - self.m) * self.m
cosine = K.dot(x_normalize, k_normalize) # W.Txの内積
sine = K.sqrt(1.0 - K.square(cosine))
phi = cosine * cos_m - sine * sin_m #cos(θ+m)の加法定理
if self.easy_margin:
phi = tf.where(cosine > 0, phi, cosine)
else:
phi = tf.where(cosine > th, phi, cosine - mm)
# 正解クラス:cos(θ+m) 他のクラス:cosθ
output = (y * phi) + ((1.0 - y) * cosine)
output *= self.s
return output
def compute_output_shape(self, input_shape):
return (input_shape[0][0], self.output_dim) #入力[x,y]のためx[0]はinput_shape[0][0]
# adacosとmobilenetV2を接合して学習
def train_adacos(x, y, classes):
print("Adacos training...")
base_model=MobileNetV2(input_shape=x.shape[1:],alpha=0.5,
weights='imagenet',
include_top=False)
#add new layers
c = base_model.output
yinput = Input(shape=(classes,)) #Adacosで使用
# stock hidden model
hidden = GlobalAveragePooling2D()(c)
c = Adacoslayer(classes, np.sqrt(2)*np.log(classes-1), 0.0)([hidden,yinput]) #outputをクラス数と同じ数に
prediction = Activation('softmax')(c)
model = Model(inputs=[base_model.input, yinput], outputs=prediction)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=0.0001, amsgrad=True),
metrics=['accuracy'])
#学習
hist = model.fit([x, y], y, batch_size=128, epochs=10, verbose = False)
#plt.figure()
#plt.plot(hist.history['acc'],label="train_acc")
#plt.legend(loc="lower right")
#plt.show()
return model
def get_score_arc(model, train, test):
"""
テスト用
model: 特徴抽出用モデル(predict)
"""
model = Model(model.get_layer(index=0).input, model.get_layer(index=-4).output) # Adacosを削除
# 正常vectorの呼び出し
hold_vector = model.predict(train)# shape(len(train), 1280)
# test_vectorの呼び出し
predict_vector = model.predict(test)# shape(len(test), 1280)
score = []
for i in range(len(predict_vector)):
cos_similarity = cosine_similarity(predict_vector[i], hold_vector) # shape(len(test), len(train))
score.append(np.max(cos_similarity))
return np.array(score)
# コサイン類似度の計算
def cosine_similarity(x1, x2):
if x1.ndim == 1:
x1 = x1[np.newaxis]
if x2.ndim == 1:
x2 = x2[np.newaxis]
x1_norm = np.linalg.norm(x1, axis=1)
x2_norm = np.linalg.norm(x2, axis=1)
cosine_sim = np.dot(x1, x2.T)/(x1_norm*x2_norm+1e-10)
return cosine_sim
```
#Experiment(fashion-MNIST)
```
DATA = data(8000)
result = []
for i in range(10):
print(i+1,"/10")
x_train_normal, x_ref, y_ref, x_test_normal, x_test_anomaly = DATA.get_fashion_data()
normal_label = np.zeros((len(x_train_normal), y_ref.shape[1]))
normal_label[:,6] = 1
#AdaCos
model = train_adacos(np.vstack((x_train_normal, x_ref)), np.vstack((normal_label, y_ref)), y_ref.shape[1])
# 異常スコアの算出(異常が高いほど、スコアが高くなるように符号反転)
Z1_arc = -get_score_arc(model, x_train_normal, x_test_normal)
Z2_arc = -get_score_arc(model, x_train_normal, x_test_anomaly)
#auc
result.append(auc(Z1_arc, Z2_arc))
print(result)
```
#Experiment(cifar-10)
```
DATA = data(8000)
result = []
for i in range(10):
print(i+1,"/10")
x_train_normal, x_ref, y_ref, x_test_normal, x_test_anomaly = DATA.get_cifar_data()
normal_label = np.zeros((len(x_train_normal), y_ref.shape[1]))
normal_label[:,3] = 1
# Adacos
model = train_adacos(np.vstack((x_train_normal, x_ref)), np.vstack((normal_label, y_ref)), y_ref.shape[1])
# 異常スコアの算出(異常が高いほど、スコアが高くなるように符号反転)
Z1_arc = -get_score_arc(model, x_train_normal, x_test_normal)
Z2_arc = -get_score_arc(model, x_train_normal, x_test_anomaly)
#auc
result.append(auc(Z1_arc, Z2_arc))
print(result)
```
# graph
```
import seaborn as sns
import matplotlib.pyplot as plt
x = [[0.92, 0.94, 0.91, 0.96, 0.91, 0.91, 0.92, 0.93, 0.91, 0.88],
[0.93, 0.94, 0.94, 0.92, 0.94, 0.95, 0.94, 0.91, 0.95, 0.92],
[0.95, 0.93, 0.95, 0.90, 0.89, 0.90, 0.92, 0.95, 0.95, 0.95]]
sns.set_context("notebook",font_scale=2)
sns.boxplot(x=[x[0],x[1],x[2]], y=["L2-SoftmaxLoss","ArcFace","AdaCos"])
plt.title("AUC\n(Fashion MNIST - sneaker vs boots)")
plt.xlim(0.84,1)
plt.show()
x = [[0.83, 0.84, 0.86, 0.81, 0.84, 0.82, 0.84, 0.87, 0.79, 0.83],
[0.77, 0.8, 0.83, 0.83, 0.8, 0.82, 0.83, 0.81, 0.76, 0.8],
[0.83, 0.80, 0.81, 0.80, 0.80, 0.79, 0.82, 0.83, 0.78, 0.79]]
sns.set_context("notebook",font_scale=2)
sns.boxplot(x=[x[0],x[1],x[2]], y=["L2-SoftmaxLoss","ArcFace","AdaCos"])
plt.title("AUC\n(cifar-10 - deer vs horse)")
plt.xlim(0.75,0.9)
plt.show()
```
| github_jupyter |
# IPython Notebook In Depth
Today we are going to dive into some of the interesting features of IPython and the IPython notebook, which are useful for a number of daily tasks in data-intensive science.
We will work through these features "live"; feel free to type along with me as we go!
## Outline
- IPython: command-line vs. notebook
- Input/Output History
- Tab Completion
- Getting help and accessing documentation
- Useful Keyboard Shortcuts
- Magic Commands
- Shell commands
- Interactivity with ``ipywidgets``
## IPython Command Line and Notebook
### Launching the IPython Shell
If you have installed IPython correctly, you should be able to type ``ipython`` in your command prompt and see something like this:
```
IPython 4.0.1 -- An enhanced Interactive Python.
? -> Introduction and overview of IPython's features.
%quickref -> Quick reference.
help -> Python's own help system.
object? -> Details about 'object', use 'object??' for extra details.
In [1]:
```
With that, you're ready to follow along.
### Launching the IPython Notebook
The IPython notebook is a browser-based graphical interface to the IPython shell, and builds on it a rich set of dynamic display capabilities.
As well as executing Python/IPython statements, the notebook allows the user to include formatted text, static and dynamic visualizations, mathematical equations, javascript widgets, and much more.
Furthermore, these documents can be saved in a way that lets other people open them and execute the code on their own systems.
Though the IPython notebook is viewed and edited through your web browser window, it must connect to a running Python process in order to execute code.
This process (known as a "kernel") can be started by running the following command in your system shell:
```
$ ipython notebook
```
This command will launch a local web server which will be visible to your browser.
It immediately spits out a log showing what it is doing; that log will look something like this:
```
$ ipython notebook
[NotebookApp] Using existing profile dir: '/home/jake/.ipython/profile_default'
[NotebookApp] Serving notebooks from local directory: /home/jake/notebooks/
[NotebookApp] The IPython Notebook is running at: http://localhost:8888/
[NotebookApp] Use Control-C to stop this server and shut down all kernels (twice to skip confirmation)
```
At the command, your default browser should automatically open and navigate to the listed local URL;
the exact address will depend on your system.
If the browser does not open automatically, you can open a window and copy this address (here ``http://localhost:8888/``) manually.
## Input/Output History
A useful feature of IPython is the storage of input and output history.
### Terminal Only
There are a few useful shortcuts that can be used only in the IPython terminal.
We will demonstrate the following in the terminal:
- up arrow for history
- partial completion with up arrow
- reverse search with ctrl-r
### Terminal and Notebook
- Previous results can be obtained using underscores: number of underscores is the number of the previous command:
```
1 + 1
_ * 100
_ + __
_ + __ ** ___
```
Beyond three underscores, you can use underscore followed by a number to indicate the result of a particular line number:
```
_3
```
More useful often is the ``Out`` array, which stores all previous results:
```
Out[3]
Out
```
Similarly, you can access the ``In`` array to see the code history:
```
In[2]
In
```
To see all history at once, use the ``%history`` magic command (more on magic commands below):
```
%history
```
## Tab Completion
The feature of IPython that I use the most frequently is perhaps the tab completion functionality.
Tab completion works for finishing built-in commands:
```python
In [20]: import matp<TAB>
```
will be completed to
```python
In [20]: import matplotlib
```
It works for variables that you have defined:
```
my_variable = 4
```
```python
In [22]: my<TAB>
```
will be completed to
```python
In [22]: my_variable
```
This also works for strings which represent filenames, or pandas columns, etc.
It works for importing packages:
```python
In [25]: import num<TAB>
```
will be completed to
```python
In [25]: import numpy
```
It works for finding attributes of packages and other objects:
```python
In [25]: numpy.ran<TAB>
```
will be completed to
```python
In [25]: numpy.random
```
## Accessing Help and Documentation
After tab completion, I think the next most useful feature of the notebook is the help functionality.
One question mark after any valid object gives you access to its documentation string:
```
numpy.random?
def myfunc(x):
return x ** 2
myfunc?
```
Two question marks gives you access to its source code (if the object is implemented in Python):
```
myfunc??
```
In addition, you can use a single question mark with asterisks to do a wildcard match:
```
numpy.*exp*?
```
If you are curious about the call signature for a funciton, you can type ``shift tab`` within the open-closed parentheses to see its argument list:
Hitting ``shift tab`` multiple times will give you progressively more information about the function:
Using a combination of these, you can quickly remind yourself of how to use various funcitons without ever leaving the terminal/notebook.
## Useful Keyboard Shortcuts
One of the keys to working effectively with IPython is learning your way around the keyboard.
Note: some of the shortcuts below will only work on Linux and Mac; many will work on windows as well
### Terminal shortcuts
If you are familiar with emacs, vim, and similar tools, many of the terminal-based keyboard shortcuts will feel very familiar to you:
#### Navigation
| Keystroke | Action |
|-------------------------------|--------------------------------------------|
| ``Ctrl-a`` | Move cursor to the beginning of the line |
| ``Ctrl-e`` | Move cursor to the end of the line |
| ``Ctrl-b`` or ``left-arrow`` | Move cursor back one character |
| ``Ctrl-f`` or ``right-arrow`` | Move cursor forward one character |
#### Text Entry
| Keystroke | Action |
|-------------------------------|-------------------------------------------------|
| ``backspace`` | Delete previous character in line |
| ``Ctrl-d`` | Delete next character in line |
| ``Ctrl-k`` | Cut text from cursor to end of line |
| ``Ctrl-u`` | Cut all text in line |
| ``Ctrl-y`` | Yank (i.e. Paste) text which was previously cut |
| ``Ctrl-t`` | Transpose (i.e. switch) previous two characters |
#### Command History
| Keystroke | Action |
|-------------------------------|--------------------------------------------|
| ``Ctrl-p`` or ``up-arrow`` | Access previous command in history |
| ``Ctrl-n`` or ``down-arrow`` | Access next command in history |
| ``Ctrl-r`` | Reverse-search through command history |
#### Miscellaneous
| Keystroke | Action |
|-------------------------------|--------------------------------------------|
| ``Ctrl-l`` | Clear terminal screen |
| ``Ctrl-c`` | Interrupt current Python command |
| ``Ctrl-d`` | Exit IPython session |
### Notebook Shortcuts
Depending on your operating system and browser, many of the navigation and text-entry shortcuts will work in the notebook as well. In addition, the notebook has many of its own shortcuts.
First, though, we must mention that the notebook has two "modes" of operation: command mode and edit mode.
- In **command mode**, you are doing operations that affect entire cells. You can enable command mode by pressing the escape key (or pressing ``ctrl m``). For example, in command mode, the up and down arrows will navigate from cell to cell.
- In **edit mode**, you can do operations that affect the contents of a single cell. You can enable edit mode by pressing enter from the command mode. For example, in edit mode, the up and down arrows will navigate lines within the cell
To get a listing of all available shortcuts, enter command mode and press "h"
## Magic Commands
IPython extends the functionality of Python with so-called "magic" commands: these are marked with a ``%`` sign.
We saw one of these above; the ``%history`` command.
Magic commands come in two flavors: *line magics* start with one percent sign, and *cell magics* start with two percent signs.
We'll go through a few examples of magic commands here, but first, using what you've seen above, how do you think you might get a list of all available magic commands? How do you think you might get help on any particular command?
```
%timeit?
```
### Profiling with ``timeit``
For example, here's the ``%timeit``/``%%timeit`` magic, which can be very useful for quick profiling of your code:
```
import numpy as np
x = np.random.rand(1000000)
%timeit x.sum()
L = list(x)
%timeit sum(L)
%%timeit
y = x + 1
z = y ** 2
q = z.sum()
```
### Interpreter: ``paste`` and ``cpaste``
Try pasting this Python code into your IPython interpreter:
```python
>>> def donothing(x):
return x
```
You'll likely get an error.
Now try typing ``%paste`` in your interpreter: what happens?
Next try typing ``%cpaste`` and then use cmd-v to paste your test: what happens?
### Creating a file with ``%%file``
Sometimes it's useful to create a file programatically from within the notebook
```
%%file myscript.py
def foo(x):
return x ** 2
z = foo(12)
print(foo(14))
```
### Running a script with ``%run``
```
%run myscript.py
z
foo(2)
```
### Controlling figures: ``%matplotlib``
You can use the ``%matplotlib`` function to specify the matplotlib *backend* you would like to use.
For example:
- ``%matplotlib`` by itself uses the default system backend
- ``%matplotlib inline`` creates inline, static figures (great for publication and/or sharing)
- ``%matplotlib notebook`` creates inline, interactive figures (though in my experience it can be a bit unstable)
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.plot(np.random.rand(100));
```
### Help functions and more info
- The ``%magic`` function will tell you all about magic commands
- The ``%lsmagic`` function will list all available magic commands
- Remember that the ``?`` can be used to get documentation!
- Though we won't cover it here, it is possible to [create and activate your own magic commands](https://ipython.org/ipython-doc/stable/config/custommagics.html)
```
%lsmagic
%magic
%debug?
```
## Shell Commands
IPython is meant to be an all-purpose scientific computing environment, and access to the shell is critical.
Any command that starts with an exclamation point will be passed to the shell.
Note that because windows has a different kind of shell than Linux/OSX, shell commands will be different from operating system to operating system.
All the commands you have learned previously will work here:
```
!ls
!pwd
```
You can even seamlessly pass values to and from the Python interpreter.
For example, we can store the result of a directory listing:
```
contents = !ls
contents
```
We can inject Python variables into a shell command with ``{}``:
```
!cat {contents[4]}
for filename in contents:
if filename.endswith('.py'):
print(filename)
!head -10 {filename}
```
With these tools, you should never have to switch from IPython to a terminal to run a command.
## IPython Widgets
One incredibly useful feature of the notebook is the interactivity provided by the [``ipywidgets`` package](https://github.com/ipython/ipywidgets). You'll have to install this using, e.g.
$ conda install ipywidgets
You can find a full set of documentation notebooks [here](https://github.com/ipython/ipywidgets/blob/master/examples/notebooks/Index.ipynb).
We're going to walk through a quick demonstration of the functionality in [WidgetsDemo.ipynb](WidgetsDemo.ipynb)
| github_jupyter |
# Mining Input Grammars
So far, the grammars we have seen have been mostly specified manually – that is, you (or the person knowing the input format) had to design and write a grammar in the first place. While the grammars we have seen so far have been rather simple, creating a grammar for complex inputs can involve quite some effort. In this chapter, we therefore introduce techniques that _automatically mine grammars from programs_ – by executing the programs and observing how they process which parts of the input. In conjunction with a grammar fuzzer, this allows us to
1. take a program,
2. extract its input grammar, and
3. fuzz it with high efficiency and effectiveness, using the concepts in this book.
```
from bookutils import YouTubeVideo
YouTubeVideo("KsqwszWnAmM")
```
**Prerequisites**
* You should have read the [chapter on grammars](Grammars.ipynb).
* The [chapter on configuration fuzzing](ConfigurationFuzzer.ipynb) introduces grammar mining for configuration options, as well as observing variables and values during execution.
* We use the tracer from the [chapter on coverage](Coverage.ipynb).
* The concept of parsing from the [chapter on parsers](Parser.ipynb) is also useful.
## Synopsis
<!-- Automatically generated. Do not edit. -->
To [use the code provided in this chapter](Importing.ipynb), write
```python
>>> from fuzzingbook.GrammarMiner import <identifier>
```
and then make use of the following features.
This chapter provides a number of classes to mine input grammars from existing programs. The function `recover_grammar()` could be the easiest to use. It takes a function and a set of inputs, and returns a grammar that describes its input language.
We apply `recover_grammar()` on a `url_parse()` function that takes and decomposes URLs:
```python
>>> url_parse('https://www.fuzzingbook.org/')
>>> URLS
['http://user:pass@www.google.com:80/?q=path#ref',
'https://www.cispa.saarland:80/',
'http://www.fuzzingbook.org/#News']
```
We extract the input grammar for `url_parse()` using `recover_grammar()`:
```python
>>> grammar = recover_grammar(url_parse, URLS, files=['urllib/parse.py'])
>>> grammar
{'<start>': ['<urlsplit@437:url>'],
'<urlsplit@437:url>': ['<urlparse@394:scheme>:<_splitnetloc@411:url>'],
'<urlparse@394:scheme>': ['https', 'http'],
'<_splitnetloc@411:url>': ['//<urlparse@394:netloc>/',
'//<urlparse@394:netloc><urlsplit@481:url>'],
'<urlparse@394:netloc>': ['www.cispa.saarland:80',
'user:pass@www.google.com:80',
'www.fuzzingbook.org'],
'<urlsplit@481:url>': ['/#<urlparse@394:fragment>',
'<urlsplit@486:url>#<urlparse@394:fragment>'],
'<urlsplit@486:url>': ['/?<urlparse@394:query>'],
'<urlparse@394:query>': ['q=path'],
'<urlparse@394:fragment>': ['News', 'ref']}
```
The names of nonterminals are a bit technical; but the grammar nicely represents the structure of the input; for instance, the different schemes (`"http"`, `"https"`) are all identified:
```python
>>> syntax_diagram(grammar)
start
```

```
urlsplit@437:url
```

```
urlparse@394:scheme
```

```
_splitnetloc@411:url
```

```
urlparse@394:netloc
```

```
urlsplit@481:url
```

```
urlsplit@486:url
```

```
urlparse@394:query
```

```
urlparse@394:fragment
```

The grammar can be immediately used for fuzzing, producing arbitrary combinations of input elements, which are all syntactically valid.
```python
>>> from GrammarCoverageFuzzer import GrammarCoverageFuzzer
>>> fuzzer = GrammarCoverageFuzzer(grammar)
>>> [fuzzer.fuzz() for i in range(5)]
['http://www.cispa.saarland:80/?q=path#ref',
'https://www.fuzzingbook.org/',
'http://user:pass@www.google.com:80/#News',
'http://user:pass@www.google.com:80/#ref',
'https://www.cispa.saarland:80/#News']
```
Being able to automatically extract a grammar and to use this grammar for fuzzing makes for very effective test generation with a minimum of manual work.
## A Grammar Challenge
Consider the `process_inventory()` method from the [chapter on parsers](Parser.ipynb):
```
import bookutils
from typing import List, Tuple, Callable, Any
from collections.abc import Iterable
from Parser import process_inventory, process_vehicle, process_car, process_van, lr_graph # minor dependency
```
It takes inputs of the following form.
```
INVENTORY = """\
1997,van,Ford,E350
2000,car,Mercury,Cougar
1999,car,Chevy,Venture\
"""
print(process_inventory(INVENTORY))
```
We found from the [chapter on parsers](Parser.ipynb) that coarse grammars do not work well for fuzzing when the input format includes details expressed only in code. That is, even though we have the formal specification of CSV files ([RFC 4180](https://tools.ietf.org/html/rfc4180)), the inventory system includes further rules as to what is expected at each index of the CSV file. The solution of simply recombining existing inputs, while practical, is incomplete. In particular, it relies on a formal input specification being available in the first place. However, we have no assurance that the program obeys the input specification given.
One of the ways out of this predicament is to interrogate the program under test as to what its input specification is. That is, if the program under test is written in a style such that specific methods are responsible for handling specific parts of the input, one can recover the parse tree by observing the process of parsing. Further, one can recover a reasonable approximation of the grammar by abstraction from multiple input trees.
_We start with the assumption (1) that the program is written in such a fashion that specific methods are responsible for parsing specific fragments of the program -- This includes almost all ad hoc parsers._
The idea is as follows:
* Hook into the Python execution and observe the fragments of input string as they are produced and named in different methods.
* Stitch the input fragments together in a tree structure to retrieve the **Parse Tree**.
* Abstract common elements from multiple parse trees to produce the **Context Free Grammar** of the input.
## A Simple Grammar Miner
Say we want to obtain the input grammar for the function `process_vehicle()`. We first collect the sample inputs for this function.
```
VEHICLES = INVENTORY.split('\n')
```
The set of methods responsible for processing inventory are the following.
```
INVENTORY_METHODS = {
'process_inventory',
'process_vehicle',
'process_van',
'process_car'}
```
We have seen from the chapter on [configuration fuzzing](ConfigurationFuzzer.ipynb) that one can hook into the Python runtime to observe the arguments to a function and any local variables created. We have also seen that one can obtain the context of execution by inspecting the `frame` argument. Here is a simple tracer that can return the local variables and other contextual information in a traced function. We reuse the `Coverage` tracing class.
### Tracer
```
from Coverage import Coverage
import inspect
class Tracer(Coverage):
def traceit(self, frame, event, arg):
method_name = inspect.getframeinfo(frame).function
if method_name not in INVENTORY_METHODS:
return
file_name = inspect.getframeinfo(frame).filename
param_names = inspect.getargvalues(frame).args
lineno = inspect.getframeinfo(frame).lineno
local_vars = inspect.getargvalues(frame).locals
print(event, file_name, lineno, method_name, param_names, local_vars)
return self.traceit
```
We run the code under trace context.
```
with Tracer() as tracer:
process_vehicle(VEHICLES[0])
```
The main thing that we want out of tracing is a list of assignments of input fragments to different variables. We can use the tracing facility `settrace()` to get that as we showed above.
However, the `settrace()` function hooks into the Python debugging facility. When it is in operation, no debugger can hook into the program. That is, if there is a problem with our grammar miner, we will not be able to attach a debugger to it to understand what is happening. This is not ideal. Hence, we limit the tracer to the simplest implementation possible, and implement the core of grammar mining in later stages.
The `traceit()` function relies on information from the `frame` variable which exposes Python internals. We define a `context` class that encapsulates the information that we need from the `frame`.
### Context
The `Context` class provides easy access to the information such as the current module, and parameter names.
```
class Context:
def __init__(self, frame, track_caller=True):
self.method = inspect.getframeinfo(frame).function
self.parameter_names = inspect.getargvalues(frame).args
self.file_name = inspect.getframeinfo(frame).filename
self.line_no = inspect.getframeinfo(frame).lineno
def _t(self):
return (self.file_name, self.line_no, self.method,
','.join(self.parameter_names))
def __repr__(self):
return "%s:%d:%s(%s)" % self._t()
```
Here we add a few convenience methods that operate on the `frame` to `Context`.
```
class Context(Context):
def extract_vars(self, frame):
return inspect.getargvalues(frame).locals
def parameters(self, all_vars):
return {k: v for k, v in all_vars.items() if k in self.parameter_names}
def qualified(self, all_vars):
return {"%s:%s" % (self.method, k): v for k, v in all_vars.items()}
```
We hook printing the context to our `traceit()` to see it in action. First we define a `log_event()` for displaying events.
```
def log_event(event, var):
print({'call': '->', 'return': '<-'}.get(event, ' '), var)
```
And use the `log_event()` in the `traceit()` function.
```
class Tracer(Tracer):
def traceit(self, frame, event, arg):
log_event(event, Context(frame))
return self.traceit
```
Running `process_vehicle()` under trace prints the contexts encountered.
```
with Tracer() as tracer:
process_vehicle(VEHICLES[0])
```
The trace produced by executing any function can get overwhelmingly large. Hence, we need to restrict our attention to specific modules. Further, we also restrict our attention exclusively to `str` variables since these variables are more likely to contain input fragments. (We will show how to deal with complex objects later in exercises.)
The `Context` class we developed earlier is used to decide which modules to monitor, and which variables to trace.
We store the current *input string* so that it can be used to determine if any particular string fragments came from the current input string. Any optional arguments are processed separately.
```
class Tracer(Tracer):
def __init__(self, my_input, **kwargs):
self.options(kwargs)
self.my_input, self.trace = my_input, []
```
We use an optional argument `files` to indicate the specific source files we are interested in, and `methods` to indicate which specific methods are of interest. Further, we also use `log` to specify whether verbose logging should be enabled during trace. We use the `log_event()` method we defined earlier for logging.
The options processing is as below.
```
class Tracer(Tracer):
def options(self, kwargs):
self.files = kwargs.get('files', [])
self.methods = kwargs.get('methods', [])
self.log = log_event if kwargs.get('log') else lambda _evt, _var: None
```
The `files` and `methods` are checked to determine, if a particular event should be traced or not
```
class Tracer(Tracer):
def tracing_context(self, cxt, event, arg):
fres = not self.files or any(
cxt.file_name.endswith(f) for f in self.files)
mres = not self.methods or any(cxt.method == m for m in self.methods)
return fres and mres
```
Similar to the context of events, we also want to restrict our attention to specific variables. For now, we want to focus only on strings. (See the Exercises at the end of the chapter on how to extend it to other kinds of objects).
```
class Tracer(Tracer):
def tracing_var(self, k, v):
return isinstance(v, str)
```
We modify the `traceit()` to call an `on_event()` function with the context information only on the specific events we are interested in.
```
class Tracer(Tracer):
def on_event(self, event, arg, cxt, my_vars):
self.trace.append((event, arg, cxt, my_vars))
def create_context(self, frame):
return Context(frame)
def traceit(self, frame, event, arg):
cxt = self.create_context(frame)
if not self.tracing_context(cxt, event, arg):
return self.traceit
self.log(event, cxt)
my_vars = {
k: v
for k, v in cxt.extract_vars(frame).items()
if self.tracing_var(k, v)
}
self.on_event(event, arg, cxt, my_vars)
return self.traceit
```
The `Tracer` class can now focus on specific kinds of events on specific files. Further, it provides a first level filter for variables that we find interesting. For example, we want to focus specifically on variables from `process_*` methods that contain input fragments. Here is how our updated `Tracer` can be used.
```
with Tracer(VEHICLES[0], methods=INVENTORY_METHODS, log=True) as tracer:
process_vehicle(VEHICLES[0])
```
The execution produced the following trace.
```
for t in tracer.trace:
print(t[0], t[2].method, dict(t[3]))
```
Since we are saving the input already in `Tracer`, it is redundant to specify it separately again as an argument.
```
with Tracer(VEHICLES[0], methods=INVENTORY_METHODS, log=True) as tracer:
process_vehicle(tracer.my_input)
```
### DefineTracker
We define a `DefineTracker` class that processes the trace from the `Tracer`. The idea is to store different variable definitions which are input fragments.
The tracker identifies string fragments that are part of the input string, and stores them in a dictionary `my_assignments`. It saves the trace, and the corresponding input for processing. Finally it calls `process()` to process the `trace` it was given. We will start with a simple tracker that relies on certain assumptions, and later see how these assumptions can be relaxed.
```
class DefineTracker:
def __init__(self, my_input, trace, **kwargs):
self.options(kwargs)
self.my_input = my_input
self.trace = trace
self.my_assignments = {}
self.process()
```
One of the problems of using substring search is that short string sequences tend to be included in other string sequences even though they may not have come from the original string. That is, say the input fragment is `v`, it could have equally come from either `van` or `chevy`. We rely on being able to predict the exact place in the input where a given fragment occurred. Hence, we define a constant `FRAGMENT_LEN` such that we ignore strings up to that length. We also incorporate a logging facility as before.
```
FRAGMENT_LEN = 3
class DefineTracker(DefineTracker):
def options(self, kwargs):
self.log = log_event if kwargs.get('log') else lambda _evt, _var: None
self.fragment_len = kwargs.get('fragment_len', FRAGMENT_LEN)
```
Our tracer simply records the variable values as they occur. We next need to check if the variables contain values from the **input string**. Common ways to do this is to rely on symbolic execution or at least dynamic tainting, which are powerful, but also complex. However, one can obtain a reasonable approximation by simply relying on substring search. That is, we consider any value produced that is a substring of the original input string to have come from the original input.
We define an `is_input_fragment()` method that relies on string inclusion to detect if the string came from the input.
```
class DefineTracker(DefineTracker):
def is_input_fragment(self, var, value):
return len(value) >= self.fragment_len and value in self.my_input
```
We can use `is_input_fragment()` to select only a subset of variables defined, as implemented below in `fragments()`.
```
class DefineTracker(DefineTracker):
def fragments(self, variables):
return {k: v for k, v in variables.items(
) if self.is_input_fragment(k, v)}
```
The tracker processes each event, and at each event, it updates the dictionary `my_assignments` with the current local variables that contain strings that are part of the input. Note that there is a choice here with respect to what happens during reassignment. We can either discard all the reassignments, or keep only the last assignment. Here, we choose the latter. If you want the former behavior, check whether the value exists in `my_assignments` before storing a fragment.
```
class DefineTracker(DefineTracker):
def track_event(self, event, arg, cxt, my_vars):
self.log(event, (cxt.method, my_vars))
self.my_assignments.update(self.fragments(my_vars))
def process(self):
for event, arg, cxt, my_vars in self.trace:
self.track_event(event, arg, cxt, my_vars)
```
Using the tracker, we can obtain the input fragments. For example, say we are only interested in strings that are at least `5` characters long.
```
tracker = DefineTracker(tracer.my_input, tracer.trace, fragment_len=5)
for k, v in tracker.my_assignments.items():
print(k, '=', repr(v))
```
Or strings that are `2` characters long (the default).
```
tracker = DefineTracker(tracer.my_input, tracer.trace)
for k, v in tracker.my_assignments.items():
print(k, '=', repr(v))
class DefineTracker(DefineTracker):
def assignments(self):
return self.my_assignments.items()
```
### Assembling a Derivation Tree
```
from Grammars import START_SYMBOL, syntax_diagram, \
is_nonterminal, Grammar
from GrammarFuzzer import GrammarFuzzer, display_tree, \
DerivationTree
```
The input fragments from the `DefineTracker` only tell half the story. The fragments may be created at different stages of parsing. Hence, we need to assemble the fragments to a derivation tree of the input. The basic idea is as follows:
Our input from the previous step was:
```python
"1997,van,Ford,E350"
```
We start a derivation tree, and associate it with the start symbol in the grammar.
```
derivation_tree: DerivationTree = (START_SYMBOL, [("1997,van,Ford,E350", [])])
display_tree(derivation_tree)
```
The next input was:
```python
vehicle = "1997,van,Ford,E350"
```
Since vehicle covers the `<start>` node's value completely, we replace the value with the vehicle node.
```
derivation_tree: DerivationTree = (START_SYMBOL,
[('<vehicle>', [("1997,van,Ford,E350", [])],
[])])
display_tree(derivation_tree)
```
The next input was:
```python
year = '1997'
```
Traversing the derivation tree from `<start>`, we see that it replaces a portion of the `<vehicle>` node's value. Hence we split the `<vehicle>` node's value to two children, where one corresponds to the value `"1997"` and the other to `",van,Ford,E350"`, and replace the first one with the node `<year>`.
```
derivation_tree: DerivationTree = (START_SYMBOL,
[('<vehicle>', [('<year>', [('1997', [])]),
(",van,Ford,E350", [])], [])])
display_tree(derivation_tree)
```
We perform similar operations for
```python
company = 'Ford'
```
```
derivation_tree: DerivationTree = (START_SYMBOL,
[('<vehicle>', [('<year>', [('1997', [])]),
(",van,", []),
('<company>', [('Ford', [])]),
(",E350", [])], [])])
display_tree(derivation_tree)
```
Similarly for
```python
kind = 'van'
```
and
```python
model = 'E350'
```
```
derivation_tree: DerivationTree = (START_SYMBOL,
[('<vehicle>', [('<year>', [('1997', [])]),
(",", []),
("<kind>", [('van', [])]),
(",", []),
('<company>', [('Ford', [])]),
(",", []),
("<model>", [('E350', [])])
], [])])
display_tree(derivation_tree)
```
We now develop the complete algorithm with the above described steps.
The derivation tree `TreeMiner` is initialized with the input string, and the variable assignments, and it converts the assignments to the corresponding derivation tree.
```
class TreeMiner:
def __init__(self, my_input, my_assignments, **kwargs):
self.options(kwargs)
self.my_input = my_input
self.my_assignments = my_assignments
self.tree = self.get_derivation_tree()
def options(self, kwargs):
self.log = log_call if kwargs.get('log') else lambda _i, _v: None
def get_derivation_tree(self):
return (START_SYMBOL, [])
```
The `log_call()` is as follows.
```
def log_call(indent, var):
print('\t' * indent, var)
```
The basic idea is as follows:
* **For now, we assume that the value assigned to a variable is stable. That is, it is never reassigned. In particular, there are no recursive calls, or multiple calls to the same function from different parts.** (We will show how to overcome this limitation later).
* For each pair _var_, _value_ found in `my_assignments`:
1. We search for occurrences of _value_ `val` in the derivation tree recursively.
2. If an occurrence was found as a value `V1` of a node `P1`, we partition the value of the node `P1` into three parts, with the central part matching the _value_ `val`, and the first and last part, the corresponding prefix and suffix in `V1`.
3. Reconstitute the node `P1` with three children, where prefix and suffix mentioned earlier are string values, and the matching value `val` is replaced by a node `var` with a single value `val`.
First, we define a wrapper to generate a nonterminal from a variable name.
```
def to_nonterminal(var):
return "<" + var.lower() + ">"
```
The `string_part_of_value()` method checks whether the given `part` value was part of the whole.
```
class TreeMiner(TreeMiner):
def string_part_of_value(self, part, value):
return (part in value)
```
The `partition_by_part()` splits the `value` by the given part if it matches, and returns a list containing the first part, the part that was replaced, and the last part. This is a format that can be used as a part of the list of children.
```
class TreeMiner(TreeMiner):
def partition(self, part, value):
return value.partition(part)
class TreeMiner(TreeMiner):
def partition_by_part(self, pair, value):
k, part = pair
prefix_k_suffix = [
(k, [[part, []]]) if i == 1 else (e, [])
for i, e in enumerate(self.partition(part, value))
if e]
return prefix_k_suffix
```
The `insert_into_tree()` method accepts a given tree `tree` and a `(k,v)` pair. It recursively checks whether the given pair can be applied. If the pair can be applied, it applies the pair and returns `True`.
```
class TreeMiner(TreeMiner):
def insert_into_tree(self, my_tree, pair):
var, values = my_tree
k, v = pair
self.log(1, "- Node: %s\t\t? (%s:%s)" % (var, k, repr(v)))
applied = False
for i, value_ in enumerate(values):
value, arr = value_
self.log(2, "-> [%d] %s" % (i, repr(value)))
if is_nonterminal(value):
applied = self.insert_into_tree(value_, pair)
if applied:
break
elif self.string_part_of_value(v, value):
prefix_k_suffix = self.partition_by_part(pair, value)
del values[i]
for j, rep in enumerate(prefix_k_suffix):
values.insert(j + i, rep)
applied = True
self.log(2, " > %s" % (repr([i[0] for i in prefix_k_suffix])))
break
else:
continue
return applied
```
Here is how `insert_into_tree()` is used.
```
tree: DerivationTree = (START_SYMBOL, [("1997,van,Ford,E350", [])])
m = TreeMiner('', {}, log=True)
```
First, we have our input string as the only node.
```
display_tree(tree)
```
Inserting the `<vehicle>` node.
```
v = m.insert_into_tree(tree, ('<vehicle>', "1997,van,Ford,E350"))
display_tree(tree)
```
Inserting `<model>` node.
```
v = m.insert_into_tree(tree, ('<model>', 'E350'))
display_tree((tree))
```
Inserting `<company>`.
```
v = m.insert_into_tree(tree, ('<company>', 'Ford'))
display_tree(tree)
```
Inserting `<kind>`.
```
v = m.insert_into_tree(tree, ('<kind>', 'van'))
display_tree(tree)
```
Inserting `<year>`.
```
v = m.insert_into_tree(tree, ('<year>', '1997'))
display_tree(tree)
```
To make life simple, we define a wrapper function `nt_var()` that will convert a token to its corresponding nonterminal symbol.
```
class TreeMiner(TreeMiner):
def nt_var(self, var):
return var if is_nonterminal(var) else to_nonterminal(var)
```
Now, we need to apply a new definition to an entire grammar.
```
class TreeMiner(TreeMiner):
def apply_new_definition(self, tree, var, value):
nt_var = self.nt_var(var)
return self.insert_into_tree(tree, (nt_var, value))
```
This algorithm is implemented as `get_derivation_tree()`.
```
class TreeMiner(TreeMiner):
def get_derivation_tree(self):
tree = (START_SYMBOL, [(self.my_input, [])])
for var, value in self.my_assignments:
self.log(0, "%s=%s" % (var, repr(value)))
self.apply_new_definition(tree, var, value)
return tree
```
The `TreeMiner` is used as follows:
```
with Tracer(VEHICLES[0]) as tracer:
process_vehicle(tracer.my_input)
assignments = DefineTracker(tracer.my_input, tracer.trace).assignments()
dt = TreeMiner(tracer.my_input, assignments, log=True)
dt.tree
```
The obtained derivation tree is as below.
```
display_tree(TreeMiner(tracer.my_input, assignments).tree)
```
Combining all the pieces:
```
trees = []
for vehicle in VEHICLES:
print(vehicle)
with Tracer(vehicle) as tracer:
process_vehicle(tracer.my_input)
assignments = DefineTracker(tracer.my_input, tracer.trace).assignments()
trees.append((tracer.my_input, assignments))
for var, val in assignments:
print(var + " = " + repr(val))
print()
```
The corresponding derivation trees are below.
```
csv_dt = []
for inputstr, assignments in trees:
print(inputstr)
dt = TreeMiner(inputstr, assignments)
csv_dt.append(dt)
display_tree(dt.tree)
```
### Recovering Grammars from Derivation Trees
We define a class `Miner` that can combine multiple derivation trees to produce the grammar. The initial grammar is empty.
```
class GrammarMiner:
def __init__(self):
self.grammar = {}
```
The `tree_to_grammar()` method converts our derivation tree to a grammar by picking one node at a time, and adding it to the grammar. The node name becomes the key, and any list of children it has becomes another alternative for that key.
```
class GrammarMiner(GrammarMiner):
def tree_to_grammar(self, tree):
node, children = tree
one_alt = [ck for ck, gc in children]
hsh = {node: [one_alt] if one_alt else []}
for child in children:
if not is_nonterminal(child[0]):
continue
chsh = self.tree_to_grammar(child)
for k in chsh:
if k not in hsh:
hsh[k] = chsh[k]
else:
hsh[k].extend(chsh[k])
return hsh
gm = GrammarMiner()
gm.tree_to_grammar(csv_dt[0].tree)
```
The grammar being generated here is `canonical`. We define a function `readable()` that takes in a canonical grammar and returns it in a readable form.
```
def readable(grammar):
def readable_rule(rule):
return ''.join(rule)
return {k: list(set(readable_rule(a) for a in grammar[k]))
for k in grammar}
syntax_diagram(readable(gm.tree_to_grammar(csv_dt[0].tree)))
```
The `add_tree()` method gets a combined list of non-terminals from current grammar, and the tree to be added to the grammar, and updates the definitions of each non-terminal.
```
import itertools
class GrammarMiner(GrammarMiner):
def add_tree(self, t):
t_grammar = self.tree_to_grammar(t.tree)
self.grammar = {
key: self.grammar.get(key, []) + t_grammar.get(key, [])
for key in itertools.chain(self.grammar.keys(), t_grammar.keys())
}
```
The `add_tree()` is used as follows:
```
inventory_grammar_miner = GrammarMiner()
for dt in csv_dt:
inventory_grammar_miner.add_tree(dt)
syntax_diagram(readable(inventory_grammar_miner.grammar))
```
Given execution traces from various inputs, one can define `update_grammar()` to obtain the complete grammar from the traces.
```
class GrammarMiner(GrammarMiner):
def update_grammar(self, inputstr, trace):
at = self.create_tracker(inputstr, trace)
dt = self.create_tree_miner(inputstr, at.assignments())
self.add_tree(dt)
return self.grammar
def create_tracker(self, *args):
return DefineTracker(*args)
def create_tree_miner(self, *args):
return TreeMiner(*args)
```
The complete grammar recovery is implemented in `recover_grammar()`.
```
def recover_grammar(fn: Callable, inputs: Iterable[str],
**kwargs: Any) -> Grammar:
miner = GrammarMiner()
for inputstr in inputs:
with Tracer(inputstr, **kwargs) as tracer:
fn(tracer.my_input)
miner.update_grammar(tracer.my_input, tracer.trace)
return readable(miner.grammar)
```
Note that the grammar could have been retrieved directly from the tracker, without the intermediate derivation tree stage. However, going through the derivation tree allows one to inspect the inputs being fragmented and verify that it happens correctly.
#### Example 1. Recovering the Inventory Grammar
```
inventory_grammar = recover_grammar(process_vehicle, VEHICLES)
inventory_grammar
```
#### Example 2. Recovering URL Grammar
Our algorithm is robust enough to recover grammar from real world programs. For example, the `urlparse` function in the Python `urlib` module accepts the following sample URLs.
```
URLS = [
'http://user:pass@www.google.com:80/?q=path#ref',
'https://www.cispa.saarland:80/',
'http://www.fuzzingbook.org/#News',
]
```
The urllib caches its intermediate results for faster access. Hence, we need to disable it using `clear_cache()` after every invocation.
```
from urllib.parse import urlparse, clear_cache # type: ignore
```
We use the sample URLs to recover grammar as follows. The `urlparse` function tends to cache its previous parsing results. Hence, we define a new method `url_parse()` that clears the cache before each call.
```
def url_parse(url):
clear_cache()
urlparse(url)
trees = []
for url in URLS:
print(url)
with Tracer(url) as tracer:
url_parse(tracer.my_input)
assignments = DefineTracker(tracer.my_input, tracer.trace).assignments()
trees.append((tracer.my_input, assignments))
for var, val in assignments:
print(var + " = " + repr(val))
print()
url_dt = []
for inputstr, assignments in trees:
print(inputstr)
dt = TreeMiner(inputstr, assignments)
url_dt.append(dt)
display_tree(dt.tree)
```
Let us use `url_parse()` to recover the grammar:
```
url_grammar = recover_grammar(url_parse, URLS, files=['urllib/parse.py'])
syntax_diagram(url_grammar)
```
The recovered grammar describes the URL format reasonably well.
### Fuzzing
We can now use our recovered grammar for fuzzing as follows.
First, the inventory grammar.
```
f = GrammarFuzzer(inventory_grammar)
for _ in range(10):
print(f.fuzz())
```
Next, the URL grammar.
```
f = GrammarFuzzer(url_grammar)
for _ in range(10):
print(f.fuzz())
```
What this means is that we can now take a program and a few samples, extract its grammar, and then use this very grammar for fuzzing. Now that's quite an opportunity!
### Problems with the Simple Miner
One of the problems with our simple grammar miner is the assumption that the values assigned to variables are stable. Unfortunately, that may not hold true in all cases. For example, here is a URL with a slightly different format.
```
URLS_X = URLS + ['ftp://freebsd.org/releases/5.8']
```
The grammar generated from this set of samples is not as nice as what we got earlier
```
url_grammar = recover_grammar(url_parse, URLS_X, files=['urllib/parse.py'])
syntax_diagram(url_grammar)
```
Clearly, something has gone wrong.
To investigate why the `url` definition has gone wrong, let us inspect the trace for the URL.
```
clear_cache()
with Tracer(URLS_X[0]) as tracer:
urlparse(tracer.my_input)
for i, t in enumerate(tracer.trace):
if t[0] in {'call', 'line'} and 'parse.py' in str(t[2]) and t[3]:
print(i, t[2]._t()[1], t[3:])
```
Notice how the value of `url` changes as the parsing progresses? This violates our assumption that the value assigned to a variable is stable. We next look at how this limitation can be removed.
## Grammar Miner with Reassignment
One way to uniquely identify different variables is to annotate them with *line numbers* both when they are defined and also when their value changes. Consider the code fragment below
### Tracking variable assignment locations
```
def C(cp_1):
c_2 = cp_1 + '@2'
c_3 = c_2 + '@3'
return c_3
def B(bp_7):
b_8 = bp_7 + '@8'
return C(b_8)
def A(ap_12):
a_13 = ap_12 + '@13'
a_14 = B(a_13) + '@14'
a_14 = a_14 + '@15'
a_13 = a_14 + '@16'
a_14 = B(a_13) + '@17'
a_14 = B(a_13) + '@18'
```
Notice how all variables are either named corresponding to either where they are defined, or the value is annotated to indicate that it was changed.
Let us run this under the trace.
```
with Tracer('____') as tracer:
A(tracer.my_input)
for t in tracer.trace:
print(t[0], "%d:%s" % (t[2].line_no, t[2].method), t[3])
```
Each variables were referenced first as follows:
* `cp_1` -- *call* `1:C`
* `c_2` -- *line* `3:C` (but the previous event was *line* `2:C`)
* `c_3` -- *line* `4:C` (but the previous event was *line* `3:C`)
* `bp_7` -- *call* `7:B`
* `b_8` -- *line* `9:B` (but the previous event was *line* `8:B`)
* `ap_12` -- *call* `12:A`
* `a_13` -- *line* `14:A` (but the previous event was *line* `13:A`)
* `a_14` -- *line* `15:A` (the previous event was *return* `9:B`. However, the previous event in `A()` was *line* `14:A`)
* reassign `a_14` at *15* -- *line* `16:A` (the previous event was *line* `15:A`)
* reassign `a_13` at *16* -- *line* `17:A` (the previous event was *line* `16:A`)
* reassign `a_14` at *17* -- *return* `17:A` (the previous event in `A()` was *line* `17:A`)
* reassign `a_14` at *18* -- *return* `18:A` (the previous event in `A()` was *line* `18:A`)
So, our observations are that, if it is a call, the current location is the right one for any new variables being defined. On the other hand, if the variable being referenced for the first time (or reassigned a new value), then the right location to consider is the previous location *in the same method invocation*. Next, let us see how we can incorporate this information into variable naming.
Next, we need a way to track the individual method calls as they are being made. For this we define the class `CallStack`. Each method invocation gets a separate identifier, and when the method call is over, the identifier is reset.
### CallStack
```
class CallStack:
def __init__(self, **kwargs):
self.options(kwargs)
self.method_id = (START_SYMBOL, 0)
self.method_register = 0
self.mstack = [self.method_id]
def enter(self, method):
self.method_register += 1
self.method_id = (method, self.method_register)
self.log('call', "%s%s" % (self.indent(), str(self)))
self.mstack.append(self.method_id)
def leave(self):
self.mstack.pop()
self.log('return', "%s%s" % (self.indent(), str(self)))
self.method_id = self.mstack[-1]
```
A few extra functions to make life simpler.
```
class CallStack(CallStack):
def options(self, kwargs):
self.log = log_event if kwargs.get('log') else lambda _evt, _var: None
def indent(self):
return len(self.mstack) * "\t"
def at(self, n):
return self.mstack[n]
def __len__(self):
return len(mstack) - 1
def __str__(self):
return "%s:%d" % self.method_id
def __repr__(self):
return repr(self.method_id)
```
We also define a convenience method to display a given stack.
```
def display_stack(istack):
def stack_to_tree(stack):
current, *rest = stack
if not rest:
return (repr(current), [])
return (repr(current), [stack_to_tree(rest)])
display_tree(stack_to_tree(istack.mstack), graph_attr=lr_graph)
```
Here is how we can use the `CallStack`.
```
cs = CallStack()
display_stack(cs)
cs
cs.enter('hello')
display_stack(cs)
cs
cs.enter('world')
display_stack(cs)
cs
cs.leave()
display_stack(cs)
cs
cs.enter('world')
display_stack(cs)
cs
cs.leave()
display_stack(cs)
cs
```
In order to account for variable reassignments, we need to have a more intelligent data structure than a dictionary for storing variables. We first define a simple interface `Vars`. It acts as a container for variables, and is instantiated at `my_assignments`.
### Vars
The `Vars` stores references to variables as they occur during parsing in its internal dictionary `defs`. We initialize the dictionary with the original string.
```
class Vars:
def __init__(self, original):
self.defs = {}
self.my_input = original
```
The dictionary needs two methods: `update()` that takes a set of key-value pairs to update itself, and `_set_kv()` that updates a particular key-value pair.
```
class Vars(Vars):
def _set_kv(self, k, v):
self.defs[k] = v
def __setitem__(self, k, v):
self._set_kv(k, v)
def update(self, v):
for k, v in v.items():
self._set_kv(k, v)
```
The `Vars` is a proxy for the internal dictionary. For example, here is how one can use it.
```
v = Vars('')
v.defs
v['x'] = 'X'
v.defs
v.update({'x': 'x', 'y': 'y'})
v.defs
```
### AssignmentVars
We now extend the simple `Vars` to account for variable reassignments. For this, we define `AssignmentVars`.
The idea for detecting reassignments and renaming variables is as follows: We keep track of the previous reassignments to particular variables using `accessed_seq_var`. It contains the last rename of any particular variable as its corresponding value. The `new_vars` contains a list of all new variables that were added on this iteration.
```
class AssignmentVars(Vars):
def __init__(self, original):
super().__init__(original)
self.accessed_seq_var = {}
self.var_def_lines = {}
self.current_event = None
self.new_vars = set()
self.method_init()
```
The `method_init()` method takes care of keeping track of method invocations using records saved in the `call_stack`. `event_locations` is for keeping track of the locations accessed *within this method*. This is used for line number tracking of variable definitions.
```
class AssignmentVars(AssignmentVars):
def method_init(self):
self.call_stack = CallStack()
self.event_locations = {self.call_stack.method_id: []}
```
The `update()` is now modified to track the changed line numbers if any, using `var_location_register()`. We reinitialize the `new_vars` after use for the next event.
```
class AssignmentVars(AssignmentVars):
def update(self, v):
for k, v in v.items():
self._set_kv(k, v)
self.var_location_register(self.new_vars)
self.new_vars = set()
```
The variable name now incorporates an index of how many reassignments it has gone through, effectively making each reassignment a unique variable.
```
class AssignmentVars(AssignmentVars):
def var_name(self, var):
return (var, self.accessed_seq_var[var])
```
While storing variables, we need to first check whether it was previously known. If it is not, we need to initialize the rename count. This is accomplished by `var_access`.
```
class AssignmentVars(AssignmentVars):
def var_access(self, var):
if var not in self.accessed_seq_var:
self.accessed_seq_var[var] = 0
return self.var_name(var)
```
During a variable reassignment, we update the `accessed_seq_var` to reflect the new count.
```
class AssignmentVars(AssignmentVars):
def var_assign(self, var):
self.accessed_seq_var[var] += 1
self.new_vars.add(self.var_name(var))
return self.var_name(var)
```
These methods can be used as follows
```
sav = AssignmentVars('')
sav.defs
sav.var_access('v1')
sav.var_assign('v1')
```
Assigning to it again increments the counter.
```
sav.var_assign('v1')
```
The core of the logic is in `_set_kv()`. When a variable is being assigned, we get the sequenced variable name `s_var`. If the sequenced variable name was previously unknown in `defs`, then we have no further concerns. We add the sequenced variable to `defs`.
If the variable is previously known, then it is an indication of a possible reassignment. In this case, we look at the value the variable is holding. We check if the value changed. If it has not, then it is not.
If the value has changed, it is a reassignment. We first increment the variable usage sequence using `var_assign`, retrieve the new name, update the new name in `defs`.
```
class AssignmentVars(AssignmentVars):
def _set_kv(self, var, val):
s_var = self.var_access(var)
if s_var in self.defs and self.defs[s_var] == val:
return
self.defs[self.var_assign(var)] = val
```
Here is how it can be used. Assigning a variable the first time initializes its counter.
```
sav = AssignmentVars('')
sav['x'] = 'X'
sav.defs
```
If the variable is assigned again with the same value, it is probably not a reassignment.
```
sav['x'] = 'X'
sav.defs
```
However, if the value changed, it is a reassignment.
```
sav['x'] = 'Y'
sav.defs
```
There is a subtlety here. It is possible for a child method to be called from the middle of a parent method, and for both to use the same variable name with different values. In this case, when the child returns, parent will have the old variable with old value in context. With our implementation, we consider this as a reassignment. However, this is OK because adding a new reassignment is harmless, but missing one is not. Further, we will discuss later how this can be avoided.
We also define book keeping codes for `register_event()` `method_enter()` and `method_exit()` which are the methods responsible for keeping track of the method stack. The basic idea is that, each `method_enter()` represents a new method invocation. Hence it merits a new method id, which is generated from the `method_register`, and saved in the `method_id`. Since this is a new method, the method stack is extended by one element with this id. In the case of `method_exit()`, we pop the method stack, and reset the current `method_id` to what was below the current one.
```
class AssignmentVars(AssignmentVars):
def method_enter(self, cxt, my_vars):
self.current_event = 'call'
self.call_stack.enter(cxt.method)
self.event_locations[self.call_stack.method_id] = []
self.register_event(cxt)
self.update(my_vars)
def method_exit(self, cxt, my_vars):
self.current_event = 'return'
self.register_event(cxt)
self.update(my_vars)
self.call_stack.leave()
def method_statement(self, cxt, my_vars):
self.current_event = 'line'
self.register_event(cxt)
self.update(my_vars)
```
For each of the method events, we also register the event using `register_event()` which keeps track of the line numbers that were referenced in *this* method.
```
class AssignmentVars(AssignmentVars):
def register_event(self, cxt):
self.event_locations[self.call_stack.method_id].append(cxt.line_no)
```
The `var_location_register()` keeps the locations of newly added variables. The definition location of variables in a `call` is the *current* location. However, for a `line`, it would be the previous event in the current method.
```
class AssignmentVars(AssignmentVars):
def var_location_register(self, my_vars):
def loc(mid):
if self.current_event == 'call':
return self.event_locations[mid][-1]
elif self.current_event == 'line':
return self.event_locations[mid][-2]
elif self.current_event == 'return':
return self.event_locations[mid][-2]
else:
assert False
my_loc = loc(self.call_stack.method_id)
for var in my_vars:
self.var_def_lines[var] = my_loc
```
We define `defined_vars()` which returns the names of variables annotated with the line numbers as below.
```
class AssignmentVars(AssignmentVars):
def defined_vars(self, formatted=True):
def fmt(k):
v = (k[0], self.var_def_lines[k])
return "%s@%s" % v if formatted else v
return [(fmt(k), v) for k, v in self.defs.items()]
```
Similar to `defined_vars()` we define `seq_vars()` which annotates different variables with the number of times they were used.
```
class AssignmentVars(AssignmentVars):
def seq_vars(self, formatted=True):
def fmt(k):
v = (k[0], self.var_def_lines[k], k[1])
return "%s@%s:%s" % v if formatted else v
return {fmt(k): v for k, v in self.defs.items()}
```
### AssignmentTracker
The `AssignmentTracker` keeps the assignment definitions using the `AssignmentVars` we defined previously.
```
class AssignmentTracker(DefineTracker):
def __init__(self, my_input, trace, **kwargs):
self.options(kwargs)
self.my_input = my_input
self.my_assignments = self.create_assignments(my_input)
self.trace = trace
self.process()
def create_assignments(self, *args):
return AssignmentVars(*args)
```
To fine-tune the process, we define an optional parameter called `track_return`. During tracing a method return, Python produces a virtual variable that contains the result of the returned value. If the `track_return` is set, we capture this value as a variable.
* `track_return` -- if true, add a *virtual variable* to the Vars representing the return value
```
class AssignmentTracker(AssignmentTracker):
def options(self, kwargs):
self.track_return = kwargs.get('track_return', False)
super().options(kwargs)
```
There can be different kinds of events during a trace, which includes `call` when a function is entered, `return` when the function returns, `exception` when an exception is thrown and `line` when a statement is executed.
The previous `Tracker` was too simplistic in that it did not distinguish between the different events. We rectify that and define `on_call()`, `on_return()`, and `on_line()` respectively, which get called on their corresponding events.
Note that `on_line()` is called also for `on_return()`. The reason is, that Python invokes the trace function *before* the corresponding line is executed. Hence, effectively, the `on_return()` is called with the binding produced by the execution of the previous statement in the environment. Our processing in effect is done on values that were bound by the previous statement. Hence, calling `on_line()` here is appropriate as it provides the event handler a chance to work on the previous binding.
```
class AssignmentTracker(AssignmentTracker):
def on_call(self, arg, cxt, my_vars):
my_vars = cxt.parameters(my_vars)
self.my_assignments.method_enter(cxt, self.fragments(my_vars))
def on_line(self, arg, cxt, my_vars):
self.my_assignments.method_statement(cxt, self.fragments(my_vars))
def on_return(self, arg, cxt, my_vars):
self.on_line(arg, cxt, my_vars)
my_vars = {'<-%s' % cxt.method: arg} if self.track_return else {}
self.my_assignments.method_exit(cxt, my_vars)
def on_exception(self, arg, cxt, my_vara):
return
def track_event(self, event, arg, cxt, my_vars):
self.current_event = event
dispatch = {
'call': self.on_call,
'return': self.on_return,
'line': self.on_line,
'exception': self.on_exception
}
dispatch[event](arg, cxt, my_vars)
```
We can now use `AssignmentTracker` to track the different variables. To verify that our variable line number inference works, we recover definitions from the functions `A()`, `B()` and `C()` (with data annotations removed so that the input fragments are correctly identified).
```
def C(cp_1): # type: ignore
c_2 = cp_1
c_3 = c_2
return c_3
def B(bp_7): # type: ignore
b_8 = bp_7
return C(b_8)
def A(ap_12): # type: ignore
a_13 = ap_12
a_14 = B(a_13)
a_14 = a_14
a_13 = a_14
a_14 = B(a_13)
a_14 = B(a_14)[3:]
```
Running `A()` with sufficient input.
```
with Tracer('---xxx') as tracer:
A(tracer.my_input)
tracker = AssignmentTracker(tracer.my_input, tracer.trace, log=True)
for k, v in tracker.my_assignments.seq_vars().items():
print(k, '=', repr(v))
print()
for k, v in tracker.my_assignments.defined_vars(formatted=True):
print(k, '=', repr(v))
```
As can be seen, the line numbers are now correctly identified for each variables.
Let us try retrieving the assignments for a real world example.
```
traces = []
for inputstr in URLS_X:
clear_cache()
with Tracer(inputstr, files=['urllib/parse.py']) as tracer:
urlparse(tracer.my_input)
traces.append((tracer.my_input, tracer.trace))
tracker = AssignmentTracker(tracer.my_input, tracer.trace, log=True)
for k, v in tracker.my_assignments.defined_vars():
print(k, '=', repr(v))
print()
```
The line numbers of variables can be verified from the source code of [urllib/parse.py](https://github.com/python/cpython/blob/3.6/Lib/urllib/parse.py).
### Recovering a Derivation Tree
Does handling variable reassignments help with our URL examples? We look at these next.
```
class TreeMiner(TreeMiner):
def get_derivation_tree(self):
tree = (START_SYMBOL, [(self.my_input, [])])
for var, value in self.my_assignments:
self.log(0, "%s=%s" % (var, repr(value)))
self.apply_new_definition(tree, var, value)
return tree
```
#### Example 1: Recovering URL Derivation Tree
First we obtain the derivation tree of the URL 1
##### URL 1 derivation tree
```
clear_cache()
with Tracer(URLS_X[0], files=['urllib/parse.py']) as tracer:
urlparse(tracer.my_input)
sm = AssignmentTracker(tracer.my_input, tracer.trace)
dt = TreeMiner(tracer.my_input, sm.my_assignments.defined_vars())
display_tree(dt.tree)
```
Next, we obtain the derivation tree of URL 4
##### URL 4 derivation tree
```
clear_cache()
with Tracer(URLS_X[-1], files=['urllib/parse.py']) as tracer:
urlparse(tracer.my_input)
sm = AssignmentTracker(tracer.my_input, tracer.trace)
dt = TreeMiner(tracer.my_input, sm.my_assignments.defined_vars())
display_tree(dt.tree)
```
The derivation trees seem to belong to the same grammar. Hence, we obtain the grammar for the complete set. First, we update the `recover_grammar()` to use `AssignTracker`.
### Recover Grammar
```
class GrammarMiner(GrammarMiner):
def update_grammar(self, inputstr, trace):
at = self.create_tracker(inputstr, trace)
dt = self.create_tree_miner(inputstr, at.my_assignments.defined_vars())
self.add_tree(dt)
return self.grammar
def create_tracker(self, *args):
return AssignmentTracker(*args)
def create_tree_miner(self, *args):
return TreeMiner(*args)
```
Next, we use the modified `recover_grammar()` on derivation trees obtained from URLs.
```
url_grammar = recover_grammar(url_parse, URLS_X, files=['urllib/parse.py'])
```
The recovered grammar is below.
```
syntax_diagram(url_grammar)
```
Let us fuzz a little to see if the produced values are sane.
```
f = GrammarFuzzer(url_grammar)
for _ in range(10):
print(f.fuzz())
```
Our modifications does seem to help. Next, we check whether we can still retrieve the grammar for inventory.
#### Example 2: Recovering Inventory Grammar
```
inventory_grammar = recover_grammar(process_vehicle, VEHICLES)
syntax_diagram(inventory_grammar)
```
Using fuzzing to produce values from the grammar.
```
f = GrammarFuzzer(inventory_grammar)
for _ in range(10):
print(f.fuzz())
```
### Problems with the Grammar Miner with Reassignment
One of the problems with our grammar miner is that it doesn't yet account for the current context. That is, when replacing, a variable can replace tokens that it does not have access to (and hence, it is not a fragment of). Consider this example.
```
with Tracer(INVENTORY) as tracer:
process_inventory(tracer.my_input)
sm = AssignmentTracker(tracer.my_input, tracer.trace)
dt = TreeMiner(tracer.my_input, sm.my_assignments.defined_vars())
display_tree(dt.tree, graph_attr=lr_graph)
```
As can be seen, the derivation tree obtained is not quite what we expected. The issue is easily seen if we enable logging in the `TreeMiner`.
```
dt = TreeMiner(tracer.my_input, sm.my_assignments.defined_vars(), log=True)
```
Look at the last statement. We have a value `1999,car,` where only the `year` got replaced. We no longer have a `'car'` variable to continue the replacement here. This happens because the `'car'` value in `'1999,car,Chevy,Venture'` is not treated as a new value because the value `'car'` had occurred for `'vehicle'` variable in the exact same location for a *different* method call (for `'2000,car,Mercury,Cougar'`).
## A Grammar Miner with Scope
We need to incorporate inspection of the variables in the current context. We already have a stack of method calls so that we can obtain the current method at any point. We need to do the same for variables.
For that, we extend the `CallStack` to a new class `InputStack` which holds the method invoked as well as the parameters observed. It is essentially the record of activation of the method. We start with the original input at the base of the stack, and for each new method-call, we push the parameters of that call into the stack as a new record.
### Input Stack
```
class InputStack(CallStack):
def __init__(self, i, fragment_len=FRAGMENT_LEN):
self.inputs = [{START_SYMBOL: i}]
self.fragment_len = fragment_len
super().__init__()
```
In order to check if a particular variable be saved, we define `in_current_record()` which checks only the variables in the current scope for inclusion (rather than the original input string).
```
class InputStack(InputStack):
def in_current_record(self, val):
return any(val in var for var in self.inputs[-1].values())
my_istack = InputStack('hello my world')
my_istack.in_current_record('hello')
my_istack.in_current_record('bye')
my_istack.inputs.append({'greeting': 'hello', 'location': 'world'})
my_istack.in_current_record('hello')
my_istack.in_current_record('my')
```
We define the method `ignored()` that returns true if either the variable is not a string, or the variable length is less than the defined `fragment_len`.
```
class InputStack(InputStack):
def ignored(self, val):
return not (isinstance(val, str) and len(val) >= self.fragment_len)
my_istack = InputStack('hello world')
my_istack.ignored(1)
my_istack.ignored('a')
my_istack.ignored('help')
```
We can now define the `in_scope()` method that checks whether the variable needs to be ignored, and if it is not to be ignored, whether the variable value is present in the current scope.
```
class InputStack(InputStack):
def in_scope(self, k, val):
if self.ignored(val):
return False
return self.in_current_record(val)
```
Finally, we update `enter()` that pushes relevant variables in the current context to the stack.
```
class InputStack(InputStack):
def enter(self, method, inputs):
my_inputs = {k: v for k, v in inputs.items() if self.in_scope(k, v)}
self.inputs.append(my_inputs)
super().enter(method)
```
When a method returns, we also need a corresponding `leave()` to pop out the inputs and unwind the stack.
```
class InputStack(InputStack):
def leave(self):
self.inputs.pop()
super().leave()
```
### ScopedVars
We need to update our `AssignmentVars` to include information about which scope the variable was defined in. We start by updating `method_init()`.
```
class ScopedVars(AssignmentVars):
def method_init(self):
self.call_stack = self.create_call_stack(self.my_input)
self.event_locations = {self.call_stack.method_id: []}
def create_call_stack(self, i):
return InputStack(i)
```
Similarly, the `method_enter()` now initializes the `accessed_seq_var` for the current method call.
```
class ScopedVars(ScopedVars):
def method_enter(self, cxt, my_vars):
self.current_event = 'call'
self.call_stack.enter(cxt.method, my_vars)
self.accessed_seq_var[self.call_stack.method_id] = {}
self.event_locations[self.call_stack.method_id] = []
self.register_event(cxt)
self.update(my_vars)
```
The `update()` method now saves the context in which the value is defined. In the case of a parameter to a function, the context should be the context in which the function was called. On the other hand, a value defined during a statement execution would have the current context.
Further, we annotate on value rather than key because we do not want to duplicate variables when parameters are in context in the next line. They will have same value, but different context because they are present in a statement execution.
```
class ScopedVars(ScopedVars):
def update(self, v):
if self.current_event == 'call':
context = -2
elif self.current_event == 'line':
context = -1
else:
context = -1
for k, v in v.items():
self._set_kv(k, (v, self.call_stack.at(context)))
self.var_location_register(self.new_vars)
self.new_vars = set()
```
We also need to save the current method invocation so as to determine which variables are in scope. This information is now incorporated in the variable name as `accessed_seq_var[method_id][var]`.
```
class ScopedVars(ScopedVars):
def var_name(self, var):
return (var, self.call_stack.method_id,
self.accessed_seq_var[self.call_stack.method_id][var])
```
As before, `var_access` simply initializes the corresponding counter, this time in the context of `method_id`.
```
class ScopedVars(ScopedVars):
def var_access(self, var):
if var not in self.accessed_seq_var[self.call_stack.method_id]:
self.accessed_seq_var[self.call_stack.method_id][var] = 0
return self.var_name(var)
```
During a variable reassignment, we update the `accessed_seq_var` to reflect the new count.
```
class ScopedVars(ScopedVars):
def var_assign(self, var):
self.accessed_seq_var[self.call_stack.method_id][var] += 1
self.new_vars.add(self.var_name(var))
return self.var_name(var)
```
We now update `defined_vars()` to account for the new information.
```
class ScopedVars(ScopedVars):
def defined_vars(self, formatted=True):
def fmt(k):
method, i = k[1]
v = (method, i, k[0], self.var_def_lines[k])
return "%s[%d]:%s@%s" % v if formatted else v
return [(fmt(k), v) for k, v in self.defs.items()]
```
Updating `seq_vars()` to account for new information.
```
class ScopedVars(ScopedVars):
def seq_vars(self, formatted=True):
def fmt(k):
method, i = k[1]
v = (method, i, k[0], self.var_def_lines[k], k[2])
return "%s[%d]:%s@%s:%s" % v if formatted else v
return {fmt(k): v for k, v in self.defs.items()}
```
### Scope Tracker
With the `InputStack` and `Vars` defined, we can now define the `ScopeTracker`. The `ScopeTracker` only saves variables if the value is present in the current scope.
```
class ScopeTracker(AssignmentTracker):
def __init__(self, my_input, trace, **kwargs):
self.current_event = None
super().__init__(my_input, trace, **kwargs)
def create_assignments(self, *args):
return ScopedVars(*args)
```
We define a wrapper for checking whether a variable is present in the scope.
```
class ScopeTracker(ScopeTracker):
def is_input_fragment(self, var, value):
return self.my_assignments.call_stack.in_scope(var, value)
```
We can use the `ScopeTracker` as follows.
```
vehicle_traces = []
with Tracer(INVENTORY) as tracer:
process_inventory(tracer.my_input)
sm = ScopeTracker(tracer.my_input, tracer.trace)
vehicle_traces.append((tracer.my_input, sm))
for k, v in sm.my_assignments.seq_vars().items():
print(k, '=', repr(v))
```
### Recovering a Derivation Tree
The main difference in `apply_new_definition()` is that we add a second condition that checks for scope. In particular, variables are only allowed to replace portions of string fragments that were in scope.
The variable scope is indicated by `scope`. However, merely accounting for scope is not sufficient. For example, consider the fragment below.
```python
def my_fn(stringval):
partA, partB = stringval.split('/')
return partA, partB
svalue = ...
v1, v2 = my_fn(svalue)
```
Here, `v1` and `v2` get their values from a previous function call. Not from their current context. That is, we have to provide an exception for cases where an internal child method call may have generated a large fragment as we showed above. To account for that, we define `mseq()` that retrieves the method call sequence. In the above case, the `mseq()` of the internal child method call would be larger than the current `mseq()`. If so, we allow the replacement to proceed.
```
class ScopeTreeMiner(TreeMiner):
def mseq(self, key):
method, seq, var, lno = key
return seq
```
The `nt_var()` method needs to take the tuple and generate a non-terminal symbol out of it. We skip the method sequence because it is not relevant for the grammar.
```
class ScopeTreeMiner(ScopeTreeMiner):
def nt_var(self, key):
method, seq, var, lno = key
return to_nonterminal("%s@%d:%s" % (method, lno, var))
```
We now redefine the `apply_new_definition()` to account for context and scope. In particular, a variable is allowed to replace a part of a value only if the variable is in *scope* -- that is, it's scope (method sequence number of either its calling context in case it is a parameter or the current context in case it is a statement) is same as that of the value's method sequence number. An exception is made when the value's method sequence number is greater than the variable's method sequence number. In that case, the value may have come from an internal call. We allow the replacement to proceed in that case.
```
class ScopeTreeMiner(ScopeTreeMiner):
def partition(self, part, value):
return value.partition(part)
def partition_by_part(self, pair, value):
(nt_var, nt_seq), (v, v_scope) = pair
prefix_k_suffix = [
(nt_var, [(v, [], nt_seq)]) if i == 1 else (e, [])
for i, e in enumerate(self.partition(v, value))
if e]
return prefix_k_suffix
def insert_into_tree(self, my_tree, pair):
var, values, my_scope = my_tree
(nt_var, nt_seq), (v, v_scope) = pair
applied = False
for i, value_ in enumerate(values):
key, arr, scope = value_
self.log(2, "-> [%d] %s" % (i, repr(value_)))
if is_nonterminal(key):
applied = self.insert_into_tree(value_, pair)
if applied:
break
else:
if v_scope != scope:
if nt_seq > scope:
continue
if not v or not self.string_part_of_value(v, key):
continue
prefix_k_suffix = [(k, children, scope) for k, children
in self.partition_by_part(pair, key)]
del values[i]
for j, rep in enumerate(prefix_k_suffix):
values.insert(j + i, rep)
applied = True
self.log(2, " > %s" % (repr([i[0] for i in prefix_k_suffix])))
break
return applied
```
The `apply_new_definition()` is now modified to carry additional contextual information `mseq`.
```
class ScopeTreeMiner(ScopeTreeMiner):
def apply_new_definition(self, tree, var, value_):
nt_var = self.nt_var(var)
seq = self.mseq(var)
val, (smethod, mseq) = value_
return self.insert_into_tree(tree, ((nt_var, seq), (val, mseq)))
```
We also modify `get_derivation_tree()` so that the initial node carries the context.
```
class ScopeTreeMiner(ScopeTreeMiner):
def get_derivation_tree(self):
tree = (START_SYMBOL, [(self.my_input, [], 0)], 0)
for var, value in self.my_assignments:
self.log(0, "%s=%s" % (var, repr(value)))
self.apply_new_definition(tree, var, value)
return tree
```
#### Example 1: Recovering URL Parse Tree
We verify that our URL parse tree recovery still works as expected.
```
url_dts = []
for inputstr in URLS_X:
clear_cache()
with Tracer(inputstr, files=['urllib/parse.py']) as tracer:
urlparse(tracer.my_input)
sm = ScopeTracker(tracer.my_input, tracer.trace)
for k, v in sm.my_assignments.defined_vars(formatted=False):
print(k, '=', repr(v))
dt = ScopeTreeMiner(
tracer.my_input,
sm.my_assignments.defined_vars(
formatted=False))
display_tree(dt.tree, graph_attr=lr_graph)
url_dts.append(dt)
```
#### Example 2: Recovering Inventory Parse Tree
Next, we look at recovering the parse tree from `process_inventory()` which failed last time.
```
with Tracer(INVENTORY) as tracer:
process_inventory(tracer.my_input)
sm = ScopeTracker(tracer.my_input, tracer.trace)
for k, v in sm.my_assignments.defined_vars():
print(k, '=', repr(v))
inventory_dt = ScopeTreeMiner(
tracer.my_input,
sm.my_assignments.defined_vars(
formatted=False))
display_tree(inventory_dt.tree, graph_attr=lr_graph)
```
The recovered parse tree seems reasonable.
One of the things that one might notice from our Example (2) is that the three subtrees -- `vehicle[2:1]`, `vehicle[4:1]` and `vehicle[6:1]` are quite alike. We will examine how this can be exploited to generate a grammar directly, next.
### Grammar Mining
The `tree_to_grammar()` is now redefined as follows, to account for the extra scope in nodes.
```
class ScopedGrammarMiner(GrammarMiner):
def tree_to_grammar(self, tree):
key, children, scope = tree
one_alt = [ckey for ckey, gchildren, cscope in children if ckey != key]
hsh = {key: [one_alt] if one_alt else []}
for child in children:
(ckey, _gc, _cscope) = child
if not is_nonterminal(ckey):
continue
chsh = self.tree_to_grammar(child)
for k in chsh:
if k not in hsh:
hsh[k] = chsh[k]
else:
hsh[k].extend(chsh[k])
return hsh
```
The grammar is in canonical form, which needs to be massaged to display. First, the recovered grammar for inventory.
```
si = ScopedGrammarMiner()
si.add_tree(inventory_dt)
syntax_diagram(readable(si.grammar))
```
The recovered grammar for URLs.
```
su = ScopedGrammarMiner()
for t in url_dts:
su.add_tree(t)
syntax_diagram(readable(su.grammar))
```
One might notice that the grammar is not entirely human readable, with a number of single token definitions.
Hence, the last piece of the puzzle is the cleanup method `clean_grammar()`, which cleans up such definitions. The idea is to look for single token definitions such that a key is defined exactly by another key (single alternative, single token, nonterminal).
```
class ScopedGrammarMiner(ScopedGrammarMiner):
def get_replacements(self, grammar):
replacements = {}
for k in grammar:
if k == START_SYMBOL:
continue
alts = grammar[k]
if len(set([str(i) for i in alts])) != 1:
continue
rule = alts[0]
if len(rule) != 1:
continue
tok = rule[0]
if not is_nonterminal(tok):
continue
replacements[k] = tok
return replacements
```
Once we have such a list, iteratively replace the original key where ever it is used with the token we found earlier. Repeat until none is left.
```
class ScopedGrammarMiner(ScopedGrammarMiner):
def clean_grammar(self):
replacements = self.get_replacements(self.grammar)
while True:
changed = set()
for k in self.grammar:
if k in replacements:
continue
new_alts = []
for alt in self.grammar[k]:
new_alt = []
for t in alt:
if t in replacements:
new_alt.append(replacements[t])
changed.add(t)
else:
new_alt.append(t)
new_alts.append(new_alt)
self.grammar[k] = new_alts
if not changed:
break
for k in changed:
self.grammar.pop(k, None)
return readable(self.grammar)
```
The `clean_grammar()` is used as follows:
```
si = ScopedGrammarMiner()
si.add_tree(inventory_dt)
syntax_diagram(readable(si.clean_grammar()))
```
We update the `update_grammar()` to use the right tracker and miner.
```
class ScopedGrammarMiner(ScopedGrammarMiner):
def update_grammar(self, inputstr, trace):
at = self.create_tracker(inputstr, trace)
dt = self.create_tree_miner(
inputstr, at.my_assignments.defined_vars(
formatted=False))
self.add_tree(dt)
return self.grammar
def create_tracker(self, *args):
return ScopeTracker(*args)
def create_tree_miner(self, *args):
return ScopeTreeMiner(*args)
```
The `recover_grammar()` uses the right miner, and returns a cleaned grammar.
```
def recover_grammar(fn, inputs, **kwargs): # type: ignore
miner = ScopedGrammarMiner()
for inputstr in inputs:
with Tracer(inputstr, **kwargs) as tracer:
fn(tracer.my_input)
miner.update_grammar(tracer.my_input, tracer.trace)
return readable(miner.clean_grammar())
url_grammar = recover_grammar(url_parse, URLS_X, files=['urllib/parse.py'])
syntax_diagram(url_grammar)
f = GrammarFuzzer(url_grammar)
for _ in range(10):
print(f.fuzz())
inventory_grammar = recover_grammar(process_inventory, [INVENTORY])
syntax_diagram(inventory_grammar)
f = GrammarFuzzer(inventory_grammar)
for _ in range(10):
print(f.fuzz())
```
We see how tracking scope helps us to extract an even more precise grammar.
Notice that we use *String* inclusion testing as a way of determining whether a particular string fragment came from the original input string. While this may seem rather error-prone compared to dynamic tainting, we note that numerous tracing tools such as `dtrace()` and `ptrace()` allow one to obtain the information we seek from execution of binaries directly in different platforms. However, methods for obtaining dynamic taints almost always involve instrumenting the binaries before they can be used. Hence, this method of string inclusion can be more generally applied than dynamic tainting approaches. Further, dynamic taints are often lost due to implicit transmission, or at the boundary between *Python* and *C* code. String inclusion has not such problems. Hence, our approach can often obtain better results than relying on dynamic tainting.
## Synopsis
This chapter provides a number of classes to mine input grammars from existing programs. The function `recover_grammar()` could be the easiest to use. It takes a function and a set of inputs, and returns a grammar that describes its input language.
We apply `recover_grammar()` on a `url_parse()` function that takes and decomposes URLs:
```
url_parse('https://www.fuzzingbook.org/')
URLS
```
We extract the input grammar for `url_parse()` using `recover_grammar()`:
```
grammar = recover_grammar(url_parse, URLS, files=['urllib/parse.py'])
grammar
```
The names of nonterminals are a bit technical; but the grammar nicely represents the structure of the input; for instance, the different schemes (`"http"`, `"https"`) are all identified:
```
syntax_diagram(grammar)
```
The grammar can be immediately used for fuzzing, producing arbitrary combinations of input elements, which are all syntactically valid.
```
from GrammarCoverageFuzzer import GrammarCoverageFuzzer
fuzzer = GrammarCoverageFuzzer(grammar)
[fuzzer.fuzz() for i in range(5)]
```
Being able to automatically extract a grammar and to use this grammar for fuzzing makes for very effective test generation with a minimum of manual work.
## Lessons Learned
* Given a set of sample inputs for a program, we can learn an input grammar by examining variable values during execution if the program relies on handwritten parsers.
* Simple string inclusion checks are sufficient to obtain reasonably accurate grammars from real world programs.
* The resulting grammars can be directly used for fuzzing, and can have a multiplier effect on any samples you have.
## Next Steps
* Learn how to use [information flow](InformationFlow.ipynb) to further improve mapping inputs to states.
## Background
Recovering the language from a _set of samples_ (i.e., not taking into account a possible program that might process them) is a well researched topic. The excellent reference by Higuera \cite{higuera2010grammatical} covers all the classical approaches. The current state of the art in black box grammar mining is described by Clark \cite{clark2013learning}.
Learning an input language from a _program_, with or without samples, is yet a emerging topic, despite its potential for fuzzing. The pioneering work in this area was done by Lin et al. \cite{Lin2008} who invented a way to retrieve the parse trees from top down and bottom up parsers. The approach described in this chapter is based directly on the AUTOGRAM work of Hoschele et al. \cite{Hoschele2017}.
## Exercises
### Exercise 1: Flattening complex objects
Our grammar miners only check for string fragments. However, programs may often pass containers or custom objects containing input fragments. For example, consider the plausible modification for our inventory processor, where we use a custom object `Vehicle` to carry fragments.
```
class Vehicle:
def __init__(self, vehicle: str):
year, kind, company, model, *_ = vehicle.split(',')
self.year, self.kind, self.company, self.model = year, kind, company, model
def process_inventory_with_obj(inventory: str) -> str:
res = []
for vehicle in inventory.split('\n'):
ret = process_vehicle(vehicle)
res.extend(ret)
return '\n'.join(res)
def process_vehicle_with_obj(vehicle: str) -> List[str]:
v = Vehicle(vehicle)
if v.kind == 'van':
return process_van_with_obj(v)
elif v.kind == 'car':
return process_car_with_obj(v)
else:
raise Exception('Invalid entry')
def process_van_with_obj(vehicle: Vehicle) -> List[str]:
res = [
"We have a %s %s van from %s vintage." % (vehicle.company,
vehicle.model, vehicle.year)
]
iyear = int(vehicle.year)
if iyear > 2010:
res.append("It is a recent model!")
else:
res.append("It is an old but reliable model!")
return res
def process_car_with_obj(vehicle: Vehicle) -> List[str]:
res = [
"We have a %s %s car from %s vintage." % (vehicle.company,
vehicle.model, vehicle.year)
]
iyear = int(vehicle.year)
if iyear > 2016:
res.append("It is a recent model!")
else:
res.append("It is an old but reliable model!")
return res
```
We recover the grammar as before.
```
vehicle_grammar = recover_grammar(
process_inventory_with_obj,
[INVENTORY],
methods=INVENTORY_METHODS)
```
The new vehicle grammar is missing in details, especially as to the different models and company for a van and car.
```
syntax_diagram(vehicle_grammar)
```
The problem is that, we are looking specifically for string objects that contain fragments of the input string during tracing. Can you modify our grammar miner to correctly account for the complex objects too?
**Solution.**
The problem can be understood if we execute the tracer under verbose logging.
```
with Tracer(INVENTORY, methods=INVENTORY_METHODS, log=True) as tracer:
process_inventory(tracer.my_input)
print()
print('Traced values:')
for t in tracer.trace:
print(t)
```
You can see that we lose track of string fragments as soon as they are incorporated into the `Vehicle` object. The way out is to trace these variables separately.
For that, we develop the `flatten()` method that given any custom complex object and its key, returns a list of flattened *key*,*value* pairs that correspond to the object passed in.
The `MAX_DEPTH` parameter controls the maximum flattening limit.
```
MAX_DEPTH = 10
def set_flatten_depth(depth):
global MAX_DEPTH
MAX_DEPTH = depth
def flatten(key, val, depth=MAX_DEPTH):
tv = type(val)
if depth <= 0:
return [(key, val)]
if isinstance(val, (int, float, complex, str, bytes, bytearray)):
return [(key, val)]
elif isinstance(val, (set, frozenset, list, tuple, range)):
values = [(i, e) for i, elt in enumerate(val) for e in flatten(i, elt, depth-1)]
return [("%s.%d" % (key, i), v) for i, v in values]
elif isinstance(val, dict):
values = [e for k, elt in val.items() for e in flatten(k, elt, depth-1)]
return [("%s.%s" % (key, k), v) for k, v in values]
elif isinstance(val, str):
return [(key, val)]
elif hasattr(val, '__dict__'):
values = [e for k, elt in val.__dict__.items()
for e in flatten(k, elt, depth-1)]
return [("%s.%s" % (key, k), v) for k, v in values]
else:
return [(key, val)]
```
Next, we hook the `flatten()` into the `Context` class so that the parameters we obtain are flattened.
```
class Context(Context):
def extract_vars(self, frame):
vals = inspect.getargvalues(frame).locals
return {k1: v1 for k, v in vals.items() for k1, v1 in flatten(k, v)}
def parameters(self, all_vars):
def check_param(k):
return any(k.startswith(p) for p in self.parameter_names)
return {k: v for k, v in all_vars.items() if check_param(k)}
def qualified(self, all_vars):
return {"%s:%s" % (self.method, k): v for k, v in all_vars.items()}
```
With this change, we have the following trace output.
```
with Tracer(INVENTORY, methods=INVENTORY_METHODS, log=True) as tracer:
process_inventory(tracer.my_input)
print()
print('Traced values:')
for t in tracer.trace:
print(t)
```
Our change seems to have worked. Let us derive the grammar.
```
vehicle_grammar = recover_grammar(
process_inventory,
[INVENTORY],
methods=INVENTORY_METHODS)
syntax_diagram(vehicle_grammar)
```
The recovered grammar contains all the details that we were able to recover before.
### Exercise 2: Incorporating Taints from InformationFlow
We have been using *string inclusion* to check whether a particular fragment came from the input string. This is unsatisfactory as it required us to compromise on the size of the strings tracked, which was limited to those greater than `FRAGMENT_LEN`. Further, it is possible that a single method could process a string where a fragment repeats, but is part of different tokens. For example, an embedded comma in the CSV file would cause our parser to fail. One way to avoid this is to rely on *dynamic taints*, and check for taint inclusion rather than string inclusion.
The chapter on [information flow](InformationFlow.ipynb) details how to incorporate dynamic taints. Can you update our grammar miner based on scope to use *dynamic taints* instead?
**Solution.**
First, we import `ostr` to track the origins of string fragments.
```
from InformationFlow import ostr
```
Next, we define `is_fragment()` to verify that a fragment is from a given input string.
```
def is_fragment(fragment, original):
assert isinstance(original, ostr)
if not isinstance(fragment, ostr):
return False
return set(fragment.origin) <= set(original.origin)
```
Now, all that remains is to hook the tainted fragment check to our grammar miner. This is accomplished by modifying `in_current_record()` and `ignored()` methods in the `InputStack`.
```
class TaintedInputStack(InputStack):
def in_current_record(self, val):
return any(is_fragment(val, var) for var in self.inputs[-1].values())
class TaintedInputStack(TaintedInputStack):
def ignored(self, val):
return not isinstance(val, ostr)
```
We then hook in the `TaintedInputStack` to the grammar mining infrastructure.
```
class TaintedScopedVars(ScopedVars):
def create_call_stack(self, i):
return TaintedInputStack(i)
class TaintedScopeTracker(ScopeTracker):
def create_assignments(self, *args):
return TaintedScopedVars(*args)
class TaintedScopeTreeMiner(ScopeTreeMiner):
def string_part_of_value(self, part, value):
return is_fragment(part, value)
def partition(self, part, value):
begin = value.origin.index(part.origin[0])
end = value.origin.index(part.origin[-1])+1
return value[:begin], value[begin:end], value[end:]
```
<!-- **Advanced.** The *dynamic taint* approach is limited in that it can not observe implicit flows. For example, consider the fragment below.
```python
if my_fragment == 'begin':
return 'begin'
```
In this case, we lose track of the string `begin` that is returned even though it is dependent on the value of `my_fragment`. For such cases, a better (but costly) alternative is to rely on concolic execution and capture the constraints as it relates to input characters on each variable.
The chapter on [concolic fuzzing](ConcolicFuzzer.ipynb) details how to incorporate concolic symbolic execution to program execution. Can you update our grammar miner to use *concolic execution* to track taints instead?
-->
```
class TaintedScopedGrammarMiner(ScopedGrammarMiner):
def create_tracker(self, *args):
return TaintedScopeTracker(*args)
def create_tree_miner(self, *args):
return TaintedScopeTreeMiner(*args)
```
Finally, we define `recover_grammar_with_taints()` to recover the grammar.
```
def recover_grammar_with_taints(fn, inputs, **kwargs):
miner = TaintedScopedGrammarMiner()
for inputstr in inputs:
with Tracer(ostr(inputstr), **kwargs) as tracer:
fn(tracer.my_input)
miner.update_grammar(tracer.my_input, tracer.trace)
return readable(miner.clean_grammar())
```
Here is how one can use it.
```
inventory_grammar = recover_grammar_with_taints(
process_inventory, [INVENTORY],
methods=[
'process_inventory', 'process_vehicle', 'process_car', 'process_van'
])
syntax_diagram(inventory_grammar)
url_grammar = recover_grammar_with_taints(
url_parse, URLS_X + ['ftp://user4:pass1@host4/?key4=value3'],
methods=['urlsplit', 'urlparse', '_splitnetloc'])
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Array/decorrelation_stretch.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Array/decorrelation_stretch.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Array/decorrelation_stretch.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Array/decorrelation_stretch.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
# Decorrelation Stretch
def dcs(image, region, scale):
# function dcs(image, region, scale) {
bandNames = image.bandNames()
# The axes are numbered, so to make the following code more
# readable, give the axes names.
imageAxis = 0
bandAxis = 1
# Compute the mean of each band in the region.
means = image.reduceRegion(ee.Reducer.mean(), region, scale)
# Create a constant array image from the mean of each band.
meansArray = ee.Image(means.toArray())
# Collapse the bands of the image into a 1D array per pixel,
# with images along the first axis and bands along the second.
arrays = image.toArray()
# Perform element-by-element subtraction, which centers the
# distribution of each band within the region.
centered = arrays.subtract(meansArray)
# Compute the covariance of the bands within the region.
covar = centered.reduceRegion(**{
'reducer': ee.Reducer.centeredCovariance(),
'geometry': region,
'scale': scale
})
# Get the 'array' result and cast to an array. Note this is a
# single array, not one array per pixel, and represents the
# band-to-band covariance within the region.
covarArray = ee.Array(covar.get('array'))
# Perform an eigen analysis and slice apart the values and vectors.
eigens = covarArray.eigen()
eigenValues = eigens.slice(bandAxis, 0, 1)
eigenVectors = eigens.slice(bandAxis, 1)
# Rotate by the eigenvectors, scale to a variance of 30, and rotate back.
i = ee.Array.identity(bandNames.length())
variance = eigenValues.sqrt().matrixToDiag()
scaled = i.multiply(30).divide(variance)
rotation = eigenVectors.transpose() \
.matrixMultiply(scaled) \
.matrixMultiply(eigenVectors)
# Reshape the 1-D 'normalized' array, so we can left matrix multiply
# with the rotation. This requires embedding it in 2-D space and
# transposing.
transposed = centered.arrayRepeat(bandAxis, 1).arrayTranspose()
# Convert rotated results to 3 RGB bands, and shift the mean to 127.
return transposed.matrixMultiply(ee.Image(rotation)) \
.arrayProject([bandAxis]) \
.arrayFlatten([bandNames]) \
.add(127).byte()
# }
image = ee.Image('MODIS/006/MCD43A4/2002_07_04')
Map.setCenter(-52.09717, -7.03548, 7)
# View the original bland image.
rgb = [0, 3, 2]
Map.addLayer(image.select(rgb), {'min': -100, 'max': 2000}, 'Original Image')
# Stretch the values within an interesting region.
region = ee.Geometry.Rectangle(-57.04651, -8.91823, -47.24121, -5.13531)
Map.addLayer(dcs(image, region, 1000).select(rgb), {}, 'DCS Image')
# Display the region in which covariance stats were computed.
Map.addLayer(ee.Image().paint(region, 0, 2), {}, 'Region')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
## Exercise 3
In the videos you looked at how you would improve Fashion MNIST using Convolutions. For your exercise see if you can improve MNIST to 99.8% accuracy or more using only a single convolutional layer and a single MaxPooling 2D. You should stop training once the accuracy goes above this amount. It should happen in less than 20 epochs, so it's ok to hard code the number of epochs for training, but your training must end once it hits the above metric. If it doesn't, then you'll need to redesign your layers.
I've started the code for you -- you need to finish it!
When 99.8% accuracy has been hit, you should print out the string "Reached 99.8% accuracy so cancelling training!"
```
import tensorflow as tf
from os import path, getcwd, chdir
# DO NOT CHANGE THE LINE BELOW. If you are developing in a local
# environment, then grab mnist.npz from the Coursera Jupyter Notebook
# and place it inside a local folder and edit the path to that location
path = f"{getcwd()}/../tmp2/mnist.npz"
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('acc')>0.998):
print("\nReached 99.8% accuracy so cancelling training!")
self.model.stop_training = True
callbacks = myCallback()
# GRADED FUNCTION: train_mnist_conv
def train_mnist_conv():
# Please write your code only where you are indicated.
# please do not remove model fitting inline comments.
# YOUR CODE STARTS HERE
# YOUR CODE ENDS HERE
mnist = tf.keras.datasets.mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data(path=path)
training_images=training_images.reshape(60000, 28, 28, 1)
training_images=training_images / 255.0
test_images = test_images.reshape(10000, 28, 28, 1)
test_images=test_images/255.0
model = tf.keras.models.Sequential([
# YOUR CODE STARTS HERE
tf.keras.layers.Conv2D(32, (3,3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(2, 2),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
# YOUR CODE ENDS HERE
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# model fitting
history = model.fit(
# YOUR CODE STARTS HERE
training_images,
training_labels,
epochs=10,
callbacks=[callbacks]
# YOUR CODE ENDS HERE
)
# model fitting
return history.epoch, history.history['acc'][-1]
_, _ = train_mnist_conv()
# Now click the 'Submit Assignment' button above.
# Once that is complete, please run the following two cells to save your work and close the notebook
%%javascript
<!-- Save the notebook -->
IPython.notebook.save_checkpoint();
%%javascript
<!-- Shutdown and close the notebook -->
window.onbeforeunload = null
window.close();
IPython.notebook.session.delete();
```
| github_jupyter |
# Concept Drift
In the context of data streams, it is assumed that data can change over time. The change in the relationship between the data (features) and the target to learn is known as **Concept Drift**. As examples we can mention, the electricity demand across the year, the stock market, and the likelihood of a new movie to be successful. Let's consider the movie example: Two movies can have similar features such as popular actors/directors, storyline, production budget, marketing campaigns, etc. yet it is not certain that both will be similarly successful. What the target audience *considers* worth watching (and their money) is constantly changing and production companies must adapt accordingly to avoid "box office flops".
## Impact of drift on learning
Concept drift can have a significant impact on predictive performance if not handled properly. Most batch learning models will fail in the presence of concept drift as they are essentially trained on different data. On the other hand, stream learning methods continuously update themselves and adapt to new concepts. Furthermore, drift-aware methods use change detection methods (a.k.a. drift detectors) to trigger *mitigation mechanisms* if a change in performance is detected.
## Detecting concept drift
Multiple drift detection methods have been proposed. The goal of a drift detector is to signal an alarm in the presence of drift. A good drift detector maximizes the number of true positives while keeping the number of false positives to a minimum. It must also be resource-wise efficient to work in the context of infinite data streams.
For this example, we will generate a synthetic data stream by concatenating 3 distributions of 1000 samples each:
- $dist_a$: $\mu=0.8$, $\sigma=0.05$
- $dist_b$: $\mu=0.4$, $\sigma=0.02$
- $dist_c$: $\mu=0.6$, $\sigma=0.1$.
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import gridspec
# Generate data for 3 distributions
random_state = np.random.RandomState(seed=42)
dist_a = random_state.normal(0.8, 0.05, 1000)
dist_b = random_state.normal(0.4, 0.02, 1000)
dist_c = random_state.normal(0.6, 0.1, 1000)
# Concatenate data to simulate a data stream with 2 drifts
stream = np.concatenate((dist_a, dist_b, dist_c))
# Auxiliary function to plot the data
def plot_data(dist_a, dist_b, dist_c, drifts=None):
fig = plt.figure(figsize=(7,3), tight_layout=True)
gs = gridspec.GridSpec(1, 2, width_ratios=[3, 1])
ax1, ax2 = plt.subplot(gs[0]), plt.subplot(gs[1])
ax1.grid()
ax1.plot(stream, label='Stream')
ax2.grid(axis='y')
ax2.hist(dist_a, label=r'$dist_a$')
ax2.hist(dist_b, label=r'$dist_b$')
ax2.hist(dist_c, label=r'$dist_c$')
if drifts is not None:
for drift_detected in drifts:
ax1.axvline(drift_detected, color='red')
plt.show()
plot_data(dist_a, dist_b, dist_c)
```
### Drift detection test
We will use the ADaptive WINdowing (`ADWIN`) drift detection method. Remember that the goal is to indicate that drift has occurred after samples **1000** and **2000** in the synthetic data stream.
```
from river import drift
drift_detector = drift.ADWIN()
drifts = []
for i, val in enumerate(stream):
drift_detector.update(val) # Data is processed one sample at a time
if drift_detector.change_detected:
# The drift detector indicates after each sample if there is a drift in the data
print(f'Change detected at index {i}')
drifts.append(i)
drift_detector.reset() # As a best practice, we reset the detector
plot_data(dist_a, dist_b, dist_c, drifts)
```
We see that `ADWIN` successfully indicates the presence of drift (red vertical lines) close to the begining of a new data distribution.
---
We conclude this example with some remarks regarding concept drift detectors and their usage:
- In practice, drift detectors provide stream learning methods with robustness against concept drift. Drift detectors monitor the model usually through a performance metric.
- Drift detectors work on univariate data. This is why they are used to monitor a model's performance and not the data itself. Remember that concept drift is defined as a change in the relationship between data and the target to learn (in supervised learning).
- Drift detectors define their expectations regarding input data. It is important to know these expectations to feed a given drift detector with the correct data.
| github_jupyter |
```
# Import required packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import nltk
import keras
from keras.preprocessing.text import Tokenizer
from keras.layers import Dense, Dropout, Embedding, LSTM, Bidirectional,InputLayer
from sklearn.model_selection import train_test_split
# Download all ntlk resources
nltk.download('all')
# Import Sentiment140 dataset
# Update file location of Sentiment140.csv according to your working environment
data_path = "Sentiment140.csv"
twitter_data = pd.read_csv(data_path,names=['target','id','date','flag','user','text'],
encoding = "ISO-8859-1")
# Create NumPy array of unprocessed input text and target
X=np.array(twitter_data['text'])
Y=np.array(twitter_data['target'])
# Set Y=1 for Positive Tweets
Y[Y==4]=1
# Visualize Dataset
index = 123 # index in range [0,1599999]
print(X[index])
# Define Preprocessing functions
def tokenize(X):
"""
Tokenize the data using nltk
"""
treebank = nltk.tokenize.TreebankWordTokenizer()
X_tokenized=[treebank.tokenize(sentence) for sentence in X]
return X_tokenized
def remove_stopwords(X):
"""
Remove Stopwords using nltk
"""
stopwords=nltk.corpus.stopwords.words('english') + ['@']
X_without_stopwords = []
for sentence in X:
temp = [word for word in sentence if not word in stopwords]
X_without_stopwords.append(temp)
return X_without_stopwords
def stem(X,type='porter'):
"""
Perform Stemming using nltk
type = 'Porter','Snowball','Lancaster'
"""
if type == 'porter':
stemmer= nltk.stem.PorterStemmer()
elif type == 'snowball':
stemmer = nltk.stem.SnowballStemmer()
elif type == 'lancaster':
stemmer = nltk.stem.LancasterStemmer()
X_stemmed = []
for sentence in X:
temp = [stemmer.stem(word) for word in sentence]
X_stemmed.append(temp)
return X_stemmed
def get_wordnet_pos(treebank_tag):
"""
return WORDNET POS compliance to WORDENT lemmatization (a,n,r,v)
"""
if treebank_tag.startswith('J'):
return 'a'
elif treebank_tag.startswith('V'):
return 'v'
elif treebank_tag.startswith('N'):
return 'n'
elif treebank_tag.startswith('R'):
return 'r'
else:
return 'n'
def lemmatize(X):
"""
Lemmatize words using corresponding POS tag
"""
lemmatizer = nltk.stem.WordNetLemmatizer()
X_pos = []
X_lemmatized = []
for sentence in X :
temp = nltk.pos_tag(sentence)
X_pos.append(temp)
for sentence in X_pos :
temp = [ lemmatizer.lemmatize(word[0],pos=get_wordnet_pos(word[1])) for word in sentence]
X_lemmatized.append(temp)
return X_lemmatized
```
# **Training on Unprocessed data**
```
# Count total no. of distinct tokens
tokenizer = Tokenizer(filters='@')
tokenizer.fit_on_texts(X)
print('No. of distinct tokens = '+str(len(tokenizer.word_index)))
# Define Vocabulary size (no. of most frequent tokens) to consider
max_vocab=50000
# Reload Twitter dataset with new Vocabulary
tokenizer = Tokenizer(num_words=max_vocab,filters='@')
tokenizer.fit_on_texts(X)
# Vectorize input text using Vocabulary
X_vectorized=tokenizer.texts_to_sequences(X)
# Count average length of tweets
sum=0
for sentence in X_vectorized:
sum+=len(sentence)
print('Average length of tweets = '+str(sum/len(X_vectorized)))
# Define Maximum input length of the Model
max_length=20
# Pad or Trim data to defined input length
X_pad = keras.preprocessing.sequence.pad_sequences(X_vectorized,max_length,padding='post',
truncating='post')
print(X_pad.shape)
# Visualize unprocessed data
index = 123 # index in range [0,1599999]
print('\nOriginal :')
print(X[index])
print('\nTokenized :')
print(X_tokenized[index])
print('\nVectorized :')
print(X_vectorized[index])
print('\nPadded :')
print(X_pad[index])
# Define Classification model
np.random.seed(123)
model_1 = keras.models.Sequential(name='Model_1(Sentiment140_Unprocessed)')
model_1.add(InputLayer(input_shape=(20,),name = 'Integer_Encoding'))
model_1.add(Embedding(max_vocab, 100,input_length=max_length,mask_zero=True,
name='100D_Encoding',trainable=False))
model_1.add(Bidirectional(LSTM(64,name='LSTM'),name='Bidirectional_RNN'))
model_1.add(Dropout(0.5,name='Regularizer'))
model_1.add(Dense(1, activation='sigmoid',name='Sigmoid_Classifier'))
# Compile the Model
model_1.compile(optimizer=keras.optimizers.adam(lr=0.001),loss='binary_crossentropy',
metrics=['accuracy'])
# Model details
model_1.summary()
keras.utils.plot_model(model_1,to_file='model_1.png')
# Perform train-test split
np.random.seed(123)
X_train, X_test, Y_train, Y_test = train_test_split (X_pad,Y.reshape(Y.shape[0],1),test_size=0.05)
print(X_train.shape)
print(X_test.shape)
print(Y_train.shape)
print(Y_test.shape)
# Train the model
history_model_1 = model_1.fit(X_train,Y_train,batch_size=1024,epochs=15,validation_split=0.05)
# Evaluate model on Test data
model_1.evaluate(X_test,Y_test)
```
# **Training on Pre-Processed data with GloVe Word Embeddings**
```
# Preprocess the data
X_tokenized = tokenize (X)
X_without_stopwords = remove_stopwords ( X_tokenized )
X_lemmatized = lemmatize ( X_without_stopwords )
X_clean = []
for sentence in X_lemmatized:
temp = " ".join(sentence)
X_clean.append(temp)
# Count total no. of distinct tokens
tokenizer = Tokenizer(filters='@')
tokenizer.fit_on_texts(X_clean)
print('No. of distinct tokens = '+str(len(tokenizer.word_index)))
# Define Vocabulary size (no. of most frequent tokens) to consider
max_vocab=50000
# Reload Twitter dataset with new Vocabulary
tokenizer = Tokenizer(num_words=max_vocab,filters='@')
tokenizer.fit_on_texts(X_clean)
# Vectorize input text using Vocabulary
X_clean_vectorized=tokenizer.texts_to_sequences(X_clean)
# Count average length of tweets
length=[]
for sentence in X_clean_vectorized:
length.append(len(sentence))
print('Average length of tweets = '+str(np.mean(length)))
# Define Maximum input length of the Model
max_length=20
# Pad or Trim data to defined input length
X_clean_pad = keras.preprocessing.sequence.pad_sequences(X_clean_vectorized,max_length,padding='post',
truncating='post')
print(X_clean_pad.shape)
# Visualize pre-processed data
index = 123 # index in range [0,1599999]
print('\nOriginal :')
print(X[index])
print('\nTokenized :')
print(X_tokenized[index])
print('\nStopwords removed :')
print(X_without_stopwords[index])
print('\nPOS tagged :')
print(nltk.pos_tag(X_without_stopwords[index]))
print('\nLemmatized :')
print(X_lemmatized[index])
print('\nClean :')
print(X_clean[index])
print('\nVectorized :')
print(X_clean_vectorized[index])
print('\nPadded :')
print(X_clean_pad[index])
# Load Pre-trained Word embeddings
# Update file location of glove.6B.100d.txt according to your working environment
embedding_path = 'glove.6B.100d.txt'
embeddings_index = dict()
f = open(embedding_path)
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
# Create Word Embedding Matrix
embedding_matrix = np.zeros((max_vocab, 100))
for i in range(1,max_vocab):
embedding_vector = embeddings_index.get(tokenizer.index_word[i])
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# Define Classification model
np.random.seed(123)
model_2 = keras.models.Sequential(name='Model_2(Sentiment140_Preprocessed + GloVe 100D)')
model_2.add(InputLayer(input_shape=(20,),name='Integer_Encoding_after_Preprocessing'))
model_2.add(Embedding(max_vocab, 100, weights=[embedding_matrix], input_length=max_length,mask_zero=True,
name='Pretrained_GloVe_100D', trainable=False))
model_2.add(Bidirectional(LSTM(64,name='LSTM'),name='Bidirectional_RNN'))
model_2.add(Dropout(0.5,name='Regularizer'))
model_2.add(Dense(1, activation='sigmoid',name='Sigmoid_Classifier'))
# Compile the Model
model_2.compile(optimizer=keras.optimizers.adam(lr=0.001),loss='binary_crossentropy',
metrics=['accuracy'])
# Model details
model_2.summary()
keras.utils.plot_model(model_2,to_file='model_2.png')
# Perform train-test split
np.random.seed(123)
X_train, X_test, Y_train, Y_test = train_test_split (X_clean_pad,Y.reshape(Y.shape[0],1),test_size=0.05)
print(X_train.shape)
print(X_test.shape)
print(Y_train.shape)
print(Y_test.shape)
# Train the model
history_model_2 = model_2.fit(X_train,Y_train,batch_size=1024,epochs=15,validation_split=0.05)
# Evaluate model on Test data
model_2.evaluate(X_test,Y_test)
```
#**COMPARING BOTH MODELS**
###Model 1: Unprocessed data
###Model 2: Pre-processed data with GloVe Word Embeddings
```
# Calculate No. of epochs
n_epochs=len(history_model_1.history['loss'])
e = list(range(1,n_epochs+1))
# Training Loss
plt.plot(e,history_model_1.history['loss'],'r',label='Model 1')
plt.plot(e,history_model_2.history['loss'],'b',label='Model 2')
plt.xlabel('Epochs')
plt.ylabel('Training Loss')
plt.legend()
plt.show()
# Validation Loss
plt.plot(e,history_model_1.history['val_loss'],'r',label='Model 1')
plt.plot(e,history_model_2.history['val_loss'],'b',label='Model 2')
plt.xlabel('Epochs')
plt.ylabel('Validation Loss')
plt.legend()
plt.show()
# Training Accuracy
plt.plot(e,history_model_1.history['acc'],'r',label='Model 1')
plt.plot(e,history_model_2.history['acc'],'b',label='Model 2')
plt.xlabel('Epochs')
plt.ylabel('Training Accuracy')
plt.legend()
plt.show()
# Validation Accuracy
plt.plot(e,history_model_1.history['val_acc'],'r',label='Model 1')
plt.plot(e,history_model_2.history['val_acc'],'b',label='Model 2')
plt.xlabel('Epochs')
plt.ylabel('Validation Accuracy')
plt.legend()
plt.show()
```
| github_jupyter |
```
# Import all the necessary files!
import os
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras import Model
%matplotlib inline
# Download the inception v3 weights
#!wget --no-check-certificate \
# https://storage.googleapis.com/mledu-datasets/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5 \
# -O /tmp/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5
# Import the inception model
from tensorflow.keras.applications.inception_v3 import InceptionV3
# Create an instance of the inception model from the local pre-trained weights
local_weights_file = './tmp/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5'
pre_trained_model = InceptionV3(input_shape = (150, 150, 3),
include_top = False,
weights = None)
pre_trained_model.load_weights(local_weights_file)
# Make all the layers in the pre-trained model non-trainable
for layer in pre_trained_model.layers:
layer.trainable = False
# Print the model summary
pre_trained_model.summary()
# Expected Output is extremely large, but should end with:
#batch_normalization_v1_281 (Bat (None, 3, 3, 192) 576 conv2d_281[0][0]
#__________________________________________________________________________________________________
#activation_273 (Activation) (None, 3, 3, 320) 0 batch_normalization_v1_273[0][0]
#__________________________________________________________________________________________________
#mixed9_1 (Concatenate) (None, 3, 3, 768) 0 activation_275[0][0]
# activation_276[0][0]
#__________________________________________________________________________________________________
#concatenate_5 (Concatenate) (None, 3, 3, 768) 0 activation_279[0][0]
# activation_280[0][0]
#__________________________________________________________________________________________________
#activation_281 (Activation) (None, 3, 3, 192) 0 batch_normalization_v1_281[0][0]
#__________________________________________________________________________________________________
#mixed10 (Concatenate) (None, 3, 3, 2048) 0 activation_273[0][0]
# mixed9_1[0][0]
# concatenate_5[0][0]
# activation_281[0][0]
#==================================================================================================
#Total params: 21,802,784
#Trainable params: 0
#Non-trainable params: 21,802,784
last_layer = pre_trained_model.get_layer('mixed7')
print('last layer output shape: ', last_layer.output_shape)
last_output = last_layer.output
# Expected Output:
# ('last layer output shape: ', (None, 7, 7, 768))
# Define a Callback class that stops training once accuracy reaches 99.9%
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('acc')>0.999):
print("\nReached 99.9% accuracy so cancelling training!")
self.model.stop_training = True
from tensorflow.keras.optimizers import RMSprop
# Flatten the output layer to 1 dimension
x = layers.Flatten()(last_output)
# Add a fully connected layer with 1,024 hidden units and ReLU activation
x = layers.Dense(1024, activation='relu')(x)
# Add a dropout rate of 0.2
x = layers.Dropout(0.2)(x)
# Add a final sigmoid layer for classification
x = layers.Dense(1, activation='sigmoid')(x)
model = Model(pre_trained_model.input, x)
model.compile(optimizer = RMSprop(lr=0.0001),
loss = 'binary_crossentropy',
metrics = ['acc'])
model.summary()
# Expected output will be large. Last few lines should be:
# mixed7 (Concatenate) (None, 7, 7, 768) 0 activation_248[0][0]
# activation_251[0][0]
# activation_256[0][0]
# activation_257[0][0]
# __________________________________________________________________________________________________
# flatten_4 (Flatten) (None, 37632) 0 mixed7[0][0]
# __________________________________________________________________________________________________
# dense_8 (Dense) (None, 1024) 38536192 flatten_4[0][0]
# __________________________________________________________________________________________________
# dropout_4 (Dropout) (None, 1024) 0 dense_8[0][0]
# __________________________________________________________________________________________________
# dense_9 (Dense) (None, 1) 1025 dropout_4[0][0]
# ==================================================================================================
# Total params: 47,512,481
# Trainable params: 38,537,217
# Non-trainable params: 8,975,264
# Get the Horse or Human dataset
#!wget --no-check-certificate https://storage.googleapis.com/laurencemoroney-blog.appspot.com/horse-or-human.zip -O ./tmp/horse-or-human.zip
# Get the Horse or Human Validation dataset
#!wget --no-check-certificate https://storage.googleapis.com/laurencemoroney-blog.appspot.com/validation-horse-or-human.zip -O ./tmp/validation-horse-or-human.zip
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import zipfile
local_zip = './/tmp/horse-or-human.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('./tmp/horse-or-human/training')
zip_ref.close()
local_zip = './/tmp/validation-horse-or-human.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('./tmp/horse-or-human/validation')
zip_ref.close()
train_horses_dir = './tmp/horse-or-human/training/horses'
train_humans_dir = './tmp/horse-or-human/training/humans'
validation_horses_dir = './tmp/horse-or-human/validation/horses'
validation_humans_dir = './tmp/horse-or-human/validation/humans'
train_horses_fnames = os.listdir(train_horses_dir)
train_humans_fnames = os.listdir(train_humans_dir)
validation_horses_fnames = os.listdir(validation_horses_dir)
validation_humans_fnames = os.listdir(validation_humans_dir)
print(len(train_horses_fnames))
print(len(train_humans_fnames))
print(len(validation_horses_fnames))
print(len(validation_humans_fnames))
# Expected Output:
# 500
# 527
# 128
# 128
# Define our example directories and files
train_dir = './tmp/horse-or-human/training'
validation_dir = './tmp/horse-or-human/validation'
# Add our data-augmentation parameters to ImageDataGenerator
train_datagen = ImageDataGenerator(rescale = 1./255.,
rotation_range = 40,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator( rescale = 1.0/255. )
# Flow training images in batches of 20 using train_datagen generator
train_generator = train_datagen.flow_from_directory(train_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (150, 150))
# Flow validation images in batches of 20 using test_datagen generator
validation_generator = test_datagen.flow_from_directory( validation_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (150, 150))
# Expected Output:
# Found 1027 images belonging to 2 classes.
# Found 256 images belonging to 2 classes.
# Run this and see how many epochs it should take before the callback
# fires, and stops training at 99.9% accuracy
# (It should take less than 100 epochs)
callbacks = myCallback()
history = model.fit_generator(
train_generator,
validation_data = validation_generator,
epochs = 50,
verbose = 2,
callbacks = [callbacks])
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend(loc=0)
plt.figure()
plt.show()
```
| github_jupyter |
## Using machine learning to classify phase behavior of a surfactant/oil system
#### Tongzhou Zeng
#### Hildebrand Department of Petroleum and Geosystems Engineering, Cockrell School of Engineering
### Subsurface Machine Learning Course, The University of Texas at Austin
#### Hildebrand Department of Petroleum and Geosystems Engineering, Cockrell School of Engineering
#### Department of Geological Sciences, Jackson School of Geosciences
_____________________
Workflow supervision and review by:
#### Instructor: Prof. Michael Pyrcz, Ph.D., P.Eng., Associate Professor, The Univeristy of Texas at Austin
##### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#### Course TA: Jack Xiao, Graduate Student, The University of Texas at Austin
##### [Twitter](https://twitter.com/jackxiao6) | [LinkedIn](https://www.linkedin.com/in/yuchen-jack-xiao-b1b20876/)
### Executive Summary
* Phase behavior is essential in chemical EOR process. A intermediate microemulsion layer is desired when mixing oil with surfactant solution.
