text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
## Question: *when are molecules likely to have similar phenotypes?*
```
%load_ext autoreload
%autoreload 2
%cd /Users/sabrieyuboglu/Documents/sabri/research/projects/milieu/milieu
import numpy as np
from scipy.sparse import csr_matrix
from scipy.stats import pearsonr, spearmanr, ttest_ind, ttest_rel
import seaborn as sns
import matplotlib.pyplot as plt
from milieu.data.associations import load_diseases, build_disease_matrix
from milieu.data.network_matrices import load_network_matrices
from milieu.data.network import Network
network = Network("data/networks/species_9606/huri/network.txt")
```
## Compute jaccard similarity between proteins
```
processes = load_diseases("data/associations/gene_ontology/species_9606/go_process/associations.csv")
functions = load_diseases("data/associations/gene_ontology/species_9606/go_function/associations.csv")
diseases = load_diseases("data/associations/disgenet/associations.csv")
drugs = load_diseases("data/associations/drugbank/associations.csv")
associations = {}
for dct in [diseases, functions, processes, drugs]:
associations.update(dct)
association_matrix, _ = build_disease_matrix(associations, network)
association_matrix.shape
def compute_jaccard(matrix):
"""
Computes the pairwise jaccard similarity between
:param matrix: (nd.array) an NxD matrix where N is the # of sets and D is
the maximum cardinality of the sets.
"""
intersection = (csr_matrix(matrix)
.dot(csr_matrix(matrix.T)).todense())
union = np.zeros_like(intersection)
union += matrix.sum(axis=1, keepdims=True)
union += matrix.sum(axis=1, keepdims=True).T
union -= intersection
jaccard = np.array(np.nan_to_num(intersection / union, 0))
return jaccard
association_jaccard = compute_jaccard(association_matrix.T)
np.fill_diagonal(association_jaccard, 0)
# test jaccard
from sklearn.metrics import jaccard_score
for _ in range(10000):
i = np.random.randint(0, association_jaccard.shape[0])
j = np.random.randint(0, association_jaccard.shape[0])
if i == j:
continue
computed = association_jaccard[i, j]
value = jaccard_score(association_matrix[:, i], association_matrix[:, j])
if computed != value:
raise ValueError("Failed")
print("passed")
mi_matrix = load_network_matrices({"mi": "data/networks/species_9606/huri/mutual_interactor"},
network=network)["mi"]
pearsonr(mi_matrix[np.triu_indices(mi_matrix.shape[0], k=1)],
association_jaccard[np.triu_indices(association_jaccard.shape[0], k=1)])
x = network.adj_matrix
di_matrix = x / x.sum(axis=0, keepdims = True) / x.sum(axis=1, keepdims=True)
mi_values = mi_matrix[np.triu_indices(mi_matrix.shape[0], k=1)]
di_values = di_matrix[np.triu_indices(di_matrix.shape[0], k=1)]
adj_values = network.adj_matrix[np.triu_indices(network.adj_matrix.shape[0], k=1)]
jaccard_values = association_jaccard[np.triu_indices(association_jaccard.shape[0], k=1)]
# Claim: we findg that molecules with high mutual interactor scores are more similar than molecules with high direct interactor scores.
print(jaccard_values[mi_values >= np.percentile(mi_values, 99.9)].mean())
print(jaccard_values[di_values >= np.percentile(di_values, 99.9)].mean())
k = adj_values.sum().astype(int)
ttest_rel(jaccard_values[mi_values.argsort()[-k:]],
jaccard_values[di_values.argsort()[-k:]])
k = adj_values.sum().astype(int)
ttest_ind(jaccard_values[np.argpartition(mi_values, -k)[-k:]],
jaccard_values[np.argpartition(di_values, -k)[-k:]])
k = adj_values.sum().astype(int)
print(jaccard_values[np.argpartition(mi_values, -k)[-k:]].mean())
print(jaccard_values[np.argpartition(di_values, -k)[-k:]].mean())
adj_values[np.argpartition(jaccard_values, -k)[-k:]].mean()
(jaccard_values > 0.2).sum()
len(list(network.get_interactions()))
adj_values.sum()
```
| github_jupyter |
# Goal
* Simulating fullCyc Day1 control gradients
* Not simulating incorporation (all 0% isotope incorp.)
* Don't know how much true incorporatation for emperical data
* Using parameters inferred from emperical data (fullCyc Day1 seq data), or if not available, default SIPSim parameters
* Determining whether simulated taxa show similar distribution to the emperical data
## Input parameters
* phyloseq.bulk file
* taxon mapping file
* list of genomes
* fragments simulated for all genomes
* bulk community richness
## workflow
* Creating a community file from OTU abundances in bulk soil samples
* phyloseq.bulk --> OTU table --> filter to sample --> community table format
* Fragment simulation
* simulated_fragments --> parse out fragments for target OTUs
* simulated_fragments --> parse out fragments from random genomes to obtain richness of interest
* combine fragment python objects
* Convert fragment lists to kde object
* Add diffusion
* Make incorp config file
* Add isotope incorporation
* Calculating BD shift from isotope incorp
* Simulating gradient fractions
* Simulating OTU table
* Simulating PCR
* Subsampling from the OTU table
## Init
```
import os
import glob
import re
import nestly
%load_ext rpy2.ipython
%%R
library(ggplot2)
library(dplyr)
library(tidyr)
library(gridExtra)
library(phyloseq)
## BD for G+C of 0 or 100
BD.GCp0 = 0 * 0.098 + 1.66
BD.GCp100 = 1 * 0.098 + 1.66
```
# Nestly
* assuming fragments already simulated
```
workDir = '/home/nick/notebook/SIPSim/dev/fullCyc/n1147_frag_norm_9_2.5_n5/'
buildDir = os.path.join(workDir, 'Day1_default_run')
R_dir = '/home/nick/notebook/SIPSim/lib/R/'
fragFile= '/home/nick/notebook/SIPSim/dev/bac_genome1147/validation/ampFrags.pkl'
targetFile = '/home/nick/notebook/SIPSim/dev/fullCyc/CD-HIT/target_taxa.txt'
physeqDir = '/var/seq_data/fullCyc/MiSeq_16SrRNA/515f-806r/lib1-7/phyloseq/'
physeq_bulkCore = 'bulk-core'
physeq_SIP_core = 'SIP-core_unk'
prefrac_comm_abundance = ['1e9']
richness = 2503 # chao1 estimate for bulk Day 1
seq_per_fraction = ['lognormal', 9.432, 0.5, 10000, 30000] # dist, mean, scale, min, max
bulk_days = [1]
nprocs = 24
# building tree structure
nest = nestly.Nest()
## varying params
nest.add('abs', prefrac_comm_abundance)
## set params
nest.add('bulk_day', bulk_days, create_dir=False)
nest.add('percIncorp', [0], create_dir=False)
nest.add('percTaxa', [0], create_dir=False)
nest.add('np', [nprocs], create_dir=False)
nest.add('richness', [richness], create_dir=False)
nest.add('subsample_dist', [seq_per_fraction[0]], create_dir=False)
nest.add('subsample_mean', [seq_per_fraction[1]], create_dir=False)
nest.add('subsample_scale', [seq_per_fraction[2]], create_dir=False)
nest.add('subsample_min', [seq_per_fraction[3]], create_dir=False)
nest.add('subsample_max', [seq_per_fraction[4]], create_dir=False)
### input/output files
nest.add('buildDir', [buildDir], create_dir=False)
nest.add('R_dir', [R_dir], create_dir=False)
nest.add('fragFile', [fragFile], create_dir=False)
nest.add('targetFile', [targetFile], create_dir=False)
nest.add('physeqDir', [physeqDir], create_dir=False)
nest.add('physeq_bulkCore', [physeq_bulkCore], create_dir=False)
# building directory tree
nest.build(buildDir)
# bash file to run
bashFile = os.path.join(buildDir, 'SIPSimRun.sh')
%%writefile $bashFile
#!/bin/bash
export PATH={R_dir}:$PATH
#-- making DNA pool similar to gradient of interest
echo '# Creating comm file from phyloseq'
phyloseq2comm.r {physeqDir}{physeq_bulkCore} -s 12C-Con -d {bulk_day} > {physeq_bulkCore}_comm.txt
printf 'Number of lines: '; wc -l {physeq_bulkCore}_comm.txt
echo '## Adding target taxa to comm file'
comm_add_target.r {physeq_bulkCore}_comm.txt {targetFile} > {physeq_bulkCore}_comm_target.txt
printf 'Number of lines: '; wc -l {physeq_bulkCore}_comm_target.txt
echo '# Adding extra richness to community file'
printf "1\t{richness}\n" > richness_needed.txt
comm_add_richness.r -s {physeq_bulkCore}_comm_target.txt richness_needed.txt > {physeq_bulkCore}_comm_all.txt
### renaming comm file for downstream pipeline
cat {physeq_bulkCore}_comm_all.txt > {physeq_bulkCore}_comm_target.txt
rm -f {physeq_bulkCore}_comm_all.txt
echo '## parsing out genome fragments to make simulated DNA pool resembling the gradient of interest'
## all OTUs without an associated reference genome will be assigned a random reference (of the reference genome pool)
### this is done through --NA-random
SIPSim fragment_KDE_parse {fragFile} {physeq_bulkCore}_comm_target.txt \
--rename taxon_name --NA-random > fragsParsed.pkl
echo '#-- SIPSim pipeline --#'
echo '# converting fragments to KDE'
SIPSim fragment_KDE \
fragsParsed.pkl \
> fragsParsed_KDE.pkl
echo '# adding diffusion'
SIPSim diffusion \
fragsParsed_KDE.pkl \
--np {np} \
> fragsParsed_KDE_dif.pkl
echo '# adding DBL contamination'
SIPSim DBL \
fragsParsed_KDE_dif.pkl \
--np {np} \
> fragsParsed_KDE_dif_DBL.pkl
echo '# making incorp file'
SIPSim incorpConfigExample \
--percTaxa {percTaxa} \
--percIncorpUnif {percIncorp} \
> {percTaxa}_{percIncorp}.config
echo '# adding isotope incorporation to BD distribution'
SIPSim isotope_incorp \
fragsParsed_KDE_dif_DBL.pkl \
{percTaxa}_{percIncorp}.config \
--comm {physeq_bulkCore}_comm_target.txt \
--np {np} \
> fragsParsed_KDE_dif_DBL_inc.pkl
#echo '# calculating BD shift from isotope incorporation'
#SIPSim BD_shift \
# fragsParsed_KDE_dif_DBL.pkl \
# fragsParsed_KDE_dif_DBL_inc.pkl \
# --np {np} \
# > fragsParsed_KDE_dif_DBL_inc_BD-shift.txt
echo '# simulating gradient fractions'
SIPSim gradient_fractions \
{physeq_bulkCore}_comm_target.txt \
> fracs.txt
echo '# simulating an OTU table'
SIPSim OTU_table \
fragsParsed_KDE_dif_DBL_inc.pkl \
{physeq_bulkCore}_comm_target.txt \
fracs.txt \
--abs {abs} \
--np {np} \
> OTU_abs{abs}.txt
#echo '# simulating PCR'
SIPSim OTU_PCR \
OTU_abs{abs}.txt \
> OTU_abs{abs}_PCR.txt
echo '# subsampling from the OTU table (simulating sequencing of the DNA pool)'
SIPSim OTU_subsample \
--dist {subsample_dist} \
--dist_params mean:{subsample_mean},sigma:{subsample_scale} \
--min_size {subsample_min} \
--max_size {subsample_max} \
OTU_abs{abs}_PCR.txt \
> OTU_abs{abs}_PCR_sub.txt
echo '# making a wide-formatted table'
SIPSim OTU_wideLong -w \
OTU_abs{abs}_PCR_sub.txt \
> OTU_abs{abs}_PCR_sub_w.txt
echo '# making metadata (phyloseq: sample_data)'
SIPSim OTU_sampleData \
OTU_abs{abs}_PCR_sub.txt \
> OTU_abs{abs}_PCR_sub_meta.txt
!chmod 777 $bashFile
!cd $workDir; \
nestrun --template-file $bashFile -d Day1_default_run --log-file log.txt -j 1
```
## Checking amplicon fragment BD distribution
### 'Raw' fragments
```
workDir1 = os.path.join(workDir, 'Day1_default_run/1e9/')
!cd $workDir1; \
SIPSim KDE_info \
-s fragsParsed_KDE.pkl \
> fragsParsed_KDE_info.txt
%%R -i workDir1
inFile = file.path(workDir1, 'fragsParsed_KDE_info.txt')
df = read.delim(inFile, sep='\t') %>%
filter(KDE_ID == 1)
df %>% head(n=3)
%%R -w 600 -h 300
ggplot(df, aes(median)) +
geom_histogram(binwidth=0.001) +
labs(x='Buoyant density') +
theme_bw() +
theme(
text = element_text(size=16)
)
```
### fragments w/ diffusion + DBL
```
workDir1 = os.path.join(workDir, 'Day1_default_run/1e9/')
!cd $workDir1; \
SIPSim KDE_info \
-s fragsParsed_KDE_dif_DBL.pkl \
> fragsParsed_KDE_dif_DBL_info.pkl
%%R -i workDir1
inFile = file.path(workDir1, 'fragsParsed_KDE_dif_DBL_info.pkl')
df = read.delim(inFile, sep='\t') %>%
filter(KDE_ID == 1)
df %>% head(n=3)
%%R -w 600 -h 300
ggplot(df, aes(median)) +
geom_histogram(binwidth=0.001) +
labs(x='Buoyant density') +
theme_bw() +
theme(
text = element_text(size=16)
)
```
# BD min/max
* what is the min/max BD that we care about?
```
%%R
## min G+C cutoff
min_GC = 13.5
## max G+C cutoff
max_GC = 80
## max G+C shift
max_13C_shift_in_BD = 0.036
min_BD = min_GC/100.0 * 0.098 + 1.66
max_BD = max_GC/100.0 * 0.098 + 1.66
max_BD = max_BD + max_13C_shift_in_BD
cat('Min BD:', min_BD, '\n')
cat('Max BD:', max_BD, '\n')
```
## Plotting number of taxa in each fraction
### Emperical data (fullCyc)
```
%%R
# simulated OTU table file
OTU.table.dir = '/home/nick/notebook/SIPSim/dev/fullCyc/frag_norm_9_2.5_n5/Day1_default_run/1e9/'
OTU.table.file = 'OTU_abs1e9_PCR_sub.txt'
#OTU.table.file = 'OTU_abs1e9_sub.txt'
#OTU.table.file = 'OTU_abs1e9.txt'
%%R -i physeqDir -i physeq_SIP_core -i bulk_days
# bulk core samples
F = file.path(physeqDir, physeq_SIP_core)
physeq.SIP.core = readRDS(F)
physeq.SIP.core.m = physeq.SIP.core %>% sample_data
physeq.SIP.core = prune_samples(physeq.SIP.core.m$Substrate == '12C-Con' &
physeq.SIP.core.m$Day %in% bulk_days,
physeq.SIP.core) %>%
filter_taxa(function(x) sum(x) > 0, TRUE)
physeq.SIP.core.m = physeq.SIP.core %>% sample_data
physeq.SIP.core
%%R -w 800 -h 300
## dataframe
df.EMP = physeq.SIP.core %>% otu_table %>%
as.matrix %>% as.data.frame
df.EMP$OTU = rownames(df.EMP)
df.EMP = df.EMP %>%
gather(sample, abundance, 1:(ncol(df.EMP)-1))
df.EMP = inner_join(df.EMP, physeq.SIP.core.m, c('sample' = 'X.Sample'))
df.EMP.nt = df.EMP %>%
group_by(sample) %>%
mutate(n_taxa = sum(abundance > 0)) %>%
ungroup() %>%
distinct(sample) %>%
filter(Buoyant_density >= min_BD,
Buoyant_density <= max_BD)
## plotting
p = ggplot(df.EMP.nt, aes(Buoyant_density, n_taxa)) +
geom_point(color='blue') +
geom_line(color='blue') +
#geom_vline(xintercept=c(BD.GCp0, BD.GCp100), linetype='dashed', alpha=0.5) +
labs(x='Buoyant density', y='Number of taxa') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
p
```
### w/ simulated data
```
%%R -w 800 -h 300
# loading file
F = file.path(workDir1, OTU.table.file)
df.SIM = read.delim(F, sep='\t')
## edit table
df.SIM.nt = df.SIM %>%
filter(count > 0) %>%
group_by(library, BD_mid) %>%
summarize(n_taxa = n()) %>%
filter(BD_mid >= min_BD,
BD_mid <= max_BD)
## plot
p = ggplot(df.SIM.nt, aes(BD_mid, n_taxa)) +
geom_point(color='red') +
geom_line(color='red') +
geom_point(data=df.EMP.nt, aes(x=Buoyant_density), color='blue') +
geom_line(data=df.EMP.nt, aes(x=Buoyant_density), color='blue') +
#geom_vline(xintercept=c(BD.GCp0, BD.GCp100), linetype='dashed', alpha=0.5) +
labs(x='Buoyant density', y='Number of taxa') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
p
%%R -w 800 -h 300
# normalized by max number of taxa
## edit table
df.SIM.nt = df.SIM.nt %>%
group_by() %>%
mutate(n_taxa_norm = n_taxa / max(n_taxa))
df.EMP.nt = df.EMP.nt %>%
group_by() %>%
mutate(n_taxa_norm = n_taxa / max(n_taxa))
## plot
p = ggplot(df.SIM.nt, aes(BD_mid, n_taxa_norm)) +
geom_point(color='red') +
geom_line(color='red') +
geom_point(data=df.EMP.nt, aes(x=Buoyant_density), color='blue') +
geom_line(data=df.EMP.nt, aes(x=Buoyant_density), color='blue') +
#geom_vline(xintercept=c(BD.GCp0, BD.GCp100), linetype='dashed', alpha=0.5) +
scale_y_continuous(limits=c(0, 1)) +
labs(x='Buoyant density', y='Number of taxa\n(fraction of max)') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
p
```
## Total sequence count
```
%%R -w 800 -h 300
# simulated
df.SIM.s = df.SIM %>%
group_by(library, BD_mid) %>%
summarize(total_abund = sum(count)) %>%
rename('Day' = library, 'Buoyant_density' = BD_mid) %>%
ungroup() %>%
mutate(dataset='simulated')
# emperical
df.EMP.s = df.EMP %>%
group_by(Day, Buoyant_density) %>%
summarize(total_abund = sum(abundance)) %>%
ungroup() %>%
mutate(dataset='emperical')
# join
df.j = rbind(df.SIM.s, df.EMP.s) %>%
filter(Buoyant_density >= min_BD,
Buoyant_density <= max_BD)
df.SIM.s = df.EMP.s = ""
# plot
ggplot(df.j, aes(Buoyant_density, total_abund, color=dataset)) +
geom_point() +
geom_line() +
scale_color_manual(values=c('blue', 'red')) +
labs(x='Buoyant density', y='Total sequences per sample') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
```
## Plotting Shannon diversity for each
```
%%R
shannon_index_long = function(df, abundance_col, ...){
# calculating shannon diversity index from a 'long' formated table
## community_col = name of column defining communities
## abundance_col = name of column defining taxon abundances
df = df %>% as.data.frame
cmd = paste0(abundance_col, '/sum(', abundance_col, ')')
df.s = df %>%
group_by_(...) %>%
mutate_(REL_abundance = cmd) %>%
mutate(pi__ln_pi = REL_abundance * log(REL_abundance),
shannon = -sum(pi__ln_pi, na.rm=TRUE)) %>%
ungroup() %>%
dplyr::select(-REL_abundance, -pi__ln_pi) %>%
distinct_(...)
return(df.s)
}
%%R
# calculating shannon
df.SIM.shan = shannon_index_long(df.SIM, 'count', 'library', 'fraction') %>%
filter(BD_mid >= min_BD,
BD_mid <= max_BD)
df.EMP.shan = shannon_index_long(df.EMP, 'abundance', 'sample') %>%
filter(Buoyant_density >= min_BD,
Buoyant_density <= max_BD)
%%R -w 800 -h 300
# plotting
p = ggplot(df.SIM.shan, aes(BD_mid, shannon)) +
geom_point(color='red') +
geom_line(color='red') +
geom_point(data=df.EMP.shan, aes(x=Buoyant_density), color='blue') +
geom_line(data=df.EMP.shan, aes(x=Buoyant_density), color='blue') +
scale_y_continuous(limits=c(4, 7.5)) +
labs(x='Buoyant density', y='Shannon index') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
p
```
# min/max abundances of taxa
```
%%R -h 300 -w 800
# simulated
df.SIM.s = df.SIM %>%
filter(rel_abund > 0) %>%
group_by(BD_mid) %>%
summarize(min_abund = min(rel_abund),
max_abund = max(rel_abund)) %>%
ungroup() %>%
rename('Buoyant_density' = BD_mid) %>%
mutate(dataset = 'simulated')
# emperical
df.EMP.s = df.EMP %>%
group_by(Buoyant_density) %>%
mutate(rel_abund = abundance / sum(abundance)) %>%
filter(rel_abund > 0) %>%
summarize(min_abund = min(rel_abund),
max_abund = max(rel_abund)) %>%
ungroup() %>%
mutate(dataset = 'emperical')
df.j = rbind(df.SIM.s, df.EMP.s) %>%
filter(Buoyant_density >= min_BD,
Buoyant_density <= max_BD)
# plotting
ggplot(df.j, aes(Buoyant_density, max_abund, color=dataset, group=dataset)) +
geom_point() +
geom_line() +
scale_color_manual(values=c('blue', 'red')) +
labs(x='Buoyant density', y='Maximum relative abundance') +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none'
)
```
## Plotting rank-abundance of heavy fractions
* In heavy fractions, is DBL resulting in approx. equal abundances among taxa?
```
%%R -w 900
# simulated
df.SIM.s = df.SIM %>%
select(BD_mid, rel_abund) %>%
rename('Buoyant_density' = BD_mid) %>%
mutate(dataset='simulated')
# emperical
df.EMP.s = df.EMP %>%
group_by(Buoyant_density) %>%
mutate(rel_abund = abundance / sum(abundance)) %>%
ungroup() %>%
filter(rel_abund > 0) %>%
select(Buoyant_density, rel_abund) %>%
mutate(dataset='emperical')
# join
df.j = rbind(df.SIM.s, df.EMP.s) %>%
filter(Buoyant_density > 1.73) %>%
mutate(Buoyant_density = round(Buoyant_density, 3),
Buoyant_density_c = as.character(Buoyant_density))
df.j$Buoyant_density_c = reorder(df.j$Buoyant_density_c, df.j$Buoyant_density)
ggplot(df.j, aes(Buoyant_density_c, rel_abund)) +
geom_boxplot() +
scale_color_manual(values=c('blue', 'red')) +
labs(x='Buoyant density', y='Maximum relative abundance') +
facet_grid(dataset ~ .) +
theme_bw() +
theme(
text = element_text(size=16),
axis.text.x = element_text(angle=60, hjust=1),
legend.position = 'none'
)
```
# BD range where an OTU is detected
* Do the simulated OTU BD distributions span the same BD range of the emperical data?
### Simulated
```
%%R
# loading comm file
F = file.path(workDir1, 'bulk-core_comm_target.txt')
df.comm = read.delim(F, sep='\t') %>%
dplyr::select(library, taxon_name, rel_abund_perc) %>%
rename('bulk_abund' = rel_abund_perc) %>%
mutate(bulk_abund = bulk_abund / 100)
## joining
df.SIM.j = inner_join(df.SIM, df.comm, c('library' = 'library',
'taxon' = 'taxon_name')) %>%
filter(BD_mid >= min_BD,
BD_mid <= max_BD)
df.SIM.j %>% head(n=3)
```
### Emperical
```
%%R
bulk_days = c(1)
%%R
physeq.dir = '/var/seq_data/fullCyc/MiSeq_16SrRNA/515f-806r/lib1-7/phyloseq/'
physeq.bulk = 'bulk-core'
physeq.file = file.path(physeq.dir, physeq.bulk)
physeq.bulk = readRDS(physeq.file)
physeq.bulk.m = physeq.bulk %>% sample_data
physeq.bulk = prune_samples(physeq.bulk.m$Exp_type == 'microcosm_bulk' &
physeq.bulk.m$Day %in% bulk_days, physeq.bulk)
physeq.bulk.m = physeq.bulk %>% sample_data
physeq.bulk
%%R
physeq.bulk.n = transform_sample_counts(physeq.bulk, function(x) x/sum(x))
physeq.bulk.n
%%R
# making long format of each bulk table
bulk.otu = physeq.bulk.n %>% otu_table %>% as.data.frame
ncol = ncol(bulk.otu)
bulk.otu$OTU = rownames(bulk.otu)
bulk.otu = bulk.otu %>%
gather(sample, abundance, 1:ncol)
bulk.otu = inner_join(physeq.bulk.m, bulk.otu, c('X.Sample' = 'sample')) %>%
dplyr::select(OTU, abundance) %>%
rename('bulk_abund' = abundance)
bulk.otu %>% head(n=3)
%%R
# joining tables
df.EMP.j = inner_join(df.EMP, bulk.otu, c('OTU' = 'OTU')) %>%
filter(Buoyant_density >= min_BD,
Buoyant_density <= max_BD)
df.EMP.j %>% head(n=3)
%%R -h 400
# filtering & combining emperical w/ simulated data
## emperical
max_BD_range = max(df.EMP.j$Buoyant_density) - min(df.EMP.j$Buoyant_density)
df.EMP.j.f = df.EMP.j %>%
filter(abundance > 0) %>%
group_by(OTU) %>%
summarize(mean_rel_abund = mean(bulk_abund),
min_BD = min(Buoyant_density),
max_BD = max(Buoyant_density),
BD_range = max_BD - min_BD,
BD_range_perc = BD_range / max_BD_range * 100) %>%
ungroup() %>%
mutate(dataset = 'emperical')
## simulated
max_BD_range = max(df.SIM.j$BD_mid) - min(df.SIM.j$BD_mid)
df.SIM.j.f = df.SIM.j %>%
filter(count > 0) %>%
group_by(taxon) %>%
summarize(mean_rel_abund = mean(bulk_abund),
min_BD = min(BD_mid),
max_BD = max(BD_mid),
BD_range = max_BD - min_BD,
BD_range_perc = BD_range / max_BD_range * 100) %>%
ungroup() %>%
rename('OTU' = taxon) %>%
mutate(dataset = 'simulated')
## join
df.j = rbind(df.EMP.j.f, df.SIM.j.f) %>%
filter(BD_range_perc > 0,
mean_rel_abund > 0)
## plotting
ggplot(df.j, aes(mean_rel_abund, BD_range_perc, color=dataset)) +
geom_point(alpha=0.5, shape='O') +
#stat_density2d() +
#scale_fill_gradient(low='white', high='red', na.value='grey50') +
#scale_x_log10(limits=c(min(df.j$mean_rel_abund, na.rm=T), 1e-2)) +
#scale_y_continuous(limits=c(90, 100)) +
scale_x_log10() +
scale_y_continuous() +
scale_color_manual(values=c('blue', 'red')) +
labs(x='Pre-fractionation abundance', y='% of total BD range') +
#geom_vline(xintercept=0.001, linetype='dashed', alpha=0.5) +
facet_grid(dataset ~ .) +
theme_bw() +
theme(
text = element_text(size=16),
panel.grid = element_blank(),
legend.position = 'none'
)
```
## BD span of just overlapping taxa
* taxa overlapping between emperical data and genomes in dataset
```
%%R -i targetFile
df.target = read.delim(targetFile, sep='\t')
df.target %>% nrow %>% print
df.target %>% head(n=3)
%%R
# filtering to just target taxa
df.j.t = df.j %>%
filter(OTU %in% df.target$OTU)
## plotting
ggplot(df.j.t, aes(mean_rel_abund, BD_range_perc, color=dataset)) +
geom_point(alpha=0.5, shape='O') +
#stat_density2d() +
#scale_fill_gradient(low='white', high='red', na.value='grey50') +
#scale_x_log10(limits=c(min(df.j$mean_rel_abund, na.rm=T), 1e-2)) +
#scale_y_continuous(limits=c(90, 100)) +
scale_x_log10() +
scale_y_continuous() +
scale_color_manual(values=c('blue', 'red')) +
labs(x='Pre-fractionation abundance', y='% of total BD range') +
#geom_vline(xintercept=0.001, linetype='dashed', alpha=0.5) +
facet_grid(dataset ~ .) +
theme_bw() +
theme(
text = element_text(size=16),
panel.grid = element_blank(),
legend.position = 'none'
)
```
# Plotting abundance distributions
```
%%R
## emperical
df.EMP.j.f = df.EMP.j %>%
filter(abundance > 0) %>%
dplyr::select(OTU, sample, abundance, Buoyant_density, bulk_abund) %>%
mutate(dataset = 'emperical')
## simulated
df.SIM.j.f = df.SIM.j %>%
filter(count > 0) %>%
dplyr::select(taxon, fraction, count, BD_mid, bulk_abund) %>%
rename('OTU' = taxon,
'sample' = fraction,
'Buoyant_density' = BD_mid,
'abundance' = count) %>%
mutate(dataset = 'simulated')
df.j = rbind(df.EMP.j.f, df.SIM.j.f) %>%
group_by(sample) %>%
mutate(rel_abund = abundance / sum(abundance))
df.j %>% head(n=3) %>% as.data.frame
%%R -w 800 -h 400
# plotting absolute abundances of subsampled
## plot
p = ggplot(df.j, aes(Buoyant_density, abundance, fill=OTU)) +
geom_area(stat='identity', position='dodge', alpha=0.5) +
#geom_vline(xintercept=c(BD.GCp0, BD.GCp100), linetype='dashed', alpha=0.5) +
labs(x='Buoyant density', y='Subsampled community\n(absolute abundance)') +
facet_grid(dataset ~ .) +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none',
axis.title.y = element_text(vjust=1),
axis.title.x = element_blank(),
plot.margin=unit(c(0.1,1,0.1,1), "cm")
)
p
%%R -w 800 -h 400
# plotting relative abundances of subsampled
p = ggplot(df.j, aes(Buoyant_density, rel_abund, fill=OTU)) +
geom_area(stat='identity', position='dodge', alpha=0.5) +
#geom_vline(xintercept=c(BD.GCp0, BD.GCp100), linetype='dashed', alpha=0.5) +
labs(x='Buoyant density', y='Subsampled community\n(relative abundance)') +
facet_grid(dataset ~ .) +
theme_bw() +
theme(
text = element_text(size=16),
legend.position = 'none',
axis.title.y = element_text(vjust=1),
axis.title.x = element_blank(),
plot.margin=unit(c(0.1,1,0.1,1), "cm")
)
p
```
***
***
# --OLD--
# Determining the pre-fractionation abundances of taxa in each gradient fraction
* emperical data
* low-abundant taxa out at the tails?
* OR broad distributions of high abundant taxa
```
%%R
physeq.SIP.core.n = transform_sample_counts(physeq.SIP.core, function(x) x/sum(x))
physeq.SIP.core.n
%%R
physeq.dir = '/var/seq_data/fullCyc/MiSeq_16SrRNA/515f-806r/lib1-7/phyloseq/'
physeq.bulk = 'bulk-core'
physeq.file = file.path(physeq.dir, physeq.bulk)
physeq.bulk = readRDS(physeq.file)
physeq.bulk.m = physeq.bulk %>% sample_data
physeq.bulk = prune_samples(physeq.bulk.m$Exp_type == 'microcosm_bulk' &
physeq.bulk.m$Day %in% bulk_days, physeq.bulk)
physeq.bulk.m = physeq.bulk %>% sample_data
physeq.bulk
%%R
physeq.bulk.n = transform_sample_counts(physeq.bulk, function(x) x/sum(x))
physeq.bulk.n
%%R
# making long format of SIP OTU table
SIP.otu = physeq.SIP.core.n %>% otu_table %>% as.data.frame
ncol = ncol(SIP.otu)
SIP.otu$OTU = rownames(SIP.otu)
SIP.otu = SIP.otu %>%
gather(sample, abundance, 1:ncol)
SIP.otu = inner_join(physeq.SIP.core.m, SIP.otu, c('X.Sample' = 'sample')) %>%
select(-core_dataset, -Sample_location, -Sample_date, -Sample_treatment,
-Sample_subtreatment, -library, -Sample_type)
SIP.otu %>% head(n=3)
%%R
# making long format of each bulk table
bulk.otu = physeq.bulk.n %>% otu_table %>% as.data.frame
ncol = ncol(bulk.otu)
bulk.otu$OTU = rownames(bulk.otu)
bulk.otu = bulk.otu %>%
gather(sample, abundance, 1:ncol)
bulk.otu = inner_join(physeq.bulk.m, bulk.otu, c('X.Sample' = 'sample')) %>%
select(OTU, abundance) %>%
rename('bulk_abund' = abundance)
bulk.otu %>% head(n=3)
%%R
# joining tables
SIP.otu = inner_join(SIP.otu, bulk.otu, c('OTU' = 'OTU'))
SIP.otu %>% head(n=3)
%%R -w 900 -h 900
# for each gradient, plotting gradient rel_abund vs bulk rel_abund
ggplot(SIP.otu, aes(bulk_abund, abundance)) +
geom_point(alpha=0.2) +
geom_point(shape='O', alpha=0.6) +
facet_wrap(~ Buoyant_density) +
labs(x='Pre-fractionation relative abundance',
y='Fraction relative abundance') +
theme_bw() +
theme(
text = element_text(size=16)
)
%%R -w 900 -h 900
# for each gradient, plotting gradient rel_abund vs bulk rel_abund
ggplot(SIP.otu, aes(bulk_abund, abundance)) +
geom_point(alpha=0.2) +
geom_point(shape='O', alpha=0.6) +
scale_x_continuous(limits=c(0,0.01)) +
scale_y_continuous(limits=c(0,0.01)) +
facet_wrap(~ Buoyant_density) +
labs(x='Pre-fractionation relative abundance',
y='Fraction relative abundance') +
theme_bw() +
theme(
text = element_text(size=16),
axis.text.x = element_text(angle=90, hjust=1, vjust=0.5)
)
```
## Plotting the abundance distribution of top 10 most abundant taxa (bulk samples)
```
%%R -w 500 -h 300
# checking bulk rank-abundance
tmp = bulk.otu %>%
mutate(rank = row_number(-bulk_abund))
ggplot(tmp, aes(rank, bulk_abund)) +
geom_point()
%%R -w 900
top.n = filter(tmp, rank <= 10)
SIP.otu.f = SIP.otu %>%
filter(OTU %in% top.n$OTU)
ggplot(SIP.otu.f, aes(Buoyant_density, abundance, group=OTU, fill=OTU)) +
#geom_point() +
#geom_line() +
geom_area(position='dodge', alpha=0.4) +
labs(y='Relative abundance', x='Buoyant density') +
theme_bw() +
theme(
text = element_text(size=16)
)
%%R -w 600 -h 400
# Number of gradients that each OTU is found in
max_BD_range = max(SIP.otu$Buoyant_density) - min(SIP.otu$Buoyant_density)
SIP.otu.f = SIP.otu %>%
filter(abundance > 0) %>%
group_by(OTU) %>%
summarize(bulk_abund = mean(bulk_abund),
min_BD = min(Buoyant_density),
max_BD = max(Buoyant_density),
BD_range = max_BD - min_BD,
BD_range_perc = BD_range / max_BD_range * 100) %>%
ungroup()
ggplot(SIP.otu.f, aes(bulk_abund, BD_range_perc, group=OTU)) +
geom_point() +
scale_x_log10() +
labs(x='Pre-fractionation abundance', y='% of total BD range') +
geom_vline(xintercept=0.001, linetype='dashed', alpha=0.5) +
theme_bw() +
theme(
text = element_text(size=16)
)
```
#### Notes
* Currently, simulated taxa with an abundance of >0.1% are detected in approx. all fractions.
| github_jupyter |
# String Key Hash Table
### Problem Statement
In this quiz, you'll write your own hash table and hash function that uses string keys. Your table will store strings in the buckets. The (bucket) index is calculated by the first two letters of the string, according to the formula below:
Hash Value = (ASCII Value of First Letter * 100) + ASCII Value of Second Letter
In the formula above, the generated hash value is the (bucket) index.
**Example**: For a string "UDACITY", the ASCII value for letters 'U' and 'D' are 85 and 68 respectively. The hash value would be: `(85 *100) + 68 = 8568`.
You can use the Python function `ord()` to get the ASCII value of a letter, and `chr()` to get the letter associated with an ASCII value.
**Assumptions**
1. The string will have at least two letters,
2. The first two characters are uppercase letters (ASCII values from 65 to 90).
**Rules**
- Do not use a Python dictionary—only lists!
- Store `lists` at each bucket, and not just the string itself. For example, you can store "UDACITY" at index 8568 as ["UDACITY"].
### Instructions
Create a `HashTable` class, with the following functions:
- `store()` - a function that takes a string as input, and stores it into the hash table.
- `lookup()` - a function that checks if a string is already available in the hash table. If yes, return the hash value, else return -1.
- `calculate_hash_value()` - a helper function to calculate a hash value of a given string.
### Exercise - Try building a string hash table!
```
"""Write a HashTable class that stores strings
in a hash table, where keys are calculated
using the first two letters of the string."""
class HashTable(object):
def __init__(self):
# Table size
# ord('Z') * 100 + ord('Z') -> 9090
self.table = [None]*10000
def store(self, string):
"""TODO: Input a string that's stored in
the table."""
hash_code = self.calculate_hash_value(string)
if self.table[hash_code]:
self.table[hash_code].append(string)
else:
self.table[hash_code] = [string]
def lookup(self, string):
"""TODO: Return the hash value if the
string is already in the table.
Return -1 otherwise."""
hash_code = self.calculate_hash_value(string)
if self.table[hash_code]:
if string in self.table[hash_code]:
return hash_code
return -1
def calculate_hash_value(self, string):
"""TODO: Helper function to calulate a
hash value from a string."""
hash_code = ord(string[0])*100 + ord(string[1])
return hash_code
```
### Test Cases - Let's test your function
```
# Setup
hash_table = HashTable()
# Test calculate_hash_value
print (hash_table.calculate_hash_value('UDACITY')) # Should be 8568
# Test lookup edge case
print (hash_table.lookup('UDACITY')) # Should be -1
# Test store
hash_table.store('UDACITY')
print (hash_table.lookup('UDACITY')) # Should be 8568
# Test store edge case
hash_table.store('UDACIOUS')
print (hash_table.lookup('UDACIOUS')) # Should be 8568
```
<span class="graffiti-highlight graffiti-id_53gqd1t-id_55iwcxy"><i></i><button>Show Solution</button></span>
| github_jupyter |
Deep Learning
=============
Assignment 2
------------
Previously in `1_notmnist.ipynb`, we created a pickle with formatted datasets for training, development and testing on the [notMNIST dataset](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html).
The goal of this assignment is to progressively train deeper and more accurate models using TensorFlow.
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
```
First reload the data we generated in `1_notmnist.ipynb`.
```
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
```
Reformat into a shape that's more adapted to the models we're going to train:
- data as a flat matrix,
- labels as float 1-hot encodings.
```
image_size = 28
num_labels = 10
def reformat(dataset, labels):
dataset = dataset.reshape((-1, image_size * image_size)).astype(np.float32)
# Map 0 to [1.0, 0.0, 0.0 ...], 1 to [0.0, 1.0, 0.0 ...]
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
```
We're first going to train a multinomial logistic regression using simple gradient descent.
TensorFlow works like this:
* First you describe the computation that you want to see performed: what the inputs, the variables, and the operations look like. These get created as nodes over a computation graph. This description is all contained within the block below:
with graph.as_default():
...
* Then you can run the operations on this graph as many times as you want by calling `session.run()`, providing it outputs to fetch from the graph that get returned. This runtime operation is all contained in the block below:
with tf.Session(graph=graph) as session:
...
Let's load all the data into TensorFlow and build the computation graph corresponding to our training:
```
# With gradient descent training, even this much data is prohibitive.
# Subset the training data for faster turnaround.
train_subset = 10000
graph = tf.Graph()
with graph.as_default():
# Input data.
# Load the training, validation and test data into constants that are
# attached to the graph.
tf_train_dataset = tf.constant(train_dataset[:train_subset, :])
tf_train_labels = tf.constant(train_labels[:train_subset])
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
# These are the parameters that we are going to be training. The weight
# matrix will be initialized using random values following a (truncated)
# normal distribution. The biases get initialized to zero.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
# We multiply the inputs with the weight matrix, and add biases. We compute
# the softmax and cross-entropy (it's one operation in TensorFlow, because
# it's very common, and it can be optimized). We take the average of this
# cross-entropy across all training examples: that's our loss.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
# We are going to find the minimum of this loss using gradient descent.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
# These are not part of training, but merely here so that we can report
# accuracy figures as we train.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
```
Let's run this computation and iterate:
```
num_steps = 801
with tf.Session(graph=graph) as session:
# This is a one-time operation which ensures the parameters get initialized as
# we described in the graph: random weights for the matrix, zeros for the
# biases.
tf.global_variables_initializer().run()
print('Initialized')
for step in range(num_steps):
# Run the computations. We tell .run() that we want to run the optimizer,
# and get the loss value and the training predictions returned as numpy
# arrays.
_, l, predictions = session.run([optimizer, loss, train_prediction])
if (step % 100 == 0):
print('Loss at step %d: %f' % (step, l))
print('Training accuracy: %.1f%%' % accuracy(
predictions, train_labels[:train_subset, :]))
# Calling .eval() on valid_prediction is basically like calling run(), but
# just to get that one numpy array. Note that it recomputes all its graph
# dependencies.
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
```
Let's now switch to stochastic gradient descent training instead, which is much faster.
The graph will be similar, except that instead of holding all the training data into a constant node, we create a `Placeholder` node which will be fed actual data at every call of `session.run()`.
```
batch_size = 128
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
```
Let's run it:
```
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
```
---
Problem
-------
Turn the logistic regression example with SGD into a 1-hidden layer neural network with rectified linear units [nn.relu()](https://www.tensorflow.org/versions/r0.7/api_docs/python/nn.html#relu) and 1024 hidden nodes. This model should improve your validation / test accuracy.
---
```
batch_size = 128
hidden_layer_size = 1024
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights_hidden = tf.Variable(
tf.truncated_normal([image_size * image_size, hidden_layer_size]))
biases_hidden = tf.Variable(tf.zeros([hidden_layer_size]))
weights = tf.Variable(
tf.truncated_normal([hidden_layer_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
hidden_layer_output = tf.nn.relu(tf.matmul(tf_train_dataset, weights_hidden)
+ biases_hidden)
logits = tf.matmul(hidden_layer_output, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf.nn.relu(tf.matmul(tf_valid_dataset, weights_hidden)
+ biases_hidden),weights) + biases)
test_prediction = tf.nn.softmax(
tf.matmul(tf.nn.relu(tf.matmul(tf_test_dataset, weights_hidden)
+ biases_hidden), weights) + biases)
num_steps = 3001
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
```
| github_jupyter |
```
#IMPORT SEMUA LIBRARY DISINI
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY POSTGRESQL
import psycopg2
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY PDF
from fpdf import FPDF
#IMPORT LIBRARY BASEPATH
import io
#IMPORT LIBRARY BASE64 IMG
import base64
#IMPORT LIBRARY NUMPY
import numpy as np
#IMPORT LIBRARY EXCEL
import xlsxwriter
#IMPORT LIBRARY SIMILARITAS
import n0similarities as n0
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
def uploadToPSQL(host, username, password, database, port, table, judul, filePath, name, subjudul, dataheader, databody):
#TEST KONEKSI KE DATABASE
try:
for t in range(0, len(table)):
#DATA DIJADIKAN LIST
rawstr = [tuple(x) for x in zip(dataheader, databody[t])]
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=database)
cursor = connection.cursor()
connection.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT);
#CEK TABLE
cursor.execute("SELECT * FROM information_schema.tables where table_name=%s", (table[t],))
exist = bool(cursor.rowcount)
#KALAU ADA DIHAPUS DULU, TERUS DICREATE ULANG
if exist == True:
cursor.execute("DROP TABLE "+ table[t] + " CASCADE")
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#KALAU GA ADA CREATE DATABASE
else:
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#MASUKAN DATA KE DATABASE YANG TELAH DIBUAT
cursor.execute('INSERT INTO '+table[t]+'(tanggal, total) values ' +str(rawstr)[1:-1])
#JIKA BERHASIL SEMUA AKAN MENGHASILKAN KELUARAN BENAR (TRUE)
return True
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
return error
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
def makeChart(host, username, password, db, port, table, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath):
try:
datarowsend = []
for t in range(0, len(table)):
#TEST KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBIL DATA DARI DATABASE DENGAN LIMIT YANG SUDAH DIKIRIMKAN DARI VARIABLE DIBAWAH
postgreSQL_select_Query = "SELECT * FROM "+table[t]+" ORDER BY tanggal DESC LIMIT " + str(limitdata)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MENYIMPAN DATA DARI DATABASE KE DALAM VARIABLE
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
lengthy.append(row[2])
datarowsend.append(mobile_records)
#JUDUL CHART
judulgraf = A2 + " " + wilayah[t]
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#DATA CHART DIMASUKAN DISINI
ax.bar(uid, lengthy, align='center')
#JUDUL CHART
ax.set_title(judulgraf)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#BUAT CHART MENJADI FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART DIJADIKAN BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#line
#DATA CHART DIMASUKAN DISINI
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#JUDUL CHART
plt.title(judulgraf)
plt.grid(True)
l = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(l, format='png', bbox_inches="tight")
#GAMBAR DIJADIKAN BAS64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#pie
#JUDUL CHART
plt.title(judulgraf)
#DATA CHART DIMASUKAN DISINI
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.plot(legend=None)
plt.axis('equal')
p = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(p, format='png', bbox_inches="tight")
#CHART DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#CHART DISIMPAN KE DIREKTORI DIJADIKAN FORMAT PNG
#BARCHART
bardata = base64.b64decode(barChart)
barname = basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[t]+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINECHART
linedata = base64.b64decode(lineChart)
linename = basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[t]+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIECHART
piedata = base64.b64decode(pieChart)
piename = basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[t]+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#MEMANGGIL FUNGSI EXCEL
makeExcel(datarowsend, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, name, limitdata, table, wilayah, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(datarowsend, judul, barChart, lineChart, pieChart, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, table, wilayah, basePath)
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
print (error)
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
def makePDF(datarow, judul, bar, line, pie, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, lengthPDF, table, wilayah, basePath):
#PDF DIATUR DENGAN SIZE A4 DAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#TAMBAH HALAMAN PDF
pdf.add_page()
#SET FONT DAN JUGA PADDING
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#TAMPILKAN JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#SET FONT DAN JUGA PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#TAMPILKAN SUB JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#BUAT GARIS DIBAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
pdf.set_font('Times','B',11.0)
pdf.ln(0.5)
th1 = pdf.font_size
#BUAT TABLE DATA DATA DI DPF
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, A2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Region", border=1, align='C')
pdf.cell(177, 2*th1, B2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Frekuensi", border=1, align='C')
pdf.cell(177, 2*th1, C2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Unit", border=1, align='C')
pdf.cell(177, 2*th1, D2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Sumber", border=1, align='C')
pdf.cell(177, 2*th1, E2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Status", border=1, align='C')
pdf.cell(177, 2*th1, F2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "ID Seri", border=1, align='C')
pdf.cell(177, 2*th1, G2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Kode SR", border=1, align='C')
pdf.cell(177, 2*th1, H2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Pertama", border=1, align='C')
pdf.cell(177, 2*th1, str(I2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Terakhir ", border=1, align='C')
pdf.cell(177, 2*th1, str(J2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Waktu pembaruan terakhir", border=1, align='C')
pdf.cell(177, 2*th1, str(K2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.set_xy(17.0, 125.0)
pdf.set_font('Times','B',11.0)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
pdf.ln(0.5)
th = pdf.font_size
#HEADER TABLE DATA F2
pdf.cell(col_width, 2*th, str("Wilayah"), border=1, align='C')
#TANGAL HEADER DI LOOPING
for row in datarow[0]:
pdf.cell(col_width, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#ISI TABLE F2
for w in range(0, len(table)):
data=list(datarow[w])
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(col_width, 2*th, wilayah[w], border=1, align='C')
#DATA BERDASARKAN TANGGAL
for row in data:
pdf.cell(col_width, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#PEMANGGILAN GAMBAR
for s in range(0, len(table)):
col = pdf.w - 2*pdf.l_margin
pdf.ln(2*th)
widthcol = col/3
#TAMBAH HALAMAN
pdf.add_page()
#DATA GAMBAR BERDASARKAN DIREKTORI DIATAS
pdf.image(basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[s]+'-bar.png', link='', type='',x=8, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[s]+'-line.png', link='', type='',x=103, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/15. Sektor Pertanian/img/'+name+''+table[s]+'-pie.png', link='', type='',x=195, y=80, w=widthcol)
pdf.ln(4*th)
#PDF DIBUAT
pdf.output(basePath+'jupyter/CEIC/15. Sektor Pertanian/pdf/'+A2+'.pdf', 'F')
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
def makeExcel(datarow, A2, B2, C2, D2, E2, F2, G2, H2, I2, J2, K2, name, limit, table, wilayah, basePath):
#BUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/CEIC/15. Sektor Pertanian/excel/'+A2+'.xlsx')
#BUAT WORKSHEET EXCEL
worksheet = workbook.add_worksheet('sheet1')
#SETTINGAN UNTUK BORDER DAN FONT BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#HEADER UNTUK TABLE EXCEL F2
header = ["Wilayah", "Kategori","Region","Frekuensi","Unit","Sumber","Status","ID Seri","Kode SR","Tanggal Obs. Pertama","Tanggal Obs. Terakhir ","Waktu pembaruan terakhir"]
#DATA DATA DITAMPUNG PADA VARIABLE
for rowhead2 in datarow[0]:
header.append(str(rowhead2[1]))
#DATA HEADER DARI VARIABLE DIMASUKAN KE SINI UNTUK DITAMPILKAN BERDASARKAN ROW DAN COLUMN
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
#DATA ISI TABLE F2 DITAMPILKAN DISINI
for w in range(0, len(table)):
data=list(datarow[w])
body = [wilayah[w], A2, B2, C2, D2, E2, F2, G2, H2, str(I2.date()), str(J2.date()), str(K2.date())]
for rowbody2 in data:
body.append(str(rowbody2[2]))
for col_num, data in enumerate(body):
worksheet.write(w+1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
#BASE PATH UNTUK NANTINYA MENGCREATE FILE ATAU MEMANGGIL FILE
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE SIMILARITY WILAYAH
filePathwilayah = basePath+'data mentah/CEIC/allwilayah.xlsx';
#BACA FILE EXCEL DENGAN PANDAS
readexcelwilayah = pd.read_excel(filePathwilayah)
dfwilayah = list(readexcelwilayah.values)
readexcelwilayah.fillna(0)
allwilayah = []
#PEMILIHAN JENIS DATA, APA DATA ITU PROVINSI, KABUPATEN, KECAMATAN ATAU KELURAHAN
tipewilayah = 'prov'
if tipewilayah == 'prov':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][1])
elif tipewilayah=='kabkot':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][3])
elif tipewilayah == 'kec':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][5])
elif tipewilayah == 'kel':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][7])
semuawilayah = list(set(allwilayah))
#SETTING VARIABLE UNTUK DATABASE DAN DATA YANG INGIN DIKIRIMKAN KE FUNGSI DISINI
name = "02. Pertanian Produksi (RIB001-RIB044) Part 1"
host = "localhost"
username = "postgres"
password = "1234567890"
port = "5432"
database = "ceic"
judul = "Produk Domestik Bruto (AA001-AA007)"
subjudul = "Badan Perencanaan Pembangunan Nasional"
filePath = basePath+'data mentah/CEIC/15. Sektor Pertanian/'+name+'.xlsx';
limitdata = int(8)
readexcel = pd.read_excel(filePath)
tabledata = []
wilayah = []
databody = []
#DATA EXCEL DIBACA DISINI DENGAN MENGGUNAKAN PANDAS
df = list(readexcel.values)
head = list(readexcel)
body = list(df[0])
readexcel.fillna(0)
#PILIH ROW DATA YANG INGIN DITAMPILKAN
rangeawal = 106
rangeakhir = 107
rowrange = range(rangeawal, rangeakhir)
#INI UNTUK MEMFILTER APAKAH DATA YANG DIPILIH MEMILIKI SIMILARITAS ATAU TIDAK
#ISIKAN 'WILAYAH' UNTUK SIMILARITAS
#ISIKAN BUKAN WILAYAH JIKA BUKAN WILAYAH
jenisdata = "Indonesia"
#ROW DATA DI LOOPING UNTUK MENDAPATKAN SIMILARITAS WILAYAH
#JIKA VARIABLE JENISDATA WILAYAH AKAN MASUK KESINI
if jenisdata == 'Wilayah':
for x in rowrange:
rethasil = 0
big_w = 0
for w in range(0, len(semuawilayah)):
namawilayah = semuawilayah[w].lower().strip()
nama_wilayah_len = len(namawilayah)
hasil = n0.get_levenshtein_similarity(df[x][0].lower().strip()[nama_wilayah_len*-1:], namawilayah)
if hasil > rethasil:
rethasil = hasil
big_w = w
wilayah.append(semuawilayah[big_w].capitalize())
tabledata.append('produkdomestikbruto_'+semuawilayah[big_w].lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#JIKA BUKAN WILAYAH MASUK KESINI
else:
for x in rowrange:
wilayah.append(jenisdata.capitalize())
tabledata.append('produkdomestikbruto_'+jenisdata.lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#HEADER UNTUK PDF DAN EXCEL
A2 = "Data Migas"
B2 = df[rangeawal][1]
C2 = df[rangeawal][2]
D2 = df[rangeawal][3]
E2 = df[rangeawal][4]
F2 = df[rangeawal][5]
G2 = df[rangeawal][6]
H2 = df[rangeawal][7]
I2 = df[rangeawal][8]
J2 = df[rangeawal][9]
K2 = df[rangeawal][10]
#DATA ISI TABLE F2
dataheader = []
for listhead in head[11:]:
dataheader.append(str(listhead))
#FUNGSI UNTUK UPLOAD DATA KE SQL, JIKA BERHASIL AKAN MAMANGGIL FUNGSI UPLOAD CHART
sql = uploadToPSQL(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, dataheader, databody)
if sql == True:
makeChart(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath)
else:
print(sql)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Terrain/us_ned_chili.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/us_ned_chili.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Datasets/Terrain/us_ned_chili.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/us_ned_chili.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for this first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
dataset = ee.Image('CSP/ERGo/1_0/US/CHILI')
usChili = dataset.select('constant')
usChiliVis = {
'min': 0.0,
'max': 255.0,
}
Map.setCenter(-105.8636, 40.3439, 11)
Map.addLayer(usChili, usChiliVis, 'US CHILI')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Federated Learning for Image Classification
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/federated/tutorials/federated_learning_for_image_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/federated/blob/master/docs/tutorials/federated_learning_for_image_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/federated/blob/master/docs/tutorials/federated_learning_for_image_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
**NOTE**: This colab has been verified to work with the [latest released version](https://github.com/tensorflow/federated#compatibility) of the `tensorflow_federated` pip package, but the Tensorflow Federated project is still in pre-release development and may not work on `master`.
In this tutorial, we use the classic MNIST training example to introduce the
Federated Learning (FL) API layer of TFF, `tff.learning` - a set of
higher-level interfaces that can be used to perform common types of federated
learning tasks, such as federated training, against user-supplied models
implemented in TensorFlow.
This tutorial, and the Federated Learning API, are intended primarily for users
who want to plug their own TensorFlow models into TFF, treating the latter
mostly as a black box. For a more in-depth understanding of TFF and how to
implement your own federated learning algorithms, see the tutorials on the FC Core API - [Custom Federated Algorithms Part 1](custom_federated_algorithms_1.ipynb) and [Part 2](custom_federated_algorithms_2.ipynb).
For more on `tff.learning`, continue with the
[Federated Learning for Text Generation](federated_learning_for_text_generation.ipynb),
tutorial which in addition to covering recurrent models, also demonstrates loading a
pre-trained serialized Keras model for refinement with federated learning
combined with evaluation using Keras.
## Before we start
Before we start, please run the following to make sure that your environment is
correctly setup. If you don't see a greeting, please refer to the
[Installation](../install.md) guide for instructions.
```
#@test {"skip": true}
!pip install --quiet --upgrade tensorflow_federated_nightly
!pip install --quiet --upgrade nest_asyncio
import nest_asyncio
nest_asyncio.apply()
%load_ext tensorboard
import collections
import numpy as np
import tensorflow as tf
import tensorflow_federated as tff
np.random.seed(0)
tff.federated_computation(lambda: 'Hello, World!')()
```
## Preparing the input data
Let's start with the data. Federated learning requires a federated data set,
i.e., a collection of data from multiple users. Federated data is typically
non-[i.i.d.](https://en.wikipedia.org/wiki/Independent_and_identically_distributed_random_variables),
which poses a unique set of challenges.
In order to facilitate experimentation, we seeded the TFF repository with a few
datasets, including a federated version of MNIST that contains a version of the [original NIST dataset](https://www.nist.gov/srd/nist-special-database-19) that has been re-processed using [Leaf](https://github.com/TalwalkarLab/leaf) so that the data is keyed by the original writer of the digits. Since each writer has a unique style, this dataset exhibits the kind of non-i.i.d. behavior expected of federated datasets.
Here's how we can load it.
```
emnist_train, emnist_test = tff.simulation.datasets.emnist.load_data()
```
The data sets returned by `load_data()` are instances of
`tff.simulation.ClientData`, an interface that allows you to enumerate the set
of users, to construct a `tf.data.Dataset` that represents the data of a
particular user, and to query the structure of individual elements. Here's how
you can use this interface to explore the content of the data set. Keep in mind
that while this interface allows you to iterate over clients ids, this is only a
feature of the simulation data. As you will see shortly, client identities are
not used by the federated learning framework - their only purpose is to allow
you to select subsets of the data for simulations.
```
len(emnist_train.client_ids)
emnist_train.element_type_structure
example_dataset = emnist_train.create_tf_dataset_for_client(
emnist_train.client_ids[0])
example_element = next(iter(example_dataset))
example_element['label'].numpy()
from matplotlib import pyplot as plt
plt.imshow(example_element['pixels'].numpy(), cmap='gray', aspect='equal')
plt.grid(False)
_ = plt.show()
```
### Exploring heterogeneity in federated data
Federated data is typically non-[i.i.d.](https://en.wikipedia.org/wiki/Independent_and_identically_distributed_random_variables), users typically have different distributions of data depending on usage patterns. Some clients may have fewer training examples on device, suffering from data paucity locally, while some clients will have more than enough training examples. Let's explore this concept of data heterogeneity typical of a federated system with the EMNIST data we have available. It's important to note that this deep analysis of a client's data is only available to us because this is a simulation environment where all the data is available to us locally. In a real production federated environment you would not be able to inspect a single client's data.
First, let's grab a sampling of one client's data to get a feel for the examples on one simulated device. Because the dataset we're using has been keyed by unique writer, the data of one client represents the handwriting of one person for a sample of the digits 0 through 9, simulating the unique "usage pattern" of one user.
```
## Example MNIST digits for one client
figure = plt.figure(figsize=(20, 4))
j = 0
for example in example_dataset.take(40):
plt.subplot(4, 10, j+1)
plt.imshow(example['pixels'].numpy(), cmap='gray', aspect='equal')
plt.axis('off')
j += 1
```
Now let's visualize the number of examples on each client for each MNIST digit label. In the federated environment, the number of examples on each client can vary quite a bit, depending on user behavior.
```
# Number of examples per layer for a sample of clients
f = plt.figure(figsize=(12, 7))
f.suptitle('Label Counts for a Sample of Clients')
for i in range(6):
client_dataset = emnist_train.create_tf_dataset_for_client(
emnist_train.client_ids[i])
plot_data = collections.defaultdict(list)
for example in client_dataset:
# Append counts individually per label to make plots
# more colorful instead of one color per plot.
label = example['label'].numpy()
plot_data[label].append(label)
plt.subplot(2, 3, i+1)
plt.title('Client {}'.format(i))
for j in range(10):
plt.hist(
plot_data[j],
density=False,
bins=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
```
Now let's visualize the mean image per client for each MNIST label. This code will produce the mean of each pixel value for all of the user's examples for one label. We'll see that one client's mean image for a digit will look different than another client's mean image for the same digit, due to each person's unique handwriting style. We can muse about how each local training round will nudge the model in a different direction on each client, as we're learning from that user's own unique data in that local round. Later in the tutorial we'll see how we can take each update to the model from all the clients and aggregate them together into our new global model, that has learned from each of our client's own unique data.
```
# Each client has different mean images, meaning each client will be nudging
# the model in their own directions locally.
for i in range(5):
client_dataset = emnist_train.create_tf_dataset_for_client(
emnist_train.client_ids[i])
plot_data = collections.defaultdict(list)
for example in client_dataset:
plot_data[example['label'].numpy()].append(example['pixels'].numpy())
f = plt.figure(i, figsize=(12, 5))
f.suptitle("Client #{}'s Mean Image Per Label".format(i))
for j in range(10):
mean_img = np.mean(plot_data[j], 0)
plt.subplot(2, 5, j+1)
plt.imshow(mean_img.reshape((28, 28)))
plt.axis('off')
```
User data can be noisy and unreliably labeled. For example, looking at Client #2's data above, we can see that for label 2, it is possible that there may have been some mislabeled examples creating a noisier mean image.
### Preprocessing the input data
Since the data is already a `tf.data.Dataset`, preprocessing can be accomplished using Dataset transformations. Here, we flatten the `28x28` images
into `784`-element arrays, shuffle the individual examples, organize them into batches, and renames the features
from `pixels` and `label` to `x` and `y` for use with Keras. We also throw in a
`repeat` over the data set to run several epochs.
```
NUM_CLIENTS = 10
NUM_EPOCHS = 5
BATCH_SIZE = 20
SHUFFLE_BUFFER = 100
PREFETCH_BUFFER= 10
def preprocess(dataset):
def batch_format_fn(element):
"""Flatten a batch `pixels` and return the features as an `OrderedDict`."""
return collections.OrderedDict(
x=tf.reshape(element['pixels'], [-1, 784]),
y=tf.reshape(element['label'], [-1, 1]))
return dataset.repeat(NUM_EPOCHS).shuffle(SHUFFLE_BUFFER).batch(
BATCH_SIZE).map(batch_format_fn).prefetch(PREFETCH_BUFFER)
```
Let's verify this worked.
```
preprocessed_example_dataset = preprocess(example_dataset)
sample_batch = tf.nest.map_structure(lambda x: x.numpy(),
next(iter(preprocessed_example_dataset)))
sample_batch
```
We have almost all the building blocks in place to construct federated data
sets.
One of the ways to feed federated data to TFF in a simulation is simply as a
Python list, with each element of the list holding the data of an individual
user, whether as a list or as a `tf.data.Dataset`. Since we already have
an interface that provides the latter, let's use it.
Here's a simple helper function that will construct a list of datasets from the
given set of users as an input to a round of training or evaluation.
```
def make_federated_data(client_data, client_ids):
return [
preprocess(client_data.create_tf_dataset_for_client(x))
for x in client_ids
]
```
Now, how do we choose clients?
In a typical federated training scenario, we are dealing with potentially a very
large population of user devices, only a fraction of which may be available for
training at a given point in time. This is the case, for example, when the
client devices are mobile phones that participate in training only when plugged
into a power source, off a metered network, and otherwise idle.
Of course, we are in a simulation environment, and all the data is locally
available. Typically then, when running simulations, we would simply sample a
random subset of the clients to be involved in each round of training, generally
different in each round.
That said, as you can find out by studying the paper on the
[Federated Averaging](https://arxiv.org/abs/1602.05629) algorithm, achieving convergence in a system with randomly sampled
subsets of clients in each round can take a while, and it would be impractical
to have to run hundreds of rounds in this interactive tutorial.
What we'll do instead is sample the set of clients once, and
reuse the same set across rounds to speed up convergence (intentionally
over-fitting to these few user's data). We leave it as an exercise for the
reader to modify this tutorial to simulate random sampling - it is fairly easy to
do (once you do, keep in mind that getting the model to converge may take a
while).
```
sample_clients = emnist_train.client_ids[0:NUM_CLIENTS]
federated_train_data = make_federated_data(emnist_train, sample_clients)
print('Number of client datasets: {l}'.format(l=len(federated_train_data)))
print('First dataset: {d}'.format(d=federated_train_data[0]))
```
## Creating a model with Keras
If you are using Keras, you likely already have code that constructs a Keras
model. Here's an example of a simple model that will suffice for our needs.
```
def create_keras_model():
return tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(784,)),
tf.keras.layers.Dense(10, kernel_initializer='zeros'),
tf.keras.layers.Softmax(),
])
```
**Note:** we do not compile the model yet. The loss, metrics, and optimizers are introduced later.
In order to use any model with TFF, it needs to be wrapped in an instance of the
`tff.learning.Model` interface, which exposes methods to stamp the model's
forward pass, metadata properties, etc., similarly to Keras, but also introduces
additional elements, such as ways to control the process of computing federated
metrics. Let's not worry about this for now; if you have a Keras model like the
one we've just defined above, you can have TFF wrap it for you by invoking
`tff.learning.from_keras_model`, passing the model and a sample data batch as
arguments, as shown below.
```
def model_fn():
# We _must_ create a new model here, and _not_ capture it from an external
# scope. TFF will call this within different graph contexts.
keras_model = create_keras_model()
return tff.learning.from_keras_model(
keras_model,
input_spec=preprocessed_example_dataset.element_spec,
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=[tf.keras.metrics.SparseCategoricalAccuracy()])
```
## Training the model on federated data
Now that we have a model wrapped as `tff.learning.Model` for use with TFF, we
can let TFF construct a Federated Averaging algorithm by invoking the helper
function `tff.learning.build_federated_averaging_process`, as follows.
Keep in mind that the argument needs to be a constructor (such as `model_fn`
above), not an already-constructed instance, so that the construction of your
model can happen in a context controlled by TFF (if you're curious about the
reasons for this, we encourage you to read the follow-up tutorial on
[custom algorithms](custom_federated_algorithms_1.ipynb)).
One critical note on the Federated Averaging algorithm below, there are **2**
optimizers: a _client_optimizer_ and a _server_optimizer_. The
_client_optimizer_ is only used to compute local model updates on each client.
The _server_optimizer_ applies the averaged update to the global model at the
server. In particular, this means that the choice of optimizer and learning rate
used may need to be different than the ones you have used to train the model on
a standard i.i.d. dataset. We recommend starting with regular SGD, possibly with
a smaller learning rate than usual. The learning rate we use has not been
carefully tuned, feel free to experiment.
```
iterative_process = tff.learning.build_federated_averaging_process(
model_fn,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=0.02),
server_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=1.0))
```
What just happened? TFF has constructed a pair of *federated computations* and
packaged them into a `tff.templates.IterativeProcess` in which these computations
are available as a pair of properties `initialize` and `next`.
In a nutshell, *federated computations* are programs in TFF's internal language
that can express various federated algorithms (you can find more about this in
the [custom algorithms](custom_federated_algorithms_1.ipynb) tutorial). In this
case, the two computations generated and packed into `iterative_process`
implement [Federated Averaging](https://arxiv.org/abs/1602.05629).
It is a goal of TFF to define computations in a way that they could be executed
in real federated learning settings, but currently only local execution
simulation runtime is implemented. To execute a computation in a simulator, you
simply invoke it like a Python function. This default interpreted environment is
not designed for high performance, but it will suffice for this tutorial; we
expect to provide higher-performance simulation runtimes to facilitate
larger-scale research in future releases.
Let's start with the `initialize` computation. As is the case for all federated
computations, you can think of it as a function. The computation takes no
arguments, and returns one result - the representation of the state of the
Federated Averaging process on the server. While we don't want to dive into the
details of TFF, it may be instructive to see what this state looks like. You can
visualize it as follows.
```
str(iterative_process.initialize.type_signature)
```
While the above type signature may at first seem a bit cryptic, you can
recognize that the server state consists of a `model` (the initial model
parameters for MNIST that will be distributed to all devices), and
`optimizer_state` (additional information maintained by the server, such as the
number of rounds to use for hyperparameter schedules, etc.).
Let's invoke the `initialize` computation to construct the server state.
```
state = iterative_process.initialize()
```
The second of the pair of federated computations, `next`, represents a single
round of Federated Averaging, which consists of pushing the server state
(including the model parameters) to the clients, on-device training on their
local data, collecting and averaging model updates, and producing a new updated
model at the server.
Conceptually, you can think of `next` as having a functional type signature that
looks as follows.
```
SERVER_STATE, FEDERATED_DATA -> SERVER_STATE, TRAINING_METRICS
```
In particular, one should think about `next()` not as being a function that runs on a server, but rather being a declarative functional representation of the entire decentralized computation - some of the inputs are provided by the server (`SERVER_STATE`), but each participating device contributes its own local dataset.
Let's run a single round of training and visualize the results. We can use the
federated data we've already generated above for a sample of users.
```
state, metrics = iterative_process.next(state, federated_train_data)
print('round 1, metrics={}'.format(metrics))
```
Let's run a few more rounds. As noted earlier, typically at this point you would
pick a subset of your simulation data from a new randomly selected sample of
users for each round in order to simulate a realistic deployment in which users
continuously come and go, but in this interactive notebook, for the sake of
demonstration we'll just reuse the same users, so that the system converges
quickly.
```
NUM_ROUNDS = 11
for round_num in range(2, NUM_ROUNDS):
state, metrics = iterative_process.next(state, federated_train_data)
print('round {:2d}, metrics={}'.format(round_num, metrics))
```
Training loss is decreasing after each round of federated training, indicating
the model is converging. There are some important caveats with these training
metrics, however, see the section on *Evaluation* later in this tutorial.
##Displaying model metrics in TensorBoard
Next, let's visualize the metrics from these federated computations using Tensorboard.
Let's start by creating the directory and the corresponding summary writer to write the metrics to.
```
#@test {"skip": true}
logdir = "/tmp/logs/scalars/training/"
summary_writer = tf.summary.create_file_writer(logdir)
state = iterative_process.initialize()
```
Plot the relevant scalar metrics with the same summary writer.
```
#@test {"skip": true}
with summary_writer.as_default():
for round_num in range(1, NUM_ROUNDS):
state, metrics = iterative_process.next(state, federated_train_data)
for name, value in metrics.train._asdict().items():
tf.summary.scalar(name, value, step=round_num)
```
Start TensorBoard with the root log directory specified above. It can take a few seconds for the data to load.
```
#@test {"skip": true}
%tensorboard --logdir /tmp/logs/scalars/ --port=0
#@test {"skip": true}
# Run this this cell to clean your directory of old output for future graphs from this directory.
!rm -R /tmp/logs/scalars/*
```
In order to view evaluation metrics the same way, you can create a separate eval folder, like "logs/scalars/eval", to write to TensorBoard.
## Customizing the model implementation
Keras is the [recommended high-level model API for TensorFlow](https://medium.com/tensorflow/standardizing-on-keras-guidance-on-high-level-apis-in-tensorflow-2-0-bad2b04c819a), and we encourage using Keras models (via
`tff.learning.from_keras_model`) in TFF whenever possible.
However, `tff.learning` provides a lower-level model interface, `tff.learning.Model`, that exposes the minimal functionality necessary for using a model for federated learning. Directly implementing this interface (possibly still using building blocks like `tf.keras.layers`) allows for maximum customization without modifying the internals of the federated learning algorithms.
So let's do it all over again from scratch.
### Defining model variables, forward pass, and metrics
The first step is to identify the TensorFlow variables we're going to work with.
In order to make the following code more legible, let's define a data structure
to represent the entire set. This will include variables such as `weights` and
`bias` that we will train, as well as variables that will hold various
cumulative statistics and counters we will update during training, such as
`loss_sum`, `accuracy_sum`, and `num_examples`.
```
MnistVariables = collections.namedtuple(
'MnistVariables', 'weights bias num_examples loss_sum accuracy_sum')
```
Here's a method that creates the variables. For the sake of simplicity, we
represent all statistics as `tf.float32`, as that will eliminate the need for
type conversions at a later stage. Wrapping variable initializers as lambdas is
a requirement imposed by
[resource variables](https://www.tensorflow.org/api_docs/python/tf/enable_resource_variables).
```
def create_mnist_variables():
return MnistVariables(
weights=tf.Variable(
lambda: tf.zeros(dtype=tf.float32, shape=(784, 10)),
name='weights',
trainable=True),
bias=tf.Variable(
lambda: tf.zeros(dtype=tf.float32, shape=(10)),
name='bias',
trainable=True),
num_examples=tf.Variable(0.0, name='num_examples', trainable=False),
loss_sum=tf.Variable(0.0, name='loss_sum', trainable=False),
accuracy_sum=tf.Variable(0.0, name='accuracy_sum', trainable=False))
```
With the variables for model parameters and cumulative statistics in place, we
can now define the forward pass method that computes loss, emits predictions,
and updates the cumulative statistics for a single batch of input data, as
follows.
```
def mnist_forward_pass(variables, batch):
y = tf.nn.softmax(tf.matmul(batch['x'], variables.weights) + variables.bias)
predictions = tf.cast(tf.argmax(y, 1), tf.int32)
flat_labels = tf.reshape(batch['y'], [-1])
loss = -tf.reduce_mean(
tf.reduce_sum(tf.one_hot(flat_labels, 10) * tf.math.log(y), axis=[1]))
accuracy = tf.reduce_mean(
tf.cast(tf.equal(predictions, flat_labels), tf.float32))
num_examples = tf.cast(tf.size(batch['y']), tf.float32)
variables.num_examples.assign_add(num_examples)
variables.loss_sum.assign_add(loss * num_examples)
variables.accuracy_sum.assign_add(accuracy * num_examples)
return loss, predictions
```
Next, we define a function that returns a set of local metrics, again using TensorFlow. These are the values (in addition to model updates, which are handled automatically) that are eligible to be aggregated to the server in a federated learning or evaluation process.
Here, we simply return the average `loss` and `accuracy`, as well as the
`num_examples`, which we'll need to correctly weight the contributions from
different users when computing federated aggregates.
```
def get_local_mnist_metrics(variables):
return collections.OrderedDict(
num_examples=variables.num_examples,
loss=variables.loss_sum / variables.num_examples,
accuracy=variables.accuracy_sum / variables.num_examples)
```
Finally, we need to determine how to aggregate the local metrics emitted by each
device via `get_local_mnist_metrics`. This is the only part of the code that isn't written in TensorFlow - it's a *federated computation* expressed in TFF. If you'd like to
dig deeper, skim over the [custom algorithms](custom_federated_algorithms_1.ipynb)
tutorial, but in most applications, you won't really need to; variants of the
pattern shown below should suffice. Here's what it looks like:
```
@tff.federated_computation
def aggregate_mnist_metrics_across_clients(metrics):
return collections.OrderedDict(
num_examples=tff.federated_sum(metrics.num_examples),
loss=tff.federated_mean(metrics.loss, metrics.num_examples),
accuracy=tff.federated_mean(metrics.accuracy, metrics.num_examples))
```
The input `metrics` argument corresponds to the `OrderedDict` returned by `get_local_mnist_metrics` above, but critically the values are no longer `tf.Tensors` - they are "boxed" as `tff.Value`s, to make it clear you can no longer manipulate them using TensorFlow, but only using TFF's federated operators like `tff.federated_mean` and `tff.federated_sum`. The returned
dictionary of global aggregates defines the set of metrics which will be available on the server.
### Constructing an instance of `tff.learning.Model`
With all of the above in place, we are ready to construct a model representation
for use with TFF similar to one that's generated for you when you let TFF ingest
a Keras model.
```
class MnistModel(tff.learning.Model):
def __init__(self):
self._variables = create_mnist_variables()
@property
def trainable_variables(self):
return [self._variables.weights, self._variables.bias]
@property
def non_trainable_variables(self):
return []
@property
def local_variables(self):
return [
self._variables.num_examples, self._variables.loss_sum,
self._variables.accuracy_sum
]
@property
def input_spec(self):
return collections.OrderedDict(
x=tf.TensorSpec([None, 784], tf.float32),
y=tf.TensorSpec([None, 1], tf.int32))
@tf.function
def forward_pass(self, batch, training=True):
del training
loss, predictions = mnist_forward_pass(self._variables, batch)
num_exmaples = tf.shape(batch['x'])[0]
return tff.learning.BatchOutput(
loss=loss, predictions=predictions, num_examples=num_exmaples)
@tf.function
def report_local_outputs(self):
return get_local_mnist_metrics(self._variables)
@property
def federated_output_computation(self):
return aggregate_mnist_metrics_across_clients
```
As you can see, the abstract methods and properties defined by
`tff.learning.Model` corresponds to the code snippets in the preceding section
that introduced the variables and defined the loss and statistics.
Here are a few points worth highlighting:
* All state that your model will use must be captured as TensorFlow variables,
as TFF does not use Python at runtime (remember your code should be written
such that it can be deployed to mobile devices; see the
[custom algorithms](custom_federated_algorithms_1.ipynb) tutorial for a more
in-depth commentary on the reasons).
* Your model should describe what form of data it accepts (`input_spec`), as
in general, TFF is a strongly-typed environment and wants to determine type
signatures for all components. Declaring the format of your model's input is
an essential part of it.
* Although technically not required, we recommend wrapping all TensorFlow
logic (forward pass, metric calculations, etc.) as `tf.function`s,
as this helps ensure the TensorFlow can be serialized, and removes the need
for explicit control dependencies.
The above is sufficient for evaluation and algorithms like Federated SGD.
However, for Federated Averaging, we need to specify how the model should train
locally on each batch. We will specify a local optimizer when building the Federated Averaging algorithm.
### Simulating federated training with the new model
With all the above in place, the remainder of the process looks like what we've
seen already - just replace the model constructor with the constructor of our
new model class, and use the two federated computations in the iterative process
you created to cycle through training rounds.
```
iterative_process = tff.learning.build_federated_averaging_process(
MnistModel,
client_optimizer_fn=lambda: tf.keras.optimizers.SGD(learning_rate=0.02))
state = iterative_process.initialize()
state, metrics = iterative_process.next(state, federated_train_data)
print('round 1, metrics={}'.format(metrics))
for round_num in range(2, 11):
state, metrics = iterative_process.next(state, federated_train_data)
print('round {:2d}, metrics={}'.format(round_num, metrics))
```
To see these metrics within TensorBoard, refer to the steps listed above in "Displaying model metrics in TensorBoard".
## Evaluation
All of our experiments so far presented only federated training metrics - the
average metrics over all batches of data trained across all clients in the
round. This introduces the normal concerns about overfitting, especially since
we used the same set of clients on each round for simplicity, but there is an
additional notion of overfitting in training metrics specific to the Federated
Averaging algorithm. This is easiest to see if we imagine each client had a
single batch of data, and we train on that batch for many iterations (epochs).
In this case, the local model will quickly exactly fit to that one batch, and so
the local accuracy metric we average will approach 1.0. Thus, these training
metrics can be taken as a sign that training is progressing, but not much more.
To perform evaluation on federated data, you can construct another *federated
computation* designed for just this purpose, using the
`tff.learning.build_federated_evaluation` function, and passing in your model
constructor as an argument. Note that unlike with Federated Averaging, where
we've used `MnistTrainableModel`, it suffices to pass the `MnistModel`.
Evaluation doesn't perform gradient descent, and there's no need to construct
optimizers.
For experimentation and research, when a centralized test dataset is available,
[Federated Learning for Text Generation](federated_learning_for_text_generation.ipynb)
demonstrates another evaluation option: taking the trained weights from
federated learning, applying them to a standard Keras model, and then simply
calling `tf.keras.models.Model.evaluate()` on a centralized dataset.
```
evaluation = tff.learning.build_federated_evaluation(MnistModel)
```
You can inspect the abstract type signature of the evaluation function as follows.
```
str(evaluation.type_signature)
```
No need to be concerned about the details at this point, just be aware that it
takes the following general form, similar to `tff.templates.IterativeProcess.next`
but with two important differences. First, we are not returning server state,
since evaluation doesn't modify the model or any other aspect of state - you can
think of it as stateless. Second, evaluation only needs the model, and doesn't
require any other part of server state that might be associated with training,
such as optimizer variables.
```
SERVER_MODEL, FEDERATED_DATA -> TRAINING_METRICS
```
Let's invoke evaluation on the latest state we arrived at during training. In
order to extract the latest trained model from the server state, you simply
access the `.model` member, as follows.
```
train_metrics = evaluation(state.model, federated_train_data)
```
Here's what we get. Note the numbers look marginally better than what was
reported by the last round of training above. By convention, the training
metrics reported by the iterative training process generally reflect the
performance of the model at the beginning of the training round, so the
evaluation metrics will always be one step ahead.
```
str(train_metrics)
```
Now, let's compile a test sample of federated data and rerun evaluation on the
test data. The data will come from the same sample of real users, but from a
distinct held-out data set.
```
federated_test_data = make_federated_data(emnist_test, sample_clients)
len(federated_test_data), federated_test_data[0]
test_metrics = evaluation(state.model, federated_test_data)
str(test_metrics)
```
This concludes the tutorial. We encourage you to play with the
parameters (e.g., batch sizes, number of users, epochs, learning rates, etc.), to modify the code above to simulate training on random samples of users in
each round, and to explore the other tutorials we've developed.
| github_jupyter |
## Configurations for Colab
```
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!apt-get install -y xvfb python-opengl > /dev/null 2>&1
!pip install gym pyvirtualdisplay > /dev/null 2>&1
!pip install JSAnimation==0.1
from pyvirtualdisplay import Display
# Start virtual display
dis = Display(visible=0, size=(400, 400))
dis.start()
```
# 07. N-Step Learning
[R. S. Sutton, "Learning to predict by the methods of temporal differences." Machine learning, 3(1):9–44, 1988.](http://incompleteideas.net/papers/sutton-88-with-erratum.pdf)
Q-learning accumulates a single reward and then uses the greedy action at the next step to bootstrap. Alternatively, forward-view multi-step targets can be used (Sutton 1988). We call it Truncated N-Step Return
from a given state $S_t$. It is defined as,
$$
R^{(n)}_t = \sum_{k=0}^{n-1} \gamma_t^{(k)} R_{t+k+1}.
$$
A multi-step variant of DQN is then defined by minimizing the alternative loss,
$$
(R^{(n)}_t + \gamma^{(n)}_t \max_{a'} q_{\theta}^{-}
(S_{t+n}, a')
− q_{\theta}(S_t, A_t))^2.
$$
Multi-step targets with suitably tuned $n$ often lead to faster learning (Sutton and Barto 1998).
```
import os
from collections import deque
from typing import Deque, Dict, List, Tuple
import gym
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
from IPython.display import clear_output
from torch.nn.utils import clip_grad_norm_
```
## Replay buffer for N-step learning
There are a little bit changes in Replay buffer for N-step learning. First, we use `deque` to store the most recent n-step transitions.
```python
self.n_step_buffer = deque(maxlen=n_step)
```
You can see it doesn't actually store a transition in the buffer, unless `n_step_buffer` is full.
```
# in store method
if len(self.n_step_buffer) < self.n_step:
return False
```
When the length of `n_step_buffer` becomes equal to N, it eventually stores the N-step transition, which is calculated by `_get_n_step_info` method.
(Please see *01.dqn.ipynb* for detailed description of the basic replay buffer.)
```
class ReplayBuffer:
"""A simple numpy replay buffer."""
def __init__(
self,
obs_dim: int,
size: int,
batch_size: int = 32,
n_step: int = 3,
gamma: float = 0.99,
):
self.obs_buf = np.zeros([size, obs_dim], dtype=np.float32)
self.next_obs_buf = np.zeros([size, obs_dim], dtype=np.float32)
self.acts_buf = np.zeros([size], dtype=np.float32)
self.rews_buf = np.zeros([size], dtype=np.float32)
self.done_buf = np.zeros(size, dtype=np.float32)
self.max_size, self.batch_size = size, batch_size
self.ptr, self.size, = 0, 0
# for N-step Learning
self.n_step_buffer = deque(maxlen=n_step)
self.n_step = n_step
self.gamma = gamma
def store(
self,
obs: np.ndarray,
act: np.ndarray,
rew: float,
next_obs: np.ndarray,
done: bool
) -> bool:
transition = (obs, act, rew, next_obs, done)
self.n_step_buffer.append(transition)
# single step transition is not ready
if len(self.n_step_buffer) < self.n_step:
return False
# make a n-step transition
rew, next_obs, done = self._get_n_step_info(
self.n_step_buffer, self.gamma
)
obs, action = self.n_step_buffer[0][:2]
self.obs_buf[self.ptr] = obs
self.next_obs_buf[self.ptr] = next_obs
self.acts_buf[self.ptr] = act
self.rews_buf[self.ptr] = rew
self.done_buf[self.ptr] = done
self.ptr = (self.ptr + 1) % self.max_size
self.size = min(self.size + 1, self.max_size)
return True
def sample_batch(self) -> Dict[str, np.ndarray]:
indices = np.random.choice(
self.size, size=self.batch_size, replace=False
)
return dict(
obs=self.obs_buf[indices],
next_obs=self.next_obs_buf[indices],
acts=self.acts_buf[indices],
rews=self.rews_buf[indices],
done=self.done_buf[indices],
# for N-step Learning
indices=indices,
)
def sample_batch_from_idxs(
self, indices: np.ndarray
) -> Dict[str, np.ndarray]:
# for N-step Learning
return dict(
obs=self.obs_buf[indices],
next_obs=self.next_obs_buf[indices],
acts=self.acts_buf[indices],
rews=self.rews_buf[indices],
done=self.done_buf[indices],
)
def _get_n_step_info(
self, n_step_buffer: Deque, gamma: float
) -> Tuple[np.int64, np.ndarray, bool]:
"""Return n step rew, next_obs, and done."""
# info of the last transition
rew, next_obs, done = n_step_buffer[-1][-3:]
for transition in reversed(list(n_step_buffer)[:-1]):
r, n_o, d = transition[-3:]
rew = r + gamma * rew * (1 - d)
next_obs, done = (n_o, d) if d else (next_obs, done)
return rew, next_obs, done
def __len__(self) -> int:
return self.size
```
## Network
We are going to use a simple network architecture with three fully connected layers and two non-linearity functions (ReLU).
```
class Network(nn.Module):
def __init__(self, in_dim: int, out_dim: int):
"""Initialization."""
super(Network, self).__init__()
self.layers = nn.Sequential(
nn.Linear(in_dim, 128),
nn.ReLU(),
nn.Linear(128, 128),
nn.ReLU(),
nn.Linear(128, out_dim)
)
def forward(self, x: torch.Tensor) -> torch.Tensor:
"""Forward method implementation."""
return self.layers(x)
```
## DQN Agent + N-step learning Agent
Here is a summary of DQNAgent class.
| Method | Note |
| --- | --- |
|select_action | select an action from the input state. |
|step | take an action and return the response of the env. |
|compute_dqn_loss | return dqn loss. |
|update_model | update the model by gradient descent. |
|target_hard_update| hard update from the local model to the target model.|
|train | train the agent during num_frames. |
|test | test the agent (1 episode). |
|plot | plot the training progresses. |
We use two buffers: `memory` and `memory_n` for 1-step transitions and n-step transitions respectively. It guarantees that any paired 1-step and n-step transitions have the same indices (See `step` method for more details). Due to the reason, we can sample pairs of transitions from the two buffers once we have indices for samples.
```python
def update_model(self) -> torch.Tensor:
...
samples = self.memory.sample_batch()
indices = samples["indices"]
...
# N-step Learning loss
if self.use_n_step:
samples = self.memory_n.sample_batch_from_idxs(indices)
...
```
One thing to note that we are gonna combine 1-step loss and n-step loss so as to control high-variance / high-bias trade-off.
(Search the comments with *N-step Leaning* to see any difference from DQN.)
```
class DQNAgent:
"""DQN Agent interacting with environment.
Attribute:
env (gym.Env): openAI Gym environment
memory (ReplayBuffer): replay memory to store transitions
batch_size (int): batch size for sampling
epsilon (float): parameter for epsilon greedy policy
epsilon_decay (float): step size to decrease epsilon
max_epsilon (float): max value of epsilon
min_epsilon (float): min value of epsilon
target_update (int): period for target model's hard update
gamma (float): discount factor
dqn (Network): model to train and select actions
dqn_target (Network): target model to update
optimizer (torch.optim): optimizer for training dqn
transition (list): transition information including
state, action, reward, next_state, done
use_n_step (bool): whether to use n_step memory
n_step (int): step number to calculate n-step td error
memory_n (ReplayBuffer): n-step replay buffer
"""
def __init__(
self,
env: gym.Env,
memory_size: int,
batch_size: int,
target_update: int,
epsilon_decay: float,
max_epsilon: float = 1.0,
min_epsilon: float = 0.1,
gamma: float = 0.99,
# N-step Learning
n_step: int = 3,
):
"""Initialization.
Args:
env (gym.Env): openAI Gym environment
memory_size (int): length of memory
batch_size (int): batch size for sampling
target_update (int): period for target model's hard update
epsilon_decay (float): step size to decrease epsilon
lr (float): learning rate
max_epsilon (float): max value of epsilon
min_epsilon (float): min value of epsilon
gamma (float): discount factor
n_step (int): step number to calculate n-step td error
"""
obs_dim = env.observation_space.shape[0]
action_dim = env.action_space.n
self.env = env
self.batch_size = batch_size
self.epsilon = max_epsilon
self.epsilon_decay = epsilon_decay
self.max_epsilon = max_epsilon
self.min_epsilon = min_epsilon
self.target_update = target_update
self.gamma = gamma
# memory for 1-step Learning
self.memory = ReplayBuffer(
obs_dim, memory_size, batch_size, n_step=1
)
# memory for N-step Learning
self.use_n_step = True if n_step > 1 else False
if self.use_n_step:
self.n_step = n_step
self.memory_n = ReplayBuffer(
obs_dim, memory_size, batch_size, n_step=n_step, gamma=gamma
)
# device: cpu / gpu
self.device = torch.device(
"cuda" if torch.cuda.is_available() else "cpu"
)
print(self.device)
# networks: dqn, dqn_target
self.dqn = Network(obs_dim, action_dim).to(self.device)
self.dqn_target = Network(obs_dim, action_dim).to(self.device)
self.dqn_target.load_state_dict(self.dqn.state_dict())
self.dqn_target.eval()
# optimizer
self.optimizer = optim.Adam(self.dqn.parameters())
# transition to store in memory
self.transition = list()
# mode: train / test
self.is_test = False
def select_action(self, state: np.ndarray) -> np.ndarray:
"""Select an action from the input state."""
# epsilon greedy policy
if self.epsilon > np.random.random():
selected_action = self.env.action_space.sample()
else:
selected_action = self.dqn(
torch.FloatTensor(state).to(self.device)
).argmax()
selected_action = selected_action.detach().cpu().numpy()
if not self.is_test:
self.transition = [state, selected_action]
return selected_action
def step(self, action: np.ndarray) -> Tuple[np.ndarray, np.float64, bool]:
"""Take an action and return the response of the env."""
next_state, reward, done, _ = self.env.step(action)
if not self.is_test:
self.transition += [reward, next_state, done]
# add N-step transition
if self.use_n_step:
is_n_step_stored = self.memory_n.store(*self.transition)
# add a single step transition
if not self.use_n_step or is_n_step_stored:
self.memory.store(*self.transition)
return next_state, reward, done
def update_model(self) -> torch.Tensor:
"""Update the model by gradient descent."""
samples = self.memory.sample_batch()
indices = samples["indices"]
loss = self._compute_dqn_loss(samples, self.gamma)
# N-step Learning loss
# we are gonna combine 1-step loss and n-step loss so as to
# prevent high-variance.
if self.use_n_step:
samples = self.memory_n.sample_batch_from_idxs(indices)
gamma = self.gamma ** self.n_step
n_loss = self._compute_dqn_loss(samples, gamma)
loss += n_loss
self.optimizer.zero_grad()
loss.backward()
# gradient clipping
# https://pytorch.org/docs/stable/nn.html#torch.nn.utils.clip_grad_norm_
clip_grad_norm_(self.dqn.parameters(), 1.0, norm_type=1)
self.optimizer.step()
return loss.item()
def train(self, num_frames: int, plotting_interval: int = 200):
"""Train the agent."""
self.is_test = False
state = self.env.reset()
update_cnt = 0
epsilons = []
losses = []
scores = []
score = 0
for frame_idx in range(1, num_frames + 1):
action = self.select_action(state)
next_state, reward, done = self.step(action)
state = next_state
score += reward
# if episode ends
if done:
state = env.reset()
scores.append(score)
score = 0
# if training is ready
if len(self.memory) >= self.batch_size:
loss = self.update_model()
losses.append(loss)
update_cnt += 1
# linearly decrease epsilon
self.epsilon = max(
self.min_epsilon, self.epsilon - (
self.max_epsilon - self.min_epsilon
) * self.epsilon_decay
)
epsilons.append(self.epsilon)
# if hard update is needed
if update_cnt % self.target_update == 0:
self._target_hard_update()
# plotting
if frame_idx % plotting_interval == 0:
self._plot(frame_idx, scores, losses, epsilons)
self.env.close()
def test(self) -> List[np.ndarray]:
"""Test the agent."""
self.is_test = True
state = self.env.reset()
done = False
score = 0
frames = []
while not done:
frames.append(self.env.render(mode="rgb_array"))
action = self.select_action(state)
next_state, reward, done = self.step(action)
state = next_state
score += reward
print("score: ", score)
self.env.close()
return frames
def _compute_dqn_loss(
self,
samples: Dict[str, np.ndarray],
gamma: float
) -> torch.Tensor:
"""Return dqn loss."""
device = self.device # for shortening the following lines
state = torch.FloatTensor(samples["obs"]).to(device)
next_state = torch.FloatTensor(samples["next_obs"]).to(device)
action = torch.LongTensor(samples["acts"].reshape(-1, 1)).to(device)
reward = torch.FloatTensor(samples["rews"].reshape(-1, 1)).to(device)
done = torch.FloatTensor(samples["done"].reshape(-1, 1)).to(device)
# G_t = r + gamma * v(s_{t+1}) if state != Terminal
# = r otherwise
curr_q_value = self.dqn(state).gather(1, action)
next_q_value = self.dqn_target(next_state).max(
dim=1, keepdim=True
)[0].detach()
mask = 1 - done
target = (reward + gamma * next_q_value * mask).to(self.device)
# calculate dqn loss
loss = ((target - curr_q_value).pow(2)).mean()
return loss
def _target_hard_update(self):
"""Hard update: target <- local."""
self.dqn_target.load_state_dict(self.dqn.state_dict())
def _plot(
self,
frame_idx: int,
scores: List[float],
losses: List[float],
epsilons: List[float],
):
"""Plot the training progresses."""
clear_output(True)
plt.figure(figsize=(20, 5))
plt.subplot(131)
plt.title('frame %s. score: %s' % (frame_idx, np.mean(scores[-10:])))
plt.plot(scores)
plt.subplot(132)
plt.title('loss')
plt.plot(losses)
plt.subplot(133)
plt.title('epsilons')
plt.plot(epsilons)
plt.show()
```
## Environment
You can see the [code](https://github.com/openai/gym/blob/master/gym/envs/classic_control/cartpole.py) and [configurations](https://github.com/openai/gym/blob/master/gym/envs/__init__.py#L53) of CartPole-v0 from OpenAI's repository.
```
# environment
env_id = "CartPole-v0"
env = gym.make(env_id)
```
## Set random seed
```
seed = 777
def seed_torch(seed):
torch.manual_seed(seed)
if torch.backends.cudnn.enabled:
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
np.random.seed(seed)
seed_torch(seed)
env.seed(seed)
```
## Initialize
```
# parameters
num_frames = 10000
memory_size = 1000
batch_size = 32
target_update = 200
epsilon_decay = 1 / 2000
# train
agent = DQNAgent(env, memory_size, batch_size, target_update, epsilon_decay)
```
## Train
```
agent.train(num_frames)
```
## Test
Run the trained agent (1 episode).
```
frames = agent.test()
```
## Render
```
# Imports specifically so we can render outputs in Colab.
from matplotlib import animation
from JSAnimation.IPython_display import display_animation
from IPython.display import display
def display_frames_as_gif(frames):
"""Displays a list of frames as a gif, with controls."""
patch = plt.imshow(frames[0])
plt.axis('off')
def animate(i):
patch.set_data(frames[i])
anim = animation.FuncAnimation(
plt.gcf(), animate, frames = len(frames), interval=50
)
display(display_animation(anim, default_mode='loop'))
# display
display_frames_as_gif(frames)
```
| github_jupyter |
```
# Copyright (c) Facebook, Inc. and its affiliates. All rights reserved.
```
# Camera position optimization using differentiable rendering
In this tutorial we will learn the [x, y, z] position of a camera given a reference image using differentiable rendering.
We will first initialize a renderer with a starting position for the camera. We will then use this to generate an image, compute a loss with the reference image, and finally backpropagate through the entire pipeline to update the position of the camera.
This tutorial shows how to:
- load a mesh from an `.obj` file
- initialize a `Camera`, `Shader` and `Renderer`,
- render a mesh
- set up an optimization loop with a loss function and optimizer
## 0. Install and import modules
If `torch`, `torchvision` and `pytorch3d` are not installed, run the following cell:
```
!pip install torch torchvision
!pip install 'git+https://github.com/facebookresearch/pytorch3d.git@stable'
import os
import torch
import numpy as np
from tqdm.notebook import tqdm
import imageio
import torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
from skimage import img_as_ubyte
# io utils
from pytorch3d.io import load_obj
# datastructures
from pytorch3d.structures import Meshes, Textures
# 3D transformations functions
from pytorch3d.transforms import Rotate, Translate
# rendering components
from pytorch3d.renderer import (
OpenGLPerspectiveCameras, look_at_view_transform, look_at_rotation,
RasterizationSettings, MeshRenderer, MeshRasterizer, BlendParams,
SoftSilhouetteShader, HardPhongShader, PointLights
)
```
## 1. Load the Obj
We will load an obj file and create a **Meshes** object. **Meshes** is a unique datastructure provided in PyTorch3D for working with **batches of meshes of different sizes**. It has several useful class methods which are used in the rendering pipeline.
If you are running this notebook locally after cloning the PyTorch3D repository, the mesh will already be available. **If using Google Colab, fetch the mesh and save it at the path `data/`**:
```
!mkdir -p data
!wget -P data https://dl.fbaipublicfiles.com/pytorch3d/data/teapot/teapot.obj
# Set the cuda device
if torch.cuda.is_available():
device = torch.device("cuda:0")
torch.cuda.set_device(device)
else:
device = torch.device("cpu")
# Load the obj and ignore the textures and materials.
verts, faces_idx, _ = load_obj("./data/teapot.obj")
faces = faces_idx.verts_idx
# Initialize each vertex to be white in color.
verts_rgb = torch.ones_like(verts)[None] # (1, V, 3)
textures = Textures(verts_rgb=verts_rgb.to(device))
# Create a Meshes object for the teapot. Here we have only one mesh in the batch.
teapot_mesh = Meshes(
verts=[verts.to(device)],
faces=[faces.to(device)],
textures=textures
)
```
## 2. Optimization setup
### Create a renderer
A **renderer** in PyTorch3D is composed of a **rasterizer** and a **shader** which each have a number of subcomponents such as a **camera** (orthgraphic/perspective). Here we initialize some of these components and use default values for the rest.
For optimizing the camera position we will use a renderer which produces a **silhouette** of the object only and does not apply any **lighting** or **shading**. We will also initialize another renderer which applies full **phong shading** and use this for visualizing the outputs.
```
# Initialize an OpenGL perspective camera.
cameras = OpenGLPerspectiveCameras(device=device)
# To blend the 100 faces we set a few parameters which control the opacity and the sharpness of
# edges. Refer to blending.py for more details.
blend_params = BlendParams(sigma=1e-4, gamma=1e-4)
# Define the settings for rasterization and shading. Here we set the output image to be of size
# 256x256. To form the blended image we use 100 faces for each pixel. We also set bin_size and max_faces_per_bin to None which ensure that
# the faster coarse-to-fine rasterization method is used. Refer to rasterize_meshes.py for
# explanations of these parameters. Refer to docs/notes/renderer.md for an explanation of
# the difference between naive and coarse-to-fine rasterization.
raster_settings = RasterizationSettings(
image_size=256,
blur_radius=np.log(1. / 1e-4 - 1.) * blend_params.sigma,
faces_per_pixel=100,
)
# Create a silhouette mesh renderer by composing a rasterizer and a shader.
silhouette_renderer = MeshRenderer(
rasterizer=MeshRasterizer(
cameras=cameras,
raster_settings=raster_settings
),
shader=SoftSilhouetteShader(blend_params=blend_params)
)
# We will also create a phong renderer. This is simpler and only needs to render one face per pixel.
raster_settings = RasterizationSettings(
image_size=256,
blur_radius=0.0,
faces_per_pixel=1,
)
# We can add a point light in front of the object.
lights = PointLights(device=device, location=((2.0, 2.0, -2.0),))
phong_renderer = MeshRenderer(
rasterizer=MeshRasterizer(
cameras=cameras,
raster_settings=raster_settings
),
shader=HardPhongShader(device=device, cameras=cameras, lights=lights)
)
```
### Create a reference image
We will first position the teapot and generate an image. We use helper functions to rotate the teapot to a desired viewpoint. Then we can use the renderers to produce an image. Here we will use both renderers and visualize the silhouette and full shaded image.
The world coordinate system is defined as +Y up, +X left and +Z in. The teapot in world coordinates has the spout pointing to the left.
We defined a camera which is positioned on the positive z axis hence sees the spout to the right.
```
# Select the viewpoint using spherical angles
distance = 3 # distance from camera to the object
elevation = 50.0 # angle of elevation in degrees
azimuth = 0.0 # No rotation so the camera is positioned on the +Z axis.
# Get the position of the camera based on the spherical angles
R, T = look_at_view_transform(distance, elevation, azimuth, device=device)
# Render the teapot providing the values of R and T.
silhouete = silhouette_renderer(meshes_world=teapot_mesh, R=R, T=T)
image_ref = phong_renderer(meshes_world=teapot_mesh, R=R, T=T)
silhouete = silhouete.cpu().numpy()
image_ref = image_ref.cpu().numpy()
plt.figure(figsize=(10, 10))
plt.subplot(1, 2, 1)
plt.imshow(silhouete.squeeze()[..., 3]) # only plot the alpha channel of the RGBA image
plt.grid(False)
plt.subplot(1, 2, 2)
plt.imshow(image_ref.squeeze())
plt.grid(False)
```
### Set up a basic model
Here we create a simple model class and initialize a parameter for the camera position.
```
class Model(nn.Module):
def __init__(self, meshes, renderer, image_ref):
super().__init__()
self.meshes = meshes
self.device = meshes.device
self.renderer = renderer
# Get the silhouette of the reference RGB image by finding all the non zero values.
image_ref = torch.from_numpy((image_ref[..., :3].max(-1) != 0).astype(np.float32))
self.register_buffer('image_ref', image_ref)
# Create an optimizable parameter for the x, y, z position of the camera.
self.camera_position = nn.Parameter(
torch.from_numpy(np.array([3.0, 6.9, +2.5], dtype=np.float32)).to(meshes.device))
def forward(self):
# Render the image using the updated camera position. Based on the new position of the
# camer we calculate the rotation and translation matrices
R = look_at_rotation(self.camera_position[None, :], device=self.device) # (1, 3, 3)
T = -torch.bmm(R.transpose(1, 2), self.camera_position[None, :, None])[:, :, 0] # (1, 3)
image = self.renderer(meshes_world=self.meshes.clone(), R=R, T=T)
# Calculate the silhouette loss
loss = torch.sum((image[..., 3] - self.image_ref) ** 2)
return loss, image
```
## 3. Initialize the model and optimizer
Now we can create an instance of the **model** above and set up an **optimizer** for the camera position parameter.
```
# We will save images periodically and compose them into a GIF.
filename_output = "./teapot_optimization_demo.gif"
writer = imageio.get_writer(filename_output, mode='I', duration=0.3)
# Initialize a model using the renderer, mesh and reference image
model = Model(meshes=teapot_mesh, renderer=silhouette_renderer, image_ref=image_ref).to(device)
# Create an optimizer. Here we are using Adam and we pass in the parameters of the model
optimizer = torch.optim.Adam(model.parameters(), lr=0.05)
```
### Visualize the starting position and the reference position
```
plt.figure(figsize=(10, 10))
_, image_init = model()
plt.subplot(1, 2, 1)
plt.imshow(image_init.detach().squeeze().cpu().numpy()[..., 3])
plt.grid(False)
plt.title("Starting position")
plt.subplot(1, 2, 2)
plt.imshow(model.image_ref.cpu().numpy().squeeze())
plt.grid(False)
plt.title("Reference silhouette");
```
## 4. Run the optimization
We run several iterations of the forward and backward pass and save outputs every 10 iterations. When this has finished take a look at `./teapot_optimization_demo.gif` for a cool gif of the optimization process!
```
loop = tqdm(range(200))
for i in loop:
optimizer.zero_grad()
loss, _ = model()
loss.backward()
optimizer.step()
loop.set_description('Optimizing (loss %.4f)' % loss.data)
if loss.item() < 200:
break
# Save outputs to create a GIF.
if i % 10 == 0:
R = look_at_rotation(model.camera_position[None, :], device=model.device)
T = -torch.bmm(R.transpose(1, 2), model.camera_position[None, :, None])[:, :, 0] # (1, 3)
image = phong_renderer(meshes_world=model.meshes.clone(), R=R, T=T)
image = image[0, ..., :3].detach().squeeze().cpu().numpy()
image = img_as_ubyte(image)
writer.append_data(image)
plt.figure()
plt.imshow(image[..., :3])
plt.title("iter: %d, loss: %0.2f" % (i, loss.data))
plt.grid("off")
plt.axis("off")
writer.close()
```
## 5. Conclusion
In this tutorial we learnt how to **load** a mesh from an obj file, initialize a PyTorch3D datastructure called **Meshes**, set up an **Renderer** consisting of a **Rasterizer** and a **Shader**, set up an optimization loop including a **Model** and a **loss function**, and run the optimization.
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
%matplotlib inline
import seaborn as sns
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import gensim
from gensim.models.word2vec import Word2Vec
def tsnescatterplot(model, word, list_names,curr_pos):
""" Plot in seaborn the results from the t-SNE dimensionality reduction algorithm of the vectors of a query word,
its list of most similar words, and a list of words.
"""
arrays = np.empty((0, 300), dtype='f')
word_labels = [word]
color_list = ['red']
# adds the vector of the query word
arrays = np.append(arrays, model.wv.__getitem__([word]), axis=0)
# gets list of most similar words
close_words = model.wv.most_similar([word])
# adds the vector for each of the closest words to the array
for wrd_score in close_words:
wrd_vector = model.wv.__getitem__([wrd_score[0]])
word_labels.append(wrd_score[0].replace('T','U'))
color_list.append('blue')
arrays = np.append(arrays, wrd_vector, axis=0)
# adds the vector for each of the words from list_names to the array
for wrd in list_names:
wrd_vector = model.wv.__getitem__([wrd])
word_labels.append(wrd.replace('T','U'))
color_list.append('green')
arrays = np.append(arrays, wrd_vector, axis=0)
# Reduces the dimensionality from 300 to 50 dimensions with PCA
reduc = PCA(n_components=20).fit_transform(arrays)
# Finds t-SNE coordinates for 2 dimensions
np.set_printoptions(suppress=True)
Y = TSNE(n_components=2, random_state=0, perplexity=15).fit_transform(reduc)
# Sets everything up to plot
df = pd.DataFrame({'x': [x for x in Y[:, 0]],
'y': [y for y in Y[:, 1]],
'words': word_labels,
'color': color_list})
# fig, ax = plt.subplot()
# fig.set_size_inches(9, 9)
ax = plt.subplot(1,4,curr_pos)
# Basic plot
p1 = sns.regplot(data=df,
x="x",
y="y",
fit_reg=False,
marker="o",
scatter_kws={'s': 40,
'facecolors': df['color']
},
ax=ax)
# Adds annotations one by one with a loop
for line in range(0, df.shape[0]):
p1.text(df["x"][line],
df['y'][line],
' ' + df["words"][line],
horizontalalignment='left',
verticalalignment='bottom', size='medium',
color=df['color'][line],
weight='normal'
).set_size(12)
ax.set_xlim(Y[:, 0].min()-50, Y[:, 0].max()+50)
ax.set_ylim(Y[:, 1].min()-50, Y[:, 1].max()+50)
ax.set_title('t-SNE visualization for {}'.format(word))
#plt.savefig('./Figs/t-SNE_visualization_for_{}'.format(word))
return ax
model = Word2Vec.load('../Embeddings/Gen2vec')
fig, axes = plt.subplots(1,4)
fig.set_size_inches(24, 4)
axes[0] = tsnescatterplot(model, 'CCA', [i[0] for i in model.wv.most_similar(negative=["CCA"])],1)
axes[1] = tsnescatterplot(model, 'GGA', [i[0] for i in model.wv.most_similar(negative=["GGA"])],2)
axes[2] = tsnescatterplot(model, 'ACC', [i[0] for i in model.wv.most_similar(negative=["ACC"])],3)
axes[3] = tsnescatterplot(model, 'CAA', [i[0] for i in model.wv.most_similar(negative=["CAA"])],4)
```
| github_jupyter |
## Dependencies
```
import os
import sys
import cv2
import shutil
import random
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import multiprocessing as mp
import matplotlib.pyplot as plt
from tensorflow import set_random_seed
from sklearn.utils import class_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, cohen_kappa_score
from keras import backend as K
from keras.models import Model
from keras.utils import to_categorical
from keras import optimizers, applications
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Dense, Dropout, GlobalAveragePooling2D, Input
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, Callback, LearningRateScheduler, ModelCheckpoint
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(0)
seed = 0
seed_everything(seed)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
sys.path.append(os.path.abspath('../input/efficientnet/efficientnet-master/efficientnet-master/'))
from efficientnet import *
```
## Load data
```
fold_set = pd.read_csv('../input/aptos-split-oldnew-balanced/5-fold.csv')
X_train = fold_set[fold_set['fold_0'] == 'train']
X_val = fold_set[fold_set['fold_0'] == 'validation']
test = pd.read_csv('../input/aptos2019-blindness-detection/test.csv')
# Preprocecss data
test["id_code"] = test["id_code"].apply(lambda x: x + ".png")
print('Number of train samples: ', X_train.shape[0])
print('Number of validation samples: ', X_val.shape[0])
print('Number of test samples: ', test.shape[0])
display(X_train.head())
```
# Model parameters
```
# Model parameters
model_path = '../working/effNetB5_img224_noBen_fold1.h5'
FACTOR = 4
BATCH_SIZE = 8 * FACTOR
WARMUP_EPOCHS = 5
EPOCHS = 20
LEARNING_RATE = 1e-4 * FACTOR
WARMUP_LEARNING_RATE = 1e-3 * FACTOR
HEIGHT = 224
WIDTH = 224
CHANNELS = 3
TTA_STEPS = 5
ES_PATIENCE = 5
LR_WARMUP_EPOCHS = 3
STEP_SIZE = len(X_train) // BATCH_SIZE
TOTAL_STEPS = EPOCHS * STEP_SIZE
WARMUP_STEPS = LR_WARMUP_EPOCHS * STEP_SIZE
```
# Pre-procecess images
```
old_data_base_path = '../input/diabetic-retinopathy-resized/resized_train/resized_train/'
new_data_base_path = '../input/aptos2019-blindness-detection/train_images/'
test_base_path = '../input/aptos2019-blindness-detection/test_images/'
train_dest_path = 'base_dir/train_images/'
validation_dest_path = 'base_dir/validation_images/'
test_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
# Creating train, validation and test directories
os.makedirs(train_dest_path)
os.makedirs(validation_dest_path)
os.makedirs(test_dest_path)
def crop_image(img, tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
img = np.stack([img1,img2,img3],axis=-1)
return img
def circle_crop(img):
img = crop_image(img)
height, width, depth = img.shape
largest_side = np.max((height, width))
img = cv2.resize(img, (largest_side, largest_side))
height, width, depth = img.shape
x = width//2
y = height//2
r = np.amin((x, y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x, y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image(img)
return img
def preprocess_image(image_id, base_path, save_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = circle_crop(image)
image = cv2.resize(image, (HEIGHT, WIDTH))
# image = cv2.addWeighted(image, 4, cv2.GaussianBlur(image, (0,0), sigmaX), -4 , 128)
cv2.imwrite(save_path + image_id, image)
def preprocess_data(df, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
item = df.iloc[i]
image_id = item['id_code']
item_set = item['fold_0']
item_data = item['data']
if item_set == 'train':
if item_data == 'new':
preprocess_image(image_id, new_data_base_path, train_dest_path)
if item_data == 'old':
preprocess_image(image_id, old_data_base_path, train_dest_path)
if item_set == 'validation':
if item_data == 'new':
preprocess_image(image_id, new_data_base_path, validation_dest_path)
if item_data == 'old':
preprocess_image(image_id, old_data_base_path, validation_dest_path)
def preprocess_test(df, base_path=test_base_path, save_path=test_dest_path, HEIGHT=HEIGHT, WIDTH=WIDTH, sigmaX=10):
df = df.reset_index()
for i in range(df.shape[0]):
image_id = df.iloc[i]['id_code']
preprocess_image(image_id, base_path, save_path)
n_cpu = mp.cpu_count()
train_n_cnt = X_train.shape[0] // n_cpu
val_n_cnt = X_val.shape[0] // n_cpu
test_n_cnt = test.shape[0] // n_cpu
# Pre-procecss old data train set
pool = mp.Pool(n_cpu)
dfs = [X_train.iloc[train_n_cnt*i:train_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_train.iloc[train_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss validation set
pool = mp.Pool(n_cpu)
dfs = [X_val.iloc[val_n_cnt*i:val_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = X_val.iloc[val_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_data, [x_df for x_df in dfs])
pool.close()
# Pre-procecss test set
pool = mp.Pool(n_cpu)
dfs = [test.iloc[test_n_cnt*i:test_n_cnt*(i+1)] for i in range(n_cpu)]
dfs[-1] = test.iloc[test_n_cnt*(n_cpu-1):]
res = pool.map(preprocess_test, [x_df for x_df in dfs])
pool.close()
```
# Data generator
```
datagen=ImageDataGenerator(rescale=1./255,
rotation_range=360,
horizontal_flip=True,
vertical_flip=True)
train_generator=datagen.flow_from_dataframe(
dataframe=X_train,
directory=train_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
valid_generator=datagen.flow_from_dataframe(
dataframe=X_val,
directory=validation_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
test_generator=datagen.flow_from_dataframe(
dataframe=test,
directory=test_dest_path,
x_col="id_code",
batch_size=1,
class_mode=None,
shuffle=False,
target_size=(HEIGHT, WIDTH),
seed=seed)
def classify(x):
if x < 0.5:
return 0
elif x < 1.5:
return 1
elif x < 2.5:
return 2
elif x < 3.5:
return 3
return 4
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
def plot_confusion_matrix(train, validation, labels=labels):
train_labels, train_preds = train
validation_labels, validation_preds = validation
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 7))
train_cnf_matrix = confusion_matrix(train_labels, train_preds)
validation_cnf_matrix = confusion_matrix(validation_labels, validation_preds)
train_cnf_matrix_norm = train_cnf_matrix.astype('float') / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
validation_cnf_matrix_norm = validation_cnf_matrix.astype('float') / validation_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=labels, columns=labels)
validation_df_cm = pd.DataFrame(validation_cnf_matrix_norm, index=labels, columns=labels)
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues",ax=ax1).set_title('Train')
sns.heatmap(validation_df_cm, annot=True, fmt='.2f', cmap=sns.cubehelix_palette(8),ax=ax2).set_title('Validation')
plt.show()
def plot_metrics(history, figsize=(20, 14)):
fig, (ax1, ax2) = plt.subplots(2, 1, sharex='col', figsize=figsize)
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['acc'], label='Train accuracy')
ax2.plot(history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
def apply_tta(model, generator, steps=10):
step_size = generator.n//generator.batch_size
preds_tta = []
for i in range(steps):
generator.reset()
preds = model.predict_generator(generator, steps=step_size)
preds_tta.append(preds)
return np.mean(preds_tta, axis=0)
def evaluate_model(train, validation):
train_labels, train_preds = train
validation_labels, validation_preds = validation
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds, train_labels, weights='quadratic'))
print("Validation Cohen Kappa score: %.3f" % cohen_kappa_score(validation_preds, validation_labels, weights='quadratic'))
print("Complete set Cohen Kappa score: %.3f" % cohen_kappa_score(np.append(train_preds, validation_preds), np.append(train_labels, validation_labels), weights='quadratic'))
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueError('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
```
# Model
```
def create_model(input_shape):
input_tensor = Input(shape=input_shape)
base_model = EfficientNetB5(weights=None,
include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/efficientnet-keras-weights-b0b5/efficientnet-b5_imagenet_1000_notop.h5')
x = GlobalAveragePooling2D()(base_model.output)
final_output = Dense(1, activation='linear', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
```
# Train top layers
```
model = create_model(input_shape=(HEIGHT, WIDTH, CHANNELS))
for layer in model.layers:
layer.trainable = False
for i in range(-2, 0):
model.layers[i].trainable = True
metric_list = ["accuracy"]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history_warmup = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
verbose=2).history
```
# Fine-tune the model
```
for layer in model.layers:
layer.trainable = True
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True, save_weights_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
cosine_lr = WarmUpCosineDecayScheduler(learning_rate_base=LEARNING_RATE,
total_steps=TOTAL_STEPS,
warmup_learning_rate=0.0,
warmup_steps=WARMUP_STEPS,
hold_base_rate_steps=(2 * STEP_SIZE))
callback_list = [checkpoint, es, cosine_lr]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=EPOCHS,
callbacks=callback_list,
verbose=2).history
fig, ax = plt.subplots(1, 1, sharex='col', figsize=(20, 4))
ax.plot(cosine_lr.learning_rates)
ax.set_title('Fine-tune learning rates')
plt.xlabel('Steps')
plt.ylabel('Learning rate')
sns.despine()
plt.show()
```
# Model loss graph
```
plot_metrics(history)
# Create empty arays to keep the predictions and labels
df_preds = pd.DataFrame(columns=['label', 'pred', 'set'])
train_generator.reset()
valid_generator.reset()
# Add train predictions and labels
for i in range(STEP_SIZE_TRAIN + 1):
im, lbl = next(train_generator)
preds = model.predict(im, batch_size=train_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'train']
# Add validation predictions and labels
for i in range(STEP_SIZE_VALID + 1):
im, lbl = next(valid_generator)
preds = model.predict(im, batch_size=valid_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'validation']
df_preds['label'] = df_preds['label'].astype('int')
# Classify predictions
df_preds['predictions'] = df_preds['pred'].apply(lambda x: classify(x))
train_preds = df_preds[df_preds['set'] == 'train']
validation_preds = df_preds[df_preds['set'] == 'validation']
```
# Model Evaluation
## Confusion Matrix
### Original thresholds
```
plot_confusion_matrix((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Quadratic Weighted Kappa
```
evaluate_model((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
## Apply model to test set and output predictions
```
preds = apply_tta(model, test_generator, TTA_STEPS)
predictions = [classify(x) for x in preds]
results = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions})
results['id_code'] = results['id_code'].map(lambda x: str(x)[:-4])
# Cleaning created directories
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
```
# Predictions class distribution
```
fig = plt.subplots(sharex='col', figsize=(24, 8.7))
sns.countplot(x="diagnosis", data=results, palette="GnBu_d").set_title('Test')
sns.despine()
plt.show()
results.to_csv('submission.csv', index=False)
display(results.head())
```
| github_jupyter |
# Data Exploration
In this part of the homework, the data is going to be inspected in terms of the data samples present and the distributons of these classes in their respective sets (train or test). This notebook makes use of the Dataset class which is implemented on a seperate file named **dataset.py**, which includes the wrapper class for retrieving the data from the provided data files.
## Necessary Imports
This notebook makes use of the following packages in addition to the *dataset.py* file which is a personal effort:
- Pandas (to structure the imported data)
- Numpy (to perform mathematical operations on matrices)
- Matplotlib (to plot necessary figures)
```
# Necessary Imports
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from dataset import Dataset
```
### Location of data files
In the current configuraton, the files containing the data for the assignment are included in the **data/** directory. This folder is located at the home directory where this notebook is located.
```
# File Name Constants
TRAIN_DATA_FILE = "data/ann-train.data"
TEST_DATA_FILE = "data/ann-test.data"
FEATURE_COST_FILE = "data/ann-thyroid.cost"
```
### The Dataset Class
Dataset class is a personal implementation which helps managing the data files provided for the assignment. It reads the data line by line and returns two DataFrames that include the testing data and training data respectively. The dataframe that represents the testing data is named *test_data* and dataframe represents training data is named as *train_data*.
```
dataset = Dataset(TRAIN_DATA_FILE, TEST_DATA_FILE, FEATURE_COST_FILE)
```
#### Dataset details
In order to be able to inspect the details of the given dataset, the following dataframes are outputted. As it can be seen clearly, the $1^{st}, 17^{th}, 18^{th}, 19^{th}, 20^{th}, 21^{st}$ are continuous, whereas the other features are binary values.
```
dataset.train_data.head()
dataset.train_data.describe()
dataset.test_data.head()
dataset.test_data.describe()
```
### Cost values for different features
The cost values that are used in the third part of teh assignment are organized in a dictionary structure. This is done for easy access to feature names and cost values. The corresponding dictinary is given below.
```
dataset.cost_data
```
## Class Imbalance in Dataset
In order to observe the class imbalance problem in the dataset, the frequencies of different classes (3 classes to be specific) are attempted to be visualied, using a histogram. Here it is observed that class 3 is the majority class in the dataset where class 1 and class 2 are much less in terms of frequency. When class 1 and class 2 are compared, the class 2 is the majority class but this difference is not comparable with the difference with class 3.
```
def plot_dist(dataset):
class_instances = {
"train": np.unique(dataset.train_data["y"].to_numpy(), return_counts=True),
"test": np.unique(dataset.test_data["y"].to_numpy(), return_counts=True),
}
bar_width = 0.35
x = np.arange(class_instances["train"][0].size)
fig, ax = plt.subplots(figsize=(10,6))
ax.bar(x - bar_width/2, class_instances["train"][1], width=bar_width, label="Training Data", color="b")
ax.bar(x + bar_width/2, class_instances["test"][1], width=bar_width, label="Testing Data", color="g")
ax.set_ylabel("Class Frequencies")
ax.set_xlabel("Classes")
ax.set_title("Frequencies of data samples belonging to different classes")
ax.set_xticks(x)
ax.set_xticklabels(["Class 1", "Class 2", "Class 3"])
ax.legend()
fig.savefig("figures/class_balance.png")
plot_dist(dataset)
```
For a numeric comparison of distributions the class frequencies in terms of numeric values ar also identified below. These values just makes the statement clearer with the help of numeric percenatges.
```
def report_class_dist(data):
classes, counts = np.unique(data, return_counts=True)
prob_dist = counts / np.sum(counts)
for class_idx in range(len(classes)):
print("\t {}% of instances belong to class {}".format(round(prob_dist[class_idx] * 100, 2), int(classes[class_idx])))
print("Class distributions for training set and test set: ")
print("\tTraining Set:")
report_class_dist(dataset.train_data.to_numpy()[:, -1])
print("\tTest Set:")
report_class_dist(dataset.test_data.to_numpy()[:, -1])
```
To be able to observe the linear seperability of the dataset, for training data the distribution of the features is observed. After this observation, in some plots is is visible that for some thresholds they seperate the training instances. The scatter plots for the features are given below:
```
var_list = dataset.train_data.columns
wanted_cols = dataset.train_data.copy()
fig, axs = plt.subplots(3, 7, figsize=(40,15), constrained_layout=True)
for var_idx in range(len(var_list) - 1):
axs[var_idx % 3, var_idx % 7].scatter(wanted_cols["y"].to_numpy(dtype=int), wanted_cols[var_list[var_idx]].to_numpy())
axs[var_idx % 3, var_idx % 7].set_xlabel("y", fontsize=15)
axs[var_idx % 3, var_idx % 7].set_ylabel(var_list[var_idx], fontsize=15)
fig.suptitle("Varibale distributions for corresponding feature", fontsize=30)
fig.savefig("figures/scatter_dist.png")
```
| github_jupyter |
---
layout: page
title: Teorema Central do Limite
nav_order: 7
---
[<img src="./colab_favicon_small.png" style="float: right;">](https://colab.research.google.com/github/icd-ufmg/icd-ufmg.github.io/blob/master/_lessons/07-tcl.ipynb)
# Teorema Central do Limite
{: .no_toc .mb-2 }
O teorema base para os nossos testes de hipóteses
{: .fs-6 .fw-300 }
{: .no_toc .text-delta }
Resultados Esperados
1. Revisar conceitos de Probabilidade ligados a distribuição normal
1. Revisar o teorema central do limite
1. Entendimento do teorema central do limite
1. Simular médias de qualquer distribuição
1. Mostrar como a distribuição de média segue uma normal
---
**Sumário**
1. TOC
{:toc}
---
```
# -*- coding: utf8
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
plt.style.use('seaborn-colorblind')
plt.rcParams['figure.figsize'] = (16, 10)
plt.rcParams['axes.labelsize'] = 20
plt.rcParams['axes.titlesize'] = 20
plt.rcParams['legend.fontsize'] = 20
plt.rcParams['xtick.labelsize'] = 20
plt.rcParams['ytick.labelsize'] = 20
plt.rcParams['lines.linewidth'] = 4
plt.ion()
def despine(ax=None):
if ax is None:
ax = plt.gca()
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
```
## Introdução
Uma razão pela qual a distribuição normal é tão útil é o teorema central do limite, que diz (em essência) que uma variável aleatória definida como a média (ou soma) de um grande número de variáveis aleatórias independentes e identicamente distribuídas é aproximadamente distribuída normalmente. Em outras palavras, a distribuição amostral de médias segue uma normal.
Em detalhes, se $X_1, ..., X_n$ são variáveis aleatórias. Em particular, todas as VAs foram amostradas de uma mesma população com média $\mu$ (finita), desvio padrão $\sigma$ (finito). Além do mais, a geração de cada VA é independente da outra, sendo toas identicamente distribuídas. Quando $n$ é grande, então
$$\frac{1}{n}(X_1 + \cdots + X_n)$$
é aproximadamente distribuído por uma Normal com média $\mu$ e desvio padrão $\sigma/\sqrt{n}$. De forma equivalente (mas muitas vezes mais útil),
$$Z = \frac{(X_1 + \cdots + X_n) - \mu }{\sigma / \sqrt{n}}$$
é aproximadamente uma normal com média 0 e desvio padrão 1.
$$Z \sim Normal(0, 1).$$
### Como transformar VAs?
Lemebre-se da aula passada que sabemos estimar:
$$\bar{x} \approx \mu$$
e
$$s^2 \approx \sigma$$
Além do mais, sabemos que a variância do estimador da média é:
$$Var(\hat{\mu}) = \frac{\sigma^2}{n}$$
Assim:
\begin{align}
\bar{X} \sim Normal(\mu, \frac{\sigma^2}{n}) \\
\bar{X}- \mu \sim Normal(0, \frac{\sigma^2}{n}) \\
\frac{\bar{X}- \mu}{\sigma / \sqrt{n}} \sim Normal(0, 1) \\
\end{align}
## Exemplo das Moedas
Considere o caso de uma moeda sendo jogada para cima. Agora, escolhe um número `n` (tamanho da amostra), e gere amostras da mesma. Ou seja, jogue uma moeda para cima `n` vezes. Por fim, some quantas vezes a moeda cai em `cara` (ou `coroa`). Isto é uma soma para uma amostra de tamanho `n`.
O processo de geração destes dados é bem capturado por uma distribuição Binomial. Variáveis aleatórias binomiais, que possuem dois parâmetros $n$ e $p$. A distribuição binomial é útil para contar o número de sucessos $n$ dada uma probabilidade $p$. Por exemplo, quantas vezes ($n$) uma moeda ($p$) gera um o valor cara. Formalmente, uma variável aleatória Binomial($n, p$) é simplesmente a soma de $n$ variáveis aleatórias independentes de Bernoulli($p$), cada uma delas igual a $1$ com probabilidade $p$ e $0$ com probabilidade $1 - p$.
Ao gerar um valor de uma Binomial estamos falando "Jogue uma moeda para cima n vezes e conte quantas caras!". No caso abaixo, jogue uma moeda para cima 5 vezes e conte quantas caras!
```
num_caras = np.random.binomial(5, 0.5)
num_caras
```
Vamos repetir o processo várias! Jogue uma moeda para cima 5 vezes, pare, respire, depois jogue mais 5. Por aí vai. Note que temos a contagem para cada experimento de tamanho 5.
```
np.random.binomial(5, 0.5, size=10)
```
Agora, vamos ver o gráfico de tal experimento!
```
num_caras_a_cada_5 = np.random.binomial(5, 0.5, size=10000)
plt.hist(num_caras_a_cada_5, bins=[-0.5, 0.5, 1.5, 2.5, 3.5, 4.5, 5.5], edgecolor='k')
plt.xlabel('Jogadas de moedas')
plt.ylabel('Número de caras')
plt.title('1000 jogadas de moedas em grupos de 5')
despine()
```
Mesma coisa com 10 moedas
```
num_caras_a_cada_5 = np.random.binomial(10, 0.5, size=1000000)
ticks = np.linspace(-0.5, 10.5, num=12)
print(ticks)
plt.hist(num_caras_a_cada_5, bins=ticks, edgecolor='k')
plt.xlabel('Jogadas de moedas')
plt.ylabel('Número de caras')
plt.title('1000 jogadas de moedas em grupos de 5')
despine()
```
Lembrando da sua aula de probabilidade, uma população que segue uma variável Binomial($n, p$) tem média $\mu = np$ e desvio padrão $\sigma = \sqrt{np(1 - p)}$. Se plotarmos ambos, você pode facilmente ver a semelhança. Obseve o plot abaixo da PDF com os parâmetros que listamos (média, desvio padrão).
```
import scipy.stats as ss
mean = 10 * 0.5
std = np.sqrt(10 * 0.5 *(1 - 0.5))
x = np.linspace(1, 11, 1000)
y = ss.distributions.norm.pdf(loc=mean, scale=std, x=x)
plt.xlim(0, 10)
plt.plot(x, y)
plt.xlabel('Número caras - x')
plt.ylabel('P(X = x)')
despine()
mean = 10 * 0.5
std = np.sqrt(10 * 0.5 *(1 - 0.5))
x = np.linspace(1, 11, 1000)
y = ss.distributions.norm.pdf(loc=mean, scale=std, x=x)
plt.xlim(0, 10)
plt.plot(x, y, label='Aproximação Normal')
num_caras_a_cada_5 = np.random.binomial(10, 0.5, size=1000000)
ticks = np.linspace(-0.5, 10.5, num=12)
plt.hist(num_caras_a_cada_5, bins=ticks, edgecolor='k', label='Dados',
density=True)
plt.plot(x, y)
plt.xlabel('Jogadas de moedas')
plt.ylabel('Número de caras')
plt.title('1000 jogadas de moedas em grupos de 5')
plt.legend()
despine()
```
## Exemplo com Dados Sintéticos de Matrículas
Para exemplificar com dados, considere o exemplo abaixo onde geramos uma distribuição sintética de 25 mil alunos da UFMG. A distribuição captura o número de matéria que um aluno se matrícula no ano. Note que diferente da moeda, que gera apenas cara ou cora, cada aluno pode se matricular entre [min, max] matérias. No exemplo, vamos suport que todo aluno precisa se matricular em pelo menos uma matéria `min=1` e pode ser matricular em no máximo 10 matérias `max=10`. Agora, vamos suport que cada número de matéria tem probabilidade $p_i$. Ou seja, a chance se matricular em uma matéria é $p_1$ e por aí vai.
Dados deste tipo são modelados por distribuições multinomiais. Generalizando a Binomial, uma Multinomial conta a quantidade de sucessos (matrículas) em cada $p_i$. A mesma é definida por $n > 0$, número de amostras ou matrículas, e $p_1, \ldots, p_k$ probabilidade de se matrícular em $i$ matérias. A pmf de uma multinomial é dada por:
$$P(X = x) = \frac{n!}{x_1!\cdots x_k!} p_1^{x_1} \cdots p_k^{x_k}$$
Inicialmente observe os valores de $p_i$.
```
num_materias = np.arange(10) + 1
prob_materias = np.array([6, 7, 16, 25, 25, 25, 10, 12, 2, 11])
prob_materias = prob_materias / prob_materias.sum()
plt.bar(num_materias, prob_materias, edgecolor='k')
plt.xlabel('Número de Matérias no Semestre')
plt.ylabel('Fração de Alunos')
plt.title('Distribuição do número de matérias no semestre')
despine()
```
Agora vamos responder a pergunta: **Quantas matérias, em média, um aluno se matrícula?!**. Note que a nossa pergunta aqui é **em média!!**. Então, vamos considerar que temos 25 mil discente na ufmg. Para cada um destes alunos, vamos amostrar de $p_i$ o número de matérias que tal aluno está matrículado no atual semestre.
```
amostras = 25000
mats = np.arange(10) + 1
print(mats)
dados = []
for i in range(25000):
n_mat = np.random.choice(mats, p=prob_materias)
dados.append(n_mat)
dados = np.array(dados)
dados
```
Agora vamos responder nossa pergunta. **Quantas matérias, em média, um aluno se matrícula?!**. Para tirar uma média precisamos de uma amostra. Vamos definir amostras de tamanho 100. Então, vamos amostrar 100 alunos, **com repetição**, dos nossos 25 mil alunos.
```
n_amostra = 100
soma = 0
for i in range(n_amostra):
aluno = np.random.randint(0, len(dados))
num_mat = dados[aluno]
soma += num_mat
media = soma / n_amostra
print(media)
```
Vamos repetir o processo algumas vezes. Tipo, 10000 vezes.
```
n_amostra = 100
medias = []
for _ in range(10000):
soma = 0
for i in range(n_amostra):
aluno = np.random.randint(0, len(dados))
num_mat = dados[aluno]
soma += num_mat
media = soma / n_amostra
medias.append(media)
medias = np.array(medias)
```
Agora vamos ver os resultados!
```
plt.hist(medias, bins=20, edgecolor='k')
plt.ylabel('P(X = x)')
plt.xlabel('Média das matérias - x')
plt.title('CLT na Prática')
despine()
```
Agora, vamos comparar com a nossa Normal, para isto podemos usar a média das médias e o desvio padrão das médias.
```
mean = np.mean(medias)
# ddof=1 faz dividir por n-1
std = np.std(medias, ddof=1)
# pegue 1000 números entre o minimo e o max
x = np.linspace(np.min(medias), np.max(medias), 1000)
y = ss.distributions.norm.pdf(loc=mean, scale=std, x=x)
plt.plot(x, y, label='Aproximação Normal')
plt.hist(medias, bins=20, edgecolor='k', density=True)
plt.ylabel('P(X = x)')
plt.xlabel('Média das matérias - x')
plt.title('CLT na Prática')
despine()
```
## Com Dados
```
df = pd.read_csv('https://media.githubusercontent.com/media/icd-ufmg/material/master/aulas/03-Tabelas-e-Tipos-de-Dados/nba_salaries.csv')
df.head()
df['SALARY'].sort_values().plot.hist(bins=20, edgecolor='k')
plt.xlabel('Salário na NBA')
plt.ylabel('Número de Linhas')
despine()
N = 10000
data = df['SALARY']
medias = []
for i in range(N):
mean = np.random.choice(data, 100).mean()
medias.append(mean)
mean = np.mean(medias)
# ddof=1 faz dividir por n-1
std = np.std(medias, ddof=1)
# pegue 1000 números entre o minimo e o max
x = np.linspace(np.min(medias), np.max(medias), 1000)
y = ss.distributions.norm.pdf(loc=mean, scale=std, x=x)
plt.plot(x, y, label='Aproximação Normal')
plt.hist(medias, bins=20, edgecolor='k', density=True)
plt.ylabel('P(X = x)')
plt.xlabel('Salário da NBA - x')
plt.title('CLT na Prática')
despine()
```
## Condições para o TCL
Existem algumas condições para garantir que o TCL seja válido.
1. Dados independentes e identicamente distribuídos.
1. Variância finita.
1. Pelo menos umas 30 amostras
Observe do wikipedia que uma distribuição Pareto(1) tem variância infinita. Quebramos nossa condição. Olhe que o plot abaixo não parece em nada com uma Normal.
https://en.wikipedia.org/wiki/Pareto_distribution
```
data = []
for _ in range(10000):
m = np.random.pareto(1, size=100).mean()
data.append(m)
plt.hist(data, bins=100, edgecolor='k')
despine()
```
Podemos quebrar também com amostras muito pequenas, tipo na Beta(3, 2, size=2) abaixo.
Observe como é muito perto de uma Normal mas tem um certo viés para a direita.
```
data = []
for _ in range(10000):
m = np.random.beta(3, 2, size=2).mean()
data.append(m)
plt.hist(data, edgecolor='k')
despine()
mean = np.mean(data)
# ddof=1 faz dividir por n-1
std = np.std(data, ddof=1)
# pegue 1000 números entre o minimo e o max
x = np.linspace(np.min(data), np.max(data), 1000)
y = ss.distributions.norm.pdf(loc=mean, scale=std, x=x)
plt.plot(x, y, label='Aproximação Normal')
plt.hist(data, bins=20, edgecolor='k', density=True)
plt.ylabel('P(X = x)')
plt.title('CLT na Prática')
despine()
```
| github_jupyter |
# Assignment
- Kannada MNIST를 이용한 미니 Competetion
```
import numpy as np
import pandas as pd
import torch
from torch.autograd import Variable as Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.model_zoo as model_zoo
from torch.optim import lr_scheduler
from collections import OrderedDict
import torchvision
from torch.utils.data import DataLoader, Dataset
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import time
import copy
import os
plt.ion()
train = pd.read_csv("train_df.csv")
train.shape
test = pd.read_csv("test_df.csv")
test.head()
test.shape
X = train.iloc[:,1:]
X = X / 255
Y =train.iloc[:,0]
from sklearn.model_selection import train_test_split
X_train,X_val,Y_train,Y_val = train_test_split(X, Y, train_size=0.85,random_state=10, stratify = train.label)
pd.DataFrame(pd.concat([X_train,Y_train],axis=1)).to_csv("train.csv", index =False)
pd.DataFrame(pd.concat([X_val,Y_val],axis=1)).to_csv("val.csv", index = False)
from torch.utils.data import DataLoader
class BengaliDataset(Dataset):
def __init__(self, csv, transform=None):
# super(BengaliDataset, self).__init__(transform=transform)
self.csv = pd.read_csv(csv)
self.image_ids = self.csv.index
self.transform = transform
def __len__(self):
return len(self.csv)
def __getitem__(self, index):
image = self.csv.iloc[index,:-1]
label = self.csv.iloc[index,-1]
return (torch.tensor(image, dtype=torch.float), torch.tensor(label, dtype=torch.long))
trainset = BengaliDataset("train.csv")
valset = BengaliDataset("val.csv")
train_loader = DataLoader(trainset,batch_size = 128, shuffle= True)
test_loader = DataLoader(valset,batch_size = 128, shuffle= True) # validation
```
### 모델 설명
MINIST로 학습된 pretrained 모델을 불러와서 fine tunning을 해보자.
MLP로 구축된 모델로, MINIST에서 약 98%의 정확도를 보여줌.
http://ml.cs.tsinghua.edu.cn/~chenxi/pytorch-models/mnist-b07bb66b.pth' 에 저장되어 있는 weight을 가져와서 layer에 initailization.
```
class YEJI(nn.Module):
def __init__(self, input_dims, n_hiddens, n_class):
super(YEJI, self).__init__()
self.input_dims = input_dims
current_dims = input_dims
layers = OrderedDict()
if isinstance(n_hiddens, int):
n_hiddens = [n_hiddens]
else:
n_hiddens = list(n_hiddens)
for i, n_hidden in enumerate(n_hiddens):
layers['fc{}'.format(i+1)] = nn.Linear(current_dims, n_hidden)
layers['relu{}'.format(i+1)] = nn.ReLU()
layers['drop{}'.format(i+1)] = nn.Dropout(0.2)
current_dims = n_hidden
layers['out'] = nn.Linear(current_dims, n_class)
self.model= nn.Sequential(layers)
print(self.model)
def forward(self, input):
return self.model.forward(input)
def TOBIGS(input_dims=784, n_hiddens=[256, 256], n_class=10):
model = YEJI(input_dims, n_hiddens, n_class)
model_urls = {'mnist': 'http://ml.cs.tsinghua.edu.cn/~chenxi/pytorch-models/mnist-b07bb66b.pth'}
m = model_zoo.load_url(model_urls['mnist'])
state_dict = m.state_dict() if isinstance(m, nn.Module) else m
assert isinstance(state_dict, (dict, OrderedDict)), type(state_dict)
model.load_state_dict(state_dict)
return model
model = TOBIGS(input_dims=784, n_hiddens=[256, 256], n_class=10)
model = torch.nn.DataParallel(model)
model.cuda()
# optimizer
optimizer = optim.Adam(model.parameters(), lr=0.0001)
```
- 초기화된 parameters 확인
```
for i in model.parameters():
print(i)
def expand_user(path):
return os.path.abspath(os.path.expanduser(path))
def model_snapshot(model, new_file, old_file=None, verbose = True):
if isinstance(model, torch.nn.DataParallel):
model = model.module
if old_file and os.path.exists(expand_user(old_file)):
print("Removing old model {}".format(expand_user(old_file)))
os.remove(expand_user(old_file))
if verbose:
print("Saving model to {}".format(expand_user(new_file)))
state_dict = OrderedDict()
for k, v in model.state_dict().items():
if v.is_cuda:
v = v.cpu()
state_dict[k] = v
torch.save(state_dict, expand_user(new_file))
best_acc = 0.0
old_file = None
for epoch in range(10):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
indx_target = target.clone()
data, target = data.cuda(), target.cuda()
data, target = Variable(data), Variable(target)
optimizer.zero_grad()
output = model(data)
# output = nn.Softmax(output)
# loss = F.mse_loss(output, target)
loss=F.cross_entropy(output, target)
loss.backward()
optimizer.step()
if batch_idx % 10 ==0 and batch_idx > 0:
pred = output.data.max(1)[1] # get the index of the max log-probability
correct = pred.cpu().eq(indx_target).sum()
acc = correct * 1.0 / len(data)
print('Train Epoch: {} [{}/{}] Loss: {:.6f} Acc: {:.4f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
loss, acc))
model.eval()
test_loss = 0
correct = 0
for data, target in test_loader:
indx_target = target.clone()
data, target = data.cuda(), target.cuda()
data, target = Variable(data, volatile=True), Variable(target)
output = model(data)
test_loss += F.cross_entropy(output, target)
pred = output.data.max(1)[1]
correct += pred.cpu().eq(indx_target).sum()
test_loss = test_loss / len(test_loader)
acc = 100. * correct / float(len(test_loader.dataset))
print('\tTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)'.format(
test_loss, correct, len(test_loader.dataset), acc))
if acc > best_acc:
new_file = os.path.join('./', 'best-{}.pth'.format(epoch))
model_snapshot(model, new_file, old_file=old_file)
best_acc = acc
old_file = new_file
for epoch in range(20):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
indx_target = target.clone()
data, target = data.cuda(), target.cuda()
data, target = Variable(data), Variable(target)
optimizer.zero_grad()
output = model(data)
# output = nn.Softmax(output)
# loss = F.mse_loss(output, target)
loss=F.cross_entropy(output, target)
loss.backward()
optimizer.step()
if batch_idx % 10 ==0 and batch_idx > 0:
pred = output.data.max(1)[1] # get the index of the max log-probability
correct = pred.cpu().eq(indx_target).sum()
acc = correct * 1.0 / len(data)
print('Train Epoch: {} [{}/{}] Loss: {:.6f} Acc: {:.4f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
loss, acc))
model.eval()
test_loss = 0
correct = 0
for data, target in test_loader:
indx_target = target.clone()
data, target = data.cuda(), target.cuda()
data, target = Variable(data, volatile=True), Variable(target)
output = model(data)
test_loss += F.cross_entropy(output, target)
pred = output.data.max(1)[1]
correct += pred.cpu().eq(indx_target).sum()
test_loss = test_loss / len(test_loader)
acc = 100. * correct / float(len(test_loader.dataset))
print('\tTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)'.format(
test_loss, correct, len(test_loader.dataset), acc))
if acc > best_acc:
new_file = os.path.join('./', 'best3-{}.pth'.format(epoch))
model_snapshot(model, new_file, old_file=old_file)
best_acc = acc
old_file = new_file
test
X_test = torch.tensor(test.iloc[:,1:].values / 255,dtype=torch.float)
X_test
best_model = TOBIGS(input_dims=784, n_hiddens=[256, 256], n_class=10)
best_model.load_state_dict(torch.load('./best3-14.pth'))
best_model.cuda()
best_model.eval()
X_test = Variable(X_test.cuda())
pred = best_model(X_test)
sub = pd.read_csv("sample_submission.csv")
sub["Category"] = pred.argmax(axis=1).cpu().numpy()
sub.to_csv("torch_pred.csv")
```
| github_jupyter |

Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Logging
_**This notebook showcases various ways to use the Azure Machine Learning service run logging APIs, and view the results in the Azure portal.**_
---
---
## Table of Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. Validate Azure ML SDK installation
1. Initialize workspace
1. Set experiment
1. [Logging](#Logging)
1. Starting a run
1. Viewing a run in the portal
1. Viewing the experiment in the portal
1. Logging metrics
1. Logging string metrics
1. Logging numeric metrics
1. Logging vectors
1. Logging tables
1. Uploading files
1. [Analyzing results](#Analyzing-results)
1. Tagging a run
1. [Next steps](#Next-steps)
## Introduction
Logging metrics from runs in your experiments allows you to track results from one run to another, determining trends in your outputs and understand how your inputs correspond to your model and script performance. Azure Machine Learning services (AzureML) allows you to track various types of metrics including images and arbitrary files in order to understand, analyze, and audit your experimental progress.
Typically you should log all parameters for your experiment and all numerical and string outputs of your experiment. This will allow you to analyze the performance of your experiments across multiple runs, correlate inputs to outputs, and filter runs based on interesting criteria.
The experiment's Run History report page automatically creates a report that can be customized to show the KPI's, charts, and column sets that are interesting to you.
|  |  |
|:--:|:--:|
| *Run Details* | *Run History* |
---
## Setup
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, go through the [configuration](../../../configuration.ipynb) Notebook first if you haven't already to establish your connection to the AzureML Workspace. Also make sure you have tqdm and matplotlib installed in the current kernel.
```
(myenv) $ conda install -y tqdm matplotlib
```
### Validate Azure ML SDK installation and get version number for debugging purposes
```
from azureml.core import Experiment, Workspace, Run
import azureml.core
import numpy as np
from tqdm import tqdm
# Check core SDK version number
print("This notebook was created using SDK version 1.3.0, you are currently running version", azureml.core.VERSION)
```
### Initialize workspace
Initialize a workspace object from persisted configuration.
```
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep='\n')
```
### Set experiment
Create a new experiment (or get the one with the specified name). An *experiment* is a container for an arbitrary set of *runs*.
```
experiment = Experiment(workspace=ws, name='logging-api-test')
```
---
## Logging
In this section we will explore the various logging mechanisms.
### Starting a run
A *run* is a singular experimental trial. In this notebook we will create a run directly on the experiment by calling `run = exp.start_logging()`. If you were experimenting by submitting a script file as an experiment using ``experiment.submit()``, you would call `run = Run.get_context()` in your script to access the run context of your code. In either case, the logging methods on the returned run object work the same.
This cell also stores the run id for use later in this notebook. The run_id is not necessary for logging.
```
# start logging for the run
run = experiment.start_logging()
# access the run id for use later
run_id = run.id
# change the scale factor on different runs to see how you can compare multiple runs
scale_factor = 2
# change the category on different runs to see how to organize data in reports
category = 'Red'
```
#### Viewing a run in the Portal
Once a run is started you can see the run in the portal by simply typing ``run``. Clicking on the "Link to Portal" link will take you to the Run Details page that shows the metrics you have logged and other run properties. You can refresh this page after each logging statement to see the updated results.
```
run
```
### Viewing an experiment in the portal
You can also view an experiement similarly by typing `experiment`. The portal link will take you to the experiment's Run History page that shows all runs and allows you to analyze trends across multiple runs.
```
experiment
```
## Logging metrics
Metrics are visible in the run details page in the AzureML portal and also can be analyzed in experiment reports. The run details page looks as below and contains tabs for Details, Outputs, Logs, and Snapshot.
* The Details page displays attributes about the run, plus logged metrics and images. Metrics that are vectors appear as charts.
* The Outputs page contains any files, such as models, you uploaded into the "outputs" directory from your run into storage. If you place files in the "outputs" directory locally, the files are automatically uploaded on your behald when the run is completed.
* The Logs page allows you to view any log files created by your run. Logging runs created in notebooks typically do not generate log files.
* The Snapshot page contains a snapshot of the directory specified in the ''start_logging'' statement, plus the notebook at the time of the ''start_logging'' call. This snapshot and notebook can be downloaded from the Run Details page to continue or reproduce an experiment.
### Logging string metrics
The following cell logs a string metric. A string metric is simply a string value associated with a name. A string metric String metrics are useful for labelling runs and to organize your data. Typically you should log all string parameters as metrics for later analysis - even information such as paths can help to understand how individual experiements perform differently.
String metrics can be used in the following ways:
* Plot in hitograms
* Group by indicators for numerical plots
* Filtering runs
String metrics appear in the **Tracked Metrics** section of the Run Details page and can be added as a column in Run History reports.
```
# log a string metric
run.log(name='Category', value=category)
```
### Logging numerical metrics
The following cell logs some numerical metrics. Numerical metrics can include metrics such as AUC or MSE. You should log any parameter or significant output measure in order to understand trends across multiple experiments. Numerical metrics appear in the **Tracked Metrics** section of the Run Details page, and can be used in charts or KPI's in experiment Run History reports.
```
# log numerical values
run.log(name="scale factor", value = scale_factor)
run.log(name='Magic Number', value=42 * scale_factor)
```
### Logging vectors
Vectors are good for recording information such as loss curves. You can log a vector by creating a list of numbers, calling ``log_list()`` and supplying a name and the list, or by repeatedly logging a value using the same name.
Vectors are presented in Run Details as a chart, and are directly comparable in experiment reports when placed in a chart.
**Note:** vectors logged into the run are expected to be relatively small. Logging very large vectors into Azure ML can result in reduced performance. If you need to store large amounts of data associated with the run, you can write the data to file that will be uploaded.
```
fibonacci_values = [0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89]
scaled_values = (i * scale_factor for i in fibonacci_values)
# Log a list of values. Note this will generate a single-variable line chart.
run.log_list(name='Fibonacci', value=scaled_values)
for i in tqdm(range(-10, 10)):
# log a metric value repeatedly, this will generate a single-variable line chart.
run.log(name='Sigmoid', value=1 / (1 + np.exp(-i)))
```
### Logging tables
Tables are good for recording related sets of information such as accuracy tables, confusion matrices, etc.
You can log a table in two ways:
* Create a dictionary of lists where each list represents a column in the table and call ``log_table()``
* Repeatedly call ``log_row()`` providing the same table name with a consistent set of named args as the column values
Tables are presented in Run Details as a chart using the first two columns of the table
**Note:** tables logged into the run are expected to be relatively small. Logging very large tables into Azure ML can result in reduced performance. If you need to store large amounts of data associated with the run, you can write the data to file that will be uploaded.
```
# create a dictionary to hold a table of values
sines = {}
sines['angle'] = []
sines['sine'] = []
for i in tqdm(range(-10, 10)):
angle = i / 2.0 * scale_factor
# log a 2 (or more) values as a metric repeatedly. This will generate a 2-variable line chart if you have 2 numerical columns.
run.log_row(name='Cosine Wave', angle=angle, cos=np.cos(angle))
sines['angle'].append(angle)
sines['sine'].append(np.sin(angle))
# log a dictionary as a table, this will generate a 2-variable chart if you have 2 numerical columns
run.log_table(name='Sine Wave', value=sines)
```
### Logging images
You can directly log _matplotlib_ plots and arbitrary images to your run record. This code logs a _matplotlib_ pyplot object. Images show up in the run details page in the Azure ML Portal.
```
%matplotlib inline
# Create a plot
import matplotlib.pyplot as plt
angle = np.linspace(-3, 3, 50) * scale_factor
plt.plot(angle,np.tanh(angle), label='tanh')
plt.legend(fontsize=12)
plt.title('Hyperbolic Tangent', fontsize=16)
plt.grid(True)
# Log the plot to the run. To log an arbitrary image, use the form run.log_image(name, path='./image_path.png')
run.log_image(name='Hyperbolic Tangent', plot=plt)
```
### Uploading files
Files can also be uploaded explicitly and stored as artifacts along with the run record. These files are also visible in the *Outputs* tab of the Run Details page.
```
file_name = 'outputs/myfile.txt'
with open(file_name, "w") as f:
f.write('This is an output file that will be uploaded.\n')
# Upload the file explicitly into artifacts
run.upload_file(name = file_name, path_or_stream = file_name)
```
### Completing the run
Calling `run.complete()` marks the run as completed and triggers the output file collection. If for any reason you need to indicate the run failed or simply need to cancel the run you can call `run.fail()` or `run.cancel()`.
```
run.complete()
```
---
## Analyzing results
You can refresh the run in the Azure portal to see all of your results. In many cases you will want to analyze runs that were performed previously to inspect the contents or compare results. Runs can be fetched from their parent Experiment object using the ``Run()`` constructor or the ``experiment.get_runs()`` method.
```
fetched_run = Run(experiment, run_id)
fetched_run
```
Call ``run.get_metrics()`` to retrieve all the metrics from a run.
```
fetched_run.get_metrics()
```
Call ``run.get_metrics(name = <metric name>)`` to retrieve a metric value by name. Retrieving a single metric can be faster, especially if the run contains many metrics.
```
fetched_run.get_metrics(name = "scale factor")
```
See the files uploaded for this run by calling ``run.get_file_names()``
```
fetched_run.get_file_names()
```
Once you know the file names in a run, you can download the files using the ``run.download_file()`` method
```
import os
os.makedirs('files', exist_ok=True)
for f in run.get_file_names():
dest = os.path.join('files', f.split('/')[-1])
print('Downloading file {} to {}...'.format(f, dest))
fetched_run.download_file(f, dest)
```
### Tagging a run
Often when you analyze the results of a run, you may need to tag that run with important personal or external information. You can add a tag to a run using the ``run.tag()`` method. AzureML supports valueless and valued tags.
```
fetched_run.tag("My Favorite Run")
fetched_run.tag("Competition Rank", 1)
fetched_run.get_tags()
```
## Next steps
To experiment more with logging and to understand how metrics can be visualized, go back to the *Start a run* section, try changing the category and scale_factor values and going through the notebook several times. Play with the KPI, charting, and column selection options on the experiment's Run History reports page to see how the various metrics can be combined and visualized.