* However, investigating phase behavior is very tedious and the selection of chemical for testing is 100% experience based.
* For this workflow, a machine learning classifier is trained to predict the phase behavior result (show microemulsion vs non-microemulsion) given oil and surfactant compositions, with a P-R AUC around 0.8.
* We can use this model to get an idea of possible outcome before performing the tedious lab screening process.
### Import Packages
```
# first import machine learning libraries used in almost any workflows
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from sklearn.decomposition import PCA # PCA for feature building
from sklearn.cluster import KMeans # K-mean clustering for feature building
from sklearn.model_selection import train_test_split # split training/test set
from sklearn.linear_model import LogisticRegression # LR method
from sklearn.ensemble import RandomForestClassifier # RF method
from sklearn.preprocessing import StandardScaler, MinMaxScaler # normalization
from sklearn.model_selection import GridSearchCV # for hyperparameter tuning
from sklearn import metrics # for plotting the model results
```
### Functions
for plotting the results of a model.
For this work, two evaluation metrics are mainly considered: the **accuracy** and the **PR AUC**. Since this is a binary classification, the accuracy is what I care. However, even though my y-label is not very unbalanced (13:7), accuracy will be biased as long as the y-label is not perfectly balanced. Therefore, the AUC of the precision-recall curve is also taken into account. Here I choose PR AUC over ROC AUC, because I do have a preference of true positives over true negatives in this context. I choose PR AUC over precision only or recall only, because I do care both type I and type II errors.
```
def present_result(y_true, y_pred, y_pred_proba):
# print out the results of evaluation metrics
# y_true is the true label
# y_pred is the predicted label
# y_pred_proba is the probability of predicting 1
cm = metrics.confusion_matrix(y_true, y_pred)
#print(cm)
fig,ax=plt.subplots()
sns.heatmap(cm,
annot=True, fmt='g',
cmap="Blues")
ax.set_ylim([0,2])
ax.set_xlabel('Predicted')
ax.set_ylabel('True label')
print('accuracy: ', np.round((cm[0][0] + cm[1][1]) / np.sum(np.sum(cm)),3))
#print('precision: ', cm[1][1]/(cm[0][1] + cm[1][1]) )
#print('recall: ', cm[1][1]/(cm[1][0] + cm[1][1]) )
#fpr, tpr, thresholds = metrics.roc_curve(y_vali, y_predprob)
#print('ROC AUC: ', metrics.auc(fpr, tpr))
precision, recall, thresholds = metrics.precision_recall_curve(y_true, y_pred_proba)
print('PR AUC: ', np.round(metrics.auc(recall, precision),3))
```
### Load Data
The data set used for the following workflow is parsed from the phase behavior database build by Leonard Chang. The dataset will NOT be released.
We will work with the following tables:
* **ML_FinalProject_F20_Tongzhou Zeng_data_1.csv** - Parsed phase behavior data, just including oil name, surfactant composition and concentrations.
* **ML_FinalProject_F20_Tongzhou Zeng_data_2.csv** - The property for each of the oil
* **ML_FinalProject_F20_Tongzhou Zeng_data_3.csv** - The property for each of the surfactant/co-solvents
Note: In this workflow, all three tables are going to be further processed. The processed tables are also included when submitting this project for easy access, but they will be generated during the workflow.
```
df_pb = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_1.csv',index_col = 0)
df_oil = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_2.csv',index_col = 0)
df_chem = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_3.csv', index_col = 0)
```
We first visualize each of the tables.
For **chemical_properties** table, it contains 263 rows and 10 columns.
* **issurf** - 1 for surfactant, 0 for co-solvents
* **NC** - number of carbon in the chain
* **NEO** - number of EO group in the molecule
* **NPO** - number of PO group in the molecule
* **HG_XXX** - One hot encoded for headgroup
* **cnt** - occurance frequency
```
df_chem.head()
```
For **oil_property** table, it contains 161 rows and 6 columns.
* **MW** - Molar weight of oil, g/mol
* **EACN** - Effective ACN for oil
* **Vm** - molar volume, cm3/mol
* **density** - density, g/cm3
* **TAN** - Total acid number
* **active** - 1 for active, 0 for non-active
```
df_oil.head()
```
For **phase_behavior** table, it contains 1990 rows and 5 columns.
* **Cat** - Column for response variable, 1 for showing microemulsion, 0 for not showing microemulsion
* **Formulation** - chemical composition and their individual weight concentration, separated by #.
* **Oil** - name of oil
```
df_pb.head()
```
We need to look at the distribution of y-labels.
```
df_pb['Cat'].value_counts()
```
We see that the negative to positive ratio is 13:7. It is relatively balanced, so NO up-sampling or down-sampling will be performed.
The dataset has no features in it, so I have to manually create features. The workflow contains 4 parts, which are:
* **Creating oil features**
* **Creating chemical features**
* **Train models**
* **Hyperparameter tuning**
## 1. Creating oil features
First of all, there are missing values in the oil property table. Since I cannot find any clue of the missing ones, I will just use the mean value of the column to impute it.
```
df_oil=df_oil.fillna(df_oil.mean())
df_oil.describe()
```
Then we plot the correlation matrix to see if the features are correlated or not.
```
corr_matrix = np.corrcoef(df_oil, rowvar = False)
print(np.around(corr_matrix,2)) # print the correlation matrix to 2 decimals
```
We can see that the features are highly correlated, also we noticed that the Vm column is simply the ratio of MW over density. So to avoid multicolinearlity, I am going to use **PCA** to reduce its dimension. The data will be normalized prior to PCA.
```
# normalizing features
SC = StandardScaler()
x = SC.fit_transform(df_oil)
#performing PCA
n_components = 6
pca = PCA(n_components=n_components)
pca.fit(x)
```
Now we plot the variance explained for each principal component
```
var_exp = np.round(pca.explained_variance_ratio_,3)
plt.bar(range(1, 7) ,var_exp,
color="g", align="center")
plt.xticks(range(1, 7))
plt.title("Variance explained for i-th principal component")
print('Variance explained 6 principle components =', var_exp)
```
We can see that the first 3 components explained 90% variance, so the first three components will be used as oil features.
```
x_trans = pca.transform(x)
x_pca_origin = SC.inverse_transform(x_trans)
for i in range(3):
df_oil['PCA_' + str(i + 1)] = x_pca_origin[:, i]
df_oil.to_csv('ML_FinalProject_F20_Tongzhou Zeng_data_2_processed.csv')
```
## 2. Creating chemical features
For the chemicals, the features are all related to the molecular structure of the particles. These features are controlled by manufacturing processes, so these features are relatively independent. We first plot the histogram of the frequency of each chemical.
```
sns.displot(df_chem['cnt'], bins=20)
```
We see that the distribution is highly skewed, which means that half of the chemicals occured less than 5 times in the dataset, while several chemicals are extremely popular. I noticed that the molecue structure difference between some chemicals are small, often within the error range of the manufacturing process.
Given the two reasons above, small difference between items but relative independent features, I will use **K-means clustering** to decrease the dimensions of chemicals. Also, by domain knowledge, surfactant and co-solvents will be treated separately
```
df_surf = df_chem[df_chem['issurf']==1]
df_cosolv = df_chem[df_chem['issurf']==0]
```
Like above, the features are first normalized
```
norm_scaler = MinMaxScaler()
X_surf = norm_scaler.fit_transform(df_surf.iloc[:,1:])
df_surf.iloc[:,1:] = X_surf
df_surf.describe()
```
Now we perform K-means clustering, and plot the inertia vs number of clusters
```
center_num = list(range(10,100,5))
inertia = []
for i in center_num:
kmeans = KMeans(n_clusters=i).fit(X_surf)
inertia.append(kmeans.inertia_)
plt.plot(center_num, inertia)
plt.title('K-Mean clustering for surfactant')
plt.xlabel('Number of clusters')
plt.ylabel('Inertia')
```
There is no obvious elbow in this plot. Here I pick K=80 because it gives low enough inertia, so that I do not lose too much resolution.
```
center_num=80
kmeans = KMeans(n_clusters=center_num).fit(X_surf)
df_surf['label'] = kmeans.labels_
df_surf.head()
```
Similar for co-solvents. For cosolvents, only NC and EO have non-zero values, so I only consider those two columns.
```
norm_scaler = MinMaxScaler()
X_cosolv = norm_scaler.fit_transform(df_cosolv.iloc[:,[1,2]])
center_num = list(range(2,30,2))
inertia = []
for i in center_num:
kmeans = KMeans(n_clusters=i).fit(X_cosolv)
inertia.append(kmeans.inertia_)
plt.plot(center_num, inertia)
plt.title('K-Mean clustering for co-solvents')
plt.xlabel('Number of clusters')
plt.ylabel('Inertia')
```
Here I see an elbow point around K=12, so K=12 will be used.
```
center_num=12
kmeans = KMeans(n_clusters=center_num).fit(X_cosolv)
df_cosolv['label'] = kmeans.labels_ + 80
df_cosolv.head()
```
Then we concatenate this two dataframes, and save the clustered features.
```
df_chem = pd.concat([df_surf, df_cosolv])
df_chem.head()
df_chem.to_csv('ML_FinalProject_F20_Tongzhou Zeng_data_3_processed.csv')
```
## 3. Train models
### 3.1 Fill in features
We successfully extract the features, so now we are going to build the dataset for training. We first reload the processed property tables, and view their first 5 rows.
```
df_pb = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_1.csv',index_col = 0)
df_oil = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_2_processed.csv',index_col = 0)
df_chem = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_3_processed.csv', index_col = 0)
df_chem.head()
df_oil.head()
df_pb.head()
```
Next for the main phase behavior dataframe, we need to allocate space for features. My designed features are 3 dense features describing the propeties of oil, and 92 sparse features charaterizing the concentration of each chemical cluster.
```
for i in range(3): # for oil we have 3 features
df_pb['oil_pca_' + str(i + 1)] = 0
for j in range(92): # for chemical we have 80+12=92 clusters
df_pb['chemical_cluster_' + str(j)] = 0
df_pb.head()
```
Then we iterate each row to fill in blanks. For chemicals, we put the total concentration for each cluster.
```
for i in range(len(df_pb)):
# for oil
oil_name = df_pb.loc[i, 'Oil'] # find the name of the oil
for j in range(3):
# for each principal component, find the cell in the oil_pca table with the oil name
df_pb.loc[i, 'oil_pca_' + str(j + 1)] = df_oil.loc[oil_name, 'PCA_' + str(j + 1)]
# for chemical
chem_list = df_pb.loc[i, 'Formulation'].split('#') # it will be a list of [conc1, name1, conc2, name2....]
num = len(chem_list)//2 # find out how many different kind of chemicals
flag = False
for j in range(1, num + 1): # for each of the chemical included
idx1 = 2 * (j - 1) # index for its concentration in the chem_list
idx2 = 2 * (j - 1) + 1 # index for its name in the chem_list
concentrationj = float(chem_list[idx1][:-1]) # extract its concentration, [:-1] means getting rid of "%"
chemj = chem_list[idx2] # extract its name
clus = df_chem.loc[chemj, 'label'] # use the name to find which cluster it belongs to
df_pb.loc[i, 'chemical_cluster_' + str(int(clus))] += concentrationj # we add the concentration in the cell
df_pb.head()
```
we save the data frame in case we do not want to redo it.
```
df_pb.to_csv('ML_FinalProject_F20_Tongzhou Zeng_data_1_processed.csv')
```
### 3.2 Model Training
Finally its time to train model! We load the dataset with features.
```
df = pd.read_csv('ML_FinalProject_F20_Tongzhou Zeng_data_1_processed.csv',index_col = 0)
df.head()
```
Separate the predictor and response
```
X = df.iloc[:, 3:]
y = df.iloc[:, 0]
```
Train test split
```
X_tr, X_test, y_tr, y_test = train_test_split(X, y, test_size = 0.2, random_state = 19961012)
#X_train, X_vali, y_train, y_vali = train_test_split(X_tr, y_tr, test_size = 0.25, random_state = 19961012)
```
We first try a logistic regression model as our base case.
```
classifier = LogisticRegression()
# for logistic regression we need to normalize the features
norm_scaler = MinMaxScaler()
X_tr_n = norm_scaler.fit_transform(X_tr)
X_test_n = norm_scaler.transform(X_test)
classifier.fit(X_tr_n, y_tr)
y_pred = classifier.predict(X_test_n)
y_predprob = classifier.predict_proba(X_test_n)[:,1]
present_result(y_test, y_pred, y_predprob)
```
We see it does not work well. It almost predict every thing 0. But logistic regression is just a basecase. The simple linear classifier will underfit this complicated dataset without tedious and detailed feature engineering process. Next I will try random forest, another popular classification algorithm. Random forest generally works good for the data size on this order by experience. The tree model generally have low bias and its double randomness (bootstrap + random feature selection) will reduce the variance.
```
classifier = RandomForestClassifier()
# for random forest, we do not need to normalize features
classifier.fit(X_tr, y_tr)
y_pred = classifier.predict(X_test)
y_predprob = classifier.predict_proba(X_test)[:,1]
present_result(y_test, y_pred, y_predprob)
```
We can see random forest works much better than logistic regression.
## 4. Hyperparameter Tuning
Here I tried hyperparameter tuning using cross validation (GridSearchCV package) on the training set. The process is tedious, the code is pretty repeatitive, and the execution time is long. Here I just show the final model after hyperparameter tuning.
```
# after CV hyperparameter tuning on training set
classifier = RandomForestClassifier(criterion = 'entropy', max_depth=19, min_samples_split=4, n_estimators=125, oob_score=True)
# for random forest, we do not need to normalize features
classifier.fit(X_tr, y_tr)
y_pred = classifier.predict(X_test)
y_predprob = classifier.predict_proba(X_test)[:,1]
present_result(y_test, y_pred, y_predprob)
```
After hyperparameter tuning, the accuracy and PR AUC are around 0.8, which is much better than random guess.
In this project, the raw data is not as nice as that of homework problems, which already give numerical values of every predictor variable. For this project, the raw data is in a "relational database"-like structure. This workflow demonstrates the process of solving this problem from scratch, including data cleaning, data preprocessing, feature building, model training, metrics selection, and hyperparameter tuning. For the final random forest model, the accuracy and PR AUC are around 0.8, which is pretty good for my purpose.
I hope this was helpful,
*Tongzhou Zeng*
___________________
#### Work Supervised by:
### Michael Pyrcz, Associate Professor, University of Texas at Austin
*Novel Data Analytics, Geostatistics and Machine Learning Subsurface Solutions*
With over 17 years of experience in subsurface consulting, research and development, Michael has returned to academia driven by his passion for teaching and enthusiasm for enhancing engineers' and geoscientists' impact in subsurface resource development.
For more about Michael check out these links:
#### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#### Want to Work Together?
I hope this content is helpful to those that want to learn more about subsurface modeling, data analytics and machine learning. Students and working professionals are welcome to participate.
* Want to invite me to visit your company for training, mentoring, project review, workflow design and / or consulting? I'd be happy to drop by and work with you!
* Interested in partnering, supporting my graduate student research or my Subsurface Data Analytics and Machine Learning consortium (co-PIs including Profs. Foster, Torres-Verdin and van Oort)? My research combines data analytics, stochastic modeling and machine learning theory with practice to develop novel methods and workflows to add value. We are solving challenging subsurface problems!
* I can be reached at mpyrcz@austin.utexas.edu.
I'm always happy to discuss,
*Michael*
Michael Pyrcz, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
| github_jupyter |
Licensed under the MIT License.
# Activity2Vec: Learning ADL Embeddings from Sensor Data with a Sequence-to-Sequence Model
This notebook provides an example of how to train an Act2Vec model to extract features. We will train an Act2Vec based model on the describe in the paper refrence blow.
In this notebook, we will train an Act2Vec model on UCI-HAR data, and then employ a random forest as our classifier.
Reference: https://arxiv.org/abs/1907.05597
```
# Import packages
import os
import sys
sys.path.insert(0, os.path.abspath('../'))
import tensorflow as tf
import numpy as np
from datasets.har import load_data
from model.act2vec import Act2Vec
from util.utils import plot_latent_space
from util.utils import rf
from util.utils import plot_confusion_matrix
from util.utils import print_result
# allow tenserflow to use GPU
physical_devices = tf.config.experimental.list_physical_devices('GPU')
assert len(physical_devices) > 0, "Not enough GPU hardware devices available"
tf.config.experimental.set_memory_growth(physical_devices[0], True)
os.environ['AUTOGRAPH_VERBOSITY'] = '10'
tf.autograph.set_verbosity(0)
```
## UCI-HAR dataset
Let's start with downloading dataset, and load the datasets.
```
(x_train, y_train), (x_test, y_test) = load_data()
print('x_train shape is: ', x_train.shape)
print('x_test shape is: ', x_test.shape)
print('Number of classes: ', len(np.unique(y_train)))
```
## Encode the data by employing Act2Vec model
### Building an Act2Vec Model
```
act2vec = Act2Vec(units=128, input_dim=x_train.shape)
opt = tf.keras.optimizers.Adam(lr=2e-5, decay=2e-11)
act2vec.compile(loss='mse',
optimizer=opt,
metrics=['mse'])
# train the act2vec moedl
act2vec.fit(x_train,x_train,
batch_size=32,
epochs=4000)
act2vec.save_weights('HAR-act2vec_model.h5')
```
### After taining the act2vec model, we can use it to extract fetures
```
X_train = act2vec.encoder(x_train)
X_test = act2vec.encoder(x_test)
print('Encoded X_train shape is: ', X_train.shape)
print('Encoded X_test shape is: ', X_test.shape)
plot_latent_space(X_train, y_train, 'UCI_HAR_Xtrain_latentspace')
```
### Train random forest on encoded data
```
random_forest_en = rf(X_train, y_train, n_estimators=100)
print_result(random_forest_en, X_train, y_train, X_test, y_test)
LABELS = ['WALKING', 'WALKING_UPSTAIRS', 'WALKING_DOWNSTAIRS', 'SITTING', 'STANDING', 'LAYING']
plot_confusion_matrix(random_forest_en, X_test, y_test, class_names=LABELS, file_name='HAR_confusionmatrix_en', normalize=True)
```
### Train random forest on raw data
```
random_forest_raw = rf(x_train.reshape(X_train.shape[0], np.prod(x_train.shape[1:])), y_train, n_estimators=100)
print_result(random_forest_raw, x_train.reshape(x_train.shape[0], np.prod(x_train.shape[1:])), y_train, x_test.reshape(x_test.shape[0], np.prod(x_test.shape[1:])), y_test)
LABELS = ['WALKING', 'WALKING_UPSTAIRS', 'WALKING_DOWNSTAIRS', 'SITTING', 'STANDING', 'LAYING']
plot_confusion_matrix(random_forest_raw, x_test.reshape(x_test.shape[0], np.prod(x_test.shape[1:])), y_test, class_names=LABELS, file_name='HAR_confusionmatrix_raw', normalize=True)
```
| github_jupyter |
# [Module 3.1] Train Script 생성
이 노트북은 추후 사용할 학습 스크립트를 작성 합니다.
## Write a train script
```
%%writefile tf_script_bert_tweet.py
import time
import random
import pandas as pd
from glob import glob
import pprint
import argparse
import json
import subprocess
import sys
import os
import tensorflow as tf
#subprocess.check_call([sys.executable, '-m', 'pip', 'install', 'tensorflow==2.1.0'])
subprocess.check_call([sys.executable, '-m', 'pip', 'install', 'transformers==2.8.0'])
subprocess.check_call([sys.executable, '-m', 'pip', 'install', 'sagemaker-tensorflow==2.1.0.1.0.0'])
subprocess.check_call([sys.executable, '-m', 'pip', 'install', 'scikit-learn==0.23.1'])
from transformers import DistilBertTokenizer
from transformers import TFDistilBertForSequenceClassification
from transformers import TextClassificationPipeline
from transformers.configuration_distilbert import DistilBertConfig
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.models import load_model
####################
# 기존 소스에 변경 사항
# Define 10 labels
####################
CLASSES = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
###################
# TF 에 입력될 Data Set 생성 함수
# BERT-related Function
####################
def select_data_and_label_from_record(record):
x = {
'input_ids': record['input_ids'],
'input_mask': record['input_mask'],
'segment_ids': record['segment_ids']
}
y = record['label_ids']
return (x, y)
# BERT-related Function
def file_based_input_dataset_builder(channel,
input_filenames,
pipe_mode,
is_training,
drop_remainder,
batch_size,
epochs,
steps_per_epoch,
max_seq_length):
# For training, we want a lot of parallel reading and shuffling.
# For eval, we want no shuffling and parallel reading doesn't matter.
if pipe_mode:
print('***** Using pipe_mode with channel {}'.format(channel))
from sagemaker_tensorflow import PipeModeDataset
dataset = PipeModeDataset(channel=channel,
record_format='TFRecord')
else:
print('***** Using input_filenames {}'.format(input_filenames))
dataset = tf.data.TFRecordDataset(input_filenames)
dataset = dataset.repeat(epochs * steps_per_epoch)
dataset = dataset.prefetch(tf.data.experimental.AUTOTUNE)
name_to_features = {
"input_ids": tf.io.FixedLenFeature([max_seq_length], tf.int64),
"input_mask": tf.io.FixedLenFeature([max_seq_length], tf.int64),
"segment_ids": tf.io.FixedLenFeature([max_seq_length], tf.int64),
"label_ids": tf.io.FixedLenFeature([], tf.int64),
}
def _decode_record(record, name_to_features):
"""Decodes a record to a TensorFlow example."""
record = tf.io.parse_single_example(record, name_to_features)
# TODO: wip/bert/bert_attention_head_view/train.py
# Convert input_ids into input_tokens with DistilBert vocabulary
# if hook.get_collections()['all'].save_config.should_save_step(modes.EVAL, hook.mode_steps[modes.EVAL]):
# hook._write_raw_tensor_simple("input_tokens", input_tokens)
return record
dataset = dataset.apply(
tf.data.experimental.map_and_batch(
lambda record: _decode_record(record, name_to_features),
batch_size=batch_size,
drop_remainder=drop_remainder,
num_parallel_calls=tf.data.experimental.AUTOTUNE))
dataset.cache()
if is_training:
dataset = dataset.shuffle(seed=42,
buffer_size=1000,
reshuffle_each_iteration=True)
return dataset
def load_checkpoint_model(checkpoint_path):
'''
Load the last checkpoint file
'''
import glob
import os
glob_pattern = os.path.join(checkpoint_path, '*.h5')
print('glob pattern {}'.format(glob_pattern))
list_of_checkpoint_files = glob.glob(glob_pattern)
print('List of checkpoint files {}'.format(list_of_checkpoint_files))
latest_checkpoint_file = max(list_of_checkpoint_files)
print('Latest checkpoint file {}'.format(latest_checkpoint_file))
initial_epoch_number_str = latest_checkpoint_file.rsplit('_', 1)[-1].split('.h5')[0]
initial_epoch_number = int(initial_epoch_number_str)
loaded_model = TFDistilBertForSequenceClassification.from_pretrained(
latest_checkpoint_file,
config=config)
print('loaded_model {}'.format(loaded_model))
print('initial_epoch_number {}'.format(initial_epoch_number))
return loaded_model, initial_epoch_number
def parse_args():
'''
커멘드 라인 변수를 읽음. 상당수는 환경 변수의 값을 디폴트로 설정 함
'''
parser = argparse.ArgumentParser()
parser.add_argument('--train_data',
type=str,
default=os.environ['SM_CHANNEL_TRAIN'])
parser.add_argument('--validation_data',
type=str,
default=os.environ['SM_CHANNEL_VALIDATION'])
parser.add_argument('--test_data',
type=str,
default=os.environ['SM_CHANNEL_TEST'])
parser.add_argument('--output_dir',
type=str,
default=os.environ['SM_OUTPUT_DIR'])
parser.add_argument('--hosts',
type=list,
default=json.loads(os.environ['SM_HOSTS']))
parser.add_argument('--current_host',
type=str,
default=os.environ['SM_CURRENT_HOST'])
parser.add_argument('--num_gpus',
type=int,
default=os.environ['SM_NUM_GPUS'])
parser.add_argument('--checkpoint_base_path',
type=str,
default='/opt/ml/checkpoints')
parser.add_argument('--use_xla',
type=eval,
default=False)
parser.add_argument('--use_amp',
type=eval,
default=False)
parser.add_argument('--max_seq_length',
type=int,
default=128)
parser.add_argument('--train_batch_size',
type=int,
default=128)
parser.add_argument('--validation_batch_size',
type=int,
default=256)
parser.add_argument('--test_batch_size',
type=int,
default=256)
parser.add_argument('--epochs',
type=int,
default=1)
parser.add_argument('--learning_rate',
type=float,
default=0.00003)
parser.add_argument('--epsilon',
type=float,
default=0.00000001)
parser.add_argument('--train_steps_per_epoch',
type=int,
default=1)
parser.add_argument('--validation_steps',
type=int,
default=1)
parser.add_argument('--test_steps',
type=int,
default=1)
parser.add_argument('--freeze_bert_layer',
type=eval,
default=False)
parser.add_argument('--run_validation',
type=eval,
default=False)
parser.add_argument('--run_test',
type=eval,
default=False)
parser.add_argument('--run_sample_predictions',
type=eval,
default=False)
parser.add_argument('--enable_checkpointing',
type=eval,
default=False)
parser.add_argument('--model_dir',
type=str,
default=os.environ['SM_MODEL_DIR'])
return parser.parse_args()
if __name__ == '__main__':
# This points to the S3 location - this should not be used by our code
# We should use /opt/ml/model/ instead
# parser.add_argument('--model_dir',
# type=str,
# default=os.environ['SM_MODEL_DIR'])
args = parse_args()
print("################ Args: #######################")
print(args)
env_var = os.environ
print("################ Environment Variables: ################")
pprint.pprint(dict(env_var), width = 1)
print('SM_TRAINING_ENV {}'.format(env_var['SM_TRAINING_ENV']))
sm_training_env_json = json.loads(env_var['SM_TRAINING_ENV'])
is_master = sm_training_env_json['is_master']
print("################ Extract varaibles from Command Args #######################")
train_data = args.train_data
print('train_data {}'.format(train_data))
validation_data = args.validation_data
print('validation_data {}'.format(validation_data))
test_data = args.test_data
print('test_data {}'.format(test_data))
# local_model_dir = args.model_dir
local_model_dir = os.environ['SM_MODEL_DIR']
print('local_model_dir {}'.format(local_model_dir))
output_dir = args.output_dir
print('output_dir {}'.format(output_dir))
hosts = args.hosts
print('hosts {}'.format(hosts))
current_host = args.current_host
print('current_host {}'.format(current_host))
num_gpus = args.num_gpus
print('num_gpus {}'.format(num_gpus))
job_name = os.environ['SAGEMAKER_JOB_NAME']
print('job_name {}'.format(job_name))
use_xla = args.use_xla
print('use_xla {}'.format(use_xla))
use_amp = args.use_amp
print('use_amp {}'.format(use_amp))
max_seq_length = args.max_seq_length
print('max_seq_length {}'.format(max_seq_length))
train_batch_size = args.train_batch_size
print('train_batch_size {}'.format(train_batch_size))
validation_batch_size = args.validation_batch_size
print('validation_batch_size {}'.format(validation_batch_size))
test_batch_size = args.test_batch_size
print('test_batch_size {}'.format(test_batch_size))
epochs = args.epochs
print('epochs {}'.format(epochs))
learning_rate = args.learning_rate
print('learning_rate {}'.format(learning_rate))
epsilon = args.epsilon
print('epsilon {}'.format(epsilon))
train_steps_per_epoch = args.train_steps_per_epoch
print('train_steps_per_epoch {}'.format(train_steps_per_epoch))
validation_steps = args.validation_steps
print('validation_steps {}'.format(validation_steps))
test_steps = args.test_steps
print('test_steps {}'.format(test_steps))
freeze_bert_layer = args.freeze_bert_layer
print('freeze_bert_layer {}'.format(freeze_bert_layer))
run_validation = args.run_validation
print('run_validation {}'.format(run_validation))
run_test = args.run_test
print('run_test {}'.format(run_test))
run_sample_predictions = args.run_sample_predictions
print('run_sample_predictions {}'.format(run_sample_predictions))
enable_checkpointing = args.enable_checkpointing
print('enable_checkpointing {}'.format(enable_checkpointing))
checkpoint_base_path = args.checkpoint_base_path
print('checkpoint_base_path {}'.format(checkpoint_base_path))
print(" ################ Set Checkpoint path: ################")
if is_master:
checkpoint_path = checkpoint_base_path
else:
checkpoint_path = '/tmp/checkpoints'
print('checkpoint_path {}'.format(checkpoint_path))
# Determine if PipeMode is enabled
pipe_mode_str = os.environ.get('SM_INPUT_DATA_CONFIG', '')
pipe_mode = (pipe_mode_str.find('Pipe') >= 0)
print('Using pipe_mode: {}'.format(pipe_mode))
print ("################ Mirrored distributed_strategy ################")
#distributed_strategy = tf.distribute.MirroredStrategy()
# Comment out when using smdebug as smdebug does not support MultiWorkerMirroredStrategy() as of smdebug 0.8.0
distributed_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
with distributed_strategy.scope():
tf.config.optimizer.set_jit(use_xla)
tf.config.optimizer.set_experimental_options({"auto_mixed_precision": use_amp})
train_data_filenames = glob(os.path.join(train_data, '*.tfrecord'))
print('train_data_filenames {}'.format(train_data_filenames))
train_dataset = file_based_input_dataset_builder(
channel='train',
input_filenames=train_data_filenames,
pipe_mode=pipe_mode,
is_training=True,
drop_remainder=False,
batch_size=train_batch_size,
epochs=epochs,
steps_per_epoch=train_steps_per_epoch,
max_seq_length=max_seq_length).map(select_data_and_label_from_record)
tokenizer = None
config = None
model = None
# This is required when launching many instances at once... the urllib request seems to get denied periodically
successful_download = False
retries = 0
while (retries < 5 and not successful_download):
try:
tokenizer = DistilBertTokenizer.from_pretrained('distilbert-base-uncased')
config = DistilBertConfig.from_pretrained('distilbert-base-uncased',
num_labels=len(CLASSES))
model = TFDistilBertForSequenceClassification.from_pretrained('distilbert-base-uncased',
config=config)
successful_download = True
print('Sucessfully downloaded after {} retries.'.format(retries))
except:
retries = retries + 1
random_sleep = random.randint(1, 30)
print('Retry #{}. Sleeping for {} seconds'.format(retries, random_sleep))
time.sleep(random_sleep)
callbacks = []
initial_epoch_number = 0
if enable_checkpointing:
print('***** Checkpoint enabled *****')
os.makedirs(checkpoint_path, exist_ok=True)
if os.listdir(checkpoint_path):
print('***** Found checkpoint *****')
print(checkpoint_path)
model, initial_epoch_number = load_checkpoint_model(checkpoint_path)
print('***** Using checkpoint model {} *****'.format(model))
checkpoint_callback = ModelCheckpoint(
filepath=os.path.join(checkpoint_path, 'tf_model_{epoch:05d}.h5'),
save_weights_only=False,
verbose=1,
monitor='val_accuracy')
print('*** CHECKPOINT CALLBACK {} ***'.format(checkpoint_callback))
callbacks.append(checkpoint_callback)
if not tokenizer or not model or not config:
print('Not properly initialized...')
optimizer = tf.keras.optimizers.Adam(learning_rate=learning_rate, epsilon=epsilon)
print('** use_amp {}'.format(use_amp))
if use_amp:
# loss scaling is currently required when using mixed precision
optimizer = tf.keras.mixed_precision.experimental.LossScaleOptimizer(optimizer, 'dynamic')
print('*** OPTIMIZER {} ***'.format(optimizer))
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
metric = tf.keras.metrics.SparseCategoricalAccuracy('accuracy')
model.compile(optimizer=optimizer, loss=loss, metrics=[metric])
print('Compiled model {}'.format(model))
model.layers[0].trainable = not freeze_bert_layer
print(model.summary())
# Validation 이 True 이면 Train and Validation 진행
if run_validation:
validation_data_filenames = glob(os.path.join(validation_data, '*.tfrecord'))
print('validation_data_filenames {}'.format(validation_data_filenames))
validation_dataset = file_based_input_dataset_builder(
channel='validation',
input_filenames=validation_data_filenames,
pipe_mode=pipe_mode,
is_training=False,
drop_remainder=False,
batch_size=validation_batch_size,
epochs=epochs,
steps_per_epoch=validation_steps,
max_seq_length=max_seq_length).map(select_data_and_label_from_record)
print('Starting Training and Validation...')
validation_dataset = validation_dataset.take(validation_steps)
train_and_validation_history = model.fit(train_dataset,
shuffle=True,
epochs=epochs,
initial_epoch=initial_epoch_number,
steps_per_epoch=train_steps_per_epoch,
validation_data=validation_dataset,
validation_steps=validation_steps,
callbacks=callbacks)
print(train_and_validation_history)
else: # Not running validation
print('Starting Training (Without Validation)...')
train_history = model.fit(train_dataset,
shuffle=True,
epochs=epochs,
initial_epoch=initial_epoch_number,
steps_per_epoch=train_steps_per_epoch,
callbacks=callbacks)
print(train_history)
# run_test가 True 이면 진행
if run_test:
test_data_filenames = glob(os.path.join(test_data, '*.tfrecord'))
print('test_data_filenames {}'.format(test_data_filenames))
test_dataset = file_based_input_dataset_builder(
channel='test',
input_filenames=test_data_filenames,
pipe_mode=pipe_mode,
is_training=False,
drop_remainder=False,
batch_size=test_batch_size,
epochs=epochs,
steps_per_epoch=test_steps,
max_seq_length=max_seq_length).map(select_data_and_label_from_record)
print('Starting test...')
test_history = model.evaluate(test_dataset,
steps=test_steps,
callbacks=callbacks)
print('Test history {}'.format(test_history))
# Model Output
transformer_fine_tuned_model_path = os.path.join(local_model_dir, 'transformers/fine-tuned/')
os.makedirs(transformer_fine_tuned_model_path, exist_ok=True)
# SavedModel Output
tensorflow_saved_model_path = os.path.join(local_model_dir, 'tensorflow/saved_model/0')
os.makedirs(tensorflow_saved_model_path, exist_ok=True)
# Save the Fine-Yuned Transformers Model as a New "Pre-Trained" Model
print("######## Save the Fine-Yuned Transformers Model as a New Pre-Trained Model ##########")
print('transformer_fine_tuned_model_path {}'.format(transformer_fine_tuned_model_path))
model.save_pretrained(transformer_fine_tuned_model_path)
# Save the TensorFlow SavedModel for Serving Predictions
print("######## Save the TensorFlow SavedModel for Serving Predictions Model ##########")
print('tensorflow_saved_model_path {}'.format(tensorflow_saved_model_path))
model.save(tensorflow_saved_model_path, save_format='tf')
```
| github_jupyter |
*best viewed in [nbviewer](https://nbviewer.jupyter.org/github/CambridgeSemiticsLab/BH_time_collocations/blob/master/results/notebooks/2_function_distribution.ipynb)*
# Time Adverbial Distribution and Collocations
## Function Distribution
### Cody Kingham
<a href="../../../docs/sponsors.md"><img height=200px width=200px align="left" src="../../../docs/images/CambridgeU_BW.png"></a>
```
! echo "last updated:"; date
```
## Introduction
This notebook will produce data for my in-progress article on time adverbial components.
<hr>
# Python
Now we import the modules and data needed for the analysis.
```
# see config.py for variables
# config.py is used across the notebooks
# for the article project
from config import *
# load broad dataset for distribution analysis
phrase_df = pd.read_csv(broad_dataset)
phrase_df.set_index('node', inplace=True)
phrase_df['parsed'] = phrase_df.index.isin(functs_df.index) * 1
# define sample of phrase functions for distribution
# analysis
functions_sample = sorted([
'Adju',
'Cmpl',
'Loca',
'Objc',
'Subj',
'Time',
])
funct_sample = phrase_df[phrase_df.s_function.isin(functions_sample)]
funct_sample.shape
```
## Deviation of Proportions
Now we'll look to see how representative the datasample is across the<br>
Hebrew Bible. Have the selection requirements negatively affected any of the books?
```
# make a cross-tabulation from function to
# book, n_pa, n_heads, and n_daughters to
# enable selection of data as needed
by_book = pd.pivot_table(
funct_sample,
index='s_function',
columns=['n_phrase_atoms', 'n_heads', 'n_relas', 'parsed', 'book'],
aggfunc='size',
fill_value=0
)
# result is a multi-index df
by_book.head()
```
### Does the sample selection affect book-by-book representation of functions?
```
# exp_prop is a table of ratios which state
# what proportion of a function's total uses is represented within a given book
# the observed proportion will be that subset of phrases that fit our selection
# criteria, i.e. na=1, nh=1, nd=0
exp_freq = by_book.groupby(axis=1, level='book').sum() # group by book column
exp_prop = exp_freq.div(exp_freq.sum(1), 0)
# observed proportion is the ratio of the selected sample function's
# total representation within a given book: in this case only functions
# which have 1 atom, 1 head, 0 relas, and parse value of 1
obv_freq = by_book[(1, 1, 0, 1)] # select multindex
obv_prop = obv_freq.div(obv_freq.sum(1), 0)
# the deviation of proportions will be the DIFFERENCE between the expected ratios
# (calculated across the whole dataset) and the ratios across the selected subset
# of phrases
deviation_prop = round(exp_prop - obv_prop, 2)
exp_freq
exp_prop
obv_freq
obv_prop
deviation_prop
deviation_prop.abs().sum(0).sort_values(ascending=False)[:15]
deviation_prop.mean(0)
```
**Based on this data, we can say that the filtered selection of phrase functions<br>
does not negatively affect representation across the Hebrew Bible. The deviation<br>
of proportions is very small across both book and function categories.**
### How evenly distributed is Time compared to the other key functions? (DP)
We can answer this question using DP (Gries 2008), the Degree of Dispersion.<br>
For a given sample (e.g. Hebrew Bible), divide the sample into its various parts (books).<br>
Use a baseline (e.g. n-phrases) to calculate what proportion each part *should* account<br>
for in the dataset. And compare the observed proportion for a given variable (in this case<br>
function).
```
exp_freq.sum().sum()
# expected proportional representation of total number
# of phrases for any given book; retrieved by dividing
# a book's total phrases by the marginal total of all books' n-phrases
book_exp_props = exp_freq.sum(0) / exp_freq.sum(0).sum()
# observed proportional representation for any given book
# of a given function's total occurrences
book_obv_prop = exp_freq.div(exp_freq.sum(1), 0)
# calculate the deviation of proportion of the observed
# proportional representation from the expected proportion
book_dev_prop = book_obv_prop.sub(book_exp_props, 1)
# calculate DP score following Gries 2008 by summing absolute
# value of all deviations per function and dividing by 2
# the resulting score ranges from 0 to 1 where 0 is the
# hypothetical perfectly distributed function and 1 is the
# hypothetical worst distributed function
book_DP = abs(book_dev_prop).sum(1) / 2
book_DP = book_DP.sort_values()
book_exp_props.sort_values(ascending=False)
book_dev_prop
book_DP
```
**Adjunct**
```
dev_min = book_dev_prop.loc[['Adju', 'Loca', 'Time']].min().min() - 0.005
dev_max = book_dev_prop.loc[['Adju', 'Loca', 'Time']].max().max() + 0.005
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20, 6))
ax = ax1
book_dev_prop.loc['Adju'].sort_values().plot(kind='bar', ax=ax, color='lightgray', edgecolor='black')
ax.axhline(0, color='black', linewidth=0.8)
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_ylabel('deviation of observed % from expected %', size=10)
ax.set_title('Adju')
ax.set_ylim(dev_min, dev_max)
ax=ax2
book_dev_prop.loc['Loca'].sort_values().plot(kind='bar', ax=ax, color='lightgray', edgecolor='black')
ax.axhline(0, color='black', linewidth=0.8)
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_ylabel('deviation of observed % from expected %', size=10)
ax.set_title('Loca')
ax.set_ylim(dev_min, dev_max)
savefig('devProp_book_Adju_Loca')
plt.show()
book_dev_prop.loc['Adju'].abs().sort_values(ascending=False)
book_dev_prop.loc['Loca'].sort_values()
fig, ax = plt.subplots(figsize=(10, 6))
book_dev_prop.loc['Time'].sort_values().plot(kind='bar', ax=ax, color='lightgray', edgecolor='black')
ax.axhline(0, color='black', linewidth=0.8)
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_ylabel('deviation of observed % from expected %', size=10)
ax.set_title('Time')
ax.set_ylim(dev_min, dev_max)
savefig('devProp_book_Time')
plt.show()
book_dev_prop.loc['Time'].abs().sort_values(ascending=False)
book_dev_prop.loc['Loca'].sum()
```
### DP by chapter
```
funct_sample2 = funct_sample.copy()
funct_sample2['chapter_node'] = [L.u(ph,'chapter')[0] for ph in funct_sample.index]
chapt_counts = pd.pivot_table(
funct_sample2,
index='s_function',
columns=['chapter_node'],
aggfunc='size',
fill_value=0,
)
chapt_counts
ch_exp_prop = chapt_counts.sum(0) / chapt_counts.sum(0).sum()
ch_obv_prop = chapt_counts.div(chapt_counts.sum(1), 0)
ch_dev_prop = ch_obv_prop.sub(ch_exp_prop, 1)
ch_DP = abs(ch_dev_prop).sum(1) / 2
ch_DP = ch_DP.sort_values()
ch_DP
```
### DP determined using clause clusters
```
# build clusters of clauses into a column that can be added DF
cluster_size = 10
cl_clusters = []
this_cluster = []
for cl in F.otype.s('clause'):
this_cluster.append(cl)
if len(this_cluster) == cluster_size:
cl_clusters.append(this_cluster)
this_cluster = []
cl_clusters.append(this_cluster) # finish the last short cluster
print(len(cl_clusters), 'clusters of size', cluster_size, 'built')
print(len(cl_clusters[-1]), 'short cluster added')
# map clustesr to sets of clauses
# that can be used for membership checks
clauses_2_id = []
for i, cluster in enumerate(cl_clusters):
clauses_2_id.append((set(cluster), i))
# make a new column for the DF that maps a phrase
# to its clause cluster group
phrase_clust_group = []
for ph in funct_sample.index:
cl = L.u(ph, 'clause')[0]
for clauses, cluster in clauses_2_id:
if cl in clauses:
phrase_clust_group.append(cluster)
continue
print(len(phrase_clust_group), 'phrases assigned to clusters')
funct_sample3 = funct_sample.copy()
funct_sample3['cl_cluster'] = phrase_clust_group
cl_counts = pd.pivot_table(
funct_sample3,
index='s_function',
columns=['cl_cluster'],
aggfunc='size',
fill_value=0,
)
cl_counts
cl_exp_prop = cl_counts.sum(0) / cl_counts.sum(0).sum()
cl_obv_prop = cl_counts.div(cl_counts.sum(1), 0)
cl_dev_prop = cl_obv_prop.sub(cl_exp_prop, 1)
cl_DP = abs(cl_dev_prop).sum(1) / 2
cl_DP = cl_DP.sort_values()
cl_DP
```
### DP with Genre
```
genre_counts = pd.pivot_table(
funct_sample2,
index='s_function',
columns=['genre'],
aggfunc='size',
fill_value=0,
)
genre_counts
genre_exp_prop = genre_counts.sum(0) / genre_counts.sum(0).sum()
genre_obv_prop = genre_counts.div(genre_counts.sum(1), 0)
genre_dev_prop = genre_obv_prop.sub(genre_exp_prop, 1)
genre_DP = abs(genre_dev_prop).sum(1) / 2
genre_DP = genre_DP.sort_values()
genre_DP
genre_dev_prop.loc['Time']
fig, axs = plt.subplots(2, 3, figsize=(15, 8))
axs = axs.ravel()
plt.subplots_adjust(wspace=0.3, hspace=0.2)
dev_max = genre_dev_prop.max().max() + 0.02
dev_min = genre_dev_prop.min().min() - 0.02
funct_order = genre_dev_prop.abs().sum(1).sort_values()
genre_order = genre_counts.sum().sort_values(ascending=False).index
for i, funct in enumerate(funct_order.index):
ax = axs[i]
genre_dev_prop.loc[funct][genre_order].plot(ax=ax, kind='bar', color='white', edgecolor='black')
ax.set_ylim(dev_min, dev_max)
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_title(funct)
ax.set_xlabel('')
ax.set_xticklabels(ax.get_xticklabels(), size=8, rotation=0)
ax.set_ylabel('deviation of %', size=9)
ax.axhline(0, color='black', linewidth=0.8)
#ax.ax_hline(0)
savefig('devProp_genre')
plt.show()
genre_dev_prop.loc['Time']
fig, ax = plt.subplots(figsize=(7, 5))
genre_dev_prop.loc['Time'].sort_values().plot(kind='bar', ax=ax, color='white', edgecolor='black', linewidth=1.1)
ax.set_ylim(-0.12, 0.15)
ax.axhline(0, color='black', linewidth=0.8)
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_ylabel('deviation of observed % from expected %', size=10)
ax.set_title('Time')
savefig('devProp_Time_genre')
plt.show()
```
### subplots of book, chapter, and cluster DP
```
fig, axs = plt.subplots(1, 4, figsize=(18,4))
axs = axs.ravel()
titles = ['DP by genre', 'DP by book', 'DP by chapter', 'DP by clause clusters of 10', ]
plt.subplots_adjust(wspace=0.3)
all_dps = [genre_DP, book_DP, ch_DP, cl_DP]
for i, dp in enumerate(all_dps):
ax=axs[i]
dp.plot(kind='bar', ax=ax, color='lightgrey', edgecolor='black')
ax.set_xlabel('function')
ax.set_ylabel('DP score')
ax.grid(axis='y')
ax.set_axisbelow(True)
ax.set_title(titles[i])
savefig('DP_functions')
DP_book_ch_cl = pd.concat(all_dps, 1, sort=True)
DP_book_ch_cl.columns = ['genre_DP', 'book DP', 'chapter DP', 'clause cl. DP']
DP_book_ch_cl = DP_book_ch_cl.loc[DP_book_ch_cl.sum(1).sort_values().index]
DP_book_ch_cl.round(2).to_clipboard()
DP_book_ch_cl
```
| github_jupyter |
<h1> Preprocessing using Dataflow </h1>
This notebook illustrates:
<ol>
<li> Creating datasets for Machine Learning using Dataflow
</ol>
<p>
While Pandas is fine for experimenting, for operationalization of your workflow, it is better to do preprocessing in Apache Beam. This will also help if you need to preprocess data in flight, since Apache Beam also allows for streaming.
```
!pip install apache-beam[gcp]
```
After doing a pip install, click **"Reset Session"** on the notebook so that the Python environment picks up the new packages.
```
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
%%bash
if ! gsutil ls | grep -q gs://${BUCKET}/; then
gsutil mb -l ${REGION} gs://${BUCKET}
fi
```
<h2> Create ML dataset using Dataflow </h2>
Let's use Cloud Dataflow to read in the BigQuery data, do some preprocessing, and write it out as CSV files.
In this case, I want to do some preprocessing, modifying data so that we can simulate what is known if no ultrasound has been performed.
Note that after you launch this, the actual processing is happening on the cloud. Go to the GCP webconsole to the Dataflow section and monitor the running job. It took about 20 minutes for me.
<p>
If you wish to continue without doing this step, you can copy my preprocessed output:
<pre>
gsutil -m cp -r gs://cloud-training-demos/babyweight/preproc gs://your-bucket/
</pre>
But if you do this, you also have to use my TensorFlow model since yours might expect the fields in a different order
```
import apache_beam as beam
import datetime, os
def to_csv(rowdict):
import hashlib
import copy
# TODO #1:
# Pull columns from BQ and create line(s) of CSV input
CSV_COLUMNS = None
# Create synthetic data where we assume that no ultrasound has been performed
# and so we don't know sex of the baby. Let's assume that we can tell the difference
# between single and multiple, but that the errors rates in determining exact number
# is difficult in the absence of an ultrasound.
no_ultrasound = copy.deepcopy(rowdict)
w_ultrasound = copy.deepcopy(rowdict)
no_ultrasound['is_male'] = 'Unknown'
if rowdict['plurality'] > 1:
no_ultrasound['plurality'] = 'Multiple(2+)'
else:
no_ultrasound['plurality'] = 'Single(1)'
# Change the plurality column to strings
w_ultrasound['plurality'] = ['Single(1)', 'Twins(2)', 'Triplets(3)', 'Quadruplets(4)', 'Quintuplets(5)'][rowdict['plurality'] - 1]
# Write out two rows for each input row, one with ultrasound and one without
for result in [no_ultrasound, w_ultrasound]:
data = ','.join([str(result[k]) if k in result else 'None' for k in CSV_COLUMNS])
key = hashlib.sha224(data).hexdigest() # hash the columns to form a key
yield str('{},{}'.format(data, key))
def preprocess(in_test_mode):
import shutil, os, subprocess
job_name = 'preprocess-babyweight-features' + '-' + datetime.datetime.now().strftime('%y%m%d-%H%M%S')
if in_test_mode:
print 'Launching local job ... hang on'
OUTPUT_DIR = './preproc'
shutil.rmtree(OUTPUT_DIR, ignore_errors=True)
os.makedirs(OUTPUT_DIR)
else:
print 'Launching Dataflow job {} ... hang on'.format(job_name)
OUTPUT_DIR = 'gs://{0}/babyweight/preproc/'.format(BUCKET)
try:
subprocess.check_call('gsutil -m rm -r {}'.format(OUTPUT_DIR).split())
except:
pass
options = {
'staging_location': os.path.join(OUTPUT_DIR, 'tmp', 'staging'),
'temp_location': os.path.join(OUTPUT_DIR, 'tmp'),
'job_name': job_name,
'project': PROJECT,
'teardown_policy': 'TEARDOWN_ALWAYS',
'no_save_main_session': True
}
opts = beam.pipeline.PipelineOptions(flags = [], **options)
if in_test_mode:
RUNNER = 'DirectRunner'
else:
RUNNER = 'DataflowRunner'
p = beam.Pipeline(RUNNER, options = opts)
query = """
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
ABS(FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING)))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
AND weight_pounds > 0
AND mother_age > 0
AND plurality > 0
AND gestation_weeks > 0
AND month > 0
"""
if in_test_mode:
query = query + ' LIMIT 100'
for step in ['train', 'eval']:
if step == 'train':
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) < 3'.format(query)
else:
selquery = 'SELECT * FROM ({}) WHERE MOD(ABS(hashmonth),4) = 3'.format(query)
(p
## TODO Task #2: Modify the Apache Beam pipeline such that the first part of the pipe reads the data from BigQuery
| '{}_read'.format(step) >> None
| '{}_csv'.format(step) >> beam.FlatMap(to_csv)
| '{}_out'.format(step) >> beam.io.Write(beam.io.WriteToText(os.path.join(OUTPUT_DIR, '{}.csv'.format(step))))
)
job = p.run()
if in_test_mode:
job.wait_until_finish()
print "Done!"
# TODO Task #3: Once you have verified that the files produced locally are correct, change in_test_mode to False
# to execute this in Cloud Dataflow
preprocess(in_test_mode = True)
%bash
gsutil ls gs://${BUCKET}/babyweight/preproc/*-00000*
```
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# Amazon SageMaker Studio Walkthrough
_**Using Gradient Boosted Trees to Predict Mobile Customer Departure**_
---
This notebook walks you through some of the main features of Amazon SageMaker Studio.
* [Amazon SageMaker Experiments](https://docs.aws.amazon.com/sagemaker/latest/dg/experiments.html)
* Manage multiple trials
* Experiment with hyperparameters and charting
* [Amazon SageMaker Debugger](https://docs.aws.amazon.com/sagemaker/latest/dg/train-debugger.html)
* Debug your model
* [Model hosting](https://docs.aws.amazon.com/sagemaker/latest/dg/how-it-works-hosting.html)
* Set up a persistent endpoint to get predictions from your model
* [SageMaker Model Monitor](https://docs.aws.amazon.com/sagemaker/latest/dg/model-monitor.html)
* Monitor the quality of your model
* Set alerts for when model quality deviates
You must run this walkthrough in Amazon SageMaker Studio. For Studio onboarding and setup instructions, see the [README](README.md).
---
## Contents
1. [Background](#Background) - Predict customer churn with XGBoost
1. [Data](#Data) - Prep the dataset and upload it to Amazon S3
1. [Train](#Train) - Train with the Amazon SageMaker XGBoost algorithm
- [Amazon SageMaker Experiments](#Amazon-SageMaker-Experiments)
- [Amazon SageMaker Debugger](#Amazon-SageMaker-Debugger)
1. [Host](#Host)
1. [SageMaker Model Monitor](#SageMaker-Model-Monitor)
---
## Background
_This notebook has been adapted from an [AWS blog post](https://aws.amazon.com/blogs/ai/predicting-customer-churn-with-amazon-machine-learning/).
Losing customers is costly for any business. Identifying unhappy customers early on gives you a chance to offer them incentives to stay. This notebook describes using machine learning (ML) for automated identification of unhappy customers, also known as customer churn prediction. It uses Amazon SageMaker features for managing experiments, training and debugging the model, and monitoring it after it has been deployed.
Import the Python libraries we'll need for this walkthrough.
```
import sys
!{sys.executable} -m pip install -qU awscli boto3 "sagemaker>=1.71.0,<2.0.0"
!{sys.executable} -m pip install sagemaker-experiments
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import os
import sys
import time
import json
from IPython.display import display
from time import strftime, gmtime
import boto3
import re
import sagemaker
from sagemaker import get_execution_role
from sagemaker.predictor import csv_serializer
from sagemaker.debugger import rule_configs, Rule, DebuggerHookConfig
from sagemaker.model_monitor import DataCaptureConfig, DatasetFormat, DefaultModelMonitor
from sagemaker.s3 import S3Uploader, S3Downloader
from smexperiments.experiment import Experiment
from smexperiments.trial import Trial
from smexperiments.trial_component import TrialComponent
from smexperiments.tracker import Tracker
sess = boto3.Session()
sm = sess.client('sagemaker')
role = sagemaker.get_execution_role()
```
---
## Data
Mobile operators have historical records that tell them which customers ended up churning and which continued using the service. We can use this historical information to train an ML model that can predict customer churn. After training the model, we can pass the profile information of an arbitrary customer (the same profile information that we used to train the model) to the model to have the model predict whether this customer will churn.
The dataset we use is publicly available and was mentioned in [Discovering Knowledge in Data](https://www.amazon.com/dp/0470908742/) by Daniel T. Larose. It is attributed by the author to the University of California Irvine Repository of Machine Learning Datasets. The `data` folder that came with this notebook contains the dataset, which we've already preprocessed for this walkthrough. The dataset has been split into training and validation sets. To see how the dataset was preprocessed, see this notebook: [XGBoost customer churn notebook that starts with the original dataset](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_applying_machine_learning/xgboost_customer_churn/xgboost_customer_churn.ipynb).
We'll train on a .csv file without the header. But for now, the following cell uses `pandas` to load some of the data from a version of the training data that has a header.
Explore the data to see the dataset's features and the data that will be used to train the model.
```
# Set the path we can find the data files that go with this notebook
%cd /root/amazon-sagemaker-examples/aws_sagemaker_studio/getting_started
local_data_path = './data/training-dataset-with-header.csv'
data = pd.read_csv(local_data_path)
pd.set_option('display.max_columns', 500) # Make sure we can see all of the columns
pd.set_option('display.max_rows', 10) # Keep the output on one page
data
```
Now we'll upload the files to S3 for training but first we will create an S3 bucket for the data if one does not already exist.
```
account_id = sess.client('sts', region_name=sess.region_name).get_caller_identity()["Account"]
bucket = 'sagemaker-studio-{}-{}'.format(sess.region_name, account_id)
prefix = 'xgboost-churn'
try:
if sess.region_name == "us-east-1":
sess.client('s3').create_bucket(Bucket=bucket)
else:
sess.client('s3').create_bucket(Bucket=bucket,
CreateBucketConfiguration={'LocationConstraint': sess.region_name})
except Exception as e:
print("Looks like you already have a bucket of this name. That's good. Uploading the data files...")
# Return the URLs of the uploaded file, so they can be reviewed or used elsewhere
s3url = S3Uploader.upload('data/train.csv', 's3://{}/{}/{}'.format(bucket, prefix,'train'))
print(s3url)
s3url = S3Uploader.upload('data/validation.csv', 's3://{}/{}/{}'.format(bucket, prefix,'validation'))
print(s3url)
```
---
## Train
We'll use the XGBoost library to train a class of models known as gradient boosted decision trees on the data that we just uploaded.
Because we're using XGBoost, we first need to specify the locations of the XGBoost algorithm containers.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
docker_image_name = get_image_uri(boto3.Session().region_name, 'xgboost', repo_version='0.90-2')
```
Then, because we're training with the CSV file format, we'll create `s3_input`s that our training function can use as a pointer to the files in S3.
```
s3_input_train = sagemaker.s3_input(s3_data='s3://{}/{}/train'.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data='s3://{}/{}/validation/'.format(bucket, prefix), content_type='csv')
```
### Amazon SageMaker Experiments
Amazon SageMaker Experiments allows us to keep track of model training; organize related models together; and log model configuration, parameters, and metrics to reproduce and iterate on previous models and compare models. We'll create a single experiment to keep track of the different approaches we'll try to train the model.
Each approach or block of training code that we run will be an experiment trial. Later, we'll be able to compare different trials in Amazon SageMaker Studio.
Let's create the experiment.
```
sess = sagemaker.session.Session()
create_date = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
customer_churn_experiment = Experiment.create(experiment_name="customer-churn-prediction-xgboost-{}".format(create_date),
description="Using xgboost to predict customer churn",
sagemaker_boto_client=boto3.client('sagemaker'))
```
#### Hyperparameters
Now we can specify our XGBoost hyperparameters. Among them are these key hyperparameters:
- `max_depth` Controls how deep each tree within the algorithm can be built. Deeper trees can lead to better fit, but are more computationally expensive and can lead to overfitting. Typically, you need to explore some trade-offs in model performance between a large number of shallow trees and a smaller number of deeper trees.
- `subsample` Controls sampling of the training data. This hyperparameter can help reduce overfitting, but setting it too low can also starve the model of data.
- `num_round` Controls the number of boosting rounds. This value specifies the models that are subsequently trained using the residuals of previous iterations. Again, more rounds should produce a better fit on the training data, but can be computationally expensive or lead to overfitting.
- `eta` Controls how aggressive each round of boosting is. Larger values lead to more conservative boosting.
- `gamma` Controls how aggressively trees are grown. Larger values lead to more conservative models.
- `min_child_weight` Also controls how aggresively trees are grown. Large values lead to a more conservative model.
For more information about these hyperparameters, see [XGBoost's hyperparameters GitHub page](https://github.com/dmlc/xgboost/blob/master/doc/parameter.rst).
```
hyperparams = {"max_depth":5,
"subsample":0.8,
"num_round":600,
"eta":0.2,
"gamma":4,
"min_child_weight":6,
"silent":0,
"objective":'binary:logistic'}
```
#### Trial 1 - XGBoost in algorithm mode
For our first trial, we'll use the built-in XGBoost algorithm to train a model without supplying any additional code. This way, we can use XGBoost to train and deploy a model as we would with other Amazon SageMaker built-in algorithms.
We'll create a new `Trial` object and associate the trial with the experiment that we created earlier. To train the model, we'll create an estimator and specify a few parameters, like the type of training instances we'd like to use and how many, and where the artifacts of the trained model should be stored.
We'll also associate the training job with the experiment trial that we just created when we call the `fit` method of the `estimator`.
```
trial = Trial.create(trial_name="algorithm-mode-trial-{}".format(strftime("%Y-%m-%d-%H-%M-%S", gmtime())),
experiment_name=customer_churn_experiment.experiment_name,
sagemaker_boto_client=boto3.client('sagemaker'))
xgb = sagemaker.estimator.Estimator(image_name=docker_image_name,
role=role,
hyperparameters=hyperparams,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
base_job_name="demo-xgboost-customer-churn",
sagemaker_session=sess)
xgb.fit({'train': s3_input_train,
'validation': s3_input_validation},
experiment_config={
"ExperimentName": customer_churn_experiment.experiment_name,
"TrialName": trial.trial_name,
"TrialComponentDisplayName": "Training",
}
)
```
#### Review the results
After the training job completes successfully, you can view metrics, logs, and graphs related to the trial on the **Experiments** tab in Amazon SageMaker Studio.
To see the experiments, choose the **Experiments** button.

To see an experiment's components, in the **Experiments** list, double-click it. To see the componenets for multiple experiments, hold down the Crtl key and choose the experiments, then open the context menu (by right-clicking on an experiment). To see the compenents together, choose "Open in trial component list". This enables charting across experiments.

The components are sorted so that the best model is at the top.

#### Download the model
You can find and download the model in Amazon SageMaker Studio. To find the model, choose the **Experiments** button in the left tray, and keep drilling down through the experiment, the most recent trial listed, and its most recent component until you see the **Describe Trial Components** page. Choose the **Artifacts** tab. It contains links to the training and validation data in the **Input Artifacts** section, and the generated model artifact in the **Output Artifacts** section.

#### Trying other hyperparameter values
To improve a model, you typically try other hyperparameter values to see if they affect the final validation error. Let's vary the `min_child_weight` parameter and start other training jobs with those different values to see how they affect the validation error. For each value, we'll create a separate trial so that we can compare the results in Amazon SageMaker Studio later.
```
min_child_weights = [1, 2, 4, 8, 10]
for weight in min_child_weights:
hyperparams["min_child_weight"] = weight
trial = Trial.create(trial_name="algorithm-mode-trial-{}-weight-{}".format(strftime("%Y-%m-%d-%H-%M-%S", gmtime()), weight),
experiment_name=customer_churn_experiment.experiment_name,
sagemaker_boto_client=boto3.client('sagemaker'))
t_xgb = sagemaker.estimator.Estimator(image_name=docker_image_name,
role=role,
hyperparameters=hyperparams,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
base_job_name="demo-xgboost-customer-churn",
sagemaker_session=sess)
t_xgb.fit({'train': s3_input_train,
'validation': s3_input_validation},
wait=False,
experiment_config={
"ExperimentName": customer_churn_experiment.experiment_name,
"TrialName": trial.trial_name,
"TrialComponentDisplayName": "Training",
}
)
```
#### Create charts
To create a chart, multi-select the components. Because this is a sample training run and the data is sparse, there's not much to chart in a time series. However, we can create a scatter plot for the parameter sweep. The following image is an example.

##### How to create a scatter plot
Multi-select components, then choose **Add chart**. In the **Chart Properties** panel, choose **Summary Statistics** as the **Data type**. For **Chart type**, choose **Scatter plot**. Choose the hyperparameter `min_child_weight` as the X-axis (because this is the hyperparameter that you are iterating on in this notebook). For Y-axis metrics, choose either `validation:error_last` or `validation:error_avg`. Color them by choosing `trialComponentName`.

You can also adjust the chart at any time by changing the components that are selected. And you can zoom in and out. Each item on the graph displays contextual information.

### Amazon SageMaker Debugger
Amazon SageMaker Debugger lets you debug a model during training. As you train, Debugger periodicially saves tensors, which fully specify the state of the model at that point in time. Debugger saves the tensors to an Amazon S3 bucket. You can then use Amazon SageMaker Studio for analysis and visualization to diagnose training issues.
#### Specify SageMaker Debugger Rules
To enable automated detection of common issues during training, Amazon SageMaker Debugger also allows you to attach a list of rules to evaluate the training job against.
Some rules that apply to XGBoost include `AllZero`, `ClassImbalance`, `Confusion`, `LossNotDecreasing`, `Overfit`, `Overtraining`, `SimilarAcrossRuns`, `TensorVariance`, `UnchangedTensor`, and `TreeDepth`.
We'll use the `LossNotDecreasing` rule--which is triggered if the loss doesn't decrease monotonically at any point during training--the `Overtraining` rule, and the `Overfit` rule. Let's create the rules.
```
debug_rules = [Rule.sagemaker(rule_configs.loss_not_decreasing()),
Rule.sagemaker(rule_configs.overtraining()),
Rule.sagemaker(rule_configs.overfit())
]
```
#### Trial 2 - XGBoost in framework mode
For the next trial, we'll train a similar model, but use XGBoost in framework mode. If you've worked with the open source XGBoost, using XGBoost this way will be familiar to you. Using XGBoost as a framework provides more flexibility than using it as a built-in algorithm because it enables more advanced scenarios that allow incorporating preprocessing and post-processing scripts into your training script. Specifically, we'll be able to specify a list of rules that we want Amazon SageMaker Debugger to evaluate our training against.
#### Fit estimator
To use XGBoost as a framework, you need to specify an entry-point script that can incorporate additional processing into your training jobs.
We've made a couple of simple changes to enable the Amazon SageMaker Debugger `smdebug` library. We created a `SessionHook`, which we pass as a callback function when creating a `Booster`. We passed a `SaveConfig` object that tells the hook to save the evaluation metrics, feature importances, and SHAP values at regular intervals. (Debugger is highly configurable. You can choose exactly what to save.) We describe the changes in more detail after we train this example. For even more detail, see the [Developer Guide for XGBoost](https://github.com/awslabs/sagemaker-debugger/tree/master/docs/xgboost).
```
!pygmentize xgboost_customer_churn.py
```
Let's create our framwork estimator and call `fit` to start the training job. As before, we'll create a separate trial for this run so that we can use Amazon SageMaker Studio to compare it with other trials later. Because we are running in framework mode, we also need to pass additional parameters, like the entry point script and the framework version, to the estimator.
As training progresses, you'll be able to see Amazon SageMaker Debugger logs that evaluate the rule against the training job.
```
entry_point_script = "xgboost_customer_churn.py"
trial = Trial.create(trial_name="framework-mode-trial-{}".format(strftime("%Y-%m-%d-%H-%M-%S", gmtime())),
experiment_name=customer_churn_experiment.experiment_name,
sagemaker_boto_client=boto3.client('sagemaker'))
framework_xgb = sagemaker.xgboost.XGBoost(image_name=docker_image_name,
entry_point=entry_point_script,
role=role,
framework_version="0.90-2",
py_version="py3",
hyperparameters=hyperparams,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
base_job_name="demo-xgboost-customer-churn",
sagemaker_session=sess,
rules=debug_rules
)
framework_xgb.fit({'train': s3_input_train,
'validation': s3_input_validation},
experiment_config={
"ExperimentName": customer_churn_experiment.experiment_name,
"TrialName": trial.trial_name,
"TrialComponentDisplayName": "Training",
})
```
After the training has been running for a while you can view debug info in the Debugger panel. To get to this panel you must click through the experiment, trial, and then component.

---
## Host the model
Now that we've trained the model, let's deploy it to a hosted endpoint. To monitor the model after it's hosted and serving requests, we'll also add configurations to capture data that is being sent to the endpoint.
```
data_capture_prefix = '{}/datacapture'.format(prefix)
endpoint_name = "demo-xgboost-customer-churn-" + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print("EndpointName = {}".format(endpoint_name))
xgb_predictor = xgb.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge',
endpoint_name=endpoint_name,
data_capture_config=DataCaptureConfig(enable_capture=True,
sampling_percentage=100,
destination_s3_uri='s3://{}/{}'.format(bucket, data_capture_prefix)
)
)
```
### Invoke the deployed model
Now that we have a hosted endpoint running, we can make real-time predictions from our model by making an http POST request. But first, we need to set up serializers and deserializers for passing our `test_data` NumPy arrays to the model behind the endpoint.
```
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
xgb_predictor.deserializer = None
```
Now, we'll loop over our test dataset and collect predictions by invoking the XGBoost endpoint:
```
print("Sending test traffic to the endpoint {}. \nPlease wait for a minute...".format(endpoint_name))
with open('data/test_sample.csv', 'r') as f:
for row in f:
payload = row.rstrip('\n')
response = xgb_predictor.predict(data=payload)
time.sleep(0.5)
```
### Verify that data is captured in Amazon S3
When we made some real-time predictions by sending data to our endpoint, we should have also captured that data for monitoring purposes.
Let's list the data capture files stored in Amazon S3. Expect to see different files from different time periods organized based on the hour in which the invocation occurred. The format of the Amazon S3 path is:
`s3://{destination-bucket-prefix}/{endpoint-name}/{variant-name}/yyyy/mm/dd/hh/filename.jsonl`
```
from time import sleep
current_endpoint_capture_prefix = '{}/{}'.format(data_capture_prefix, endpoint_name)
for _ in range(12): # wait up to a minute to see captures in S3
capture_files = S3Downloader.list("s3://{}/{}".format(bucket, current_endpoint_capture_prefix))
if capture_files:
break
sleep(5)
print("Found Data Capture Files:")
print(capture_files)
```
All the data captured is stored in a SageMaker specific json-line formatted file. Next, Let's take a quick peek at the contents of a single line in a pretty formatted json so that we can observe the format a little better.
```
capture_file = S3Downloader.read_file(capture_files[-1])
print("=====Single Data Capture====")
print(json.dumps(json.loads(capture_file.split('\n')[0]), indent=2)[:2000])
```
As you can see, each inference request is captured in one line in the jsonl file. The line contains both the input and output merged together. In our example, we provided the ContentType as `text/csv` which is reflected in the `observedContentType` value. Also, we expose the enconding that we used to encode the input and output payloads in the capture format with the `encoding` value.
To recap, we have observed how you can enable capturing the input and/or output payloads to an Endpoint with a new parameter. We have also observed how the captured format looks like in S3. Let's continue to explore how SageMaker helps with monitoring the data collected in S3.
---
## Amazon SageMaker Model Monitor
Amazon SageMaker Model Monitor lets you monitor and evaluate the data observed by endpoints. It works like this:
1. We need to create a baseline that we can use to compare real-time traffic against.
1. When a baseline is ready, we can set up a schedule to continously evaluate and compare against the baseline.
1. We can send synthetic traffic to trigger alarms.
**Important**: It takes an hour or more to complete this section because the shortest monitoring polling time is one hour. The following graphic shows how the monitoring results look after running for a few hours and some of the errors triggered by synthetic traffic.

### Baselining and continous monitoring
#### 1. Constraint suggestion with the baseline (training) dataset
The training dataset that you use to train a model is usually a good baseline dataset. Note that the training dataset data schema and the inference dataset schema must match exactly (for example, they should have the same number and type of features).
Using our training dataset, let's ask Amazon SageMaker Model Monitor to suggest a set of baseline `constraints` and generate descriptive `statistics` so we can explore the data. For this example, let's upload the training dataset, which we used to train model. We'll use the dataset file with column headers so we have descriptive feature names.
```
baseline_prefix = prefix + '/baselining'
baseline_data_prefix = baseline_prefix + '/data'
baseline_results_prefix = baseline_prefix + '/results'
baseline_data_uri = 's3://{}/{}'.format(bucket,baseline_data_prefix)
baseline_results_uri = 's3://{}/{}'.format(bucket, baseline_results_prefix)
print('Baseline data uri: {}'.format(baseline_data_uri))
print('Baseline results uri: {}'.format(baseline_results_uri))
baseline_data_path = S3Uploader.upload("data/training-dataset-with-header.csv", baseline_data_uri)
```
##### Create a baselining job with the training dataset
Now that we have the training data ready in S3, let's start a job to `suggest` constraints. To generate the constraints, the convenient helper starts a `ProcessingJob` using a ProcessingJob container provided by Amazon SageMaker.
```
my_default_monitor = DefaultModelMonitor(role=role,
instance_count=1,
instance_type='ml.m5.xlarge',
volume_size_in_gb=20,
max_runtime_in_seconds=3600,
)
baseline_job = my_default_monitor.suggest_baseline(baseline_dataset=baseline_data_path,
dataset_format=DatasetFormat.csv(header=True),
output_s3_uri=baseline_results_uri,
wait=True
)
```
Once the job succeeds, we can explore the `baseline_results_uri` location in s3 to see what files where stored there.
```
print("Found Files:")
S3Downloader.list("s3://{}/{}".format(bucket, baseline_results_prefix))
```
We have a`constraints.json` file that has information about suggested constraints. We also have a `statistics.json` which contains statistical information about the data in the baseline.
```
baseline_job = my_default_monitor.latest_baselining_job
schema_df = pd.io.json.json_normalize(baseline_job.baseline_statistics().body_dict["features"])
schema_df.head(10)
constraints_df = pd.io.json.json_normalize(baseline_job.suggested_constraints().body_dict["features"])
constraints_df.head(10)
```
#### 2. Analyzing subsequent captures for data quality issues
Now that we've generated a baseline dataset and processed it to get baseline statistics and constraints, let's monitor and analyze the data being sent to the endpoint with monitoring schedules.
##### Create a schedule
First, let's create a monitoring schedule for the endpoint. The schedule specifies the cadence at which we want to run a new processing job so that we can compare recent data captures to the baseline.
```
# First, copy over some test scripts to the S3 bucket so that they can be used for pre and post processing
code_prefix = '{}/code'.format(prefix)
pre_processor_script = S3Uploader.upload('preprocessor.py', 's3://{}/{}'.format(bucket,code_prefix))
s3_code_postprocessor_uri = S3Uploader.upload('postprocessor.py', 's3://{}/{}'.format(bucket,code_prefix))
```
We are ready to create a model monitoring schedule for the Endpoint created before and also the baseline resources (constraints and statistics) which were generated above.
```
from sagemaker.model_monitor import CronExpressionGenerator
from time import gmtime, strftime
reports_prefix = '{}/reports'.format(prefix)
s3_report_path = 's3://{}/{}'.format(bucket,reports_prefix)
mon_schedule_name = 'demo-xgboost-customer-churn-model-schedule-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
my_default_monitor.create_monitoring_schedule(monitor_schedule_name=mon_schedule_name,
endpoint_input=xgb_predictor.endpoint,
#record_preprocessor_script=pre_processor_script,
post_analytics_processor_script=s3_code_postprocessor_uri,
output_s3_uri=s3_report_path,
statistics=my_default_monitor.baseline_statistics(),
constraints=my_default_monitor.suggested_constraints(),
schedule_cron_expression=CronExpressionGenerator.hourly(),
enable_cloudwatch_metrics=True,
)
```
#### 3. Start generating some artificial traffic
The following block starts a thread to send some traffic to the endpoint. This allows us to continue to send traffic to the endpoint so that we'll have data continually captured for analysis. If there is no traffic, the monitoring jobs will start to fail later.
To terminate this thread, you need to stop the kernel.
```
from threading import Thread
runtime_client = boto3.client('runtime.sagemaker')
# (just repeating code from above for convenience/ able to run this section independently)
def invoke_endpoint(ep_name, file_name, runtime_client):
with open(file_name, 'r') as f:
for row in f:
payload = row.rstrip('\n')
response = runtime_client.invoke_endpoint(EndpointName=ep_name,
ContentType='text/csv',
Body=payload)
response['Body'].read()
sleep(1)
def invoke_endpoint_forever():
while True:
invoke_endpoint(endpoint_name, 'data/test-dataset-input-cols.csv', runtime_client)
thread = Thread(target = invoke_endpoint_forever)
thread.start()
# Note that you need to stop the kernel to stop the invocations
```
##### List executions
Once the schedule is set up, jobs start at the specified intervals. The following code lists the last five executions. If you run this code soon after creating the hourly schedule, you might not see any executions listed. To see executions, you might have to wait until you cross the hour boundary (in UTC). The code includes the logic for waiting.
```
mon_executions = my_default_monitor.list_executions()
if len(mon_executions) == 0:
print("We created a hourly schedule above and it will kick off executions ON the hour.\nWe will have to wait till we hit the hour...")
while len(mon_executions) == 0:
print("Waiting for the 1st execution to happen...")
time.sleep(60)
mon_executions = my_default_monitor.list_executions()
```
##### Evaluate the latest execution and list the generated reports
```
latest_execution = mon_executions[-1]
latest_execution.wait()
print("Latest execution result: {}".format(latest_execution.describe()['ExitMessage']))
report_uri = latest_execution.output.destination
print("Found Report Files:")
S3Downloader.list(report_uri)
```
##### List violations
If there are any violations compared to the baseline, they will be generated here. Let's list the violations.
```
violations = my_default_monitor.latest_monitoring_constraint_violations()
pd.set_option('display.max_colwidth', -1)
constraints_df = pd.io.json.json_normalize(violations.body_dict["violations"])
constraints_df.head(10)
```
You can plug in the processing job arn for a single execution of the monitoring into [this notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/sagemaker_model_monitor/visualization/SageMaker-Model-Monitor-Visualize.ipynb) to see more detailed visualizations of the violations and distribution statistics of the data captue that was processed in that execution
```
latest_execution.describe()['ProcessingJobArn']
```
## Clean up
If you no longer need this notebook, clean up your environment by running the following cell. It removes the hosted endpoint that you created for this walkthrough and prevents you from incurring charges for running an instance that you no longer need. It also cleans up all artifacts related to the experiments.
You might also want to delete artifacts stored in the S3 bucket used in this notebook. To do so, open the Amazon S3 console, find the `sagemaker-studio-<region-name>-<account-name>` bucket, and delete the files associated with this notebook.
```
try:
sess.delete_monitoring_schedule(mon_schedule_name)
except:
pass
while True:
try:
print("Waiting for schedule to be deleted")
sess.describe_monitoring_schedule(mon_schedule_name)
sleep(15)
except:
print("Schedule deleted")
break
sess.delete_endpoint(xgb_predictor.endpoint)
def cleanup(experiment):
'''Clean up everything in the given experiment object'''
for trial_summary in experiment.list_trials():
trial = Trial.load(trial_name=trial_summary.trial_name)
for trial_comp_summary in trial.list_trial_components():
trial_step=TrialComponent.load(trial_component_name=trial_comp_summary.trial_component_name)
print('Starting to delete TrialComponent..' + trial_step.trial_component_name)
sm.disassociate_trial_component(TrialComponentName=trial_step.trial_component_name, TrialName=trial.trial_name)
trial_step.delete()
time.sleep(1)
trial.delete()
experiment.delete()
cleanup(customer_churn_experiment)
```
| github_jupyter |
TSG045 - The maximum number of data disks allowed to be attached to a VM of this size (AKS)
===========================================================================================
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
first_run = True
rules = None
debug_logging = False
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# apply expert rules (to run follow-on notebooks), based on output
#
if rules is not None:
apply_expert_rules(line_decoded)
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
"""Load a json file from disk and return the contents"""
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
"""Load any 'expert rules' from the metadata of this notebook (.ipynb) that should be applied to the stderr of the running executable"""
try:
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
j = load_json("tsg045-max-number-data-disks-allowed.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
"""Determine if the stderr line passed in, matches the regular expressions for any of the 'expert rules', if so
inject a 'HINT' to the follow-on SOP/TSG to run"""
global rules
for rule in rules:
# rules that have 9 elements are the injected (output) rules (the ones we want). Rules
# with only 8 elements are the source (input) rules, which are not expanded (i.e. TSG029,
# not ../repair/tsg029-nb-name.ipynb)
if len(rule) == 9:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
if debug_logging:
print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
if debug_logging:
print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['no such host', 'TSG011 - Restart sparkhistory server', '../repair/tsg011-restart-sparkhistory-server.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb']}
```
### Instantiate Kubernetes client
```
# Instantiate the Python Kubernetes client into 'api' variable
import os
try:
from kubernetes import client, config
from kubernetes.stream import stream
if "KUBERNETES_SERVICE_PORT" in os.environ and "KUBERNETES_SERVICE_HOST" in os.environ:
config.load_incluster_config()
else:
try:
config.load_kube_config()
except:
display(Markdown(f'HINT: Use [TSG112 - App-Deploy Proxy Nginx Logs](../log-analyzers/tsg112-get-approxy-nginx-logs.ipynb) to resolve this issue.'))
raise
api = client.CoreV1Api()
print('Kubernetes client instantiated')
except ImportError:
from IPython.display import Markdown
display(Markdown(f'HINT: Use [SOP059 - Install Kubernetes Python module](../install/sop059-install-kubernetes-module.ipynb) to resolve this issue.'))
raise
```
### Get the namespace for the big data cluster
Get the namespace of the Big Data Cluster from the Kuberenetes API.
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = api.list_namespace(label_selector='MSSQL_CLUSTER').items[0].metadata.name
except IndexError:
from IPython.display import Markdown
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print('The kubernetes namespace for your big data cluster is: ' + namespace)
```
### Set the text to look for in pod events
```
name="master-0"
kind="Pod"
precondition_text="The maximum number of data disks allowed to be attached to a VM of this size"
```
### Get events for a kubernetes resources
Get the events for a kubernetes named space resource:
```
V1EventList=api.list_namespaced_event(namespace)
for event in V1EventList.items:
if (event.involved_object.kind==kind and event.involved_object.name==name):
print(event.message)
```
### PRECONDITION CHECK
```
precondition=False
for event in V1EventList.items:
if (event.involved_object.kind==kind and event.involved_object.name==name):
if event.message.find(precondition_text) != -1:
precondition=True
if not precondition:
raise Exception("PRECONDITION NON-MATCH: 'tsg045-max-number-data-disks-allowed' is not a match for an active problem")
print("PRECONDITION MATCH: 'tsg045-max-number-data-disks-allowed' is a match for an active problem in this cluster")
```
Resolution
----------
Kubernetes may have got stuck trying to get this pod to start on a node
which doesn’t allow enough Persistent Volumes to be attached to it.
You can list the Persistent volumes:
```
run('kubectl get pv')
```
To workaround this issue delete the master-0 pod, and the Kubernetes
statefulset will automatically create a new pod, hopefully on another
node!
```
run(f'kubectl delete pod/master-0 -n {namespace}')
```
Now watch the events from the new master-0 pod, to verify the issue is
resolved
### Get events for a kubernetes resources
Get the events for a kubernetes named space resource:
```
V1EventList=api.list_namespaced_event(namespace)
for event in V1EventList.items:
if (event.involved_object.kind==kind and event.involved_object.name==name):
print(event.message)
```
### View the pods
Verify the master-0 pod is now running
```
run(f'kubectl get pods -n {namespace}')
```
### Describe the master-0
Verify the master pod looks healthy
```
run('kubectl describe pod/master-0 -n {namespace}')
print('Notebook execution complete.')
```
| github_jupyter |
# Demonstration on a toy problem
```
import numpy as np
import hypothesis
import torch
import os
import matplotlib.pyplot as plt
import warnings
from hypothesis.visualization.util import make_square
from toy import allocate_prior
from toy import load_estimator
from toy import Simulator
from toy import RatioEstimator
from toy import JointTrain
from toy import JointTest
```
## Setting and setup
```
prior = allocate_prior()
simulator = Simulator() # The likelihood model is normal
# Draw a sample from the joint.
truth = prior.sample()
x_o = simulator(truth) # Draw x_o ~ N(truth, 1)
# Cleanup the old data files if they are present
!rm -r estimators
```
## Training
We train the ratio estimator using the binary provided by the [`hypothesis`](https://github.com/montefiore-ai/hypothesis) package.
```
!python -u -m hypothesis.bin.ratio_estimation.train -h
```
### Hyperparameters
```
batch_size = 4096
epochs = 10
learning_rate = 0.0001
weight_decay = 0.0
num_estimators = 3
```
### Training procedure
```console
python -u -m hypothesis.bin.ratio_estimation.train
--data-paralle # Automatically train the model on multiple GPU's when the resources are available
--out $path # Store the trained models and other metadata in the specified folder
--show # Show the progress of the training procedure in `stdout`
--denominator "inputs|outputs" # Denominator of the training criterion '|' denotes an independence relation.
# -> p(inputs)p(outputs) -> so the ratio it will learn is p(inputs,outputs)/(p(inputs)p(outputs))
# -> pointwise mutual information
--batch-size $batch_size # Batch size
--conservativeness 0.0 # Conservativeness criterion, how much weight should be given to the prior.
# -> Convenient way to impose a conservative estimation.
# -> Setting this value to 1 will result in a posterior that is indistinghuisable from the prior.
--epochs $epochs # Number of epochs
--lr $learning_rate # Learning rate
--lrsched # Enables learning rate scheduling
--lrsched-every 5 # Schedule every 5 epochs
--lrsched-gamma 0.5 # Decay learning rate every 5 epochs by 0.5
--weight-decay $weight_decay # Weight decay, automatically enables the AdamW optimizer.
--workers 8 # Number of concurrent data loaders
--data-train "toy.JointTrain" # Class of the training dataset
--data-test "toy.JointTest" # Class of the test dataset
--estimator "toy.RatioEstimator" # Class of the ratio estimator, should accept the keywords defined in `--denominator`.
# -> forward(inputs=inputs, outputs=outputs).
```
```
!mkdir -p estimators
for index in range(num_estimators):
path = "estimators/" + str(index).zfill(5)
# Check if the model has been trained
if not os.path.exists(path + "/best-model.th"):
print("\nTraining started!")
!python -u -m hypothesis.bin.ratio_estimation.train \
--data-parallel \
--out $path \
--show \
--denominator "inputs|outputs" \
--batch-size $batch_size \
--conservativeness 0.0 \
--epochs $epochs \
--lr $learning_rate \
--lrsched \
--lrsched-every 5 \
--lrsched-gamma 0.5 \
--weight-decay $weight_decay \
--workers 8 \
--data-train "toy.JointTrain" \
--data-test "toy.JointTest" \
--estimator "toy.RatioEstimator"
```
The loss curves:
```
import glob
paths = "estimators/*/losses-test.npy"
losses = np.vstack([np.load(p).reshape(1, -1) for p in glob.glob(paths)])
epochs = np.arange(1, len(losses[0]) + 1)
m = np.mean(losses, axis=0)
s = np.std(losses, axis=0)
plt.plot(epochs, m, color="black", lw=2)
plt.fill_between(epochs, m - s, m + s, color="black", alpha=.1)
plt.ylabel("Test loss")
plt.xlabel("Epochs")
make_square(plt.gca())
plt.show()
```
## Inference
```
# Load the trained ratio estimator, and prepare the inputs.
resolution = 100
r = load_estimator("estimators/*/best-model.th")
inputs = torch.linspace(prior.low, prior.high, resolution).view(-1, 1)
outputs = x_o.repeat(resolution).view(-1, 1)
```
We now have all components to compute
$$
\log\hat{p}(\vartheta\vert x) = \log p(\vartheta) + \log\hat{r}(x\vert\vartheta)
$$
```
# Compute the posterior
log_posterior = prior.log_prob(inputs) + r.log_ratio(inputs=inputs, outputs=outputs)
```
As you can tell, we compute the posterior density function by evaluating the input space for the observable $x_o$.
```
# Plot the posterior
plt.plot(inputs.detach().numpy(), log_posterior.exp().detach().numpy(), lw=2, color="black")
plt.axvline(truth, lw=2, color="C0", label="Truth")
plt.axvline(x_o, lw=2, color="red", label="True MAP")
plt.ylabel("Posterior density")
make_square(plt.gca())
plt.legend()
plt.show()
# Plot the posterior
plt.plot(inputs.detach().numpy(), log_posterior.detach().numpy(), lw=2, color="black")
plt.axvline(truth, lw=2, color="C0", label="Truth")
plt.axvline(x_o, lw=2, color="red", label="True MAP")
plt.ylabel("Posterior density")
make_square(plt.gca())
plt.legend()
plt.show()
```
Due to the amortization, we can simply evaluate the posterior for a different observable without retraining! This enables fast likelihood-free inference when confronted with many observables.
```
# Draw a sample from the joint.
truth = prior.sample()
x_o = simulator(truth)
# Prepare the inputs for the ratio estimator
inputs = torch.linspace(prior.low, prior.high, resolution).view(-1, 1)
outputs = x_o.repeat(resolution).view(-1, 1)
# Compute the posterior
log_posterior = prior.log_prob(inputs) + r.log_ratio(inputs=inputs, outputs=outputs)
# Plot the posterior
plt.plot(inputs.detach().numpy(), log_posterior.exp().detach().numpy(), lw=2, color="black")
plt.axvline(truth, lw=2, color="C0", label="Truth")
plt.axvline(x_o, lw=2, color="red", label="True MAP")
plt.ylabel("Posterior density")
make_square(plt.gca())
plt.legend()
plt.show()
# Plot the log posterior
plt.plot(inputs.detach().numpy(), log_posterior.detach().numpy(), lw=2, color="black")
plt.axvline(truth, lw=2, color="C0", label="Truth")
plt.axvline(x_o, lw=2, color="red", label="True MAP")
plt.ylabel("Posterior density")
make_square(plt.gca())
plt.legend()
plt.show()
```
| github_jupyter |
# Smart signatures
#### 06.1 Writing Smart Contracts
##### Peter Gruber (peter.gruber@usi.ch)
2022-01-12
* Write and deploy smart Signatures
## Setup
See notebook 04.1, the lines below will always automatically load functions in `algo_util.py`, the five accounts and the Purestake credentials
```
# Loading shared code and credentials
import sys, os
codepath = '..'+os.path.sep+'..'+os.path.sep+'sharedCode'
sys.path.append(codepath)
from algo_util import *
cred = load_credentials()
# Shortcuts to directly access the 3 main accounts
MyAlgo = cred['MyAlgo']
Alice = cred['Alice']
Bob = cred['Bob']
Charlie = cred['Charlie']
Dina = cred['Dina']
from algosdk import account, mnemonic
from algosdk.v2client import algod
from algosdk.future import transaction
from algosdk.future.transaction import PaymentTxn
from algosdk.future.transaction import AssetConfigTxn, AssetTransferTxn, AssetFreezeTxn
from algosdk.future.transaction import LogicSig, LogicSigTransaction
import algosdk.error
import json
import base64
import hashlib
from pyteal import *
# Initialize the algod client (Testnet or Mainnet)
algod_client = algod.AlgodClient(algod_token='', algod_address=cred['algod_test'], headers=cred['purestake_token'])
print(Alice['public'])
print(Bob['public'])
print(Charlie['public'])
```
#### Check Purestake API
```
last_block = algod_client.status()["last-round"]
print(f"Last committed block is: {last_block}")
```
## The Donation Escrow
A classical smart contact. **Alice** donates to **Bob** (and only to **Bob**).
**Bob** can decide when to ask for the money.
##### Step 1: The programmer writes down the conditions as a PyTeal program
```
escrow_pyteal = (
Txn.receiver() == Addr(Bob["public"]) # Receipient must be Bob
) # Encode addresses using Addr()
# Security missing ... do not copy-paste
```
##### Step 2: Compile PyTeal -> Teal
```
escrow_teal = compileTeal(escrow_pyteal, Mode.Signature, version=3)
print(escrow_teal)
```
##### Step 3: Compile Teal -> Bytecode for AVM
```
# compile Teal -> Bytecode
Escrow = algod_client.compile(escrow_teal)
Escrow
```
##### Step 4: Alice funds and deploys the smart signature
```
# Step 1: prepare transaction
sp = algod_client.suggested_params()
amt = int(1.5*1e6)
txn = transaction.PaymentTxn(sender=Alice['public'], sp=sp, receiver=Escrow['hash'], amt=amt)
# Step 2+3: sign and sen
stxn = txn.sign(Alice['private'])
txid = algod_client.send_transaction(stxn)
# Step 4: wait for confirmation
txinfo = wait_for_confirmation(algod_client, txid)
# Smart Signature is now funded
print('https://testnet.algoexplorer.io/address/'+ Escrow['hash'])
```
##### Step 5: Alice informs Bob
```
print("Alice communicates to Bob the following")
print("Compiled smart signature:", Escrow['result'])
print("Address of smart signature: ", Escrow['hash'])
```
#### Step 6: Bob proposes a transaction to the smart signature
* Again he proposes a payment from the cashmachine to **himself**
* The payment transaction is signed by the smart signature, **if the conditions are fullfilled** (correct recipient)
```
# Step 1: prepare TX
sp = algod_client.suggested_params()
withdrawal_amt = int(1.298*1e6) - int(0.001*1e6) - int(0.1*1e6)
txn = PaymentTxn(sender=Escrow['hash'], sp=sp,
receiver=Bob['public'], amt=withdrawal_amt)
# Step 2: sign TX <---- This step is different!
encodedProg = Escrow['result'].encode()
program = base64.decodebytes(encodedProg)
lsig = LogicSig(program)
stxn = LogicSigTransaction(txn, lsig)
# Step 3: send
txid = algod_client.send_transaction(stxn)
# Step 4: wait for confirmation
txinfo = wait_for_confirmation(algod_client, txid)
```
### Exercise
* Repeat Step 6. Can Bob withdraw several times?
* Write the transaction(s) for Charlie to try to withdraw from the Escrow
```
# Python code goes here
```
| github_jupyter |
```
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import KFold
from sklearn.model_selection import ShuffleSplit
from sklearn.metrics import accuracy_score
from keras.layers import Dense
from keras.models import Sequential
from keras.optimizers import SGD
from matplotlib import pyplot as plt
import matplotlib as mpl
import seaborn as sns
import numpy as np
import pandas as pd
import category_encoders as ce
import os
import pickle
import gc
from tqdm import tqdm
import pickle
from sklearn.svm import SVR
from sklearn.linear_model import LinearRegression
from sklearn import linear_model
from sklearn.neighbors import KNeighborsRegressor
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn import ensemble
import xgboost as xgb
from numpy import loadtxt
from keras.models import Sequential
from keras.layers import Dense
def encode_text_features(encode_decode, data_frame, encoder_isa=None, encoder_mem_type=None):
# Implement Categorical OneHot encoding for ISA and mem-type
if encode_decode == 'encode':
encoder_isa = ce.one_hot.OneHotEncoder(cols=['isa'])
encoder_mem_type = ce.one_hot.OneHotEncoder(cols=['mem-type'])
encoder_isa.fit(data_frame, verbose=1)
df_new1 = encoder_isa.transform(data_frame)
encoder_mem_type.fit(df_new1, verbose=1)
df_new = encoder_mem_type.transform(df_new1)
encoded_data_frame = df_new
else:
df_new1 = encoder_isa.transform(data_frame)
df_new = encoder_mem_type.transform(df_new1)
encoded_data_frame = df_new
return encoded_data_frame, encoder_isa, encoder_mem_type
def absolute_percentage_error(Y_test, Y_pred):
error = 0
for i in range(len(Y_test)):
if(Y_test[i]!= 0 ):
error = error + (abs(Y_test[i] - Y_pred[i]))/Y_test[i]
error = error/ len(Y_test)
return error
def create_model(name = 'dnn_1'):
input_dim = 22
# define the keras model
# DNN 1
if name == 'dnn_1':
dnn_1 = Sequential()
dnn_1.add(Dense(512, input_dim=input_dim, activation='relu'))
dnn_1.add(Dense(1, activation='linear'))
# print('Model : DNN 1', dnn_1.summary())
# compile the keras model
return dnn_1
# DNN 2
# define the keras model
elif name == 'dnn_2':
dnn_2 = Sequential()
dnn_2.add(Dense(512, input_dim=input_dim, activation='relu'))
dnn_2.add(Dense(512, activation='relu'))
dnn_2.add(Dense(512, activation='relu'))
dnn_2.add(Dense(1, activation='linear'))
# print('Model : DNN 2', dnn_2.summary())
return dnn_2
# compile the keras model
# DNN 3
# define the keras model
elif name == 'dnn_3':
dnn_3 = Sequential()
dnn_3.add(Dense(256, input_dim=input_dim, activation='relu'))
dnn_3.add(Dense(64, activation='relu'))
dnn_3.add(Dense(16, activation='relu'))
dnn_3.add(Dense(4, activation='relu'))
dnn_3.add(Dense(1, activation='linear'))
# print('Model : DNN 3', dnn_3.summary())
# compile the keras model
return dnn_3
# DNN 4
# define the keras model
else:
dnn_4 = Sequential()
dnn_4.add(Dense(512, input_dim=input_dim, activation='relu'))
dnn_4.add(Dense(128, activation='relu'))
dnn_4.add(Dense(32, activation='relu'))
dnn_4.add(Dense(8, activation='relu'))
dnn_4.add(Dense(2, activation='relu'))
dnn_4.add(Dense(1, activation='linear'))
# print('Model : DNN 4', dnn_4.summary())
# compile the keras model
return dnn_4
def process_all(dataset_path, dataset_name, path_for_saving_data):
################## Data Preprocessing ######################
df = pd.read_csv(dataset_path)
encoded_data_frame, encoder_isa, encoder_mem_type = encode_text_features('encode', df,
encoder_isa = None, encoder_mem_type=None)
# total_data = encoded_data_frame.drop(columns = ['arch', 'arch1'])
total_data = encoded_data_frame.drop(columns = ['arch', 'sys'])
total_data = total_data.fillna(0)
X_columns = total_data.drop(columns = 'runtime').columns
X = total_data.drop(columns = ['runtime']).to_numpy()
Y = total_data['runtime'].to_numpy()
# X_columns = total_data.drop(columns = 'PS').columns
# X = total_data.drop(columns = ['runtime','PS']).to_numpy()
# Y = total_data['runtime'].to_numpy()
print('Data X and Y shape', X.shape, Y.shape)
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
print('Train Test Split:', X_train.shape, X_test.shape, Y_train.shape, Y_test.shape)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
################## Data Preprocessing ######################
k = 0
# best_models = [dnn_1, dnn_2, dnn_3, dnn_4]
best_models_name = ['dnn_1', 'dnn_2', 'dnn_3', 'dnn_4']
# best_models_name = ['dnn_4']
df = pd.DataFrame(columns = ['model_name', 'dataset_name', 'r2', 'mse', 'mape', 'mae' ])
for model in best_models_name:
print('Running model number:', k+1, 'with Model Name: ', best_models_name[k])
print('####################################################################')
r2_scores = []
mse_scores = []
mape_scores = []
mae_scores = []
# cv = KFold(n_splits = 10, random_state = 42, shuffle = True)
cv = ShuffleSplit(n_splits=10, random_state=0, test_size = 0.2)
# print(cv)
fold = 1
for train_index, test_index in cv.split(X):
model_orig = create_model(best_models_name[k])
# print("Train Index: ", train_index, "\n")
# print("Test Index: ", test_index)
X_train_fold, X_test_fold, Y_train_fold, Y_test_fold = X[train_index], X[test_index], Y[train_index], Y[test_index]
# print(X_train_fold.shape, X_test_fold.shape, Y_train_fold.shape, Y_test_fold.shape)
model_orig.compile(loss='mae', optimizer='adam', metrics=['mae'])
model_orig.fit(X_train_fold, Y_train_fold, epochs=100, batch_size=10, verbose = 0)
Y_pred_fold = model_orig.predict(X_test_fold)
# save the folds to disk
# data = [X_train_fold, X_test_fold, Y_train_fold, Y_test_fold]
# filename = path_for_saving_data + '/folds_data/' + best_models_name[k] +'_'+ str(fold) + '.pickle'
# pickle.dump(data, open(filename, 'wb'))
# save the model to disk
# serialize model to JSON
# filename_1 = path_for_saving_data + '/models_data/' + best_models_name[k] + '_' + str(fold) + '.json'
# filename_2 = path_for_saving_data + '/models_data/' + best_models_name[k] + '_' + str(fold) + '.h5'
fold = fold + 1
# model_json = model_orig.to_json()
#with open(filename_1, "w") as json_file:
#json_file.write(model_json)
# serialize weights to HDF5
#model_orig.save_weights(filename_2)
# print("Saved model to disk")
# later...
'''
# load json and create model
json_file = open('model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("model.h5")
print("Loaded model from disk")
'''
# some time later...
'''
# load the model from disk
loaded_model = pickle.load(open(filename, 'rb'))
result = loaded_model.score(X_test, Y_test)
print(result)
'''
# scores.append(best_svr.score(X_test, y_test))
'''
plt.figure()
plt.plot(Y_test_fold, 'b')
plt.plot(Y_pred_fold, 'r')
'''
# print('Accuracy =',accuracy_score(Y_test, Y_pred))
r2_scores.append(r2_score(Y_test_fold, Y_pred_fold))
mse_scores.append(mean_squared_error(Y_test_fold, Y_pred_fold))
mape_scores.append(absolute_percentage_error(Y_test_fold, Y_pred_fold))
mae_scores.append(mean_absolute_error(Y_test_fold, Y_pred_fold))
df = df.append({'model_name': best_models_name[k], 'dataset_name': dataset_name
, 'r2': r2_scores, 'mse': mse_scores, 'mape': mape_scores, 'mae': mae_scores }, ignore_index=True)
k = k + 1
print(df.head())
df.to_csv(r'Results_80_20.csv')
```
# Main Function Called
```
dataset_name = 'matmul_lab_omp_physical'
dataset_path = 'C:\\Users\\Rajat\\Desktop\\DESKTOP_15_05_2020\\Evaluating-Machine-Learning-Models-for-Disparate-Computer-Systems-Performance-Prediction\\Dataset_CSV\\PhysicalSystems\\matmul_lab_omp.csv'
path_for_saving_data = 'data\\' + dataset_name
process_all(dataset_path, dataset_name, path_for_saving_data)
```
| github_jupyter |
### Recommendations with MovieTweetings: Getting to Know The Data
Throughout this lesson, you will be working with the [MovieTweetings Data](https://github.com/sidooms/MovieTweetings/tree/master/recsyschallenge2014). To get started, you can read more about this project and the dataset from the [publication here](http://crowdrec2013.noahlab.com.hk/papers/crowdrec2013_Dooms.pdf).
**Note:** There are solutions to each of the notebooks available by hitting the orange jupyter logo in the top left of this notebook. Additionally, you can watch me work through the solutions on the screencasts that follow each workbook.
To get started, read in the libraries and the two datasets you will be using throughout the lesson using the code below.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tests as t
%matplotlib inline
# Read in the datasets
movies = pd.read_csv('https://raw.githubusercontent.com/sidooms/MovieTweetings/master/latest/movies.dat', delimiter='::', header=None, names=['movie_id', 'movie', 'genre'], dtype={'movie_id': object}, engine='python')
reviews = pd.read_csv('https://raw.githubusercontent.com/sidooms/MovieTweetings/master/latest/ratings.dat', delimiter='::', header=None, names=['user_id', 'movie_id', 'rating', 'timestamp'], dtype={'movie_id': object, 'user_id': object, 'timestamp': object}, engine='python')
```
#### 1. Take a Look At The Data
Take a look at the data and use your findings to fill in the dictionary below with the correct responses to show your understanding of the data.