After learning about all of the logging options, go to the [train on remote vm](..\train-on-remote-vm\train-on-remote-vm.ipynb) notebook and experiment with logging from remote compute contexts.
| github_jupyter |
# Train
```
!pip install keras_applications==1.0.8
!pip install image-classifiers==1.0.0
!pip install efficientnet==1.0.0
!pip install segmentation_models==1.0.1
!pip install tensorflow==2.3.1
!pip install keras==2.4.3
!pip install tensorflow_addons
!pip install albumentations
!pip install imagecodecs
%env SM_FRAMEWORK=tf.keras
import warnings
warnings.filterwarnings('ignore')
import os
import gc
import cv2
import json
import time
import random
import numpy as np
import pandas as pd
import tifffile as tiff
import tensorflow as tf
import tensorflow_addons as tfa
import matplotlib.pyplot as plt
import albumentations as albu
from sklearn.model_selection import train_test_split, KFold, GroupKFold
import tensorflow.keras.backend as K
from tensorflow.keras import Model, Sequential
from tensorflow.keras.models import load_model
from tensorflow.keras.utils import Sequence
from tensorflow.keras.losses import binary_crossentropy
from tensorflow.keras.layers import *
from tensorflow.keras.optimizers import Adam, SGD
from tensorflow.keras.callbacks import *
import segmentation_models as sm
from segmentation_models import Unet, FPN
from segmentation_models.losses import bce_jaccard_loss
print('tensorflow version:', tf.__version__)
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
if gpu_devices:
for gpu_device in gpu_devices:
print('device available:', gpu_device)
#policy = tf.keras.mixed_precision.experimental.Policy('mixed_float16')
#tf.keras.mixed_precision.experimental.set_policy(policy)
pd.set_option('display.max_columns', None)
VER = 'v2'
PARAMS = {
'version': VER,
'folds': 4,
'img_size': 256,
'resize': 1,
'batch_size': 20,
'epochs': 1000,
'patience': 5,
'decay': False,
'backbone': 'efficientnetb2', # efficientnetbX, resnet34/50, resnext50, seresnet34, seresnext50
'bce_weight': 1.,
'loss': 'bce_jaccard_loss', # bce_jaccard_loss bce_dice
'seed': 2020,
'split': 'group',
'mirror': False,
'aughard': True,
'umodel': 'unet',
'pseudo': '',
'lr': .0002,
'shift': False,
'external': '', # 'None' otherwise
'comments': ''
}
DATA_PATH = '../data'
resize = PARAMS['resize']
size = PARAMS['img_size']
ext = PARAMS['external']
pseudo = PARAMS['pseudo']
if PARAMS['pseudo']:
IMGS_PATH = f'{DATA_PATH}/tiles_r{resize}_s{size}_{pseudo}/'
MSKS_PATH = f'{DATA_PATH}/masks_r{resize}_s{size}_{pseudo}/'
else:
if PARAMS['shift']:
if ext:
IMGS_PATH = f'{DATA_PATH}/tiles_r{resize}_s{size}_shft_{ext}/'
MSKS_PATH = f'{DATA_PATH}/masks_r{resize}_s{size}_shft_{ext}/'
else:
IMGS_PATH = f'{DATA_PATH}/tiles_r{resize}_s{size}_shft/'
MSKS_PATH = f'{DATA_PATH}/masks_r{resize}_s{size}_shft/'
else:
if ext:
IMGS_PATH = f'{DATA_PATH}/tiles_r{resize}_s{size}_{ext}/'
MSKS_PATH = f'{DATA_PATH}/masks_r{resize}_s{size}_{ext}/'
else:
IMGS_PATH = f'{DATA_PATH}/tiles_v2_1500_to256/'
MSKS_PATH = f'{DATA_PATH}/masks_v2_1500_t0256/'
MDLS_PATH = f'../models_{VER}'
if not os.path.exists(MDLS_PATH):
os.mkdir(MDLS_PATH)
with open(f'{MDLS_PATH}/params.json', 'w') as file:
json.dump(PARAMS, file)
if not PARAMS['mirror']:
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
STRATEGY = tf.distribute.get_strategy()
else:
STRATEGY = tf.distribute.MirroredStrategy()
def seed_all(seed):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
tf.random.set_seed(seed)
seed_all(PARAMS['seed'])
start_time = time.time()
```
# Utils
```
if PARAMS['aughard']:
aug = albu.Compose([
albu.OneOf([
albu.RandomBrightness(limit=.2, p=1),
albu.RandomContrast(limit=.2, p=1),
albu.RandomGamma(p=1)
], p=.5),
albu.OneOf([
albu.Blur(blur_limit=3, p=1),
albu.MedianBlur(blur_limit=3, p=1)
], p=.1),
albu.RandomRotate90(p=.5),
albu.HorizontalFlip(p=.5),
albu.VerticalFlip(p=.5),
albu.ShiftScaleRotate(p=.25)
])
else:
aug = albu.Compose([
albu.OneOf([
albu.RandomBrightness(limit=.2, p=1),
albu.RandomContrast(limit=.2, p=1),
albu.RandomGamma(p=1)
], p=.5),
albu.RandomRotate90(p=.25),
albu.HorizontalFlip(p=.25),
albu.VerticalFlip(p=.25)
])
class DataGenKid(Sequence):
def __init__(self, imgs_path, msks_path, imgs_idxs, img_size,
batch_size=32, mode='fit', shuffle=False,
aug=None, resize=None):
self.imgs_path = imgs_path
self.msks_path = msks_path
self.imgs_idxs = imgs_idxs
self.img_size = img_size
self.batch_size = batch_size
self.mode = mode
self.shuffle = shuffle
self.aug = aug
self.resize = resize
self.on_epoch_end()
# self.imgs_lst = os.listdir(imgs_path).sort()
# self.msks_lst = os.listdir(msks_path).sort()
def __len__(self):
return int(np.floor(len(self.imgs_idxs) / self.batch_size))
def on_epoch_end(self):
self.indexes = np.arange(len(self.imgs_idxs))
if self.shuffle:
np.random.shuffle(self.indexes)
def __getitem__(self, index):
batch_size = min(self.batch_size, len(self.imgs_idxs) - index*self.batch_size)
X = np.zeros((batch_size, self.img_size, self.img_size, 3), dtype=np.float32)
imgs_batch = self.imgs_idxs[index * self.batch_size : (index+1) * self.batch_size]
if self.mode == 'fit':
y = np.zeros((batch_size, self.img_size, self.img_size), dtype=np.float32)
for i, img_idx in enumerate(imgs_batch):
X[i, ], y[i] = self.get_tile(img_idx)
return X, y
elif self.mode == 'predict':
for i, img_idx in enumerate(imgs_batch):
X[i, ] = self.get_tile(img_idx)
return X
else:
raise AttributeError('fit mode parameter error')
def get_tile(self, img_idx):
img_path = f'{self.imgs_path}/{img_idx}'
img = np.load(img_path)
if img is None:
print('error load image:', img_path)
# if self.resize:
# img = cv2.resize(img, (int(img.shape[1] / self.resize), int(img.shape[0] / self.resize)))
img = img.astype(np.float32)
if self.mode == 'fit':
msk_path = f'{self.msks_path}/{img_idx}'
msk = np.load(msk_path)
if msk is None:
print('error load mask:', msk_path)
# if self.resize:
# msk = cv2.resize(msk, (int(msk.shape[1] / self.resize), int(msk.shape[0] / self.resize)))
msk = msk.astype(np.float32)
if self.aug:
augmented = self.aug(image=img, mask=msk)
img = augmented['image']
msk = augmented['mask']
return img, msk
else:
if self.aug:
img = self.aug(image=img)['image']
return img
imgs_idxs = os.listdir(IMGS_PATH)
train_datagen = DataGenKid(
imgs_path=IMGS_PATH,
msks_path=MSKS_PATH,
imgs_idxs=imgs_idxs,
img_size=PARAMS['img_size'],
batch_size=PARAMS['batch_size'],
mode='fit',
shuffle=True,
aug=aug,
resize=None
)
val_datagen = DataGenKid(
imgs_path=IMGS_PATH,
msks_path=MSKS_PATH,
imgs_idxs=imgs_idxs,
img_size=PARAMS['img_size'],
batch_size=PARAMS['batch_size'],
mode='fit',
shuffle=False,
aug=None,
resize=None
)
bsize = min(8, PARAMS['batch_size'])
Xt, yt = train_datagen.__getitem__(0)
print('test X: ', Xt.shape)
print('test y: ', yt.shape)
fig, axes = plt.subplots(figsize=(16, 4), nrows=2, ncols=bsize)
for j in range(bsize):
axes[0, j].imshow(Xt[j])
axes[0, j].set_title(j)
axes[0, j].axis('off')
axes[1, j].imshow(yt[j])
axes[1, j].axis('off')
plt.show()
def dice_coef(y_true, y_pred, smooth=1):
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
return (2 * intersection + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
def dice_loss(y_true, y_pred, smooth=1):
return (1 - dice_coef(y_true, y_pred, smooth))
def bce_dice_loss(y_true, y_pred):
return PARAMS['bce_weight'] * binary_crossentropy(y_true, y_pred) + \
(1 - PARAMS['bce_weight']) * dice_loss(y_true, y_pred)
def get_model(backbone, input_shape, loss_type='bce_dice',
umodel='unet', classes=1, lr=.001):
with STRATEGY.scope():
if loss_type == 'bce_dice':
loss = bce_dice_loss
elif loss_type == 'bce_jaccard_loss':
loss = bce_jaccard_loss
else:
raise AttributeError('loss mode parameter error')
if umodel == 'unet':
model = Unet(backbone_name=backbone, encoder_weights='imagenet',
input_shape=input_shape,
classes=classes, activation='sigmoid')
elif umodel == 'fpn':
model = FPN(backbone_name=backbone, encoder_weights='imagenet',
input_shape=input_shape,
classes=classes, activation='sigmoid')
else:
raise AttributeError('umodel mode parameter error')
model.compile(
optimizer=tfa.optimizers.Lookahead(
tf.keras.optimizers.Adam(learning_rate=lr),
sync_period=max(6, int(PARAMS['patience'] / 4))
),
loss=loss,
metrics=[dice_coef]
)
return model
def get_lr_callback(batch_size=10, epochs=100, warmup=5, plot=False):
lr_start = 1e-5
lr_max = 1e-3
lr_min = lr_start / 100
lr_ramp_ep = warmup
lr_sus_ep = 0
lr_decay = .95
def lr_scheduler(epoch):
if epoch < lr_ramp_ep:
lr = (lr_max - lr_start) / lr_ramp_ep * epoch + lr_start
elif epoch < lr_ramp_ep + lr_sus_ep:
lr = lr_max
else:
lr = (lr_max - lr_min) * lr_decay ** (epoch - lr_ramp_ep - lr_sus_ep) + lr_min
return lr
if not plot:
lr_callback = tf.keras.callbacks.LearningRateScheduler(lr_scheduler, verbose=False)
return lr_callback
else:
return lr_scheduler
if PARAMS['decay']:
lr_scheduler_plot = get_lr_callback(
batch_size=PARAMS['batch_size'],
epochs=PARAMS['epochs'],
plot=True
)
xs = [i for i in range(PARAMS['epochs'])]
y = [lr_scheduler_plot(x) for x in xs]
plt.plot(xs, y)
plt.title(f'lr schedule from {y[0]:.5f} to {max(y):.3f} to {y[-1]:.8f}')
plt.show()
```
# Train
```
def train_model(mparams, n_fold, train_datagen, val_datagen):
model = get_model(
mparams['backbone'],
input_shape=(mparams['img_size'], mparams['img_size'], 3),
loss_type=mparams['loss'],
umodel=mparams['umodel'],
lr=mparams['lr']
)
checkpoint_path = f'{MDLS_PATH}/model_{n_fold}.hdf5'
earlystopper = EarlyStopping(
monitor='val_dice_coef',
patience=mparams['patience'],
verbose=0,
restore_best_weights=True,
mode='max'
)
lrreducer = ReduceLROnPlateau(
monitor='val_dice_coef',
factor=.1,
patience=int(mparams['patience'] / 2),
verbose=0,
min_lr=1e-7,
mode='max'
)
checkpointer = ModelCheckpoint(
checkpoint_path,
monitor='val_dice_coef',
verbose=0,
save_best_only=True,
save_weights_only=True,
mode='max'
)
callbacks = [earlystopper, checkpointer]
if mparams['decay']:
callbacks.append(get_lr_callback(mparams['batch_size']))
print('lr warmup and decay')
else:
callbacks.append(lrreducer)
print('lr reduce on plateau')
history = model.fit(
train_datagen,
validation_data=val_datagen,
callbacks=callbacks,
epochs=mparams['epochs'],
verbose=1
)
history_file = f'{MDLS_PATH}/history_{n_fold}.json'
dict_to_save = {}
for k, v in history.history.items():
dict_to_save.update({k: [np.format_float_positional(x) for x in history.history[k]]})
with open(history_file, 'w') as file:
json.dump(dict_to_save, file)
model.load_weights(checkpoint_path)
return model, history
for iname in list(set([x[:9] for x in imgs_idxs])):
print('img name:', iname,
'| imgs number:', len([x for x in imgs_idxs if x[:9] == iname]))
if PARAMS['split'] == 'kfold':
kfold = KFold(n_splits=PARAMS['folds'],
random_state=PARAMS['seed'],
shuffle=True).split(imgs_idxs)
elif PARAMS['split'] == 'group':
grps = [x[:9] for x in imgs_idxs]
kfold = GroupKFold(n_splits=PARAMS['folds']).split(imgs_idxs, imgs_idxs, grps)
elif PARAMS['split'] == 'twos':
grps = []
for x in imgs_idxs:
if x[:9] in ['e79de561c', '1e2425f28']: grps.append(0)
elif x[:9] in ['cb2d976f4', 'aaa6a05cc']: grps.append(1)
elif x[:9] in ['095bf7a1f', '54f2eec69']: grps.append(2)
else: grps.append(3)
kfold = GroupKFold(n_splits=PARAMS['folds']).split(imgs_idxs, imgs_idxs, grps)
else:
raise AttributeError('split mode parameter error')
epoch_by_folds = []
loss_by_folds = []
dice_coef_by_folds = []
passed = 0
if passed:
epoch_by_folds.extend([0])
loss_by_folds.extend([0])
dice_coef_by_folds.extend([0])
for n, (tr, te) in enumerate(kfold):
print('=' * 10, f'FOLD {n}', '=' * 10)
X_tr = [imgs_idxs[i] for i in tr]; X_val = [imgs_idxs[i] for i in te]
print('train:', len(X_tr), '| test:', len(X_val))
print('groups train:', set([x[:9] for x in X_tr]),
'\ngroups test:', set([x[:9] for x in X_val]))
if passed > 0:
print('fold done already')
passed -= 1
else:
train_datagen = DataGenKid(
imgs_path=IMGS_PATH,
msks_path=MSKS_PATH,
imgs_idxs=X_tr,
img_size=PARAMS['img_size'],
batch_size=PARAMS['batch_size'],
mode='fit',
shuffle=True,
aug=aug,
resize=None
)
val_datagen = DataGenKid(
imgs_path=IMGS_PATH,
msks_path=MSKS_PATH,
imgs_idxs=X_val,
img_size=PARAMS['img_size'],
batch_size=PARAMS['batch_size'],
mode='fit',
shuffle=False,
aug=None,
resize=None
)
model, history = train_model(PARAMS, n, train_datagen, val_datagen)
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.legend()
plt.show()
plt.plot(history.history['dice_coef'], label='dice_coef')
plt.plot(history.history['val_dice_coef'], label='val_dice_coef')
plt.legend()
plt.show()
best_epoch = np.argmax(history.history['val_dice_coef'])
best_loss = history.history['val_loss'][best_epoch]
best_dice_coef = history.history['val_dice_coef'][best_epoch]
print('best epoch:', best_epoch,
'| best loss:', best_loss,
'| best dice coef:', best_dice_coef)
epoch_by_folds.append(best_epoch)
loss_by_folds.append(best_loss)
dice_coef_by_folds.append(best_dice_coef)
del train_datagen, val_datagen, model; gc.collect()
elapsed_time = time.time() - start_time
print(f'time elapsed: {elapsed_time // 60:.0f} min {elapsed_time % 60:.0f} sec')
result = PARAMS.copy()
result['bavg_epoch'] = np.mean(epoch_by_folds)
result['bavg_loss'] = np.mean(loss_by_folds)
result['bavg_dice_coef'] = np.mean(dice_coef_by_folds)
result['dice_by_folds'] = ' '.join([f'{x:.4f}' for x in dice_coef_by_folds])
with open(f'{MDLS_PATH}/params.json', 'w') as file:
json.dump(result, file)
if not os.path.exists('results.csv'):
df_save = pd.DataFrame(result, index=[0])
df_save.to_csv('results.csv', sep='\t')
else:
df_old = pd.read_csv('results.csv', sep='\t', index_col=0)
df_save = pd.DataFrame(result, index=[df_old.index.max() + 1])
df_save = df_old.append(df_save, ignore_index=True)
df_save.to_csv('results.csv', sep='\t')
pd.read_csv('results.csv', sep='\t', index_col=0)
```
# Predict
```
larger = 4
test_models = []
for n_fold in list(range(PARAMS['folds'])):
checkpoint_path = f'{MDLS_PATH}/model_{n_fold}.hdf5'
print(checkpoint_path)
model_lrg = get_model(
PARAMS['backbone'],
input_shape=(PARAMS['img_size'] * larger, PARAMS['img_size'] * larger, 3),
loss_type=PARAMS['loss'],
umodel=PARAMS['umodel']
)
model_lrg.load_weights(checkpoint_path) # or .set_weights(model.get_weights()) from smaller model
test_models.append(model_lrg)
!ls ../data
img_num = 0
resize = PARAMS['resize']
shft = .6
wnd = PARAMS['img_size'] * larger
img = plt.imread('../data/patches/S 050001807_6_PAS_ind_00_X0_Y0_005727_025687_img.png')
if len(img.shape) == 5: img = np.transpose(img.squeeze(), (1, 2, 0))
mask = plt.imread('../data/patches/S 050001807_6_PAS_ind_00_X0_Y0_006179_026048_mask.png')
print(img.shape, mask.shape)
img = cv2.resize(img,
(img.shape[1] // resize, img.shape[0] // resize),
interpolation=cv2.INTER_AREA)
mask = cv2.resize(mask,
(mask.shape[1] // resize, mask.shape[0] // resize),
interpolation=cv2.INTER_NEAREST)
img = img[int(img.shape[0]*shft) : int(img.shape[0]*shft)+wnd,
int(img.shape[1]*shft) : int(img.shape[1]*shft)+wnd,
:]
mask = mask[int(mask.shape[0]*shft) : int(mask.shape[0]*shft)+wnd,
int(mask.shape[1]*shft) : int(mask.shape[1]*shft)+wnd]
plt.figure(figsize=(4, 4))
plt.axis('off')
plt.imshow(img)
plt.imshow(mask, alpha=.4)
plt.show()
fig, axes = plt.subplots(figsize=(16, 4), nrows=1, ncols=len(test_models))
for j in range(len(test_models)):
mask_lrg = test_models[j].predict(img[np.newaxis, ] / 255)
axes[j].imshow(np.squeeze(mask_lrg))
axes[j].set_title(f'img {j}: {np.min(mask_lrg):.2f}-{np.max(mask_lrg):.2f}')
axes[j].axis('off')
plt.show()
plt.figure(figsize=(14, 4))
plt.hist(mask_lrg.flatten(), bins=100)
plt.show()
plt.figure(figsize=(14, 4))
plt.hist(np.where(mask_lrg < 10e-4, np.nan, mask_lrg).flatten(), bins=100)
plt.show()
```
| github_jupyter |
# Randomized Benchmarking
## Introduction
One of the main challenges in building a quantum information processor is the non-scalability of completely
characterizing the noise affecting a quantum system via process tomography. In addition, process tomography is sensitive to noise in the pre- and post rotation gates plus the measurements (SPAM errors). Gateset tomography can take these errors into account, but the scaling is even worse. A complete characterization
of the noise is useful because it allows for the determination of good error-correction schemes, and thus
the possibility of reliable transmission of quantum information.
Since complete process tomography is infeasible for large systems, there is growing interest in scalable
methods for partially characterizing the noise affecting a quantum system. A scalable (in the number $n$ of qubits comprising the system) and robust algorithm for benchmarking the full set of Clifford gates by a single parameter using randomization techniques was presented in [1]. The concept of using randomization methods for benchmarking quantum gates is commonly called **Randomized Benchmarking
(RB)**.
## The Randomized Benchmarking Protocol
We should first import the relevant qiskit classes for the demonstration:
```
#Import general libraries (needed for functions)
import numpy as np
import matplotlib.pyplot as plt
from IPython import display
#Import the RB Functions
import qiskit.ignis.verification.randomized_benchmarking as rb
#Import Qiskit classes
import qiskit
from qiskit.providers.aer.noise import NoiseModel
from qiskit.providers.aer.noise.errors.standard_errors import depolarizing_error, thermal_relaxation_error
```
A RB protocol (see [1,2]) consists of the following steps:
### Step 1: Generate RB sequences
The RB sequences consist of random Clifford elements chosen uniformly from the Clifford group on $n$-qubits,
including a computed reversal element,
that should return the qubits to the initial state.
More precisely, for each length $m$, we choose $K_m$ RB sequences.
Each such sequence contains $m$ random elements $C_{i_j}$ chosen uniformly from the Clifford group on $n$-qubits, and the $m+1$ element is defined as follows: $C_{i_{m+1}} = (C_{i_1}\cdot ... \cdot C_{i_m})^{-1}$. It can be found efficiently by the Gottesmann-Knill theorem.
For example, we generate below several sequences of 2-qubit Clifford circuits.
```
#Generate RB circuits (2Q RB)
#number of qubits
nQ=2
rb_opts = {}
#Number of Cliffords in the sequence
rb_opts['length_vector'] = [1, 10, 20, 50, 75, 100, 125, 150, 175, 200]
#Number of seeds (random sequences)
rb_opts['nseeds'] = 5
#Default pattern
rb_opts['rb_pattern'] = [[0, 1]]
rb_circs, xdata = rb.randomized_benchmarking_seq(**rb_opts)
```
As an example, we print the circuit corresponding to the first RB sequence
```
rb_circs[0][0].draw()
```
One can verify that the Unitary representing each RB circuit should be the identity (with a global phase).
We simulate this using Aer unitary simulator.
```
# Create a new circuit without the measurement
qregs = rb_circs[0][-1].qregs
cregs = rb_circs[0][-1].cregs
qc = qiskit.QuantumCircuit(*qregs, *cregs)
for i in rb_circs[0][-1][0:-nQ]:
qc.data.append(i)
# The Unitary is an identity (with a global phase)
backend = qiskit.Aer.get_backend('unitary_simulator')
basis_gates = ['u1','u2','u3','cx'] # use U,CX for now
job = qiskit.execute(qc, backend=backend, basis_gates=basis_gates)
from qiskit_textbook.tools import array_to_latex
array_to_latex(np.around(job.result().get_unitary(),3), pretext="\\text{Unitary} = ")
```
### Step 2: Execute the RB sequences (with some noise)
We can execute the RB sequences either using Qiskit Aer Simulator (with some noise model) or using IBMQ provider, and obtain a list of results.
By assumption each operation $C_{i_j}$ is allowed to have some error, represented by $\Lambda_{i_j,j}$, and each sequence can be modeled by the operation:
$$\textit{S}_{i_m} = \bigcirc_{j=1}^{m+1} (\Lambda_{i_j,j} \circ C_{i_j})$$
where ${i_m} = (i_1,...,i_m)$ and $i_{m+1}$ is uniquely determined by ${i_m}$.
```
# Run on a noisy simulator
noise_model = NoiseModel()
# Depolarizing error on the gates u2, u3 and cx (assuming the u1 is virtual-Z gate and no error)
p1Q = 0.002
p2Q = 0.01
noise_model.add_all_qubit_quantum_error(depolarizing_error(p1Q, 1), 'u2')
noise_model.add_all_qubit_quantum_error(depolarizing_error(2 * p1Q, 1), 'u3')
noise_model.add_all_qubit_quantum_error(depolarizing_error(p2Q, 2), 'cx')
backend = qiskit.Aer.get_backend('qasm_simulator')
```
### Step 3: Get statistics about the survival probabilities
For each of the $K_m$ sequences the survival probability $Tr[E_\psi \textit{S}_{\textbf{i}_\textbf{m}}(\rho_\psi)]$
is measured.
Here $\rho_\psi$ is the initial state taking into account preparation errors and $E_\psi$ is the
POVM element that takes into account measurement errors.
In the ideal (noise-free) case $\rho_\psi = E_\psi = | \psi {\rangle} {\langle} \psi |$.
In practice one can measure the probability to go back to the exact initial state, i.e. all the qubits in the ground state $ {|} 00...0 {\rangle}$ or just the probability for one of the qubits to return back to the ground state. Measuring the qubits independently can be more convenient if a correlated measurement scheme is not possible. Both measurements will fit to the same decay parameter according to the properties of the *twirl*.
### Step 4: Find the averaged sequence fidelity
Average over the $K_m$ random realizations of the sequence to find the averaged sequence **fidelity**,
$$F_{seq}(m,|\psi{\rangle}) = Tr[E_\psi \textit{S}_{K_m}(\rho_\psi)]$$
where
$$\textit{S}_{K_m} = \frac{1}{K_m} \sum_{\textbf{i}_\textbf{m}} \textit{S}_{\textbf{i}_\textbf{m}}$$
is the average sequence operation.
### Step 5: Fit the results
Repeat Steps 1 through 4 for different values of $m$ and fit the results for the averaged sequence fidelity to the model:
$$ \textit{F}_{seq}^{(0)} \big(m,{|}\psi {\rangle} \big) = A_0 \alpha^m +B_0$$
where $A_0$ and $B_0$ absorb state preparation and measurement errors as well as an edge effect from the
error on the final gate.
$\alpha$ determines the average error-rate $r$, which is also called **Error per Clifford (EPC)**
according to the relation
$$ r = 1-\alpha-\frac{1-\alpha}{2^n} = \frac{2^n-1}{2^n}(1-\alpha)$$
(where $n=nQ$ is the number of qubits).
As an example, we calculate the average sequence fidelity for each of the RB sequences, fit the results to the exponential curve, and compute the parameters $\alpha$ and EPC.
```
# Create the RB fitter
backend = qiskit.Aer.get_backend('qasm_simulator')
basis_gates = ['u1','u2','u3','cx']
shots = 200
transpiled_circs_list = []
rb_fit = rb.RBFitter(None, xdata, rb_opts['rb_pattern'])
for rb_seed, rb_circ_seed in enumerate(rb_circs):
print('Compiling seed %d'%rb_seed)
new_rb_circ_seed = qiskit.compiler.transpile(rb_circ_seed, basis_gates=basis_gates)
transpiled_circs_list.append(new_rb_circ_seed)
print('Simulating seed %d'%rb_seed)
job = qiskit.execute(new_rb_circ_seed, backend, shots=shots,
noise_model=noise_model,
backend_options={'max_parallel_experiments': 0})
# Add data to the fitter
rb_fit.add_data(job.result())
print('After seed %d, alpha: %f, EPC: %f'%(rb_seed,rb_fit.fit[0]['params'][1], rb_fit.fit[0]['epc']))
```
### Extra Step: Plot the results
```
plt.figure(figsize=(8, 6))
ax = plt.subplot(1, 1, 1)
# Plot the essence by calling plot_rb_data
rb_fit.plot_rb_data(0, ax=ax, add_label=True, show_plt=False)
# Add title and label
ax.set_title('%d Qubit RB'%(nQ), fontsize=18)
plt.show()
```
## The Intuition Behind RB
The depolarizing quantum channel has a parameter $\alpha$, and works like this: with probability $\alpha$, the state remains the same as before; with probability $1-\alpha$, the state becomes the totally mixed state, namely:
$$\rho_f = \alpha \rho_i + \frac{1-\alpha}{2^n} * \mathbf{I}$$
Suppose that we have a sequence of $m$ gates, not necessarily Clifford gates,
where the error channel of the gates is a depolarizing channel with parameter $\alpha$
(same $\alpha$ for all the gates).
Then with probability $\alpha^m$ the state is correct at the end of the sequence,
and with probability $1-\alpha^m$ it becomes the totally mixed state, therefore:
$$\rho_f^m = \alpha^m \rho_i + \frac{1-\alpha^m}{2^n} * \mathbf{I}$$
Now suppose that in addition we start with the ground state;
that the entire sequence amounts to the identity;
and that we measure the state at the end of the sequence with the standard basis.
We derive that the probability of success at the end of the sequence is:
$$\alpha^m + \frac{1-\alpha^m}{2^n} = \frac{2^n-1}{2^n}\alpha^m + \frac{1}{2^n} = A_0\alpha^m + B_0$$
It follows that the probability of success, aka fidelity, decays exponentially with the sequence length, with exponent $\alpha$.
The last statement is not necessarily true when the channel is other than the depolarizing channel. However, it turns out that if the gates are uniformly-randomized Clifford gates, then the noise of each gate behaves on average as if it was the depolarizing channel, with some parameter that can be computed from the channel, and we obtain the exponential decay of the fidelity.
Formally, taking an average over a finite group $G$ (like the Clifford group) of a quantum channel $\bar \Lambda$ is also called a *twirl*:
$$ W_G(\bar \Lambda) \frac{1}{|G|} \sum_{u \in G} U^{\dagger} \circ \bar \Lambda \circ U$$
Twirling over the entire unitary group yields exactly the same result as the Clifford group. The Clifford group is a *2-design* of the unitary group.
## Simultaneous Randomized Benchmarking
RB is designed to address fidelities in multiqubit systems in two ways. For one, RB over the full $n$-qubit space
can be performed by constructing sequences from the $n$-qubit Clifford group. Additionally, the $n$-qubit space
can be subdivided into sets of qubits $ \{n_i\} $ and $n_i$-qubit RB performed in each subset simultaneously [4].
Both methods give metrics of fidelity in the $n$-qubit space.
For example, it is common to perform 2Q RB on the subset of two-qubits defining a CNOT gate while the other qubits are quiescent. As explained in [4], this RB data will not necessarily decay exponentially because the other qubit subspaces are not twirled. Subsets are more rigorously characterized by simultaneous RB, which also measures some level of crosstalk error since all qubits are active.
An example of simultaneous RB (1Q RB and 2Q RB) can be found in:
https://github.com/Qiskit/qiskit-tutorials/blob/master/tutorials/noise/4_randomized_benchmarking.ipynb
## Predicted Gate Fidelity
If we know the errors on the underlying gates (the gateset) we can predict the EPC without running RB experiment. This calculation verifies that your RB experiment followed by fitting yields correct EPC value. First we need to count the number of these gates per Clifford.
Then, the two qubit Clifford gate error function <code>calculate_2q_epc</code> gives the error per 2Q Clifford. It assumes that the error in the underlying gates is depolarizing. This function is derived in the supplement to [5].
```
# count the number of single and 2Q gates in the 2Q Cliffords
qubits = rb_opts['rb_pattern'][0]
gate_per_cliff = rb.rb_utils.gates_per_clifford(
transpiled_circuits_list=transpiled_circs_list,
clifford_lengths=xdata[0],
basis=basis_gates,
qubits=qubits)
for basis_gate in basis_gates:
print("Number of %s gates per Clifford: %f"%(
basis_gate,
np.mean([gate_per_cliff[qubit][basis_gate] for qubit in qubits])))
# convert from depolarizing error to epg (1Q)
epg_q0 = {'u1': 0, 'u2': p1Q/2, 'u3': 2 * p1Q/2}
epg_q1 = {'u1': 0, 'u2': p1Q/2, 'u3': 2 * p1Q/2}
# convert from depolarizing error to epg (2Q)
epg_q01 = 3/4 * p2Q
# calculate the predicted epc from underlying gate errors
pred_epc = rb.rb_utils.calculate_2q_epc(
gate_per_cliff=gate_per_cliff,
epg_2q=epg_q01,
qubit_pair=qubits,
list_epgs_1q=[epg_q0, epg_q1])
print("Predicted 2Q Error per Clifford: %e (qasm simulator result: %e)" % (pred_epc, rb_fit.fit[0]['epc']))
```
On the other hand, we can calculate the errors on the underlying gates (the gateset) from the experimentally obtained EPC. Given that we know the errors on the every single-qubit gates in the RB sequence, we can predict 2Q gate error from the EPC of two qubit RB experiment.
The two qubit gate error function ``calculate_2q_epg`` gives the estimate of error per 2Q gate. In this section we prepare single-qubit errors using the deporalizing error model. If the error model is unknown, EPGs of those gates, for example [``u1``, ``u2``, ``u3``], can be estimated with a separate 1Q RB experiment with the utility function ``calculate_1q_epg``.
```
# use 2Q EPC from qasm simulator result and 1Q EPGs from depolarizing error model
pred_epg = rb.rb_utils.calculate_2q_epg(
gate_per_cliff=gate_per_cliff,
epc_2q=rb_fit.fit[0]['epc'],
qubit_pair=qubits,
list_epgs_1q=[epg_q0, epg_q1])
print("Predicted 2Q Error per gate: %e (gate error model: %e)" % (pred_epg, epg_q01))
```
## References
1. Easwar Magesan, J. M. Gambetta, and Joseph Emerson, *Robust randomized benchmarking of quantum processes*,
https://arxiv.org/pdf/1009.3639
2. Easwar Magesan, Jay M. Gambetta, and Joseph Emerson, *Characterizing Quantum Gates via Randomized Benchmarking*,
https://arxiv.org/pdf/1109.6887
3. A. D. C'orcoles, Jay M. Gambetta, Jerry M. Chow, John A. Smolin, Matthew Ware, J. D. Strand, B. L. T. Plourde, and M. Steffen, *Process verification of two-qubit quantum gates by randomized benchmarking*, https://arxiv.org/pdf/1210.7011
4. Jay M. Gambetta, A. D. C´orcoles, S. T. Merkel, B. R. Johnson, John A. Smolin, Jerry M. Chow,
Colm A. Ryan, Chad Rigetti, S. Poletto, Thomas A. Ohki, Mark B. Ketchen, and M. Steffen,
*Characterization of addressability by simultaneous randomized benchmarking*, https://arxiv.org/pdf/1204.6308
5. David C. McKay, Sarah Sheldon, John A. Smolin, Jerry M. Chow, and Jay M. Gambetta, *Three Qubit Randomized Benchmarking*, https://arxiv.org/pdf/1712.06550
```
import qiskit
qiskit.__qiskit_version__
```
| github_jupyter |
# Biblioteca NumPy
Este tutorial é sobre a utilização da biblioteca NumPy, uma das bibliotecas fundamentais do Python para computação científica, utilizada para trabalhar com **matrizes**, cuja designação em inglês e ao longo deste tutorial é **n-dimensional array** ou simplesmente **ndarrays**.
Esta bilbioteca faz parte do package de computação científica [SciPy](https://www.scipy.org/).
A biblioteca NumPy foi pensada para lidar com computação vetorizada possuindo funcionalidades para manipulação de vetores/matrizes muito mais eficientes do que a utilização de ciclos de repetição como o **for**.
É a base de muitas outras bibliotecas como o Pandas, MatplotLib, Sickit-learn, entre outros.
O primeiro passo para trabalhar com uma biblioteca no Python é importá-la com a utilização do comando import. Para além de importá-la podemos dar-lhe um nome (alias) mais curto para escrever menos texto. No caso da numpy utiliza-se comumente a sigla np.
```
#Para importar bilbiotecas em python é necessário utilizar o comando import
import numpy as np
#Caso o numpy não esteja instalado deverá correr o comando "pip install numpy"
a = np.array([1, 2, 3]) # Cria uma matriz com três elementos
print(type(a)) # Imprime "<class 'numpy.ndarray'>"
print(a.shape) # Imprime o formato da matriz "(3,)"
print(a[0], a[1], a[2]) # Imprime "1 2 3"
a[0] = 5 # Altera o valor de um elemento da matriz
print(a) # Imprime "[5, 2, 3]"
b = np.array([[1,2,3],[4,5,6]]) # Cria uma matriz bidimensional
print(b.shape) # Imprime o formato da matriz "(2, 3)"
print(b[0, 0], b[0, 1], b[1, 0]) # Imprime "1 2 4"
c = np.array([[23,1.75,'antonio'],[23,1.75,'antonio'],[23,1.75,'miguel'],[23,1.75,'antonio'],[23,1.75,'antonio']])
print(c.shape) # Imprime o formato da matriz "(2, 3)"
print(c[2,2]) # Imprime o elemento da matriz na posicao (2,2) neste caso o valor 'miguel'
```
## Tipos de dados
No numpy todos os elementos de uma matriz/ndarray são de um determinado tipo de dados.
```
#Depois de importar a biblioteca uma primeira vez para o notebook não é preciso voltar a importar.
#Neste notebook importa-se em cada uma das células por uma questão de demonstração
import numpy as np
x = np.array([1, 2])
print(x.dtype)
x = np.array([1.0, 2.0])
print(x.dtype)
x = np.array([1, 2], dtype=np.float64)
print(x.dtype)
```
## Criação de matrizes/ndarray
A bilioteca NumPy fornece muitas formas para criar matrizes/ndarray.
```
import numpy as np
print("Vetor de 5 elementos seguidos: ")
print(np.arange(5))
print("Vetor de 5 elementos (igualmente espaçados), entre 1 e 2: ")
print(np.linspace(1,2,5))
print("Vetor de 5 elementos iguais a 0s:")
print(np.zeros(5))
print("Matriz 2x2 de zeros: ")
print(np.zeros((2,2)))
print("Matriz 2x3 de uns:")
print(np.ones((2,3)))
print("Matriz 4x2 de setes(7):")
print(np.full((4,2), 7))
print("Matriz identidade 2x2: ")
print(np.eye(2))
```
### Criação matrizes com valores aleatórios
O NumPy permite a criação de matrizes com valores aleatórios que podem seguir uma das muitas distribuições disponíveis no NumPy (beta, binomial, chisquare, dirichlet, exponential,
f, gamma, geometric, gumbel, hypergeometric, laplace, logistic, lognormal, logseries, multinomial, multivariate_normal, negative_binomial, noncentral_chisquare, noncentral_f,
normal, pareto, poisson, power, rayleigh, standard_cauchy, standard_exponential, standard_gamma(shape[, size]), standard_normal([size]), standard_t, triangular, uniform,
vonmises, wald, weibull, zipf)
Ver mais em [Random sampling](https://docs.scipy.org/doc/numpy-1.14.0/reference/routines.random.html).
De seguida apresentam-se alguns exemplos.
```
import numpy as np
#Geracao de vetores com valores aleatorios
#Utilizando uma distribuico uniforme
v1 = np.random.random(5000)
print("Vetor v1:")
print(v1)
print("Media: " + str(np.mean(v1)))
print("Desvio padrao: " + str(np.std(v1)))
#Utilizando uma distribuição normal de média 0 e desvio padrão 0
v2 = np.random.normal(0,1,5000)
print("Vetor v2:")
print(v2)
print("Media: " + str(np.mean(v2)))
print("Desvio padrao: " + str(np.std(v2)))
#Geração de uma matriz/ndarray (3x5) com valores aleatórios com base na distribuição uniforme
#contínua com valores entre [0,1)
m1 = np.random.random((3,5))
print("Matriz de valores aleatórios entre 0, 1 com base numa distribuição uniforme: ")
print(m1)
#Para verificar pode-se calcular a média e o desvio padrão dos valores
print("Média (a+b)/2, em que a = 0 e b = 1: " + str(np.mean(m1)))
print("Desvio padrão (Raiz Quadrada ((b-a)^2/12)): " + str(np.std(m1)))
print()
#Geração de uma matriz/ndarray (3x5) com valores aleatórios com base na distribuição normal
#com média 0 e desivo padrão 1
#O primeiro parametro é a média, o segundo o desvio padrão e o terceiro a matriz
m2 = np.random.normal(0,1,(3,5))
print("Matriz de valores aleatórios entre 0, 1 com base numa distribuição normal: ")
print(m2)
#Para verificar pode-se calcular a média e o desvio padrão dos valores
print("Média (0): " + str(np.mean(m2)))
print("Desvio padrão (1): " + str(np.std(m2)))
print("Nota: com poucos valores é provável que a média e o desvio padrão não sejam 0 e 1. Correr para uma matriz de maiores dimensões.")
print()
#Geração de uma matriz/ndarray (3x5) com valores aleatórios com base na distribuição exponencial
#O primeiro padrão é a escala (inverso da taxa, escala = 1/lambda) e o segundo a matriz
lambda1 = 5
g = np.random.exponential(1/lambda1,(3,5))
print("Matriz de valores aleatórios entre 0,1 com base numa distribuição exponencial: ")
print(g)
#Para verificar que está certo pode-se calcular a média e a variância dos valores
#print("Lambda: " + str(lambda1))
print("Média (1/lambda): " + str(np.mean(g))) #Deverá ser igual a 1/
print("Desvio Padrão (Raiz Quadrada (1/lambda^2)): " + str(np.std(g)))
print("Nota: caso a média e o desvio padrão não sejam próximos dos valores esperados, correr para uma matriz de maiores dimensões.")
#Por vezes é necessário colocar os valores de vetor/matriz entre 0 e 1
#Este processo designa-se de normalização
#Para tal é necessário subtrair cada um dos elementos pelo menor elemento e dividir pela amplitude dos elementos
# do vetor/matriz
#Exemplo:
v3 = np.arange(1,5)
print(v3)
#Normalizar
v3 = (v3-min(v3))/(max(v3)-min(v3))
print(v3)
```
## Slicing de matrizes/ndarray
Também permite o slicing, visto no notebook "Introdução ao Python"
```
import numpy as np
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
b = a[:2, 1:3]
print(a)
print(b)
import numpy as np
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
l1_d1 = a[1, :] # Retira a segunda linha para uma matriz unidimensional
l1_d2 = a[1:2, :] # Retira a segunda linha para uma matriz bidimensional
print(l1_d1, l1_d1.shape) # Imprime os valores da segunda linha e respetivo formato
print(l1_d2, l1_d2.shape) # Imprime os valores da segunda linha e respetivo formato
c1_d1 = a[:, 1]
c1_d2 = a[:, 1:2]
print(c1_d1, c1_d1.shape)
print(c1_d2, c1_d2.shape)
```
## Indexação de matrizes/ndarray
A biblioteca NumPy permite selecionar elementos de uma matriz/ndarray tendo por base outras matrizes e expressões booleanas.
```
#Indexação de matrizes utilizando outras matrizes
import numpy as np
a = np.array([[1,2], [3, 4], [5, 6]])
print("Matriz a de 3x2:")
print(a)
#Indexação utilizando uma matriz de posições a mostrar
print(a[[0, 1, 2], [0, 1, 0]]) #mostrar os elementos que estão na posição 0,0->1; 1,1->4; 2,0->5
#O exemplo acima pode ser conseguido através da indicação de cada um dos elementos
print(np.array([a[0, 0], a[1, 1], a[2, 0]]))
#Ao utilizar uma matriz de elementos inteiros para indexação podemos reutilizar elementos
print(a[[0, 0], [1, 1]])
#O exemplo acima com outra notação
print(np.array([a[0, 1], a[0, 1]]))
#Alterar os elementos de posições da matriz
import numpy as np
a = np.array([[1,2,3], [4,5,6], [7,8,9], [10, 11, 12]])
print(a)
b = np.array([0, 2, 0, 1])
# Selecionar um elemento de cada linha utilizando os indices em b
print(a[np.arange(4), b])
# Alterar um elemento de cada linha utilizando os indicies em b
a[np.arange(4), b] += 10
print(a)
print(np.arange(4))
#Indexação de matrizes utilizando expressões booleanas
#Utilizado para selecionar os lementos de uma matriz que
#satisfaçam uma determinada condição
import numpy as np
a = np.array([[1,2], [3, 4], [5, 6]])
bool_idx = (a > 2)
print(bool_idx)
print(a[bool_idx])
print(a[a > 2])
#Utilizando a função where
print(a[np.where(a>2)])
```
## Aritmética de matrizes/ndarray
De seguida aprentam-se algumas operações aritméticas para a manipulação dos elementos de uma matriz/ndarray.
```
#Operacoes ao nível dos elementos das matrizes
import numpy as np
a = np.array([[1,2],[3,4]], dtype=np.float64)
b = np.array([[5,6],[7,8]], dtype=np.float64)
print("Soma dos elementos de x com y")
print(a + b)
print(np.add(a, b))
print("Diferenca dos elementos de x com y")
print(a - b)
print(np.subtract(a, b))
print("Multiplicacao dos elementos de x com y")
print(a * b)
print(np.multiply(a, b))
print("Divisao dos elementos de x por y")
print(a / b)
print(np.divide(a, b))
print("Raiz quadrada dos elementos de x")
print(np.sqrt(a))
#Operacoes ao nível das matrizes
import numpy as np
x = np.array([[1,2],[3,4]])
y = np.array([[5,6],[7,8]])
v = np.array([9,10])
w = np.array([11, 12])
print("Produto da matrizes v e w")
print(v.dot(w))
print(np.dot(v, w))
print("Produto das matrizes x e v")
print(x.dot(v))
print(np.dot(x, v))
print("Produto das matrizes x e y")
print(x.dot(y))
print(np.dot(x, y))
```
## Soma de elementos da matriz/ndarray
```
import numpy as np
a = np.array([[1,2],[3,4]])
print('Matriz a:')
print(a)
print("Soma todos os elementos de x")
print(np.sum(a))
print("Soma todos os elementos das colunas de x")
print(np.sum(a, axis=0))
print("Soma todos os elementos das linhas de x")
print(np.sum(a, axis=1))
```
## Estatística sobre elementos das matrizes
A biblioteca NumPy disponibiliza o cálculo de algumas estatítisticas como a média, o desvio padrão, etc.
Permite que o cálculo das estatíticas poassa ser feito sobre os elementos todos da matriz/ndarray ou só sobre as linhas ou só sobre as colunas.
De seguida apresentam-se alguns exemplos.
Mais informação disponível em (https://docs.scipy.org/doc/numpy/reference/routines.statistics.html)
```
import numpy as np
a = np.array([[4, 3, 5], [1, 2, 1]])
print("Matriz a:")
print(a)
print("Média de todos os valores: ")
print(np.mean(a))
print("Média dos valores nas linhas: ")
print(np.mean(a,axis=1))
print("Média dos valores nas colunas: ")
print(np.mean(a,axis=0))
print("Desvio padrão de todos os valores: ")
print(np.std(a))
print("Desvio padrão dos valores nas linhas: ")
print(np.std(a,axis=1))
print("Desvio padrão dos valores nas colunas: ")
print(np.std(a,axis=0))
print("Valor minimo: ")
print(np.amin(a))
print("Valor minimo dos valores nas linhas: ")
print(np.amin(a,axis=1))
print("Valor minimo dos valores nas colunas: ")
print(np.amin(a,axis=0))
print("Vetor b:")
b = [1,2,3,1,1,3]
print(b)
print("Valores únicos:")
print(np.unique(b))
print("Contagem dos valores únicos:")
print(np.unique(b, return_counts=True))
#print(np.bincount(b))
print("Histograma a partir dos valores de um vetor:")
c = np.array([1,1.5,2.0,2.0,3.0,3.5,3.5])
print("Vetor c:")
print(c)
print("Histograma: contagem/intervalos da classe:")
print(np.histogram(b))
print("Histograma: gráfico:")
#Mais à frente ver-se-á como trabalhar com as bibliotecas gráficas mas fica aqui um exemplo
import matplotlib.pyplot as plt
#Caso esta bilbioteca não esteja instalada correr "pip install matplotlib"
plt.hist(b)
plt.show()
#Exemplo 1. Criar um histograma das alturas dos alunos de um turma
alturas = np.random.randint(160,190,60) #Geracao de uma turma de 60 alunos com alturas entre 160..190
print(alturas)
print(np.mean(alturas))
print(np.bincount(alturas)[160:]) #Contar o numero de observacoes em cada uma das alturas
print(np.unique(alturas)) #Para validar o histograma
#Histograma
import matplotlib.pyplot as plt
plt.hist(alturas,bins=np.arange(160,191))
plt.show()
```
## Ordenação de elementos da matriz
```
import numpy as np
a = np.array([[4, 3, 5], [1, 2, 1]])
print("Matriz a:")
print(a)
print("Ordenar os elementos das linhas:")
b = np.sort(a, axis=1)
print(b)
```
## Broadcasting
Mecanismo que permite ao NumPy trabalhar com matrizes/ndarray de diferentes dimensões.
No exemplo seguinte mostra-se como somar os valores de um vetor a uma matriz/ndarray, da forma tradicional e utilizando o mecanismo de broadcasting.
```
import numpy as np
# Adicionar o vetor a cada linha da matriz x guardando na matriz y
a = np.array([[1,2,3], [4,5,6], [7,8,9], [10, 11, 12]])
v = np.array([1, 0, 1])
print("Metodo 1:")
b = np.empty_like(a) #Cria um vetor b igual ao vetor a com valores todos a zero
# Adcionar à matriz b os valores de a e v
for i in range(4):
b[i, :] = a[i, :] + v
print(b)
print("Metodo 2:")
b = a + v
print(b)
```
## Alterar o formato das matrizes/ndarray
A bilbioteca NumPy permite alterar o formato das matrizes/ndarray.
Para isso possui os seguintes métodos:
* transpose, que apresenta a matriz/ndarray transposta
* ravel, que transforma uma matriz/ndarray numa lista de elementos
* reshape, que dá a forma da matriz/ndarray desejada a uma lista de elementos
* newaxis, que permite adicionar uma nova dimensão à matriz/ndarray
* resize, que permite adicionar elementos à matriz/ndarray
Exemplos:
```
import numpy as np
a = np.array([[1, 2, 3], [4, 5, 6]])
print('Matriz a:')
print(a)
print(a.shape)
print('Matriz a transposta:')
print(np.transpose(a))
print('Elementos da matriz transformados numa lista:')
print(a.ravel())
print(np.ravel(a))
print('Matriz b:')
b = a.reshape((3, 2)) #Da outra estrutura ao vetor a, tem de ter os mesmos elementos e segue a ordem natural
print(b)
print(b.shape)
print(b.ravel())
print('Redimensionar uma matriz:')
c = np.array([[1, 2, 3], [4, 5, 6]])
print(c)
c.resize((5,2))
print(c)
#Não se pode redimensionar uma matriz que já tenha sido referenciada anteriormente
print('Adicionar uma dimensão a uma matriz:')
d = np.arange(4)
print(d)
e = d[:,np.newaxis]
print(e)
```
## Utilização de funções lambda e list comprehension em matrizes/ndarray
As funções lambda, apresentadas no notebook [pytrigo-1-intro.ipynb](pytrigo-1-intro.ipynb), permitem aplicar a todos os elementos de uma matriz/ndarray um determinada função, ao contrário do processo normal que seria o de passar uma matriz/ndarray por parametro para a execução de uma determinada função.
De seguida apresentam-se alguns exemplos de utilização de funções lambdas em matrizes/arrays.
```
import numpy as np
a = np.array([1, 2, 3])
b = np.array([[1, 2, 3], [4, 5, 6]])
print("Array a:")
print(a)
#Mostrar todos os valores superiores a 3
print("Array a (>2): ")
print(a[a>2])
print([x for x in a if x > 2])
print(np.array(list(filter(lambda x : x > 2, a))))
print("Array b:")
print(b)
#Mostrar todos os valores superiores a 3
print("Array b (>2): ")
print(b[b>2])
print([[j for j in i if j>2] for i in b])
#print(np.array(list(filter(lambda x : x > 2, b))))
#a instrução acima dá erro, é preciso converter primeiro o array 2d para 1d
print(np.array(list(filter(lambda x : x > 2, b.reshape(6,1)))))
#Mostrar o quadrado de todos os valores
print("Array b^2: ")
print(b**2)
print(np.array([x**2 for x in b]))
print(np.array(list(map(lambda x : x**2, b))))
```
## Leitura e escrita de ficheiros
A leitura e escrita de ficheiros na biblioteca NumPy faz-se recorrendo aos métodos:
* loadtxt(fname[, dtype, comments, delimiter, ...])
* savetxt(fname, X[, fmt, delimiter, newline, ...])
* genfromtxt(fname[, dtype, comments, ...])
* fromregex(file, regexp, dtype)
* fromstring(string[, dtype, count, sep])
* ndarray.tofile(fid[, sep, format])
* ndarray.tolist()
De seguida apresentam-se dois exemplos:
* o primeiro le um ficheiro csv para um array, divide os elementos por 2 e grava num novo ficheiro.
* o segundo le um ficheiro csv para um array, mas com a especificação do tipo de dados (inteiros)
```
import numpy as np
#Exemplo 1
print('Exemplo 1:')
a = np.genfromtxt('PyTrigo-V2-3-NumPy-csv1.csv', delimiter=',')
print('Matriz a:')
print(a)
b = a*2
print('Matriz 2*a:')
print(b)
np.savetxt('PyTrigo-V2-3-NumPy-csv2.csv', b, delimiter=',')
print('Exemplo 2:')
a = np.genfromtxt('PyTrigo-V2-3-NumPy-csv1.csv', delimiter=',',dtype=int)
print('Matriz a (inteiros):')
print(a)
# Existem mais opcoes para a leitura / escrita de dados
# A funcao genfromtxt tambem permite indicar o tratamento a dar aos missing values
```
### Exercícios sobre NumPy
1. Crie um vetor com 10 elementos todos a 0
2. Crie uma matriz de zeros com a dimensão de 2X5
3. Crie um vetor de 10 elementos com valores entre 10 e 49
4. Crie uma matriz 2X5 com valores entre 10 e 19
5. Crie uma matriz identidade com a dimensão 5x5
6. Gere uma matriz 5x5 com valores aleatorios
7. Normalize os valores de uma matriz 5x5
8. Crie um vetor com 10 elementos aleatórios e ordene-o.
9. Descubra a posicao num vetor do elemento com o valor mais próximo de um valor gerado aleatoriamente.
## Exercicios sobre os conceitos anteriores
```
#Exercicio 1. Crie um vetor de 10 elementos a 0
v0001 = np.zeros(10)
print(v0001)
v0002 = np.array([0 for x in range(10)])
print(v0002)
#Exercicio 2. Crie uma matriz de zeros com a dimensão de 2X5
m0001 = np.zeros(shape=(2,5))
print(m0001)
#Exercicio 3. Crie um vetor de 10 elementos com valores entre 10 e 49 (espaçados igualmente)
v0002 = np.linspace(10,49,10,dtype='int32')
print(v0002)
#Exercicio 4. Crie uma matriz 2X5 com valores entre 10 e 19
m0002 = np.arange(10,20).reshape(2,5)
print(m0002)
#Exercicio 5. Crie uma matriz identidade com com 5x5
m0003 = np.eye(5)
print(m0003)
#Exercicio 6. Gere uma matriz 5x5 com valores aleatorios
m0004 = np.random.random((5,5))
print(m0004)
#Exercicio 7. Normalize os valores de uma matriz 5x5
m0005 = np.random.random((5,5))
print(m0005)
mmax, mmin = m0005.max(), m0005.min()
m = (m0005 - mmin)/(mmax - mmin)
print(m0005)
#Exercicio 8. Crie um vetor com 100 elementos aleatorios e ordene-os
v0003 = np.random.random(100)
print(np.sort(v0003))
print(np.sort(v0003)[::-1])
#Exemplo 9. Apresente a transposta de uma matriz
m0006 = np.random.randint(20,60,(3,7))
print(m0006)
print(np.transpose(m0006))
#Exemplo 10. Descubra a posicao num vetor do elemento com o valor mais proximo de um valor gerado aleatoriamente
#Solucao1
v0004 = np.random.randint(10,100,15)
print(v0004)
a = np.random.randint(10,100)
print(a)
print(np.argmin(np.abs(v0004-a)))
#print(np.abs(v0006-a).argmin())
#Solucao 2
v0005 = np.arange(100)
valor = np.random.uniform(0,100)
indice = (np.abs(v0005-valor)).argmin()
print(v0005)
print(valor)
print('Posicao: ', indice,' Valor: ', v0005[indice])
```
Mais exercícios disponíveis em: http://www.labri.fr/perso/nrougier/teaching/numpy.100/
## Resolver um sistema de duas equações lineares recorrendo a matrizes
```
#Resolver um sistema de duas equações lineares recorrendo a matrizes
#Exemplo:
#4x + 3y = 20
#-5x + 9y = 26
# Matriz A com os coeficentes do sistema (variaveis dependentes)
# Matriz X com as incógnitas do sistema
# Matriz B com as soluções (termos independentes)
# Para resolver o sistema temos de calcular a inversa da matriz A
# (se esta for quadrada e não singular (determinante != 0)) e multiplicar por B
A = np.array([[4,3],[-5,9]])
B = np.array([20,26])
#testar o determiante caso não consigam faze-lo visulamente det(m[[a,b][c,d]]) = ad - bc
print(np.linalg.det(A))
inv_A = np.linalg.inv(A)
X = inv_A.dot(B)
print("Metodo1 - solucao: " + str(X))
print("Metodo2 (utilizando o solve) - solucao: " + str(np.linalg.solve(A,B)))
```
### Velocidade de execução: Python vs NumPy
No dominio da Data Science a velcocidade de execução do código tem um impacto muito grande quando se analisa grandes quantidades de dados, pelo que deverá sempre procurar a forma mais eficiente de o fazer, preferindo a utilização de bibliotecas como o NumPy.
Para medir a execução do código que implementa pode utilizar a biblioteca **time** do Python e registar o tempo antes e depois da execução de um pedaço de código. Outra forma de o fazer é utilizar a biblioteca **timeit**, que corre o bloco de código x vezes, que devolve a média e desvio padrão do tempo de execução desse código.
De seguida apresenta-se o tempo de execução de um código para fazer o cálculo da distância entre todas as linhas da matriz, retirado de
http://nbviewer.jupyter.org/github/ogrisel/notebooks/blob/master/Numba%20Parakeet%20Cython.ipynb.
No seguinte link - http://arogozhnikov.github.io/2015/01/06/benchmarks-of-speed-numpy-vs-all.html - poderá obter mais informação sobre outras bibliotecas, criadas com o bjetivo de serem mais eficientes que a biblioteca NumPy, como a Numba ou a Cython.
```
import numpy as np
from timeit import timeit
X = np.random.random((1000 ,3))
#X_wide = np.random.random((1000, 100))
#Python
def pairwise_python(X):
M = X.shape[0]
N = X.shape[1]
D = np.empty((M, M), dtype=np.float)
for i in range(M):
for j in range(M):
d = 0.0
for k in range(N):
tmp = X[i, k] - X[j, k]
d += tmp * tmp
D[i, j] = np.sqrt(d)
return D
%timeit pairwise_python(X)
#NumPy
def pairwise_numpy(X):
return np.sqrt(((X[:, None, :] - X) ** 2).sum(-1))
%timeit pairwise_numpy(X)
print('A segunda implementação recorrendo à biblioteca NumPy é 500x mais rápida!')
```
| github_jupyter |
```
from fastText.FastText import train_supervised, fasttext, load_model
import numpy as np
import pandas as pd
import re
import subprocess
# from sklearn.feature_extraction.text import TfidfVectorizer
# from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score, StratifiedKFold, KFold
from scipy.sparse import hstack
from sklearn.metrics import roc_auc_score, classification_report
from datetime import datetime
class_names = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
train = pd.read_csv('data/train_preprocessed.csv').fillna(' ')
test = pd.read_csv('data/test_preprocessed.csv').fillna(' ')
tr_ids = train[['id']]
train[class_names] = train[class_names].astype(np.int8)
target = train[class_names]
def get_probs(rez):
probs = []
for r, prob in zip(rez[0], rez[1]):
if r[0][-1] == '1':
probs.append(prob[0])
else:
probs.append(1 - prob[0])
return probs
def training(train_data, train_indices, val_indices, target, test_data):
df_train = train_data.loc[train_indices]
df_val = train_data.loc[val_indices]
df_train[target + '_ft'] = df_train[target].apply(lambda x: '__label__1 ' if x == 1 else '__label__0 ')
df_train[[target + '_ft', 'comment_text']].to_csv('train_fastText.csv', index=False, header=False)
d = subprocess.Popen("/home/ladmin/fastText-0.1.0/fasttext supervised -input /home/ladmin/toxic_comments/train_fastText.csv -output /home/ladmin/toxic_comments/fasttext_model -pretrainedVectors /home/ladmin/toxic_comments/embeddings/crawl-300d-2M.vec -loss hs -minCount 5 -dim 300".split())
d.communicate()
classifier = load_model('fasttext_model.bin')
val_proba = np.array(get_probs(classifier.predict(list(df_val['comment_text']))))
sub_proba = np.array(get_probs(classifier.predict(list(test_data['comment_text']))))
# train_score = roc_auc_score(df_train[target], train_proba)
val_score = roc_auc_score(df_val[target], val_proba)
return val_score, val_proba, sub_proba, val_indices
submission = pd.DataFrame.from_dict({'id': test['id']})
train_submission = pd.DataFrame.from_dict({'id': train['id']})
predictors = 5
scores = []
for i, class_name in enumerate(class_names):
print('Class: %s' % class_name)
sub_probas = np.zeros(shape=(len(test), ))
train_probas = np.zeros(shape=(len(train), ))
kf = KFold(n_splits=predictors, shuffle=True, random_state=42)
train_scores, val_scores = [], []
for train_indices, val_indices in kf.split(train):
val_score, val_proba, sub_proba, val_indices = training(train, train_indices, val_indices, class_name, test)
val_scores.append(val_score)
train_probas[val_indices] += val_proba
sub_probas += sub_proba / predictors
scores.append(np.mean(val_scores))
print('\tVal ROC-AUC: %s' % np.mean(val_scores))
submission[class_name] = sub_probas
train_submission[class_name] = train_probas
print('Total: %s' % np.mean(scores))
submission.head()
submission.to_csv('data/submission_fasttext.csv', index=False)
train_submission.to_csv('data/train_fasttext.csv', index=False)
```
| github_jupyter |
# Character-level Language Modeling with LSTMs
This notebook is adapted from [Keras' lstm_text_generation.py](https://github.com/fchollet/keras/blob/master/examples/lstm_text_generation.py).
Steps:
- Download a small text corpus and preprocess it.
- Extract a character vocabulary and use it to vectorize the text.
- Train an LSTM-based character level language model.
- Use the trained model to sample random text with varying entropy levels.
- Implement a beam-search deterministic decoder.
**Note**: fitting language models is very computation intensive. It is **recommended to do this notebook on a server with a GPU or powerful CPUs** that you can leave running for several hours at once.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
```
## Loading some text data
Let's use some publicly available philosopy:
```
from keras.utils.data_utils import get_file
URL = "https://s3.amazonaws.com/text-datasets/nietzsche.txt"
corpus_path = get_file('nietzsche.txt', origin=URL)
text = open(corpus_path).read().lower()
print('Corpus length: %d characters' % len(text))
print(text[:600], "...")
text = text.replace("\n", " ")
split = int(0.9 * len(text))
train_text = text[:split]
test_text = text[split:]
```
## Building a vocabulary of all possible symbols
To simplify things, we build a vocabulary by extracting the list all possible characters from the full datasets (train and validation).
In a more realistic setting we would need to take into account that the test data can hold symbols never seen in the training set. This issue is limited when we work at the character level though.
Let's build the list of all possible characters and sort it to assign a unique integer to each possible symbol in the corpus:
```
chars = sorted(list(set(text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
```
`char_indices` is a mapping to from characters to integer identifiers:
```
len(char_indices)
sorted(char_indices.items())[:15]
```
`indices_char` holds the reverse mapping:
```
len(indices_char)
indices_char[52]
```
While not strictly required to build a language model, it's a good idea to have a look at the distribution of relative frequencies of each symbol in the corpus:
```
from collections import Counter
counter = Counter(text)
chars, counts = zip(*counter.most_common())
indices = np.arange(len(counts))
plt.figure(figsize=(14, 3))
plt.bar(indices, counts, 0.8)
plt.xticks(indices, chars);
```
Let's cut the dataset into fake sentences at random with some overlap. Instead of cutting at random we could use a English specific sentence tokenizer. This is explained at the end of this notebook. In the mean time random substring will be good enough to train a first language model.
```
max_length = 40
step = 3
def make_sequences(text, max_length=max_length, step=step):
sequences = []
next_chars = []
for i in range(0, len(text) - max_length, step):
sequences.append(text[i: i + max_length])
next_chars.append(text[i + max_length])
return sequences, next_chars
sequences, next_chars = make_sequences(train_text)
sequences_test, next_chars_test = make_sequences(test_text, step=10)
print('nb train sequences:', len(sequences))
print('nb test sequences:', len(sequences_test))
```
Let's shuffle the sequences to break some of the dependencies:
```
from sklearn.utils import shuffle
sequences, next_chars = shuffle(sequences, next_chars,
random_state=42)
sequences[0]
next_chars[0]
```
## Converting the training data to one-hot vectors
Unfortunately the LSTM implementation in Keras does not (yet?) accept integer indices to slice columns from an input embedding by it-self. Let's use one-hot encoding. This is slightly less space and time efficient than integer coding but should be good enough when using a small character level vocabulary.
**Exercise:**
One hot encoded the training `data sequences` as `X` and `next_chars` as `y`:
```
n_sequences = len(sequences)
n_sequences_test = len(sequences_test)
voc_size = len(chars)
X = np.zeros((n_sequences, max_length, voc_size),
dtype=np.float32)
y = np.zeros((n_sequences, voc_size), dtype=np.float32)
X_test = np.zeros((n_sequences_test, max_length, voc_size),
dtype=np.float32)
y_test = np.zeros((n_sequences_test, voc_size), dtype=np.float32)
# TODO
# %load solutions/language_model_one_hot_data.py
X.shape
y.shape
X[0]
y[0]
```
## Measuring per-character perplexity
The NLP community measures the quality of probabilistic model using [perplexity](https://en.wikipedia.org/wiki/Perplexity).
In practice perplexity is just a base 2 exponentiation of the average negative log2 likelihoods:
$$perplexity_\theta = 2^{-\frac{1}{n} \sum_{i=1}^{n} log_2 (p_\theta(x_i))}$$
**Note**: here we define the **per-character perplexity** (because our model naturally makes per-character predictions). **It is more common to report per-word perplexity**. Note that this is not as easy to compute the per-world perplexity as we would need to tokenize the strings into a sequence of words and discard whitespace and punctuation character predictions. In practice the whitespace character is the most frequent character by far making our naive per-character perplexity lower than it should be if we ignored those.
**Exercise**: implement a Python function that computes the per-character perplexity with model predicted probabilities `y_pred` and `y_true` for the encoded ground truth:
```
def perplexity(y_true, y_pred):
"""Compute the per-character perplexity of model predictions.
y_true is one-hot encoded ground truth.
y_pred is predicted likelihoods for each class.
2 ** -mean(log2(p))
"""
# TODO
return 1.
# %load solutions/language_model_perplexity.py
y_true = np.array([
[0, 1, 0],
[0, 0, 1],
[0, 0, 1],
])
y_pred = np.array([
[0.1, 0.9, 0.0],
[0.1, 0.1, 0.8],
[0.1, 0.2, 0.7],
])
perplexity(y_true, y_pred)
```
A perfect model has a minimal perplexity of 1.0 bit (negative log likelihood of 0.0):
```
perplexity(y_true, y_true)
```
## Building recurrent model
Let's build a first model and train it on a very small subset of the data to check that it works as expected:
```
from keras.models import Sequential
from keras.layers import LSTM, Dense
from keras.optimizers import RMSprop
model = Sequential()
model.add(LSTM(128, input_shape=(max_length, voc_size)))
model.add(Dense(voc_size, activation='softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(optimizer=optimizer, loss='categorical_crossentropy')
```
Let's measure the perplexity of the randomly initialized model:
```
def model_perplexity(model, X, y, verbose=1):
predictions = model.predict(X, verbose=verbose)
return perplexity(y, predictions)
model_perplexity(model, X_test, y_test)
```
Let's train the model for one epoch on a very small subset of the training set to check that it's well defined:
```
small_train = slice(0, None, 40)
model.fit(X[small_train], y[small_train], validation_split=0.1,
batch_size=128, epochs=1)
model_perplexity(model, X[small_train], y[small_train])
model_perplexity(model, X_test, y_test)
```
## Sampling random text from the model
Recursively generate one character at a time by sampling from the distribution parameterized by the model:
$$
p_{\theta}(c_n | c_{n-1}, c_{n-2}, \ldots, c_0) \cdot p_{\theta}(c_{n-1} | c_{n-2}, \ldots, c_0) \cdot \ldots \cdot p_{\theta}(c_{0})
$$
This way of parametrizing the joint probability of a set of random-variables that are structured sequentially is called **auto-regressive modeling**.
```
def sample_one(preds, temperature=1.0):
"""Sample the next character according to the network output.
Use a lower temperature to force the model to output more
confident predictions: more peaky distribution.
"""
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
# Draw a single sample (size=1) from a multinoulli distribution
# parameterized by the output of the softmax layer of our
# network. A multinoulli distribution is a multinomial
# distribution with a single trial with n_classes outcomes.
probs = np.random.multinomial(1, preds, size=1)
return np.argmax(probs)
def generate_text(model, seed_string, length=300, temperature=1.0):
"""Recursively sample a sequence of chars, one char at a time.
Each prediction is concatenated to the past string of predicted
chars so as to condition the next prediction.
Feed seed string as a sequence of characters to condition the
first predictions recursively. If seed_string is lower than
max_length, pad the input with zeros at the beginning of the
conditioning string.
"""
generated = seed_string
prefix = seed_string
for i in range(length):
# Vectorize prefix string to feed as input to the model:
x = np.zeros((1, max_length, voc_size))
shift = max_length - len(prefix)
for t, char in enumerate(prefix):
x[0, t + shift, char_indices[char]] = 1.
preds = model.predict(x, verbose=0)[0]
next_index = sample_one(preds, temperature)
next_char = indices_char[next_index]
generated += next_char
prefix = prefix[1:] + next_char
return generated
```
The temperature parameter makes it possible to increase or decrease the entropy into the multinouli distribution parametrized by the output of the model.
Temperature lower than 1 will yield very regular text (biased towards the most frequent patterns of the training set). Temperatures higher than 1 will render the model "more creative" but also noisier (with a large fraction of meaningless words). A temperature of 1 is neutral (the noise of the generated text only stems from the imperfection of the model).
```
generate_text(model, 'philosophers are ', temperature=0.1)
generate_text(model, 'atheism is the root of ', temperature=0.8)
```
## Training the model
Let's train the model and monitor the perplexity after each epoch and sample some text to qualitatively evaluate the model:
```
nb_epoch = 30
seed_strings = [
'philosophers are ',
'atheism is the root of ',
]
for epoch in range(nb_epoch):
print("# Epoch %d/%d" % (epoch + 1, nb_epoch))
print("Training on one epoch takes ~90s on a K80 GPU")
model.fit(X, y, validation_split=0.1, batch_size=128, epochs=1,
verbose=2)
print("Computing perplexity on the test set:")
test_perplexity = model_perplexity(model, X_test, y_test)
print("Perplexity: %0.3f\n" % test_perplexity)
for temperature in [0.1, 0.5, 1]:
print("Sampling text from model at %0.2f:\n" % temperature)
for seed_string in seed_strings:
print(generate_text(model, seed_string, temperature=temperature))
print()
```
## Beam search for deterministic decoding
**Optional exercise**: adapt the sampling decoder to implement a deterministic decoder with a beam of k=5 sequences that are the most likely sequences based on the model predictions.
## Better handling of sentence boundaries
To simplify things we used the lower case version of the text and we ignored any sentence boundaries. This prevents our model to learn when to stop generating characters. If we want to train a model that can start generating text at the beginning of a sentence and stop at the end of a sentence, we need to provide it with sentence boundary markers in the training set and use those special markers when sampling.
The following give an example of how to use NLTK to detect sentence boundaries in English text.
This could be used to insert an explicit "end_of_sentence" (EOS) symbol to mark separation between two consecutive sentences. This should make it possible to train a language model that explicitly generates complete sentences from start to end.
Use the following command (in a terminal) to install nltk before importing it in the notebook:
```
$ pip install nltk
```
```
with open(corpus_path, 'rb') as f:
text_with_case = f.read().decode('utf-8').replace("\n", " ")
import nltk
nltk.download('punkt')
from nltk.tokenize import sent_tokenize
sentences = sent_tokenize(text_with_case)
plt.hist([len(s.split()) for s in sentences], bins=30);
plt.title('Distribution of sentence lengths')
plt.xlabel('Approximate number of words');
```
The first few sentences detected by NLTK are too short to be considered real sentences. Let's have a look at short sentences with at least 20 characters:
```
sorted_sentences = sorted([s for s in sentences if len(s) > 20], key=len)
for s in sorted_sentences[:5]:
print(s)
```
Some long sentences:
```
for s in sorted_sentences[-3:]:
print(s)
```
The NLTK sentence tokenizer seems to do a reasonable job despite the weird casing and '--' signs scattered around the text.
Note that here we use the original case information because it can help the NLTK sentence boundary detection model make better split decisions. Our text corpus is probably too small to train a good sentence aware language model though, especially with full case information. Using larger corpora such as a large collection of [public domain books](http://www.gutenberg.org/) or Wikipedia dumps. The NLTK toolkit also comes from [corpus loading utilities](http://www.nltk.org/book/ch02.html).
The following loads a selection of famous books from the Gutenberg project archive:
```
import nltk
nltk.download('gutenberg')
book_selection_text = nltk.corpus.gutenberg.raw().replace("\n", " ")
print(book_selection_text[:300])
print("Book corpus length: %d characters" % len(book_selection_text))
```
Let's do an arbitrary split. Note the training set will have a majority of text that is not authored by the author(s) of the validation set:
```
split = int(0.9 * len(book_selection_text))
book_selection_train = book_selection_text[:split]
book_selection_validation = book_selection_text[split:]
```
## Bonus exercises
- Adapt the previous language model to handle explicitly sentence boundaries with a special EOS character.
- Train a new model on the random sentences sampled from the book selection corpus with full case information.
- Adapt the random sampling code to start sampling at the beginning of sentence and stop when the sentence ends.
- Train a deep GRU (e.g. two GRU layers instead of a single LSTM) to see if you can improve the validation perplexity.
- Git clone the source code of the [Linux kernel](https://github.com/torvalds/linux) and train a C programming language model on it. Instead of sentence boundary markers, we could use source file boundary markers for this exercise. Compare your resutls with Andrej Karpathy's https://karpathy.github.io/2015/05/21/rnn-effectiveness/.
- Try to increase the vocabulary size to 256 using a [Byte Pair Encoding](https://arxiv.org/abs/1508.07909) strategy.
## Why build a language model?
Building a language model is not very useful by it-self. However language models have recently been shown to be useful for **transfer learning** to build **contextualized word embeddings** as a better alternative to word2vec or GloVe.
Using language-model based word representations makes it possible to reach the **state-of-the-art at many natural language understanding problems**.
The workflow is the following:
- **train** a (bi-directional) **deep language model on a very large, unlabeled corpus** (e.g. 1 billion words or more);
- plug the resulting **language model as the input layer** (and sometimes also the output layer) **of a task specific architecture**, for instance: text classification, semantic role labeling for knowledge extraction, logical entailment, question answering and reading comprehension;
- **train the task specific parameters** of the new architecture on the **smaller task-labeled corpus**;
- optionally fine-tune the full architecture on the task-labeled corpus if it's big enough not to overfit.
More information on this approach:
- Deep contextualized word representations, https://arxiv.org/abs/1802.05365
- [Pytorch implementation of ELMo](https://github.com/allenai/allennlp/blob/master/allennlp/modules/elmo.py) as part of the AllenNLP project: https://github.com/allenai/allennlp
- Fine-tuned Language Models for Text Classification https://arxiv.org/abs/1801.06146 (FitLaM)
| github_jupyter |
# Sleep stage classification: SVC
____
This model aims to classify sleep stages based on two EEG channel. We will use the features extracted in the `pipeline.ipynb` notebook as the input to a support vector classification (SVC).
We will only be looking at `LinearSVC`, because the `SVC` model implemented by scikitlearn has a quadratic time complexity. As it is stated in the docs, it may be impractical beyond tens of thousands of samples, which corresponds to our sample numbers.
```
%load_ext autoreload
%autoreload 2
import os
import sys
# Ensure parent folder is in PYTHONPATH
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
%matplotlib inline
from itertools import groupby
import matplotlib.pyplot as plt
import numpy as np
import joblib
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler
from sklearn.svm import LinearSVC
from sklearn.model_selection import (GridSearchCV,
RandomizedSearchCV,
GroupKFold,
cross_validate)
from sklearn.metrics import (accuracy_score,
confusion_matrix,
classification_report,
f1_score,
cohen_kappa_score,
make_scorer)
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.decomposition import PCA
from sklearn.ensemble import AdaBoostClassifier
from constants import (SLEEP_STAGES_VALUES,
N_STAGES,
EPOCH_DURATION)
from model_utils import (print_hypnogram,
train_test_split_one_subject,
train_test_split_according_to_age,
evaluate_hyperparams_grid,
print_results_cv,
print_results_cv_scores,
get_pipeline)
```
## Load the features
___
```
# position of the subject information and night information in the X matrix
SUBJECT_IDX = 0
NIGHT_IDX = 1
USE_CONTINUOUS_AGE = False
DOWNSIZE_SET = False
TEST_SET_SUBJECTS = [0.0, 24.0, 49.0, 71.0]
if USE_CONTINUOUS_AGE:
X_file_name = "../data/x_features-age-continuous.npy"
y_file_name = "../data/y_observations-age-continuous.npy"
else:
X_file_name = "../data/x_features.npy"
y_file_name = "../data/y_observations.npy"
X_init = np.load(X_file_name, allow_pickle=True)
y_init = np.load(y_file_name, allow_pickle=True)
X_init = np.vstack(X_init)
y_init = np.hstack(y_init)
print(X_init.shape)
print(y_init.shape)
print("Number of subjects: ", np.unique(X_init[:,SUBJECT_IDX]).shape[0]) # Some subject indexes are skipped, thus total number is below 83 (as we can see in https://physionet.org/content/sleep-edfx/1.0.0/)
print("Number of nights: ", len(np.unique([f"{int(x[0])}-{int(x[1])}" for x in X_init[:,SUBJECT_IDX:NIGHT_IDX+1]])))
print("Subjects available: ", np.unique(X_init[:,SUBJECT_IDX]))
X_test, X_train_valid, y_test, y_train_valid = train_test_split_according_to_age(
X_init,
y_init,
use_continuous_age=USE_CONTINUOUS_AGE,
subjects_test=TEST_SET_SUBJECTS)
print(X_test.shape, X_train_valid.shape, y_test.shape, y_train_valid.shape)
```
## LinearSVC validation
____
```
NB_KFOLDS = 5
NB_CATEGORICAL_FEATURES = 2
NB_FEATURES = 48
CLASSIFIER_PIPELINE_KEY = 'classifier'
RANDOM_STATE = 42
def get_cv_iterator(n_splits=2):
return GroupKFold(n_splits=n_splits).split(
X_train_valid, groups=X_train_valid[:,SUBJECT_IDX]
)
def cross_validate_with_confusion_matrix(pipeline, n_fold):
accuracies = []
macro_f1_scores = []
weighted_f1_scores = []
kappa_agreements = []
for train_index, valid_index in get_cv_iterator(n_splits=n_fold):
# We drop the subject and night indexes
X_train, X_valid = X_train_valid[train_index, 2:], X_train_valid[valid_index, 2:]
y_train, y_valid = y_train_valid[train_index], y_train_valid[valid_index]
# Scaling features and model training
training_pipeline = pipeline
training_pipeline.fit(X_train, y_train)
# Validation
y_valid_pred = training_pipeline.predict(X_valid)
print("----------------------------- FOLD RESULTS --------------------------------------\n")
current_kappa = cohen_kappa_score(y_valid, y_valid_pred)
print("TRAIN:", train_index, "VALID:", valid_index, "\n\n")
print(confusion_matrix(y_valid, y_valid_pred), "\n")
print(classification_report(y_valid, y_valid_pred, target_names=SLEEP_STAGES_VALUES.keys()), "\n")
print("Agreement score (Cohen Kappa): ", current_kappa, "\n")
accuracies.append(round(accuracy_score(y_valid, y_valid_pred),2))
macro_f1_scores.append(f1_score(y_valid, y_valid_pred, average="macro"))
weighted_f1_scores.append(f1_score(y_valid, y_valid_pred, average="weighted"))
kappa_agreements.append(current_kappa)
print_results_cv(accuracies, macro_f1_scores, weighted_f1_scores, kappa_agreements)
%%time
cross_validate_with_confusion_matrix(get_pipeline(
classifier=LinearSVC(
dual=False, # Prefer dual=False when n_samples > n_features. (documentation)
class_weight="balanced",
random_state=RANDOM_STATE
)
), n_fold=NB_KFOLDS)
```
CV Results:
```
Mean accuracy : 0.70 ± 0.026
Mean macro F1-score : 0.63 ± 0.025
Mean weighted F1-score : 0.69 ± 0.026
Mean Kappa's agreement : 0.59 ± 0.039
```
## Validation results
___
### Dimension reduction
___
*Definitions from scikit learn*:
Principal Component Analysis (PCA) applied to this data identifies the combination of attributes (principal components, or directions in the feature space) that account for the most variance in the data.
Linear Discriminant Analysis (LDA) tries to identify attributes that account for the most variance between classes. In particular, LDA, in contrast to PCA, is a supervised method, using known class labels.
We will compare each method and keep the one with the best results.
```
def cross_validate_with_dim_reduction(dim_reduction):
scores = cross_validate(
estimator=get_pipeline(
classifier=LinearSVC(
dual=False, # Prefer dual=False when n_samples > n_features. (documentation)
class_weight="balanced",
random_state=RANDOM_STATE
),
dimension_reduction=dim_reduction
),
X=X_train_valid,
y=y_train_valid,
groups=X_train_valid[:,SUBJECT_IDX],
scoring={
"agreement": make_scorer(cohen_kappa_score),
"accuracy": 'accuracy',
"f1-score-macro": 'f1_macro',
"f1-score-weighted": 'f1_weighted',
},
cv=get_cv_iterator(n_splits=5),
return_train_score=True,
verbose=1,
n_jobs=-1
)
print_results_cv_scores(scores)
```
#### 1. Linear discriminant analysis
___
LDA only allows `n_components` between 1 and `min(n_classes - 1, n_features)`. We will then **reduce our 48 features to 4 components**.
A particularity of LDA is that it has pratically no hyperparameters to fix (except the number of components, which is limited).
```
%%time
cross_validate_with_dim_reduction(LinearDiscriminantAnalysis())
```
#### 2. Principal component analysis
___
It is a non supervised dimension reduction method. In that sense, it can lead to worst results than LDA. On the other side, we can set the `n_components` to any value between 1 and `n_features`, which allows us to control the information that we leave out.
We've already looked at explained ratio by nb of components in the RF_HMM notebook. The results were:
```
1 components included: 0.9253
2 components included: 0.9405
4 components included: 0.9591
10 components included: 0.9807
16 components included: 0.9906
20 components included: 0.9943
30 components included: 0.9987
```
We will look at 4, 16 and 30 components.
```
%%time
cross_validate_with_dim_reduction(PCA(n_components=4))
%%time
cross_validate_with_dim_reduction(PCA(n_components=16))
%%time
cross_validate_with_dim_reduction(PCA(n_components=30))
%%time
cross_validate_with_dim_reduction(PCA(n_components=35))
```
If we resume all our results, which we tested upon 5 folds, we have the following:
| Score | without | LDA | PCA (n_comp=4) | PCA (n_comp=16) | PCA (n_comp=30) | PCA (n_comp=35) |
|--------------------|---------------|---------------|----------------|-----------------|-----------------|-----------------|
| accuracy | 0.70 ± 0.030 | 0.69 ± 0.033 | 0.53 ± 0.018 | 0.66 ± 0.028 | 0.69 ± 0.028 | 0.70 ± 0.030 |
| macro F1-score | 0.63 ± 0.027 | 0.60 ± 0.030 | 0.46 ± 0.011 | 0.58 ± 0.022 | 0.62 ± 0.027 | 0.62 ± 0.027 |
| weighted F1-score | 0.69 ± 0.028 | 0.67 ± 0.030 | 0.51 ± 0.011 | 0.64 ± 0.025 | 0.68 ± 0.028 | 0.69 ± 0.028 |
| Kappa's agreement | 0.59 ± 0.042 | 0.57 ± 0.044 | 0.37 ± 0.025 | 0.53 ± 0.037 | 0.58 ± 0.038 | 0.59 ± 0.041 |
| Time | 57.1 s | 12 s | 12.3 s | 16.2 s | 30 s | 36 s |
We will keep the PCA dimension reduction, with n_components=35, because it keeps the same accuracy, weighted F1-score and Cohen's Kappa as the model that doesn't not have dimension reduction.
### Hyperparameters tuning
___
#### 1. `C`: Regularization parameter
___
Inverse of regularization strength. Like in support vector machines, smaller values specify stronger regularization. It controls the trade off between smooth decision boundary and classifying the training points correctly. Increasing C values may lead to overfitting the training data.
```
%%time
evaluate_hyperparams_grid(
params={
f"{CLASSIFIER_PIPELINE_KEY}__C": np.linspace(1.95, 2.3, 10)
},
estimator=get_pipeline(
classifier=LinearSVC(
dual=False,
class_weight="balanced",
random_state=RANDOM_STATE
),
dimension_reduction=PCA(n_components=35)
),
X=X_train_valid,
y=y_train_valid,
cv=get_cv_iterator(n_splits=5),
use_randomized=False
)
```
**Results**:
|Rank| C | Test score |
|----|------------------|----------------|
|1 | 2.236842105263158| 0.5903 ± 0.039 |
|2 | 1.931578947368421| 0.5902 ± 0.039 |
|3 | 2.389473684210526| 0.5902 ± 0.039 |
...