```
# number of movies
print("The number of movies is {}.".format(movies.shape[0]))
# number of ratings
print("The number of ratings is {}.".format(reviews.shape[0]))
# unique users
print("The number of unique users is {}.".format(reviews.user_id.nunique()))
# missing ratings
print("The number of missing reviews is {}.".format(int(reviews.rating.isnull().mean()*reviews.shape[0])))
# the average, min, and max ratings given
print("The average, minimum, and max ratings given are {}, {}, and {}, respectively.".format(np.round(reviews.rating.mean(), 0), reviews.rating.min(), reviews.rating.max()))
# number of different genres
genres = []
for val in movies.genre:
try:
genres.extend(val.split('|'))
except AttributeError:
pass
# we end up needing this later
genres = set(genres)
print("The number of genres is {}.".format(len(genres)))
# Use your findings to match each variable to the correct statement in the dictionary
a = 53968
b = 10
c = 7
d = 31245
e = 15
f = 0
g = 4
h = 712337
i = 28
dict_sol1 = {
'The number of movies in the dataset': d,
'The number of ratings in the dataset': h,
'The number of different genres': i,
'The number of unique users in the dataset': a,
'The number missing ratings in the reviews dataset': f,
'The average rating given across all ratings': c,
'The minimum rating given across all ratings': f,
'The maximum rating given across all ratings': b
}
# Check your solution
t.q1_check(dict_sol1)
```
#### 2. Data Cleaning
Next, we need to pull some additional relevant information out of the existing columns.
For each of the datasets, there are a couple of cleaning steps we need to take care of:
#### Movies
* Pull the date from the title and create new column
* Dummy the date column with 1's and 0's for each century of a movie (1800's, 1900's, and 2000's)
* Dummy column the genre with 1's and 0's for each genre
#### Reviews
* Create a date out of time stamp
You can check your results against the header of my solution by running the cell below with the **show_clean_dataframes** function.
```
# pull date if it exists
create_date = lambda val: val[-5:-1] if val[-1] == ')' else np.nan
# apply the function to pull the date
movies['date'] = movies['movie'].apply(create_date)
# Return century of movie as a dummy column
def add_movie_year(val):
if val[:2] == yr:
return 1
else:
return 0
# Apply function
for yr in ['18', '19', '20']:
movies[str(yr) + "00's"] = movies['date'].apply(add_movie_year)
# Function to split and return values for columns
def split_genres(val):
try:
if val.find(gene) >-1:
return 1
else:
return 0
except AttributeError:
return 0
# Apply function for each genre
for gene in genres:
movies[gene] = movies['genre'].apply(split_genres)
movies.head() #Check what it looks like
import datetime
change_timestamp = lambda val: datetime.datetime.fromtimestamp(int(val)).strftime('%Y-%m-%d %H:%M:%S')
reviews['date'] = reviews['timestamp'].apply(change_timestamp)
# now reviews and movies are the final dataframes with the necessary columns
reviews.to_csv('./reviews_clean.csv')
movies.to_csv('./movies_clean.csv')
```
| github_jupyter |
# VGG16
This notebook will recreate the VGG16 model from [FastAI Lesson 1](http://course.fast.ai/lessons/lesson1.html) ([wiki](http://wiki.fast.ai/index.php/Lesson_1_Notes)) and [FastAI Lesson 2](http://course.fast.ai/lessons/lesson2.html) ([wiki](http://wiki.fast.ai/index.php/Lesson_2_Notes))
The [Oxford Visual Geometry Group](http://www.robots.ox.ac.uk/~vgg/research/very_deep/) created a 16 layer deep ConvNet which placed first in certain aspects of the 2014 Image-Net competition. Their NN was trained on 1000's of images from the image-net database for all sorts of objects. Instead of retraining the NN ourselves, it is possible to download the weights from their trained NN. By using a pretained model, we can create their work and also adapt it to our own classification task.
## Repurposing the Pretrained VGG16 Model
The pretrained VGG16 model was trained using [image-net data](http://image-net.org/explore). This data is made up of thousands of categories of "things" which each have many framed, well-lit, and focused photos. Knowing these characteristics of the training images will help us understand how this model can and can't work for our dogs vs. cats task.
The image-net data is more specific than just dogs and cats, it has been trained on specific breeds of each. **One hot encoding** is used to label images. This is where the label is a vector of 0's of size equal to the number of categories, but has a 1 where the category is true. So for [dogs, cats] a label of [0, 1] would mean it is a cat.
By repurposing the image-net VGG16 to look for just cats and dogs, we are **Finetuning** the mode. This is where we start with a model that already solved a similar problem. Many of the parameters should be the same, so we only select a subset of them to re-train. Finetuning will replace the 1000's of image-net categories with the 2 it found in our directory structure (dogs and cats). It does this by removing the last layer (with the keras .pop method) and then adding a new output layer with size 2. This will leave us with a pretrained VGG16 model specifically made for categorizing just cats and dogs.
Why do Finetuning instead of training our own network?
Image-net NN has already learned a lot about what the world looks like. The first layer of a NN looks for basic shapes, patterns, or gradients ... which are known as **gabor filters**. These images come from this paper ([Visualizing and Understanding Convolutional Networks](https://arxiv.org/pdf/1311.2901.pdf)):
<img src="images/Layer1.png" alt="Drawing" style="width: 600px;"/>
The second layer combines layer 1 filters to create newer, more complex filters. So it turns multiple line filters into corner filters, and combines lines into curved edges, for example.
<img src="images/Layer2.png" alt="Drawing" style="width: 600px;"/>
Further into the hidden layers of a NN, filters start to find more complex shapes, repeating geometric patterns, faces, etc.
<img src="images/Layer3.png" alt="Drawing" style="width: 600px;"/>
<img src="images/Layer4-5.png" alt="Drawing" style="width: 600px;"/>
VGG16 has ... 16 ... layers, so there are tons of filters created at each layer. Finetuning keeps these lower level filters which have been created already and then combines them in a different way to address different inputs (i.e. cats and dogs instead of 1000's of categories). Neural networks pretrained on HUGE datasets have already found all of these lower level filters, so we don't need to spend weeks doing that part ourselves. Finetuning usually works best on the second to last layer, but it's also a good idea to try it at every layer.
Additional information on fine-tuning (aka transfer-learning) can be found on Stanford's CS231n website [here](http://cs231n.github.io/transfer-learning/).
## VGG Detailed Sizing
A rough calculation for the memory requirements of running VGG16 can be calculated, as was done in the [Stanford CS231n CNN Course](http://cs231n.#GBs required for 16 image mini-batch
size = ((15184000 + 3*4096000) * 4 * 2 * 16) / (1024**3)
print(str(round(size,2)) + 'GB')
This makes sense when tested with my 6GB GTX980ti. A mini-batch size of 32 ran out of VRAM. The GPU has to run other stuff too and has a normal load of around 0.7GB.github.io/convolutional-networks/). At each layer, we can find the size of the memory required and weights. Notice that most of the memory (and compute time) is used in the first layers, while most of the parameters are in the last FC layers. Notice that the POOL layers reduce the spatial dimensions by 50% (don't effect depth) and do not introduce any new parameters.
| Layer | Size/Memory | Weights |
|:--- |:--- |:--- |
| INPUT | 224x224x3 = 150K | 0 |
| CONV3-64 | 224x224x64 = 3.2M | (3x3x3)x64 = 1,728 |
| CONV3-64 | 224x224x64 = 3.2M | (3x3x3)x64 = 36,864 |
| POOL2 | 112x112x64 = 800K | 0 |
| CONV3-128 | 112x112x128 = 1.6M | (3x3x64)x128 = 73,728 |
| CONV3-128 | 112x112x128 = 1.6M | (3x3x128)x128 = 147,456 |
| POOL2 | 56x56x128 = 400K | 0 |
| CONV3-256 | 56x56x256 = 800K | (3x3x128)x256 = 294,912 |
| CONV3-256 | 56x56x256 = 800K | (3x3x256)x256 = 589,824 |
| CONV3-256 | 56x56x256 = 800K | (3x3x256)x256 = 589,824 |
| POOL2 | 28x28x256 = 200K | 0 |
| CONV3-512 | 28x28x512 = 400K | (3x3x256)x512 = 1,179,648 |
| CONV3-512 | 28x28x512 = 400K | (3x3x512)x512 = 2,359,296 |
| CONV3-512 | 28x28x512 = 400K | (3x3x512)x512 = 2,359,296 |
| POOL2 | 14x14x512 = 100K | 0 |
| CONV3-512 | 14x14x512 = 100K | (3x3x512)x512 = 2,359,296 |
| CONV3-512 | 14x14x512 = 100K | (3x3x512)x512 = 2,359,296 |
| CONV3-512 | 14x14x512 = 100K | (3x3x512)x512 = 2,359,296 |
| POOL2 | 7x7x512 = 25K | 0 |
| FC | 1x1x4096 = 4K | 7x7x512x4096 = 102,760,448 |
| FC | 1x1x4096 = 4K | 4096x4096 = 16,777,216 |
| FC | 1x1x1000 = 1K | 4096x1000 = 4,096,000 |
TOTAL MEMORY = (LayerSizes + 3\*Weights) \* 4 Bytes \* 2 (fwd and bkwd passes) \* images/batch
```
#GBs required for 16 image mini-batch
size = ((15184000 + 3*4096000) * 4 * 2 * 16) / (1024**3)
print(str(round(size,2)) + 'GB')
```
This makes sense when tested with my 6GB GTX980ti. A mini-batch size of 32 ran out of VRAM. The GPU has to run other stuff too and has a normal load of around 0.7GB.
## Custom Written VGG16 Model
This section will go step by step through the process of recreating the VGG16 model from scratch, using python and Keras.
### Prepare the Workspace
Here we will set matplotlib plots to load directly in this notebook, load all of the python packages needed, check where our data directory is saved, as well as save the pre-trained VGG16 model from the web url.
```
%matplotlib inline
import json
from matplotlib import pyplot as plt
import numpy as np
from numpy.random import random, permutation
from scipy import misc, ndimage
from scipy.ndimage.interpolation import zoom
import keras
from keras import backend as K
from keras.utils.data_utils import get_file
from keras.models import Sequential, Model
from keras.layers.core import Flatten, Dense, Dropout, Lambda
from keras.layers import Input
from keras.layers.convolutional import Convolution2D, MaxPooling2D, ZeroPadding2D
from keras.optimizers import SGD, RMSprop
from keras.preprocessing import image
data_path = "../../fastAI/deeplearning1/nbs/data/dogscats/"
!ls $data_path
```
The Keras 'get_file' function will download a file from a URL if it's not already in the cache. The !ls command shows that the file is in the .keras/models/ directory which we specified as our cache location:
```
FILE_URL = "http://files.fast.ai/models/";
FILE_CLASS = "imagenet_class_index.json";
fpath = get_file(FILE_CLASS, FILE_URL+FILE_CLASS, cache_subdir='models')
!ls ~/.keras/models
```
The class file itself is a dictionary where keys are strings from 0 to 1000 and the values are names of everyday objects. Let's open the file using 'json.load' and convert it to a 'classes' array:
```
with open(fpath) as f:
class_dict = json.load(f)
classes = [class_dict[str(i)][1] for i in range(len(class_dict))]
print(class_dict['809'])
print(class_dict['809'][1])
```
Check how many objects are in the 'classes' array and then print the first 5:
```
print(len(classes))
print(classes[:5])
```
### Build the Model
We need to define the NN model architecture and then load the pre-trained weights (that we downloaded) into it. The VGG model has 1 type of convolutional block and 1 type of fully-connected block. We'll create functions to define each of these blocks and then call them later to actually instantiate the VGG model:
```
def ConvBlock(layers, model, filters):
for i in range(layers):
model.add(ZeroPadding2D((1,1)))
model.add(Convolution2D(filters, 3, 3, activation='relu'))
model.add(MaxPooling2D((2,2), strides=(2,2)))
def FullyConnectedBlock(model):
model.add(Dense(4096, activation='relu'))
model.add(Dropout(0.5))
```
### Preprocessing
The original VGG model has a mean of zero for each channel, obtained by subtracting the average of each RGB channel. It also expects data in the BGR order, so we need to do some preprocessing:
```
vgg_mean = np.array([123.68, 116.779, 103.939]).reshape((3,1,1))
def vgg_preprocess(x):
x = x - vgg_mean #subtract mean
return x[:, ::-1] #RGB -> BGR
```
### Instantiate the Model
The convolutional layers help find patterns in the images, while the fully connected (Dense) layers combine patterns across an image. The following function calls the other functions written above. It will instantiate a 16 layer VGG model:
```
def VGG16():
model = Sequential()
model.add(Lambda(vgg_preprocess, input_shape=(3,224,224)))
ConvBlock(2, model, 64)
ConvBlock(2, model, 128)
ConvBlock(3, model, 256)
ConvBlock(3, model, 512)
ConvBlock(3, model, 512)
model.add(Flatten())
FullyConnectedBlock(model)
FullyConnectedBlock(model)
model.add(Dense(1000, activation='softmax'))
return model
model = VGG16()
```
### Load Pretrained Weights
Now that a VGG16 model has been created, we can load it up with the pretrained weights we downloaded earlier. This step prevents us from having to train the NN on the 1000's of image-net samples:
```
fweights = get_file('vgg16.h5', FILE_URL+'vgg16.h5', cache_subdir='models')
model.load_weights(fweights)
```
### Grab Batches of Images
Now the NN is setup to use, so we can grab batches of images and start using the NN to predict their output classes:
```
batch_size = 4
```
The following helper function will use the Keras [image.ImageDataGenerator](https://keras.io/preprocessing/image/) object with its **flow_from_directory()** method to start pulling batches of images from the directory we tell it to. It returns an Iterator which we can call with next to get the next *batch_size* amount of image/label pairs:
```
def get_batches(dirname, gen=image.ImageDataGenerator(), shuffle=True, batch_size=batch_size, class_mode='categorical'):
return gen.flow_from_directory(data_path+dirname, target_size=(224,224), class_mode=class_mode, shuffle=shuffle, batch_size=batch_size)
batches = get_batches('sample/train', batch_size=batch_size)
val_batches = get_batches('sample/valid', batch_size=batch_size)
def show_plots(ims, figsize=(12,6), rows=1, interp=False, titles=None):
if type(ims[0]) is np.ndarray:
ims = np.array(ims).astype(np.uint8)
if (ims.shape[-1] != 3):
ims = ims.transpose((0,2,3,1))
f = plt.figure(figsize=figsize)
cols = len(ims)//rows if len(ims) % 2 == 0 else len(ims)//rows + 1
for i in range(len(ims)):
sp = f.add_subplot(rows, cols, i+1)
sp.axis('Off')
if titles is not None:
sp.set_title(titles[i], fontsize=16)
plt.imshow(ims[i], interpolation=None if interp else 'none')
```
Checking the shape of imgs, we can see that this array holds 4 images, each with 3 channels (BGR) and are of size 224x224 pixels
```
imgs,labels = next(batches)
print(imgs.shape)
print(labels[0])
show_plots(imgs, titles=labels)
```
### Predict
Now we will call the predict method on our Sequential Keras model. This returns a vector of size 1000 with probabilities that each image belongs to one of the 1000 image-net categories. The function below is written to find the highest probability for each image in our batch:
```
def pred_batch(imgs):
preds = model.predict(imgs)
idxs = np.argmax(preds, axis=1)
print('Shape: {}'.format(preds.shape))
print('Predictions prob/class: ')
for i in range(len(idxs)):
idx = idxs[i]
print (' {:.4f}/{}'.format(preds[i, idx], classes[idx]))
pred_batch(imgs)
model.summary()
```
## Fine Tuning
#### New Output Layer
Retrain the Last Layer, keeping everything else the same. Let's replace the final layer with a 2-node softmax activation). Now the NN will use everything it learned on the whole dataset to only classify things into 1 of 2 categories.
```
model.pop()
for layer in model.layers: layer.trainable=False
model.add(Dense(2, activation='softmax'))
```
#### Freezing Layers
Setting all the layers in the model to "trainable=False" means that their weights will not be updated during training. If all of them were untrainable, then training wouldn't actually do anything! Adding a new Dense layer after these **frozen** layers will make it trainable. Generally, if we are adding a completely new layer and initializing it from scratch, we would only want to train that layer for at least 1-2 epochs. This allows the final layer to have weights which are closer to what they should be (at least compared to random init values). Once they are set, **unfreeze** earlier Dense layers so their weights can be updated during training. Because the final layer is now "closer" to the previous ones, the training process won't have vastly differing weights which would have made drastic updates to the previous layers. Learning will proceed much more smoothly.
#### Pre-calculate CONV Layer Outputs
It is usually best to freeze the convolutional layers and not retrain them. This means that the weights of the filters in the CONV layers do not change. If the original weights were from a huge dataset like ImageNet, this is probably a good thing because that dataset is so vast that it already contains small, medium, and large complexity filters (like edges, corners, and objects) that can be reused on our new dataset. Our dataset is unlikely to contain any edges, colors, etc. which has not already been seen from the ImageNet data. Because we are not going to update the CONV layer weights (filters) it is best to **pre-calculate the CONV layer outputs**.
Way back at the beginning of time, the NN was created from scratch. All of the weights are each layer were randomly initialized and it was trained on a dataset. All the data was passed through once (an epoch) and the weights were updated. Then it was passed over many many more times (10's or 100's of epochs) to really train the NN and lock in the ideal weights. Therefore, **the weights of the CONV layers start to represent the training set**. When we start adding new Dense layers at the output, we don't need to run through all of the data again, the CONV layers already represent that data as seen for XX epochs. We can just compute the output from all the CONV layers and treat that as an input to our new Dense layers ... this will save a lot of time training. Now we train the NN just using the CONV outputs fed into our new Dense layers, once it is trained we can load those updated last layer weights onto the original entire CNN and have a completely updated model.
| github_jupyter |
Kernelized support vector machines are powerful models and perform well on a variety of datasets.
1. SVMs allow for complex decision boundaries, even if the data has only a few features.
2. They work well on low-dimensional and high-dimensional data (i.e., few and many features), but don’t scale very well with the number of samples.
> **Running an SVM on data with up to 10,000 samples might work well, but working with datasets of size 100,000 or more can become challenging in terms of runtime and memory usage.**
3. SVMs requires careful preprocessing of the data and tuning of the parameters. This is why, these days, most people instead use tree-based models such as random forests or gradient boosting (which require little or no preprocessing) in many applications.
4. SVM models are hard to inspect; it can be difficult to understand why a particular prediction was made, and it might be tricky to explain the model to a nonexpert.
### Important Parameters
The important parameters in kernel SVMs are the regularization parameter C, the choice of the kernel, and the kernel-specific parameters. gamma and C both control the complexity of the model, with large values in either resulting in a more complex model. Therefore, good settings for the two parameters are usually strongly correlated, and C and gamma should be adjusted together.
```
#load libraries
import pandas as pd
import numpy as np
#Supervised learning
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
#Load data set
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
cancer =pd.DataFrame(cancer.data)
cancer.head()
#Split data set in train 75% and test 25%
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, test_size=0.25, stratify=cancer.target, random_state=66)
print("X_train shape: {}".format(X_train.shape))
print("y_train shape: {}".format(y_train.shape))
print("X_test shape: {}".format(X_test.shape))
print("y_test shape: {}".format(y_test.shape))
list(cancer.target_names)
list(cancer.feature_names)
## Create an SVM classifier and train it on 75% of the data set.
svc =SVC(probability=True)
svc.fit(X_train, y_train)
## Create an SVM classifier and train it on 70% of the data set.
#clf = SVC(probability=True)
#clf.fit(X_train, y_train)
# Analyze accuracy of predictions on 25% of the holdout test sample.
classifier_score_test = svc.score(X_test, y_test)
classifier_score_train = svc.score(X_train, y_train)
print 'The classifier accuracy on the test set is {:.2f}'.format(classifier_score_test)
print 'The classifier accuracy on the training set is {:.2f}'.format(classifier_score_train)
print("Accuracy on training set: {:.2f}".format(svc.score(X_train, y_train)))
#print("Accuracy on test set: {:.2f}".format(svc.score(X_test, y_test)))
```
> **The model overfits quite substantially, with a perfect score on the training set and only 63% accuracy on the test set.**
While SVMs often perform quite well, they are very sensitive to the settings of the parameters and to the scaling of the data. In particular, they require all the features to vary on a similar scale. Let’s look at the minimum and maximum values for each feature, plotted in log-spac
```
# import Matplotlib (scientific plotting library)
import matplotlib.pyplot as plt
# allow plots to appear within the notebook
%matplotlib inline
plt.plot(X_train.min(axis=0), 'o', label="min")
plt.plot(X_train.max(axis=0), '^', label="max")
plt.legend(loc=4)
plt.xlabel("Feature index")
plt.ylabel("Feature magnitude")
plt.yscale("log")
```
## Preprocessing data for SVM -Rescaling the data
SVMs,is very sensitive to the scaling of the data. Therefore, a common practice is to adjust the features so that the data representation is more suitable for these algorithms. Often, this is a simple per-feature rescaling and shift of the data
One way to resolve model overfitting problem is by rescaling each feature so that they are all approximately on the same scale. A common rescaling method for kernel SVMs is to scale the data such that all features are between 0 and 1. We will see how to do this using the MinMaxScaler
```
# compute the minimum value per feature on the training set
min_on_training = X_train.min(axis=0)
# compute the range of each feature (max - min) on the training set
range_on_training = (X_train - min_on_training).max(axis=0)
# subtract the min, and divide by range
# afterward, min=0 and max=1 for each feature
X_train_scaled = (X_train - min_on_training) / range_on_training
print("Minimum for each feature\n{}".format(X_train_scaled.min(axis=0)))
print("Maximum for each feature\n {}".format(X_train_scaled.max(axis=0)))
# use THE SAME transformation on the test set,
# using min and range of the training set (see Chapter 3 for details)
X_test_scaled = (X_test - min_on_training) / range_on_training
svc = SVC()
svc.fit(X_train_scaled, y_train)
print("Accuracy on training set: {:.3f}".format(
svc.score(X_train_scaled, y_train)))
print("Accuracy on test set: {:.3f}".format(svc.score(X_test_scaled, y_test)))
```
### MinMaxScaler
```
from sklearn.preprocessing import MinMaxScaler
# preprocessing using 0-1 scaling
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# learning an SVM on the scaled training data
svm =SVC()
svm.fit(X_train_scaled, y_train)
# scoring on the scaled test set
print("Scaled test set accuracy: {:.2f}".format(
svm.score(X_test_scaled, y_test)))
# preprocessing using zero mean and unit variance scaling
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# learning an SVM on the scaled training data
svm.fit(X_train_scaled, y_train)
# scoring on the scaled test set
print("SVM test accuracy: {:.2f}".format(svm.score(X_test_scaled, y_test)))
```
### Tuning the parameters
The gamma parameter is the one shown in the formula given in the previous section, which controls the width of the Gaussian kernel. It determines the scale of what it means for points to be close together. The C parameter is a regularization parameter, similar to that used in the linear models. It limits the importance of each point
```
from sklearn.grid_search import GridSearchCV
from sklearn import cross_validation
from sklearn.cross_validation import KFold, cross_val_score
from sklearn.preprocessing import StandardScaler
# Test options and evaluation metric
num_folds = 10
num_instances = len(X_train)
seed = 7
scoring = 'accuracy'
# Tune scaled SVM
scaler = StandardScaler().fit(X_train)
rescaledX = scaler.transform(X_train)
c_values = [0.1, 0.3, 0.5, 0.7, 0.9, 1.0, 1.3, 1.5, 1.7, 2.0]
kernel_values = [ 'linear' , 'poly' , 'rbf' , 'sigmoid' ]
param_grid = dict(C=c_values, kernel=kernel_values)
model = SVC()
kfold = cross_validation.KFold(n=num_instances, n_folds=num_folds, random_state=seed)
grid = GridSearchCV(estimator=model, param_grid=param_grid, scoring=scoring, cv=kfold)
grid_result = grid.fit(rescaledX, y_train)
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
for params, mean_score, scores in grid_result.grid_scores_:
print("%f (%f) with: %r" % (scores.mean(), scores.std(), params))
```
Scaling the data made a huge difference! Now we are actually in an underfitting regime, where training and test set performance are quite similar but less close to 100% accuracy. From here, we can try increasing either C or gamma to fit a more complex model. For example:
### Decision Boundaries
1. What are decision boundaries?
>Demonstrate the classification result by ploting the decision boundery
Sample usage of Nearest Neighbors classification. It will plot the decision boundaries for each class:
```
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn import svm, datasets
def decision_plot(X_train, y_train, n_neighbors, weights):
h = .02 # step size in the mesh
Xtrain = X_train[:, :2] # we only take the first two features.
#================================================================
# Create color maps
#================================================================
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
#================================================================
# we create an instance of SVM and fit out data.
# We do not scale ourdata since we want to plot the support vectors
#================================================================
C = 1.0 # SVM regularization parameter
svm = SVC(kernel='linear', random_state=0, gamma=0.2, C=C).fit(Xtrain, y_train)
rbf_svc = SVC(kernel='rbf', gamma=0.7, C=C).fit(Xtrain, y_train)
poly_svc = SVC(kernel='poly', degree=3, C=C).fit(Xtrain, y_train)
#lin_svc = svm.LinearSVC(C=C).fit(Xtrain, y_train)
#================================================================
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, m_max]x[y_min, y_max].
#================================================================
x_min, x_max = Xtrain[:, 0].min() - 1, Xtrain[:, 0].max() + 1
y_min, y_max = Xtrain[:, 1].min() - 1, Xtrain[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1),
np.arange(y_min, y_max, 0.1))
Z = svm.predict(np.c_[xx.ravel(), yy.ravel()])
#================================================================
# Put the result into a color plot
#================================================================
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot also the training points
plt.scatter(Xtrain[:, 0], Xtrain[:, 1], c=y_train, cmap=cmap_bold)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
#plt.title("2-Class classification (k = %i, weights = '%s')"
# % (n_neighbors, weights))
plt.show()
%matplotlib inline
plt.rcParams['figure.figsize'] = (15, 9)
plt.rcParams['axes.titlesize'] = 'large'
# create a mesh to plot in
x_min, x_max = Xtrain[:, 0].min() - 1, Xtrain[:, 0].max() + 1
y_min, y_max = Xtrain[:, 1].min() - 1, Xtrain[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.1),
np.arange(y_min, y_max, 0.1))
# title for the plots
titles = ['SVC with linear kernel',
'LinearSVC (linear kernel)',
'SVC with RBF kernel',
'SVC with polynomial (degree 3) kernel']
for i, clf in enumerate((svm, rbf_svc, poly_svc)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(2, 2, i + 1)
plt.subplots_adjust(wspace=0.4, hspace=0.4)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.coolwarm, alpha=0.8)
# Plot also the training points
plt.scatter(Xtrain[:, 0], Xtrain[:, 1], c=y_train, cmap=plt.cm.coolwarm)
plt.xlabel('mean radius')
plt.ylabel('mean texture')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xticks(())
plt.yticks(())
plt.title(titles[i])
plt.show()
```
</div>
<div class="section" id="classification-accuracy using-confusion-matrix">
<h1 style="color:blue;">Classification Accuracy Using Confusion Matrix <a class="headerlink" href="#classification-accuracy using-confusion-matrix" title="Permalink to this headline">¶</a></h1>
Confusion matrix describes the performance of a classification table.
Every observation in the testing set is represented in the matrix. For a 2 classification problem with 2 responses is a 2 by 2 matrix.
~~~~
Model says "+" Model says "-"
Actual: "+" True positive | False negative
----------------------------------
Actual: "-" False positive | True negative
~~~~
* **Basic Terminology**
* True Positive Model correctly predicts tumor is benign (1- response
* True Negative Model correctly predicts tumor is Malignant (0-response)
* False Positive Model incorrectly predicts that the tumor is benign while it is not (Type 1 error)
* False Negative Model incorrectly predicts that the tumor is benign, while it is not (Type II error)
```
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler
# prepare the model
scaler = StandardScaler().fit(X_train)
rescaledX = scaler.transform(X_train)
model = SVC(kernel='linear', random_state=0, gamma=0.2, C=0.3, probability=False)
model.fit(rescaledX, y_train)
# estimate accuracy on validation dataset
rescaledtestX = scaler.transform(X_test)
predictions = model.predict(rescaledtestX)
n_classes = cancer.target_names.shape[0]
print(accuracy_score(y_test, predictions))
print(confusion_matrix(y_test, predictions, labels=range(n_classes)))
print(classification_report(y_test, predictions, target_names=cancer.target_names ))
#print the first 25 true and predicted responses
#print 'True:', (y_test.values)[0:25]
print 'Pred:', predictions[0:25]
```
## Matrics computed from Confusion matrix
~~~~
Model says "+" Model says "-"
Actual: "+" 50 (TP ) | 3 (FN)
----------------------------------
Actual: "-" 1 (FP) | 89 (TN)
~~~~
### Description of Results
1. **Classification Accuracy** - Answers the question how often is the classifier correct?? TP+TN/Total
* An accuracy score of 97% indicates that svm classifier is 97% correct in predicting the tumor is
2. **Confusion Matrix**
- Majority of the prediction fall on the diagonal line of the matrix, of the 53 Malignant diagnosis, 50 were correctly classified as malignant tumor, and of the 90 Benign predictions, 89 were classified as benign tumor.
3. **Summary of classification report**.
* Precision/positive predicted value(PPV) - Statistical measure that describes the performance of a diagnostic test.
* When a positive value is predicted, how often is the predicition correct?(How precise is the classifier when predicting positives instances. In this case, SVM Classifier is 97% precise in predicting a malignat(cancerous) tumor
* Recall/Sensitivity/True Positive rate (TPR) - Quantifies avoidance of false negatives- When the actual value is positive, how often is the value correct..."How sensitivy the classifier is in detecting a positive instance?
* **94% (50/53)of the patients classified as having a malignant tumor, while 99% (89/90)classified as having a benign tumor**
* In general the SVM classifier is 97% correct, in predicting when the actual value is positive
* f1-Score - F1 score (also F-score or F-measure) is a measure of a test's accuracy. It considers both the precision p and the recall r of the test to compute the score. f1-Score = 2x((Precision* Recall)/(Precision+Recall)) (http://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html)
### Conclusion
* Confusion matrix gives you a more compplete picture of how well your classier is performaing
* Allows you to compute various calassification metrics and these metrics can guide your model selction
### Which Metrics should you focus on?
* Choice of metric depends on **business objective**
* **Spam filter** (positve class is spam); Optimize for precision or specificity because false negatives (spam goes into inbox) are more acceptable than false positives (non-spam is caught by spam filter)
* **Fradulent transaction dector** (positive class is "fraud). Optimize for sensitivity because false positives (normal transactions that are flagged as possible frauds) are more acceptable that false negatives (fradulent transactions that are not detected)
### Adjusting The Classification Threshold /Optimizing parameters
```
#print the first 10 predicted response
svm.predict(X_test)[0:10]
```
### Receiver operating characteristic (ROC) curve.
In statistical modeling and machine learning, a commonly-reported performance measure of model accuracy is Area Under the Curve (AUC), where, by “curve”, the ROC curve is implied. ROC is a term that originated from the Second World War and was used by radar engineers (it had nothing to do with machine learning or pattern recognition).
```
# Plot the receiver operating characteristic curve (ROC).
from sklearn.metrics import roc_curve, auc
plt.figure(figsize=(20,10))
probas_ = model.predict_proba(X_test)
fpr, tpr, thresholds = roc_curve(y_test, probas_[:, 1])
roc_auc = auc(fpr, tpr)
plt.plot(fpr, tpr, lw=1, label='ROC fold (area = %0.2f)' % (roc_auc))
plt.plot([0, 1], [0, 1], '--', color=(0.6, 0.6, 0.6), label='Random')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
#plt.axes().set_aspect(1)
```
### Reference
https://github.com/InesdeSantiago/machine_learning_cca1/blob/master/Module2.py
| github_jupyter |
```
import pandas as pd
import numpy as np
from tokenizer import tokenize
stopwords = ['i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've", "you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', 'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their', 'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', 'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', 'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', 'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after', 'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further', 'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more', 'most', 'other', 'some', 'such', 'no', 'nor', 'not', 'only', 'own', 'same', 'so', 'than', 'too', 'very', 's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', 've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn', "hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'mustn', "mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", 'won', "won't", 'wouldn', "wouldn't"]
paragraphs = ["""So, keep working. Keep striving. Never give up. Fall down seven times, get up eight.
Ease is a greater threat to progress than hardship. Ease is a greater threat to progress
than hardship. So, keep moving, keep growing, keep learning. See you at work."""]
text = ' '.join(paragraphs)
print(text)
df = pd.DataFrame()
df['sentences'] = list(map(lambda x: x.replace('\n','').replace(' ',''), text.split('. ')))
df.head()
def token(sentence):
sentence_result = []
for token in list(tokenize(sentence)):
word = token[1]
if word and len(word)>1 and word.lower() not in stopwords:
sentence_result.append(word.lower())
return ' '.join(sentence_result)
df['tokenized_sentences'] = df['sentences'].map(token)
df.head()
tokenized_words = [word for sentence in df['tokenized_sentences'] for word in sentence.split()]
frequency_words = {}
for word in set(tokenized_words):
frequency_words[word] = tokenized_words.count(word)
max_frequency = max(frequency_words.values())
for word in frequency_words.keys():
frequency_words[word] /= max_frequency
def get_sentence_weight(sentence, frequency_words=frequency_words):
return np.sum([frequency_words[word] for word in sentence.split()])
df['sentence_weight'] = df['tokenized_sentences'].map(get_sentence_weight)
df.sort_values(by = ['sentence_weight'], ascending = False, inplace = True)
df.head()
k = 2
summary = [sentence+'.' for sentence in df.iloc[0:k]['sentences']]
' '.join(summary)
```
| github_jupyter |
# Read DEIMOS metadata from XML file
Extract relevant information about the DEIMOS bands into dataframes.
**NOTE**: DEIMOS bands are provided as `NIR-R-G-B`, while we store them in `EOPatches` as `B-G-R-NIR` as in Sentinel-2 datasets. This means that we will have to swap the info read from XML files in `split_per_band`.
```
import os
from xml.etree import ElementTree as ET
import pandas as pd
from fs_s3fs import S3FS
```
## Config
```
instance_id = ''
aws_access_key_id = ''
aws_secret_access_key = ''
filesystem = S3FS(bucket_name='',
aws_access_key_id=aws_access_key_id,
aws_secret_access_key=aws_secret_access_key)
tiles_folder = ''
def tag_parser(el_iterator, vals_dict, attr='text', attrib_key=None):
for sub in el_iterator:
if attr == 'attrib':
vals_dict[sub.tag] = getattr(sub, attr)[attrib_key]
else:
vals_dict[sub.tag] = getattr(sub, attr)
def multitag_parser(el_iterator, vals_dict, attr='text'):
children = []
for sub in el_iterator:
tag_name = sub.tag
x = {}
tag_parser(sub.getchildren(), x)
children.append(x)
vals_dict[tag_name] = children
def parse_bbox(el_iterator, vals_dict, outname, use_xy=True):
appendices = ['X', 'Y'] if use_xy else ['LAT', 'LON']
vertex_dict = {f'FRAME_{appendix}': [] for appendix in appendices}
for vertex in el_iterator:
for appendix in appendices:
vertex_dict[f'FRAME_{appendix}'].append(vertex.find(f'./FRAME_{appendix}').text)
if use_xy:
vals_dict[outname] = [min(vertex_dict['FRAME_X']), min(vertex_dict['FRAME_Y']),
max(vertex_dict['FRAME_X']), max(vertex_dict['FRAME_Y'])]
else:
vals_dict[outname] = [min(vertex_dict['FRAME_LAT']), min(vertex_dict['FRAME_LON']),
max(vertex_dict['FRAME_LAT']), max(vertex_dict['FRAME_LON'])]
def split_per_band(columns, column, query_keys, revert_bands=True,
index_col='BAND_INDEX', n_bands=4):
for valdict in columns[column]:
if all([key in set(valdict.keys()) for key in query_keys]):
for key in query_keys:
idx = int(valdict[index_col])
if revert_bands:
idx = n_bands-idx+1
columns[f'{key}_{idx}'] = valdict[key]
columns.pop(column, None)
def parse_deimos_metadata_file(metadata_file, filesystem):
tree = ET.parse(filesystem.open(metadata_file))
root = tree.getroot()
columns = {}
tag_parser(root.findall('./Dataset_Id/'), columns)
tag_parser(root.findall('./Production/'), columns)
tag_parser(root.findall('./Data_Processing/'), columns)
tag_parser(root.findall('./Raster_CS/'), columns)
parse_bbox(root.findall('./Dataset_Frame/'), columns, 'bbox')
tag_parser(root.findall('./Raster_Encoding/'), columns)
tag_parser(root.findall('./Data_Access/'), columns)
tag_parser(root.findall('./Data_Access/Data_File/'), columns, attr='attrib', attrib_key='href')
tag_parser(root.findall('./Raster_Dimensions/'), columns)
multitag_parser(root.findall('./Image_Interpretation/'), columns)
multitag_parser(root.findall('./Image_Display/'), columns)
tag_parser(root.findall('./Dataset_Sources/Source_Information/Coordinate_Reference_System/'), columns)
tag_parser(root.findall('./Dataset_Sources/Source_Information/Scene_Source/'), columns)
multitag_parser(root.findall('./Dataset_Sources/Source_Information/Quality_Assessment/'), columns)
parse_bbox(root.findall('./Dataset_Sources/Source_Information/Source_Frame/'),
columns, 'source_frame_bbox_latlon', use_xy=False)
split_per_band(columns,
'Band_Statistics',
['STX_STDV', 'STX_MEAN', 'STX_MIN', 'STX_MAX'])
split_per_band(columns,
'Spectral_Band_Info',
['PHYSICAL_GAIN', 'PHYSICAL_BIAS', 'PHYSICAL_UNIT', 'ESUN'])
return pd.DataFrame([columns])
ms4_dfs = []
pan_dfs = []
tiles = filesystem.listdir(tiles_folder)
for tile in tiles:
# this is needed because folder was copied from somewhere else
if not filesystem.exists(f'{tiles_folder}/{tile}'):
filesystem.makedirs(f'{tiles_folder}/{tile}')
metadata = filesystem.listdir(f'{tiles_folder}/{tile}')
metadata = [meta for meta in metadata if os.path.splitext(meta)[-1] == '.dim']
metadata_file_ms4 = metadata[0] if '_MS4_' in metadata[0] else metadata[1]
metadata_file_pan = metadata[0] if '_PAN_' in metadata[0] else metadata[1]
ms4_dfs.append(parse_deimos_metadata_file(f'{tiles_folder}/{tile}/{metadata_file_ms4}',
filesystem))
pan_dfs.append(parse_deimos_metadata_file(f'{tiles_folder}/{tile}/{metadata_file_pan}',
filesystem))
ms4_metadata = pd.concat(ms4_dfs)
pan_metadata = pd.concat(pan_dfs)
ms4_metadata.to_parquet(filesystem.openbin('metadata/deimos_ms4_metadata.pq', 'wb'))
pan_metadata.to_parquet(filesystem.openbin('metadata/deimos_pan_metadata.pq', 'wb'))
```
| github_jupyter |
# Import statements
```
from google.colab import drive
drive.mount('/content/drive')
from my_ml_lib import MetricTools, PlotTools
import os
import numpy as np
import matplotlib.pyplot as plt
import pickle
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import json
import datetime
import copy
from PIL import Image as im
import joblib
from sklearn.model_selection import train_test_split
# import math as Math
import random
import torch.optim
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
from torch.utils.data import DataLoader
from torch.utils.data import Dataset
import torchvision
import cv2
```
# Saving and Loading code
```
# Saving and Loading models using joblib
def save(filename, obj):
with open(filename, 'wb') as handle:
joblib.dump(obj, handle, protocol=pickle.HIGHEST_PROTOCOL)
def load(filename):
with open(filename, 'rb') as handle:
return joblib.load(filename)
```
# Importing Dataset
```
p = "/content/drive/MyDrive/A3/"
data_path = p + "dataset/train.pkl"
x = load(data_path)
# save_path = "/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/dataset/"
# # saving the images and labels array
# save(save_path + "data_image.pkl",data_image)
# save(save_path + "data_label.pkl",data_label)
# # dict values where labels key and image arrays as vlaues in form of list
# save(save_path + "my_dict.pkl",my_dict)
save_path = p + "dataset/"
# saving the images and labels array
data_image = load(save_path + "data_image.pkl")
data_label = load(save_path + "data_label.pkl")
# dict values where labels key and image arrays as vlaues in form of list
my_dict = load(save_path + "my_dict.pkl")
len(data_image) , len(data_label), my_dict.keys()
```
# Data Class and Data Loaders and Data transforms
```
len(x['names']) ,x['names'][4999] , data_image[0].shape
```
## Splitting the data into train and val
```
X_train, X_test, y_train, y_test = train_test_split(data_image, data_label, test_size=0.10, random_state=42,stratify=data_label )
len(X_train) , len(y_train) , len(X_test) ,len(y_test)
pd.DataFrame(y_test).value_counts()
```
## Data Class
```
class myDataClass(Dataset):
"""Custom dataset class"""
def __init__(self, images, labels , transform=None):
"""
Args:
images : Array of all the images
labels : Correspoing labels of all the images
"""
self.images = images
self.labels = labels
self.transform = transform
def __len__(self):
return len(self.images)
def __getitem__(self, idx):
# converts image value between 0 and 1 and returns a tensor C,H,W
img = torchvision.transforms.functional.to_tensor(self.images[idx])
target = self.labels[idx]
if self.transform:
img = self.transform(img)
return img,target
```
## Data Loaders
```
batch = 64
train_dataset = myDataClass(X_train, y_train)
test_dataset = myDataClass(X_test, y_test)
train_dataloader = DataLoader(train_dataset, batch_size= batch, shuffle=True)
test_dataloader = DataLoader(test_dataset, batch_size= batch, shuffle=True)
# next(iter(train_dataloader))[0].shape
len(train_dataloader) , len(test_dataloader)
```
# Train and Test functions
```
def load_best(all_models,model_test):
FILE = all_models[-1]
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model_test.parameters(), lr=0)
checkpoint = torch.load(FILE)
model_test.load_state_dict(checkpoint['model_state'])
optimizer.load_state_dict(checkpoint['optim_state'])
epoch = checkpoint['epoch']
model_test.eval()
return model_test
def train(save_path,epochs,train_dataloader,model,test_dataloader,optimizer,criterion,basic_name):
model_no = 1
c = 1
all_models = []
valid_loss_min = np.Inf
train_losses = []
val_losses = []
for e in range(epochs):
train_loss = 0.0
valid_loss = 0.0
model.train()
for idx, (images,labels) in enumerate(train_dataloader):
images, labels = images.to(device) , labels.to(device)
optimizer.zero_grad()
log_ps= model(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
train_loss += ((1 / (idx + 1)) * (loss.data - train_loss))
else:
accuracy = 0
correct = 0
model.eval()
with torch.no_grad():
for idx, (images,labels) in enumerate(test_dataloader):
images, labels = images.to(device) , labels.to(device)
log_ps = model(images)
_, predicted = torch.max(log_ps.data, 1)
loss = criterion(log_ps, labels)
# correct += (predicted == labels).sum().item()
equals = predicted == labels.view(*predicted.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor))
valid_loss += ((1 / (idx + 1)) * (loss.data - valid_loss))
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
e+1,
train_loss,
valid_loss
), "Test Accuracy: {:.3f}".format(accuracy/len(test_dataloader)))
train_losses.append(train_loss)
val_losses.append(valid_loss)
if valid_loss < valid_loss_min:
print('Saving model..' + str(model_no))
valid_loss_min = valid_loss
checkpoint = {
"epoch": e+1,
"model_state": model.state_dict(),
"optim_state": optimizer.state_dict(),
"train_losses": train_losses,
"test_losses": val_losses,
}
FILE = save_path + basic_name +"_epoch_" + str(e+1) + "_model_" + str(model_no)
all_models.append(FILE)
torch.save(checkpoint, FILE)
model_no = model_no + 1
save(save_path + basic_name + "_all_models.pkl", all_models)
return model, train_losses, val_losses, all_models
def plot(train_losses,val_losses,title='Training Validation Loss with CNN'):
plt.plot(train_losses, label='Training loss')
plt.plot(val_losses, label='Validation loss')
plt.xlabel('Iterations')
plt.ylabel('Loss')
plt.legend()
_ = plt.ylim()
plt.title(title)
# plt.savefig('plots/Training Validation Loss with CNN from scratch.png')
plt.show()
def test(loader, model, criterion, device, name):
test_loss = 0.
correct = 0.
total = 0.
y = None
y_hat = None
model.eval()
for batch_idx, (images, labels) in enumerate(loader):
# move to GPU or CPU
images, labels = images.to(device) , labels.to(device)
target = labels
# forward pass: compute predicted outputs by passing inputs to the model
output = model(images)
# calculate the loss
loss = criterion(output,labels)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
if y is None:
y = target.cpu().numpy()
y_hat = pred.data.cpu().view_as(target).numpy()
else:
y = np.append(y, target.cpu().numpy())
y_hat = np.append(y_hat, pred.data.cpu().view_as(target).numpy())
correct += np.sum(pred.view_as(labels).cpu().numpy() == labels.cpu().numpy())
total = total + images.size(0)
# if batch_idx % 20 == 0:
# print("done till batch" , batch_idx+1)
print(name + ' Loss: {:.6f}\n'.format(test_loss))
print(name + ' Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
return y, y_hat
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# def train(save_path,epochs,train_dataloader,model,test_dataloader,optimizer,criterion,basic_name)
# def plot(train_losses,val_losses,title='Training Validation Loss with CNN')
# def test(loader, model, criterion, device)
```
# Relu [ X=2 Y=3 Z=1 ]
## CNN-Block-123
### model
```
cfg3 = {
'B123': [16,16,'M','D',32,32,32,'M','D',64,'M','D'],
}
def make_layers3(cfg, batch_norm=True):
layers = []
in_channels = 3
for v in cfg:
if v == 'M':
layers += [nn.AvgPool2d(kernel_size=2, stride=2)]
elif v == 'M1':
layers += [nn.AvgPool2d(kernel_size=4, stride=3)]
elif v == 'D':
layers += [nn.Dropout(p=0.5)]
else:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3)
if batch_norm:
layers += [conv2d, nn.BatchNorm2d(v), nn.ReLU(inplace=True)]
else:
layers += [conv2d, nn.ReLU(inplace=True)]
in_channels = v
return nn.Sequential(*layers)
class Model_B123(nn.Module):
'''
Model
'''
def __init__(self, features):
super(Model_B123, self).__init__()
self.features = features
self.classifier = nn.Sequential(
# nn.Linear(1600, 512),
# nn.ReLU(True),
# nn.Linear(512, 256),
# nn.ReLU(True),
# nn.Linear(256, 64),
# nn.ReLU(True),
nn.Linear(64, 10),
)
def forward(self, x):
x = self.features(x)
# print(x.shape)
x = x.view(x.size(0), -1)
x = self.classifier(x)
return x
# m = Model_B123(make_layers3(cfg3['B123']))
# for i,l in train_dataloader:
# o = m(i)
model3 = Model_B123(make_layers3(cfg3['B123'])).to(device)
learning_rate = 0.001
criterion3 = nn.CrossEntropyLoss()
optimizer3 = optim.Adam(model3.parameters(), lr=learning_rate)
print(model3)
```
### train
```
# !rm '/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/models_saved_Q1/1_3/bw_blocks/Dropout(0.5)/cnn_block123/'*
# !ls '/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/models_saved_Q1/1_3/bw_blocks/Dropout(0.5)/cnn_block123/'
save_path3 = p + "models_saved_Q1/1_4/colab_notebooks /Batchnorm_and_pooling/models/"
m, train_losses, val_losses,m_all_models = train(save_path3,100,train_dataloader,model3,test_dataloader,optimizer3,criterion3,"cnn_b123_x2_y3_z1_with_BN_avg_pool_dp_without_WD_0.001")
```
### Tests and Plots
```
plot(train_losses,val_losses,'Training Validation Loss with CNN-block1')
all_models3 = load(save_path3 + "cnn_b123_x2_y3_z1_with_BN_avg_pool_dp_without_WD_0.001_all_models.pkl")
FILE = all_models3[-1]
m3 = Model_B123(make_layers3(cfg3['B123'])).to(device)
m3 = load_best(all_models3,m3)
train_y, train_y_hat = test(train_dataloader, m3, criterion3, device, "TRAIN")
cm = MetricTools.confusion_matrix(train_y, train_y_hat, nclasses=10)
PlotTools.confusion_matrix(cm, [i for i in range(10)], title='',
filename='Confusion Matrix with CNN', figsize=(6,6))
test_y, test_y_hat = test(test_dataloader, m3, criterion3, device,"TEST")
cm = MetricTools.confusion_matrix(test_y, test_y_hat, nclasses=10)
PlotTools.confusion_matrix(cm, [i for i in range(10)], title='',
filename='Confusion Matrix with CNN', figsize=(6,6))
```
| github_jupyter |
# Create Athena Database Schema
Amazon Athena is an interactive query service that makes it easy to analyze data in Amazon S3 using standard SQL. Athena is serverless, so there is no infrastructure to manage, and you pay only for the queries that you run.
Athena is based on Presto, and supports various standard data formats, including CSV, JSON, Avro or columnar data formats such as Apache Parquet and Apache ORC.
Presto is an open source, distributed SQL query engine, developed for fast analytic queries against data of any size. It can query data where it is stored, without the need to move the data. Query execution runs in parallel over a pure memory-based architecture which makes Presto extremely fast.
<img src="img/athena_setup.png" width="60%" align="left">
```
import boto3
import sagemaker
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
ingest_create_athena_db_passed = False
%store -r s3_public_path_tsv
try:
s3_public_path_tsv
except NameError:
print("*****************************************************************************")
print("[ERROR] PLEASE RE-RUN THE PREVIOUS COPY TSV TO S3 NOTEBOOK ******************")
print("[ERROR] THIS NOTEBOOK WILL NOT RUN PROPERLY. ********************************")
print("*****************************************************************************")
print(s3_public_path_tsv)
%store -r s3_private_path_tsv
try:
s3_private_path_tsv
except NameError:
print("*****************************************************************************")
print("[ERROR] PLEASE RE-RUN THE PREVIOUS COPY TSV TO S3 NOTEBOOK ******************")
print("[ERROR] THIS NOTEBOOK WILL NOT RUN PROPERLY. ********************************")
print("*****************************************************************************")
print(s3_private_path_tsv)
```
# Import PyAthena
[PyAthena](https://pypi.org/project/PyAthena/) is a Python DB API 2.0 (PEP 249) compliant client for Amazon Athena.
```
from pyathena import connect
```
# Create Athena Database
```
database_name = "dsoaws"
```
Note: The databases and tables that we create in Athena use a data catalog service to store the metadata of your data. For example, schema information consisting of the column names and data type of each column in a table, together with the table name, is saved as metadata information in a data catalog.
Athena natively supports the AWS Glue Data Catalog service. When we run `CREATE DATABASE` and `CREATE TABLE` queries in Athena with the AWS Glue Data Catalog as our source, we automatically see the database and table metadata entries being created in the AWS Glue Data Catalog.
```
# Set S3 staging directory -- this is a temporary directory used for Athena queries
s3_staging_dir = "s3://{0}/athena/staging".format(bucket)
conn = connect(region_name=region, s3_staging_dir=s3_staging_dir)
statement = "CREATE DATABASE IF NOT EXISTS {}".format(database_name)
print(statement)
import pandas as pd
pd.read_sql(statement, conn)
```
# Verify The Database Has Been Created Succesfully
```
statement = "SHOW DATABASES"
df_show = pd.read_sql(statement, conn)
df_show.head(5)
if database_name in df_show.values:
ingest_create_athena_db_passed = True
%store ingest_create_athena_db_passed
```
# Store Variables for the Next Notebooks
```
%store
```
# Release Resources
```
%%html
<p><b>Shutting down your kernel for this notebook to release resources.</b></p>
<button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
<script>
try {
els = document.getElementsByClassName("sm-command-button");
els[0].click();
}
catch(err) {
// NoOp
}
</script>
%%javascript
try {
Jupyter.notebook.save_checkpoint();
Jupyter.notebook.session.delete();
}
catch(err) {
// NoOp
}
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn.manifold import MDS, TSNE
from deslib.util.diversity import double_fault
import matplotlib.lines as mlines
from umap import UMAP
```
# Get the pairwise diversity matrix
```
def compute_pairwise_diversity_matrix(targets, prediction_matrix, diversity_func):
"""Computes the pairwise diversity matrix.
Parameters
----------
targets : array of shape (n_samples):
Class labels of each sample in X.
prediction_matrix : array of shape (n_samples, n_classifiers):
Predicted class labels for each classifier in the pool
diversity_func : Function
Function used to estimate the pairwise diversity
Returns
-------
diversity : array of shape = [n_classifiers]
The average pairwise diversity matrix calculated for the pool of
classifiers
"""
n_classifiers = prediction_matrix.shape[1]
diversity = np.zeros((n_classifiers, n_classifiers))
for clf_index in range(n_classifiers):
for clf_index2 in range(clf_index + 1, n_classifiers):
this_diversity = diversity_func(targets,
prediction_matrix[:, clf_index],
prediction_matrix[:, clf_index2])
diversity[clf_index, clf_index2] = this_diversity
diversity[clf_index2, clf_index] = this_diversity
return diversity
def load_predictions(dataset_name):
path = './Saved Predict and Proba/' + dataset_name.upper() + '/pred_val_' + dataset_name.lower() + '.csv'
table_pred = pd.read_csv(path)
if dataset_name == 'zw':
label = table_pred['norm']
methods = table_pred.drop('norm', axis=1)
else:
label = table_pred['class']
methods = table_pred.drop('class', axis=1)
methods = methods.drop('Unnamed: 0', axis=1)
return label, methods
def compute_matrix_embedding(dataset_name, n_classifiers, n_features, method='mds', n_neighbors=5, min_dist=0.7
):
D = compute_pairwise_diversity_matrix(label.to_numpy(), methods.to_numpy(), double_fault)
D = 1/D
D[D==np.inf] = 0
if method == 'mds':
method = MDS(dissimilarity='precomputed', random_state=123456987, n_init=20, max_iter=100000)
elif method == 'tsne':
method = TSNE(perplexity=25, init='pca', random_state=42, early_exaggeration=50,
learning_rate=200, n_iter=2500, angle=0.5)
else:
method = UMAP(n_neighbors=2, metric='euclidean', random_state=42, min_dist=1, n_components=2,)
D_tilde = method.fit_transform(D)
D_tilde = D_tilde.reshape(n_classifiers, n_features, 2)
return D_tilde
```
# Plotting
```
from matplotlib import pyplot as plt
def plot_diversity(D_changed, method_tmp, dataset_name, title):
s = 100
colors = {}
colors[0] = '#DAA520'
colors[1] = '#FF0000'
colors[2] = '#0000FF'
colors[3] = '#228B22'
colors[4] = '#000000'
markers = {}
markers[0] = 'X'
markers[1] = 'd'
markers[2] = '*'
markers[3] = "^"
markers[4] = 'o'
n_classifiers, n_features, _ = D_changed.shape
plt.figure(figsize=(15,10))
method=0
for idx in range(n_classifiers):
for idx2 in range(n_features):
x, y = D_changed[idx,idx2, 0], D_changed[idx,idx2, 1]
plt.scatter(x, y, color=colors[idx2], s=s, lw=0, marker=markers[idx2])
plt.annotate(method_tmp[method], xy=(x, y), textcoords='offset points', xytext=(5, 15), ha='right', va='top')
method += 1
m1 = mlines.Line2D([], [], color=colors[0], marker=markers[0], linestyle='None', markersize=10, label='CV')
m2 = mlines.Line2D([], [], color=colors[1], marker=markers[1], linestyle='None', markersize=10, label='TFIDF')
m3 = mlines.Line2D([], [], color=colors[2], marker=markers[2], linestyle='None', markersize=10, label='Glove')
m4 = mlines.Line2D([], [], color=colors[3], marker=markers[3], linestyle='None', markersize=10, label='Word2Vec')
m5 = mlines.Line2D([], [], color=colors[4], marker=markers[4], linestyle='None', markersize=10, label='FastText')
plt.title(title + " " + dataset_name.upper()+' dataset')
plt.legend(handles=[m1, m2, m3, m4, m5])
plt.tight_layout()
plt.savefig(title + " " + dataset_name.upper()+' dataset.pdf', dpi=450)
n_classifiers = 8
method = 'umap'
n_features = 5
for dataset_name in ['td', 'zw', 'td_zw']:
label, methods = load_predictions(dataset_name)
methods_name = [name.split('-', 1)[0] for name in methods.columns]
D_tilde = compute_matrix_embedding(dataset_name, n_classifiers, n_features, method)
plot_diversity(D_tilde, methods_name, dataset_name, method)
```
| github_jupyter |
# Viral Transmission Model
```
import pandas as pd
import numpy as np
import pylab as plt
import datetime
import scipy.optimize as spo
import scipy.integrate as spi
```
## Data Preprocessing
The dataset we will be using is the Global Coronavirus (COVID-19) Data (Corona Data Scraper) provided by Enigma.
[AWS product link](https://aws.amazon.com/marketplace/pp/prodview-vtnf3vvvheqzw?qid=1597409751562&sr=0-1&ref_=brs_res_product_title#overview)
[Corona Data Scraper page](https://coronadatascraper.com/#home)
We are only interested in the state-level data in United States. To save time from opening a super large dataset, we save each state's data into small files.
```
df = pd.read_csv("datasets/timeseries.csv")
df_US = df[(df["country"]=="United States") & (df["level"]=="state")]
df_US = df_US[["state", "population", "cases", "deaths", "recovered", "tested", "hospitalized", "date"]]
states = np.unique(df_US["state"])
for state in states:
df_US[df_US["state"]==state].to_csv("datasets/timeseries_states/"+state+".csv", index=0)
```
## State Data Snapshot
```
def chooseState(state, output=True):
df_state = pd.read_csv("datasets/timeseries_states/"+state+".csv")
### data cleaning
# some col is missing in some state's report
na_cols = []
for col in df_state.columns.values:
if df_state[col].isna().all():
na_cols.append(col)
for col in na_cols:
df_state = df_state.drop(col, axis=1)
# some data is missing at the beginning of the outbreak
suggest_startdate = df_state.iloc[0]["date"]
for i in range(len(df_state.index)):
if df_state.iloc[i].notna().all():
suggest_startdate = df_state.iloc[i]["date"]
break
# regulate the data
df_state = df_state.fillna(0)
df_state["date"] = pd.to_datetime(df_state["date"])
if output:
### snapshot and mark data inconsistency
plt.figure()
plt.title(state)
for line, style in [ ("cases", "-b"), ("deaths", "-k"), ("recovered", "-g")]:
if line in df_state.columns.values:
plt.plot(df_state[line], style, label=line)
for i in range(1, len(df_state.index)):
if abs(df_state[line][i] - df_state[line][i-1]) > (max(df_state[line]) - min(df_state[line])) / 6:
plt.plot(i, df_state[line][i], "xr")
plt.legend()
### print out data-cleaning message
if na_cols == []:
print("Data complete, ready to analyze.")
else:
print("Data incomplete, cannot analyze.")
print("NA cols: ", na_cols)
print("Suggest choosing start date after", suggest_startdate)
### discard the outliers
for line in [ "cases", "deaths", "recovered", "hospitalized"]:
if line in df_state.columns.values:
col_idx = list(df_state.columns).index(line)
for i in range(1, len(df_state.index)-1):
if (df_state.iloc[i, col_idx] - df_state.iloc[i-1, col_idx]) * (df_state.iloc[i+1, col_idx] - df_state.iloc[i, col_idx]) < -((max(df_state[line]) - min(df_state[line])) /4)**2:
df_state.iloc[i, col_idx] = (df_state.iloc[i-1, col_idx] + df_state.iloc[i+1, col_idx]) / 2
if output:
plt.plot(i, df_state[line][i], "or")
return df_state, na_cols, suggest_startdate
```
For example, we are interested in New York state.
```
state = "New York"
df_state, _, _ = chooseState(state)
df_state.sample(5).sort_values("date")
```
For example, we are interested in the first two weeks in April.
```
def chooseTime(df_state, start_date, end_date):
start = np.datetime64(start_date)
end = np.datetime64(end_date)
# The time period of interest
sample = df_state[ (df_state["date"] >= start) & (df_state["date"] <= end) ]
# The future period to determine "exposed"
sample_future = df_state[ (df_state["date"] >= start + np.timedelta64(14,'D')) & (df_state["date"] <= end + np.timedelta64(14,'D')) ]
return sample, sample_future
start_date, end_date = "2020-04-01", "2020-04-14"
sample, sample_future = chooseTime(df_state, start_date, end_date)
```
## SEIR Infection Model

We can use an SEIR model to describe the transmission dynamics of Covid19 as above.
We Assume...
- Susceptible (S): healthy people, will be infected and turn into E after close contact with E or Q.
- Exposed (E): infected but have no symptoms yet, infectious with a rate of $\lambda$. E will turn into I after the virus incubation period, which is 14 days on average. So we assume $\sigma = 1/14$, dE/dt (t) = dI/dt (t+14).
- Infectious (I): infected and have symptoms. We will take the data of test_positive or cases_reported as the data of I. The severe cases will be hospitalized (H), the mild cases will be in self quarantine (Q). I may recover or die after some time.
- Self Quarantine (Q): have symptoms, may still have some contact with others, thus infectious with a different rate of $c\lambda$ ($0 \le c \le 1$). We also assume $Q = kI$, where $k = 1 - avg(\frac{\Delta hospitalized}{\Delta test\_pos}) $
- Hospitalized (H): have symptoms, kept in hospitals, assume no contact with S.
- Recovered (R): recovered and immune, may turn into S again (immunity lost or virus not cleared)
- Dead (X): dead unfortunately :(
Therefore, we have a set of differential equations to describe this process:
$\begin{aligned}
&\frac{dS}{dt}&
&=& - \lambda \frac{S}{N} E - c\lambda \frac{S}{N} Q + \alpha R ~~~
&=& - \lambda \frac{S}{N} E - c\lambda \frac{S}{N} kI + \alpha R
\\
&\frac{dE}{dt}&
&=& \lambda \frac{S}{N} E + c\lambda \frac{S}{N} Q - \sigma E ~~~
&=& \lambda \frac{S}{N} E + c\lambda \frac{S}{N} kI - \sigma E
\\
&\frac{dI}{dt}&
&=& \sigma E - \mu I - \omega I
\\
&\frac{dX}{dt}&
&=& \omega I
\\
&\frac{dR}{dt}&
&=& \mu I - \alpha R
\end{aligned}$
$S + E + I + R + X = N,~ I = Q + H$
Apply to our datasets, we have:
$ R = recovered,~ X = deaths,~ I = test\_pos - deaths - recovered,\\
E(t) = I(t+14) - I(t),~ S = N - E - I - R - X,\\
k = 1 - avg(\frac{\Delta hospitalized}{\Delta test\_pos})
$
```
### run SEIR model on sample data
def SEIR(sample, sample_future, output=True):
### differential equations for spi.odeint, INP - initial point, t - time range
# dS/dt = - lamda*S/N*E - c*lamda*S/N*k*I + alpha*R
# dE/dt = lamda*S/N*E + c*lamda*S/N*k*I - sigma*E
# dI/dt = sigma*E - miu*I - omega*I
# dX/dt = omega*I
# dR/dt = miu*I - alpha*R
def diff_eqs(INP, t, lamda_p, c_p, alpha_p, omega_p, miu_p):
Y = np.zeros((5))
V = INP
Y[0] = - lamda_p*V[0]/N*V[1] - c_p*lamda_p*V[0]/N*k*V[2] + alpha_p*V[4]
Y[1] = lamda_p*V[0]/N*V[1] + c_p*lamda_p*V[0]/N*k*V[2] - sigma*V[1]
Y[2] = sigma*V[1] - miu_p*V[2] - omega_p*V[2]
Y[3] = omega_p*V[2]
Y[4] = miu_p*V[2] - alpha_p*V[4]
return Y
### cost function for optimization
def MSE(params):
INP = (S[0], E[0], I[0], X[0], R[0])
t_range = np.arange(0, len(S), 1)
RES = spi.odeint(diff_eqs, INP, t_range, args=tuple(params))
mse = 0
for i in range(len(S)):
mse += ( (RES[i,0] - S[i]) ) **2
mse += ( (RES[i,1] - E[i]) ) **2
mse += ( (RES[i,2] - I[i]) ) **2
mse += ( (RES[i,3] - X[i]) ) **2
mse += ( (RES[i,4] - R[i]) ) **2
mse = mse / len(S)
return mse
### get necessary data from dataset
cases = np.array(list(sample["cases"])) # test_positive
cases_future = np.array(list(sample_future["cases"])) # to calculate exposed
hospitalized = np.array(list(sample["hospitalized"])) # to calculate k
deaths = np.array(list(sample["deaths"])) # X
recovered = np.array(list(sample["recovered"])) # R
N = np.mean(df_state["population"])
X = deaths
R = recovered
I = cases - deaths - recovered
E = cases_future - cases
S = N - E - I - X - R
dS = S[1:] - S[:-1]
dE = E[1:] - E[:-1]
dI = I[1:] - I[:-1]
dX = X[1:] - X[:-1]
dR = R[1:] - R[:-1]
S = S[:-1]
E = E[:-1]
I = I[:-1]
X = X[:-1]
R = R[:-1]
### guess params
# By experience: k, sigma
k = 1 - np.mean( (hospitalized[1:]-hospitalized[0:-1] +1e-5) / (cases[1:]-cases[:-1] +1e-5) ) # k = deltaH / deltaCases
sigma = 1/14 # virus incubation period = 14 days
# From optimization: lamda, c, alpha, omega, miu
alpha0 = 0.006
omega0 = np.mean((dX+1e-5) / (I+1e-5)) # dx/dt = omega*i
c0 = 0.4
lamda0 = np.mean(- (dS - alpha0*R +1e-5) / (S/N * (E+c0*k*I) +1e-5) ) # dS/dt = - lamda*S/N*(E+ckI) + alpha*R
miu0 = np.mean((dR + alpha0*R +1e-5) / (I+1e-5)) # dr/dt = miu*i - alpha*r
### Optimization to find best params
params0 = (lamda0, c0, alpha0, omega0, miu0) # lamda, c, alpha, omega, miu
ret = spo.minimize(MSE, params0, bounds=[(0,1), (0,1), (0,1), (0,1), (0,1)])
params = ret.x
params0 = [round(i,8) for i in params0]
params = [round(i,8) for i in params]
k = round(k,8)
sigma = round(sigma,8)
if output:
print("Estimated k: ", k, "; sigma: ", sigma)
print("Optimization for lamda, c, alpha, omega, miu")
print("params0: ", params0)
print("params: ", params)
### solve ode and plot
INP = (S[0], E[0], I[0], X[0], R[0])
t_range = np.arange(0, len(S)*5, 1)
RES = spi.odeint(diff_eqs, INP, t_range, args=tuple(params))
plt.figure(figsize=(10,6))
plt.plot(RES[:,1], '-b', label='E_predict')
plt.plot(RES[:,2], '-r', label='I_predict')
plt.plot(RES[:,3], '-k', label='X_predict')
plt.plot(RES[:,4], '-g', label='R_predict' )
plt.plot(E, "xb", label="E_real")
plt.plot(I, "xr", label="I_real")
plt.plot(X, "xk", label="X_real")
plt.plot(R, "xg", label="R_real")
plt.grid()
plt.xlabel('Time', fontsize = 12)
plt.ylabel('number of people', fontsize = 12)
plt.legend(loc=0)
plt.title(state + ": " + str(start_date) + " - " + str(end_date), fontsize = 14)
plt.show();
return params, k, sigma
params = SEIR(sample, sample_future)
```
### Example1: California
```
state = "New York"
df_state, _, _ = chooseState(state)
# df_state.sample(5).sort_values("date")
```
Everything seems fine (except that jump), then we choose an appropriate time period to analyze
```
start_date, end_date = "2020-04-15", "2020-04-28"
sample, sample_future = chooseTime(df_state, start_date, end_date)
params = SEIR(sample, sample_future)
```
### Example2: New York
```
state = "New York"
df_state, _, _ = chooseState(state)
# df_state.sample(5).sort_values("date")
```
Everything seems fine, then we choose an appropriate time period to analyze.
```
start_date, end_date = "2020-05-01", "2020-05-14"
sample, sample_future = chooseTime(df_state, start_date, end_date)
params = SEIR(sample, sample_future)
```
### Example3: Illinois
```
state = "Illinois"
df_state, _, _ = chooseState(state)
# df_state.sample(5).sort_values("date")
```
There are too many missing data (no recovered) for Illinois, we can not analyze now.
### Example4: Texas
```
state = "Texas"
df_state, _, _ = chooseState(state)
# df_state.sample(5).sort_values("date")
```
The hospitalized data is missing, we may use the average k instead. But we can not analyze now.
### Generate state-time-params data
Now that our SEIR infection model is working, we choose some states of interest and compute the corresponding optimal parameters for further modeling.
```
StateOfInterest = ["Arizona", "California", "Minnesota", "New Mexico", "New York",
"Oklahoma", "South Carolina", "Tennessee", "Utah", "Virginia",
"West Virginia", "Wisconsin"]
df_SOI = pd.DataFrame(columns = ["state", "startdate", "enddate", "k", "sigma", "lamda", "c", "alpha", "omega", "miu"])
for state in StateOfInterest:
df_state, _, suggest_startdate = chooseState(state, False)
# choose 15 days as a period
if np.datetime64(suggest_startdate,'D') - np.datetime64(suggest_startdate,'M') <= np.timedelta64(14,'D'):
real_startdate = np.datetime64(suggest_startdate,'M') + np.timedelta64(15,'D')
else:
real_startdate = np.datetime64(np.datetime64(suggest_startdate,'M') + np.timedelta64(1,'M'), 'D')
stopdate = np.datetime64("2020-07-01")
startdate = real_startdate
while True:
if startdate > stopdate:
break
enddate = startdate + np.timedelta64(14,'D')
sample, sample_future = chooseTime(df_state, startdate, enddate)
params = SEIR(sample, sample_future, False)
df_SOI = df_SOI.append([{"state":state, "startdate":startdate, "enddate":enddate, "k":params[1], "sigma":params[2], "lamda":params[0][0], "c":params[0][1], "alpha":params[0][2], "omega":params[0][3], "miu":params[0][4]}], ignore_index=True)
if np.datetime64(startdate,'D') - np.datetime64(startdate,'M') <= np.timedelta64(0,'D'):
startdate = np.datetime64(startdate,'M') + np.timedelta64(15,'D')
else:
startdate = np.datetime64(np.datetime64(startdate,'M') + np.timedelta64(1,'M'), 'D')
df_SOI.to_csv("datasets/model_out.csv", index=0)
df_SOI.sample(5)
```
| github_jupyter |
<table width="100%">
<tr style="border-bottom:solid 2pt #009EE3">
<td style="text-align:left" width="10%">
<a href="open_signals_after_acquisition.dwipynb" download><img src="../../images/icons/download.png"></a>
</td>
<td style="text-align:left" width="10%">
<a href="https://mybinder.org/v2/gh/biosignalsnotebooks/biosignalsnotebooks/biosignalsnotebooks_binder?filepath=biosignalsnotebooks_environment%2Fcategories%2FLoad%2Fopen_signals_after_acquisition.dwipynb" target="_blank"><img src="../../images/icons/program.png" title="Be creative and test your solutions !"></a>
</td>
<td></td>
<td style="text-align:left" width="5%">
<a href="../MainFiles/biosignalsnotebooks.ipynb"><img src="../../images/icons/home.png"></a>
</td>
<td style="text-align:left" width="5%">
<a href="../MainFiles/contacts.ipynb"><img src="../../images/icons/contacts.png"></a>
</td>
<td style="text-align:left" width="5%">
<a href="https://github.com/biosignalsnotebooks/biosignalsnotebooks" target="_blank"><img src="../../images/icons/github.png"></a>
</td>
<td style="border-left:solid 2pt #009EE3" width="15%">
<img src="../../images/ost_logo.png">
</td>
</tr>
</table>
<link rel="stylesheet" href="../../styles/theme_style.css">
<!--link rel="stylesheet" href="../../styles/header_style.css"-->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/font-awesome/4.7.0/css/font-awesome.min.css">
<table width="100%">
<tr>
<td id="image_td" width="15%" class="header_image_color_1"><div id="image_img"
class="header_image_1"></div></td>
<td class="header_text"> Load Signals after Acquisition [OpenSignals] </td>
</tr>
</table>
<div id="flex-container">
<div id="diff_level" class="flex-item">
<strong>Difficulty Level:</strong> <span class="fa fa-star checked"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
</div>
<div id="tag" class="flex-item-tag">
<span id="tag_list">
<table id="tag_list_table">
<tr>
<td class="shield_left">Tags</td>
<td class="shield_right" id="tags">load☁open☁opensignals☁post-acquisition</td>
</tr>
</table>
</span>
<!-- [OR] Visit https://img.shields.io in order to create a tag badge-->
</div>
</div>
<a href="https://www.biosignalsplux.com/en/software" target="_blank"><span class="color1"><strong>OpenSignals <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a> is the Plux's software dedicated to acquire, store and process physiological signals acquired with electronic devices developed and commercialized by the company (such as <a href="https://www.biosignalsplux.com/en/" target="_blank"><span class="color2"><strong>biosignalsplux <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a> and <a href="https://bitalino.com/en/" target="_blank"><span class="color4"><strong>bitalino <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a>).
Two brief tutorials about "signal recording" and "file storage" with <span class="color1"><strong>OpenSignals</strong></span> are available in the notebooks <strong><a href="../Record/record_data.ipynb">Signal Acquisition [OpenSignals] <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></strong> and <strong><a href="../Record/store_signals_after_acquisition.ipynb">Store Files after Acquisition [OpenSignals] <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></strong>.
In the current <span class="color4"><strong>Jupyter Notebook</strong></span> we continue the interaction with <span class="color1"><strong>OpenSignals</strong></span>, demonstrating how the previously acquired/stored files can be loaded.
<hr>
<p class="steps">1 - Execute <span class="color1"><strong>OpenSignals</strong></span> application in order to get access to the main page</p>
<img src="../../images/open/load_sig_opensignals_main.png">
<p class="steps">2 - Press the <strong>"Play"</strong> icon for entering in the offline visualization environment</p>
<img src="../../images/open/load_sig_opensignals_play.gif">
<i>In normal conditions there is a file in memory (the last acquisition made or the last file opened in the offline visualization environment). As you can see, our file in memory is a respiratory signal</i>
<p class="steps">3 - However we may want to load a older file. For achieving this purpose, in the visualization environment window click on the <strong>"Folder"</strong> icon near the bottom-right corner of the screen</p>
<i>This action will cause the opening of a file exploration window ! </i>
<img src="../../images/open/load_sig_opensignals_file.gif">
<i>Now, you should navigate through the directories until reaching the file you want to open</i>
<p class="steps">4 - Select the file and press "OK" button</p>
<img src="../../images/open/load_sig_opensignals_signal.gif">
Finally, we load the desired ECG signal !
<img src="../../images/open/load_sig_opensignals_vis_env_signal.png">
Like you saw, opening the acquired signals is quick and easy with <span class="color1"><strong>OpenSignals</strong></span> !
<strong><span class="color7">We hope that you have enjoyed this guide. </span><span class="color2">biosignalsnotebooks</span><span class="color4"> is an environment in continuous expansion, so don't stop your journey and learn more with the remaining <a href="../MainFiles/biosignalsnotebooks.ipynb">Notebooks <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></span></strong> !
<hr>
<table width="100%">
<tr>
<td style="border-right:solid 3px #009EE3" width="20%">
<img src="../../images/ost_logo.png">
</td>
<td width="40%" style="text-align:left">
<a href="../MainFiles/aux_files/biosignalsnotebooks_presentation.pdf" target="_blank">☌ Project Presentation</a>
<br>
<a href="https://github.com/biosignalsnotebooks/biosignalsnotebooks" target="_blank">☌ GitHub Repository</a>
<br>
<a href="https://pypi.org/project/biosignalsnotebooks/" target="_blank">☌ How to install biosignalsnotebooks Python package ?</a>
<br>
<a href="../MainFiles/signal_samples.ipynb">☌ Signal Library</a>
</td>
<td width="40%" style="text-align:left">
<a href="../MainFiles/biosignalsnotebooks.ipynb">☌ Notebook Categories</a>
<br>
<a href="../MainFiles/by_diff.ipynb">☌ Notebooks by Difficulty</a>
<br>
<a href="../MainFiles/by_signal_type.ipynb">☌ Notebooks by Signal Type</a>
<br>
<a href="../MainFiles/by_tag.ipynb">☌ Notebooks by Tag</a>
</td>
</tr>
</table>
```
from biosignalsnotebooks.__notebook_support__ import css_style_apply
css_style_apply()
%%html
<script>
// AUTORUN ALL CELLS ON NOTEBOOK-LOAD!
require(
['base/js/namespace', 'jquery'],
function(jupyter, $) {
$(jupyter.events).on("kernel_ready.Kernel", function () {
console.log("Auto-running all cells-below...");
jupyter.actions.call('jupyter-notebook:run-all-cells-below');
jupyter.actions.call('jupyter-notebook:save-notebook');
});
}
);
</script>
```
| github_jupyter |
# Neural Networks
[Neural networks](https://en.wikipedia.org/wiki/Artificial_neural_network) are models inspired by biological brains for learning arbitrary functions.
# Function Approximation
To start with, let's train a neural network to learn a simple mathematical function such as `z = sin(x) + 2*y`:
```
import numpy as np
from sklearn import preprocessing
from sklearn.cross_validation import train_test_split
np.random.seed(0)
# Number of samples
n = 100
# Choose n random numbers for x and y
x = np.random.rand(n)
y = np.random.rand(n)
# Create an array of [x,y] scaled:
# We scale the data because neural networks perform better when all inputs are
# in a similar value range
data = preprocessing.scale(np.stack([x,y], axis=1))
# Create z. We reshape it to an array of 1-element arrays for pyBrain
target = (np.sin(x) + 2*y).reshape(n,1)
# Create train/test split
data_train, data_test, target_train, target_test = train_test_split(
data, target, test_size=.25, random_state=1
)
```
Now let's create a neural network:
```
from pybrain.tools.shortcuts import buildNetwork
from pybrain.structure import TanhLayer
network = buildNetwork(2, 5, 1, hiddenclass=TanhLayer)
```
The above creates a network with 2 inputs (x and y) has 5 hidden units and 1 output (z).
Now we need to create datasets that PyBrain can use.
```
from pybrain.datasets.classification import SupervisedDataSet
# Create a dataset with 2 inputs and 1 output
ds_train = SupervisedDataSet(2,1)
# add our data to the dataset
ds_train.setField('input', data_train)
ds_train.setField('target', target_train)
# Do the same for the test set
ds_test = SupervisedDataSet(2,1)
ds_test.setField('input', data_test)
ds_test.setField('target', target_test)
```
Finally, let's train our network and report it's accuracy:
```
from pybrain.supervised.trainers import BackpropTrainer
from pybrain.tools.validation import Validator
# Create a trainer for the network and training dataset
trainer = BackpropTrainer(network, ds_train)
# Train for a number of epochs and report accuracy:
for i in range(10):
# Train 10 epochs
trainer.trainEpochs(10)
# Report mean squared error for training and testing sets
# `network.activateOnDataset` will return the predicted values for each input in the dataset passed to it.
# Then `Validator.MSE` returns the mean squared error of the returned value with the actual value.
print("Train MSE:", Validator.MSE(network.activateOnDataset(ds_train), target_train))
print("Test MSE:", Validator.MSE(network.activateOnDataset(ds_test), target_test))
```
# Classification with Neural Networks
Neural networks can also be used for classification.
# Digits Sample Data
We'll be using the same dataset we used for the [classification notebook](classification.ipynb) - a dataset of 8x8 pixel handwritten digits such as the following:

The data is in the form of a 64 element array of integers representing grayscale values for each pixel.
```
from sklearn.datasets import load_digits
# Load all the samples for all digits 0-9
digits = load_digits()
# Assign the matrices to a variable `data`
data = digits.data
# Assign the labels to a variable `target`
target = digits.target.reshape((len(digits.target), 1))
# Split the data into 75% train, 25% test
data_train, data_test, target_train, target_test = train_test_split(
data, target, test_size=.25, random_state=0
)
```
Now let's build our network again. This time we have 64 inputs and 10 outputs (1 output for each digit). We can also create multiple hidden layers. For example two layers of eight hidden units each:
```
from pybrain.structure import SoftmaxLayer
# Create a network with 64 inputs, 2 layers of 16 hidden units and 10 outputs (for each digit 0-9)
network = buildNetwork(data.shape[1], 16, 16, 10, hiddenclass=SoftmaxLayer)
```
This time we'll use a dataset specifically for classification. And we will create dummy variables for each of the labels instead of using the numerical value directly.
```
from pybrain.datasets.classification import ClassificationDataSet
# Create a dataset with 64 inputs
ds_train = ClassificationDataSet(data_train.shape[1])
# Set the input data
ds_train.setField('input', data_train)
ds_train.setField('target', target_train)
# The convert to dummy variables
# For instance, this will represent 0 as (1,0,0,0,0,0,0,0,0,0)
# 1 as (0,1,0,0,0,0,0,0,0,0), 4 as (0,0,0,0,1,0,0,0,0,0) and so on.
ds_train._convertToOneOfMany()
# Do the same for the test set
ds_test = ClassificationDataSet(data_test.shape[1])
ds_test.setField('input', data_test)
ds_test.setField('target', target_test)
ds_test._convertToOneOfMany()
```
Now we can train as we did before. We'll need to train this network for longer because the model is more complicated as is the function.
```
from sklearn.metrics import accuracy_score
trainer = BackpropTrainer(network, ds_train)
for i in range(10):
trainer.trainEpochs(50)
# We use `argmax(1)` to convert the results back from the 10 outputs to a single label
print("Training Accuracy:", accuracy_score(target_train, network.activateOnDataset(ds_train).argmax(1)))
print("Testing Accuracy:", accuracy_score(target_test, network.activateOnDataset(ds_test).argmax(1)))
```
# Further Reading
* [PyBrain Official Documentation](http://pybrain.org/docs/index.html)
| github_jupyter |
IN:GET_LOCATION is the second most frequent intent. Split data such that 80% of this intent will be in a different subset.
```
import pandas as pd
data = pd.read_table('../data/top-dataset-semantic-parsing/train.tsv', names=['text', 'tokens', 'schema'])
data
get_loc_mask = ['IN:GET_LOCATION' in text for text in data.schema]
n_get_loc = sum(get_loc_mask)
print(n_get_loc)
n_leave = int(n_get_loc * 0.2)
n_take = n_get_loc - n_leave
print('Take : ', n_take)
print('Leave: ', n_leave)
get_loc_mask_ids = [i for i, m in enumerate(get_loc_mask) if m]
get_loc_mask_ids[:10]
from random import shuffle
shuffle(get_loc_mask_ids)
get_loc_mask_ids[:10]
small_subset_ids = get_loc_mask_ids[:n_take]
small_subset = data.iloc[small_subset_ids]
big_subset_ids = listsub(set(range(len(data))) - set(small_subset_ids))
big_subset = data.iloc[big_subset_ids]
all_schema_tokens = {'IN:COMBINE',
'IN:GET_CONTACT',
'IN:GET_DIRECTIONS',
'IN:GET_DISTANCE',
'IN:GET_ESTIMATED_ARRIVAL',
'IN:GET_ESTIMATED_DEPARTURE',
'IN:GET_ESTIMATED_DURATION',
'IN:GET_EVENT',
'IN:GET_EVENT_ATTENDEE',
'IN:GET_EVENT_ATTENDEE_AMOUNT',
'IN:GET_EVENT_ORGANIZER',
'IN:GET_INFO_ROAD_CONDITION',
'IN:GET_INFO_ROUTE',
'IN:GET_INFO_TRAFFIC',
'IN:GET_LOCATION',
'IN:GET_LOCATION_HOME',
'IN:GET_LOCATION_HOMETOWN',
'IN:GET_LOCATION_SCHOOL',
'IN:GET_LOCATION_WORK',
'IN:NEGATION',
'IN:UNINTELLIGIBLE',
'IN:UNSUPPORTED',
'IN:UNSUPPORTED_EVENT',
'IN:UNSUPPORTED_NAVIGATION',
'IN:UPDATE_DIRECTIONS',
'SL:AMOUNT',
'SL:ATTENDEE_EVENT',
'SL:ATTRIBUTE_EVENT',
'SL:CATEGORY_EVENT',
'SL:CATEGORY_LOCATION',
'SL:COMBINE',
'SL:CONTACT',
'SL:CONTACT_RELATED',
'SL:DATE_TIME',
'SL:DATE_TIME_ARRIVAL',
'SL:DATE_TIME_DEPARTURE',
'SL:DESTINATION',
'SL:GROUP',
'SL:LOCATION',
'SL:LOCATION_CURRENT',
'SL:LOCATION_MODIFIER',
'SL:LOCATION_USER',
'SL:LOCATION_WORK',
'SL:METHOD_TRAVEL',
'SL:NAME_EVENT',
'SL:OBSTRUCTION',
'SL:OBSTRUCTION_AVOID',
'SL:ORDINAL',
'SL:ORGANIZER_EVENT',
'SL:PATH',
'SL:PATH_AVOID',
'SL:POINT_ON_MAP',
'SL:ROAD_CONDITION',
'SL:ROAD_CONDITION_AVOID',
'SL:SEARCH_RADIUS',
'SL:SOURCE',
'SL:TYPE_RELATION',
'SL:UNIT_DISTANCE',
'SL:WAYPOINT',
'SL:WAYPOINT_ADDED',
'SL:WAYPOINT_AVOID'}
from collections import Counter
from tqdm.auto import tqdm
import matplotlib.pyplot as plt
frequencies_big = Counter()
for schema_str in tqdm(big_subset.schema):
for token in all_schema_tokens:
if token in schema_str:
frequencies_big[token] += 1
frequencies_small = Counter()
for schema_str in tqdm(small_subset.schema):
for token in all_schema_tokens:
if token in schema_str:
frequencies_small[token] += 1
labels, values_big = zip(*frequencies_big.most_common())
values_small = [frequencies_small[l] for l in labels]
_plot_df = pd.DataFrame({'token': labels, 'big_subset': values_big, 'small_subset': values_small})
_plot_df.plot('token', ['big_subset', 'small_subset'], kind='bar', figsize=(18, 10))
small_subset_classes = [l for l, v_big, v_small in zip(labels, values_big, values_small) if v_small > 1.5 * v_big]
small_subset_classes
import os
os.makedirs('../data/splits/get_location_80_20/subset80')
os.makedirs('../data/splits/get_location_80_20/subset20')
big_subset.to_csv('../data/splits/get_location_80_20/subset80/train.tsv', sep='\t', index=False)
small_subset.to_csv('../data/splits/get_location_80_20/subset20/train.tsv', sep='\t', index=False)
len(big_subset), len(small_subset)
```
| github_jupyter |
# Welcome to Pynq Audio
This notebook shows the basic recording and playback features of the board.
It uses the audio jack to play back recordings from the built-in microphone, as well as a pre-recorded audio sample. Moreover, visualization with matplotlib and playback with IPython.Audio are shown.
## Create new audio object
```
from pynq.overlays.base import BaseOverlay
base = BaseOverlay("base.bit")
pAudio = base.audio
```
## Record and play
Record a 3-second sample and save it into a file.
```
pAudio.record(3)
pAudio.save("Recording_1.pdm")
```
## Load and play
Load a sample and play the loaded sample.
```
pAudio.load("/home/xilinx/pynq/lib/tests/pynq_welcome.pdm")
pAudio.play()
```
## Play in notebook
Users can also play the audio directly in notebook. To do this, the file format has to be converted from Pulse Density Modulation (PDM) to Pulse Code Modulation (PCM).
For more information, please refer to: https://en.wikipedia.org/wiki/Pulse-density_modulation.
### Step 1: Preprocessing
In this step, we first convert the 32-bit integer buffer to 16-bit. Then we divide 16-bit words (16 1-bit samples each) into 8-bit words with 1-bit sample each.
```
import time
import numpy as np
start = time.time()
af_uint8 = np.unpackbits(pAudio.buffer.astype(np.int16)
.byteswap(True).view(np.uint8))
end = time.time()
print("Time to convert {:,d} PDM samples: {:0.2f} seconds"
.format(np.size(pAudio.buffer)*16, end-start))
print("Size of audio data: {:,d} Bytes"
.format(af_uint8.nbytes))
```
### Step 2: Converting PDM to PCM
We now convert PDM to PCM by decimation. The sample rate is reduced from 3MHz to 32kHz.
We will remove the first and last 10 samples in case there are outliers introduced by decimation. We will also remove the DC offset from the waveform.
```
import time
from scipy import signal
start = time.time()
af_dec = signal.decimate(af_uint8,8,zero_phase=True)
af_dec = signal.decimate(af_dec,6,zero_phase=True)
af_dec = signal.decimate(af_dec,2,zero_phase=True)
af_dec = (af_dec[10:-10]-af_dec[10:-10].mean())
end = time.time()
print("Time to convert {:,d} Bytes: {:0.2f} seconds"
.format(af_uint8.nbytes, end-start))
print("Size of audio data: {:,d} Bytes"
.format(af_dec.nbytes))
del af_uint8
```
### Step 3: Audio Playback in Web Browser
```
from IPython.display import Audio as IPAudio
IPAudio(af_dec, rate=32000)
```
## Plotting PCM data
Users can display the audio data in notebook:
1. Plot the audio signal's amplitude over time.
2. Plot the spectrogram of the audio signal.
#### Amplitude over time
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.figure(num=None, figsize=(15, 5))
time_axis = np.arange(0,((len(af_dec))/32000),1/32000)
plt.title('Audio Signal in Time Domain')
plt.xlabel('Time in s')
plt.ylabel('Amplitude')
plt.plot(time_axis, af_dec)
plt.show()
```
#### Frequency spectrum
```
from scipy.fftpack import fft
yf = fft(af_dec)
yf_2 = yf[1:len(yf)//2]
xf = np.linspace(0.0, 32000//2, len(yf_2))
plt.figure(num=None, figsize=(15, 5))
plt.plot(xf, abs(yf_2))
plt.title('Magnitudes of Audio Signal Frequency Components')
plt.xlabel('Frequency in Hz')
plt.ylabel('Magnitude')
plt.show()
```
#### Frequency spectrum over time
Use the `classic` plot style for better display.
```
import matplotlib
np.seterr(divide='ignore',invalid='ignore')
matplotlib.style.use("classic")
plt.figure(num=None, figsize=(15, 4))
plt.title('Audio Signal Spectogram')
plt.xlabel('Time in s')
plt.ylabel('Frequency in Hz')
_ = plt.specgram(af_dec, Fs=32000)
```
| github_jupyter |
```
from __future__ import print_function
import cv2
import imutils
import numpy as np
import torch
import os
import torch.nn as nn
from collections import Counter, OrderedDict
from detection.layers.functions.prior_box import PriorBox
from detection.data import cfg
from detection.models.faceboxes import FaceBoxes
from detection.utils.box_utils import decode
from detection.utils.timer import Timer
from detection.utils.nms.py_cpu_nms import py_cpu_nms
from torchvision import models, transforms
import torch.backends.cudnn as cudnn
import torch.utils.data as data
device = 'cpu'
if torch.cuda.is_available():
torch.cuda.current_device()
device = 'cuda'
class_names = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I',
'J', 'K', 'L', 'M', 'N', 'O', 'P', 'Q', 'R',
'S', 'T', 'U', 'V', 'W', 'X', 'Y', 'Z',
'del', 'nothing', 'space']
special_classes = {'space': ' ', 'nothing': ''}
```
### Load FaceBoxes - model for detection
```
# Load weights for FaceBoxes model which is trained on hands
pretrained_dict = torch.load('weights/hand_boxes.pt')
# For some reason in downloaded weights there is prefix "module", so get rid of them
weights = OrderedDict((k.replace('module.', ''), v) for k, v in pretrained_dict.items())
torch.set_grad_enabled(False)
model = FaceBoxes(phase='test', size=None, num_classes=2)
model.load_state_dict(weights)
model.eval()
# print(model)
model = model.to(device)
```
### Load GoogLeNet - model for classification
```
asl = models.googlenet(pretrained=True)
num_ftrs = asl.fc.in_features
asl.fc = nn.Linear(num_ftrs, len(class_names))
asl.load_state_dict(torch.load('weights/asl_recognition.pt'))
asl.eval()
# print(asl)
asl = asl.to(device)
hand_transforms = transforms.Compose([
transforms.ToPILImage(),
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
def pre_detection_trans(img):
scale = torch.Tensor([img.shape[1], img.shape[0], img.shape[1], img.shape[0]])
img -= (104, 117, 123)
img = img.transpose(2, 0, 1)
img = torch.from_numpy(img).unsqueeze(0)
img = img.to(device)
scale = scale.to(device)
return img, scale
def post_detection_trans(out, im_size, treshold = {'acc': .7, 'nms': .6}):
priorbox = PriorBox(cfg, out[2], im_size, phase='test')
priors = priorbox.forward()
priors = priors.to(device)
loc, conf, _ = out
prior_data = priors.data
boxes = decode(loc.data.squeeze(0), prior_data, cfg['variance'])
boxes = boxes * scale / resize
boxes = boxes.cpu().numpy()
scores = conf.data.cpu().numpy()[:, 1]
# ignore low scores
inds = np.where(scores > treshold['acc'])[0]
boxes = boxes[inds]
scores = scores[inds]
# keep top-K before NMS
order = scores.argsort()[::-1][:5000]
boxes = boxes[order]
scores = scores[order]
# do NMS
dets = np.hstack((boxes, scores[:, np.newaxis])).astype(np.float32, copy=False)
keep = py_cpu_nms(dets, treshold['nms'])
return dets[keep, :]
resize = 2
_t = {'detect': Timer(), 'misc': Timer(), 'letter_pred': Timer()}
word_buf = ''
letter_buf = []
cap = cv2.VideoCapture(0)
try:
while(True):
ret, frame = cap.read()
frame = cv2.flip(frame, 1)
to_show = frame.copy()
img = np.float32(to_show)
if resize != 1:
img = cv2.resize(img, None, None, fx=resize, fy=resize, interpolation=cv2.INTER_LINEAR)
im_height, im_width, _ = img.shape
img, scale = pre_detection_trans(img)
_t['detect'].tic()
out = model(img)
_t['detect'].toc()
_t['misc'].tic()
dets = post_detection_trans(out, (im_height, im_width), {'acc': .7, 'nms': .6})
# keep top-K faster NMS
dets = dets[:750, :]
_t['misc'].toc()
for i in range(dets.shape[0]):
# get coordinates of rectangle corners
min_x = max([int(dets[i][0]) - 50, 0])
min_y = max([int(dets[i][1]) - 50, 0])
max_x = min([int(dets[i][2]) + 50, frame.shape[1]])
max_y = min([int(dets[i][3]) + 50, frame.shape[0]])
cv2.rectangle(to_show, (min_x, max_y), (max_x, min_y), [0, 0, 255], 3)
# make a prediction of letter
_t['letter_pred'].tic()
hand_img = frame[min_y:max_y, min_x:max_x]
hand_img = torch.stack([hand_transforms(hand_img)])
hand_img = hand_img.to(device)
asl_outputs = asl(hand_img)
_, preds = torch.max(asl_outputs, 1)
_t['letter_pred'].toc()
# store some last predictions and choose the most frequent one
# to decrease level of failures
letter_buf.append(class_names[preds[0]])
if len(letter_buf) == 10:
pred_letter = max(Counter(letter_buf).items(), key=lambda x: x[1])[0]
if pred_letter in special_classes:
word_buf += special_classes[pred_letter]
elif pred_letter == 'del':
word_buf = word_buf[:-1]
else:
word_buf += pred_letter
letter_buf = []
# clear buffer
if len(word_buf) > 20:
word_buf = word_buf[-20:]
font = cv2.FONT_HERSHEY_SIMPLEX
cv2.putText(to_show, word_buf,
(frame.shape[0] - 30 * len(word_buf), 50),
font, 2, (255,255,255), 2, cv2.LINE_AA)
cv2.imshow('image', to_show)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
except Exception as ex:
print(ex)
finally:
cap.release()
cv2.destroyAllWindows()
print('Average time for detection: ', _t['detect'].average_time)
print('Average time for post processing of detector output : ', _t['misc'].average_time)
print('Average time for classification: ', _t['letter_pred'].average_time)
```
| github_jupyter |
## import 需要的库
```
import pandas as pd
import os
import fnmatch
import numpy as np
import matplotlib.pyplot as plt
import shutil
import utils
import importlib
importlib.reload(utils)
# TODO:show some images for every category after categorizing
```
## Categorize图片
- 读取categories from file categories.txt
- 将训练数据按其对应的category分好,如果一个sample有多个category,则在每个category目录下这个sample都要出现
```E:\MY_STUDY_PLACE\PYTHON\JUPYTER\KAGGLE
├─.ipynb_checkpoints
└─categorizedTrain
├─Actin filaments
├─Aggresome
├─Cell junctions
├─Centrosome
├─Cytokinetic bridge
├─Cytoplasmic bodies
├─Cytosol
├─Endoplasmic reticulum
├─Endosomes
├─Focal adhesion sites
├─Golgi apparatus
├─Intermediate filaments
├─Lipid droplets
├─Lysosomes
├─Microtubule ends
├─Microtubule organizing center
├─Microtubules
├─Mitochondria
├─Mitotic spindle
├─Nuclear bodies
├─Nuclear membrane
├─Nuclear speckles
├─Nucleoli
├─Nucleoli fibrillar center
├─Nucleoplasm
├─Peroxisomes
├─Plasma membrane
└─Rods & rings
```
```
# test
categories = utils.readCategories()
print(categories)
# Test
labels = utils.readLabels(r"H:\all\train.csv")
def categorizeTrainingData(categories, trainDataPath="./train", trainLabelFile="./train.csv", rootPath=".\categorizedTrain", maxSize=-1):
"""
# Description: categorize the training data based on categories
# Arguments:
categories: dict of categories: {index:{'name':value}}
trainDataPath: the path that the training data are located
trainLabelFile: the file in which the training data's labels are saved
rootPath: rootPath where save the training data to, default is ".\categorizedTrain".
maxSize: the maxSize that are categorized for the following work.
# Returns:
None
# Raises:
None
"""
# create the category directory
for key,category in categories.items():
categoryPath = os.path.join(rootPath, category['name'])
utils.rebuildDir(categoryPath)
# read the trainLabels from trainLebelFile
labels = utils.readLabels(trainLabelFile)
count = 0;
for label in labels:
# print(label)
sampleId = label['Id']
targetList = label['targetList']
for targetId in targetList:
categoryName = categories[int(targetId)]['name']
categorizedDir = os.path.join(rootPath, categoryName)
# print(sampleId, targetId, categoryName)
# move file to categorized dir
try:
shutil.copy(os.path.join(trainDataPath, sampleId + "_green.png"), categorizedDir)
shutil.copy(os.path.join(trainDataPath, sampleId + "_red.png"), categorizedDir)
shutil.copy(os.path.join(trainDataPath, sampleId + "_blue.png"), categorizedDir)
shutil.copy(os.path.join(trainDataPath, sampleId + "_yellow.png"), categorizedDir)
except FileNotFoundError:
pass
count +=1
if maxSize >=0 and count >= maxSize:
break
# test
trainSize = 5000
categorizeTrainingData(categories, trainDataPath=r"H:\all\train", trainLabelFile=r"H:\all\train.csv", maxSize=trainSize)
```
## Expore the training data
```
# print the training data size and count the sample numbers based on labels
print("total # of training data is: ", trainSize)
def getDataSizeByLabel(dataPath="./"):
"""
# Description: get number of data based on different labels
# Arguments:
dataPath: the path that the training data are located, must categorized by labels
# Return:
dataSize: dictionary of data Size {label:size}
# Raises:
None
"""
dataSize = {}
for fileName in os.listdir(dataPath):
numberOfSample = len(fnmatch.filter(os.listdir(os.path.join(dataPath, fileName)), '*_green.png'))
dataSize[fileName] = numberOfSample
return dataSize
def plotDataSizeByLabel(dataSize):
"""
# Description: plot a bar chart of the dataSize by Label
# Arguments:
dataSize:dictionary of data Size {label:size}
# Return:
None
# Raises:
None
"""
# this is for plotting purpose
index = np.arange(len(dataSize))
labelNames = dataSize.keys()
labelValues = dataSize.values()
print(labelNames)
print(labelValues)
plt.figure(figsize=(20,16))
plt.tick_params(labelsize=16)
plt.barh(index, labelValues, align="center")
plt.ylabel('Label', fontsize=20)
plt.xlabel('No of Samples', fontsize=20)
plt.yticks(index, labelNames, fontsize=16, rotation=0)
plt.title('DataSize By Label')
plt.show()
dataSize = getDataSizeByLabel("./categorizedTrain")
print(dataSize)
plotDataSizeByLabel(dataSize)
```
we can see the distribution of the sample size is skew badly
| github_jupyter |
# Udacity Computer Vision Nano Degree
## Project 1 - Facial Keypoint Detection
Margaret Maynard-Reid, 6/11/2019
## Define the Convolutional Neural Network
After you've looked at the data you're working with and, in this case, know the shapes of the images and of the keypoints, you are ready to define a convolutional neural network that can *learn* from this data.
In this notebook and in `models.py`, you will:
1. Define a CNN with images as input and keypoints as output
2. Construct the transformed FaceKeypointsDataset, just as before
3. Train the CNN on the training data, tracking loss
4. See how the trained model performs on test data
5. If necessary, modify the CNN structure and model hyperparameters, so that it performs *well* **\***
**\*** What does *well* mean?
"Well" means that the model's loss decreases during training **and**, when applied to test image data, the model produces keypoints that closely match the true keypoints of each face. And you'll see examples of this later in the notebook.
---
## CNN Architecture
Recall that CNN's are defined by a few types of layers:
* Convolutional layers
* Maxpooling layers
* Fully-connected layers
You are required to use the above layers and encouraged to add multiple convolutional layers and things like dropout layers that may prevent overfitting. You are also encouraged to look at literature on keypoint detection, such as [this paper](https://arxiv.org/pdf/1710.00977.pdf), to help you determine the structure of your network.
### TODO: Define your model in the provided file `models.py` file
This file is mostly empty but contains the expected name and some TODO's for creating your model.
---
## PyTorch Neural Nets
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the feedforward behavior of a network that employs those initialized layers in the function `forward`, which takes in an input image tensor, `x`. The structure of this Net class is shown below and left for you to fill in.
Note: During training, PyTorch will be able to perform backpropagation by keeping track of the network's feedforward behavior and using autograd to calculate the update to the weights in the network.
#### Define the Layers in ` __init__`
As a reminder, a conv/pool layer may be defined like this (in `__init__`):
```
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
```
#### Refer to Layers in `forward`
Then referred to in the `forward` function like this, in which the conv1 layer has a ReLu activation applied to it before maxpooling is applied:
```
x = self.pool(F.relu(self.conv1(x)))
```
Best practice is to place any layers whose weights will change during the training process in `__init__` and refer to them in the `forward` function; any layers or functions that always behave in the same way, such as a pre-defined activation function, should appear *only* in the `forward` function.
#### Why models.py
You are tasked with defining the network in the `models.py` file so that any models you define can be saved and loaded by name in different notebooks in this project directory. For example, by defining a CNN class called `Net` in `models.py`, you can then create that same architecture in this and other notebooks by simply importing the class and instantiating a model:
```
from models import Net
net = Net()
```
```
# load the data if you need to; if you have already loaded the data, you may comment this cell out
# -- DO NOT CHANGE THIS CELL -- #
!mkdir /data
!wget -P /data/ https://s3.amazonaws.com/video.udacity-data.com/topher/2018/May/5aea1b91_train-test-data/train-test-data.zip
!unzip -n /data/train-test-data.zip -d /data
```
<div class="alert alert-info">**Note:** Workspaces automatically close connections after 30 minutes of inactivity (including inactivity while training!). Use the code snippet below to keep your workspace alive during training. (The active_session context manager is imported below.)
</div>
```
from workspace_utils import active_session
with active_session():
train_model(num_epochs)
```
```
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# import utilities to keep workspaces alive during model training
from workspace_utils import active_session
# watch for any changes in model.py, if it changes, re-load it automatically
%load_ext autoreload
%autoreload 2
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import Net
net = Net()
print(net)
```
## Transform the dataset
To prepare for training, create a transformed dataset of images and keypoints.
### TODO: Define a data transform
In PyTorch, a convolutional neural network expects a torch image of a consistent size as input. For efficient training, and so your model's loss does not blow up during training, it is also suggested that you normalize the input images and keypoints. The necessary transforms have been defined in `data_load.py` and you **do not** need to modify these; take a look at this file (you'll see the same transforms that were defined and applied in Notebook 1).
To define the data transform below, use a [composition](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) of:
1. Rescaling and/or cropping the data, such that you are left with a square image (the suggested size is 224x224px)
2. Normalizing the images and keypoints; turning each RGB image into a grayscale image with a color range of [0, 1] and transforming the given keypoints into a range of [-1, 1]
3. Turning these images and keypoints into Tensors
These transformations have been defined in `data_load.py`, but it's up to you to call them and create a `data_transform` below. **This transform will be applied to the training data and, later, the test data**. It will change how you go about displaying these images and keypoints, but these steps are essential for efficient training.
As a note, should you want to perform data augmentation (which is optional in this project), and randomly rotate or shift these images, a square image size will be useful; rotating a 224x224 image by 90 degrees will result in the same shape of output.
```
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='/data/training_frames_keypoints.csv',
root_dir='/data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## Batching and loading data
Next, having defined the transformed dataset, we can use PyTorch's DataLoader class to load the training data in batches of whatever size as well as to shuffle the data for training the model. You can read more about the parameters of the DataLoader, in [this documentation](http://pytorch.org/docs/master/data.html).
#### Batch size
Decide on a good batch size for training your model. Try both small and large batch sizes and note how the loss decreases as the model trains. Too large a batch size may cause your model to crash and/or run out of memory while training.
**Note for Windows users**: Please change the `num_workers` to 0 or you may face some issues with your DataLoader failing.
```
# load training data in batches
batch_size = 16
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
```
## Before training
Take a look at how this model performs before it trains. You should see that the keypoints it predicts start off in one spot and don't match the keypoints on a face at all! It's interesting to visualize this behavior so that you can compare it to the model after training and see how the model has improved.
#### Load in the test dataset
The test dataset is one that this model has *not* seen before, meaning it has not trained with these images. We'll load in this test data and before and after training, see how your model performs on this set!
To visualize this test data, we have to go through some un-transformation steps to turn our images into python images from tensors and to turn our keypoints back into a recognizable range.
```
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='/data/test_frames_keypoints.csv',
root_dir='/data/test/',
transform=data_transform)
# load test data in batches
batch_size = 16
test_loader = DataLoader(test_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
```
## Apply the model on a test sample
To test the model on a test sample of data, you have to follow these steps:
1. Extract the image and ground truth keypoints from a sample
2. Wrap the image in a Variable, so that the net can process it as input and track how it changes as the image moves through the network.
3. Make sure the image is a FloatTensor, which the model expects.
4. Forward pass the image through the net to get the predicted, output keypoints.
This function test how the network performs on the first batch of test data. It returns the images, the transformed images, the predicted keypoints (produced by the model), and the ground truth keypoints.
```
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
```
#### Debugging tips
If you get a size or dimension error here, make sure that your network outputs the expected number of keypoints! Or if you get a Tensor type error, look into changing the above code that casts the data into float types: `images = images.type(torch.FloatTensor)`.
```
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
```
## Visualize the predicted keypoints
Once we've had the model produce some predicted output keypoints, we can visualize these points in a way that's similar to how we've displayed this data before, only this time, we have to "un-transform" the image/keypoint data to display it.
Note that I've defined a *new* function, `show_all_keypoints` that displays a grayscale image, its predicted keypoints and its ground truth keypoints (if provided).
```
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
```
#### Un-transformation
Next, you'll see a helper function. `visualize_output` that takes in a batch of images, predicted keypoints, and ground truth keypoints and displays a set of those images and their true/predicted keypoints.
This function's main role is to take batches of image and keypoint data (the input and output of your CNN), and transform them into numpy images and un-normalized keypoints (x, y) for normal display. The un-transformation process turns keypoints and images into numpy arrays from Tensors *and* it undoes the keypoint normalization done in the Normalize() transform; it's assumed that you applied these transformations when you loaded your test data.
```
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=10):
for i in range(batch_size):
plt.figure(figsize=(20,10))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's Variable wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts)
```
## Training
#### Loss function
Training a network to predict keypoints is different than training a network to predict a class; instead of outputting a distribution of classes and using cross entropy loss, you may want to choose a loss function that is suited for regression, which directly compares a predicted value and target value. Read about the various kinds of loss functions (like MSE or L1/SmoothL1 loss) in [this documentation](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html).
### TODO: Define the loss and optimization
Next, you'll define how the model will train by deciding on the loss function and optimizer.
---
```
## TODO: Define the loss and optimization
import torch.optim as optim
criterion = nn.MSELoss()
optimizer = optim.Adam(net.parameters(), lr=0.0005)
```
## Training and Initial Observation
Now, you'll train on your batched training data from `train_loader` for a number of epochs.
To quickly observe how your model is training and decide on whether or not you should modify it's structure or hyperparameters, you're encouraged to start off with just one or two epochs at first. As you train, note how your the model's loss behaves over time: does it decrease quickly at first and then slow down? Does it take a while to decrease in the first place? What happens if you change the batch size of your training data or modify your loss function? etc.
Use these initial observations to make changes to your model and decide on the best architecture before you train for many epochs and create a final model.