CPU times: user 13.1 s, sys: 998 ms, total: 14.1 s
Wall time: 5min 28s
Other values gave the same test score (`0.5902 ± 0.039`): `0.557894, 1.626315, 1.473684, 1.015789, ... 2.847368`. We then looked at the range between 1.95 and 2.3:
|Rank| C | Test score |
|----|------------------|----------------|
|1 | 2.105 | 0.5903 ± 0.039 |
|2 | 2.144 | 0.5902 ± 0.039 |
|3 | 2.183 | 0.5902 ± 0.039 |
...
CPU times: user 13.4 s, sys: 1.19 s, total: 14.6 s
Wall time: 5min 53s
We then fixed the value to `2.105`.
## Meta-estimators
___
```
%%time
adaboosted_pipeline = Pipeline([
('scaling', ColumnTransformer([
('pass-through-categorical', 'passthrough', list(range(NB_CATEGORICAL_FEATURES))),
('scaling-continuous', StandardScaler(copy=False), list(range(NB_CATEGORICAL_FEATURES,NB_FEATURES)))
])),
("dim_reduction", PCA(n_components=35)),
(CLASSIFIER_PIPELINE_KEY, AdaBoostClassifier(
LinearSVC(
C=2.105,
dual=False,
class_weight="balanced",
# random_state=RANDOM_STATE
),
algorithm='SAMME',
n_estimators=100,
random_state=RANDOM_STATE
))
])
scores = cross_validate(
estimator=adaboosted_pipeline,
X=X_train_valid,
y=y_train_valid,
groups=X_train_valid[:,SUBJECT_IDX],
scoring={
"agreement": make_scorer(cohen_kappa_score),
"accuracy": 'accuracy',
"f1-score-macro": 'f1_macro',
"f1-score-weighted": 'f1_weighted',
},
cv=get_cv_iterator(n_splits=2),
return_train_score=True,
verbose=1,
n_jobs=-1
)
print_results_cv_scores(scores)
```
Results:
```
Mean accuracy : 0.67 ± 0.003
Mean macro F1-score : 0.60 ± 0.005
Mean weighted F1-score : 0.66 ± 0.003
Mean Kappa's agreement : 0.55 ± 0.001
CPU times: user 49.7 ms, sys: 57 ms, total: 107 ms
Wall time: 30.5 s
```
The results have not been improved compared to the same model without the `AdaBoost` meta-estimator.
## SVM testing
___
```
%%time
testing_pipeline = get_pipeline(
classifier=LinearSVC(
dual=False,
C=2.105,
class_weight="balanced",
random_state=RANDOM_STATE
),
dimension_reduction=PCA(n_components=35)
)
testing_pipeline.fit(X_train_valid[:, 2:], y_train_valid);
y_test_pred = testing_pipeline.predict(X_test[:,2:])
print(confusion_matrix(y_test, y_test_pred))
print(classification_report(y_test, y_test_pred, target_names=SLEEP_STAGES_VALUES.keys()))
print("Agreement score (Cohen Kappa): ", cohen_kappa_score(y_test, y_test_pred))
```
### Test results
___
#### a) Without PCA and without tuning (C=1)
___
```
[[1443 48 7 24 102]
[ 244 244 298 4 193]
[ 71 89 3037 318 88]
[ 4 0 16 591 0]
[ 97 161 240 3 801]]
precision recall f1-score support
W 0.78 0.89 0.83 1624
N1 0.45 0.25 0.32 983
N2 0.84 0.84 0.84 3603
N3 0.63 0.97 0.76 611
REM 0.68 0.62 0.64 1302
accuracy 0.75 8123
macro avg 0.68 0.71 0.68 8123
weighted avg 0.74 0.75 0.74 8123
Agreement score (Cohen Kappa): 0.6557078634244826
```
#### b) Without PCA and with tuning (C=2.105)
___
```
precision recall f1-score support
W 0.78 0.89 0.83 1624
N1 0.45 0.25 0.32 983
N2 0.84 0.84 0.84 3603
N3 0.63 0.97 0.76 611
REM 0.68 0.62 0.64 1302
accuracy 0.75 8123
macro avg 0.67 0.71 0.68 8123
weighted avg 0.74 0.75 0.74 8123
Agreement score (Cohen Kappa): 0.6555409806957044
```
#### c) With PCA and tuning (C=2.105)
___
```
precision recall f1-score support
W 0.77 0.88 0.82 1624
N1 0.44 0.24 0.31 983
N2 0.84 0.84 0.84 3603
N3 0.62 0.97 0.76 611
REM 0.68 0.62 0.64 1302
accuracy 0.75 8123
macro avg 0.67 0.71 0.68 8123
weighted avg 0.74 0.75 0.73 8123
Agreement score (Cohen Kappa): 0.6506513702343493
```
```
print("Test subjects are subjects: ", np.unique(X_test[:,0]))
plt.rcParams["figure.figsize"] = (20,5)
for test_subject in np.unique(X_test[:,0]):
test_subject_indexes = [idx for idx, elem in enumerate(X_test) if elem[0] == test_subject]
for night_idx in np.unique(X_test[test_subject_indexes,1]):
test_night_subject_indexes = [
idx for idx, elem in enumerate(X_test)
if elem[0] == test_subject and elem[1] == night_idx]
hypnograms = [
y_test[test_night_subject_indexes],
y_test_pred[test_night_subject_indexes]
]
print_hypnogram(hypnograms,
labels=["scored", "predicted"],
subject=test_subject,
night=night_idx)
```
## Saving trained model
___
```
SAVED_DIR = "../trained_model"
if not os.path.exists(SAVED_DIR):
os.mkdir(SAVED_DIR);
if USE_CONTINUOUS_AGE:
joblib.dump(testing_pipeline, f"{SAVED_DIR}/classifier_SVC_age_continuous.joblib")
else:
joblib.dump(testing_pipeline, f"{SAVED_DIR}/classifier_SVC.joblib")
```
| github_jupyter |
# Create a Dataset for Sentiment Analysis
> TL;DR In this tutorial, you'll learn how to create a dataset for Sentiment Analysis by scraping user reviews for Android apps. You'll convert the app and review information into Data Frames and save that to CSV files.
- [Read the tutorial](https://www.curiousily.com/posts/create-dataset-for-sentiment-analysis-by-scraping-google-play-app-reviews-using-python/)
- [Run the notebook in your browser (Google Colab)](https://colab.research.google.com/drive/1GDJIpz7BXw55jl9wTOMQDool9m8DIOyp)
- [Read the `Getting Things Done with Pytorch` book](https://github.com/curiousily/Getting-Things-Done-with-Pytorch)
You'll learn how to:
- Set a goal and inclusion criteria for your dataset
- Get real-world user reviews by scraping Google Play
- Use Pandas to convert and save the dataset into CSV files
## Setup
Let's install the required packages and setup the imports:
```
!pip install -qq google-play-scraper
!pip install -qq -U watermark
%reload_ext watermark
%watermark -v -p pandas,matplotlib,seaborn,google_play_scraper
import json
import pandas as pd
from tqdm import tqdm
import seaborn as sns
import matplotlib.pyplot as plt
from pygments import highlight
from pygments.lexers import JsonLexer
from pygments.formatters import TerminalFormatter
from google_play_scraper import Sort, reviews, app
%matplotlib inline
%config InlineBackend.figure_format='retina'
sns.set(style='whitegrid', palette='muted', font_scale=1.2)
```
## The Goal of the Dataset
You want to get feedback for your app. Both negative and positive are good. But the negative one can reveal critical features that are missing or downtime of your service (when it is much more frequent).
Lucky for us, Google Play has plenty of apps, reviews, and scores. We can scrape app info and reviews using the [google-play-scraper](https://github.com/JoMingyu/google-play-scraper) package.
You can choose plenty of apps to analyze. But different app categories contain different audiences, domain-specific quirks, and more. We'll start simple.
We want apps that have been around some time, so opinion is collected organically. We want to mitigate advertising strategies as much as possible. Apps are constantly being updated, so the time of the review is an important factor.
Ideally, you would want to collect every possible review and work with that. However, in the real world data is often limited (too large, inaccessible, etc). So, we'll do the best we can.
Let's choose some apps that fit the criteria from the *Productivity* category. We'll use [AppAnnie](https://www.appannie.com/apps/google-play/top-chart/?country=US&category=29&device=&date=2020-04-05&feed=All&rank_sorting_type=rank&page_number=1&page_size=100&table_selections=) to select some of the top US apps:
```
app_packages = [
'com.anydo',
'com.todoist',
'com.ticktick.task',
'com.habitrpg.android.habitica',
'cc.forestapp',
'com.oristats.habitbull',
'com.levor.liferpgtasks',
'com.habitnow',
'com.microsoft.todos',
'prox.lab.calclock',
'com.gmail.jmartindev.timetune',
'com.artfulagenda.app',
'com.tasks.android',
'com.appgenix.bizcal',
'com.appxy.planner'
]
```
## Scraping App Information
Let's scrape the info for each app:
```
app_infos = []
for ap in tqdm(app_packages):
info = app(ap, lang='en', country='us')
del info['comments']
app_infos.append(info)
```
We got the info for all 15 apps. Let's write a helper function that prints JSON objects a bit better:
```
def print_json(json_object):
json_str = json.dumps(
json_object,
indent=2,
sort_keys=True,
default=str
)
print(highlight(json_str, JsonLexer(), TerminalFormatter()))
```
Here is a sample app information from the list:
```
print_json(app_infos[0])
```
This contains lots of information including the number of ratings, number of reviews and number of ratings for each score (1 to 5). Let's ignore all of that and have a look at their beautiful icons:
```
def format_title(title):
sep_index = title.find(':') if title.find(':') != -1 else title.find('-')
if sep_index != -1:
title = title[:sep_index]
return title[:10]
fig, axs = plt.subplots(2, len(app_infos) // 2, figsize=(14, 5))
for i, ax in enumerate(axs.flat):
ai = app_infos[i]
img = plt.imread(ai['icon'])
ax.imshow(img)
ax.set_title(format_title(ai['title']))
ax.axis('off')
```
We'll store the app information for later by converting the JSON objects into a Pandas dataframe and saving the result into a CSV file:
```
app_infos_df = pd.DataFrame(app_infos)
app_infos_df.to_csv('apps.csv', index=None, header=True)
```
## Scraping App Reviews
In an ideal world, we would get all the reviews. But there are lots of them and we're scraping the data. That wouldn't be very polite. What should we do?
We want:
- Balanced dataset - roughly the same number of reviews for each score (1-5)
- A representative sample of the reviews for each app
We can satisfy the first requirement by using the scraping package option to filter the review score. For the second, we'll sort the reviews by their helpfulness, which are the reviews that Google Play thinks are most important. Just in case, we'll get a subset from the newest, too:
```
app_reviews = []
for ap in tqdm(app_packages):
for score in list(range(1, 6)):
for sort_order in [Sort.MOST_RELEVANT, Sort.NEWEST]:
rvs, _ = reviews(
ap,
lang='en',
country='us',
sort=sort_order,
count= 200 if score == 3 else 100,
filter_score_with=score
)
for r in rvs:
r['sortOrder'] = 'most_relevant' if sort_order == Sort.MOST_RELEVANT else 'newest'
r['appId'] = ap
app_reviews.extend(rvs)
```
Note that we're adding the app id and sort order to each review. Here's an example for one:
```
print_json(app_reviews[0])
```
`repliedAt` and `replyContent` contain the developer response to the review. Of course, they can be missing.
How many app reviews did we get?
```
len(app_reviews)
```
Let's save the reviews to a CSV file:
```
app_reviews_df = pd.DataFrame(app_reviews)
app_reviews_df.to_csv('reviews.csv', index=None, header=True)
```
## Summary
Well done! You now have a dataset with more than 15k user reviews from 15 productivity apps. Of course, you can go crazy and get much much more.
- [Read the tutorial](https://www.curiousily.com/posts/create-dataset-for-sentiment-analysis-by-scraping-google-play-app-reviews-using-python/)
- [Run the notebook in your browser (Google Colab)](https://colab.research.google.com/drive/1GDJIpz7BXw55jl9wTOMQDool9m8DIOyp)
- [Read the `Getting Things Done with Pytorch` book](https://github.com/curiousily/Getting-Things-Done-with-Pytorch)
You learned how to:
- Set goals and expectations for your dataset
- Scrape Google Play app information
- Scrape user reviews for Google Play apps
- Save the dataset to CSV files
Next, we're going to use the reviews for sentiment analysis with BERT. But first, we'll have to do some text preprocessing!
## References
- [Google Play Scraper for Python](https://github.com/JoMingyu/google-play-scraper)
| github_jupyter |
```
import pickle
import pandas as pd
import numpy as np
from collections import defaultdict
import math
import matplotlib.pyplot as plt
import networkx as nx
```
# 1. Load input
```
lines = 500000
```
Load episode set $E$ with the users that retweeted each original tweet in the trace.
```
E = pickle.load(open("./extracted/E"+ str(lines) + ".p", "rb"))
for tweet in E:
E[tweet] = list(dict.fromkeys(E[tweet]))
D = pickle.load(open("./extracted/D"+ str(lines) + ".p", "rb"))
```
Load the set of original tweets denoted by $S$.
The set of original tweets is denoted by $S$, where $S$ = S is the total number of original tweets
```
S = pickle.load(open("./extracted/S"+ str(lines) + ".p", "rb"))
```
Load $U$ set xith unique users
```
U = pickle.load(open("./extracted/U"+ str(lines) + ".p", "rb"))
U = list(U)
```
Load $M_{ij}$ variables that count number of episodes where the ordered pair (i,j) appears
```
M = pickle.load(open("./extracted/M"+ str(lines) + ".p", "rb"))
```
Load $Q_{ij}$ results for the ordered pair (i,j) derived from the consrained algorithm
```
Q = pickle.load(open("./extracted/Q_constrained_"+ str(lines) + ".p", "rb"))
```
Load $s_{ij}$ derived from the consrained algorithm
```
s = pickle.load(open("./extracted/s_constrained_"+ str(lines) + ".p", "rb"))
```
Load $k_{ij}$ derived from Saito
```
k = pickle.load(open("./extracted/k_saito_"+ str(lines) + ".p", "rb"))
```
Load $Q_{ij}$ derived from Newman's
```
Q_newman = pickle.load(open("./extracted/Q_newman_"+ str(lines) + ".p", "rb"))
```
## Necessary functions
```
def flatten(obj):
if type(obj) == list:
return [l for L in obj for l in L]
if type(obj) == dict:
return [l for i in obj for l in obj[i].values()]
if type(obj) == defaultdict:
return [l for i in obj for l in obj[i].values()]
def create_chain_graph(U,D):
G = nx.DiGraph()
G.add_nodes_from(U)
for tweet in D:
for time in D[tweet]:
ind = list(D[tweet].keys()).index(time)
if ind+1==len(D[tweet]): break
next_time = list(D[tweet].keys())[ind+1]
for u1 in D[tweet][time]:
for u2 in D[tweet][next_time]:
G.add_edge(u1,u2)
return G
def create_saito_graph(U,k):
G = nx.DiGraph()
G.add_nodes_from(U)
for i in k:
for j in k[i]:
if k[i][j] > 0.5:
G.add_edge(i,j)
return G
def create_star_graph(U,E):
G = nx.DiGraph()
G.add_nodes_from(U)
for s in E:
for j in E[s][1:]:
G.add_edge(E[s][0],j)
return G
def create_our_graph(U,Q):
G = nx.DiGraph()
G.add_nodes_from(U)
for i in Q:
for j in Q[i]:
if Q[i][j] > 0.5:
G.add_edge(i,j)
return G
def create_newman_graph(U,Q):
G = nx.DiGraph()
G.add_nodes_from(U)
for i in Q:
for j in Q[i]:
if Q[i][j] > 0.5:
G.add_edge(i,j)
return G
def f_check(E, Q):
'''
Function that checks feasibility of graph.
'''
retweets = 0 # minimum existing edges
infeasible_episodes = 0
total_feasible_edges = []
total_inf = 0
for s in E:
feasible_edges = 0
for j in E[s]:
indx = E[s].index(j)
if indx!=0:
u_before = E[s][:indx]
for i in u_before:
if i in Q and j in Q[i] and Q[i][j] > 0.5:
feasible_edges +=1
total_feasible_edges.append((i,j))
infeasible = (len(E[s]) - 1) - feasible_edges
if infeasible > 0:
total_inf+=infeasible
infeasible_episodes+=1
retweets += len(E[s])-1
total_feasible_edges = list(set(total_feasible_edges))
return infeasible_episodes
def longest_path(E, U, Q, k, graph_type):
max_l = 0
max_path = 0
if graph_type=='ours' or graph_type=='newman':
for s in E:
G = nx.DiGraph()
G.add_nodes_from(U)
for j in E[s][1:]:
indx = E[s].index(j)
u_before = E[s][:indx]
for i in u_before:
if j in Q[i] and Q[i][j] > 0.5:
G.add_edge(i,j)
if len(nx.dag_longest_path(G))>max_l:
max_l = len(nx.dag_longest_path(G))
max_path = nx.dag_longest_path(G)
if graph_type=='star':
for s in E:
G = nx.DiGraph()
G.add_nodes_from(U)
for j in E[s][1:]:
G.add_edge(E[s][0],j)
if len(nx.dag_longest_path(G))>max_l:
max_l = len(nx.dag_longest_path(G))
max_path = nx.dag_longest_path(G)
if graph_type=='saito':
for s in E:
G = nx.DiGraph()
G.add_nodes_from(U)
for j in E[s][1:]:
indx = E[s].index(j)
u_before = E[s][:indx]
for i in u_before:
if j in k[i] and k[i][j] > 0.5:
G.add_edge(i,j)
if len(nx.dag_longest_path(G))>max_l:
max_l = len(nx.dag_longest_path(G))
max_path = nx.dag_longest_path(G)
return max_l, max_path
def plot_tweetgraph(s, E, S, Q, k, pos, graph_type):
G = nx.DiGraph()
if graph_type=='ours' or graph_type=='newman':
t = 0
for j in E[s][1:]:
G.add_node(j)
indx = E[s].index(j)
u_before = E[s][:indx]
for i in u_before:
G.add_node(i)
if j in Q[i] and Q[i][j] > 0.5:
G.add_edge(i,j, label=f'{t}')
t+=1
elif graph_type=='star':
G.add_node(E[s][0])
for j in E[s][1:]:
G.add_edge(E[s][0],j)
G.add_node(j)
elif graph_type=='saito':
t = 0
for j in E[s][1:]:
G.add_node(j)
indx = E[s].index(j)
u_before = E[s][:indx]
for i in u_before:
G.add_node(i)
if j in k[i] and k[i][j] > 0.5:
G.add_edge(i,j, label=f'{t}')
t+=1
elif graph_type=='chain':
for time in D[s]:
ind = list(D[tweet].keys()).index(time)
if ind+1==len(D[tweet]): break
next_time = list(D[tweet].keys())[ind+1]
for u1 in D[tweet][time]:
for u2 in D[tweet][next_time]:
G.add_edge(u1,u2)
color_map = []
cmap = plt.get_cmap('Greens')
for node in G:
if node == S[tweet]:
color_map.append('green')
else:
color_map.append('yellow')
pos = nx.spring_layout(G)
nx.draw_networkx_nodes(G, pos, node_color = color_map, cmap=plt.get_cmap('jet'), node_size = 300)
nx.draw_networkx_edges(G, pos, edge_color='r', arrows=True)
nx.draw_networkx_labels(G, pos, font_size=8)
if graph_type=='ours' or graph_type=='saito':
nx.draw_networkx_edge_labels(G, pos, font_size=8)
plt.show()
```
# Statistics
## 1. number of infeasible episodes
```
data = dict()
data['Graph Type with Lines: ' + str(lines)] = ['Ours','Saito','Star','Chain', 'Newman']
inf_ep_ours = f_check(E, Q)
inf_ep_saito = f_check(E, k)
inf_ep_newman = f_check(E, Q_newman)
data['Infeasible Episodes'] = [inf_ep_ours, inf_ep_saito, 0, 0, inf_ep_newman]
```
## 2. Number of edges
```
G_star = create_star_graph(U,E)
G_ours = create_our_graph(U,Q)
G_saito = create_saito_graph(U,k)
G_newman = create_newman_graph(U,Q_newman)
G_chain = create_chain_graph(U,D)
edges_ours = len(G_ours.edges())
edges_saito = len(G_saito.edges())
edges_star = len(G_star.edges())
edges_chain = len(G_chain.edges())
edges_newman = len(G_newman.edges())
data['Number of edges'] = [edges_ours, edges_saito, edges_star, edges_chain, edges_newman]
```
## 3. Average out degree
```
av_ours = sum(d[1] for d in G_ours.out_degree())/float(len(G_ours))
av_saito = sum(d[1] for d in G_saito.out_degree())/float(len(G_saito))
av_star = sum(d[1] for d in G_star.out_degree())/float(len(G_star))
av_chain = sum(d[1] for d in G_chain.out_degree())/float(len(G_chain))
av_newman = sum(d[1] for d in G_newman.out_degree())/float(len(G_newman))
data['Average out degree'] = [av_ours, av_saito, av_star, av_chain, av_newman]
degree_sequence = list(G_ours.out_degree())
max_degree_our = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_saito.out_degree())
max_degree_saito = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_star.out_degree())
max_degree_star = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_chain.out_degree())
max_degree_chain = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_newman.out_degree())
max_degree_newman = max(np.array(degree_sequence)[:,1])
data['Max out degree'] = [max_degree_our, max_degree_saito, max_degree_star, max_degree_chain, max_degree_newman]
degree_sequence = list(G_ours.in_degree())
max_degree_our = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_saito.in_degree())
max_degree_saito = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_star.in_degree())
max_degree_star = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_chain.in_degree())
max_degree_chain = max(np.array(degree_sequence)[:,1])
degree_sequence = list(G_newman.in_degree())
max_degree_newman = max(np.array(degree_sequence)[:,1])
data['Max in degree'] = [max_degree_our, max_degree_saito, max_degree_star, max_degree_chain, max_degree_newman]
```
## 4. Graph diameter
The maximum among all the distances between a vertex to all other vertices is considered as the diameter of the Graph G.
```
def avg_sh_path(G, graph_type):
lst = dict(nx.all_pairs_shortest_path_length(G))
sum_lst = sum(lst[i][j] for i in lst for j in lst[i])
l_lst = sum(1 for i in lst for j in lst[i] if i!=j)
avg_spl = sum_lst/l_lst
diameter = [max(val.values()) for key, val in lst.items()]
diameter = max(diameter)
return avg_spl, diameter
avg_ours, d_ours = avg_sh_path(G_ours, 'Ours')
avg_saito, d_saito = avg_sh_path(G_saito, 'Saito')
avg_star, d_star = avg_sh_path(G_star, 'Star')
avg_chain, d_chain= avg_sh_path(G_chain, 'Chain')
avg_newman, d_newman = avg_sh_path(G_newman, 'Newman')
data['Graph diameter'] = [d_ours, d_saito, d_star, d_chain, d_newman]
data['Average shortest path'] = [avg_ours, avg_saito, avg_star, avg_chain, avg_newman]
```
# 5. Number of connected components
```
def number_cc(G, graph_type):
scc = 0
wcc = 0
for C in nx.strongly_connected_components(G):
C = G.subgraph(C)
if len(C)>1: # skip one nodes
scc+=1
for C in nx.weakly_connected_components(G):
C = G.subgraph(C)
if len(C)>1: # skip one nodes
wcc+=1
return scc, wcc
scc_ours, wcc_ours = number_cc(G_ours, 'Ours')
scc_saito, wcc_saito = number_cc(G_saito, 'Saito')
scc_star, wcc_star = number_cc(G_star, 'Star')
scc_chain, wcc_chain = number_cc(G_chain, 'Chain')
scc_newman, wcc_newman = number_cc(G_newman, 'Newman')
data['Number of scc'] = [scc_ours, scc_saito, scc_star, scc_chain, scc_newman]
data['Number of wcc'] = [wcc_ours, wcc_saito, wcc_star, wcc_chain, wcc_newman]
```
# Save results
```
df = pd.DataFrame(data)
df.to_csv('./Results.csv', mode='a', index = False, header=True)
```
| github_jupyter |
# OrionExplorer - CSV Processing
This is a demo notebook showing how to use the `lstm_dynamic_threshold.json` pipeline to analyze a collection of signal CSV files and later on retrieve the list of Events found.
## 1. Create an OrionExlorer Instance
In this first step, we setup the environment, import the `OrionExplorer` and create
an instance passing the name of the database which we want to connect to.
```
import logging;
logging.basicConfig(level=logging.ERROR)
logging.getLogger().setLevel(level=logging.ERROR)
import warnings
warnings.simplefilter("ignore")
from orion.explorer import OrionExplorer
explorer = OrionExplorer(database='orion-process-csvs')
```
In this case we will drop the database before starting to make sure that we are working
on a clean environment.
**WARNING**: This will remove all the data that exists in this database!
```
explorer.drop_database()
```
## 2. Add the pipeline that we will be using
The second step is to register the pipeline that we are going to use.
For this, we will enter:
* a pipeline name.
* the path to the `lstm_dynamic_threshold` json.
```
pipeline = explorer.add_pipeline(
'lstm_dynamic_threshold',
'../orion/pipelines/lstm_dynamic_threshold.json'
)
```
Afterwards, we can obtain the list of pipelines to see if it has been properly registered
```
explorer.get_pipelines()
```
## 3. Get the list of CSV files
In this example we will use the `os` module to find the list of CSV files that exist inside the directory
`data` that we have created inside this `notebooks` folder.
Another way to do it would be to provide an explicit list of filenames
```
import os
CSVS_FOLDER = './data'
csvs = os.listdir(CSVS_FOLDER)
csvs
```
## 3. Register the new datasets
We will execute a loop in which, for each CSV file, we will register a new Dataset in the Database.
For each CSV, the name that we will use for dataset and the signal will be name of the file without the `.csv` extension, and will be leaving the satellite_id blank.
In this case we need no additional arguments, such as timestamp_column or value_column, but if they were required
we would add them to the `add_dataset` call.
We will also capture the output of the `add_dataset` call in a list, so we can use these datasets later on.
```
datasets = list()
cwd = os.getcwd()
for path in csvs:
name = os.path.basename(path)[:-4]
location = os.path.join(CSVS_FOLDER, path)
print('Adding dataset {} for CSV {}'.format(name, location))
dataset = explorer.add_dataset(
name,
name,
location=location,
timestamp_column=None, # Replace if needed
value_column=None, # Replace if needed
)
datasets.append(dataset)
```
Afterwards we can check that the datasets were properly registered
```
explorer.get_datasets()
```
## 4. Run the pipeline on the datasets
Once the pipeline and the datasets are registered, we can start the processing loop.
```
for dataset in datasets:
print('Analyzing dataset {}'.format(dataset.name))
explorer.analyze(dataset.name, pipeline.name)
```
## 5. Analyze the results
Once the execution has finished, we can explore the Dataruns and the detected Events.
```
explorer.get_dataruns()
explorer.get_events()
```
| github_jupyter |
# 1. PubMed
Search PubMed for papers
https://www.ncbi.nlm.nih.gov/pubmed/
https://www.ncbi.nlm.nih.gov/books/NBK25499/
```
import os
import lcp.reuse as reuse
from Bio import Entrez
from IPython.display import display
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
PROJECT = 'eICU' # 'MIMIC'
def full_query(base_query, restriction_query):
return ' AND '.join([base_query, restriction_query])
entrez_email = 'mimic-support@physionet.org'
if PROJECT == 'eICU':
# Retrieves publications which cite this reference
PMID = '30204154'
mimic_query = '(mimic-ii OR mimic-iii OR mimicii OR mimiciii OR mimic-2 OR mimic-3 OR mimic2 OR mimic3)'
if PROJECT == 'MIMIC':
base_query = mimic_query
elif PROJECT == 'eICU':
base_query = f'(eicu-crd OR "eICU Collaborative Research Database" OR (eicu AND ({mimic_query} OR database OR MIT OR Phillips)))'
# Other terms added to remove false positives. The more terms added without increasing FPs, the better.
if PROJECT == 'MIMIC':
restriction_query = '(physionet OR icu OR "intensive care" OR "critical care")'
elif PROJECT == 'eICU':
restriction_query = '("intensive care" OR "critical care")'
# more restriction keyword ideas: clinical, database, waveform (not suitable on their own due to general mimic term)
full_query = full_query(base_query, restriction_query)
if PMID:
search_strings = [
'PMID'
]
else:
search_strings = [
base_query#,
# full_query
]
# search_results = reuse.search_list(search_strings, entrez_email)
if PMID:
citations = []
pmc_ids = Entrez.read(Entrez.elink(dbfrom='pubmed', db='pmc', LinkName='pubmed_pmc_refs', from_uid=PMID))
pmc_ids = [list(i.values())[0] for i in pmc_ids[0]['LinkSetDb'][0]['Link']]
pm_ids = Entrez.read(Entrez.elink(dbfrom='pmc', db='pubmed', LinkName='pmc_pubmed', from_uid=','.join(pmc_ids)))
for pm_id in [list(i.values())[0] for i in pm_ids[0]['LinkSetDb'][0]['Link']]:
citations.append(pm_id)
search_results_all = reuse.search_list(entrez_email, ids=citations, all=True)
else:
search_results_all = reuse.search_list(entrez_email, search_strings=search_strings, all=True)
# Display number of results
if PMID:
result = search_results_all['PMID']
print(f'PMID:\n - Count: {len(result.index)}')
else:
for ss in search_strings:
result = search_results_all[ss]
print(f'{ss}:\n - Count: {len(result.index)}')
# Write the titles to files
write_dir = os.path.join('search_results', PROJECT, 'pubmed')
os.makedirs(write_dir, exist_ok=True)
if PMID:
search_results_all['PMID'].to_csv(os.path.join(write_dir, 'PMID-citations-all.csv'), index=False)
else:
search_results_all[base_query].to_csv(os.path.join(write_dir, 'without-constraints-all.csv'), index=False)
search_results_all[full_query].to_csv(os.path.join(write_dir, 'with-constraints-all.csv'), index=False)
# base_query_file = os.path.join(write_dir, 'without-constraints.txt')
# with open(base_query_file, 'w') as f:
# for line in search_results[base_query].paper_titles:
# f.write(line+'\n')
# full_query_file = os.path.join(write_dir, 'with-constraints.txt')
# with open(full_query_file, 'w') as f:
# for line in search_results[full_query].paper_titles:
# f.write(line+'\n')
# Create a new one for each year
if PMID:
all_years = sorted(set(list(search_results_all['PMID']['Publication Year'])))
for year in all_years:
year_df = search_results_all['PMID'][search_results_all['PMID']['Publication Year'] == year]
year_df.to_csv(os.path.join(write_dir, f'PMID-citations-all_{year}.csv'), index=False)
else:
all_years = sorted(set(list(search_results_all[full_query]['Publication Year'])))
for year in all_years:
year_df_without = search_results_all[base_query][search_results_all[base_query]['Publication Year'] == year]
year_df_with = search_results_all[full_query][search_results_all[full_query]['Publication Year'] == year]
year_df_without.to_csv(os.path.join(write_dir, f'without-constraints-all_{year}.csv'), index=False)
year_df_with.to_csv(os.path.join(write_dir, f'with-constraints-all_{year}.csv'), index=False)
# Create a histogram of number of publications each year
if PMID:
all_years = sorted(set(list(search_results_all['PMID']['Publication Year'])))
years = []
for year in all_years:
year_df = search_results_all['PMID'][search_results_all['PMID']['Publication Year'] == year]
for _ in range(len(year_df.index)):
years.append(year)
plt.figure(figsize=(10,6))
plt.hist(years, facecolor='k', edgecolor='w', bins=np.arange(len(all_years)+1)-0.5)
plt.savefig(os.path.join(write_dir, 'PMID-citations_histogram.jpg'))
plt.xlabel('Year', fontsize=12)
plt.ylabel('Number of Publications', fontsize=12)
plt.show()
else:
all_years = sorted(set(list(search_results_all[full_query]['Publication Year'])))
year_without = []
year_with = []
for year in all_years:
year_df_without = search_results_all[base_query][search_results_all[base_query]['Publication Year'] == year]
year_df_with = search_results_all[full_query][search_results_all[full_query]['Publication Year'] == year]
for _ in range(len(year_df_without.index)):
year_without.append(year)
for _ in range(len(year_df_with.index)):
year_with.append(year)
plt.figure(figsize=(10,6))
plt.hist(year_without, facecolor='k', edgecolor='w', bins=np.arange(len(all_years)+1)-0.5)
plt.savefig(os.path.join(write_dir, 'without-constraints_histogram.jpg'))
plt.xlabel('Year', fontsize=12)
plt.ylabel('Number of Publications', fontsize=12)
plt.show()
plt.figure(figsize=(10,6))
plt.hist(year_with, facecolor='k', edgecolor='w', bins=np.arange(len(all_years)+1)-0.5)
plt.savefig(os.path.join(write_dir, 'with-constraints_histogram.jpg'))
plt.xlabel('Year', fontsize=12)
plt.ylabel('Number of Publications', fontsize=12)
plt.show()
# DO: Create a file called without-constraints-inspected.tsv and mark the second column with T/F for true/false positives.
# Look at the effect of restricting the search by additional criteria.
# The differences show that many false positives, and a few true positives, are removed.
reuse.showdiff(search_results[search_strings[0]],
search_results[search_strings[1]])
# Read in the labelled results for the general unconstrained search query
labelled_results = pd.read_csv(os.path.join(write_dir, 'without-constraints-inspected.tsv'), delimiter='\t', header=None)
false_positives = labelled_results.loc[labelled_results[1]=='F'][0].values
true_positives = labelled_results.loc[labelled_results[1]=='T'][0].values
print('Number of results found using the unconstrained search term:', len(labelled_results))
print('Number of false positives:',len(false_positives))
print('Number of true positives:',len(true_positives))
constrained_titles = search_results[search_strings[1]].paper_titles
print('\nCompare ^ true positives with:')
print('Number of results from the constrained search term:', len(constrained_titles))
missed_papers = set(true_positives) - set(constrained_titles)
print('Number of missed true positives:', len(missed_papers))
# Take a look at some True positives missed by the constrained search term to figure out what else you can add.
# Write to a file to label comments.
write_dir = os.path.join('search_results', 'pubmed')
missed_papers_file = os.path.join(write_dir, 'missed-papers.tsv')
with open(missed_papers_file, 'w') as f:
for line in missed_papers:
f.write(line+'\n')
display(missed_papers)
```
# 2. Web of Science
```
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import re
import time
from IPython.display import display
base_mimic_query = '(mimic-ii OR mimic-iii OR mimicii OR mimiciii OR mimic-2 OR mimic-3 OR mimic2 OR mimic3)'
restriction_query = '(physionet OR icu OR “intensive care” OR “critical care”)'
def full_query(base_query, restriction_query):
return ' AND '.join([base_query, restriction_query])
full_mimic_query = full_query(base_mimic_query, restriction_query)
#base_search_url = 'https://apps.webofknowledge.com/WOS_GeneralSearch_input.do?product=WOS&search_mode=GeneralSearch&SID=2F46AeWkMQBRAZlzDWm&preferencesSaved='
base_search_url = 'https://apps.webofknowledge.com/WOS_GeneralSearch_input.do?product=WOS&search_mode=GeneralSearch&SID=1AnC2UMojuKrtrl7T5R&preferencesSaved='
all_titles = []
# Get to the search page
driver = webdriver.Firefox()
driver.get(base_search_url)
# Input the query string
time.sleep(2.5)
searchbox = driver.find_element_by_id('value(input1)')
searchbox.send_keys(full_mimic_query)
# Search
time.sleep(1)
searchbutton = driver.find_element_by_css_selector('.standard-button.primary-button.large-search-button')
searchbutton.click()
# Get the total number of pages
npages = int(driver.find_element_by_id('pageCount.top').text)
# Get the titles!!!
while True:
# Get the current page number
pagenum = int(driver.find_element_by_class_name('goToPageNumber-input').get_property('value'))
# Get the titles. This also captures the journals. So every second value is not a title.
elements = driver.find_elements_by_class_name('smallV110')
for e in elements[::2]:
all_titles.append(e.find_element_by_tag_name('value').text)
if pagenum < npages:
nextbutton = driver.find_element_by_class_name('paginationNext')
nextbutton.click()
else:
print('Got all paper titles!')
driver.close()
break
all_titles = set(all_titles)
#all_titles.remove('')
all_titles = [t.lower() for t in list(all_titles)]
display(all_titles)
# Write the titles to files
write_dir = os.path.join('search_results/wos')
full_query_file = os.path.join(write_dir, 'with-constraints.txt')
with open(full_query_file, 'w') as f:
for line in all_titles:
f.write(line+'\n')
```
# 3. SCOPUS
Shit search
# 4. IEEE
```
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import re
import time
from IPython.display import display
import os
base_mimic_query = '(mimic-ii OR mimic-iii OR mimicii OR mimiciii OR mimic-2 OR mimic-3 OR mimic2 OR mimic3)'
restriction_query = '(physionet OR icu OR “intensive care” OR “critical care”)'
def full_query(base_query, restriction_query):
return ' AND '.join([base_query, restriction_query])
full_mimic_query = full_query(base_mimic_query, restriction_query)
base_search_url = 'http://ieeexplore.ieee.org/search/advsearch.jsp?expression-builder'
all_titles = []
# Get to the search page
driver = webdriver.Firefox()
driver.get(base_search_url)
# Input the query string
searchbox = driver.find_element_by_id('expression-textarea')
searchbox.send_keys(full_mimic_query)
# Select the 'full text and metadata' box
radiobutton = driver.find_element_by_id('Search_All_Text')
radiobutton.click()
# Search
time.sleep(1)
searchbutton = driver.find_element_by_class_name('stats-Adv_Command_search')
searchbutton.click()
# Get the total number of pages
#npages = int(driver.find_element_by_id('pageCount.top').text)
# Get the titles!!!
while True:
# let the page load
time.sleep(2)
# Get scroll height
last_height = driver.execute_script("return document.body.scrollHeight")
while True:
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
time.sleep(0.5)
new_height = driver.execute_script("return document.body.scrollHeight")
if new_height == last_height:
break
last_height = new_height
# Get the titles.
# They are in: <h2 class="result-item-title"><a class="ng-binding ng-scope">title</a></h2>
elements = driver.find_elements_by_class_name('result-item-title')
for e in elements:
# Text may appear with "[::sometext::]"
all_titles.append(e.find_element_by_tag_name('a').get_attribute('text').replace('[::', '').replace('::]', ''))
# New line separated journal info and such
#all_titles.append(e.text.split('\n')[0])
# Click next page if any
e = driver.find_element_by_class_name('next')
if 'disabled' in e.get_attribute('class'):
print('Got all paper titles!')
driver.close()
break
else:
nextbutton = driver.find_element_by_link_text('>')
nextbutton.click()
all_titles = set(all_titles)
all_titles = [t.lower() for t in list(all_titles)]
print(len(all_titles))
display(all_titles)
# Write the titles to files
write_dir = os.path.join('search_results/ieee')
full_query_file = os.path.join(write_dir, 'with-constraints.txt')
with open(full_query_file, 'w') as f:
for line in all_titles:
f.write(line+'\n')
```
# Combining Results - pubmed, wos, ieee
```
result_dir = 'search_results'
combined_results = []
for service in ['pubmed', 'wos', 'ieee']:
# For pubmed, get the curated true positives from the unconstrained search instead
if service == 'pubmed':
df = pd.read_csv(os.path.join(result_dir, service, 'without-constraints-inspected.tsv'), delimiter='\t', header=None)
service_results = list(df.loc[df[1]=='T'][0].values)
# For other services, get the constrained search results
else:
with open(os.path.join(result_dir, service, 'with-constraints.txt')) as f:
service_results = f.readlines()
print('Number of results from service '+service+': '+str(len(service_results)))
combined_results = combined_results + [r.strip() for r in service_results]
print('\nTotal number of non-unique results: ', len(combined_results))
combined_results = sorted(list(set(combined_results)))
print('Total number of unique results: ', len(combined_results))
with open(os.path.join(result_dir, 'combined', 'with-constraints.txt'), 'w') as f:
for r in combined_results:
f.write(r+'\n')
# may 21 2018
Number of results from service pubmed: 155
Number of results from service wos: 152
Number of results from service ieee: 322
Total number of non-unique results: 629
Total number of unique results: 456
```
# Attempting to parse GS automatically failed. Below is evidence of failure. Can ignore...
# N. Search Google Scholar
Packages found online:
- https://github.com/ckreibich/scholar.py
- https://github.com/venthur/gscholar
- https://github.com/adeel/google-scholar-scraper
- http://code.activestate.com/recipes/523047-search-google-scholar/
- https://github.com/erdiaker/torrequest
- https://github.com/NikolaiT/GoogleScraper
- https://stackoverflow.com/questions/8049520/web-scraping-javascript-page-with-python
Query: `("mimic ii" OR "mimic iii") AND ("database" OR "clinical" OR "waveform" OR ICU)`
https://scholar.google.com/scholar?q=%28mimic-ii+OR+mimic-iii%29&btnG=&hl=en&as_sdt=1%2C22&as_vis=1
https://scholar.google.com/scholar/help.html
https://superuser.com/questions/565722/how-to-config-tor-to-use-a-http-socks-proxy
## Requirements
1. Browse with JS enabled. requests library uses http. Otherwise google will think (correctly) that you are a robot.
2. Change IP every time, or google will block.
```
from scholarly import scholarly
PROJECT = 'eICU'
# Search by Google Scholar publication ID
search_results = []
if PROJECT == 'eICU':
try:
search_results.append(scholarly.search_citedby(11878669525996073977))
except Exception as e:
print(e)
total_results = search_results[0].total_results
# all_pubs = list(search_results)
all_pubs = []
for i in range(total_results):
print(100*i/total_results)
try:
ct = next(search_results)
all_pubs.append({
'title': ct['bib']['title'],
'pub_year': ct['bib']['pub_year']
})
except:
pass
# print(all_pubs)
```
| github_jupyter |
# Plotting Colormaps
You can use colormaps to colour your network plots in order to get a quick and simple overview of line loadings and bus voltages. The plotting module provides functions to easily modify your bus and line collections. It also provides colourbars to match the colourcodes to the according numerical values.
### Continuous Colormaps
First, we load the network and run a loadflow to retrieve results:
```
import pandapower as pp
import pandapower.networks as nw
import pandapower.plotting as plot
%matplotlib inline
net = nw.mv_oberrhein()
pp.runpp(net)
```
The pandapower plotting package contains convenience functions to create common colorbars and norms. We use the cmap_continuous function to get a linear colormap with color centers green at 20%, yellow at 50% and red at 60% line loading:
```
cmap_list=[(20, "green"), (50, "yellow"), (60, "red")]
cmap, norm = plot.cmap_continuous(cmap_list)
```
The colormap and norm are now passend to the create_bus_collection function and the collection is plotted with draw_collections:
```
lc = plot.create_line_collection(net, net.line.index, zorder=1, cmap=cmap, norm=norm, linewidths=2)
plot.draw_collections([lc], figsize=(8,6))
cmap_list=[(0.975, "blue"), (1.0, "green"), (1.03, "red")]
cmap, norm = plot.cmap_continuous(cmap_list)
bc = plot.create_bus_collection(net, net.bus.index, size=80, zorder=2, cmap=cmap, norm=norm)
plot.draw_collections([lc, bc], figsize=(8,6))
```
### Discrete Colormaps
Discrete colormaps can be used in the same way as continuous colormaps using the cmap_voltage_discrete and cmap_loading_discrete functions. For discrete colormaps, each color has to be assigned a range instead of a center:
```
net = nw.mv_oberrhein()
pp.runpp(net)
cmap_list=[((0.975, 0.985), "blue"), ((0.985, 1.0), "green"), ((1.0, 1.03), "red")]
cmap, norm = plot.cmap_discrete(cmap_list)
bc = plot.create_bus_collection(net, net.bus.index, size=80, zorder=2, cmap=cmap, norm=norm)
cmap_list=[((10, 40), "green"), ((40, 55), "yellow"), ((55, 60), "red")]
cmap, norm = plot.cmap_discrete(cmap_list)
lc = plot.create_line_collection(net, net.line.index, zorder=1, cmap=cmap, norm=norm, linewidths=2)
plot.draw_collections([lc, bc], figsize=(8,6))
```
### Logarithmic Colormaps
This option can be used to create logarithmic colormaps. The intermediate values of the logarithmic scale are created automatically based on the minimum an maximum given values in analogy to the LogNorm. The colormap itself has a linear segmentation of the given colors. Also, it can only be used with at least 3 colors an increasing values which have to be above 0.
```
net = nw.mv_oberrhein()
pp.runpp(net)
min_value = 1.0
max_value = 1.03
colors = ["blue", "green", "red"]
cmap, norm = plot.cmap_logarithmic(min_value, max_value, colors)
bc = plot.create_bus_collection(net, size=100, cmap=cmap, norm=norm, zorder=2)
min_value = 10
max_value = 60
colors = ["green", "yellow", "red"]
cmap, norm = plot.cmap_logarithmic(min_value, max_value, colors)
lc = plot.create_line_collection(net, net.line.index, zorder=1, cmap=cmap, norm=norm, linewidth=2)
plot.draw_collections([bc, lc])
```
## Custom Colormaps and Colorbars
The functions to create colormaps and norms are merely convenience functions. You can individually create any colormap you like and pass it to the create_collection functions.
For example, for the colorbar "PuBu_r" from matplotlib:
```
from matplotlib.pyplot import get_cmap
from matplotlib.colors import Normalize
cmap = get_cmap('PuBu_r')
lc = plot.create_line_collection(net, net.line.index, zorder=1, color="grey", linewidths=2,
cmap=cmap)
bc = plot.create_bus_collection(net, net.bus.index, size=80, zorder=2)
plot.draw_collections([lc, bc], figsize=(8,6))
```
Plotting without a norm maps the colorbar to the range of the data points (here: line loadings). Normalizing to values between 20 and 100 yields:
```
cmap = get_cmap('PuBu_r')
norm = Normalize(vmin=20, vmax=100)
lc = plot.create_line_collection(net, net.line.index, zorder=1, color="grey", linewidths=2,
cmap=cmap, norm=norm)
bc = plot.create_bus_collection(net, net.bus.index, size=80, zorder=2)
plot.draw_collections([lc, bc], figsize=(8,6))
```
The colorbar can be customized by disabling the automatic plotting of the colorbar in draw_collections and plotting the colorbar directly with the desired parameters:
```
from matplotlib.pyplot import colorbar
plot.draw_collections([lc, bc], figsize=(8,6), plot_colorbars=False)
cbar = colorbar(lc, extend="max")
cbar.set_ticks([50, 70, 100])
cbar.ax.set_ylabel("This is a individual colorbar title")
```
| github_jupyter |
# Generalized Latency Performance Estimation for Once-For-All Neural Architecture Search
## High Level Overview
<img src="latency_predictor/nas_overview.png" width=800 height=800 />
## Architecture
<img src="latency_predictor/lnas_architecture.png" width=900 height=900 />
In this notebook, we will be demonstrating:
- Latency predictor to predict latency of a given model architecture
- Generalization of the Latency predictor over different hardware
- Effects of using Latency predictors in Neural Architecture Search paradigm (OFA)
## Initialization
Initialize OFA Code:
```
import torch
import numpy as np
import time
import random
import os
from collections import *
from accuracy_predictor import AccuracyPredictor
from latency_table import LatencyTable
from evolution_finder import EvolutionFinder
import csv
import pandas as pd
import sys
sys.path.append("..")
from ofa.model_zoo import ofa_net
import matplotlib.pyplot as plt
# set random seed
random_seed = 10291284
random.seed(random_seed)
np.random.seed(random_seed)
torch.manual_seed(random_seed)
print('Successfully imported all packages and configured random seed to %d!'%random_seed)
cuda_available = torch.cuda.is_available()
if cuda_available:
torch.backends.cudnn.enabled = True
torch.backends.cudnn.benchmark = True
torch.cuda.manual_seed(random_seed)
print('Using GPU.')
else:
print('Using CPU.')
ofa_network = ofa_net('ofa_mbv3_d234_e346_k357_w1.0', pretrained=True)
print('The OFA Network is ready.')
data_loader = None
# Accuracy Predictor
accuracy_predictor = AccuracyPredictor(
pretrained=True,
device='cuda:0' if cuda_available else 'cpu'
)
print('The accuracy predictor is ready!')
print(accuracy_predictor.model)
# Latency Predictor
target_hardware = 'note10'
latency_table = LatencyTable(device=target_hardware, use_latency_table=True)
print('The Latency lookup table on %s is ready!' % target_hardware)
""" Hyper-parameters for the evolutionary search process
You can modify these hyper-parameters to see how they influence the final ImageNet accuracy of the search sub-net.
"""
latency_constraint = 25 # ms, suggested range [15, 33] ms
P = 100 # The size of population in each generation
N = 500 # How many generations of population to be searched
r = 0.25 # The ratio of networks that are used as parents for next generation
params = {
'constraint_type': target_hardware, # Let's do FLOPs-constrained search
'efficiency_constraint': latency_constraint,
'mutate_prob': 0.1, # The probability of mutation in evolutionary search
'mutation_ratio': 0.5, # The ratio of networks that are generated through mutation in generation n >= 2.
'efficiency_predictor': latency_table, # To use a predefined efficiency predictor.
'accuracy_predictor': accuracy_predictor, # To use a predefined accuracy_predictor predictor.
'population_size': P,
'max_time_budget': N,
'parent_ratio': r,
'arch': 'compofa', ## change
}
# build the evolution finder
finder = EvolutionFinder(**params)
```
## Dataset Creation
Create dataset:
```
# time to create dataset
start = time.time()
# create latency dataset
number_of_datapoints = 10
latency_dataset = finder.create_latency_dataset(search_space='compofa', num_of_samples=number_of_datapoints)
# create dataset csv file
curr_hardware = 'note10_lookuptable' #'note10_lookuptable' #'Intel_Xeon_CPU' #RTX_2080_Ti_GPU'
filename = 'latency_predictor/datasets/' + curr_hardware + '_fixedkernelcompofa_test.csv'
with open(filename, 'w') as csv_file:
w = csv.writer(csv_file)
w.writerow(['child_arch', 'latency'])
for i in range(len(latency_dataset['child_arch'])):
child_arch = latency_dataset['child_arch'][i]
latency = latency_dataset['latency'][i]
w.writerow([child_arch, latency])
end = time.time()
print('Wrote Latency Dataset to File: {}'.format(filename))
print('Time to Create Dataset of {} points: {}'.format(number_of_datapoints, end-start))
df = pd.read_csv('latency_predictor/datasets/note10_lookuptable_fixedkernelcompofa_test.csv', usecols=["child_arch", "latency"])
df
```
## Latency Prediction Model
<img src="latency_predictor/Latency_predictor_model.png" width=400 height=400 />
```
if '/latency_predictor' not in os.getcwd():
os.chdir('./latency_predictor')
from latency_predictor import LatencyPredictor, RMSELoss, data_preprocessing, dataset_creation
```
### Evaluate Note10 OFA Model
```
from latency_encoding import latency_encoding
note10_checkpt = torch.load('../checkpoints/latency_prediction_model/Individual_trained_models/ofa/checkpoint_note10_ofa.pt')
note10_model = LatencyPredictor()
note10_model.load_state_dict(note10_checkpt)
criterion = RMSELoss()
#print(note10_model.model)
```
Load Note10 Dataset
```
dataset_path = 'datasets/Note10_LookupTable/note10_lookuptable_ofa.csv'#note10_lookuptable_elastickernelcompofa.csv'
df = pd.read_csv(dataset_path, usecols=["child_arch", "latency"])
```
Lets look at the data
```
df.head()
df.iloc[0]['child_arch']
print("One hot encoding of the child architecture:")
latency_encoding(eval(df.iloc[0]['child_arch']))
print("Latency based on Look up Table:", df.iloc[0]['latency'], "ms")
print("Latency based on Latency predictor:",note10_model.predict_efficiency(eval(df.iloc[0]['child_arch'])), "ms")
training_data, validation_data, test_data = data_preprocessing(dataset_path, 5000)
#test_data.head()
```
Vectoring child architecture via one hot encoding:
```
train_loader, validation_loader, test_dataset = dataset_creation(training_data, validation_data, test_data)
print("size of the test data set:", len(test_dataset))
test_loss = []
for input_arch, target in test_dataset:
prediction = note10_model(input_arch)[0][0]
loss = criterion(prediction, target)
test_loss.append(loss)
print("RMSE Loss over test data set:", torch.mean(torch.tensor(test_loss)), "ms")
```
### Evaluate RTX_2080_Ti_GPU OFA Model
```
RTX_checkpt = torch.load('../checkpoints/latency_prediction_model/Individual_trained_models/ofa/checkpoint_RTX_2080_Ti_GPU_ofa.pt')
RTX_model = LatencyPredictor()
RTX_model.load_state_dict(RTX_checkpt)
criterion = RMSELoss()
#print(RTX_model.model)
```
Load RTX Dataset
```
dataset_path = 'datasets/RTX_2080_Ti_GPU/RTX_2080_Ti_GPU_ofa.csv'
df_RTX = pd.read_csv(dataset_path, usecols=["child_arch", "latency"])
#df_RTX.head()
df_RTX.iloc[0]['child_arch']
sample_child_arch = eval(df_RTX.iloc[0]['child_arch'])
sample_latency = df_RTX.iloc[0]['latency']
print("Latency from look up table:", sample_latency, "ms")
print("Latency from latency predictor: ", RTX_model.predict_efficiency(sample_child_arch), "ms")
training_data, validation_data, test_data = data_preprocessing(dataset_path, 5000)
train_loader, validation_loader, test_dataset = dataset_creation(training_data, validation_data, test_data)
#len(test_dataset)
test_loss = []
for input_arch, target in test_dataset:
prediction = RTX_model(input_arch)[0][0]
loss = criterion(prediction, target)
test_loss.append(loss)
print("RMSE loss over test dataset: ", torch.mean(torch.tensor(test_loss)), "ms")
```
### Evaluate Finetuned RTX_2080_Ti_GPU Model using Note10 Base
```
RTX_checkpt = torch.load('../checkpoints/latency_prediction_model/Finetune_Note_10/checkpoint_finetune_RTX_2080_Ti_GPU_ofa.pt')
finetune=True
if finetune:
key_names = ["model.0.weight", "model.0.bias", "model.2.weight", "model.2.bias", "model.4.weight", "model.4.bias", "model.6.weight", "model.6.bias"]
new_state_dict = OrderedDict()
i = 0
for k, v in RTX_checkpt.items():
new_state_dict[key_names[i]] = v
i = i + 1
RTX_checkpt = new_state_dict
RTX_model = LatencyPredictor()
RTX_model.load_state_dict(RTX_checkpt)
criterion = RMSELoss()
dataset_path = 'datasets/RTX_2080_Ti_GPU/RTX_2080_Ti_GPU_ofa.csv'
df_RTX = pd.read_csv(dataset_path, usecols=["child_arch", "latency"])
#df_RTX.head()
print(df_RTX.iloc[0]['child_arch'])
sample_child_arch = eval(df_RTX.iloc[0]['child_arch'])
sample_latency = df_RTX.iloc[0]['latency']
print("Latency based on look up table", sample_latency, "ms")
print("Latency based on fine-tuned latency predictor:", RTX_model.predict_efficiency(sample_child_arch), "ms")
training_data, validation_data, test_data = data_preprocessing(dataset_path, 5000)
train_loader, validation_loader, test_dataset = dataset_creation(training_data, validation_data, test_data)
#len(test_dataset)
test_loss = []
for input_arch, target in test_dataset:
prediction = RTX_model(input_arch)[0][0]
loss = criterion(prediction, target)
test_loss.append(loss)
print("RMSE loss over test dataset:", torch.mean(torch.tensor(test_loss)), "ms")
```
# Inference Time Analysis
```
from latency_encoding import latency_encoding
sample_child_arch = eval(df.iloc[0]['child_arch'])
sample_latency = df.iloc[0]['latency']
print('Child_Arch: {}, \n\nLatency: {}'.format(sample_child_arch, sample_latency))
sample_child_arch_encoded = latency_encoding(sample_child_arch)
sample_child_arch_encoded
```
Estimation Time using Latency Prediction Model:
With encoding and memoization:
```
times = []
for i in range(1):
start = time.time()
prediction = RTX_model.predict_efficiency(sample_child_arch)
end = time.time()
model_time = end - start
times.append(model_time)
#print(model_time)
print(sum(times)/len(times))
```
Estimation Time using Note10 Lookup Table (Previous Method):
```
times = []
for i in range(1):
start = time.time()
prediction = finder.efficiency_predictor.predict_efficiency(sample_child_arch)
end = time.time()
lookup_time = end - start
times.append(lookup_time)
print(sum(times)/len(times))
```
# Neural Architecture Search
Using Note10 Lookup Table as the Performance Estimation
```
""" Hyper-parameters for the evolutionary search process
You can modify these hyper-parameters to see how they influence the final ImageNet accuracy of the search sub-net.
"""
latency_constraint = 25 # ms, suggested range [15, 33] ms
P = 100 # The size of population in each generation
N = 500 # How many generations of population to be searched
r = 0.25 # The ratio of networks that are used as parents for next generation
target_hardware = 'note10'
params = {
'constraint_type': target_hardware, # Let's do FLOPs-constrained search
'efficiency_constraint': latency_constraint,
'mutate_prob': 0.1, # The probability of mutation in evolutionary search
'mutation_ratio': 0.5, # The ratio of networks that are generated through mutation in generation n >= 2.
'efficiency_predictor': latency_table, # To use a predefined efficiency predictor.
'accuracy_predictor': accuracy_predictor, # To use a predefined accuracy_predictor predictor.
'population_size': P,
'max_time_budget': N,
'parent_ratio': r,
'arch': 'compofa', ## change
}
# build the evolution finder
finder = EvolutionFinder(**params)
result_lis1 = []
latency_constraint = 20
st = time.time()
finder.set_efficiency_constraint(latency_constraint)
best_valids, best_info = finder.run_evolution_search()
ed = time.time()
result_lis1.append(best_info)
print("Done! Time Taken: {} seconds".format(ed-st))
result_lis1
```
Using Latency Prediction Model as the Performance Estimation
```
""" Hyper-parameters for the evolutionary search process
You can modify these hyper-parameters to see how they influence the final ImageNet accuracy of the search sub-net.
"""
latency_constraint = 25 # ms, suggested range [15, 33] ms
P = 100 # The size of population in each generation
N = 500 # How many generations of population to be searched
r = 0.25 # The ratio of networks that are used as parents for next generation
target_hardware = 'note10'
params = {
'constraint_type': target_hardware, # Let's do FLOPs-constrained search
'efficiency_constraint': latency_constraint,
'mutate_prob': 0.1, # The probability of mutation in evolutionary search
'mutation_ratio': 0.5, # The ratio of networks that are generated through mutation in generation n >= 2.
'efficiency_predictor': note10_model, # To use a predefined efficiency predictor.
'accuracy_predictor': accuracy_predictor, # To use a predefined accuracy_predictor predictor.
'population_size': P,
'max_time_budget': N,
'parent_ratio': r,
'arch': 'compofa', ## change
}
# build the evolution finder
finder = EvolutionFinder(**params)
result_lis2 = []
latency_constraint = 20
st = time.time()
finder.set_efficiency_constraint(latency_constraint)
best_valids, best_info = finder.run_evolution_search()
ed = time.time()
result_lis2.append(best_info)
print("Done! Time Taken: {} seconds".format(ed-st))
result_lis2
accuracy_predictor.predict_accuracy([result_lis1[0][1], result_lis2[0][1]])
```
### Application in NAS Experiments
#### Comparison of pareto frontier of NAS (look up) vs NAS (latency predictor)
#### Note10
Search Space: Fixed Kernel CompOFA
5 constraints: 15, 20, 25, 30, 35
Target Hardware: Note10
Note10 Lookup Table Results:
Top1 Accuracy:
[72.80000114440918, 75.20000076293945, 75.70000076293945, 75.80000305175781, 75.9000015258789]
Top5 Accuracy:
[91.1, 92.1, 93.2, 93.3, 93.5]
Latency:
[14.69766178609321, 19.88688251867145, 24.83390000854306, 29.512724161788654, 34.79207886302561]
Time for NAS: 90.72894263267517
Note10_OFA Model Latency Prediction Results:
Top1 Accuracy:
[73.40000343322754, 76.0, 75.30000305175781, 76.10000038146973, 76.00000190734863]
Top5 Accuracy:
[91.4, 92.2, 92.8, 93.3, 93.3]
Latency:
[14.7567, 18.6498, 24.8855, 29.4332, 32.4759]
Time for NAS: 175.0260510444641
Note10_OFA Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[75.80000114440918, 76.80000114440918, 76.80000305175781, 77.30000114440918, 77.20000267028809]
Top5 Accuracy:
[92.7, 93.1, 93.8, 94.5, 94.4]
Latency:
[14.7567, 18.6498, 24.8855, 29.4332, 32.4759]
Time for NAS: 59.26314926147461
```
lookup_table_top1 = [72.8, 75.2, 75.7, 75.8, 75.9]
lookup_table_top5 = [91.1, 92.1, 93.2, 93.3, 93.5]
lookup_table_latency = [14.69, 19.88, 24.83, 29.51, 34.79]
model_top1 = [73.4, 76.0, 75.3, 76.1, 76.0]
model_top5 = [91.4, 92.2, 92.8, 93.3, 93.3]
model_latency = [14.75, 18.64, 24.88, 29.43, 32.47]
model_memo_top1 = [75.8, 76.8, 76.8, 77.3, 77.2]
model_memo_top5 = [92.7, 93.1, 93.8, 94.5, 94.4]
model_memo_latency = [14.75, 18.64, 24.88, 29.43, 32.47]
plt.figure(figsize=(4,4))
plt.plot(lookup_table_latency, lookup_table_top1, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Lookup_Table')
plt.plot(model_memo_latency, model_memo_top1, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='Model_Memo')
plt.plot(model_latency, model_top1, '--', marker='+', linewidth=2, markersize=8, label='Model')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-1 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
plt.figure(figsize=(4,4))
plt.plot(lookup_table_latency, lookup_table_top5, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Lookup_Table')
plt.plot(model_memo_latency, model_memo_top5, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='Model_Memo')
plt.plot(model_latency, model_top5, '--', marker='+', linewidth=2, markersize=8, label='Model')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-5 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
```
#### Note10
Search Space: Elastic Kernel CompOFA
5 constraints: 15, 20, 25, 30, 35
Target Hardware: Note10
Note10 Lookup Table Results:
Top1 Accuracy:
[73.30000305175781, 75.80000114440918, 75.70000267028809, 76.30000305175781, 76.4000015258789]
Top5 Accuracy:
[90.2, 91.9, 93.0, 93.2, 93.8]
Latency:
[14.883577352241502, 19.9517505771064, 24.523907686903634, 29.628799671866894, 34.987063535187986]
Time for NAS: 105.40677523612976
Note10_OFA Model Latency Prediction Results:
Top1 Accuracy:
[73.4000015258789, 75.80000114440918, 75.30000114440918, 77.70000076293945, 76.60000228881836]
Top5 Accuracy:
[91.3, 92.6, 93.3, 93.4, 93.7]
Latency:
[14.845640182495117, 19.97205352783203, 24.33823585510254, 29.964521408081055, 34.266754150390625]
Time for NAS: 223.74719524383545
Note10_OFA Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[76.00000190734863, 76.40000343322754, 76.80000114440918, 77.4000015258789, 77.30000305175781]
Top5 Accuracy:
[92.7, 93.6, 94.3, 94.9, 94.6]
Latency:
[14.730718612670898, 19.913232803344727, 23.94869613647461, 29.688018798828125, 31.26772689819336]
Time for NAS: 173.9402687549591
```
lookup_table_top1 = [73.3, 75.8, 75.7, 76.3, 76.4]
lookup_table_top5 = [90.2, 91.9, 93.0, 93.2, 93.8]
lookup_table_latency = [14.88, 19.95, 24.52, 29.62, 34.98]
model_top1 = [73.4, 75.8, 75.3, 77.7, 76.6]
model_top5 = [91.3, 92.6, 93.3, 93.4, 93.7]
model_latency = [14.84, 19.97, 24.33, 29.96, 34.26]
model_memo_top1 = [76.0, 76.4, 76.8, 77.4, 77.3]
model_memo_top5 = [92.7, 93.6, 94.3, 94.9, 94.6]
model_memo_latency = [14.73, 19.91, 23.94, 29.68, 31.26]
plt.figure(figsize=(4,4))
plt.plot(lookup_table_latency, lookup_table_top1, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Lookup_Table')
plt.plot(model_memo_latency, model_memo_top1, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='Model_Memo')
plt.plot(model_latency, model_top1, '--', marker='+', linewidth=2, markersize=8, label='Model')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-1 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
plt.figure(figsize=(4,4))
plt.plot(lookup_table_latency, lookup_table_top5, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Lookup_Table')
plt.plot(model_memo_latency, model_memo_top5, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='Model_Memo')
plt.plot(model_latency, model_top5, '--', marker='+', linewidth=2, markersize=8, label='Model')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-5 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
```
#### RTX 2080 Ti GPU
Search Space: CompOFA
5 constraints: 15, 20, 25, 30, 35
Target Hardware: gpu
Latency Measurement Results:
Top1 Accuracy:
[76.9000015258789, 76.70000076293945, 76.79999923706055, 77.20000267028809, 76.80000305175781]
Top5 Accuracy:
[94.2, 94.2, 94.5, 94.4, 94.2]
Latency:
[14.194464683532715, 17.401909828186035, 17.401909828186035, 17.401909828186035, 33.16640853881836]
Time for NAS: 202.69879007339478
RTX_OFA Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[77.39999961853027, 77.00000190734863, 77.4000015258789, 77.00000190734863, 76.5]
Top5 Accuracy:
[94.0, 94.2, 94.2, 94.3, 94.5]
Latency:
[13.162400245666504, 15.776034355163574, 15.776034355163574, 15.776034355163574, 32.576904296875]
Time for NAS: 57.54925608634949
RTX_OFA Note10_FineTuned Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[76.20000076293945, 77.70000267028809, 77.00000190734863, 76.70000076293945, 77.20000267028809]
Top5 Accuracy:
[94.2, 94.4, 94.2, 94.3, 94.2]
Latency:
[14.683911323547363, 15.258056640625, 15.258056640625, 29.110185623168945, 31.35794448852539]
Time for NAS: 57.574804067611694
```
measure_top1 = [76.90, 76.70, 76.79, 77.20, 76.80]
measure_top5 = [94.2, 94.2, 94.5, 94.4, 94.2]
measure_latency = [14.19, 17.40, 17.40, 17.40, 33.16]
model_top1 = [77.39, 77.00, 77.40, 77.00, 76.50]
model_top5 = [94.0, 94.2, 94.2, 94.3, 94.5]
model_latency = [13.16, 15.77, 15.77, 15.77, 32.57]
finetuned_top1 = [76.20, 77.70, 77.00, 76.70, 77.20]
finetuned_top5 = [94.2, 94.4, 94.2, 94.3, 94.2]
finetuned_latency = [14.68, 15.25, 15.25, 29.11, 31.35]
target_hardware = 'RTX_2080_Ti_GPU'
plt.figure(figsize=(4,4))
plt.plot(measure_latency, measure_top1, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Latency_Measure')
plt.plot(model_latency, model_top1, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='RTX_Model')
plt.plot(finetuned_latency, finetuned_top1, '--', marker='+', linewidth=2, markersize=8, label='Finetuned')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-1 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
target_hardware = 'RTX_2080_Ti_GPU'
plt.figure(figsize=(4,4))
plt.plot(measure_latency, measure_top5, 'x-', marker='o', color='red', linewidth=2, markersize=8, label='Latency_Measure')
plt.plot(model_latency, model_top5, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='RTX_Model')
plt.plot(finetuned_latency, finetuned_top5, '--', marker='+', linewidth=2, markersize=8, label='Finetuned')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-5 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
```
#### RTX 2080 Ti GPU
#### Comparison of Finetuned Model with Hardware specific model
Search Space: Elastic Kernel CompOFA
5 constraints: 15, 20, 25, 30, 35
Target Hardware: gpu
RTX_OFA Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[77.10000228881836, 77.40000343322754, 77.30000114440918, 77.20000076293945, 77.90000343322754]
Top5 Accuracy:
[94.5, 94.8, 94.8, 95.0, 94.7]
Latency:
[14.033023834228516, 17.6427001953125, 24.21096420288086, 22.906673431396484, 23.46289825439453]
Time for NAS: 147.10737991333008
RTX_OFA Note10_FineTuned Model Latency Prediction WITH Memoization Results:
Top1 Accuracy:
[76.70000076293945, 78.10000038146973, 77.10000038146973, 78.20000076293945, 76.80000114440918]
Top5 Accuracy:
[93.8, 94.5, 94.6, 94.6, 94.8]
Latency:
[14.805492401123047, 19.535118103027344, 22.045217514038086, 26.886178970336914, 34.85415267944336]
Time for NAS: 139.161794424057
```
model_top1 = [77.10, 77.40, 77.30, 77.20, 77.90]
model_top5 = [94.5, 94.8, 94.8, 95.0, 94.7]
model_latency = [14.03, 17.64, 24.21, 22.90, 23.46]
finetuned_top1 = [76.70, 78.10, 77.10, 78.20, 76.80]
finetuned_top5 = [93.8, 94.5, 94.6, 94.6, 94.8]
finetuned_latency = [14.80, 19.53, 22.04, 26.88, 34.85]
target_hardware = 'RTX_2080_Ti_GPU'
plt.figure(figsize=(4,4))
plt.plot(model_latency, model_top1, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='RTX_Model')
plt.plot(finetuned_latency, finetuned_top1, '--', marker='+', linewidth=2, markersize=8, label='Finetuned')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-1 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
target_hardware = 'RTX_2080_Ti_GPU'
plt.figure(figsize=(4,4))
plt.plot(model_latency, model_top5, 'x-', marker='*', color='darkred', linewidth=2, markersize=8, label='RTX_Model')
plt.plot(finetuned_latency, finetuned_top5, '--', marker='+', linewidth=2, markersize=8, label='Finetuned')
plt.xlabel('%s Latency (ms)' % target_hardware, size=12)
plt.ylabel('ImageNet Top-5 Accuracy (%)', size=12)
plt.legend(loc='lower right')
plt.grid(True)
plt.show()
```
| github_jupyter |
# Pandas basics
Hi! In this programming assignment you need to refresh your `pandas` knowledge. You will need to do several [`groupby`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html)s and [`join`]()`s to solve the task.
```
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
%matplotlib inline
DATA_FOLDER = './data/'
transactions = pd.read_csv(os.path.join(DATA_FOLDER, 'sales_train.csv.gz'))
items = pd.read_csv(os.path.join(DATA_FOLDER, 'items.csv'))
item_categories = pd.read_csv(os.path.join(DATA_FOLDER, 'item_categories.csv'))
shops = pd.read_csv(os.path.join(DATA_FOLDER, 'shops.csv'))
```
The dataset we are going to use is taken from the competition, that serves as the final project for this course. You can find complete data description at the [competition web page](https://www.kaggle.com/c/competitive-data-science-final-project/data). To join the competition use [this link](https://www.kaggle.com/t/1ea93815dca248e99221df42ebde3540).
# Task
Let's start with a simple task.
<ol start="0">
<li><b>Print the shape of the loaded dataframes and use [`df.head`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.head.html) function to print several rows. Examine the features you are given.</b></li>
</ol>
```
print(f"Transaction shape: {transactions.shape}, head is:")
transactions.head()
print(f"Items shape: {items.shape}, head is:")
items.head()
print(f"Item categories shape: {item_categories.shape}, head is:")
item_categories.head()
print(f"Shops shape: {shops.shape}, head is:")
shops.head()
```
Now use your `pandas` skills to get answers for the following questions.
The first question is:
1. ** What was the maximum total revenue among all the shops in September, 2014?**
* Hereinafter *revenue* refers to total sales minus value of goods returned.
*Hints:*
* Sometimes items are returned, find such examples in the dataset.
* It is handy to split `date` field into [`day`, `month`, `year`] components and use `df.year == 14` and `df.month == 9` in order to select target subset of dates.
* You may work with `date` feature as with strings, or you may first convert it to `pd.datetime` type with `pd.to_datetime` function, but do not forget to set correct `format` argument.
```
transactions['item_value'] = transactions.item_price * transactions.item_cnt_day
transactions.datetime = pd.to_datetime(transactions.date, format="%d.%m.%Y")
mask = (transactions.datetime >= '2014-09-01') & (transactions.datetime <= '2014-09-30')
max_revenue = transactions.loc[mask].groupby("shop_id").item_value.sum().max()
max_revenue
```
Great! Let's move on and answer another question:
<ol start="2">
<li><b>What item category generated the highest revenue in summer 2014?</b></li>
</ol>
* Submit `id` of the category found.
* Here we call "summer" the period from June to August.
*Hints:*
* Note, that for an object `x` of type `pd.Series`: `x.argmax()` returns **index** of the maximum element. `pd.Series` can have non-trivial index (not `[1, 2, 3, ... ]`).
```
transactions.datetime = pd.to_datetime(transactions.date, format="%d.%m.%Y")
mask = (transactions.datetime >= '2014-06-01') & (transactions.datetime <= '2014-08-31')
june_august_transactions = transactions.loc[mask]
joined_df = june_august_transactions[["item_id", "item_value"]] \
.join(items.set_index("item_id"), on="item_id", how='left')
revenue_per_cat = joined_df.groupby(by="item_category_id")['item_value'].sum().reset_index()
category_id_with_max_revenue = revenue_per_cat.loc[revenue_per_cat['item_value'].argmax()].item_category_id
category_id_with_max_revenue
```
<ol start="3">
<li><b>How many items are there, such that their price stays constant (to the best of our knowledge) during the whole period of time?</b></li>
</ol>
* Let's assume, that the items are returned for the same price as they had been sold.
```
unique_price = transactions.groupby("item_id")["item_price"].nunique()
num_items_constant_price = unique_price.loc[unique_price == 1].count()
num_items_constant_price
```
Remember, the data can sometimes be noisy.
<ol start="4">
<li><b>What was the variance of the number of sold items per day sequence for the shop with `shop_id = 25` in December, 2014? Do not count the items, that were sold but returned back later.</b></li>
</ol>
* Fill `total_num_items_sold` and `days` arrays, and plot the sequence with the code below.
* Then compute variance. Remember, there can be differences in how you normalize variance (biased or unbiased estimate, see [link](https://math.stackexchange.com/questions/496627/the-difference-between-unbiased-biased-estimator-variance)). Compute ***unbiased*** estimate (use the right value for `ddof` argument in `pd.var` or `np.var`).