```
def train_net(n_epochs):
# prepare the net for training
net.train()
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image']
key_pts = data['keypoints']
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.FloatTensor)
images = images.type(torch.FloatTensor)
# forward pass to get outputs
output_pts = net(images)
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts)
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
# print loss statistics
running_loss += loss.item()
if batch_i % 10 == 9: # print every 10 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/10))
running_loss = 0.0
print('Finished Training')
# train your network
n_epochs = 5 # start small, and increase when you've decided on your model structure and hyperparams
%%time
# this is a Workspaces-specific context manager to keep the connection
# alive while training your model, not part of pytorch
with active_session():
train_net(n_epochs)
```
## Test data
See how your model performs on previously unseen, test data. We've already loaded and transformed this data, similar to the training data. Next, run your trained model on these images to see what kind of keypoints are produced. You should be able to see if your model is fitting each new face it sees, if the points are distributed randomly, or if the points have actually overfitted the training data and do not generalize.
```
# get a sample of test data again
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts)
```
Once you've found a good model (or two), save your model so you can load it and use it later!
Save your models but please **delete any checkpoints and saved models before you submit your project** otherwise your workspace may be too large to submit.
```
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'keypoints_model_20190611.pt'
# after training, save your model parameters in the dir 'saved_models'
torch.save(net.state_dict(), model_dir+model_name)
```
After you've trained a well-performing model, answer the following questions so that we have some insight into your training and architecture selection process. Answering all questions is required to pass this project.
### Question 1: What optimization and loss functions did you choose and why?
**Answer**:
* I chose MSELoss since the keypoint detection is a regression problem.
* I chose Adam optimizer with learning rate of 0.001 as the NaimishNet paper suggested
### Question 2: What kind of network architecture did you start with and how did it change as you tried different architectures? Did you decide to add more convolutional layers or any layers to avoid overfitting the data?
**Answer**: I started out with implementing the same network architecture as in [the NaimishNet paper](https://arxiv.org/pdf/1710.00977.pdf). Training in the Udacity workspace or Colab with GPU doesn't allow the same parameters as the paper (i.e. 128 batch size with many epochs). My training loss doesn't seem to trend down. Then I used batch normalization instead of dropout layers which seemed to result in loss trending down.
### Question 3: How did you decide on the number of epochs and batch_size to train your model?
**Answer**:
* **Number of Epochs** - I first started at 1 then gradually increasd to 5. It trains fairly slow since this Udacity workspace doesn't have GPU. So I kept the epoch numbers small.
* **Batch size** - I started with 10 as what was in the notebook. It was too slow so I increased to 128 as the paper suggested, got error "DataLoader worker (pid 1219) is killed by signal: Bus error." So I tried a few other lower numbers and looks like 16 is the highest I can go with training in the workspace. I could train with batch_size = 64 when training in Colab with GPU.
## Feature Visualization
Sometimes, neural networks are thought of as a black box, given some input, they learn to produce some output. CNN's are actually learning to recognize a variety of spatial patterns and you can visualize what each convolutional layer has been trained to recognize by looking at the weights that make up each convolutional kernel and applying those one at a time to a sample image. This technique is called feature visualization and it's useful for understanding the inner workings of a CNN.
In the cell below, you can see how to extract a single filter (by index) from your first convolutional layer. The filter should appear as a grayscale grid.
```
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.numpy()
filter_index = 0
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
```
## Feature maps
Each CNN has at least one convolutional layer that is composed of stacked filters (also known as convolutional kernels). As a CNN trains, it learns what weights to include in it's convolutional kernels and when these kernels are applied to some input image, they produce a set of **feature maps**. So, feature maps are just sets of filtered images; they are the images produced by applying a convolutional kernel to an input image. These maps show us the features that the different layers of the neural network learn to extract. For example, you might imagine a convolutional kernel that detects the vertical edges of a face or another one that detects the corners of eyes. You can see what kind of features each of these kernels detects by applying them to an image. One such example is shown below; from the way it brings out the lines in an the image, you might characterize this as an edge detection filter.
<img src='images/feature_map_ex.png' width=50% height=50%/>
Next, choose a test image and filter it with one of the convolutional kernels in your trained CNN; look at the filtered output to get an idea what that particular kernel detects.
### TODO: Filter an image to see the effect of a convolutional kernel
---
```
##TODO: load in and display any image from the transformed test dataset
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
import cv2
original = cv2.imread('./images/straw_hat.png')
original = cv2.cvtColor(original, cv2.COLOR_BGR2RGB)
plt.imshow(original)
filtered = cv2.filter2D(original, -1,w[filter_index][0])
plt.imshow(filtered, cmap='gray')
```
### Question 4: Choose one filter from your trained CNN and apply it to a test image; what purpose do you think it plays? What kind of feature do you think it detects?
**Answer**: the feature seems to be detecting horizontal lines.
---
## Moving on!
Now that you've defined and trained your model (and saved the best model), you are ready to move on to the last notebook, which combines a face detector with your saved model to create a facial keypoint detection system that can predict the keypoints on *any* face in an image!
| github_jupyter |
## Problema
Tenemos pelotitas de 3 colores: Rojas, Verdes y Azules. Queremos contar la cantidad de formas en las que las podemos ordenar, sin tener dos pelotas seguidas del mismo color.
- Tenemos $R$ pelotitas rojas.
- Tenemos $G$ pelotitas verdes.
- Tenemos $B$ pelotitas azules.
### Ejemplos:
Entrada: ${r = 2, g = 2, b = 0}$.
$$ [r, g, r, g] $$
$$ [g, r, g, r] $$
### Como resolver este problema
Vamos a hacer una solución por fuerza bruta, probando todas las formas en las cuales podemos ordenar la secuencia de pelotitas.
### Caso base
- Si me queda la última bola sin ubicar, tengo una única opcion.
- Si me quedan mas que una bola de un único color, no voy a tener forma de ordenarlas sin que sean consecutivas. Tengo 0 opciones.
$$
f(r, 0, 0) = \left\{\begin{matrix}
1 \mathrm{\, si\, } r = 1 \\
0 \mathrm{\, si\, } r > 1
\end{matrix}\right.
$$
$$
f(0, g, 0) = \left\{\begin{matrix}
1 \mathrm{\, si\, } g = 1 \\
0 \mathrm{\, si\, } g > 1
\end{matrix}\right.
$$
$$
f(0, 0, b) = \left\{\begin{matrix}
1 \mathrm{\, si\, } b = 1 \\
0 \mathrm{\, si\, } b > 1
\end{matrix}\right.
$$
### Caso general
Para el caso general, tengo que sumar la cantidad de combinaciones que surgen de elegir cada color.
- Teniendo en cuenta que no podemos repetir el color anterior.
```
%time
def f(r, g, b, prev =None):
if g == 0 and b == 0:
return 1 if r == 1 else 0
elif r == 0 and b == 0:
return 1 if g == 1 else 0
elif r == 0 and g == 0:
return 1 if b == 1 else 0
# Caso gral:
ans = 0
if prev != 'r' and 0 < r:
ans += f(r - 1, g, b, 'r')
if prev != 'g' and 0 < g:
ans += f(r, g - 1, b, 'g')
if prev != 'b' and 0 < b:
ans += f(r, g, b - 1, 'b')
return ans
f(4, 4, 4)
```
## Memoizado al rescate
Esto no escala para input grandes...
Pero los sub-problemas se repiten. podemos guardarnos los resultados para ahorrarnos computarlos otra vez.
```
%time
memoria = {}
def cantidad_secuencias(r, g, b, prev = None):
global memoria
input = (r, g, b, prev)
if input in memoria:
return memoria[input]
# Caso base
if g == 0 and b ==0:
return 1 if r == 1 else 0
elif r == 0 and b ==0:
return 1 if g == 1 else 0
elif r == 0 and g ==0:
return 1 if b == 1 else 0
ans = 0
# Probé poniendo uno rojo y cuento todas las combinaciones de colas de esa seq
if prev != 'r' and r > 0:
ans += cantidad_secuencias(r - 1, g, b, prev='r')
# Probé poniendo uno verde y cuento todas las combinaciones de colas de esa seq
if prev != 'g' and g > 0:
ans += cantidad_secuencias(r, g - 1, b, prev='g')
# Probé poniendo uno azul y cuento todas las combinaciones de colas de esa seq
if prev != 'b' and b > 0:
ans += cantidad_secuencias(r, g, b - 1, prev='b')
memoria[input] = ans
return ans
cantidad_secuencias(100, 100, 100)
```
- Tenemos un algoritmo recursivo
- Los inputs de cada llamado recursivo se repien
- En vez de calcular varias veces lo mismo, hacemos un lookup en un diccionario, un array o lo que nos convenga.
1. declaro una `memoria`
2. Entro a un llamado recursivo
3. Me fijo si ya conozco la respuesta
1. Si ya conozco la respuesta, la devuelvo
2. Si no la conozco, la computo recursivamente
4. Finalmente obtengo una respuesta
5. Guardo esa respuesta
6. Devuelvo.
$$
(r, g, b, prev) \in \{R, G, B, 3\} \in O(3RGB) \in O(N^3)
$$
donde $n= max(R, G, B)$
| github_jupyter |
# Navigation
---
In this notebook, you will learn how to use the Unity ML-Agents environment for the first project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893).
### 1. Start the Environment
We begin by importing some necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
```
from unityagents import UnityEnvironment
import numpy as np
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
import pandas as pd
%matplotlib inline
```
Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
- **Mac**: `"path/to/Banana.app"`
- **Windows** (x86): `"path/to/Banana_Windows_x86/Banana.exe"`
- **Windows** (x86_64): `"path/to/Banana_Windows_x86_64/Banana.exe"`
- **Linux** (x86): `"path/to/Banana_Linux/Banana.x86"`
- **Linux** (x86_64): `"path/to/Banana_Linux/Banana.x86_64"`
- **Linux** (x86, headless): `"path/to/Banana_Linux_NoVis/Banana.x86"`
- **Linux** (x86_64, headless): `"path/to/Banana_Linux_NoVis/Banana.x86_64"`
For instance, if you are using a Mac, then you downloaded `Banana.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
```
env = UnityEnvironment(file_name="Banana.app")
```
```
env = UnityEnvironment(file_name=".\Banana_Windows_x86_64\Banana.exe")
```
Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
```
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
print("brain_name:", brain_name)
print("brain: ", brain)
```
### 2. Examine the State and Action Spaces
The simulation contains a single agent that navigates a large environment. At each time step, it has four actions at its disposal:
- `0` - walk forward
- `1` - walk backward
- `2` - turn left
- `3` - turn right
The state space has `37` dimensions and contains the agent's velocity, along with ray-based perception of objects around agent's forward direction. A reward of `+1` is provided for collecting a yellow banana, and a reward of `-1` is provided for collecting a blue banana.
Run the code cell below to print some information about the environment.
```
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.vector_observations[0]
print('States look like:', state)
state_size = len(state)
print('States have length:', state_size)
```
### 3. Take Random Actions in the Environment
In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
Once this cell is executed, you will watch the agent's performance, if it selects an action (uniformly) at random with each time step. A window should pop up that allows you to observe the agent, as it moves through the environment.
Of course, as part of the project, you'll have to change the code so that the agent is able to use its experience to gradually choose better actions when interacting with the environment!
```
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = np.random.randint(action_size) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
```
When finished, you can close the environment.
```
#env.close()
```
### 4. It's Your Turn!
Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
```python
env_info = env.reset(train_mode=True)[brain_name]
```
```
from dqn_agent import Agent
agent = Agent(state_size, action_size,seed=0)
def dqn(n_episodes=2000, eps_start=1.0, eps_end=0.01, eps_decay=0.99):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
N=100 # Contains size of scoring window
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=N) # last 100 scores
eps = eps_start # initialize epsilon
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
state = env_info.vector_observations[0] # get the current state
score = 0
while True:
action = int(agent.act(state, eps)) # agent takes action
#action = np.random.randint(action_size)
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state, action, reward, next_state, done) # update agent with experience tuple
state = next_state # update state
score += reward # update total score
if done:
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
# Print Score while training
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % N == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
# Stop when required average score of 13 (+ a little extra buffer) is reached
if np.mean(scores_window) >= 13.:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-N, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), 'model_weights.pth') #save the weights
break
return scores
scores = dqn()
fig, ax = plt.subplots(1, 1, figsize=[10, 5])
plt.rcParams.update({'font.size': 14})
scores_rolling = pd.Series(scores).rolling(100).mean()
ax.plot(scores, "-", c="blue", alpha=0.25)
ax.plot(scores_rolling, "-", c="red", linewidth=2)
ax.set_xlabel("Episode")
ax.set_ylabel("Score")
ax.grid(which="major")
ax.axhline(13, c="green", linewidth=3, alpha=0.5)
ax.legend(["Reward per Episode", "Average of last 100 Episodes", "Target Score"])
fig.tight_layout()
fig.savefig("plot.jpg")
env.close()
```
| github_jupyter |
# Federated Learning Training Plan: Host Plan & Model
Here we load Plan and Model params created earlier in "Create Plan" notebook
and host them on PyGrid.
After that it should be possible to run FL worker using
SwiftSyft, KotlinSyft, syft.js, or FL python worker
and train the hosted model using local worker's data.
```
%load_ext autoreload
%autoreload 2
import warnings
warnings.filterwarnings("ignore")
import websockets
import json
import requests
import torch
import syft as sy
from syft.grid.grid_client import GridClient
from syft.serde import protobuf
from syft_proto.execution.v1.plan_pb2 import Plan as PlanPB
from syft_proto.execution.v1.state_pb2 import State as StatePB
sy.make_hook(globals())
# force protobuf serialization for tensors
hook.local_worker.framework = None
async def sendWsMessage(data):
async with websockets.connect('ws://' + gatewayWsUrl) as websocket:
await websocket.send(json.dumps(data))
message = await websocket.recv()
return json.loads(message)
def deserializeFromBin(worker, filename, pb):
with open(filename, "rb") as f:
bin = f.read()
pb.ParseFromString(bin)
return protobuf.serde._unbufferize(worker, pb)
```
## Step 4a: Host in PyGrid
Here we load "ops list" Plan.
PyGrid should translate it to other types (e.g. torchscript) automatically.
```
# Load files with protobuf created in "Create Plan" notebook.
training_plan = deserializeFromBin(hook.local_worker, "tp_full.pb", PlanPB())
model_params_state = deserializeFromBin(hook.local_worker, "model_params.pb", StatePB())
```
Follow PyGrid README.md to build `openmined/grid-gateway` image from the latest `dev` branch
and spin up PyGrid using `docker-compose up --build`.
```
# Default gateway address when running locally
gatewayWsUrl = "127.0.0.1:5000"
grid = GridClient(id="test", address=gatewayWsUrl, secure=False)
grid.connect()
```
Define name, version, configs.
```
# These name/version you use in worker
name = "mnist"
version = "1.0.0"
client_config = {
"name": name,
"version": version,
"batch_size": 64,
"lr": 0.005,
"max_updates": 100 # custom syft.js option that limits number of training loops per worker
}
server_config = {
"min_workers": 3, # temporarily this plays role "min # of worker's diffs" for triggering cycle end event
"max_workers": 3,
"pool_selection": "random",
"num_cycles": 5,
"do_not_reuse_workers_until_cycle": 4,
"cycle_length": 28800,
"minimum_upload_speed": 0,
"minimum_download_speed": 0
}
```
Shoot!
If everything's good, success is returned.
If the name/version already exists in PyGrid, change them above or cleanup PyGrid db by re-creating docker containers (e.g. `docker-compose up --force-recreate`).
```
response = grid.host_federated_training(
model=model_params_state,
client_plans={'training_plan': training_plan},
client_protocols={},
server_averaging_plan=None,
client_config=client_config,
server_config=server_config
)
print("Host response:", response)
```
Let's double-check that data is loaded by requesting a cycle.
(Request is made directly, will be methods on grid client in the future)
```
auth_request = {
"type": "federated/authenticate",
"data": {}
}
auth_response = await sendWsMessage(auth_request)
print('Auth response: ', json.dumps(auth_response, indent=2))
cycle_request = {
"type": "federated/cycle-request",
"data": {
"worker_id": auth_response['data']['worker_id'],
"model": name,
"version": version,
"ping": 1,
"download": 10000,
"upload": 10000,
}
}
cycle_response = await sendWsMessage(cycle_request)
print('Cycle response:', json.dumps(cycle_response, indent=2))
worker_id = auth_response['data']['worker_id']
request_key = cycle_response['data']['request_key']
model_id = cycle_response['data']['model_id']
training_plan_id = cycle_response['data']['plans']['training_plan']
```
Let's download model and plan (both versions) and check they are actually workable.
```
# Model
req = requests.get(f"http://{gatewayWsUrl}/federated/get-model?worker_id={worker_id}&request_key={request_key}&model_id={model_id}")
model_data = req.content
pb = StatePB()
pb.ParseFromString(req.content)
model_params_downloaded = protobuf.serde._unbufferize(hook.local_worker, pb)
print("Params shapes:", [p.shape for p in model_params_downloaded.tensors()])
# Plan "list of ops"
req = requests.get(f"http://{gatewayWsUrl}/federated/get-plan?worker_id={worker_id}&request_key={request_key}&plan_id={training_plan_id}&receive_operations_as=list")
pb = PlanPB()
pb.ParseFromString(req.content)
plan_ops = protobuf.serde._unbufferize(hook.local_worker, pb)
print(plan_ops.code)
print(plan_ops.torchscript)
# Plan "torchscript"
req = requests.get(f"http://{gatewayWsUrl}/federated/get-plan?worker_id={worker_id}&request_key={request_key}&plan_id={training_plan_id}&receive_operations_as=torchscript")
pb = PlanPB()
pb.ParseFromString(req.content)
plan_ts = protobuf.serde._unbufferize(hook.local_worker, pb)
print(plan_ts.code)
print(plan_ts.torchscript.code)
```
## Step 5a: Train
To train hosted model, use one of the existing FL workers:
* Python FL Client: see "[Execute Plan with Python FL Client](Execute%20Plan%20with%20Python%20FL%20Client.ipynb)" notebook that
has example of using python FL worker.
* [SwiftSyft](https://github.com/OpenMined/SwiftSyft)
* [KotlinSyft](https://github.com/OpenMined/KotlinSyft)
* [syft.js](https://github.com/OpenMined/syft.js)
| github_jupyter |
# Completing the Picture II
*Review, missing pieces and projects*
## Today
1. Theory
- Classification
- Categorical data
- Supplements
1. Exercises and questions
- General feedback on submissions
- Dicussion and solution
1. Group projects
- Documents and deadlines
- Group allocation
- Checklist
# Classification
Dealing with *qualitative* response variables.
---
Based on ISL.
## Why Not Linear Regression?
Encoding should not matter:
- Numerical/quantitative representation of categories should be irrelevant
- Linear regression implies ordering
We are often interested in probabilities, i.e. values $\in [0,1]$:
- How likely is a certain outcome?
- Output of linear regression is unbound (cf. discussion of advertising data)
## Logistic Regression
Idea:
- Transform output of linear regression using a sigmoid function
<img src="../images/isl/isl_4.2_logistic_regression.png" alt="Accuracy, precision and bias" width="600">
Comments:
- Despite its name, logistic regression is a classification method
- Logistic regression is a special case of [Generalized linear models](https://en.wikipedia.org/wiki/Generalized_linear_model#Link_function)
- In scikit-learn: [Logisitic Regression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html#sklearn.linear_model.LogisticRegression)
```
import numpy as np
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
X, y = load_iris(return_X_y=True)
print('X:', X[:5, :], sep='\n')
print('y:', y[:5], sep='\n')
print('np.unique(y):', np.unique(y))
# create model
clf = LogisticRegression(solver='lbfgs', multi_class='multinomial', max_iter=500)
# fit
clf.fit(X, y)
# predict
clf.predict(X[:2])
# predict class probabilities
clf.predict_proba(X[:2])
# compute score
clf.score(X, y)
```
## Categorical Data
Dealing with *categorical* data.
---
Group projects: it may be necessary to *preprocess* the data
### Describing non-numerical values
Q:
- How do we compute a summary statistics for non-numerical values?
A:
- We treat them differently.
```
import pandas as pd
import seaborn as sns
%matplotlib inline
df = pd.read_csv('https://raw.githubusercontent.com/mwaskom/seaborn-data/master/tips.csv')
df
# default summary statistics only covers numerical variables
df.describe()
# we can include all by setting the corresponding paramter
df.describe(include='all')
# however, to obtain a better undestanding, it may make sense to look at them
# separately
print('sex:', df['sex'].value_counts(), sep='\n')
print()
print('smoker:', df['smoker'].value_counts(), sep='\n')
print()
print('day:', df['day'].value_counts(), sep='\n')
print()
print('time:', df['time'].value_counts(), sep='\n')
axs = pd.plotting.scatter_matrix(df, figsize=(10, 10))
```
### Categorical plots
Q:
- How do we deal with categorical values in visualizations?
A:
- Different options, very common to use categories as grouping variables.
See [seaborn](https://seaborn.pydata.org/api.html) for statistical data visualizations.
```
import seaborn as sns
sns.boxplot(data=df, y='tip', x='sex')
sns.boxplot(data=df, y='tip', x='day', order=['Fri', 'Thur', 'Sat', 'Sun'])
```
### Transforming non-numerical values
Q:
- How do we deal with non-numerical values?
A:
- We transform, i.e. encode them.
---
*Every continuous variable can be discretized/vectorized/categorized (with loss of information).*
```
# turn categories into numerical values (encoding)
print('original:')
print(df.head())
print()
columns_to_transform = ['sex', 'smoker', 'day', 'time']
df_transformed = df.copy() # we leave the original data untouched
for column in columns_to_transform:
labels, unique = pd.factorize(df_transformed[column])
df_transformed[column] = labels
print('transformed:')
print(df_transformed.head())
axs = pd.plotting.scatter_matrix(df_transformed, figsize=(10, 10))
# you can also replace values if you want to provide a custom mapping
# it may be easier to remember the encoding when it is done explicitely like here
custom_mapping = {'Male': 0, 'Female': 1}
df_transformed['sex'] = df['sex'].replace(custom_mapping)
df_transformed['sex']
```
### Using categorical data in a model
Q:
- Can we directly feed numerical categorical data into a model?
A: Depends on the model:
- Some models (like tree-based models) support categorical variables out of the box.
- Others (like the classical linear model) require variable transformations.
See also the [scikit-learn user guide](https://scikit-learn.org/stable/modules/preprocessing.html).
Q:
- Why/when do we (usually) not pass categorical data directly into e.g. a linear regression model?
A:
- Encoding and ordering should not matter
Idea:
- One-hot or dummy encode variables
#### Categorical to integer code
[`OrdinalEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OrdinalEncoder.html#sklearn.preprocessing.OrdinalEncoder) can be used to convert categorical features to integer codes.
This is similar to `pd.factorize`.
---
Group projects: use `pd.factorize` as first preprocessing step.
```
# choose a more interesting example displaying several categories at once
df_example = df.iloc[[0, 1, 200, 100, 240, 241, 243]]
df_example
from sklearn.preprocessing import OrdinalEncoder
X = df_example[['sex', 'day']].head()
print('original:')
print(X)
print()
encoder = OrdinalEncoder()
X_transformed = encoder.fit_transform(X)
print('transformed:')
print(X_transformed)
```
#### Categorical to one-hot or dummy encoding
[`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html#sklearn.preprocessing.OneHotEncoder) is used to convert categorical features into one-hot or dummy encoded variables.
---
Group projects: do not forget to one-hot encode variables with more than two categories if your model does not support categorical input
```
from sklearn.preprocessing import OneHotEncoder
X = df_example[['sex', 'day']].head()
print('original:')
print(X)
print()
encoder = OneHotEncoder()
X_transformed = encoder.fit_transform(X)
print('transformed (without dropping first):')
# print(X_transformed)
print(X_transformed.toarray()) # observe the co-linearity we created
encoder = OneHotEncoder(drop='first') # to encode each column into (n_categories - 1) columns
X_transformed = encoder.fit_transform(X)
print('transformed:')
# print(X_transformed)
print(X_transformed.toarray())
```
#### Preprocessing in a pipeline
[`ColumnTransformer`](https://scikit-learn.org/stable/modules/generated/sklearn.compose.ColumnTransformer.html#sklearn.compose.ColumnTransformer) is used to apply several (different) transformations to a DataFrame at once.
---
Group projects: if necessary, create a pipeline to simplify matters
```
from sklearn.compose import ColumnTransformer
ct = ColumnTransformer(
[
('encoder', OneHotEncoder(drop='first'), ['sex', 'day']),
# we could add an additional transformer here as well, e.g. to
# perform feature scaling
],
remainder='passthrough', # to not drop but pass through variables
# that are not transformed
)
X = df[['sex', 'day', 'total_bill']]
print('original:')
print(X)
print()
ct.fit(X) # we can call fit and transform independently
X_transformed = ct.transform(X)
print('transformed:')
print(X_transformed)
# example of how to use ColumnTransformer in a pipeline
from sklearn.linear_model import LinearRegression
from sklearn.pipeline import Pipeline
X = df[['sex', 'day', 'total_bill']]
y = df['tip']
regressor = LinearRegression()
model = Pipeline(
[
('preprocessor', ct),
('regressor', regressor)
]
)
model.fit(X, y)
model.score(X, y)
```
## Multiclass Classification
Multiclass: more than 2 response classes
Q:
- Which models support multiclass classification?
A:
- Some models natively support multiclass classification (e.g. tree-based methods).
- Models can be combined in a *one vs. all* fashion to produce a multiclass classifier.
# Supplements
## Ensemble Methods
> Combine multiple models to obtain better predictive performance.
Requirements:
- Different models
- Aggregation of results
---
*In practical applications (with a focus on predictive performance) ensemble methods are widely used.*
Two approaches:
- Averaging: build several independent models
- Boosting: build models sequentially
Comments:
- [Scikit-learn provides models for both](https://scikit-learn.org/stable/modules/ensemble.html#ensemble)
- [RandomForests](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html#sklearn.ensemble.RandomForestClassifier) are a classical example (reduce variance by aggregating many models)
- Ensemble methods may scale to very large data sets
## Bias Variance Trade-Off
> Trade-off between prediction accuracy and model interpretability
<a href="http://faculty.marshall.usc.edu/gareth-james/ISL/">
<img src="../images/isl/isl_2.7_bias_variance_trade_off.png" alt="Bias Variance Trade-Off" width="80%">
</a>
---
*In general, you cannot have both. Depends on use case what is more important to you.*
## Variable Importance
Q:
- Which variable importance measures do you know already?
A:
- Coefficients in linear regression if variable have same scale
- RandomForest feature importance
### Permutation Importance
Idea:
- The output of a model depends on different variables
- We *destroy* the information in one variable
- We observe how model performance then deteriorates
- The larger the effect, the more important the variable must be
Notes:
- This is independent of the model used
- *destroy information* e.g. through permuting values
See also:
- [Interpretable ML](https://christophm.github.io/interpretable-ml-book/)
- [ELI5](https://eli5.readthedocs.io/en/latest/overview.html)
- Example in "california housing" exercise
### Further Reading
Scikit-learn user guide:
- [Model selection and evaluation](https://scikit-learn.org/stable/model_selection.html)
- [Dataset transformations](https://scikit-learn.org/stable/data_transforms.html)
The [tutorial](https://scikit-learn.org/stable/tutorial/index.html) also lists a lot of
helpful examples.
---
[10 minutes to pandas](https://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html) is a good wrap up of how some common operations are carried out in pandas.
---
[matplotlib](https://matplotlib.org/index.html) comes with a large gallery of example plots in case you need something we have not yet covered.
---
For easy-to-create statistical data visualizations, take a look at [seaborn](https://seaborn.pydata.org/).
# Projects
See [here](https://github.com/caichinger/MLMNC2020/projects).
# Exercises
## Let's review a few submissions together!
## Before we start:
> We have received some very nice solutions! Congratulations on the progress you have made already! \o/
Over all exercises and students:
| | total_score |
|:------|--------------:|
| count | 90 |
| mean | 0.860766 |
| std | 0.159342 |
| min | 0.138889 |
| 25% | 0.808333 |
| 50% | 0.883333 |
| 75% | 0.9825 |
| max | 1 |
### Examples
*What do you like? What can be improved?*
Q:
2. Compare the decision boundaries between the `RandomForestClassifier` and the `KNeighborsClassifier`. How do they differ?
A:
The RandomForestClassifier exhibits linear and horizontal decision boundaries whereas the KNeighborsClassifier does not assume any specific form (apart from forming neighborhoods). the kneighborsclassifier ignores outliers, thus the line is much smoother when and thus it is easier to interpret however, the random forest classifier is more accurate because it includes the outliers but it makes it harder to interpret
Q:
1/b) Setting a lower but fixed value for `noise` and varying `n_neighbors`.
A:
If we choose n_neighbors too small (=1) the boundary becomes too flaky; the model is too flexible. There seems to be a good choice that roughly captures the shape of the expected boundary. If we choose n_neighbors too large (=50) we loose details; the model is no longer flexible enough.
Q:
2. What does `pd.plotting.scatter_matrix()` tell us about the data?
A:
It creates creates nxn plots where off the diagonal: scatter plots are used to visualize correlation between variables on the diagonal: the distribution of a variable is visualized This command shows a great overview of the correlation between the given input (TV, Radio, Newspaper) variable and the output (Sales) variable. The computed matrix of scatter plots visualizes the relationship between the variables and thereby complements df.corr() smoothly. Again, the abovementioned relationships/correlations between the variables can be observed. E.g., TV ads have the strongest linear and positive influence on sales, as can be seen in the upper-right corner matrix.
Q:
1. Why did we decide to scale the inputs?
A:
We wanted to infer variable importance by comparing the coefficients. Therefore, the variables need to share the same scale. We decided to scale the inputs to be able to accurately compare the correlation coefficients of TV, radio and newspaper advertising spending and sales in the model. Beforehand, the correlation coefficients were not comparable between the three advertising variables, as they depend on the respective scales.
Q:
1. Why did we decide to scale the inputs?
A:
We wanted to infer variable importance by comparing the coefficients. Therefore, the variables need to share the same scale. We decided to scale the input variables to account for their different ranges and hence the effect on the coefficients. As a result, we achieve a higher comparability of the coefficients due to scaling and the model performs better.
Q:
3/b) Using one of the models, describe and interpret the effect on sales when spending "an ever increasing amount on advertising". Is this behaviour reasonable?
A:
This behavior is not reasonable, cf. concept of diminishing returns. Using the original model, an ever increasing amount of ad spending also leads to ever rising sales - the effect is especially strong for TV and radio. It can be assumed that this does not reflect reality, as the law of diminishing returns would imply that there is a point where sales would virtually remain the same, even if ad spendings are further increased. Therefore, the second model was created to account for these dynamics.
Q:
3. How do the decision regions differ between the default and the cross-validated model?
A:
Less fragmented, fewer details, coarser boundary. Maybe too coarse. after cross validating, the decision regions become clearer and the model fit is better. Therefore we should cross validate the data first.
Q:
3/b) Using one of the models, describe and interpret the effect on sales when spending "an ever increasing amount on advertising". Is this behaviour reasonable?
A:
This behavior is not reasonable, cf. concept of diminishing returns. much_ads = [[100000, 100000, 100000]] much_ads_sales = model.predict(much_ads) print(much_ads_sales) [-5.18937105e+09] This behaviour is not reasonable as sales cannot turn negative. In "simple" linear model (without scaled inputs) sales would be predicted to become eternally high, which is also not reasonable.
Q:
3/b) Using one of the models, describe and interpret the effect on sales when spending "an ever increasing amount on advertising". Is this behaviour reasonable?
A:
This behavior is not reasonable, cf. concept of diminishing returns. 2) An ever increasing amount of money: Code: from sklearn.linear_model import LinearRegression regressor = LinearRegression() regressor.fit(X_train, y_train) print(regressor.coef_) --- [0.04405928 0.1992875 0.00688245] The coefficients show that when we
* increase TV spending by 1, then sales increase by 0.0441
* increase radio spending by 1, then sales increase by 0.1993
* increase newspaper spending by 1, then sales increase by 0.0069. I believe that this behavior is not reasonable as this means that we can increase spending forever and sales will always grow by these.
In practice, this is not true because we will reach a point where additional spending will not (or only slightly) increase sales. Hence, we can conclude that the model only performs well with realistic advertising spending (should be within the span of our test data set).
Q:
3. Based on your assessment, do you recommend to use the ML system or not? Provide an explanation.
A:
Based on this assessment, I do not recommend the ML system, because the probability of the model falsely predicting a positive outcome is quite high (0.29). Moreover, the loss for this case is -2000, which is rather high as well.
Q:
3. Based on your assessment, do you recommend to use the ML system or not? Provide an explanation.
A:
Yes, using the ML system is this case appears fairly accurate. The expected value is positive; that is, above zero. (This exercise was tricky due to axis labeling questions: changes on both axis from 0->1 (negative/positive) or vice versa had profound effect on result). If my calculations are done right, the expected value of 84 means the predicted values of true positives (TP) and true negatives (TN), 300 and 100 respectively, are correctly represented by the EV of 84. The ML system therefore works, and can be used.
Q:
3. Based on your assessment, do you recommend to use the ML system or not? Provide an explanation.
A:
In this case, I would not recommend to use this ML system for evaluation because, according to the confusion matrix, the model has only a 84% accuracy rate [(True Postive+True Negative)/Total], which should be improved to improve the confidence of the model. We have a 14% chance of getting a false positive (type 1 error). When we also take a look at the system's ability to detect events in the positive class, which is the sensitivity, we get a 71% rate (True positve + False Positive)/ Total Positives. Having these metrics, I assert that the performance of the system is insufficient to have a reliable prediction. Still, we have to check the context of our problem to understand the significance of having those test errors. When we take the current corona crisis as an example, a false negative would be detrimental as someone who is infected would not be put in quarantine, leading to further infections. If we take false positives as an example, it is problematic when someone is diagnosed with cancer and receives a treatment even though the person is healthy.
Q:
3. Based on your assessment, do you recommend to use the ML system or not? Provide an explanation.
A:
I would argue that the model does not perform well, as the expected value of an enitrely accurate model would be 228. Therefore, the current model's classification deviates by 50% from the correct classification of the underlying data. This also becomes apparent when plotting the decision boundaries of the model, where it seems that the data is overfitted.
Q:
4. List three specific tasks you consider important in tackling this problem.
A:
* Feature Scaling, Data exploration, Choosing model variables
Q:
5. Perform three specific tasks (not necessarily the same) to approach a solution and describe your findings.
A:
1. Latitude and longitude variables logically don't make any sense to put in a correlation unless they are converted into some other datapoints that tell us more about the region the unit is in. Besides, a quick correlation analysis shows that the house age also barely correlates with housing prices (unless they are really really old). All three variables can therefore be omitted from our model.
2. For correlation analysis, I decided to try both LinearRegression() as well as GridSearchCV().
3. Both models were measured using mean_squared_error. Using the test_dataset (20% of the whole dataset), GridSearchCV seems to perform better than LinearRegression.
Q:
5. Perform three specific tasks (not necessarily the same) to approach a solution and describe your findings.
A:
* From the descriptive statistics table I got to know the minimum/maximum value, mean, etc. of each feature.
* From the histogram, I discovered that the attributes have different scales and this will probably have to be fixed.
* Median income, average number of rooms and house age had the biggest three coefficient with price. Therefore, these three has the largest correlation with the value of house.
* I created a scatter plot with geographical information(longitude, latitude) and this provided me an overall picture of high-density areas, i.e. Bay area, LA, etc. And i also created a color map that helped me see that the prices are higher in locations closer to ocean.
* Finding these patterns or information can be used to create more relevant features like 'ocean proximity' to better predict the price. As the next step, I will select the most appropriate model to predict the house value and then split the data into test and train set.
Q:
5. Perform three specific tasks (not necessarily the same) to approach a solution and describe your findings.
A:
* First, I tried linear regression and got poor r2 score of 0.595770232606166,
* I then tried to optimize the model by adding polynomial features, but cross-validation showed that "1" is the best feature, so it turned me back to the simple linear model that I already analyzed,
* I then chose RandomForestClassifier model and applied cross-validation (grid search cv), which gave me r2 score of 0.8072513477374939.
Q:
5. Perform three specific tasks (not necessarily the same) to approach a solution and describe your findings.
A:
* By exploring correlations between the various attributes and the median house value, I found the strongest correlations with median income, total rooms and median age. It is difficult to eliminate overfitting of the data. Linear regression is not the best choice for predicting the median house value.
Q:
5. Perform three specific tasks (not necessarily the same) to approach a solution and describe your findings.
A:
* It is a supervised learning task and a multiple regression problem, but more specifically a univariate regression problem because the goal is to use multiple features to make one prediction for each district.
* When I plotted the Price data using sns.distplot and noticed that the frequency for houses over 500000 is very high, so my assumption is that these are either outliers or the data is wrong. Because of this, I attempted to remove the outliers, but it didn’t work. Something in my code is wrong, but I believe that removing the outliers would make the model more accurate when testing the model with test data.
* In order to see if some variables had significant correlations, I created a heat map correlation matrix. From this matrix I can see that the top 2 positive correlations are average bedrooms – average rooms (85% correlation) and price – media income (69%). However, both of these correlations were to be expected, so it is not helpful in the analysis. Although, there was a strong negative correlation between block population and median house age in block. This leads me to wonder whether the population on the block decreases as the median age of the houses in a block increases, which could lead to the area being less desirable hence have a lower median house price, but an analysis on causation would need to be done to determine this.
## Takeaway Messages and Remarks
### Approaching and Discussing a Problem
1. Expectation: What do you expect based on your current knowledge? Write this down before you work on a problem.
1. Observation: What do you see? Describe your findings.
1. Interpretation: What does it mean and why does it matter? Explain cause and effect of your findings.
---
- Without clearly defined expectations, it is all too easy to fool yourself.
- Separating observation and interpretation facilitates the discussion. It makes it easier to review und improve. Your observations could be correct even if your interpretations are not.
### Making Assertions
When you make an assertion, check it's plausibility if you have not done so already.
*Code allows you to play around and try things out - do that.*
### Wording / Concepts
Q:
- What does linear mean?
A:
- Linear function in one variable $f(x) = ax + b$
- Linear relationship between $x$ and $y$ if it can be expressed as $y = f(x) = ax + b$
- Similarily in more dimension
- Graphically: A straight line (or a (hyper)plane)
---
*If something looks like a curve, it is not a linear relationship.*
Q:
- What is an outlier?
A:
- Outlier refers to a data point/record that differs significantly from other observations.
---
- By definition, an outlier is a rare event.
- If you are not explicitely interested in rare events, you do not want these events to
influence your model.
- Outliers may cause problems.
- Different ranges do not per se constitute outliers.
- Sometimes one is explicitely interested in outliers (outlier detection).
---
*The "california housing" data set contains a few outliers.*
Q:
- Model parameter vs. hyperparameter?
A:
- Hyperparameter is part of the model specification, defines family of possible models
- Model parameters are learned from the data (using `fit`)
- Hyperparameters are estimated using `GridSearch`*
---
*Additional hyperparameter search methods exist (random, model based, ...).
Examples:
- Coefficients in linear regression: model parameter
- Number of degrees in polynomial features fed into linear regression: hyperparameter
- Split-rules in decision tree: model parameter
- Max depth of tree: hyperparameter
- Number of estimators in random forest: hyperparameter
Q:
- What is (statistical) bias?
- Is bias a problem?
A:
- In general: consistent/structural deviation of result caused by systematic error in procedure
- In linear regression: intercept term (constant) is called bias
- Bias is considered problematic (often not known/considered)
---
- ML system may introduce or learn basis from data
- ML systems are not per se unbiased
---
**https://catalogofbias.org/**
[FAO](http://fao.org)'s basic statistical tools I
<img src="http://www.fao.org/3/w7295e2m.gif" alt="Accuracy" width="600">
[FAO](http://fao.org)'s basic statistical tools II
<img src="http://www.fao.org/3/w7295e2n.gif" alt="Accuracy, precision and bias" width="400">
Q:
- Train-test-split vs. cross validation (CV) vs. grid search vs. CV grid search?
A:
- Train-test-split:
- split data into train and test set
- different strategies: random subset, stratified split, ...
- CV:
- perform train-test-split several times
- different strategies: random, n-folds, ...
- Grid search:
- Define parameter grid
- For every parameter combination:
1. Fit model on train
2. Compute score on test
- Choose best parameter combination
- CV grid search: combine both grid search and CV into one procedure
| github_jupyter |
# Data Cleaning
```
# import the library
%matplotlib inline
import pandas as pd
import numpy as np
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# convert scientific notation to decimals
pd.set_option('display.float_format', lambda x: '%.2f' % x)
```
# 1. Load Datasets
```
#Source file: https://www.kaggle.com/usdot/flight-delays#flights.csv
#Main dataset
df_delayed_flights = pd.read_csv('data/flights.csv', low_memory=False)
#Complementary datasets
df_airports = pd.read_csv('data/airports.csv', low_memory=False)
df_airlines = pd.read_csv('data/airlines.csv', low_memory=False)
```
# 2. Summarize the data
```
print('------- Main Dataset, Flights -------')
print(df_delayed_flights.shape)
print(df_delayed_flights.columns)
print(df_delayed_flights.head())
print(df_delayed_flights.describe())
print('\n ------- Airports -------')
print(df_airports.shape)
print(df_airports.columns)
print(df_airports.head())
print(df_airports.describe())
print('\n ------- Airlines -------')
print(df_airlines.shape)
print(df_airlines.columns)
print(df_airlines.head())
print(df_airlines.describe())
```
# 3. Data Cleaning
## 3.1. Merge Columns - DATE
```
df_delayed_flights['DATE'] = pd.to_datetime(df_delayed_flights[['YEAR','MONTH', 'DAY']])
print(df_delayed_flights.columns)
```
## 3.2. Remove not relevant and duplicated columns
```
df_delayed_flights = df_delayed_flights.drop(['YEAR','DAY','DAY_OF_WEEK'], axis=1)
df_delayed_flights = df_delayed_flights.drop(['TAXI_OUT','TAXI_IN','WHEELS_OFF', 'WHEELS_ON','AIR_TIME','ELAPSED_TIME'], axis=1)
print(df_delayed_flights.columns)
df_airports = df_airports[['IATA_CODE','AIRPORT']]
print(df_airports.columns)
```
## 3.3. Rename Columns
```
df_airports.columns = ['AIRPORT_CODE','AIRPORT_NAME']
df_airlines.columns = ['AIRLINE_CODE','AIRLINE_NAME']
```
## 3.4. Change type
```
# Function to convert from 'HHMM' string to datetime.time
# Reference: https://www.kaggle.com/fabiendaniel/predicting-flight-delays-tutorial
import datetime
def format_time(time_string):
if isinstance(time_string, datetime.time):
return time_string
if pd.isnull(time_string):
return np.nan
else:
if time_string == 2400: #12 midnight
time_string = 0
time_string = "{0:04d}".format(int(time_string)) #Format the string to be as 4 decimals (hh:mm)
formated_time = datetime.time(int(time_string[0:2]), int(time_string[2:4])) #Split the 4 digits into 2 parts
return formated_time
# Call the Function:
df_delayed_flights['SCHEDULED_DEPARTURE'] = df_delayed_flights['SCHEDULED_DEPARTURE'].apply(format_time)
df_delayed_flights['DEPARTURE_TIME'] = df_delayed_flights['DEPARTURE_TIME'].apply(format_time)
df_delayed_flights['SCHEDULED_ARRIVAL'] =df_delayed_flights['SCHEDULED_ARRIVAL'].apply(format_time)
df_delayed_flights['ARRIVAL_TIME'] = df_delayed_flights['ARRIVAL_TIME'].apply(format_time)
#Print a sample..
df_delayed_flights.loc[:5, ['SCHEDULED_DEPARTURE', 'SCHEDULED_ARRIVAL', 'DEPARTURE_TIME',
'ARRIVAL_TIME', 'DEPARTURE_DELAY', 'ARRIVAL_DELAY']]
```
## 3.5. Change Values
```
#Replace cancellation reason with meaningful values
df_delayed_flights["CANCELLATION_REASON"].replace({'A':'Airline',
'B':'Weather',
'C':'National Air System',
'D':'Security'}, inplace=True)
df_delayed_flights["CANCELLATION_REASON"].value_counts()
```
## 3.6. Missing values
```
print("Delayed Flights Missing Values:\n", df_delayed_flights.isnull().sum())
print("Airlines Missing Values:\n", df_airlines.isnull().sum())
print("Airports Missing Values:\n", df_airports.isnull().sum())
```
### CANCELLATION_REASON
```
print("Total number of delayed flights: ", len(df_delayed_flights))
print("Cancelled flights= ", sum(df_delayed_flights['CANCELLED']))
print(df_delayed_flights['CANCELLATION_REASON'].value_counts())
```
#### Conclusion: Number of missing data under the variable "CANCELLATION_REASON" is large since when the flight was not cancelled, no reason code was assigned. We can replace no values with "Not Cancelled":
```
df_delayed_flights['CANCELLATION_REASON'] = df_delayed_flights['CANCELLATION_REASON'].fillna('Not_Cancelled')
df_delayed_flights['CANCELLATION_REASON'].isnull().sum()
print(df_delayed_flights['CANCELLATION_REASON'].value_counts())
len(df_delayed_flights)
df_delay = df_delayed_flights[(df_delayed_flights['DEPARTURE_DELAY']!=0)&(df_delayed_flights['ARRIVAL_DELAY']!=0)]
df_delay['target'] = -1
len(df_delay)
df_cancel = df_delayed_flights[df_delayed_flights['CANCELLED']==1]
df_cancel['target'] = 1
len(df_cancel)
df_ontime = df_delayed_flights[(~df_delayed_flights.index.isin(df_delay.index))&(~df_delayed_flights.index.isin(df_cancel.index))]
df_ontime['target'] = 0
len(df_ontime)
df_join = pd.concat([df_delay, df_cancel, df_ontime], axis=0)
df_delayed_flights.head()
len(df_join), len(df_delayed_flights)
delayed = []
for row in df_delayed_flights['DEPARTURE_DELAY'].isnull():
if row != 0:
delayed.append(df_delayed_flights['DEPARTURE_DELAY'].mean())
df_delayed_flights['delayed'] = delayed
for idx in df_delayed_flights.index[df_delayed_flights['CANCELLED'].notnull()]:
# checking only non-cancelled flights
if (df_delayed_flights.loc[idx,'CANCELLED'] == 0):
# print(df_delayed_flights['CANCELLED'][idx])
if (df_delayed_flights.loc[idx,'DEPARTURE_DELAY'] != 0 or df_delayed_flights.loc[idx,'ARRIVAL_DELAY'] != 0):
df_delayed_flights.loc[idx,'CANCELLED'] = -1
# print(df_delayed_flights['CANCELLED'][idx])
return df_delayed_flights
print(df_delayed_flights['CANCELLED'].value_counts())
df_delayed_flights
```
# 4. Remove Outliers
```
df_delayed_flights[["DEPARTURE_DELAY","ARRIVAL_DELAY"]].plot.box()
plt.show()
plt.hist(df_delayed_flights['ARRIVAL_DELAY'], bins=150)
plt.title("Arrival Delays")
plt.show()
plt.hist(df_delayed_flights['DEPARTURE_DELAY'], bins=150)
plt.title("Departure Delays")
plt.show()
#Determine Outliers
mean_arrival_delays = np.mean(df_delayed_flights['ARRIVAL_DELAY'] )
sd_arrival_delays = np.std(df_delayed_flights['ARRIVAL_DELAY'])
mean_departure_delays = np.mean(df_delayed_flights['DEPARTURE_DELAY'])
sd_departure_delays = np.std(df_delayed_flights['DEPARTURE_DELAY'])
print('Arrival Delays:\t \t Mean = {0} \t SD = {1}'.format(mean_arrival_delays, sd_arrival_delays))
print('Departure Delays:\t Mean = {0} \t SD = {1}'.format(mean_departure_delays, sd_departure_delays))
#Arrrival_delay or Departure_delay != 0
arrival_delays = df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] != 0.00]['ARRIVAL_DELAY']
departure_delays = df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] != 0.00]['DEPARTURE_DELAY']
print(arrival_delays.shape)
mean_ad = np.mean(arrival_delays)
sd_ad = np.std(arrival_delays)
mean_dd = np.mean(departure_delays)
sd_dd = np.std(departure_delays)
print("With removing on-time flights:")
print('Arrival Delays:\t \t Mean = {0} \t SD = {1}'.format(mean_ad, sd_ad))
print('Departure Delays:\t Mean = {0} \t SD = {1}'.format(mean_dd, sd_dd))
# Removing
flights_to_remove = []
# remove based on arrival and departure delays (normal distribution)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] > mean_ad + 3.0* sd_ad].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['ARRIVAL_DELAY'] < mean_ad - 3.0* sd_ad].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] > mean_dd + 3.0* sd_dd].index)
flights_to_remove = flights_to_remove + list(df_delayed_flights[df_delayed_flights['DEPARTURE_DELAY'] < mean_dd - 3.0* sd_dd].index)
print('')
print('# Flights to remove', len(flights_to_remove))
new_delayed_flights = df_delayed_flights[~df_delayed_flights.index.isin(flights_to_remove)]
print("Was: ", df_delayed_flights.shape, " Now: ", new_delayed_flights.shape)
plt.hist(new_delayed_flights['ARRIVAL_DELAY'], bins=150)
plt.title("Arrival Delays")
plt.show()
plt.hist(new_delayed_flights['DEPARTURE_DELAY'], bins=150)
plt.title("Departure Delays")
plt.show()
```
## 5. Merging datasets
```
df_merge_v1 = new_delayed_flights.copy()
#Merge Airlines and Flights dfs
df_merge_v1 = pd.merge(new_delayed_flights, df_airlines, left_on='AIRLINE', right_on='AIRLINE_CODE', how='left')
#Merge Airports and Flights dfs on Origin_Airport and Airport_Code
df_merge_v1 = pd.merge(df_merge_v1, df_airports, left_on='ORIGIN_AIRPORT', right_on='AIRPORT_CODE', how='left')
df_merge_v1.rename(columns={'ORIGIN_AIRPORT':'ORIGIN_AC', #Origin Airport Code
'AIRPORT_NAME':'ORIGIN_AIRPORT', #Origin Airport Name
'DESTINATION_AIRPORT': 'DESTINATION_AC'}, inplace=True) #Dest Airport Code
df_merge_v1.drop(['AIRLINE','AIRPORT_CODE'], axis=1, inplace=True)
#Merge Airports and Flights dfs on Destination_Airport and Airport_Code
df_merge_v1 = pd.merge(df_merge_v1, df_airports, left_on='DESTINATION_AC', right_on='AIRPORT_CODE', how='left')
df_merge_v1.rename(columns={'AIRPORT_NAME':'DESTINATION_AIRPORT'}, inplace=True) #Dest Airport Name
df_merge_v1.drop('AIRPORT_CODE', axis=1, inplace=True)
print("Merged Dataframe Columns: \n", df_merge_v1.columns)
df_merge_v1[['ORIGIN_AIRPORT', 'ORIGIN_AC','DESTINATION_AIRPORT', 'DESTINATION_AC']]
```
## Save file
```
df_merge_v1.to_csv('data/flightsmerged.csv', index=False)
```
| github_jupyter |
```
import time
import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from xgboost import XGBClassifier
%matplotlib inline
plt.style.use('seaborn-dark-palette')
import warnings
warnings.filterwarnings('ignore')
file = glob.iglob('*.csv')
df = pd.read_csv(*file)
print(f'The dimension of the data is - {df.shape}')
df.head()
df.tail()
X = df.iloc[:, :-1].values
Y = df.iloc[:, -1].values
X
Y
print("Size of X: {}".format(X.shape))
print("Size of Y: {}".format(Y.shape))
X_train, X_test, Y_train, Y_test = train_test_split(X,
Y,
test_size=0.25,
random_state=0,
shuffle=True)
print("Size of X_train: {}".format(X_train.shape))
print("Size of X_test: {}".format(X_test.shape))
print("Size of Y_train: {}".format(Y_train.shape))
print("Size of Y_test: {}".format(Y_test.shape))
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
classifier = XGBClassifier()
classifier.fit(X_train, Y_train)
y_pred = classifier.predict(X_test)
y_pred
cm = confusion_matrix(Y_test, y_pred)
cm
acc = accuracy_score(Y_test, y_pred)
print(f"The accuracy in percentage - {acc*100}%")
report = classification_report(Y_test, y_pred)
print(report)
acc = cross_val_score(estimator = classifier,
X = X_train,
y = Y_train,
n_jobs = -1,
verbose = 0,
cv = 10)
print(f"Accuracy Score: {acc.mean()*100:.3f}%")
print(f"Standard Deviation: {acc.std()*100:.2f} %")
# start = time.time()
# parameters = [{'C': [0.25, 0.5, 0.75, 1],
# 'kernel': ['linear']},
# {'C': [0.25, 0.5, 0.75, 1],
# 'kernel': ['rbf'],
# 'gamma': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]}]
# grid_search = GridSearchCV(estimator = classifier,
# param_grid = parameters,
# scoring = 'accuracy',
# n_jobs = -1,
# cv = 10,
# verbose = 1
# )
# grid_search.fit(X_train, Y_train)
# best_accuracy = grid_search.best_score_
# best_parameters = grid_search.best_params_
# print(f"Accuracy Score: {best_accuracy*100:.3f}%")
# print(f"Best Parameters: {best_parameters}")
# end = time.time()
# print(f"Total Time Taken {end - start}")
# # Training Set
# figure = plt.figure(figsize = (10,10))
# x_set, y_set = X_train, Y_train
# X1, X2 = np.meshgrid(np.arange(start = x_set[:, 0].min() - 1,
# stop = x_set[:, 0].max() + 1,
# step = 0.01),
# np.arange(start = x_set[:, 1].min() - 1,
# stop = x_set[:, 1].max() + 1,
# step = 0.01))
# plt.contourf(X1,
# X2,
# classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
# alpha = 0.4,
# cmap = ListedColormap(('red', 'green')))
# for i, j in enumerate(np.unique(y_set)):
# plt.scatter(x_set[y_set == j, 0],
# x_set[y_set == j, 1],
# color = ListedColormap(('red', 'green'))(i),
# s = 15,
# marker = '*',
# label = j
# )
# plt.xlim(X1.min(), X1.max())
# plt.ylim(X2.min(), X2.max())
# plt.title('Kernel - SVM Classifier (Training Set)')
# plt.xlabel('Age')
# plt.ylabel('Estimated Salary')
# plt.legend()
# # Visuaizing the test case result
# figure = plt.figure(figsize = (10,10))
# x_set, y_set = X_test, Y_test
# X1, X2 = np.meshgrid(np.arange(start = x_set[:, 0].min() - 1,
# stop = x_set[:, 0].max() + 1,
# step = 0.01),
# np.arange(start = x_set[:, 1].min() - 1,
# stop = x_set[:, 1].max() + 1,
# step = 0.01))
# plt.contourf(X1,
# X2,
# classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
# cmap = ListedColormap(('red', 'green')),
# alpha = 0.4
# )
# for i, j in enumerate(np.unique(y_set)):
# plt.scatter(x_set[y_set == j, 0],
# x_set[y_set == j, 1 ],
# color = ListedColormap(('red', 'green'))(i),
# s = 15,
# label = j,
# marker = '^'
# )
# plt.xlim(X1.min(), X1.max())
# plt.ylim(X2.min(), X2.max())
# plt.title("Kernel SVM - Test Case")
# plt.xlabel('Age')
# plt.ylabel('Estimated Salary')
# plt.legend()
```
| github_jupyter |
<img src="http://landlab.github.io/assets/Landlab-logo.png"
style="float:left;width:150px;padding:0px">
<br />
# Reuse a published regional Landlab shallow landslide model to explore changes in forest cover at a subcatchment within the study area<br />
### You will explore how to reuse model code and data in a subregion within a larger region where the model was developed
* Load data from a regional Landlab landslide model (Strauch et al., 2018) developed for the North Cascades National Park, WA USA, published on HydroShare. <br />
* Define a geographic subset (Thunder Creek watershed) within the study region.
* Explore landslide probability sensitivity to fire by adjusting the cohesion parameter for Thunder Creek.
* Save results to a new HydroShare resource. <br />
The shallow landslide model you will is based on a spatially distributed Monte Carlo solution of the infinite slope stability model. Detailes of the model and the study site are described in Strauch et al. (2018). Please see the end of this Notebook for Acknowledgements and Citation information.
<br /> <img src="https://www.washington.edu/brand/files/2014/09/W-Logo_Purple_Hex.png" style="float:right;width:120px;padding:10px">
## To run this notebook:
Click in each shaded cell below and "shift + enter" to run each code block. Alternatively, you can run groups of cells by clicking "Cell" on the menu above and selecting your run options from the pull-down menu. This is also where you can clear outputs from previous runs.
If an error occurs, click on *Kernal* and *Restart and Clear Outputs* in the menu above.
## 1. Introduction
** 1.1 Infinite Slope Factor of Safety Equation **
This equation predicts the ratio of stabilizing to destabilizing forces on a hillslope plane. The properties are assumed to represent an infinte plane, neglecting the boundary conditions around the landslide location. When FS<1, the slope is instable.


C* can be calculated by the ratio of the sum of root cohesion, Cr, and soil cohesion, Cs, to the product of soil depth, density, and gravity.

Relative wetness is the ratio of depth of water subsurface flow above an impervious layer to the depth of soil.

R: recharge rate to water table (m/d)
T: soil transmissivity (m^2/d)
a: specific catchment area (m).
** 1.2 Monte Carlo solution of the FS equation **
Below the Monte Carlo approach used by Strauch et al. (2018) is illustrated in a figure. At each node of the model grid variables for soil and vegetation are generated from triangular distributions. Recharge to water table is obtained from hydrologic model simulations by selecting the largest daily value for each year. Probability of shallow landslide initiation is defined as the ratio of number of times FS<1 to the total sample size of FS calculations.

### Module overview for a Landlab Modeling Toolkit Landslide Introduction
This Jupyter Notebook runs the Landlab LandslideProbability component on a 30-m digital elevation model (DEM) for Thunder Creek, a subwaterhsed in the larger study domain developed by Strauch et al. (2018), using _data driven spatial_ recharge distribution as described in the paper (https://www.earth-surf-dynam.net/6/1/2018/).
To run a landslide demonstration using the paper data and approach we will:<br >
1) Import data from North Cascades National Park (NOCA: study area)<br />
2) Review data needed as input for the landslide model<br />
3) Create a RasterModelGrid based on a 30-m DEM - subset to Thunder Creek watershed<br />
4) Assign data fields used to calculate landslide probability<br />
5) Specify recharge option as _data driven spatial_ and access Python dictionaries to generate recharge distributions<br />
6) Set Number of iterations to run Monte Carlo simulation<br />
7) Run Landlab LandslideProbability component<br />
8) Run the model again to simulate post-fire conditions,<br />
9) Display and visualize results of stability analysis<br />
10) Save Notebook and Results back to HydroShare<br />
<br />
The estimated time to run this Notebook is 30-60 minutes with 20 minutes of computation time.
## 2.0 Data Science & Cyberinfrastructure Methods
Update or add libraries that are not installed on the CUAHSI JupyterHub server. At the time of publication, the Landlab library was dependent on a different numpy library than installed in the software environment. The code block below was added as an example of how to update a library to a new version and restart the notebook kernel. If this update is not needed, comment with a <#>. If other updates are needed, add them below.
```
# comment this out if this update is not needed
!conda update -y numpy
from IPython.display import display_html
display_html("<script>Jupyter.notebook.kernel.restart()</script>",raw=True)
```
### 2.1 Data Science Methods
### 2.1.1 Import Landlab components, functions for importing data, plotting tools, and HydroShare utilities
To run this notebook, we must import several libraries.
The hs_utils library provides functions for interacting with HydroShare, including resource querying, dowloading. and creation. Additional libraries support the functions of Landlab. The CUAHSI JupyterHub server provides many Python packages and libraries, but to add additional libraries to your personal user space, use the cell below. To request an Installation to the server, visit https://github.com/hydroshare/hydroshare-jupyterhub, create a New Issue, and add the label 'Installation Request'.
### 2.1.2 Import Python libraries
```
#Import standard Python utilities for calculating and plotting
import six
import os
import matplotlib as mpl
mpl.use('agg')
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import pickle as pickle
from datetime import datetime, timedelta
import geopandas as gpd
# Import Landlab libraries
from landlab.components import LandslideProbability
from landlab import imshow_grid_at_node
from landlab.io import read_esri_ascii
from landlab.io import write_esri_ascii
from landlab.plot import imshow_grid
from landlab import CORE_NODE, CLOSED_BOUNDARY
#Import utilities for importing and exporting to HydroShare
from hstools import hydroshare
# Import general tools
import time
from collections import defaultdict
st = time.time()
%matplotlib inline
```
### 2.1.3 Connection with HydroShare
After importing libraires, we now establish a secure connection with HydroShare by instantiating the hydroshare class that is defined within hs_utils. In addition to connecting with HydroShare, this command also sets and prints environment variables for several parameters that will be useful for saving work back to HydroShare.
```
hs=hydroshare.hydroshare()
homedir=os.getcwd()
#homedir = os.path.join('/home/jovyan/work/notebooks/data', str(homeresid),str(homeresid),'data/contents/')
#print('This is a basic Unix folder structure.')
#print('Data will be loaded from and saved to:'+homedir)
```
If you are curious about where the data is being downloaded, click on the Jupyter Notebook dashboard icon in upper rigth corner to see a File System view. The homedir directory location printed above is where you can find the data and contents you will download to a HydroShare JupyterHub server. At the end of this work session, you can migrate this data to the HydroShare iRods server as a Generic Resource.
### 2.2. Cyberinfrastructure methods
Strauch et al (2018) pre-processed the data for the North Cascades National Park Complex case study and is on HydroShare as [Regional landslide hazard using Landlab - NOCA Data](https://www.hydroshare.org/resource/a5b52c0e1493401a815f4e77b09d352b/). Here the first task is find this resource on Hydrohsare. We will click on the link to see the published data repository on HydroShare and collect the resource ID of the data. The resource ID can be found in the "How to cite" box, and it will be the series of numbers and letters following "hs.". Here's the copied resource: http://dx.doi.org/10.4211/hs.a5b52c0e1493401a815f4e77b09d352b citation. Now we copy this ID and introduce it as "Data_ResourceID=" in the code below.
#### 2.2.1 Set DEM data download variable name
To learn more about this data visit [Regional landslide hazard using Landlab - NOCA Data](https://www.hydroshare.org/resource/a5b52c0e1493401a815f4e77b09d352b/)
```
Data_ResourceID='a5b52c0e1493401a815f4e77b09d352b'
```
#### 2.2.2 Download Data
We will execute the next cell to download data from HydroShare iRods database to your personal user space - this may take a few minutes.
```
hs.getResourceFromHydroShare(Data_ResourceID)
data_folder = os.path.join(os.getcwd(), hs.getContentPath(Data_ResourceID), 'Data files')
print('This HydroShare resource has been downloaded to: %s' % data_folder)
```
### 2.3. Review data needed as input for the landslide model
This section loads metadata associated with the Landlab component.
To view the code source for this component, visit [Landlab on Github](https://github.com/landlab/) or [Download the landslide_probability.py python file](https://github.com/landlab/landlab/blob/master/landlab/components/landslides/landslide_probability.py)
Check the list of data inputs that the component needs.
```
sorted(LandslideProbability.input_var_names)
```
Review the details of what each variable represents.
```
LandslideProbability._var_doc
```
Check the units of each variable.
```
LandslideProbability._var_units
```
Now we will establish a RasterModelGrid based on a DEM for assigning our variables to.
Nodes are the center point of grid cells or pixels that are 30 m by 30 m in this example.
### 2.4. Create a watershed subset using a regional RasterModelGrid
The shapefile that you will download is a GIS point file with a table containing a grid_code for each 30m DEM cell in the Thunder Creek watershed. We import this table to generate a list of locations we want to keep "Open" as active nodes in this Landlab application for Thunder Creek.
Here we will establish the watershed domain for our modeling study. In step 2.2.2, we have downloaded the DEM of the entire NOCA study region of Strauch et al. (2018) as well as other input data. This includes NoData resource that was created using a mask of Thunder Creek will be downladed and used to set no data nodes as inactive nodes (e.g., -9999). This step will establish boundary conditions. In your final run when you output modeled probability of landslide initiation, you will notice gaps in the model results at some ridge tops and peaks. This results from excluding glaciated areas and bedrock. <br />
This might take a few minutes as the park is large (2,757 km2).
This shapefile resource was uploaded to HydroShare to generate an interoperable Notebook. To learn more about this data visit [Thunder Creek Landlab Node ID Point Shapefile
](https://www.hydroshare.org/resource/8bf8de77227c4dcba4816dbe15e55687/)
#### 2.4.1 Get shapefile of ThunderCreek watershed
The shapefile table contains Landlab Gridcode value and Albers Conical X and Y values, spatial projection is WGS84.
```
Data_ResourceID='8bf8de77227c4dcba4816dbe15e55687'
hs.getResourceFromHydroShare(Data_ResourceID)
Node_path = os.path.join(hs.getContentPath(Data_ResourceID), 'Thunder_node_id_WGS.shp')
NodeID_shpfile=gpd.GeoDataFrame.from_file(Node_path)
```
Print a sample portion (bottom) of the table
* Col 1 = FID or POINTID
* Col 2 = landlab grid code for NOCA region
* Col 3 Point_X - the Alber Conical Latitude (original dataset)
* Col 4 Point_Y - the Alber Conical Longitude (original dataset)
* Geometry of lat/lon in WGS coordinate system of this GIS point file for launching in HydroShareGIS App
```
NodeID_shpfile.tail()
```
Make an array using Grid_Code column to develop a node based mask. No output from this command.
```
filtercriteria = np.array(NodeID_shpfile.GRID_CODE)
```
#### Load spatial landslide model inputs from ASCII textfile (ArcGIS raster conversion) into Landlab grid
Create Landlab RasterModelGrid using DEM grid with elevation - this takes approximately 60 sec for the North Cascades National Park (NOCA).
```
(grid, z) = read_esri_ascii(data_folder+'/elevation.txt',name='topographic__elevation')
grid.at_node.keys()
```
The original NOCA dem dataset has nodes closed outside of NOCA.
All nodes insides of NOCA are open. To close all all nodes except those inside Thunder Creek:
* Close all nodes in the landlab RasterModelGrid.
* Open the Landlab RasterModelGrid only within the Thunder Creek watershed.
* Assign Core Nodes (these are the only nodes where computation will be executed).
Plot the elevation grid of NOCA
```
imshow_grid(grid, 'topographic__elevation', limits=(0, 3000))
```
Plot where the watershed is witin NOCA and zoom in.
```
grid.status_at_node[grid.nodes.flatten()] = CLOSED_BOUNDARY
grid.status_at_node[filtercriteria] = CORE_NODE
# In case the DEM has no data values inside the subset area, set boundary conditions closed.
grid.set_nodata_nodes_to_closed(grid.at_node['topographic__elevation'], -9999)
fig = plt.figure('Limit NOCA computational nodes to Thunder Creek extent')
xticks = np.arange(-0.1, 0.8, 0.4)
ax1 = fig.add_subplot(221)
ax1.xaxis.set_visible(False)
imshow_grid(grid, 'topographic__elevation', limits=(0, 3000),plot_name='NOCA extent',
allow_colorbar=False,color_for_closed='white')
ax2 = fig.add_subplot(222)
ax2.xaxis.set_visible(False)
imshow_grid(grid, 'topographic__elevation', limits=(0, 3000),plot_name='Thunder Creek extent',color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
```
Confirm the size of the grid, nodes located every 30 m.
```
grid.number_of_nodes
```
Confirm the size of the core nodes where we'll run our model, a subset of the nodes for the watershed.
```
grid.number_of_core_nodes
```
### 2.5. Attach data (e.g., soil and vegetation variables) to the Landlab rastermodelgrid
This will be used to calculate shallow landslide probability and set boundary conditions. THe data we attach in this step was downloaded in Step 2.2.2.
#### For each input below
1. Load data from ascii text file
2. Add this data as node variable to the Thunder Creek grid
3. Set boundary conditions
For the entire NOCA extent, this takes ~60 sec to load each file using the CUAHSI JupyterHub server.
#### 2.5.1 Load slope
```
(grid1, slope) = read_esri_ascii(data_folder+'/slope_tang17d.txt')
grid.add_field('node', 'topographic__slope', slope)
grid.set_nodata_nodes_to_closed(grid.at_node['topographic__slope'], -9999)
grid.set_nodata_nodes_to_closed(grid.at_node['topographic__slope'], 0.0)
print(np.max(grid.at_node['topographic__slope'][grid.core_nodes]))
print(np.min(grid.at_node['topographic__slope'][grid.core_nodes]))
```
#### 2.5.2 Load contributing area
```
(grid1, ca) = read_esri_ascii(data_folder+'/cont_area.txt')
grid.add_field('node', 'topographic__specific_contributing_area', ca)
grid.set_nodata_nodes_to_closed(grid.at_node['topographic__specific_contributing_area'], -9999)
```
#### 2.5.3 Load transmissivity
```
(grid1, T) = read_esri_ascii(data_folder+'/transmis.txt')
grid.add_field('node', 'soil__transmissivity', T)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__transmissivity'], -9999)
```
#### 2.5.4 Load cohesion (mode, min, and max)
This takes ~3 minutes because 3 cohesion fields are provided to create more flexibility in how cohesion is distributed on the landscape with different vegetation.
```
(grid1, C) = read_esri_ascii(data_folder+'/cohesion_mode.txt')
C[C == 0.0] = 1.0 # ensure minimum is >0 Pa for use in distributions generation
grid.add_field('node', 'soil__mode_total_cohesion', C)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__mode_total_cohesion'], -9999)
(grid1, C_min) = read_esri_ascii(data_folder+'/cohesion_min.txt')
grid.add_field('node', 'soil__minimum_total_cohesion', C_min)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__minimum_total_cohesion'], -9999)
(grid1, C_max) = read_esri_ascii(data_folder+'/cohesion_max.txt')
grid.add_field('node', 'soil__maximum_total_cohesion', C_max)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__maximum_total_cohesion'], -9999)
```
#### 2.5.5 Load internal angle of friction
```
(grid1, phi) = read_esri_ascii(data_folder+'/frict_angle.txt')
grid.add_field('node', 'soil__internal_friction_angle', phi)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__internal_friction_angle'], -9999)
```
#### 2.5.6 Set soil density values
In this example, we assign all nodes a constant value.
```
grid['node']['soil__density'] = 2000*np.ones(grid.number_of_nodes)
```
#### 2.5.7 Load soil thickness
```
(grid1, hs) = read_esri_ascii(data_folder+'/soil_depth.txt')
grid.add_field('node', 'soil__thickness', hs)
grid.set_nodata_nodes_to_closed(grid.at_node['soil__thickness'], -9999)
```
#### 2.5.8 Load observed landslide inventory. Class 1-5 are landslides, 8 is no landslide mapped for later plotting
```
(grid1, slides) = read_esri_ascii(data_folder+'/landslide_type.txt')
grid.add_field('node', 'landslides', slides)
```
### 2.6. Specify recharge option as _data driven spatial_ and access Python dictionaries to generate recharge distributions
Recharge in this model represents the annual maximum recharge in mm/day generated within the upslope contributing area of each model element. This corresponds to the wettest conditions expected annually, which would provide the highest pore-water pressure in a year. Details of this approach can be found in Strauch et al. (2018).
#### 2.6.1 Select recharge method from the component
```
distribution = 'data_driven_spatial'
```
#### 2.6.2 Load pre-processed routed flows dictionaries
These contain HSD_id and fractional drainage at each node and recharge dictionaries. HSD is the Hydrologic Source Domain, which is the VIC hydrologic model data in this case study at ~5x6 km2 grid size. The 'pickle' utility loads existing dictionaries.
```
# dict of node id (key) and HSD_ids (values)
HSD_id_dict = pickle.load(open(data_folder+'/dict_uniq_ids.p', 'rb'),encoding='latin1')
# dict of node id (key) and fractions (values)
fract_dict = pickle.load(open(data_folder+'/dict_coeff.p', 'rb'),encoding='latin1')
# dict of HSD id (key) with arrays of recharge (values)
HSD_dict = pickle.load(open(data_folder+'/HSD_dict.p', 'rb'),encoding='latin1')
```
#### 2.6.3 Combine dictionaries into __ordered__ parameters
This sequence of parameters is required for _data driven spatial_ distribution in the component. Recharge is this model is unique to each node represented by an array.
```
HSD_inputs = [HSD_dict, HSD_id_dict, fract_dict]
```
### 2.7. Set Number of iterations to run Monte Carlo simulation
The landslide component employes the infinite slope model to calculate factor-of-safety index values using a Monte Carlo simulation, which randomly selects input values from parameter distributions. You can specify the number of Monte Carlo samples, but the default is 250. The larger the Monte Carlos sample size, the longer the program runs, but the more precise the probability of failure results become. Strauch et al. (2018) sampled 3,000 times for each parameter in each model grid.
```
iterations = 250
```
## 3. Results
### 3.1. Run the Landlab LandslideProbability Component in Thunder Creek for pre-fire conditions
#### 3.1.1 Initialize Pre-fire
To run the landslide model, we first instantiate the LandslideProbability component with the above parameters, as well as the grid and number of samples we specified before. Instantiation creates an instance of a class called LS_prob.
No outputs are generated by this command as it is setting up the recharge and instantiating the component.
```
LS_prob = LandslideProbability(grid,
number_of_iterations=iterations,
groudwater__recharge_distribution=distribution,
groudwater__recharge_HSD_inputs=HSD_inputs)
```
#### 3.1.2 Run the Pre-fire model
Once the component has been instantiated, we generate outputs from running the component by calling the component's 'calculate_landslide_probability' method using the class instance (e.g., LS_prob). The cell below runs the model; in the following section we will assessing the results. These calculations will take a few minutes given the size of the modeling domain represented by core nodes.
```
LS_prob.calculate_landslide_probability()
print('Landslide probability successfully calculated')
```
What is the maximum probabilty of failure we found?
```
np.max(grid.at_node['landslide__probability_of_failure'][grid.core_nodes])
```
The outputs of landslide model simulation are:
```
sorted(LS_prob.output_var_names)
```
This simulation generates a probability value for each core node.
```
grid.at_node['landslide__probability_of_failure']
```
This simulation generates a probability of saturation value for each core node as well.
```
grid.at_node['soil__probability_of_saturation']
```
-9999 means there is no data for that cell, it is a closed node.
### 3.2. Run the Landlab LandslideProbability Component in Thunder Creek for fire conditions
Make a 'grid_fire' version of 'grid' for which we will give post-fire cohesion parameter values 30% of the original cohesion. This is a crude estimation of the lowest root cohesion after tree removal based on a combined decay and regrowth model (e.g., Sidle, 1992).
Sidle, R. C. (1992), A theoretical model of the effects of timber harvesting
on slope stability, Water Resour. Res., 28(7), 1897–1910.
#### 3.2.1 Make a Post-fire copy of the grid
```
import copy
grid_fire=copy.deepcopy(grid)
grid_fire.at_node['soil__mode_total_cohesion']=grid.at_node['soil__mode_total_cohesion']*0.3
grid_fire.at_node['soil__minimum_total_cohesion']=grid.at_node['soil__minimum_total_cohesion']*0.3
grid_fire.at_node['soil__maximum_total_cohesion']=grid.at_node['soil__maximum_total_cohesion']*0.3
```
#### 3.2.2 Change the Post-fire cohesion values
```
print("this is the highest mode value for coehsion before fire across the domain")
print(np.max(grid.at_node['soil__mode_total_cohesion'][grid.core_nodes]))
print("this is the lowest mode value for coehsion before fire across the domain")
print(np.min(grid.at_node['soil__mode_total_cohesion'][grid.core_nodes]))
print("this is the highest mode value for coehsion after fire across the domain")
print(np.max(grid_fire.at_node['soil__mode_total_cohesion'][grid.core_nodes]))
print("this is the lowest mode value for coehsion after fire across the domain")
print(np.min(grid_fire.at_node['soil__mode_total_cohesion'][grid.core_nodes]))
```
#### 3.2.3 Initialize the Post-fire model
Now we'll run the landslide component with the adjusted cohesion, everything else kept constant.
```
LS_probFire = LandslideProbability(grid_fire,number_of_iterations=iterations,
groudwater__recharge_distribution=distribution,
groudwater__recharge_HSD_inputs=HSD_inputs)
print('Post-fire cohesion successfully instantiated')
```
#### 3.2.4 Run the Post-fire model
```
LS_probFire.calculate_landslide_probability()
print('Landslide probability successfully calculated')
np.max(grid_fire.at_node['landslide__probability_of_failure'][grid.core_nodes])
```
The outputs of landslide model simulation are:
```
sorted(LS_probFire.output_var_names)
```
This simulation generates a probability value for each core node.
```
grid_fire.at_node['landslide__probability_of_failure']
```
### 3.3. Display and visualize results of stability analysis
Set plotting parameters
```
mpl.rcParams['xtick.labelsize'] = 15
mpl.rcParams['ytick.labelsize'] = 15
mpl.rcParams['lines.linewidth'] = 1
mpl.rcParams['axes.labelsize'] = 18
mpl.rcParams['legend.fontsize'] = 15
```
Plot elevation
```
plt.figure('Elevations from the DEM [m]')
imshow_grid_at_node(grid, 'topographic__elevation', cmap='terrain',
grid_units=('coordinates', 'coordinates'),
shrink=0.75, var_name='Elevation', var_units='m',color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
plt.savefig('NOCA_elevation.png')
```
Excluded areas from the analysis are shown in white, including outside the park and inside the park areas that are water bodies, snow, glaciers, wetlands, exposed bedrock, and slopes <= 17 degrees.
Plot slope overlaid with mapped landslide types. Takes about a few minutes.
```
plt.figure('Landslides')
ls_mask1 = grid.at_node['landslides'] != 1.0
ls_mask2 = grid.at_node['landslides'] != 2.0
ls_mask3 = grid.at_node['landslides'] != 3.0
ls_mask4 = grid.at_node['landslides'] != 4.0
overlay_landslide1 = np.ma.array(grid.at_node['landslides'], mask=ls_mask1)
overlay_landslide2 = np.ma.array(grid.at_node['landslides'], mask=ls_mask2)
overlay_landslide3 = np.ma.array(grid.at_node['landslides'], mask=ls_mask3)
overlay_landslide4 = np.ma.array(grid.at_node['landslides'], mask=ls_mask4)
imshow_grid_at_node(grid, 'topographic__slope', cmap='pink',
grid_units=('coordinates', 'coordinates'), vmax=2.,
shrink=0.75, var_name='Slope', var_units='m/m',color_for_closed='white',)
imshow_grid_at_node(grid, overlay_landslide1, color_for_closed='None',
allow_colorbar=False, cmap='cool')
imshow_grid_at_node(grid, overlay_landslide2, color_for_closed='None',
allow_colorbar=False, cmap='autumn')
imshow_grid_at_node(grid, overlay_landslide3, color_for_closed='None',
allow_colorbar=False, cmap='winter')
imshow_grid_at_node(grid, overlay_landslide4, color_for_closed='None',
allow_colorbar=False,cmap='summer')
#plt.savefig('NOCA_Landslides_on_Slope.png')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
```
Legend to mapped landslides: blue - debris avalanches, cyan - falls/topples, red - debris torrents, and green - slumps/creeps
Plot of soil depth (m)
```
plt.figure('Soil Thickness')
imshow_grid_at_node(grid, 'soil__thickness', cmap='copper_r',
grid_units=('coordinates', 'coordinates'), shrink=0.75,
var_name='Soil Thickness', var_units='m', color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
#plt.savefig('NOCA_SoilDepth.png')
```
Plot probability of saturation
```
fig = plt.figure('Probability of Saturation')
xticks = np.arange(-0.1, 0.8, 0.4)
ax1 = fig.add_subplot(221)
ax1.xaxis.set_visible(True)
imshow_grid(grid, 'soil__probability_of_saturation',cmap='YlGnBu',
limits=((0), (1)),plot_name='Pre-Fire',
allow_colorbar=False,grid_units=('coordinates', 'coordinates'), color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
ax2 = fig.add_subplot(222)
ax2.xaxis.set_visible(True)
ax2.yaxis.set_visible(False)
imshow_grid(grid_fire, 'soil__probability_of_saturation',cmap='YlGnBu',
limits=((0), (1)),plot_name='Post-Fire',grid_units=('coordinates', 'coordinates'), color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
```
This map shows the probability of saturation as high throughout much of the area because we modeled the annual maximum recharge, which is esssentially the worst case conditions that might lead to instability.
Plot probability of failure; Compare this with the elevation and slope maps.
```
fig = plt.figure('Probability of Failure')
xticks = np.arange(-0.1, 0.8, 0.4)
ax1 = fig.add_subplot(221)
#ax1.xaxis.set_visible(False)
imshow_grid(grid, 'landslide__probability_of_failure',cmap='OrRd',
limits=((0), (1)),plot_name='Pre-fire',
allow_colorbar=False,grid_units=('coordinates', 'coordinates'), color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
ax2 = fig.add_subplot(222)
ax2.yaxis.set_visible(False)
imshow_grid(grid_fire, 'landslide__probability_of_failure',cmap='OrRd',
limits=((0), (1)),plot_name='Post-fire',grid_units=('coordinates', 'coordinates'), color_for_closed='white')
plt.xlim(25000, 55000)
plt.ylim(30000, 55000)
#plt.savefig('Probability_of_Failure_Original_Fire.png')
```
The map of probability of failure shows higher probabilities in a post-fire scenario with reduced cohesion.
We can compare the range of probabilities before and after the simulated fire using a cumulative distribution plot.
```
X1 = np.sort(np.array(grid.at_node['landslide__probability_of_failure'][grid.core_nodes]))
Y1 = np.arange(1, len(X1)+1)/len(X1)
X2 = np.sort(np.array(grid_fire.at_node['landslide__probability_of_failure'][grid.core_nodes]))
Y2 = np.arange(1, len(X2)+1)/len(X2)
fig = plt.figure('Empirical CDF of Probability of Failure')
plt.plot(X1,Y1, marker=' ', color='black', linewidth=2, label='Pre-fire')
plt.plot(X2,Y2, marker=' ', color='red', linewidth=2, linestyle='dashed', label='Post-fire')
plt.xlabel('Probability of Failure')
plt.ylabel('Fraction of Area')
plt.legend()
```
We can interpret this figure as a higher fraction of the landscape has a higher probability of failure (red) after fire than before fire (black).
To review the fields assigned to the grid, simply execute the following command.
```
grid.at_node
```
Export data from model run: FS probability, mean Reletive wetness, probability of saturation
```
import pandas as pd
core_nodes = grid.core_nodes
data_extracted = {'Prob_fail_std': np.array(
grid.at_node['landslide__probability_of_failure'][grid.core_nodes]),
'mean_RW_std': np.array(grid.at_node['soil__mean_relative_wetness']
[grid.core_nodes]),'prob_sat_std': np.array(
grid.at_node['soil__probability_of_saturation'][grid.core_nodes]),
'Prob_fail_fire': np.array(
grid_fire.at_node['landslide__probability_of_failure'][grid_fire.core_nodes]),
'mean_RW_fire': np.array(grid_fire.at_node['soil__mean_relative_wetness']
[grid_fire.core_nodes]),'prob_sat_fire': np.array(
grid_fire.at_node['soil__probability_of_saturation'][grid_fire.core_nodes])}
headers = ['Prob_fail_std','mean_RW_std','prob_sat_std','Prob_fail_fire','mean_RW_fire','prob_sat_fire']
df = pd.DataFrame(data_extracted, index=core_nodes, columns=(headers))
df.to_csv('Landslide_std_fire.csv')
```
Make ascii files for raster creation in GIS
```
write_esri_ascii('prob_fail_std.txt',grid,names='landslide__probability_of_failure')
write_esri_ascii('mean_RW_std.txt',grid,names='soil__mean_relative_wetness')
write_esri_ascii('prob_sat_std.txt',grid,names='soil__probability_of_saturation')
write_esri_ascii('prob_fail_fire.txt',grid_fire,names='landslide__probability_of_failure')
write_esri_ascii('mean_RW_fire.txt',grid_fire,names='soil__mean_relative_wetness')
write_esri_ascii('prob_sat_fire.txt',grid_fire,names='soil__probability_of_saturation')
```
How long did the code above take to run?
```
print('Elapsed time is %3.2f seconds' % (time.time() - st))
```
## 4.0. Save the results back into HydroShare
<a name="creation"></a>
Using the `hs_utils` library, the results of the Geoprocessing steps above can be saved back into HydroShare. First, define all of the required metadata for resource creation, i.e. *title*, *abstract*, *keywords*, *content files*. In addition, we must define the type of resource that will be created, in this case *genericresource*.
***Note:*** Make sure you save the notebook at this point, so that all notebook changes will be saved into the new HydroShare resource.
***Option A*** : define the resource from which this "NEW" content has been derived. This is one method for tracking resource provenance.
Create list of files to save to HydroShare. Verify location and names.
```
ThisNotebook='replicate_landslide_model_for_fire.ipynb' #check name for consistency
files=[os.path.join(homedir, ThisNotebook),
os.path.join(homedir, 'Landslide_std_fire.csv'),
os.path.join(homedir, 'prob_fail_std.txt'),
os.path.join(homedir, 'mean_RW_std.txt'),
os.path.join(homedir, 'prob_sat_std.txt'),
os.path.join(homedir, 'prob_fail_fire.txt'),
os.path.join(homedir, 'mean_RW_fire.txt'),
os.path.join(homedir, 'prob_sat_fire.txt')]
for f in files:
if not os.path.exists(f):
print('\n[ERROR] invalid path: %s' % f)
help(hs)
# Save the results back to HydroShare
# title for the new resource
title = 'Landslide Model run with Monte Carlo from NOCA Observatory - Thunder Creek with Fire'
# abstract for the new resource
abstract = 'This a reproducible demonstration of the landslide modeling results from eSurf paper: Strauch et al. (2018) '
# keywords for the new resource
keywords = ['landslide', 'climate', 'VIC','saturation','relative wetness','fire','Geohackweek']
# Hydroshare resource type
rtype = 'compositeresource'
# create the new resource
resource_id = hs.createHydroShareResource(abstract,
title,
keywords=keywords,
resource_type=rtype,
content_files=files,
public=False)
```
### Acknowledgements
[](https://zenodo.org/badge/latestdoi/187289993)
This Notebooks serves as content for:
Bandaragoda, C. J., A. Castronova, E. Istanbulluoglu, R. Strauch, S. S. Nudurupati, J. Phuong, J. M. Adams, et al. “Enabling Collaborative Numerical Modeling in Earth Sciences Using Knowledge Infrastructure.” Environmental Modelling & Software, April 24, 2019. https://doi.org/10.1016/j.envsoft.2019.03.020.
and
Bandaragoda, C., A. M. Castronova, J. Phuong, E. Istanbulluoglu, S. S. Nudurupati, R. Strauch, N. Lyons, K. Barnhart (2019). Enabling Collaborative Numerical Modeling in Earth Sciences using Knowledge Infrastructure: Landlab Notebooks, HydroShare, http://www.hydroshare.org/resource/fdc3a06e6ad842abacfa5b896df73a76
This notebook was developed from code written by Ronda Strauch as part of her Ph.D. disseration at the University of Washington.
Use or citation of this notebook should also reference:
Strauch R., Istanbulluoglu E., Nudurupati S.S., Bandaragoda C., Gasparini N.M., and G.E. Tucker (2018). A hydro-climatological approach to predicting regional landslide probability using Landlab. Earth Surf. Dynam., 6, 1–26.
https://www.earth-surf-dynam.net/6/1/2018/
| github_jupyter |
## Time-series based analysis of Earthquake Risk Factors
### Part 4: Performing linear regression
Importing the required libraries:
```
import requests
import json
import pandas as pd
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler, PolynomialFeatures
from sklearn.model_selection import train_test_split, cross_val_score
import datetime as dt
import seaborn as sns
import pickle
```
### Part 2.1: Prepping Loma Prieta EQ's:
```
file_path = "../datasets/" + "eq_loma_prieta" + "_clean" + ".csv"
df_lp = pd.read_csv(file_path)
df_lp.drop(columns = "Unnamed: 0", inplace = True)
df_lp["time"] = pd.to_datetime(df_lp["time"])
df_lp.sort_values(by = "time", inplace=True)
df_lp.reset_index(drop = True, inplace=True)
df_lp.drop(columns = ["index", "place", "status", "tsunami", "net", "nst", "type"], inplace=True)
df_lp.head()
```
## Finding the main failure time:
```
# Source: https://stackoverflow.com/questions/43601868/converting-an-array-datatime-datetime-to-float
epoch = dt.datetime(1970, 1, 1)
time = []
for t in [(d - epoch).total_seconds() for d in df_lp["time"]]:
time.append(float('%.6f' % t))
(float(time[0]))
df_lp["time_seconds"] = time
failure_event = df_lp[df_lp["mag"] == df_lp["mag"].max()]
df_lp["time_to_failure_sec"] = np.array(failure_event["time_seconds"]) - np.array(df_lp["time_seconds"])
df_lp.drop(columns=["time", "time_seconds"], inplace=True)
df_lp.head()
```
Checking if failure is correctly zero:
```
failure_event = df_lp[df_lp["mag"] == df_lp["mag"].max()]
failure_event
df_lp.head(2)
ax = plt.gca()
df_lp.plot(kind="scatter", x="longitude", y="latitude",
s=df_lp['mag']/0.05, label="EQ",
c=df_lp.index, cmap=plt.get_cmap("jet"),
colorbar=True, alpha=0.4, figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout()
plt.plot(df_lp["time_to_failure_sec"])
```
# Building the model:
```
df_corr = df_lp.corr()
#plt.figure(figsize=(6,6))
plt.figure(figsize=(8,6))
sns.heatmap(df_corr[['time_to_failure_sec']].sort_values(by=['time_to_failure_sec'],ascending=False),
vmin=-1,
cmap='coolwarm',
annot=True);
plt.savefig("../plots/Loma_prieta_EDA_corr_plots.png")
```
# LR model
### LINE Assumptions
The assumptions necessary to conduct a proper linear regression are easily remembered by the "LINE" acronym:
* L - Linearity: there is a linear relationship between x and y (fix: apply non-linear transformation)
* I - Independence of residuals (usually assumed)
* N - Normality: residuals are normally distributed (fix: log y variable)
* E - Equality of variance: residuals have constant variance (fix: check outliers, non-linear transformation)
```
df = df_lp.copy()
df.info()
# Step 3: Instantiate the model
lr = LinearRegression()
# Step 1: Assemble our X and y variables
# We need an X matrix that is n-by-p (in this case, p = 1)
X = df_lp[["mag", "sig", "depth", "longitude", "latitude"]]
# We need a y vector that is length n
y = df_lp["time_to_failure_sec"]
lr.fit(X, y)
y_pred = lr.predict(X)
plt.scatter(y_pred, y)
# Creat residuals
resids = y - y_pred
# N assumption:
plt.hist(resids, bins=50);
# L and E assumption
# our actual residuals
plt.scatter(y_pred, resids)
plt.axhline(0, color='orange');
lr.score(X, y)
# Via sklearn.metrics
from sklearn import metrics
metrics.mean_absolute_error(y, y_pred)
import statsmodels.api as sm
X_sm = sm.add_constant(X)
ols = sm.OLS(y, X_sm).fit()
ols.summary()
```
## The Linear Regression was not very good, now, trying the clustering idea.
# Clustering the AE events:
```
ax = plt.gca()
df_lp_eq.plot(kind="scatter", x="depth", y="longitude",
s=df_lp_eq['mag']/0.05, label="EQ",
c=df_lp_eq.index, cmap=plt.get_cmap("jet"),
colorbar=True, alpha=0.4, figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout();
ax = plt.gca()
df_lp_eq.plot(kind="scatter", x="longitude", y="latitude",
s=df_lp_eq['mag']/0.05, label="EQ",
c=df_lp_eq.index, cmap=plt.get_cmap("jet"),
colorbar=True, alpha=0.4, figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout()
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import silhouette_score
from sklearn.datasets import load_iris, load_wine
from sklearn.cluster import DBSCAN
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
```
# 2D clusgering within long-lat : DBSCAN
```
df = df_lp_eq[["longitude", "latitude"]]
df.head()
```
Standardizing the data:
```
ss = StandardScaler()
X_scaled = ss.fit_transform(df)
```
Instantitating and fitting a DBSCAN:
```
dbscan = DBSCAN()
dbscan.fit(X_scaled);
```
model evaluation:
```
silhouette_score(X_scaled, dbscan.labels_)
df['cluster'] = dbscan.labels_
plt.figure(figsize = (20, 10))
sns.pairplot(df, hue='cluster')
df.head()
ax = plt.gca()
df.plot(kind="scatter", x="longitude", y="latitude",
s=[10] * len(df),
label="EQ",
c=df["cluster"],
cmap=plt.get_cmap("jet"),
colorbar=True,
alpha=0.4,
figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout()
```
# 3D clustering with long-lat-depth
```
df = df_lp_eq[["longitude", "latitude", "depth"]]
df.head()
```
Standardizing the data:
```
ss = StandardScaler()
X_scaled = ss.fit_transform(df)
```
Instantitating and fitting a DBSCAN:
```
dbscan = DBSCAN()
dbscan.fit(X_scaled);
```
model evaluation:
```
silhouette_score(X_scaled, dbscan.labels_)
df['cluster'] = dbscan.labels_
plt.figure(figsize = (20, 10))
sns.pairplot(df, hue='cluster')
df.head()
ax = plt.gca()
df.plot(kind="scatter", x="longitude", y="latitude",
s=[10] * len(df),
label="EQ",
c=df["cluster"],
cmap=plt.get_cmap("jet"),
colorbar=True,
alpha=0.4,
figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout()
ax = plt.gca()
df.plot(kind="scatter", x="depth", y="latitude",
s=[10] * len(df),
label="EQ",
c=df["cluster"],
cmap=plt.get_cmap("jet"),
colorbar=True,
alpha=0.4,
figsize=(10,7), ax = ax
)
plt.legend()
plt.tight_layout()
print("Hello World!")
```
| github_jupyter |
```
%matplotlib inline
```
Analytic 300-hPa Trough
=======================
```
import matplotlib.pyplot as plt
import metpy.calc as mpcalc
from metpy.units import units
import numpy as np
```
Below are three definitions to create an analytic 300-hPa trough roughly
based on the Sanders Analytic Model with modified coefficients to create
different style waves.
```
def single_300hPa_trough(parameter='hght'):
""" Single trough with heights and Temperatures based on Sanders Analytic Model
"""
X = np.linspace(.25, .75, 101)
Y = np.linspace(.25, .75, 101)
x, y = np.meshgrid(X, Y)
p = 4
q = 2
if parameter == 'hght':
return (9240 + 100 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 200 * np.cos(y * np.pi) + 300 * y * np.cos(x * np.pi + np.pi / 2))
elif parameter == 'temp':
return (-50 + 2 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 2 * np.cos(y * np.pi) + 0.5 * y * np.cos(x * np.pi + np.pi / 2))
def lifting_300hPa_trough(parameter='hght'):
""" Lifting trough with heights and Temperatures based on Sanders Analytic Model
"""
X = np.linspace(.25, .75, 101)
Y = np.linspace(.25, .75, 101)
x, y = np.meshgrid(X, Y)
p = 4
q = 2
if parameter == 'hght':
return (9240 + 150 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 200 * np.cos(y * np.pi) + 400 * y * np.cos(x * np.pi + np.pi))
elif parameter == 'temp':
return (-50 + 2 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 2 * np.cos(y * np.pi) + 5 * y * np.cos(x * np.pi + np.pi))
def digging_300hPa_trough(parameter='hght'):
""" Digging trough with heights and Temperatures based on Sanders Analytic Model
"""
X = np.linspace(.25, .75, 101)
Y = np.linspace(.25, .75, 101)
x, y = np.meshgrid(X, Y)
p = 4
q = 2
if parameter == 'hght':
return (9240 + 150 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 200 * np.cos(y * np.pi) + 400 * y * np.sin(x * np.pi + 5 * np.pi / 2))
elif parameter == 'temp':
return (-50 + 2 * np.cos(p * x * np.pi) * np.cos(q * y * np.pi)
+ 2 * np.cos(y * np.pi) + 5 * y * np.sin(x * np.pi + np.pi / 2))
```
Call the appropriate definition to develop the desired wave.
```
# Single Trough
Z = single_300hPa_trough(parameter='hght')
T = single_300hPa_trough(parameter='temp')
# Lifting Trough
# Z = lifting_300hPa_trough(parameter='hght')
# T = lifting_300hPa_trough(parameter='temp')
# Digging Trough
# Z = digging_300hPa_trough(parameter='hght')
# T = digging_300hPa_trough(parameter='temp')
```
Set geographic parameters for analytic grid to then
```
lats = np.linspace(35, 50, 101)
lons = np.linspace(260, 290, 101)
lon, lat = np.meshgrid(lons, lats)
# Calculate Geostrophic Wind from Analytic Heights
f = mpcalc.coriolis_parameter(lat * units('degrees'))
dx, dy = mpcalc.lat_lon_grid_deltas(lons, lats)
ugeo, vgeo = mpcalc.geostrophic_wind(Z*units.meter, f, dx, dy, dim_order='yx')
# Get the wind direction for each point
wdir = mpcalc.wind_direction(ugeo, vgeo)
# Compute the Gradient Wind via an approximation
dydx = mpcalc.first_derivative(Z, delta=dx, axis=1)
d2ydx2 = mpcalc.first_derivative(dydx, delta=dx, axis=1)
R = ((1 + dydx.m**2)**(3. / 2.)) / d2ydx2.m
geo_mag = mpcalc.wind_speed(ugeo, vgeo)
grad_mag = geo_mag.m - (geo_mag.m**2) / (f.magnitude * R)
ugrad, vgrad = mpcalc.wind_components(grad_mag * units('m/s'), wdir)
# Calculate Ageostrophic wind
uageo = ugrad - ugeo
vageo = vgrad - vgeo
# Compute QVectors
uqvect, vqvect = mpcalc.q_vector(ugeo, vgeo, T * units.degC, 500 * units.hPa, dx, dy)
# Calculate divergence of the ageostrophic wind
div = mpcalc.divergence(uageo, vageo, dx, dy, dim_order='yx')
# Calculate Relative Vorticity Advection
relvor = mpcalc.vorticity(ugeo, vgeo, dx, dy, dim_order='yx')
adv = mpcalc.advection(relvor, (ugeo, vgeo), (dx, dy), dim_order='yx')
```
Create figure containing Geopotential Heights, Temperature, Divergence
of the Ageostrophic Wind, Relative Vorticity Advection (shaded),
geostrphic wind barbs, and Q-vectors.
```
fig = plt.figure(figsize=(10, 10))
ax = plt.subplot(111)
# Plot Geopotential Height Contours
cs = ax.contour(lons, lats, Z, range(0, 12000, 120), colors='k')
plt.clabel(cs, fmt='%d')
# Plot Temperature Contours
cs2 = ax.contour(lons, lats, T, range(-50, 50, 2), colors='r', linestyles='dashed')
plt.clabel(cs2, fmt='%d')
# Plot Divergence of Ageo Wind Contours
cs3 = ax.contour(lons, lats, div*10**9, np.arange(-25, 26, 3), colors='grey',
linestyles='dotted')
plt.clabel(cs3, fmt='%d')
# Plot Rel. Vor. Adv. colorfilled
cf = ax.contourf(lons, lats, adv*10**9, np.arange(-20, 21, 1), cmap=plt.cm.bwr)
cbar = plt.colorbar(cf, orientation='horizontal', pad=0.05, aspect=50)
cbar.set_label('Rel. Vor. Adv.')
# Plot Geostrophic Wind Barbs
wind_slice = slice(5, None, 10)
ax.barbs(lons[wind_slice], lats[wind_slice],
ugeo[wind_slice, wind_slice].to('kt').m, vgeo[wind_slice, wind_slice].to('kt').m)
# Plot Ageostrophic Wind Vectors
# ageo_slice = slice(None, None, 10)
# ax.quiver(lons[ageo_slice], lats[ageo_slice],
# uageo[ageo_slice, ageo_slice].m, vageo[ageo_slice, ageo_slice].m,
# color='blue', pivot='mid')
# Plot QVectors
qvec_slice = slice(None, None, 10)
ax.quiver(lons[qvec_slice], lats[qvec_slice],
uqvect[qvec_slice, qvec_slice].m, vqvect[qvec_slice, qvec_slice].m,
color='darkcyan', pivot='mid')
plt.title('300-hPa Geo Heights (black), Q-Vector (dark cyan), Divergence (grey; dashed)')
plt.show()
```
| github_jupyter |
## 사전 설치
* gensim, BeautifulSoup, nltk 가 설치되지 않았다면 설치가 필요합니다.
### gensim
* 최근버전에서 pickle을 로드하지 못 하는 문제로 이전 버전을 사용합니다.
* pip사용시 : `pip install --upgrade gensim==3.2.0`
* conda 사용시 : `conda install -c conda-forge gensim==3.2.0`
### nltk
* pip사용시 : `pip install -U nltk`
* conda 사용시 : `conda install -c anaconda nltk`
## [Bag of Words Meets Bags of Popcorn | Kaggle](https://www.kaggle.com/c/word2vec-nlp-tutorial#part-3-more-fun-with-word-vectors)
# 튜토리얼 파트 3, 4
* [DeepLearningMovies/KaggleWord2VecUtility.py at master · wendykan/DeepLearningMovies](https://github.com/wendykan/DeepLearningMovies/blob/master/KaggleWord2VecUtility.py)
* 캐글에 링크 되어 있는 github 튜토리얼을 참고하여 만들었으며 파이썬2로 되어있는 소스를 파이썬3에 맞게 일부 수정하였다.
### 첫 번째 시도(average feature vectors)
- 튜토리얼2의 코드로 벡터의 평균을 구한다.
### 두 번째 시도(K-means)
- Word2Vec은 의미가 관련있는 단어들의 클러스터를 생성하기 때문에 클러스터 내의 단어 유사성을 이용하는 것이다.
- 이런식으로 벡터를 그룹화 하는 것을 "vector quantization(벡터 양자화)"라고 한다.
- 이를 위해서는 K-means와 같은 클러스터링 알고리즘을 사용하여 클러스터라는 단어의 중심을 찾아야 한다.
- 비지도학습인 K-means를 통해 클러스터링을 하고 지도학습인 랜덤포레스트로 리뷰가 추천인지 아닌지를 예측한다.
```
import pandas as pd
import numpy as np
from gensim.models import Word2Vec
from sklearn.cluster import KMeans
from sklearn.ensemble import RandomForestClassifier
from bs4 import BeautifulSoup
import re
import time
from nltk.corpus import stopwords
import nltk.data
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
model = Word2Vec.load('300features_40minwords_10text')
model
# 숫자로 단어를 표현
# Word2Vec 모델은 어휘의 각 단어에 대한 feature 벡터로 구성되며
# 'syn0'이라는 넘파이 배열로 저장된다.
# syn0의 행 수는 모델 어휘의 단어 수
# 컬럼 수는 2 부에서 설정 한 피처 벡터의 크기
type(model.wv.syn0)
# syn0의 행 수는 모델 어휘의 단어 수
# 열 수는 2부에서 설정한 특징 벡터의 크기
model.wv.syn0.shape
# 개별 단어 벡터 접근
model.wv['flower'].shape
model.wv['flower'][:10]
```
## K-means (K평균)클러스터링으로 데이터 묶기
* [K-평균 알고리즘 - 위키백과, 우리 모두의 백과사전](https://ko.wikipedia.org/wiki/K-%ED%8F%89%EA%B7%A0_%EC%95%8C%EA%B3%A0%EB%A6%AC%EC%A6%98)
- 클러스터링은 비지도 학습 기법
- 클러스터링은 유사성 등 개념에 기초해 몇몇 그룹으로 분류하는 기법
- 클러스터링의 목적은 샘플(실수로 구성된 n차원의 벡터)을 내부적으로는 비슷하지만 외부적으로 공통 분모가 없는 여러 그룹으로 묶는 것
- 특정 차원의 범위가 다른 차원과 차이가 크면 클러스터링 하기 전에 스케일을 조정해야 한다.
1. 최초 센트로이드(centroid)(중심점)로 k개의 벡터를 무작위로 선정한다.
2. 각 샘플을 그 위치에서 가장 가까운 센트로이드에 할당한다.
3. 센트로이드의 위치를 재계산한다.
4. 센트로이드가 더 이상 움직이지 않을 때까지 2와 3을 반복한다.
참고 : [책] 모두의 데이터 과학(with 파이썬)
```
# 단어 벡터에서 k-means를 실행하고 일부 클러스터를 찍어본다.
start = time.time() # 시작시간
# 클러스터의 크기 "k"를 어휘 크기의 1/5 이나 평균 5단어로 설정한다.
word_vectors = model.wv.syn0 # 어휘의 feature vector
num_clusters = word_vectors.shape[0] / 5
num_clusters = int(num_clusters)
num_clusters
# K means 를 정의하고 학습시킨다.
kmeans_clustering = KMeans( n_clusters = num_clusters )
idx = kmeans_clustering.fit_predict( word_vectors )
# 끝난시간에서 시작시간을 빼서 걸린 시간을 구한다.
end = time.time()
elapsed = end - start
print("Time taken for K Means clustering: ", elapsed, "seconds.")
# 각 어휘 단어를 클러스터 번호에 매핑되게 word/Index 사전을 만든다.
idx = list(idx)
names = model.wv.index2word
word_centroid_map = {names[i]: idx[i] for i in range(len(names))}
# word_centroid_map = dict(zip( model.wv.index2word, idx ))
# 첫번째 클러스터의 처음 10개를 출력
for cluster in range(0,10):
# 클러스터 번호를 출력
print("\nCluster {}".format(cluster))
# 클러스터번호와 클러스터에 있는 단어를 찍는다.
words = []
for i in range(0,len(list(word_centroid_map.values()))):
if( list(word_centroid_map.values())[i] == cluster ):
words.append(list(word_centroid_map.keys())[i])
print(words)
"""
판다스로 데이터프레임 형태의 데이터로 읽어온다.
QUOTE_MINIMAL (0), QUOTE_ALL (1),
QUOTE_NONNUMERIC (2) or QUOTE_NONE (3).
그리고 이전 튜토리얼에서 했던 것처럼 clean_train_reviews 와
clean_test_reviews 로 텍스트를 정제한다.
"""
train = pd.read_csv('data/labeledTrainData.tsv',
header=0, delimiter="\t", quoting=3)
test = pd.read_csv('data/testData.tsv',
header=0, delimiter="\t", quoting=3)
# unlabeled_train = pd.read_csv( 'data/unlabeledTrainData.tsv', header=0, delimiter="\t", quoting=3 )
from KaggleWord2VecUtility import KaggleWord2VecUtility
# 학습 리뷰를 정제한다.
clean_train_reviews = []
for review in train["review"]:
clean_train_reviews.append(
KaggleWord2VecUtility.review_to_wordlist( review, \
remove_stopwords=True ))
# 테스트 리뷰를 정제한다.
clean_test_reviews = []
for review in test["review"]:
clean_test_reviews.append(
KaggleWord2VecUtility.review_to_wordlist( review, \
remove_stopwords=True ))
# bags of centroids 생성
# 속도를 위해 centroid 학습 세트 bag을 미리 할당 한다.
train_centroids = np.zeros((train["review"].size, num_clusters), \
dtype="float32" )
print(train_centroids.shape)
train_centroids[:5]
# centroid 는 두 클러스터의 중심점을 정의 한 다음 중심점의 거리를 측정한 것
def create_bag_of_centroids( wordlist, word_centroid_map ):
# 클러스터의 수는 word / centroid map에서 가장 높은 클러스트 인덱스와 같다.
num_centroids = max( word_centroid_map.values() ) + 1
# 속도를 위해 bag of centroids vector를 미리 할당한다.
bag_of_centroids = np.zeros( num_centroids, dtype="float32" )
# 루프를 돌며 단어가 word_centroid_map에 있다면
# 해당되는 클러스터의 수를 하나씩 증가시켜 준다.
for word in wordlist:
if word in word_centroid_map:
index = word_centroid_map[word]
bag_of_centroids[index] += 1
# bag of centroids를 반환한다.
return bag_of_centroids
# 학습 리뷰를 bags of centroids 로 변환한다.
counter = 0
for review in clean_train_reviews:
train_centroids[counter] = create_bag_of_centroids( review, \
word_centroid_map )
counter += 1
# 테스트 리뷰도 같은 방법으로 반복해 준다.
test_centroids = np.zeros(( test["review"].size, num_clusters), \
dtype="float32" )
counter = 0
for review in clean_test_reviews:
test_centroids[counter] = create_bag_of_centroids( review, \
word_centroid_map )
counter += 1
# 랜덤포레스트를 사용하여 학습시키고 예측
forest = RandomForestClassifier(n_estimators = 100)
# train 데이터의 레이블을 통해 학습시키고 예측한다.
# 시간이 좀 소요되기 때문에 %time을 통해 걸린 시간을 찍도록 함
print("Fitting a random forest to labeled training data...")