* If there were no sales at a given day, ***do not*** impute missing value with zero, just ignore that day
```
from datetime import datetime
shop_id = 25
transactions['datetime'] = pd.to_datetime(transactions.date, format="%d.%m.%Y")
date_mask = (transactions.datetime >= '2014-12-01') & (transactions.datetime <= '2014-12-31')
shop_mask = transactions.shop_id == shop_id
december_shop_trans = transactions.loc[date_mask & shop_mask]
#sold_items = december_shop_trans.loc[december_shop_trans["item_cnt_day"] > 0]
#returned_items = december_shop_trans.loc[december_shop_trans["item_cnt_day"] < 0]
#joined_df = sold_items.join(returned_items.set_index(["datetime", "item_id"])["item_cnt_day"],
# on=["datetime","item_id"], how="left",
# lsuffix="_sold", rsuffix="_returned")
#joined_df["total_cnt"] = joined_df["item_cnt_day_sold"] + joined_df["item_cnt_day_returned"].fillna(0)
#item_sold_per_day = joined_df.reset_index().groupby(by="datetime")["total_cnt"].sum()
total_num_items_sold = december_shop_trans.groupby("datetime")["item_cnt_day"].sum()
days = [d.astype(datetime) for d in total_num_items_sold .index.values]
# Plot it
plt.plot(days, total_num_items_sold)
plt.ylabel('Num items')
plt.xlabel('Day')
plt.title("Daily revenue for shop_id = 25")
plt.show()
total_num_items_sold_var = np.var(total_num_items_sold.values, ddof=1)
total_num_items_sold_var
```
Well done! :)
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
import seaborn as sn
import math
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.optimizers import Adam
from sklearn.preprocessing import MinMaxScaler, normalize
from functools import reduce
import random
from numpy.linalg import norm
import os
val_errors1 = []
test_errors1 = []
SEED = 1000
random.seed(SEED)
np.random.seed(SEED)
tf.random.set_seed(SEED)
os.environ['PYTHONHASHSEED'] = str(SEED)
df = pd.read_csv('Data\\X_train_raw.csv').T
df_valid = pd.read_csv('Data\\X_valid_raw.csv').T
df_test = pd.concat((pd.read_csv('Data\\X_test_raw_A.txt').T, pd.read_csv('Data\\X_test_raw_B.txt').T)).iloc[[0, 1, 2, 4, 5], :]
rach_clusters = pd.read_csv('Data\\X_train_clusters.csv')
Y_data = df.iloc[1:, -1].astype('float64')
Y_copy = Y_data
Y_valid_data = df_valid.iloc[1:, -1].astype('float64')
Y_valid_copy = Y_valid_data
common_IDs = reduce(np.intersect1d, (df.iloc[0, :-1].values, df_valid.iloc[0, :-1].values, df_test.iloc[0, :].values))
idx = np.where(df.iloc[0, :].isin(common_IDs))[0]
df = df.iloc[:, idx]
idx_valid = np.where(df_valid.iloc[0, :].isin(common_IDs))[0]
df_valid = df_valid.iloc[:, idx_valid]
idx_test = np.where(df_test.iloc[0, :].isin(common_IDs))[0]
df_test = df_test.iloc[:, idx_test]
X_data = df.iloc[1:, :].astype('float64')
X_ID = df.iloc[0, :]
X_valid_data = df_valid.iloc[1:, :].astype('float64')
X_valid_ID = df_valid.iloc[0, :]
X_test_data = df_test.iloc[1:, :].astype('float64')
X_test_ID = df_test.iloc[0, :]
X_ID1 = np.argsort(X_ID)
X_ID = X_ID.iloc[X_ID1]
X_data = X_data.iloc[:, X_ID1]
X_data.columns = X_ID
X_ID1 = np.argsort(X_valid_ID)
X_valid_ID = X_valid_ID.iloc[X_ID1]
X_valid_data = X_valid_data.iloc[:, X_ID1]
X_valid_data.columns = X_valid_ID
X_ID1 = np.argsort(X_test_ID)
X_test_ID = X_test_ID.iloc[X_ID1]
X_test_data = X_test_data.iloc[:, X_ID1]
X_test_data.columns = X_test_ID
genes = ['AT1G13650.1',
'AT3G55450.1',
'AT1G02930.2',
'AT1G79500.3',
'AT5G24850.1',
'AT5G06870.1',
'AT5G41460.1',
'AT5G01820.1',
'AT4G08870.1',
'AT1G75100.1',
'AT2G29650.2',
'AT5G06690.1',
'AT3G17609.2',
'AT4G15690.1',
'AT1G06040.1'
]
X_data = X_data.loc[:, genes]
X_valid_data = X_valid_data.loc[:, genes]
X_test_data = X_test_data.loc[:, genes]
n_folds = Y_data.shape[0]
folds = KFold(n_splits=n_folds, random_state=SEED, shuffle=True)
y_cos = -np.cos((2 * np.pi * Y_data.astype('float64') / 24)+(np.pi/2))
y_sin = np.sin((2 * np.pi * Y_data.astype('float64') / 24)+(np.pi/2))
Y_valid_cos = -np.cos((2 * np.pi * Y_valid_data.astype('float64') / 24)+(np.pi/2))
Y_valid_sin = np.sin((2 * np.pi * Y_valid_data.astype('float64') / 24)+(np.pi/2))
scaler = MinMaxScaler()
scaler.fit(X_data)
X_data = scaler.transform(X_data)
X_valid_data = scaler.transform(X_valid_data)
X_test_data = scaler.transform(X_test_data)
def cyclical_loss(y_true, y_pred):
error = 0
for i in range(y_pred.shape[0]):
error += np.arccos((y_true[i, :] @ y_pred[i, :]) / (norm(y_true[i, :]) * norm(y_pred[i, :])))
return error
def custom_loss(y_true, y_pred):
return tf.reduce_mean((tf.math.acos(tf.matmul(y_true, tf.transpose(y_pred)) / ((tf.norm(y_true) * tf.norm(y_pred)) + tf.keras.backend.epsilon()))**2))
adam = Adam(lr=0.00001, beta_1=0.9, beta_2=0.999, amsgrad=False)
def larger_model():
# create model
model = Sequential()
model.add(Dense(32, kernel_initializer='normal', activation='relu'))
model.add(Dense(128, kernel_initializer='normal', activation='relu'))
model.add(Dense(512, kernel_initializer='normal', activation='relu'))
model.add(Dense(2, kernel_initializer='normal'))
# Compile model
model.compile(loss=custom_loss, optimizer=adam)
return model
Y_data = np.concatenate((y_cos.values.reshape(-1, 1), y_sin.values.reshape(-1, 1)), axis=1)
Y_valid_data = np.concatenate((Y_valid_cos.values.reshape(-1, 1), Y_valid_sin.values.reshape(-1, 1)), axis=1)
error = 0 # Initialise error
all_preds = np.zeros((Y_data.shape[0], 2)) # Create empty array
all_valid_preds = np.zeros((Y_valid_data.shape[0], 2)) # Create empty array
early_stop = EarlyStopping(patience=50, restore_best_weights=True, monitor='val_loss', mode='min')
def reset_seeds(reset_graph_with_backend=None, seed=0):
if reset_graph_with_backend is not None:
K = reset_graph_with_backend
K.clear_session()
tf.compat.v1.reset_default_graph()
np.random.seed(seed)
random.seed(seed)
tf.compat.v1.set_random_seed(seed)
valid_preds = []
test_preds = []
for n_fold, (train_idx, valid_idx) in enumerate(folds.split(X_data, Y_data)):
X_train, Y_train = X_data[train_idx], Y_data[train_idx] # Define training data for this iteration
X_valid, Y_valid = X_data[valid_idx], Y_data[valid_idx]
model = larger_model()
model.fit(X_train.astype('float64'), Y_train.astype('float64'), validation_data=(X_valid.astype('float64'), Y_valid.astype('float64')),
batch_size=1, epochs=5000, callbacks=[early_stop]) # Fit the model on the training data
preds = normalize(model.predict(X_valid)) # Predict on the validation data
all_preds[valid_idx] = normalize(model.predict(X_valid))
all_valid_preds += (normalize(model.predict(X_valid_data)) / n_folds)
valid_preds.append(normalize(model.predict(X_valid_data)))
test_preds.append(normalize(model.predict(X_test_data)))
error += cyclical_loss(Y_valid.astype('float64'), preds.astype('float64')) # Evaluate the predictions
print(cyclical_loss(Y_valid.astype('float64'), preds.astype('float64')) / Y_valid.shape[0])
angles = []
for i in range(all_preds.shape[0]):
angles.append(math.atan2(all_preds[i, 0], all_preds[i, 1]) / math.pi * 12)
for j in range(len(angles)):
if angles[j] < 0:
angles[j] = angles[j] + 24
ax = sn.scatterplot(Y_data[:, 0], Y_data[:, 1])
ax = sn.scatterplot(all_preds[:, 0], all_preds[:, 1])
plt.show()
angles_arr = np.vstack(angles)
hour_pred = angles_arr
plt.figure(dpi=500)
ax = sn.lineplot(np.arange(Y_copy.shape[0]), Y_copy)
ax = sn.lineplot(np.arange(Y_copy.shape[0]), angles_arr.ravel())
plt.show()
angles = []
for i in range(all_preds.shape[0]):
angles.append(math.atan2(all_preds[i, 0], all_preds[i, 1]) / math.pi * 12)
for j in range(len(angles)):
if angles[j] < 0:
angles[j] = angles[j] + 24
valid_angles = []
valid_preds = np.mean(valid_preds, axis=0)
for i in range(valid_preds.shape[0]):
valid_angles.append(math.atan2(valid_preds[i, 0], valid_preds[i, 1]) / math.pi * 12)
for j in range(len(valid_angles)):
if valid_angles[j] < 0:
valid_angles[j] = valid_angles[j] + 24
valid_preds = normalize(valid_preds)
ax = sn.scatterplot(Y_valid_data[:, 0], Y_valid_data[:, 1])
ax = sn.scatterplot(valid_preds[:, 0], valid_preds[:, 1])
plt.show()
angles_arr_valid = np.vstack(valid_angles)
hour_pred_valid = angles_arr_valid
plt.figure(dpi=500)
ax = sn.lineplot(np.arange(Y_valid_copy.shape[0]), Y_valid_copy)
ax = sn.lineplot(np.arange(Y_valid_copy.shape[0]), angles_arr_valid.ravel())
plt.show()
print("Average training error = {} minutes".format(60 * 12 * cyclical_loss(Y_data.astype('float64'), all_preds.astype('float64')) / (Y_data.shape[0] * np.pi)))
print("Average validation error = {} minutes".format(60 * 12 * cyclical_loss(Y_valid_data.astype('float64'), valid_preds.astype('float64')) / (Y_valid_data.shape[0] * np.pi)))
Y_copy1 = np.array([2, 5, 8, 11, 14, 17, 20, 23, 2, 5, 8, 11, 14, 17, 20, 23])
test_angles = []
test_preds_copy = test_preds
test_preds = np.mean(test_preds, axis=0)
for j in range(len(test_preds_copy)):
for i in range(test_preds.shape[0]):
test_preds_copy[j][i, 0] = math.atan2(test_preds_copy[j][i, 0], test_preds_copy[j][i, 1]) / math.pi * 12
if test_preds_copy[j][i, 0] < 0:
test_preds_copy[j][i, 0] += 24
test_preds_copy[j] = np.delete(test_preds_copy[j], 1, 1)
for i in range(test_preds.shape[0]):
test_angles.append(math.atan2(test_preds[i, 0], test_preds[i, 1]) / math.pi * 12)
for j in range(len(test_angles)):
if test_angles[j] < 0:
test_angles[j] = test_angles[j] + 24
test_preds = normalize(test_preds)
angles_arr_test = np.vstack(test_angles)
hour_pred_test = angles_arr_test
Y_test = np.array([12, 0, 12, 0])
Y_test_cos = -np.cos((2 * np.pi * Y_test.astype('float64') / 24) + (np.pi / 2))
Y_test_sin = np.sin((2 * np.pi * Y_test.astype('float64') / 24) + (np.pi / 2))
Y_test_ang = np.concatenate((Y_test_cos.reshape(-1, 1), Y_test_sin.reshape(-1, 1)), axis=1)
print("Average test error = {} minutes".format(60 * 12 * cyclical_loss(Y_test_ang.astype('float64'), test_preds.astype('float64')) / (Y_test_ang.shape[0] * np.pi)))
val_errors1.append(60 * 12 * cyclical_loss(Y_valid_data.astype('float64'), all_valid_preds.astype('float64')) / (Y_valid_data.shape[0] * np.pi))
test_errors1.append(60 * 12 * cyclical_loss(Y_test_ang.astype('float64'), test_preds.astype('float64')) / (Y_test_ang.shape[0] * np.pi))
model.save('ArabidopsisTrainedModel')
```
| github_jupyter |
```
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
```
В папке subsid (shared/homeworks/python_ds_miniprojects/5_subsid) находятся файлы (tm_sales_1, tm_sales_2, ...) с продажами продуктов через телемаркетинг. Каждый файл содержит, как минимум, 4 колонки (поля): FILIAL_ID, SUBS_ID, PROD_ID, ACT_DTTM.
Суть задачи в том, чтобы проверить подключения продуктов определенным пользователям, соединив файлы о продажах с логами по подключениям в системе.
Особенности данных:
сотрудники телемаркетинга не всегда указывают полный id, если 'id' нет в начале SUBS_ID, то нужно его добавить
поля в файлах могут быть расположены абсолютно случайным образом, но названия полей статичны
продажа не засчитывается, если отключение (END_DTTM) произошло меньше чем через 5 минут после подключения (START_DTTM)
если в файле с продажами встречается строка без указанного SUBS_ID, она пропускается
Сохраните результат в датафрэйм с разделителем ;, содержащий корректные подключения.
Note: обратите внимание на то, как pandas переводит дату из строки, возможно вам понадобится параметр format
Пример содержания итогового файла (колонка difference это разница между START_DTTM и END_DTTM):
```
prod_activations_logs = pd.read_csv('/home/jupyter-l.fedoseeva-12/Lesson_5/Dop_zadanie/prod_activations_logs.csv', sep=';')
tm_sales_1 = pd.read_csv('/home/jupyter-l.fedoseeva-12/Lesson_5/Dop_zadanie/tm_sales_1.csv', sep=';')
tm_sales_2 = pd.read_csv('/home/jupyter-l.fedoseeva-12/Lesson_5/Dop_zadanie/tm_sales_2.csv', sep=';')
tm_sales_3 = pd.read_csv('/home/jupyter-l.fedoseeva-12/Lesson_5/Dop_zadanie/tm_sales_3.csv', sep=';')
tm_sales_1.dtypes
tm_sales_2
prod_activations_logs.dtypes
tm_sales_2
tm_sales_3
tm_sales_1
all_tm_sales_1 = pd.concat([tm_sales_1, tm_sales_2, tm_sales_3])
all_tm_sales_1 = all_tm_sales_1.reset_index(drop=True)
all_tm_sales_1 = all_tm_sales_1.dropna()
all_tm_sales_1
def result(cell):
if cell.startswith('id'):
return cell
return 'id{}'.format(cell)
all_tm_sales_1['SUBS_ID'] = all_tm_sales_1.SUBS_ID.apply(result)
all_tm_sales_1
all_tm_sales_1 = all_tm_sales_1.merge(prod_activations_logs, how='inner', on=['SUBS_ID', 'PROD_ID'])
all_tm_sales_1
all_tm_sales_1.START_DTTM = pd.to_datetime(all_tm_sales_1['START_DTTM'], format='%d-%m-%Y %H:%M')
all_tm_sales_1.END_DTTM = pd.to_datetime(all_tm_sales_1['END_DTTM'], format='%d-%m-%Y %H:%M')
all_tm_sales_1['raznitsa'] = all_tm_sales_1.END_DTTM - all_tm_sales_1.START_DTTM
all_tm_sales_1
greater_than_5min = all_tm_sales_1['raznitsa'] > '5m'
greater_than_5min
final_data = all_tm_sales_1.loc[greater_than_5min]
final_data
final_data.SUBS_ID.sort_values().str.cat(sep=', ')
final_data[['FILIAL_ID', 'ACT_DTTM', 'raznitsa']]
final_data[['ACT_DTTM', 'FILIAL_ID', 'raznitsa']] \
.to_csv('final_data_correction.csv', sep=';', index=False)
```
| github_jupyter |
# Fine-Tuning
:label:`sec_fine_tuning`
In earlier chapters, we discussed how to train models on the Fashion-MNIST training dataset with only 60000 images. We also described ImageNet, the most widely used large-scale image dataset in academia, which has more than 10 million images and 1000 objects. However, the size of the dataset that we usually encounter is between those of the two datasets.
Suppose that we want to recognize different types of chairs from images, and then recommend purchase links to users.
One possible method is to first identify
100 common chairs,
take 1000 images of different angles for each chair,
and then train a classification model on the collected image dataset.
Although this chair dataset may be larger than the Fashion-MNIST dataset,
the number of examples is still less than one-tenth of
that in ImageNet.
This may lead to overfitting of complicated models
that are suitable for ImageNet on this chair dataset.
Besides, due to the limited amount of training examples,
the accuracy of the trained model
may not meet practical requirements.
In order to address the above problems,
an obvious solution is to collect more data.
However, collecting and labeling data can take a lot of time and money.
For example, in order to collect the ImageNet dataset, researchers have spent millions of dollars from research funding.
Although the current data collection cost has been significantly reduced, this cost still cannot be ignored.
Another solution is to apply *transfer learning* to transfer the knowledge learned from the *source dataset* to the *target dataset*.
For example, although most of the images in the ImageNet dataset have nothing to do with chairs, the model trained on this dataset may extract more general image features, which can help identify edges, textures, shapes, and object composition.
These similar features may
also be effective for recognizing chairs.
## Steps
In this section, we will introduce a common technique in transfer learning: *fine-tuning*. As shown in :numref:`fig_finetune`, fine-tuning consists of the following four steps:
1. Pretrain a neural network model, i.e., the *source model*, on a source dataset (e.g., the ImageNet dataset).
1. Create a new neural network model, i.e., the *target model*. This copies all model designs and their parameters on the source model except the output layer. We assume that these model parameters contain the knowledge learned from the source dataset and this knowledge will also be applicable to the target dataset. We also assume that the output layer of the source model is closely related to the labels of the source dataset; thus it is not used in the target model.
1. Add an output layer to the target model, whose number of outputs is the number of categories in the target dataset. Then randomly initialize the model parameters of this layer.
1. Train the target model on the target dataset, such as a chair dataset. The output layer will be trained from scratch, while the parameters of all the other layers are fine-tuned based on the parameters of the source model.

:label:`fig_finetune`
When target datasets are much smaller than source datasets, fine-tuning helps to improve models' generalization ability.
## Hot Dog Recognition
Let us demonstrate fine-tuning via a concrete case:
hot dog recognition.
We will fine-tune a ResNet model on a small dataset,
which was pretrained on the ImageNet dataset.
This small dataset consists of
thousands of images with and without hot dogs.
We will use the fine-tuned model to recognize
hot dogs from images.
```
%matplotlib inline
import os
from mxnet import gluon, init, np, npx
from mxnet.gluon import nn
from d2l import mxnet as d2l
npx.set_np()
```
### Reading the Dataset
[**The hot dog dataset we use was taken from online images**].
This dataset consists of
1400 positive-class images containing hot dogs,
and as many negative-class images containing other foods.
1000 images of both classes are used for training and the rest are for testing.
After unzipping the downloaded dataset,
we obtain two folders `hotdog/train` and `hotdog/test`. Both folders have `hotdog` and `not-hotdog` subfolders, either of which contains images of
the corresponding class.
```
#@save
d2l.DATA_HUB['hotdog'] = (d2l.DATA_URL + 'hotdog.zip',
'fba480ffa8aa7e0febbb511d181409f899b9baa5')
data_dir = d2l.download_extract('hotdog')
```
We create two instances to read all the image files in the training and testing datasets, respectively.
```
train_imgs = gluon.data.vision.ImageFolderDataset(
os.path.join(data_dir, 'train'))
test_imgs = gluon.data.vision.ImageFolderDataset(
os.path.join(data_dir, 'test'))
```
The first 8 positive examples and the last 8 negative images are shown below. As you can see, [**the images vary in size and aspect ratio**].
```
hotdogs = [train_imgs[i][0] for i in range(8)]
not_hotdogs = [train_imgs[-i - 1][0] for i in range(8)]
d2l.show_images(hotdogs + not_hotdogs, 2, 8, scale=1.4);
```
During training, we first crop a random area of random size and random aspect ratio from the image,
and then scale this area
to a $224 \times 224$ input image.
During testing, we scale both the height and width of an image to 256 pixels, and then crop a central $224 \times 224$ area as input.
In addition,
for the three RGB (red, green, and blue) color channels
we *standardize* their values channel by channel.
Concretely,
the mean value of a channel is subtracted from each value of that channel and then the result is divided by the standard deviation of that channel.
[~~Data augmentations~~]
```
# Specify the means and standard deviations of the three RGB channels to
# standardize each channel
normalize = gluon.data.vision.transforms.Normalize(
[0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
train_augs = gluon.data.vision.transforms.Compose([
gluon.data.vision.transforms.RandomResizedCrop(224),
gluon.data.vision.transforms.RandomFlipLeftRight(),
gluon.data.vision.transforms.ToTensor(),
normalize])
test_augs = gluon.data.vision.transforms.Compose([
gluon.data.vision.transforms.Resize(256),
gluon.data.vision.transforms.CenterCrop(224),
gluon.data.vision.transforms.ToTensor(),
normalize])
```
### [**Defining and Initializing the Model**]
We use ResNet-18, which was pretrained on the ImageNet dataset, as the source model. Here, we specify `pretrained=True` to automatically download the pretrained model parameters.
If this model is used for the first time,
Internet connection is required for download.
```
pretrained_net = gluon.model_zoo.vision.resnet18_v2(pretrained=True)
```
The pretrained source model instance contains two member variables: `features` and `output`. The former contains all layers of the model except the output layer, and the latter is the output layer of the model.
The main purpose of this division is to facilitate the fine-tuning of model parameters of all layers but the output layer. The member variable `output` of source model is shown below.
```
pretrained_net.output
```
As a fully-connected layer, it transforms ResNet's final global average pooling outputs into 1000 class outputs of the ImageNet dataset.
We then construct a new neural network as the target model. It is defined in the same way as the pretrained source model except that
its number of outputs in the final layer
is set to
the number of classes in the target dataset (rather than 1000).
In the following code, the model parameters in the member variable features of the target model instance finetune_net are initialized to the model parameters of the corresponding layer of the source model. Since the model parameters in the features are pre-trained on the ImageNet data set and are good enough, generally only a small learning rate is needed to fine-tune these parameters.
The model parameters in the member variable output are initialized randomly, and generally require a larger learning rate to train from scratch. Assuming that the learning rate in the Trainer instance is η, we set the learning rate of the model parameters in the member variable output to be 10η in the iteration.
In the code below, the model parameters before the output layer of the target model instance `finetune_net` are initialized to model parameters of the corresponding layers from the source model.
Since these model parameters were obtained via pretraining on ImageNet,
they are effective.
Therefore, we can only use
a small learning rate to *fine-tune* such pretrained parameters.
In contrast, model parameters in the output layer are randomly initialized and generally require a larger learning rate to be learned from scratch.
Let the base learning rate be $\eta$, a learning rate of $10\eta$ will be used to iterate the model parameters in the output layer.
```
finetune_net = gluon.model_zoo.vision.resnet18_v2(classes=2)
finetune_net.features = pretrained_net.features
finetune_net.output.initialize(init.Xavier())
# The model parameters in the output layer will be iterated using a learning
# rate ten times greater
finetune_net.output.collect_params().setattr('lr_mult', 10)
```
### [**Fine-Tuning the Model**]
First, we define a training function `train_fine_tuning` that uses fine-tuning so it can be called multiple times.
```
def train_fine_tuning(net, learning_rate, batch_size=128, num_epochs=5):
train_iter = gluon.data.DataLoader(
train_imgs.transform_first(train_augs), batch_size, shuffle=True)
test_iter = gluon.data.DataLoader(
test_imgs.transform_first(test_augs), batch_size)
devices = d2l.try_all_gpus()
net.collect_params().reset_ctx(devices)
net.hybridize()
loss = gluon.loss.SoftmaxCrossEntropyLoss()
trainer = gluon.Trainer(net.collect_params(), 'sgd', {
'learning_rate': learning_rate, 'wd': 0.001})
d2l.train_ch13(net, train_iter, test_iter, loss, trainer, num_epochs,
devices)
```
We [**set the base learning rate to a small value**]
in order to *fine-tune* the model parameters obtained via pretraining. Based on the previous settings, we will train the output layer parameters of the target model from scratch using a learning rate ten times greater.
```
train_fine_tuning(finetune_net, 0.01)
```
[**For comparison,**] we define an identical model, but (**initialize all of its model parameters to random values**). Since the entire model needs to be trained from scratch, we can use a larger learning rate.
```
scratch_net = gluon.model_zoo.vision.resnet18_v2(classes=2)
scratch_net.initialize(init=init.Xavier())
train_fine_tuning(scratch_net, 0.1)
```
As we can see, the fine-tuned model tends to perform better for the same epoch
because its initial parameter values are more effective.
## Summary
* Transfer learning transfers knowledge learned from the source dataset to the target dataset. Fine-tuning is a common technique for transfer learning.
* The target model copies all model designs with their parameters from the source model except the output layer, and fine-tunes these parameters based on the target dataset. In contrast, the output layer of the target model needs to be trained from scratch.
* Generally, fine-tuning parameters uses a smaller learning rate, while training the output layer from scratch can use a larger learning rate.
## Exercises
1. Keep increasing the learning rate of `finetune_net`. How does the accuracy of the model change?
2. Further adjust hyperparameters of `finetune_net` and `scratch_net` in the comparative experiment. Do they still differ in accuracy?
3. Set the parameters before the output layer of `finetune_net` to those of the source model and do *not* update them during training. How does the accuracy of the model change? You can use the following code.
```
finetune_net.features.collect_params().setattr('grad_req', 'null')
```
4. In fact, there is a "hotdog" class in the `ImageNet` dataset. Its corresponding weight parameter in the output layer can be obtained via the following code. How can we leverage this weight parameter?
```
weight = pretrained_net.output.weight
hotdog_w = np.split(weight.data(), 1000, axis=0)[713]
hotdog_w.shape
```
[Discussions](https://discuss.d2l.ai/t/368)
| github_jupyter |
```
%pdb
%matplotlib inline
import os
os.chdir('..')
import tensorflow as tf
import numpy as np
import pandas as pd
EAGER=False
RUN_TESTS=False
PRINT_ALPHABET=False
if EAGER:
tf.enable_eager_execution()
SAMPLING_RATE=16000
def to_path(filename):
return './data/' + filename
def random_stretch(audio, params):
"""
Stretches randomly the input audio
"""
rate = random.uniform(params['random_stretch_min'], params['random_stretch_max'])
return librosa.effects.time_stretch(audio, rate)
import random
def random_shift(audio, params):
"""
Shifts randomly the input audio to the left or the right
"""
_shift = random.randrange(params['random_shift_min'], params['random_shift_max'])
if _shift < 0:
pad = (_shift * -1, 0)
else:
pad = (0, _shift)
return np.pad(audio, pad, mode='constant')
import glob
noise_files = glob.glob('./data/*.wav')
noises = {}
def random_noise(audio, params):
_factor = random.uniform(
params['random_noise_factor_min'],
params['random_noise_factor_max']
)
if params['random_noise'] > random.uniform(0, 1):
_path = random.choice(noise_files)
if _path in noises:
wave = noises[_path]
else:
if os.path.isfile(_path + '.wave.hkl'):
wave = hkl.load(_path + '.wave.hkl').astype(np.float32)
noises[_path] = wave
else:
wave, _ = librosa.load(_path, sr=SAMPLING_RATE)
hkl.dump(wave, _path + '.wave.hkl')
noises[_path] = wave
noise = random_shift(
wave,
{
'random_shift_min': -16000,
'random_shift_max': 16000
}
)
max_noise = np.max(noise[0:len(audio)])
max_wave = np.max(audio)
noise = noise * (max_wave / max_noise)
return _factor * noise[0:len(audio)] + (1.0 - _factor) * audio
else:
return audio
import librosa
import hickle as hkl
import os.path
def load_wave(example, absolute=False):
row, params = example
_path = row.filename if absolute else to_path(row.filename)
if os.path.isfile(_path + '.wave.hkl'):
wave = hkl.load(_path + '.wave.hkl').astype(np.float32)
else:
wave, _ = librosa.load(_path, sr=SAMPLING_RATE)
hkl.dump(wave, _path + '.wave.hkl')
if len(wave) <= params['max_wave_length']:
if params['augment'] and row.filename.split('/')[0] != 'voxforge':
wave = random_noise(
random_stretch(
random_shift(
wave,
params
),
params
),
params
)
else:
wave = None
return wave, row
#from IPython.display import Audio
#
#params = {
# 'augment': True,
# 'random_shift_min': -4000,
# 'random_shift_max': 4000,
# 'random_noise': 1.0,
# 'random_noise_factor_min': 0.1,
# 'random_noise_factor_max': 0.15,
# 'random_stretch_min': 0.8,
# 'random_stretch_max': 1.2
#}
#
#_, row = next(train_data.iterrows())
#
#wave, _ = load_wave((row, params))
#
#Audio(data=wave, rate=16000)
#import os.path
#import glob
#
#with_length_filename = 'with_lengths.csv'
#
#if not os.path.isfile(with_length_filename):
# train_eval_data = pd.read_csv('./data/cv_corpus_v1/cv-valid-train.csv')
#
# def get_len(row):
# wave, _ = load_wave(row)
#
# return len(wave)
#
# lengths = [
# get_len(row)
# for _, row in train_eval_data.iterrows()
# ]
#
# hkl.dump(lengths, 'lengths.hkl')
#
# train_eval_data['length'] = lengths
#
# train_eval_data.to_csv('with_lengths.csv')
#else:
# train_eval_data = pd.read_csv(with_length_filename)
train_eval_data = pd.read_csv('./data/cv_corpus_v1/cv-valid-train.csv')
#train_eval_data = train_eval_data[train_eval_data.length <= 80000]
if not os.path.isfile('train.csv'):
eval_data = train_eval_data.sample(n=int(len(train_eval_data) * 0.1 ))
train_data = train_eval_data[~train_eval_data.isin(eval_data)]
train_data = train_data[train_data.filename.notnull()]
train_data.to_csv('train.csv')
eval_data.to_csv('eval.csv')
else:
train_data = pd.read_csv('train.csv')
eval_data = pd.read_csv('eval.csv')
#import seaborn as sb
#
#lens = []
#libri_count = 0
#
#for _, row in libri.iterrows():
# wave, _ = load_wave((row, { 'augment': False, 'max_wave_length': 320000 }))
# if wave is not None:
# libri_count += 1
# lens.append(len(wave))
#
#print(libri_count)
#sb.distplot(lens, kde = False)
#import seaborn as sb
#
#lens = []
#vox_count = 0
#
#for _, row in vox.iterrows():
# wave, _ = load_wave((row, { 'augment': False, 'max_wave_length': 320000 }))
# if wave is not None:
# vox_count += 1
# lens.append(len(wave))
#
#print(vox_count)
#sb.distplot(lens, kde = False)
if not os.path.isfile('libri.csv'):
def libri_texts():
for path in glob.glob('./data/LibriSpeech/**/**/**/*.txt'):
file_prefix = '/'.join(path.split('/')[0:-1])
with open(path, 'r') as file:
lines = file.readlines()
for line in lines:
elements = line.split(' ')
prefix = elements[0]
text = ' '.join(elements[1:]).replace('\n', '').lower()
yield '{}/{}.flac'.format(file_prefix.replace('./data/', ''), prefix), text
libri_frame = pd.DataFrame([ (a, b) for a, b in libri_texts() ], columns=['filename', 'text'])
libri_frame.to_csv('libri.csv')
else:
libri_frame = pd.read_csv('libri.csv')
if not os.path.isfile('vox.csv'):
def vox_texts():
for path in glob.glob('./data/voxforge/**/**/PROMPTS'):
file_prefix = '/'.join(path.split('/')[0:-2])
with open(path, 'r') as file:
lines = file.readlines()
if 'wav' in [ s.split('/')[-1] for s in glob.glob('{}/*'.format(file_prefix))]:
form = 'wav'
else:
form = 'flac'
for line in lines:
elements = line.split(' ')
prefix = elements[0].split('/')[-1]
text = ' '.join(elements[1:]).replace('\n', '').lower()
fn = '{}/{}/{}.{}'.format(file_prefix.replace('./data/', ''), form, prefix, form)
#import pdb; pdb.set_trace()
if os.path.isfile('./data/{}'.format(fn)):
yield fn, text
vox_frame = pd.DataFrame([ (a, b) for a, b in vox_texts() ], columns=['filename', 'text'])
vox_frame.to_csv('vox.csv')
else:
vox_frame = pd.read_csv('vox.csv')
test_data = pd.read_csv('./data/cv_corpus_v1/cv-valid-test.csv')
train_data['filename'] = train_data['filename'].apply(lambda f: 'cv_corpus_v1/{}'.format(f))
eval_data['filename'] = eval_data['filename'].apply(lambda f: 'cv_corpus_v1/{}'.format(f))
test_data['filename'] = test_data['filename'].apply(lambda f: 'cv_corpus_v1/{}'.format(f))
if not os.path.isfile('full_train.csv'):
train_data = train_data[['filename', 'text']].append(libri_frame).append(vox_frame)
train_data.to_csv('full_train.csv')
else:
train_data = pd.read_csv('full_train.csv')
train_data
# we need to gather the alphabet to use:
def print_alphabet():
uniques = set()
data_frames = [
train_data,
test_data
]
for ix, data_frame in enumerate(data_frames):
print("Gathering unique characters for set no. " + str(ix + 1) + " (of " + str(len(data_frames)) + " all)")
data_frame.apply(lambda row: [ c for c in row.text ], axis=1).apply(uniques.update)
print(sorted(uniques))
print(''.join(sorted(uniques)))
if PRINT_ALPHABET:
print_alphabet()
print_alphabet()
def compute_lengths(original_lengths, params):
"""
Computes the length of data for CTC
"""
return tf.cast(
tf.floor(
(tf.cast(original_lengths, dtype=tf.float32) - params['n_fft']) /
params['frame_step']
) + 1,
tf.int32
)
def encode_labels(labels, params):
characters = list(params['alphabet'])
table = tf.contrib.lookup.HashTable(
tf.contrib.lookup.KeyValueTensorInitializer(
characters,
list(range(len(characters)))
),
-1,
name='char2id'
)
return table.lookup(
tf.string_split(labels, delimiter='')
)
def decode_codes(codes, params):
characters = list(params['alphabet'])
table = tf.contrib.lookup.HashTable(
tf.contrib.lookup.KeyValueTensorInitializer(
list(range(len(characters))),
characters
),
'',
name='id2char'
)
return table.lookup(codes)
def decode_logits(logits, lengths, params):
if len(tf.shape(lengths).shape) == 1:
lengths = tf.reshape(lengths, [1])
else:
lengths = tf.squeeze(lengths)
predicted_codes, _ = tf.nn.ctc_beam_search_decoder(
tf.transpose(logits, (1, 0, 2)),
lengths,
merge_repeated=True
)
codes = tf.cast(predicted_codes[0], tf.int32)
text = decode_codes(codes, params)
return text, codes
class LogMelSpectrogram(tf.layers.Layer):
def __init__(self,
sampling_rate,
n_fft,
frame_step,
lower_edge_hertz,
upper_edge_hertz,
num_mel_bins,
**kwargs):
super(LogMelSpectrogram, self).__init__(**kwargs)
self.sampling_rate = sampling_rate
self.n_fft = n_fft
self.frame_step = frame_step
self.lower_edge_hertz = lower_edge_hertz
self.upper_edge_hertz = upper_edge_hertz
self.num_mel_bins = num_mel_bins
def call(self, inputs, training=True):
stfts = tf.contrib.signal.stft(
inputs,
frame_length=self.n_fft,
frame_step=self.frame_step,
fft_length=self.n_fft,
pad_end=False
)
power_spectrograms = tf.real(stfts * tf.conj(stfts))
num_spectrogram_bins = power_spectrograms.shape[-1].value
linear_to_mel_weight_matrix = tf.constant(
np.transpose(
librosa.filters.mel(
sr=self.sampling_rate,
n_fft=self.n_fft + 1,
n_mels=self.num_mel_bins,
fmin=self.lower_edge_hertz,
fmax=self.upper_edge_hertz
)
),
dtype=tf.float32
)
mel_spectrograms = tf.tensordot(
power_spectrograms,
linear_to_mel_weight_matrix,
1
)
mel_spectrograms.set_shape(
power_spectrograms.shape[:-1].concatenate(
linear_to_mel_weight_matrix.shape[-1:]
)
)
return tf.log(mel_spectrograms + 1e-6)
class AtrousConv1D(tf.layers.Layer):
def __init__(self,
filters,
kernel_size,
dilation_rate,
use_bias=True,
kernel_initializer=tf.glorot_normal_initializer(),
causal=True
):
super(AtrousConv1D, self).__init__()
self.filters = filters
self.kernel_size = kernel_size
self.dilation_rate = dilation_rate
self.causal = causal
self.conv1d = tf.layers.Conv1D(
filters=filters,
kernel_size=kernel_size,
dilation_rate=dilation_rate,
padding='valid' if causal else 'same',
use_bias=use_bias,
kernel_initializer=kernel_initializer
)
def call(self, inputs):
if self.causal:
padding = (self.kernel_size - 1) * self.dilation_rate
inputs = tf.pad(inputs, tf.constant([(0, 0,), (1, 0), (0, 0)]) * padding)
return self.conv1d(inputs)
class ResidualBlock(tf.layers.Layer):
def __init__(self, filters, kernel_size, dilation_rate, causal, **kwargs):
super(ResidualBlock, self).__init__(**kwargs)
self.dilated_conv1 = AtrousConv1D(
filters=filters,
kernel_size=kernel_size,
dilation_rate=dilation_rate,
causal=causal
)
self.dilated_conv2 = AtrousConv1D(
filters=filters,
kernel_size=kernel_size,
dilation_rate=dilation_rate,
causal=causal
)
self.out = tf.layers.Conv1D(
filters=filters,
kernel_size=1
)
def call(self, inputs, training=True):
data = tf.layers.batch_normalization(
inputs,
training=training
)
filters = self.dilated_conv1(data)
gates = self.dilated_conv2(data)
filters = tf.nn.tanh(filters)
gates = tf.nn.sigmoid(gates)
out = tf.nn.tanh(
self.out(
filters * gates
)
)
return out + inputs, out
class ResidualStack(tf.layers.Layer):
def __init__(self, filters, kernel_size, dilation_rates, causal, **kwargs):
super(ResidualStack, self).__init__(**kwargs)
self.blocks = [
ResidualBlock(
filters=filters,
kernel_size=kernel_size,
dilation_rate=dilation_rate,
causal=causal
)
for dilation_rate in dilation_rates
]
def call(self, inputs, training=True):
data = inputs
skip = 0
for block in self.blocks:
data, current_skip = block(data, training=training)
skip += current_skip
return skip
class SpeechNet(tf.layers.Layer):
def __init__(self, params, **kwargs):
super(SpeechNet, self).__init__(**kwargs)
self.to_log_mel = LogMelSpectrogram(
sampling_rate=params['sampling_rate'],
n_fft=params['n_fft'],
frame_step=params['frame_step'],
lower_edge_hertz=params['lower_edge_hertz'],
upper_edge_hertz=params['upper_edge_hertz'],
num_mel_bins=params['num_mel_bins']
)
self.expand = tf.layers.Conv1D(
filters=params['stack_filters'],
kernel_size=1,
padding='same'
)
self.stacks = [
ResidualStack(
filters=params['stack_filters'],
kernel_size=params['stack_kernel_size'],
dilation_rates=params['stack_dilation_rates'],
causal=params['causal_convolutions']
)
for _ in range(params['stacks'])
]
self.out = tf.layers.Conv1D(
filters=len(params['alphabet']) + 1,
kernel_size=1,
padding='same'
)
def call(self, inputs, training=True):
data = self.to_log_mel(inputs)
data = tf.layers.batch_normalization(
data,
training=training
)
if len(data.shape) == 2:
data = tf.expand_dims(data, 0)
data = self.expand(data)
for stack in self.stacks:
data = stack(data, training=training)
data = tf.layers.batch_normalization(
data,
training=training
)
return self.out(data) + 1e-8
from multiprocessing import Pool
def input_fn(input_dataset, params, load_wave_fn=load_wave):
def _input_fn():
"""
Returns raw audio wave along with the label
"""
dataset = input_dataset
print(params)
if 'max_text_length' in params and params['max_text_length'] is not None:
print('Constraining dataset to the max_text_length')
dataset = input_dataset[input_dataset.text.str.len() < params['max_text_length']]
if 'min_text_length' in params and params['min_text_length'] is not None:
print('Constraining dataset to the min_text_length')
dataset = input_dataset[input_dataset.text.str.len() >= params['min_text_length']]
if 'max_wave_length' in params and params['max_wave_length'] is not None:
print('Constraining dataset to the max_wave_length')
print('Resulting dataset length: {}'.format(len(dataset)))
def generator_fn():
pool = Pool()
buffer = []
for epoch in range(params['epochs']):
if params['shuffle']:
dataset = dataset.sample(frac=1)
else:
dataset = input_dataset
for _, row in dataset.iterrows():
buffer.append((row, params))
if len(buffer) >= params['batch_size']:
if params['parallelize']:
audios = pool.map(
load_wave_fn,
buffer
)
else:
audios = map(
load_wave_fn,
buffer
)
for audio, row in audios:
if audio is not None:
if np.isnan(audio).any():
print('SKIPPING! NaN coming from the pipeline!')
else:
#print(row.text)
yield (audio, len(audio)), row.text.encode()
buffer = []
return tf.data.Dataset.from_generator(
generator_fn,
output_types=((tf.float32, tf.int32), (tf.string)),
output_shapes=((None,()), (()))
) \
.padded_batch(
batch_size=params['batch_size'],
padded_shapes=(
(tf.TensorShape([None]), tf.TensorShape(())),
tf.TensorShape(())
)
)
return _input_fn
def model_fn(features, labels, mode, params):
if isinstance(features, dict):
audio = features['audio']
original_lengths = features['length']
else:
audio, original_lengths = features
lengths = compute_lengths(original_lengths, params)
if labels is not None:
codes = encode_labels(labels, params)
network = SpeechNet(params)
is_training = mode==tf.estimator.ModeKeys.TRAIN
print('Is training? {}'.format(is_training))
logits = network(audio, training=is_training)
text, predicted_codes = decode_logits(logits, lengths, params)
if mode == tf.estimator.ModeKeys.PREDICT:
predictions = {
'logits': logits,
'text': tf.sparse_tensor_to_dense(
text,
''
)
}
export_outputs = {
'predictions': tf.estimator.export.PredictOutput(predictions)
}
return tf.estimator.EstimatorSpec(
mode,
predictions=predictions,
export_outputs=export_outputs
)
else:
loss = tf.reduce_mean(
tf.nn.ctc_loss(
labels=codes,
inputs=logits,
sequence_length=lengths,
time_major=False,
ignore_longer_outputs_than_inputs=True
)
)
mean_edit_distance = tf.reduce_mean(
tf.edit_distance(
tf.cast(predicted_codes, tf.int32),
codes
)
)
distance_metric = tf.metrics.mean(mean_edit_distance)
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(
mode,
loss=loss,
eval_metric_ops={ 'edit_distance': distance_metric }
)
elif mode == tf.estimator.ModeKeys.TRAIN:
global_step = tf.train.get_or_create_global_step()
tf.summary.text(
'train_predicted_text',
tf.sparse_tensor_to_dense(text, '')
)
tf.summary.scalar('train_edit_distance', mean_edit_distance)
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=global_step,
learning_rate=params['lr'],
optimizer=(params['optimizer']),
update_ops=update_ops,
clip_gradients=params['clip_gradients'],
summaries=[
"learning_rate",
"loss",
"global_gradient_norm",
]
)
return tf.estimator.EstimatorSpec(
mode,
loss=loss,
train_op=train_op
)
def experiment_name(params, excluded_keys=['alphabet', 'data', 'lr', 'clip_gradients']):
def represent(key, value):
if key in excluded_keys:
return None
else:
if isinstance(value, list):
return '{}_{}'.format(key, '_'.join([str(v) for v in value]))
else:
return '{}_{}'.format(key, value)
parts = filter(
lambda p: p is not None,
[
represent(k, params[k])
for k in sorted(params.keys())
]
)
return '/'.join(parts)
def dataset_params(batch_size=32,
epochs=50000,
parallelize=True,
max_text_length=None,
min_text_length=None,
max_wave_length=80000,
shuffle=True,
random_shift_min=-4000,
random_shift_max= 4000,
random_stretch_min=0.7,
random_stretch_max= 1.3,
random_noise=0.75,
random_noise_factor_min=0.2,
random_noise_factor_max=0.5,
augment=False):
return {
'parallelize': parallelize,
'shuffle': shuffle,
'max_text_length': max_text_length,
'min_text_length': min_text_length,
'max_wave_length': max_wave_length,
'random_shift_min': random_shift_min,
'random_shift_max': random_shift_max,
'random_stretch_min': random_stretch_min,
'random_stretch_max': random_stretch_max,
'random_noise': random_noise,
'random_noise_factor_min': random_noise_factor_min,
'random_noise_factor_max': random_noise_factor_max,
'epochs': epochs,
'batch_size': batch_size,
'augment': augment
}
def experiment_params(data,
optimizer='Adam',
lr=1e-4,
alphabet=" 'abcdefghijklmnopqrstuvwxyz",
causal_convolutions=True,
stack_dilation_rates= [1, 3, 9, 27, 81],
stacks=2,
stack_kernel_size= 3,
stack_filters= 32,
sampling_rate=16000,
n_fft=160*4,
frame_step=160,
lower_edge_hertz=0,
upper_edge_hertz=8000,
num_mel_bins=160,
clip_gradients=None,
codename='regular',
**kwargs):
params = {
'optimizer': optimizer,
'lr': lr,
'data': data,
'alphabet': alphabet,
'causal_convolutions': causal_convolutions,
'stack_dilation_rates': stack_dilation_rates,
'stacks': stacks,
'stack_kernel_size': stack_kernel_size,
'stack_filters': stack_filters,
'sampling_rate': sampling_rate,
'n_fft': n_fft,
'frame_step': frame_step,
'lower_edge_hertz': lower_edge_hertz,
'upper_edge_hertz': upper_edge_hertz,
'num_mel_bins': num_mel_bins,
'clip_gradients': clip_gradients,
'codename': codename
}
#import pdb; pdb.set_trace()
if kwargs is not None and 'data' in kwargs:
params['data'] = { **params['data'], **kwargs['data'] }
del kwargs['data']
if kwargs is not None:
params = { **params, **kwargs }
return params
import copy
def experiment(data_params=dataset_params(), **kwargs):
params = experiment_params(
data_params,
**kwargs
)
print(params)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir='stats/{}'.format(experiment_name(params)),
params=params
)
#import pdb; pdb.set_trace()
train_spec = tf.estimator.TrainSpec(
input_fn=input_fn(
train_data,
params['data']
)
)
features = {
"audio": tf.placeholder(dtype=tf.float32, shape=[None]),
"length": tf.placeholder(dtype=tf.int32, shape=[])
}
serving_input_receiver_fn = tf.estimator.export.build_raw_serving_input_receiver_fn(
features
)
best_exporter = tf.estimator.BestExporter(
name="best_exporter",
serving_input_receiver_fn=serving_input_receiver_fn,
exports_to_keep=5
)
eval_params = copy.deepcopy(params['data'])
eval_params['augment'] = False
eval_spec = tf.estimator.EvalSpec(
input_fn=input_fn(
eval_data,
eval_params
),
throttle_secs=60*30,
exporters=best_exporter
)
tf.estimator.train_and_evaluate(
estimator,
train_spec,
eval_spec
)
def test(data_params=dataset_params(), **kwargs):
params = experiment_params(
data_params,
**kwargs
)
print(params)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir='stats/{}'.format(experiment_name(params)),
params=params
)
eval_params = copy.deepcopy(params['data'])
eval_params['augment'] = False
eval_params['epochs'] = 1
eval_params['shuffle'] = False
estimator.evaluate(
input_fn=input_fn(
test_data,
eval_params
)
)
def predict_test(**kwargs):
params = experiment_params(
dataset_params(
augment=False,
shuffle=False,
batch_size=1,
epochs=1,
parallelize=False
),
**kwargs
)
print(len(test_data))
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir='stats/{}'.format(experiment_name(params)),
params=params
)
return list(
estimator.predict(
input_fn=input_fn(
test_data,
params['data']
)
)
)
def predict(filepath, **kwargs):
params = experiment_params(
dataset_params(
augment=False,
shuffle=False,
batch_size=1,
epochs=1,
parallelize=False
),
**kwargs
)
dataset = pd.DataFrame(columns=['filename', 'text'])
dataset['filename'] = [filepath]
dataset['text'] = ['']
print(len(dataset))
estimator = tf.estimator.Estimator(
model_fn=model_fn,
model_dir='stats/{}'.format(experiment_name(params)),
params=params
)
return list(
estimator.predict(
input_fn=input_fn(
dataset,
params['data']
)
)
)
def eager_experiment(data_params=dataset_params(), **kwargs):
params = experiment_params(
data_params,
**kwargs
)
print(params)
dataset=input_fn(
train_data,
params['data']
)()
features, labels = dataset.make_one_shot_iterator().get_next()
model_fn(features, labels, tf.estimator.ModeKeys.TRAIN, params)
import unittest
from hypothesis import given, settings, note, assume, reproduce_failure
import hypothesis.strategies as st
import hypothesis.extra.numpy as npst
# assuming test path will look like: 1/file.wav
def dummy_load_wave(example):
row, params = example
path = row.filename
return np.ones((SAMPLING_RATE)) * float(path.split('/')[0]), row
class TestNotebook(unittest.TestCase):
@given(
st.sampled_from([22000, 16000, 8000]),
st.sampled_from([1024, 512]),
st.sampled_from([1024, 512]),
npst.arrays(
np.float32,
(4, 16000),
elements=st.floats(-1, 1)
)
)
@settings(max_examples=5)
def test_log_mel_conversion_works(self, sampling_rate, n_fft, frame_step, audio):
lower_edge_hertz=0.0
upper_edge_hertz=sampling_rate / 2.0
num_mel_bins=64
def librosa_melspectrogram(audio_item):
spectrogram = np.abs(
librosa.core.stft(
audio_item,
n_fft=n_fft,
hop_length=frame_step,
center=False
)
)**2
return np.log(
librosa.feature.melspectrogram(
S=spectrogram,
sr=sampling_rate,
n_mels=num_mel_bins,
fmin=lower_edge_hertz,
fmax=upper_edge_hertz,
) + 1e-6
)
audio_ph = tf.placeholder(tf.float32, (4, 16000))
librosa_log_mels = np.transpose(
np.stack([
librosa_melspectrogram(audio_item)
for audio_item in audio
]),
(0, 2, 1)
)
log_mel_op = tf.check_numerics(
LogMelSpectrogram(
sampling_rate=sampling_rate,
n_fft=n_fft,
frame_step=frame_step,
lower_edge_hertz=lower_edge_hertz,
upper_edge_hertz=upper_edge_hertz,
num_mel_bins=num_mel_bins
)(audio_ph),
message="log mels"
)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
log_mels = session.run(
log_mel_op,
{
audio_ph: audio
}
)
np.testing.assert_allclose(
log_mels,
librosa_log_mels,
rtol=1e-1,
atol=0
)
@given(
npst.arrays(
np.float32,
(4, 16000),
elements=st.floats(-1, 1)
),
st.sampled_from([64, 32]),
st.sampled_from([7, 3]),
st.sampled_from([1, 4]),
)
@settings(max_examples=10)
def test_residual_block_works(self, audio_waves, filters, size, dilation_rate):
with tf.Graph().as_default() as g:
audio_ph = tf.placeholder(tf.float32, (4, None))
log_mel_op = LogMelSpectrogram(
sampling_rate=16000,
n_fft=512,
frame_step=256,
lower_edge_hertz=0,
upper_edge_hertz=8000,
num_mel_bins=10
)(audio_ph)
expanded_op = tf.layers.Dense(filters)(log_mel_op)
_, block_op = ResidualBlock(
filters=filters,
kernel_size=size,
causal=True,
dilation_rate=dilation_rate
)(expanded_op, training=True)
# really dumb loss function just for the sake
# of testing:
loss_op = tf.reduce_sum(block_op)
variables = tf.trainable_variables()
self.assertTrue(any(["batch_normalization" in var.name for var in variables]))
grads_op = tf.gradients(
loss_op,
variables
)
for grad, var in zip(grads_op, variables):
if grad is None:
note(var)
self.assertTrue(grad is not None)
with tf.Session(graph=g) as session:
session.run(tf.global_variables_initializer())
result, expanded, grads, _ = session.run(
[block_op, expanded_op, grads_op, loss_op],
{
audio_ph: audio_waves
}
)
self.assertFalse(np.array_equal(result, expanded))
self.assertEqual(result.shape, expanded.shape)
self.assertEqual(len(grads), len(variables))
self.assertFalse(any([np.isnan(grad).any() for grad in grads]))
@given(
npst.arrays(
np.float32,
(4, 16000),
elements=st.floats(-1, 1)
),
st.sampled_from([64, 32]),
st.sampled_from([7, 3])
)
@settings(max_examples=10)
def test_residual_stack_works(self, audio_waves, filters, size):
dilation_rates = [1,2,4]
with tf.Graph().as_default() as g:
audio_ph = tf.placeholder(tf.float32, (4, None))
log_mel_op = LogMelSpectrogram(
sampling_rate=16000,
n_fft=512,
frame_step=256,
lower_edge_hertz=0,
upper_edge_hertz=8000,
num_mel_bins=10
)(audio_ph)
expanded_op = tf.layers.Dense(filters)(log_mel_op)
stack_op = ResidualStack(
filters=filters,
kernel_size=size,
causal=True,
dilation_rates=dilation_rates
)(expanded_op, training=True)
# really dumb loss function just for the sake
# of testing:
loss_op = tf.reduce_sum(stack_op)
variables = tf.trainable_variables()
self.assertTrue(any(["batch_normalization" in var.name for var in variables]))
grads_op = tf.gradients(
loss_op,
variables
)
for grad, var in zip(grads_op, variables):
if grad is None:
note(var)
self.assertTrue(grad is not None)
with tf.Session(graph=g) as session:
session.run(tf.global_variables_initializer())
result, expanded, grads, _ = session.run(
[stack_op, expanded_op, grads_op, loss_op],
{
audio_ph: audio_waves
}
)
self.assertFalse(np.array_equal(result, expanded))
self.assertEqual(result.shape, expanded.shape)
self.assertEqual(len(grads), len(variables))
self.assertFalse(any([np.isnan(grad).any() for grad in grads]))
def test_causal_conv1d_works(self):
conv_size2_dilation_1 = AtrousConv1D(
filters=1,
kernel_size=2,
dilation_rate=1,
kernel_initializer=tf.ones_initializer(),
use_bias=False
)
conv_size3_dilation_1 = AtrousConv1D(
filters=1,
kernel_size=3,
dilation_rate=1,
kernel_initializer=tf.ones_initializer(),
use_bias=False
)
conv_size2_dilation_2 = AtrousConv1D(
filters=1,
kernel_size=2,
dilation_rate=2,
kernel_initializer=tf.ones_initializer(),
use_bias=False
)
conv_size2_dilation_3 = AtrousConv1D(
filters=1,
kernel_size=2,
dilation_rate=3,
kernel_initializer=tf.ones_initializer(),
use_bias=False
)
data = np.array(list(range(1, 31)))
data_ph = tf.placeholder(tf.float32, (1, 30, 1))
size2_dilation_1_1 = conv_size2_dilation_1(data_ph)
size2_dilation_1_2 = conv_size2_dilation_1(size2_dilation_1_1)
size3_dilation_1_1 = conv_size3_dilation_1(data_ph)
size3_dilation_1_2 = conv_size3_dilation_1(size3_dilation_1_1)
size2_dilation_2_1 = conv_size2_dilation_2(data_ph)
size2_dilation_2_2 = conv_size2_dilation_2(size2_dilation_2_1)
size2_dilation_3_1 = conv_size2_dilation_3(data_ph)
size2_dilation_3_2 = conv_size2_dilation_3(size2_dilation_3_1)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
outputs = session.run(
[
size2_dilation_1_1,
size2_dilation_1_2,
size3_dilation_1_1,
size3_dilation_1_2,
size2_dilation_2_1,
size2_dilation_2_2,
size2_dilation_3_1,
size2_dilation_3_2
],
{
data_ph: np.reshape(data, (1, 30, 1))
}
)
for ix, out in enumerate(outputs):
out = np.squeeze(out)
outputs[ix] = out
self.assertEqual(out.shape[0], len(data))
np.testing.assert_equal(
outputs[0],
np.array([1, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57, 59], dtype=np.float32)
)
np.testing.assert_equal(
outputs[1],
np.array([1, 4, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48, 52, 56, 60, 64, 68, 72, 76, 80, 84, 88, 92, 96, 100, 104, 108, 112, 116], dtype=np.float32)
)
np.testing.assert_equal(
outputs[2],
np.array([1, 3, 6, 9, 12, 15, 18, 21, 24, 27, 30, 33, 36, 39, 42, 45, 48, 51, 54, 57, 60, 63, 66, 69, 72, 75, 78, 81, 84, 87], dtype=np.float32)
)
np.testing.assert_equal(
outputs[3],
np.array([1, 4, 10, 18, 27, 36, 45, 54, 63, 72, 81, 90, 99, 108, 117, 126, 135, 144, 153, 162, 171, 180, 189, 198, 207, 216, 225, 234, 243, 252], dtype=np.float32)
)
np.testing.assert_equal(
outputs[4],
np.array([1, 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50, 52, 54, 56, 58], dtype=np.float32)
)
np.testing.assert_equal(
outputs[5],
np.array([1, 2, 5, 8, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48, 52, 56, 60, 64, 68, 72, 76, 80, 84, 88, 92, 96, 100, 104, 108, 112], dtype=np.float32)
)
np.testing.assert_equal(
outputs[6],
np.array([1, 2, 3, 5, 7, 9, 11, 13, 15, 17, 19, 21, 23, 25, 27, 29, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53, 55, 57], dtype=np.float32)
)
np.testing.assert_equal(
outputs[7],
np.array([1, 2, 3, 6, 9, 12, 16, 20, 24, 28, 32, 36, 40, 44, 48, 52, 56, 60, 64, 68, 72, 76, 80, 84, 88, 92, 96, 100, 104, 108], dtype=np.float32)
)
@given(
npst.arrays(
np.float32,
(4, 30, len('abcdefghijk1234!@#$%^&*')),
elements=st.floats(0, 1)
)
)
@settings(max_examples=10)
def test_decode_logits_doesnt_crash(self, logits):
params = { 'alphabet': 'abcdefghijk1234!@#$%^&*' }
lengths = np.array([15, 15, 15, 15], dtype=np.int32)
logits_ph = tf.placeholder(
tf.float32,
shape=(4, 30, len(params['alphabet']))
)
lengths_ph = tf.placeholder(
tf.int32,
shape=(4)
)
decode_op, codes_op = decode_logits(
logits_ph,
lengths_ph,
params
)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
session.run(tf.tables_initializer(name='init_all_tables'))
codes, decoded = session.run(
[codes_op, decode_op],
{
logits_ph: logits,
lengths_ph: lengths
}
)
results = np.ones(codes.dense_shape) * -1
for ix, value in zip(codes.indices, codes.values):
results[ix[0], ix[1]] = value
for row in results:
self.assertLessEqual(len(row[row != -1]), 15)
@given(st.text(alphabet="!\"&',-.01234:;\abcdefghijklmnopqrstuvwxyz", max_size=10))
@settings(max_examples=10)
def test_encode_and_decode_work(self, text):
assume(text != '')
params = { 'alphabet': '!"&\',-.01234:;\abcdefghijklmnopqrstuvwxyz' }
label_ph = tf.placeholder(tf.string, shape=(1), name='text')
codes_op = encode_labels(label_ph, params)
decode_op = decode_codes(codes_op, params)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
session.run(tf.tables_initializer(name='init_all_tables'))
codes, decoded = session.run(
[codes_op, decode_op],
{
label_ph: np.array([text])
}
)
note(codes)
note(decoded)
self.assertEqual(text, ''.join(map(lambda s: s.decode('UTF-8'), decoded.values)))
self.assertEqual(codes.values.dtype, np.int32)
self.assertEqual(len(codes.values), len(text))
@given(
npst.arrays(
np.float32,
(st.integers(min_value=16000, max_value=16000*5)),
elements=st.floats(-1, 1)
),
st.sampled_from([22000, 16000, 8000]),
st.sampled_from([1024, 512, 640]),
st.sampled_from([1024, 512, 160]),
)
@settings(max_examples=10)
def test_compute_lengths_works(self,
audio_wave,
sampling_rate,
n_fft,
frame_step
):
assume(n_fft >= frame_step)
original_wave_length = audio_wave.shape[0]
audio_waves_ph = tf.placeholder(tf.float32, (None, None), name="audio_waves")
original_lengths_ph = tf.placeholder(tf.int32, (None), name="original_lengths")
lengths_op = compute_lengths(
original_lengths_ph,
{
'frame_step': frame_step,
'n_fft': n_fft
}
)
self.assertEqual(lengths_op.dtype, tf.int32)
log_mel_op = LogMelSpectrogram(
sampling_rate=sampling_rate,
n_fft=n_fft,
frame_step=frame_step,
lower_edge_hertz=0.0,
upper_edge_hertz=8000.0,
num_mel_bins=13
)(audio_waves_ph)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
lengths, log_mels = session.run(
[lengths_op, log_mel_op],
{
audio_waves_ph: np.array([audio_wave]),
original_lengths_ph: np.array([original_wave_length])
}
)
note(original_wave_length)
note(lengths)
note(log_mels.shape)
self.assertEqual(lengths[0], log_mels.shape[1])
@given(
npst.arrays(
np.float32,
(4, 16000),
elements=st.floats(-1, 1)
)
)
@settings(max_examples=10)
def test_speech_net_works(self, audio_waves):
with tf.Graph().as_default() as g:
audio_ph = tf.placeholder(tf.float32, (4, None))
logits_op = SpeechNet(
experiment_params(
{},
stack_dilation_rates= [1, 2, 4],
stack_kernel_size= 3,
stack_filters= 32,
alphabet= 'abcd'
)
)(audio_ph)
# really dumb loss function just for the sake
# of testing:
loss_op = tf.reduce_sum(logits_op)
variables = tf.trainable_variables()
self.assertTrue(any(["batch_normalization" in var.name for var in variables]))
grads_op = tf.gradients(
loss_op,
variables
)
for grad, var in zip(grads_op, variables):
if grad is None:
note(var)
self.assertTrue(grad is not None)
with tf.Session(graph=g) as session:
session.run(tf.global_variables_initializer())
result, grads, _ = session.run(
[logits_op, grads_op, loss_op],
{
audio_ph: audio_waves
}
)
self.assertEqual(result.shape[2], 5)
self.assertEqual(len(grads), len(variables))
self.assertFalse(any([np.isnan(grad).any() for grad in grads]))
@given(
npst.arrays(
np.float32,
(4, 16000),
elements=st.floats(-1, 1)
)
)
@settings(max_examples=10)
def test_batch_normalization_when_not_training_doesnt_have_gradients(self, audio_waves):
with tf.Graph().as_default() as g:
audio_ph = tf.placeholder(tf.float32, (4, None))
logits_op = SpeechNet(
experiment_params(
{},
stack_dilation_rates= [1, 2, 4],
stack_kernel_size= 3,
stack_filters= 32,
alphabet= 'abcd'
)
)(audio_ph, training=False)
# really dumb loss function just for the sake
# of testing:
loss_op = tf.reduce_sum(logits_op)
variables = tf.trainable_variables()
grads_op = tf.gradients(
loss_op,
variables
)
for grad, var in zip(grads_op, variables):
if grad is None:
note(var)
self.assertTrue(grad is not None)
with tf.Session(graph=g) as session:
session.run(tf.global_variables_initializer())
result, grads, _ = session.run(
[logits_op, grads_op, loss_op],
{
audio_ph: audio_waves
}
)
no_batch_norms = list(
filter(
lambda var: 'batch_normaslization' not in var.name,
variables
)
)
self.assertEqual(len(grads), len(no_batch_norms))
self.assertFalse(any([np.isnan(grad).any() for grad in grads]))
def test_dataset_returns_data_in_order(self):
params = experiment_params(
dataset_params(
batch_size=2,
epochs=1,
augment=False
)
)
data = pd.DataFrame(
data={
'text': [ str(i) for i in range(10)],
'filename': [ '{}/wav'.format(i) for i in range(10)]
}
)
dataset = input_fn(data, params['data'], dummy_load_wave)()
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
with tf.Session() as session:
try:
while True:
audio, label = session.run(next_element)
audio, length = audio
for _audio, _label in zip(list(audio), list(label)):
self.assertEqual(_audio[0], float(_label))
for _length in length:
self.assertEqual(_length, SAMPLING_RATE)
except tf.errors.OutOfRangeError:
pass
if __name__ == '__main__' and RUN_TESTS:
import doctest
doctest.testmod()
unittest.main(
argv=['first-arg-is-ignored', 'TestNotebook.test_encode_and_decode_work'],
failfast=True,
exit=False
)
#import glob
#
#def get_len(filename):
# return len(hkl.load(filename))
#
#lengths = [
# get_len(filename)
# for filename in glob.glob('data/cv_corpus_v1/cv-valid-train/*.wave.hkl')
#]
#np.min(lengths)
#np.max(lengths)
#import seaborn as sb
#
#sb.distplot(lengths, kde = False)
```
### Experiments
```
experiment(
dataset_params(
batch_size=18,
epochs=10,
max_wave_length=320000,
augment=True,
random_noise=0.75,
random_noise_factor_min=0.1,
random_noise_factor_max=0.15,
random_stretch_min=0.8,
random_stretch_max=1.2
),
codename='deep_max_20_seconds',
alphabet=' !"&\',-.01234:;\\abcdefghijklmnopqrstuvwxyz', # !"&',-.01234:;\abcdefghijklmnopqrstuvwxyz
causal_convolutions=False,
stack_dilation_rates=[1, 3, 9, 27],
stacks=6,
stack_kernel_size=7,
stack_filters=3*128,
n_fft=160*8,
frame_step=160*4,
num_mel_bins=160,
optimizer='Momentum',
lr=0.00001,
clip_gradients=20.0
)
test(
dataset_params(
batch_size=18,
epochs=10,
max_wave_length=320000,
augment=True,
random_noise=0.75,
random_noise_factor_min=0.1,
random_noise_factor_max=0.15,
random_stretch_min=0.8,
random_stretch_max=1.2
),
codename='deep_max_20_seconds',
alphabet=' !"&\',-.01234:;\\abcdefghijklmnopqrstuvwxyz', # !"&',-.01234:;\abcdefghijklmnopqrstuvwxyz
causal_convolutions=False,
stack_dilation_rates=[1, 3, 9, 27],
stacks=6,
stack_kernel_size=7,
stack_filters=3*128,
n_fft=160*8,
frame_step=160*4,
num_mel_bins=160,
optimizer='Momentum',
lr=0.00001,
clip_gradients=20.0
)
results = predict(
'cv_corpus_v1/test-me.m4a',
codename='deep_max_20_seconds',
alphabet=' !"&\',-.01234:;\\abcdefghijklmnopqrstuvwxyz', # !"&',-.01234:;\abcdefghijklmnopqrstuvwxyz
causal_convolutions=False,
stack_dilation_rates=[1, 3, 9, 27],
stacks=6,
stack_kernel_size=7,
stack_filters=3*128,
n_fft=160*8,
frame_step=160*4,
num_mel_bins=160,
optimizer='Momentum',
lr=0.00001,
clip_gradients=20.0
)
b''.join(results[0]['text'])
test_results = predict_test(
codename='deep_max_20_seconds',
alphabet=' !"&\',-.01234:;\\abcdefghijklmnopqrstuvwxyz', # !"&',-.01234:;\abcdefghijklmnopqrstuvwxyz
causal_convolutions=False,
stack_dilation_rates=[1, 3, 9, 27],
stacks=6,
stack_kernel_size=7,
stack_filters=3*128,
n_fft=160*8,
frame_step=160*4,
num_mel_bins=160,
optimizer='Momentum',
lr=0.00001,
clip_gradients=20.0
)
[ b''.join(t['text']) for t in test_results ]
```
| github_jupyter |
----
<img src="../../../files/refinitiv.png" width="20%" style="vertical-align: top;">
# Data Library for Python
----
## Content layer - Environmental Social and Governance (ESG)
This notebook demonstrates how to retrieve Environmental Social and Governance data.
#### Learn more
To learn more about the Refinitiv Data Library for Python please join the Refinitiv Developer Community. By [registering](https://developers.refinitiv.com/iam/register) and [logging](https://developers.refinitiv.com/content/devportal/en_us/initCookie.html) into the Refinitiv Developer Community portal you will have free access to a number of learning materials like
[Quick Start guides](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/quick-start),
[Tutorials](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/learning),
[Documentation](https://developers.refinitiv.com/en/api-catalog/refinitiv-data-platform/refinitiv-data-library-for-python/docs)
and much more.
#### Getting Help and Support
If you have any questions regarding using the API, please post them on
the [Refinitiv Data Q&A Forum](https://community.developers.refinitiv.com/spaces/321/index.html).
The Refinitiv Developer Community will be happy to help.
## Set the configuration file location
For a better ease of use, you have the option to set initialization parameters of the Refinitiv Data Library in the _refinitiv-data.config.json_ configuration file. This file must be located beside your notebook, in your user folder or in a folder defined by the _RD_LIB_CONFIG_PATH_ environment variable. The _RD_LIB_CONFIG_PATH_ environment variable is the option used by this series of examples. The following code sets this environment variable.
```
import os
os.environ["RD_LIB_CONFIG_PATH"] = "../../../Configuration"
```
## Some Imports to start with
```
import refinitiv.data as rd
from refinitiv.data.content import esg
```
## Open the data session
The open_session() function creates and open sessions based on the information contained in the refinitiv-data.config.json configuration file. Please edit this file to set the session type and other parameters required for the session you want to open.
```
rd.open_session('platform.rdp')
```
## Retrieve data
### Get all universe
```
response = esg.universe.Definition().get_data()
response.data.df
```
### Basic Overview for one instrument
```
response = esg.basic_overview.Definition("MSFT.O").get_data()
response.data.df
```
### Standard Scores
```
response = esg.standard_scores.Definition("6758.T").get_data()
response.data.df
response = esg.standard_scores.Definition('AAPL.O').get_data()
response.data.df
```
### Full Scores
```
response = esg.full_scores.Definition(
universe="4295904307",
start=0,
end=-5
).get_data()
response.data.df
```
### Standard Measures
```
response = esg.standard_measures.Definition("BNPP.PA").get_data()
response.data.df
```
### Full Measures
```
response = esg.full_measures.Definition("BNPP.PA").get_data()
response.data.df
```
## Close the session
```
rd.close_session()
```
| github_jupyter |
```
# Install tf-transformers from github
```
# Roberta2Roberta + Summarization + Xsum
This tutorial contains code to fine-tune an Roberta2Roberta Encoder Decoder Model for Summarization
In this notebook:
- Load the data + create ```tf.data.Dataset``` using TFWriter
- Load and warmstart Roberta base and use it to create a Summarization Model
- Train using ```tf.keras.Model.fit``` and ```Custom Trainer```
- Minimze LM loss
- Evaluate ROUGE score
- In production using faster ```tf.SavedModel``` + no architecture code
```
import datasets
import json
import os
import glob
import time
from tf_transformers.models import EncoderDecoderModel
from transformers import RobertaTokenizer
from tf_transformers.data import TFWriter, TFReader, TFProcessor
from tf_transformers.losses import cross_entropy_loss
from tf_transformers.core import optimization, SimpleTrainer
from absl import logging
logging.set_verbosity("INFO")
```
### Load Tokenizer
```
tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
```
### Load XSum sumarization data using Huggingface datasets
```
examples = datasets.load_from_disk("/mnt/home/PRE_MODELS/HuggingFace_models/datasets/xsum/")
train_examples = examples["train"]
```
### Parse train data
```
encoder_max_length=512
decoder_max_length=64
def parse_train():
result = {}
for f in train_examples:
input_ids = [tokenizer.cls_token] + tokenizer.tokenize(f['document'])[: encoder_max_length-2] + [tokenizer.sep_token] # -2 to add CLS and SEP
input_ids = tokenizer.convert_tokens_to_ids(input_ids)
input_mask = [1] * len(input_ids)
input_type_ids = [0] * len(input_ids)
decoder_input_ids = [tokenizer.cls_token] + tokenizer.tokenize(f['summary'])[: decoder_max_length-2] + [tokenizer.sep_token]
decoder_input_ids = tokenizer.convert_tokens_to_ids(decoder_input_ids)
decoder_input_type_ids = [0] * len(decoder_input_ids)
result = {}
result['encoder_input_ids'] = input_ids
result['encoder_input_mask'] = input_mask
result['encoder_input_type_ids'] = input_type_ids
result['decoder_input_ids'] = decoder_input_ids[:-1] # except last word
result['decoder_input_type_ids'] = decoder_input_type_ids[:-1] # except last word
result['labels'] = decoder_input_ids[1:] # not including first word
result['labels_mask'] = [1] * len(decoder_input_ids[1:])
# Decoder doesnt need input_mask because by default decoder has causal mask mode
yield result
# Lets write using TF Writer
# Use TFProcessor for smalled data
schema = {
"encoder_input_ids": ("var_len", "int"),
"encoder_input_mask": ("var_len", "int"),
"encoder_input_type_ids": ("var_len", "int"),
"decoder_input_ids": ("var_len", "int"),
"decoder_input_type_ids": ("var_len", "int"),
"labels": ("var_len", "int"),
"labels_mask": ("var_len", "int"),
}
tfrecord_train_dir = '../OFFICIAL_TFRECORDS/bbc_xsum/roberta/train'
tfrecord_filename = 'bbc_xsum'
tfwriter = TFWriter(schema=schema,
file_name=tfrecord_filename,
model_dir=tfrecord_train_dir,
tag='train',
overwrite=True
)
tfwriter.process(parse_fn=parse_train())
```
### Read TFRecords using TFReader
```
# Read Data
schema = json.load(open("{}/schema.json".format(tfrecord_train_dir)))
all_files = glob.glob("{}/*.tfrecord".format(tfrecord_train_dir))
tf_reader = TFReader(schema=schema,
tfrecord_files=all_files)
x_keys = ['encoder_input_ids', 'encoder_input_type_ids', 'encoder_input_mask', 'decoder_input_ids', 'decoder_input_type_ids']
y_keys = ['labels', 'labels_mask']
batch_size = 8
train_dataset = tf_reader.read_record(auto_batch=True,
keys=x_keys,
batch_size=batch_size,
x_keys = x_keys,
y_keys = y_keys,
shuffle=True,
drop_remainder=True
)
# Look at inputs, labels
for (batch_inputs, batch_labels) in train_dataset.take(1):
print(batch_inputs, batch_labels)
```
### Load Roberta2Roberta (Encoder Decoder Model)
```
import tensorflow as tf
tf.keras.backend.clear_session()
model_layer, model, config = EncoderDecoderModel(model_name='roberta-base',
is_training=True,
encoder_checkpoint_dir='/mnt/home/PRE_MODELS/LegacyAI_models/checkpoints/roberta-base/')
```
### Define Loss
Loss function is simple.
* labels: 2D (batch_size x sequence_length)
* logits: 3D (batch_size x sequence_length x vocab_size)
* label_weights: 2D (batch_size x sequence_length) # we don't want all words in the sequence to have loss so, we mask them and don't calculate for loss
```
def lm_loss(y_true_dict, y_pred_dict):
return cross_entropy_loss(labels=y_true_dict['labels'],
logits=y_pred_dict['token_logits'],
label_weights=y_true_dict['labels_mask'])
```
### Define Optimizer
**PRO TIP**: These models are very sensitive to optimizer, especially learning rates. So, make sure you play around to find a good combination
```
train_data_size = 204045
learning_rate = 1e-05
steps_per_epoch = int(train_data_size / batch_size)
EPOCHS = 3
num_train_steps = steps_per_epoch * EPOCHS
warmup_steps = int(0.1 * num_train_steps)
# creates an optimizer with learning rate schedule
optimizer_type = 'adamw'
adam_beta2=0.997
adam_epsilon=1e-09
optimizer, learning_rate_fn = optimization.create_optimizer(learning_rate,
steps_per_epoch * EPOCHS,
warmup_steps,
optimizer_type = optimizer_type,
learning_rate_type = 'linear',
adam_beta_2 = adam_beta2,
adam_epsilon = adam_epsilon)
```
### Train Using Keras :-)
- ```compile2``` allows you to have directly use model outputs as well batch dataset outputs into the loss function, without any further complexity.
Note: For ```compile2```, loss_fn must be None, and custom_loss_fn must be active. Metrics are not supprted for time being.
```
# Keras Fit
# Change epochs and steps_per_epoch for full training
# If steps_per_epoch is not familiar, dont use it, provide only epochs
keras_loss_fn = {'token_logits': lm_loss
}
model.compile2(optimizer=optimizer,
loss=None,
custom_loss=keras_loss_fn
)
history = model.fit(train_dataset, epochs=2, steps_per_epoch=5)
```
### Train using SimpleTrainer (part of tf-transformers)
```
# Custom training
# You can provide gradient_accumulation_steps if required
# I find it hurting the performance, don't know why
history = SimpleTrainer(model = model,
optimizer = optimizer,
loss_fn = lm_loss,
dataset = train_dataset.repeat(EPOCHS+1), # This is important
epochs = EPOCHS,
num_train_examples = train_data_size,
batch_size = batch_size,
steps_per_call=100,
gradient_accumulation_steps=None)
model.save_checkpoint("../OFFICIAL_MODELS/bbc_xsum/roberta2roberta")
```
### Save Models
You can save models as checkpoints using ```.save_checkpoint``` attribute, which is a part of all ```LegacyModels```
```
model_save_dir = "../OFFICIAL_MODELS/bbc_xsum/roberta2roberta"
model.save_checkpoint(model_save_dir)
```
### Load the model for Text Genration (Auto-Regressive)
1. For any model to use for auto-regressive tasks we have to provide **"pipeline_mode='auto-regressive'"**
tf-transformers will handle everything for you internally
```
# Load the model by disabling dropout and add pipeline_mode = 'auto-regressive'
import tensorflow as tf
tf.keras.backend.clear_session()
model_layer, model, config = EncoderDecoderModel(model_name='roberta-base',
is_training=False,
pipeline_mode='auto-regressive'
)
model.load_checkpoint(model_save_dir)
```
### Save the model as serialized version
This is very important, because serialized model is significantly faster.
tf-transfomers provide **save_as_serialize_module**
```
# tf-transformers provide "save_as_serialize_module" for this
model.save_as_serialize_module("{}/saved_model".format(model_save_dir))
loaded = tf.saved_model.load("{}/saved_model".format(model_save_dir))
```
### Parse validation data
We use ```TFProcessor``` to create validation data, because dev data is small
```
examples = datasets.load_from_disk("/mnt/home/PRE_MODELS/HuggingFace_models/datasets/xsum/")
dev_examples = examples['validation']
encoder_max_length=512
decoder_max_length=64
def parse_dev():
result = {}
for f in dev_examples:
input_ids = [tokenizer.cls_token] + tokenizer.tokenize(f['document'])[: encoder_max_length-2] + [tokenizer.sep_token] # -2 to add CLS and SEP
input_ids = tokenizer.convert_tokens_to_ids(input_ids)
input_mask = [1] * len(input_ids)
input_type_ids = [0] * len(input_ids)
result = {}
result['encoder_input_ids'] = input_ids
result['encoder_input_mask'] = input_mask
result['encoder_input_type_ids'] = input_type_ids
yield result
tf_processor = TFProcessor()
dev_dataset = tf_processor.process(parse_fn=parse_dev())
dev_dataset = tf_processor.auto_batch(dev_dataset, batch_size=32)
for (batch_inputs) in dev_datasetset.take(1):
print(batch_inputs)
```
### Text-Generation for dev dataset
1. For **EncoderDecoder** models like Roberta2Roberts, Bert2GPT, t5, BART use **TextDecoderSeq2Seq**
2. For **Encoder** only models like GPT2, BERT, Roberta use **TextDecoder**
```
from tf_transformers.text import TextDecoderSeq2Seq
# You can pass model = (saved_model or keras model)
# Saved model take 1200 seconds
# Keras Model take 2300 seconds
# Thats why always choose saved_model for faster inference in production
decoder = TextDecoderSeq2Seq(model=loaded,
decoder_start_token_id=tokenizer.cls_token_id, # Decoder always expect a start_token_id
decoder_input_type_ids=0 # If you have input_type_ids
)
# Greedy Decoding
start_time = time.time()
predicted_summaries = []
for batch_inputs in dev_dataset:
model_outputs = decoder.decode(batch_inputs,
mode='greedy',
max_iterations=64,
eos_id=tokenizer.sep_token_id)
output_summaries = tokenizer.batch_decode(tf.squeeze(model_outputs['predicted_ids'], 1), skip_special_tokens=True)
predicted_summaries.extend(output_summaries)
end_time = time.time()
print("Time taken is {}".format(end_time-start_time))
```
### Evaluate ROUGE score using Huggingface datasets metric
```
original_summaries = [item['summary'] for item in dev_examples]
rouge = datasets.load_metric("rouge")
rouge_output2 = rouge.compute(predictions=predicted_summaries, references=original_summaries, rouge_types=["rouge2"])["rouge2"].mid
rouge_output1 = rouge.compute(predictions=predicted_summaries, references=original_summaries, rouge_types=["rouge1"])["rouge1"].mid
rouge_outputL = rouge.compute(predictions=predicted_summaries, references=original_summaries, rouge_types=["rougeL"])["rougeL"].mid
print("Rouge1", rouge_output1)
print("Rouge2", rouge_output2)
print("RougeL", rouge_outputL)
Rouge1 Score(precision=0.4030931388183621, recall=0.36466254213804195, fmeasure=0.37530511219642493)
Rouge2 Score(precision=0.16782261295821255, recall=0.15203057122838165, fmeasure=0.1561568579032115)
RougeL Score(precision=0.327351036176015, recall=0.2969254630660535, fmeasure=0.30522124104859427)
```
### Evaluate ROUGE score using Google rouge_score library
```
from rouge_score import rouge_scorer
from rouge_score import scoring
scorer = rouge_scorer.RougeScorer(["rouge1", "rouge2", "rougeLsum"], use_stemmer=True)
aggregator = scoring.BootstrapAggregator()
for i in range(len(original_summaries)):
score = scorer.score(original_summaries[i], predicted_summaries[i])
aggregator.add_scores(score)
print("Aggregated scores", aggregator.aggregate())
{'rouge1': AggregateScore(low=Score(precision=0.4140161871825283, recall=0.37430234257946526, fmeasure=0.38546441367482415), mid=Score(precision=0.4170448579572875, recall=0.377157245705547, fmeasure=0.3882117449428194), high=Score(precision=0.4201437437963473, recall=0.37995542626327267, fmeasure=0.39091534567006364)),
'rouge2': AggregateScore(low=Score(precision=0.1691630552674417, recall=0.1529611842223162, fmeasure=0.1573395483096894),mid=Score(precision=0.17186887531131723, recall=0.1556336324896489, fmeasure=0.15993720424744032), high=Score(precision=0.17473545118625014, recall=0.1582221241761583, fmeasure=0.16259483307298692)),
'rougeLsum': AggregateScore(low=Score(precision=0.33240856683967057, recall=0.301583552325229, fmeasure=0.3101081853974168), mid=Score(precision=0.33521687141849427, recall=0.3040523188928863, fmeasure=0.3125425409744611), high=Score(precision=0.33844188019812493, recall=0.3066532822640133, fmeasure=0.31522376216006714))}
```
### In Production
1. Lets see how we can deploy this model in production
```
from tf_transformers.text import TextDecoderSeq2Seq
from tf_transformers.data import pad_dataset
# 1. Load Saved Model
loaded = tf.saved_model.load("{}/saved_model".format(model_save_dir))
# 2. Initiate a decode object
decoder = TextDecoderSeq2Seq(model=loaded,
decoder_start_token_id=tokenizer.cls_token_id, # Decoder always expect a start_token_id
decoder_input_type_ids=0 # If you have input_type_ids
)
# 3. Convert text to inputs
# Tokenizer fn convert text -> model inputs
# Make sure you return dict with key-> list of list
# pad_dataset is a decorator, hich will automatically taken care of padding
# If you want to write your own function, please. model expect inputs in a specifed format thats all.
@pad_dataset
def tokenizer_fn(texts):
input_ids = []
input_mask = []
input_type_ids = []
for text in texts:
input_ids_ex = [tokenizer.cls_token] + tokenizer.tokenize(text)[: encoder_max_length-2] + [tokenizer.sep_token] # -2 to add CLS and SEP
input_ids_ex = tokenizer.convert_tokens_to_ids(input_ids_ex)
input_mask_ex = [1] * len(input_ids_ex)
input_type_ids_ex = [0] * len(input_ids_ex)
input_ids.append(input_ids_ex)
input_mask.append(input_mask_ex)
input_type_ids.append(input_type_ids_ex)
result = {}
result['encoder_input_ids'] = input_ids
result['encoder_input_mask'] = input_mask
result['encoder_input_type_ids'] = input_type_ids
return result
# 4. Examples
text1 = '''Tulips (Tulipa) form a genus of spring-blooming perennial herbaceous bulbiferous geophytes (having bulbs as storage organs). The flowers are usually large, showy and brightly colored, generally red, pink, yellow, or white (usually in warm colors). They often have a different colored blotch at the base of the tepals (petals and sepals, collectively), internally. Because of a degree of variability within the populations, and a long history of cultivation, classification has been complex and controversial. The tulip is a member of the lily family, Liliaceae, along with 14 other genera, where it is most closely related to Amana, Erythronium and Gagea in the tribe Lilieae. There are about 75 species, and these are divided among four subgenera. The name "tulip" is thought to be derived from a Persian word for turban, which it may have been thought to resemble. Tulips originally were found in a band stretching from Southern Europe to Central Asia, but since the seventeenth century have become widely naturalised and cultivated (see map). In their natural state they are adapted to steppes and mountainous areas with temperate climates. Flowering in the spring, they become dormant in the summer once the flowers and leaves die back, emerging above ground as a shoot from the underground bulb in early spring.