%time forest = forest.fit(train_centroids, train["sentiment"])
temp = pd.DataFrame(test_centroids)
print(temp.shape)
temp.sum()
from sklearn.model_selection import cross_val_score
%time score = np.mean(cross_val_score(\
forest, train_centroids, train['sentiment'], cv=10,\
scoring='roc_auc'))
%time result = forest.predict(test_centroids)
score
# 결과를 csv로 저장
output = pd.DataFrame(data={"id":test["id"], "sentiment":result})
output.to_csv("data/submit_BagOfCentroids_{0:.5f}.csv".format(score), index=False, quoting=3)
fig, axes = plt.subplots(ncols=2)
fig.set_size_inches(12,5)
sns.countplot(train['sentiment'], ax=axes[0])
sns.countplot(output['sentiment'], ax=axes[1])
output_sentiment = output['sentiment'].value_counts()
print(output_sentiment[0] - output_sentiment[1])
output_sentiment
# 캐글 점수 0.84908
print(330/528)
```
### 왜 이 튜토리얼에서는 Bag of Words가 더 좋은 결과를 가져올까?
벡터를 평균화하고 centroids를 사용하면 단어 순서가 어긋나며 Bag of Words 개념과 매우 비슷하다. 성능이 (표준 오차의 범위 내에서) 비슷하기 때문에 튜토리얼 1, 2, 3이 동등한 결과를 가져온다.
첫째, Word2Vec을 더 많은 텍스트로 학습시키면 성능이 좋아진다. Google의 결과는 10 억 단어가 넘는 코퍼스에서 배운 단어 벡터를 기반으로 한다. 학습 레이블이 있거나 레이블이 없는 학습 세트는 단지 대략 천팔백만 단어 정도다. 편의상 Word2Vec은 Google의 원래 C도구에서 출력되는 사전 학습 된 모델을 로드하는 기능을 제공하기 때문에 C로 모델을 학습 한 다음 Python으로 가져올 수도 있다.
둘째, 출판 된 자료들에서 분산 워드 벡터 기술은 Bag of Words 모델보다 우수한 것으로 나타났다. 이 논문에서는 IMDB 데이터 집합에 단락 벡터 (Paragraph Vector)라는 알고리즘을 사용하여 현재까지의 최첨단 결과 중 일부를 생성한다. 단락 벡터는 단어 순서 정보를 보존하는 반면 벡터 평균화 및 클러스터링은 단어 순서를 잃어 버리기 때문에 여기에서 시도하는 방식보다 부분적으로 더 좋다.
* 더 공부하기 : 스탠포드 NLP 강의 : [Lecture 1 | Natural Language Processing with Deep Learning - YouTube](https://www.youtube.com/watch?v=OQQ-W_63UgQ&list=PL3FW7Lu3i5Jsnh1rnUwq_TcylNr7EkRe6)
| github_jupyter |
```
# pandas
import pandas as pd
from pandas import Series,DataFrame
# numpy, matplotlib, seaborn
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
%matplotlib inline
import xgboost as xgb #GBM algorithm
from xgboost import XGBRegressor
from sklearn import cross_validation, metrics #Additional scklearn functions
from sklearn.grid_search import GridSearchCV # Perforing grid search
from IPython.display import display
# remove warnings
import warnings
warnings.filterwarnings('ignore')
train_data = pd.read_csv('data/new_train.csv')
test_data = pd.read_csv('data/new_test.csv')
print train_data.shape
display(train_data.head(1))
# display(train_data.info())
print test_data.shape
display(test_data.head(1))
# display(test_data.info())
train_length = train_data.shape[0]
import math
def common_num_range(start,stop,step):
startlen = stoplen = steplen = 0
if '.' in str(start):
startlen = len(str(start)) - str(start).index('.') - 1
if '.' in str(stop):
stoplen = len(str(stop)) - str(stop).index('.') - 1
if '.' in str(step):
steplen = len(str(step)) - str(step).index('.') - 1
maxlen = startlen
if stoplen > maxlen:
maxlen = stoplen
if steplen > maxlen:
maxlen = steplen
power = math.pow(10, maxlen)
if startlen == 0 and stoplen == 0 and steplen == 0:
return range(start, stop, step)
else:
return [num / power for num in range(int(start*power), int(stop*power), int(step*power))]
train_id = train_data['Id']
train_Y = train_data['SalePrice']
train_data.drop(['Id', 'SalePrice'], axis=1, inplace=True)
train_X = train_data
test_Id = test_data['Id']
test_data.drop('Id', axis=1, inplace=True)
test_X = test_data
# formatting for xgb
dtrain = xgb.DMatrix(train_X, label=train_Y)
dtest = xgb.DMatrix(test_X)
```
# XGBoost & Parameter Tuning
Ref: [Complete Guide to Parameter Tuning in XGBoost](https://www.analyticsvidhya.com/blog/2016/03/complete-guide-parameter-tuning-xgboost-with-codes-python/)
## Parameters Tuning Plan
The overall parameters can be divided into 3 categories:
1. General Parameters: Guide the overall functioning
2. Booster Parameters: Guide the individual booster (tree/regression) at each step
3. Learning Task Parameters: Guide the optimization performed
In `XGBRegressor`:
```
class xgboost.XGBRegressor(max_depth=3, learning_rate=0.1, n_estimators=100, silent=True, objective='reg:linear', nthread=-1, gamma=0, min_child_weight=1, max_delta_step=0, subsample=1, colsample_bytree=1, colsample_bylevel=1, reg_alpha=0, reg_lambda=1, scale_pos_weight=1, base_score=0.5, seed=0, missing=None)
```
```
# The error metric: RMSE on the log of the sale prices.
from sklearn.metrics import mean_squared_error
def rmse(y_true, y_pred):
return np.sqrt(mean_squared_error(y_true, y_pred))
def model_cross_validate(xgb_regressor, cv_paramters, dtrain, cv_folds = 5,
early_stopping_rounds = 50, perform_progress=False):
"""
xgb model cross validate to choose best param from giving cv_paramters.
@param cv_paramters:dict,where to choose best param. {'param':[1,2,3]}
@param dtrain:xgboost.DMatrix, training data formatted for xgb
@param early_stopping_rounds: Activates early stopping.Stop when perfomance
does not improve for some rounds
"""
# get initial parameters
xgb_param = xgb_regressor.get_xgb_params()
# save best param
best_params = {}
best_cvresult = None
min_mean_rmse = float("inf")
for param, values in cv_paramters.items():
print '===========Tuning paramter:',param,'==========='
best_param = values[0]
for value in values:
# set the param's value
xgb_param[param] = value
# cv to tune param from values
cvresult = xgb.cv(xgb_param, dtrain, num_boost_round=xgb_param['n_estimators'],
nfold=cv_folds, metrics='rmse',
early_stopping_rounds=early_stopping_rounds)
# calcuate the mean of several final rmses
round_count = cvresult.shape[0]
mean_rmse = cvresult.loc[round_count-11:round_count-1,'test-rmse-mean'].mean()
if perform_progress:
std_rmse = cvresult.loc[round_count-11:round_count-1,'test-rmse-std'].mean()
if isinstance(value, int):
print "%s=%d CV RMSE : Mean = %.7g | Std = %.7g" % (param, value, mean_rmse, std_rmse)
else:
print "%s=%f CV RMSE : Mean = %.7g | Std = %.7g" % (param, value, mean_rmse, std_rmse)
if mean_rmse < min_mean_rmse:
best_param = value
best_cvresult = cvresult
min_mean_rmse = mean_rmse
best_params[param] = best_param
# set best param value for xgb params, important
xgb_param[param] = best_param
print "best ", param, " = ", best_params[param]
return best_params, min_mean_rmse, best_cvresult
def model_fit(xgb_regressor, train_x, train_y, performCV=True,
printFeatureImportance=True, cv_folds=5):
# Perform cross-validation
if performCV:
xgb_param = xgb_regressor.get_xgb_params()
cvresult = xgb.cv(xgb_param, dtrain, num_boost_round=xgb_param['n_estimators'],
nfold=cv_folds, metrics='rmse',
early_stopping_rounds=50)
round_count = cvresult.shape[0]
mean_rmse = cvresult.loc[round_count-11:round_count-1,'test-rmse-mean'].mean()
std_rmse = cvresult.loc[round_count-11:round_count-1,'test-rmse-std'].mean()
print "CV RMSE : Mean = %.7g | Std = %.7g" % (mean_rmse, std_rmse)
# fir the train data
xgb_regressor.fit(train_x, train_y)
# Predict training set
train_predictions = xgb_regressor.predict(train_x)
mse = rmse(train_y, train_predictions)
print("Train RMSE: %.7f" % mse)
# Print Feature Importance
if printFeatureImportance:
feature_importances = pd.Series(xgb_regressor.feature_importances_, train_x.columns.values)
feature_importances = feature_importances.sort_values(ascending=False)
feature_importances= feature_importances.head(40)
feature_importances.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
return xgb_regressor, feature_importances
```
Baseline XGBRegressor
```
xgb_regressor = XGBRegressor(seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
```
### 1. Choose a relatively high learning_rate,optimum n_estimators
```
param_test = {'n_estimators':range(300,400,10)}
xgb_regressor = XGBRegressor(
learning_rate =0.05,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
best_param, min_mean_rmse, best_cvresult = \
model_cross_validate(xgb_regressor, param_test, dtrain, perform_progress=True)
print 'cross-validate best params:', best_param
print 'cross-validate min_mean_rmse:', min_mean_rmse
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
```
### 2.Fix learning rate and number of estimators for tuning tree-based parameters
Tune `max_depth` and `min_child_weight`
```
param_test = {'max_depth':range(1,6,1),
'min_child_weight':common_num_range(1,2,0.1)}
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
best_param, min_mean_rmse, best_cvresult = \
model_cross_validate(xgb_regressor, param_test, dtrain, perform_progress=True)
print 'cross-validate best params:', best_param
print 'cross-validate min_mean_rmse:', min_mean_rmse
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
```
Tune `gamma`,Minimum loss reduction required to make a further partition on a leaf node of the tree.
```
param_test = {'gamma':[0, 0.1, 0.01, 0.001,0.0001, 0.00001]}
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
best_param, min_mean_rmse, best_cvresult = \
model_cross_validate(xgb_regressor, param_test, dtrain, perform_progress=True)
print 'cross-validate best params:', best_param
print 'cross-validate min_mean_rmse:', min_mean_rmse
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.8,
colsample_bytree=0.8,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
```
Tune `subsample` and `colsample_bytree`
- subsample : Subsample ratio of the training instance.
- colsample_bytree : Subsample ratio of columns when constructing each tree
```
param_test = {'subsample':common_num_range(0.6, 0.9, 0.01),
'colsample_bytree':common_num_range(0.6, 0.9, 0.01)}
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
best_param, min_mean_rmse, best_cvresult = \
model_cross_validate(xgb_regressor, param_test, dtrain, perform_progress=True)
print 'cross-validate best params:', best_param
print 'cross-validate min_mean_rmse:', min_mean_rmse
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.72,
colsample_bytree=0.89,
reg_lambda = 0.1,
reg_alpha = 0.1,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
param_test2 = {'reg_lambda':common_num_range(0.55, 0.65, 0.01),
'reg_alpha':common_num_range(0.45, 0.6, 0.01)}
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.72,
colsample_bytree=0.89,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
best_param, min_mean_rmse, best_cvresult = \
model_cross_validate(xgb_regressor, param_test2, dtrain, perform_progress=True)
print 'cross-validate best params:', best_param
print 'cross-validate min_mean_rmse:', min_mean_rmse
xgb_regressor = XGBRegressor(
learning_rate =0.05,
n_estimators = 300,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.72,
colsample_bytree=0.89,
reg_lambda = 0.61,
reg_alpha = 0.53,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
xgb_regressor = XGBRegressor(
learning_rate =0.01,
n_estimators = 4000,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.72,
colsample_bytree=0.89,
reg_lambda = 0.61,
reg_alpha = 0.53,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
xgb_regressor, feature_importances = model_fit(xgb_regressor,train_X, train_Y)
```
Final paramters:
```
xgb_regressor = XGBRegressor(
learning_rate =0.01,
n_estimators = 4000,
max_depth=3,
min_child_weight=1.1,
gamma=0.01,
subsample=0.72,
colsample_bytree=0.89,
reg_lambda = 0.61,
reg_alpha = 0.53,
scale_pos_weight=1,
objective= 'reg:linear',
seed=10)
```
```
xgb_predictions = xgb_regressor.predict(test_X)
xgb_predictions = np.expm1(xgb_predictions)
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": xgb_predictions
})
submission.to_csv("result/xgb_param_tune_predictions_2_13.csv", index=False)
print "Done."
```
# Model Voting
Ridge, ElasticNet, Lasso, XGBRegressor model voting.
```
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(train_X, train_Y,
test_size=0.4, random_state=0)
from sklearn.linear_model import Ridge, ElasticNet, Lasso
def simple_model_cross_validate(alphas, Model, model_name):
min_rmse = float('inf')
best_alpha = None
for alpha in alphas:
model = Model(alpha, max_iter=50000).fit(X_train, y_train)
model_rmse = rmse(model.predict(X_test), y_test)
if model_rmse < min_rmse:
best_alpha = alpha
min_rmse = model_rmse
print model_name, 'best_alpha = ', best_alpha, 'min_rmse = ', min_rmse
alphas = common_num_range(0.0001, 0.002, 0.0001)
simple_model_cross_validate(alphas, Lasso, 'Lasso')
simple_model_cross_validate(alphas, ElasticNet, 'ElasticNet')
alphas = common_num_range(25, 50, 1)
simple_model_cross_validate(alphas, Ridge, 'Ridge')
lasso_model = Lasso(alpha=0.0009, max_iter=50000).fit(X_train, y_train)
elastic_net_model = ElasticNet(alpha=0.0019, max_iter=50000).fit(X_train, y_train)
ridge_model = Ridge(alpha=41, max_iter=50000).fit(X_train, y_train)
lasso_predictions = lasso_model.predict(test_X)
lasso_predictions = np.expm1(lasso_predictions)
ridge_predictions = ridge_model.predict(test_X)
ridge_predictions = np.expm1(ridge_predictions)
elastic_net_predictions = elastic_net_model.predict(test_X)
elastic_net_predictions = np.expm1(elastic_net_predictions)
predictions = (lasso_predictions + ridge_predictions + elastic_net_predictions + xgb_predictions) / 4
plt.subplot(221)
plt.plot(lasso_predictions, c="blue") # 0.12818
plt.title('lasso 0.12818')
plt.subplot(222)
plt.plot(elastic_net_predictions, c="yellow") # 0.12908
plt.title('elastic_net 0.12908')
plt.subplot(223)
plt.plot(ridge_predictions, c="pink") # 0.13161
plt.title('ridge 0.13161')
plt.subplot(224)
plt.plot(xgb_predictions, c="green") # 0.12167
plt.title('xgb 0.12167')
plt.subplots_adjust(wspace=0.5, hspace=0.5)
plt.show()
plt.subplot(111)
plt.plot(predictions, c="red") # 0.12419
plt.title('4 model vote 0.12419')
# outlier data
np.argwhere(lasso_predictions == lasso_predictions[lasso_predictions > 700000])
# convert outlier data to xgb_predictions[1089]
lasso_predictions[1089] = xgb_predictions[1089]
ridge_predictions[1089] = xgb_predictions[1089]
elastic_net_predictions[1089] = xgb_predictions[1089]
lasso_score = 1-0.12818
ridge_score = 1-0.13161
elastic_net_score = 1-0.12908
xgb_score = 1-0.12167
total_score = lasso_score + ridge_score + elastic_net_score + xgb_score
predictions = (lasso_score / total_score) * lasso_predictions + \
(ridge_score / total_score) * ridge_predictions + \
(elastic_net_score / total_score) * elastic_net_predictions + \
(xgb_score / total_score) * xgb_predictions
plt.subplot(221)
plt.plot(lasso_predictions, c="blue") # 0.12818
plt.title('lasso 0.12818')
plt.subplot(222)
plt.plot(elastic_net_predictions, c="yellow") # 0.12908
plt.title('elastic_net 0.12908')
plt.subplot(223)
plt.plot(ridge_predictions, c="pink") # 0.13161
plt.title('ridge 0.13161')
plt.subplot(224)
plt.plot(xgb_predictions, c="green") # 0.12167
plt.title('xgb 0.12167')
plt.subplots_adjust(wspace=0.5, hspace=0.5)
plt.show()
plt.subplot(111)
plt.plot(predictions, c="red") # 0.12417
plt.title('4 model vote 0.12417')
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": lasso_predictions
})
submission.to_csv("result/lasso_predictions_2_13.csv", index=False)
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": ridge_predictions
})
submission.to_csv("result/ridge_predictions_2_13.csv", index=False)
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": elastic_net_predictions
})
submission.to_csv("result/elastic_net_predictions_2_13.csv", index=False)
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": xgb_predictions
})
submission.to_csv("result/xgb_predictions_2_13.csv", index=False)
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": predictions
})
submission.to_csv("result/4_model_vote_predictions_2_13.csv", index=False)
print "Done."
```
# Best Vote Score
```
from sklearn.linear_model import LassoCV
model_lasso = LassoCV(alphas = [1, 0.1, 0.001, 0.0005]).fit(train_X, train_Y)
print rmse(model_lasso.predict(train_X), train_Y)
lasso_predictions = model_lasso.predict(test_X)
lasso_predictions = np.expm1(lasso_predictions)
predictions = (lasso_predictions + xgb_predictions) / 2
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": predictions
})
submission.to_csv("result/lasso_xgb_vote_predictions_2_13.csv", index=False)
print "Done."
lasso_score = 1-0.12818
xgb_score = 1-0.12167
total_score = lasso_score + xgb_score
predictions = (lasso_score / total_score) * lasso_predictions + \
(xgb_score / total_score) * xgb_predictions
submission = pd.DataFrame({
"Id": test_Id,
"SalePrice": predictions
})
submission.to_csv("result/lasso_xgb_weighted_vote_predictions_2_13.csv", index=False)
print "Done."
```
| github_jupyter |
# Multigraph Network Styling for visJS2jupyter
------------
Authors: Brin Rosenthal (sbrosenthal@ucsd.edu), Mikayla Webster (m1webste@ucsd.edu)
-----------
## Import packages
```
import matplotlib as mpl
import networkx as nx
import pandas as pd
import random
import numpy as np
from visJS2jupyter import visJS_module
reload(visJS_module)
```
We start by creating a randomized, single-edged graph, and convert that to a multigraph
```
G = nx.connected_watts_strogatz_graph(30,5,.2)
G = nx.MultiGraph(G)
edges = G.edges(keys = True) # for multigraphs every edge has to be represented by a three-tuple (source, target, key)
```
We duplicate every edge in the graph to make it a true multigraph.
Note: NetworkX does not support duplicate edges with opposite directions. NetworkX will flip any backwards edges you try to add to your graph. For example, if your graph currently contains the edges [(0,1), (1,2)] and you add the edge (1,0) to your graph, your graph will now contain edges [(0,1), (0,1), (1,2)]
```
sources = list(zip(*edges)[0])
targets = list(zip(*edges)[1])
backward_edges = list(zip(targets, sources)) # demonstarting adding backwards edges
G.add_edges_from(backward_edges)
edges = list(G.edges(data = True))
nodes = list(G.nodes()) # type cast to list in order to make compatible with networkx 1.11 and 2.0
edges = list(G.edges(keys = True)) # for nx 2.0, returns an "EdgeView" object rather than an iterable
```
## Multigraph Node and Edge Styling
There is no difference between multigraph and single-edged-graph styling. Just map the node and edge attributes to some visual properties, and style the nodes and edges according to these properties (like usual!)
```
# add some node attributes to color-code by
degree = dict(G.degree()) # nx 2.0 returns a "DegreeView" object. Cast to dict to maintain compatibility with nx 1.11
bc = nx.betweenness_centrality(G)
nx.set_node_attributes(G, name = 'degree', values = degree) # between networkx 1.11 and 2.0, therefore we must
nx.set_node_attributes(G, name = 'betweenness_centrality', values = bc) # explicitly pass our arguments
# (not implicitly through position)
# add the edge attribute 'weight' to color-code by
weights = []
for i in range(len(edges)):
weights.append(float(random.randint(1,5)))
w_dict = dict(zip(edges, weights))
nx.set_edge_attributes(G, name = 'weight', values = w_dict)
# map the betweenness centrality to the node color, using matplotlib spring_r colormap
node_to_color = visJS_module.return_node_to_color(G,field_to_map='betweenness_centrality',cmap=mpl.cm.spring_r,alpha = 1,
color_max_frac = .9,color_min_frac = .1)
# map weight to edge color, using default settings
edge_to_color = visJS_module.return_edge_to_color(G,field_to_map='weight')
```
## Interactive network
Note that this example is simply the multigraph version of our "Complex Parameters" notebook.
```
# set node initial positions using networkx's spring_layout function
pos = nx.spring_layout(G)
nodes_dict = [{"id":n,"color":node_to_color[n],
"degree":nx.degree(G,n),
"x":pos[n][0]*1000,
"y":pos[n][1]*1000} for n in nodes
]
node_map = dict(zip(nodes,range(len(nodes)))) # map to indices for source/target in edges
edges_dict = [{"source":node_map[edges[i][0]], "target":node_map[edges[i][1]],
"color":edge_to_color[(edges[i][0],edges[i][1],edges[i][2])],"title":'test'} # remember (source, target, key)
for i in range(len(edges))]
# set some network-wide styles
visJS_module.visjs_network(nodes_dict,edges_dict,
node_size_multiplier=3,
node_size_transform = '',
node_color_highlight_border='red',
node_color_highlight_background='#D3918B',
node_color_hover_border='blue',
node_color_hover_background='#8BADD3',
node_font_size=25,
edge_arrow_to=True,
physics_enabled=True,
edge_color_highlight='#8A324E',
edge_color_hover='#8BADD3',
edge_width=3,
max_velocity=15,
min_velocity=1,
edge_smooth_enabled = True)
```
| github_jupyter |
# 네이버 영화리뷰 감정 분석 문제에 SentencePiece 적용해 보기
## Step 1. SentencePiece 설치하기
SentencePiece는 SentencePiece는 Google에서 제공하는 오픈소스 기반 Sentence Tokenizer/Detokenizer 로서, BPE와 unigram 2가지 subword 토크나이징 모델 중 하나를 선택해서 사용할 수 있도록 패키징한 것입니다. 아래 링크의 페이지에서 상세한 내용을 파악할 수 있습니다.
google/sentencepiece (https://github.com/google/sentencepiece)
위 페이지의 서두에서도 언급하고 있듯, SentencePiece는 딥러닝 자연어처리 모델의 앞부분에 사용할 목적으로 최적화되어 있는데, 최근 pretrained model들이 거의 대부분 SentencePiece를 tokenizer로 채용하면서 사실상 표준의 역할을 하고 있습니다. 앞으로의 실습 과정에서 자주 만나게 될 것이므로 꼭 친숙해지시기를 당부드립니다.
다음과 같이 설치를 진행합니다. SentencePiece는 python에서 쓰라고 만들어진 라이브러리는 아니지만 편리한 파이썬 wrapper를 아래와 같이 제공하고 있습니다.
```
!pip install sentencepiece
import os
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
%matplotlib inline
import os
path_to_file = os.getenv('HOME')+'/aiffel/GD_02_sp_tokenizer/data/korean-english-park.train.ko'
with open(path_to_file, "r") as f:
raw = f.read().splitlines()
print("Data Size:", len(raw))
print("Example:")
for sen in raw[0:100][::20]: print(">>", sen)
min_len = 999
max_len = 0
sum_len = 0
for sen in raw:
length = len(sen)
if min_len > length: min_len = length
if max_len < length: max_len = length
sum_len += length
print("문장의 최단 길이:", min_len)
print("문장의 최장 길이:", max_len)
print("문장의 평균 길이:", sum_len // len(raw))
sentence_length = np.zeros((max_len), dtype=np.int)
for sen in raw:
sentence_length[len(sen)-1] += 1
plt.bar(range(max_len), sentence_length, width=1.0)
plt.title("Sentence Length Distribution")
plt.show()
def check_sentence_with_length(raw, length):
count = 0
for sen in raw:
if len(sen) == length:
print(sen)
count += 1
if count > 100: return
check_sentence_with_length(raw, 1)
for idx, _sum in enumerate(sentence_length):
# 문장의 수가 1500을 초과하는 문장 길이를 추출합니다.
if _sum > 1500:
print("Outlier Index:", idx+1)
check_sentence_with_length(raw, 11)
min_len = 999
max_len = 0
sum_len = 0
cleaned_corpus = list(set(raw)) # set를 사용해서 중복을 제거합니다.
print("Data Size:", len(cleaned_corpus))
for sen in cleaned_corpus:
length = len(sen)
if min_len > length: min_len = length
if max_len < length: max_len = length
sum_len += length
print("문장의 최단 길이:", min_len)
print("문장의 최장 길이:", max_len)
print("문장의 평균 길이:", sum_len // len(cleaned_corpus))
sentence_length = np.zeros((max_len), dtype=np.int)
for sen in cleaned_corpus: # 중복이 제거된 코퍼스 기준
sentence_length[len(sen)-1] += 1
plt.bar(range(max_len), sentence_length, width=1.0)
plt.title("Sentence Length Distribution")
plt.show()
max_len = 150
min_len = 10
# 길이 조건에 맞는 문장만 선택합니다.
filtered_corpus = [s for s in cleaned_corpus if (len(s) < max_len) & (len(s) >= min_len)]
# 분포도를 다시 그려봅니다.
sentence_length = np.zeros((max_len), dtype=np.int)
for sen in filtered_corpus:
sentence_length[len(sen)-1] += 1
plt.bar(range(max_len), sentence_length, width=1.0)
plt.title("Sentence Length Distribution")
plt.show()
```
## Step 2. SentencePiece 모델 학습¶
앞서 배운 tokenize() 함수를 기억하나요? 다시 한번 상기시켜드릴게요!
```
def tokenize(corpus): # corpus: Tokenized Sentence's List
tokenizer = tf.keras.preprocessing.text.Tokenizer(filters='')
tokenizer.fit_on_texts(corpus)
tensor = tokenizer.texts_to_sequences(corpus)
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor, padding='post')
return tensor, tokenizer
import sentencepiece as spm
import os
temp_file = os.getenv('HOME')+'/aiffel/GD_02_sp_tokenizer/data/korean-english-park.train.ko.temp'
vocab_size = 8000
with open(temp_file, 'w') as f:
for row in filtered_corpus: # 이전 스텝에서 정제했던 corpus를 활용합니다.
f.write(str(row) + '\n')
# spm.SentencePieceTrainer.Train(
# '--input={} --model_prefix=korean_spm --vocab_size={}'.format(temp_file, vocab_size)
# )
# #위 Train에서 --model_type = 'unigram'이 디폴트 적용되어 있습니다. --model_type = 'bpe' 로 옵션을 주어 변경할 수 있습니다.
# # 왜인지 오류가 나서 학습이 되지 않음
vocab_size_8k = 8000
vocab_size_16k = 16000
# vocab size=8000, model_type=unigram
spm.SentencePieceTrainer.Train(
'--input={} --model_prefix=kor_spm_unigram_8k --vocab_size={}'.format(temp_file, vocab_size_8k)
)
# vocab size=16000, model_type=unigram
spm.SentencePieceTrainer.Train(
'--input={} --model_prefix=kor_spm_uingram_4k --vocab_size={}'.format(temp_file, vocab_size_8k/2)
)
# vocab size=8000, model_type=bpe
spm.SentencePieceTrainer.Train(
'--input={} --model_prefix=kor_spm_bpe_8k --model_type=bpe --vocab_size={}'.format(temp_file, vocab_size_8k)
)
# vocab size = 16000, model_type=bpe
spm.SentencePieceTrainer.Train(
'--input={} --model_prefix=kor_spm_bpe_16k --model_type=bpe --vocab_size={}'.format(temp_file, vocab_size_8k*2)
)
#위 Train에서 --model_type = 'unigram'이 디폴트 적용되어 있습니다. --model_type = 'bpe' 로 옵션을 주어 변경할 수 있습니다.
!ls -l korean_spm*
s = spm.SentencePieceProcessor()
s.Load('korean_spm.model')
# SentencePiece를 활용한 sentence -> encoding
tokensIDs = s.EncodeAsIds('아버지가방에들어가신다.')
print(tokensIDs)
# SentencePiece를 활용한 sentence -> encoded pieces
print(s.SampleEncodeAsPieces('아버지가방에들어가신다.',1, 0.0))
# SentencePiece를 활용한 encoding -> sentence 복원
print(s.DecodeIds(tokensIDs))
```
## Step 3. Tokenizer 함수 작성
우리는 위에서 훈련시킨 SentencePiece를 활용하여 위 함수와 유사한 기능을 하는 sp_tokenize() 함수를 정의할 겁니다. 하지만 SentencePiece가 동작하는 방식이 단순 토큰화와는 달라 완전히 동일하게는 정의하기 어렵습니다. 그러니 아래 조건을 만족하는 함수를 정의하도록 하습니다.
1) 매개변수로 토큰화된 문장의 list를 전달하는 대신 온전한 문장의 list 를 전달합니다.
2) 생성된 vocab 파일을 읽어와 { : } 형태를 가지는 word_index 사전과 { : } 형태를 가지는 index_word 사전을 생성하고 함께 반환합니다.
3) 리턴값인 tensor 는 앞의 함수와 동일하게 토큰화한 후 Encoding된 문장입니다. 바로 학습에 사용할 수 있게 Padding은 당연히 해야겠죠?
```
def sp_tokenize(s, corpus):
tensor = []
for sen in corpus:
tensor.append(s.EncodeAsIds(sen))
with open("./korean_spm.vocab", 'r') as f:
vocab = f.readlines()
word_index = {}
index_word = {}
for idx, line in enumerate(vocab):
word = line.split("\t")[0]
word_index.update({idx:word})
index_word.update({word:idx})
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor, padding='post')
return tensor, word_index, index_word
#sp_tokenize(s, corpus) 사용예제
my_corpus = ['나는 새싹 캠퍼스에 출근해서 일을 했습니다.', '하지만 ㅠㅠ 월급을 받지 못해서 밥을 굶었습니다...']
tensor, word_index, index_word = sp_tokenize(s, my_corpus)
print(tensor)
```
## Step 4. 네이버 영화리뷰 감정 분석 문제에 SentencePiece 적용해 보기
아마 여러분들은 네이버 영화리뷰 감정 분석 태스크(https://github.com/e9t/nsmc/)를 한 번쯤은 다루어 보았을 것입니다. 한국어로 된 corpus를 다루어야 하므로 주로 KoNLPy에서 제공하는 형태소 분석기를 사용하여 텍스트를 전처리해서 RNN 모델을 분류기로 사용했을 것입니다.
만약 이 문제에서 tokenizer를 SentencePiece로 바꾸어 다시 풀어본다면 더 성능이 좋아질까요? 비교해 보는 것도 흥미로울 것입니다.
네이버 영화리뷰 감정 분석 코퍼스에 SentencePiece를 적용시킨 모델 학습하기
학습된 모델로 sp_tokenize() 메소드 구현하기
구현된 토크나이저를 적용하여 네이버 영화리뷰 감정 분석 모델을 재학습하기
KoNLPy 형태소 분석기를 사용한 모델과 성능 비교하기
(보너스) SentencePiece 모델의 model_type, vocab_size 등을 변경해 가면서 성능 개선 여부 확인하기
Word Vector는 활용할 필요가 없습니다. 활용이 가능하지도 않을 것입니다. 머지않아 SentencePiece와 BERT 등의 pretrained 모델을 함께 활용하는 태스크를 다루게 될 것입니다.
### Reference
1) https://github.com/miinkang/AI_Project_AIFFEL/blob/main/%5BGD-02%5DSentencePiece.ipynb
2) https://piaojian.tistory.com/35
## 1) 데이터 준비와 확인
```
import pandas as pd
import urllib.request
%matplotlib inline
import matplotlib.pyplot as plt
import re
from konlpy.tag import Okt
from tensorflow import keras
from tensorflow.keras.preprocessing.text import Tokenizer
import numpy as np
from tensorflow.keras.preprocessing.sequence import pad_sequences
from collections import Counter
# 데이터를 읽어봅시다.
train_data_org = pd.read_table('~/aiffel/GD_02_sp_tokenizer/data/nsmc/ratings_train.txt')
test_data = pd.read_table('~/aiffel/GD_02_sp_tokenizer/data/nsmc/ratings_test.txt')
train_data.head()
```
## 2) 데이터로더 구성
실습 때 다루었던 IMDB 데이터셋은 텍스트를 가공하여 imdb.data_loader() 메소드를 호출하면 숫자 인덱스로 변환된 텍스트와 word_to_index 딕셔너리까지 친절하게 제공합니다. 그러나 이번에 다루게 될 nsmc 데이터셋은 전혀 가공되지 않은 텍스트 파일로 이루어져 있습니다. 이것을 읽어서 imdb.data_loader()와 동일하게 동작하는 자신만의 data_loader를 만들어 보는 것으로 시작합니다. data_loader 안에서는 다음을 수행해야 합니다.
데이터의 중복 제거
NaN 결측치 제거
한국어 토크나이저로 토큰화
불용어(Stopwords) 제거
사전word_to_index 구성
텍스트 스트링을 사전 인덱스 스트링으로 변환
X_train, y_train, X_test, y_test, word_to_index 리턴
```
!pip install --upgrade pip
!pip install --upgrade gensim==3.8.3
!pip install konlpy
!sudo apt-get install curl git
!bash <(curl -s https://raw.githubusercontent.com/konlpy/konlpy/master/scripts/mecab.sh)
## LMS Exploration 06 Movie Sentiment Analysis codes
# from konlpy.tag import Mecab
# tokenizer = Mecab()
# stopwords = ['의','가','이','은','들','는','좀','잘','걍','과','도','를','으로','자','에','와','한','하다']
# def load_data(train_data, test_data, num_words=10000):
# train_data.drop_duplicates(subset=['document'], inplace=True)
# train_data = train_data.dropna(how = 'any')
# test_data.drop_duplicates(subset=['document'], inplace=True)
# test_data = test_data.dropna(how = 'any')
# X_train = []
# for sentence in train_data['document']:
# temp_X = tokenizer.morphs(sentence) # 토큰화
# temp_X = [word for word in temp_X if not word in stopwords] # 불용어 제거
# X_train.append(temp_X)
# X_test = []
# for sentence in test_data['document']:
# temp_X = tokenizer.morphs(sentence) # 토큰화
# temp_X = [word for word in temp_X if not word in stopwords] # 불용어 제거
# X_test.append(temp_X)
# words = np.concatenate(X_train).tolist()
# counter = Counter(words)
# counter = counter.most_common(10000-4)
# vocab = ['', '', '', ''] + [key for key, _ in counter]
# word_to_index = {word:index for index, word in enumerate(vocab)}
# def wordlist_to_indexlist(wordlist):
# return [word_to_index[word] if word in word_to_index else word_to_index[''] for word in wordlist]
# X_train = list(map(wordlist_to_indexlist, X_train))
# X_test = list(map(wordlist_to_indexlist, X_test))
# return X_train, np.array(list(train_data['label'])), X_test, np.array(list(test_data['label'])), word_to_index
# X_train, y_train, X_test, y_test, word_to_index = load_data(train_data, test_data)
print("Unique training data : ", train_data_org['document'].nunique())
print("Unique target data : ", train_data_org['label'].nunique())
# delete the duplicate data
train_data_org.drop_duplicates(subset=['document'], inplace=True)
# use the drop_duplicate method
# check the number of the trainig data again
print('number of total sample : ',len(train_data_org))
# check the null value in the document (label)
print(train_data_org_.isnull().values.any())
# check the location of the null data
train_data_org.loc[train_data_org.document.isnull()]
# remove the NULL data
train_data_none = train_data_org.dropna(how = 'any')
print(train_data_none.isnull().values.any())
# data preprocessing
document = train_data_none['document']
naver_review = document
min_len = 999
max_len = 0
sum_len = 0
for sen in document:
length = len(sen)
if min_len > length:
min_len = length
if max_len < length:
max_len = length
sum_len += length
print("문장의 최단 길이:", min_len)
print("문장의 최장 길이:", max_len)
print("문장의 평균 길이:", sum_len // len(document))
sentence_length = np.zeros((max_len), dtype=np.int)
for sen in document:
sentence_length[len(sen)-1] += 1
plt.bar(range(max_len), sentence_length, width=1.0)
plt.title("Sentence Length Distribution")
plt.show()
```
최장 길이 140에 가까운 데이터가 꽤 많습니다. 어떤 문장인지 확인해봅니다.
```
def check_sentence_with_length(raw, length):
count = 0
for sen in raw:
if len(sen) == length:
print(sen)
count += 1
if count > 100: return
check_sentence_with_length(document, 135)
check_sentence_with_length(document, 1)
check_sentence_with_length(document, 8)
```
### 최적의 max_len을 찾는 함수
```
def below_threshold_len(max_len, nested_list):
cnt = 0
for s in nested_list:
if(len(s) <= max_len):
cnt = cnt + 1
print('전체 샘플 중 길이가 %s 이하인 샘플의 비율: %s'%(max_len, (cnt / len(nested_list))*100))
below_threshold_len(36, document) # 평균
below_threshold_len(140, document) # 최장
below_threshold_len(70, document) # 중앙값
below_threshold_len(50, document)
below_threshold_len(60, document)
max_len = 55
min_len = 0
# 길이 조건에 맞는 문장만 선택합니다.
filtered_corpus = [s for s in naver_review if (len(s) < max_len)]
# 데이터프레임 조건부 슬라이싱
# str.len() 두 함수에 대해 알 수 있었다.
train_data = naver_review[naver_review.str.len() < max_len]
sentence_length = np.zeros((max_len), dtype=np.int)
naver_review.head()
for sen in train_data: # 중복이 제거된 코퍼스 기준
sentence_length[len(sen)-1] += 1
plt.bar(range(max_len), sentence_length, width=1.0)
plt.title("Sentence Length Distribution")
plt.show()
plt.figure(figsize=(12,5))
plt.boxplot([sentence_length], showmeans=True)
plt.show()
```
## 전처리 하다가 몇개 데이터 날려먹어서 LSTM 모델 돌릴 때 사이즈가 안 맞아서 다시 EDA 처음부터 시작!
```
train_data = pd.read_table("data/nsmc/"+"ratings_train.txt")
test_data = pd.read_table("data/nsmc/"+"ratings_test.txt")
display(train_data.head())
display(test_data.head())
print("학습데이터 유니크 :",train_data['document'].nunique()) # 약 4,000개의 중복 샘플이 존재
print("타겟데이터 유니크 : ",train_data['label'].nunique())
# 중복데이터 제거
train_data.drop_duplicates(subset=['document'], inplace=True) # drop_duplicates 활용
# 학습데이터 개수 다시 확인
print('총 샘플의 수 :',len(train_data))
# document에 null 값이 있는지 확인
print(train_data.isnull().values.any())
# 어느 위치에 누락값이 있는지 확인
train_data.loc[train_data.document.isnull()]
# Null값 제거
train_data = train_data.dropna(how = 'any') # Null 값이 존재하는 행 제거
print(train_data.isnull().values.any()) # Null 값이 존재하는지 확인
# 데이터 전처리
train_data['document'] = train_data['document'].str.replace("[^ㄱ-ㅎㅏ-ㅣ가-힣 ]","") # 한글과 공백을 제외하고 모두 제거
train_data[:5]
# 바뀐 데이터 내부에 null값이 있는지 다시 확인
train_data['document'] = train_data['document'].str.replace('^ +', "") # white space 데이터를 empty value로 변경
train_data['document'].replace('', np.nan, inplace=True)
print(train_data.isnull().sum())
# null 데이터 삭제
train_data = train_data.dropna(how = 'any')
print(len(train_data))
# 테스트데이터에도 동일한 과정 진행
test_data.drop_duplicates(subset = ['document'], inplace=True) # document 열에서 중복인 내용이 있다면 중복 제거
test_data['document'] = test_data['document'].str.replace("[^ㄱ-ㅎㅏ-ㅣ가-힣 ]","") # 정규 표현식 수행
test_data['document'] = test_data['document'].str.replace('^ +', "") # 공백은 empty 값으로 변경
test_data['document'].replace('', np.nan, inplace=True) # 공백은 Null 값으로 변경
test_data = test_data.dropna(how='any') # Null 값 제거
print('전처리 후 테스트용 샘플의 개수 :',len(test_data))
# 데이터 확인
train_data.info()
print("="*50)
test_data.info()
# 데이터 확인
display(train_data.head())
display(test_data.head())
print("훈련데이터 : ",len(train_data))
print("테스트데이터 : ",len(test_data))
# 데이터 길이 분포 확인하기
import matplotlib.pyplot as plt
min_len = 999
max_len = 0
sum_len = 0
for sen in train_data['document']:
length = len(sen)
# 문장 최소 길이 찾기
if min_len > length:
min_len = length
# 문장 최대 길이 찾기
if max_len < length:
max_len = length
# 전체 문장을 합치면 길이가 얼마나 될까요?
sum_len += length
print("문장의 최단 길이:", min_len)
print("문장의 최장 길이:", max_len)
print("문장의 평균 길이:", sum_len // len(train_data))
# 전체 길이만큼 0벡터 ==> 길이에 따른 문장의 수를 저장하기 위해 먼저 0으로 이루어진 리스트를 만든다!!
sentence_length = np.zeros((max_len), dtype=np.int)
print("sentence_length : ",sentence_length)
print("="*60)
# raw는 위에서 다운로드한 데이터셋!! 전체 길이와 상관없음
for sen in train_data['document']:
sentence_length[len(sen)-1] += 1 # 0으로 이루어진 벡터에 문장 count를 더한 뒤 넣는다.
plt.bar(range(max_len), sentence_length, width=1.0) # 너비는 1.0씩 늘어나도록 설정
plt.title("Sentence Length Distribution")
plt.show()
# 길이 45이하로 한정해 데이터를 만듭니다. ==> 너무 짧은 리뷰는 의미가 없음
# 이렇게 하면 document 내용만 list로 전달됨 ==> 나는 데이터프레임을 원한다!!
train_list = [s for s in train_data['document'] if (len(s) <= 45)]
test_list = [s for s in test_data['document'] if (len(s) <= 45)]
train_list = list(set(train_list))
test_list = list(set(test_list))
train_df = pd.DataFrame(train_list)
test_df = pd.DataFrame(test_list)
# 길이 40이하인 데이터를 기존 데이터와 병합합니다.
new_train_df = pd.merge(train_data, train_df, how='inner', left_on='document', right_on=0)
new_test_df = pd.merge(test_data, test_df, how='inner', left_on='document', right_on=0)
train_data = new_train_df[['id', 'document', 'label']]
test_data = new_test_df[['id', 'document', 'label']]
display(train_data.head())
display(test_data.head())
print("훈련데이터 : ",len(train_data))
print("테스트데이터 : ",len(test_data))
```
## Sentence Piece subwords 점검
```
!ls -l kor_spm_unigram_8k*
!ls -l kor_spm_unigram_4k*
!ls -l kor_spm_bpe_8k*
!ls -l kor_spm_bpe_16k*
#왜인지 unigram_4k 가 없다.
import csv
vocab_list_uni_8k = pd.read_csv('kor_spm_unigram_8k.vocab', sep='\t', header=None, quoting=csv.QUOTE_NONE)
vocab_list_uni_8k[:10]
print('subword의 개수 :', len(vocab_list_uni_8k))
vocab_list_uni_8k.sample(10)
vocab_list_bpe_8k = pd.read_csv('kor_spm_bpe_8k.vocab', sep='\t', header=None, quoting=csv.QUOTE_NONE)
vocab_list_bpe_8k[:10]
print('subword의 개수 :', len(vocab_list_bpe_8k))
vocab_list_bpe_8k.sample(10)
vocab_list_bpe_16k = pd.read_csv('kor_spm_bpe_16k.vocab', sep='\t', header=None, quoting=csv.QUOTE_NONE)
vocab_list_bpe_16k[:10]
print('subword의 개수 :', len(vocab_list_bpe_16k))
vocab_list_bpe_16k.sample(10)
```
## SentencePiece 성능 비교평가
- vocab size = 8k, 16k
- model type = unigram, bpe
- kor_spm_unigram_8k
- kor_spm_unigram_16k
- kor_spm_bpe_8k
- kor_spm_bpe_16k
```
def test_performance(model):
s = spm.SentencePieceProcessor()
s.Load(model)
# SentencePiece를 활용한 sentence -> encoding
tokensIDs = s.EncodeAsIds('아버지가방에들어가신다.')
print(tokensIDs)
# SentencePiece를 활용한 sentence -> encoded pieces
print(s.SampleEncodeAsPieces('아버지가방에들어가신다.',1, 0.0))
# SentencePiece를 활용한 encoding -> sentence 복원
print(s.DecodeIds(tokensIDs))
print('kor_spm_unigram_8k')
test_performance(model='kor_spm_unigram_8k.model')
# # unigram은 오류가 나서 학습이 되지 않았다.
# print('kor_spm_unigram_4k')
# test_performance(model='kor_spm_unigram_4k.model')
print('kor_spm_bpe_8k')
test_performance(model='kor_spm_bpe_8k.model')
print('kor_spm_bpe_16k')
test_performance(model='kor_spm_bpe_16k.model')
```
## 3. SPM을 받을 수 있는 Tokenizer 함수 작성
#### 훈련시킨 SentencePiece를 활용해 tokenizer역할을 하는 함수를 정의합니다.
함수의 조건
1) 매개변수로 토큰화된 문장의 list를 전달하는 대신 온전한 문장의 list 를 전달합니다.
2) 생성된 vocab 파일을 읽어와 { \ : \ } 형태를 가지는 word_index 사전과 { \ : \} 형태를 가지는 index_word 사전을 생성하고 함께 반환합니다.
3) 리턴값인 tensor는 앞의 함수와 동일하게 토큰화한 후 Encoding된 문장입니다. 바로 학습에 사용할 수 있게 Padding은 당연히 해야겠죠?
```
def sp_tokenize(s, corpus, spm):
tensor = []
for sen in corpus:
tensor.append(s.EncodeAsIds(sen))
with open("./{}.vocab".format(spm), 'r') as f:
vocab = f.readlines()
word_index = {}
index_word = {}
for idx, line in enumerate(vocab):
word = line.split("\t")[0]
word_index.update({idx:word})
index_word.update({word:idx})
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor, padding='pre', maxlen=max_len)
return tensor, word_index, index_word
# sp_tokenize(s, corpus) 사용 예제
s = spm.SentencePieceProcessor()
# s.Load('kor_spm_bpe_16k.model') # lstm 학습시 오류
s.Load('kor_spm_unigram_8k.model')
my_corpus = ['나는 새싹 캠퍼스에 출근해서 일을 했습니다.', '하지만 ㅠㅠ 월급을 받지 못해서 밥을 굶었습니다...']
tensor, word_index, index_word = sp_tokenize(s, my_corpus, 'kor_spm_unigram_8k')
print(tensor)
# print(word_index)
# print(index_word)
model_uni_8k='kor_spm_unigram_8k'
model_uni_4k='kor_spm_unigram_4k'
model_bpe_8k='kor_spm_bpe_8k'
model_bpe_16k='kor_spm_bpe_16k'
```
## 훈련데이터 / 테스트 데이터 나누기
```
# document와 label을 나눠봅시다.
x_train,x_train_word_index,x_train_index_word = sp_tokenize(s, train_data['document'], model_uni_8k)
x_test,x_test_word_index, x_test_index_word = sp_tokenize(s, test_data['document'], model_uni_8k)
# 현재 list 상태 ==> ndarray로 바꿔주기
y_train = np.array(list(train_data['label']))
y_test = np.array(list(test_data['label']))
print(x_train[:3])
print(x_test[:3])
print(y_train[:3])
print(y_test[:3])
print("학습데이터 :",len(x_train))
print("타겟데이터 :",len(y_train))
from sklearn.model_selection import train_test_split
train_input, val_input, train_target, val_target = train_test_split(X_train, y_train, test_size=0.2, random_state=42)
print(train_input[:3])
print(val_input[:3])
```
## 4. lstm으로 감정분석 모델 학습
SentencePiece 토크나이저가 적용된 모델을 학습 후 수렴하는 것을 확인합니다.
SentencePiece 토크나이저 성능 평가
```
tensor, word_index, index_word = sp_tokenize(s, naver_review, 'kor_spm_unigram_8k')
from tensorflow.keras.initializers import Constant
vocab_size = vocab_size
word_vector_dim = 32
model_LSTM = keras.Sequential()
model_LSTM.add(keras.layers.Embedding(vocab_size,
word_vector_dim,
))
model_LSTM.add(keras.layers.LSTM(128))
# model_LSTM.add(keras.layers.Dense(128, activation='relu'))
model_LSTM.add(keras.layers.Dense(1, activation='sigmoid')) # 최종 출력은 긍정/부정을 나타내는 1dim 입니다.
model_LSTM.summary()
```
#### Earlystopping & Model Checkpoint
```
from tensorflow.keras.models import load_model
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
es = keras.callbacks.EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=4)
mc = ModelCheckpoint('best_model.h5', monitor='val_acc', mode='max', verbose=1, save_best_only=True)
model_LSTM.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
batch_size=64
history_LSTM = model_LSTM.fit(train_input,
train_target,
epochs=20,
batch_size=batch_size,
validation_data=(val_input, val_target),
callbacks=[es, mc],
verbose=1
)
results = model_LSTM.evaluate(x_test, y_test, verbose=2)
print(results)
def visualize_train(train_history, param):
# summarize history for accuracy
plt.plot(train_history.history['accuracy'])
plt.plot(train_history.history['val_accuracy'])
plt.title('{}_accuracy'.format(param))
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(train_history.history['loss'])
plt.plot(train_history.history['val_loss'])
plt.title('{}_loss'.format(param))
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
visualize_train(history_LSTM, param='unigram_8k_dim_8')
```
## 5. 모델 성능 비교 평가
비교 대상
- LSTM의 input x_data를 토크나이징한 SentencePiece 모델
- LSTM모델의 Word_vector_dimension
```
def performance_test_spm(spm_model, vocab_size, word_vector_dim):
s = spm.SentencePieceProcessor()
s.Load("{}.model".format(spm_model))
# document와 label을 나눠봅시다.
x_train,x_train_word_index,x_train_index_word = sp_tokenize(s, train_data['document'], model_uni_8k)
x_test,x_test_word_index, x_test_index_word = sp_tokenize(s, test_data['document'], model_uni_8k)
# 현재 list 상태 ==> ndarray로 바꿔주기
y_train = np.array(list(train_data['label']))
y_test = np.array(list(test_data['label']))
train_input, val_input, train_target, val_target = train_test_split(x_train, y_train, test_size=0.2, random_state=42)
model_LSTM = keras.Sequential()
model_LSTM.add(keras.layers.Embedding(vocab_size,
word_vector_dim,
))
model_LSTM.add(keras.layers.LSTM(128))
model_LSTM.add(keras.layers.Dense(1, activation='sigmoid')) # 최종 출력은 긍정/부정을 나타내는 1dim 입니다.
model_LSTM.summary()
model_LSTM.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
epochs=10
batch_size=64
es = keras.callbacks.EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=4)
mc = ModelCheckpoint('best_model.h5', monitor='val_acc', mode='max', verbose=1, save_best_only=True)
history_LSTM = model_LSTM.fit(train_input,
train_target,
epochs=20,
batch_size=batch_size,
validation_data=(val_input, val_target),
callbacks=[es, mc],
verbose=1
)
results = model_LSTM.evaluate(x_test, y_test, verbose=2)
print(results)
return history_LSTM
## 모델에서 변수들의 이름이 섞여서 (train_target 과 x_train 등) 애를 많이 먹었다. 결국 이를 고쳐준 나의 승리!
history_unigram_8k_dim_8 = performance_test_spm('kor_spm_unigram_8k', vocab_size, word_vector_dim=8)
history_unigram_8k_dim_64 = performance_test_spm('kor_spm_unigram_8k', vocab_size, word_vector_dim=64)
visualize_train(history_unigram_8k_dim_8, param='unigram_8k_dim_8')
# 학습을 시킬 때 에러가 났다.
# history_bpe_16k_dim_32 = performance_test_spm('kor_spm_bpe_16k', vocab_size, word_vector_dim=32)
# 아래는 에러코드입니다.
# InvalidArgumentError: 2 root error(s) found.
# (0) Invalid argument: indices[60,129] = 15311 is not in [0, 8000)
# [[node sequential_7/embedding_7/embedding_lookup (defined at <ipython-input-213-39b87160ea35>:44) ]]
# [[sequential_7/embedding_7/embedding_lookup/_20]]
# (1) Invalid argument: indices[60,129] = 15311 is not in [0, 8000)
# [[node sequential_7/embedding_7/embedding_lookup (defined at <ipython-input-213-39b87160ea35>:44) ]]
# 0 successful operations.
# 0 derived errors ignored. [Op:__inference_train_function_226789]
# Errors may have originated from an input operation.
# Input Source operations connected to node sequential_7/embedding_7/embedding_lookup:
# sequential_7/embedding_7/embedding_lookup/225557 (defined at /opt/conda/lib/python3.7/contextlib.py:112)
# Input Source operations connected to node sequential_7/embedding_7/embedding_lookup:
# sequential_7/embedding_7/embedding_lookup/225557 (defined at /opt/conda/lib/python3.7/contextlib.py:112)
# Function call stack:
# train_function -> train_function
visualize_train(history_unigram_8k_dim_64, param='unigram_8k_dim_64')
visualize_train(history_bpe_8k_dim_8, param='bpe_8k_dim_8')
# 왠지 모르지만 16k는 학습할 때 에러가 났다.
# visualize_train(history_bpe_16k_dim_64, param='bpe_8k_dim_64')
```
## 회고 및 결론
original unigram_8k
- 1245/1245 - 10s - loss: 0.4736 - accuracy: 0.8194
- [0.47356146574020386, 0.8193917870521545]
#### Best result
uningram_8k, vocab = 8
- 1245/1245 - 10s - loss: 0.4242 - accuracy: 0.8188
- [0.42419102787971497, 0.8188393115997314]
unigram_8k, vocab = 64
- 1245/1245 - 10s - loss: 0.4325 - accuracy: 0.8247
- [0.43249306082725525, 0.8247156143188477]
uningram_8k, vocab = 8의 모델이 가장 학습이 잘 되었다.
위에서 공유한 두 개의 링크
- https://piaojian.tistory.com/35,
- https://github.com/miinkang/AI_Project_AIFFEL/blob/main/%5BGD-02%5DSentencePiece.ipynb)
를 참고하면서 코드를 작성해봤다. 첫 번재의 링크는 모델을 간결하게 쓰고, callbacks, modelcheckpoint등을 사용해서 좋았고, 두 분째의 링크는 모델의 종류(unigram, bpe)와 workd_vector dimension을 여러가지로 바꾸어 보는 시도와 여러가지 전처리등을 활용했기에 둘 다 합치면 더 좋은 결과가 나올거라고 생각했다. 하지만... 나의 한계인지 bpe모델들은 무언가가 계속 에러가 났다.
다음 기회에는 이 sentence piece를 활용해서 챗봇에 적용해보고 싶다.
흥미로웠던 점은 word_vector의 차원이 올라간다고 해서 무조건 모델의 정확도가 올라가는 것은 아니라는 것이다.
또한, 전처리 과정에서 너무 길거나 짧은 글자들을 제거해준다는 점 (EXP07에서도 활용)을 보았을시에 전처리를 어떻게 하는가, 그리고 어떠한 토크나이저를 사용하는가가 중요한 것을 알 수 있었다.
| github_jupyter |
# Verify Bayesian optimization standard deviation reflects plot size fairly
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from numpy.random import seed
# GPR
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, Matern, DotProduct, ConstantKernel, WhiteKernel
%matplotlib inline
# Definition of a black-box function to test
def f_y(x) :
y = -1 * (x * np.sin(x * np.pi/30.0) + 100 + np.random.normal(0,1,1))
return y
# Draw black-box function
x = np.arange(0, 180, 0.1)
y = f_y(x)
plt.plot(x, y, c = 'r')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
# Make samples to make predisction model within the range from 1 to 180, while missing samples from 60 to 100, with the step of 5
```
x1 = np.arange(0, 60, 5)
x2 = np.arange(100, 180, 5)
x = np.hstack((x1, x2))
x
# Draw sample distribution
y = f_y(x)
plt.scatter(x, y, c = 'r')
#plt.axvline(x = 0, ymin = -250, ymax = 80)
#plt.axvline(x = 60, ymin = -250, ymax = 80)
#plt.axvline(x = 100, ymin = -250, ymax = 80)
#plt.axvline(x = 180, ymin = -250, ymax = 80)
plt.axvspan(-20, 0, color = "gray", alpha = 0.3) # samples are missing in the gray are
plt.axvspan(60, 100, color = "gray", alpha = 0.3)
plt.axvspan(180, 200, color = "gray", alpha = 0.3)
plt.xlabel('x')
plt.ylabel('y')
plt.xlim(-20, 200)
plt.show()
```
# Make a model using Gaussian process regression
```
# kernel
kernel = ConstantKernel() * RBF() + WhiteKernel() + ConstantKernel() * DotProduct()
# Make a model
x = x.reshape(-1,1)
y = y.reshape(-1,1)
model = GaussianProcessRegressor(kernel = kernel).fit(x, y)
model
```
# Overlay graph
```
plt.scatter(x, model.predict(x), s = 100, alpha = 0.3, label = 'prediction')
plt.scatter(x, y, c = 'r', marker = 'x', label = 'original data')
plt.legend(loc = 'best')
plt.show()
```
# Plot prediction over the true curve including extrapolation renge, reflecting standard deviation as a circle size (large circle = small standard deviation) at the sampling points
```
x_test = np.arange(-100, 300, 3).reshape(-1,1)
pred_y_mean, pred_y_std = model.predict(x_test, return_std = True)
fig = plt.figure(figsize=(12, 5))
std_max = pred_y_std.max()
std_min = pred_y_std.min()
m_size = 10 + 300 * (std_max - pred_y_std) / (std_max - std_min)
plt.scatter(x_test, pred_y_mean, c = 'r', marker = 'o', alpha = 0.3, s = m_size)
plt.scatter(x_test, f_y(x_test), c = 'b', marker = 'o', alpha = 0.5)
plt.xlabel('x')
plt.ylabel('y')
plt.axvspan(-120, 0, color = "gray", alpha = 0.3) # samples are missing in the gray are
plt.axvspan(60, 100, color = "gray", alpha = 0.3)
plt.axvspan(180, 320, color = "gray", alpha = 0.3)
plt.xlim(-120, 320)
plt.show()
fig = plt.figure(figsize=(6, 6))
slp_1_begin = 1.1 * pred_y_mean.min()
slp_1_end = 1.1 * pred_y_mean.max()
plt.scatter(f_y(x_test), pred_y_mean, c = 'r', s = m_size, alpha = 0.5)
plt.plot([slp_1_begin, slp_1_end], [slp_1_begin, slp_1_end], c = 'b')
plt.title('Fitting check')
plt.xlabel('Observed value')
plt.ylabel('Predicted value')
plt.show()
```
# Circle size is determined by a mini-max concept using standard deviation from GPR, and it looks working well.
| github_jupyter |
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
from IPython.display import HTML, IFrame
display(IFrame(src = "http://pandas.pydata.org/_static/pydata_cover.jpg", width=600, height=700))
```
## Material in this notebook is from the Appendix of:
Python for Data Analysis
Data Wrangling with Pandas, NumPy, and IPython
By Wes McKinney
Publisher: O'Reilly Media
Exercises:
http://www.w3resource.com/python-exercises/
http://www.practicepython.org/
http://codingbat.com/python
# Python Introduction
Python is an interpreted language.
The Python interpreter runs a program by executing
one statement at a time. The standard interactive Python interpreter can be invoked on
the command line with the python command:
The >>> you see is the prompt where you’ll type expressions. To exit the Python interpreter
and return to the command prompt, you can either type exit() or press Ctrl-D.
Running Python programs is as simple as calling python with a .py file as its first argument.
Suppose we had created hello_world.py with these contents:
This can be run from the terminal simply as:
While many Python programmers execute all of their Python code in this way, many
scientific Python programmers make use of IPython, an enhanced interactive Python
interpreter.
By using the %run command,
IPython executes the code in the specified file in the same process, enabling you to
explore the results interactively when it’s done.
The default IPython prompt adopts the numbered In [2]: style compared with the
standard >>> prompt.
# Jupyter Notebook Environment
One considerable advantage that Python offers is a user-friendly interactive development environment. This environment is called the Jupyter Notebook. We will mostly be working in this environment during or lectures/workshops and you will also be completing your assignments in this environment.
In order to start an Jupyter Notebook session, you will simply need to double click one of your notebooks (file extension .ipynb) and this will launch your default browser if 'jupyter-notebook.exe' is on the path. Otherwise, you can also drag the .ipynb file on to the executable.
The Jupyter Notebook is a remarkable piece of technology. It is essentially an **interactive computational environment accessed through a web browser** (running on a local machine), which allows executable code to be combined with plots, images, mathematics and rich media.
Its primary aim is to facilitate agile data analysis and exploratory computation. Its exceptional achievement is found in its ability to **support reproducible and verifiable research**, since all inputs and outputs may be stored in notebook documents.
The notebook application runs as a lightweight server process on the command line. The actual notebooks running in a web browser are divided into 'cells'. Cells which have 'In []:' in front of them are as below are 'executable'. This means that if you press down the 'Shift' button and press 'Enter', the code that is in the adjacent sell will be executed and the output of that code will be displayed in a cell bellow.
<img src=http://jupyter.org/assets/main-logo.svg width=400 height=500>
The notebook server process can be started on the command line by running:
On most platforms, your primary web browser will automatically open up to the notebook
dashboard. In some cases you may have to navigate to the listed URL. From there,
you can create a new notebook and start exploring.
## Basics
## Language semantics
The Python language design is distinguished by its emphasis on readability, simplicity,
and explicitness. Some people go so far as to liken it to “executable pseudocode”.
### **Indentation, not braces**
Python uses whitespace indentation (tabs or spaces)to delimit blocks; this is termed the **off-side rule**.
Take the for loop in the below quicksort algorithm:
A colon denotes the start of an indented code block after which all of the code must be
indented by the same amount until the end of the block. In another language, you might
instead have something like:
### **Everything is an object**
An important characteristic of the Python language is the consistency of its object
model. **Every number, string, data structure, function, class, module, and so on exists
in the Python interpreter in its own “box” which is referred to as a Python object**. Each
object has an associated type (for example, string or function) and internal data. In
practice this makes the language very flexible, as even functions can be treated just like
any other object.
# **Comments**
Any text preceded by the hash mark (pound sign) # is ignored by the Python interpreter.
This is often used to add comments to code. At times you may also want to exclude
certain blocks of code without deleting them. An easy solution is to comment out the
code:
# Arithmetic Operators
<table class="src">
<tbody><tr>
<th style="width:5%">Operator</th><th>Description</th><th>Example (If a = 10 and b = 20)</th>
</tr>
<tr>
<td>+</td><td>Addition - Adds values on either side of the operator</td><td> a + b will give 30</td>
</tr>
<tr>
<td>-</td><td>Subtraction - Subtracts right hand operand from left hand operand</td><td> a - b will give -10</td>
</tr>
<tr>
<td>\*</td><td>Multiplication - Multiplies values on either side of the operator</td><td> a \* b will give 200</td>
</tr>
<tr>
<td>/</td><td>Division - Divides left hand operand by right hand operand</td><td> b / a will give 2</td>
</tr>
<tr>
<td>%</td><td>Modulus - Divides left hand operand by right hand operand and returns remainder</td><td> b % a will give 0</td>
</tr>
<tr>
<td> \*\* </td><td>Exponent - Performs exponential (power) calculation on operators</td><td> a\**b will give 10 to the power 20</td>
</tr>
<tr>
<td>//</td><td>Floor Division - The division of operands where the result is the quotient in which the digits
after the decimal point are removed.</td><td> 9//2 is equal to 4 and 9.0//2.0 is equal to 4.0</td>
</tr>
</tbody></table>
# Scalar Types
Python has a small set of built-in types for handling numerical data, strings, boolean
(True or False) values, and dates and time. See Table A-2 for a list of the main scalar
types. Date and time handling will be discussed separately as these are provided by the
datetime module in the standard library.
## **Numeric types**
The primary Python types for numbers are int and float. The size of the integer which
can be stored as an int is dependent on your platform (whether 32 or 64-bit), but Python
will transparently convert a very large integer to long, which can store arbitrarily large
integers.
```
ival = 17239871
ival ** 6
```
Floating point numbers are represented with the Python float type. Under the hood
each one is a double-precision (64 bits) value.
Careful with division. In Python 2 - two ints in a division will return an int. In Python 3, two ints in a division will return a float:
```
3 / 2
```
In Python 2.7 and below (which some readers will likely be using), you can enable floating point division using /
by default, by putting the following cryptic-looking statement at the top of
your module:
```
from __future__ import division
3 / 2
```
Without this in place, you can always explicitly convert the denominator into a floating point number:
```
3 / float(2)
```
To get C-style integer division (which drops the fractional part if the result is not a
whole number), use the floor division operator //:
```
3 // 2.0
```
## **Dynamic references, strong types**
Python is a **dynamically-typed language**. In contrast with many compiled languages, such as Java and C++, object references in
Python have no type associated with them.
An assignment statement in Python creates new variables and gives them values.
There is no problem with the following:
```
a = 5
```
In Python the assignment, using the above, translates to "(generic) name *a* receives a reference to a separate, dynamically allocated object of numeric (int) type of value 5."
This is referred to as "binding" the name to the object.
```
type(a)
a = 'blah'
type(a)
vv = 0o10
vv
```
Variables are names for objects within a particular namespace; the type information is
stored in the object itself.
Some observers might hastily conclude that Python is not a
**“typed language”**. This is not true; consider this example:
```
'5' + 5
```
In some languages, such as Visual Basic, the string '5' might get implicitly converted
(or casted) to an integer, thus yielding 10. Yet in other languages, such as JavaScript,
the integer 5 might be casted to a string, yielding the concatenated string '55'.
In this
regard **Python is also considered a strongly-typed language**, which means that every object
has a specific type (or class), and implicit conversions will occur only in certain obvious
circumstances, such as the following:
```
a = 4.5
b = 2
# String formatting, to be visited later
print ('a is %s, b is %s' % (type(a), type(b)))
a / b
```
Knowing the type of an object is important, and it’s useful to be able to write functions that can handle many different kinds of input. You can check that an object is an instance of a particular type using the isinstance function:
```
a = 5
isinstance(a, int)
```
isinstance can accept a tuple (more on tuples later) of types if you want to check that an object’s type is
among those present in the tuple:
```
a = 5; b = 4.5
isinstance(a, (int, float))
isinstance(b, (int, float))
```
# Functions
Functions are the primary and most important method of code organization and reuse
in Python.
There probably isn't such a thing as having too many functions. In fact, I would
argue that most programmers doing data analysis don’t write enough functions!
Functions are declared using the **def** keyword
and returned from using the **return** keyword:
```
def my_function(x, y, z=1.5):
if z > 1:
return z * (x + y)
else:
return z / (x + y)
```
There is no issue with having **multiple return statements**. If the end of a function is
reached without encountering a return statement, **None** is returned.
Each function can have some number of **positional arguments** and some number of
**keyword arguments**. Keyword arguments are most commonly used to specify default
values or optional arguments. In the above function, x and y are positional arguments
while z is a keyword argument. This means that it can be called in either of these
equivalent ways:
The main restriction on function arguments it that the **keyword arguments must follow
the positional arguments** (if any).
**You can specify keyword arguments in any order**;
this frees you from having to remember which order the function arguments were
specified in and only what their names are.
Almost every object in Python has attached functions, known as methods, that have
access to the object’s internal contents.
They can be called using the syntax:
### **Exercise**:
Write a function that finds the Max of three numbers.
```
#your code here:
def my_max(a, b, c): return max(a, b, c)
my_max(4,5,2)
```
# Functions and pass-by-reference Python style: pass-by-object-reference
When assigning a variable (or name) in Python, you are creating a reference to the object on the right hand side of the equals sign.
In practical terms, consider a list (more on lists later) of integers:
```
a = [1, 2, 3]
a
```
Suppose we assign *a* to a new variable *b*:
```
b = a
```
In some languages, this assignment would cause the data [1, 2, 3] to be copied. In
Python, *a* and *b* actually now refer to the same object, the original list [1, 2, 3]
You can prove this to yourself by appending an element to
*a* and then examining b:
```
a.append(4)
b
b.append(5)
a
```
What if we decide to change the object reference by assigning something different to b?
```
b = 'hmmm...what does this do to a?'
a
```
**In this instance, a new memory location is created for *b***.
Understanding the semantics of references in Python and when, how, and why data is
copied is especially critical when working with larger data sets in Python.
**When you pass objects as arguments to a function, you are only passing object references; no
copying occurs**.
This means that **a function can mutate the internals of its arguments**.
Suppose we had the following function:
```
def append_element(some_list, element):
some_list.append(element)
```
Then given what’s been said, this should not come as a surprise:
```
data = [1, 2, 3]
append_element(data, 4)
data
```
However, **attempting to changing the actual reference object in the function will result in a new memory location being created** and the callee object remaining unaltered:
```
def change_object_reference(some_list):
some_list = [10, 9, 8]
data = [1, 2, 3]
change_object_reference(data)
data
```
**Object state/contents is mutable when passed to functions, but the memory location is not.**
If you need object references changed, then wrap them up in another object and pass them to a function.
Python's argument passing model is sometimes called pass-by-object-reference, but is in essence pass-by-reference whereby the reference is passed by value.
## Namespaces, Scope, and Local Functions
Functions can access variables in two different scopes: **global** and **local**.
An alternate
and more descriptive name describing a variable scope in Python is a **namespace**.
Any
variables that are assigned within a function by default are assigned to the local namespace.
The local namespace is created when the function is called and immediately
populated by the function’s arguments.
After the function is finished, the local namespace
is destroyed (with some exceptions, see section on closures below). Consider the
following function:
```
def func():
a = []
for i in [1,2,3,4,5]:
a.append(i)
```
Upon calling func(), the empty list a is created, 5 elements are appended, then *a* is
destroyed when the function exits.
Suppose instead we had declared *a* outside of the function:
```
a = []
def func():
for i in [1,2,3,4,5]:
a.append(i)
func()
a
```
Now suppose we tried to change the object reference for *a*
inside the function:
```
a = []
def func2():
a = 'interesting...'
print(a)
func2()
print(a)
```
Assigning to global variables within a function is possible, but those variables must be
declared as global using the global keyword:
```
a = None
print(a)
def bind_a_variable():
global a
a = [1,2]
bind_a_variable()
print(a)
```
I generally discourage people from using the global keyword frequently.
Typically global variables are used to store some kind of state in a system.
If you find yourself using a lot of them, it’s probably a sign that
some object-oriented programming (using classes) is in order.
Functions can be declared anywhere, and there is no problem with having local functions
that are dynamically created when a function is called:
```
def outer_function(x, y, z):
def inner_function(a, b, c):
pass
pass
```
In the above code, the inner_function will not exist until outer_function is called. As
soon as outer_function is done executing, the inner_function is destroyed.
Nested inner functions can access the local namespace of the enclosing function, but
they cannot bind new variables in it. (See
closures.)
In a strict sense, all functions are local to some scope, that scope may just be the module
level scope.
## Returning Multiple Values
When I first programmed in Python after having programmed in C and C++, one of
my favorite features was the ability to return multiple values from a function. Here’s a
simple example:
```
def f():
a = 5
b = 6
c = 7
return a, b, c
a, b, c = f()
a
```
In data analysis and other scientific applications, you will likely find yourself doing this
very often as **many functions may have multiple outputs**, whether those are data structures
or other auxiliary data computed inside the function.
If you think about tuple
packing and unpacking, you will later realize that what’s happening
here is that the function is actually just returning one object, namely a tuple,
which is then being unpacked into the result variables.
In the above example, we could
have done instead:
```
return_value = f()
return_value
```
In this case, return_value would be, as you may guess, a 3-tuple with the three returned
variables. A potentially attractive alternative to returning multiple values like above
might be to return a dict (more on dictionaries later) instead:
```
def f():
a = 5
b = 6
c = 7
return {'a' : a, 'b' : b, 'c' : c}
```
# **Attributes and methods** (Object-Oriented Programming)
Objects in Python typically have both **attributes** (other Python objects stored “inside”
the object) and **methods**, functions associated with an object which can have access to
the object’s internal data.
Both of them are accessed via the syntax obj.attribute_name:
## Rules for naming variables
Every language has rules for naming variables. The rules in Python are the following:
A valid variable name is a non-empty sequence of characters of any length with:
1. the start character can be the underscore "_" or a capital or lower case letter
2. the letters following the start character can be anything which is permitted as a start character plus the digits.
3. Just a warning for Windows-spoilt users: variable names are case-sensitive
4. Python keywords are not allowed as variable names
<p>The following list shows the keywords in Python. These reserved words may not be used as constant or variable or any other identifier names.</p>
<table class="src">
<th>
<tr><td>and</td><td>exec</td><td>not</td></tr>
<tr><td>assert</td><td>finally</td><td>or</td></tr>
<tr><td>break</td><td>for</td><td>pass</td></tr>
<tr><td>class</td><td>from</td><td>print</td></tr>
<tr><td>continue</td><td>global</td><td>raise</td></tr>
<tr><td>def</td><td>if</td><td>return</td></tr>
<tr><td>del</td><td>import</td><td>try</td></tr>
<tr><td>elif</td><td>in</td><td>while</td></tr>
<tr><td>else</td><td>is</td><td>with </td></tr>
<tr><td>except</td><td>lambda</td><td>yield</td></tr>
<th>
</table>
# Control Flow and Logic Operators
<img src=http://www.principledpolicy.com/wp-content/uploads/2010/10/logic.jpg width=500>
<table class="src">
<tbody><tr>
<th style="width:5%">Operator</th><th>Description</th><th>Example (If a = 10 and b = 20)</th>
</tr>
<tr>
<td>==</td><td> Checks if the value of two operands are equal or not, if yes then condition becomes true.</td><td> (a == b) is not true. </td>
</tr>
<tr>
<td>!=</td><td> Checks if the value of two operands are equal or not, if values are not equal then condition becomes true.</td><td> (a != b) is true. </td>
</tr>
<tr>
<td><></td><td> Checks if the value of two operands are equal or not, if values are not equal then condition becomes true.</td><td> (a <> b) is true. This is similar to != operator.</td>
</tr>
<tr>
<td>></td><td> Checks if the value of left operand is greater than the value of right operand, if yes then condition becomes true.</td><td> (a > b) is not true. </td>
</tr>
<tr>
<td><</td><td> Checks if the value of left operand is less than the value of right operand, if yes then condition becomes true.</td><td> (a < b) is true. </td>
</tr>
<tr>
<td>>=</td><td> Checks if the value of left operand is greater than or equal to the value of right operand, if yes then condition becomes true.</td><td> (a >= b) is not true. </td>
</tr>
<tr>
<td><=</td><td> Checks if the value of left operand is less than or equal to the value of right operand, if yes then condition becomes true.</td><td> (a <= b) is true. </td>
</tr>
<th>
</tbody></table>
<img src=http://calwatchdog.com/wp-content/uploads/2013/05/Spock-logic-300x240.jpg >
Logical operators enable us to construct more complex boolean expressions.
<table class="src">
<tbody><tr>
<th style="width:5%">Operator</th><th>Description</th><th>Example (If a = True and b = True)</th>
</tr>
<tr>
<td>and</td><td> Called Logical AND operator. If both the operands are true then then condition becomes true.</td><td> (a and b) is true.</td>
</tr>
<tr>
<td>or</td><td>Called Logical OR Operator. If any of the two operands are non zero then then condition becomes true.</td><td> (a or b) is true.</td>
</tr>
<tr><td>not</td><td>Called Logical NOT Operator. Use to reverses the logical state of its operand. If a condition is true then Logical NOT operator will make false.</td><td> not(a and b) is false. </td>
</tr>
<th>
</tbody>
</table>
---
# **Binary operators and comparisons**
Most of the binary math operations and comparisons are as you might expect.
```
5 >= 2
```
To check if two references refer to the same object, use the **is** keyword. **is not** is also
perfectly valid if you want to check that two objects are not the same:
```
a = [1, 2, 3]
b = a
a is b
# Note, the list function always creates a new list
c = list(a)
print(c)
a is not c
```
Note this is not the same thing is comparing with ==, because in this case we have:
```
a == c
```
In the above, the actual contents are being compared, which are in this case the same, though the objects are different (two different memory locations are involved).
A very common use of **is** and **is not** is to check if a variable is **None**, since there is only one instance of **None**:
```
a = None
a is None
```
# **if, elif, and else**
The if statement is one of the most well-known control flow statement types. It checks
a condition which, if True, evaluates the code in the block that follows:
An if statement can be optionally followed by one or more elif (contraction for *else if*) blocks and a catch-all
else block if all of the conditions are False:
If any of the conditions is True, no further elif or else blocks will be reached. With a
compound condition using and or or, conditions are evaluated left-to-right and will
short circuit:
```
a = 5; b = 7
c = 8; d = 4
if a < b or c > d:
print('Made it')
```
In this example, the comparison c > d never gets evaluated because the first comparison
was True.
### **Exercise**:
Write a function that takes three integers and returns their sum. However, if two values are equal sum will be zero.
```
def sum_it(x,y,z):
if x == y or x == z or y == z:
return 0
return x + y + z
```
# **for loops**
for loops are for iterating over a collection (like a list or tuple) or an iterator. The
standard syntax for a for loop is:
A for loop can be advanced to the next iteration, skipping the remainder of the block,
using the continue keyword. Consider this code which sums up integers in a list and
skips None values:
```
sequence = [1, 2, None, 4, None, 5]
total = 0
for value in sequence:
print (value)
if value is None:
continue
total += value
print ("total:", total)
```
A for loop can be exited altogether using the break keyword. This code sums elements
of the list until a 5 is reached:
```
sequence = [1, 2, 0, 4, 6, 5, 2, 1]
total_until_5 = 0
for value in sequence:
if value == 5:
break
total_until_5 += value
```
As we will see in more detail, if the elements in the collection or iterator are sequences
(tuples or lists, say), they can be conveniently unpacked into variables in the for loop
statement:
### **Exercise**:
Given an array of ints and a target int, return the number of specified ints in the argument that are found in the array.
eg. array_count([1, 9, 9, 3, 9], 9) → 3
```
def array_count(my_array, target):
count = 0
for n in my_array:
if n == target:
count += 1
return count
array_count([1, 9, 9, 3, 9], 9)
```
# **range**
The range function produces a list of evenly-spaced integers:
```
range(5)
```
This can be seen more readily if we cast the range object to a list:
```
list(range(5))
list(range(0, 10, 3))
```
As you can see, range produces integers up to but not including the endpoint. A common
use of range is for iterating through sequences by index:
```
seq = [1, 2, 3, 4]
for i in range(len(seq)):
val = seq[i]
print (val)
```
Range in python 3 behaves the same as xrange in python 2 - it returns an iterator that generates integers one by one rather than generating all of them up-front and storing them in a (potentially very large) list.
### Exercise:
Generate a range of numbers from 100 to 200 at intervals of 5:
### **Exercise:**
Given an array of ints, return True if the sequence.. 1, 2, 3, .. appears in the array somewhere.
array123([1, 1, 2, 3, 1]) → True
```
def array123(nums):
array123([1, 2, 3])
array123([1, 1, 2, 3, 1])
```
### **Exercise**:
Write function to creates a histogram for * characters for each int from a given list of integers in the format below.
```
def histogram( items ):
histogram([2, 3, 6, 10, 5, 2])
```
# **while loops**
A while loop specifies a condition and a block of code that is to be executed until the
condition evaluates to False or the loop is explicitly ended with break:
```
x = 256
total = 0
while x > 0:
if total > 500:
break
total += x
x = x // 2
```
# **pass**
pass is the “no-op” statement in Python. It can be used in blocks where no action is to
be taken; it is only required because Python uses whitespace to delimit blocks:
```
if x < 0:
print ('negative!')
elif x == 0:
# TODO: put something smart here
pass
else:
print ('positive!')
```
It’s common to use pass as a place-holder in code while working on a new piece of
functionality:
```
def f(x, y, z):
# TODO: implement this function!
pass
```
# **Strings**
Many people use Python for its powerful and flexible built-in string processing capabilities. **You can write string literals using either single quotes ' or double quotes "**:
```
a = 'one way of writing a string'
b = "another way"
```
For multiline strings with line breaks, you can use triple quotes, either ''' or """:
```
c = """
This is a longer string that
spans multiple lines
"""
c
```
Many Python objects can be converted to a string using the str function:
```
a = 5.6
s = str(a)
s
```
Strings are a sequence of characters and therefore can be treated like other sequences,
such as lists and tuples:
```
s = 'python'
list(s)
```
The backslash character \ is an escape character, meaning that it is used to specify
special characters like newline \n or unicode characters. To write a string literal with backslashes, you need to escape them:
```
s = '12\445'
print (s)
print (type(s))
s = '12\\445'
print (s)
print (type(s))
```
If you have a string with a lot of backslashes and no special characters, you might find this a bit annoying. Fortunately you can preface the leading quote of the string with **r** which means that the characters should be interpreted as is:
```
s = r'this\has\no\special\characters'
print (s)
print (type(s))
```
Adding two strings together concatenates them and produces a new string:
```
a = 'this is the first half '
b = 'and this is the second half'
a + b
```
String templating or formatting is another useful topic. The number of ways to do
so has expanded with the advent of Python 3, here I will briefly describe the mechanics
of one of the main interfaces.
Strings with a % followed by one or more format characters is a target for inserting a value into that string (this is quite similar to the printf function in C).
As an example, consider this string:
```
template = '%.2f %s are worth $%d'
```
In this string, %s means to format an argument as a string, %.2f a number with 2 decimal places, and %d an integer. To substitute arguments for these format parameters, use the binary operator % with a tuple of values:
```
template % (4.5560, 'Argentine Pesos', 1)
```
String formatting is a broad topic; there are multiple methods and numerous options
and tweaks available to control how values are formatted in the resulting string.
To learn more, I recommend you seek out more information on the web.
Almost all built-in Python constructs and any class defining the \__nonzero__ magic method which have a True or False interpretation in an if statement:
```
a = [1, 2, 3]
if a:
print ('I found something!')
b = []
if not b:
print ('Empty!')
```
Most objects in Python have a notion of true- or falseness. For example, empty sequences (lists, dicts, tuples, etc.) are treated as False if used in control flow (as above with the empty list b).
You can see exactly what boolean value an object coerces to by invoking bool on it:
```
bool([]), bool([1, 2, 3])
bool('Hello world!'), bool('')
bool(0), bool(1)
```
# **Type casting**
The str, bool, int and float types are also functions which can be used to cast values
to those types:
```
s = '3.14159'
type(s)
fval = float(s)
type(fval)
```
# **None**
None is the Python null value type. If a function does not explicitly return a value, it implicitly returns None.
```
a = None
a is None
b = 5; b is not None
```
None is also a common default value for optional function arguments:
```
def add_and_maybe_multiply(a, b, c=None):
result = a + b
if c is not None:
result = result * c
return result
```
While a technical point, it’s worth bearing in mind that None is not a reserved keyword
but rather a unique instance of NoneType.
# **Dates and times**
The built-in Python datetime module provides datetime, date, and time types.
The
datetime type as you may imagine combines the information stored in date and time
and is the most commonly used:
```
from datetime import datetime, date, time
dt = datetime(2011, 10, 29, 20, 30, 21)
dt.day
dt.minute
```
Given a datetime instance, you can extract the equivalent date and time objects by
calling methods on the datetime of the same name:
```
dt.date()
dt.time()
```
The strftime method formats a datetime as a string:
```
dt.strftime('%m/%d/%Y %H:%M')
```
Strings can be converted (parsed) into datetime objects using the strptime function:
```
datetime.strptime('20091031', '%Y%m%d')
```
When aggregating or otherwise grouping time series data, it will occasionally be useful
to replace fields of a series of datetimes, for example replacing the minute and second
fields with zero, producing a new object:
```
print (dt)
print(dt.replace(minute=0, second=0))
```
The difference of two datetime objects produces a datetime.timedelta type:
```
dt2 = datetime(2011, 11, 15, 22, 30)
delta = dt2 - dt
print (type(delta))
print (delta)
```
Adding a timedelta to a datetime produces a new shifted datetime:
```
dt
dt + delta
```
### Exercise:
Write a script to output exactly how long ago Christmas last year was.
```
diff =
diff
print (diff.days)
print (diff.seconds)
print (diff.microseconds)
print (diff.total_seconds())
```
# **Mutable and immutable objects**
Most objects in Python are mutable, such as lists, dicts, NumPy arrays, or most userdefined types (classes). This means that the object or values that they contain can be modified. Others, like strings and tuples, are immutable:
```
a_tuple = (3, 5, (4, 5))
print (a_tuple[1])
a_tuple[1] = 'four'
a = 'this is a string'
a[10] = 'f'
```
# **Imports**
In Python a module is simply a .py file containing function and variable definitions
along with such things imported from other .py files. Suppose that we had the following
module:
If we wanted to access the variables and functions defined in some_module.py, from
another file in the same directory we could do:
Or equivalently:
By using the **as** keyword you can give imports different variable names:
| github_jupyter |
```
import nltk
import string
import numpy as np
%matplotlib inline
from nltk import word_tokenize
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
from sklearn import metrics
import warnings
warnings.filterwarnings("ignore")
enstop = stopwords.words('english')
punct = string.punctuation
def tokenizer(sent):
sent = sent.lower()
tmp = word_tokenize(sent)
res = []
for word in tmp:
if word not in enstop and word not in punct:
res.append(word)
return res
```
### IMDB data read:
```
import torch
import torch.nn as nn
from torchtext import data
from torchtext import vocab
text_field = data.Field(tokenize=tokenizer, lower=True, include_lengths=True, fix_length=256)
label_field = data.Field(sequential=False, use_vocab=False, dtype=torch.long)
train, valid, test = data.TabularDataset.splits(path='',
train='./PHEME/2018_train.csv',
validation='./PHEME/2018_val.csv',
test='./PHEME/2018_test.csv',
format='csv', skip_header=True,
fields=[('id', None),('sentence', text_field), ('label', label_field)])
```
### Read the GloVe word vector:
```
vec = vocab.Vectors(name='glove.6B.300d.txt')
text_field.build_vocab(train, valid, test, max_size=250000, vectors=vec,
unk_init=torch.Tensor.normal_)
label_field.build_vocab(train, valid, test)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iter, valid_iter, test_iter = data.BucketIterator.splits((train, valid, test), batch_sizes=(128, 128, 128),
sort_key=lambda x: len(x.sentence),
sort_within_batch=True,
repeat=False, shuffle=True,
device=device)
```
### Model training function (standard training and DropAttack adversarial training)
```
def train(model, train_iter, dev_iter, num_epoch, opt, criterion, eva, out_model_file):
print("Training begin!")
model.train()
loss_list = []
dev_acc = []
train_acc = []
best_dev_acc = 0.
for epoch in range(num_epoch):
total_loss = 0.
for batch in train_iter:
output = model(batch.sentence)
loss = criterion(output, batch.label)
opt.zero_grad()
loss.backward()
opt.step()
total_loss += loss.item()
loss_list.append(total_loss)
dev_acc.append(eva(model, dev_iter))
train_acc.append(eva(model,train_iter))
print(f"Epoch: {epoch+1}/{num_epoch}. Total loss: {total_loss:.3f}.Train_Acc: {train_acc[-1]:.3%}. Validation Set Acc: {dev_acc[-1]:.3%}.")
if dev_acc[-1] > best_dev_acc:
best_dev_acc = dev_acc[-1]
torch.save(model.state_dict(), out_model_file)
return loss_list, dev_acc
# import torch
class DropAttack():
def __init__(self, model):
self.model = model
self.param_backup = {}
self.grad_backup = {}
self.mask_backup = {}
def attack(self, epsilon=5.0, p_attack =0.5, param_name='embed.weight', is_first_attack=False):
# The emb_name parameter should be replaced with the name of the parameter to be attacked in your model
for name, param in self.model.named_parameters():
if param.requires_grad and param_name == name:
if is_first_attack:
self.param_backup[name] = param.data.clone()
mask = np.random.binomial(n=1, p=p_attack, size= param.grad.shape)
mask = torch.from_numpy(mask).float() # attack mask
self.mask_backup['mask'] = mask.clone()
else: mask = self.mask_backup['mask']
norm = torch.norm(param.grad)
if norm != 0 and not torch.isnan(norm):
r_at = epsilon * param.grad / norm
r_at *= mask.cuda() # Randomly attack some of the parameters
param.data.add_(r_at)
def restore(self, param_name='embed.weight'):
for name, param in self.model.named_parameters():
if param.requires_grad and param_name == name:
assert name in self.param_backup
param.data = self.param_backup[name]
param_backup = {}
def backup_grad(self):
for name, param in self.model.named_parameters():
if param.requires_grad:
self.grad_backup[name] = param.grad.clone()
def restore_grad(self):
for name, param in self.model.named_parameters():
if param.requires_grad:
param.grad = self.grad_backup[name]
grad_backup = {}
def train_DA(model, train_iter, dev_iter, num_epoch, opt, criterion, eva, out_model_file):
K = 1
print(f'Adversarial training begin! (DropAttack-{K})')
model.train()
dropattack = DropAttack(model)
loss_list = []
dev_acc = []
best_dev_acc = 0.
for epoch in range(num_epoch):
total_loss = 0.
model.train()
for batch in train_iter:
output = model(batch.sentence)
loss = criterion(output, batch.label)
loss.backward(retain_graph=True) # Calculate the original gradient
dropattack.backup_grad() # Backup the initial gradient
# Attack the embedding layer
for t in range(K):
dropattack.attack(5, 0.5, 'embed.weight', is_first_attack=(t==0)) # Add adversarial disturbance to the parameters, backup param.data for the first attack
output = model(batch.sentence)
loss_adv1 = criterion(output, batch.label)/K
loss_adv1.backward(retain_graph=True) # # Backpropagation, and accumulate the gradient of the adversarial training based on the normal grad
loss += loss_adv1
dropattack.restore('embed.weight') # # Restore the disturbed parameters
dropattack.restore_grad()
# Attack the hidden layer
for t in range(K):
dropattack.attack(5, 0.5, 'rnn.rnn.weight_hh_l0', is_first_attack=(t==0)) # Add adversarial disturbance to the parameters, backup param.data for the first attack
output = model(batch.sentence)
loss_adv2 = criterion(output, batch.label)/K
loss_adv2.backward(retain_graph=True) # Backpropagation, and accumulate the gradient of the adversarial training based on the normal grad
loss += loss_adv2
dropattack.restore('rnn.rnn.weight_hh_l0') # Restore the disturbed parameters
opt.zero_grad()
loss.backward()
opt.step() # Update parameters
# loss = loss + loss_adv1 + loss_adv2
total_loss += loss.item()
opt.zero_grad()
loss_list.append(total_loss)
dev_acc.append(eva(model, dev_iter))
print(f"Epoch: {epoch+1}/{num_epoch}. Total loss: {total_loss:.3f}. Validation Set Acc: {dev_acc[-1]:.3%}.")
if dev_acc[-1] > best_dev_acc:
best_dev_acc = dev_acc[-1]
torch.save(model.state_dict(), out_model_file)
return loss_list, dev_acc
```
### Model evaluation function
```
def eva(model, data_iter):
correct, count = 0, 0
with torch.no_grad():
for batch in data_iter:
pred = model(batch.sentence)
pred = torch.argmax(pred, dim=-1)
correct += (pred == batch.label).sum().item()
count += len(pred)
return correct / count
```
### LSTM model
```
class LSTM(nn.Module):
def __init__(self, input_size, hidden_size, num_layers, bidirectional):
super(LSTM, self).__init__()
self.rnn = nn.LSTM(input_size=input_size, hidden_size=hidden_size,
num_layers=num_layers, bidirectional=bidirectional)
def forward(self, x, length):
packed_x = nn.utils.rnn.pack_padded_sequence(x, length)
packed_output, (hidden, cell) = self.rnn(packed_x)
output, output_lengths = nn.utils.rnn.pad_packed_sequence(packed_output)
return hidden, output
class GRU(nn.Module):
def __init__(self, input_size, hidden_size, num_layers, bidirectional):
super(GRU, self).__init__()
self.rnn = nn.GRU(input_size=input_size, hidden_size=hidden_size,
num_layers=num_layers, bidirectional=bidirectional)
def forward(self, x, length):
packed_x = nn.utils.rnn.pack_padded_sequence(x, length)
packed_output, hidden = self.rnn(packed_x)
output, output_lengths = nn.utils.rnn.pad_packed_sequence(packed_output)
return hidden, output
class TextRNN(nn.Module):
def __init__(self, embed_size, hidden_size, num_layers, bidirectional, out_dim,
pretrained_embed, use_gru=False, freeze=True,
random_embed=False, vocab_size=None):
super(TextRNN, self).__init__()
if random_embed:
self.embed = nn.Embedding(vocab_size, embed_size)
else:
self.embed = nn.Embedding.from_pretrained(pretrained_embed, freeze=False)
if use_gru:
self.rnn = GRU(embed_size, hidden_size, num_layers, bidirectional)
else:
self.rnn = LSTM(embed_size, hidden_size, num_layers, bidirectional)
self.proj = nn.Linear(2*hidden_size, out_dim)
def forward(self, x):
text, text_length = x # text: [seq_len, bs]
text = text.permute(1, 0) # text: [bs, seq_len]
embed_x = self.embed(text) # embed_x: [bs, seq_len, embed_dim]
embed_x = embed_x.permute(1, 0, 2) # embed_x: [seq_len, bs, embed_dim]
hidden, _ = self.rnn(embed_x, text_length) # hidden: [2*num_layers, bs, hidden_size]
hidden = torch.cat((hidden[-1,:,:], hidden[-2,:,:]), dim=1)
return self.proj(hidden)
```
### Training
```
embed_size = 300
hidden_size = 300
num_layers = 1
bidirectional = True
out_dim = 2
pretrained_embed = text_field.vocab.vectors
lr = 0.001
num_epoch = 5
freeze = False
use_gru = True
random_embed = False
vocab_size = len(text_field.vocab.stoi)
out_model_file = 'textrnn_PHEME_DA.pt'
# ————————————————————————————————————————————————————————
use_dropattack = True # Whether to use DropAttack
# ————————————————————————————————————————————————————————
model = TextRNN(embed_size, hidden_size, num_layers, bidirectional, out_dim,
pretrained_embed, use_gru=use_gru, freeze=freeze,
random_embed=random_embed, vocab_size=None).to(device)
opt = torch.optim.Adam(model.parameters(), lr=lr)
criterion = nn.CrossEntropyLoss()
if use_dropattack:
loss_list, dev_acc_list = train_DA(model, train_iter, valid_iter, num_epoch, opt, criterion, eva, out_model_file)
else:
loss_list, dev_acc_list = train(model, train_iter, valid_iter, num_epoch, opt, criterion, eva, out_model_file)
model = TextRNN(embed_size, hidden_size, num_layers, bidirectional, out_dim,
pretrained_embed, use_gru=use_gru, freeze=freeze,
random_embed=random_embed, vocab_size=None).to(device)
model.load_state_dict(torch.load('textrnn_PHEME_DA.pt'))
print(f"Test set acc: {eva(model, test_iter):.3%}")
```
| github_jupyter |
## Delta Hedging
### Summary
Being short volatility hasn't been profitable in this period of extreme implied and realized volatility movements but may be an interesting entry point for some.
In this note I take a further look at this strategy and extend it with delta hedging to understand how it can impact performance.
Each day I sell a 1m10y straddle (like last time) - but this time I also trade a swap with a matched effective date and termination date to hedge my delta. Each day I unwind the previous day's swap and trade into a new one.
I examine premium collected at inception, payout on option expiry and mark-to-market over the life of the trade to compare the two strategies.
Look out for future publications where I will build on this analysis further by adding transaction costs and analyzing performance accross strategies.
The content of this notebook is split into:
* [1 - Let's get started with gs quant](#1---Let's-get-started-with-gs-quant)
* [2 - Create portfolio](#2---Create-portfolio)
* [3 - Grab the data](#3---Grab-the-data)
* [4 - Putting it all together](#4---Putting-it-all-together)
### 1 - Let's get started with gs quant
Start every session with authenticating with your unique client id and secret. If you don't have a registered app, create one [here](https://marquee.gs.com/s/developer/myapps/register). `run_analytics` scope is required for the functionality covered in this example. Below produced using gs-quant version 0.8.108.
```
from gs_quant.session import GsSession
GsSession.use(client_id=None, client_secret=None, scopes=('run_analytics',))
use_batch = True
```
### 2 - Create portfolio
Just like in our last analysis, let's start by creating a portfolio with a rolling strip of straddles. For each date in our date range (start of 2019 through today), we will construct a 1m10y straddle and include it in our portfolio.
```
from gs_quant.markets import HistoricalPricingContext, PricingContext
from gs_quant.markets.portfolio import Portfolio
from gs_quant.common import Currency, PayReceive
from gs_quant.instrument import IRSwaption
import datetime as dt
start_date = dt.datetime(2020, 12, 1).date()
end_date = dt.datetime.today().date()
# create and resolve a new straddle on every day of the pricing context
with HistoricalPricingContext(start=start_date, end=end_date, show_progress=True):
f = IRSwaption(PayReceive.Straddle, '10y', Currency.USD, expiration_date='1m',
notional_amount=1e8, buy_sell='Sell').resolve(in_place=False)
# put resulting swaptions in a portfolio
result = f.result().items()
portfolio = Portfolio([v[1] for v in sorted(result)])
```
I will now convert the portfolio to a dataframe, extend it with trade dates and remove any instruments with a premium payment date after today.
```
frame = portfolio.to_frame()
frame.index = frame.index.droplevel(0)
# extend dataframe with trade dates
trade_dates = {value:key for key, value in result}
frame['trade_date'] = frame.apply(lambda x: trade_dates[x.name], axis=1)
# filter any swaptions with premium date larger than today
frame = frame[frame.premium_payment_date < dt.datetime.today().date()]
frame.head(2)
```
### 3 - Grab the Data
Now the fun part - we need to calculate a lot of datapoints for this backtest.
For each straddle, we need to define a new swap every day and price it the following day when we unwind it. This means about 36,000 points (~300 instruments * 30 days * 4 measures (swaption price, swaption delta, swap price, swap delta)).
Like last time I will compute as much as I can asyncrously and keep track of the futures for each measure.
Introducing a high-level `PricingContext` to batch requests can improve speed as well. Note just using `PricingContext` will improve speed but `batch=True` can add efficiency.
To learn more about async and other compute controls and how to use them, please see our [pricing context guide](https://developer.gs.com/docs/gsquant/guides/Pricing-and-Risk/pricing-context/).
I'll start by getting the prices and delta for the swaptions first.
```
from gs_quant.risk import IRDeltaParallel
# insert columns in our frame to track the futures
frame['so_price_f'] = len(frame) * [None]
frame['so_delta_f'] = len(frame) * [None]
with PricingContext(is_batch=use_batch, show_progress=True):
for inst, row in frame.iterrows():
with HistoricalPricingContext(start=row.trade_date,
end=min(row.expiration_date, dt.datetime.today().date()),
is_async=True):
so_price = inst.price()
so_delta = inst.calc(IRDeltaParallel)
frame.at[inst, 'so_price_f'] = so_price
frame.at[inst, 'so_delta_f'] = so_delta
```
Easy enough. I will now do the same for the swaps which I will use to delta hedge. Note instead of pricing the same already resolved swaption each day, here I create and price a new swap each day which will reflect that's day's ATM rate and matches the effective date and termination date of the corresponding swaption.
```
from gs_quant.instrument import IRSwap
# insert columns in our frame to track the futures
frame['s_f'] = len(frame) * [None]
frame['s_delta_f'] = len(frame) * [None]
with PricingContext(is_batch=use_batch, show_progress=True):
for inst, row in frame.iterrows():
swap = IRSwap(PayReceive.Pay, row.termination_date, Currency.USD,
effective_date=row.effective_date, fixed_rate='ATMF', notional_amount=1e8)
with HistoricalPricingContext(start=row.trade_date,
end=min(row.expiration_date, dt.datetime.today().date()),
is_async=True):
# track the resolved swap - we will need to price it when we unwind following day
s = swap.resolve(in_place=False)
s_delta = swap.calc(IRDeltaParallel)
frame.at[inst, 's_f'] = s
frame.at[inst, 's_delta_f'] = s_delta
```
In the above request, we created a new resolved swaption for each day but we still need to price it the following day when we unwind it. In the below, I collect the resolved swaps from the previous request and price lagged 1 day - that is, the following day.
```
from gs_quant.markets import PricingContext
import pandas as pd
swaps = pd.concat([pd.Series(row.s_f.result(), name=row.name) for _, row in frame.iterrows()],
axis=1, sort=True).shift(periods=1)
g = {}
with PricingContext(is_batch=use_batch, show_progress=True):
for date, row in swaps.iterrows():
with PricingContext(date, is_async=True):
prices = {k: p if isinstance(p, float) else p.price() for k, p in row.iteritems()}
g[date] = prices
swap_prices = pd.DataFrame(g).T
```
Finally, let's collect all the points and do some arithmetic to create a timeseries for each swaption. I will create two frames - one for the simple vol selling strategy and one taking into account the changing delta hedge.
```
from gs_quant.timeseries import *
not_delta_hedged = []
delta_hedged = []
for inst, row in frame.iterrows():
# collect all the results
total_result = pd.concat([row.so_price_f.result(), row.so_delta_f.result(),
pd.Series({k: v.result() for k, v in swap_prices[inst].iteritems()
if not isinstance(v, float)}),
row.s_delta_f.result()], axis=1, sort=True)
total_result.columns = ['swaption_prices', 'swaption_delta', 'swap_bought_prices', 'swap_sold_delta']
# today's hedge notional will be the ratio of prior day's swaption/swap delta ratio - that's
# how much of the swap we bought to hedge so will use it to scale unwind PV of the swap today
total_result['hedge_notional'] = -(total_result.swaption_delta/total_result.swap_sold_delta).shift(periods=1)
total_result = total_result.fillna(0)
# scale the umwind PV of prior day's swap hedge
total_result['swap_pos'] = total_result['hedge_notional'] * total_result['swap_bought_prices']
# add to swaption price to get total performance cutting off last time due to the lag
swaption_pl = diff(total_result['swaption_prices'], 1).fillna(0)
total_result['total_pv'] = swaption_pl + total_result['swap_pos']
not_delta_hedged.append(pd.Series(swaption_pl[:-1], name=inst))
delta_hedged.append(pd.Series(total_result['total_pv'][:-1], name=inst))
```
### 4 - Putting it all together
Now, let's combine all the results to look at the impact delta hedging makes on the strategy. Unsurprisingly, the delta hedged version provides protection to tail events like March 2020.
```
not_dh = pd.concat(not_delta_hedged, axis=1, sort=True).fillna(0).sum(axis=1).cumsum()
dh = pd.concat(delta_hedged, axis=1, sort=True).fillna(0).sum(axis=1).cumsum()
comp = pd.DataFrame([dh, not_dh]).T
comp.columns = ['Delta Hedged', 'Not Delta Hedged']
comp.plot(figsize=(10, 6), title='Hedged vs Not Hedged')
```
Note that this backtesting doesn't include transaction costs and the implementation is different from how one might hedge in practice (unwinding and trading a new swap every day) but is economically equivalent to layering the hedges (and is cleaner from a calculation perspective).
Look out for future publications for added transaction costs and ways to quantitatively compare these strategies.
### Generic Backtesting Framework
```
from gs_quant.backtests.triggers import PeriodicTrigger, PeriodicTriggerRequirements
from gs_quant.backtests.actions import AddTradeAction, HedgeAction
from gs_quant.backtests.generic_engine import GenericEngine
from gs_quant.backtests.strategy import Strategy
from gs_quant.risk import IRDeltaParallelLocalCcy, Price
# dates on which actions will be triggered
trig_req = PeriodicTriggerRequirements(start_date=start_date, end_date=end_date, frequency='B')
# instrument that will be added on AddTradeAction
irswaption = IRSwaption('Straddle', '10y', 'USD', expiration_date='1m', notional_amount=1e8,
buy_sell='Sell', name='1m10y')
swap_hedge = IRSwap(PayReceive.Pay, '10y', 'USD', fixed_rate='ATMF', notional_amount=1e8, name='10y_swap')
action_trade = AddTradeAction(irswaption, 'expiration_date')
action_hedge = HedgeAction(IRDeltaParallelLocalCcy, swap_hedge, '1b')
# starting with empty portfolio (first arg to Strategy), apply actions on trig_req
triggers = PeriodicTrigger(trig_req, [action_trade, action_hedge])
strategy = Strategy(None, triggers)
# run backtest
GE = GenericEngine()
backtest = GE.run_backtest(strategy, start=start_date, end=end_date, frequency='B', show_progress=True)
```
Note, these strategies won't be exactly the same since the original series. This is because the original series is matching each swaption with a swap and holding both positions while resizing the swap each day. In the generic backtester snippet above, we passed '1b' for hedge holding period. This means that the delta from ALL swaption positions on a given date will be hedged with a single swap which will be replaced each day. Note if instead of '1b', we put '10y' for example (to match swaption tail), the swap hedge will be 'layered' - that is, we will add positive or negative notional each day but not unwind the existing swap hedges.