Originally growing wild in the valleys of the Tian Shan Mountains, tulips were cultivated in Constantinople as early as 1055. By the 15th century, tulips were among the most prized flowers; becoming the symbol of the Ottomans.[2] While tulips had probably been cultivated in Persia from the tenth century, they did not come to the attention of the West until the sixteenth century, when Western diplomats to the Ottoman court observed and reported on them. They were rapidly introduced into Europe and became a frenzied commodity during Tulip mania. Tulips were frequently depicted in Dutch Golden Age paintings, and have become associated with the Netherlands, the major producer for world markets, ever since. In the seventeenth century Netherlands, during the time of the Tulip mania, an infection of tulip bulbs by the tulip breaking virus created variegated patterns in the tulip flowers that were much admired and valued. While truly broken tulips do not exist anymore, the closest available specimens today are part of the group known as the Rembrandts – so named because Rembrandt painted some of the most admired breaks of his time.[3]'''
text2 = '''By any yardstick, the 2013 blockbuster Drishyam is a hard act to follow. Writer-director Jeethu Joseph’s crime thriller starring Mohanlal, Meena, Asha Sharath and Siddique was so well-rounded in the writing and execution of its murder-and-subsequent-cover-up mystery and such a box-office superhit that it was remade in Tamil, Telugu, Hindi and Kannada, headlined by some of the biggest male stars of those industries, in addition to foreign revisitations in Sinhalese and Mandarin.
At the time, Jeethu was questioned about his script drawing on Japanese novelist Keigo Higashino’s The Devotion of Suspect X, but he denied the charge and said he was inspired instead by a real-life incident. Be that as it may, Drishyam 2: The Resumption is all the redemption he needs. In a country that does not have a great track record with whodunnits, pulling off a brilliant howdunnit and howhegotawaywithit like Drishyam was an achievement. Returning with a howhe’sstillgettingawaywithit and actually pulling it off is nothing short of incredible.
Drishyam 2 is a surprisingly satisfying sequel to a spectacular first film.
Jeethu Joseph’s new crime drama is set in the same Kerala town where the events of its precursor took place. Georgekutty (Mohanlal) is now the owner of a cinema theatre. His prosperity is reflected in the larger, posher house he currently occupies with his wife Rani (Meena) and their daughters Anju (Ansiba) and Anu (Esther Anil) on the same land where they earlier lived. He is still movie crazy. Rani and he are still a committed couple yet constantly sniping at each other as before. And they are still a rock-solid team in the upbringing of their girls.
The difference between then and now is twofold. First, the townsfolk had backed the family when IG Geetha Prabhakar (Asha Sharath) got after them on the suspicion that they killed her son. They are not so supportive any more, driven as they are by jealousy at Georgekutty’s rise in life.
Second, the experiences of Drishyam have had a deep psychological impact on both Rani and Anju. Rani is tormented by Georgekutty’s refusal to ever discuss what happened back then. The first half of Drishyam 2 constructs their continuing trauma and gradually establishes the fact that the police never gave up on the case. The second half is about the resumed investigation.'''
# 5. Choose the type of decoding
batch_inputs = tokenizer_fn([text1, text2])
model_outputs = decoder.decode(batch_inputs,
mode='greedy',
max_iterations=64,
eos_id=tokenizer.sep_token_id)
output_summaries = tokenizer.batch_decode(tf.squeeze(model_outputs['predicted_ids'], 1), skip_special_tokens=True)
output_summaries
```
### Advanced
**TextDecoderSerializable** internally uses for loop.
Can we do better. If we could use ```tf.while_loop```, we can save the whole model as serialized.
which no only improves speed, but also make life much easier in production
```
# Save the end-to-end decoder as seriazed model
from tf_transformers.text import TextDecoderSerializableSeq2Seq
from tf_transformers.core import LegacyModule
decoder_layer = TextDecoderSerializableSeq2Seq(model=model,
decoder_start_token_id=tokenizer.cls_token_id, # Decoder always expect a start_token_id
decoder_input_type_ids=0, # If you have input_type_ids
mode="greedy",
max_iterations=64,
eos_id=tokenizer.sep_token_id
)
decoder_model = decoder_layer.get_model()
decoder_module = LegacyModule(decoder_model)
decoder_module.save("{}/saved_decoder_model".format(model_save_dir))
```
### In Production (Advanced) - Just 2 lines of code.
```
# 1. Load serialized model
decoder_serialized = tf.saved_model.load("{}/saved_decoder_model".format(model_save_dir))
# 2. text to inputs
model_outputs2 = decoder_serialized(**batch_inputs)
output_summaries2 = tokenizer.batch_decode(tf.squeeze(model_outputs2['predicted_ids'], 1), skip_special_tokens=True)
# Output summaries matches with TextDecoderSerializableSeq2Seq
assert(output_summaries == output_summaries2)
# Succesful :-)
```
| github_jupyter |
# Explanation of the fibertree union operator
## Introduction
The following cells provide a series of examples of the ```union``` operation on fibers. In general, the ```union``` operation creates a new fiber with an element for each coordinate that exists in either input fiber (i.e., the **union** of their coordinates) and a payload that is a tuple of the corresponding payloads from the input fibers.
First, we include some libraries and provide some dropdown lists to select the display style and type of animation.
```
#
# Run boilerplate code to set up environment
#
%run ../prelude.py --style=tree --animation=movie
```
## Fibertree union operator
One can union the contents of two fibers using the ``or`` (|) operator. That operator takes two fibers as operands, and returns a fiber that has a element for each coordinate that appears in **either** input fiber and a payload that consists of a triple (three element tuple). The first element of the triple is a mask (indicating whether the rest of the triple contains non-empty payloads from only-A, only-B or both-A-and-B. The next two elements of the triple contain the corresponding payloads from the two input fibers. If an input fiber doesn't have a particular coordinate the **default** payload value (typically zero) of that fiber is used in the triple. However, if there is no payload at a particular coordinate in either input fiber then that coordinate will not appear in the output - note the absense of coordinate 1 in result for the example below.
```
#
# Create two rank-1 tensors
#
a_M = Tensor.fromUncompressed(["M"], [1, 0, 3, 0, 5, 0, 7])
b_M = Tensor.fromUncompressed(["M"], [2, 0, 4, 5])
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
#
# Calculate the union of the two fibers
#
z1_m = a_m | b_m
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber a_m | b_m")
displayTensor(z1_m)
print(f"{z1_m:n*}")
```
The **default** payload of a the result of ```union``` operation is a tuple of the form of the payloads in the result fiber. This default payload will be used for operations such as an insertion on a new coordinate into the fiber (e.g., using the ```<<``` operator). It also is the shape of the payload returned when the fiber is iterated over. The **default** value for payloads in the fiber ```z1_m``` created above is shown in the cell below, and is an empty string and two scalar zeros.
```
#
# Obtain the default payload of the result and print it
#
z1_m_default = z1_m.getDefault()
print(z1_m_default)
```
## Traversing the result of a union
Traversing the result of a union using a ```for``` loop is like traversing any other fiber, except the payload must match the shape of the **default** payload of the fiber. See below how the payload returned by the interaction is a three element tuple:
```
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
#
# Traverse the elements of a union operation
#
for c, (mask, a_val, b_val) in a_m | b_m:
print(f"Coordinate: {c}")
print(f"Mask: {mask}")
print(f"A_val: {a_val}")
print(f"B_val: {b_val}")
```
## Union of complex payloads
Note that one can take the union of a fiber whose payloads have a more complex type, such as a fiber (e.g., from the top rank of a multirank tensor). This is shown in the example below. Note that neither fiber a or fiber b has a payload at coordinate 3, so the output has no output at that coordinate. Unfortunatly, the image view of a tensor whose payload are tuples with Fibers as an element are sort of messy. So a textual print of the result is also shown.
```
#
# Create two rank-2 tensors
#
a_MK = Tensor.fromUncompressed(["M", "K"], [[1, 0, 3, 0, 5, 0, 7],
[2, 2, 0, 3, 0, 0, 8],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[4, 0, 5, 0, 8, 0, 9]])
b_MK = Tensor.fromUncompressed(["M", "K"], [[2, 0, 4, 5],
[0, 0, 0, 0],
[3, 4, 6, 0],
[0, 0, 0, 0],
[1, 2, 3, 4]]
)
#
# Get the root fibers of the tensors
#
a_m = a_MK.getRoot()
b_m = b_MK.getRoot()
#
# Calculate the union of the two fibers
#
z2_m = a_m | b_m
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber a_m | b_m")
displayTensor(z2_m)
print(f"{z2_m:n*}")
```
The default payload for the above union of fibers with more complex payloads is also more complex. In this case, it is a tuple containing an empty string and two fibers (actually a reference to the constructor for a fiber).
```
#
# Obtain the default payload of the result and print it
#
z2_m_default = z2_m.getDefault()
print(z2_m_default)
```
## Union of asymetric complex payloads
Note that one can take the union of a fibers whose payloads are different types. In this example we union a fiber with fibers as its payloads (e.g., the top rank of a multirank tensor) with a fiber whose payloads are scalars.
```
#
# Create another rank-2 tensor (is this the same as above)
#
a_MK = Tensor.fromUncompressed(["M", "K"], [[1, 0, 3, 0, 5, 0, 7],
[0, 0, 0, 0, 0, 0, 0],
[2, 2, 0, 3, 0, 0, 8],
[0, 0, 0, 0, 0, 0, 0],
[4, 0, 5, 0, 8, 0, 9]])
#
# Get the root fibers of the tensors
#
a_m = a_MK.getRoot()
b_m = b_M.getRoot()
#
# Calculate the union of the two fibers
#
z3_m = a_m | b_m
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber a_m | b_m")
displayTensor(z3_m)
print(f"{z3_m:n*}")
```
The default payload for the above union of fibers with more complex payloads is also more complex. In this case, it is a tuple containing an empty string and a fiber and a zero.
```
#
# Obtain the default payload of the result and print it
#
z3_m_default = z3_m.getDefault()
print(z3_m_default)
```
## Unions of unions
We can take the union of fiber with a fiber that was already a union of two fibers. This is illustrated in the cell below
Note; That the coordinate 1 of the result has an empty "A" element of the tuple, which was generated from the **default** payload for the result of ```a_m | b_m```. Also there is no coordinate 5 in the result, so **no** input fiber had a non-empty payload for coordinate 5.
```
#
# Create another rank-1 tensor
#
c_M = Tensor.fromUncompressed(["M"], [1, 2, 3])
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
c_m = c_M.getRoot()
#
# Calculate the union of the three fibers
#
z4_m = (a_m | b_m) | c_m
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber c_m")
displayTensor(c_m)
print("Fiber (a_m | b_m) | c_m")
displayTensor(z4_m)
# Fiber is too complex to print!
#print(f"{z4_m:n*}")
for m, (mask1, (mask2, a_val, b_val), c_val) in z4_m:
print(f"mask1: {mask1}")
print(f" mask2: {mask2}")
print(f" a_val: {a_val}")
print(f" b_val: {b_val}")
print(f" c_val: {c_val}")
```
Note the shape of the default payload for the result fiber which contains a nested tuple.
```
#
# Obtain the default payload of the result and print it
#
z4_m_default = z4_m.getDefault()
print(z4_m_default)
```
## Different assocation
Associating the unions differently produces a result that differs in the nesting of the payloads. Note how coordinates 4 and 6 have a payload that contains the **default** payload for the union of ```b_m``` and ```c_m```.
```
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
c_m = c_M.getRoot()
#
# Calculate the union of the three fibers
#
z5_m = a_m | ( b_m | c_m )
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber c_m")
displayTensor(c_m)
print("Fiber a_m | (b_m | c_m)")
displayTensor(z5_m)
# Fiber is too complex to print!
#print(f"{z4_m:n*}")
for m, (mask1, a_val, (mask2, b_val, c_val)) in z5_m:
print(f"mask1: {mask1}")
print(f" a_val: {a_val}")
print(f" mask2: {mask2}")
print(f" b_val: {b_val}")
print(f" c_val: {c_val}")
```
Note the shape of the default payload for the result fiber, which again is nested tuples, but in this case the third element is a tuple.
```
#
# Obtain the default payload of the result and print it
#
z5_m_default = z5_m.getDefault()
print(z5_m_default)
```
## Union multi-argument operator
To allow a cleaner union of multiple operands the library includes a union operator that takes an arbitrary number of arguments (signature ```Fiber.union(*args)```). The payloads of the result of such a multi-argument union is a payload for each coordinate that exists in **any** of the inpupt arguments and that payload is a tuple comtaining a mask (with letters A-Z) and a entry sourced from the corresponding payload in each input argument fiber (input argument fibers with nothing at that coordinate will use the **default** payload from that fiber). This is illustrated below:
```
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
c_m = c_M.getRoot()
#
# Calculate the union of the three fibers
#
z6_m = Fiber.union(a_m, b_m, c_m)
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber c_m")
displayTensor(c_m)
print("Fiber union(a_m, b_m, c_m")
displayTensor(z6_m)
# Fiber is too complex to print!
#print(f"{z4_m:n*}")
for m, (mask1, a_val, b_val, c_val) in z6_m:
print(f"mask1: {mask1}")
print(f" a_val: {a_val}")
print(f" b_val: {b_val}")
print(f" c_val: {c_val}")
```
Note the shape of the default payload for the result fiber
```
#
# Obtain the default payload of the result and print it
#
z6_m_default = z6_m.getDefault()
print(z6_m_default)
```
## Multi-argument union with complex payloads
The ```Fiber.union()``` operator also works with more complex payloads
```
#
# Create one more rank-2 tensor
#
d_MK = Tensor.fromUncompressed(["M", "K"], [[8, 0, 6, 0],
[0, 0, 0, 0],
[5, 0, 7, 0],
[4, 8, 1, 2],
[1, 2, 3, 4]])
#
# Get the root fibers of the tensors
#
a_m = a_M.getRoot()
b_m = b_M.getRoot()
c_m = c_M.getRoot()
d_m = d_MK.getRoot()
#
# Calculate the union of the three fibers
#
z7_m = Fiber.union(a_m, b_m, c_m, d_m)
#
# Print the inputs and outputs
#
print("Fiber a_m")
displayTensor(a_m)
print("Fiber b_m")
displayTensor(b_m)
print("Fiber c_m")
displayTensor(c_m)
print("Fiber d_m")
displayTensor(d_m)
print("Fiber union(a_m, b_m, c_m, d_m)")
displayTensor(z7_m)
# Fiber is too complex to print!
#print(f"{z4_m:n*}")
for m, (mask1, a_val, b_val, c_val, d_val) in z7_m:
print(f"mask1: {mask1}")
print(f" a_val: {a_val}")
print(f" b_val: {b_val}")
print(f" c_val: {c_val}")
print(f" d_val: {d_val}")
```
Note the shape of the default payload for the resulting fiber from ```Fiber.union()```, where one of the elements of the default payload is a Fiber.
```
#
# Obtain the default payload of the result and print it
#
z7_m_default = z7_m.getDefault()
print(z7_m_default)
```
## Testing area
For running alternative algorithms
| github_jupyter |
# Instance Segmentation with Mask R-CNN with TPU support
*by Georgios K. Ouzounis, June 22nd, 2021*
In this notebook we will experiment with **instance segmentation** in still images using the **Mask R-CNN** model, trained on Cloud TPU.
This is a slightly altered version of the original notebook posted by Google Research that can be found [here](https://colab.research.google.com/github/tensorflow/tpu/blob/master/models/official/mask_rcnn/mask_rcnn_demo.ipynb#scrollTo=t_iHs_wm2Mhh)
For each test image, the output set of predictions includes bounding boxes, labels and instance masks that are overlayed on the input.
## Instructions
<h3><a href="https://cloud.google.com/tpu/"><img valign="middle" src="https://raw.githubusercontent.com/GoogleCloudPlatform/tensorflow-without-a-phd/master/tensorflow-rl-pong/images/tpu-hexagon.png" width="50"></a> Use a free Cloud TPU</h3>
On the main menu, click Runtime and select **Change runtime type**. Set "TPU" as the hardware accelerator.
## Download the source code
Download the source code of the Mask R-CNN model from the **tensorflow/tpu/** github repo.
```
!git clone https://github.com/tensorflow/tpu/
```
## Import libraries
```
import numpy as np
import cv2
%tensorflow_version 1.x
import tensorflow as tf
import sys
sys.path.insert(0, 'tpu/models/official')
sys.path.insert(0, 'tpu/models/official/mask_rcnn')
import coco_metric
from mask_rcnn.object_detection import visualization_utils
```
## Load the COCO index mapping
This Colab uses a pretrained checkpoint of the Mask R-CNN model that is trained using the COCO dataset. Below is the mapping between the indices that the model predicts and the categories in text.
```
ID_MAPPING = {
1: 'person',
2: 'bicycle',
3: 'car',
4: 'motorcycle',
5: 'airplane',
6: 'bus',
7: 'train',
8: 'truck',
9: 'boat',
10: 'traffic light',
11: 'fire hydrant',
13: 'stop sign',
14: 'parking meter',
15: 'bench',
16: 'bird',
17: 'cat',
18: 'dog',
19: 'horse',
20: 'sheep',
21: 'cow',
22: 'elephant',
23: 'bear',
24: 'zebra',
25: 'giraffe',
27: 'backpack',
28: 'umbrella',
31: 'handbag',
32: 'tie',
33: 'suitcase',
34: 'frisbee',
35: 'skis',
36: 'snowboard',
37: 'sports ball',
38: 'kite',
39: 'baseball bat',
40: 'baseball glove',
41: 'skateboard',
42: 'surfboard',
43: 'tennis racket',
44: 'bottle',
46: 'wine glass',
47: 'cup',
48: 'fork',
49: 'knife',
50: 'spoon',
51: 'bowl',
52: 'banana',
53: 'apple',
54: 'sandwich',
55: 'orange',
56: 'broccoli',
57: 'carrot',
58: 'hot dog',
59: 'pizza',
60: 'donut',
61: 'cake',
62: 'chair',
63: 'couch',
64: 'potted plant',
65: 'bed',
67: 'dining table',
70: 'toilet',
72: 'tv',
73: 'laptop',
74: 'mouse',
75: 'remote',
76: 'keyboard',
77: 'cell phone',
78: 'microwave',
79: 'oven',
80: 'toaster',
81: 'sink',
82: 'refrigerator',
84: 'book',
85: 'clock',
86: 'vase',
87: 'scissors',
88: 'teddy bear',
89: 'hair drier',
90: 'toothbrush',
}
#create a dictionary with class IDs mapped to the COCO labels
category_index = {k: {'id': k, 'name': ID_MAPPING[k]} for k in ID_MAPPING}
```
## Get a sample image
Use the **wget** command to download locally an image of your liking or mount your Google Drive and copy one locally
```
# sample image used in the original Colab Notebook
!wget https://upload.wikimedia.org/wikipedia/commons/thumb/0/08/Kitano_Street_Kobe01s5s4110.jpg/2560px-Kitano_Street_Kobe01s5s4110.jpg -O test.jpg
image_path = 'test.jpg'
# sample image from the author's github repo
!wget https://github.com/georgiosouzounis/instance-segmentation-mask-rcnn/raw/main/data/newyork.jpg -O test.jpg
image_path = 'test.jpg'
# read the image both as 3D numpy array (openCV) and as a serialized string
# for model compatibility
image = cv2.imread(image_path)
# convert the BGR order to RGB
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# get the image width and height
width, height = image.shape[1], image.shape[0]
# serialization
with open(image_path, 'rb') as f:
np_image_string = np.array([f.read()])
# view the selected image
from google.colab.patches import cv2_imshow
cv2_imshow(image)
```
## Create a Tensorflow session
Create a Tensorflow session to run the inference. You can either connect to a TPU or a normal CPU backend.
```
use_tpu = False #@param {type:"boolean"}
# if using the TPU runtime:
if use_tpu:
import os
import pprint
# assert the TPU address
assert 'COLAB_TPU_ADDR' in os.environ, 'ERROR: Not connected to a TPU runtime; please see the first cell in this notebook for instructions!'
TPU_ADDRESS = 'grpc://' + os.environ['COLAB_TPU_ADDR']
print('TPU address is', TPU_ADDRESS)
# initialize a session
session = tf.Session(TPU_ADDRESS, graph=tf.Graph())
print('TPU devices:')
pprint.pprint(session.list_devices())
# else if using the CPU runtime:
else:
# initialize a session
session = tf.Session(graph=tf.Graph())
```
## Load the pretrained model
Load the COCO pretrained model from the public GCS bucket. Ignore the deprecation warnings as there is no immediate fix for using tensorflow2 togetehr with Mask R-CNN
```
# set the model directory here or on the line to the right
saved_model_dir = 'gs://cloud-tpu-checkpoints/mask-rcnn/1555659850' #@param {type:"string"}
# load the model
# underscore _ is considered as "I don't Care" or "Throwaway" variable in Python.
_ = tf.saved_model.loader.load(session, ['serve'], saved_model_dir)
```
## Compute instance segmentation
Run the inference and process the predictions returned by the model.
```
# get the predictions by running the session created earlier
num_detections, detection_boxes, detection_classes, detection_scores, detection_masks, image_info = session.run(
['NumDetections:0', 'DetectionBoxes:0', 'DetectionClasses:0', 'DetectionScores:0', 'DetectionMasks:0', 'ImageInfo:0'],
feed_dict={'Placeholder:0': np_image_string})
# remove axes of length 1 in each of the numpy arrays returned at the end of the session.
num_detections = np.squeeze(num_detections.astype(np.int32), axis=(0,))
detection_boxes = np.squeeze(detection_boxes * image_info[0, 2], axis=(0,))[0:num_detections]
detection_scores = np.squeeze(detection_scores, axis=(0,))[0:num_detections]
detection_classes = np.squeeze(detection_classes.astype(np.int32), axis=(0,))[0:num_detections]
instance_masks = np.squeeze(detection_masks, axis=(0,))[0:num_detections]
# extract the bounding box endpoints from the detection_boxes array
ymin, xmin, ymax, xmax = np.split(detection_boxes, 4, axis=-1)
# convert each bbox endpoint array to the desired format [x_start, y_start, width, height]
processed_boxes = np.concatenate([xmin, ymin, xmax - xmin, ymax - ymin], axis=-1)
# generates the segmentation result from an instance mask and its bbox for each detection
segmentations = coco_metric.generate_segmentation_from_masks(instance_masks, processed_boxes, height, width)
```
## Visualize the detection results
```
# set the max number of boxes to draw and the detection confidence threshold
max_boxes_to_draw = 50 #@param {type:"integer"}
min_score_thresh = 0.5 #@param {type:"slider", min:0, max:1, step:0.01}
# create the ouput image with bboxes, labels and segments imprinted
image_with_detections = visualization_utils.visualize_boxes_and_labels_on_image_array(
image,
detection_boxes,
detection_classes,
detection_scores,
category_index,
instance_masks=segmentations,
use_normalized_coordinates=False,
max_boxes_to_draw=max_boxes_to_draw,
min_score_thresh=min_score_thresh)
# dispaly the resulting image
cv2_imshow(image_with_detections)
```
## Training Mask R-CNN on Cloud TPU
To train the Mask R-CNN on custom data on Cloud TPU you may wish to consult [this tutorial](https://cloud.google.com/tpu/docs/tutorials/mask-rcnn) from Google Research. Please do note that the tutorial uses billable components of Google Cloud, including:
- Compute Engine
- Cloud TPU
- Cloud Storage
| github_jupyter |
# 模型选择、欠拟合和过拟合
通过多项式拟合来交互地探索这些概念
```
import sys
sys.path.append('..')
import math
import numpy as np
import mindspore
from mindspore import nn
from d2l import mindspore as d2l
```
使用以下三阶多项式来生成训练和测试数据的标签:
$$y = 5 + 1.2x - 3.4\frac{x^2}{2!} + 5.6 \frac{x^3}{3!} + \epsilon \text{ where }
\epsilon \sim \mathcal{N}(0, 0.1^2)$$
```
max_degree = 20
n_train, n_test = 100, 100
true_w = np.zeros(max_degree)
true_w[0:4] = np.array([5, 1.2, -3.4, 5.6])
features = np.random.normal(size=(n_train + n_test, 1))
np.random.shuffle(features)
poly_features = np.power(features, np.arange(max_degree).reshape(1, -1))
for i in range(max_degree):
poly_features[:, i] /= math.gamma(i + 1)
labels = np.dot(poly_features, true_w)
labels += np.random.normal(scale=0.1, size=labels.shape)
```
看一下前2个样本
```
# true_w, features, poly_features, labels = [mindspore.Tensor(x, dtype=
# mindspore.float32) for x in [true_w, features, poly_features, labels]]
features[:2], poly_features[:2, :], labels[:2]
```
实现一个函数来评估模型在给定数据集上的损失
```
def evaluate_loss(net, dataset, loss):
"""评估给定数据集上模型的损失。"""
metric = d2l.Accumulator(2)
for X, y in dataset.create_tuple_iterator():
out = net(X)
y = y.reshape(out.shape)
l = loss(out, y)
metric.add(l.sum().asnumpy(), l.size)
return metric[0] / metric[1]
```
定义训练函数
```
def train(train_features, test_features, train_labels, test_labels,
num_epochs=400):
loss = nn.MSELoss()
input_shape = train_features.shape[-1]
net = nn.SequentialCell([nn.Dense(input_shape, 1, has_bias=False)])
batch_size = min(10, train_labels.shape[0])
train_dataset = d2l.load_array((train_features, train_labels.reshape(-1,1)),
batch_size)
test_dataset = d2l.load_array((test_features, test_labels.reshape(-1,1)),
batch_size, is_train=False)
optim = nn.SGD(net.trainable_params(), learning_rate=0.01)
animator = d2l.Animator(xlabel='epoch', ylabel='loss', yscale='log',
xlim=[1, num_epochs], ylim=[1e-3, 1e2],
legend=['train', 'test'])
for epoch in range(num_epochs):
d2l.train_epoch_ch3(net, train_dataset, loss, optim)
if epoch == 0 or (epoch + 1) % 20 == 0:
animator.add(epoch + 1, (evaluate_loss(net, train_dataset, loss),
evaluate_loss(net, test_dataset, loss)))
print('weight:', net[0].weight.data.asnumpy())
```
三阶多项式函数拟合(正态)
```
train(poly_features[:n_train, :4], poly_features[n_train:, :4],
labels[:n_train], labels[n_train:])
```
线性函数拟合(欠拟合)
```
train(poly_features[:n_train, :2], poly_features[n_train:, :2],
labels[:n_train], labels[n_train:])
```
高阶多项式函数拟合(过拟合)
```
train(poly_features[:n_train, :], poly_features[n_train:, :],
labels[:n_train], labels[n_train:], num_epochs=1500)
```
| github_jupyter |
## Exploratory Data Analysis - Sports
- Problem Statement: Perform Exploratory Data Analysis on 'Indian Premiere League' <br>
- As a sports analysts, find out the most successful teams, players and factors contributing win or loss of a team.<br>
- Suggest teams or players a company should endorse for its products.<br>
### Importing LIBRARIES:
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
### Loading 1st Dataset
```
matches = pd.read_csv("matches.csv")
matches.head()
matches.tail()
```
### Data information:
```
matches.info()
matches.shape
matches.describe()
```
### Loading 2nd Dataset
```
deliveries = pd.read_csv("deliveries.csv")
deliveries.head()
deliveries.tail()
```
### Data information:
```
deliveries.info()
deliveries.shape
deliveries.describe()
```
### Now, We will merge the 2 datasets for better insights from the data
```
#merging the 2 datasets
merge = pd.merge(deliveries,matches, left_on='match_id', right_on ='id')
merge.head(2)
merge.info()
merge.describe()
matches.id.is_unique
```
- Note:<br>
- since id is unique we can set this as our index
```
matches.set_index('id', inplace=True)
#Summary statistics of matches data
matches.describe(include = 'all')
```
### Data Preprocessing <br>
- Here we will perform Data Preprocessing on our matches dataset first, to make the data usable for EDA.
```
matches.head()
```
### From Pre profiling, we found that:
- city has missing values
- team1 and team2 columns have 14 distinct values but winner has 15 distinct values
- umpire1 and umpire2 have 1 missing value each
- umpire3 has 94% missing values
- city has 33 distinct values while venue has 35 distinct values
### Filling in the missing values of city column
### First let's find the venues corresponding to which the values of city are empty
```
matches[matches.city.isnull()][['city','venue']]
```
- So, missing values can be filled with Dubai
```
matches.city = matches.city.fillna('Dubai')
```
- umpire1 and umpire2 columns have one missing value each.
```
matches[(matches.umpire1.isnull()) | (matches.umpire2.isnull())]
```
- Umpire3 column has close to 92% missing values. hence dropping that column
```
matches = matches.drop('umpire3', axis = 1)
#city has 33 distinct values while we have 35 venues.
#Let's find out venues grouped by cities to see which cities have multiple venues
city_venue = matches.groupby(['city','venue']).count()['season']
city_venue_df = pd.DataFrame(city_venue)
city_venue_df
```
### Observations
- Bengaluru and Bangalore both are in the data when they are same. So we need to keep one of them
- Chandigarh and Mohali are same and there is just one stadium Punjab Cricket Association IS Bindra Stadium, Mohali whose value has not been entered correctly. We need to have either Chandigarh or Mohali as well as correct name of the stadium there
- Mumbai has 3 stadiums/venues used for IPL
- Pune has 2 venues for IPL<br>
### Visual representation of number of venues in each city .
```
#Plotting venues along with cities
v = pd.crosstab(matches['city'],matches['venue'])
v.replace(v[v!=0],1, inplace = True)
#Adding a column by summing each columns
v['count'] = v.sum(axis = 'columns')
#We will just keep last column = 'count'
b = v['count']
#Plotting
plt.figure(figsize = (20,7))
b.plot(kind = 'bar')
plt.title("Number of stadiums in different cities", fontsize = 25, fontweight = 'bold')
plt.xlabel("City", size = 30)
plt.ylabel("Frequency", size = 30)
plt.xticks(size = 20)
plt.yticks(size = 20)
```
### Exploratory Data Analysis:<br>
### Number of matches played in each season
```
plt.figure(figsize=(15,5))
sns.countplot('season', data = matches)
plt.title("Number of matches played each season",fontsize=18,fontweight="bold")
plt.ylabel("Count", size = 25)
plt.xlabel("Season", size = 25)
plt.xticks(size = 20)
plt.yticks(size = 20)
```
- 2011-2013 have more matches being played than other seasons
- All other seasons have approximately 58-60 matches while 2011-2013 have more than 70 matches.
### How many teams played in each season?
```
matches.groupby('season')['team1'].nunique().plot(kind = 'bar', figsize=(15,5))
plt.title("Number of teams participated each season ",fontsize=18,fontweight="bold")
plt.ylabel("Count of teams", size = 25)
plt.xlabel("Season", size = 25)
plt.xticks(size = 15)
plt.yticks(size = 15)
```
- 10 teams played in 2011 and 9 teams each in 2012 and 2013
- This explains why 2011-2013 have seen more matches being played than other seasons
### Venue which has hosted most number of IPL matches .
```
matches.venue.value_counts().sort_values(ascending = True).tail(10).plot(kind = 'barh',figsize=(12,8), fontsize=15, color='green')
plt.title("Venue which has hosted most number of IPL matches",fontsize=18,fontweight="bold")
plt.ylabel("Venue", size = 25)
plt.xlabel("Frequency", size = 25)
```
- M Chinnaswamy Stadium in Bengaluru has hosted the highest number of matches so far in IPL followed by Eden Gardens in Kolkata
### Which team has maximum wins in IPL so far?
```
#creating a dataframe with season and winner columns
winning_teams = matches[['season','winner']]
#dictionaries to get winners to each season
winners_team = {}
for i in sorted(winning_teams.season.unique()):
winners_team[i] = winning_teams[winning_teams.season == i]['winner'].tail(1).values[0]
winners_of_IPL = pd.Series(winners_team)
winners_of_IPL = pd.DataFrame(winners_of_IPL, columns=['team'])
winners_of_IPL['team'].value_counts().plot(kind = 'barh', figsize = (15,5), color = 'darkblue')
plt.title("Winners of IPL across 11 seasons",fontsize=18,fontweight="bold")
plt.ylabel("Teams", size = 25)
plt.xlabel("Frequency", size = 25)
plt.xticks(size = 15)
plt.yticks(size = 15)
```
- MI and CSK have both won 3 times each followed by KKR who has won 2 times.
- Hyderabad team has also won 2 matches under 2 franchise name - Deccan Chargers and Sunrisers Hyderabad
### Does teams choosed to bat or field first, after winning toss?
```
matches['toss_decision'].value_counts().plot(kind='pie', fontsize=14, autopct='%3.1f%%',
figsize=(10,7), shadow=True, startangle=135, legend=True, cmap='Oranges')
plt.ylabel('Toss Decision')
plt.title('Decision taken by captains after winning tosses')
```
- Close to 60% times teams who have won tosses have decided to chase down
### How toss decision affects match results?
```
matches['toss_win_game_win'] = np.where((matches.toss_winner == matches.winner),'Yes','No')
plt.figure(figsize = (15,5))
sns.countplot('toss_win_game_win', data=matches, hue = 'toss_decision')
plt.title("How Toss Decision affects match result", fontsize=18,fontweight="bold")
plt.xticks(size = 15)
plt.yticks(size = 15)
plt.xlabel("Winning Toss and winning match", fontsize = 25)
plt.ylabel("Frequency", fontsize = 25)
```
- Teams winning tosses and electng to field first have won most number of times.
### Individual teams decision to choose bat or field after winning toss.
```
plt.figure(figsize = (25,10))
sns.countplot('toss_winner', data = matches, hue = 'toss_decision')
plt.title("Teams decision to bat first or second after winning toss", size = 30, fontweight = 'bold')
plt.xticks(size = 10)
plt.yticks(size = 15)
plt.xlabel("Toss Winner", size = 35)
plt.ylabel("Count", size = 35)
```
- Most teams field first after winning toss except for Chennai Super Kings who has mostly opted to bat first. Deccan Chargers and Pune Warriors also show the same trend.
### Which player's performance has mostly led team's win?
```
MoM= matches['player_of_match'].value_counts()
MoM.head(10).plot(kind = 'bar',figsize=(12,8), fontsize=15, color='black')
plt.title("Top 10 players with most MoM awards",fontsize=18,fontweight="bold")
plt.ylabel("Frequency", size = 25)
plt.xlabel("Players", size = 25)
```
- Chris Gayle has so far won the most number of MoM awards followed by AB de Villiers.
- Also, all top 10 are batsmen which kind of hints that in IPL batsmen have mostly dictated the matches
### How winning matches by fielding first varies across venues?
```
new_matches = matches[matches['result'] == 'normal'] #taking all those matches where result is normal and creating a new dataframe
new_matches['win_batting_first'] = np.where((new_matches.win_by_runs > 0), 'Yes', 'No')
new_matches.groupby('venue')['win_batting_first'].value_counts().unstack().plot(kind = 'barh', stacked = True,
figsize=(15,15))
plt.title("How winning matches by fielding first varies across venues?", fontsize=18,fontweight="bold")
plt.xticks(size = 15)
plt.yticks(size = 15)
plt.xlabel("Frequency", fontsize = 25)
plt.ylabel("Venue", fontsize = 25)
```
- Batting second has been more rewarding in almost all the venues
### Is batting second advantageous across all years?
```
plt.figure(figsize = (15,5))
sns.countplot('season', data = new_matches, hue = 'win_batting_first')
plt.title("Is batting second advantageous across all years", fontsize=20,fontweight="bold")
plt.xticks(size = 15)
plt.yticks(size = 15)
plt.xlabel("Season", fontsize = 25)
plt.ylabel("Count", fontsize = 25)
```
- Exceptt for 2010 and 2015, in all other years it can be seen that teams batting second have won more matches
### Teams total scoring runs, over the years?
```
merge.groupby('season')['batsman_runs'].sum().plot(kind = 'line', linewidth = 3, figsize =(15,5),
color = 'blue')
plt.title("Runs over the years",fontsize= 25, fontweight = 'bold')
plt.xlabel("Season", size = 25)
plt.ylabel("Total Runs Scored", size = 25)
plt.xticks(size = 12)
plt.yticks(size = 12)
```
- Run scoring has gone up from the start of the IPL in 2008.
### Top Run Getters of IPL.
```
#let's plot the top 10 run getter so far in IPL
merge.groupby('batsman')['batsman_runs'].sum().sort_values(ascending = False).head(10).plot(kind = 'bar', color = 'red',
figsize = (15,5))
plt.title("Top Run Getters of IPL", fontsize = 20, fontweight = 'bold')
plt.xlabel("Batsmen", size = 25)
plt.ylabel("Total Runs Scored", size = 25)
plt.xticks(size = 12)
plt.yticks(size = 12)
```
- Except for MS Dhoni, all other top run getters are either openers or come in 3rd or 4th positions to bat
- Suresh Raina is the highest run getter in IPL.
### Which batsman has been most consistent among top 10 run getters?
```
consistent_batsman = merge[merge.batsman.isin(['SK Raina', 'V Kohli','RG Sharma','G Gambhir',
'RV Uthappa', 'S Dhawan','CH Gayle', 'MS Dhoni',
'DA Warner', 'AB de Villiers'])][['batsman','season','total_runs']]
consistent_batsman.groupby(['season','batsman'])['total_runs'].sum().unstack().plot(kind = 'box', figsize = (15,8))
plt.title("Most Consistent batsmen of IPL", fontsize = 20, fontweight = 'bold')
plt.xlabel("Batsmen", size = 25)
plt.ylabel("Total Runs Scored each season", size = 25)
plt.xticks(size = 15)
plt.yticks(size = 15)
```
- Median score for Raina is above all the top 10 run getters. He has the highest lowest run among all the batsmen across 11 seasons. Considering the highest and lowest season totals and spread of runs, it seems Raina has been most consistent among all.
### Which bowlers have performed the best?
```
merge.groupby('bowler')['player_dismissed'].count().sort_values(ascending = False).head(10).plot(kind = 'bar',
color = 'purple', figsize = (15,5))
plt.title("Top Wicket Takers of IPL", fontsize = 20, fontweight = 'bold')
plt.xlabel("Bowler", size = 25)
plt.ylabel("Total Wickets Taken", size = 25)
plt.xticks(size = 12)
plt.yticks(size = 12)
```
- Malinga has taken the most number of wickets in IPL followed by Bravo and Amit Mishra
- In top 10 bowlers, 5 are fast and medium pacers while the other 5 are spinners
- All 5 spinners are right arm spinners and 2 are leg spinners while 3 are off spinners
- All 5 pacers are right arm pacers
### Batsmen with the best strike rates over the years .
```
#We will consider players who have played 10 or more seasons
no_of_balls = pd.DataFrame(merge.groupby('batsman')['ball'].count()) #total number of matches played by each batsman
runs = pd.DataFrame(merge.groupby('batsman')['batsman_runs'].sum()) #total runs of each batsman
seasons = pd.DataFrame(merge.groupby('batsman')['season'].nunique()) #season = 1 implies played only 1 season
batsman_strike_rate = pd.DataFrame({'balls':no_of_balls['ball'],'run':runs['batsman_runs'],'season':seasons['season']})
batsman_strike_rate.reset_index(inplace = True)
batsman_strike_rate['strike_rate'] = batsman_strike_rate['run']/batsman_strike_rate['balls']*100
highest_strike_rate = batsman_strike_rate[batsman_strike_rate.season.isin([10,11])][['season','batsman','strike_rate']].sort_values(by = 'strike_rate',
ascending = False)
highest_strike_rate.head(10)
plt.figure(figsize = (15,6))
sns.barplot(x='batsman', y='strike_rate', data = highest_strike_rate.head(10), hue = 'season')
plt.title("Highest strike rates in IPL",fontsize= 30, fontweight = 'bold')
plt.xlabel("Player", size = 25)
plt.ylabel("Strike Rate", size = 25)
plt.xticks(size = 14)
plt.yticks(size = 14)
```
- AB de Villiers, Gayle have the highest strike rates in IPL. They are the big hitters and can win any match on their day
- One surprise here is that Harbhajan Singh who is a bowler has a strike rate of 130+ and comes before Rohit Sharma in ranking
### Bowlers with maximum number of extras.
```
extra = deliveries[deliveries['extra_runs']!=0]['bowler'].value_counts()[:10]
extra.plot(kind='bar', figsize=(11,6), title='Bowlers who have bowled maximum number of Extra balls')
plt.xlabel('BOWLER')
plt.ylabel('BALLS')
plt.show()
extra = pd.DataFrame(extra)
extra.T
```
### Which bowlers have picked up wickets more frequently?
```
#strike_rate = balls bowled by wickets taken
balls_bowled = pd.DataFrame(merge.groupby('bowler')['ball'].count())
wickets_taken = pd.DataFrame(merge[merge['dismissal_kind'] != 'no dismissal'].groupby('bowler')['dismissal_kind'].count())
seasons_played = pd.DataFrame(merge.groupby('bowler')['season'].nunique())
bowler_strike_rate = pd.DataFrame({'balls':balls_bowled['ball'],'wickets':wickets_taken['dismissal_kind'],
'season':seasons_played['season']})
bowler_strike_rate.reset_index(inplace = True)
bowler_strike_rate['strike_rate'] = bowler_strike_rate['balls']/bowler_strike_rate['wickets']
def highlight_cols(s):
color = 'skyblue'
return 'background-color: %s' % color
#Strike rate for bowlers who have taken more than 50 wickets
best_bowling_strike_rate = bowler_strike_rate[bowler_strike_rate['wickets'] > 50].sort_values(by = 'strike_rate', ascending = True)
best_bowling_strike_rate.head().style.applymap(highlight_cols, subset=pd.IndexSlice[:, ['bowler', 'wickets','strike_rate']])
```
### Q1. As a sports analysts, The most successful teams, players & factors contributing win or loss of a team:
- Mumbai Indians is the most successful team in IPL and has won the most number of toss.
- There were more matches won by chasing the total(419 matches) than defending(350 matches).
- When defending a total, the biggest victory was by 146 runs(Mumbai Indians defeated Delhi Daredevils by 146 runs on 06 May 2017 at Feroz Shah Kotla stadium, Delhi).
- When chasing a target, the biggest victory was by 10 wickets(without losing any wickets) and there were 11 such instances.
- The Mumbai city has hosted the most number of IPL matches.
- Chris Gayle has won the maximum number of player of the match title.
- S. Ravi(Sundaram Ravi) has officiated the most number of IPL matches on-field.
- Eden Gardens has hosted the maximum number of IPL matches.
- If a team wins a toss choose to field first as it has highest probablity of winning
### Q2. Teams or Players a company should endorse for its products.
- If the franchise is looking for a consistant batsman who needs to score good amount of runs then go for V Kohli, S Raina, Rohit Sharma , David Warner...
- If the franchise is looking for a game changing batsman then go for Chris Gayle, AB deVillers, R Sharma , MS Dhoni...
- If the franchise is looking for a batsman who could score good amount of runs every match the go for DA Warner, CH Gayle, V Kohli,AB de Villiers,S Dhawan
- If the franchise needs the best finisher in lower order having good strike rate then go for CH Gayle,KA Pollard, DA Warner,SR Watson,BB McCullum
- If the franchise need a experienced bowler then go for Harbhajan Singh ,A Mishra,PP Chawla ,R Ashwin,SL Malinga,DJ Bravo
- If the franchise need a wicket taking bowler then go for SL Malinga,DJ Bravo,A Mishra ,Harbhajan Singh, PP Chawla
- If the franchise need a bowler bowling most number of dot balls then go for Harbhajan Singh,SL Malinga,B Kumar,A Mishra,PP Chawla
- If the franchise need a bowler with good economy then go for DW Steyn ,M Muralitharan ,R Ashwin,SP Narine ,Harbhajan Singh
| github_jupyter |
This notebook is part of https://github.com/AudioSceneDescriptionFormat/splines, see also https://splines.readthedocs.io/.
[back to rotation splines](index.ipynb)
# Naive 4D Quaternion Interpolation
This method for interpolating rotations is normally not recommended.
But it might still be interesting to try it out ...
Since quaternions form a vector space (albeit a four-dimensional one),
all methods for [Euclidean splines](../euclidean/index.ipynb) can be applied.
However, even though rotations can be represented by *unit* quaternions,
which are a subset of all quaternions,
this subset is *not* a Euclidean space.
All *unit* quaternions form the unit hypersphere $S^3$
(which is a curved space),
and each point on this hypersphere uniquely corresponds to a rotation.
When we convert our desired rotation "control points"
to quaternions and naively interpolate in 4D quaternion space,
the interpolated quaternions are in general *not* unit quaternions,
i.e. they are not part of the unit hypersphere and
they don't correspond to a rotation.
In order to force them onto the unit hypersphere,
we can normalize them, though,
which projects them onto the unit hypersphere.
Note that this is a very crude form of interpolation
and it might result in unexpected curve shapes.
Especially the temporal behavior might be undesired.
> If, for some application, more speed is essential,
> non-spherical quaternion splines will undoubtedly be faster
> than angle interpolation,
> while still free of axis bias and gimbal lock.
>
> --<cite data-cite="shoemake1985animating">Shoemake (1985)</cite>, section 5.4
> Abandoning the unit sphere,
> one could work with the four-dimensional Euclidean space
> of arbitrary quaternions.
> How do standard interpolation methods applied there
> behave when mapped back to matrices?
> Note that we now have little guidance in picking the inverse image for a matrix,
> and that cusp-free $\mathbf{R}^4$ paths
> do not always project to cusp-free $S^3$ paths.
>
> --<cite data-cite="shoemake1985animating">Shoemake (1985)</cite>, section 6
```
import numpy as np
import splines
from splines.quaternion import Quaternion
```
As always, we use a few helper functions from [helper.py](helper.py):
```
from helper import angles2quat, animate_rotations, display_animation
rotations = [
angles2quat(0, 0, 0),
angles2quat(0, 0, 45),
angles2quat(90, 90, 0),
angles2quat(180, 0, 90),
]
```
We use `xyzw` coordinate order here
(because it is more common),
but since the 4D coordinates are independent,
we could as well use `wxyz` order
(or any order, for that matter) with identical results
(apart from rounding errors).
However, for illustrating the non-normalized case,
we rely on the implicit conversion from `xyzw` coordinates
in the function `animate_rotations()`.
```
rotations_xyzw = [q.xyzw for q in rotations]
```
As an example we use
[splines.CatmullRom](../python-module/splines.rst#splines.CatmullRom)
here, but any Euclidean spline could be used.
```
s = splines.CatmullRom(rotations_xyzw, endconditions='closed')
times = np.linspace(s.grid[0], s.grid[-1], 100)
interpolated_xyzw = s.evaluate(times)
normalized = [
Quaternion(w, (x, y, z)).normalized()
for x, y, z, w in interpolated_xyzw]
```
For comparison, we also create a
[splines.quaternion.CatmullRom](../python-module/splines.quaternion.rst#splines.quaternion.CatmullRom)
instance:
```
spherical_cr = splines.quaternion.CatmullRom(rotations, endconditions='closed')
ani = animate_rotations({
'normalized 4D interp.': normalized,
'spherical interp.': spherical_cr.evaluate(times),
}, figsize=(5, 2))
display_animation(ani, default_mode='loop')
```
In case you are wondering what would happen
if you forget to normalize the results,
let's also show the non-normalized data:
```
ani = animate_rotations({
'normalized': normalized,
'not normalized': interpolated_xyzw,
}, figsize=(5, 2))
display_animation(ani, default_mode='loop')
```
Obviously, the non-normalized values are not pure rotations.
To get a different temporal behavior,
let's try using [centripetal parameterization](../euclidean/catmull-rom-properties.ipynb#Centripetal-Parameterization).
Note that this guarantees the absence
of cusps and self-intersections
in the 4D curve,
but this guarantee doesn't extend to
the projection onto the unit hypersphere.
```
s2 = splines.CatmullRom(rotations_xyzw, alpha=0.5, endconditions='closed')
times2 = np.linspace(s2.grid[0], s2.grid[-1], len(times))
normalized2 = [
Quaternion(w, (x, y, z)).normalized()
for x, y, z, w in s2.evaluate(times2)]
ani = animate_rotations({
'uniform': normalized,
'centripetal': normalized2,
}, figsize=(5, 2))
display_animation(ani, default_mode='loop')
```
Let's also try *arc-length parameterization* with the
[ConstantSpeedAdapter](../python-module/splines.rst#splines.ConstantSpeedAdapter):
```
s3 = splines.ConstantSpeedAdapter(s2)
times3 = np.linspace(s3.grid[0], s3.grid[-1], len(times))
normalized3 = [
Quaternion(w, (x, y, z)).normalized()
for x, y, z, w in s3.evaluate(times3)]
```
The arc-length parameterized spline
has a constant speed in 4D quaternion space,
but that doesn't mean it has a constant angular speed!
For comparison,
we also create a rotation spline with constant angular speed:
```
s4 = splines.ConstantSpeedAdapter(
splines.quaternion.CatmullRom(
rotations, alpha=0.5, endconditions='closed'))
times4 = np.linspace(s4.grid[0], s4.grid[-1], len(times))
ani = animate_rotations({
'constant 4D speed': normalized3,
'constant angular speed': s4.evaluate(times4),
}, figsize=(5, 2))
display_animation(ani, default_mode='loop')
```
The difference is subtle, but it is definitely visible.
More extreme examples can certainly be found.
| github_jupyter |
# Parametrized Quantum Circuits
In this section, we introduce parameterised quantum circuits, then describe their properties and implement some examples used in quantum machine learning.
## Contents
1. [Introduction](#Introduction)
1. [Parameterized Quantum Circuit Properties](#Parametrized-Quantum-Circuit-Properties)
1. [Expressibility](#Expressibility)
1. [Entangling-Capability](#Entangling-Capability)
1. [Parameterized Quantum Circuits for Machine Learning](#Parameterized-Quantum-Circuits-for-Machine-Learning)
1. [References](#References)
## Introduction
As we saw in the [variational quantum classifier](vqc.ipynb) example in the [introduction](introduction.ipynb), parameterized quantum circuits, where the gates are defined through tunable parameters, are a fundamental building block of near-term quantum machine learning algorithms. In the literature, depending on the context, parameterized quantum circuits are also called parameterised trial states, variational forms or ansatze.
Here is an example of a simple parameterised circuit, with two parameterised gates, a single-qubit $z$-rotation gate, with a variable rotation $\theta$, followed by a two-qubit controlled $z$-rotation gate, with the same variable rotation $\theta$:
<figure>
<img src="images/pqc.png" style="width:150px">
</figure>
The parameterized circuit above can be created in Qiskit using the `QuantumCircuit` and `Parameter` classes. This allows us to create a circuit with rotation gates, without having to specify what the angles are yet:
```
from qiskit.circuit import QuantumCircuit, Parameter
theta = Parameter('θ')
qc = QuantumCircuit(2)
qc.rz(theta,0)
qc.crz(theta,0,1)
qc.draw(output="mpl")
```
If we want the gates to have different parameters, we can use two `Parameters`, or we create a `ParameterVector`, which acts like a list of `Parameters`:
```
from qiskit.circuit import ParameterVector
theta_list = ParameterVector('θ', length=2)
qc = QuantumCircuit(2)
qc.rz(theta_list[0],0)
qc.crz(theta_list[1],0,1)
qc.draw(output="mpl")
```
As all quantum gates used in a quantum circuit are [unitary](gloss:unitary), a parametrized circuit itself can be described as a unitary operation on $n$ qubits, $\mathbf{U_\theta}$, acting on some initial state $|\phi_0\rangle$, often set to $|0\rangle^{\otimes n}$. The resulting parameterized quantum state is $|\psi_\mathbf{\theta}\rangle = \mathbf{U_\theta} |\phi_0\rangle$ where $\mathbf{\theta}$ is a set of tunable parameters.
<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-pqc-0") -->
<!-- ::: .question -->
Which of these Qiskit gates can be parametrized?
<!-- ::: -->
<!-- ::: .option -->
1. <code>XGate</code>
<!-- ::: -->
<!-- ::: .option(correct) -->
2. <code>PhaseGate</code>
<!-- ::: -->
<!-- ::: .option -->
3. <code>HGate</code>
<!-- ::: -->
<!-- ::: -->
<!-- ::: -->
## Parametrized Quantum Circuit Properties
How do we choose one parameterized circuit over the other? To use parameterised quantum circuits as a machine learning model, we need them to generalize well. This means that the selected circuit should be able to generate a significant subset of the states within the output Hilbert space. To avoid being easy to simulate on a classical computer, the selected circuit should ideally also entangle qubits.
In Reference [1](#references), the authors propose the measures of *expressibility* and *entangling capability* to discriminate between different parameterised quantum circuits. The *expressibility* of a circuit can be understood as the extent to which a parametrized circuit is able to generate states within the Hilbert space, while the *entangling capability* of a circuit describes its ability to generate entangled states.
### *Expressibility*
*Expressibility* of a parameterizes quantum circuit is quantified in Reference [1](#references) by computing the extent to which the states generated from the circuit deviate from the [uniform distribution](gloss:uniform_dist).
Let's compare the two following single-qubit circuits to get a sense of what it means for a circuit to be expressible or not, by sampling 2000 output states for each circuit and plotting them on a Bloch sphere.
<figure>
<img src="images/expr.png" style="width:400px">
</figure>
```
import numpy as np
import matplotlib.pyplot as plt
import qiskit.visualization as qv
import qiskit.quantum_info as qi
```
First, we need to define the circuits:
```
theta_param = Parameter('θ')
phi_param = Parameter('Φ')
# Circuit A
qc_A = QuantumCircuit(1)
qc_A.h(0)
qc_A.rz(theta_param,0)
qc_A.draw(output="mpl")
# Circuit B
qc_B=QuantumCircuit(1)
qc_B.h(0)
qc_B.rz(theta_param,0)
qc_B.rx(phi_param,0)
qc_B.draw(output="mpl")
```
Next we uniformly sample the parameter space for the two parameters $\theta$ and $\phi$:
```
np.random.seed(0)
num_param = 2000
theta = [2*np.pi*np.random.uniform() for i in range(num_param)]
phi = [2*np.pi*np.random.uniform() for i in range(num_param)]
```
Then we take the parameter value lists, build the state vectors corresponding to each circuit and plot them on the Bloch sphere:
```
# Converts state vectors to points on the Bloch sphere
def state_to_bloch(state_vec):
phi = np.angle(state_vec.data[1])-np.angle(state_vec.data[0])
theta = 2*np.arccos(np.abs(state_vec.data[0]))
return [np.sin(theta)*np.cos(phi),np.sin(theta)*np.sin(phi),np.cos(theta)]
# Bloch sphere plot formatting
width, height = plt.figaspect(1/2)
fig=plt.figure(figsize=(width, height))
ax1, ax2 = fig.add_subplot(1, 2, 1, projection='3d'), fig.add_subplot(1, 2, 2, projection='3d')
b1,b2 = qv.bloch.Bloch(axes=ax1), qv.bloch.Bloch(axes=ax2)
b1.point_color, b2.point_color = ['tab:blue'],['tab:blue']
b1.point_marker, b2.point_marker= ['o'],['o']
b1.point_size, b2.point_size=[2],[2]
# Calculate state vectors for circuit A and circuit B for each set of sampled parameters
# and add to their respective Bloch sphere
for i in range(num_param):
state_1=qi.Statevector.from_instruction(qc_A.bind_parameters({theta_param:theta[i]}))
state_2=qi.Statevector.from_instruction(qc_B.bind_parameters({theta_param:theta[i], phi_param:phi[i]}))
b1.add_points(state_to_bloch(state_1))
b2.add_points(state_to_bloch(state_2))
b1.show()
b2.show()
```
For Circuit A, with a Hadamard gate followed by a parameterizes $z$-rotation gate, we observe output states distributed about the equator of the Bloch sphere (left). With Circuit B, with the additional parameterised $z$-rotation gate, we are able to cover all the Bloch sphere with output states, but the coverage is not uniform; there are concentrations of points on the $+X$ and $-X$ poles of the Bloch sphere.
Expressibility is a way to quantify what we just observed for these single-qubit circuits. Circuit A would have a low expressibility score due to the limited set of states it can explore, whereas Circuit B would have a high expressibility score.
### *Entangling Capability*
Entanglement is often seen as a key resource in quantum computing. The Meyer-Wallach measure is one of many metrics that can be used quantify how entangled a given state is. The Meyer-Wallach measure of an unentangled [product state](gloss:product_state) is 0, while the Meyer-Wallach measure of a highly entangled state, such as the Bell state, is 1. In Reference [1](#references), the authors define the entangling capability of a parameterized quantum circuit as the average Meyer-Wallach measure for the states it can generate.
For example, consider the following multi-qubit parameterized circuits:
<figure>
<img src="images/ent.png" style="width:300px">
</figure>
Circuit A has no entangling operations, that is, no two-qubit gates, so it has no entangling capability. Thus, this circuit would produce states that have Meyer-Wallach measures of 0, leading to an averaged value of 0. Circuit B has several two-qubit gates and thus is able to generate quantum states with some entanglement. Therefore, the average Meyer-Wallach measure would be greater than 0.
Using these two parameterized quantum circuit descriptors, expressibility and entangling capability, we can identify circuits that have limited capabilities. We expect that limited circuits are poor candidates for quantum machine learning applications. This was investigated in Reference [2](#references) for the variational quantum classifier; and for their studied datasets and parameterized circuits, it was found that there was a strong correlation between classification accuracy and expressibility, and a weak correlation between classification accuracy entangling capability.
<!-- ::: q-block.exercise -->
### Quick quiz
<!-- ::: q-quiz(goal="qml-pqc-1") -->
<!-- ::: .question -->
Why is entangling capabilty desirable in a quantum circuit?
<!-- ::: -->
<!-- ::: .option(correct) -->
1. Circuits that don't entangle qubits are easy to simulate classically.
<!-- ::: -->
<!-- ::: .option -->
2. Entangling qubits reduces errors in the hardware.
<!-- ::: -->
<!-- ::: -->
<!-- ::: -->
## Parameterized Quantum Circuits for Machine Learning
Here we will give some examples of how to build parameterized quantum circuits used in the quantum machine learning literature in Qiskit.
In Reference [3](#references), the authors introduce the following parameterized circuit, which we will study in detail [later](kernel.ipynb):
$$\mathcal{U}_{\Phi(\mathbf{x})}=\prod_d U_{\Phi(\mathbf{x})}H^{\otimes n},\ U_{\Phi(\mathbf{x})}=\exp\left(i\sum_{S\subseteq[n]}\phi_S(\mathbf{x})\prod_{k\in S} P_i\right), $$
which contains layers of Hadamard gates interleaved with entangling blocks, $U_{\Phi(\mathbf{x})}$. Within the entangling blocks, $U_{\Phi(\mathbf{x})}$: $P_i \in \{ I, X, Y, Z \}$ denotes the Pauli matrices, the index $S$ describes connectivities between different qubits: $S \in \{\binom{n}{k}\ \text{combinations},\ k = 1,... n \}$, and the data mapping function $\phi_S(\mathbf{x})$ is
$$\phi_S:\mathbf{x}\mapsto \Bigg\{\begin{array}{ll}
x_i & \mbox{if}\ S=\{i\} \\
(\pi-x_i)(\pi-x_j) & \mbox{if}\ S=\{i,j\}
\end{array}$$
Specifically, they use $k = 2, P_0 = Z, P_1 = ZZ$, which in Qiskit is the `ZZFeatureMap` circuit:
```
from qiskit.circuit.library import ZZFeatureMap
qc_zz = ZZFeatureMap(3, reps=1, insert_barriers=True)
print(qc_zz.parameters)
qc_zz.draw(output='mpl')
```
In Reference [3](#references), the authors also use a hardware efficient circuit consisting of alternating layers of single-qubit rotation gates, followed by two-qubit gates. In particular, they use $y$-and-$z$-rotation gates, and controlled-$z$ gates, which in Qiskit, can be built using the `TwoLocal` circuit:
```
from qiskit.circuit.library import TwoLocal
qc_twolocal = TwoLocal(num_qubits=3, reps=2, rotation_blocks=['ry','rz'],
entanglement_blocks='cz', skip_final_rotation_layer=True,
insert_barriers=True)
print(qc_twolocal.parameters)
qc_twolocal.draw('mpl')
```
The `TwoLocal` circuit in Qiskit can be used to make many parameterised circuits, such as circuit 13 in Reference [1](#references):
```
qc_13 = TwoLocal(4, rotation_blocks='ry', entanglement_blocks='crz', entanglement='sca',
reps=3, skip_final_rotation_layer=True, insert_barriers=True)
print(qc_13.parameters)
qc_13.draw('mpl')
```
The `NLocal` circuit can also be used to create more general parameterized circuits with alternating rotation and entanglement layers.
Here is a `NLocal` circuit, with a rotation block on 2 qubits and an entanglement block on 4 qubits using linear entanglement:
```
from qiskit.circuit.library import NLocal
# rotation block:
rot = QuantumCircuit(2)
params = ParameterVector('r', 2)
rot.ry(params[0], 0)
rot.rz(params[1], 1)
# entanglement block:
ent = QuantumCircuit(4)
params = ParameterVector('e', 3)
ent.crx(params[0], 0, 1)
ent.crx(params[1], 1, 2)
ent.crx(params[2], 2, 3)
qc_nlocal = NLocal(num_qubits=6, rotation_blocks=rot, entanglement_blocks=ent,
entanglement='linear', skip_final_rotation_layer=True, insert_barriers=True)
print(qc_nlocal.parameters)
qc_nlocal.draw('mpl')
```
<!-- ::: q-block.exercise -->
### Try it yourself
Create a circuit that [estimates the phase](https://qiskit.org/textbook/ch-algorithms/quantum-phase-estimation.html) introduced by a parametrized RZGate acting on the state $|1\rangle$. Can you plot a graph of the parameter value vs the measured phase?
Try this in the [IBM Quantum Lab](https://quantum-computing.ibm.com/)
<!-- ::: -->
## References <a id="references"></a>
1. Sukin Sim, Peter D. Johnson and Alan Aspuru-Guzik, *Expressibility and entangling capability of parameterized quantum circuits for hybrid quantum-classical algorithms*, Advanced Quantum Technology 2 (2019) 1900070, [doi:10.1002/qute.201900070](https://onlinelibrary.wiley.com/doi/abs/10.1002/qute.201900070), [arXiv:1905.10876](https://arxiv.org/abs/1905.10876).
2. Thomas Hubregtsen, Josef Pichlmeier, and Koen Bertels, *Evaluation of Parameterized Quantum Circuits: On the Design, and the Relation between Classification Accuracy, Expressibility and Entangling Capability* 2020, [arXiv:2003.09887](https://arxiv.org/abs/2003.09887).
3. Vojtech Havlicek, Antonio D. Córcoles, Kristan Temme, Aram W. Harrow, Abhinav Kandala, Jerry M. Chow and Jay M. Gambetta, *Supervised learning with quantum enhanced feature spaces*, Nature 567, 209-212 (2019), [doi.org:10.1038/s41586-019-0980-2](https://doi.org/10.1038/s41586-019-0980-2), [arXiv:1804.11326](https://arxiv.org/abs/1804.11326).
```
import qiskit.tools.jupyter
%qiskit_version_table
```
| github_jupyter |
# Convolutional Networks
So far we have worked with deep fully-connected networks, using them to explore different optimization strategies and network architectures. Fully-connected networks are a good testbed for experimentation because they are very computationally efficient, but in practice all state-of-the-art results use convolutional networks instead.
First you will implement several layer types that are used in convolutional networks. You will then use these layers to train a convolutional network on the CIFAR-10 dataset.
```
# As usual, a bit of setup
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from nnlib.classifiers.cnn import *
from nnlib.data_utils import get_CIFAR10_data
from nnlib.gradient_check import eval_numerical_gradient_array, eval_numerical_gradient
from nnlib.layers import *
from nnlib.fast_layers import *
from nnlib.solver import Solver
%matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading external modules
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
%load_ext autoreload
%autoreload 2
def rel_error(x, y):
""" returns relative error """
return np.max(np.abs(x - y) / (np.maximum(1e-8, np.abs(x) + np.abs(y))))
# Load the (preprocessed) CIFAR10 data.
data = get_CIFAR10_data()
for k, v in data.items():
print('%s: ' % k, v.shape)
```
# Convolution: Naive forward pass
The core of a convolutional network is the convolution operation. In the file `nnlib/layers.py`, implement the forward pass for the convolution layer in the function `conv_forward_naive`.
You don't have to worry too much about efficiency at this point; just write the code in whatever way you find most clear.
You can test your implementation by running the following:
```
x_shape = (2, 3, 4, 4)
w_shape = (3, 3, 4, 4)
x = np.linspace(-0.1, 0.5, num=np.prod(x_shape)).reshape(x_shape)
w = np.linspace(-0.2, 0.3, num=np.prod(w_shape)).reshape(w_shape)
b = np.linspace(-0.1, 0.2, num=3)
conv_param = {'stride': 2, 'pad': 1}
out, _ = conv_forward_naive(x, w, b, conv_param)
correct_out = np.array([[[[-0.08759809, -0.10987781],
[-0.18387192, -0.2109216 ]],
[[ 0.21027089, 0.21661097],
[ 0.22847626, 0.23004637]],
[[ 0.50813986, 0.54309974],
[ 0.64082444, 0.67101435]]],
[[[-0.98053589, -1.03143541],
[-1.19128892, -1.24695841]],
[[ 0.69108355, 0.66880383],
[ 0.59480972, 0.56776003]],
[[ 2.36270298, 2.36904306],
[ 2.38090835, 2.38247847]]]])
# Compare your output to ours; difference should be around 2e-8
print('Testing conv_forward_naive')
print('difference: ', rel_error(out, correct_out))
```
# Aside: Image processing via convolutions
As fun way to both check your implementation and gain a better understanding of the type of operation that convolutional layers can perform, we will set up an input containing two images and manually set up filters that perform common image processing operations (grayscale conversion and edge detection). The convolution forward pass will apply these operations to each of the input images. We can then visualize the results as a sanity check.
```
from scipy.misc import imread, imresize
kitten, puppy = imread('kitten.jpg'), imread('puppy.jpg')
# kitten is wide, and puppy is already square
d = kitten.shape[1] - kitten.shape[0]
kitten_cropped = kitten[:, d//2:-d//2, :]
img_size = 200 # Make this smaller if it runs too slow
x = np.zeros((2, 3, img_size, img_size))
x[0, :, :, :] = imresize(puppy, (img_size, img_size)).transpose((2, 0, 1))
x[1, :, :, :] = imresize(kitten_cropped, (img_size, img_size)).transpose((2, 0, 1))
# Set up a convolutional weights holding 2 filters, each 3x3
w = np.zeros((2, 3, 3, 3))
# The first filter converts the image to grayscale.
# Set up the red, green, and blue channels of the filter.
w[0, 0, :, :] = [[0, 0, 0], [0, 0.3, 0], [0, 0, 0]]
w[0, 1, :, :] = [[0, 0, 0], [0, 0.6, 0], [0, 0, 0]]
w[0, 2, :, :] = [[0, 0, 0], [0, 0.1, 0], [0, 0, 0]]
# Second filter detects horizontal edges in the blue channel.
w[1, 2, :, :] = [[1, 2, 1], [0, 0, 0], [-1, -2, -1]]
# Vector of biases. We don't need any bias for the grayscale
# filter, but for the edge detection filter we want to add 128
# to each output so that nothing is negative.
b = np.array([0, 128])
# Compute the result of convolving each input in x with each filter in w,
# offsetting by b, and storing the results in out.
out, _ = conv_forward_naive(x, w, b, {'stride': 1, 'pad': 1})
def imshow_noax(img, normalize=True):
""" Tiny helper to show images as uint8 and remove axis labels """
if normalize:
img_max, img_min = np.max(img), np.min(img)
img = 255.0 * (img - img_min) / (img_max - img_min)
plt.imshow(img.astype('uint8'))
plt.gca().axis('off')
# Show the original images and the results of the conv operation
plt.subplot(2, 3, 1)
imshow_noax(puppy, normalize=False)
plt.title('Original image')
plt.subplot(2, 3, 2)
imshow_noax(out[0, 0])
plt.title('Grayscale')
plt.subplot(2, 3, 3)
imshow_noax(out[0, 1])
plt.title('Edges')
plt.subplot(2, 3, 4)
imshow_noax(kitten_cropped, normalize=False)
plt.subplot(2, 3, 5)
imshow_noax(out[1, 0])
plt.subplot(2, 3, 6)
imshow_noax(out[1, 1])
plt.show()
```
# Convolution: Naive backward pass
Implement the backward pass for the convolution operation in the function `conv_backward_naive` in the file `nnlib/layers.py`. Again, you don't need to worry too much about computational efficiency.
When you are done, run the following to check your backward pass with a numeric gradient check.
```
np.random.seed(231)
x = np.random.randn(4, 3, 5, 5)
w = np.random.randn(2, 3, 3, 3)
b = np.random.randn(2,)
dout = np.random.randn(4, 2, 5, 5)
conv_param = {'stride': 1, 'pad': 1}
dx_num = eval_numerical_gradient_array(lambda x: conv_forward_naive(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_forward_naive(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_forward_naive(x, w, b, conv_param)[0], b, dout)
out, cache = conv_forward_naive(x, w, b, conv_param)
dx, dw, db = conv_backward_naive(dout, cache)
# Your errors should be around 1e-8'
print('Testing conv_backward_naive function')
print('dx error: ', rel_error(dx, dx_num))
print('dw error: ', rel_error(dw, dw_num))
print('db error: ', rel_error(db, db_num))
```
# Max pooling: Naive forward
Implement the forward pass for the max-pooling operation in the function `max_pool_forward_naive` in the file `nnlib/layers.py`. Again, don't worry too much about computational efficiency.
Check your implementation by running the following:
```
x_shape = (2, 3, 4, 4)
x = np.linspace(-0.3, 0.4, num=np.prod(x_shape)).reshape(x_shape)
pool_param = {'pool_width': 2, 'pool_height': 2, 'stride': 2}
out, _ = max_pool_forward_naive(x, pool_param)
correct_out = np.array([[[[-0.26315789, -0.24842105],
[-0.20421053, -0.18947368]],
[[-0.14526316, -0.13052632],
[-0.08631579, -0.07157895]],
[[-0.02736842, -0.01263158],
[ 0.03157895, 0.04631579]]],
[[[ 0.09052632, 0.10526316],
[ 0.14947368, 0.16421053]],
[[ 0.20842105, 0.22315789],
[ 0.26736842, 0.28210526]],
[[ 0.32631579, 0.34105263],
[ 0.38526316, 0.4 ]]]])
# Compare your output with ours. Difference should be around 1e-8.
print('Testing max_pool_forward_naive function:')
print('difference: ', rel_error(out, correct_out))
```
# Max pooling: Naive backward
Implement the backward pass for the max-pooling operation in the function `max_pool_backward_naive` in the file `nnlib/layers.py`. You don't need to worry about computational efficiency.
Check your implementation with numeric gradient checking by running the following:
```
np.random.seed(231)
x = np.random.randn(3, 2, 8, 8)
dout = np.random.randn(3, 2, 4, 4)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
dx_num = eval_numerical_gradient_array(lambda x: max_pool_forward_naive(x, pool_param)[0], x, dout)
out, cache = max_pool_forward_naive(x, pool_param)
dx = max_pool_backward_naive(dout, cache)
# Your error should be around 1e-12
print('Testing max_pool_backward_naive function:')
print('dx error: ', rel_error(dx, dx_num))
```
# Fast layers
Making convolution and pooling layers fast can be challenging. To spare you the pain, we've provided fast implementations of the forward and backward passes for convolution and pooling layers in the file `nnlib/fast_layers.py`.
The fast convolution implementation depends on a Cython extension; to compile it you need to run the following from the `nnlib` directory:
```bash
python setup.py build_ext --inplace
```
The API for the fast versions of the convolution and pooling layers is exactly the same as the naive versions that you implemented above: the forward pass receives data, weights, and parameters and produces outputs and a cache object; the backward pass recieves upstream derivatives and the cache object and produces gradients with respect to the data and weights.
**NOTE:** The fast implementation for pooling will only perform optimally if the pooling regions are non-overlapping and tile the input. If these conditions are not met then the fast pooling implementation will not be much faster than the naive implementation.
You can compare the performance of the naive and fast versions of these layers by running the following:
```
from nnlib.fast_layers import conv_forward_fast, conv_backward_fast
from time import time, clock
np.random.seed(231)
x = np.random.randn(100, 3, 31, 31)
w = np.random.randn(25, 3, 3, 3)
b = np.random.randn(25,)
dout = np.random.randn(100, 25, 16, 16)
conv_param = {'stride': 2, 'pad': 1}
t0 = clock()
out_naive, cache_naive = conv_forward_naive(x, w, b, conv_param)
t1 = clock()
out_fast, cache_fast = conv_forward_fast(x, w, b, conv_param)
t2 = clock()
print('Testing conv_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('Difference: ', rel_error(out_naive, out_fast))
t0 = clock()
dx_naive, dw_naive, db_naive = conv_backward_naive(dout, cache_naive)
t1 = clock()
dx_fast, dw_fast, db_fast = conv_backward_fast(dout, cache_fast)
t2 = clock()
print('\nTesting conv_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('Fast: %fs' % (t2 - t1))
print('Speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
print('dw difference: ', rel_error(dw_naive, dw_fast))
print('db difference: ', rel_error(db_naive, db_fast))
from nnlib.fast_layers import max_pool_forward_fast, max_pool_backward_fast
np.random.seed(231)
x = np.random.randn(100, 3, 32, 32)
dout = np.random.randn(100, 3, 16, 16)
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
t0 = clock()
out_naive, cache_naive = max_pool_forward_naive(x, pool_param)
t1 = clock()
out_fast, cache_fast = max_pool_forward_fast(x, pool_param)
t2 = clock()
print('Testing pool_forward_fast:')
print('Naive: %fs' % (t1 - t0))
print('fast: %fs' % (t2 - t1))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('difference: ', rel_error(out_naive, out_fast))
t0 = clock()
dx_naive = max_pool_backward_naive(dout, cache_naive)
t1 = clock()
dx_fast = max_pool_backward_fast(dout, cache_fast)
t2 = clock()
print('\nTesting pool_backward_fast:')
print('Naive: %fs' % (t1 - t0))
print('speedup: %fx' % ((t1 - t0) / (t2 - t1)))
print('dx difference: ', rel_error(dx_naive, dx_fast))
```
# Convolutional "sandwich" layers
Previously we introduced the concept of "sandwich" layers that combine multiple operations into commonly used patterns. In the file `nnlib/layer_utils.py` you will find sandwich layers that implement a few commonly used patterns for convolutional networks.
```
from nnlib.layer_utils import conv_relu_pool_forward, conv_relu_pool_backward
np.random.seed(231)
x = np.random.randn(2, 3, 16, 16)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
pool_param = {'pool_height': 2, 'pool_width': 2, 'stride': 2}
out, cache = conv_relu_pool_forward(x, w, b, conv_param, pool_param)
dx, dw, db = conv_relu_pool_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_pool_forward(x, w, b, conv_param, pool_param)[0], b, dout)
print('Testing conv_relu_pool')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
from nnlib.layer_utils import conv_relu_forward, conv_relu_backward
np.random.seed(231)
x = np.random.randn(2, 3, 8, 8)
w = np.random.randn(3, 3, 3, 3)
b = np.random.randn(3,)
dout = np.random.randn(2, 3, 8, 8)
conv_param = {'stride': 1, 'pad': 1}
out, cache = conv_relu_forward(x, w, b, conv_param)
dx, dw, db = conv_relu_backward(dout, cache)
dx_num = eval_numerical_gradient_array(lambda x: conv_relu_forward(x, w, b, conv_param)[0], x, dout)
dw_num = eval_numerical_gradient_array(lambda w: conv_relu_forward(x, w, b, conv_param)[0], w, dout)
db_num = eval_numerical_gradient_array(lambda b: conv_relu_forward(x, w, b, conv_param)[0], b, dout)
print('Testing conv_relu:')
print('dx error: ', rel_error(dx_num, dx))
print('dw error: ', rel_error(dw_num, dw))
print('db error: ', rel_error(db_num, db))
```
# Three-layer ConvNet
Now that you have implemented all the necessary layers, we can put them together into a simple convolutional network.
Open the file `nnlib/classifiers/cnn.py` and complete the implementation of the `ThreeLayerConvNet` class. Run the following cells to help you debug:
## Sanity check loss
After you build a new network, one of the first things you should do is sanity check the loss. When we use the softmax loss, we expect the loss for random weights (and no regularization) to be about `log(C)` for `C` classes. When we add regularization this should go up.
```
model = ThreeLayerConvNet()
N = 50
X = np.random.randn(N, 3, 32, 32)
y = np.random.randint(10, size=N)
loss, grads = model.loss(X, y)
print('Initial loss (no regularization): ', loss)
model.reg = 0.5
loss, grads = model.loss(X, y)
print('Initial loss (with regularization): ', loss)
```
## Gradient check
After the loss looks reasonable, use numeric gradient checking to make sure that your backward pass is correct. When you use numeric gradient checking you should use a small amount of artifical data and a small number of neurons at each layer. Note: correct implementations may still have relative errors up to 1e-2.
```
num_inputs = 2
input_dim = (3, 16, 16)
reg = 0.0
num_classes = 10
np.random.seed(231)
X = np.random.randn(num_inputs, *input_dim)
y = np.random.randint(num_classes, size=num_inputs)
model = ThreeLayerConvNet(num_filters=3, filter_size=3,
input_dim=input_dim, hidden_dim=7,
dtype=np.float64)
loss, grads = model.loss(X, y)
for param_name in sorted(grads):
f = lambda _: model.loss(X, y)[0]
param_grad_num = eval_numerical_gradient(f, model.params[param_name], verbose=False, h=1e-6)
e = rel_error(param_grad_num, grads[param_name])
print('%s max relative error: %e' % (param_name, rel_error(param_grad_num, grads[param_name])))
```
## Overfit small data
A nice trick is to train your model with just a few training samples. You should be able to overfit small datasets, which will result in very high training accuracy and comparatively low validation accuracy.
```
np.random.seed(231)
num_train = 100
small_data = {
'X_train': data['X_train'][:num_train],
'y_train': data['y_train'][:num_train],
'X_val': data['X_val'],
'y_val': data['y_val'],
}
model = ThreeLayerConvNet(weight_scale=1e-2)
solver = Solver(model, small_data,
num_epochs=15, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=1)
solver.train()
```
Plotting the loss, training accuracy, and validation accuracy should show clear overfitting:
```
plt.subplot(2, 1, 1)
plt.plot(solver.loss_history, 'o')
plt.xlabel('iteration')
plt.ylabel('loss')
plt.subplot(2, 1, 2)
plt.plot(solver.train_acc_history, '-o')
plt.plot(solver.val_acc_history, '-o')
plt.legend(['train', 'val'], loc='upper left')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.show()
```
## Train the net
By training the three-layer convolutional network for one epoch, you should achieve greater than 40% accuracy on the training set:
```
model = ThreeLayerConvNet(weight_scale=0.001, hidden_dim=500, reg=0.001)
solver = Solver(model, data,
num_epochs=1, batch_size=50,
update_rule='adam',
optim_config={
'learning_rate': 1e-3,
},
verbose=True, print_every=20)
solver.train()
```
## Visualize Filters
You can visualize the first-layer convolutional filters from the trained network by running the following:
```
from nnlib.vis_utils import visualize_grid
grid = visualize_grid(model.params['W1'].transpose(0, 2, 3, 1))
plt.imshow(grid.astype('uint8'))
plt.axis('off')
plt.gcf().set_size_inches(5, 5)
plt.show()
```
# Spatial Batch Normalization
We already saw that batch normalization is a very useful technique for training deep fully-connected networks. Batch normalization can also be used for convolutional networks, but we need to tweak it a bit; the modification will be called "spatial batch normalization."
Normally batch-normalization accepts inputs of shape `(N, D)` and produces outputs of shape `(N, D)`, where we normalize across the minibatch dimension `N`. For data coming from convolutional layers, batch normalization needs to accept inputs of shape `(N, C, H, W)` and produce outputs of shape `(N, C, H, W)` where the `N` dimension gives the minibatch size and the `(H, W)` dimensions give the spatial size of the feature map.
If the feature map was produced using convolutions, then we expect the statistics of each feature channel to be relatively consistent both between different imagesand different locations within the same image. Therefore spatial batch normalization computes a mean and variance for each of the `C` feature channels by computing statistics over both the minibatch dimension `N` and the spatial dimensions `H` and `W`.
## Spatial batch normalization: forward
In the file `nnlib/layers.py`, implement the forward pass for spatial batch normalization in the function `spatial_batchnorm_forward`. Check your implementation by running the following:
```
np.random.seed(231)
# Check the training-time forward pass by checking means and variances
# of features both before and after spatial batch normalization
N, C, H, W = 2, 3, 4, 5
x = 4 * np.random.randn(N, C, H, W) + 10
print('Before spatial batch normalization:')
print(' Shape: ', x.shape)
print(' Means: ', x.mean(axis=(0, 2, 3)))
print(' Stds: ', x.std(axis=(0, 2, 3)))
# Means should be close to zero and stds close to one
gamma, beta = np.ones(C), np.zeros(C)
bn_param = {'mode': 'train'}
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization:')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
# Means should be close to beta and stds close to gamma
gamma, beta = np.asarray([3, 4, 5]), np.asarray([6, 7, 8])
out, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
print('After spatial batch normalization (nontrivial gamma, beta):')
print(' Shape: ', out.shape)
print(' Means: ', out.mean(axis=(0, 2, 3)))
print(' Stds: ', out.std(axis=(0, 2, 3)))
np.random.seed(231)
# Check the test-time forward pass by running the training-time
# forward pass many times to warm up the running averages, and then
# checking the means and variances of activations after a test-time
# forward pass.
N, C, H, W = 10, 4, 11, 12
bn_param = {'mode': 'train'}
gamma = np.ones(C)
beta = np.zeros(C)
for t in range(50):
x = 2.3 * np.random.randn(N, C, H, W) + 13
spatial_batchnorm_forward(x, gamma, beta, bn_param)
bn_param['mode'] = 'test'
x = 2.3 * np.random.randn(N, C, H, W) + 13
a_norm, _ = spatial_batchnorm_forward(x, gamma, beta, bn_param)
# Means should be close to zero and stds close to one, but will be
# noisier than training-time forward passes.
print('After spatial batch normalization (test-time):')
print(' means: ', a_norm.mean(axis=(0, 2, 3)))
print(' stds: ', a_norm.std(axis=(0, 2, 3)))
```
## Spatial batch normalization: backward
In the file `nnlib/layers.py`, implement the backward pass for spatial batch normalization in the function `spatial_batchnorm_backward`. Run the following to check your implementation using a numeric gradient check:
```
np.random.seed(231)
N, C, H, W = 2, 3, 4, 5
x = 5 * np.random.randn(N, C, H, W) + 12
gamma = np.random.randn(C)
beta = np.random.randn(C)
dout = np.random.randn(N, C, H, W)
bn_param = {'mode': 'train'}
fx = lambda x: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fg = lambda a: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
fb = lambda b: spatial_batchnorm_forward(x, gamma, beta, bn_param)[0]
dx_num = eval_numerical_gradient_array(fx, x, dout)
da_num = eval_numerical_gradient_array(fg, gamma, dout)
db_num = eval_numerical_gradient_array(fb, beta, dout)
_, cache = spatial_batchnorm_forward(x, gamma, beta, bn_param)
dx, dgamma, dbeta = spatial_batchnorm_backward(dout, cache)
print('dx error: ', rel_error(dx_num, dx))
print('dgamma error: ', rel_error(da_num, dgamma))
print('dbeta error: ', rel_error(db_num, dbeta))
```
# Extra Credit Description
If you implement any additional features for extra credit, clearly describe them here with pointers to any code in this or other files if applicable.
| github_jupyter |
CER020 - Create Management Proxy certificate
============================================
This notebook creates a certificate for the Management Proxy endpoint.
Steps
-----
### Parameters
```
import getpass
app_name = "mgmtproxy"
scaledset_name = "mgmtproxy"
container_name = "service-proxy"
prefix_keyfile_name = "service-proxy"
common_name = "mgmtproxy-svc"
country_name = "US"
state_or_province_name = "Illinois"
locality_name = "Chicago"
organization_name = "Contoso"
organizational_unit_name = "Finance"
email_address = f"{getpass.getuser().lower()}@contoso.com"
ssl_configuration_file = "service.openssl.cnf"
days = "398" # the number of days to certify the certificate for
test_cert_store_root = "/var/opt/secrets/test-certificates"
extendedKeyUsage = ""
```
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
first_run = True
rules = None
debug_logging = False
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# apply expert rules (to run follow-on notebooks), based on output
#
if rules is not None:
apply_expert_rules(line_decoded)
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
"""Load a json file from disk and return the contents"""
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
"""Load any 'expert rules' from the metadata of this notebook (.ipynb) that should be applied to the stderr of the running executable"""
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
try:
j = load_json("cer020-create-management-service-proxy-cert.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"expanded_rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["expanded_rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
"""Determine if the stderr line passed in, matches the regular expressions for any of the 'expert rules', if so
inject a 'HINT' to the follow-on SOP/TSG to run"""
global rules
for rule in rules:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
if debug_logging:
print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
if debug_logging:
print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond'], 'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']], 'azdata': [['azdata login', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Error processing command: "ApiError', 'TSG110 - Azdata returns ApiError', '../repair/tsg110-azdata-returns-apierror.ipynb'], ['Error processing command: "ControllerError', 'TSG036 - Controller logs', '../log-analyzers/tsg036-get-controller-logs.ipynb'], ['ERROR: 500', 'TSG046 - Knox gateway logs', '../log-analyzers/tsg046-get-knox-logs.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ["Can't open lib 'ODBC Driver 17 for SQL Server", 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb'], 'azdata': ['SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb']}
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Create a temporary directory to stage files
```
# Create a temporary directory to hold configuration files
import tempfile
temp_dir = tempfile.mkdtemp()
print(f"Temporary directory created: {temp_dir}")
```
### Helper function to save configuration files to disk
```
# Define helper function 'save_file' to save configuration files to the temporary directory created above
import os
import io
def save_file(filename, contents):
with io.open(os.path.join(temp_dir, filename), "w", encoding='utf8', newline='\n') as text_file:
text_file.write(contents)
print("File saved: " + os.path.join(temp_dir, filename))
print("Function `save_file` defined successfully.")
```
### Get endpoint hostname
```
import json
import urllib
endpoint = run(f'azdata bdc endpoint list --endpoint="{app_name}"', return_output=True)
endpoint = json.loads(endpoint)
endpoint = endpoint['endpoint']
print(f"endpoint: {endpoint}")
hostname = urllib.parse.urlparse(endpoint).hostname
print(f"hostname: {hostname}")
```
### Get name of the ‘Running’ `controller` `pod`
```
# Place the name of the 'Running' controller pod in variable `controller`
controller = run(f'kubectl get pod --selector=app=controller -n {namespace} -o jsonpath={{.items[0].metadata.name}} --field-selector=status.phase=Running', return_output=True)
print(f"Controller pod name: {controller}")
```
### Create the DNS alt\_names for data plane in secure clusters
Get the cluster configuration from the Big Data Cluster using
`azdata bdc config`, and pull the Active Directory DNS names out of it,
and place them into the certificate configuration file as DNS alt\_names
```
import json
alt_names = ""
bdc_config = run("azdata bdc config show", return_output=True)
bdc_config = json.loads(bdc_config)
dns_counter = 3 # DNS.1 and DNS.2 are already in the certificate template.
# Add entry for "DNS.3 = {common_name}.<your_ad_domain_name>.local"
#
if "security" in bdc_config["spec"] and "activeDirectory" in bdc_config["spec"]["security"]:
domain_dns_name = bdc_config["spec"]["security"]["activeDirectory"]["domainDnsName"]
alt_names += f"DNS.{str(dns_counter)} = {common_name}.{domain_dns_name}\n"
dns_counter = dns_counter + 1
# Add entry for "DNS.4 = <your_endpoint_name>.<your_ad_domain_name>.local"
#
if app_name in bdc_config["spec"]["resources"]:
app_name_endpoints = bdc_config["spec"]["resources"][app_name]["spec"]["endpoints"]
for endpoint in app_name_endpoints:
if "dnsName" in endpoint:
alt_names += f'DNS.{str(dns_counter)} = {endpoint["dnsName"]}\n'
dns_counter = dns_counter + 1
# Special case for the controller certificate
#
if app_name == "controller":
alt_names += f"DNS.{str(dns_counter)} = localhost\n"
dns_counter = dns_counter + 1
print("DNS alt_names (data plane):")
print(alt_names)
```
### Create the DNS alt\_names for control plane in secure clusters
Get the cluster configuration from the Big Data Cluster using
`azdata bdc endpoint list`, and pull the Active Directory DNS names out
of it for the control plane expternal endpoints (Controller and
Management Proxy), and place them into the certificate configuration
file as DNS alt\_names
```
import json
from urllib.parse import urlparse
if app_name == "controller" or app_name == "mgmtproxy":
bdc_endpoint_list = run("azdata bdc endpoint list", return_output=True)
bdc_endpoint_list = json.loads(bdc_endpoint_list)
# Parse the DNS host name from:
#
# "endpoint": "https://monitor.aris.local:30777"
#
for endpoint in bdc_endpoint_list:
if endpoint["name"] == app_name:
url = urlparse(endpoint["endpoint"])
alt_names += f"DNS.{str(dns_counter)} = {url.hostname}\n"
dns_counter = dns_counter + 1
print("DNS alt_names (control plane):")
print(alt_names)
```
### Create alt\_names
If the Kuberenetes service is of “NodePort” type, then the IP address
needed to validate the cluster certificate could be for any node in the
Kubernetes cluster, so here all node IP addresses in the Big Data
Cluster are added as alt\_names. Otherwise (if not NodePort, and
therefore LoadBalancer), add just the hostname as returned from
`azdata bdc endpoint list` above.
```
service_type = run(f"kubectl get svc {common_name}-external -n {namespace} -o jsonpath={{.spec.type}}", return_output=True)
print(f"Service type for '{common_name}-external' is: '{service_type}'")
print("")
if service_type == "NodePort":
nodes_ip_address = run("kubectl ""get nodes -o jsonpath={.items[*].status.addresses[0].address}""", return_output=True)
nodes_ip_address = nodes_ip_address.split(' ')
counter = 1
for ip in nodes_ip_address:
alt_names += f"IP.{counter} = {ip}\n"
counter = counter + 1
else:
alt_names += f"IP.1 = {hostname}\n"
print("All (DNS and IP) alt_names:")
print(alt_names)
```
### Generate Certificate Configuration file
NOTE: There is a special case for the `controller` certificate, that
needs to be generated in PKCS\#1 format.
```
certificate = f"""
[ req ]
# Options for the `req` tool (`man req`).
default_bits = 2048
default_keyfile = {test_cert_store_root}/{app_name}/{prefix_keyfile_name}-privatekey{".pkcs8" if app_name == "controller" else ""}.pem
distinguished_name = req_distinguished_name
string_mask = utf8only
# SHA-1 is deprecated, so use SHA-2 instead.
default_md = sha256
req_extensions = v3_req
[ req_distinguished_name ]
countryName = Country Name (2 letter code)
countryName_default = {country_name}
stateOrProvinceName = State or Province Name (full name)
stateOrProvinceName_default = {state_or_province_name}
localityName = Locality Name (eg, city)
localityName_default = {locality_name}
organizationName = Organization Name (eg, company)
organizationName_default = {organization_name}
organizationalUnitName = Organizational Unit (eg, division)
organizationalUnitName_default = {organizational_unit_name}
commonName = Common Name (e.g. server FQDN or YOUR name)
commonName_default = {common_name}
emailAddress = Email Address
emailAddress_default = {email_address}
[ v3_req ]
subjectAltName = @alt_names
subjectKeyIdentifier = hash
basicConstraints = CA:FALSE
keyUsage = digitalSignature, keyEncipherment
{extendedKeyUsage}
[ alt_names ]
DNS.1 = {common_name}
DNS.2 = {common_name}.{namespace}.svc.cluster.local # Use the namespace applicable for your cluster
{alt_names}
"""
save_file(ssl_configuration_file, certificate)
```
### Copy certificate configuration to `controller` `pod`
```
import os
cwd = os.getcwd()
os.chdir(temp_dir) # Use chdir to workaround kubectl bug on Windows, which incorrectly processes 'c:\' on kubectl cp cmd line
run(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c "mkdir -p {test_cert_store_root}/{app_name}"')
run(f'kubectl cp {ssl_configuration_file} {controller}:{test_cert_store_root}/{app_name}/{ssl_configuration_file} -c controller -n {namespace}')
os.chdir(cwd)
```
### Generate certificate
Use openssl req to generate a certificate in PKCS\#10 format. See:
- https://www.openssl.org/docs/man1.0.2/man1/req.html
```
cmd = f"openssl req -config {test_cert_store_root}/{app_name}/service.openssl.cnf -newkey rsa:2048 -sha256 -nodes -days {days} -out {test_cert_store_root}/{app_name}/{prefix_keyfile_name}-signingrequest.csr -outform PEM -subj '/C={country_name}/ST={state_or_province_name}/L={locality_name}/O={organization_name}/OU={organizational_unit_name}/CN={common_name}'"
run(f'kubectl exec {controller} -n {namespace} -c controller -- bash -c "{cmd}"')
```
### Clean up temporary directory for staging configuration files
```
# Delete the temporary directory used to hold configuration files
import shutil
shutil.rmtree(temp_dir)
print(f'Temporary directory deleted: {temp_dir}')
print('Notebook execution complete.')
```
Related
-------
- [CER021 - Create Knox
certificate](../cert-management/cer021-create-knox-cert.ipynb)
- [CER030 - Sign Management Proxy certificate with generated
CA](../cert-management/cer030-sign-service-proxy-generated-cert.ipynb)
- [CER040 - Install signed Management Proxy
certificate](../cert-management/cer040-install-service-proxy-cert.ipynb)
| github_jupyter |
# Licence
Copyright (c) 2019 Marcus D. Bloice <marcus.bloice@medunigraz.at>
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# Patch Augmentation
Patch augmentation is a novel technique used to attempt to create more efficient decision boundaries in neural networks in order to mitigate against adversarial attacks.
**Patch Augmentation is a data-independent approach that creates new image data based on image/label pairs, where a patch from one of the two images in the pair is superimposed on to the other image, creating a new augmented sample.**
This notebook contains an implementaton of the technique, and demonstrates its application to the CIFAR-10 and CIFAR-100 datasets, comparing it to baseline accuracies.
Portions of this code (code relating to the creation of the ResNetv1 and ResNetv2 networks) are from the Keras documentation.
Further experiments will follow and will be added to this notebook.
## Imports
A few imports are required at first:
```
import keras
from keras.layers import Dense, Conv2D, BatchNormalization, Activation
from keras.layers import AveragePooling2D, Input, Flatten
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, LearningRateScheduler
from keras.callbacks import ReduceLROnPlateau
from keras.preprocessing.image import ImageDataGenerator
from keras.regularizers import l2
from keras import backend as K
from keras.models import Model
from keras.datasets import cifar10
from keras.utils import Sequence
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
import math
import random
import os
%matplotlib inline
np.random.seed(0)
```
## Define the PatchAugmentation Class
The `PatchAugmentation` class is a sub-class of Keras' `Sequence` class. According to the Keras documentation:
*This structure guarantees that the network will only train once on each sample per epoch which is not the case with generators.*
See <https://keras.io/utils/#sequence>
This means that Keras will ensure we are passing all image data through the network per epoch, and that this does not need to be implemented by us.
Once we have defined the class we can perform some visual verificiations that the class works as intended.
```
class PatchAugmentation(Sequence):
def __init__(self, batch_size, x, y, probability=1.0, image_area=1024, patch_area=0.25):
self.batch_size = batch_size
self.x_train = x
self.y_train = y
self.probability = probability
self.image_area = image_area
self.patch_area = patch_area
# Calculate various dimensions for the patch placement.
self.dim = int(round(math.sqrt(self.image_area)))
self.crop_area_in_pixels = self.image_area * self.patch_area
self.crop_dim = int(round(math.sqrt(self.crop_area_in_pixels)))
self.max_horizontal_shift = (math.sqrt(self.image_area)) - self.crop_dim
self.max_vertical_shift = (math.sqrt(self.image_area)) - self.crop_dim
def __len__(self):
return int(np.ceil(len(self.x_train) / float(self.batch_size)))
def __getitem__(self, idx):
batch_x = np.copy(self.x_train[idx * self.batch_size:(idx+1) * self.batch_size])
batch_y = np.copy(self.y_train[idx * self.batch_size:(idx+1) * self.batch_size])
for i in range(len(batch_x)):
if np.random.uniform(0, 1) <= self.probability:
r_i = np.random.randint(0, len(self.x_train))
x1 = np.random.randint(0, self.dim - self.crop_dim)
x2 = x1 + self.crop_dim
y1 = np.random.randint(0, self.dim - self.crop_dim)
y2 = y1 + self.crop_dim
batch_x[i][x1:x2, y1:y2, :] = self.x_train[r_i][x1:x2, y1:y2, :]
lambda_value = self.patch_area
batch_y[i] = (1- lambda_value) * batch_y[i] + lambda_value * self.y_train[r_i]
return batch_x, batch_y
```
## Visual Verification of Patch Augmentation Class
### Create A Dataset
We can create a quick dataset to visually confirm the `PatchAugmentation` class functions as intended.
Sample cat, dog, and bird images are CC0 (Creative Commons Zero) licensed, sourced from: <https://www.pexels.com/>
First, we load some sample data:
```
cat = Image.open('DemoImages/cat.jpg').resize((400,400), Image.LANCZOS)
dog = Image.open('DemoImages/dog.jpg').resize((400,400), Image.LANCZOS)
bird = Image.open('DemoImages/bird.jpg').resize((400,400), Image.LANCZOS)
cat = np.asarray(cat, dtype=np.uint16)
dog = np.asarray(dog, dtype=np.uint16)
bird = np.asarray(bird, dtype=np.uint16)
plt.imshow(cat);
```
We now create a a dataset with 1,000 samples based on the three images loaded above:
```
num_of_examples = 1000
x_train_exp = np.empty((num_of_examples, 400, 400, 3), dtype=np.uint16)
y_train_exp = []
for i in range(0, num_of_examples):
r = random.choice([0,1,2])
if r == 0:
x_train_exp[i,:,:,:] = cat
y_train_exp.append([1.0, 0.0, 0.0])
elif r == 1:
x_train_exp[i,:,:,:] = dog
y_train_exp.append([0.0, 1.0, 0.0])
else:
x_train_exp[i,:,:,:] = bird
y_train_exp.append([0.0, 0.0, 1.0])
y_train_exp = np.array(y_train_exp)
# Confirm some numbers
print("Image array shape: %s, length of label vector: %s" % (x_train_exp.shape, len(y_train_exp)))
cat_count, dog_count, bird_count = 0, 0, 0
for i in range(0, len(y_train_exp)):
if np.argmax(y_train_exp[i]) == 0:
cat_count += 1
elif np.argmax(y_train_exp[i]) == 1:
dog_count += 1
else:
bird_count += 1
print("Number of cats: %s, number of dogs: %s, number of birds: %s." % (cat_count, dog_count, bird_count))
assert (cat_count + dog_count + bird_count) == num_of_examples
```
We have created a dataset containing 1,000 images and 1,000 labels, with three classes.
We can now choose a random example from the dataset and confirm it is correctly labelled.
Note that the labels for the three classes are as follows:
```
Cat = [1. 0. 0.]
Dog = [0. 1. 0.]
Bird = [0. 0. 1.]
```
```
idx = np.random.randint(0, len(x_train_exp))
print("Random index: %s" % idx)
plt.imshow(x_train_exp[idx])
print("Label vector: %s" % y_train_exp[idx])
```
The label vector should match with the class labels printed above for the class cat, dog, or bird.
### Create Augmented Dataset
Now we use the `PatchAugmentation` class to generate an augmented data set using the Patch Augmentation technique.
As mentioned previously, the `PatchAugmentation` class is a subclass of Keras' `Sequence`, hence we will create a generator to view some of the data created by it:
```
b_size = 100
m = PatchAugmentation(b_size, x=x_train_exp, y=y_train_exp, probability=1.0, image_area=400*400, patch_area=0.25)
```
Get the generator to produce a batch of augmented samples (this will be equal to `b_size` defined above):
```
r_g = np.random.randint(0, len(m))
x, y = m.__getitem__(r_g)
print("Random batch index %s" % (r_g))
```
Let's look at some random images returned by the patch generator, which should contain augmented images and labels. Remeber the original one-hot-encoded labels are as follows:
```
Cat = [1. 0. 0.]
Dog = [0. 1. 0.]
Bird = [0. 0. 1.]
```
```
idex = np.random.randint(0, len(x))
print("Index: %s, Batch: %s, Label vector: %s" % (idex, r_g, y[idex]))
plt.imshow(x[idex]);
idex = np.random.randint(0, len(x))
print("Index: %s, Batch: %s, Label vector: %s" % (idex, r_g, y[idex]))
plt.imshow(x[idex]);
idex = np.random.randint(0, len(x))
print("Index: %s, Batch: %s, Label vector: %s" % (idex, r_g, y[idex]))
plt.imshow(x[idex]);
```
Now that we have visually confirmed the `PatchAugmentation` class we can define the networks we will use for the experiments.
## Setting Up ResNetv1 and ResNetv2
Here we define some functions for creating ResNet v1 and v2, etc.
Learning rate scheduling is used by most implementations of CIFAR, so we will also use it:
```
def lr_schedule(epoch):
"""Learning Rate Schedule
Learning rate is scheduled to be reduced after 100, 140, 180, and 190 epochs.
Called automatically every epoch as part of callbacks during training.
# Arguments
epoch (int): The number of epochs
# Returns
lr (float32): learning rate
"""
lr = 1e-3
if epoch > 190:
lr *= 0.5e-3
elif epoch > 180:
lr *= 1e-3
elif epoch > 140:
lr *= 1e-2
elif epoch > 100:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
```
Define a RESNET layer:
```
def resnet_layer(inputs,
num_filters=16,
kernel_size=3,
strides=1,
activation='relu',
batch_normalization=True,
conv_first=True):
"""2D Convolution-Batch Normalization-Activation stack builder
# Arguments
inputs (tensor): input tensor from input image or previous layer
num_filters (int): Conv2D number of filters
kernel_size (int): Conv2D square kernel dimensions
strides (int): Conv2D square stride dimensions
activation (string): activation name
batch_normalization (bool): whether to include batch normalization
conv_first (bool): conv-bn-activation (True) or
bn-activation-conv (False)
# Returns
x (tensor): tensor as input to the next layer
"""
conv = Conv2D(num_filters,
kernel_size=kernel_size,
strides=strides,
padding='same',
kernel_initializer='he_normal',
kernel_regularizer=l2(1e-4))
x = inputs
if conv_first:
x = conv(x)
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
else:
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
x = conv(x)
return x
```
Now we define our ResNet networks, using the `resnet_layer()` function defined above. We start with ResNetv1:
```
def resnet_v1(input_shape, depth, num_classes=10):
"""ResNet Version 1 Model builder [a]
Stacks of 2 x (3 x 3) Conv2D-BN-ReLU
Last ReLU is after the shortcut connection.
At the beginning of each stage, the feature map size is halved (downsampled)
by a convolutional layer with strides=2, while the number of filters is
doubled. Within each stage, the layers have the same number filters and the
same number of filters.
Features maps sizes:
stage 0: 32x32, 16
stage 1: 16x16, 32
stage 2: 8x8, 64
The Number of parameters is approx the same as Table 6 of [a]:
ResNet20 0.27M
ResNet32 0.46M
ResNet44 0.66M
ResNet56 0.85M
ResNet110 1.7M
# Arguments
input_shape (tensor): shape of input image tensor
depth (int): number of core convolutional layers
num_classes (int): number of classes (CIFAR10 has 10)
# Returns
model (Model): Keras model instance
"""
if (depth - 2) % 6 != 0:
raise ValueError('depth should be 6n+2 (eg 20, 32, 44 in [a])')
# Start model definition.
num_filters = 16
num_res_blocks = int((depth - 2) / 6)
inputs = Input(shape=input_shape)
x = resnet_layer(inputs=inputs)
# Instantiate the stack of residual units
for stack in range(3):
for res_block in range(num_res_blocks):
strides = 1
if stack > 0 and res_block == 0: # first layer but not first stack
strides = 2 # downsample
y = resnet_layer(inputs=x,
num_filters=num_filters,
strides=strides)
y = resnet_layer(inputs=y,
num_filters=num_filters,
activation=None)
if stack > 0 and res_block == 0: # first layer but not first stack
# linear projection residual shortcut connection to match
# changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters,
kernel_size=1,
strides=strides,
activation=None,
batch_normalization=False)
x = keras.layers.add([x, y])
x = Activation('relu')(x)
num_filters *= 2
# Add classifier on top.
# v1 does not use BN after last shortcut connection-ReLU
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
# Instantiate model.
model = Model(inputs=inputs, outputs=outputs)
return model
```
Followed by ResNetv2, also using the `resnet_layer()` function defined previously:
```
def resnet_v2(input_shape, depth, num_classes=10):
"""ResNet Version 2 Model builder [b]
Stacks of (1 x 1)-(3 x 3)-(1 x 1) BN-ReLU-Conv2D or also known as
bottleneck layer
First shortcut connection per layer is 1 x 1 Conv2D.
Second and onwards shortcut connection is identity.
At the beginning of each stage, the feature map size is halved (downsampled)
by a convolutional layer with strides=2, while the number of filter maps is
doubled. Within each stage, the layers have the same number filters and the
same filter map sizes.
Features maps sizes:
conv1 : 32x32, 16
stage 0: 32x32, 64
stage 1: 16x16, 128
stage 2: 8x8, 256
# Arguments
input_shape (tensor): shape of input image tensor
depth (int): number of core convolutional layers
num_classes (int): number of classes (CIFAR10 has 10)
# Returns
model (Model): Keras model instance
"""
if (depth - 2) % 9 != 0:
raise ValueError('depth should be 9n+2 (eg 56 or 110 in [b])')
# Start model definition.
num_filters_in = 16
num_res_blocks = int((depth - 2) / 9)
inputs = Input(shape=input_shape)
# v2 performs Conv2D with BN-ReLU on input before splitting into 2 paths
x = resnet_layer(inputs=inputs,
num_filters=num_filters_in,
conv_first=True)
# Instantiate the stack of residual units
for stage in range(3):
for res_block in range(num_res_blocks):
activation = 'relu'
batch_normalization = True
strides = 1
if stage == 0:
num_filters_out = num_filters_in * 4
if res_block == 0: # first layer and first stage
activation = None
batch_normalization = False
else:
num_filters_out = num_filters_in * 2
if res_block == 0: # first layer but not first stage
strides = 2 # downsample
# bottleneck residual unit
y = resnet_layer(inputs=x,
num_filters=num_filters_in,
kernel_size=1,
strides=strides,
activation=activation,
batch_normalization=batch_normalization,
conv_first=False)
y = resnet_layer(inputs=y,
num_filters=num_filters_in,
conv_first=False)
y = resnet_layer(inputs=y,
num_filters=num_filters_out,
kernel_size=1,
conv_first=False)
if res_block == 0:
# linear projection residual shortcut connection to match
# changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters_out,
kernel_size=1,
strides=strides,
activation=None,
batch_normalization=False)
x = keras.layers.add([x, y])
num_filters_in = num_filters_out
# Add classifier on top.
# v2 has BN-ReLU before Pooling
x = BatchNormalization()(x)
x = Activation('relu')(x)
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
# Instantiate model.
model = Model(inputs=inputs, outputs=outputs)
return model
```
Define a number of hyper-parameters:
```
# Subtracting the pixel mean helps with the final accuracy. For all experiments we leave this as True.
subtract_pixel_mean = True
# Depth of the ResNet network can be defined here.
n = 3
# Model version
# Orig paper: version = 1 (ResNet v1), Improved ResNet: version = 2 (ResNet v2)
version = 1
```
Here we prepare the CIFAR10 data set using some of the parameters above defined above:
```
cifar_10_labels = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
num_classes = len(cifar_10_labels)
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
input_shape = x_train.shape[1:]
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
if subtract_pixel_mean:
x_train_mean = np.mean(x_train, axis=0)
x_train -= x_train_mean
x_test -= x_train_mean
```
Now we are ready to create the network, using the parameters set above (such as ResNet version and depth):
```
# Computed depth from supplied model parameter n
if version == 1:
depth = n * 6 + 2
elif version == 2:
depth = n * 9 + 2
# Model name, depth and version
model_type = 'ResNet%dv%d' % (depth, version)
if version == 2:
model = resnet_v2(input_shape=input_shape, depth=depth)
else:
model = resnet_v1(input_shape=input_shape, depth=depth)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=lr_schedule(0)),
metrics=['accuracy'])
save_dir = os.path.join('/tmp/', 'saved_models')
model_name = 'cifar10_%s_model.{epoch:03d}.h5' % model_type
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
filepath = os.path.join(save_dir, model_name)
# Prepare callbacks for model saving and for learning rate adjustment.
checkpoint = ModelCheckpoint(filepath=filepath,
monitor='val_acc',
verbose=1,
save_best_only=True)
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
callbacks = [lr_reducer, lr_scheduler] # Add checkpoint to this list to save model file
print(model_type)
```
The type of network is printed above for confirmation.
## Train The Network
Now we have prepared a network, we can train it using a `PatchAugmentation` generator:
```
patch_swap_generator = PatchAugmentation(batch_size=128,
x=x_train,
y = y_train,
probability=0.5,
patch_area=0.25)
```
Then start training the network with the generator:
```
history = model.fit_generator(patch_swap_generator,
epochs=200,
validation_data=(x_test, y_test),
shuffle=True,
callbacks=callbacks,
verbose=1)
```
Plotting the accuracy and loss:
```
# Plot Accuracy
acc_peak = np.max(history.history['val_acc'])
plt.plot(history.history['acc'], color='tab:red')
plt.plot(history.history['val_acc'], color='tab:blue')
plt.title('Patch Augmentation Accuracy (Peak %s) - %s' % (acc_peak, model_type))
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot Loss
plt.plot(history.history['loss'], color='tab:red')
plt.plot(history.history['val_loss'], color='tab:blue')
plt.title('Patch Augmentation Loss - %s' % (model_type))
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
```
## Results
From a baseline accuracy of approximately 80.5%, Patch Augmentation improves accuracy to 86.8%.
| github_jupyter |
# Backwards Compatibility Examples with Different Protocols
## Prerequisites
* A kubernetes cluster with kubectl configured
* curl
* grpcurl
* pygmentize
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html) to setup Seldon Core with an ingress - either Ambassador or Istio.
Then port-forward to that ingress on localhost:8003 in a separate terminal either with:
* Ambassador: `kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080`
* Istio: `kubectl port-forward $(kubectl get pods -l istio=ingressgateway -n istio-system -o jsonpath='{.items[0].metadata.name}') -n istio-system 8003:8080`
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
import json
import time
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, "w") as f:
f.write(cell.format(**globals()))
VERSION=!cat ../version.txt
VERSION=VERSION[0]
VERSION
```
## Model with Old REST Wrapper Upgraded
We will deploy a REST model that uses the SELDON Protocol namely by specifying the attribute `protocol: seldon`
```
%%writetemplate resources/model_seldon.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: example-seldon
spec:
protocol: seldon
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier_rest:1.4.0
name: classifier
graph:
name: classifier
type: MODEL
name: model
replicas: 1
!kubectl apply -f resources/model_seldon.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-seldon -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep example-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/example-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
%%writetemplate resources/model_seldon.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: example-seldon
spec:
protocol: seldon
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
name: classifier
graph:
name: classifier
type: MODEL
name: model
replicas: 1
!kubectl apply -f resources/model_seldon.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-seldon -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep example-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/example-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
X=!cd ../executor/proto && grpcurl -d '{"data":{"ndarray":[[1.0,2.0,5.0]]}}' \
-rpc-header seldon:example-seldon -rpc-header namespace:seldon \
-plaintext \
-proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict
d=json.loads("".join(X))
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
!kubectl delete -f resources/model_seldon.yaml
```
## Model with Old GRPC Wrapper Upgraded
We will deploy a gRPC model that uses the SELDON Protocol namely by specifying the attribute `protocol: seldon`
```
%%writefile resources/model_seldon_grpc.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: grpc-seldon
spec:
name: grpcseldon
protocol: seldon
transport: grpc
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier_grpc:1.3
name: classifier
graph:
name: classifier
type: MODEL
endpoint:
type: GRPC
name: model
replicas: 1
!kubectl apply -f resources/model_seldon_grpc.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=grpc-seldon -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep grpc-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!cd ../executor/proto && grpcurl -d '{"data":{"ndarray":[[1.0,2.0,5.0,6.0]]}}' \
-rpc-header seldon:grpc-seldon -rpc-header namespace:seldon \
-plaintext \
-proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict
d=json.loads("".join(X))
print(d)
!kubectl delete -f resources/model_seldon_grpc.yaml
```
## Old Operator and Model Upgraded
```
!helm delete seldon seldon-core-operator \
--namespace seldon-system
!helm install seldon seldon-core-operator \
--repo https://storage.googleapis.com/seldon-charts \
--version 1.4.0 \
--namespace seldon-system \
--wait
%%writetemplate resources/model_seldon.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: example-seldon
spec:
protocol: seldon
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier_rest:1.4.0
name: classifier
graph:
name: classifier
type: MODEL
name: model
replicas: 1
%%writefile ../servers/sklearnserver/samples/iris.yaml
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
name: sklearn
spec:
name: iris
predictors:
- graph:
children: []
implementation: SKLEARN_SERVER
modelUri: gs://seldon-models/sklearn/iris
name: classifier
name: default
replicas: 1
svcOrchSpec:
env:
- name: SELDON_LOG_LEVEL
value: DEBUG
!kubectl apply -f resources/model_seldon.yaml
!kubectl apply -f ../servers/sklearnserver/samples/iris.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-seldon -o jsonpath='{.items[0].metadata.name}')
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=sklearn -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep example-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/example-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0, 6.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/sklearn/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
!helm upgrade seldon \
../helm-charts/seldon-core-operator \
--namespace seldon-system \
--wait
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-seldon -o jsonpath='{.items[0].metadata.name}')
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=sklearn -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep example-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
```
Only REST calls will be available as image is still old python wrapper
```
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/example-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
```
Rest and gRPC calls will work with new server as image will have been updated.
```
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0, 6.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/sklearn/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
X=!cd ../executor/proto && grpcurl -d '{"data":{"ndarray":[[1.0,2.0,5.0,6.0]]}}' \
-rpc-header seldon:sklearn -rpc-header namespace:seldon \
-plaintext \
-proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict
d=json.loads("".join(X))
print(d)
%%writetemplate resources/model_seldon.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: example-seldon
spec:
protocol: seldon
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
name: classifier
graph:
name: classifier
type: MODEL
name: model
replicas: 1
!kubectl apply -f resources/model_seldon.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example-seldon -o jsonpath='{.items[0].metadata.name}')
for i in range(60):
state=!kubectl get sdep example-seldon -o jsonpath='{.status.state}'
state=state[0]
print(state)
if state=="Available":
break
time.sleep(1)
assert(state=="Available")
X=!curl -s -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
-X POST http://localhost:8003/seldon/seldon/example-seldon/api/v1.0/predictions \
-H "Content-Type: application/json"
d=json.loads(X[0])
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
X=!cd ../executor/proto && grpcurl -d '{"data":{"ndarray":[[1.0,2.0,5.0]]}}' \
-rpc-header seldon:example-seldon -rpc-header namespace:seldon \
-plaintext \
-proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict
d=json.loads("".join(X))
print(d)
assert(d["data"]["ndarray"][0][0] > 0.4)
!kubectl delete -f resources/model_seldon.yaml
!kubectl delete -f ../servers/sklearnserver/samples/iris.yaml
```
| github_jupyter |
# Who Can Tell What Harald Uhlig Said Six Years Ago in Front of 60-Plus Witnesses?
Yes, it is John Cochrane blogging once again. Once again, some of the plain people of the internet who wish me ill tell me I need to go and read John Cochrane—presumably to make me ill.
## Should Harald Uhlig Be Lead Editor of the _Journal of Political Economy_?
And I do. And it does: **John Cochrane** <https://tinyurl.com/2020-06-16-a> on why Harald Uhlig deserves to still be lead editor of the _Journal of Political Economy_:
>The _J[ournal of ]P[olitical ]E[conomy]_ advisory board (Robert Shimer, Lars Peter Hansen, Steve Levitt, and Philip J. Reny, all good friends and great economists, so my pain here is deep at having to criticize their action) carefully say nothing about the tweets, media coverage, and protest. They cite instead the accusation of discriminatory conduct in a classroom setting. Now, such conduct is a very serious charge...
Which Cochrane then dismisses:
>But this account of events does not hold water. Ba, now a professor at U.C. Irvine, was sitting in—not taking for credit—a class in 2014, six years ago. At the University of Chicago, there was always the issue for classes that meet on Mondays, how do you reschedule the class that would normally take place on Martin Luther King day? It was always a mess...
One might think that one sucks it up: plans for the fact that there is a holiday.
Or one might think that one would make the holiday class up during reading period, if one has failed to plan one's syllabus according to the university calender.
And one might think this is not a problem unique to Martin Luther King, Jr. Day: that it applies to Washington & Lincoln's Birthday, Memorial Day, and Thanksgiving as well.
But, apparently, that is not what University of Chicago economics professor Harald Uhlig did. **Bocar A. Ba** <https://tinyurl.com/2020-06-16-b>:
>@haralduhlig: I sat in your class in Winter 2014:
>1. You talked about scheduling a class on MLK Day
>2. You made fun of Dr. King and people honoring him
>3. You sarcastically asked me in front of everyone whether I was offended
**chitownprof** <https://tinyurl.com/2020-06-16-f>: "I was one of the 60+ fellow PhD students in that course. Everything you say is true. Uhlig had to reschedule a class and suggested the following Monday. A classmate said it was MLK day, to which he looked for you (the only Black student), and asked: 'YOU don't mind, do you?' And, to be clear, at the time MLK was a University of Chicago observed holiday in which classes were cancelled."
**Alejandro Hoyos** <https://tinyurl.com/2020-06-16-c>: "I was in the same class with Bocar and each and every word is true, this happened exactly as he described it."
**Jordan Rosenthal-Kay** <https://tinyurl.com/2020-06-16-d>: "For the record: similar comments were repeated in class this year"
**Corinne Low** <https://tinyurl.com/2020-06-16-e>: "Yes, I have heard HU also said the same thing this year, although perhaps without a black student to single out and humiliate. It also gives lie to the idea he's criticizing BLM bc of how much he loves MLK and peaceful protest."
**fluffynuffy** <https://tinyurl.com/2020-06-16-g>: "Uhlig makes weird MLKjr Day 'jokes' every year in his classes apparently. This is probably acceptable yet bad free expression. But calling on black students specifically to ask how they feel about this isn't free expression at all."
Ah. So it wasn't that Harald Uhlig had failed to plan for MLK Day. It was that he had made an outside commitment during what was supposed to be his lecture time, and wanted to reschedule the class he wanted to miss for MLK Day. The "mess" is one created by Uhlig's scheduling outside commitments during class time. Yet somehow Cochrane does not find space to say this.
Cochrane continues:
>...In that discussion, Harald said something that Ba found offensive—that much is undeniable. What "fun" did Harald make of Dr. King? Precise words would help. Clearly in this interaction the tone fo voice—whether Harald's inquiry as to offense was "sarcastic" or well-intended—mattered as much as what was actually said. Yes, this merits investigation, to the extent that one can investigate comments made in classes six years ago reported via tweet.
>But the JPE, on Friday, was clearly responding to Harald's tweets, not just this accusation, and the JPE's contrary assertion a poor excuse. There is no way on this green earth that a tweet made on Thursday about a comment made in class six years ago leads to being fired from the JPE on Friday, absent a mob demanding just that head for previous tweets about defunding the police. An allegation of misbehavior in class would justify suspending Harald from teaching classes, maybe.
>I spent much of my last few years of teaching afraid that I would say something that could be misunderstood and thus be offensive to someone. Many of my colleagues report the same worries. It is not good for open and honest communication in the classroom if a tweet about a comment six years ago can destroy you.
Yes, plain people of the internet. You have made me ill. Again. Satisfied?
## Fear of Reporters
So to recover my equilibrium, let's talk about John Cochrane's Fear of Reporters. Immediately after the last quote above, he continues:
>I also learned the hard way, don't talk to reporters who are out to destroy you, and that includes anyone from the _New York Times_ or the _New Yorker_. Compare the _Times_ article to the unedited transcript of Uhlig's interview <https://tinyurl.com/2020-06-16-i>. I think Harald just learned another sharp lesson.
Here are the paragraphs in the _New York Times_ article from Harald Uhlig's email interview: **Ben Casselman** & **Tim Tankersley** <https://tinyurl.com/2020-06-16-h>:
>[Uhlig] said in an email interview on Tuesday night that his “flat earther” comparison “appears to have caused irritation” but disagreed with critics who say his comments “hurt and marginalize people of color and their allies in the economics profession; call into question his impartiality in assessing academic work on this and related topics; and damage the standing of the economics discipline in society.” The reference to the Klan, he said, was a case where “I chose an extreme example” to make a point about free speech.
>“Discrimination and racism is wrong,” Mr. Uhlig wrote in an email. Later, he added: “I would love to have more black economists (or is it ‘Afro-American economists’?) among our undergraduate students, Ph.D. students and faculty. It is my impression that the good ones are highly sought after. We also have very few American Indians among our colleagues. We need to find good way to change these numbers.”
I genuinely did not see why Cochrane thinks Uhlig has a beef here.
## "We Should Have a Recession... Pounding Nails in Nevada..."
But then I remembered the moment when I first concluded that John Cochrane had gone bats--t—that he has simply completely lost it. It was on December 23, 2008, when I read a Bloomberg News article by **John Lippert** <https://tinyurl.com/2020-06-16-j>:
>John Cochrane was steaming as word of U.S. Treasury Secretary Henry Paulson’s plan to buy 700 billion in troubled mortgage assets rippled across the University of Chicago in September.... During a lunch held on a balcony with a view of Rockefeller Memorial Chapel, Cochrane, son-in-law of Chicago efficient-market theorist Eugene Fama, and some colleagues made their stand. They wrote a petition... collected 230 signatures. Republican Senator Richard Shelby of Alabama waved the document as he scorned the rescue. When Congress rejected it on Sept. 29, Cochrane fired off congratulatory e-mails. The victory was short-lived. Lawmakers approved the plan four days later, swayed by what Cochrane calls a pinata of pork-barrel amendments. “We should have a recession,” Cochrane said in November, speaking to students and investors in a conference room that looks out on Lake Michigan. “People who spend their lives pounding nails in Nevada need something else to do.”
Now we need to look at some data. So we need to start by initializing our environment:
```
%matplotlib inline
# ensure graphs show up inside rather than
# outside the notebook
# initialize libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
```
Now we can grab the time series for the share of nonfarm workers in America in residential building—the people "pounding nails". Let us look at this share each year from 2000 to 2012 in October, before cold weather in the north has started to shut down construction:
```
pounding_nails_df = pd.read_csv('https://delong.typepad.com/files/2020-06-16-pounding-nails-in-nevada-3.csv')
slice_df = pounding_nails_df[(pounding_nails_df['month'] == 10) &
(pounding_nails_df['year'] > 1999) &
(pounding_nails_df['year'] < 2013)
]
format_dict = { 'construction_share': '{0:,.2%}'}
slice_df[['date','construction_share']].style.format(format_dict)
```
In the small table above we can clearly see the housing boom of the mid-2000s and its unwinding: the share of American workers working in residential construction rises from 5.15% in the early 2000s up to 5.6% at the peak of the housing boom in 2006, and then declines. By October 2008 the share of American workers in construction was back to its pre-boom level, and around its long-term average.
Before Cochrane stood up and said "We should have a recession. People who spend their lives pounding nails in Nevada need something else to do", the excess employment in construction—those people who would need to find something else to do in the long run—had already exited the sector. So why, then, did we need to have a recession? We didn't. Cochrane's argument was incoherent. And illiterate. The task of structural adjustment to the end of the housing boom had already been accomplished. And did not require a recession.
So what was Cochrane's reaction when Paul Krugman called him out on his "we should have a recession...pounding nails in Nevada" claim?
It was not very pretty <https://tinyurl.com/2020-06-16-k>:
>The level of personal attack in the [Krugman] _New York Times_ article, and the fudging of the facts to achieve it, is simply amazing. As one little example, take my quotation about carpenters in Nevada.... I did not write this. It is an attribution, taken out of context, from a bloomberg.com article, written by a reporter [John Lippert] with whom I spent about 10 hours patiently trying to explain some basics, and who also turned out only to be on a hunt for embarrassing quotes. Nevertheless, I was trying to explain how sectoral shifts contribute to unemployment. I never asserted that ‘it takes mass unemployment across the whole nation to get carpenters to move out of Nevada’. You cannot even dredge up an out-of-context quote for that monstrously made-up opinion.
Cochrane is correct. He did not write "We should have a recession. People who spend their lives pounding nails in Nevada need something else to do". But he is incorrect in leaving readers with the impression that it is an "attribution" from a patient discussion with Bloomberg reporter John Lippert.
Cochrane made the "we should have a recession... pounding nails in Nevada" claim in the keynote speech he gave at the 2008 CRSP Forum <https://tinyurl.com/2020-06-16-l> at the Gleacher Center in downtown Chicago. He made the claim in front of several hundred people, each of whom the University of Chicago had charged 650 (350 for academics) to attend.
And he said something very similar to what Lippert heard to the _New Yorker's_ John Cassidy <https://tinyurl.com/2020-06-16-m>:
>When we discover we made too many houses in Nevada some people are going to have to move to different jobs, and it is going to take them a while of looking to find the right job for them. There will be some unemployment.... Some component of unemployment is people searching for better fits after shifts that have to happen.... Is ten per cent [unemployment right now] the right number? Now we are talking opinions.... But what we need is models, data, predictions... not my opinion versus your opinion...
As I wrote to reporter John Lippert about Cochrane's paragraph explaining away his "pounding nails in Nevada": "Something that’s a keynote speech at a conference that’s an important part of your school’s public intellectual/fundraising outreach effort hardly seems to be well-characterized as: 'I did not write this. It is an attribution, taken out of context, from an article, written by a reporter with whom I spent about 10 hours patiently trying to explain some basics, and who also turned out only to be on a hunt for embarrassing quotes…' I mean: why do this? Is the belief that nobody will check?"
John Lippert responded:
>Thanks for your note.
>Professor Cochrane's complaint is something of which I became aware... after Cochrane responded to something Paul Krugman had written.... Bloomberg did not respond to Cochrane's comments. He never sent them to us, despite my request that he do so.
>When we became aware of his complaint, we saw no reason to make a correction.
Bloomberg's institutional view was and is: since we are happy with our story, there is no need for us on our own initiative to put forward any correction; and in the absence of a complaint, there is no need for us to expand on our story.
And I think I am done here.
----
>nbviewer: <https://nbviewer.jupyter.org/github/braddelong/weblog-support/blob/master/2020-06-16-pounding-nails-in-nevada.ipynb>
>github: <https://github.com/braddelong/weblog-support/blob/master/2020-06-16-pounding-nails-in-nevada.ipynb>
| github_jupyter |
# Time series forecasting using ARIMA
### Import necessary libraries
```
%matplotlib notebook
import numpy
import pandas
import datetime
import sys
import time
import matplotlib.pyplot as ma
import statsmodels.tsa.seasonal as st
import statsmodels.tsa.arima_model as arima
import statsmodels.tsa.stattools as tools
```
### Load necessary CSV file
```
try:
ts = pandas.read_csv('../../datasets/srv-1-ntt-1h.csv')
except:
print("I am unable to connect to read .csv file", sep=',', header=1)
ts.index = pandas.to_datetime(ts['ts'])
# delete unnecessary columns
del ts['id']
del ts['ts']
del ts['min']
del ts['max']
del ts['sum']
del ts['cnt']
# print table info
ts.info()
```
### Get values from specified range
```
ts = ts['2018-06-16':'2018-07-15']
```
### Remove possible zero and NA values (by interpolation)
We are using MAPE formula for counting the final score, so there cannot occure any zero values in the time series. Replace them with NA values. NA values are later explicitely removed by linear interpolation.
```
def print_values_stats():
print("Zero Values:\n",sum([(1 if x == 0 else 0) for x in ts.values]),"\n\nMissing Values:\n",ts.isnull().sum(),"\n\nFilled in Values:\n",ts.notnull().sum(), "\n")
idx = pandas.date_range(ts.index.min(), ts.index.max(), freq="1h")
ts = ts.reindex(idx, fill_value=None)
print("Before interpolation:\n")
print_values_stats()
ts = ts.replace(0, numpy.nan)
ts = ts.interpolate(limit_direction="both")
print("After interpolation:\n")
print_values_stats()
```
### Plot values
```
# Idea: Plot figure now and do not wait on ma.show() at the end of the notebook
ma.ion()
ma.show()
fig1 = ma.figure(1)
ma.plot(ts, color="blue")
ma.draw()
try:
ma.pause(0.001) # throws NotImplementedError, ignore it
except:
pass
```
### Ignore timestamps, make the time series single dimensional
Since now the time series is represented by continuous single-dimensional Python list. ARIMA does not need timestamps or any irrelevant data.
```
dates = ts.index # save dates for further use
ts = [x[0] for x in ts.values]
```
### Split time series into train and test series
We have decided to split train and test time series by two weeks.
```
train_data_length = 24*7
ts_train = ts[:train_data_length]
ts_test = ts[train_data_length+1:]
```
### Estimate integrated (I) parameter
Check time series stationarity and estimate it's integrated parameter (maximum integration value is 2). The series itself is highly seasonal, so we can assume that the time series is not stationary.
```
def check_stationarity(ts, critic_value=0.05):
try:
result = tools.adfuller(ts)
return result[0] < 0.0 and result[1] < critic_value
except:
# Program may raise an exception when there are NA values in TS
return False
integrate_param = 0
ts_copy = pandas.Series(ts_train, copy=True) # Create copy for stationarizing
while not check_stationarity(ts_copy) and integrate_param < 2:
integrate_param += 1
ts_copy = ts_copy - ts_copy.shift()
ts_copy.dropna(inplace=True) # Remove initial NA values
print("Estimated integrated (I) parameter: ", integrate_param, "\n")
```
### Print ACF and PACF graphs for AR(p) and MA(q) order estimation
AutoCorellation and Parcial AutoCorellation Functions are necessary for ARMA order estimation. Configure the *NLagsACF* and *NlagsPACF* variables for number of lagged values in ACF and PACF graphs.
```
def plot_bar(ts, horizontal_line=None):
ma.bar(range(0, len(ts)), ts, width=0.5)
ma.axhline(0)
if horizontal_line != None:
ma.axhline(horizontal_line, linestyle="-")
ma.axhline(-horizontal_line, linestyle="-")
ma.draw()
try:
ma.pause(0.001) # throws NotImplementedError, ignore it
except:
pass
NlagsACF = 200
NLagsPACF = 40
# ACF
ma.figure(2)
plot_bar(tools.acf(ts_train, nlags=NlagsACF), 1.96 / numpy.sqrt(len(ts)))
# PACF
ma.figure(3)
plot_bar(tools.pacf(ts_train, nlags=NLagsPACF), 1.96 / numpy.sqrt(len(ts)))
```
### ARIMA order estimation and prediction configuration
According to the Box-Jenkins model (https://www.itl.nist.gov/div898/handbook/pmc/section4/pmc446.htm) we assumed that this time series is an AR(p) model. In the ACF graph we can see that the values are positively and negatively altering (not regularly). Stationarity test suggested that the series is already stationarized, but since there are high-valued spikes its better to difference it.
In the PACF graph we can see that there is one significant spike at index 0 and also the data consist of high, irregular anomalies, that's why it's better to pick AR(1) model.
You can specify how many values you want to use for ARIMA model fitting (by setting *N_train_data* variable) and how many new values you want to predict in single step (by setting *N_values_to_forecast* variable).
```
ARIMA_order = (1,1,0)
M_train_data = sys.maxsize
N_values_to_forecast = 1
```
### Forecast new values
Unexpectedly, we have a very large time series (over 8 thousand samples), so the forecasting takes much time.
```
predictions = []
confidence = []
print("Forecasting started...")
start_time = time.time()
ts_len = len(ts)
for i in range(train_data_length+1, ts_len, N_values_to_forecast):
try:
start = i-M_train_data if i-M_train_data >= 0 else 0
arima_model = arima.ARIMA(ts[start:i], order=ARIMA_order).fit(disp=0)
forecast = arima_model.forecast(steps=N_values_to_forecast)
for j in range(0, N_values_to_forecast):
predictions.append(forecast[0][j])
confidence.append(forecast[2][j])
except:
print("Error during forecast: ", i, i+N_values_to_forecast)
# Push back last successful predictions
for j in range(0, N_values_to_forecast):
predictions.append(predictions[-1] if len(predictions) > 0 else 0)
confidence.append(confidence[-1] if len(confidence) > 0 else 0)
print("Forecasting finished")
print("Time elapsed: ", time.time() - start_time)
```
### Count mean absolute percentage error
We use MAPE (https://www.forecastpro.com/Trends/forecasting101August2011.html) instead of MSE because the result of MAPE does not depend on size of values.
```
values_sum = 0
for value in zip(ts_test, predictions):
actual = value[0]
predicted = value[1]
values_sum += abs((actual - predicted) / actual)
values_sum *= 100/len(predictions)
print("MAPE: ", values_sum, "%\n")
```
### Plot forecasted values
```
fig2 = ma.figure(4)
ma.plot(ts_test, color="blue")
ma.plot(predictions, color="red")
ts_len = len(ts)
date_offset_indices = ts_len // 6
num_date_ticks = ts_len // date_offset_indices + 1
ma.xticks(range(0, ts_len, date_offset_indices), [x.date().strftime('%Y-%m-%d') for x in dates[::date_offset_indices]])
ma.draw()
```
| github_jupyter |
```
import os
import re
import sklearn
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from collections import Counter
from sklearn.metrics import *
from sklearn.linear_model import *
from sklearn.model_selection import *
pd.set_option('display.max_columns', None)
all_games_df = pd.read_csv('all_games_df.csv')
all_games_df = all_games_df[['is_tourney','Season','HTeamID','RTeamID','Hwin']]
all_games_df.head()
all_games_df.tail()
kp_df = pd.read_csv('kp_all.csv')
kp_df.sort_index()
kp_df.tail()
regseason_df = pd.read_csv('regseason_df.csv')
regseason_df = regseason_df[['TeamID','Season','wins_top25','PointMargin','FG','FG3']]
regseason_df.head()
regseason_df.tail()
massey_df = pd.read_csv('C:/Users/FLUXNATURE/Desktop/New Kaggle world/NCAAM/MMasseyOrdinals.csv')
POM_df = massey_df[massey_df['SystemName'].str.contains("POM")]
POM_end_df = POM_df.loc[POM_df['RankingDayNum'] == 133]
POM_end_df.rename(columns={'OrdinalRank': 'RankPOM'}, inplace=True)
POM_end_df = POM_end_df[['Season','TeamID','RankPOM']]
POM_end_df.head()
POM_end_df.tail()
```
Create test data set
```
#Test set (this sets the data up in the format Kaggle needs for scoring)
df_seeds = pd.read_csv('C:/Users/FLUXNATURE/Desktop/New Kaggle world/NCAAM/MNCAATourneySeeds.csv')
df_seeds = df_seeds[df_seeds['Season']>=2015]
df_19_tourney = df_seeds.merge(df_seeds, how='inner', on='Season')
df_19_tourney = df_19_tourney[df_19_tourney['TeamID_x'] < df_19_tourney['TeamID_y']]
df_19_tourney['ID'] = df_19_tourney['Season'].astype(str) + '_' \
+ df_19_tourney['TeamID_x'].astype(str) + '_' \
+ df_19_tourney['TeamID_y'].astype(str)
df_19_tourney['SeedInt_x'] = [int(x[1:3]) for x in df_19_tourney['Seed_x']]
df_19_tourney['SeedInt_y'] = [int(x[1:3]) for x in df_19_tourney['Seed_y']]
#Make home team lower seed (consistent with training data)
df_19_tourney.loc[df_19_tourney['SeedInt_x']<df_19_tourney['SeedInt_y'], 'HTeamID'] = df_19_tourney['TeamID_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']>df_19_tourney['SeedInt_y'], 'HTeamID'] = df_19_tourney['TeamID_y']
df_19_tourney.loc[df_19_tourney['SeedInt_x']<df_19_tourney['SeedInt_y'], 'RTeamID'] = df_19_tourney['TeamID_y']
df_19_tourney.loc[df_19_tourney['SeedInt_x']>df_19_tourney['SeedInt_y'], 'RTeamID'] = df_19_tourney['TeamID_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']==df_19_tourney['SeedInt_y'], 'HTeamID'] = df_19_tourney['TeamID_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']==df_19_tourney['SeedInt_y'], 'RTeamID'] = df_19_tourney['TeamID_y']
df_19_tourney.loc[df_19_tourney['SeedInt_x']<df_19_tourney['SeedInt_y'], 'HSeed'] = df_19_tourney['SeedInt_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']>df_19_tourney['SeedInt_y'], 'HSeed'] = df_19_tourney['SeedInt_y']
df_19_tourney.loc[df_19_tourney['SeedInt_x']<df_19_tourney['SeedInt_y'], 'RSeed'] = df_19_tourney['SeedInt_y']
df_19_tourney.loc[df_19_tourney['SeedInt_x']>df_19_tourney['SeedInt_y'], 'RSeed'] = df_19_tourney['SeedInt_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']==df_19_tourney['SeedInt_y'], 'HSeed'] = df_19_tourney['SeedInt_x']
df_19_tourney.loc[df_19_tourney['SeedInt_x']==df_19_tourney['SeedInt_y'], 'RSeed'] = df_19_tourney['SeedInt_y']
df_19_tourney['is_tourney'] = 1
df_19_tourney = df_19_tourney.drop(['Seed_x','Seed_y','TeamID_x','TeamID_y','SeedInt_x','SeedInt_y'], axis=1)
df_19_tourney.sort_index()
home_road = ['H','R']
for hr in home_road:
df_19_tourney = pd.merge(df_19_tourney, regseason_df, left_on=['Season',hr+'TeamID'], right_on = ['Season','TeamID'], how='left')
df_19_tourney.rename(columns={'wins_top25': hr+'wins_top25'}, inplace=True)
df_19_tourney.rename(columns={'PointMargin': hr+'PointMargin'}, inplace=True)
df_19_tourney.rename(columns={'FG': hr+'FG'}, inplace=True)
df_19_tourney.rename(columns={'FG3': hr+'FG3'}, inplace=True)
df_19_tourney = df_19_tourney.drop(['TeamID'], axis=1)
df_19_tourney.sort_index()
for hr in home_road:
df_19_tourney = pd.merge(df_19_tourney, POM_end_df, left_on=['Season',hr+'TeamID'], right_on = ['Season','TeamID'], how='left')
df_19_tourney.rename(columns={'RankPOM': hr+'RankPOM'}, inplace=True)
df_19_tourney = df_19_tourney.drop(['TeamID'], axis=1)
df_19_tourney.sort_index()
efficiency_list = ['conf','adjem','adjo','adjd','luck', 'TeamID']
for hr in home_road:
df_19_tourney = pd.merge(df_19_tourney, kp_df, left_on=[hr+'TeamID','Season'], right_on = ['TeamID','Season'], how='left')
df_19_tourney = df_19_tourney.drop(['TeamID'], axis=1)
for metric in efficiency_list:
df_19_tourney.rename(columns={metric: hr+metric}, inplace=True)
if hr == 'H':
df_19_tourney.rename(columns={'team': 'home'}, inplace=True)
if hr == 'R':
df_19_tourney.rename(columns={'team': 'road'}, inplace=True)
df_19_tourney.sort_index()
df_19_tourney['Htourny20plus'] = 0
df_19_tourney['Rtourny20plus'] = 0
experienced_teams = ['kansas','north carolina','kentucky','duke','michigan st.','wisconsin','florida','villanova','gonzaga','louisville','arizona','xavier','connecticut','syracuse','butler','ohio st.','ucla','west virginia','texas','michigan','pittsburgh','memphis','oregon']
for team in experienced_teams:
df_19_tourney.loc[df_19_tourney['home']==team, 'Htourny20plus'] = 1
df_19_tourney.loc[df_19_tourney['road']==team, 'Rtourny20plus'] = 1
df_19_tourney.sort_index()
df_19_tourney['HBig4Conf'] = 0
df_19_tourney['RBig4Conf'] = 0
conferences = ['ACC','B10','B12','SEC']
for conf in conferences:
df_19_tourney.loc[df_19_tourney['Hconf']==conf, 'HBig4Conf'] = 1
df_19_tourney.loc[df_19_tourney['Rconf']==conf, 'RBig4Conf'] = 1
list(df_19_tourney)
df_19_tourney.sort_index()
df_19_tourney = df_19_tourney.fillna(df_19_tourney.mean())
df_19_tourney.to_csv('test_combos_df_2015.csv', index=False)
```
| github_jupyter |
```
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import higher
from transformers import GPT2Tokenizer, GPT2LMHeadModel
import utils
%load_ext autoreload
%autoreload 2
```
## Load Model, Data
```
device = 'cuda' if torch.cuda.is_available() else 'cpu'
```
Try using GPT2 vs distilgpt2 (KL div actually goes up though)
```
def loadOTSModel():
model = GPT2LMHeadModel.from_pretrained("gpt2")
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
tokenizer.pad_token = tokenizer.eos_token
return model, tokenizer
model, tokenizer = loadOTSModel()
dataloader = utils.retrieveDataloader(
tokenizer,
bs=1,
dataset='train'
)
len(dataloader)
for train_step, (lm_data, edit_example, _) in enumerate(dataloader):
lm_tokens, lm_mask = lm_data
lm_tokens, lm_mask = lm_tokens.to(device), lm_mask.to(device)
edit_tokens, edit_mask = edit_example
edit_tokens, edit_mask = edit_tokens.to(device), edit_mask.to(device)
lm_labels = lm_tokens.masked_fill(lm_mask == 0, -100)
edit_labels = edit_tokens.masked_fill(edit_mask == 0, -100)
break
```
## Double Check Data
```
lm_tokens
lm_mask
lm_labels
tokenizer.decode(lm_tokens[lm_labels != -100])
tokenizer.decode(lm_labels[lm_labels != -100])
tokenizer.decode(lm_tokens[lm_labels != -100]) == tokenizer.decode(lm_labels[lm_labels != -100])
```
## KL Divergence on same model, same data, model.train()
```
model.train()
model.to(device)
model_out1 = model(lm_tokens, attention_mask=lm_mask, labels=lm_labels)
model_out2 = model(lm_tokens, attention_mask=lm_mask, labels=lm_labels)
model_out1.loss
model_out2.loss
model_out1.logits
model_out1.logits - model_out2.logits
model_out1.logits.shape
```
This seems like the wrong dimension
```
torch.sum(F.softmax(model_out1.logits, dim=1), dim=1)
torch.sum(F.softmax(model_out1.logits, dim=1), dim=1).shape
F.kl_div(
F.log_softmax(model_out1.logits, dim=1),
F.log_softmax(model_out2.logits, dim=1),
reduction='batchmean',
log_target=True
)
```
This seems like the right dimension - softmax over the 50257 words in vocab for each position in sequence 1-200
```
torch.sum(F.softmax(model_out1.logits, dim=-1), dim=-1)
torch.sum(F.softmax(model_out1.logits, dim=-1), dim=-1).shape
F.kl_div(
F.log_softmax(model_out1.logits, dim=-1),
F.log_softmax(model_out2.logits, dim=-1),
reduction='batchmean',
log_target=True
)
kl_loss = nn.KLDivLoss(reduction = 'batchmean')
l_loc = kl_loss(
F.log_softmax(model_out1.logits, dim=-1),
F.softmax(model_out2.logits, dim=-1)
)
l_loc
```
## KL Divergence on same model, same data, model.eval()
```
model.eval()
model_eval1 = model(lm_tokens, attention_mask=lm_mask, labels=lm_labels)
model_eval2 = model(lm_tokens, attention_mask=lm_mask, labels=lm_labels)
F.kl_div(
F.log_softmax(model_eval1.logits, dim=-1),
F.log_softmax(model_eval2.logits, dim=-1),
reduction='batchmean',
log_target=True
)
```
Large KL divergence coming from dropout?
## From editable code:
```
def edit(self, inputs, targets, max_steps=None, model_kwargs=None, loss_kwargs=None, opt_kwargs=None, **kwargs):
"""
Attempts to edit model (out-of-place) and return an edited copy
:param inputs: data that is fed into the model
:param targets: reference answers that are fed into loss function
:param max_steps: after this many gradient steps the process is terminated
:param model_kwargs: optional extra model inputs, used as model(inputs, **model_params)
:param loss_kwargs: optional extra loss parameters, self.loss_function(model(inputs), targets, **loss_params)
:param opt_kwargs: optional overrides for optimizer.get_initial_state
:param kwargs: extra parameters passed to optimizer.step
:returns: edited_model, is_edit_successful, final_loss, gradients_steps
:rtype: Editable.EditResult
"""
model_kwargs, loss_kwargs, opt_kwargs = model_kwargs or {}, loss_kwargs or {}, opt_kwargs or {}
optimizer_state = self.optimizer.get_initial_state(self, **opt_kwargs)
editable = self
for step in count():
prediction = editable(inputs, **model_kwargs)
loss = self.loss_function(prediction, targets, **loss_kwargs)
if self.is_edit_finished(**locals()):
return self.EditResult(editable, success=True, loss=loss, complexity=step)
elif step >= (max_steps or self.max_steps):
return self.EditResult(editable, success=False, loss=loss, complexity=step)
optimizer_state, editable = self.optimizer.step(
optimizer_state, editable, loss, parameters=editable.get_editable_parameters(editable.module), **kwargs)
def train_on_batch(self, x_batch, y_batch, x_edit, y_edit, prefix='train/', is_train=True, **kwargs):
""" Performs a single gradient update and reports metrics """
x_batch, y_batch = map(torch.as_tensor, (x_batch, y_batch))
self.opt.zero_grad()
with training_mode(self.model, is_train=is_train):
logits = self.model(x_batch)
main_loss = self.loss_function(logits, y_batch).mean()
with training_mode(self.model, is_train=False):
model_edited, success, editability_loss, complexity = self.model.edit(x_edit, y_edit, **kwargs)
logits_updated = model_edited(x_batch)
stability_loss = - (F.softmax(logits.detach(), dim=1) * F.log_softmax(logits_updated, dim=1)).sum(dim=1).mean()
final_loss = main_loss + self.stability_coeff * stability_loss + self.editability_coeff * editability_loss
metrics = dict(
final_loss=final_loss.item(), stability_loss=stability_loss.item(),
editability_loss=editability_loss.item(), main_loss=main_loss.item(),
)
final_loss.backward()
if self.max_norm is not None:
metrics['grad_norm'] = torch.nn.utils.clip_grad_norm_(self.model.parameters(), max_norm=self.max_norm)
self.opt.step()
return self.record(**metrics, prefix=prefix)
```
Note stability loss:
```
-(F.softmax(model_out2.logits.detach(), dim=1) * F.log_softmax(model_out1.logits, dim=1)).sum(dim=1).mean()
```
Might need to change dimensions though:
```
-(F.softmax(model_out2.logits.detach(), dim=-1) * F.log_softmax(model_out1.logits, dim=-1)).sum(dim=-1).mean()
P = F.log_softmax(model_out2.logits.detach(), dim=-1)
Q = F.log_softmax(model_out1.logits, dim=-1)
(P * (P / Q).log()).sum()
stability_loss = (
F.softmax(base_out.logits.detach(), dim=-1)
* (F.log_softmax(base_out.logits.detach(), dim=-1) - F.log_softmax(edited_base_out.logits, dim=-1))
).sum(-1).mean()
(P * (P / Q).log()).sum()
(P * (P / Q).log()).sum(-1).mean()
F.kl_div(
F.log_softmax(model_out1.logits, dim=-1),
F.log_softmax(model_out2.logits, dim=-1),
reduction='mean',
log_target=True
)
(P * -Q.log()).sum(dim=-1).mean()
# F.kl_div(Q.log(), P, None, None, 'sum')
```
Conclusions:
* Use different stability loss
* Double check on softmax dimensions in KL div
* Take original model logits in training mode
* Performs edit function on model in eval mode (`with training_mode(self.model, is_train=False):`)
## Checking Errors
```
errpath = 'errors/errors_20210311.19.03.1615490257'
!ls errors/errors_20210311.19.03.1615490257
edit_tokens = torch.load(f"{errpath}/edit_tokens_14")
ent_tokens = torch.load(f"{errpath}/ent_tokens_14")
edit_tokens
tokenizer.decode(edit_tokens.squeeze())
tokenizer.decode(ent_tokens.squeeze())
torch.tensor(tokenizer.encode('Baker'))
ent_tokens = torch.tensor(tokenizer.encode('Baker'))
edit_tokens.squeeze().unsqueeze(0)
edit_tokens.shape
ent_tokens.shape
torch.gather(edit_tokens, 0, ent_tokens.unsqueeze(1))
max(np.in1d(edit_tokens, ent_tokens))
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import higher
from transformers import GPT2Tokenizer, GPT2LMHeadModel
import utils
import data_process as dp
%load_ext autoreload
%autoreload 2
dataloader = utils.retrieveDataloader(
tokenizer,
bs=1,
dataset='train'
)
import glob
writtenFiles = (
glob.glob("../data/permuted*")
)
max_obs = float("inf")
fileIndex = max(map(lambda x: int(x.split(".")[-1]), writtenFiles))
limitIndex = min(max_obs, fileIndex)
limitIndex
dataset = dp.TorchDataset(list(range(limitIndex)), tokenizer, 'train')
# for idx, (raw, perm, ent) in enumerate(dataset):
# print(idx, ent[0])
# if idx > 15:
# break
for train_step, (lm_data, edit_example, ent) in enumerate(dataloader):
lm_tokens, lm_mask = lm_data
lm_labels = lm_tokens.masked_fill(lm_mask == 0, -100)
edit_tokens, edit_mask = edit_example
ent_tokens = ent[0].flatten()
ent_tokens = ent_tokens[ent_tokens != 50256]
# edit_locs = locateEntityEdit(edit_tokens, ent_tokens)
if train_step == 14:
print(ent_tokens)
print(edit_tokens)
print(lm_tokens)
test = max(np.in1d(edit_tokens, ent_tokens))
print(train_step,test)
if not test:
print(ent_tokens)
if train_step > 20:
break
```
| github_jupyter |
# Matplotlib
---
```
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
arr = np.tan(np.linspace(-np.pi, np.pi, num=50))
arr
plt.plot(arr)
np.info(plt.figure)
fig = plt.figure(edgecolor='red')
dir(fig)
fig.show()
fig = plt.figure(edgecolor='blue')
fig.add_subplot(111)
plt.plot(np.linspace(1, 99))
fig.show()
np.info(fig.add_subplot)
arr = np.linspace(-np.pi, np.pi, num=50)
plt.plot(arr, np.sin(arr))
plt.plot(arr, np.cos(arr))
plt.show()
np.info(plt.plot)
plt.plot(arr, np.sin(arr), color='c', linewidth=2)
plt.plot(arr, np.cos(arr), color='m', linewidth=2)
plt.show()
plt.plot(arr, linestyle='--', linewidth=2)
plt.plot(np.linspace(0, 100, 10), linestyle='--', marker='o', linewidth=2)
plt.plot(np.linspace(0, 100, 10), linestyle='--', marker='x', linewidth=1)
plt.plot(arr, np.sin(arr), color='c', linewidth=2)
plt.xlim(-3.5, 3.5)
plt.ylim(-1.5, 1.5)
plt.show()
y_sin = np.sin(arr)
plt.plot(arr, y_sin, color='c', linewidth=2)
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.show()
y_sin = np.sin(arr)
plt.plot(arr, y_sin, color='c', linewidth=2)
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi])
plt.show()
y_sin = np.sin(arr)
plt.plot(arr, y_sin, color='c', linewidth=2)
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],
[r'$-\pi$', r'$-\pi/2$', r'$0$', r'$\pi/2$', r'$\pi$'])
plt.show()
np.info(plt.xticks)
y_sin = np.sin(arr)
plt.plot(arr, y_sin, color='c', linewidth=2, label="sin(x)")
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],
[r'$-\pi$', r'$-\pi/2$', r'$0$', r'$\pi/2$', r'$\pi$'])
plt.yticks([-1, 0, 1])
plt.legend()
plt.show()
np.info(plt.legend)
y_sin = np.sin(arr)
y_cos = np.cos(arr)
plt.plot(arr, y_sin, color='c', linewidth=2, label="sin(x)")
plt.plot(arr, y_cos, color='m', linewidth=2, label="cos(x)")
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],
[r'$-\pi$', r'$-\pi/2$', r'$0$', r'$\pi/2$', r'$\pi$'])
plt.yticks([-1, 0, 1])
plt.legend(loc='lower right', frameon=False, fontsize='large')
plt.show()
plt.figure(1)
plt.plot(np.linspace(0, 100, num=10), label='Hello')
plt.show()
np.info(plt.subplot)
plt.figure(1)
# number of rows, number of columns, number of plot
plt.subplot(211) # plt.subplot(2, 1, 1)
plt.plot(arr, y_sin, color='c', linewidth=2, label="sin(x)")
plt.subplot(212)
plt.plot(arr, y_cos, color='m', linewidth=2, label="cos(x)")
plt.show()
plt.figure(1)
# number of rows, number of columns, number of plot
plt.subplot(2, 2, 2) # plt.subplot(2, 1, 1)
plt.plot(arr, y_sin, color='c', linewidth=2, label="sin(x)")
plt.subplot(2, 2, 3)
plt.plot(arr, y_cos, color='m', linewidth=2, label="cos(x)")
plt.show()
y_sin = np.sin(arr)
y_cos = np.cos(arr)
plt.plot(arr, y_sin, color='c', linewidth=2, label="sin(x)")
plt.plot(arr, y_cos, color='m', linewidth=2, label="cos(x)")
plt.xlim(arr.min()*1.1, arr.max()*1.1)
plt.ylim(y_sin.min()*1.1, y_sin.max()*1.1)
plt.xticks([-np.pi, -np.pi/2, 0, np.pi/2, np.pi],
[r'$-\pi$', r'$-\pi/2$', r'$0$', r'$\pi/2$', r'$\pi$'])
plt.yticks([-1, 0, 1])
plt.legend(loc='lower right', frameon=False, fontsize='large')
plt.show()
np.info(plt.gca)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', 500)
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm
color = sns.color_palette()
%matplotlib inline
tr_queries = pd.read_csv('data/train_queries.csv')
te_queries = pd.read_csv('data/test_queries.csv')
tr_plans = pd.read_csv('data/train_plans.csv')
te_plans = pd.read_csv('data/test_plans.csv')
tr_click = pd.read_csv('data/train_clicks.csv')
# t_order = 't_order.csv'
# t_user = 't_user.csv'
from tqdm import tqdm_notebook
import json
train_clicks.tail()
print(train_clicks['click_mode'].value_counts())
# plt.hist(df_click['pid'],rwidth = 0.2)
train_clicks['click_mode'].hist(bins=10) #Series
train_plans.tail()
train_queries.tail()
tr_data = tr_queries.merge(tr_click, on='sid', how='left')
tr_data = tr_data.merge(tr_plans, on='sid', how='left')
tr_data = tr_data.drop(['click_time'], axis=1)
tr_data['click_mode'] = tr_data['click_mode'].fillna(0)
te_data = te_queries.merge(te_plans, on='sid', how='left')
te_data['click_mode'] = -1
data = pd.concat([tr_data, te_data], axis=0)
data = data.drop(['plan_time'], axis=1)
data = data.reset_index(drop=True)
print('total data size: {}'.format(data.shape))
print('raw data columns: {}'.format(', '.join(data.columns)))
tr_data.tail()
te_data.tail()
data.tail()
def gen_od_feas(data):
data['o1'] = data['o'].apply(lambda x: float(x.split(',')[0]))
data['o2'] = data['o'].apply(lambda x: float(x.split(',')[1]))
data['d1'] = data['d'].apply(lambda x: float(x.split(',')[0]))
data['d2'] = data['d'].apply(lambda x: float(x.split(',')[1]))
data = data.drop(['o', 'd'], axis=1)
return data
data = gen_od_feas(data)
data.tail()
data['plans'][0]
n = data.shape[0]
mode_list_feas = np.zeros((n, 22))
speed, pricePerDis = np.zeros((n,11)), np.zeros((n,11))
for i, plan in tqdm_notebook(enumerate(data['plans'].values)):
# if i == 5:
# break
try:
cur_plan_list = json.loads(plan)
except:
cur_plan_list = []
if len(cur_plan_list) == 0:
speed[i] = speed[i]-1
pricePerDis[i] = pricePerDis[i]-1
else:
for tmp_dit in cur_plan_list:
# print(tmp_dit['distance'])
# print(tmp_dit['eta'])
_speed = round(int(tmp_dit['distance'])/int(tmp_dit['eta']) , 2)
if tmp_dit['price'] == '':
tmp_dit['price'] = 0
_pricePerDis = round(int(tmp_dit['price'])/int(tmp_dit['distance']), 2)
speed[i][int(tmp_dit['transport_mode'])-1] = _speed
pricePerDis[i][int(tmp_dit['transport_mode'])-1] = _pricePerDis
feature_data = pd.DataFrame(mode_list_feas)
feature_data.columns = ['fea_{}'.format(i) for i in range(22)]
for i in range(22):
if i%2 == 0:
feature_data['fea_{}'.format(i)] = speed[:,int(i/2)]
else:
feature_data['fea_{}'.format(i)] = pricePerDis[:,int(i/2)]
data = pd.concat([data, feature_data], axis=1)
data = data.drop(['plans'], axis=1)
from sklearn.decomposition import TruncatedSVD
def read_profile_data():
profile_data = pd.read_csv('data/profiles.csv')
profile_na = np.zeros(67)
profile_na[0] = -1
profile_na = pd.DataFrame(profile_na.reshape(1, -1))
profile_na.columns = profile_data.columns
profile_data = profile_data.append(profile_na)
return profile_data
profile_data = read_profile_data()
x = profile_data.drop(['pid'], axis=1).values
svd = TruncatedSVD(n_components=20, n_iter=20, random_state=2019)
svd_x = svd.fit_transform(x)
svd_feas = pd.DataFrame(svd_x)
svd_feas.columns = ['svd_fea_{}'.format(i) for i in range(20)]
svd_feas['pid'] = profile_data['pid'].values
data['pid'] = data['pid'].fillna(-1)
data = data.merge(svd_feas, on='pid', how='left')
data['req_time'] = pd.to_datetime(data['req_time'])
data['weekday'] = data['req_time'].dt.dayofweek
data['hour'] = data['req_time'].dt.hour
data = data.drop(['req_time'], axis=1)
data.to_csv("data.csv")
train_data = data[data['click_mode'] != -1]
test_data = data[data['click_mode'] == -1]
submit = test_data[['sid']].copy()
train_data = train_data.drop(['sid', 'pid'], axis=1)
test_data = test_data.drop(['sid', 'pid'], axis=1)
test_data = test_data.drop(['click_mode'], axis=1)
train_y = train_data['click_mode'].values
train_x = train_data.drop(['click_mode'], axis=1)
train_x.columns
feature_columns_to_use = ['o1', 'o2', 'd1', 'd2', 'fea_0', 'fea_1', 'fea_2', 'fea_3', 'fea_4',
'fea_5', 'fea_6', 'fea_7', 'fea_8', 'fea_9', 'fea_10', 'fea_11',
'fea_12', 'fea_13', 'fea_14', 'fea_15', 'fea_16', 'fea_17', 'fea_18',
'fea_19', 'fea_20', 'fea_21', 'svd_fea_0', 'svd_fea_1', 'svd_fea_2',
'svd_fea_3', 'svd_fea_4', 'svd_fea_5', 'svd_fea_6', 'svd_fea_7',
'svd_fea_8', 'svd_fea_9', 'svd_fea_10', 'svd_fea_11', 'svd_fea_12',
'svd_fea_13', 'svd_fea_14', 'svd_fea_15', 'svd_fea_16', 'svd_fea_17',
'svd_fea_18', 'svd_fea_19', 'weekday', 'hour']
train_for_matrix = train_x[feature_columns_to_use]
test_for_matrix = test_data[feature_columns_to_use]
train_X = train_for_matrix.as_matrix()
test_X = test_for_matrix.as_matrix()
train_y
import xgboost as xgb
from sklearn import preprocessing
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import f1_score
from time import gmtime, strftime
gbm = xgb.XGBClassifier(silent=1, max_depth=10, n_estimators=1000, learning_rate=0.05)
gbm.fit(train_X, train_y)
predictions = gbm.predict(test_X)
now_time = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
submit['mode'] = predictions
submit.to_csv(
'submit/{}_result_{}.csv'.format(model_name, now_time), index=False)
```
| github_jupyter |
# Example of the `aitlas` toolbox in the context of image segmentation
This notebook shows a sample implementation of a image segmentation using the `aitlas` toolbox.