```
pd.DataFrame({'Generic backtester': backtest.result_summary['Cash'].cumsum() + backtest.result_summary[Price],
'Original series': dh}).plot(figsize=(10, 6), title='Delta hedged backtest comparison')
```
| github_jupyter |
# 量子振幅増幅
量子振幅増幅は、重ね合わせ状態におけるある特定の状態の振幅を増幅させるアルゴリズムです。
後に説明するグローバーのアルゴリズムの中核をなします。
## アルゴリズムの説明
### Amplitude amplification (振幅増幅)
振幅という言葉が出てきましたが、各状態の係数のことを**振幅**、各状態が測定される確率のことを**確率振幅**と言います。
以下の式を見てみます。
$$
H \lvert 0 \rangle \otimes H \lvert 0 \rangle = \frac{1}{2} (\lvert 00 \rangle + \lvert 01 \rangle + \lvert10 \rangle + \lvert 11 \rangle) \xrightarrow{\text{振幅増幅}} \lvert 00 \rangle
$$
この場合、矢印の左側では確率振幅は全て $1/4$ となり、
右側の状態は $00$ の確率振幅が $1$ となっています。
Amplitude amplification は言葉の通りこの確率振幅を上のように増幅させるアルゴリズムです。
以下の図を用いて具体的な計算を説明します。
<img width="30%" src="https://upload.wikimedia.org/wikipedia/commons/1/16/Grovers_algorithm_geometry.png">
参考: https://en.wikipedia.org/wiki/Grover%27s_algorithm
まずは記号の説明をします。
$\lvert s\rangle$ は任意の初期状態です。
ここでは具体例として、全状態の重ね合わせとして考えます。この初期状態は後に説明するグローバーのアルゴリズムで用いられます。
$$
\lvert s \rangle = \otimes^n H \lvert 0 \rangle =\frac{1}{\sqrt{2^n}}\sum^{2^n}_{x\in \{0, 1\}^n}\lvert x \rangle
$$
このうち、振幅増幅させたい状態を $\omega$ としています。
$$
\lvert \omega \rangle = \frac{1}{\sqrt{2^n}} \lvert 00...010...00\rangle
$$
$x$ 番目に1が入っているとします。
$\lvert s'\rangle$ は $\lvert s\rangle$ から $\omega$ を除いたベクトルです。
$$
\lvert s' \rangle = \lvert s \rangle - \lvert \omega \rangle = \lvert \omega^{\perp} \rangle
$$
$\lvert \omega \rangle$ に垂直なことがわかります。
$U_{\omega}$ は $\lvert s' \rangle$ を軸に $\psi$ を反転させる行列です。
すなわち $\psi$ を $-\phi$ 回転させます。
$$
U_{\omega} \lvert \psi \rangle = \cos(-\phi) \tilde{\lvert s' \rangle} + \sin(-\phi) \tilde{\lvert \omega \rangle} = \cos(\phi) \tilde{\lvert s' \rangle} - \sin(\phi) \tilde{\lvert \omega \rangle}
$$
$U_s$ は $\psi$ を $\lvert s \rangle$ を軸に反転させる行列です。
$$
U_s \lvert \psi \rangle = \cos \bigg\{ \frac{\theta}{2} + \big(\frac{\theta}{2} - \phi \big) \bigg\} \tilde{\lvert s' \rangle} + \sin \bigg\{ \frac{\theta}{2} + \big(\frac{\theta}{2} - \phi \big) \bigg\}\tilde{\lvert \omega \rangle} = \cos(\theta - \phi)\tilde{\lvert s' \rangle} + \sin(\theta - \phi) \tilde{\lvert \omega \rangle}
$$
### 概要
アルゴリズムの概要は以下のようになります。
1. $\lvert s \rangle$ を $U_{\omega}$ を用いて $\lvert s' \rangle$ で折り返す。
2. $U_{\omega}\lvert s \rangle$ を $U_s$ を用いて $\lvert s \rangle$ で折り返す。
以上の流れを詳しく説明していきます。
#### 1. $\lvert s' \rangle$ に関する折り返し
$\lvert s \rangle$ について上の図のように $\theta$ を用いて
$$
\lvert s \rangle = \cos\bigl(\frac{\theta}{2}\bigr) \tilde{\lvert s' \rangle} - \sin\bigl(\frac{\theta}{2}\bigr) \tilde{\lvert \omega \rangle}
$$
と定義します。
このとき
$$
\cos\bigl( \frac{\theta}{2} \bigr) = \sqrt{\frac{2^n-1}{2^n}},\ \ \ \ \sin\bigl( \frac{\theta}{2} \bigr) = \sqrt{\frac{1}{2^n}}
$$
と表せます。
$U_{\omega}$ で $\lvert s \rangle$ を $\lvert s' \rangle$ を軸に折り返します。
上の図から以下のようにかけます。
$$
U_{\omega} \lvert s \rangle = \cos\bigl(-\frac{\theta}{2}\bigr)\tilde{\lvert s' \rangle} + \sin\bigl(-\frac{\theta}{2}\bigr)\tilde{\lvert \omega \rangle} = \cos\bigl(\frac{\theta}{2}\bigr)\tilde{\lvert s' \rangle} - \sin\bigl(\frac{\theta}{2}\bigr)\tilde{\lvert \omega \rangle}
$$
この操作に関しては $\lvert \omega \rangle$ のみに作用しているので $U_{\omega}$ は上で述べた Oracle を表していることがわかります。
#### 2. $\lvert s \rangle$ に関する折り返し
$U_s$ で $U_{\omega}\lvert s \rangle$ を $\lvert s \rangle$ を軸に折り返します。
$$
U_s U_{\omega} \lvert s\rangle = U_s\biggl( \cos\bigl(-\frac{\theta}{2}\bigr)\tilde{\lvert s' \rangle} + \sin\bigl(-\frac{\theta}{2}\bigr)\tilde{\lvert \omega \rangle} \biggr)
$$
ここで $2\theta$ 回転させれば良いので
$$
U_s U_{\omega} \lvert s\rangle = \cos\bigl(\frac{3}{2}\theta\bigr)\tilde{\lvert s' \rangle} + \sin\bigl(\frac{3}{2}\theta\bigr)\tilde{\lvert \omega \rangle}
$$
具体的に $\cos$ と $\sin$ を求めると加法定理から
$$
\cos \frac{3}{2}\theta = \bigl( 1-\frac{4}{2^n} \bigr) \sqrt{\frac{2^n-1}{2^n}},\ \ \ \ \sin \frac{3}{2}\theta = \bigl( 3-\frac{4}{2^n} \bigr) \sqrt{\frac{1}{2^n}}
$$
よって、$\lvert s' \rangle$, $\lvert \omega \rangle$ を用いると
$$
U_s U_{\omega} \lvert s\rangle = \bigl(1 - \frac{4}{2^n}\bigr)\lvert s' \rangle + \bigl(3 - \frac{4}{2^n}\bigr)\lvert \omega \rangle
$$
この操作によって $2^n$ 個の振幅のうち $\lvert \omega \rangle$ が他のよりも約3倍大きくなりました。
以上で振幅増幅させることができました。
## 回路の実装
これをblueqatで実装してみましょう。
2量子ビットの量子振幅増幅を考えます。
2量子ビットの状態は、00,01,10,11の4通りです。その中から特定の状態を増幅したいと思います。
増幅したい状態は上記の $\lvert \omega \rangle$ に対応します。
#### 1. $\lvert s \rangle$ を $U_{\omega}$ を用いて $\lvert s' \rangle$ で折り返す。
まず、 $\lvert \omega \rangle$ と垂直な $\lvert s' \rangle$ を軸に、状態を反転させるゲート $U_{\omega}$ を実現する必要があります。
そのためにはゲート操作をつかって、「解に対応する状態ベクトルだけに-1がかかる対角行列」を数学的に作ります。
ここではHゲート、Sゲート、CZゲートを用いれば可能です。
各回路を一つ一つ見ていきましょう。.run_with_sympy_unitary()を実行することで、回路のユニタリ行列を確認することができます。
CZゲートの対角行列から始めて、それを変更しながら回路を作っていきます。
```
from blueqat import Circuit
'''
#marking on 11
-------*-----
-------Z-----
'''
Circuit(2).cz[0,1].run_with_sympy_unitary()
'''
#marking on 01
----S--*--S---
-------Z-------
'''
Circuit(2).s[0].cz[0,1].s[0].run_with_sympy_unitary()
'''
#marking on 10
--------*------
----S--Z--S---
'''
Circuit(2).s[1].cz[0,1].s[1].run_with_sympy_unitary()
'''
#00
----S--*--S--
----S--Z--S--
'''
Circuit(2).s[:].cz[0,1].s[:].run_with_sympy_unitary()
```
最後の結果は符号が反転しています。このような場合には、すべてのマイナス記号をプラスに、プラス記号をマイナスに反転させるグローバル位相を考えることができます。
#### 2. $U_{\omega}\lvert s \rangle$ を $U_s$ を用いて $\lvert s \rangle$ で折り返す。
$U_s$ については以下のように考えます。
$\lvert s \rangle$ の定義から $\lvert s \rangle = \lvert s' \rangle + \lvert \omega \rangle$ と分けて考えると、$U_{\omega}$ はシンプルに
$$
U_{\omega} (\lvert s' \rangle + \lvert \omega \rangle) = \lvert s' \rangle - \lvert \omega \rangle
$$
となります。
つまりZ、CZゲートなどのように特定の状態だけ符号を変えるゲートを用いれば良いことがわかります。
$U_s$ は上の図から幾何的に考えると以下のようにかけます。
$$
\begin{align}
U_s U_{\omega} \lvert s \rangle &= 2(\langle s \lvert U_{\omega} \rvert s \rangle \rvert s \rangle - U_{\omega}\lvert s \rangle) + U_{\omega}\lvert s \rangle \\
&= 2\lvert s \rangle \langle s \lvert U_{\omega} \rvert s \rangle - U_{\omega}\lvert s \rangle \\
&= (2\lvert s\rangle \langle s\rvert - I) U_{\omega}\lvert s \rangle
\end{align}
$$
よって $U_s = 2\lvert s\rangle \langle s\rvert - I$ となります。
さらに初期状態 $\lvert s \rangle = \otimes^n H \lvert 0 \rangle$ とした場合、 $U_s$ は以下のように分解できます。
$$
2\lvert s\rangle \langle s\rvert - I = 2H^{\otimes n}\lvert 0^n\rangle \langle 0^n\rvert H^{\otimes n} - I = H^{\otimes n} (2\lvert 0^n\rangle \langle 0^n\rvert - I) H^{\otimes n}\ \ \ \ (\lvert 0^n \rangle = \lvert 00\cdots 00 \rangle)
$$
ここで $2\lvert 0^n\rangle \langle 0^n\rvert - I$ に関して
$$
2\lvert 0^n\rangle \langle 0^n\rvert - I =
\begin{pmatrix}
-1 & 0 & 0 & \ldots & 0 & 0 \\
0 & 1 & 0 & \ldots & 0 & 0 \\
0 & 0 & 1 & \ldots & 0 & 0 \\
\vdots & \vdots & \vdots & \ddots & \vdots & \vdots \\
0 & 0 & 0 & \ldots & 1 & 0 \\
0 & 0 & 0 & \ldots & 0 & 1
\end{pmatrix}
$$
と表せます。
これは
$$
XZX =
\begin{pmatrix}
-1 & 0 \\
0 & 1
\end{pmatrix}
$$
の性質から
$$
2\lvert 0^n\rangle \langle 0^n\rvert - I = X^{\otimes n}C^n ZX^{\otimes n}
$$
とかけます。
こちらをBlueqatに直してみます。
```
'''
--H-X-*-X-H--
--H-X-Z-X-H--
'''
Circuit(2).h[:].x[:].cz[0,1].x[:].h[:].run_with_sympy_unitary()
```
以上から振幅増幅用オラクルをゲートで書き直すことができました。
では、実際の回路の実装です。
```
#振幅増幅反転
a = Circuit(2).h[:].x[:].cz[0,1].x[:].h[:].m[:]
'''
#00回路
--H--S--*--S----H-X-*-X-H--
--H--S--Z--S----H-X-Z-X-H--
'''
(Circuit(2).h[:].s[:].cz[0,1].s[:] + a).run(shots=100)
'''
#01回路
--H-----*-------H-X-*-X-H--
--H--S--Z--S---H-X-Z-X-H--
'''
(Circuit(2).h[:].s[1].cz[0,1].s[1] + a).run(shots=100)
'''
#10回路
--H--S--*--S----H-X-*-X-H--
--H-----Z--------H-X-Z-X-H--
'''
(Circuit(2).h[:].s[0].cz[0,1].s[0] + a).run(shots=100)
'''
#11回路
--H-----*-------H-X-*-X-H--
--H-----Z-------H-X-Z-X-H--
'''
(Circuit(2).h[:].cz[0,1] + a).run(shots=100)
```
以上より、特定の状態が測定される確率を増幅できました。
| github_jupyter |
## 8.2 UNET을 이용한 컬러 복원 처리
UNET을 이용하여 흑백 이미지를 컬러 이미지로 복원하는 예제를 다룸
```
# set to use CPU
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
```
### 8.2.1 패키지 임포트
```
#from keras import models, backend
#from keras.layers import Input, Conv2D, MaxPooling2D, Dropout, Activation
#from keras.layers import UpSampling2D, BatchNormalization, Concatenate
from tensorflow.keras import models, backend
from tensorflow.keras.layers import Input, Conv2D, MaxPooling2D, Dropout, Activation
from tensorflow.keras.layers import UpSampling2D, BatchNormalization, Concatenate
```
### 8.2.2 UNET 모델링
```
class UNET(models.Model):
def __init__(self, org_shape, n_ch):
ic = 3 if backend.image_data_format() == 'channels_last' else 1
def conv(x, n_f, mp_flag=True):
x = MaxPooling2D((2, 2), padding='same')(x) if mp_flag else x
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
x = Dropout(0.05)(x)
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
return x
def deconv_unet(x, e, n_f):
x = UpSampling2D((2, 2))(x)
x = Concatenate(axis=ic)([x, e])
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
return x
# Input
original = Input(shape=org_shape)
# Encoding
c1 = conv(original, 16, mp_flag=False)
c2 = conv(c1, 32)
# Encoder
encoded = conv(c2, 64)
# Decoding
x = deconv_unet(encoded, c2, 32)
x = deconv_unet(x, c1, 16)
decoded = Conv2D(n_ch, (3, 3), activation='sigmoid', padding='same')(x)
#decoded = Conv2D(n_ch, (3, 3), padding='same')(x)
super().__init__(original, decoded)
self.compile(optimizer='adadelta', loss='mse')
```
### 8.2.3 데이터 준비
```
###########################
# 데이타 불러오기
###########################
from tensorflow.keras import datasets, utils
class DATA():
def __init__(self, in_ch=None):
(x_train, y_train), (x_test, y_test) = datasets.cifar10.load_data()
if x_train.ndim == 4:
if backend.image_data_format() == 'channels_first':
n_ch, img_rows, img_cols = x_train.shape[1:]
else:
img_rows, img_cols, n_ch = x_train.shape[1:]
else:
img_rows, img_cols = x_train.shape[1:]
n_ch = 1
# in_ch can be 1 for changing BW to color image using UNet
in_ch = n_ch if in_ch is None else in_ch
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
def RGB2Gray(X, fmt):
if fmt == 'channels_first':
R = X[:, 0:1]
G = X[:, 1:2]
B = X[:, 2:3]
else: # "channels_last
R = X[..., 0:1]
G = X[..., 1:2]
B = X[..., 2:3]
return 0.299 * R + 0.587 * G + 0.114 * B
def RGB2RG(x_train_out, x_test_out, fmt):
if fmt == 'channels_first':
x_train_in = x_train_out[:, :2]
x_test_in = x_test_out[:, :2]
else:
x_train_in = x_train_out[..., :2]
x_test_in = x_test_out[..., :2]
return x_train_in, x_test_in
if backend.image_data_format() == 'channels_first':
x_train_out = x_train.reshape(x_train.shape[0], n_ch, img_rows, img_cols)
x_test_out = x_test.reshape(x_test.shape[0], n_ch, img_rows, img_cols)
input_shape = (in_ch, img_rows, img_cols)
else:
x_train_out = x_train.reshape(x_train.shape[0], img_rows, img_cols, n_ch)
x_test_out = x_test.reshape(x_test.shape[0], img_rows, img_cols, n_ch)
input_shape = (img_rows, img_cols, in_ch)
if in_ch == 1 and n_ch == 3:
x_train_in = RGB2Gray(x_train_out, backend.image_data_format())
x_test_in = RGB2Gray(x_test_out, backend.image_data_format())
elif in_ch == 2 and n_ch == 3:
# print(in_ch, n_ch)
x_train_in, x_test_in = RGB2RG(x_train_out, x_test_out, backend.image_data_format())
else:
x_train_in = x_train_out
x_test_in = x_test_out
self.input_shape = input_shape
self.x_train_in, self.x_train_out = x_train_in, x_train_out
self.x_test_in, self.x_test_out = x_test_in, x_test_out
self.n_ch = n_ch
self.in_ch = in_ch
```
### 8.2.4 UNET 처리 그래프 그리기
```
###########################
# UNET 검증
###########################
from keraspp.skeras import plot_loss
import matplotlib.pyplot as plt
###########################
# UNET 동작 확인
###########################
import numpy as np
from sklearn.preprocessing import minmax_scale
def show_images(data, unet):
x_test_in = data.x_test_in
x_test_out = data.x_test_out
decoded_imgs_org = unet.predict(x_test_in)
decoded_imgs = decoded_imgs_org
if backend.image_data_format() == 'channels_first':
print(x_test_out.shape)
x_test_out = x_test_out.swapaxes(1, 3).swapaxes(1, 2)
print(x_test_out.shape)
decoded_imgs = decoded_imgs.swapaxes(1, 3).swapaxes(1, 2)
if data.in_ch == 1:
x_test_in = x_test_in[:, 0, ...]
elif data.in_ch == 2:
print(x_test_out.shape)
x_test_in_tmp = np.zeros_like(x_test_out)
x_test_in = x_test_in.swapaxes(1, 3).swapaxes(1, 2)
x_test_in_tmp[..., :2] = x_test_in
x_test_in = x_test_in_tmp
else:
x_test_in = x_test_in.swapaxes(1, 3).swapaxes(1, 2)
else:
# x_test_in = x_test_in[..., 0]
if data.in_ch == 1:
x_test_in = x_test_in[..., 0]
elif data.in_ch == 2:
x_test_in_tmp = np.zeros_like(x_test_out)
x_test_in_tmp[..., :2] = x_test_in
x_test_in = x_test_in_tmp
n = 10
plt.figure(figsize=(20, 6))
for i in range(n):
ax = plt.subplot(3, n, i + 1)
if x_test_in.ndim < 4:
plt.imshow(x_test_in[i], cmap='gray')
else:
plt.imshow(x_test_in[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.imshow(decoded_imgs[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n * 2)
plt.imshow(x_test_out[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
```
### 8.2.5 UNET 학습 및 결과 확인
```
###########################
# 학습 및 확인
###########################
def main(in_ch=1, epochs=10, batch_size=128, fig=True):
data = DATA(in_ch=in_ch)
print(data.input_shape, data.x_train_in.shape)
unet = UNET(data.input_shape, data.n_ch)
history = unet.fit(data.x_train_in, data.x_train_out,
epochs=epochs,
batch_size=batch_size,
shuffle=True,
validation_split=0.2)
if fig:
plot_loss(history)
show_images(data, unet)
from distutils import util
class parser:
def __init__(self):
self.input_channels = 1
self.epochs = 10
self.batch_size = 512
self.fig = True
args = parser()
print(args.fig)
main(args.input_channels, args.epochs, args.batch_size, args.fig)
```
---
### 8.2.6 전체 코드
```
#######################################################################################
# unet_conv_cifar10rgb_mc.py
# Convlutional Layer UNET with RGB Cifar10 dataset and Class with Keras Model approach
#######################################################################################
#import matplotlib
#matplotlib.use("TkAgg")
###########################
# AE 모델링
###########################
import matplotlib.pyplot as plt
from keras import models, backend
from keras.layers import Input, Conv2D, MaxPooling2D, Dropout, \
UpSampling2D, BatchNormalization, Concatenate, Activation
# backend.set_image_data_format('channels_first')
class UNET(models.Model):
def __init__(self, org_shape, n_ch):
ic = 3 if backend.image_data_format() == 'channels_last' else 1
def conv(x, n_f, mp_flag=True):
x = MaxPooling2D((2, 2), padding='same')(x) if mp_flag else x
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
x = Dropout(0.05)(x)
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
return x
def deconv_unet(x, e, n_f):
x = UpSampling2D((2, 2))(x)
x = Concatenate(axis=ic)([x, e])
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
x = Conv2D(n_f, (3, 3), padding='same')(x)
x = BatchNormalization()(x)
x = Activation('tanh')(x)
return x
# Input
original = Input(shape=org_shape)
# Encoding
c1 = conv(original, 16, mp_flag=False)
c2 = conv(c1, 32)
# Encoder
encoded = conv(c2, 64)
# Decoding
x = deconv_unet(encoded, c2, 32)
x = deconv_unet(x, c1, 16)
decoded = Conv2D(n_ch, (3, 3), activation='sigmoid', padding='same')(x)
#decoded = Conv2D(n_ch, (3, 3), padding='same')(x)
super().__init__(original, decoded)
self.compile(optimizer='adadelta', loss='mse')
###########################
# 데이타 불러오기
###########################
from tensorflow.keras import datasets, utils
class DATA():
def __init__(self, in_ch=None):
(x_train, y_train), (x_test, y_test) = datasets.cifar10.load_data()
if x_train.ndim == 4:
if backend.image_data_format() == 'channels_first':
n_ch, img_rows, img_cols = x_train.shape[1:]
else:
img_rows, img_cols, n_ch = x_train.shape[1:]
else:
img_rows, img_cols = x_train.shape[1:]
n_ch = 1
# in_ch can be 1 for changing BW to color image using UNet
in_ch = n_ch if in_ch is None else in_ch
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
def RGB2Gray(X, fmt):
if fmt == 'channels_first':
R = X[:, 0:1]
G = X[:, 1:2]
B = X[:, 2:3]
else: # "channels_last
R = X[..., 0:1]
G = X[..., 1:2]
B = X[..., 2:3]
return 0.299 * R + 0.587 * G + 0.114 * B
def RGB2RG(x_train_out, x_test_out, fmt):
if fmt == 'channels_first':
x_train_in = x_train_out[:, :2]
x_test_in = x_test_out[:, :2]
else:
x_train_in = x_train_out[..., :2]
x_test_in = x_test_out[..., :2]
return x_train_in, x_test_in
if backend.image_data_format() == 'channels_first':
x_train_out = x_train.reshape(x_train.shape[0], n_ch, img_rows, img_cols)
x_test_out = x_test.reshape(x_test.shape[0], n_ch, img_rows, img_cols)
input_shape = (in_ch, img_rows, img_cols)
else:
x_train_out = x_train.reshape(x_train.shape[0], img_rows, img_cols, n_ch)
x_test_out = x_test.reshape(x_test.shape[0], img_rows, img_cols, n_ch)
input_shape = (img_rows, img_cols, in_ch)
if in_ch == 1 and n_ch == 3:
x_train_in = RGB2Gray(x_train_out, backend.image_data_format())
x_test_in = RGB2Gray(x_test_out, backend.image_data_format())
elif in_ch == 2 and n_ch == 3:
# print(in_ch, n_ch)
x_train_in, x_test_in = RGB2RG(x_train_out, x_test_out, backend.image_data_format())
else:
x_train_in = x_train_out
x_test_in = x_test_out
self.input_shape = input_shape
self.x_train_in, self.x_train_out = x_train_in, x_train_out
self.x_test_in, self.x_test_out = x_test_in, x_test_out
self.n_ch = n_ch
self.in_ch = in_ch
###########################
# UNET 검증
###########################
from keraspp.skeras import plot_loss
import matplotlib.pyplot as plt
###########################
# UNET 동작 확인
###########################
import numpy as np
from sklearn.preprocessing import minmax_scale
def show_images(data, unet):
x_test_in = data.x_test_in
x_test_out = data.x_test_out
decoded_imgs_org = unet.predict(x_test_in)
decoded_imgs = decoded_imgs_org
if backend.image_data_format() == 'channels_first':
print(x_test_out.shape)
x_test_out = x_test_out.swapaxes(1, 3).swapaxes(1, 2)
print(x_test_out.shape)
decoded_imgs = decoded_imgs.swapaxes(1, 3).swapaxes(1, 2)
if data.in_ch == 1:
x_test_in = x_test_in[:, 0, ...]
elif data.in_ch == 2:
print(x_test_out.shape)
x_test_in_tmp = np.zeros_like(x_test_out)
x_test_in = x_test_in.swapaxes(1, 3).swapaxes(1, 2)
x_test_in_tmp[..., :2] = x_test_in
x_test_in = x_test_in_tmp
else:
x_test_in = x_test_in.swapaxes(1, 3).swapaxes(1, 2)
else:
# x_test_in = x_test_in[..., 0]
if data.in_ch == 1:
x_test_in = x_test_in[..., 0]
elif data.in_ch == 2:
x_test_in_tmp = np.zeros_like(x_test_out)
x_test_in_tmp[..., :2] = x_test_in
x_test_in = x_test_in_tmp
n = 10
plt.figure(figsize=(20, 6))
for i in range(n):
ax = plt.subplot(3, n, i + 1)
if x_test_in.ndim < 4:
plt.imshow(x_test_in[i], cmap='gray')
else:
plt.imshow(x_test_in[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.imshow(decoded_imgs[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n * 2)
plt.imshow(x_test_out[i])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
###########################
# 학습 및 확인
###########################
def main(in_ch=1, epochs=10, batch_size=512, fig=True):
data = DATA(in_ch=in_ch)
print(data.input_shape, data.x_train_in.shape)
unet = UNET(data.input_shape, data.n_ch)
history = unet.fit(data.x_train_in, data.x_train_out,
epochs=epochs,
batch_size=batch_size,
shuffle=True,
validation_split=0.2)
if fig:
plot_loss(history)
show_images(data, unet)
if __name__ == '__main__':
import argparse
from distutils import util
parser = argparse.ArgumentParser(description='UNET for Cifar-10: Gray to RGB')
parser.add_argument('--input_channels', type=int, default=1,
help='input channels (default: 1)')
parser.add_argument('--epochs', type=int, default=10,
help='training epochs (default: 10)')
parser.add_argument('--batch_size', type=int, default=512,
help='batch size (default: 1000)')
parser.add_argument('--fig', type=lambda x: bool(util.strtobool(x)),
default=True, help='flag to show figures (default: True)')
args = parser.parse_args()
print("Aargs:", args)
print(args.fig)
main(args.input_channels, args.epochs, args.batch_size, args.fig)
```
| github_jupyter |
```
from tqdm import tqdm
import os
from io import BytesIO
import ast
import numpy as np
import pickle
import torch
import boto3
from scipy.spatial.distance import cdist
import networkx as nx
import matplotlib.pyplot as plt
from matplotlib.offsetbox import OffsetImage, AnnotationBbox
from itertools import combinations
import umap.umap_ as umap
```
### 1. Import the feature vectors from S3
```
# https://alexwlchan.net/2017/07/listing-s3-keys/
def get_all_s3_keys(bucket):
"""Get a list of all keys in an S3 bucket."""
keys = []
kwargs = {"Bucket": bucket}
while True:
resp = s3.list_objects_v2(**kwargs)
for obj in resp["Contents"]:
keys.append(obj["Key"])
try:
kwargs["ContinuationToken"] = resp["NextContinuationToken"]
except KeyError:
break
return keys
bucket_name = "miro-images-feature-vectors"
s3 = boto3.client("s3")
keys = get_all_s3_keys(bucket_name)
len(keys)
folder_name = "reduced_feature_vectors_20_dims"
keys = [k for k in keys if k.split("/")[0] == folder_name]
len(keys)
feature_vectors = {}
for key in tqdm(keys):
obj = s3.get_object(Bucket=bucket_name, Key=key)
read_obj = obj["Body"].read()
feature_vectors[key] = np.frombuffer(read_obj, dtype=np.float)
```
### 2. Get the distances between feature vectors
```
feature_vectors_list = list(feature_vectors.values())
[v for f in feature_vectors_list for v in f if "nan" in str(v)]
dist_mat = cdist(feature_vectors_list, feature_vectors_list, metric="cosine")
dist_mat
dist_mat_top = np.zeros_like(dist_mat)
dist_mat_top[:] = None
n = 3
# Find the top n neighbours for each image
for i, _ in tqdm(enumerate(keys)):
arr = dist_mat[i].argsort()
top_args = arr[arr != i]
dist_mat_top[i][top_args[0:n]] = dist_mat[i][top_args[0:n]]
for j in top_args[0:n]:
dist_mat_top[j][i] = dist_mat[j][i]
```
### 3a. Load images for plotting
```
with open("../data/processed_images_sample.pkl", "rb") as handle:
images_original = pickle.load(handle)
# Put in the same order as the feature vectors
images = []
for key in feature_vectors.keys():
image_key = os.path.basename(key)
images.append(images_original[image_key])
images[0]
```
### 3. Plot the network of images connected to their closest neighbours
```
def inv_rel_norm(value, min_val, max_val):
value = (value - min_val) / (max_val - min_val)
value = 1 / (value + 1e-8)
return value
def create_graph(dist_mat_top):
min_val = np.nanmin(dist_mat_top)
max_val = np.nanmax(dist_mat_top)
nodes = list(range(0, len(dist_mat_top[0])))
G = nx.Graph()
G.add_nodes_from(nodes)
# Put the weights in as the distances
# only inc nodes if they are in the closest related neighbours
for start, end in list(combinations(nodes, 2)):
if ~np.isnan(dist_mat_top[start, end]):
# Since in the plot a higher weight makes the nodes closer,
# but a higher value in the distance matrix means the images are further away,
# we need to inverse the weight (so higher = closer)
G.add_edge(
start,
end,
weight=inv_rel_norm(dist_mat_top[start, end], min_val, max_val),
)
return G
def plot_graph(G, image_names=None):
pos = nx.spring_layout(G)
plt.figure(3, figsize=(10, 10))
nx.draw(G, pos, node_size=10)
for p in pos: # raise text positions
pos[p][1] += 0.06
if image_names:
image_names_dict = {k: str(k) + " " + v for k, v in enumerate(image_names)}
nx.draw_networkx_labels(G, pos, labels=image_names_dict)
plt.show()
G = create_graph(dist_mat_top)
plot_graph(G)
```
### 4. Visualise the clusters by reducing dimensions
```
reducer = umap.UMAP()
embedding_fv = reducer.fit_transform(feature_vectors_list)
embedding_fv.shape
# from https://www.kaggle.com/gaborvecsei/plants-t-sne
def visualize_scatter_with_images(X_2d_data, images, figsize=(45, 45), image_zoom=1):
fig, ax = plt.subplots(figsize=figsize)
artists = []
for xy, i in zip(X_2d_data, images):
x0, y0 = xy
img = OffsetImage(i, zoom=image_zoom)
ab = AnnotationBbox(img, (x0, y0), xycoords="data", frameon=False)
artists.append(ax.add_artist(ab))
ax.update_datalim(X_2d_data)
ax.autoscale()
plt.axis("off")
plt.show()
x_data = [[a, b] for (a, b) in zip(embedding_fv[:, 0], embedding_fv[:, 1])]
visualize_scatter_with_images(x_data, images=images, image_zoom=0.3)
```
### 5a. Pick 2 images and look at the route between them
```
image_names_dict = {k: v for k, v in enumerate(images)}
node1 = np.random.choice(list(image_names_dict))
node2 = np.random.choice(list(image_names_dict))
node_path = nx.dijkstra_path(G, node1, node2, weight=None)
print(node_path)
show_images = [images[i] for i in node_path]
fig = plt.figure(figsize=(20, 10))
columns = len(show_images)
for i, image in enumerate(show_images):
ax = plt.subplot(len(show_images) / columns + 1, columns, i + 1)
ax.set_axis_off()
plt.imshow(image)
```
### 5b. User sets number of images in pathway
```
path_size = 10
```
### 5c. Pick 3 images and look at the paths between them
```
node1 = np.random.choice(list(image_names_dict))
node2 = np.random.choice(list(image_names_dict))
node3 = np.random.choice(list(image_names_dict))
node_path_a = nx.dijkstra_path(G, node1, node2, weight=None)
node_path_b = nx.dijkstra_path(G, node2, node3, weight=None)
node_path_c = nx.dijkstra_path(G, node3, node1, weight=None)
node_path_3 = node_path_a[:-1] + node_path_b[:-1] + node_path_c
print(node_path_3)
show_images = [images[i] for i in node_path_3]
fig = plt.figure(figsize=(20, 10))
columns = len(show_images)
for i, image in enumerate(show_images):
ax = plt.subplot(len(show_images) / columns + 1, columns, i + 1)
if node_path_3[i] in [node1, node2, node3]:
ax.set(title="NODE")
ax.set_axis_off()
plt.imshow(image)
```
### 6. Plot route on graph
```
# from https://www.kaggle.com/gaborvecsei/plants-t-sne
def visualize_scatter_pathway_with_images(
X_2d_data, pathway, images, figsize=(45, 45), image_zoom=1
):
fig, ax = plt.subplots(figsize=figsize)
x_path = [x_data[c][0] for c in node_path]
y_path = [x_data[c][1] for c in node_path]
artists = []
for num, (xy, i) in enumerate(zip(X_2d_data, images)):
x0, y0 = xy
if num in pathway:
img = OffsetImage(i, zoom=image_zoom * 2, alpha=0.8)
else:
img = OffsetImage(i, zoom=image_zoom, alpha=0.2)
ab = AnnotationBbox(img, (x0, y0), xycoords="data", frameon=False)
artists.append(ax.add_artist(ab))
ax.update_datalim(X_2d_data)
ax.autoscale()
plt.plot(x_path, y_path, "ro-", linewidth=5)
plt.axis("off")
plt.show()
visualize_scatter_pathway_with_images(
x_data, node_path, images=images, figsize=(30, 30), image_zoom=0.3
)
```
| github_jupyter |
```
from os import path
from utils import fix_path
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from statsmodels import robust
from statsmodels.stats.stattools import medcouple
import json
def fx_unsupervised(v_uk_id):
#v_uk_id= 123456
my_path = 'db/data/' + v_uk_id + "/"
v_input_json_file = my_path + "details.json"
v_output_json_file = my_path + "result_desc.json"
v_output_result_csv = my_path+"result_individual_columns.csv"
v_output_result_csv = my_path+"result.csv"
with open(v_input_json_file, encoding="utf-8") as data_file:
input_json = json.load(data_file)
v_input_csv = input_json['filename']
df = pd.read_csv(v_input_csv)
df['anomalydetectid']=df.index
######################
v_id = 'anomalydetectid'
v_analysis_columns_list = input_json['dimension']
df_outliers = pd.DataFrame({})
#v_analysis_columns = ''.join(v_analysis_columns_list)
for my_analysis_column in v_analysis_columns_list:
v_df_temp = fx_ThreeSigmaRule(df[v_id], df[my_analysis_column], 2, 1)
v_df_temp['Outlier Type'] = 'Three Sigma Rule'
v_df_temp['Analysis Column'] = my_analysis_column
df_outliers = pd.concat([df_outliers, v_df_temp])
######################
df_outliers_out = pd.merge(df_outliers, df, on='anomalydetectid')
df_outliers_out.to_csv(v_output_result_csv, encoding="utf-8", index=False, header=True)
x_groupby_type = df_outliers.groupby(['Analysis Column'])
df2 = x_groupby_type.count()
df2.reset_index(inplace=True)
df3 = df2.sort_values(['data'], ascending=True).head(4)
df3.reset_index(inplace=True)
for i, r in df3.iterrows():
v_filename = 'image'+str(i)
v_list = df.loc[:, [r['Analysis Column']]]
v_title = 'Outliers for '+r['Analysis Column']
v_column=r['Analysis Column']
#fx_box_plot (v_uk_id,v_list , v_title,v_filename )
#plt.scatter(df['anomalydetectid'].values,df[v_column].values)
fx_scatter_plot(v_uk_id,df['anomalydetectid'].values,df[v_column].values, v_title,v_filename)
##################################json code
v_output_json_contents = {
"image_title1": "nish1",
"image_title2": "jfbgcjshhgsj",
"image_title3": "jfbgcjshhgsj",
"image_title4": "jfbgcjshhgsj",
"image_name1": "image1.png",
"image_name2": "image2.png",
"image_name3": "image3.png",
"image_name4": "image4.png",
"image_desc1": "jfbgcjshhgsj",
"image_desc2": "jfbgcjshhgsj",
"image_desc3": "jfbgcjshhgsj",
"image_desc4": "jfbgcjshhgsj",
"model": [{"model_desc": "ssss", "model_file": "ssss"}, {"model_desc": "ssss", "model_file": "ssss"}]
}
with open(v_output_json_file, 'w') as outfile:
json.dump(v_output_json_contents, outfile)
##################################
return df_outliers_out
def fx_scatter_plot (v_uk_id , v_id_series,v_column_series , v_title,v_filename ):
my_path = "db/data/" + v_uk_id + "/"
fig, ax = plt.subplots()
#jpg_filename = my_path + v_filename+'.jpg'
png_filename = my_path + v_filename
ax.scatter(v_id_series,v_column_series)
ax.set_title(v_title)
fig.tight_layout()
#fig.show()
#fig.savefig(jpg_filename, dpi=1000)
fig.savefig(png_filename+'.png', dpi=1000)
def fx_box_plot (v_uk_id,v_list , v_title,v_filename ):
my_path = "db/data/" + v_uk_id + "/"
fig, ax = plt.subplots()
jpg_filename = my_path + v_filename+'.jpg'
png_filename = my_path + v_filename+'.png'
x = range(len(v_list))
ax.boxplot(v_list, patch_artist=True)
ax.set_title(v_title)
fig.tight_layout()
#fig.show()
fig.savefig(jpg_filename, dpi=1000)
fig.savefig(png_filename+'.png', dpi=1000)
def fx_ThreeSigmaRule(series_id, series_data, v_number_of_std, v_masking_Iteration):
'''
value should be in +- range of -->mean + n * std
Probability
n =1 --> 68
n =2 --> 95
n= 3 --> 99.7
Usage :
df_outliers= fx_ThreeSigmaRule(df['myid'],df['series1'], 2,2)
df_outliers
Good for normal distributed data ---series or sequences
Drawback :
Not good when data is not normal distribution
Sensitive to extreme points ---Masking effect. If extreme value is too large then it overshadow other values
-->iteration solves this issue
'''
v_df = pd.DataFrame({})
v_df_outliers_final = pd.DataFrame({})
v_df['anomalydetectid'] = series_id
v_df['data'] = series_data
v_Iteration = 0
while (v_Iteration < v_masking_Iteration):
#############################
print(str(v_masking_Iteration))
v_masking_Iteration = v_masking_Iteration - 1
v_threshold = np.std(v_df['data']) * v_number_of_std
v_mean = np.mean(v_df['data'])
print(str(v_mean - v_threshold))
print(str(v_threshold + v_mean))
where_tuple = (np.abs(v_df['data'] - v_mean) > v_threshold)
v_df_outliers = v_df[where_tuple]
# v_outliersList = [ [r[0] , r[1]] for i,r in v_df.iterrows() if np.abs(r[1]) > v_threshold + v_mean]
if (len(v_df_outliers) > 0):
v_df_outliers_final = pd.concat([v_df_outliers_final, v_df_outliers])
# Update data - remove otliers from the list
# list1 = [x for x in list1 if x not in v_outliersList[1]]
where_tuple = (np.abs(v_df['data'] - v_mean) <= v_threshold)
v_df = v_df[where_tuple]
else:
break
############################
if len(v_df_outliers_final) > 0:
return (v_df_outliers_final)
else:
return (pd.DataFrame({}))
print("Three are No Outliers")
###############################################################################
def fx_mad_Rule(series_id, series_data, v_number_of_std):
'''
value should be in +- range of -->median + n * MAD/.6745
Usage :
df_outliers= fx_mad_Rule(df['myid'],df['series1'], 2)
df_outliers
Good for not normal distributed data
No issue with extreme points
Drawback :
Not good when data is normal distribution
Too agressive
'''
# warning ignore for verylarge values
# np.seterr(invalid='ignore')
# np.errstate(invalid='ignore')
# np.warnings.filterwarnings('ignore')
v_df = pd.DataFrame({})
v_df_outliers_final = pd.DataFrame({})
v_df['id'] = series_id
v_df['data'] = series_data
#############################
v_threshold = robust.mad(v_df['data'], c=1) * v_number_of_std / 0.6745
v_median = np.median(v_df['data'])
print(str(v_median - v_threshold))
print(str(v_threshold + v_median))
where_tuple = (np.abs(v_df['data'] - v_median) > v_threshold)
v_df_outliers_final = v_df[where_tuple]
############################
if len(v_df_outliers_final) > 0:
return (v_df_outliers_final)
else:
print("Three are No Outliers")
###############################################################################
def fx_boxplot_Rule(series_id, series_data):
'''
value should be in +- range of -->(y > q3 + 1.5 * iqr) or (y < q1 - 1.5 * iqr
Usage :
df_outliers= fx_boxplot_Rule(df['myid'],df['series1'])
df_outliers
For presense of outliers --less sensitive than 3 sigma but more sensitive to MAD test
No depenedence of median and mean
better for moderately asymmetric distribution
Drawback :
Too agressive
'''
# warning ignore for verylarge values
# np.seterr(invalid='ignore')
# np.errstate(invalid='ignore')
# np.warnings.filterwarnings('ignore')
v_df = pd.DataFrame({})
v_df_outliers_final = pd.DataFrame({})
v_df['id'] = series_id
v_df['data'] = series_data
#############################
q1 = np.percentile(v_df['data'], 25)
q3 = np.percentile(v_df['data'], 75)
iqr = q3 - q1
print(str(q1 - 1.5 * iqr))
print(str(q3 + 1.5 * iqr))
where_tuple1 = (v_df['data'] > q3 + 1.5 * iqr)
where_tuple2 = (v_df['data'] < q1 - 1.5 * iqr)
v_df_outliers_final = v_df[where_tuple1 | where_tuple2]
############################
if len(v_df_outliers_final) > 0:
return (v_df_outliers_final)
else:
print("Three are No Outliers")
###############################################################################
def fx_adjusted_boxplot_Rule(series_id, series_data):
'''
value should be in +- range of -->(y > q3 + 1.5 * iqr) or (y < q1 - 1.5 * iqr
Usage :
df_outliers= fx_boxplot_Rule(df['myid'],df['series1'])
df_outliers
For presense of outliers --less sensitive than 3 sigma but more sensitive to MAD test
No depenedence of median and mean
better for moderately asymmetric distribution
Drawback :
Too agressive
'''
# warning ignore for verylarge values
# np.seterr(invalid='ignore')
# np.errstate(invalid='ignore')
# np.warnings.filterwarnings('ignore')
v_df = pd.DataFrame({})
v_df_outliers_final = pd.DataFrame({})
v_df['id'] = series_id
v_df['data'] = series_data
#############################
q1 = np.percentile(v_df['data'], 25)
q3 = np.percentile(v_df['data'], 75)
iqr = q3 - q1
mc = medcouple(v_df['data'])
if (mc >= 0):
lr = q1 - 1.5 * iqr * np.exp(-4 * mc)
ur = q3 + 1.5 * iqr * np.exp(3 * mc)
else:
lr = q1 - 1.5 * iqr * np.exp(-3 * mc)
ur = q3 + 1.5 * iqr * np.exp(4 * mc)
print(str(lr))
print(str(ur))
where_tuple1 = (v_df['data'] > ur)
where_tuple2 = (v_df['data'] < lr)
v_df_outliers_final = v_df[where_tuple1 | where_tuple2]
############################
if len(v_df_outliers_final) > 0:
return (v_df_outliers_final)
else:
print("Three are No Outliers")
###############################################################################
###############################################################################
###############################################################################
###############################################################################
###############################################################################
fx_unsupervised('123456')
```
| github_jupyter |
# EmbeddingRandomForest
## Random Forest with Embeddings
So far we have created both a random forest and a NN to do tabular modeling. One thing intresting about a NN is that it contains embeddings. Why don't we try to use these embeddings from the Neural Network in Random Forests? Will it improve the random forest? Lets find out!
```
#hide
!pip install -Uqq fastbook kaggle waterfallcharts treeinterpreter dtreeviz
import fastbook
fastbook.setup_book()
#hide
from fastbook import *
from kaggle import api
from pandas.api.types import is_string_dtype, is_numeric_dtype, is_categorical_dtype
from fastai.tabular.all import *
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from dtreeviz.trees import *
from IPython.display import Image, display_svg, SVG
pd.options.display.max_rows = 20
pd.options.display.max_columns = 8
```
## Unzipping data
```
import zipfile
z= zipfile.ZipFile('bluebook-for-bulldozers.zip') #unzip first
z.extractall() #extract
```
## Grabbing the Data
Similer to what we did in lesson 9, we will grab our data, set the ordinal var, and feature engineer the date.
```
df_nn = pd.read_csv(Path()/'TrainAndValid.csv', low_memory=False) #Data
#Set ordinal variables using our order
sizes = 'Large','Large / Medium','Medium','Small','Mini','Compact'
df_nn['ProductSize'] = df_nn['ProductSize'].astype('category')
df_nn['ProductSize'].cat.set_categories(sizes, ordered=True, inplace=True)
dep_var = 'SalePrice'
df_nn[dep_var] = np.log(df_nn[dep_var]) #remember we need to take log of the label (Kaggle requires)
df_nn = add_datepart(df_nn, 'saledate') #Also remember that we used feature engineering on date
```
### Continous and Categorical columns
```
cont_nn,cat_nn = cont_cat_split(df_nn, max_card=9000, dep_var=dep_var) #Max_card makes it so that any col with more than
# 9000 lvls, it will be treated as cont
cont_nn
```
> Notice that it's missing saleElpased from the cont_nn. We need to add this as we want this col to be treated as cont.
```
cont_nn.append('saleElapsed')
cont_nn
cat_nn.remove('saleElapsed')
df_nn.dtypes['saleElapsed'] #must change to int as an object type will cause error
df_nn['saleElapsed'] = df_nn['saleElapsed'].astype('int')
```
### Split
We want to split our data by date, not randomly.
```
cond = (df_nn.saleYear<2011) | (df_nn.saleMonth<10)
train_idx = np.where( cond)[0]
valid_idx = np.where(~cond)[0]
splits = (list(train_idx),list(valid_idx))
```
## Tabular object
Now that we have everything we need, lets create our tabular object.
```
procs_nn = [Categorify, FillMissing, Normalize]
to_nn = TabularPandas(df_nn, procs_nn, cat_nn, cont_nn, splits=splits, y_names=dep_var)
dls = to_nn.dataloaders(1024) #minibatches
y = to_nn.train.y
y.min(),y.max()
```
## Training
```
learn = tabular_learner(dls, y_range=(8,12), layers=[500,250],
n_out=1, loss_func=F.mse_loss)
learn.lr_find() #find best lr
learn.fit_one_cycle(5, 1e-2) #train
#Functions will be using to calc error
def r_mse(pred,y): return round(math.sqrt(((pred-y)**2).mean()), 6)
def m_rmse(m, xs, y): return r_mse(m.predict(xs), y)
preds,targs = learn.get_preds()
r_mse(preds,targs)
```
> This is actually very good
## Random Forest
Lets now create our random forest and compare it to our NN
### Tabular object
```
procs = [Categorify, FillMissing]
rf_to = TabularPandas(df_nn, procs, cat_nn, cont_nn, y_names=dep_var, splits=splits)
#Grab our x,y
xs,y = rf_to.train.xs,rf_to.train.y
valid_xs,valid_y = rf_to.valid.xs,rf_to.valid.y
```
### Random Forest
```
#Method below creates our random forest and fits it
def rf(xs, y, n_estimators=40, max_samples=200_000, max_features=0.5, min_samples_leaf=5, **kwargs):
return RandomForestRegressor(n_jobs=-1, n_estimators=n_estimators,
max_samples=max_samples, max_features=max_features,
min_samples_leaf=min_samples_leaf, oob_score=True).fit(xs, y)
m = rf(xs, y) #Fitting
m_rmse(m, xs, y), m_rmse(m, valid_xs, valid_y)
```
> So it seems our random forest preformed worse in comparison to the NN. Let's improve this by adding the NN embeddings!
# Adding embeddings
```
learn.model.embeds[:5] #These are just some of the embedding within the NN
```
**The function below extracts the embeddings from the model**
```
def embed_features(learner, xs):
xs = xs.copy()
for i, feature in enumerate(learn.dls.cat_names):
emb = learner.model.embeds[i].cpu()
new_feat = pd.DataFrame(emb(tensor(xs[feature], dtype=torch.int64)), index=xs.index, columns=[f'{feature}_{j}' for j in range(emb.embedding_dim)])
xs.drop(columns=feature, inplace=True)
xs = xs.join(new_feat)
return xs
#Extracting embeddings
embeded_xs = embed_features(learn, learn.dls.train.xs)
xs_valid = embed_features(learn, learn.dls.valid.xs)
embeded_xs.shape, xs_valid.shape
```
### Fitting embeddings
Now that we have our embeddings, lets fit it into the random forest.
```
m = rf(embeded_xs, y) #Fitting
m_rmse(m, embeded_xs, y), m_rmse(m, xs_valid, valid_y)
```
> It seems that adding the NN embeddings improves the random forest!
| github_jupyter |
# 1-7.2 Intro Python Practice
## `while()` loops & increments
<font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
- create forever loops using `while` and `break`
- use incrementing variables in a while loop
- control while loops using Boolean operators
```
# [ ] use a "forever" while loop to get user input of integers to add to sum,
# until a non-digit is entered, then break the loop and print sum
sum = 0
while True:
addition = input("enter an integer: ")
if addition.isdigit():
sum += int(addition)
else:
print(sum)
break
# [ ] use a while True loop (forever loop) to give 4 chances for input of a correct color in a rainbow
# rainbow = "red orange yellow green blue indigo violet"
rainbow = "red orange yellow green blue indigo violet"
cnt = 1
get_rainbow = input("enter rainbow colors: ")
while True:
if get_rainbow == rainbow:
print("Great!")
break
elif cnt < 4:
print("Wrong, try again!")
cnt += 1
else:
break
get_rainbow = input("enter rainbow colors: ")
# [ ] Get input for a book title, keep looping while input is Not in title format (title is every word capitalized)
title = input("enter a book title: ")
while title != title.title():
title = input("enter a book title: ")
# [ ] create a math quiz question and ask for the solution until the input is correct
while True:
answer = input("How many yards are in one mile: ")
if answer == '1760':
break
```
### Fix the Error
```
# [ ] review the code, run, fix the error
tickets = int(input("enter tickets remaining (0 to quit): "))
while tickets > 0:
# if tickets are multiple of 3 then "winner"
if int(tickets/3) == tickets/3:
print("you win!")
else:
print("sorry, not a winner.")
tickets = int(input("enter tickets remaining (0 to quit): "))
print("Game ended")
```
### create a function: quiz_item() that asks a question and tests if input is correct
- quiz_item()has 2 parameter **strings**: question and solution
- shows question, gets answer input
- returns True if `answer == solution` or continues to ask question until correct answer is provided
- use a while loop
create 2 or more quiz questions that call quiz_item()
**Hint**: provide multiple choice or T/F answers
```
# Create quiz_item() and 2 or more quiz questions that call quiz_item()
def quiz_item(question, solution):
print(question)
while True:
answer = input("enter your answer: ")
if answer == solution:
print("Yes!")
break
return True
else:
print("Sorry, wrong answer. Try again!")
question = "When Python 2 support is end?"
solution = "2020"
quiz_item(question, solution)
question = "How Python style guide is called?"
solution = "PEP8"
quiz_item(question, solution)
```
[Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| github_jupyter |
# Importing MIMIC IV data into a graph database
### About the data
The data were obtaind from Physionet, where the following abstract describes the data: "The Medical Information Mart for Intensive Care (MIMIC)-III database provided critical care data for over 40,000 patients admitted to intensive care units at the Beth Israel Deaconess Medical Center (BIDMC). Importantly, MIMIC-III was deidentified, and patient identifiers were removed according to the Health Insurance Portability and Accountability Act (HIPAA) Safe Harbor provision. MIMIC-III has been integral in driving large amounts of research in clinical informatics, epidemiology, and machine learning. Here we present MIMIC-IV, an update to MIMIC-III, which incorporates contemporary data and improves on numerous aspects of MIMIC-III. MIMIC-IV adopts a modular approach to data organization, highlighting data provenance and facilitating both individual and combined use of disparate data sources. MIMIC-IV is intended to carry on the success of MIMIC-III and support a broad set of applications within healthcare."
## Import MIMIC-IV Data to into a graph database
---
Data downloaded from https://physionet.org/content/mimiciv/0.4/ on 11 February, 2021. This is version 0.4 of the MIMIC-IV data, which was published 13 Aug 2020.
The downloaded data consisted of the following 27 CSV files, which collectively used 66.5 GB of memory:
- admissions.csv
- chartevents.csv
- datetimeevents.csv
- d_hcpcs.csv
- diagnoses_icd.csv
- d_icd_diagnoses.csv
- d_icd_procedures.csv
- d_items.csv
- d_labitems.csv
- drgcodes.csv
- emar.csv
- emar_detail.csv
- hcpcsevents.csv
- icustays.csv
- inputevents.csv
- labevents.csv
- microbiologyevents.csv
- outputevents.csv
- patients.csv
- pharmacy.csv
- poe.csv
- poe_detail.csv
- prescriptions.csv
- procedureevents.csv
- procedures_icd.csv
- services.csv
- transfers.csv
These files were placed in the Import folder of the MIMIC-IV Neo4j database to make them readily available for import into the graph.
```
import pandas as pd
path = '/home/tim/.local/share/neo4j-relate/dbmss/dbms-3077a569-80a7-4968-9e81-743773698121/import/'
# Create a list of all CSV files to import
csv_files = ['admissions.csv', 'chartevents.csv', 'datetimeevents.csv', 'd_hcpcs.csv', 'diagnoses_icd.csv', 'd_icd_diagnoses.csv', 'd_icd_procedures.csv', 'd_items.csv', 'd_labitems.csv', 'drgcodes.csv', 'emar.csv', 'emar_detail.csv', 'hcpcsevents.csv', 'icustays.csv', 'inputevents.csv', 'labevents.csv', 'microbiologyevents.csv', 'outputevents.csv', 'patients.csv', 'pharmacy.csv', 'poe.csv', 'poe_detail.csv', 'prescriptions.csv', 'procedureevents.csv', 'procedures_icd.csv', 'services.csv', 'transfers.csv']
# Create a dictionary with file names as keys and the list of headers for each file as values
headers_dict = {}
for file in csv_files:
headers = pd.read_csv(path+file, nrows=1)
headers = headers.columns.tolist()
if not file in headers_dict:
headers_dict[file] = headers
# Inspect an example item in the dictionary
print(headers_dict['admissions.csv'])
# Create a function that writes the string for a cypher command
# to create nodes from each CSV file
def csv_to_node(csv_file):
# Create the node label based on the CSV file name. Place it in title case and remove the '.csv' suffix
label= csv_file[:-4].title()
# Convert the CSV's headers into node properties
properties = '{'
col_index = 0
for header in headers_dict[csv_file]:
properties = properties+header+':COLUMN['+str(col_index)+'], '
col_index += 1
properties = properties[:-2]+'}' # Delete last comma of the list and add the ending curly bracket
# Compile the complete cypher command
cypher = '''USING PERIODIC COMMIT 100000 LOAD CSV FROM "file:///{csv_file}" AS COLUMN CREATE (n:Mimic4:{label} {properties})'''.format(csv_file=csv_file, label=label, properties=properties)
return cypher
# Generate the cypher code for a single csv file to test in the Neo4j browser
csv_to_node('d_labitems.csv')
```
### Initialize a connection to the neo4j database.
```
import getpass
password = getpass.getpass("\nPlease enter the Neo4j database password to continue \n")
from neo4j import GraphDatabase
driver=GraphDatabase.driver(uri="bolt://localhost:7687", auth=('neo4j',password))
session=driver.session()
# Create all nodes
for csv_name in csv_files:
query = csv_to_node(csv_name)
session.run(query)
```
### Create relationships
```
# Create all relationships
query_list = ['MATCH (n:Admissions), (m:Chartevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Datetimeevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Diagnoses_Icd) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Drgcodes) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Icustays) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Inputevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Labevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Microbiologyevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Outputevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Prescriptions) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Procedureevents) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Procedures_Icd) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Services) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Admissions), (m:Transfers) WHERE m.hadm_id = n.hadm_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Chartevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Datetimeevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Inputevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Microbiologyevents) WHERE m.spec_itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Microbiologyevents) WHERE m.org_itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Microbiologyevents) WHERE m.ab_itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Outputevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Items), (m:Procedureevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:D_Labitems), (m:Labevents) WHERE m.itemid = n.itemid MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Diagnoses_Icd), (m:D_Icd_Diagnoses) WHERE m.icd_code = n.icd_code MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Emar_Detail), (m:Emar) WHERE m.emar_id = n.emar_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Hcpcsevents), (m:D_Hcpcs) WHERE m.code = n.hcpcs_cd MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Chartevents) WHERE m.stay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Datetimeevents) WHERE m.stay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Inputevents) WHERE m.stay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Outputevents) WHERE m.stay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Procedureevents) WHERE m.stay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Icustays), (m:Transfers) WHERE m.icustay_id = n.stay_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Admissions) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Chartevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Datetimeevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Diagnoses_Icd) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Drgcodes) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Icustays) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Inputevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Labevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Microbiologyevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Outputevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Prescriptions) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Procedureevents) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Procedures_Icd) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Services) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Patients), (m:Transfers) WHERE m.subject_id = n.subject_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Pharmacy), (m:Emar) WHERE m.pharmacy_id = n.pharmacy_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Pharmacy), (m:Emar_detail) WHERE m.pharmacy_id = n.pharmacy_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Poe), (m:Emar) WHERE m.poe_id = n.poe_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Poe), (m:Pharmacy) WHERE m.poe_id = n.poe_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Poe_Detail), (m:Poe) WHERE m.poe_id = n.poe_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Prescriptions), (m:Pharmacy) WHERE m.pharmacy_id = n.pharmacy_id MERGE (n)<-[:SQL_CHILD_OF ]-(m)', 'MATCH (n:Procedures_Icd), (m:D_Icd_Procedures) WHERE m.icd_code = n.icd_code MERGE (n)<-[:SQL_CHILD_OF ]-(m)']
count = 0
for command in query_list:
session.run(command)
count += 1
print(count+'of 54')
```
### Close the connection to the neo4j database
```
session.close()
```
### Data references:
MIMIC IV:
Johnson, A., Bulgarelli, L., Pollard, T., Horng, S., Celi, L. A., & Mark, R. (2020). MIMIC-IV (version 0.4). PhysioNet. https://doi.org/10.13026/a3wn-hq05
Physionet:
Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., ... & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation. 101(23), pp. e215-e220.
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import os
import networkx as nx
import matplotlib.pyplot as plt
import caselawnet
filepath = '/media/sf_VBox_Shared/CaseLaw/2018-01-29-lido/derived/'
links_sub = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-to-case-links.csv'))
cases_sub = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-nodes-sub.csv'))
links_sub_ext = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-to-case-links-ext.csv'))
cases_sub_ext = pd.read_csv(os.path.join(filepath, 'subnetwork', 'case-nodes-ext.csv'))
print('Subnetwork: {} nodes, {} links'.format(len(cases_sub), len(links_sub)))
print('Extended subnetwork: {} nodes, {} links'.format(len(cases_sub_ext), len(links_sub_ext)))
cases_sub.head()
cases_sub_ext.head()
def add_network_statistics(nodes, links):
if len(nodes)==0:
return nodes
graph = nx.readwrite.json_graph.node_link_graph({'nodes': nodes, 'links': links}, directed=True, multigraph=False)
degree = nx.degree(graph)
if max(dict(degree).values()) > 0:
hubs, authorities = caselawnet.network_analysis.get_hits(graph)
statistics = {
'degree': degree,
'in_degree': graph.in_degree(),
'out_degree': graph.out_degree(),
'degree_centrality': nx.degree_centrality(graph),
'in_degree_centrality': nx.in_degree_centrality(graph),
'out_degree_centrality': nx.out_degree_centrality(graph),
'betweenness_centrality': nx.betweenness_centrality(graph),
'closeness_centrality': nx.closeness_centrality(graph),
'pagerank': caselawnet.network_analysis.get_pagerank(graph),
'hubs': hubs,
'authorities': authorities
}
else:
statistics = {}
for i, node in enumerate(nodes):
nodeid = node['id']
for var in statistics.keys():
node[var] = statistics[var][nodeid]
caselawnet.network_analysis.get_community(graph, nodes)
return graph, nodes
cases_sub_list = cases_sub.rename_axis({'link_id':'id'}, axis=1).to_dict(orient='records')
links_sub_list = links_sub.rename_axis({'link_id':'id'}, axis=1).to_dict(orient='records')
cases_ext_list = cases_sub_ext.rename_axis({'link_id':'id'}, axis=1).to_dict(orient='records')
links_ext_list = links_sub_ext.rename_axis({'link_id':'id'}, axis=1).to_dict(orient='records')
g_sub, nodes_sub = add_network_statistics(cases_sub_list, links_sub_list)
g_ext, nodes_ext = add_network_statistics(cases_ext_list, links_ext_list)
nodes_sub_out = [n for n in nodes_sub if n['degree']>0]
nodes_ext_out = [n for n in nodes_ext if n['degree']>0]
for n in nodes_sub_out:
n['subject'] = 'unknown'
n['creator'] = n['court']
n['title'] = n['ecli']
for n in nodes_ext_out:
n['subject'] = 'unknown'
n['creator'] = n['court']
n['title'] = n['ecli']
caselawnet.to_sigma_json(nodes_sub_out, links_sub_list, 'Employer liability', os.path.join(filepath, 'subnetwork', 'network_sub.json'))
caselawnet.to_sigma_json(nodes_ext_out, links_ext_list, 'Employer liability, extended', os.path.join(filepath, 'subnetwork', 'network_sub_ext.json'))
nodes_sub[0]
nodes_sub_df = pd.DataFrame(nodes_sub)
nodes_ext_df = pd.DataFrame(nodes_ext)
nodes_sub_df.describe()
nodes_sub_df.sort_values('authorities', ascending=False)[['ecli', 'authorities']].head(30)
nodes_sub_df[nodes_sub_df['ecli']=='ECLI:NL:HR:2008:BD1847']
# What are the highest ranked nodes?
for att in ['in_degree', 'authorities', 'betweenness_centrality']:
print(att)
print("subnetwork")
print(nodes_sub_df.sort_values(att, ascending=False)[['ecli', att]].head())
print('extended')
print(nodes_ext_df.sort_values(att, ascending=False)[['ecli', att]].head())
# What are the highest ranked nodes?
nodes_ext_df.sort_values('in_degree', ascending=False).head()
def networks_per_year(nodes_df, links_list):
networks = {}
for year in range(int(min(nodes_df['year'])), int(max(nodes_df['year']))+1):
nodes_filtered_df = nodes_df[nodes_df['year']<=year]
nodes_filtered = nodes_filtered_df.to_dict(orient='records')
links_filtered = [l for l in links_list
if l['source'] in nodes_filtered_df['id'].as_matrix()
and l['target'] in nodes_filtered_df['id'].as_matrix()]
if len(links_filtered) > 0:
g, nodes_year = add_network_statistics(nodes_filtered.copy(), links_filtered.copy())
networks[year] = {'nodes': nodes_year, 'edges': links_filtered}
years = list(networks.keys())
years.sort()
nodes_total_df = pd.concat(
[pd.DataFrame(networks[year]['nodes']).set_index('ecli') for year in years], keys=years)
year_stats = pd.DataFrame(index=years)
year_stats['nr_nodes'] = nodes_total_df.groupby(level=0)['id'].count()
year_stats['nr_links'] = [len(networks[year]['edges']) for year in year_stats.index]
for var in ['in_degree', 'out_degree', 'degree',
'in_degree_centrality', 'out_degree_centrality', 'degree_centrality',
'authorities', 'hubs', 'pagerank', 'betweenness_centrality', 'closeness_centrality']:
year_stats['avg_'+var] = nodes_total_df.groupby(level=0)[var].mean()
return networks, nodes_total_df, year_stats
networks_sub, nodes_years_sub, year_stats_sub = networks_per_year(cases_sub, links_sub_list)
networks_ext, nodes_years_ext, year_stats_ext = networks_per_year(cases_sub_ext, links_ext_list)
fig, axes = plt.subplots(2, 2, figsize=(15,10), sharey='row')
year_stats_sub[['nr_nodes', 'nr_links']].plot(kind='bar', ax=axes[0,0])
axes[0,0].set_title('Subnetwork')
year_stats_ext[['nr_nodes', 'nr_links']].plot(kind='bar', ax=axes[0,1])
axes[0,1].set_title('Subnetwork extended with citing cases')
pd.DataFrame({'subnetwork': year_stats_sub['avg_degree'], 'extended': year_stats_ext['avg_degree']}).plot(ax=axes[1,0], kind='bar', cmap='binary', edgecolor='black')
axes[1,0].set_title('Average degree of complete network')
pd.DataFrame({'subnetwork': nodes_sub_df.groupby('year')['out_degree'].mean(),
'extended': nodes_ext_df.groupby('year')['out_degree'].mean()}).plot(ax=axes[1,1], kind='bar', cmap='binary', edgecolor='black')
axes[1,1].set_title('Average out-degree of nodes per year')
plt.show()
fig, axes = plt.subplots(2)
for i, nodes in enumerate([nodes_sub_df, nodes_ext_df]):
values = nodes.groupby(['year', 'out_degree']).size().unstack(fill_value=0)
values.plot(kind='bar', stacked=True, cmap='Blues', linewidth=1, edgecolor='black', ax=axes[i])
axes[i].legend(bbox_to_anchor=(1.1, 1), loc='upper left')
```
| github_jupyter |
# Introduction to SQLite &
Selecting Sources from the Sloan Digital Sky Survey
========
#### Version 0.1
***
By AA Miller 2019 Mar 25
As noted earlier, there will be full lectures on databases over the remainder of this week.
This notebook provides a quick introduction to [`SQLite`](https://sqlite.org/index.html) a lightweight implementation of a Structured Query Language (SQL) database. One of the incredibly nice things about `SQLite` is the low overhead needed to set up a database (as you will see in a minute). We will take advantage of this low overhead to build a database later in the week.
```
import matplotlib.pyplot as plt
%matplotlib notebook
```
At the most basic level - databases *store your bytes*, and later *return those bytes* (or a subset of them) when queried.
They provide a highly efficient means for filtering your bytes (there are many different strategies that the user can employ).
The backend for most databases is the **Structured Query Language** or SQL, which is a standard declarative language.
There are many different libraries that implement SQL: MySQL, PostgreSQL, Greenplum, Microsoft SQL server, IBM DB2, Oracle Database, etc.
## Problem 1) Basic SQL Operations with SQLite
The most basic implementation is [`SQLite`](https://www.sqlite.org) a self-contained, SQL database engine. We will discuss `SQLite` further later in the week, but in brief - it is a nice stand alone package that works really well for small problems (such as the example that we are about to encounter).
```
import sqlite3
```
Without diving too much into the weeds (we'll investigate this further later this week), we need to establish a [`connection`](https://docs.python.org/2/library/sqlite3.html#sqlite3.Connection) to the database. From the `connection` we create a [`cursor`](https://docs.python.org/2/library/sqlite3.html#sqlite3.Connection), which allows us to actually interact with the database.
```
conn = sqlite3.connect("intro.db")
cur = conn.cursor()
```
And just like that - we have now created a new database `intro.db`, with which we can "store bytes" or later "retrieve bytes" once we have added some data to the database.
*Aside* - note that unlike many SQL libraries, `SQLite` does not require a server and creates an actual database file on your hard drive. This improves portability, but also creates some downsides as well.
Now we need to create a table and insert some data. We will interact with the database via the [`execute()`](https://docs.python.org/2/library/sqlite3.html#sqlite3.Cursor.execute) method for the `cursor` object.
Recall that creating a table requires a specification of the table name, the columns in the table, and the data type for each column. Here's an example where I create a table to store info on my pets:
cur.execute("""create table PetInfo(
Name text,
Species text,
Age tinyint,
FavoriteFood text
)""")
**Problem 1a**
Create a new table in the database called `DSFPstudents` with columns `Name`, `Institution`, and `Year`, where `Year` is the year in graduate school.
```
cur.execute( # complete
```
Once a table is created, we can use the database to store bytes. If I were to populate my `PetInfo` table I would do the following:
cur.execute("""insert into PetInfo(Name, Species, Age, FavoriteFood)
values ("Rocky", "Dog", 12, "Bo-Nana")""")
cur.execute("""insert into PetInfo(Name, Species, Age, FavoriteFood)
values ("100 Emoji-Flames Emoji", "Red Panda", 2, "bamboo leaves")""")
*Note* - column names do not need to be explicitly specified, but for clarity this is always preferred.
**Problem 1b**
Insert data for yourself, and the two people sitting next to you into the database.
```
cur.execute( # complete
```
Now that we have bytes in the database, we can retrieve those bytes with one (or several) queries. There are 3 basic building blocks to a query:
SELECT...
FROM...
WHERE...
Where `SELECT` specifies the information we want to retrieve from the database, `FROM` specifies the tables being queried in the database, and `WHERE` specifies the conditions for the query.
**Problem 1c**
Select the institutions for all students in the `DSFPstudents` table who have been in grad school for more than 2 years.
*Hint* - to display the results of your query run `cur.fetchall()`.
```
cur.execute( # complete
cur.fetchall()
```
In closing this brief introduction to databases, note that good databases follow the 4 ACID properties:
1. Atomicity
2. Consistency
3. Isolation
4. Durability
In closing this brief introduction to databases, note that good databases follow the 4 ACID properties:
1. Atomicity - all parts of transaction succeed, or rollback state of database
2. Consistency
3. Isolation
4. Durability
In closing this brief introduction to databases, note that good databases follow the 4 ACID properties:
1. Atomicity - all parts of transaction succeed, or rollback state of database
2. Consistency - data always meets validation rules
3. Isolation
4. Durability
In closing this brief introduction to databases, note that good databases follow the 4 ACID properties:
1. Atomicity - all parts of transaction succeed, or rollback state of database
2. Consistency - data always meets validation rules
3. Isolation - no interference across transactions (even if concurrent)
4. Durability
In closing this brief introduction to databases, note that good databases follow the 4 ACID properties:
1. Atomicity - all parts of transaction succeed, or rollback state of database
2. Consistency - data always meets validation rules
3. Isolation - no interference across transactions (even if concurrent)
4. Durability - a committed transaction remains committed (even if there's a power outage, etc)
## Problem 2) Complex Queries with SDSS
Above we looked at the most basic operations possible with a database (recall - databases are unnecessary, and possibly cumbersome, with small data sets). A typical database consists of many tables, and these tables may be joined together to unlock complex questions for the data.
As a reminder on (some of) this functionality, we are now going to go through some problems using the SDSS database. The full [SDSS schema](http://skyserver.sdss.org/dr13/en/help/browser/browser.aspx) explains all of the tables, columns, views and functions for querying the database. We will keep things relatively simple in that regard.
```
# you may need to run conda install -c astropy astroquery
from astroquery.sdss import SDSS
```
[`astroquery`](http://astroquery.readthedocs.io/en/latest/) enables seemless connections to the SDSS database via the Python shell.
**Problem 2a**
Select 20 random sources from the [`PhotoObjAll`](https://skyserver.sdss.org/dr13/en/help/docs/tabledesc.aspx?name=PhotoObjAll) table and return all columns in the table.
*Hint* - while this would normally be accomplished by starting the query `select limit 20 ...`, SDSS CasJobs uses Microsoft's SQL Server, which adopts `select top 20 ...` to accomplish an identical result.
```
SDSS.query_sql( # complete
```
That's more columns than we will likely ever need. Instead, let's focus on `objID`, a unique identifier, `cModelMag_u`, `cModelMag_g`, `cModelMag_r`, `cModelMag_i`, and `cModelMag_z`, the source magnitude in $u', g', r', i', z'$, respectively.
We will now introduce the concept of joining two tables.
The most common operation is known as an `inner join` (which is often referred to as just `join`). An `inner join` returns records that have matching sources in both tables in the join.
Less, but nevertheless still powerful, is the `outer join`. An outer join returns *all* records in either table, with `NULL` values for columns in a table in which the record does not exist.
Specialized versions of the `outer join` include the `left join` and `right join`, whereby *all* records in either the left or right table, respectively, are returned along with their counterparts.
**Problem 2b**
Select `objid` and $u'g'r'i'z'$ from `PhotoObjAll` and the corresponding `class` from [`specObjAll`](https://skyserver.sdss.org/dr13/en/help/docs/tabledesc.aspx?name=SpecObjAll) for 20 random sources.
There are multiple columns you could use to join the tables, in this case match `objid` to `bestobjid` from `specObjAll` and use an `inner join`.
```
SDSS.query_sql( # complete
```
**Problem 2c**
Perform an identical query to the one above, but this time use a `left outer join` (or `left join`).
How do your results compare to the previous query?
```
SDSS.query_sql( # complete
```
**Problem 2d**
This time use a `right outer join` (or `right join`).
How do your results compare to the previous query?
```
SDSS.query_sql( # complete
```
## Challenge Problem
To close the notebook we will perform a nested query. In brief, the idea is to join the results of one query with a separate query.
Here, we are going to attempt to identify bright AGN that don't have SDSS spectra. To do so we will need the `photoObjAll` table, the `specObjAll` table, and the `rosat` table, which includes all cross matches between SDSS sources and X-ray sources detected by the [Rosat satellite](https://heasarc.gsfc.nasa.gov/docs/rosat/rosat3.html).
Create a nested query that selects all *Rosat* sources that don't have SDSS spectra with `cModelFlux_u + cModelFlux_g + cModelFlux_r + cModelFlux_i + cModelFlux_z > 10000` (this flux contraint ensures the source is bright without making any cuts on color) and `type = 3`, this last constraint means the source is extended in SDSS images.
*Hint* - you may run into timeout issues in which case you should run the query on CasJobs.
```
SDSS.query_sql( # complete
```
| github_jupyter |
<a href="https://colab.research.google.com/github/airctic/icevision/blob/master/notebooks/custom_parser.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Custom Parser - Simple
This tutorial uses the [Global Wheat Detection](http://www.global-wheat.com/) dataset, you can download it from Kaggle [here](https://www.kaggle.com/c/global-wheat-detection/data).
## Instaling icevision
```
!pip install icevision[all]
```
## Imports
As always, let's import everything from `icevision`. Additionally, we will also need `pandas` (you might need to install it with `pip install pandas`).
```
from icevision.all import *
import pandas as pd
```
## Understand the data format
In this task we were given a `.csv` file with annotations, let's take a look at that.
!!! danger "Important"
Replace `source` with your own path for the dataset directory.
```
source = Path("/home/lgvaz/data/wheat")
df = pd.read_csv(source / "train.csv")
df.head()
```
At first glance, we can make the following assumptions:
* Multiple rows with the **same** object_id, width, height
* A **different** bbox for each row
* source doesn't seem relevant right now
Once we know what our data provides we can create our custom `Parser`.
## Create the Parser
When creating a `Parser` we inherit from smaller building blocks that provides the functionallity we want:
* `parsers.FasterRCNN`: Since we only need to predict bboxes we will use a `FasterRCNN` model, this will parse all the requirements for using such a model.
* `parsers.FilepathMixin`: Provides the requirements for parsing images filepaths.
* `parsers.SizeMixin`: Provides the requirements for parsing the image dimensions.
The first step is to create a class that inherits from these smaller building blocks:
```
class WheatParser(parsers.Parser, parsers.FilepathMixin, parsers.LabelsMixin, parsers.BBoxesMixin):
pass
```
We now use a method `generate_template` that will print out all the necessary methods we have to implement.
```
WheatParser.generate_template()
```
With this, we know what methods we have to implement and what each one should return (thanks to the type annotations)!
Defining the `__init__` is completely up to you, normally we have to pass our data (the `df` in our case) and the folder where our images are contained (`source` in our case).
We then override `__iter__`, telling our parser how to iterate over our data. In our case we call `df.itertuples` to iterate over all `df` rows.
`__len__` is not obligatory but will help visualizing the progress when parsing.
And finally we override all the other methods, they all receive a single argument `o`, which is the object returned by `__iter__` (a single `DataFrame` row here).
!!! danger "Important"
Be sure to return the correct type on all overriden methods!
```
class WheatParser(parsers.FasterRCNN, parsers.FilepathMixin, parsers.SizeMixin):
def __init__(self, df, source):
self.df = df
self.source = source
def __iter__(self):
yield from self.df.itertuples()
def __len__(self):
return len(self.df)
def imageid(self, o) -> Hashable:
return o.image_id
def filepath(self, o) -> Union[str, Path]:
return self.source / f"{o.image_id}.jpg"
def image_width_height(self, o) -> Tuple[int, int]:
return get_image_size(self.filepath(o))
def labels(self, o) -> List[int]:
return [1]
def bboxes(self, o) -> List[BBox]:
return [BBox.from_xywh(*np.fromstring(o.bbox[1:-1], sep=","))]
```
Let's randomly split the data and parser with `Parser.parse`:
```
parser = WheatParser(df, source / "train")
train_rs, valid_rs = parser.parse()
```
Let's take a look at one record:
```
show_record(train_rs[0], display_label=False)
```
## Conclusion
And that's it! Now that you have your data in the standard library record format, you can use it to create a `Dataset`, visualize the image with the annotations and basically use all helper functions that IceVision provides!
## Happy Learning!
If you need any assistance, feel free to join our [forum](https://discord.gg/JDBeZYK).
| github_jupyter |
# 1. SETTINGS
```
# libraries
import pandas as pd
import numpy as np
import scipy.stats
from sklearn.preprocessing import LabelEncoder
import matplotlib.pyplot as plt
# pandas options
pd.set_option("display.max_columns", None)
# ignore warnings
import warnings
warnings.filterwarnings("ignore")
# garbage collection
import gc
gc.enable()
```
# 2. FUNCTIONS
```
##### FUNCTION FOR COUNTING MISSINGS
def count_missings(data):
total = data.isnull().sum().sort_values(ascending = False)
percent = (data.isnull().sum() / data.isnull().count() * 100).sort_values(ascending = False)
table = pd.concat([total, percent], axis = 1, keys = ["Total", "Percent"])
table = table[table["Total"] > 0]
return table
##### FUNCTION FOR CONVERTING DATES
def convert_days(data, features, t = 12, rounding = True, replace = False):
for var in features:
if replace == True:
if rounding == True:
data[var] = round(-data[var]/t)
else:
data[var] = -data[var]/t
data[var][data[var] < 0] = None
else:
if rounding == True:
data["CONVERTED_" + str(var)] = round(-data[var]/t)
else:
data["CONVERTED_" + str(var)] = -data[var]/t
data["CONVERTED_" + str(var)][data["CONVERTED_" + str(var)] < 0] = None
return data
##### FUNCTION FOR CREATING LOGARITHMS
def create_logs(data, features, replace = False):
for var in features:
if replace == True:
data[var] = np.log(data[var].abs() + 1)
else:
data["LOG_" + str(var)] = np.log(data[var].abs() + 1)
return data
##### FUNCTION FOR CREATING FLAGS FOR MISSINGS
def create_null_flags(data, features = None):
if features == None:
features = data.columns
for var in features:
num_null = data[var].isnull() + 0
if num_null.sum() > 0:
data["ISNULL_" + str(var)] = num_null
return data
##### FUNCTION FOR TREATING FACTORS
def treat_factors(data, method = "label"):
# label encoding
if method == "label":
factors = [f for f in data.columns if data[f].dtype == "object"]
for var in factors:
data[var], _ = pd.factorize(data[var])
# dummy encoding
if method == "dummy":
data = pd.get_dummies(data, drop_first = True)
# dataset
return data
##### FUNCTION FOR COMPUTING ACCEPT/REJECT RATIOS
def compute_accept_reject_ratio(data, lags = [1, 3, 5]):
# preparations
dec_prev = data[["SK_ID_CURR", "SK_ID_PREV", "DAYS_DECISION", "NAME_CONTRACT_STATUS"]]
dec_prev["DAYS_DECISION"] = -dec_prev["DAYS_DECISION"]
dec_prev = dec_prev.sort_values(by = ["SK_ID_CURR", "DAYS_DECISION"])
dec_prev = pd.get_dummies(dec_prev)
# compuatation
for t in lags:
# acceptance ratios
tmp = dec_prev[["SK_ID_CURR", "NAME_CONTRACT_STATUS_Approved"]].groupby(["SK_ID_CURR"]).head(1)
tmp = tmp.groupby(["SK_ID_CURR"], as_index = False).mean()
tmp.columns = ["SK_ID_CURR", "APPROVE_RATIO_" + str(t)]
data = data.merge(tmp, how = "left", on = "SK_ID_CURR")
# rejection ratios
tmp = dec_prev[["SK_ID_CURR", "NAME_CONTRACT_STATUS_Refused"]].groupby(["SK_ID_CURR"]).head(1)
tmp = tmp.groupby(["SK_ID_CURR"], as_index = False).mean()
tmp.columns = ["SK_ID_CURR", "REJECT_RATIO_" + str(t)]
data = data.merge(tmp, how = "left", on = "SK_ID_CURR")
# dataset
return data
##### FUNCTION FOR AGGREGATING DATA
def aggregate_data(data, id_var, label = None):
### SEPARATE FEATURES
# display info
print("- Preparing the dataset...")
# find factors
data_factors = [f for f in data.columns if data[f].dtype == "object"]
# partition subsets
num_data = data[list(set(data.columns) - set(data_factors))]
fac_data = data[[id_var] + data_factors]
# display info
num_facs = fac_data.shape[1] - 1
num_nums = num_data.shape[1] - 1
print("- Extracted %.0f factors and %.0f numerics..." % (num_facs, num_nums))
##### AGGREGATION
# aggregate numerics
if (num_nums > 0):
print("- Aggregating numeric features...")
num_data = num_data.groupby(id_var).agg(["mean", "std", "min", "max"])
num_data.columns = ["_".join(col).strip() for col in num_data.columns.values]
num_data = num_data.sort_index()
# aggregate factors
if (num_facs > 0):
print("- Aggregating factor features...")
fac_data = fac_data.groupby(id_var).agg([("mode", lambda x: scipy.stats.mode(x)[0][0]),
("unique", lambda x: x.nunique())])
fac_data.columns = ["_".join(col).strip() for col in fac_data.columns.values]
fac_data = fac_data.sort_index()
##### MERGER
# merge numerics and factors
if ((num_facs > 0) & (num_nums > 0)):
agg_data = pd.concat([num_data, fac_data], axis = 1)
# use factors only
if ((num_facs > 0) & (num_nums == 0)):
agg_data = fac_data
# use numerics only
if ((num_facs == 0) & (num_nums > 0)):
agg_data = num_data
##### LAST STEPS
# update labels
if label != None:
agg_data.columns = [label + "_" + str(col) for col in agg_data.columns]
# impute zeros for SD
#stdevs = agg_data.filter(like = "_std").columns
#for var in stdevs:
# agg_data[var].fillna(0, inplace = True)
# display info
print("- Final dimensions:", agg_data.shape)
# return dataset
return agg_data
```
# 3. DATA IMPORT
```
# import data
train = pd.read_csv("../data/raw/application_train.csv")
test = pd.read_csv("../data/raw/application_test.csv")
buro = pd.read_csv("../data/raw/bureau.csv")
bbal = pd.read_csv("../data/raw/bureau_balance.csv")
prev = pd.read_csv("../data/raw/previous_application.csv")
card = pd.read_csv("../data/raw/credit_card_balance.csv")
poca = pd.read_csv("../data/raw/POS_CASH_balance.csv")
inst = pd.read_csv("../data/raw/installments_payments.csv")
# check dimensions
print("Application:", train.shape, test.shape)
print("Buro:", buro.shape)
print("Bbal:", bbal.shape)
print("Prev:", prev.shape)
print("Card:", card.shape)
print("Poca:", poca.shape)
print("Inst:", inst.shape)
# extract target
y = train[["SK_ID_CURR", "TARGET"]]
del train["TARGET"]
```
# 4. PREPROCESSING
## 4.1. APPLICATION DATA
```
# concatenate application data
appl = pd.concat([train, test])
del train, test
### FEATURE ENGINEERING
# income ratios
appl["CREDIT_BY_INCOME"] = appl["AMT_CREDIT"] / appl["AMT_INCOME_TOTAL"]
appl["ANNUITY_BY_INCOME"] = appl["AMT_ANNUITY"] / appl["AMT_INCOME_TOTAL"]
appl["GOODS_PRICE_BY_INCOME"] = appl["AMT_GOODS_PRICE"] / appl["AMT_INCOME_TOTAL"]
appl["INCOME_PER_PERSON"] = appl["AMT_INCOME_TOTAL"] / appl["CNT_FAM_MEMBERS"]
# career ratio
appl["PERCENT_WORKED"] = appl["DAYS_EMPLOYED"] / appl["DAYS_BIRTH"]
appl["PERCENT_WORKED"][appl["PERCENT_WORKED"] < 0] = None
# number of adults
appl["CNT_ADULTS"] = appl["CNT_FAM_MEMBERS"] - appl["CNT_CHILDREN"]
appl['CHILDREN_RATIO'] = appl['CNT_CHILDREN'] / appl['CNT_FAM_MEMBERS']
# number of overall payments
appl['ANNUITY LENGTH'] = appl['AMT_CREDIT'] / appl['AMT_ANNUITY']
# external sources
#appl["EXT_SOURCE_MIN"] = appl[["EXT_SOURCE_1", "EXT_SOURCE_2", "EXT_SOURCE_3"]].min(axis = 1)
#appl["EXT_SOURCE_MAX"] = appl[["EXT_SOURCE_1", "EXT_SOURCE_2", "EXT_SOURCE_3"]].max(axis = 1)
appl["EXT_SOURCE_MEAN"] = appl[["EXT_SOURCE_1", "EXT_SOURCE_2", "EXT_SOURCE_3"]].mean(axis = 1)
#appl["EXT_SOURCE_SD"] = appl[["EXT_SOURCE_1", "EXT_SOURCE_2", "EXT_SOURCE_3"]].std(axis = 1)
appl["NUM_EXT_SOURCES"] = 3 - (appl["EXT_SOURCE_1"].isnull().astype(int) +
appl["EXT_SOURCE_2"].isnull().astype(int) +
appl["EXT_SOURCE_3"].isnull().astype(int))
# number of documents
doc_vars = ["FLAG_DOCUMENT_2", "FLAG_DOCUMENT_3", "FLAG_DOCUMENT_4", "FLAG_DOCUMENT_5", "FLAG_DOCUMENT_6",
"FLAG_DOCUMENT_7", "FLAG_DOCUMENT_8", "FLAG_DOCUMENT_9", "FLAG_DOCUMENT_10", "FLAG_DOCUMENT_11",
"FLAG_DOCUMENT_12", "FLAG_DOCUMENT_13", "FLAG_DOCUMENT_14", "FLAG_DOCUMENT_15", "FLAG_DOCUMENT_16",
"FLAG_DOCUMENT_17", "FLAG_DOCUMENT_18", "FLAG_DOCUMENT_19", "FLAG_DOCUMENT_20", "FLAG_DOCUMENT_21"]
appl["NUM_DOCUMENTS"] = appl[doc_vars].sum(axis = 1)
# application date
appl["DAY_APPR_PROCESS_START"] = "Working day"
appl["DAY_APPR_PROCESS_START"][(appl["WEEKDAY_APPR_PROCESS_START"] == "SATURDAY") |
(appl["WEEKDAY_APPR_PROCESS_START"] == "SUNDAY")] = "Weekend"
# logarithms
log_vars = ["AMT_CREDIT", "AMT_INCOME_TOTAL", "AMT_GOODS_PRICE", "AMT_ANNUITY"]
appl = create_logs(appl, log_vars, replace = True)
# convert days
day_vars = ["DAYS_BIRTH", "DAYS_REGISTRATION", "DAYS_ID_PUBLISH", "DAYS_EMPLOYED", "DAYS_LAST_PHONE_CHANGE"]
appl = convert_days(appl, day_vars, t = 30, rounding = True, replace = True)
# age ratios
appl["OWN_CAR_AGE_RATIO"] = appl["OWN_CAR_AGE"] / appl["DAYS_BIRTH"]
appl["DAYS_ID_PUBLISHED_RATIO"] = appl["DAYS_ID_PUBLISH"] / appl["DAYS_BIRTH"]
appl["DAYS_REGISTRATION_RATIO"] = appl["DAYS_REGISTRATION"] / appl["DAYS_BIRTH"]
appl["DAYS_LAST_PHONE_CHANGE_RATIO"] = appl["DAYS_LAST_PHONE_CHANGE"] / appl["DAYS_BIRTH"]
##### FEATURE REMOVAL
drops = ['APARTMENTS_MEDI', 'BASEMENTAREA_MEDI', 'COMMONAREA_MEDI', 'ELEVATORS_MEDI', 'ENTRANCES_MEDI',
'FLOORSMAX_MEDI', 'FLOORSMIN_MEDI', 'LANDAREA_MEDI', 'LIVINGAPARTMENTS_MEDI', 'LIVINGAREA_MEDI',
'NONLIVINGAPARTMENTS_MEDI', 'NONLIVINGAREA_MEDI','YEARS_BEGINEXPLUATATION_MEDI', 'YEARS_BUILD_MEDI',
'APARTMENTS_MODE', 'BASEMENTAREA_MODE', 'COMMONAREA_MODE','ELEVATORS_MODE', 'ENTRANCES_MODE',
'FLOORSMAX_MODE', 'FLOORSMIN_MODE', 'LANDAREA_MODE', 'LIVINGAPARTMENTS_MODE', 'LIVINGAREA_MODE',
'NONLIVINGAPARTMENTS_MODE', 'NONLIVINGAREA_MODE', 'TOTALAREA_MODE', 'YEARS_BEGINEXPLUATATION_MODE']
appl = appl.drop(columns = drops)
# rename features
appl.columns = ["SK_ID_CURR"] + ["app_" + str(col) for col in appl.columns if col not in "SK_ID_CURR"]
# check data
appl.head()
# count missings
nas = count_missings(appl)
nas.head()
```
## 4.2. CREDIT BUREAU DATA
### 4.2.1. BBAL DATA
```
# check bbal data
bbal.head()
### FEATURE ENGINEERING
# loan default score
bbal["NUM_STATUS"] = 0
bbal["NUM_STATUS"][bbal["STATUS"] == "X"] = None
bbal["NUM_STATUS"][bbal["STATUS"] == "1"] = 1
bbal["NUM_STATUS"][bbal["STATUS"] == "2"] = 2
bbal["NUM_STATUS"][bbal["STATUS"] == "3"] = 3
bbal["NUM_STATUS"][bbal["STATUS"] == "4"] = 4
bbal["NUM_STATUS"][bbal["STATUS"] == "5"] = 5
bbal["LOAN_SCORE"] = bbal["NUM_STATUS"] / (abs(bbal["MONTHS_BALANCE"]) + 1)
loan_score = bbal.groupby("SK_ID_BUREAU", as_index = False).LOAN_SCORE.sum()
del bbal["NUM_STATUS"]
del bbal["LOAN_SCORE"]
# dummy encoding for STATUS
bbal = pd.get_dummies(bbal, columns = ["STATUS"], prefix = "STATUS")
# count missings
nas = count_missings(bbal)
nas.head()
### AGGREGATIONS
# total month count
cnt_mon = bbal[["SK_ID_BUREAU", "MONTHS_BALANCE"]].groupby("SK_ID_BUREAU").count()
del bbal["MONTHS_BALANCE"]
# aggregate data
agg_bbal = bbal.groupby("SK_ID_BUREAU").mean()
# add total month count
agg_bbal["MONTH_COUNT"] = cnt_mon
# add loan score
agg_bbal = agg_bbal.merge(loan_score, how = "left", on = "SK_ID_BUREAU")
# count missings
nas = count_missings(agg_bbal)
nas.head()
# check data
agg_bbal.head()
# clear memory
del bbal
```
### 4.2.2. BURO DATA
```
# check buro data
buro.head()
### MERGE
buro = buro.merge(right = agg_bbal.reset_index(), how = "left", on = "SK_ID_BUREAU")
##### FEATURE ENGINEERING
# number of buro loans
cnt_buro = buro[["SK_ID_CURR", "SK_ID_BUREAU"]].groupby(["SK_ID_CURR"], as_index = False).count()
cnt_buro.columns = ["SK_ID_CURR", "CNT_BURO_LOANS"]
buro = buro.merge(cnt_buro, how = "left", on = "SK_ID_CURR")
# amount ratios
buro["AMT_SUM_OVERDUE_RATIO_1"] = buro["AMT_CREDIT_SUM_OVERDUE"] / buro["AMT_ANNUITY"]
buro["AMT_SUM_OVERDUE_RATIO_2"] = buro["AMT_CREDIT_SUM_OVERDUE"] / buro["AMT_CREDIT_SUM"]
buro["AMT_MAX_OVERDUE_RATIO_1"] = buro["AMT_CREDIT_MAX_OVERDUE"] / buro["AMT_ANNUITY"]
buro["AMT_MAX_OVERDUE_RATIO_2"] = buro["AMT_CREDIT_MAX_OVERDUE"] / buro["AMT_CREDIT_SUM"]
buro["AMT_SUM_DEBT_RATIO_1"] = buro["AMT_CREDIT_SUM_DEBT"] / buro["AMT_CREDIT_SUM"]
buro["AMT_SUM_DEBT_RATIO_2"] = buro["AMT_CREDIT_SUM_DEBT"] / buro["AMT_CREDIT_SUM_LIMIT"]
# logarithms
log_vars = ["AMT_CREDIT_SUM", "AMT_CREDIT_SUM_DEBT", "AMT_CREDIT_SUM_LIMIT", "AMT_CREDIT_SUM_OVERDUE", "AMT_ANNUITY"]
buro = create_logs(buro, log_vars, replace = True)
# convert days
day_vars = ["DAYS_CREDIT", "CREDIT_DAY_OVERDUE", "DAYS_CREDIT_ENDDATE", "DAYS_ENDDATE_FACT", "DAYS_CREDIT_UPDATE"]
buro = convert_days(buro, day_vars, t = 1, rounding = False, replace = True)
# recency-weighted loan score
buro["WEIGHTED_LOAN_SCORE"] = buro["LOAN_SCORE"] / (buro["DAYS_CREDIT"] / 12)
# day differences
buro["DAYS_END_DIFF_1"] = buro["DAYS_ENDDATE_FACT"] - buro["DAYS_CREDIT_ENDDATE"]
buro["DAYS_END_DIFF_2"] = buro["DAYS_CREDIT_UPDATE"] - buro["DAYS_CREDIT_ENDDATE"]
buro["DAYS_DURATION_1"] = buro["DAYS_CREDIT_ENDDATE"] - buro["DAYS_CREDIT"]
buro["DAYS_DURATION_2"] = buro["DAYS_ENDDATE_FACT"] - buro["DAYS_CREDIT"]
# number of active buro loans
cnt_buro = buro[["SK_ID_CURR", "CREDIT_ACTIVE"]]
cnt_buro.columns = ["SK_ID_CURR", "CNT_BURO_ACTIVE"]
cnt_buro = cnt_buro[cnt_buro["CNT_BURO_ACTIVE"] == "Active"]
cnt_buro = cnt_buro[["SK_ID_CURR", "CNT_BURO_ACTIVE"]].groupby(["SK_ID_CURR"], as_index = False).count()
buro = buro.merge(cnt_buro, how = "left", on = "SK_ID_CURR")
buro["CNT_BURO_ACTIVE"].fillna(0, inplace = True)
# number of closed buro loans
cnt_buro = buro[["SK_ID_CURR", "CREDIT_ACTIVE"]]
cnt_buro.columns = ["SK_ID_CURR", "CNT_BURO_CLOSED"]
cnt_buro = cnt_buro[cnt_buro["CNT_BURO_CLOSED"] == "Closed"]
cnt_buro = cnt_buro[["SK_ID_CURR", "CNT_BURO_CLOSED"]].groupby(["SK_ID_CURR"], as_index = False).count()
buro = buro.merge(cnt_buro, how = "left", on = "SK_ID_CURR")
buro["CNT_BURO_CLOSED"].fillna(0, inplace = True)
# number of defaulted buro loans
cnt_buro = buro[["SK_ID_CURR", "CREDIT_ACTIVE"]]
cnt_buro.columns = ["SK_ID_CURR", "CNT_BURO_BAD"]
cnt_buro = cnt_buro[cnt_buro["CNT_BURO_BAD"] == "Bad debt"]
cnt_buro = cnt_buro[["SK_ID_CURR", "CNT_BURO_BAD"]].groupby(["SK_ID_CURR"], as_index = False).count()
buro = buro.merge(cnt_buro, how = "left", on = "SK_ID_CURR")
buro["CNT_BURO_BAD"].fillna(0, inplace = True)
# dummy encodnig for factors
buro = pd.get_dummies(buro, drop_first = True)
# count missings
nas = count_missings(buro)
nas.head()
### AGGREGATIONS
# count previous buro loans
cnt_buro = buro[["SK_ID_CURR", "SK_ID_BUREAU"]].groupby("SK_ID_CURR").count()
del buro["SK_ID_BUREAU"]
# aggregate data
agg_buro = aggregate_data(buro, id_var = "SK_ID_CURR", label = "buro")
# add buro loan count
agg_buro["buro_BURO_COUNT"] = cnt_buro
# clean up
omits = ["WEIGHTED_LOAN_SCORE"]
for var in omits:
del agg_buro["buro_" + str(var) + "_std"]
del agg_buro["buro_" + str(var) + "_min"]
del agg_buro["buro_" + str(var) + "_max"]
# count missings
nas = count_missings(agg_buro)
nas.head()
# check data
agg_buro.head()
# clear memory
del buro
```
## 4.3. PREVIOUS LOAN DATA
### 4.3.1. INST DATA
```
# check inst data
inst.head()
### FEATURE ENGINEERING
# days past due and days before due (no negative values)
inst['DPD'] = inst['DAYS_ENTRY_PAYMENT'] - inst['DAYS_INSTALMENT']
inst['DBD'] = inst['DAYS_INSTALMENT'] - inst['DAYS_ENTRY_PAYMENT']
inst['DPD'] = inst['DPD'].apply(lambda x: x if x > 0 else 0)
inst['DBD'] = inst['DBD'].apply(lambda x: x if x > 0 else 0)
# percentage and difference paid in each installment
inst['PAYMENT_PERC'] = inst['AMT_PAYMENT'] / inst['AMT_INSTALMENT']
inst['PAYMENT_DIFF'] = inst['AMT_INSTALMENT'] - inst['AMT_PAYMENT']
# logarithms
log_vars = ["AMT_INSTALMENT", "AMT_PAYMENT"]
inst = create_logs(inst, log_vars, replace = True)
# dummy encodnig for factors
inst = pd.get_dummies(inst, drop_first = True)
# count missings
nas = count_missings(inst)
nas.head()
### AGGREGATIONS
# count instalments
cnt_inst = inst[["SK_ID_PREV", "NUM_INSTALMENT_NUMBER"]].groupby("SK_ID_PREV").count()
del inst["NUM_INSTALMENT_NUMBER"]
# delete ID_CURR
inst_id = inst[["SK_ID_CURR", "SK_ID_PREV"]]
del inst["SK_ID_CURR"]
# aggregate data
agg_inst = aggregate_data(inst, id_var = "SK_ID_PREV")
# add instalment count
agg_inst["inst_INST_COUNT"] = cnt_inst
# put back ID_CURR
inst_id = inst_id.drop_duplicates()
agg_inst = inst_id.merge(right = agg_inst.reset_index(), how = "right", on = "SK_ID_PREV")
del agg_inst["SK_ID_PREV"]
# aggregate data (round 2)
agg_inst = aggregate_data(agg_inst, id_var = "SK_ID_CURR", label = "inst")
# count missings
nas = count_missings(agg_inst)
nas.head()
# check data
agg_inst.head()
# clear memory
del inst
```
### 4.3.2. POCA DATA
```
# check poca data
poca.head()
### FEATURE ENGINEERING
# installments percentage
poca["INSTALLMENTS_PERCENT"] = poca["CNT_INSTALMENT_FUTURE"] / poca["CNT_INSTALMENT"]
# dummy encodnig for factors
poca = pd.get_dummies(poca, drop_first = True)
# count missings
nas = count_missings(poca)
nas.head()
### AGGREGATIONS
# count months
cnt_mon = poca[["SK_ID_PREV", "MONTHS_BALANCE"]].groupby("SK_ID_PREV").count()
del poca["MONTHS_BALANCE"]
# delete ID_CURR
poca_id = poca[["SK_ID_CURR", "SK_ID_PREV"]]
del poca["SK_ID_CURR"]
# aggregate data
agg_poca = aggregate_data(poca, id_var = "SK_ID_PREV")
# add month count
agg_poca["poca_MON_COUNT"] = cnt_mon
# put back ID_CURR
poca_id = poca_id.drop_duplicates()
agg_poca = poca_id.merge(right = agg_poca.reset_index(), how = "right", on = "SK_ID_PREV")
del agg_poca["SK_ID_PREV"]
# aggregate data (round 2)
agg_poca = aggregate_data(agg_poca, id_var = "SK_ID_CURR", label = "poca")
# count missings
nas = count_missings(agg_poca)
nas.head()
# check data
agg_poca.head()
# clear memory
del poca
```
### 4.3.3. CARD DATA
```
# check card data
card.head()
### FEATURE ENGINEERING
# logarithms
log_vars = ["AMT_BALANCE", "AMT_CREDIT_LIMIT_ACTUAL", "AMT_DRAWINGS_ATM_CURRENT", "AMT_DRAWINGS_CURRENT",
"AMT_DRAWINGS_OTHER_CURRENT", "AMT_DRAWINGS_POS_CURRENT", "AMT_INST_MIN_REGULARITY",
"AMT_PAYMENT_CURRENT", "AMT_PAYMENT_TOTAL_CURRENT", "AMT_RECEIVABLE_PRINCIPAL",
"AMT_RECIVABLE", "AMT_TOTAL_RECEIVABLE"]
card = create_logs(card, log_vars, replace = True)
# dummy encodnig for factors
card = pd.get_dummies(card, drop_first = True)
# count missings
nas = count_missings(card)
nas.head()
### AGGREGATIONS
# count months
cnt_mon = card[["SK_ID_PREV", "MONTHS_BALANCE"]].groupby("SK_ID_PREV").count()
del card["MONTHS_BALANCE"]
# delete ID_CURR
card_id = card[["SK_ID_CURR", "SK_ID_PREV"]]
del card["SK_ID_CURR"]
# aggregate data
agg_card = aggregate_data(card, id_var = "SK_ID_PREV")
# add month count
agg_card["card_MON_COUNT"] = cnt_mon
# put back ID_CURR
card_id = card_id.drop_duplicates()
agg_card = card_id.merge(right = agg_card.reset_index(), how = "right", on = "SK_ID_PREV")
del agg_card["SK_ID_PREV"]
# aggregate data (round 2)
agg_card = aggregate_data(agg_card, id_var = "SK_ID_CURR", label = "card")
# count missings
nas = count_missings(agg_card)
nas.head()
# check data
agg_card.head()
# clear memory
del card
```
### 4.3.4. PREV DATA
```
# check card data
prev.head()
### FEATURE ENGINEERING
# amount ratios
prev["AMT_GIVEN_RATIO_1"] = prev["AMT_CREDIT"] / prev["AMT_APPLICATION"]
prev["AMT_GIVEN_RATIO_2"] = prev["AMT_GOODS_PRICE"] / prev["AMT_APPLICATION"]
prev["DOWN_PAYMENT_RATIO"] = prev["AMT_DOWN_PAYMENT"] / prev["AMT_APPLICATION"]
# logarithms
log_vars = ["AMT_CREDIT", "AMT_ANNUITY", "AMT_APPLICATION", "AMT_DOWN_PAYMENT", "AMT_GOODS_PRICE"]
prev = create_logs(prev, log_vars, replace = True)
# convert days
day_vars = ["DAYS_FIRST_DRAWING", "DAYS_FIRST_DUE", "DAYS_LAST_DUE_1ST_VERSION",
"DAYS_LAST_DUE", "DAYS_TERMINATION", "DAYS_DECISION"]
prev = convert_days(prev, day_vars, t = 1, rounding = False, replace = True)
# number of applications
cnt_prev = prev[["SK_ID_CURR", "SK_ID_PREV"]].groupby(["SK_ID_CURR"], as_index = False).count()
cnt_prev.columns = ["SK_ID_CURR", "CNT_PREV_APPLICATIONS"]
prev = prev.merge(cnt_prev, how = "left", on = "SK_ID_CURR")
# number of contracts
cnt_prev = prev[["SK_ID_CURR", "FLAG_LAST_APPL_PER_CONTRACT"]]
cnt_prev.columns = ["SK_ID_CURR", "CNT_PREV_CONTRACTS"]
cnt_prev = cnt_prev[cnt_prev["CNT_PREV_CONTRACTS"] == "Y"]
cnt_prev = cnt_prev[["SK_ID_CURR", "CNT_PREV_CONTRACTS"]].groupby(["SK_ID_CURR"], as_index = False).count()
prev = prev.merge(cnt_prev, how = "left", on = "SK_ID_CURR")
# number ratio
prev["APPL_PER_CONTRACT_RATIO"] = prev["CNT_PREV_APPLICATIONS"] / prev["CNT_PREV_CONTRACTS"]
# loan decision ratios
prev = compute_accept_reject_ratio(prev, lags = [1, 3, 5])
# day differences
prev["DAYS_DUE_DIFF_1"] = prev["DAYS_LAST_DUE_1ST_VERSION"] - prev["DAYS_FIRST_DUE"]
prev["DAYS_DUE_DIFF_2"] = prev["DAYS_LAST_DUE"] - prev["DAYS_FIRST_DUE"]
prev["DAYS_TERMINATION_DIFF_1"] = prev["DAYS_TERMINATION"] - prev["DAYS_FIRST_DRAWING"]
prev["DAYS_TERMINATION_DIFF_2"] = prev["DAYS_TERMINATION"] - prev["DAYS_FIRST_DUE"]
prev["DAYS_TERMINATION_DIFF_3"] = prev["DAYS_TERMINATION"] - prev["DAYS_LAST_DUE"]
# application dates
prev["DAY_APPR_PROCESS_START"] = "Working day"
prev["DAY_APPR_PROCESS_START"][(prev["WEEKDAY_APPR_PROCESS_START"] == "SATURDAY") |
(prev["WEEKDAY_APPR_PROCESS_START"] == "SUNDAY")] = "Weekend"
##### FEATURE REMOVAL
drops = ["NAME_CLIENT_TYPE", "SK_ID_PREV"]
prev = prev.drop(columns = drops)
# dummy encodnig for factors
prev = pd.get_dummies(prev, drop_first = True)
# count missings
nas = count_missings(prev)
nas.head()
### AGGREGATIONS
# aggregate data
agg_prev = aggregate_data(prev, id_var = "SK_ID_CURR", label = "prev")
# clean up
omits = ["APPROVE_RATIO_1", "APPROVE_RATIO_3", "APPROVE_RATIO_5",
"REJECT_RATIO_1", "REJECT_RATIO_3", "REJECT_RATIO_5",
"FLAG_LAST_APPL_PER_CONTRACT_Y", "CNT_PREV_CONTRACTS", "CNT_PREV_APPLICATIONS",
"APPL_PER_CONTRACT_RATIO"]
for var in omits:
del agg_prev["prev_" + str(var) + "_std"]
del agg_prev["prev_" + str(var) + "_min"]
del agg_prev["prev_" + str(var) + "_max"]
# count missings
nas = count_missings(agg_prev)
nas.head()
# check data
agg_prev.head()
# clear memory
del prev
```
# 5. DATA EXPORT
```
# merge data
print(appl.shape)
appl = appl.merge(right = agg_buro.reset_index(), how = "left", on = "SK_ID_CURR")
print(appl.shape)
del agg_buro
appl = appl.merge(right = agg_prev.reset_index(), how = "left", on = "SK_ID_CURR")
print(appl.shape)
del agg_prev
appl = appl.merge(right = agg_inst.reset_index(), how = "left", on = "SK_ID_CURR")
print(appl.shape)
del agg_inst
appl = appl.merge(right = agg_poca.reset_index(), how = "left", on = "SK_ID_CURR")
print(appl.shape)
del agg_poca
appl = appl.merge(right = agg_card.reset_index(), how = "left", on = "SK_ID_CURR")
print(appl.shape)
del agg_card
##### CROSS-TABLE FEATURE ENGINEERING
# credit ratios
appl["mix_AMT_PREV_ANNUITY_RATIO"] = appl["app_AMT_ANNUITY"] / appl["prev_AMT_ANNUITY_mean"]
appl["mix_AMT_PREV_CREDIT_RATIO"] = appl["app_AMT_CREDIT"] / appl["prev_AMT_CREDIT_mean"]
appl["mix_AMT_PREV_GOODS_PRICE_RATIO"] = appl["app_AMT_GOODS_PRICE"] / appl["prev_AMT_GOODS_PRICE_mean"]
appl["mix_AMT_BURO_ANNUITY_RATIO"] = appl["app_AMT_ANNUITY"] / appl["buro_AMT_ANNUITY_mean"]
appl["mix_AMT_BURO_CREDIT_RATIO"] = appl["app_AMT_CREDIT"] / appl["buro_AMT_CREDIT_SUM_mean"]
# dummy encodnig for factors
appl = pd.get_dummies(appl, drop_first = True)
# label encoder for factors
#data_factors = [f for f in appl.columns if appl[f].dtype == "object"]
#for var in data_factors:
# appl[var], _ = pd.factorize(appl[var])
# count missings
nas = count_missings(appl)
nas.head()
# partitioning
train = appl[appl["SK_ID_CURR"].isin(y["SK_ID_CURR"]) == True]
test = appl[appl["SK_ID_CURR"].isin(y["SK_ID_CURR"]) == False]
del appl
# check dimensions
print(train.shape)
print(test.shape)
# export CSV
train.to_csv("../data/prepared/train_full_cor.csv", index = False, float_format = "%.8f")
test.to_csv("../data/prepared/test_full_cor.csv", index = False, float_format = "%.8f")
y.to_csv("../data/prepared/y_full_cor.csv", index = False, float_format = "%.8f")
```
| github_jupyter |
Run the LSTM training network for four different optimizers, five different dropouts and four layer sizes using powershell to redirect all output to a file
```
!"python lstm.py | tee out_lstm.txt"
```
Parse the output file and create a csv
```
final_result = []
content = []
#format = [loss,optimizer,lstm_len,dropoff,train_accuracy,validation_accuracy,test_accuracy]
with open('out_lstm.txt', 'rb') as file:
content = file.readlines()
content = [str(x.decode(encoding='utf8')).strip() for x in content]
current_result = ['loss','optimizer','lstm_len','dropout','train_accuracy','validation_accuracy','test_accuracy']
next_look = False
for line in content:
if 'Training new model' in line:
final_result.append(current_result)
current_result = []
t = line.split(',')
l = t[1].split(':')[1]
o = t[2].split('=')[1]
llen = t[3].split('=')[1]
d = t[4].split('=')[1]
current_result.extend([l,o,llen,d])
if next_look:
t = line.split('-')
t_acc = t[3].split(':')[1][2:-1]
v_acc = t[5].split(':')[1][2:]
current_result.extend([t_acc,v_acc])
next_look = False
if '20/20' in line:
next_look = True
if 'test score' in line:
t = line.split(':')
acc = t[1].split(',')[1][1:-2]
current_result.append(acc)
final_result.append(current_result)
with open('lstm.csv', 'wb') as file:
for res in final_result:
line = ''
for it in res:
line = line + it +","
line = line[:-1]
line = line +'\n'
file.write(line.encode('utf-8'))
!"python cnn.py | tee out_cnn.txt"
final_result = []
content = []
#format = [loss,optimizer,lstm_len,dropoff,train_accuracy,validation_accuracy,test_accuracy]
with open('out_bilstm.txt', 'rb') as file:
content = file.readlines()
content = [str(x.decode(encoding='utf8')).strip() for x in content]
current_result = ['loss','optimizer','lstm_len','dropout','train_accuracy','validation_accuracy','test_accuracy']
next_look = False
for line in content:
if 'Training new model' in line:
final_result.append(current_result)
current_result = []
t = line.split(',')
l = t[1].split(':')[1]
o = t[2].split('=')[1]
llen = t[3].split('=')[1]
d = t[4].split('=')[1]
current_result.extend([l,o,llen,d])
if next_look:
t = line.split('-')
t_acc = t[3].split(':')[1][2:-1]
v_acc = t[5].split(':')[1][2:]
current_result.extend([t_acc,v_acc])
next_look = False
if '20/20' in line:
next_look = True
if 'test score' in line:
t = line.split(':')
acc = t[1].split(',')[1][1:-2]
current_result.append(acc)
final_result.append(current_result)
with open('bilstm.csv', 'wb') as file:
for res in final_result:
line = ''
for it in res:
line = line + it +","
line = line[:-1]
line = line +'\n'
file.write(line.encode('utf-8'))
```
Repeat process for CNN
```
!"python cnn.py | tee out_cnn.txt"
final_result = []
content = []
#format = [loss,optimizer,lstm_len,dropoff,train_accuracy,validation_accuracy,test_accuracy]
with open('out_cnn.txt', 'rb') as file:
content = file.readlines()
content = [str(x.decode(encoding='utf8')).strip() for x in content]
current_result = ['loss','optimizer','cnn_len','dropout','train_accuracy','validation_accuracy','test_accuracy']
next_look = False
for line in content:
if 'Training new model' in line:
final_result.append(current_result)
current_result = []
t = line.split(',')
l = t[1].split(':')[1]
o = t[2].split('=')[1]
llen = t[3].split('=')[1]
d = t[4].split('=')[1]
current_result.extend([l,o,llen,d])
if next_look:
t = line.split('-')
t_acc = t[3].split(':')[1][2:-1]
v_acc = t[5].split(':')[1][2:]
current_result.extend([t_acc,v_acc])
next_look = False
if '20/20' in line:
next_look = True
if 'test score' in line:
t = line.split(':')
acc = t[1].split(',')[1][1:-2]
current_result.append(acc)
final_result.append(current_result)
with open('cnn.csv', 'wb') as file:
for res in final_result:
line = ''
for it in res:
line = line + it +","
line = line[:-1]
line = line +'\n'
file.write(line.encode('utf-8'))
final_result
```
| github_jupyter |
# API Server
```
from IPython.display import Javascript
display(Javascript('''google.colab.output.setIframeHeight(0, true, {maxHeight: 200})'''))
!pip install ninja opensimplex gradio moviepy==1.0.3 tts pyrebase
!apt install ffmpeg
!pip install git+https://github.com/1adrianb/face-alignment@v1.0.1
!pip install fbpca boto3 requests==2.23.0 #urllib3==1.25.11
# !git submodule update --init --recursive
!python -c "import nltk; nltk.download('wordnet')"
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/MyDrive/Colab/fyp
try: # set up path
import sys
sys.path.append('/content/drive/MyDrive/Colab/fyp/ganspace')
sys.path.append('/content/drive/MyDrive/Colab/fyp/first-order-model')
sys.path.append('/content/drive/MyDrive/Colab/fyp/iPERCore')
sys.path.append('/content/drive/MyDrive/Colab/fyp/Wav2Lip')
print('Paths added')
except Exception as e:
print(e)
pass
#@title Install impersonator dependencies
# set CUDA_HOME, here we use CUDA 10.1
from IPython.display import Javascript
display(Javascript('''google.colab.output.setIframeHeight(0, true, {maxHeight: 200})'''))
import os
os.environ["CUDA_HOME"] = "/usr/local/cuda-10.1"
!echo $CUDA_HOME
%cd /content/drive/MyDrive/Colab/fyp/iPERCore/
!python setup.py develop
%cd /content/
#@title Define Cropping Functions
import os
from moviepy.video.io.ffmpeg_tools import ffmpeg_extract_subclip
import face_alignment
import imageio
import numpy as np
from skimage.transform import resize
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from IPython.display import HTML, clear_output
import cv2
import shutil
import warnings
warnings.filterwarnings("ignore")
hog = cv2.HOGDescriptor()
hog.setSVMDetector(cv2.HOGDescriptor_getDefaultPeopleDetector())
fa = face_alignment.FaceAlignment(face_alignment.LandmarksType._2D, flip_input=True,
device='cuda')
image_size = 512
def create_bounding_box(target_landmarks, expansion_factor=1):
target_landmarks = np.array(target_landmarks)
x_y_min = target_landmarks.reshape(-1, 68, 2).min(axis=1)
x_y_max = target_landmarks.reshape(-1, 68, 2).max(axis=1)
expansion_factor = (expansion_factor-1)/2
bb_expansion_x = (x_y_max[:, 0] - x_y_min[:, 0]) * expansion_factor
bb_expansion_y = (x_y_max[:, 1] - x_y_min[:, 1]) * expansion_factor
x_y_min[:, 0] -= bb_expansion_x
x_y_max[:, 0] += bb_expansion_x
x_y_min[:, 1] -= bb_expansion_y
x_y_max[:, 1] += bb_expansion_y
return np.hstack((x_y_min, x_y_max-x_y_min))
def fix_dims(im):
if im.ndim == 2:
im = np.tile(im[..., None], [1, 1, 3])
return im[...,:3]
def get_crop(im, center_face=True, crop_face=True, expansion_factor=1, landmarks=None):
im = fix_dims(im)
if (center_face or crop_face) and not landmarks:
landmarks = fa.get_landmarks_from_image(im)
if (center_face or crop_face) and landmarks:
rects = create_bounding_box(landmarks, expansion_factor=expansion_factor)
x0,y0,w,h = sorted(rects, key=lambda x: x[2]*x[3])[-1]
if crop_face:
s = max(h, w)
x0 += (w-s)//2
x1 = x0 + s
y0 += (h-s)//2
y1 = y0 + s
else:
img_h,img_w = im.shape[:2]
img_s = min(img_h,img_w)
x0 = min(max(0, x0+(w-img_s)//2), img_w-img_s)
x1 = x0 + img_s
y0 = min(max(0, y0+(h-img_s)//2), img_h-img_s)
y1 = y0 + img_s
else:
h,w = im.shape[:2]
s = min(h,w)
x0 = (w-s)//2
x1 = x0 + s
y0 = (h-s)//2
y1 = y0 + s
return int(x0),int(x1),int(y0),int(y1)
def pad_crop_resize(im, x0=None, x1=None, y0=None, y1=None, new_h=256, new_w=256):
im = fix_dims(im)
h,w = im.shape[:2]
if x0 is None:
x0 = 0
if x1 is None:
x1 = w
if y0 is None:
y0 = 0
if y1 is None:
y1 = h
if x0<0 or x1>w or y0<0 or y1>h:
im = np.pad(im, pad_width=[(max(-y0,0),max(y1-h,0)),(max(-x0,0),max(x1-w,0)),(0,0)], mode='edge')
return resize(im[max(y0,0):y1-min(y0,0),max(x0,0):x1-min(x0,0)], (new_h, new_w))
def get_crop_body(im, center_body=True, crop_body=True, expansion_factor=1, rects=None):
im = fix_dims(im)
if (center_body or crop_body) and rects is None:
rects, _ = hog.detectMultiScale(im, winStride=(4, 4),padding=(8,8), scale=expansion_factor)
if (center_body or crop_body) and rects is not None and len(rects):
x0,y0,w,h = sorted(rects, key=lambda x: x[2]*x[3])[-1]
if crop_body:
x0 += w//2-h//2
x1 = x0+h
y1 = y0+h
else:
img_h,img_w = im.shape[:2]
x0 += (w-img_h)//2
x1 = x0+img_h
y0 = 0
y1 = img_h
else:
h,w = im.shape[:2]
x0 = (w-h)//2
x1 = (w+h)//2
y0 = 0
y1 = h
return int(x0),int(x1),int(y0),int(y1)
def crop_resize(im, size, crop=False):
if im.shape[:2] == size:
return im
if size[0]<im.shape[0] or size[1]<im.shape[1]:
interp = cv2.INTER_AREA
else:
interp = cv2.INTER_CUBIC
if not crop:
return np.clip(cv2.resize(im, size[::-1], interpolation=interp),0,1)
ratio = max(size[0]/im.shape[0], size[1]/im.shape[1])
im = np.clip(cv2.resize(im, (int(np.ceil(im.shape[1]*ratio)), int(np.ceil(im.shape[0]*ratio))), interpolation=interp),0,1)
return im[(im.shape[0]-size[0])//2:(im.shape[0]-size[0])//2+size[0], (im.shape[1]-size[1])//2:(im.shape[1]-size[1])//2+size[1]]
#@title Define GANSpace functions
from ipywidgets import fixed
# Taken from https://github.com/alexanderkuk/log-progress
def log_progress(sequence, every=1, size=None, name='Items'):
from ipywidgets import IntProgress, HTML, VBox
from IPython.display import display
is_iterator = False
if size is None:
try:
size = len(sequence)
except TypeError:
is_iterator = True
if size is not None:
if every is None:
if size <= 200:
every = 1
else:
every = int(size / 200) # every 0.5%
else:
assert every is not None, 'sequence is iterator, set every'
if is_iterator:
progress = IntProgress(min=0, max=1, value=1)
progress.bar_style = 'info'
else:
progress = IntProgress(min=0, max=size, value=0)
label = HTML()
box = VBox(children=[label, progress])
display(box)
index = 0
try:
for index, record in enumerate(sequence, 1):
if index == 1 or index % every == 0:
if is_iterator:
label.value = '{name}: {index} / ?'.format(
name=name,
index=index
)
else:
progress.value = index
label.value = u'{name}: {index} / {size}'.format(
name=name,
index=index,
size=size
)
yield record
except:
progress.bar_style = 'danger'
raise
else:
progress.bar_style = 'success'
progress.value = index
label.value = "{name}: {index}".format(
name=name,
index=str(index or '?')
)
def name_direction(sender):
if not text.value:
print('Please name the direction before saving')
return
if num in named_directions.values():
target_key = list(named_directions.keys())[list(named_directions.values()).index(num)]
print(f'Direction already named: {target_key}')
print(f'Overwriting... ')
del(named_directions[target_key])
named_directions[text.value] = [num, start_layer.value, end_layer.value]
save_direction(random_dir, text.value)
for item in named_directions:
print(item, named_directions[item])
def save_direction(direction, filename):
filename += ".npy"
np.save(filename, direction, allow_pickle=True, fix_imports=True)
print(f'Latent direction saved as {filename}')
def project_image(target_image, step, model, center=False, seed=303, use_clip=True, video=False):
if model == 'portrait':
pkl = '/content/drive/MyDrive/Colab/models/portrait-001000.pkl'
elif model == 'character':
pkl = '/content/drive/MyDrive/Colab/models/character-002600.pkl'
elif model == 'model':
pkl = '/content/drive/MyDrive/Colab/models/modelv4-001600.pkl'
elif model == 'lookbook':
pkl = '/content/drive/MyDrive/Colab/models/lookbook-001800.pkl'
else:
print('Model PKL file does not exists')
!python /content/drive/MyDrive/Colab/fyp/stylegan2-ada-pytorch/pbaylies_projector.py --network={pkl} --outdir=/content/projector_output/ --target-image={target_image} --num-steps={step} --use-clip={use_clip} --use-center={center} --seed={seed} --save-video={video}
projected_w = np.load('/content/projector_output/projected_w.npz')['w'].squeeze()
return projected_w
def mix_w(w1, w2, content, style):
for i in range(0,5):
w2[i] = w1[i] * (1 - content) + w2[i] * content
for i in range(5, 16):
w2[i] = w1[i] * (1 - style) + w2[i] * style
return w2
def display_sample_pytorch(seed=None, truncation=0.5, directions=None, distances=None, scale=1, start=0, end=14, w=None, disp=True, save=None):
# blockPrint()
model.truncation = truncation
if w is None:
w = model.sample_latent(1, seed=seed).detach().cpu().numpy()
w = [w]*model.get_max_latents() # one per layer
else:
w = [np.expand_dims(x, 0) for x in w]
if directions != None and distances != None:
for l in range(start, end):
for i in range(len(directions)):
w[l] = w[l] + directions[i] * distances[i] * scale
torch.cuda.empty_cache()
#save image and display
out = model.sample_np(w)
final_im = Image.fromarray((out * 255).astype(np.uint8)).resize((500,500),Image.LANCZOS)
if save is not None:
if disp == False:
print(save)
final_im.save(f'out/{seed}_{save:05}.png')
if disp:
display(final_im)
return final_im
def generate_mov(seed, truncation, direction_vec, scale, layers, n_frames, out_name = 'out', noise_spec = None, loop=True):
"""Generates a mov moving back and forth along the chosen direction vector"""
# Example of reading a generated set of images, and storing as MP4.
%mkdir out
movieName = f'out/{out_name}.mp4'
offset = -10
step = 20 / n_frames
imgs = []
for i in log_progress(range(n_frames), name = "Generating frames"):
print(f'\r{i} / {n_frames}', end='')
w = model.sample_latent(1, seed=seed).cpu().numpy()
model.truncation = truncation
w = [w]*model.get_max_latents() # one per layer
for l in layers:
if l <= model.get_max_latents():
w[l] = w[l] + direction_vec * offset * scale
#save image and display
out = model.sample_np(w)
final_im = Image.fromarray((out * 255).astype(np.uint8))
imgs.append(out)
#increase offset
offset += step
if loop:
imgs += imgs[::-1]
with imageio.get_writer(movieName, mode='I') as writer:
for image in log_progress(list(imgs), name = "Creating animation"):
writer.append_data(img_as_ubyte(image))
#@title Load Model for GANSpace
selected_model = 'portrait'#@param ["portrait", "character", "model", "lookbook"]
# Load model
from IPython.utils import io
import torch
import PIL
import numpy as np
import ipywidgets as widgets
from PIL import Image
import imageio
from models import get_instrumented_model
from decomposition import get_or_compute
from config import Config
from skimage import img_as_ubyte
# Speed up computation
torch.autograd.set_grad_enabled(False)
torch.backends.cudnn.benchmark = True
# Specify model to use
config = Config(
model='StyleGAN2',
layer='style',
output_class=selected_model,
components=80,
use_w=True,
batch_size=5_000, # style layer quite small
)
inst = get_instrumented_model(config.model, config.output_class,
config.layer, torch.device('cuda'), use_w=config.use_w)
path_to_components = get_or_compute(config, inst)
model = inst.model
comps = np.load(path_to_components)
lst = comps.files
latent_dirs = []
latent_stdevs = []
load_activations = False
for item in lst:
if load_activations:
if item == 'act_comp':
for i in range(comps[item].shape[0]):
latent_dirs.append(comps[item][i])
if item == 'act_stdev':
for i in range(comps[item].shape[0]):
latent_stdevs.append(comps[item][i])
else:
if item == 'lat_comp':
for i in range(comps[item].shape[0]):
latent_dirs.append(comps[item][i])
if item == 'lat_stdev':
for i in range(comps[item].shape[0]):
latent_stdevs.append(comps[item][i])
def load_model(output_class):
global config
global inst
global model
config = Config(
model='StyleGAN2',
layer='style',
output_class=output_class,
components=80,
use_w=True,
batch_size=5_000, # style layer quite small
)
inst = get_instrumented_model(config.model, config.output_class,
config.layer, torch.device('cuda'), use_w=config.use_w)
path_to_components = get_or_compute(config, inst)
model = inst.model
comps = np.load(path_to_components)
lst = comps.files
latent_dirs = []
latent_stdevs = []
load_activations = False
for item in lst:
if load_activations:
if item == 'act_comp':
for i in range(comps[item].shape[0]):
latent_dirs.append(comps[item][i])
if item == 'act_stdev':
for i in range(comps[item].shape[0]):
latent_stdevs.append(comps[item][i])
else:
if item == 'lat_comp':
for i in range(comps[item].shape[0]):
latent_dirs.append(comps[item][i])
if item == 'lat_stdev':
for i in range(comps[item].shape[0]):
latent_stdevs.append(comps[item][i])
```
### GANSpace UI
```
#@title Portrait GANSpace UI
import gradio as gr
import numpy as np
def generate_image(seed, truncation,
female, realism, greyhair, shorthair, shortchin, ponytail, blackhair,
start_layer, end_layer):
scale = 1
seed, start_layer, end_layer = int(seed), int(start_layer), int(end_layer)
w = None
params = {'female': female,
'realism': realism,
'greyhair': greyhair,
'shorthair': shorthair,
'shortchin': shortchin,
'ponytail': ponytail,
'blackhair': blackhair}
param_indexes = {'female': 1,
'realism': 4,
'greyhair': 5,
'shorthair': 6,
'shortchin': 8,
'ponytail': 9,
'blackhair': 10}
directions = []
distances = []
for k, v in params.items():
directions.append(latent_dirs[param_indexes[k]])
distances.append(v)
model.truncation = truncation
if w is None:
w = model.sample_latent(1, seed=seed).detach().cpu().numpy()
w = [w]*model.get_max_latents() # one per layer
else:
w = [np.expand_dims(x, 0) for x in w]
if directions != None and distances != None:
for l in range(start_layer, end_layer):
for i in range(len(directions)):
w[l] = w[l] + directions[i] * distances[i] * 1
torch.cuda.empty_cache()
#save image and display
out = model.sample_np(w)
final_im = Image.fromarray((out * 255).astype(np.uint8))
# final_im.save('/tmp/edit_output.jpg') # save the content to temp
return final_im
#return display_sample_pytorch(int(seed), truncation, directions, distances, scale, int(start_layer), int(end_layer), disp=False)
truncation = gr.inputs.Slider(minimum=0, maximum=1, default=0.5, label="Truncation")
start_layer = gr.inputs.Number(default=0, label="Start Layer")
end_layer = gr.inputs.Number(default=14, label="End Layer")
seed = gr.inputs.Number(default=0, label="Seed")
slider_max_val = 20
slider_min_val = -20
slider_step = 1
female = gr.inputs.Slider(label="Female", minimum=slider_min_val, maximum=slider_max_val, default=0)
realism = gr.inputs.Slider(label="Realism", minimum=slider_min_val, maximum=slider_max_val, default=0)
greyhair = gr.inputs.Slider(label="Grey Hair", minimum=slider_min_val, maximum=slider_max_val, default=0)
shorthair = gr.inputs.Slider(label="Short Hair", minimum=slider_min_val, maximum=slider_max_val, default=0)
shortchin = gr.inputs.Slider(label="Short Chin", minimum=slider_min_val, maximum=slider_max_val, default=0)
ponytail = gr.inputs.Slider(label="Ponytail", minimum=slider_min_val, maximum=slider_max_val, default=0)
blackhair = gr.inputs.Slider(label="Black Hair", minimum=slider_min_val, maximum=slider_max_val, default=0)
scale = 1
inputs = [seed, truncation, female, realism, greyhair, shorthair, shortchin, ponytail, blackhair, start_layer, end_layer]
gr.Interface(generate_image, inputs, "image", live=True, title="GAN Editing").launch(debug=True)
import hashlib
seed = 'asds'
#@title Character Gradio UI
import gradio as gr
import numpy as np
import hashlib
def generate_image(seed, truncation,
monster, female, skimpy, light, bodysuit, bulky, human_head,
start_layer, end_layer):
scale = 1
seed = int(hashlib.sha256(seed.encode('utf-8')).hexdigest(), 16) % 10**8
params = {'monster': monster,
'female': female,
'skimpy': skimpy,
'light': light,
'bodysuit': bodysuit,
'bulky': bulky,
'human_head': human_head}
param_indexes = {'monster': 0,
'female': 1,
'skimpy': 2,
'light': 4,
'bodysuit': 5,
'bulky': 6,
'human_head': 8}
directions = []
distances = []
for k, v in params.items():
directions.append(latent_dirs[param_indexes[k]])
distances.append(v)
style = {'description_width': 'initial'}
return display_sample_pytorch(int(seed), truncation, directions, distances, scale, int(start_layer), int(end_layer), disp=False)
truncation = gr.inputs.Slider(minimum=0, maximum=1, default=0.5, label="Truncation")
start_layer = gr.inputs.Number(default=0, label="Start Layer")
end_layer = gr.inputs.Number(default=14, label="End Layer")
seed = gr.inputs.Textbox(default='0')
slider_max_val = 20
slider_min_val = -20
slider_step = 1
monster = gr.inputs.Slider(label="Monster", minimum=slider_min_val, maximum=slider_max_val, default=0)
female = gr.inputs.Slider(label="Female", minimum=slider_min_val, maximum=slider_max_val, default=0)
skimpy = gr.inputs.Slider(label="Skimpy", minimum=slider_min_val, maximum=slider_max_val, default=0)
light = gr.inputs.Slider(label="Light", minimum=slider_min_val, maximum=slider_max_val, default=0)
bodysuit = gr.inputs.Slider(label="Bodysuit", minimum=slider_min_val, maximum=slider_max_val, default=0)
bulky = gr.inputs.Slider(label="Bulky", minimum=slider_min_val, maximum=slider_max_val, default=0)
human_head = gr.inputs.Slider(label="Human Head", minimum=slider_min_val, maximum=slider_max_val, default=0)
scale = 1
inputs = [seed, truncation, monster, female, skimpy, light, bodysuit, bulky, human_head, start_layer, end_layer]
gr.Interface(generate_image, inputs, "image", live=True, title="CharacterGAN").launch(debug=True)
```
### FOMM
```
#@title Load Checkpoint for FOMM
from demo import load_checkpoints
import moviepy.editor as mpe
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from IPython.display import HTML
from demo import make_animation
from skimage import img_as_ubyte
#!gdown --id 1jmcn19-c3p8mf39aYNXUhdMqzqDYZhQ_ -O vox-cpk.pth.tar
generator, kp_detector = load_checkpoints(config_path='/content/drive/MyDrive/Colab/fyp/first-order-model/config/vox-256.yaml',
checkpoint_path='/content/drive/MyDrive/Colab/fyp/first-order-model/vox-cpk.pth.tar')
#@title FOMM UI
import gradio as gr
import moviepy.editor as mpe
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from IPython.display import HTML
from demo import make_animation
from skimage import img_as_ubyte
def predict_video(image_path, video_path, relative, show_inputs,
start_time=0, end_time=-1,
center_video_to_head=True, crop_video_to_head=True, video_crop_expansion_factor=1.5,
center_image_to_head=True, crop_image_to_head=True, image_crop_expansion_factor=2.5,
speech="", speaker='p225'):
video_crop_expansion_factor = max(video_crop_expansion_factor, 1)
image_crop_expansion_factor = max(image_crop_expansion_factor, 1)
if end_time > start_time:
# cut video
print('Cutting Video...')
ffmpeg_extract_subclip(video_path, start_time, end_time, targetname='cut.mp4')
video_path = 'cut.mp4'
source_image = imageio.imread(image_path.name)
reader = imageio.get_reader(video_path)
source_image = pad_crop_resize(source_image, *get_crop(source_image, center_face=center_image_to_head, crop_face=crop_image_to_head, expansion_factor=image_crop_expansion_factor))
fps = reader.get_meta_data()['fps']
print('FPS', fps)
#fps = 8
driving_video = []
landmarks = None
try:
for i,im in enumerate(reader):
if not crop_video_to_head:
break
landmarks = fa.get_landmarks_from_image(im)
if landmarks:
break
x0,x1,y0,y1 = get_crop(im, center_face=center_video_to_head, crop_face=crop_video_to_head, expansion_factor=video_crop_expansion_factor, landmarks=landmarks)
reader.set_image_index(0)
for im in reader:
driving_video.append(pad_crop_resize(im,x0,x1,y0,y1))
except RuntimeError:
pass
# Generate animation
predictions = make_animation(source_image, driving_video, generator, kp_detector, relative=relative)
fig = plt.figure(figsize=(10 * show_inputs + 8 * (predictions is not None), 8))
ims = []
for i in range(len(driving_video)):
cols = []
if show_inputs and speech == "":
cols.append(source_image)
cols.append(driving_video[i])
if predictions is not None:
cols.append(predictions[i])
im = plt.imshow(np.concatenate(cols, axis=1), animated=True)
plt.axis('off')
ims.append([im])
plt.tight_layout()
ani = animation.ArtistAnimation(fig, ims, interval=50, repeat_delay=1000)
ani.save('output.mp4', fps=fps)
plt.close()
if speech != "":
!tts --text "{speech}" --out_path speech.wav --model_name "tts_models/en/vctk/sc-glow-tts" --speaker_idx {speaker}
!cd Wav2Lip && python inference.py --checkpoint_path checkpoints/wav2lip_gan.pth --face "../output.mp4" --audio '../speech.wav'
return 'Wav2Lip/results/result_voice.mp4'
else:
video_clip = mpe.VideoFileClip('output.mp4')
audio_clip = mpe.AudioFileClip(video_path)
final_clip = video_clip.set_audio(audio_clip)
final_clip.write_videofile("result.mp4", fps=fps)
return 'result.mp4'
image_input = gr.inputs.Image(type="file")
video_input = gr.inputs.Video(type="mp4")
relative = gr.inputs.Checkbox(default=True, label="Relative")
show_inputs = gr.inputs.Checkbox(default=True, label="Show Inputs")
start_time = gr.inputs.Number(default=0, label="Start Time")
end_time = gr.inputs.Number(default=-1, label="End Time")
center_video_to_head = gr.inputs.Checkbox(default=True, label="Center Video to Head")
crop_video_to_head = gr.inputs.Checkbox(default=True, label="Crop Video to Head")
video_crop_expansion_factor = gr.inputs.Number(default=2, label="Video Crop Expansion Factor")
center_image_to_head = gr.inputs.Checkbox(default=False, label="Center Image to Head")
crop_image_to_head = gr.inputs.Checkbox(default=False, label="Crop Image to Head")
image_crop_expansion_factor = gr.inputs.Number(default=2, label="Image Crop Expansion Factor")
speech = gr.inputs.Textbox(label="Text to Speech", default="")
speaker = gr.inputs.Dropdown(default='p225', choices=['p225', 'p226', 'p227', 'p228', 'p229', 'p230', 'p231', 'p232', 'p233', 'p234', 'p261'])
inputs = [image_input, video_input, relative, show_inputs, start_time, end_time,
center_video_to_head, crop_video_to_head, video_crop_expansion_factor,
center_image_to_head, crop_image_to_head, image_crop_expansion_factor,
speech, speaker]
gr.Interface(predict_video, inputs, "video", live=False, title="Facial Animation").launch(debug=True)
```
### Impersonator++
```
#@title Impersonator++ Gradio UI
import gradio as gr
import os.path as osp
import platform
import argparse
import time
import sys
import subprocess
from IPython.display import HTML
from base64 import b64encode
from moviepy.editor import ImageSequenceClip
def imitate(im, video, bg_image, background,
center_image_to_body, crop_image_to_body, image_crop_expansion_factor, keep_aspect_background, show_inputs, pose_fc, cam_fc):
source_image = imageio.imread(im.name)
source_image = pad_crop_resize(source_image, *get_crop_body(source_image, center_body=center_image_to_body, crop_body=crop_image_to_body, expansion_factor=image_crop_expansion_factor), new_h=800, new_w=800)
imageio.imwrite('/content/crop.png', (source_image*255).astype(np.uint8))
if bg_image != 'None':
bg_image = imageio.imread(bg_image.name)
bg_image = crop_resize(bg_image/255, source_image.shape[:2], crop=keep_aspect_background)
imageio.imwrite('/content/bg_crop.png', (bg_image*255).astype(np.uint8))
with imageio.get_reader(video, format='mp4') as reader:
fps = reader.get_meta_data()['fps']
gpu_ids = "0"
num_source = 1
assets_dir = "/content/drive/MyDrive/Colab/fyp/iPERCore/assets"
output_dir = "/content/drive/MyDrive/Colab/fyp/iPERCore/results"
shutil.rmtree(output_dir, ignore_errors=True)
# symlink from the actual assets directory to this current directory
work_asserts_dir = os.path.join("./assets")
if not os.path.exists(work_asserts_dir):
os.symlink(osp.abspath(assets_dir), osp.abspath(work_asserts_dir),
target_is_directory=(platform.system() == "Windows"))
cfg_path = osp.join(work_asserts_dir, "configs", "deploy.toml")
model_id = "mymodel"
src_path = "\"path?=/content/crop.png,name?=mymodel"
if background=='replace' and os.path.exists('/content/bg_crop.png'):
src_path += ',bg_path?=/content/bg_crop.png'
src_path += '"'
ref_path = "\"path?=%s," \
"name?=myoutput," \
"pose_fc?=%d,"\
"cam_fc?=%d,"\
"fps?=%f\""%(video,pose_fc,cam_fc,fps)
options = ''
if background=='inpaint':
options += ' --use_inpaintor'
!python -m iPERCore.services.run_imitator --gpu_ids $gpu_ids --num_source $num_source --image_size $image_size --output_dir $output_dir --model_id $model_id --cfg_path $cfg_path --src_path $src_path --ref_path $ref_path $options
if show_inputs:
return "/content/drive/MyDrive/Colab/fyp/iPERCore/results/primitives/mymodel/synthesis/imitations/mymodel-myoutput.mp4"
else:
result_dir = '/content/drive/MyDrive/Colab/fyp/iPERCore/results/primitives/mymodel/synthesis/imitations/mymodel-myoutput'
frames = [os.path.abspath(os.path.join(result_dir, p)) for p in os.listdir(result_dir) if p.endswith(('jpg', 'png'))]
frames.sort()
#fps = last_frame/10
clip = ImageSequenceClip(frames, fps = fps)
#import re
#output_file = re.compile('\.png$').sub('.mp4', args.output)
clip.write_videofile('impersonator_output.mp4')
return 'impersonator_output.mp4'
im = gr.inputs.Image(type="file", label="Source Image")
bg_image = gr.inputs.Image(type="file", label="Background Image")
background = gr.inputs.Radio(choices=['None', 'replace', 'inpaint'], label="Background")
center_image_to_body = gr.inputs.Checkbox(default=True, label="Center Image to Body")
crop_image_to_body = gr.inputs.Checkbox(default=False, label="Crop Image to Body")
image_crop_expansion_factor = gr.inputs.Number(default=1.05, label="Image Crop Expansion Factor")
keep_aspect_background = gr.inputs.Checkbox(default=True, label="Keep Background Aspect Ratio")
show_inputs = gr.inputs.Checkbox(default=False, label="Show Inputs")
pose_fc = gr.inputs.Number(default=300, label="Pose Smooth Factor")
cam_fc = gr.inputs.Number(default=100, label="Camera Smooth Factor")
inputs = [im, 'video', bg_image, background,
center_image_to_body, crop_image_to_body, image_crop_expansion_factor, keep_aspect_background, show_inputs, pose_fc, cam_fc]
iface = gr.Interface(imitate, inputs, "video", live=False, title="Full Body Animation")
iface.launch(debug=True)
```
## Server
```
from IPython.display import Javascript
display(Javascript('''google.colab.output.setIframeHeight(0, true, {maxHeight: 200})'''))
!pip install fastapi nest-asyncio pyngrok uvicorn aiofiles python-multipart firebase-admin
firebase_admin.delete_app(firebase_admin.get_app())
#@title API Functions
from fastapi import FastAPI
from fastapi.middleware.cors import CORSMiddleware
from fastapi.responses import FileResponse, StreamingResponse
from fastapi import FastAPI, File, UploadFile, Form
from pydantic import BaseModel
from typing import Optional, Dict, List
from io import StringIO, BytesIO
from pydantic import BaseModel
import shutil
import datetime
import firebase_admin
from firebase_admin import credentials, firestore, storage
# Use the application default credentials
if not firebase_admin._apps:
cred = credentials.Certificate("/content/drive/MyDrive/Colab/fyp/gancreate-firebase-adminsdk-yayqe-07b54a912d.json")
firebase_admin.initialize_app(cred, {
'storageBucket': 'gancreate.appspot.com'
})
db = firestore.client()
bucket = storage.bucket()
# Log the user in
user = "YyzhczpqEzMw6uWHuikt0Icg2Sp1"
app = FastAPI()
app.add_middleware(
CORSMiddleware,
allow_origins=['*'],
allow_credentials=True,
allow_methods=['*'],
allow_headers=['*'],
)
@app.get('/')
async def root():
return {'hello': 'world'}
def sample(seed: int = None, truncation: float = 0.5, w=None):
model.truncation = truncation
if w is None:
w = model.sample_latent(1, seed=seed).detach().cpu().numpy()
w = [w]*model.get_max_latents() # one per layer
#save image and display
out = model.sample_np(w)
return Image.fromarray((out * 255).astype(np.uint8))
@app.get('/generate')
async def generate(model_type: str = 'portrait', seed: int = None, truncation: float = 0.5, w=None, save=False):
if config.output_class != model_type:
load_model(model_type)
model.truncation = truncation
if w is None:
w = model.sample_latent(1, seed=seed).detach().cpu().numpy()
w = [w]*model.get_max_latents() # one per layer
#save image and display
out = model.sample_np(w)
final_im = Image.fromarray((out * 255).astype(np.uint8))
final_im.save('/tmp/output.jpg') # save the content to temp
return FileResponse('/tmp/output.jpg', media_type="image/jpeg")
class EditConfig(BaseModel):
model_type: Optional[str] = "portrait"
seed: Optional[int] = None
truncation: Optional[float] = 0.5
start_layer: Optional[int] = 0
end_layer: Optional[int] = 14
attributes: Dict[str, float]
latent_id: Optional[str] = ""
save: Optional[bool] = False
@app.post('/edit')
async def edit(edit_config: EditConfig):
model_type = edit_config.model_type
if config.output_class != model_type:
load_model(model_type)
seed = edit_config.seed
latent_id = edit_config.latent_id
model.truncation = edit_config.truncation
if latent_id is None or latent_id is "":
w = model.sample_latent(1, seed=seed).detach().cpu().numpy()
w = [w]*model.get_max_latents() # one per layer
else:
w = np.load(f"latents/{model_type}/{latent_id}.npy")
w = [np.expand_dims(x, 0) for x in w]
param_indexes = {
"portrait": {
'Gender': 1,
'Realism': 4,
'Gray Hair': 5,
'Hair Length': 6,
'Chin': 8,
'Ponytail': 9,
'Black Hair': 10
},
"model": {
'Gender': 4,
'Dress': 0,
'Sleveeless': 1,
'Short Skirt': 3,
'Jacket': 5,
'Darkness': 7,
'Slimness': 9
},
"character": {
'Monster': 0,
'Gender': 1,
'Skimpiness': 2,
'Light': 4,
'Bodysuit': 5,
'Bulkiness': 6,
'Human Head': 8
}
}
directions = []
distances = []
for k, v in edit_config.attributes.items():
directions.append(latent_dirs[param_indexes[model_type][k]])
distances.append(v)
if directions != None and distances != None:
for l in range(edit_config.start_layer, edit_config.end_layer):
for i in range(len(directions)):
w[l] = w[l] + directions[i] * distances[i] * 1
torch.cuda.empty_cache()
#save image and display
out = model.sample_np(w)
final_im = Image.fromarray((out * 255).astype(np.uint8))
final_im.save('/tmp/edit_output.jpg') # save the content to temp
if edit_config.save:
doc_ref = db.collection(f'{model_type}s').add({})
doc_id = doc_ref[1].id
blob = bucket.blob(f'{model_type}s/{doc_id}.jpg')
blob.upload_from_filename('/tmp/edit_output.jpg')
blob.make_public()
data = {
"url": blob.public_url,
"user": user,
"created_at": datetime.datetime.now()
}
db.collection(f'{model_type}s').document(doc_id).set(data)
np.save(f'latents/{model_type}/{doc_id}.npy', w)
return FileResponse('/tmp/edit_output.jpg', media_type="image/jpeg")
class AnimateFaceConfig(BaseModel):
center_video_to_head: Optional[bool] = True
crop_video_to_head: Optional[bool] = True
center_image_to_head: Optional[bool] = True
center_video_to_head: Optional[bool] = True
video_crop_expansion_factor: Optional[float] = 1.5
image_crop_expansion_factor: Optional[float] = 2.5
speech: Optional[str] = ""
speaker: Optional[str] = "p225"
show_inputs: Optional[bool] = False
@app.post("/animate/face")
async def animate_face(
files: List[UploadFile] = File(...),
center_video_to_head: bool = Form(True),
crop_video_to_head: bool = Form(True),
video_crop_expansion_factor: float = Form(1.5),
center_image_to_head: bool = Form(True),
crop_image_to_head: bool = Form(True),
image_crop_expansion_factor: float = Form(2.5),
relative: bool = Form(True),
speech: str = Form(""),
speaker: str = Form("p225"),
show_inputs: bool = Form(False),
):
video_crop_expansion_factor = max(video_crop_expansion_factor, 1)
image_crop_expansion_factor = max(image_crop_expansion_factor, 1)
source_image = files[0]
ref_video = files[1]
with open(source_image.filename, "wb") as buffer:
shutil.copyfileobj(source_image.file, buffer)
video_path = "ref_video.mp4"
with open(video_path, "wb") as buffer:
shutil.copyfileobj(ref_video.file, buffer)
source_image = imageio.imread(source_image.filename)
reader = imageio.get_reader(video_path)
source_image = pad_crop_resize(source_image, *get_crop(source_image, center_face=center_image_to_head, crop_face=crop_image_to_head, expansion_factor=image_crop_expansion_factor))
fps = reader.get_meta_data()['fps']
driving_video = []
landmarks = None
try:
for i,im in enumerate(reader):
if not crop_video_to_head:
break
landmarks = fa.get_landmarks_from_image(im)
if landmarks:
break
x0,x1,y0,y1 = get_crop(im, center_face=center_video_to_head, crop_face=crop_video_to_head, expansion_factor=video_crop_expansion_factor, landmarks=landmarks)
reader.set_image_index(0)
for im in reader:
driving_video.append(pad_crop_resize(im,x0,x1,y0,y1))
except RuntimeError:
pass
# Generate animation
predictions = make_animation(source_image, driving_video, generator, kp_detector, relative=relative)
fig = plt.figure(figsize=(12 * show_inputs + 6 * (predictions is not None), 8))
ims = []
for i in range(len(driving_video)):
cols = []
if show_inputs and speech == "":
cols.append(source_image)
cols.append(driving_video[i])
if predictions is not None:
cols.append(predictions[i])
im = plt.imshow(np.concatenate(cols, axis=1), animated=True)
plt.axis('off')
ims.append([im])
plt.tight_layout()
ani = animation.ArtistAnimation(fig, ims, interval=50, repeat_delay=1000)
ani.save('output.mp4', fps=fps)
plt.close()
if speech != "":
!tts --text "{speech}" --out_path speech.wav --model_name "tts_models/en/vctk/sc-glow-tts" --speaker_idx {speaker}
!cd Wav2Lip && python inference.py --checkpoint_path checkpoints/wav2lip_gan.pth --face "../output.mp4" --audio '../speech.wav'
return FileResponse('Wav2Lip/results/result_voice.mp4', media_type="video/mp4")
else:
video_clip = mpe.VideoFileClip('output.mp4')
audio_clip = mpe.AudioFileClip(video_path)
final_clip = video_clip.set_audio(audio_clip)
final_clip.write_videofile("result.mp4", fps=fps)
return FileResponse('result.mp4', media_type="video/mp4")
@app.post("/animate/body")
async def animate_body(
files: List[UploadFile] = File(...),
background: str = Form("None"),
center_image_to_body: bool = Form(True),
crop_image_to_body: bool = Form(False),
image_crop_expansion_factor: float = Form(1.05) ,
keep_aspect_background: bool = Form(True),
pose_fc: int = Form(300),
cam_fc: int = Form(100),
):
source_image = files[0]
ref_video = files[1]
bg_image = files[2]
with open(source_image.filename, "wb") as buffer:
shutil.copyfileobj(source_image.file, buffer)
video_path = "ref_video.mp4"
with open(video_path, "wb") as buffer:
shutil.copyfileobj(ref_video.file, buffer)
source_image = imageio.imread(source_image.filename)
source_image = pad_crop_resize(source_image, *get_crop_body(source_image, center_body=center_image_to_body, crop_body=crop_image_to_body, expansion_factor=image_crop_expansion_factor))
imageio.imwrite('/content/crop.png', (source_image*255).astype(np.uint8))
if bg_image != 'None':
with open(bg_image.filename, "wb") as buffer:
shutil.copyfileobj(bg_image.file, buffer)
bg_image = imageio.imread(bg_image.filename)
bg_image = crop_resize(bg_image/255, source_image.shape[:2], crop=keep_aspect_background)
imageio.imwrite('/content/bg_crop.png', (bg_image*255).astype(np.uint8))
with imageio.get_reader(video_path, format='mp4') as reader:
fps = reader.get_meta_data()['fps']
gpu_ids = "0"
num_source = 1
assets_dir = "/content/drive/MyDrive/Colab/fyp/iPERCore/assets"
output_dir = "/content/drive/MyDrive/Colab/fyp/iPERCore/results"
shutil.rmtree(output_dir, ignore_errors=True)
# symlink from the actual assets directory to this current directory
work_asserts_dir = os.path.join("./assets")
if not os.path.exists(work_asserts_dir):
os.symlink(osp.abspath(assets_dir), osp.abspath(work_asserts_dir),
target_is_directory=(platform.system() == "Windows"))
cfg_path = osp.join(work_asserts_dir, "configs", "deploy.toml")
model_id = "mymodel"
src_path = "\"path?=/content/crop.png,name?=mymodel"
if background=='replace' and os.path.exists('/content/bg_crop.png'):
src_path += ',bg_path?=/content/bg_crop.png'
src_path += '"'
ref_path = "\"path?=%s," \
"name?=myoutput," \
"pose_fc?=%d,"\
"cam_fc?=%d,"\
"fps?=%f\""%(video_path,pose_fc,cam_fc,fps)
options = ''
if background=='inpaint':
options += ' --use_inpaintor'
!python -m iPERCore.services.run_imitator --gpu_ids $gpu_ids --num_source $num_source --image_size $image_size --output_dir $output_dir --model_id $model_id --cfg_path $cfg_path --src_path $src_path --ref_path $ref_path $options
return FileResponse('./iPERCore/results/primitives/mymodel/synthesis/imitations/mymodel-myoutput.mp4', media_type="video/mp4")
import nest_asyncio
from pyngrok import ngrok
import uvicorn
ngrok_tunnel = ngrok.connect(8000)
print('Public URL:', ngrok_tunnel.public_url+'/docs')
nest_asyncio.apply()
uvicorn.run(app, port=8000)
```
| github_jupyter |
# Using SupCS for Training GLUE Tasks
## Dependencies
Install torch, tensorflow and SupCL-Seq packages using pip.
```
#!pip install datasets numpy
#!pip install -U scikit-learn
from datasets import load_dataset, load_metric
from transformers import TrainingArguments, Trainer, AutoTokenizer, AutoModel,AutoModelForSequenceClassification
#----for roberta-----#
from transformers import RobertaTokenizer, RobertaForSequenceClassification, RobertaModel
from sklearn.metrics import classification_report
import warnings
import numpy as np
from SupCL_Seq import SupCsTrainer
warnings.filterwarnings('ignore')
```
## GLUE Tasks
```
GLUE_TASKS = ["cola", "mnli", "mnli-mm", "mrpc", "qnli", "qqp", "rte", "sst2", "stsb", "wnli"]
task = "cola"
model_name = "nghuyong/ernie-2.0-en"#"roberta-base" #"bert-base-uncased"
actual_task = "mnli" if task == "mnli-mm" else task
dataset = load_dataset("glue", actual_task)
metric = load_metric('glue', actual_task)
tokenizer = AutoTokenizer.from_pretrained(model_name, use_fast=True)
model = AutoModel.from_pretrained(model_name)
task_to_keys = {
"cola": ("sentence", None),
"mnli": ("premise", "hypothesis"),
"mnli-mm": ("premise", "hypothesis"),
"mrpc": ("sentence1", "sentence2"),
"qnli": ("question", "sentence"),
"qqp": ("question1", "question2"),
"rte": ("sentence1", "sentence2"),
"sst2": ("sentence", None),
"stsb": ("sentence1", "sentence2"),
"wnli": ("sentence1", "sentence2"),
}
sentence1_key, sentence2_key = task_to_keys[task]
def preprocess_function(examples):
if sentence2_key is None:
return tokenizer(examples[sentence1_key], truncation=True)
return tokenizer(examples[sentence1_key], examples[sentence2_key], truncation=True)
encoded_dataset = dataset.map(preprocess_function, batched=True)
```
## Custom Metric
We employ a task dependent metric.
```
def compute_metrics(eval_pred):
predictions, labels = eval_pred
if task != "stsb":
predictions = np.argmax(predictions, axis=1)
else:
predictions = predictions[:, 0]
return metric.compute(predictions=predictions, references=labels)
validation_key = "validation_mismatched" if task == "mnli-mm" else "validation_matched" if task == "mnli" else "validation"
train_dataset = encoded_dataset["train"]
test_dataset = encoded_dataset[validation_key]
```
## Training Argument From Huggingface
```
CL_args = TrainingArguments(
output_dir = './results',
save_total_limit = 1,
num_train_epochs=5,
per_device_train_batch_size=12,
evaluation_strategy = 'no',
logging_steps = 200,
learning_rate = 5e-5,
warmup_steps=50,
weight_decay=0.01,
logging_dir='./logs',
)
```
## SupCL-Trainer
This works exactly similar to the trainer from huggingface. We first CS train and save the model.
```
SupCL_trainer = SupCsTrainer.SupCsTrainer(
w_drop_out=[0.1, 0.2],
temperature= 0.05,
def_drop_out=0.1,
pooling_strategy='mean',
model = model,
args = CL_args,
train_dataset=train_dataset,
eval_dataset=test_dataset,
tokenizer=tokenizer,
compute_metrics=compute_metrics
)
SupCL_trainer.train()
SupCL_trainer.save_model('./cs_baseline')
```
## Only FineTune a Linear Layer
After CS training we only add a linear layer and then finetune its weights only by freezing the pretrained model base parameters. Finally, finetune the linear layer on the data using cross entropy.
```
model_name = './cs_baseline'#"./results/checkpoint-500/"
num_labels = 3 if actual_task =='mnli' else 2
if actual_task =='stsb': num_labels = 1
#------ Add classification layer ---------#
#model = RobertaForSequenceClassification.from_pretrained(model_name,num_labels=num_labels)
model = AutoModelForSequenceClassification.from_pretrained(model_name,num_labels=num_labels)
# ---- Freeze the base model -------#
for param in model.base_model.parameters():
param.requires_grad = False
args = TrainingArguments(
output_dir = './results',
save_total_limit = 1,
num_train_epochs=5,
per_device_train_batch_size=28,
per_device_eval_batch_size=64,
evaluation_strategy = 'epoch',
logging_steps = 200,
learning_rate = 1e-04,
eval_steps = 200,
warmup_steps=50,
report_to ='tensorboard',
weight_decay=0.01,
logging_dir='./logs',
)
trainer = Trainer(
model,
args,
train_dataset=train_dataset,
eval_dataset=test_dataset,
tokenizer=tokenizer,
compute_metrics=compute_metrics
)
trainer.train()
```
| github_jupyter |
<img src="../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# Qiskit Aer: Device noise model simulations
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorials.
## Introduction
This notebook shows how to use the Qiskit Aer `noise.device` module to automatically generate a noise model for an IBMQ hardware device, and use this model to do noisy simulation of `QuantumCircuits` to study the effects of errors which occur on real devices.
Note that these automatic models are only a highly simplified approximation of the real errors that occur on actual devices. The study of quantum errors on real devices is an active area of research and we discuss the Qiskit Aer tools for configuring more detailed noise models in another notebook.
```
from qiskit import Aer, IBMQ, execute
from qiskit.providers.aer import noise
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit.tools.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
```
## Device noise module
The *Qiskit Aer* device noise module contains two functions to automatically generate a simplified noise model for a real device. This model is generated using the calibration information reported in the `BackendProperties` of a device.
### IBMQ Provider
We will use a real hardware device in the `IBMQ` provider as an example. First we must load our account credentials, and then select a backend from the provider.
```
# Load IBMQ account
IBMQ.load_accounts()
IBMQ.backends()
```
We will use the `ibmq_16_melbourne` device for this tutorial. We may get the properties of the backend using the `properties` method, the information in the returned `BackendProperties` object will be used to automatically generate a noise model for the device that can be used by the Qiskit Aer `QasmSimulator`. We will also want to get the `coupling_map` for the device from its `configuration` to use when compiling circuits for simulation to most closely mimic the gates that will be executed on a real device
```
device = IBMQ.get_backend('ibmq_16_melbourne')
properties = device.properties()
coupling_map = device.configuration().coupling_map
```
### Test circuit for device and simulation comparison
Now we construct a test circuit to compare the output of the real device with the noisy output simulated on the Qiskit Aer `QasmSimulator`. We will prepare a 3-qubit GHZ state $\frac{1}{2}(|0,0,0\rangle + |1,1,1\rangle)$ on qubits 0, 1 and 2. Before running with noise or on the device we show the ideal expected output with no noise.
```
# Construct quantum circuit
qr = QuantumRegister(3, 'qr')
cr = ClassicalRegister(3, 'cr')
circ = QuantumCircuit(qr, cr)
circ.h(qr[0])
circ.cx(qr[0], qr[1])
circ.cx(qr[1], qr[2])
circ.measure(qr, cr)
# Select the QasmSimulator from the Aer provider
simulator = Aer.get_backend('qasm_simulator')
# Execute and get counts
result = execute(circ, simulator).result()
counts = result.get_counts(circ)
plot_histogram(counts, title='Ideal counts for 3-qubit GHZ state')
```
## Generating a device noise model
Noise models in Qiskit Aer are presented using the `NoiseModel` object from the `aer.noise` module. The function we will use is the `basic_device_noise_model` function from `aer.noise.device` module which will return a basic approximate `NoiseModel` object configured from a device `BackendProperties`.
### Basic device noise model
The `basic_device_noise_model` constructs an approximate noise model consisting of:
* **Single-qubit gate errors** consisting of a single qubit depolarizing error followed by a single qubit thermal relaxation error.
* **Two-qubit gate errors** consisting of a two-qubit depolarizing error followed by single-qubit thermal relaxation errors on both qubits in the gate.
* **Single-qubit readout errors** on the classical bit value obtained from measurements on individual qubits
For the gate errors the error parameter of the thermal relaxation errors is derived using the `thermal_relaxation_error` function from `aer.noise.errors` module, along with the individual qubit $T_1$ and $T_2$ parameters, and the `gate_time` parameter from the device backend properties. The probability of the depolarizing error is then set so that the combined average gate infidelity from the depolarizing error followed by the thermal relaxaxtion is equal to the `gate_error` value from the backend properties.
For the readout errors the probability that the recorded classical bit value will be flipped from the true outcome after a measurement is given by the qubit `readout_errors`.
Let us construct the device noise model.
**Note:** *Since the devices don't currently provide the gate times for gates we will manually provide them for the gates we are interested in using the optional `gate_times` argument for `basic_device_noise_model`.*
```
# List of gate times for ibmq_14_melbourne device
# Note that the None parameter for u1, u2, u3 is because gate
# times are the same for all qubits
gate_times = [
('u1', None, 0), ('u2', None, 100), ('u3', None, 200),
('cx', [1, 0], 678), ('cx', [1, 2], 547), ('cx', [2, 3], 721),
('cx', [4, 3], 733), ('cx', [4, 10], 721), ('cx', [5, 4], 800),
('cx', [5, 6], 800), ('cx', [5, 9], 895), ('cx', [6, 8], 895),
('cx', [7, 8], 640), ('cx', [9, 8], 895), ('cx', [9, 10], 800),
('cx', [11, 10], 721), ('cx', [11, 3], 634), ('cx', [12, 2], 773),
('cx', [13, 1], 2286), ('cx', [13, 12], 1504), ('cx', [], 800)
]
# Construct the noise model from backend properties
# and custom gate times
noise_model = noise.device.basic_device_noise_model(properties, gate_times=gate_times)
print(noise_model)
```
## Simulating a quantum circuit with noise
To use this noise model we must make use of several keyword arguments in the `execute` function. These are:
* `noise_model`: This passes the noise model to the `QasmSimulator.run` method for noisy simulation.
* `basis_gates`: A noise model is defined with respect to specific gates, we must pass these basis gates to the Qiskit compiler so that it compiles the circuit to the correct gates for the noise model. The basis gates of a noise model may be obtained from the `NoiseModel.basis_gates` property.
* `coupling_map`: We also must make sure we provide the `coupling_map` for the real device so that the compiler will produce a Qobj for the simulator that will match the compiled experiment that can be executed on the real device.
```
# Get the basis gates for the noise model
basis_gates = noise_model.basis_gates
# Select the QasmSimulator from the Aer provider
simulator = Aer.get_backend('qasm_simulator')
# Execute noisy simulation and get counts
result_noise = execute(circ, simulator,
noise_model=noise_model,
coupling_map=coupling_map,
basis_gates=basis_gates).result()
counts_noise = result_noise.get_counts(circ)
plot_histogram(counts_noise, title="Counts for 3-qubit GHZ state with depolarizing noise model")
```
## Comparison to the real device
Now we will execute the circuit on the real device to see the effect of the actual noise processes on the output counts. Note that this execution may take some time to return the results.
```
# Submit job to real device and wait for results
job_device = execute(circ, device)
job_monitor(job_device)
# Get results from completed execution
result_device = job_device.result()
counts_device = result_device.get_counts(0)
plot_histogram(counts_device, title='Counts for 3-qubit GHZ state on IBMQ device: {}'.format(device.name()))
```
You might notice that the counts returned in the above histogram don't look exactly like our simulation. This is because the real errors that happen on a device can be very complicated, and characterizing them to create acurate models is an active area of quantum computing research. The *basic device noise model* we used for our simulation is a simplified error model that only takes into account *local* errors occuring on the qubits participating in each individual gate, and treats the erors as a relaxation process combined with an error operation which acts to randomize the state of the qubit with some probability *p* (the depolarizing probability). While this model is an approximation, due to its simplicity it is a useful starting point for studying the effects of noise on quantum computations.
| github_jupyter |
# Background #
This notebook goes through the off position reduction in class for the orion B map
```
from IPython.display import HTML
HTML('''
// http://lokeshdhakar.com/projects/lightbox2
<link href="./notebook_support/lightbox2-master/dist/css/lightbox.css" rel="stylesheet">
<script src="./notebook_support/lightbox2-master/dist/js/lightbox.js"></script>
<script>
function code_toggle() {
if (code_shown){
$('div.input').hide('500');
$('#toggleButton').val('Show Code')
} else {
$('div.input').show('500');
$('#toggleButton').val('Hide Code')
}
code_shown = !code_shown
}
$( document ).ready(function(){
code_shown=false;
$('div.input').hide()
});
</script>
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>''')
```
# import commands
```
## import
from astropy.io import fits
import numpy as np
import glob
import datetime
import pandas as pd
import logging
import os,re
import time
import subprocess
import cygrid
import matplotlib.pyplot as plt
from matplotlib.lines import lineStyles
from astropy.io import fits
from astropy import units as u
import jinja2
import requests
# notebook
from IPython.display import Image, display
from IPython.display import HTML
# astropy
from astropy.io import fits
from astropy import constants as const
from astropy import units as u
plt.rcParams['figure.figsize'] = (15.0, 7.0)
def PrintMessage(message):
html='''
<table style="width:100%">
<tr>
<td style='text-align:center;vertical-align:middle'>{0}</td>
</tr>
</table>
'''
display(HTML(html.format(message)))
def Change1dListTo2dList(array, column=2):
test = []
test_row = []
for item in array:
test_row.append(item)
if len(test_row)> column-1:
test.append(test_row)
test_row = []
if len(test_row)>0:
test.append(test_row)
return test
def DisplayImagesInColumns(images,image_columns=2, gallery_label=""):
popup_css='''
<style>
.thumb img {
border:1px solid #000;
margin:3px;
float:left;
}
.thumb span {
position:fixed;
visibility:hidden;
}
.thumb:hover, .thumb:hover span {
visibility:visible;
z-index:1;
}
</style>
'''
html='''
<a href={0} target="_blank">
<img style='width: {1:2.0f}%; margin: 0px; float: left; border: 0px solid black;' src='{0}?{2}' />
</a>
'''
html='''
<a href={0} target="_blank">
<img style='width: {1:2.0f}%; margin: 0px; float: left; border: 0px solid black;' src='{0}?{2}' />
</a>
'''
#html='''
#<a href={0} data-lightbox={3} data-title="{0}">
#<img style='width: {1:2.0f}%; margin: 0px; float: left; border: 0px solid black;' src='{0}?{2}' />
#</a>
#'''
list2d = Change1dListTo2dList(images, column=image_columns)
width_percent = (1/float(image_columns))*100.0
gallery_label = gallery_label.replace(" ","")
for row in list2d:
imagesList=popup_css.join( [html.format(image,width_percent-2,time.time(),gallery_label)
for image in row ])
display(HTML(imagesList))
return imagesList
# Helper Class to run class_cmd_strings and export the images that were created to the notebook
class RunClassScript():
def __init__(self, cmd, no_output=False, display_images=True, test=False):
self.cmd = cmd
self.cmd_file = "/tmp/cmd.class"
self.cmd_log = "/tmp/cmd.log"
self.images = [string.split()[1].replace("\"","").replace("//","/") for string in self.cmd.split("\n") if string[0:2] == "ha"]
if self.cmd.split("\n")[-1].strip() != "exit 0":
self.cmd = "{}\nexit 0\n".format(self.cmd)
subprocess.call(["rm", "-rf", "/tmp/cmd.class"])
fileout = open(self.cmd_file, "w")
fileout.write(self.cmd)
fileout.close()
#output = subprocess.check_output("class -nw @{0}".format(self.cmd_file), shell=True)
if test:
print(self.cmd)
try:
output = subprocess.check_call("class -nw @{0} > {1}".format(self.cmd_file,self.cmd_log), shell=True)
except subprocess.CalledProcessError as e:
print("======================================")
print("=========== ERROR in class cmd =======")
print("======================================")
print("\n")
print("======================================")
print("============= INPUT FILE =============")
print("======================================")
with open(self.cmd_file, 'r') as fin:
for line in fin.readlines():
if line[0]!=" ":
print(line.strip("\n"))
print("======================================")
print("========= OUTPUT LOGFILE =============")
print("======================================")
with open(self.cmd_log, 'r') as fin:
for line in fin.readlines():
if (len(set(line))>1):
print(line.strip("\n"))
raise subprocess.CalledProcessError(e.returncode,e.cmd, e.output)
if output!=0:
print(output)
self.output=output
#
if display_images:
for image in self.images:
if os.path.exists(image):
#display(image, Image(image),width=300,height=300)
display(image, Image(image))
def GenerateGIFAnimation(images,gif_filename):
convert_cmd = "convert -delay 50"\
" -loop 0 "\
"{0} "\
"{1}".format(images,gif_filename)
os.system(convert_cmd)
from IPython.display import HTML
def download_file(url,local_filename):
#local_filename = url.split('/')[-1]
r = requests.get(url)
f = open(local_filename, 'wb')
for chunk in r.iter_content(chunk_size=512 * 1024):
if chunk: # filter out keep-alive new chunks
f.write(chunk)
f.close()
return
def show_file_content(file, num_lines=10):
''' To show class console output '''
a_file = open(file, "r")
lines = a_file.readlines()
last_lines= lines[-num_lines:]
for line in last_lines:
print(line)
```
# Download test data from archive
```
data_folder = "./tmp/data"
if not os.path.exists(data_folder):
print("making folder {0}".format(data_folder))
#os.makedirs(data_folder)
os.makedirs(data_folder,exist_ok=True)
urls = []
#urls.append("https://irsa.ipac.caltech.edu/data/SOFIA/GREAT/OC4K/20161110_F348/proc/p8486/data/2016-11-10_GR_F348_04_0066_0010505_1900536.9.great.tar")
urls.append("https://irsa.ipac.caltech.edu/data/SOFIA/GREAT/OC5A/20170208_F371/proc/p4006/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9.great.tar")
for url in urls:
filename = "{0}/{1}".format(data_folder,url.split('/')[-1])
if not os.path.exists(filename):
print("downloading {0} to {1}".format(url, filename))
download_file(url, filename)
else:
print("{1} already downloaded".format(url, filename))
# untar
already_downloaded_great = glob.glob("tmp/data/**/*.great")
#
for url in urls:
# check for great file
tar_filename = "{0}/{1}".format(data_folder,url.split('/')[-1])
filename_great,ext = os.path.splitext(tar_filename)
full_filename_great,ext = os.path.splitext(os.path.basename(filename_great))
#
found = [filename for filename in already_downloaded_great if full_filename_great in filename]
if len(found)>0:
print(" {0} already extracted".format(full_filename_great))
else:
print(" {0} extracting".format(full_filename_great))
#cmd="tar -xvf {0} --wildcards --no-anchored '*Tmb.great' --one-top-level={1}".format(tar_filename,data_folder)
cmd="tar -xvf {0} --one-top-level={1}".format(tar_filename, data_folder)
print(cmd)
output = subprocess.check_call(cmd, shell=True)
```
# THE DATA PRODUCTS IN THE ARCHIVE #
## LEVEL 3 DATA ###
### Level 3 data contains fully calibrated spectra ###
### produced by the calibration program kalibrate from level 1 data ###
## 2 **.great** files: ##
- Scientific target spectra in units of Antenna Temperature and additional output spectra.
- Scientific target spectra only, in units of Main Beam Temperature.
### Antenna Temperature file ###
```
cmd = '''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
list /toc
'''
output = RunClassScript(cmd)
show_file_content('/tmp/cmd.log', 30)
```
## Additional Spectra ##
- Receiver temperature (TREC)
- System Temperature (TSYS)
- Sky-Hot spectra, Observed and fitted (S-H_OBS, S-H_FIT)
- Opacity (TAU_SIGNAL)
## example TREC spectrum ##
```
cmd ='''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
get 15469
pl
ha "./plots/trec.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
## example TSYS spectrum ##
```
cmd ='''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
get 15082
pl
ha "./plots/tsys.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
## example SKY-HOT_OBS spectra ##
```
cmd ='''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
set mod y -180 -140
get 14993 ! S-H_OBS
pl
get 14995 ! S-H_FIT
pen 1
spec
ha "./plots/s_h.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
## example TAU_SIGNAL spectra ##
```
cmd ='''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
get 14990
pl
ha "./plots/tau_signal.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
# Converting from T<sub>ant</sub> to T<sub>mb</sub> #
## The 2nd level 3 **.great** file ##
- Scientific target spectra only, in units of Main Beam Temperature.
```
cmd ='''
file in "./tmp/data/2017-02-08_GR_F371_04_0066_0010505_1900536.9/2017-02-08_GR_F371_04_0066_0010505_1900536.9_Tant.great"
find
file out "./tmp/data/sofia_school_Tmb.great" single /overwr
set source OMC_CENTER
find
def struct teles_all
list /toc tel /var teles_all
def real mb_values[14]
let mb_values 0.66 0.64 0.67 0.69 0.65 0.66 0.68 0.63 0.64 0.66 0.63 0.64 0.64 0.65
def int mb_counter
let mb_counter = 1
set unit v
set win 0 20
set align v
for telescope /in teles_all%tele
set tele 'telescope'
find
say 'telescope'
say 'mb_counter'
for i 1 to found
get next
modify beam_eff mb_values[mb_counter]
base 1
wr
next
let mb_counter 'mb_counter'+1
next
del /var teles_all
'''
output=RunClassScript(cmd)
```
# main beam temperature T<sub>mb</sub> file #
```
cmd='''
file in "./tmp/data/sofia_school_Tmb.great"
find
list
'''
output=RunClassScript(cmd)
show_file_content('/tmp/cmd.log', 100)
```
# LEVEL 4 DATA#
## Consists of **.great** files that are a combination of level 3 data products ##
- Averaged spectra (**.great**)
- Datacubes (**.lmv**)
### Averaged Pixels ###
### Horizontal and Vertical polarizations separated ###
```
cmd = '''
file in "./tmp/data/sofia_school_Tmb.great"
find
file out "./tmp/data/sofia_school_average_LFA_H_V.great" single /overwr
set def
find
set line CII_L
set teles SOF-LFA*
find
set align v
set nomatch
for i 0 to 6
set tele SOF-LFAH*'i'*
find
average /resample
wr
next
for i 0 to 6
set tele SOF-LFAV*'i'*
find
average /resample
wr
next
'''
output=RunClassScript(cmd)
cmd = '''
file in "./tmp/data/sofia_school_average_LFA_H_V.great"
find
list
'''
output=RunClassScript(cmd)
show_file_content("/tmp/cmd.log",14)
cmd ='''
@kosma-init
file in "./tmp/data/sofia_school_average_LFA_H_V.great"
find
list
set unit v
set mod x -40 40
set mod y -30 130
plot_group /group_by telescope /group_by line /smooth 50 /base 3
ha "./plots/average_h_v.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
### Averaged Pixels ###
### Horizontal and Vertical polarizations averaged together ###
```
cmd='''
file in "./tmp/data/sofia_school_Tmb.great"
find
file out "./tmp/data/sofia_school_average_LFA_HVtogether.great" single /overwr
set align v
set nocheck
set telescope SOF-LFA*
find
for i 0 to 6
set tele SOF-LFA*'i'*
find
average /resample
wr
next
'''
output=RunClassScript(cmd)
cmd='''
file in "./tmp/data/sofia_school_average_LFA_HVtogether.great"
find
list
'''
output=RunClassScript(cmd)
show_file_content("/tmp/cmd.log")
cmd ='''
@kosma-init
file in "./tmp/data/sofia_school_average_LFA_HVtogether.great"
find
list
set unit v
set mod x -40 40
set mod y -30 130
plot_group /group_by telescope /group_by line /smooth 50 /base 3
ha "./plots/average_h_v.png" /dev png /overwr
'''
output=RunClassScript(cmd)
```
# generate average pixel plot
```
class_script='''
set def
@kosma-init
file in "{filename}"
find
set unit v
set mode x {min_x} {max_x}
set mode y {min_y} {max_y}
set line {line}
set source {source}
set window {window}
set align v
find
go where
draw t 15 20 {plot_filename} 6
ha "{plot_filename}_go_where.png" /dev png /overwrite
plot_group /group_by telescope
draw t 15 20 {plot_filename} 6
ha "{plot_filename}" /dev png /overwrite
'''
plots_folder = "./plots"
os.makedirs(plots_folder,exist_ok=True)
#
class_cmd = {}
class_cmd['min_x'] = -40
class_cmd['max_x'] = 40
class_cmd['min_y'] = -30
class_cmd['max_y'] = 130
class_cmd['smooth'] = 10
class_cmd['line'] = "CII_L"
class_cmd['source'] = "OMC_CENTER"
class_cmd['window'] = "-20 20"
#already_downloaded_great = glob.glob("tmp/data/**/*.great")
already_downloaded_great = glob.glob("tmp/data/*Tmb.great")
images = []
for class_filename in already_downloaded_great:
print(class_filename)
class_cmd["filename"] = os.path.abspath(class_filename)
class_cmd["plot_filename"] = "{0}/plot_group_{1}.png".format(plots_folder,os.path.basename(class_filename))
print(class_cmd["plot_filename"])
cmd=class_script.format(**class_cmd)
output = RunClassScript(cmd, no_output=False,display_images=False)
images.extend(output.images)
message = "Test message"
PrintMessage(message)
DisplayImagesInColumns(images,2,'test_gallery')
```
# Converting K to Jy #
```
cmd='''
@jansky 220 11 30
! frequency 220 GHz, beam size in arcsec
'''
output=RunClassScript(cmd)
show_file_content("/tmp/cmd.log", 3)
```
# DATACUBE CREATION IN CLASS #
```
cmd='''
file in "./tmp/data/sofia_school_Tmb.great"
set tel SOF*
find
file out "./tmp/data/sofia_school_Tmb_base.great" mul /overwrite
for i 1 to found
get n
set window 0 20
base 1
smooth box 5
write
next
file in "./tmp/data/sofia_school_Tmb_base.great"
find
set weight e
table "./tmp/data/sofia_school_map" new /range 0 20 v /nocheck cal
let map%beam 15.1
let map%diam 2.5
let map%cell '-map%beam/4' 'map%beam/4'
let map%reso 'map%beam*1.05'
xy_map "./tmp/data/sofia_school_map"
let name "./tmp/data/sofia_school_map"
let type lmv
let first 40
let last 55
go bit
ha "./plots/datacube_sofia_school.png" /dev png /overwrite
'''
output=RunClassScript(cmd)
cmd='''
let name "./tmp/data/sofia_school_map"
let type lmv
let first 49
let last 49
go bit
ha "./plots/datacube_sofia_school_onechan.png" /dev png /overwrite
'''
output=RunClassScript(cmd)
```
# WORKING ON FITS FILES #
## Cygrid ##
### Creating the fits file from class ###
```
cmd='''
set angle sec
file in "./tmp/data/sofia_school_Tmb.great"
find
fits write "./tmp/data/sofia_school_Tmb_index.fits" /mode index
'''
output=RunClassScript(cmd)
class RawData:
def __init__(self, fits_file):
self.fits_file = fits_file
self.listremdframe = np.empty(0, dtype = int)
self.lint = False
self.rms = False
with fits.open(self.fits_file) as hdu:
self.data_fits = hdu[1].data
self.header = hdu[1].header
self.data = self._to_numpy()
self.ra = self.data[:, 3] + self.header['CRVAL2']
self.dec = self.data[:, 4] + self.header['CRVAL3']
self.specs = np.nan_to_num(np.array(self.data[:, -1].tolist()))
self.ra_min, self.ra_max = self.ra.min(), self.ra.max()
self.dec_min, self.dec_max = self.dec.min(), self.dec.max()
def _to_numpy(self):
return np.array(self.data_fits.tolist())
def _to_pandas(self):
self.data = pd.DataFrame(data=self.data,
columns=[self.data_fits.columns[i].name for i in range(len(self.data_fits.columns))])
def cygrid_define(self, ra, dec, header, pixsize = 0.0004, kernelsize_sigma = 0.0002):
'''cygrid params definition'''
mapcenter = header['CRVAL2'], header['CRVAL3']
dnaxis1 = int((ra.max() - ra.min())/pixsize)+1
dnaxis2 = int((dec.max() - dec.min())/pixsize)+1
crpix1 = (ra.max() - mapcenter[0])/pixsize
crpix2 = (mapcenter[1] - dec.min())/pixsize
kernel_type = 'gauss1d'
kernel_params = (kernelsize_sigma, )
kernel_support = 3.0 * kernelsize_sigma
hpx_maxres = kernelsize_sigma / 2.0
header_out = {'NAXIS': 3,
'NAXIS1': dnaxis1,
'NAXIS2': dnaxis2,
'NAXIS3': 301,
'CTYPE1': 'RA---GLS',
'CTYPE2': 'DEC--GLS',
'CUNIT1': 'deg',
'CUNIT2': 'deg',
'CDELT1': -pixsize,
'CDELT2': pixsize,
'CRPIX1': crpix1,
'CRPIX2': crpix2,
'CRVAL1': mapcenter[0],
'CRVAL2': mapcenter[1]
}
return header_out, kernel_type, kernel_params, kernel_support, hpx_maxres
def mygridder(self, data):
if isinstance(data, pd.DataFrame):
data = data.to_numpy()
ra = data[:, 3] + self.header['CRVAL2']
dec = data[:, 4] + self.header['CRVAL3']
print( self.header['CRVAL2'], self.header['CRVAL3'])
print(ra, dec)
specs = np.nan_to_num(np.array(data[:, -1].tolist()))
header_map, kernel_type, kernel_params, kernel_support, hpx_maxres = self.cygrid_define(
ra, dec, self.header, pixsize = 0.001, kernelsize_sigma = 0.0005)
mygridder = cygrid.WcsGrid(header_map)
mygridder.set_kernel(kernel_type, kernel_params, kernel_support, hpx_maxres)
mygridder.grid(ra, dec, specs[:, 100: -100])
gridded_map = mygridder.get_datacube()
target_wcs = mygridder.get_wcs()
return gridded_map, target_wcs
def my3gridder(self):
self.gridded_map, self.target_wcs = self.mygridder(self.data)
filename="./tmp/data/sofia_school_Tmb_index.fits"
data= RawData(filename)
data.my3gridder()
ax0 = plt.subplot(projection=data.target_wcs.celestial)
im0 = ax0.imshow(data.gridded_map[7700])
```
for info on the cygrid parameters check out *https://arxiv.org/pdf/1604.06667.pdf*
| github_jupyter |
```
# Imports
import dash
import dash_core_components as dcc
import dash_html_components as html
import pandas as pd
import plotly.graph_objs as go
# Load and preprocess data
df = pd.read_csv(
'https://gist.githubusercontent.com/chriddyp/'
'cb5392c35661370d95f300086accea51/raw/'
'8e0768211f6b747c0db42a9ce9a0937dafcbd8b2/'
'indicators.csv')
available_indicators = df['Indicator Name'].unique()
# Build AppViewer
from jupyterlab_dash import AppViewer
viewer = AppViewer()
# Build App
external_stylesheets = ['https://codepen.io/chriddyp/pen/bWLwgP.css']
app = dash.Dash(__name__, external_stylesheets=external_stylesheets)
app.layout = html.Div([
html.Div([
html.Div([
dcc.Dropdown(
id='xaxis-column',
options=[{'label': i, 'value': i} for i in available_indicators],
value='Fertility rate, total (births per woman)'
),
dcc.RadioItems(
id='xaxis-type',
options=[{'label': i, 'value': i} for i in ['Linear', 'Log']],
value='Linear',
labelStyle={'display': 'inline-block'}
)
],
style={'width': '48%', 'display': 'inline-block'}),
html.Div([
dcc.Dropdown(
id='yaxis-column',
options=[{'label': i, 'value': i} for i in available_indicators],
value='Life expectancy at birth, total (years)'
),
dcc.RadioItems(
id='yaxis-type',
options=[{'label': i, 'value': i} for i in ['Linear', 'Log']],
value='Linear',
labelStyle={'display': 'inline-block'}
)
],style={'width': '48%', 'float': 'right', 'display': 'inline-block'})
]),
dcc.Graph(id='indicator-graphic'),
dcc.Slider(
id='year--slider',
min=df['Year'].min(),
max=df['Year'].max(),
value=df['Year'].max(),
marks={str(year): str(year) for year in df['Year'].unique()}
)
])
# Callbacks
@app.callback(
dash.dependencies.Output('indicator-graphic', 'figure'),
[dash.dependencies.Input('xaxis-column', 'value'),
dash.dependencies.Input('yaxis-column', 'value'),
dash.dependencies.Input('xaxis-type', 'value'),
dash.dependencies.Input('yaxis-type', 'value'),
dash.dependencies.Input('year--slider', 'value')])
def update_graph(xaxis_column_name, yaxis_column_name,
xaxis_type, yaxis_type,
year_value):
dff = df[df['Year'] == year_value]
return {
'data': [go.Scatter(
x=dff[dff['Indicator Name'] == xaxis_column_name]['Value'],
y=dff[dff['Indicator Name'] == yaxis_column_name]['Value'],
text=dff[dff['Indicator Name'] == yaxis_column_name]['Country Name'],
mode='markers',
marker={
'size': 15,
'opacity': 1,
'color': 'blue',
'line': {'width': 2}
}
)],
'layout': go.Layout(
xaxis={
'title': xaxis_column_name,
'type': 'linear' if xaxis_type == 'Linear' else 'log'
},
yaxis={
'title': yaxis_column_name,
'type': 'linear' if yaxis_type == 'Linear' else 'log'
},
margin={'l': 40, 'b': 40, 't': 10, 'r': 0},
hovermode='closest',
)
}
viewer.show(app)
```
| github_jupyter |
# Example 8: Interfacing with Verilog.
While there is much more about PyRTL design to discuss, at some point somebody
might ask you to do something with your code other than have it print
pretty things out to the terminal. We provide **import from and export to
Verilog of designs**, export of **waveforms to VCD**, and a set of **transforms
that make doing netlist-level transforms and analyis directly in pyrtl easy.**
```
import random
import io
import pyrtl
pyrtl.reset_working_block()
```
## Importing From Verilog
Sometimes it is useful to pull in components written in Verilog to be used
as subcomponents of PyRTL designs or to be subject to analysis written over
the PyRTL core. One standard format supported by PyRTL is **"blif" format:**
https://www.ece.cmu.edu/~ee760/760docs/blif.pdf
Many tools support outputing hardware designs to this format, including the
free open source project "Yosys". Blif files can then be imported either
as a string or directly from a file name by the function input_from_blif.
Here is a simple example of a **1 bit full adder imported and then simulated**
from this blif format.
```
full_adder_blif = """
# Generated by Yosys 0.3.0+ (git sha1 7e758d5, clang 3.4-1ubuntu3 -fPIC -Os)
.model full_adder
.inputs x y cin
.outputs sum cout
.names $false
.names $true
1
.names y $not$FA.v:12$3_Y
0 1
.names x $not$FA.v:11$1_Y
0 1
.names cin $not$FA.v:15$6_Y
0 1
.names ind3 ind4 sum
1- 1
-1 1
.names $not$FA.v:15$6_Y ind2 ind3
11 1
.names x $not$FA.v:12$3_Y ind1
11 1
.names ind2 $not$FA.v:16$8_Y
0 1
.names cin $not$FA.v:16$8_Y ind4
11 1
.names x y $and$FA.v:19$11_Y
11 1
.names ind0 ind1 ind2
1- 1
-1 1
.names cin ind2 $and$FA.v:19$12_Y
11 1
.names $and$FA.v:19$11_Y $and$FA.v:19$12_Y cout
1- 1
-1 1
.names $not$FA.v:11$1_Y y ind0
11 1
.end
"""
pyrtl.input_from_blif(full_adder_blif)
# have to find the actual wire vectors generated from the names in the blif file
x, y, cin = [pyrtl.working_block().get_wirevector_by_name(s) for s in ['x', 'y', 'cin']]
io_vectors = pyrtl.working_block().wirevector_subset((pyrtl.Input, pyrtl.Output))
# we are only going to trace the input and output vectors for clarity
sim_trace = pyrtl.SimulationTrace(wires_to_track=io_vectors)
```
### Now simulate the logic with some random inputs
```
sim = pyrtl.Simulation(tracer=sim_trace)
for i in range(15):
# here we actually generate random booleans for the inputs
sim.step({
'x': random.choice([0, 1]),
'y': random.choice([0, 1]),
'cin': random.choice([0, 1])
})
sim_trace.render_trace(symbol_len=5, segment_size=5)
```
## Exporting to Verilog
However, not only do we want to have a method to import from Verilog, we also
want a way to **export it back out to Verilog** as well. To demonstrate PyRTL's
ability to export in Verilog, we will create a **sample 3-bit counter**. However
unlike the example in example2, we extend it to be **synchronously resetting**.
```
pyrtl.reset_working_block()
zero = pyrtl.Input(1, 'zero')
counter_output = pyrtl.Output(3, 'counter_output')
counter = pyrtl.Register(3, 'counter')
counter.next <<= pyrtl.mux(zero, counter + 1, 0)
counter_output <<= counter
```
The counter **gets 0 in the next cycle if the "zero" signal goes high, otherwise just
counter + 1.** Note that both "0" and "1" are bit extended to the proper length and
here we are making use of that native add operation.
Let's dump this bad boy out
to a verilog file and see what is looks like (here we are using StringIO just to
print it to a string for demo purposes, most likely you will want to pass a normal
open file).
```
print("--- PyRTL Representation ---")
print(pyrtl.working_block())
print()
print("--- Verilog for the Counter ---")
with io.StringIO() as vfile:
pyrtl.OutputToVerilog(vfile)
print(vfile.getvalue())
print("--- Simulation Results ---")
sim_trace = pyrtl.SimulationTrace([counter_output, zero])
sim = pyrtl.Simulation(tracer=sim_trace)
for cycle in range(15):
sim.step({'zero': random.choice([0, 0, 0, 1])})
sim_trace.render_trace()
```
We already did the *"hard" work* of generating a test input for this simulation so
we might want to reuse that work when we take this design through a verilog toolchain.
The class *__OutputVerilogTestbench__ grabs the inputs used in the simulation trace
and sets them up in a standard verilog testbench.*
```
print("--- Verilog for the TestBench ---")
with io.StringIO() as tbfile:
pyrtl.output_verilog_testbench(dest_file=tbfile, simulation_trace=sim_trace)
print(tbfile.getvalue())
```
### Now let's talk about transformations of the hardware block.
Many times when you are
doing some hardware-level analysis you might wish to ignore higher level things like
multi-bit wirevectors, adds, concatination, etc. and just thing about wires and basic
gates. **PyRTL supports "lowering" of designs** into this more restricted set of functionality
though the function *"synthesize".* Once we lower a design to this form we can then **apply
basic optimizations** like constant propgation and dead wire elimination as well. By
printing it out to verilog we can see exactly how the design changed.
```
print("--- Optimized Single-bit Verilog for the Counter ---")
pyrtl.synthesize()
pyrtl.optimize()
with io.StringIO() as vfile:
pyrtl.OutputToVerilog(vfile)
print(vfile.getvalue())
```
| github_jupyter |
# Linear Regression - Insurance Dataset
## Import
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
import statsmodels.api as sm
import scipy.stats as stats
from sklearn.preprocessing import LabelEncoder
import copy
sns.set()
```
## Check Missing Values and Outliers
```
# Read the dataset
insurance_df = pd.read_csv('data/insurance.csv')
# Display the first 10 elements
insurance_df.head()
# Info about the data, including shapes and individual columns
insurance_df.info()
# Check for the null values
insurance_df.isna().apply(pd.value_counts)
# Checking for the outliers
plt.figure(figsize= (20,15))
plt.subplot(3,1,1)
sns.boxplot(x= insurance_df.bmi, color='red')
plt.subplot(3,1,2)
sns.boxplot(x= insurance_df.age, color='blue')
plt.subplot(3,1,3)
sns.boxplot(x= insurance_df.expenses, color='yellow')
plt.show()
```
Interesting output for this analysis:
* bmi has a few extreme values;
* expenses as it is highly skewed, there are quiet a lot of extreme values.
```
# a brief summary for the dataset
insurance_df.describe().T
```
### Age field
* 50% of the people has 39 years
```
# Plots to see the distribution of the continuous features individually
plt.figure(figsize= (20,15))
plt.subplot(3,3,1)
plt.hist(insurance_df.bmi, color='lightblue', edgecolor = 'black', alpha = 0.7)
plt.xlabel('bmi')
plt.figure(figsize= (20,15))
plt.subplot(3,3,2)
plt.hist(insurance_df.bmi, color='lightblue', edgecolor = 'black', alpha = 0.7)
plt.xlabel('age')
plt.figure(figsize= (20,15))
plt.subplot(3,3,2)
plt.hist(insurance_df.bmi, color='lightblue', edgecolor = 'black', alpha = 0.7)
plt.xlabel('charges')
### Write the code to show age and charges graphs, same as we did for bmi
plt.show()
```
Output should include this Analysis:
- bmi looks normally distributed.
- Age looks uniformly distributed.
- As seen in the previous step, charges are highly skewed
```
# showing the skewness of variables
Skewness = pd.DataFrame({'Skeweness' : [stats.skew(insurance_df.bmi), stats.skew(insurance_df.age), stats.skew(insurance_df.charges)]},
index=['bmi','age','charges'])
# We Will Measure the skeweness of the required columns
Skewness
```
Output should include this Analysis:
- Skewness of bmi is very low as seen in the previous step
- age is uniformly distributed and thus not skewed
- charges are highly skewed
```
# visualizing data to make analysis
plt.figure(figsize=(20,25))
x = insurance_df.smoker.value_counts().index #Values for x-axis
y = [insurance_df['smoker'].value_counts()[i] for i in x] # Count of each class on y-axis
plt.subplot(4,2,1)
plt.bar(x,y, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) # plot a bar chart
plt.xlabel('Smoker?')
plt.ylabel('Count ')
plt.title('Smoker distribution')
x1 = insurance_df.sex.value_counts().index #Values for x-axis
y1 = [insurance_df['sex'].value_counts()[j] for j in x1] # Count of each class on y-axis
plt.subplot(4,2,2)
plt.bar(x,y, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) # plot a bar chart
plt.xlabel('Gender')
plt.ylabel('Count ')
plt.title('Gender distribution')
x2 = insurance_df.region.value_counts().index #Values for x-axis
y2 = [insurance_df['region'].value_counts()[k] for k in x2] # Count of each class on y-axis
plt.subplot(4,2,3)
plt.bar(x,y, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) # plot a bar chart
plt.xlabel('Region')
plt.ylabel('Count ')
plt.title('Region distribution')
x3 = insurance_df.children.value_counts().index #Values for x-axis
y3 = [insurance_df['children'].value_counts()[l] for l in x3] # Count of each class on y-axis
plt.subplot(4,2,4)
plt.bar(x,y, align='center',color = 'red',edgecolor = 'black',alpha = 0.7) # plot a bar chart
plt.xlabel('No. of children')
plt.ylabel('Count ')
plt.title('Children distribution')
plt.show()
```
- There are lot more non-smokers than smokers.
- Instances are distributed evenly accross all regions.
- Gender is also distributed evenly.
- Most instances have less than 3 children and very few have 4 or 5 children.
```
# Label encoding the variables before doing a pairplot because pairplot ignores strings
insurance_df_encoded = copy.deepcopy(insurance_df)
insurance_df_encoded.loc[:,['sex', 'smoker', 'region']] = insurance_df_encoded.loc[:,['sex', 'smoker', 'region']].apply(LabelEncoder().fit_transform)
sns.pairplot(insurance_df_encoded) # pairplot
plt.show()
```
Output should include this Analysis:
- There is an obvious correlation between 'charges' and 'smoker'
- Looks like smokers claimed more money than non-smokers
- There's an interesting pattern between 'age' and 'charges'. Notice that older people are charged more than the younger ones
```
# Do charges of people who smoke differ significantly from the people who don't?
print("Do charges of people who smoke differ significantly from the people who don't?")
insurance_df.smoker.value_counts()
# Scatter plot to look for visual evidence of dependency between attributes smoker and charges accross different ages
plt.figure(figsize=(8,6))
sns.scatterplot(insurance_df.age, insurance_df.charges,hue=insurance_df.smoker,palette= ['red','green'] ,alpha=0.6)
plt.title("Difference between charges of smokers and charges of non-smokers is apparent")
plt.show()
plt.figure(figsize=(8,6))
sns.scatterplot(insurance_df.age, insurance_df.charges,hue=insurance_df.sex,palette= ['pink','lightblue'] )
# T-test to check dependency of smoking on charges
Ho = "Charges of smoker and non-smoker are same" # Stating the Null Hypothesis
Ha = "Charges of smoker and non-smoker are not the same" # Stating the Alternate Hypothesis
x = np.array(insurance_df[insurance_df.smoker =="yes"].charges)
y = np.array(insurance_df[insurance_df.smoker =="no"].charges)
t, p_value = stats.ttest_ind(x,y, axis = 0) # Performing an Independent t-test
if p_value < 0.05: # Setting our significance level at 5%
print(f'{Ha} as the p_value ({p_value}) < 0.05')
else:
print(f'{Ho} as the p_value ({p_value}) > 0.05')
#Does bmi of males differ significantly from that of females?
print ("Does bmi of males differ significantly from that of females?")
insurance_df.sex.value_counts() #Checking the distribution of males and females
# T-test to check dependency of bmi on gender
Ho = "Gender has no effect on bmi" # Stating the Null Hypothesis
Ha = "Gender has an effect on bmi" # Stating the Alternate Hypothesis
x = np.array(insurance_df[insurance_df.sex =="male"].bmi)
y = np.array(insurance_df[insurance_df.sex =="female"].bmi)
t, p_value = stats.ttest_ind(x,y, axis = 0) #Performing an Independent t-test
if p_value < 0.05: # Setting our significance level at 5%
print(f'{Ha} as the p_value ({p_value.round()}) < 0.05')
else:
print(f'{Ho} as the p_value ({p_value.round(3)}) > 0.05')
#Is the proportion of smokers significantly different in different genders?
# Chi_square test to check if smoking habits are different for different genders
Ho = "Gender has no effect on smoking habits" # Stating the Null Hypothesis
Ha = "Gender has an effect on smoking habits" # Stating the Alternate Hypothesis
crosstab = pd.crosstab(insurance_df['sex'], insurance_df['smoker'])
chi, p_value, dof, expected = stats.chi2_contingency(crosstab)
if p_value < 0.05: # Setting our significance level at 5%
print(f'{Ha} as the p_value ({p_value.round(3)}) < 0.05')
else:
print(f'{Ho} as the p_value ({p_value.round(3)}) > 0.05')
crosstab
# Chi_square test to check if smoking habits are different for people of different regions
Ho = "Region has no effect on smoking habits" # Stating the Null Hypothesis
Ha = "Region has an effect on smoking habits" # Stating the Alternate Hypothesis
crosstab = pd.crosstab(insurance_df['smoker'], insurance_df['region'])
chi, p_value, dof, expected = stats.chi2_contingency(crosstab)
if p_value < 0.05: # Setting our significance level at 5%
print(f'{Ha} as the p_value ({p_value.round(3)}) < 0.05')
else:
print(f'{Ho} as the p_value ({p_value.round(3)}) > 0.05')
crosstab
# Is the distribution of bmi across women with no children, one child and two children, the same ?
# Test to see if the distributions of bmi values for females having different number of children, are significantly different
Ho = "No. of children has no effect on bmi" # Stating the Null Hypothesis
Ha = "No. of children has an effect on bmi" # Stating the Alternate Hypothesis
female_df = copy.deepcopy(insurance_df[insurance_df['sex'] == 'female'])
zero = female_df[female_df.children == 0]['bmi']
one = female_df[female_df.children == 1]['bmi']
two = female_df[female_df.children == 2]['bmi']
f_stat, p_value = stats.f_oneway(zero,one,two)
if p_value < 0.05: # Setting our significance level at 5%
print(f'{Ha} as the p_value ({p_value.round(3)}) < 0.05')
else:
print(f'{Ho} as the p_value ({p_value.round(3)}) > 0.05')
```
| github_jupyter |
# Label and feature engineering
Learning objectives:
1. Learn how to use BigQuery to build time-series features and labels for forecasting
2. Learn how to visualize and explore features.
3. Learn effective scaling and normalizing techniques to improve our modeling results
Now that we have explored the data, let's start building our features, so we can build a model.
<h3><font color="#4885ed">Feature Engineering</font> </h3>
Use the `price_history` table, we can look at past performance of a given stock, to try to predict it's future stock price. In this notebook we will be focused on cleaning and creating features from this table.
There are typically two different approaches to creating features with time-series data.
**One approach** is aggregate the time-series into "static" features, such as "min_price_over_past_month" or "exp_moving_avg_past_30_days". Using this approach, we can use a deep neural network or a more "traditional" ML model to train. Notice we have essentially removed all sequention information after aggregating. This assumption can work well in practice.
A **second approach** is to preserve the ordered nature of the data and use a sequential model, such as a recurrent neural network. This approach has a nice benefit that is typically requires less feature engineering. Although, training sequentially models typically takes longer.
In this notebook, we will build features and also create rolling windows of the ordered time-series data.
<h3><font color="#4885ed">Label Engineering</font> </h3>
We are trying to predict if the stock will go up or down. In order to do this we will need to "engineer" our label by looking into the future and using that as the label. We will be using the [`LAG`](https://cloud.google.com/bigquery/docs/reference/standard-sql/functions-and-operators#lag) function in BigQuery to do this. Visually this looks like:

## Import libraries; setup
```
PROJECT = 'your-gcp-project' # Replace with your project ID.
import pandas as pd
from google.cloud import bigquery
from IPython.core.magic import register_cell_magic
from IPython import get_ipython
bq = bigquery.Client(project = PROJECT)
# Allow you to easily have Python variables in SQL query.
@register_cell_magic('with_globals')
def with_globals(line, cell):
contents = cell.format(**globals())
if 'print' in line:
print(contents)
get_ipython().run_cell(contents)
def create_dataset():
dataset = bigquery.Dataset(bq.dataset("stock_market"))
try:
bq.create_dataset(dataset) # Will fail if dataset already exists.
print("Dataset created")
except:
print("Dataset already exists")
create_dataset()
```
## Create time-series features and determine label based on market movement
### Summary of base tables
```
%%with_globals
%%bigquery --project {PROJECT}
SELECT count(*) as cnt
FROM `stock_src.price_history`
%%with_globals
%%bigquery --project {PROJECT}
SELECT count(*) as cnt
FROM `stock_src.snp500`
```
### Label engineering
Ultimately, we need to end up with a single label for each day. The label takes on 3 values: {`down`, `stay`, `up`}, where `down` and `up` indicates the normalized price (more on this below) went down 1% or more and up 1% or more, respectively. `stay` indicates the stock remained within 1%.
The steps are:
1. Compare close price and open price
2. Compute price features using analytics functions
3. Compute normalized price change (%)
4. Join with S&P 500 table
5. Create labels (`up`, `down`, `stay`)
<h3><font color="#4885ed">Compare close price and open price</font> </h3>
For each row, get the close price of yesterday and the open price of tomorrow using the [`LAG`](https://cloud.google.com/bigquery/docs/reference/legacy-sql#lag) function. We will determine tomorrow's close - today's close.
#### Shift to get tomorrow's close price.
**Learning objective 1**
```
%%with_globals print
%%bigquery df --project {PROJECT}
CREATE OR REPLACE TABLE `stock_market.price_history_delta`
AS
(
WITH shifted_price AS
(
SELECT *,
(LAG(close, 1) OVER (PARTITION BY symbol order by Date DESC)) AS tomorrow_close
FROM `stock_src.price_history`
WHERE Close > 0
)
SELECT a.*,
(tomorrow_close - Close) AS tomo_close_m_close
FROM shifted_price a
)
%%with_globals
%%bigquery --project {PROJECT}
SELECT *
FROM stock_market.price_history_delta
ORDER by Date
LIMIT 100
```
The stock market is going up on average.
**Learning objective 2**
```
%%with_globals print
%%bigquery --project {PROJECT}
SELECT
AVG(close) AS avg_close,
AVG(tomorrow_close) AS avg_tomo_close,
AVG(tomorrow_close) - AVG(close) AS avg_change,
COUNT(*) cnt
FROM
stock_market.price_history_delta
```
### Add time series features
<h3><font color="#4885ed">Compute price features using analytics functions</font> </h3>
In addition, we will also build time-series features using the min, max, mean, and std (can you think of any over functions to use?). To do this, let's use [analytic functions]() in BigQuery (also known as window functions).
```
An analytic function is a function that computes aggregate values over a group of rows. Unlike aggregate functions, which return a single aggregate value for a group of rows, analytic functions return a single value for each row by computing the function over a group of input rows.
```
Using the `AVG` analytic function, we can compute the average close price of a given symbol over the past week (5 business days):
```python
(AVG(close) OVER (PARTITION BY symbol
ORDER BY Date
ROWS BETWEEN 5 PRECEDING AND 1 PRECEDING)) / close
AS close_avg_prior_5_days
```
**Learning objective 1**
```
def get_window_fxn(agg_fxn, n_days):
"""Generate a time-series feature.
E.g., Compute the average of the price over the past 5 days."""
SCALE_VALUE = 'close'
sql = '''
({agg_fxn}(close) OVER (PARTITION BY symbol
ORDER BY Date
ROWS BETWEEN {n_days} PRECEDING AND 1 PRECEDING))/{scale}
AS close_{agg_fxn}_prior_{n_days}_days'''.format(
agg_fxn=agg_fxn, n_days=n_days, scale=SCALE_VALUE)
return sql
WEEK = 5
MONTH = 20
YEAR = 52*5
agg_funcs = ('MIN', 'MAX', 'AVG', 'STDDEV')
lookbacks = (WEEK, MONTH, YEAR)
sqls = []
for fxn in agg_funcs:
for lookback in lookbacks:
sqls.append(get_window_fxn(fxn, lookback))
time_series_features_sql = ','.join(sqls) # SQL string.
def preview_query():
print(time_series_features_sql[0:1000])
preview_query()
%%with_globals print
%%bigquery --project {PROJECT}
CREATE OR REPLACE TABLE stock_market.price_features_delta
AS
SELECT *
FROM
(SELECT *,
{time_series_features_sql},
-- Also get the raw time-series values; will be useful for the RNN model.
(ARRAY_AGG(close) OVER (PARTITION BY symbol
ORDER BY Date
ROWS BETWEEN 260 PRECEDING AND 1 PRECEDING))
AS close_values_prior_260,
ROW_NUMBER() OVER (PARTITION BY symbol ORDER BY Date) AS days_on_market
FROM stock_market.price_history_delta)
WHERE days_on_market > {YEAR}
%%bigquery --project {PROJECT}
SELECT *
FROM stock_market.price_features_delta
ORDER BY symbol, Date
LIMIT 10
```
#### Compute percentage change, then self join with prices from S&P index.
We will also compute price change of S&P index, GSPC. We do this so we can compute the normalized percentage change.
<h3><font color="#4885ed">Compute normalized price change (%)</font> </h3>
Before we can create our labels we need to normalize the price change using the S&P 500 index. The normalization using the S&P index fund helps ensure that the future price of a stock is not due to larger market effects. Normalization helps us isolate the factors contributing to the performance of a stock_market.
Let's use the normalization scheme from by subtracting the scaled difference in the S&P 500 index during the same time period.
In Python:
```python
# Example calculation.
scaled_change = (50.59 - 50.69) / 50.69
scaled_s_p = (939.38 - 930.09) / 930.09
normalized_change = scaled_change - scaled_s_p
assert normalized_change == ~1.2%
```
```
scaled_change = (50.59 - 50.69) / 50.69
scaled_s_p = (939.38 - 930.09) / 930.09
normalized_change = scaled_change - scaled_s_p
print('''
scaled change: {:2.3f}
scaled_s_p: {:2.3f}
normalized_change: {:2.3f}
'''.format(scaled_change, scaled_s_p, normalized_change))
```
### Compute normalized price change (shown above).
Let's join scaled price change (tomorrow_close / close) with the [gspc](https://en.wikipedia.org/wiki/S%26P_500_Index) symbol (symbol for the S&P index). Then we can normalize using the scheme described above.
**Learning objective 3**
```
snp500_index = 'gspc'
%%with_globals print
%%bigquery --project {PROJECT}
CREATE OR REPLACE TABLE stock_market.price_features_norm_per_change
AS
WITH
all_percent_changes AS
(
SELECT *, (tomo_close_m_close / Close) AS scaled_change
FROM `stock_market.price_features_delta`
),
s_p_changes AS
(SELECT
scaled_change AS s_p_scaled_change,
date
FROM all_percent_changes
WHERE symbol="{snp500_index}")
SELECT all_percent_changes.*,
s_p_scaled_change,
(scaled_change - s_p_scaled_change)
AS normalized_change
FROM
all_percent_changes LEFT JOIN s_p_changes
--# Add S&P change to all rows
ON all_percent_changes.date = s_p_changes.date
```
#### Verify results
```
%%with_globals print
%%bigquery df --project {PROJECT}
SELECT *
FROM stock_market.price_features_norm_per_change
LIMIT 10
df.head()
```
<h3><font color="#4885ed">Join with S&P 500 table and Create labels: {`up`, `down`, `stay`}</font> </h3>
Join the table with the list of S&P 500. This will allow us to limit our analysis to S&P 500 companies only.
Finally we can create labels. The following SQL statement should do:
```sql
CASE WHEN normalized_change < -0.01 THEN 'DOWN'
WHEN normalized_change > 0.01 THEN 'UP'
ELSE 'STAY'
END
```
**Learning objective 1**
```
down_thresh = -0.01
up_thresh = 0.01
%%with_globals print
%%bigquery df --project {PROJECT}
CREATE OR REPLACE TABLE stock_market.percent_change_sp500
AS
SELECT *,
CASE WHEN normalized_change < {down_thresh} THEN 'DOWN'
WHEN normalized_change > {up_thresh} THEN 'UP'
ELSE 'STAY'
END AS direction
FROM stock_market.price_features_norm_per_change features
INNER JOIN `stock_src.snp500`
USING (symbol)
%%with_globals print
%%bigquery --project {PROJECT}
SELECT direction, COUNT(*) as cnt
FROM stock_market.percent_change_sp500
GROUP BY direction
%%with_globals print
%%bigquery df --project {PROJECT}
SELECT *
FROM stock_market.percent_change_sp500
LIMIT 20
df.columns
```
The dataset is still quite large and the majority of the days the market `STAY`s. Let's focus our analysis on dates where [earnings per share](https://en.wikipedia.org/wiki/Earnings_per_share) (EPS) information is released by the companies. The EPS data has 3 key columns surprise, reported_EPS, and consensus_EPS:
```
%%with_globals print
%%bigquery --project {PROJECT}
SELECT *
FROM `stock_src.eps`
LIMIT 10
```
The surprise column indicates the difference between the expected (consensus expected eps by analysts) and the reported eps. We can join this table with our derived table to focus our analysis during earnings periods:
```
%%with_globals print
%%bigquery --project {PROJECT}
CREATE OR REPLACE TABLE stock_market.eps_percent_change_sp500
AS
SELECT a.*, b.consensus_EPS, b.reported_EPS, b.surprise
FROM stock_market.percent_change_sp500 a
INNER JOIN `stock_src.eps` b
ON a.Date = b.date
AND a.symbol = b.symbol
%%with_globals print
%%bigquery --project {PROJECT}
SELECT *
FROM stock_market.eps_percent_change_sp500
LIMIT 5
%%with_globals print
%%bigquery --project {PROJECT}
SELECT direction, COUNT(*) as cnt
FROM stock_market.eps_percent_change_sp500
GROUP BY direction
```
## Feature exploration
Now that we have created our recent movements of the company’s stock price, let's visualize our features. This will help us understand the data better and possibly spot errors we may have made during our calculations.
As a reminder, we calculated the scaled prices 1 week, 1 month, and 1 year before the date that we are predicting at.
Let's write a re-usable function for aggregating our features.
**Learning objective 2**
```
def get_aggregate_stats(field, round_digit=2):
"""Run SELECT ... GROUP BY field, rounding to nearest digit."""
df = bq.query('''
SELECT {field}, COUNT(*) as cnt
FROM
(SELECT ROUND({field}, {round_digit}) AS {field}
FROM stock_market.eps_percent_change_sp500) rounded_field
GROUP BY {field}
ORDER BY {field}'''.format(field=field,
round_digit=round_digit,
PROJECT=PROJECT)).to_dataframe()
return df.dropna()
field = 'close_AVG_prior_260_days'
CLIP_MIN, CLIP_MAX = 0.1, 4.
df = get_aggregate_stats(field)
values = df[field].clip(CLIP_MIN, CLIP_MAX)
counts = 100 * df['cnt'] / df['cnt'].sum() # Percentage.
ax = values.hist(weights=counts, bins=30, figsize=(10,5))
ax.set(xlabel=field, ylabel="%");
field = 'normalized_change'
CLIP_MIN, CLIP_MAX = -0.1, 0.1
df = get_aggregate_stats(field, round_digit=3)
values = df[field].clip(CLIP_MIN, CLIP_MAX)
counts = 100 * df['cnt'] / df['cnt'].sum() # Percentage.
ax = values.hist(weights=counts, bins=50, figsize=(10, 5))
ax.set(xlabel=field, ylabel="%");
```
Let's look at results by day-of-week, month, etc.
```
VALID_GROUPBY_KEYS = ('DAYOFWEEK', 'DAY', 'DAYOFYEAR',
'WEEK', 'MONTH', 'QUARTER', 'YEAR')
DOW_MAPPING = {1: 'Sun', 2: 'Mon', 3: 'Tues', 4: 'Wed',
5: 'Thur', 6: 'Fri', 7: 'Sun'}
def groupby_datetime(groupby_key, field):
if groupby_key not in VALID_GROUPBY_KEYS:
raise Exception('Please use a valid groupby_key.')
sql = '''
SELECT {groupby_key}, AVG({field}) as avg_{field}
FROM
(SELECT {field},
EXTRACT({groupby_key} FROM date) AS {groupby_key}
FROM stock_market.eps_percent_change_sp500) foo
GROUP BY {groupby_key}
ORDER BY {groupby_key} DESC'''.format(groupby_key=groupby_key,
field=field,
PROJECT=PROJECT)
print(sql)
df = bq.query(sql).to_dataframe()
if groupby_key == 'DAYOFWEEK':
df.DAYOFWEEK = df.DAYOFWEEK.map(DOW_MAPPING)
return df.set_index(groupby_key).dropna()
field = 'normalized_change'
df = groupby_datetime('DAYOFWEEK', field)
ax = df.plot(kind='barh', color='orange', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
field = 'close'
df = groupby_datetime('DAYOFWEEK', field)
ax = df.plot(kind='barh', color='orange', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
field = 'normalized_change'
df = groupby_datetime('MONTH', field)
ax = df.plot(kind='barh', color='blue', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
field = 'normalized_change'
df = groupby_datetime('QUARTER', field)
ax = df.plot(kind='barh', color='green', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
field = 'close'
df = groupby_datetime('YEAR', field)
ax = df.plot(kind='line', color='purple', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
field = 'normalized_change'
df = groupby_datetime('YEAR', field)
ax = df.plot(kind='line', color='purple', alpha=0.7)
ax.grid(which='major', axis='y', linewidth=0)
```
BONUS: How do our features correlate with the label `direction`? Build some visualizations. What features are most important? You can visualize this and do it statistically using the [`CORR`](https://cloud.google.com/bigquery/docs/reference/standard-sql/statistical_aggregate_functions) function.
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
```
import os
from glob import glob
from pathlib import Path
import numpy as np
import pytorch_lightning as pl
import torch
import torch.nn as nn
import torchvision
from sklearn.model_selection import train_test_split
from torch.utils.data import Dataset, DataLoader
from torchsummary import summary
from tqdm.notebook import tqdm
from utils import *
device = torch.device('cuda')
_RANDOM_STATE = 42
seed_everything(_RANDOM_STATE)
video_paths = sorted(glob('../../Algonauts2021_devkit/AlgonautsVideos268_All_30fpsmax/*.mp4'))
audio_paths = sorted(glob('../../Algonauts2021_devkit/AlgonautsVideos268_All_30fpsmax/*.wav'))
if audio_paths is None:
for video_path in video_paths:
extract_audios(video_path)
print(f'Found {len(video_paths)} videos, {len(audio_paths)} audios')
def _extract_densenet_embeddings(model, video_paths, mode='mean'):
"""Extract embeddings from an ImageNet-pretrained model.
Future implementation will be more name-specific and include per-block feature extraction, e.g.:
if model_name == 'densenet':
for block_idx in len(densenet_blocks):
extract_features_per_densenet_block(model, input)
...
"""
resize_normalize = torchvision.transforms.Compose([
torchvision.transforms.Resize((224, 224)),
torchvision.transforms.ToTensor(),
torchvision.transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])])
embeddings = []
for video_path in tqdm(video_paths, desc='Extracting video features...', total=len(video_paths)):
vid, num_frames = sample_video_from_mp4(video_path)
embedding = []
for frame, img in enumerate(vid):
input_img = torch.autograd.Variable(resize_normalize(img).unsqueeze(0))
frame_embedding = model(input_img.cuda()).flatten()
embedding.append(frame_embedding.cpu().detach())
if mode == 'mean':
embeddings.append(torch.stack(embedding).mean(0).numpy())
elif mode == 'statpool':
embeddings.append(torch.cat([torch.stack(embedding).mean(0), torch.stack(embedding).std(0)]).numpy())
else:
raise ValueError('Embedding aggregation mode not found.')
embeddings = np.array(embeddings, dtype=np.float32)
return embeddings
# Simple way instead of extracting at each block
# Extract features just before classification
# Don't know whether per-block feature extraction is better, but this is a simpler solution in the meantime
image_model = torchvision.models.densenet169(pretrained=True)
image_model.features.norm5 = nn.Identity()
image_model.classifier = nn.Identity()
image_model.eval()
image_model.to(device)
def _get_densenet169_blocks(idx):
assert idx in [-1, -3, -5, -7]
layers = list(image_model.features.children())[:idx]
layers += [nn.AdaptiveAvgPool2d((1,1)), nn.Flatten(start_dim=1)]
return nn.Sequential(*layers)
block_embeddings = []
for idx in [-1, -3, -5, -7]:
model = _get_densenet169_blocks(idx)
block_embeddings.append(_extract_densenet_embeddings(model, video_paths))
for b in block_embeddings:
print(b.shape)
np.save(f'densenet169_concat_all_blocks', np.concatenate(block_embeddings, axis=-1))
for b in block_embeddings:
np.save(f'densenet169_img_embeddings_{b.shape[-1]}', b)
all_embeddings = np.load('densenet169_img_embeddings.npy')
train_embeddings, test_embeddings = np.split(img_embeddings, [1000], axis=0)
print(all_embeddings.shape)
sub_folders = sorted(os.listdir('../../Algonauts2021_devkit/participants_data_v2021/mini_track/'))
mini_track_ROIS = sorted(list(map(lambda x: Path(x).stem, os.listdir('../../Algonauts2021_devkit/participants_data_v2021/mini_track/sub01/'))))
def _load_data(sub, ROI, fmri_dir = '../../Algonauts2021_devkit/participants_data_v2021', batch_size=128, use_gpu=True):
if ROI == "WB":
track = "full_track"
else:
track = "mini_track"
fmri_dir = os.path.join(fmri_dir, track)
sub_fmri_dir = os.path.join(fmri_dir, sub)
results_dir = os.path.join('../results/', f'{image_model.__class__.__name__}', track, sub)
if track == "full_track":
fmri_train_all, voxel_mask = get_fmri(sub_fmri_dir, ROI)
else:
fmri_train_all = get_fmri(sub_fmri_dir, ROI)
return fmri_train_all
!ls ../../Algonauts2021_devkit/participants_data_v2021/mini_track/sub04/EBA.pkl
sub = 'sub04'
ROI = mini_track_ROIS[0]
train_fmri = _load_data(sub, ROI)
train_embeddings, val_embeddings, train_voxels, val_voxels = train_test_split(train_embeddings, train_fmri)
class MultiVoxelAutoEncoder(pl.LightningModule):
def __init__(self, in_features=1668, out_features=368):
super().__init__()
self.encoder = nn.Sequential(
nn.Linear(in_features, 1024),
nn.ReLU(),
nn.Linear(1024, 512),
nn.ReLU(),
nn.Linear(512, out_features)
)
self.decoder = nn.Sequential(
nn.Linear(out_features, 512),
nn.ReLU(),
nn.Linear(512, 1024),
nn.ReLU(),
nn.Linear(1024, in_features)
)
def forward(self, x):
x = self.encoder(x)
# x = self.decoder(x)
return x
def training_step(self, batch, batch_idx):
x, y = batch
x = x.view(x.size(0), -1)
z = self.encoder(x)
x_hat = self.decoder(z)
loss = F.mse_loss(x_hat, x)
self.log("train_loss", loss)
return loss
def configure_optimizers(self):
optimizer = torch.optim.Adam(self.parameters(), lr=1e-3)
return optimizer
model = MultiVoxelAutoEncoder()
summary(model.cuda(), input_size=(1,1668))
class VoxelDataset(Dataset):
def __init__(self, features, voxel_maps=None, transform=None):
self.features = features
self.voxel_maps = voxel_maps
self.transform = transform
def __len__(self):
return len(self.features)
def __getitem__(self, idx):
feat = self.features[idx]
if self.voxel_maps:
voxel_map = self.voxel_maps[idx]
if self.transform:
feat = self.transform(feat)
return feat, voxel_map
train_dataset = VoxelDataset(train_embeddings, train_voxels)
val_dataset = VoxelDataset(val_embeddings, val_voxels)
train_loader = DataLoader(train_dataset)
val_loader = DataLoader(val_dataset)
mvae = MultiVoxelAutoEncoder()
trainer = pl.Trainer(gpus=2)
trainer.fit(mvae, train_loader)
```
| github_jupyter |
```
videos = """
https://www.youtube.com/watch?v=4CSV7cIXAS8&ab_channel=TheSmartLocal
https://www.youtube.com/watch?v=-SibiwunZ14&ab_channel=SneakySushii
https://www.youtube.com/watch?v=kO9uBkW_OIA&ab_channel=SneakySushii
https://www.youtube.com/watch?v=li1bIAhRp_I&ab_channel=SneakySushii
https://www.youtube.com/watch?v=Nk9AzEx3rU8&ab_channel=WORLDOFBUZZ
https://www.youtube.com/watch?v=CceSkvU4sUM&ab_channel=JianHaoTan
https://www.youtube.com/watch?v=ieZwYjMcP20&ab_channel=TheMingThing
https://www.youtube.com/watch?v=qQyjWXTGkcM&ab_channel=BertVaux
https://www.youtube.com/watch?v=F15a-MH_Hzg&t=1290s&ab_channel=TheThirstySisters
https://www.youtube.com/watch?v=-mAdHcMGmEs&t=996s&ab_channel=JinnyboyTVHangouts
https://www.youtube.com/watch?v=WnipnOcDkx8
https://www.youtube.com/watch?v=fzuKoKTxfEs
https://www.youtube.com/watch?v=Hioy9UvH4yE
https://www.youtube.com/watch?v=98CiTQQtqak
https://www.youtube.com/watch?v=HKnXUGo_H2U
https://www.youtube.com/watch?v=3-DoQSmz5aI
https://www.youtube.com/watch?v=ViJXVHWTRz8
https://www.youtube.com/watch?v=XLV720mng6w
https://www.youtube.com/watch?v=gvPiF-zXur4
https://www.youtube.com/watch?v=mIc0MmSoaiI
https://www.youtube.com/watch?v=oW3yttdKVm0
https://www.youtube.com/watch?v=zHC1MPeSP34
https://www.youtube.com/watch?v=6i8PG2cTAUU
https://www.youtube.com/watch?v=gvPiF-zXur4
https://www.youtube.com/watch?v=mIc0MmSoaiI
https://www.youtube.com/watch?v=oW3yttdKVm0
https://www.youtube.com/watch?v=zHC1MPeSP34
https://www.youtube.com/watch?v=6i8PG2cTAUU
https://www.youtube.com/watch?v=8DMyfcjyrZQ
https://www.youtube.com/watch?v=rkwCky94IhQ
https://www.youtube.com/watch?v=Y6RYZpbANJI&t=1451s
https://www.youtube.com/watch?v=oJPtCCTzs_o
https://www.youtube.com/watch?v=O_rK97AUOIA
https://www.youtube.com/watch?v=ekKcn3QCU5E
https://www.youtube.com/watch?v=OZMsLGKDv14
https://www.youtube.com/watch?v=TxeBIVuXRHM
https://www.youtube.com/watch?v=Q54z0mutz5Q&ab_channel=JinnyboyTVHangouts
https://www.youtube.com/watch?v=ZcVRJSDCof0
https://www.youtube.com/watch?v=te2Be5GNNys
https://www.youtube.com/watch?v=jcvIfHTy1H0&t=1148s
https://www.youtube.com/watch?v=O19y2ZG8Y-k&t=1464s
https://www.youtube.com/watch?v=qVpcfmZF1oY
https://www.youtube.com/watch?v=L2W4inT6j3c
https://www.youtube.com/watch?v=g3iwaxDKQGQ&t=1940s
https://www.youtube.com/watch?v=XldSJLh8kMk&t=592s
https://www.youtube.com/watch?v=shr5NaEWQqk
https://www.youtube.com/watch?v=6Oq9APAIYLg
https://www.youtube.com/watch?v=TJjJ1MqyUpQ
https://www.youtube.com/watch?v=Ra_xW9MwRKQ&t=894s
https://www.youtube.com/watch?v=x8Gl_zIC9CQ
https://www.youtube.com/watch?v=QLz7USNFnng
https://www.youtube.com/watch?v=vshZhU5dbto
https://www.youtube.com/watch?v=skGDf57Q9xs
https://www.youtube.com/watch?v=r6f3OoHVkgU
https://www.youtube.com/watch?v=Zz3NcuK7QHM&ab_channel=AlastairPang
https://www.youtube.com/watch?v=NZXHVJEkxm8
https://www.youtube.com/watch?v=ezFAYdIwu_8&ab_channel=AlastairPang
https://www.youtube.com/watch?v=ZDloTzTfenM&ab_channel=TEAMTITAN
https://www.youtube.com/watch?v=Hioy9UvH4yE
https://www.youtube.com/watch?v=-L7y6_6Ar4s
https://www.youtube.com/watch?v=gW9sv26g4Io
https://www.youtube.com/watch?v=T218uk6pi4E
https://www.youtube.com/watch?v=281TVct4AvE
https://www.youtube.com/watch?v=luGvazsVsHc
https://www.youtube.com/watch?v=q6EyuUemq5U
https://www.youtube.com/watch?v=r8sUO3VXPks
https://www.youtube.com/watch?v=LsV8tnGSPEQ
https://www.youtube.com/watch?v=ezYaMrChz4M&ab_channel=AlastairPang
https://www.youtube.com/watch?v=U4dYpNf8oWY
https://www.youtube.com/watch?v=e4mNzKOZrJ0
https://www.youtube.com/watch?v=R7nCJrdRWyc
https://www.youtube.com/watch?v=7izYax1bCwg
https://www.youtube.com/watch?v=1EqlSuT6vUY
https://www.youtube.com/watch?v=kbOrRjEDnDU
https://www.youtube.com/watch?v=V2k9OWeTX84
https://www.youtube.com/watch?v=kmDb6Q-afPc
https://www.youtube.com/watch?v=roNgntnM3EI
https://www.youtube.com/watch?v=OYasARh1cBQ
https://www.youtube.com/watch?v=6RWRMNiBpJA
https://www.youtube.com/watch?v=nMQZmuINIDk
https://www.youtube.com/watch?v=-L7y6_6Ar4s
https://www.youtube.com/watch?v=NZXHVJEkxm8
https://www.youtube.com/watch?v=C3U2AtbBjHY
https://www.youtube.com/watch?v=9SKGSMRpymc
https://www.youtube.com/watch?v=h1v-4x0YP70
https://www.youtube.com/watch?v=TqLaI2X4jj0
https://www.youtube.com/watch?v=745rfUlGzgM
https://www.youtube.com/watch?v=BHF0ift4aRA
https://www.youtube.com/watch?v=LsV8tnGSPEQ
https://www.youtube.com/watch?v=r8sUO3VXPks
https://www.youtube.com/watch?v=b9YHjghfd2E&ab_channel=watermelon
https://www.youtube.com/watch?v=vpaZHhpn8CY&ab_channel=watermelon
https://www.youtube.com/watch?v=rjjK4aAbKnw
https://www.youtube.com/watch?v=TYL7mmbpsbo
https://www.youtube.com/watch?v=-2d7wzvNOCE
https://www.youtube.com/watch?v=f7qEqPNzGk8
https://www.youtube.com/watch?v=v-RunsFvf8o
https://www.youtube.com/watch?v=cs2LkcSpNpw
https://www.youtube.com/watch?v=vKpmE4ZFnGE
https://www.youtube.com/watch?v=YtCn4hNLk7o
https://www.youtube.com/watch?v=attPdWb6UVo
https://www.youtube.com/watch?v=8QNgzWcc7U4
https://www.youtube.com/watch?v=LXSNbZF9ag4
https://www.youtube.com/watch?v=alUMLUEUdCM
https://www.youtube.com/watch?v=sT3vpK8npYk
https://www.youtube.com/watch?v=uaSWgcKn0RM
https://www.youtube.com/watch?v=OpDeoZzKHBM
https://www.youtube.com/watch?v=jzUidmnXGu8
https://www.youtube.com/watch?v=cSxV6oacgaE
https://www.youtube.com/watch?v=myPdnyUGvUk
https://www.youtube.com/watch?v=YaeQ27BgUN0
https://www.youtube.com/watch?v=JyAEITJk1AI
https://www.youtube.com/watch?v=hi4MpQZVzYY
https://www.youtube.com/watch?v=m1kmhgWoQVE
https://www.youtube.com/watch?v=50EHWNk7aBc
https://www.youtube.com/watch?v=2hHAMAA614w&ab_channel=TEAMTITAN
https://www.youtube.com/watch?v=8QhPCqS5aBA
https://www.youtube.com/watch?v=RBShCX3-BtQ&t=1099s
https://www.youtube.com/watch?v=WnipnOcDkx8
https://www.youtube.com/watch?v=fzuKoKTxfEs
https://www.youtube.com/watch?v=Hioy9UvH4yE
https://www.youtube.com/watch?v=98CiTQQtqak
https://www.youtube.com/watch?v=HKnXUGo_H2U
https://www.youtube.com/watch?v=F15a-MH_Hzg&t=1142s
https://www.youtube.com/watch?v=3-DoQSmz5aI
https://www.youtube.com/watch?v=ViJXVHWTRz8&t=1649s
https://www.youtube.com/watch?v=XLV720mng6w
https://www.youtube.com/watch?v=gvPiF-zXur4
https://www.youtube.com/watch?v=mIc0MmSoaiI
https://www.youtube.com/watch?v=oW3yttdKVm0
https://www.youtube.com/watch?v=zHC1MPeSP34
https://www.youtube.com/watch?v=6i8PG2cTAUU
https://www.youtube.com/watch?v=8DMyfcjyrZQ&t=756s
https://www.youtube.com/watch?v=rkwCky94IhQ
https://www.youtube.com/watch?v=Y6RYZpbANJI&t=1451s
https://www.youtube.com/watch?v=oJPtCCTzs_o
https://www.youtube.com/watch?v=O_rK97AUOIA
https://www.youtube.com/watch?v=Jw7B8efygWI
https://www.youtube.com/watch?v=TxeBIVuXRHM
https://www.youtube.com/watch?v=EYBSXaK3cjI
https://www.youtube.com/watch?v=ekKcn3QCU5E
https://www.youtube.com/watch?v=OZMsLGKDv14
https://www.youtube.com/watch?v=jJN0vH42wcs
https://www.youtube.com/watch?v=Q54z0mutz5Q&t=321s
https://www.youtube.com/watch?v=ZcVRJSDCof0
https://www.youtube.com/watch?v=te2Be5GNNys&t=1915s
https://www.youtube.com/watch?v=jcvIfHTy1H0&t=1148s
https://www.youtube.com/watch?v=O19y2ZG8Y-k&t=1464s
https://www.youtube.com/watch?v=5zAsyNKfzps
https://www.youtube.com/watch?v=qVpcfmZF1oY
https://www.youtube.com/watch?v=L2W4inT6j3c
https://www.youtube.com/watch?v=g3iwaxDKQGQ&t=1940s
https://www.youtube.com/watch?v=XldSJLh8kMk&t=592s
https://www.youtube.com/watch?v=DFFHEbk_iYs&t=1158s
https://www.youtube.com/watch?v=x8Gl_zIC9CQ
https://www.youtube.com/watch?v=QLz7USNFnng
https://www.youtube.com/watch?v=vshZhU5dbto
https://www.youtube.com/watch?v=skGDf57Q9xs
https://www.youtube.com/watch?v=r6f3OoHVkgU
https://www.youtube.com/watch?v=QxVzXbS1WFE
https://www.youtube.com/watch?v=WJkSAHvYyqk
https://www.youtube.com/watch?v=FFtFXiIgsyk
https://www.youtube.com/watch?v=Cq6Yjwsu63U
https://www.youtube.com/watch?v=FcD9MpeWqEI
https://www.youtube.com/watch?v=79ax7ZQss5M
https://www.youtube.com/watch?v=Wyb6i3GoJv4
https://www.youtube.com/watch?v=I2BaN8cm6NY
https://www.youtube.com/watch?v=ZJ8jw3C9vB8
https://www.youtube.com/watch?v=PazCOj1nb3c
https://www.youtube.com/watch?v=XMx9tqwzQS4
https://www.youtube.com/watch?v=sZFykBDVjOI
https://www.youtube.com/watch?v=NqCp8Csl7PE
https://www.youtube.com/watch?v=zLEmAKdG9vY
https://www.youtube.com/watch?v=IAeN0jG9Hwc
https://www.youtube.com/watch?v=0jySfrZhs50
https://www.youtube.com/watch?v=CXnW4hJDp54
https://www.youtube.com/watch?v=Rkvd5p3I11M
https://www.youtube.com/watch?v=PRN5UrsDF6w
https://www.youtube.com/watch?v=RlejfOWnKgQ
https://www.youtube.com/watch?v=FNfbmh93DFo
https://www.youtube.com/watch?v=nGnyZ5c00Mo
https://www.youtube.com/watch?v=EHlGktcfDZc
https://www.youtube.com/watch?v=HkOQy1Nxpis&t=1571s
https://www.youtube.com/watch?v=m7mwwtsblIs
https://www.youtube.com/watch?v=-mAdHcMGmEs&t=92s
https://www.youtube.com/watch?v=IixUPM11VMY
https://www.youtube.com/watch?v=n3AR3nQEc7Y
https://www.youtube.com/watch?v=OWbPqC2i71c
https://www.youtube.com/watch?v=670FyeJhnps
https://www.youtube.com/watch?v=8LuihYdx0FQ
https://www.youtube.com/watch?v=1av_p0DPDas
https://www.youtube.com/watch?v=Jn2oeTajYCw
https://www.youtube.com/watch?v=2M8U_mxGbcw&ab_channel=JinnyboyTVHangouts
https://www.youtube.com/watch?v=7_-4TXv6pvY
https://www.youtube.com/watch?v=rMpf5j7683M
https://www.youtube.com/watch?v=lfoBmafO37A
https://www.youtube.com/watch?v=wnxWhWRk4Sc
https://www.youtube.com/watch?v=Zz3NcuK7QHM&t=175s
https://www.youtube.com/watch?v=X4QapKCCneQ
https://www.youtube.com/watch?v=281TVct4AvE
https://www.youtube.com/watch?v=T218uk6pi4E&t=1097s
https://www.youtube.com/watch?v=3pImfMhLYhA
https://www.youtube.com/watch?v=luGvazsVsHc&t=1839s
https://www.youtube.com/watch?v=6syqXC_o6gs
https://www.youtube.com/watch?v=rMpf5j7683M
https://www.youtube.com/watch?v=lfoBmafO37A
https://www.youtube.com/watch?v=wnxWhWRk4Sc
https://www.youtube.com/watch?v=LsV8tnGSPEQ
https://www.youtube.com/watch?v=BHF0ift4aRA
https://www.youtube.com/watch?v=745rfUlGzgM
https://www.youtube.com/watch?v=TqLaI2X4jj0
https://www.youtube.com/watch?v=9SKGSMRpymc
https://www.youtube.com/watch?v=C3U2AtbBjHY
https://www.youtube.com/watch?v=NZXHVJEkxm8
https://www.youtube.com/watch?v=-L7y6_6Ar4s
https://www.youtube.com/watch?v=6RWRMNiBpJA
https://www.youtube.com/watch?v=7izYax1bCwg
https://www.youtube.com/watch?v=ezYaMrChz4M&t=2265s
https://www.youtube.com/watch?v=e4mNzKOZrJ0
https://www.youtube.com/watch?v=R7nCJrdRWyc
https://www.youtube.com/watch?v=V2k9OWeTX84
https://www.youtube.com/watch?v=kbOrRjEDnDU
https://www.youtube.com/watch?v=1EqlSuT6vUY
https://www.youtube.com/watch?v=ezFAYdIwu_8&t=4264s
https://www.youtube.com/watch?v=U4dYpNf8oWY
https://www.youtube.com/watch?v=e4mNzKOZrJ0
https://www.youtube.com/watch?v=eXpirKKPBUA
https://www.youtube.com/watch?v=5cG8wHdaKIU
https://www.youtube.com/watch?v=EjzkQeWKwQw
https://www.youtube.com/watch?v=adq-khQo2eQ
https://www.youtube.com/watch?v=xsjayMqoC0Y
https://www.youtube.com/watch?v=aa5r9pws0AI
https://www.youtube.com/watch?v=NCiDUw4mQgE
https://www.youtube.com/watch?v=6K1feeFnKpQ
https://www.youtube.com/watch?v=7wiSVCBFmFc
https://www.youtube.com/watch?v=N5EfPvQKW90
https://www.youtube.com/watch?v=iugevNPFp2Y
https://www.youtube.com/watch?v=fgaCmnMdZ0c
https://www.youtube.com/watch?v=UryaihXnjBk
https://www.youtube.com/watch?v=lvIbeZ3qgsU
https://www.youtube.com/watch?v=l6bf4RFE31o
https://www.youtube.com/watch?v=7dPzh9PpLs8
https://www.youtube.com/watch?v=HDuY9UdkkaE
https://www.youtube.com/watch?v=-XwvfCTnseI
https://www.youtube.com/watch?v=jhLgzHQnAHU
https://www.youtube.com/watch?v=TzfCaQ8vUMc
https://www.youtube.com/watch?v=Pkv6d83oe8s
https://www.youtube.com/watch?v=woDYSE37_rI
https://www.youtube.com/watch?v=3pKA5NvTCOA&t=1035s
https://www.youtube.com/watch?v=shV1F1JlDzk
https://www.youtube.com/watch?v=rxmMRAzLRJ0
https://www.youtube.com/watch?v=Yv7uONh96tc
https://www.youtube.com/watch?v=H4M_ZUDAT3k
https://www.youtube.com/watch?v=dykz_vuzJh0&t=2554s
https://www.youtube.com/watch?v=UWm1ESejkzs
https://www.youtube.com/watch?v=otK3Gl_XX4c
https://www.youtube.com/watch?v=xMm23vU8_og
https://www.youtube.com/watch?v=mJuKUwFpDgI
https://www.youtube.com/watch?v=_7Un0q6a9zg&t=1931s
https://www.youtube.com/watch?v=3b0bYe36KNE
https://www.youtube.com/watch?v=RqG0tHR9D1g
https://www.youtube.com/watch?v=eWNQhDNoSsY
https://www.youtube.com/watch?v=tdcjTfzYSbk
https://www.youtube.com/watch?v=WSVo-qvu1GQ&t=1505s
https://www.youtube.com/watch?v=lPnRg9UsP0M&t=2400s
https://www.youtube.com/watch?v=vRnkHzyyBc4
https://www.youtube.com/watch?v=6R87P6YD2c4&t=1061s
https://www.youtube.com/watch?v=CSP1XPZu4YY
https://www.youtube.com/watch?v=VlovwtVYuw0&t=619s
https://www.youtube.com/watch?v=MCHDyLJDJrg&t=2610s
https://www.youtube.com/watch?v=7byI9tIJeRk
https://www.youtube.com/watch?v=gW9sv26g4Io
https://www.youtube.com/watch?v=pFBohAGNYWU
https://www.youtube.com/watch?v=8QhPCqS5aBA
https://www.youtube.com/watch?v=hi4MpQZVzYY
https://www.youtube.com/watch?v=ZDloTzTfenM&t=4143s
https://www.youtube.com/watch?v=cSxV6oacgaE
https://www.youtube.com/watch?v=myPdnyUGvUk
https://www.youtube.com/watch?v=JyAEITJk1AI
https://www.youtube.com/watch?v=YaeQ27BgUN0
https://www.youtube.com/watch?v=OpDeoZzKHBM
https://www.youtube.com/watch?v=jzUidmnXGu8
https://www.youtube.com/watch?v=3mt-_JgTm2E
https://www.youtube.com/watch?v=6yW185RJkGw
https://www.youtube.com/watch?v=QIkOzfAcsmE&t=1783s
https://www.youtube.com/watch?v=eFDcnTupdeQ
https://www.youtube.com/watch?v=sT3vpK8npYk
https://www.youtube.com/watch?v=uaSWgcKn0RM
https://www.youtube.com/watch?v=BrYHWQ4xmSc
https://www.youtube.com/watch?v=alUMLUEUdCM
https://www.youtube.com/watch?v=LXSNbZF9ag4
https://www.youtube.com/watch?v=8QNgzWcc7U4
https://www.youtube.com/watch?v=5VpSUeWPEuo
https://www.youtube.com/watch?v=YtCn4hNLk7o
https://www.youtube.com/watch?v=attPdWb6UVo
https://www.youtube.com/watch?v=Yzagxa4bn-M
https://www.youtube.com/watch?v=vKpmE4ZFnGE
https://www.youtube.com/watch?v=b9YHjghfd2E&t=4654s
https://www.youtube.com/watch?v=cs2LkcSpNpw
https://www.youtube.com/watch?v=v-RunsFvf8o
https://www.youtube.com/watch?v=f7qEqPNzGk8
https://www.youtube.com/watch?v=rjjK4aAbKnw
https://www.youtube.com/watch?v=TYL7mmbpsbo
https://www.youtube.com/watch?v=-2d7wzvNOCE
https://www.youtube.com/watch?v=1FLelqfAyZU
https://www.youtube.com/watch?v=HRd3RMQtkDs
https://www.youtube.com/watch?v=5SOInBKNP18
https://www.youtube.com/watch?v=ONUgGzCm00E
https://www.youtube.com/watch?v=5t_yyrP2OdI
"""
videos = list(set(filter(None, videos.split('\n'))))
len(videos)
import youtube_dl
import mp
from tqdm import tqdm
def loop(urls):
urls = urls[0]
ydl_opts = {
'format': 'bestaudio/best',
'postprocessors': [{
'key': 'FFmpegExtractAudio',
'preferredcodec': 'mp3',
'preferredquality': '192',
}],
'no-check-certificate': True
}
for i in tqdm(range(len(urls))):
try:
with youtube_dl.YoutubeDL(ydl_opts) as ydl:
ydl.download([urls[i]])
except:
pass
import mp
mp.multiprocessing(videos, loop, cores = 12, returned = False)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import kerasncp as kncp
import tensorflow as tf
import copy
import string
import seaborn as sns
import matplotlib.pyplot as plt
import os
import tensorflow as tf
import scipy.stats as stats
import numpy as np
import pandas as pd
from gensim import models
from tensorflow import keras
from tensorflow.keras import layers
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import train_test_split
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
from tensorflow import keras
from kerasncp.tf import LTCCell
from sklearn.model_selection import StratifiedKFold, train_test_split
from google.colab import drive
drive.mount('/content/drive/')
stfold = StratifiedKFold(n_splits=5, shuffle=True)
train_l = list()
val_l = list()
test_l = list()
maxlen = 25
cnt = 0
for train_index, test_index in stfold.split(df['comment'], df['label']):
cnt += 1
train = df.iloc[train_index]
train_x, train_y = train['comment'], train['label']
tmp = df.iloc[test_index]
val, test = train_test_split(tmp, test_size=0.5, stratify=tmp['label'])
val_x, val_y = val['comment'], val['label']
test_x, test_y = test['comment'], test['label']
train_x = vectorizer(np.array([[s] for s in train_x])).numpy()
test_x = vectorizer(np.array([[s] for s in test_x])).numpy()
val_x = vectorizer(np.array([[s] for s in val_x])).numpy()
test_x = keras.preprocessing.sequence.pad_sequences(test_x, maxlen=maxlen)
train_x = keras.preprocessing.sequence.pad_sequences(train_x, maxlen=maxlen)
val_x = keras.preprocessing.sequence.pad_sequences(val_x, maxlen=maxlen)
embed_dim = 32 # Embedding size for each token
wiring = kncp.wirings.NCP(
inter_neurons=12, # Number of inter neurons
command_neurons=8, # Number of command neurons
motor_neurons=1, # Number of motor neurons
sensory_fanout=4, # How many outgoing synapses has each sensory neuron
inter_fanout=4, # How many outgoing synapses has each inter neuron
recurrent_command_synapses=4, # Now many recurrent synapses are in the
# command neuron layer
motor_fanin=6,
)
ltc_cell = LTCCell(wiring) # Create LTC model
model = keras.Sequential(
[
keras.layers.InputLayer(input_shape=maxlen),
keras.layers.Embedding( input_dim = vocab_size,
output_dim = 300,
embeddings_initializer=keras.initializers.Constant(matrix),
trainable=True),
keras.layers.Dropout(0.1),
keras.layers.RNN(ltc_cell, return_sequences=False),
keras.layers.Dense(1, activation="sigmoid")
]
)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=0.0021),
loss="binary_crossentropy", metrics=["accuracy", tf.keras.metrics.AUC(name='auc')])
#create checkpoint to save model
#with best validation loss
PATH = 'drive/Shared drives/Magdas-PROJECT/data/save_weights/ncp_emb'
PATH2 = PATH + '/' + str(cnt)
!mkdir $PATH2
checkpoint_path = PATH2 + "/cnt/{epoch:04d}.ckpt"
if not os.path.isdir(PATH):
!mkdir $PATH
!mkdir $checkpoint_path
checkpoint_dir = os.path.dirname(checkpoint_path)
checkpoint = tf.keras.callbacks.ModelCheckpoint(checkpoint_path, monitor='val_auc', verbose=1,
save_weights_only=True, save_best_only=True, mode='max')
model.save_weights(checkpoint_path.format(epoch=0))
history = model.fit(
train_x, train_y,
batch_size=64, epochs=10,
validation_data=(val_x, val_y),
callbacks = [checkpoint]
)
latest = tf.train.latest_checkpoint(checkpoint_dir)
model.load_weights(latest)
train_eval = model.evaluate(train_x, train_y)
val_eval = model.evaluate(val_x, val_y)
test_eval = model.evaluate(test_x, test_y)
test_l.append(test_eval)
val_l.append(val_eval)
train_l.append(train_eval)
test_l = np.array(test_l)
val_l = np.array(val_l)
train_l = np.array(train_l)
print("test avg loss: ", np.mean(test_l[:, 0]), "+/-" ,np.std(test_l[:, 0]))
print("test avg acc: ", np.mean(test_l[:, 1]), "+/-" ,np.std(test_l[:, 1]))
print("test avg auc: ", np.mean(test_l[:, 2]), "+/-" ,np.std(test_l[:, 2]))
print('\n')
print("val avg loss: ", np.mean(val_l[:, 0]), "+/-" ,np.std(val_l[:, 0]))
print("val avg acc: ", np.mean(val_l[:, 1]), "+/-" ,np.std(val_l[:, 1]))
print("val avg auc: ", np.mean(val_l[:, 2]), "+/-" ,np.std(val_l[:, 2]))
print('\n')
print("train avg loss: ", np.mean(train_l[:, 0]), "+/-" ,np.std(train_l[:, 0]))
print("train avg acc: ", np.mean(train_l[:, 1]), "+/-" ,np.std(train_l[:, 1]))
print("train avg auc: ", np.mean(train_l[:, 2]), "+/-" ,np.std(train_l[:, 2]))
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
import pandas as pd
from sklearn.metrics import confusion_matrix
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 16}
matplotlib.rc('font', **font)
pred = np.array([model.predict(test_x) > 0.5], dtype=int).ravel()
cm = confusion_matrix(test_y.values, pred)
cm = pd.DataFrame(cm, range(2),range(2))
plt.figure(figsize = (10, 6))
sns.heatmap(cm,
fmt = 'd',
annot=True)
ax = plt.subplot()
ax.set_xlabel('Predicted')
ax.set_ylabel('Labels')
ax.xaxis.set_ticklabels(['non-toxic', 'toxic'])
ax.yaxis.set_ticklabels(['non-toxic', 'toxic'])
ax.set_title('NCP + PE Model')
plt.show()
sns.set_style("white")
plt.figure(figsize=(12, 12))
legend_handles = ltc_cell.draw_graph(layout='spiral',neuron_colors={"command": "tab:cyan"})
plt.legend(handles=legend_handles, loc="upper center", bbox_to_anchor=(1, 1))
sns.despine(left=True, bottom=True)
plt.tight_layout()
plt.show()
```
| github_jupyter |
# Seq2Seq Model with Attention Mechanism
this model is the basic sequence-to-sequence model with attention mechanism.
```
import numpy as np
import torch as th
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
from torch import optim
vocab_size = 256 # ascii size
x_ = list(map(ord, "hello")) # convert to list of ascii codes
y_ = list(map(ord, "hola")) # convert to list of ascii codes
print("hello -> ", x_)
print("hola -> ", y_)
x = Variable(th.LongTensor(x_))
y = Variable(th.LongTensor(y_))
class Seq2Seq(nn.Module):
def __init__(self, vocab_size, hidden_size):
super(Seq2Seq, self).__init__()
self.n_layers = 1
self.hidden_size = hidden_size
self.embedding = nn.Embedding(vocab_size, hidden_size)
self.encoder = nn.LSTM(hidden_size, hidden_size)
self.decoder = nn.LSTM(hidden_size, hidden_size)
self.project = nn.Linear(hidden_size, vocab_size)
self.attn = nn.Linear(2 * hidden_size, hidden_size)
def attention(self, M:"ixbxh", h_t:"bxh", i_size, b_size)->"[bxi,bxh]":
# bxixh * bxh -> attention [bxi]
attention = F.softmax(th.bmm(M.t(), h_t.unsqueeze(2)))
# bxixh * bxi -> context [bxh]
print("M:", M.size(), "att:", attention.size())
context = th.bmm(M.t().transpose(1, 2), attention)
return attention.squeeze(), context.squeeze()
def attention_func(self, encoder_outputs, decoder_hidden):
# (batch_size*input_len*hidden_len) * (batch_size*hidden_len) -> batch_size x seq_len
dot = torch.bmm(encoder_outputs.t(), decoder_hidden.unsqueeze(2))
attention = F.softmax(dot.squeeze())
# (batch_size*input_len*hidden_len)' * (batch_size*input_len) -> (batch_size*hidden_len)
enc_attention = torch.bmm(torch.transpose(torch.transpose(encoder_outputs, 0, 1), 1, 2), attention.unsqueeze(2))
# (batch_size*hidden_len)*(hidden_len*hidden_len) + (batch_size*hidden_len)*(hidden_len*hidden_len)
# -> (batch_size*hidden_len)
hidden = self.attn_tanh(self.attn_enc_linear(enc_attention.squeeze()) + self.attn_dec_linear(decoder_hidden))
return hidden
def attention(self, output_t, context):
# Output(t) = B x H x 1
# Context = B x T x H
# a = B x T x 1
a = torch.bmm(context, self.output_to_attn(output_t).unsqueeze(2))
a = F.softmax(a.squeeze(2)) # batch_size x seq_len
# Want to apply over context, scaled by a
# (B x 1 x T) (B x T x H) = (B x 1 x H)
a = a.view(a.size(0), 1, a.size(1))
combined = torch.bmm(a, context).squeeze(1)
combined = torch.cat([combined, output_t], 1)
combined = F.tanh(self.attn(combined))
return combined
def forward(self, inputs, targets):
# Encoder inputs and states
initial_state = self._init_state()
embedding = self.embedding(inputs).unsqueeze(1)
# embedding = [seq_len, batch_size, embedding_size]
# Encoder
encoder_output, encoder_state = self.encoder(embedding, initial_state)
# encoder_output = [seq_len, batch_size, hidden_size]
# encoder_state = [n_layers, seq_len, hidden_size]
# Decoder inputs and states
decoder_state = encoder_state
decoder_input = Variable(th.LongTensor([[0]]))
# Decoder
outputs = []
for i in range(targets.size()[0]):
decoder_input = self.embedding(decoder_input)
decoder_output, decoder_state = self.decoder(decoder_input, decoder_state)
attention, context = self.attention(encoder_output, decoder_output)
# Project to the vocabulary size
projection = self.project(decoder_output.view(1, -1)) # batch x vocab_size
# Make prediction
prediction = F.softmax(projection) # batch x vocab_size
outputs.append(prediction)
# update decoder input
_, top_i = prediction.data.topk(1) # 1 x 1
decoder_input = Variable(top_i)
outputs = th.stack(outputs).squeeze()
return outputs
def _init_state(self, batch_size=1):
weight = next(self.parameters()).data
return (
Variable(weight.new(self.n_layers, batch_size, self.hidden_size).zero_()),
Variable(weight.new(self.n_layers, batch_size, self.hidden_size).zero_())
)
seq2seq = Seq2Seq(vocab_size, 16)
print(seq2seq)
pred = seq2seq(x, y)
print(pred)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(seq2seq.parameters(), lr=1e-3)
log = []
for i in range(1000):
prediction = seq2seq(x, y)
loss = criterion(prediction, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
loss_val = loss.data[0]
log.append(loss_val)
if i % 100 == 0:
print("%d loss: %s" % (i, loss_val))
_, top1 = prediction.data.topk(1, 1)
for c in top1.squeeze().numpy().tolist():
print(chr(c), end=" ")
print()
import matplotlib.pyplot as plt
plt.plot(log)
plt.ylabel('cross entropy loss')
plt.show()
```
| github_jupyter |
## 1. The dataset
<p>Walt Disney Studios is the foundation on which The Walt Disney Company was built. The Studios has produced more than 600 films since their debut film, Snow White and the Seven Dwarfs in 1937. While many of its films were big hits, some of them were not. In this notebook, we will explore a dataset of Disney movies and analyze what contributes to the success of Disney movies.</p>
<p><img src="https://assets.datacamp.com/production/project_740/img/jorge-martinez-instagram-jmartinezz9-431078-unsplash_edited.jpg" alt=""></p>
<p>First, we will take a look at the Disney data compiled by <a href="https://data.world/kgarrett/disney-character-success-00-16">Kelly Garrett</a>. The data contains 579 Disney movies with six features: movie title, release date, genre, MPAA rating, total gross, and inflation-adjusted gross. </p>
<p>Let's load the file and see what the data looks like.</p>
```
# Import pandas library
import pandas as pd
# Read the file into gross
gross = pd.read_csv("datasets/disney_movies_total_gross.csv",
parse_dates=["release_date"])
# Print out gross
gross.head()
```
## 2. Top ten movies at the box office
<p>Let's started by exploring the data. We will check which are the 10 Disney movies that have earned the most at the box office. We can do this by sorting movies by their inflation-adjusted gross (we will call it adjusted gross from this point onward). </p>
```
# Sort data by the adjusted gross in descending order
inflation_adjusted_gross_desc = gross.sort_values(by='inflation_adjusted_gross',
ascending=False)
# Display the top 10 movies
inflation_adjusted_gross_desc.head(10)
```
## 3. Movie genre trend
<p>From the top 10 movies above, it seems that some genres are more popular than others. So, we will check which genres are growing stronger in popularity. To do this, we will group movies by genre and then by year to see the adjusted gross of each genre in each year.</p>
```
# Extract year from release_date and store it in a new column
gross['release_year'] = pd.DatetimeIndex(gross["release_date"]).year
# Compute mean of adjusted gross per genre and per year
group = gross.groupby(['genre','release_year']).mean()
# Convert the GroupBy object to a DataFrame
genre_yearly = group.reset_index()
# Inspect genre_yearly
genre_yearly.head(10)
```
## 4. Visualize the genre popularity trend
<p>We will make a plot out of these means of groups to better see how box office revenues have changed over time.</p>
```
# Import seaborn library
import seaborn as sns
# Plot the data
sns.relplot(x='release_year', y='inflation_adjusted_gross', kind='line',
hue='genre',
data=genre_yearly)
```
## 5. Data transformation
<p>The line plot supports our belief that some genres are growing faster in popularity than others. For Disney movies, Action and Adventure genres are growing the fastest. Next, we will build a linear regression model to understand the relationship between genre and box office gross. </p>
<p>Since linear regression requires numerical variables and the genre variable is a categorical variable, we'll use a technique called one-hot encoding to convert the categorical variables to numerical. This technique transforms each category value into a new column and assigns a 1 or 0 to the column. </p>
<p>For this dataset, there will be 11 dummy variables, one for each genre except the action genre which we will use as a baseline. For example, if a movie is an adventure movie, like The Lion King, the adventure variable will be 1 and other dummy variables will be 0. Since the action genre is our baseline, if a movie is an action movie, such as The Avengers, all dummy variables will be 0.</p>
```
# Convert genre variable to dummy variables
genre_dummies = pd.get_dummies(data=gross['genre'], drop_first=True)
# Inspect genre_dummies
genre_dummies.head()
```
## 6. The genre effect
<p>Now that we have dummy variables, we can build a linear regression model to predict the adjusted gross using these dummy variables.</p>
<p>From the regression model, we can check the effect of each genre by looking at its coefficient given in units of box office gross dollars. We will focus on the impact of action and adventure genres here. (Note that the intercept and the first coefficient values represent the effect of action and adventure genres respectively). We expect that movies like the Lion King or Star Wars would perform better for box office.</p>
```
# Import LinearRegression
from sklearn.linear_model import LinearRegression
# Build a linear regression model
regr = LinearRegression()
# Fit regr to the dataset
regr.fit(genre_dummies, gross["inflation_adjusted_gross"])
# Get estimated intercept and coefficient values
action = regr.intercept_
adventure = regr.coef_[[0]][0]
# Inspect the estimated intercept and coefficient values
print((action, adventure))
```
## 7. Confidence intervals for regression parameters (i)
<p>Next, we will compute 95% confidence intervals for the intercept and coefficients. The 95% confidence intervals for the intercept <b><i>a</i></b> and coefficient <b><i>b<sub>i</sub></i></b> means that the intervals have a probability of 95% to contain the true value <b><i>a</i></b> and coefficient <b><i>b<sub>i</sub></i></b> respectively. If there is a significant relationship between a given genre and the adjusted gross, the confidence interval of its coefficient should exclude 0. </p>
<p>We will calculate the confidence intervals using the pairs bootstrap method. </p>
```
# Import a module
import numpy as np
# Create an array of indices to sample from
inds = np.arange(len(gross['genre']))
# Initialize 500 replicate arrays
size = 500
bs_action_reps = np.empty(size)
bs_adventure_reps = np.empty(size)
```
## 8. Confidence intervals for regression parameters (ii)
<p>After the initialization, we will perform pair bootstrap estimates for the regression parameters. Note that we will draw a sample from a set of (genre, adjusted gross) data where the genre is the original genre variable. We will perform one-hot encoding after that. </p>
```
# Generate replicates
for i in range(size):
# Resample the indices
bs_inds = np.random.choice(inds, size=len(inds))
# Get the sampled genre and sampled adjusted gross
bs_genre = gross['genre'][bs_inds]
bs_gross = gross['inflation_adjusted_gross'][bs_inds]
# Convert sampled genre to dummy variables
bs_dummies = pd.get_dummies(data=gross['genre'] , drop_first=True)
# Build and fit a regression model
regr = LinearRegression().fit(bs_dummies, bs_gross)
# Compute replicates of estimated intercept and coefficient
bs_action_reps[i] = regr.intercept_
bs_adventure_reps[i] = regr.coef_[[0]][0]
```
## 9. Confidence intervals for regression parameters (iii)
<p>Finally, we compute 95% confidence intervals for the intercept and coefficient and examine if they exclude 0. If one of them (or both) does, then it is unlikely that the value is 0 and we can conclude that there is a significant relationship between that genre and the adjusted gross. </p>
```
# Compute 95% confidence intervals for intercept and coefficient values
confidence_interval_action = np.percentile(bs_action_reps, [2.5, 97.5])
confidence_interval_adventure = np.percentile(bs_adventure_reps, [2.5, 97.5])
# Inspect the confidence intervals
print(confidence_interval_action)
print(confidence_interval_adventure)
```
## 10. Should Disney make more action and adventure movies?
<p>The confidence intervals from the bootstrap method for the intercept and coefficient do not contain the value zero, as we have already seen that lower and upper bounds of both confidence intervals are positive. These tell us that it is likely that the adjusted gross is significantly correlated with the action and adventure genres. </p>
<p>From the results of the bootstrap analysis and the trend plot we have done earlier, we could say that Disney movies with plots that fit into the action and adventure genre, according to our data, tend to do better in terms of adjusted gross than other genres. So we could expect more Marvel, Star Wars, and live-action movies in the upcoming years!</p>
```
# should Disney studios make more action and adventure movies?
more_action_adventure_movies = True
```
| github_jupyter |
## An introduction to working with the Cadence Simulations for the AGN SC
### Initial Setup
This notebook provides a crash-course to the "official" crash-course overview. The projects' crash-course notebook will be in the following place (once you follow the directions below):
/home/mount/sims_maf_contrib/tutorials/Introduction Notebook.ipynb
See also
/home/mount/sims_maf_contrib/tutorials/Write a New Metric.ipynb
and further tutorials for more in-depth examples.
To use this notebook, you need to do the following:
- install docker (from https://www.docker.com/products/docker-desktop)
- create a folder to work in (e.g., /Users/gtr/Dropbox/LSST/DockerShared)
- in that folder, on a command line execute: `docker pull wyu16/lsst_sims:j_maf_19_11_05`
- then execute: `docker run -d -p 9000:8888 --name maf -v /local/path/:/home/mount/ wyu16/lsst_sims:j_maf_19_11_05`, where for me `/local/path/` is `/Users/gtr/Dropbox/LSST/DockerShared/`
- now open a browser (chrome ideally) and go to https://github.com/LSST-nonproject/sims_maf_contrib
- again in the folder from step 2, execute: `git clone https://github.com/LSST-nonproject/sims_maf_contrib`
- make a subfolder to store your opSim db files (e.g., /Users/gtr/Dropbox/LSST/DockerShared/dbFiles)
- go to http://astro-lsst-01.astro.washington.edu:8081/?runId=67
- download 2 "db" files to compare (and put them in the folder above)
- open jupyter lab in your browser with `http://localhost:9000/`
### Returning Setup
If you have already done the above and are coming back to this later (e.g., after shutting down your computer), then instead of executing the `docker run` command above, restart Docker with
- `docker start maf`
then
- open jupyter lab in your browser with `http://localhost:9000/`
<b>Now we can start computing metrics to use for comparing different opSims.</b>
### Software Setup (repeat each session)
```
# import matplotlib to show plots inline.
%matplotlib inline
import matplotlib.pyplot as plt
```
Import the sims_maf modules needed.
```
# import our python modules
import lsst.sims.maf.db as db
import lsst.sims.maf.metrics as metrics
import lsst.sims.maf.slicers as slicers
import lsst.sims.maf.stackers as stackers
import lsst.sims.maf.plots as plots
import lsst.sims.maf.metricBundles as metricBundles
```
### Metrics for beginners
We'd like to compare 2 or more opSims to determine which is "best" for our science.
In order to do this, we are first going to need to define a `metric`:
- a `metric` calculates some quantity that you want to measure (e.g. the mean value of an attribute such as the airmass).
Then we need to know how you want to split the data by defining a `slicer`:
- a `slicer` takes the list of visits from a simulated survey and orders or groups them (e.g. by night or by region on the sky).
Finally, we need to provide a `constraint`:
- a constraint applies a SQL selection to the visits you want to select from the simulated survey database (such as 'filter = "r"')
<br>
The combination of a <font color='green'> metric </font>, <font color='red'> slicer </font>, and <font color='blue'> constraint </font> is referred to as a `MetricBundle`.
Here are examples of some simple `MetricBundle`s:
- <font color='green'>coadded depth</font> <font color='red'> at each point in the sky</font> <font color='blue'> in r band</font> == 'Coaddm5Metric' + 'HealpixSlicer' + 'filter = "r"'
- <font color='green'>median airmass</font> <font color='red'>at each point in the sky</font> <font color='blue'> in i band </font> == 'MedianMetric(col="airmass")' + 'HealpixSlicer' + 'filter = "i"'
- <font color='green'>number of visits</font> <font color='red'>in each night</font> == 'CountMetric()'</font> + 'OneDSlicer(col='night')' + None
- <font color='green'>median skybrightness</font> <font color='red'>of all visits</font> <font color='blue'> in u band </font> = 'MedianMetric(col='skybrightness')' + 'UniSlicer' + 'filter = "u"'
### Example MetricBundle Code
The following code segment generates the inputs to a `MetricBundle` that will calculate the <font color='green'>maximum airmass value</font> <font color='red'> at each point of the sky</font> <font color='blue'>in the $g$ band</font>. We are also going to ignore the Deep Drilling Fields (DDFs) in this example.
```
# metric = the "maximum" of the "airmass" for each group of visits in the slicer
metric1 = metrics.MaxMetric('airmass')
# slicer = a grouping or subdivision of visits for the simulated survey
# based on their position on the sky (using a Healpix grid)
slicer1 = slicers.HealpixSlicer(nside=64)
# constraint = the sql query (or 'select') that selects all visits in r band
constraint1 = 'filter = "g"'
constraint1 += ' and note not like "DD%"' # added so the sky plot won't saturate (remove DDFs)
```
### Define a MetricBundle
Next we define the metric bundle itself, which includes the `metric`, `slicer`, and `constraints`.
```
# MetricBundle = combination of the metric, slicer, and sqlconstraint
maxairmassSky = metricBundles.MetricBundle(metric1, slicer1, constraint1)
```
### Combining Metrics
We can also combine multiple MetricBundles and run them all at once to calculate their metric values. We do this by combining the metrics into a dictionary, creating a `MetricBundleGroup`. For now we will use calculate the metric values for our single 'maxairmassSky' MetricBundle.
```
bundleDict = {'maxairmassSky': maxairmassSky}
```
### Choosing the simulated survey database (opSim) to analyze
The input data is queried from a database (usually a SQLite database).
```
#Read in an opSim (in this case "baseline_v1.3_10yrs.db")
opsdb1 = db.OpsimDatabase('/home/mount/dbFiles/baseline_v1.3_10yrs.db')
```
Create an output folder, 'outDir', that will track the results in another database ('resultsDb_sqlite.db').
```
outDir = '/home/mount/dbFiles/MAF_output'
resultsDb = db.ResultsDb(outDir=outDir)
```
### Running the Analysis
We generate the outputs by combining the bundle dictionary with the input database and output directories and database, then using the `MetricBundleGroup` class.
The MetricBundleGroup will query the data from the opsim database and calculate the metric values, using the 'runAll' method.
```
group = metricBundles.MetricBundleGroup(bundleDict, opsdb1, outDir=outDir, resultsDb=resultsDb)
group.runAll()
```
### Visualizing the output
We can now visualize the results of running these metrics. With a HealpixSlicer, the default is to generate 3 plots:
- the metric as a function of position on the sky (in a Healpix projection)
- a histogram of the metric distribution for the pixels in the Healpix projection
- the angular powers pectrum of the metric
Where in this example, the metric is the maximum airmass.
```
group.plotAll(closefigs=False)
```
---
### Extending the analysis: Adding more metrics, more information, and more opSims
Our goal is to answer the question of which opSim is "best" for any given science. Or perhaps simply which of two opSim is the "better" of the two.
We'll now illustrate two metrics specifically related to two AGN goals:
- Selection of $z>7$ quasars
- Using DCR to improve photo-z estimation
We could define fairly sophisticated metrics to estimate how well we'll do for each, but we'll start with simple proxies:
- Coadded y-band depth
- Maximum g-band airmass
We have already created the `MetricBundle` for the 2nd metric. Now we'll define one (`coAddY') for the 1st using the same slicer. To do this we use the 'Coaddm5Metric' and new sql constraint 'filter = "y"'.
```
metric2 = metrics.Coaddm5Metric()
constraint2 = 'filter = "y"'
constraint2 += ' and note not like "DD%"' # added so the sky plot won't saturate (remove DDFs)
coAddY = metricBundles.MetricBundle(metric2, slicer1, constraint2)
```
We can also add ["summary metrics"](https://confluence.lsstcorp.org/display/SIM/MAF+Summary+Statistics) to each MetricBundle. These metrics generate statistical summaries of the metric data values (e.g. the means of the number of visits per point on the sky).
```
summaryMetrics = [metrics.MinMetric(), metrics.MedianMetric(),
metrics.MaxMetric(), metrics.RmsMetric()]
maxairmassSky.setSummaryMetrics(summaryMetrics)
coAddY.setSummaryMetrics(summaryMetrics)
```
### Grouping MetricBundles together
We can group these metricBundles together into a dictionary, and pass this to the MetricBundleGroup, which will run them together.
```
bundleDict = {'maxairmassSky': maxairmassSky, 'coAddY': coAddY}
```
### Define Plots to Create
Now we'll define which plots are going to be produced in our analysis. In this case, we'll create both a histogram of our `metric` and a sky map (that essentially shows our `slicer`), but we'll leave off the angular power spectrum.
```
# plots to generate for this specific metric bundle
healpixhist = plots.HealpixHistogram()
healpixSky = plots.HealpixSkyMap()
plot_funcs = [healpixSky, healpixhist]
plot_dicts = {'left':1, 'right':2.8}
```
### Redefine MetricBundles and BundleDicts
To include only the above plots and adding a label for each opSim that will be useful for the comparison plots below.
```
maxairmassSky1 = metricBundles.MetricBundle(metric1, slicer1, constraint1, \
plotFuncs=plot_funcs, plotDict=plot_dicts, \
runName='baseline_v1.3_10yrs')
coAddY1 = metricBundles.MetricBundle(metric2, slicer1, constraint2, \
plotFuncs=plot_funcs, plotDict=plot_dicts, \
runName='baseline_v1.3_10yrs')
maxairmassSky1.setSummaryMetrics(summaryMetrics)
coAddY1.setSummaryMetrics(summaryMetrics)
bundleDict1 = {'maxairmassSky': maxairmassSky1, 'coAddY': coAddY1}
```
### Define MetricBundleGroup for these two MetricBundles and run our on opSim of choice
```
group1 = metricBundles.MetricBundleGroup(bundleDict1, opsdb1, \
outDir=outDir, resultsDb=resultsDb)
group1.runAll()
group1.plotAll(closefigs=False, savefig=False)
```
### Analyze a 2nd opSim
Now we'll analyze a second opSim, specifically `dcr_nham3_v1.3_10yrs.db`, which was designed to test possible changes to the cadence that would benefit DCR analysis.
```
# init new db connection
opsdb2 = db.OpsimDatabase('/home/mount/dbFiles/dcr_nham3_v1.3_10yrs.db')
maxairmassSky2 = metricBundles.MetricBundle(metric1, slicer1, constraint1, \
plotFuncs=plot_funcs, plotDict=plot_dicts, \
runName='dcr_nham3_v1.3_10yrs')
coAddY2 = metricBundles.MetricBundle(metric2, slicer1, constraint2, \
plotFuncs=plot_funcs, plotDict=plot_dicts, \
runName='dcr_nham3_v1.3_10yrs')
maxairmassSky2.setSummaryMetrics(summaryMetrics)
coAddY2.setSummaryMetrics(summaryMetrics)
bundleDict2 = {'maxairmassSky': maxairmassSky2, 'coAddY': coAddY2}
group2 = metricBundles.MetricBundleGroup(bundleDict2, opsdb2, outDir=outDir, resultsDb=resultsDb)
group2.runAll()
```
### Compare Metrics for Both opSims
Let's plot a histogram of each metric with the two opSim results overplotted.
```
# init handler
ph = plots.PlotHandler(outDir=outDir, resultsDb=resultsDb)
# set metrics to plot togehter
ph.setMetricBundles([group1.bundleDict['maxairmassSky'], group2.bundleDict['maxairmassSky']])
plotDicts = [{'label':'baseline_v1.3_10yr', 'color':'b'}, \
{'label':'dcr_nham2_v1.3_10yr', 'color':'r'}]
ph.plot(plotFunc = healpixhist, plotDicts=plotDicts)
# reset the metrics to plot togehter
ph.setMetricBundles([group1.bundleDict['coAddY'], group2.bundleDict['coAddY']])
ph.plot(plotFunc = healpixhist, plotDicts=plotDicts)
```
So, you can see that there is a pretty big difference between these opSims in terms of the DCR photo-z metric, but not much difference in terms of the $z>7$ metric. We'd need to check the other opSims to see how they compare in that regard.
We can similarly create the sky plots (not overplotted clearly, but at least plotted together).
```
metricbdList = [group1.bundleDict['maxairmassSky'], group2.bundleDict['maxairmassSky'],\
group1.bundleDict['coAddY'], group2.bundleDict['coAddY']]
for metricbd in metricbdList:
metricbd.plot(plotFunc=healpixSky)
```
---
### Accessing the data generated by a metric
The results of the metric calculation are stored as an attribute in each metricBundle, as 'metricValues' - a numpy masked array. (**The structure of 'metricValues' will depend on how the data is sliced, e.g., if 'Healpix' slicer is used, each value represent the evaluate metric in each healpix pixel, or if you slice the cadence by night, then each value in the array will be the metric evaluated per night. So care should be taken when trying to generate plots directly from data in 'metricValues'**). The results of the summary statistics are also stored in each metricBundle, in an attribute called 'summaryValues', which is a dictionary of the summary metric name and value.
For maxairmassSky, the metricValues are an array of max airmass (in g band) on the sky for each healpix pixel and the summaryValues are the mean, median etc of the these values.
```
print("Array with the max airmass per pixel:", maxairmassSky.metricValues)
```
N.B. The values of some metricValues above are shown as '--' because there were no opsim visits for those healpix points.
### Show summary statstics
Note: Since we redefined the metricbundles before adding in the summary statistics, we need to access summary statistics from the 'maxairmassSky1' metric bundle.
```
print("Summary of the max, median, min,",
"and rms of max airmass per pixel",
maxairmassSky1.summaryValues)
```
### Making your own plots
We can now generate plots directly using the data from 'metricValues' (i.e., instead of using the built-in tools above.)
```
# need to mask the pixels that have no available data
mask = maxairmassSky.metricValues.mask
data = maxairmassSky.metricValues.data[~mask]
# plot
fig = plt.figure(figsize=(10,6))
_ = plt.hist(data, bins=100, histtype='step', density=True, label='opSim: baseline_v1.3_10yrs')
plt.title('MaxAirmass metric sliced by a healpix slicer', fontsize=15)
plt.xlabel('Max airmass', fontsize=15)
plt.ylabel('Normalized Density', fontsize=15)
plt.legend(fontsize=15)
```
If you have gotten to this point, you are then ready to try creating more complicated metrics, analyzing more opSims, etc.!
As noted above, see
/home/mount/sims_maf_contrib/tutorials/Introduction Notebook.ipynb
and
/home/mount/sims_maf_contrib/tutorials/Write a New Metric.ipynb
for further tutorials for more in-depth examples.
| github_jupyter |
```
import numpy as np
import sys; sys.path.insert(0, '../')
from gaia_tools.query import make_query, make_simple_query
import warnings; warnings.filterwarnings('ignore')
```
# Simple Query
This is a simple single-level query.
## Default Columns
```
circle = """
--Selections: Cluster RA
1=CONTAINS(POINT('ICRS',gaia.ra,gaia.dec),
CIRCLE('ICRS',{ra:.4f},{dec:.4f},{rad:.2f}))
""".format(ra=230, dec=0, rad=4)
df = make_simple_query(
WHERE=circle, # The WHERE part of the SQL
random_index=1e4, # a shortcut to use the random_index in 'WHERE'
ORDERBY='gaia.parallax', # setting the data ordering
pprint=True, # print the query
do_query=True, # perform the query using gaia_tools.query
local=False, # whether to perform the query locally
units=True # to fill in missing units from 'defaults' file
)
df
```
The system also supports Pan-STARRS1 and 2MASS cross-matches using the `panstarrs1` and `twomass` keywords
```
df = make_simple_query(
WHERE=circle, # The WHERE part of the SQL
random_index=1e4, # a shortcut to use the random_index in 'WHERE'
ORDERBY='gaia.parallax', # setting the data ordering
panstarrs1=True, twomass=True,
do_query=True, # perform the query using gaia_tools.query
local=False, # whether to perform the query locally
units=True # to fill in missing units from 'defaults' file
)
df
```
## Different Defaults
If you want fewer default columns, this is an option through the `defaults` keyword.
```
df = make_simple_query(
WHERE=circle, random_index=1e4, ORDERBY='gaia.parallax',
do_query=True, local=False, units=True,
defaults='empty',
)
df
```
Likewise, there's an option for much greater detail.
```
df = make_simple_query(
WHERE=circle, random_index=1e4, ORDERBY='gaia.parallax',
do_query=True, local=False, units=True,
defaults='full'
)
df
```
<br><br><br><br>
- - -
- - -
<br><br><br><br>
# Complex Nested Query
A complex query like this one shows the real utility of this package.
Instead of keeping track of the complex SQL, we only need to pay close attention to the custom calculated columns.
This ADQL queries for data within a rectangular area on a sky rotated by a rotation matrix and specified North Galactic Pole angles. The specifics aren't important -- the real takeaway is that the sky rotation and calculation are written in a clear format, with all the parts of the query close together. Running the query is trivial after that.
```
###########
# Custom Calculations
# Innermost Level
l0cols = """
--Rotation Matrix
{K00}*cos(radians(dec))*cos(radians(ra))+
{K01}*cos(radians(dec))*sin(radians(ra))+
{K02}*sin(radians(dec)) AS cosphi1cosphi2,
{K10}*cos(radians(dec))*cos(radians(ra))+
{K11}*cos(radians(dec))*sin(radians(ra))+
{K12}*sin(radians(dec)) AS sinphi1cosphi2,
{K20}*cos(radians(dec))*cos(radians(ra))+
{K21}*cos(radians(dec))*sin(radians(ra))+
{K22}*sin(radians(dec)) AS sinphi2,
--c1, c2
{sindecngp}*cos(radians(dec)){mcosdecngp:+}*sin(radians(dec))*cos(radians(ra{mrangp:+})) as c1,
{cosdecngp}*sin(radians(ra{mrangp:+})) as c2
"""
# Inner Level
l1cols = """
gaia.cosphi1cosphi2, gaia.sinphi1cosphi2, gaia.sinphi2,
gaia.c1, gaia.c2,
atan2(sinphi1cosphi2, cosphi1cosphi2) AS phi1,
atan2(sinphi2, sinphi1cosphi2 / sin(atan2(sinphi1cosphi2, cosphi1cosphi2))) AS phi2"""
# Inner Level
l2cols = """
gaia.sinphi1cosphi2, gaia.cosphi1cosphi2, gaia.sinphi2,
gaia.phi1, gaia.phi2,
gaia.c1, gaia.c2,
( c1*pmra+c2*pmdec)/cos(phi2) AS pmphi1,
(-c2*pmra+c1*pmdec)/cos(phi2) AS pmphi2"""
# Outer Level
l3cols = """
gaia.phi1, gaia.phi2,
gaia.pmphi1, gaia.pmphi2"""
###########
# Custom Selection
l3sel = """
phi1 > {phi1min:+}
AND phi1 < {phi1max:+}
AND phi2 > {phi2min:+}
AND phi2 < {phi2max:+}
"""
###########
# Custom substitutions
l3userasdict = {
'K00': .656, 'K01': .755, 'K02': .002,
'K10': .701, 'K11': .469, 'K12': .537,
'K20': .53, 'K21': .458, 'K22': .713,
'sindecngp': -0.925, 'cosdecngp': .382, 'mcosdecngp': -.382,
'mrangp': -0,
'phi1min': -0.175, 'phi1max': 0.175,
'phi2min': -0.175, 'phi2max': 0.175}
###########
# Making Query
df = make_query(
gaia_mags=True,
panstarrs1=True, # doing a Pan-STARRS1 crossmatch
user_cols=l3cols,
use_AS=True, user_ASdict=l3userasdict,
# Inner Query
FROM=make_query(
gaia_mags=True,
user_cols=l2cols,
# Inner Query
FROM=make_query(
gaia_mags=True,
user_cols=l1cols,
# Innermost Query
FROM=make_query(
gaia_mags=True,
inmostquery=True, # telling system this is the innermost level
user_cols=l0cols,
random_index=1e4 # quickly specifying random index
)
)
),
WHERE=l3sel,
ORDERBY='gaia.source_id',
pprint=True,
# doing query
do_query=True, local=False, units=True
)
df
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# TensorFlow Addons Losses: TripletSemiHardLoss
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/addons/blob/master/docs/tutorials/losses_triplet.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/addons/blob/master/docs/tutorials/losses_triplet.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
# Overview
This notebook will demonstrate how to use the TripletSemiHardLoss function in TensorFlow Addons.
## Resources:
* [FaceNet: A Unified Embedding for Face Recognition and Clustering](https://arxiv.org/pdf/1503.03832.pdf)
* [Oliver Moindrot's blog does an excellent job of describing the alogrithm in detail](https://omoindrot.github.io/triplet-loss)
# TripletLoss
As first introduced in in the FaceNet paper, TripletLoss is a loss function that trains a neural network to closely embedd features of the same class while maximizing the distance between embeddings of different classes. To do this an anchor is chosen along with one negative and one postive sample.

**The loss function is described as a Euclidean distance function:**

Where A is our anchor input, P is the positive sample input, N is the negative sample input, and alpha is some margin we use to specify when a triplet has become too "easy" and we no longer want to adjust the weights from it.
# SemiHard Online Learning
As shown in the paper, the best results are from triplets known as "Semi-Hard". These are defined as triplets where the negative is farther from the anchor than the positive, but still produces a positive loss.To efficiently find these triplets we utilize online learning and only train from the Semi-Hard examples in each batch.
# Setup
```
!pip install -q tensorflow-gpu==2.0.0rc0
!pip install -q tensorflow-addons~=0.5
from __future__ import absolute_import, division, print_function, unicode_literals
import io
import numpy as np
import tensorflow as tf
import tensorflow_addons as tfa
import tensorflow_datasets as tfds
from matplotlib import pyplot as plt
from google.colab import files
```
# Prep the Data
```
def _normalize_img(img, label):
img = tf.cast(img, tf.float32) / 255.
return (img, label)
train_dataset, test_dataset = tfds.load(name="mnist", split=['train', 'test'], as_supervised=True)
# Build your input pipelines
train_dataset = train_dataset.shuffle(1024).batch(32)
train_dataset = train_dataset.map(_normalize_img)
test_dataset = test_dataset.batch(32)
test_dataset = test_dataset.map(_normalize_img)
```
# Build the Model
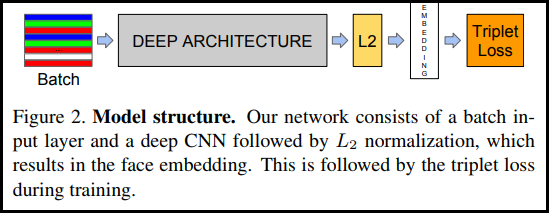
```
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(filters=64, kernel_size=2, padding='same', activation='relu', input_shape=(28,28,1)),
tf.keras.layers.MaxPooling2D(pool_size=2),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'),
tf.keras.layers.MaxPooling2D(pool_size=2),
tf.keras.layers.Dropout(0.3),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(256, activation=None), # No activation on final dense layer
tf.keras.layers.Lambda(lambda x: tf.math.l2_normalize(x, axis=1)) # L2 normalize embeddings
])
```
# Train and Evaluate
```
# Compile the model
model.compile(
optimizer=tf.keras.optimizers.Adam(0.001),
loss=tfa.losses.TripletSemiHardLoss())
# Train the network
history = model.fit(
train_dataset,
epochs=5)
# Evaluate the network
results = model.predict(test_dataset)
# Save test embeddings for visualization in projector
np.savetxt("vecs.tsv", results, delimiter='\t')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for img, labels in tfds.as_numpy(test_dataset):
[out_m.write(str(x) + "\n") for x in labels]
out_m.close()
files.download('vecs.tsv')
files.download('meta.tsv')
```
# Embedding Projector
The vector and metadata files can be loaded and visualized here: https://projector.tensorflow.org/
Youu can see the results of our embedded test data when visualized with UMAP:

```
```
| github_jupyter |
Chris Bayly:
>That is one weird functional group... a carbamate (common) but connected to an imide! A very highly toxic pesticide, basically a nerve agent, which is my excuse for being less familiar with it. Which is the problematic torsion? In the Google doc text it says "Manual inspection suggests that this transition involves a N-C-O-N torsion, but this needs to be looked at more carefully." The central bond to the N-C-O-N torsion is a single bond, but one which could have somewhat hindered rotation as esters do. Strangely connected to an imide means I do not have a good intuition for the potential barrier. The other candidate hindered rotors are the C-O-N-C (what smirnoff torsion is applied?) and obviously the O-N-C-C (which is a formal double bond).
The key issue to the sampling time must be that higher barriers take longer to sample, so any highly-delocalized acyclic system could be an issue, or sterically-hindered rotors such as 2,6-disubstituted biphenyls. Perhaps one could use some cheminformatics tricks to see when a chemical graph contains hindered rotors of either kind and then do some sampling tricks to address it. Or adaptive sampling without any cheminformatics?
Me:
>All of the following torsion angles were nearly perfectly correlated with the slowest relaxation process:
```
O1 - C2 - N2 - H14
[9, 1, 8, 25]
periodicity = 2
phase = 0.0 rad
force constant = 2.5 kcal/mol
C5 - N2 - C2 - O1
[4, 8, 1, 9]
periodicity = 2
phase = 0.0 rad
force constant = 0.625 kcal/mol
C5 - N2 - C2 - O2
[4, 8, 1, 10]
periodicity = 2
phase = 0.0 rad
force constant = 0.625 kcal/mol
N1 - O2 - C2 - N2
[7, 10, 1, 8]
periodicity = 2
phase = 0.0 rad
force constant = 2.7 kcal/mol
N1 - O2 - C2 - O1
[7, 10, 1, 9]
periodicity = 2
phase = 0.0 rad
force constant = 1.05 kcal/mol```
Chris Bayly:
>Looking at the snapshots says it's all about the carbamate... the "amide" side and the "ester" side of the carbonyl each show transitions. Would this be normative amongst structures containing linear unhindered carbamates: R1-[CX2][NH]C(=O)O[CX2]-R2? Do you have other examples containing this which are well-behaved?
David Mobley:
>the torsion for your carbamate — have you checked if this behaves similarly in GAFF? It would be helpful to know if it’s a problem with the torsion in general or if we’ve botched something in smirnoff99Frosst.
```
from bayes_implicit_solvent.solvation_free_energy import smiles_list, db, mol_top_sys_pos_list
linear_unhindered_carbamate_smarts = "[CX2][NH]C(=O)O[CX2]"
from openeye import oechem
def look_for_matches(mol, smarts):
mol_ = oechem.OEMol(mol)
# Set up query.
qmol = oechem.OEQMol()
oechem.OEParseSmarts(qmol, smarts)
unique = False
ss = oechem.OESubSearch(qmol)
matches = []
for match in ss.Match( mol_, unique):
# Compile list of atom indices that match the pattern tags
atom_indices = dict()
for ma in match.GetAtoms():
if ma.pattern.GetMapIdx() != 0:
atom_indices[ma.pattern.GetMapIdx()-1] = ma.target.GetIdx()
# Compress into list
atom_indices = [ atom_indices[index] for index in range(len(atom_indices)) ]
# Store
matches.append( tuple(atom_indices) )
return matches
all_linear_unhindered_carbamate_matches = []
for i in range(len(mol_top_sys_pos_list)):
mol, _, _, _ = mol_top_sys_pos_list[i]
all_linear_unhindered_carbamate_matches.append(look_for_matches(mol, linear_unhindered_carbamate_smarts))
sum([len(m) > 0 for m in all_linear_unhindered_carbamate_matches])
from bayes_implicit_solvent.solvation_free_energy import db, mol_top_sys_pos_list, smiles_list
mol, top, sys, pos = mol_top_sys_pos_list[0]
mol.IsValid()
[c for c in mol.GetConfs()]
len(pos)
```
[ ] TODO: Set up aldicarb simulation using GAFF
```
fname = '../../../MSKCC/Chodera Lab/feedstock/FreeSolv-0.51/mol2files_gaff/mobley_5200358.mol2'
from openeye import oeomega # Omega toolkit
omega = oeomega.OEOmega()
from openeye import oechem # OpenEye Python toolkits
from openeye import oequacpac
ifs = oechem.oemolistream()
if ifs.open(fname):
for mol_ in ifs.GetOEGraphMols():
mol = oechem.OEMol(mol_)
atoms = [a for a in mol.GetAtomIter()]
a = atoms[0]
[a.GetPartialCharge() for a in atoms]
a.GetSymmetryClass()
len(atoms)
mol
import numpy as np
coord_dict = mol.GetCoords()
positions = np.array([coord_dict[key] for key in coord_dict])
topology = generateTopologyFromOEMol(mol)
residue = list(topology.residues())[0]
len(positions)
residue
len(list(residue.atoms()))
from simtk import openmm as mm
from simtk.openmm import app
from openmoltools import forcefield_generators
forcefield = app.ForceField()
forcefield.registerTemplateGenerator(forcefield_generators.gaffTemplateGenerator)
system_generator = forcefield_generators.SystemGenerator([])
system_generator.createSystem(topology)
mol,_,_,_ = mol_top_sys_pos_list[0]
template, additional_parameters_ffxml = forcefield_generators.generateResidueTemplate(mol, gaff_version='gaff2')
forcefield.registerResidueTemplate(template)
template
system = forcefield.createSystem(topology)
import parmed
pmol = parmed.load_file(fname)
type(pmol)
pmol.coordinates
pmol.to_structure()
from openmoltools import utils
from simtk.openmm.app import ForceField
molecules = [ mol ]
from openmoltools.forcefield_generators import generateForceFieldFromMolecules
ffxml = generateForceFieldFromMolecules(molecules, gaff_version='gaff2')
gaff_xml_filename = utils.get_data_filename("parameters/gaff2.xml")
forcefield = ForceField(gaff_xml_filename)
# Parameterize the molecules.
from openmoltools.forcefield_generators import generateTopologyFromOEMol
for molecule in molecules:
# Create topology from molecule.
topology = generateTopologyFromOEMol(molecule)
# Create system with forcefield.
system = forcefield.createSystem(topology)
bond_gen = forcefield.getGenerators()[0]
bond_gen.types1
bond_gen.k
```
| github_jupyter |
```
#This will run the createQualificationType in the commandline and will create a qualification type in MTurk
#You need to find these text files in your old computer
#You should make these text files more easily editable through this python script as well (read in files, edit, output)
import os
import subprocess
os.chdir('C:\\Users\\bodil\\aws-mturk-clt-1.3.3\\bin')
args = ["createQualificationType","-properties",'properties.txt','-answer','answer_key.txt','-question','test.txt','-noretry']
args = subprocess.list2cmdline(args)
os.system(args)
for resp in qualtrics_responses:
###change this question based on the new qualtrics survey###
if qualtrics_responses[resp]['Q31'] == 1:
qualtrics_codes.append(qualtrics_responses[resp]['survey_code'])
else:
pass
#This code will get all submitted qualtrics survey codes
#Then it will get all submitted MTurk codes
#Then it compares them and approves the ones that match and rejects the ones that do not
#It also makes a list of workers and changes their qualification number so that they cannot take the survey again in the future
###########################################################
#change this value to the actual qualification type id
qualification_type_id = 'INSERT ID HERE'
###########################################################
import math
import requests
import boto.mturk.connection
connection = boto.mturk.connection.MTurkConnection(aws_access_key_id='INSERT ID HERE',
aws_secret_access_key='INSERT KEY HERE',
is_secure=True, port=None,
proxy=None, proxy_port=None,
proxy_user=None, proxy_pass=None,
host='mechanicalturk.amazonaws.com', debug=0,
https_connection_factory=None,
security_token=None, profile_name=None)
url = "https://survey.qualtrics.com/WRAPI/ControlPanel/api.php?Request=getLegacyResponseData&Version=2.0&User=[INSERT USER]&Token=[INSERT TOKEN]"
qualtrics_codes = []
qualtrics_responses = requests.get(url).json()
for resp in qualtrics_responses:
###change this question based on the new qualtrics survey###
if qualtrics_responses[resp]['Q31'] == 1:
qualtrics_codes.append(qualtrics_responses[resp]['survey_code'])
else:
pass
url = "https://survey.qualtrics.com/WRAPI/ControlPanel/api.php?Request=getLegacyResponseData&ResponsesInProgress=1&Version=2.0&User=[INSERT USER]&Token=[INSERT TOKEN]&Format=JSON&SurveyID={INSERT ID}"
qualtrics_responses = requests.get(url).json()
for resp in qualtrics_responses:
###change this question based on the new qualtrics survey###
if qualtrics_responses[resp]['Q31'] == 1:
qualtrics_codes.append(qualtrics_responses[resp]['survey_code'])
else:
pass
workers = []
approved = []
rejected = []
all_hits = [hit for hit in connection.get_all_hits()]
codes_entered_to_mturk = []
#testcode hitcode = '3CRWSLD91K4646XDRA2O9WPNTE4OM0'
for hit in all_hits:
assignments = connection.get_assignments(hit.HITId,status='Rejected',page_size=100,page_number=1)
number_of_pages = int(math.ceil(float(assignments.TotalNumResults)/100))
for number in range(2,number_of_pages+1):
assignments += connection.get_assignments(hit.HITId,status='Submitted',page_size=100,page_number=number)
for assignment in assignments:
workers.append(assignment.WorkerId)
# don't ask me why this is a 2D list
question_form_answers = assignment.answers[0]
for question_form_answer in question_form_answers:
# "user-input" is the field I created and the only one I care about
if question_form_answer.qid == "surveycode":
user_response = question_form_answer.fields[0]
user_response = user_response.replace(' ','')
codes_entered_to_mturk.append(user_response)
if user_response in qualtrics_codes:
approved.append((assignment.AssignmentId,assignment.WorkerId,user_response))
# connection.approve_assignment(assignment.AssignmentId)
else:
# connection.reject_assignment(assignment.AssignmentId, feedback="I'm sorry, your code was not found in our qualtrics database of generated codes. Please contact the email address associated with the requesters MTurk account if you feel you have been rejected in error.")
rejected.append((assignment.AssignmentId,assignment.WorkerId,user_response))
# workers = list(set(workers))
# for worker in workers:
# value = -1
# connection.update_qualification_score(qualification_type_id, worker, value)
user_response = user_response.replace(' ','')
#This script will approve a rejection due to technical difficulties (maybe we have their responses, just not on Q18...?)
import math
import requests
import boto.mturk.connection
feedback = '''
I am sorry for the rejection. There are a number of reasons this may have occurred:
1. The most common reason is if you forgot to hit submit in qualtrics after you completed the survey.
2. Another reason is the code you entered was incorrect (space before or after the code or incorrect code)
Thank you for contacting me. I have reversed the rejection. I hope you will continue to do work for me in the future.
'''
feedback = '''
I am sorry for the rejection. You were rejected because when you put the Qualtrics code into MTURK, you accidentally did
not click finish on the survey. This meant your response was still in progress, and so your HIT was automatically rejected.
We are sorry that we accidentally rejected your work. We appreciate the time you spent working on this hit.
Thank you for contacting me. I have reversed the rejection. I hope you will continue to do work for me in the future.
'''
feedback = '''
I am sorry for the rejection. Your HIT was rejected because the code you entered had a space before the code, so it did
not match with any codes in our database.
Thank you for contacting me. The code was correct (other than the space).
I have reversed the rejection. I hope you will continue to do work for me in the future.
'''
connection = boto.mturk.connection.MTurkConnection(aws_access_key_id='INSERT ID HERE',
aws_secret_access_key='INSERT KEY HERE',
is_secure=True, port=None,
proxy=None, proxy_port=None,
proxy_user=None, proxy_pass=None,
host='mechanicalturk.amazonaws.com', debug=0,
https_connection_factory=None,
security_token=None, profile_name=None)
assignment_ids_for_approval = ['INSERT ID HERE']
#Go through and approve the rejected assignments
for assignment_id in assignment_ids_for_approval:
try:
connection.approve_rejected_assignment(assignment_id, feedback=feedback)
print assignment_id
except:
print 'failure'
import requests
url = "https://survey.qualtrics.com/WRAPI/ControlPanel/api.php?Request=getLegacyResponseData&Version=2.0&User=[INSERT USER]&Token=[INSERT TOKEN]&Format=JSON&SurveyID=[INSERT ID]"
qualtrics_codes = []
qualtrics_responses = requests.get(url).json()
```
| github_jupyter |
```
import os
import glob
from Offline import utils
from sklearn.metrics import roc_curve, auc, roc_curve
from sklearn.linear_model import LogisticRegression
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats, signal
from scipy.stats import ttest_ind
from scipy.interpolate import interp1d
import mne
from mne.stats import permutation_cluster_test
from mne.evoked import _get_peak
import pandas as pd
import seaborn as sns
np.random.seed(0)
%matplotlib inline
%load_ext autoreload
%autoreload 2
sns.set(style="white")
# load tmp data
data_dir = './data/epoch_data'
f_list = glob.glob(os.path.join(data_dir, '*.npz'))
print(f_list)
data_list = [np.load(f) for f in f_list]
t_ori = (-0.3, 0.6)
t_tar = (0, 0.5)
```
## Temporal pattern
```
epochs = np.concatenate([d['epochs'] for d in data_list], axis=0)
y = np.concatenate([d['y'] for d in data_list], axis=0)
print(epochs.shape)
print(y.shape)
# clustering test
T = []
clu = []
p_clu = []
for i in range(epochs.shape[1]):
x_l = epochs[y == 1, i]
x_r = epochs[y == 2, i]
T_obs, clusters, cluster_p_values, H0 = permutation_cluster_test([x_l, x_r], n_permutations=1000, n_jobs=-1)
T.append(T_obs)
clu.append(clusters)
p_clu.append(cluster_p_values)
def fig1plot(t, y, epochs, clu, p_clu):
cls_name=('non', 'left', 'right')
ch_name= ['Pz', 'P3', 'P4', 'P7', 'P8', 'Oz', 'O1', 'O2']
ch_order = [('P3', 'P7', 'O1'), ('P4', 'P8', 'O2'), ('Pz', 'Oz')]
base = 0
ylim = 1
p_th = 0.001
ch_fontsize=14
legend_fontsize=11
ticks_fontsize=11
fig1, ax1 = plt.subplots(3, 1, figsize=(6, 9), sharex=True)
fig2, ax2 = plt.subplots(2, 1, figsize=(6, 4), sharex=True)
fig3, ax3 = plt.subplots(3, 1, figsize=(6, 9), sharex=True)
# hemispheres
for j, axes in enumerate([ax1, ax3, ax2]):
ch_ind = [ch_name.index(i) for i in ch_order[j]]
for i, ax in enumerate(axes):
ax.tick_params(axis='both', labelsize=ticks_fontsize, length=5, which='major')
ax.set_xticks([-0.3, -0.15, 0, 0.15, 0.3, 0.45, 0.6])
ax.set_yticks([-1, 0, 1])
ax.set_ylim([-ylim, ylim])
ax.set_xlim([t[0], t[-1]])
# nontarget
non_tar = epochs[y==0, ch_ind[i], :].mean(axis=0)
line1, = ax.plot(t, non_tar, c='gray', alpha=0.6, label='non-target', linewidth=2)
# leftward
line2, = ax.plot(t, epochs[y == 1, ch_ind[i], :].mean(axis=0), label='left', linewidth=2)
# rightward
line3, = ax.plot(t, epochs[y == 2, ch_ind[i], :].mean(axis=0), label='right', linewidth=2)
# shadow
# h = None
for i_c, c in enumerate(clu[ch_ind[i]]):
c = c[0]
if c.start < 300 or c.stop > 800:
continue
if p_clu[ch_ind[i]][i_c] <= p_th:
h = ax.axvspan(t[base + c.start], t[base + (c.stop - 1)],
color=(0.3, 0.3, 0.3), alpha=0.25)
ax.text(0.86, 0.84, ch_order[j][i], fontsize=ch_fontsize, transform=ax.transAxes)
if (not j) and (not i):
ax.legend((line1, line2, line3), ('Non-target', 'Leftward', 'Rightward'))
return [fig1, fig2, fig3]
t = np.linspace(-0.3, 0.6, epochs.shape[-1])
figs = fig1plot(t, y, epochs, clu, p_clu)
plt.show()
```
## ERP Analysis
```
ch_names = ['Pz', 'P3', 'P4', 'P7', 'P8', 'Oz', 'O1', 'O2']
times = np.arange(t_ori[0], t_ori[1], 1/1000)
def hemi(ch):
if ch in ['P3', 'P7', 'O1']:
return 'lh'
elif ch in ['P4', 'P8', 'O2']:
return 'rh'
else:
return 'mid'
latency = []
for d in data_list:
epochs = d['epochs']
y = d['y']
# left evoked
l_ = epochs[y==1].mean(axis=0)
# right evoked
r_ = epochs[y==2].mean(axis=0)
# compute latency
for i, ch in enumerate(ch_names):
h = hemi(ch)
loc, t, val = _get_peak(l_[[i]], times, tmin=0.1, tmax=0.3, mode='neg')
latency.append([ch, h, 'leftward', times[t], val])
loc, t, val = _get_peak(r_[[i]], times, tmin=0.1, tmax=0.3, mode='neg')
latency.append([ch, h, 'rightward', times[t], val])
lag_df = pd.DataFrame(latency, columns=['Channel', 'Hemisphere', 'Direction', 'Latency', 'Amplitude'])
lag_df.head(10)
```
### Latency
```
fig, axes = plt.subplots(1, 1, figsize=(7, 6))
sns.boxplot(x='Hemisphere',
y='Latency',
hue='Direction',
data=lag_df,
order=['lh', 'mid', 'rh'],
width=0.4,
whis=5, # no outlier points
ax=axes)
handles, _ = axes.get_legend_handles_labels()
plt.ylim([0.11, 0.33])
plt.ylabel('N200 peak latency [s]', fontsize=14)
plt.xlabel(None)
plt.xticks(range(3), ['Left-hemisphere', 'Middle', 'Right-hemisphere'], fontsize=14)
plt.legend(handles, ['Leftward', 'Rightward'], frameon=False, fontsize=13)
plt.show()
# t-test leftward vs rightward on different hemispheres
# lh
l_ = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'rightward')]
print('lh: ', ttest_ind(l_.Latency.values, r_.Latency.values))
# rh
l_ = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'rightward')]
print('rh: ', ttest_ind(l_.Latency.values, r_.Latency.values))
# mid
l_ = lag_df[(lag_df.Hemisphere == 'mid') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'mid') & (lag_df.Direction == 'rightward')]
print('mid: ', ttest_ind(l_.Latency.values, r_.Latency.values))
# t-test lh vs rh for different stimuli
# leftward
l_lh = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'leftward')]
l_rh = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'leftward')]
print('leftward lh vs rh: ', ttest_ind(l_lh.Latency.values, l_rh.Latency.values,))
# rightward
r_lh = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'rightward')]
r_rh = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'rightward')]
print('rightward lh vs rh: ', ttest_ind(r_lh.Latency.values, r_rh.Latency.values,))
```
### Amplitude
```
fig, axes = plt.subplots(1, 1, figsize=(7, 6))
sns.boxplot(x='Hemisphere',
y='Amplitude',
hue='Direction',
data=lag_df,
order=['lh', 'mid', 'rh'],
width=0.4,
whis=5, # no outlier points
ax=axes)
handles, _ = axes.get_legend_handles_labels()
plt.ylim([-2.2, 0.3])
plt.gca().invert_yaxis()
plt.ylabel('N200 peak amplitude [z-score]', fontsize=14)
plt.xlabel(None)
plt.xticks(range(3), ['Left-hemisphere', 'Middle', 'Right-hemisphere'], fontsize=14)
plt.legend(handles, ['Leftward', 'Rightward'], frameon=False, fontsize=13)
plt.show()
# t-test leftward vs rightward on different hemispheres
# lh
l_ = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'rightward')]
print('lh: ', ttest_ind(l_.Amplitude.values, r_.Amplitude.values))
# rh
l_ = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'rightward')]
print('rh: ', ttest_ind(l_.Amplitude.values, r_.Amplitude.values))
# mid
l_ = lag_df[(lag_df.Hemisphere == 'mid') & (lag_df.Direction == 'leftward')]
r_ = lag_df[(lag_df.Hemisphere == 'mid') & (lag_df.Direction == 'rightward')]
print('mid: ', ttest_ind(l_.Amplitude.values, r_.Amplitude.values))
# t-test lh vs rh for different stimuli
# leftward
l_lh = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'leftward')]
l_rh = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'leftward')]
print('leftward lh vs rh: ', ttest_ind(l_lh.Amplitude.values, l_rh.Amplitude.values,))
# rightward
r_lh = lag_df[(lag_df.Hemisphere == 'lh') & (lag_df.Direction == 'rightward')]
r_rh = lag_df[(lag_df.Hemisphere == 'rh') & (lag_df.Direction == 'rightward')]
print('rightward lh vs rh: ', ttest_ind(r_lh.Amplitude.values, r_rh.Amplitude.values,))
```
## ROC Analysis
```
fpr_interp = np.linspace(0, 1, 200)
def evaluate_multiclass(y_pred, y_true):
n_classes = len(np.unique(y_true))
y_pred_label = np.argmax(y_pred, axis=1)
# onehot label
y_true_onehot = np.eye(n_classes)[y_true]
roc = []
for i in range(n_classes):
fpr, tpr, _ = roc_curve(y_true_onehot[:, i], y_pred[:, i])
f = interp1d(fpr, tpr)
tpr = f(fpr_interp)
roc.append(tpr)
roc = np.array(roc)
return roc
def cross_val(estimator, X, y, n=2):
k_fold_rocs = []
for train_ind, test_ind in utils.uniform_kfold(y):
X_train = X[train_ind]
y_train = y[train_ind]
X_test = X[test_ind]
y_test = y[test_ind]
# normalize
mean_ = X_train.mean(axis=(0, 2), keepdims=True)
std_ = X_train.std(axis=(0, 2), keepdims=True)
X_train -= mean_
X_train /= std_
X_test -= mean_
X_test /= std_
X_train = X_train.reshape((X_train.shape[0], -1))
X_test = X_test.reshape((X_test.shape[0], -1))
estimator.fit(X_train, y_train)
y_pred = utils.pred_ave(estimator, X_test, y_test, n=n)
roc = evaluate_multiclass(y_pred, y_test)
k_fold_rocs.append(roc)
k_fold_rocs = np.array(k_fold_rocs).mean(axis=0)
return k_fold_rocs
estimator = LogisticRegression(class_weight='balanced', solver='liblinear', multi_class='ovr')
roc_subjects_classes = []
for d in data_list:
# cut epochs (0-0.5s)
epochs = utils.timewindow(t_ori, t_tar, d['epochs'])
# downsample and reshape to 2d matrix
X = epochs[..., ::25]
y = d['y']
roc = cross_val(estimator, X, y)
roc_subjects_classes.append(roc)
roc_subjects_classes = np.array(roc_subjects_classes)
# mean rocs for each classes
rocs_classes = roc_subjects_classes.mean(axis=0)
roc_classes_err = roc_subjects_classes.std(axis=0)
cmap = plt.get_cmap('tab10')
cls_names=('Non-target', 'Leftward', 'Rightward')
c = ['gray', cmap(0), cmap(1)]
auc = np.trapz(rocs_classes, fpr_interp, axis=1)
lines = []
plt.figure()
for i in range(1, 3):
line, = plt.plot(fpr_interp, rocs_classes[i], c=c[i])
lines.append(line)
plt.fill_between(fpr_interp, rocs_classes[i]-roc_classes_err[i], rocs_classes[i]+roc_classes_err[i], alpha=0.2, facecolor=c[i])
print('%s AUC: %.4f ± %.4f' % (cls_names[i], auc[i], np.std(np.trapz(roc_subjects_classes[:, i], fpr_interp, axis=-1))))
plt.plot([0, 1], [0, 1], ls="--", c=".3")
plt.legend(lines, ['%s AUC=%.2f' % (cls_names[i], auc[i]) for i in range(1, 3)], fontsize=14)
plt.xlabel('False positive rate', fontsize=14)
plt.ylabel('True positive rate', fontsize=14)
plt.gca().tick_params(axis='both', which='major', labelsize='large')
plt.show()
roc_subjects_n_trials = []
n_trials = [1, 2, 3, 5, 10]
for i in n_trials:
roc_subjects = []
for d in data_list:
# cut epochs (0-0.5s)
epochs = utils.timewindow(t_ori, t_tar, d['epochs'])
# downsample and reshape to 2d matrix
X = epochs[..., ::25]
y = d['y']
roc = cross_val(estimator, X, y, n=i)
roc_subjects.append(roc) # roc for different classes
roc_subjects_n_trials.append(np.mean(roc_subjects, axis=0))
roc_subjects_n_trials = np.array(roc_subjects_n_trials)
print(roc_subjects_n_trials.shape)
# auc for leftward
roc_subjects_n_trials_left = roc_subjects_n_trials[:, 1].copy()
auc_n_trials = np.sum(roc_subjects_n_trials_left, axis=-1) / len(fpr_interp)
plt.figure()
gray_map = plt.get_cmap('Blues')
for i in range(len(n_trials)):
plt.plot(fpr_interp, roc_subjects_n_trials_left[i], c=gray_map(0.15 * i + 0.4),
label='N=%d, AUC=%.2f' % (n_trials[i], auc_n_trials[i]))
plt.plot([0, 1], [0, 1], ls="--", c=".3")
plt.legend(fontsize=14)
plt.xlabel('False positive rate', fontsize=14)
plt.ylabel('True positive rate', fontsize=14)
plt.gca().tick_params(axis='both', which='major', labelsize='large')
plt.show()
# auc for rightward
roc_subjects_n_trials_right = roc_subjects_n_trials[:, 2].copy()
auc_n_trials = np.sum(roc_subjects_n_trials_right, axis=-1) / len(fpr_interp)
plt.figure()
gray_map = plt.get_cmap('Oranges')
for i in range(len(n_trials)):
plt.plot(fpr_interp, roc_subjects_n_trials_right[i], c=gray_map(0.15 * i + 0.4),
label='N=%d, AUC=%.2f' % (n_trials[i], auc_n_trials[i]))
plt.plot([0, 1], [0, 1], ls="--", c=".3")
plt.legend(fontsize=14)
plt.xlabel('False positive rate', fontsize=14)
plt.ylabel('True positive rate', fontsize=14)
plt.gca().tick_params(axis='both', which='major', labelsize='large')
plt.show()
```
## Online error rate
```
def cross_val_y_pred(estimator, X, y):
y_pred_n_fold = []
y_true_n_fold = []
for train_ind, test_ind in utils.uniform_kfold(y):
X_train = X[train_ind]
y_train = y[train_ind]
X_test = X[test_ind]
y_test = y[test_ind]
# normalize
mean_ = X_train.mean(axis=(0, 2), keepdims=True)
std_ = X_train.std(axis=(0, 2), keepdims=True)
X_train -= mean_
X_train /= std_
X_test -= mean_
X_test /= std_
X_train = X_train.reshape((X_train.shape[0], -1))
X_test = X_test.reshape((X_test.shape[0], -1))
estimator.fit(X_train, y_train)
y_pred = utils.pred_ave(estimator, X_test, y_test, n=1)
y_pred_n_fold.append(y_pred)
y_true_n_fold.append(y_test)
y_pred_n_fold = np.concatenate(y_pred_n_fold, axis=0)
y_true_n_fold = np.concatenate(y_true_n_fold, axis=0)
return y_pred_n_fold, y_true_n_fold
def train_estimate_online_correct_rate(data):
# cut epochs (0-0.5s)
epochs = utils.timewindow(t_ori, t_tar, data['epochs'])
# downsample and reshape to 2d matrix
X = epochs[..., ::25]
y = data['y']
y_pred, y_true = cross_val_y_pred(estimator, X, y)
pc = utils.estimate_accu_dual(y_true, y_pred, n_avg=5)
return pc
def find_n_trials(pc, expected_accu):
x = np.linspace(1, 5, 100)
interp_f = interp1d(np.arange(1,6), pc)
interp_pc = interp_f(x)
n_trials = x[np.argmin(np.abs(interp_pc - expected_accu))]
return n_trials
estimator = LogisticRegression(class_weight='balanced', solver='liblinear', multi_class='ovr')
# error rate estimation
y_pred_n_fold = []
s1_data = data_list[f_list.index(os.path.join(data_dir, 'S1_epochs.npz'))]
s6_data = data_list[f_list.index(os.path.join(data_dir, 'S6_epochs.npz'))]
s10_data = data_list[f_list.index(os.path.join(data_dir, 'S10_epochs.npz'))]
pc_s1 = train_estimate_online_correct_rate(s1_data)
pc_s6 = train_estimate_online_correct_rate(s6_data)
pc_s10 = train_estimate_online_correct_rate(s10_data)
expected_accu = 0.85
s1_n_trial = find_n_trials(pc_s1, expected_accu)
s6_n_trial = find_n_trials(pc_s6, expected_accu)
s10_n_trial = find_n_trials(pc_s10, expected_accu)
print(s1_n_trial, s6_n_trial, s10_n_trial)
fig, axes = plt.subplots(1, 3, sharey=True, figsize=(7, 5))
axes[0].plot(np.arange(1,6), pc_s1, c='k')
axes[0].scatter(1.83, 0.944, c='k')
axes[0].axhline(y=expected_accu, c='gray', linestyle='dashed')
axes[0].axvline(x=s1_n_trial, c='gray', linestyle='dashed')
axes[0].set_ylim([0, 1.05])
axes[0].set_yticks([0, 0.2, 0.4, 0.6, 0.8, 0.85, 1])
y_expected_tick = axes[0].get_yticklabels()[5]
y_expected_tick.set_color('r')
y_expected_tick.set_fontsize(13)
x = [1,s1_n_trial, 5]
axes[0].set_xticks(x)
axes[0].set_xticklabels(['%.1f' % i for i in x])
x_n_trial_tick = axes[0].get_xticklabels()[1]
x_n_trial_tick.set_color('r')
x_n_trial_tick.set_fontsize(13)
axes[0].set_title('S1')
axes[1].plot(np.arange(1,6), pc_s6, c='k')
axes[1].scatter(2.25, 0.833, c='k')
axes[1].axhline(y=expected_accu, c='gray', linestyle='dashed')
axes[1].axvline(x=s6_n_trial, c='gray', linestyle='dashed')
x = [1, s6_n_trial, 5]
axes[1].set_xticks(x)
axes[1].set_xticklabels(['%.1f' % i for i in x])
x_n_trial_tick = axes[1].get_xticklabels()[1]
x_n_trial_tick.set_color('r')
x_n_trial_tick.set_fontsize(13)
axes[1].set_title('S6')
axes[2].plot(np.arange(1,6), pc_s10, c='k', label='Estimated')
axes[2].scatter(3.4, 0.639, c='k', label='True')
axes[2].axhline(y=expected_accu, c='gray', linestyle='dashed')
x = [1,3,5]
axes[2].set_xticks(x)
axes[2].set_xticklabels(['%.1f' % i for i in x])
axes[2].set_title('S10')
axes[2].legend(fontsize=12)
fig.text(0.5, 0.01, 'Number of averaging trials', ha='center', va='center', fontsize=14)
fig.text(0.01, 0.5, 'Accuracy', ha='center', va='center', rotation='vertical', fontsize=14)
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from scipy import stats
class Stats():
def __init__(self):
self.n = 0
self.mu = 0
self.std = 0
self.M = 0
self.alpha = 0.05
self.tail_num = 2
def __calculate_zscore__(self):
se = self.std / np.sqrt(self.n)
return round((self.M - self.mu) / se, 2)
def __calculate_norm_portion__(self):
return round(stats.norm.ppf(1 - self.alpha/self.tail_num), 3)
def ztest_1samp_from_stats():
z, cr = self.__calculate_zscore__(), self.__calculate_norm_portion__()
if self.tail_num == 2:
rejection_decision = (z > cr) | (z < -1 * cr)
region = f'z > {cr} or z < -{cr}'
criteria = f'two tail, alpha {alpha}'
elif self.tail_num == 1:
if z > 0:
rejection_decision = (z > cr)
region = f'z > {cr}'
else:
rejection_decision = (z < -1 * cr)
region = f'z < -{cr}'
criteria = f'one tail, alpha {alpha}'
else:
print('Should use tail_num 1 or 2.')
return None
print(f'[{criteria}] z_statistic:{z}, critical_region:{region}\n=> null hypothesis rejection [{rejection_decision}]')
def __calculate_t_statistic__(self):
se = self.std / np.sqrt(self.n)
return round((self.M - self.mu) / se, 2)
def __calculate_t_portion__(self):
df = self.n - 1
return round(stats.t.ppf(1 - self.alpha/self.tail_num, df=df), 3)
def r_squared_from_stats():
t = self.__calculate_t_statistic__()
return t ** 2 / (t ** 2 + self.n - 1)
def ttest_1samp_from_stats():
t, cr = self.__calculate_t_statistic__(), self.__calculate_t_portion__()
if self.tail_num == 2:
rejection_decision = (t > cr) | (t < -1 * cr)
region = f't > {cr} or t < -{cr}'
criteria = f'two tail, alpha {alpha}'
elif self.tail_num == 1:
if t > 0:
rejection_decision = (t > cr)
region = f't > {cr}'
else:
rejection_decision = (t < -1 * cr)
region = f't < -{cr}'
criteria = f'one tail, alpha {alpha}'
else:
print('Should use tail_num 1 or 2.')
return None
print(f'[{criteria}] t_statistic:{t}, critical_region:{region}\n=> null hypothesis rejection [{rejection_decision}]')
def cohens_d_from_stats():
return round(abs((self.M - self.mu) / self.std), 2)
# def statistical_power_from_stats():
# se = self.std / np.sqrt(self.n)
# z = ((self.mu + 1.96 * se) - self.M) / se
# return round(1 - stats.norm.cdf(z), 4)
```
| github_jupyter |
<small><i>This notebook was put together by [Jake Vanderplas](http://www.vanderplas.com), modified by [Shannon Quinn](https://magsol.github.io). Source and license info is on [GitHub](https://github.com/eds-uga/acm-ml-workshop-2017/).</i></small>
# Introduction to Scikit-Learn: Machine Learning with Python
This session will cover the basics of Scikit-Learn, a popular package containing a collection of tools for machine learning written in Python. See more at http://scikit-learn.org.
## Outline
**Main Goal:** To introduce the central concepts of machine learning, and how they can be applied in Python using the Scikit-learn Package.
- Definition of machine learning
- Data representation in scikit-learn
- Introduction to the Scikit-learn API
## About Scikit-Learn
[Scikit-Learn](http://github.com/scikit-learn/scikit-learn) is a Python package designed to give access to **well-known** machine learning algorithms within Python code, through a **clean, well-thought-out API**. It has been built by hundreds of contributors from around the world, and is used across industry and academia.
Scikit-Learn is built upon Python's [NumPy (Numerical Python)](http://numpy.org) and [SciPy (Scientific Python)](http://scipy.org) libraries, which enable efficient in-core numerical and scientific computation within Python. As such, scikit-learn is not specifically designed for extremely large datasets, though there is [some work](https://github.com/ogrisel/parallel_ml_tutorial) in this area.
For this short introduction, I'm going to stick to questions of in-core processing of small to medium datasets with Scikit-learn.
## What is Machine Learning?
In this section we will begin to explore the basic principles of machine learning.
Machine Learning is about building programs with **tunable parameters** (typically an
array of floating point values) that are adjusted automatically so as to improve
their behavior by **adapting to previously seen data.**
Machine Learning can be considered a subfield of **Artificial Intelligence** since those
algorithms can be seen as building blocks to make computers learn to behave more
intelligently by somehow **generalizing** rather that just storing and retrieving data items
like a database system would do.
We'll take a look at two very simple machine learning tasks here.
The first is a **classification** task: the figure shows a
collection of two-dimensional data, colored according to two different class
labels. A classification algorithm may be used to draw a dividing boundary
between the two clusters of points:
```
%matplotlib inline
# set seaborn plot defaults.
# This can be safely commented out
import seaborn
# Import the example plot from the figures directory
from fig_code import plot_sgd_separator
plot_sgd_separator()
```
This may seem like a trivial task, but it is a simple version of a very important concept.
By drawing this separating line, we have learned a model which can **generalize** to new
data: if you were to drop another point onto the plane which is unlabeled, this algorithm
could now **predict** whether it's a blue or a red point.
If you'd like to see the source code used to generate this, you can either open the
code in the `fig_code` directory, or you can load the code using the `%load` magic command:
The next simple task we'll look at is a **regression** task: a simple best-fit line
to a set of data:
```
from fig_code import plot_linear_regression
plot_linear_regression()
```
Again, this is an example of fitting a model to data, such that the model can make
generalizations about new data. The model has been **learned** from the training
data, and can be used to predict the result of test data:
here, we might be given an x-value, and the model would
allow us to predict the y value. Again, this might seem like a trivial problem,
but it is a basic example of a type of operation that is fundamental to
machine learning tasks.
## Representation of Data in Scikit-learn
Machine learning is about creating models from data: for that reason, we'll start by
discussing how data can be represented in order to be understood by the computer. Along
with this, we'll show some examples of how to visualize data, and discuss the data structures required for machine learning and how they relate to the underlying mathematics.
Most machine learning algorithms implemented in scikit-learn expect data to be stored in a
**two-dimensional array or matrix**. The arrays can be
either ``numpy`` arrays, or in some cases ``scipy.sparse`` matrices.
The size of the array is expected to be `[n_samples, n_features]`
- **n_samples:** The number of samples: each sample is an item to process (e.g. classify).
A sample can be a document, a picture, a sound, a video, an astronomical object,
a row in database or CSV file,
or whatever you can describe with a fixed set of quantitative traits.
- **n_features:** The number of features or distinct traits that can be used to describe each
item in a quantitative manner. Features are generally real-valued, but may be boolean or
discrete-valued in some cases.
The number of features must be fixed in advance. However it can be very high dimensional
(e.g. millions of features) with most of them being zeros for a given sample. This is a case
where `scipy.sparse` matrices can be useful, in that they are
much more memory-efficient than numpy arrays.

(Figure from the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook))
## A Simple Example: the Iris Dataset
As an example of a simple dataset, we're going to take a look at the
iris data stored by scikit-learn.
The data consists of measurements of three different species of irises.
There are three species of iris in the dataset, which we can picture here:
```
from IPython.core.display import Image, display
display(Image(filename='images/iris_setosa.jpg'))
print("Iris Setosa\n")
display(Image(filename='images/iris_versicolor.jpg'))
print("Iris Versicolor\n")
display(Image(filename='images/iris_virginica.jpg'))
print("Iris Virginica")
```
### Quick Question:
**If we want to design an algorithm to recognize iris species, what might the data be?**
Remember: we need a 2D array of size `[n_samples x n_features]`.
- What would the `n_samples` refer to?
- What might the `n_features` refer to?
Remember that there must be a **fixed** number of features for each sample, and feature
number ``i`` must be a similar kind of quantity for each sample.
### Loading the Iris Data with Scikit-Learn
Scikit-learn has a very straightforward set of data on these iris species. The data consist of
the following:
- Features in the Iris dataset:
1. sepal length in cm
2. sepal width in cm
3. petal length in cm
4. petal width in cm
- Target classes to predict:
1. Iris Setosa
2. Iris Versicolour
3. Iris Virginica
``scikit-learn`` embeds a copy of the iris CSV file along with a helper function to load it into numpy arrays:
```
from sklearn.datasets import load_iris
iris = load_iris()
```
We can examine this data more closely, using the properties of the various Python objects storing it:
```
iris.keys()
n_samples, n_features = iris.data.shape
print((n_samples, n_features))
print(iris.data[0])
print(iris.data.shape)
print(iris.target.shape)
print(iris.target)
print(iris.target_names)
```
This data is four dimensional, but we can visualize two of the dimensions
at a time using a simple scatter-plot:
```
import numpy as np
import matplotlib.pyplot as plt
x_index = 0
y_index = 1
# this formatter will label the colorbar with the correct target names
formatter = plt.FuncFormatter(lambda i, *args: iris.target_names[int(i)])
plt.scatter(iris.data[:, x_index], iris.data[:, y_index],
c=iris.target, cmap=plt.cm.get_cmap('RdYlBu', 3))
plt.colorbar(ticks=[0, 1, 2], format=formatter)
plt.clim(-0.5, 2.5)
plt.xlabel(iris.feature_names[x_index])
plt.ylabel(iris.feature_names[y_index]);
```
### Quick Exercise:
**Change** `x_index` **and** `y_index` **in the above script
and find a combination of two parameters
which maximally separate the three classes.**
This exercise is a preview of **dimensionality reduction**, which we'll see later.
## Other Available Data
They come in three flavors:
- **Packaged Data:** these small datasets are packaged with the scikit-learn installation,
and can be downloaded using the tools in ``sklearn.datasets.load_*``
- **Downloadable Data:** these larger datasets are available for download, and scikit-learn
includes tools which streamline this process. These tools can be found in
``sklearn.datasets.fetch_*``
- **Generated Data:** there are several datasets which are generated from models based on a
random seed. These are available in the ``sklearn.datasets.make_*``
You can explore the available dataset loaders, fetchers, and generators using IPython's
tab-completion functionality. After importing the ``datasets`` submodule from ``sklearn``,
type
datasets.load_ + TAB
or
datasets.fetch_ + TAB
or
datasets.make_ + TAB
to see a list of available functions.
```
from sklearn import datasets
# Type datasets.fetch_<TAB> or datasets.load_<TAB> in IPython to see all possibilities
# datasets.fetch_
# datasets.load_
```
In the next section, we'll use some of these datasets and take a look at the basic principles of machine learning.
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import numpy as np
import lda
from data_analysis.topic_tokenizer import TopicTokenizer
from data_analysis._util import make_document_term_matrix
file_ = 'twitter.sqlite3'
tweets = pd.read_sql_table('tweets', 'sqlite:///{}'.format(file_))
tweets = tweets[tweets.is_retweet == False]
tokenizer = TopicTokenizer()
token_list = [tokenizer.tokenize(t) for t in tweets.tweet[0:2000]]
tokenizer.tokenize('@benhoff https://www.googel.com')
token_list = [t for t in token_list if t]
bool([])
document_matrix, vocabulary = make_document_term_matrix(token_list)
model = lda.LDA(n_topics=120, n_iter=1500, random_state=1)
model.fit(document_matrix)
vocab_array = np.array(list(vocabulary.keys()))
for i, topic_dist in enumerate(model.topic_word_):
temp = np.argpartition(-topic_dist, 8)
result = temp[:8]
word_result = []
for r in result:
word_result.append(vocab_array[r])
print('Topic {}: {}'.format(i, ' '.join(word_result)))
classified_data = model.fit_transform(document_matrix)
print(classified_data)
# classified_data = classified_data.argmax(1)
from collections import Counter
count = Counter(classified_data)
keys_counts = np.array(count.most_common())
keys = keys_counts[:, 0]
counts = keys_counts[:, 1]
print(keys, counts)
from matplotlib import pyplot as plt
plt.barh(np.arange(len(counts), 0, -1), counts)
def get_vocabulary_helper(topic_numbers, number=5):
vocab = np.array(list(vocabulary.keys()))
topic_models = model.topic_word_
result = []
for topic_number in topic_numbers:
words = vocab[np.argsort(topic_models[topic_number])][:-(number+1):-1]
result.append(words)
return result
word_list = get_vocabulary_helper(keys)
fig = plt.figure()
axis = fig.add_subplot(111)
X, Y = fig.get_dpi() * fig.get_size_inches()
h = Y / (20)
for row, words in enumerate(word_list):
y = Y - (row * h) - h
axis.text(0.3, y, ' '.join(words), fontsize=(h * 0.8),
horizontalalignment='left',
verticalalignment='center')
axis.set_ylim(0, Y)
axis.set_axis_off()
fig = plt.figure()
axis = fig.add_subplot(111)
X, Y = fig.get_dpi() * fig.get_size_inches()
num_print = 10
h = Y / num_print
y_s = []
for row, words in enumerate(word_list[:num_print]):
y = Y - (row * h) - h
y_s.append(y)
axis.text(1, y, ' '.join(words), fontsize=(h*.5),
horizontalalignment='left',
verticalalignment='center',
color='gold')
y_s[-1] = 0
axis.set_ylim(-20, Y)
#axis.set_xlin(0, )
axis.barh(y_s, counts[:num_print], height=30, color='midnightblue')
axis.get_yaxis().set_visible(False)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.