```
%matplotlib inline
```
Import the required packages
```
import matplotlib.pyplot as plt
from PIL import Image
import os
from aitlas.datasets import ChactunDataset
from aitlas.models import DeepLabV3
from aitlas.tasks.predict import ImageFolderDataset
from aitlas.visualizations import display_image_segmentation
```
## Visualize images and masks
```
example_image = Image.open('/media/ssd/Chactun/train/tile_1185_lidar.tif')
example_mask1 = Image.open('/media/ssd/Chactun/train/tile_1185_mask_aguada.tif')
example_mask2 = Image.open('/media/ssd/Chactun/train/tile_1185_mask_building.tif')
example_mask3 = Image.open('/media/ssd/Chactun/train/tile_1185_mask_platform.tif')
plt.rcParams["figure.figsize"] = (18, 9) # (w, h)
plt.subplot(1,4,1)
plt.imshow(example_image)
plt.title(f"Random training example")
plt.axis('off')
plt.subplot(1,4,2)
plt.imshow(example_mask1, cmap='gray')
plt.title(f"Mask for aguada")
plt.axis('off')
plt.subplot(1,4,3)
plt.imshow(example_mask2, cmap='Greys', interpolation='nearest')
plt.title(f"Mask for building")
plt.axis('off')
plt.subplot(1,4,4)
plt.imshow(example_mask3, cmap='Greys', interpolation='nearest')
plt.title(f"Mask for platform")
plt.axis('off')
plt.show()
```
## Load data
```
train_dataset_config = {
"batch_size": 4,
"shuffle": True,
"num_workers": 4,
"root": "/media/ssd/chactun_new/train",
"transforms": ["aitlas.transforms.MinMaxNormTransponse"],
"target_transforms": ["aitlas.transforms.MinMaxNorm"]
}
train_dataset = ChactunDataset(train_dataset_config)
len(train_dataset)
```
## Training
```
epochs = 5
model_directory = "/media/ssd/chactun_new/experiments/"
model_config = {"num_classes": 3, "learning_rate": 0.0001,"pretrained": True, "threshold": 0.5}
model = DeepLabV3(model_config)
model.prepare()
model.train_model(
train_dataset=train_dataset,
epochs=epochs,
model_directory=model_directory,
run_id='1'
)
```
## Evaluation
```
test_dataset_config = {
"batch_size": 4,
"shuffle": False,
"num_workers": 4,
"root": "/media/ssd/chactun_new/test",
"transforms": ["aitlas.transforms.MinMaxNormTransponse"],
"target_transforms": ["aitlas.transforms.MinMaxNorm"]
}
test_dataset = ChactunDataset(test_dataset_config)
len(test_dataset)
model_config = {"num_classes": 3, "learning_rate": 0.0001,"pretrained": True, "threshold": 0.5}
model = DeepLabV3(model_config)
model.prepare()
model_path = "/media/ssd/chactun_new/experiments/checkpoint.pth.tar"
model.evaluate(dataset=test_dataset, model_path=model_path)
model.running_metrics.get_scores(["iou"])
```
## Predictions
```
# run predictions
model_path = "/media/ssd/chactun_new/experiments/checkpoint.pth.tar"
predict_dir = "/media/ssd/chactun_new/predict/"
results_dir = "/media/ssd/chactun_new/experiments/results"
labels = ["Aguada", "Building", "Platform"]
transforms = ["aitlas.transforms.MinMaxNormTransponse"]
predict_dataset = ImageFolderDataset(predict_dir, labels, transforms)
len(predict_dataset)
# load the model
model.load_model(model_path)
# Run predictions
y_true, y_pred, y_prob = model.predict(dataset=predict_dataset)
# plot predictions
for i, image_path in enumerate(predict_dataset.data):
plot_path = os.path.join(results_dir, f"{predict_dataset.fnames[i]}_plot.png")
display_image_segmentation(
image_path,
y_true[i],
y_pred[i],
y_prob[i],
predict_dataset.labels,
plot_path,
)
```
| github_jupyter |
# Flowlines
This example details dynamic construction and plotting within a python script of synthetic flowlines generated from a plate model. For a general introduction to flowlines, see the GPlates user manual:
http://www.gplates.org/user-manual/Flowlines.html
The example goes through the case of creating flowline/motion path features within a script (assuming that we know enough to assign parameters like position of the seed points, plateid/conjugate plateid, and the time range and time sampling that we want for the feature). Then we can calculate motion paths for one (or several) rotation models, for different frames of reference.
The flowline example uses the Tasman Sea, which in the GPlates default reconstruction model opened from ~90 Ma to ~52 (Ma), with rotation parameters from the study of Gaina et al (1998), which are used in the Seton et al global compilation. The parameters we need to set to make a reasonable synthetic flowlines are as follows:
- The seed points - typically, points on a present day mid-ocean ridge or (in this case) an extinct ridge axis
- Plate IDs for the two plates that were spreading apart - in this case Australia (801) and the Lord Howe Rise (833)
- The time range for the spreading (or at least, the section of the spreading you want to visualise the flowline for)
```
import numpy as np
import pygplates
# Rotation file to be used to generate synthetic flowlines
rotation_filename = 'Data/Seton_etal_ESR2012_2012.1.rot'
# Parameters required to define a flowline (as in GPlates)
SeedPoints = zip([-40,-43],[157,157.5])
left_plate = 801
right_plate = 833
times = np.arange(0,90,1.) # using numpy to get a list of times in 1 Myr increments
```
### Create a flowline feature
The next section of codes generates a flowline feature within the script (analogous to the process in GPlates GUI of digitizing a new point and applying parameters to make it a flowline feature)
```
# CREATE FLOWLINE
# POINTS ON THE FLOWLINE
multi_point = pygplates.MultiPointOnSphere(SeedPoints)
# Create the flowline feature
flowline_feature = pygplates.Feature.create_flowline(
multi_point,
times,
valid_time=(np.max(times), np.min(times)),
left_plate=left_plate,
right_plate=right_plate)
```
### Make the flowline for a chosen reconstruction model
The flowline feature does not have any set flowline path, it is agnositic of rotation models. To make a synthetic flowline,
```
# Load a rotation model (required to generate a synthetic flowline)
rotation_model=pygplates.RotationModel(rotation_filename)
# reconstruct the flowline - we want it in present day coordinates
reconstruction_time=0
# create an empty feature in which the reconstructed flowline geometries will be placed
reconstructed_flowlines = []
# call pygplates to generate the flowline feature
pygplates.reconstruct(flowline_feature, rotation_model, reconstructed_flowlines, reconstruction_time,
anchor_plate_id=1, reconstruct_type=pygplates.ReconstructType.flowline)
```
### Visualise the results
A useful way of assessing synthetic flowlines against observations is to plot them overlain on maps showing seafloor fabric, ideally gravity anomaly maps of the oceans derived from satellite altimetry. The code below plots the flowlines over the (low resolution) etopo raster image distributed with Basemap.
The flowline features are split into 'right' and 'left' sides, so plotting them takes a few lines of code that iterate over each half of each flowline.
```
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import cartopy.io.shapereader as shpreader
import numpy as np
%matplotlib inline
# Create figure
fig = plt.figure(figsize=(12,12),dpi=300)
pmap = fig.add_subplot(111, projection=ccrs.Mercator())
pmap.set_extent([145, 170, -50, -30])
# Create land and ocean features in greyscale
pmap.coastlines(resolution='10m', color='white', linewidth=0.25)
pmap.add_feature(cfeature.LAND, color = 'grey', edgecolor='grey')
pmap.add_feature(cfeature.OCEAN, color='white')
pmap.add_feature(cfeature.LAKES, color='white', edgecolor='white')
# Iterate over the left and right sides that the flowline is split into.
for reconstructed_flowline in reconstructed_flowlines:
# Iterate over the left flowline points
# Create an empty array to hold left flowline points which we append over.
flowlinearray_left = np.empty([0,0])
# Access each left flowline from an array holding the flowline spread along the *left* plate from the
# reconstructed seed point.
for left_point in reconstructed_flowline.get_left_flowline():
# Append the initially empty flowlinearray_left array with the left points obtained from reconstructed_flowline
# which have been turned into values of a lat-lon array.
flowlinearray_left = np.append(flowlinearray_left, left_point.to_lat_lon_array())
# Repeat this iteration for the right flowline points.
flowlinearray_right = np.empty([0,0])
for right_point in reconstructed_flowline.get_right_flowline():
flowlinearray_right = np.append(flowlinearray_right, right_point.to_lat_lon_array())
x,y = flowlinearray_left[1::2],flowlinearray_left[::2]
l1=pmap.plot(x, y, 'r', transform=ccrs.PlateCarree(), zorder=2)
l3=pmap.scatter(x, y, 30, c=times, transform=ccrs.PlateCarree(),
cmap=plt.cm.gnuplot_r, edgecolor='none', zorder=1,vmin=50,vmax=90)
x,y = flowlinearray_right[1::2],flowlinearray_right[::2]
l1=pmap.plot(x, y, 'r', transform=ccrs.PlateCarree(), zorder=2)
l3=pmap.scatter(x, y, 30, c=times, transform=ccrs.PlateCarree(),
cmap=plt.cm.gnuplot_r, edgecolor='none',zorder=1,vmin=50,vmax=90)
# Set a colorbar to help visualise the passage of every millionth year.
fig.colorbar(l3, ax=pmap).set_label('Age (Ma)',fontsize=12)
plt.show()
```
| github_jupyter |
```
%%html
<link href="http://mathbook.pugetsound.edu/beta/mathbook-content.css" rel="stylesheet" type="text/css" />
<link href="https://aimath.org/mathbook/mathbook-add-on.css" rel="stylesheet" type="text/css" />
<style>.subtitle {font-size:medium; display:block}</style>
<link href="https://fonts.googleapis.com/css?family=Open+Sans:400,400italic,600,600italic" rel="stylesheet" type="text/css" />
<link href="https://fonts.googleapis.com/css?family=Inconsolata:400,700&subset=latin,latin-ext" rel="stylesheet" type="text/css" /><!-- Hide this cell. -->
<script>
var cell = $(".container .cell").eq(0), ia = cell.find(".input_area")
if (cell.find(".toggle-button").length == 0) {
ia.after(
$('<button class="toggle-button">Toggle hidden code</button>').click(
function (){ ia.toggle() }
)
)
ia.hide()
}
</script>
```
**Important:** to view this notebook properly you will need to execute the cell above, which assumes you have an Internet connection. It should already be selected, or place your cursor anywhere above to select. Then press the "Run" button in the menu bar above (the right-pointing arrowhead), or press Shift-Enter on your keyboard.
$\newcommand{\identity}{\mathrm{id}}
\newcommand{\notdivide}{\nmid}
\newcommand{\notsubset}{\not\subset}
\newcommand{\lcm}{\operatorname{lcm}}
\newcommand{\gf}{\operatorname{GF}}
\newcommand{\inn}{\operatorname{Inn}}
\newcommand{\aut}{\operatorname{Aut}}
\newcommand{\Hom}{\operatorname{Hom}}
\newcommand{\cis}{\operatorname{cis}}
\newcommand{\chr}{\operatorname{char}}
\newcommand{\Null}{\operatorname{Null}}
\newcommand{\lt}{<}
\newcommand{\gt}{>}
\newcommand{\amp}{&}
$
<div class="mathbook-content"><h2 class="heading hide-type" alt="References 13.5 References and Suggested Readings"><span class="type">Section</span><span class="codenumber">13.5</span><span class="title">References and Suggested Readings</span></h2><a href="struct-references.ipynb" class="permalink">¶</a></div>
<div class="mathbook-content"><article class="bib" id="biblio-57"><div class="bibitem">[1]</div> <div class="bibentry">Hungerford, T. W. . Springer, New York, 1974.</div></article></div>
<div class="mathbook-content"><article class="bib" id="biblio-58"><div class="bibitem">[2]</div> <div class="bibentry">Lang, S. . 3rd ed. Springer, New York, 2002.</div></article></div>
<div class="mathbook-content"><article class="bib" id="biblio-59"><div class="bibitem">[3]</div> <div class="bibentry">Rotman, J. J. . 4th ed. Springer, New York, 1995.</div></article></div>
| github_jupyter |
```
# Reading boston housing dataset
from sklearn.datasets import load_boston
from sklearn.model_selection import train_test_split
import pandas as pd
boston = load_boston()
df = pd.DataFrame(boston.data, columns=boston.feature_names)
df.describe() # describe dataset overview
# save files as csv
import os
WORK_DIRECTORY='data'
os.makedirs('{}'.format(WORK_DIRECTORY), exist_ok=True)
df.to_csv('{}/boston_housing.csv'.format(WORK_DIRECTORY), header=False, index=False)
# S3 prefix
bucket = 'sagemaker-bucket-sample-test'
prefix = 'sagemaker/sample'
# Import libraries
from sagemaker import get_execution_role
import boto3, sys, os
import sagemaker
sagemaker_session = sagemaker.Session()
# Get a SageMaker-compatible role used by this Notebook Instance.
role = get_execution_role()
my_region = boto3.session.Session().region_name # set the region of the instance
print("Execution role is " + role)
print("Success - the MySageMakerInstance is in the " + my_region + ".")
s3 = boto3.resource('s3')
try:
if my_region == 'ap-northeast-1':
s3.create_bucket(Bucket=bucket)
else:
s3.create_bucket(Bucket=bucket, CreateBucketConfiguration={'LocationConstraint': my_region})
print('S3 bucket created successfully')
except Exception as e:
print('S3 error: ', e)
# send data to S3.SageMaker will take training data from s3
training_path = sagemaker_session.upload_data(path='{}/boston_housing.csv'.format(WORK_DIRECTORY), bucket=bucket, key_prefix=prefix)
s3_train_data = 's3://{}/{}/{}'.format(bucket, prefix, WORK_DIRECTORY)
print('Uploaded training data location: {}'.format(s3_train_data))
output_location = 's3://{}/{}/output'.format(bucket, prefix)
print('Training artifacts will be uploaded to: {}'.format(output_location))
# We use the Estimator from the SageMaker Python SDK
from sagemaker.sklearn.estimator import SKLearn
script_path = 'scikit_learn_script.py'
# Initialise SDK
sklearn_estimator = SKLearn(
entry_point=script_path,
role = role,
train_instance_type="ml.c4.xlarge",
sagemaker_session=sagemaker_session,
output_path=output_location
)
print("Estimator object: {}".format(sklearn_estimator))
# Run model training job
sklearn_estimator.fit({'train': training_path})
# Deploy an estimator and endpoint
from sagemaker.predictor import csv_serializer, json_deserializer
predictor = sklearn_estimator.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge", endpoint_name="sagemaker-terraform-test")
# Specify input and output formats.
predictor.content_type = 'text/csv'
predictor.serializer = csv_serializer
predictor.deserializer = json_deserializer
# predictor.delete_endpoint()
```
| github_jupyter |
# Google Colab での使い方
## ランタイムでGPUを選択
まず、「ランタイム」→ 「ランタイムのタイプを変更」のハードウェアアクセラレーターで「GPU」を選びます
```
# GoogleColab かどうか判定
try:
from google.colab import drive
GoogleColab = True
except ModuleNotFoundError:
GoogleColab = False
```
## 推奨の追加コマンド
```
try:
import binarybrain as bb
except ModuleNotFoundError:
from google.colab import drive
!pip install pybind11
!pip install binarybrain
import binarybrain as bb
import os
drive.mount('/content/drive')
work_directory = '/content/drive/My Drive/BinaryBrain'
os.makedirs(work_directory, exist_ok=True)
os.chdir(work_directory)
```
# 以降メンテナンス用コマンド
## TestPyPIからのインストール
```
!pip install pybind11
!pip install --index-url https://test.pypi.org/simple/ binarybrain
```
## PyPIからのインストール
```
!pip install pybind11
!pip install --index-url https://test.pypi.org/simple/ binarybrain
```
## pip のアンインスール
```
!pip uninstall -y binarybrain
```
## setup.py での インストール
下記の実行でインストール可能<br>
インストール後に、一度「ランタイム」→「ランタイムの再起動」を選んで再起動が必要
```
try:
import binarybrain
except ModuleNotFoundError:
!pip install pybind11
!git clone -b ver4_release https://github.com/ryuz/BinaryBrain.git
%cd BinaryBrain
!python3 setup.py install --user
sys.exit() # please reboot runtime
```
developインストールの場合
```
try:
import binarybrain
except ModuleNotFoundError:
%cd /content
!rm -fr BinaryBrain
!pip install pybind11
!git clone -b ver4_release https://github.com/ryuz/BinaryBrain.git
%cd BinaryBrain
!python3 setup.py develop
sys.exit() # please reboot runtime
```
## GoogleDrive のマウント
```
import os
from google.colab import drive
drive.mount('/content/drive')
work_directory = '/content/drive/My Drive/BinaryBrain'
os.makedirs(work_directory, exist_ok=True)
os.chdir(work_directory)
```
## バージョン確認
```
# import
import binarybrain as bb
# バージョン表示
bb_version = bb.get_version_string()
print('BinaryBrain ver : %s'%bb_version)
# 利用可能なGPUの個数
device_count = bb.get_device_count()
print('GPU count : %d\n' % device_count)
# GPU情報の表示
for i in range(device_count):
print('[GPU<%d> Properties]'%i)
print(bb.get_device_properties_string(i))
```
## 簡単な学習確認
```
import os
import shutil
import numpy as np
from tqdm.notebook import tqdm
import torch
import torchvision
import torchvision.transforms as transforms
import binarybrain as bb
# configuration
data_path = './data/'
net_name = 'MnistDifferentiableLutSimple'
data_path = os.path.join('./data/', net_name)
rtl_sim_path = '../../verilog/mnist'
rtl_module_name = 'MnistLutSimple'
output_velilog_file = os.path.join(data_path, net_name + '.v')
sim_velilog_file = os.path.join(rtl_sim_path, rtl_module_name + '.v')
epochs = 4
mini_batch_size = 64
frame_modulation_size = 15
# dataset
dataset_path = './data/'
dataset_train = torchvision.datasets.MNIST(root=dataset_path, train=True, transform=transforms.ToTensor(), download=True)
dataset_test = torchvision.datasets.MNIST(root=dataset_path, train=False, transform=transforms.ToTensor(), download=True)
loader_train = torch.utils.data.DataLoader(dataset=dataset_train, batch_size=mini_batch_size, shuffle=True, num_workers=2)
loader_test = torch.utils.data.DataLoader(dataset=dataset_test, batch_size=mini_batch_size, shuffle=False, num_workers=2)
# define network
net = bb.Sequential([
bb.RealToBinary(frame_modulation_size=frame_modulation_size),
bb.DifferentiableLut([1024]),
bb.DifferentiableLut([420]),
bb.DifferentiableLut([70]),
bb.Reduce([10]),
bb.BinaryToReal(frame_integration_size=frame_modulation_size)
])
net.set_input_shape([1, 28, 28])
net.send_command("binary true")
loss = bb.LossSoftmaxCrossEntropy()
metrics = bb.MetricsCategoricalAccuracy()
optimizer = bb.OptimizerAdam()
optimizer.set_variables(net.get_parameters(), net.get_gradients())
# load
bb.load_networks(data_path, net)
# learning
for epoch in range(epochs):
# learning
loss.clear()
metrics.clear()
with tqdm(loader_train) as t:
for images, labels in t:
x_buf = bb.FrameBuffer.from_numpy(np.array(images).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.identity(10)[np.array(labels)].astype(np.float32))
y_buf = net.forward(x_buf, train=True)
dy_buf = loss.calculate(y_buf, t_buf)
metrics.calculate(y_buf, t_buf)
net.backward(dy_buf)
optimizer.update()
t.set_postfix(loss=loss.get(), acc=metrics.get())
# test
loss.clear()
metrics.clear()
for images, labels in loader_test:
x_buf = bb.FrameBuffer.from_numpy(np.array(images).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.identity(10)[np.array(labels)].astype(np.float32))
y_buf = net.forward(x_buf, train=False)
loss.calculate(y_buf, t_buf)
metrics.calculate(y_buf, t_buf)
print('epoch[%d] : loss=%f accuracy=%f' % (epoch, loss.get(), metrics.get()))
bb.save_networks(data_path, net, backups=3)
```
| github_jupyter |
```
import json
import re
import os
import requests
from googleapiclient.discovery import build
from google.oauth2 import service_account
from bs4 import BeautifulSoup
from IPython.display import HTML
SERVICE_ACCOUNT_FILE = 'keys.json'
SCOPES = ['https://www.googleapis.com/auth/spreadsheets.readonly']
creds = None
creds = service_account.Credentials.from_service_account_file(SERVICE_ACCOUNT_FILE, scopes=SCOPES)
SAMPLE_SPREADSHEET_ID = '1W...9k' # Not provided for privacy reasons
service = build('sheets', 'v4', credentials=creds)
sheet = service.spreadsheets().values()
class ScrapeData():
def read_from_link(self, link, replace_list=['\n','\t','\r','\u200d','\xa0']):
page = requests.get(link)
soup = BeautifulSoup(page.content, 'html.parser')
data = []
if (link.find('esamaad') >= 0):
find = soup.find_all('div', attrs={'style':'text-align: justify'})
if(not find):
find = soup.find_all('p', attrs={'style':'text-align: justify'})
if(not find):
find = soup.find_all('p')
elif(link.find('vikaspedia') >= 0):
find = soup.find_all('div', attrs={'id':'texttospeak'})
else:
find = soup.find_all('p')
for i in range(len(find)):
text = find[i].get_text()
text = re.sub("[a-zA-Z-|©&@:/^%`'$#•セ@!◌<>{}_–‘’“”=~;Ø(,+*)\[\]]*", "", text)
for j in range(len(replace_list)):
text = text.replace(replace_list[j],'')
text = re.sub("[.]+", ".", text)
text = re.sub(' +', ' ', text)
text = text.strip()
if len(text)>2:
data.append(text)
return data
def read_page(self, link=None):
if link:
text = self.read_from_link(link)
return(text)
def link_data(url):
data = []
if(url.lower().endswith('.pdf')):
return data
try:
data = scrape_tool.read_page(link=url)
data = list(set(data))
except:
data += []
return data
def fetchData(i, types, Ranges):
folder = types[i]
if not os.path.exists("jsonOutputs"):
os.mkdir("jsonOutputs")
if not os.path.exists(f"jsonOutputs/{folder}"):
os.mkdir(f"jsonOutputs/{folder}")
SAMPLE_RANGE_NAME = f'{folder.upper()}!{Ranges[i]}'
rows = sheet.get(spreadsheetId=SAMPLE_SPREADSHEET_ID, range=SAMPLE_RANGE_NAME, valueRenderOption='FORMULA').execute().get('values', [])
topic = ''
data = []
contents = []
for r in rows:
if(len(r)>0 and r[0]):
if(topic and contents):
title = ''.join(e for e in topic if e.isalnum())
with open(f'jsonOutputs/{folder}/{title}.json', 'w+', encoding='utf-8') as f:
json.dump(contents, f, ensure_ascii=False, indent=4)
topic = re.findall('"([^"]*)"',r[0])
contents = []
if(topic):
topic = topic[1]
if(not (topic and topic.strip())):
topic = r[0]
sub_topic = ''
if(len(r)>1 and r[1]):
sub_topic = re.findall('"([^"]*)"',r[1])
if(sub_topic):
sub_topic = sub_topic[1]
if(not (sub_topic and sub_topic.strip())):
sub_topic = r[1]
if(len(r)>2 and r[2]):
data = []
for c in r[2:]:
link = re.findall('"([^"]*)"',c)
if(link):
link = link[0]
if(link):
linkdata = link_data(link)
if(linkdata):
linkdata = {'url': link, 'data': linkdata}
data.append(linkdata.copy())
if(data):
out = {'topic': topic, 'sub_topic': sub_topic, 'data': data}
contents.append(out.copy())
scrape_tool = ScrapeData()
types = ['agriculture', 'finance', 'general']
Ranges = ['C3:M530', 'C3:M530', 'C3:O1120']
i = 0
# fetchData(i, types, Ranges)
i = 1
# fetchData(i, types, Ranges)
i = 2
# fetchData(i, types, Ranges)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/dvschultz/stylegan2-ada-pytorch/blob/main/SG2_ADA_PyTorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# StyleGAN2-ADA-PyTorch
**Notes**
* Training section should be fairly stable. I’ll slowly add features but it should work for most mainstream use cases
* Inference section is a work in progress. If you come across bug or have feature requests please post them in [Slack](https://ml-images.slack.com/archives/CLJGF384R) or on [Github](https://github.com/dvschultz/stylegan2-ada-pytorch/issues)
---
If you find this notebook useful, consider signing up for my [Patreon](https://www.patreon.com/bustbright) or [YouTube channel](https://www.youtube.com/channel/UCaZuPdmZ380SFUMKHVsv_AA/join). You can also send me a one-time payment on [Venmo](https://venmo.com/Derrick-Schultz).
## Setup
Let’s start by checking to see what GPU we’ve been assigned. Ideally we get a V100, but a P100 is fine too. Other GPUs may lead to issues.
```
!nvidia-smi -L
```
Next let’s connect our Google Drive account. This is optional but highly recommended.
```
from google.colab import drive
drive.mount('/content/drive')
```
## Install repo
The next cell will install the StyleGAN repository in Google Drive. If you have already installed it it will just move into that folder. If you don’t have Google Drive connected it will just install the necessary code in Colab.
```
import os
if os.path.isdir("/content/drive/MyDrive/colab-sg2-ada-pytorch"):
%cd "/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch"
elif os.path.isdir("/content/drive/"):
#install script
%cd "/content/drive/MyDrive/"
!mkdir colab-sg2-ada-pytorch
%cd colab-sg2-ada-pytorch
!git clone https://github.com/dvschultz/stylegan2-ada-pytorch
%cd stylegan2-ada-pytorch
!mkdir downloads
!mkdir datasets
!mkdir pretrained
!gdown --id 1-5xZkD8ajXw1DdopTkH_rAoCsD72LhKU -O /content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/pretrained/wikiart.pkl
else:
!git clone https://github.com/dvschultz/stylegan2-ada-pytorch
%cd stylegan2-ada-pytorch
!mkdir downloads
!mkdir datasets
!mkdir pretrained
%cd pretrained
!gdown --id 1-5xZkD8ajXw1DdopTkH_rAoCsD72LhKU
%cd ../
!pip install ninja opensimplex
```
You probably don’t need to run this, but this will update your repo to the latest and greatest.
```
%cd "/content/drive/My Drive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch"
!git config --global user.name "test"
!git config --global user.email "test@test.com"
!git fetch origin
!git pull
!git checkout origin/main -- train.py generate.py
```
## Dataset Preparation
Upload a .zip of square images to the `datasets` folder. Previously you had to convert your model to .tfrecords. That’s no longer needed :)
## Train model
Below are a series of variables you need to set to run the training. You probably won’t need to touch most of them.
* `dataset_path`: this is the path to your .zip file
* `resume_from`: if you’re starting a new dataset I recommend `'ffhq1024'` or `'./pretrained/wikiart.pkl'`
* `mirror_x` and `mirror_y`: Allow the dataset to use horizontal or vertical mirroring.
```
#required: definitely edit these!
dataset_path = './datasets/chin-morris.zip'
resume_from = './pretrained/wikiart.pkl'
aug_strength = 0.0
train_count = 0
mirror_x = True
#broken, don't use for now :(
#mirror_y = False
#optional: you might not need to edit these
gamma_value = 50.0
augs = 'bg'
config = '11gb-gpu'
snapshot_count = 4
!python train.py --gpus=1 --cfg=$config --metrics=None --outdir=./results --data=$dataset_path --snap=$snapshot_count --resume=$resume_from --augpipe=$augs --initstrength=$aug_strength --gamma=$gamma_value --mirror=$mirror_x --mirrory=False --nkimg=$train_count
```
### Resume Training
Once Colab has shutdown, you’ll need to resume your training. Reset the variables above, particularly the `resume_from` and `aug_strength` settings.
1. Point `resume_from` to the last .pkl you trained (you’ll find these in the `results` folder)
2. Update `aug_strength` to match the augment value of the last pkl file. Often you’ll see this in the console, but you may need to look at the `log.txt`. Updating this makes sure training stays as stable as possible.
3. You may want to update `train_count` to keep track of your training progress.
Once all of this has been reset, run that variable cell and the training command cell after it.
## Convert Legacy Model
If you have an older version of a model (Tensorflow based StyleGAN, or Runway downloaded .pkl file) you’ll need to convert to the newest version. If you’ve trained in this notebook you do **not** need to use this cell.
`--source`: path to model that you want to convert
`--dest`: path and file name to convert to.
```
!python legacy.py --source=/content/drive/MyDrive/runway.pkl --dest=/content/drive/MyDrive/colab-sg2-ada-pytorch/stylegan2-ada-pytorch/runway.pkl
```
## Testing/Inference
Also known as "Inference", "Evaluation" or "Testing" the model. This is the process of usinng your trained model to generate new material, usually images or videos.
### Generate Single Images
`--network`: Make sure the `--network` argument points to your .pkl file. (My preferred method is to right click on the file in the Files pane to your left and choose `Copy Path`, then paste that into the argument after the `=` sign).
`--seeds`: This allows you to choose random seeds from the model. Remember that our input to StyleGAN is a 512-dimensional array. These seeds will generate those 512 values. Each seed will generate a different, random array. The same seed value will also always generate the same random array, so we can later use it for other purposes like interpolation.
`--truncation`: Truncation, well, truncates the latent space. This can have a subtle or dramatic affect on your images depending on the value you use. The smaller the number the more realistic your images should appear, but this will also affect diversity. Most people choose between 0.5 and 1.0, but technically it's infinite.
```
!python generate.py --outdir=/content/out/images/ --trunc=1.0 --seeds=0-49 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
### Truncation Traversal
Below you can take one seed and look at the changes to it across any truncation amount. -1 to 1 will be pretty realistic images, but the further out you get the weirder it gets.
#### Options
`--network`: Again, this should be the path to your .pkl file.
`--seeds`: Pass this only one seed. Pick a favorite from your generated images.
`--start`: Starting truncation value.
`--stop`: Stopping truncation value. This should be larger than the start value. (Will probably break if its not).
`--increment`: How much each frame should increment the truncation value. Make this really small if you want a long, slow interpolation. (stop-start/increment=total frames)
```
!python generate.py --process="truncation" --outdir=out/trunc-trav/ --start=-2.0 --stop=2.0 --increment=0.01 --seeds=85 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
### Interpolations
Interpolation is the process of generating very small changes to a vector in order to make it appear animated from frame to frame.
We’ll look at different examples of interpolation below.
#### Options
`--network`: path to your .pkl file
`--interpolation`: Walk type defines the type of interpolation you want. In some cases it can also specify whether you want the z space or the w space.
`--frames`: How many frames you want to produce. Use this to manage the length of your video.
`--trunc`: truncation value
#### Linear Interpolation
```
!python generate.py --outdir=out/video1/ --trunc=1 --process="interpolation" --seeds=85,265,297,849 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
!python generate.py --outdir=out/video1-w/ --space="w" --trunc=1 --process="interpolation" --seeds=85,265,297,849 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
#### Slerp Interpolation
This gets a little heady, but technically linear interpolations are not the best in high-dimensional GANs. [This github link](https://github.com/soumith/dcgan.torch/issues/14) is one of the more popular explanations ad discussions.
In reality I do not find a huge difference between linear and spherical interpolations (the difference in z- and w-space is enough in many cases), but I’ve implemented slerp here for anyone interested.
Note: Slerp in w space currently isn’t supported. I’m working on it.
```
!python generate.py --outdir=out/video1/ --trunc=1 --process="interpolation" --interpolation="slerp" --seeds=85,265,297,849 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
#### Noise Loop
If you want to just make a random but fun interpolation of your model the noise loop is the way to go. It creates a random path thru the z space to show you a diverse set of images.
`--interpolation="noiseloop"`: set this to use the noise loop funtion
`--diameter`: This controls how "wide" the loop is. Make it smaller to show a less diverse range of samples. Make it larger to cover a lot of samples. This plus `--frames` can help determine how fast the video feels.
`--random_seed`: this allows you to change your starting place in the z space. Note: this value has nothing to do with the seeds you use to generate images. It just allows you to randomize your start point (and if you want to return to it you can use the same seed multiple times).
Noise loops currently only work in z space.
```
!python generate.py --outdir=out/video-noiseloop-0.9d/ --trunc=0.8 --process="interpolation" --interpolation="noiseloop" --diameter=0.9 --random_seed=100 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
#### Circular Loop
The noise loop is, well, noisy. This circular loop will feel much more even, while still providing a random loop.
I recommend using a higher `--diameter` value than you do with noise loops. Something between `50.0` and `500.0` alongside `--frames` can help control speed and diversity.
```
!python generate.py --outdir=out/video-circularloop/ --trunc=1 --process="interpolation" --interpolation="circularloop" --diameter=800.00 --frames=720 --random_seed=90 --network=/content/stylegan2-ada-pytorch/pretrained/wikiart.pkl
```
## Feature Extraction using Closed Form Factorization
TK!
```
!python closed_form_factorization.py --out /content/factor.pt --ckpt /content/network.pkl
!python apply_factor.py -i 0 -d 10 -s 5 --ckpt /content/network.pkl /content/factor.pt #--video --out_prefix '6k-factor'
for i in range(512):
!python apply_factor.py -i {i} -d 10 -s 5 --ckpt /content/network.pkl /content/factor.pt #--video --out_prefix '6k-factor'
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/naip_imagery.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/naip_imagery.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Datasets/naip_imagery.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/naip_imagery.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The magic command `%%capture` can be used to hide output from a specific cell. Uncomment these lines if you are running this notebook for the first time.
```
# %%capture
# !pip install earthengine-api
# !pip install geehydro
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once. Uncomment the line `ee.Authenticate()`
if you are running this notebook for the first time or if you are getting an authentication error.
```
# ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
image = ee.Image('USDA/NAIP/DOQQ/m_4609915_sw_14_1_20100629')
Map.addLayer(image, {'bands': ['N', 'R', 'G']}, 'NAIP')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
```
import open3d as o3d
import numpy as np
import sys
# monkey patches visualization and provides helpers to load geometries
sys.path.append('..')
import open3d_tutorial as o3dtut
# change to True if you want to interact with the visualization windows
o3dtut.interactive = False
```
# Mesh deformation
If we want to deform a triangle mesh according a small number of constraints, we can use mesh deformation algorithms. Open3D implements the as-rigid-as-possible method by [\[SorkineAndAlexa2007\]](../reference.html#SorkineAndAlexa2007) that optimizes the following energy functional
\begin{equation}
\sum_i \sum_{j \in \mathcal{N}(i)} w_{ij} || (\mathbf{p}'_i - \mathbf{p}'_j) - \mathbf{R}_i (\mathbf{p}_i - \mathbf{p}_j)||^2 \,,
\end{equation}
where $\mathbf{R}_i$ are rotation matrices that we want to optimize for, and $\mathbf{p}'_i$ and $\mathbf{p}_i$ are the vertex positions after and before the optimization, respectively. $\mathcal{N}(i)$ is the set of neighbours of vertex $i$. The weights $w_{ij}$ are cot weights.
Open3D implementes this method in `deform_as_rigid_as_possible`. The first argument to this method is a set of `constraint_ids` that refer to the vertices in the triangle mesh. The second argument `constrint_pos` defines at which position those vertices should be after the optimization. The optimization process is an iterative scheme. Hence, we also can define the number of iterations via `max_iter`.
```
mesh = o3dtut.get_armadillo_mesh()
vertices = np.asarray(mesh.vertices)
static_ids = [idx for idx in np.where(vertices[:, 1] < -30)[0]]
static_pos = []
for id in static_ids:
static_pos.append(vertices[id])
handle_ids = [2490]
handle_pos = [vertices[2490] + np.array((-40, -40, -40))]
constraint_ids = o3d.utility.IntVector(static_ids + handle_ids)
constraint_pos = o3d.utility.Vector3dVector(static_pos + handle_pos)
with o3d.utility.VerbosityContextManager(o3d.utility.VerbosityLevel.Debug) as cm:
mesh_prime = mesh.deform_as_rigid_as_possible(
constraint_ids, constraint_pos, max_iter=50)
print('Original Mesh')
R = mesh.get_rotation_matrix_from_xyz((0,np.pi,0))
o3d.visualization.draw_geometries([mesh.rotate(R)])
print('Deformed Mesh')
mesh_prime.compute_vertex_normals()
o3d.visualization.draw_geometries([mesh_prime.rotate(R)])
```
## Smoothed ARAP
Open3D also implements a smoothed version of the ARAP objective defined as
\begin{equation}
\sum_i \sum_{j \in \mathcal{N}(i)} w_{ij} || (\mathbf{p}'_i - \mathbf{p}'_j) - \mathbf{R}_i (\mathbf{p}_i - \mathbf{p}_j)||^2 + \alpha A ||\mathbf{R}_i - \mathbf{R}_j||^2\,,
\end{equation}
that penalizes a deviation of neighbouring rotation matrices. $\alpha$ is a trade-off parameter for the regularization term and $A$ is the surface area.
The smoothed objective can be used in `deform_as_rigid_as_possible` by using the argument `energy` with the parameter `Smoothed`.
| github_jupyter |
##### Copyright 2018 Verily Life Sciences LLC.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Default title text
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
This notebook demonstrates how one can dive deeper into QC results to explain some unexpected patterns. In this notebook, we will see that a few samples in the [Platinum Genomes](https://cloud.google.com/genomics/docs/public-datasets/illumina-platinum-genomes) have a very low number of private variants, and we will figure out why.
> Eberle, MA et al. (2017) **A reference data set of 5.4 million phased human variants validated by genetic inheritance from sequencing a three-generation 17-member pedigree.** Genome Research 27: 157-164. [doi:10.1101/gr.210500.116](http://dx.doi.org/10.1101/gr.210500.116)
# Setup
Check out the code for the various QC methods to the current working directory. Further down in the notebook we will read the SQL templates from this clone.
```
!git clone https://github.com/verilylifesciences/variant-qc.git
```
Install additional Python dependencies [plotnine](https://plotnine.readthedocs.io/en/stable/) for plotting and [jinja2](http://jinja.pocoo.org/docs/2.10/) for performing text replacements in the SQL templates.
```
!pip install --upgrade plotnine jinja2
import jinja2
import numpy as np
import os
import pandas as pd
import plotnine
from plotnine import *
plotnine.options.figure_size = (10, 6)
# Change this to be your project id.
PROJECT_ID = 'your-project-id' #@param
from google.colab import auth
auth.authenticate_user()
print('Authenticated')
def run_query(sql_template, replacements={}):
if os.path.isfile(sql_template):
sql_template = open(sql_template, "r").read()
sql = jinja2.Template(sql_template).render(replacements)
print('SQL to be executed:\n', sql)
df = pd.io.gbq.read_gbq(sql, project_id=PROJECT_ID, dialect='standard')
print('\nResult shape:\t', df.shape)
return df
```
# Get a count of private variants
## Compute the private variant counts BigQuery
Running this query is optional as this has already been done and saved to Cloud Storage. See the next section for how to retrieve these results from Cloud Storage.
```
# Read the SQL template from the cloned repository in the home directory, perform
# the variable replacements and execute the query.
df = run_query(
sql_template='variant-qc/sql/private_variants.sql',
replacements={
'GENOME_CALL_OR_MULTISAMPLE_VARIANT_TABLE': 'bigquery-public-data.human_genome_variants.platinum_genomes_deepvariant_variants_20180823',
'HIGH_QUALITY_CALLS_FILTER': 'NOT EXISTS (SELECT ft FROM UNNEST(c.FILTER) ft WHERE ft NOT IN ("PASS", "."))'
}
)
```
## Retrieve the private variant counts from Cloud Storage
We can read these values from the CSV created via [Sample-Level-QC.Rmd](https://github.com/verilylifesciences/variant-qc/blob/master/R/Sample-Level-QC.Rmd).
```
df = pd.read_csv("https://storage.googleapis.com/genomics-public-data/platinum-genomes/reports/DeepVariant_Platinum_Genomes_sample_results.csv")[["name", "private_variant_count"]]
df.shape
```
## Examine results and outliers
This small cohort does not contain enough samples to estimate the expected number of private variants. It is used here for demonstration purposes only.
```
df
```
Let's take a look at the samples who are more than one standard deviation away from the mean.
```
df.loc[abs(df.private_variant_count - df.private_variant_count.mean()) > df.private_variant_count.std(), :]
```
Next let's see if the sample metadata can be used to help explain the explain the low number of private variants that we see.
# Retrieve sample metadata
The platinum genomes samples are also members of the larger 1000 genomes dataset. We can retrieve the metadata for those samples from the 1000 genomes metadata.
```
metadata_df = run_query(
sql_template="""
SELECT
Sample AS name,
Gender AS sex,
Super_Population AS ancestry,
Relationship AS relationship
FROM
`bigquery-public-data.human_genome_variants.1000_genomes_sample_info`
WHERE
Sample IN ('NA12877', 'NA12878', 'NA12889', 'NA12890', 'NA12891', 'NA12892')
"""
)
```
# Visualize results by ancestry
```
joined_results = pd.merge(df, metadata_df, how='left', on='name')
joined_results.shape
assert(joined_results.shape == (6, 5))
p = (ggplot(joined_results) +
geom_boxplot(aes(x = 'ancestry', y = 'private_variant_count', fill = 'ancestry')) +
theme_minimal()
)
p
```
All individuals in this dataset are of the same ancestry, so that does not explain the pattern we see.
# Visualize results by relationship
We know from [the paper](https://genome.cshlp.org/content/27/1/157) that all members of this cohort are from the same family.
```
run_query("""
SELECT
*
FROM
`bigquery-public-data.human_genome_variants.1000_genomes_pedigree`
WHERE
Individual_ID IN ('NA12877', 'NA12878', 'NA12889', 'NA12890', 'NA12891', 'NA12892')
""")
p = (ggplot(joined_results) +
geom_text(aes(x = 'name', y = 'private_variant_count', label = 'relationship')) +
theme_minimal()
)
p
```
And we can see that the relationship between individuals explains the pattern we see in the private variant counts.
| github_jupyter |
# Sesion 3 :
## - Funciones Integradas y ayuda
## - Bibliotecas escritas en Python
#####################################
## Funciones
Las funciones son bloques de código que se ejecutan cuando se llaman y realizan una tarea específica. Las funciones son beneficiosas porque podemos reutilizar el código para tareas repetitivas.
Aprendimos sobre varias funciones integradas de Python en la Sesión 2:
* print(): La función de print "imprimirá" (o mostrará) datos al usuario
* len(): Encuentra la longitud de los caracteres de un string
* type(): Verificar el tipo de datos del valor que se asigna a una variable.
- Cadena de caracteres, llamado string, (str).
- Entero o intergers (int).
- Número de punto flotante (float).
* str(): En la Sesión 2, practicamos la concatenación cuando una variable se le asigna un valor entero o flotante. Como no podemos concatenar un entero o un flotante con una cadena, demostramos cómo covertimos el numero a una string, usando la función str ().
```
#Ejemplo función str() si se hace con una concatenacion
texto1 = "Mi gato tiene"
edad = 2
texto2 = "años."
frase = texto1 + " " + edad + " " + texto2
print(frase)
```
### Argumento
Un argumento son valores pasados a una función. Una función puede tener cero o más argumentos.
* print() toma cero o más argumentos y print() sin argumentos imprime una línea en blanco.
* len() toma exactamente un argumento.
* int, str, y float crean un nuevo valor a partir de uno existente.
```
# Ejemplo argumento, imprimir textos y numeros dentro de la misma linea de codigo
edad = 2
print("Mi gato tiene", edad, "años.")
# Siempre se debe usar paréntesis, incluso si están vacíos, para que Python sepa que se está llamando a una función
print('before')
print()
print('after')
```
### Las funciones integradas de uso común incluyen max, min y round.
* Usa max para encontrar el valor más grande de uno o más conjuntos de valores.
* Usa min para encontrar el valor minimo de un conjunto de valores.
* Ambos funcionan tanto en cadenas de caracteres como en números.
* Los valores “Más grande” y “más pequeño” usan (0-9, A-Z, a-z) para comparar letras,
```
# El valor máximo corresponde al último valor en orden numerico o alfabético, sin importar la longitud de la cadena.
print(max(1, 2, 3))
print(max("David", "Alicia", "Maya", "Emilio"))
# el valor mínimo corresponde al primer valor en orden alfabético, sin importar la longitud de la cadena.
print(min('a', 'A'))
print(min('a', 'A', '0'))
# Las vocales acentuadas, la letra ñ o ç se consideran posteriores al resto de vocales y consonantes
print(min("Ángeles", "Daniel"))
# Ejercicios max(), min()
print(max(4, 5, -2, 8, 3.5, -10))
print(min("David", "Alicia", "Oscar", "Emilio"))
# Las funciones pueden ejecutar solo para combinaciones de ciertos argumentos.
# max() y min() deben recibir al menos un argumento y deben recibir cosas que puedan compararse con sentido.
print(max(1, 'a', -4))
```
### La función integrada round()
* La función integrada round() redondeará un número de punto flotante.
* round() admite uno o dos argumentos numéricos.
* Por defecto, redondea a cero cifras decimales.
```
# Si sólo hay un argumento, la función devuelve el argumento redondeado al entero más próximo
print(round(-4.35))
print(round(3.712))
# Si se escriben dos argumentos, siendo el segundo un número entero y positivo, la función integrada round()
# devuelve el primer argumento redondeado en la posición indicada por el segundo argumento
# print(round(4.3527)
print(round(4.3527, 2))
print(round(4.3527, 1))
print(round(4.3527, 3))
# Si el segundo argumento es negativo, se redondea a decenas, centenas, etc.
print(round(43527, -1))
print(round(43527, -2))
print(round(43527, -3))
print(round(43527, -4))
# Ejercicio devuelve el primer argumento redondeado (tau = 6.28318) en la sgunda posición indicada por el segundo argumento.
```
### Usa la función integrada help()
* Ayuda a obtener informacion sobre una función.
```
help(round)
# consulta la informaion de la funcion max
```
### Errores frecuentes
* Python reporta un error de sintaxis cuando no puede entender la fuente de un programa.
```
# Olvidar cerrar las comillas alrededor de la cadena de caracteres.
name = 'Feng
# Un '=' adicional en la asignación.
age = = 52
# falta cerrar un parentesis
print("hola mundo"
# Python reporta un error de tiempo de ejecución cuando algo anda mal mientras un programa se está ejecutando.
age = 53
remaining = 100 - aege # mal escrito 'age'
```
# Cada función devuelve algo.
```
# Cada que llamamos a una función se produce algún resultado.
# Si la función no tiene un resultado útil que devolver, usualmente devuelve el valor especial None.
result = print('ejemplo')
print('el resultado de print es', result)
# Ejercicio
# Explica en términos simples el orden de operaciones en el siguiente programa:
# cuándo ocurre la adición y cuándo la sustracción?, cuándo es llamada cada función?, etc.
radiance = 1.0
radiance = max(2.1, 2.0 + min(radiance, 1.1 * radiance - 0.5))
print(radiance)
# 1. 1.1 * radiance = 1.1
# 2. 1.1 - 0.5 = 0.6
# 3. min(radiance, 0.6) = 0.6
# 4. 2.0 + 0.6 = 2.6
# 5. max(2.1, 2.6) = 2.6
# Al final, radiance = 2.6
```
## Bibliotecas Python
Al igual que el significado de la palabra biblioteca, una biblioteca de Python es una colección de funciones y paquetes que se pueden importar para ejecutar.
Solo imagina que necesitas un fragmento especifico de un libro para resolver un problema. Importar una función desde una biblioteca de Python es así, ¡excepto que no necesita perder tiempo buscando el libro y hojeando las páginas!
### Gran parte del poder de un lenguaje de programación está en sus bibliotecas.
* Una biblioteca es una colección de archivos (llamados módulos) que contienen funciones para ser usadas por otros programas. Pueden también contener datos (e.g. constantes numéricas) y otras cosas.
* Los contenidos de una biblioteca deberían estar relacionados, pero no hay una forma establecida para imponer esto.
* La biblioteca estándar de Python es una colección extensa de módulos que vienen con Python mismo.
* Muchas bibliotecas adicionales están disponibles en PyPI (el índice de paquetes de Python).
* Más adelante veremos cómo instalar e importar nuevas bibliotecas.
```
# Como saber que bibliotecas tengo instalado en mi entorno
!pip list
# Como ver las funciones que tenemos incorporadas en python
__builtin__.__dict__
```
### Instalacion
¿Qué es la instalación de pip?
* pip es un administrador de paquetes (también un paquete en sí mismo), que le ayuda a instalar un paquete de Python desde PyPI (índice de paquetes de Python). Es uno de los paquetes predeterminados que se incluyen al instalar Python.
* `pip install <nombre de la biblioteca>`
* `pip` usó una variable para llamar a la biblioteca pip
* `install` un comando de la biblioteca pip
Notas: "!" debe agregarse al principio para instalar la biblioteca en jupyter y derivados.
Documentación: https://pip.pypa.io/en/stable/cli/pip_install/
```
# instalar la biblioeca pandas !pip install pandas
# instalar la biblioteca statsmodels !pip install statsmodels
# desinstalar la biblioteca !pip uninstall -y statsmodels
# instalar la biblioteca statsmodels !pip install statsmodels
!pip list
```
### Dependencias entre bibliotecas de Python
* El propósito de una biblioteca de Python es simplificar el proceso, especialmente los repetitivos. La mayoría de las bibliotecas se construyen sobre bibliotecas básicas.
* Algunas bibliotecas incluso se basan en una versión específica de la biblioteca principal.
* Por lo tanto, a veces, cuando instalas una biblioteca, es posible que debas instalar primero sus dependencias y prestar un poco más de atención a la versión de la biblioteca.
### Un programa debe importar un módulo de biblioteca antes de usarlo.
Importar una biblioteca es un proceso muy sencillo
* Usa import para cargar un módulo de una biblioteca en la memoria del programa.
* Después refiérete a las cosas del módulo como nombre_del_modulo.nombre_de_la_cosa.
* Python usa . para referirse a “parte de”.
* Usando math, uno de los módulos de la biblioteca estándar:
```
# importa la biblioteca math
import math
numero = 25
raiz_cuadrada = math.sqrt(numero)
print("La raíz cuadrada de", numero, "es", raiz_cuadrada)
print('pi es', math.pi)
print('cos(pi) es', math.cos(math.pi))
```
[Biblioteca Math](https://docs.python.org/es/3.10/library/math.html)
## Crea un alias para el módulo de una biblioteca al importarlo para acortar programas.
* Usa import ... as ... para asignarle a una biblioteca un alias más corto al importarla.
* Después refiérete a los elementos de la biblioteca usando el nombre corto.
```
import math as m
print('cos(pi) es', m.cos(m.pi))
# importa la biblioteca numpy y nombralo as 'np' - import numpy as np
# importa la biblioteca pandas y nombralo as 'pd'
# Se puede importar funciones especificas dentro la biblioteca
from math import sqrt
#Ejemplo
numero = 25
raiz_cuadrada = sqrt(numero)
print("La raíz cuadrada de ", numero, "es", raiz_cuadrada)
```
* Esto se suele hacer con las librerías que se usan frecuentemente o que tienen nombres largos.
* Por ejemplo, para la biblioteca de graficación matplotlib se suele usar el alias mpl.
* Pero esto puede hacer que los programas sean más difíciles de entender, porque los lectores deberán aprender los alias de tu programa.
### Biblioteca de python - verificar versión
* Las bibliotecas de Python a menudo lanzan nuevas versiones para mejorar la finalización de la biblioteca agregando nuevas funciones o revisando ciertas funciones.
* Normalmente instalamos la última versión de una función. A veces, ciertas funciones se eliminan en la versión más reciente y pueden afectar nuestro codigo.
* Además, algunas bibliotecas de código abierto requieren una versión específica de una biblioteca para garantizar que se pueda utilizar una función específica
```
# check numpy version - np.__version__
```
### Biblioteca python - función de ayuda
* Nadie conocerá todas las funciones, todas las bibliotecas que existen y recordará lo que hacen.
* La función help () es la forma más rápida de encontrar todas las funciones en una biblioteca de Python además de buscar en Internet.
```
help(math)
```
## Lecturas adicionales para esta sesión
[How to Install a Package in Python using PIP](https://datatofish.com/install-package-python-using-pip/)
[Check the version of Python package / library](https://note.nkmk.me/en/python-package-version/)
[Python Modules](https://docs.python.org/3/tutorial/modules.html)
## Puntos clave
### Funciones Integradas
* Una función puede tomar cero o más argumentos.
* Las funciones incorporadas de uso común incluyen max, min y round.
* Las funciones solo pueden funcionar para ciertos (combinaciones de) argumentos.
* Las funciones pueden tener valores predeterminados para algunos argumentos.
* Usa la función incorporada help para obtener ayuda para una función.
* Cada función regresa algo.
* Soluciona errores de sintaxis leyendo el código fuente y errores de tiempo de ejecución rastreando la ejecución del programa
### Bibliotecas
* Gran parte del poder que puede tener un lenguaje de programación está en sus bibliotecas.
* Un programa debe importar los módulos de una biblioteca para poder usarlos.
* Usa help para aprender sobre los contenidos de un módulo de la biblioteca.
* Importa elementos específicos de una biblioteca para acortar programas.
* Crea un alias para una biblioteca al importarla para acortar programas
| github_jupyter |
```
#!pip install matplotlib
import pandas as pd
import matplotlib.pyplot as plt
from pyspark.sql import SparkSession
import pyspark.sql.functions as f
spark = SparkSession.builder.master("local[*]").getOrCreate()
```
### Load Data
```
raw_train_df = spark.read.option('header', 'true').csv('../data/raw/train.csv')\
.select('id', 'vendor_id', 'pickup_datetime', 'dropoff_datetime', 'passenger_count', 'pickup_longitude', 'pickup_latitude', 'dropoff_longitude', 'dropoff_latitude', 'trip_duration')\
.withColumn('vendor_id', f.col('vendor_id').cast('int'))\
.withColumn('pickup_datetime', f.to_timestamp(f.col('pickup_datetime'), 'YYYY-MM-dd HH:mm:ss'))\
.withColumn('dropoff_datetime', f.to_timestamp(f.col('dropoff_datetime'), 'YYYY-MM-dd HH:mm:ss'))\
.withColumn('passenger_count', f.col('passenger_count').cast('int'))\
.withColumnRenamed('pickup_longitude', 'pickup_lon')\
.withColumn('pickup_lon', f.col('pickup_lon').cast('double'))\
.withColumnRenamed('pickup_latitude', 'pickup_lat')\
.withColumn('pickup_lat', f.col('pickup_lat').cast('double'))\
.withColumnRenamed('dropoff_longitude', 'dropoff_lon')\
.withColumn('dropoff_lon', f.col('dropoff_lon').cast('double'))\
.withColumnRenamed('dropoff_latitude', 'dropoff_lat')\
.withColumn('dropoff_lat', f.col('dropoff_lat').cast('double'))\
.withColumn('trip_duration', f.col('trip_duration').cast('int'))\
raw_train_df.printSchema()
raw_train_df.show(2)
```
### Trips by date
```
raw_train_df.select(f.date_format('pickup_datetime','yyyy-MM').alias('month')).groupby('month').count().show()
```
### Trips by hour
```
trips_by_hour_df_pd = raw_train_df\
.withColumn('hour', f.hour(f.col('pickup_datetime')))\
.groupby('hour').count()\
.sort(f.asc('hour'))\
.toPandas().set_index('hour')
trips_by_hour_df_pd.plot()
```
## Distribution by trip duration
```
clean_train_df_pd = raw_train_df.groupby('trip_duration').count().sort(f.asc('trip_duration')).toPandas().set_index('trip_duration')
clean_train_df_pd[(clean_train_df_pd.index > 0) & (clean_train_df_pd.index < 6000)].plot()
clean_train_df_pd[(clean_train_df_pd.index > 0) & (clean_train_df_pd.index < 900)].plot()
len(clean_train_df_pd) , len(clean_train_df_pd[(clean_train_df_pd.index > 90) & (clean_train_df_pd.index < 6000)])
raw_train_df.where(f.col('trip_duration') > 90).where(f.col('trip_duration') < 7200).count()
raw_train_df.where(f.col('trip_duration') > 90).where(f.col('trip_duration') < 3600).count()
```
### Mean trip duration by vendor_id
```
raw_train_df.groupby('vendor_id').agg(f.mean('trip_duration')).show()
```
### Clean data
```
clean_train_df = raw_train_df\
.dropna(subset=['id', 'vendor_id', 'pickup_datetime', 'dropoff_datetime', 'passenger_count', 'pickup_lon', 'pickup_lat', 'dropoff_lon', 'dropoff_lat', 'trip_duration'])\
.where(f.col('passenger_count') > 0)\
.where(f.col('trip_duration') > 0)\
.where(f.col('trip_duration') > 90).where(f.col('trip_duration') < 7200)
clean_train_df.count()
```
### Feature engineering
```
import s2sphere
from pyspark.sql.types import StringType
def cell_id(level: int, lat: int, lng: int) -> str:
return s2sphere.CellId.from_lat_lng(s2sphere.LatLng.from_degrees(lat, lng)).parent(level).to_token()
cell_id_udf = f.udf(cell_id, StringType())
clean_train_df\
.withColumn('cell_l12', cell_id_udf(f.lit(12), f.col('pickup_lon'), f.col('pickup_lat')))\
.withColumn('cell_l18', cell_id_udf(f.lit(18), f.col('pickup_lon'), f.col('pickup_lat')))\
.drop('vendor_id')\
.show(5)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/Terrain/us_ned_landforms.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/us_ned_landforms.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/Terrain/us_ned_landforms.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
dataset = ee.Image('CSP/ERGo/1_0/US/landforms')
landforms = dataset.select('constant')
landformsVis = {
'min': 11.0,
'max': 42.0,
'palette': [
'141414', '383838', '808080', 'EBEB8F', 'F7D311', 'AA0000', 'D89382',
'DDC9C9', 'DCCDCE', '1C6330', '68AA63', 'B5C98E', 'E1F0E5', 'a975ba',
'6f198c'
],
}
Map.setCenter(-105.58, 40.5498, 11)
Map.addLayer(landforms, landformsVis, 'NED Landforms')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sqlalchemy import create_engine
import re
engine = create_engine("postgresql://postgres:postgres@localhost:5432/hr_database")
df = pd.read_csv('human_resources_dataset.csv', parse_dates=['START_DT', 'END_DT'])
df.columns = ['employee_id',
'employee_name',
'employee_email',
'hire_date',
'job',
'salary',
'department',
'manager',
'start_date',
'end_date',
'location',
'address',
'city',
'state',
'education']
df['salary'] = df['salary'].str.replace(',', '').astype('int')
df['start_date'] = df['start_date'].dt.date
df['end_date'] = df['end_date'].dt.date
df.head()
df.info()
for col in df.columns:
print(col, df[col].nunique())
df_dict = {}
for col in ['job', 'salary', 'department', 'location', 'education']:
table_df = df[[col]]\
.drop_duplicates()\
.sort_values(col)\
.reset_index()\
.drop('index', axis=1)\
.reset_index()
table_df.columns = [col+"_id", col]
df_dict[col] = table_df
table_df.to_sql(col, engine, index=False)
employee = df[['employee_id', 'employee_name', 'employee_email']]\
.drop_duplicates()\
.sort_values('employee_id')\
.reset_index()\
.drop('index', axis=1)\
.reset_index()
employee.columns = ['id', 'employee_id', 'employee_name', 'employee_email']
df_dict['employee'] = employee
employee.to_sql('employee', engine, index=False)
employee.head()
city = df[['city', 'state']]\
.drop_duplicates()\
.sort_values(['city', 'state'])\
.reset_index()\
.drop('index', axis=1)\
.reset_index()
city.columns = ['city_id', 'city', 'state']
df_dict['city'] = city
city.to_sql('city', engine, index=False)
city
address = df[['address', 'city', 'state', 'location']]\
.drop_duplicates()\
.sort_values(['state', 'city', 'address'])\
.reset_index()\
.drop('index', axis=1)\
.reset_index()
address.columns = ['address_id', 'address', 'city', 'state', 'location']
address = address.merge(df_dict['city'], on=['city', 'state']).drop(columns=['city', 'state'])
address = address.merge(df_dict['location'], on='location').drop('location', axis=1)
df_dict['address'] = address
address.to_sql('address', engine, index=False)
address
merge_df = df.drop(['employee_name', 'employee_email', 'hire_date', 'location', 'city', 'state'], axis=1)
for col in ['job', 'salary', 'department', 'education', 'address']:
merge_df = merge_df.merge(df_dict[col], on=col).drop(col, axis=1)
merge_df.head()
merge_df = merge_df.merge(df_dict['employee'],
left_on='manager',
right_on='employee_name',
how='left',
suffixes=[None,'_manager'])
merge_df = merge_df.drop(['manager','city_id', 'location_id', 'employee_name','employee_email','employee_id_manager'], axis=1)\
.rename(columns={'id':'manager_id'})
merge_df['manager_id'] = merge_df['manager_id'].fillna(18).astype(int)
merge_df.head()
merge_df = merge_df.merge(df_dict['employee'],
on='employee_id',
how='left',
suffixes=[None,'_employee'])
employee_history = merge_df.drop(['employee_id', 'employee_name', 'employee_email'], axis=1)
employee_history.head()
df_dict['employee_history'] = employee_history
employee_history = employee_history[['id',
'manager_id',
'education_id',
'job_id',
'department_id',
'address_id',
'salary_id',
'start_date',
'end_date']]
df_dict['employee_history'] = employee_history
employee_history.to_sql('employee_history', engine, index=False)
employee_history
```
| github_jupyter |
# Passive Aggressive Regressor with Robust Scaler
This Code template is for the regression analysis using a simple PassiveAggresiveRegressor based on the passive-aggressive algorithms and the feature rescaling technique used is Robust Scaler in a pipeline. Passive-aggressive algorithms are a group of algorithms for large-scale learning.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.linear_model import PassiveAggressiveRegressor
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=['']
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Model
The passive-aggressive algorithms are a family of algorithms for large-scale learning. They are similar to the Perceptron in that they do not require a learning rate. However, contrary to the Perceptron, they include a regularization parameter C
> **C** ->Maximum step size (regularization). Defaults to 1.0.
> **max_iter** ->The maximum number of passes over the training data (aka epochs). It only impacts the behavior in the fit method, and not the partial_fit method.
> **tol**->The stopping criterion. If it is not None, the iterations will stop when (loss > previous_loss - tol).
> **early_stopping**->Whether to use early stopping to terminate training when validation. score is not improving. If set to True, it will automatically set aside a fraction of training data as validation and terminate training when validation score is not improving by at least tol for n_iter_no_change consecutive epochs.
> **validation_fraction**->The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if early_stopping is True.
> **n_iter_no_change**->Number of iterations with no improvement to wait before early stopping.
> **shuffle**->Whether or not the training data should be shuffled after each epoch.
> **loss**->The loss function to be used: epsilon_insensitive: equivalent to PA-I in the reference paper. squared_epsilon_insensitive: equivalent to PA-II in the reference paper.
> **epsilon**->If the difference between the current prediction and the correct label is below this threshold, the model is not updated.
###Data Scaling
Used sklearn.preprocessing.RobustScaler
This Scaler removes the median and scales the data according to the quantile range (defaults to IQR: Interquartile Range). The IQR is the range between the 1st quartile (25th quantile) and the 3rd quartile (75th quantile).
Read more at [scikit-learn.org](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html)
```
Input=[("standard",RobustScaler()),("model",PassiveAggressiveRegressor(random_state=123))]
model = Pipeline(Input)
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),y_pred[0:20], color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Arpit Somani , Github: [Profile](https://github.com/arpitsomani8)
| github_jupyter |
## Work
1. 請自行定義一個 loss function, 為 0.3 * focal loss + 0.7 cross-entropy,訓練並比較結果
```
import os
from tensorflow import keras
# Disable GPU
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
train, test = keras.datasets.cifar10.load_data()
## 資料前處理
def preproc_x(x, flatten=True):
x = x / 255.
if flatten:
x = x.reshape((len(x), -1))
return x
def preproc_y(y, num_classes=10):
if y.shape[-1] == 1:
y = keras.utils.to_categorical(y, num_classes)
return y
x_train, y_train = train
x_test, y_test = test
# Preproc the inputs
x_train = preproc_x(x_train)
x_test = preproc_x(x_test)
# Preprc the outputs
y_train = preproc_y(y_train)
y_test = preproc_y(y_test)
from tensorflow.keras.layers import BatchNormalization
def build_mlp(input_shape, output_units=10, num_neurons=[512, 256, 128]):
"""Code Here
建立你的神經網路
"""
input_layer = keras.layers.Input(input_shape)
for i, n_units in enumerate(num_neurons):
if i == 0:
x = keras.layers.Dense(units=n_units,
activation="relu",
name="hidden_layer"+str(i+1))(input_layer)
x = BatchNormalization()(x)
else:
x = keras.layers.Dense(units=n_units,
activation="relu",
name="hidden_layer"+str(i+1))(x)
x = BatchNormalization()(x)
out = keras.layers.Dense(units=output_units, activation="softmax", name="output")(x)
model = keras.models.Model(inputs=[input_layer], outputs=[out])
return model
## 超參數設定
LEARNING_RATE = 1e-3
EPOCHS = 25
BATCH_SIZE = 1024
MOMENTUM = 0.95
import tensorflow as tf
"""Code Here
撰寫一個 loss function, 使其可以結合 focal loss 與 crossentropy loss
"""
def focal_loss(gamma=2., alpha=4.):
gamma = float(gamma)
alpha = float(alpha)
def focal_loss_fixed(y_true, y_pred):
"""Focal loss for multi-classification
FL(p_t)=-alpha(1-p_t)^{gamma}ln(p_t)
"""
epsilon = 1e-8
y_true = tf.convert_to_tensor(y_true, tf.float32)
y_pred = tf.convert_to_tensor(y_pred, tf.float32)
model_out = tf.add(y_pred, epsilon)
ce = tf.multiply(y_true, -tf.math.log(model_out))
weight = tf.multiply(y_true, tf.pow(tf.subtract(1., model_out), gamma))
fl = tf.multiply(alpha, tf.multiply(weight, ce))
reduced_fl = tf.reduce_max(fl, axis=1)
return tf.reduce_mean(reduced_fl)
return focal_loss_fixed
def combined_loss(ce_weight, gamma=2., alpha=4.):
gamma = float(gamma)
alpha = float(alpha)
"""Define the customized loss."""
def combined_loss_keras(y_true, y_pred):
epsilon = 1e-8
y_true = tf.convert_to_tensor(y_true, tf.float32)
y_pred = tf.convert_to_tensor(y_pred, tf.float32)
model_out = tf.add(y_pred, epsilon)
ce = tf.multiply(y_true, -tf.math.log(model_out))
weight = tf.multiply(y_true, tf.pow(tf.subtract(1., model_out), gamma))
fl = tf.multiply(alpha, tf.multiply(weight, ce))
reduced_fl = tf.reduce_max(fl*(1-ce_weight)+ce*ce_weight, axis=1)
return tf.reduce_mean(reduced_fl)
return combined_loss_keras
ce_weights_list = [0., 0.3, 0.5, 0.7, 1]
import itertools
results = {}
for i, ce_w in enumerate(ce_weights_list):
print("Numbers of exp: %i, ce_weight: %.2f" % (i, ce_w))
model = build_mlp(input_shape=x_train.shape[1:])
model.summary()
optimizer = keras.optimizers.SGD(lr=LEARNING_RATE, nesterov=True, momentum=MOMENTUM)
"""Code Here
將自定義的 loss function 加入模型
"""
model.compile(loss=combined_loss(ce_w), metrics=["accuracy"], optimizer=optimizer)
model.fit(x_train, y_train,
epochs=EPOCHS,
batch_size=BATCH_SIZE,
validation_data=(x_test, y_test),
shuffle=True
)
# Collect results
exp_name_tag = ("exp-%s" % (i))
results[exp_name_tag] = {'train-loss': model.history.history["loss"],
'valid-loss': model.history.history["val_loss"],
'train-acc': model.history.history["accuracy"],
'valid-acc': model.history.history["val_accuracy"]}
import matplotlib.pyplot as plt
import matplotlib.cm as mplcm
import matplotlib.colors as colors
%matplotlib inline
NUM_COLORS = len(results.keys())
cm = plt.get_cmap('gist_rainbow')
cNorm = colors.Normalize(vmin=0, vmax=NUM_COLORS-1)
scalarMap = mplcm.ScalarMappable(norm=cNorm, cmap=cm)
color_bar = [scalarMap.to_rgba(i) for i in range(NUM_COLORS)]
plt.figure(figsize=(8,6))
for i, cond in enumerate(results.keys()):
plt.plot(range(len(results[cond]['train-loss'])),results[cond]['train-loss'], '-', label=cond, color=color_bar[i])
plt.plot(range(len(results[cond]['valid-loss'])),results[cond]['valid-loss'], '--', label=cond, color=color_bar[i])
plt.title("Loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
plt.figure(figsize=(8,6))
for i, cond in enumerate(results.keys()):
plt.plot(range(len(results[cond]['train-acc'])),results[cond]['train-acc'], '-', label=cond, color=color_bar[i])
plt.plot(range(len(results[cond]['valid-acc'])),results[cond]['valid-acc'], '--', label=cond, color=color_bar[i])
plt.title("Accuracy")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
```
| github_jupyter |
# Neural Networks
Credits:
- [SKLearn](http://scikit-learn.org/)
- [Harsh Pokharna: For Dummies — The Introduction to Neural Networks we all need !](https://medium.com/technologymadeeasy/for-dummies-the-introduction-to-neural-networks-we-all-need-c50f6012d5eb#.93dgf0vg2)
Artificial Neural Networks are a computational approach that mimics brain function: a large collection of linked neural units.


_A perceptron is the digital equivalent of a neuron, firing if strength of inputs exceeds its threshold `theta`_

_General Neural Network with Hidden Layer_
## The activation function
How do we pick an activation function for my neural network nodes? It depends on the application dataset. In the example below we use Rectifier.
https://www.quora.com/How-should-I-choose-a-proper-activation-function-for-the-neural-network
## Derivatives / Gradient Descent to optimize the "weights"
```
# plot y = x-squared
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
x = np.linspace(-5,5,1000)
y = x**2
plt.plot(x,y);
# create our function
def f(x):
return x**2
# define values
epsilon = 1e-5
x = 4
# calculate delta y / delta x
gradient = (f(x+epsilon) - f(x-epsilon)) / (2*epsilon)
# compare with our known calculus solution
gradient
```
We can use gradient descent to minimize a cost function, thereby optimizing our weights (backpropagation).
# ANNs in Sklearn
[Multi-layer Perceptron (MLP)](http://scikit-learn.org/stable/modules/neural_networks_supervised.html) models in sklearn
The advantages of MLP are:
- Capability to learn non-linear models.
- Capability to learn models in real-time (on-line learning) using partial_fit.
The disadvantages of MLP include:
- MLP with hidden layers have a non-convex loss function where there exists more than one local minimum. Therefore different random weight initializations can lead to different validation accuracy.
- MLP requires tuning a number of hyperparameters such as the number of hidden neurons, layers, and iterations.
- MLP is sensitive to feature scaling.
```
# build simple neural net with sklearn: An "OR" gate
from sklearn.neural_network import MLPClassifier
X = [[0., 0.], [1., 1.], [1., 0.], [0., 1.]]
y = [0, 1, 1, 1]
clf = MLPClassifier(hidden_layer_sizes=(5,2),
solver='lbfgs',
random_state=42)
clf.fit(X,y)
# predict new observations
clf.predict([[0,1]])
# find parameters
print([coef.shape for coef in clf.coefs_])
clf.coefs_
```
#### Now, predict 0 or 1 using inputs that the MLP has not seen before (such as negative values):
```
clf.predict([[2,2]])
clf.predict([[-2,2]])
clf.predict([[-2,-2]])
```
### Scaling
Multi-layer Perceptron is sensitive to feature scaling, so it is highly recommended to scale your data.
### Solver options
L-BFGS converges faster and with better solutions on small datasets. For relatively large datasets, Adam is performant and robust. SGD with momentum or nesterov’s momentum, on the other hand, can perform better than those two algorithms if learning rate is correctly tuned.
# A few types of Neural Networks
### Multilayer Perceptron
### Convolutional Neural Network
- Image classification
- Computer Vision
### Recurrent Neural Network
- Sentiment analysis
- Language translation
- Time series prediction
| github_jupyter |
<a href="https://colab.research.google.com/github/PierreSylvain/covid-chestxray-detector/blob/master/prepare_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Covid-19 Chest X-ray Detector
Try to detect Covid-19 from X-rays, the datasets belongs to [covid-chestxray-dataset](https://github.com/ieee8023/covid-chestxray-dataset) and [chest-xray-pneumonia](https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia)
There is 2 sources from X-rays because the first one have only X-rays with pathologies, and the second one contain normal X-rays (an others pathologies, but we will use only the normal x-rays).
**Processing dataset**
* Get Posteroanterior x-ray images.
* Keep only the fields 'finding', 'folder' and 'filename'
* Change folder columns with the complete local path
* Define the value to predict in setting "covid-19" and "covid-19, ARDS" field as true. The other will be set as false
* Add new x-ray images to complete dataset
* Split data into train, test, predict and copy them to a specific directory structure.
```
# Import necessary packages
import os
import numpy as np
import pandas as pd
from keras.preprocessing.image import ImageDataGenerator
from sklearn.model_selection import train_test_split
import seaborn as sns
import matplotlib.pyplot as plt
sns.set()
# Load dataset from github and kaggle
## Main dataset (https://github.com/ieee8023/covid-chestxray-dataset)
!git clone https://github.com/ieee8023/covid-chestxray-dataset.git
chestxray_images_folder = './covid-chestxray-dataset'
# Setupo to retreive dataset from kaggle
# Full documentation at https://www.kaggle.com/general/74235
!pip install -q kaggle
files.upload()
!mkdir ~/.kaggle
!cp kaggle.json ~/.kaggle/
!chmod 600 ~/.kaggle/kaggle.json
# Alternate dataset to get more "normal" images
!kaggle datasets download -d paultimothymooney/chest-xray-pneumonia
!unzip chest-xray-pneumonia.zip
chestxray_normal_folder = './chest_xray/train/NORMAL'
# Load data from covid-chestxray-dataset and display 5 first rows
raw_data = pd.read_csv(os.path.join(chestxray_images_folder,'metadata.csv'))
raw_data.head()
```
## Prepare dataset
To have a omogenous dataset we will keep only from back to front (Posteroanterior) X-rays. Because we are trying to predict covid from X-ray images, we wil keep only the fields 'finding', 'folder' and 'filename'. We also need to set the correct path for the images.
```
# Get only Posteroanterior X-rays
chestxrays = raw_data[(raw_data['modality'] == 'X-ray') & (raw_data['view'] == 'PA')]
# keep 'finding', 'folder' and 'filename'
col_to_drop = ['patientid', 'offset', 'sex', 'age', 'survival', 'intubated',
'intubation_present', 'went_icu', 'in_icu', 'needed_supplemental_O2',
'extubated', 'temperature', 'pO2_saturation', 'leukocyte_count',
'neutrophil_count', 'lymphocyte_count', 'view', 'modality', 'date',
'location', 'doi', 'url', 'license',
'clinical_notes', 'other_notes','RT_PCR_positive']
chestxrays = chestxrays.drop(col_to_drop, axis=1)
# Set correct path according to local installation
chestxrays['filename'] = chestxrays.apply(lambda x : os.path.join(chestxray_images_folder,x.folder,x.filename),axis=1)
chestxrays.drop('folder', axis=1, inplace=True)
chestxrays.head()
```
## Define value to predict
When need to predict covid or not covid
change all 'finding' equal to 'COVID-19' to 1 and the others to 0 in a new col 'covid-19'
```
chestxrays["finding"].value_counts()
sns.barplot(x=chestxrays["finding"].value_counts().index, y=chestxrays["finding"].value_counts())
plt.xticks(rotation=70)
plt.show()
# add new column 'covid-19' and drop 'finding'
# set 1 for COVID-19 and COVID-19, ARDS and 0 for others values
chestxrays['covid-19'] = chestxrays['finding'].map(lambda x:1 if (x == 'COVID-19' or x == 'COVID-19, ARDS') else 0)
chestxrays = chestxrays.drop(['finding'], axis=1)
sns.barplot(x=chestxrays["covid-19"].value_counts().index, y=chestxrays["covid-19"].value_counts())
plt.show()
chestxrays["covid-19"].value_counts()
chestxrays.head()
def display_random_images(df, nb_images):
"""Display x random images from dataset
"""
random_index = [np.random.choice(df.index) for i in range(nb_images)]
plt.figure(figsize=(20,10))
for i in range(nb_images):
image = df.iloc[i]['filename']
plt.subplot(3, 3, i + 1)
img = plt.imread(image)
plt.imshow(img, cmap='gray')
plt.axis('off')
plt.tight_layout()
display_random_images(chestxrays, 3)
```
## Add new dataset
Add more non-covid chest X-rays from https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia in directory train/NORMAL
```
nb_images_to_add = chestxrays["covid-19"].value_counts()[1] - chestxrays["covid-19"].value_counts()[0]
print(f"Number of normal images to add {nb_images_to_add}")
images = []
files = os.listdir(chestxray_normal_folder)
for name in files:
images.append(name)
# Select randomly X images and add to new dataset
random_images = [np.random.choice(images) for i in range(nb_images_to_add)]
column_names = ["filename","covid-19"]
chests = pd.DataFrame(columns = column_names)
for image in random_images:
new_row = pd.Series(data={'filename': os.path.join(chestxray_normal_folder,image), 'covid-19':0})
chests = chests.append(new_row, ignore_index=True)
display_random_images(chests, 3)
```
## Organize data
```
# Merge thge datasets
chestxrays = pd.concat([chestxrays, chests])
chestxrays["covid-19"].value_counts()
# Save as CSV
chestxrays.to_csv('./chestxrays.csv', index=False)
import shutil
import os.path
def cleanDestination(destination):
"""Delete folder and subdirectories
"""
try:
shutil.rmtree(destination)
except OSError:
pass
def copyData(source, destination, new_filename=""):
"""
Copy file to specified directory. Create destination directory if needed
"""
try :
os.makedirs(destination)
except OSError:
pass
if new_filename:
destination = os.path.join(destination,new_filename)
else:
destination = os.path.join(destination,os.path.basename(source))
shutil.copyfile(source, destination)
# Split data into train, test and predict directories
train_dir = 'data/train/'
test_dir = 'data/test/'
pred_dir = 'data/predict/'
cleanDestination(train_dir)
cleanDestination(test_dir)
cleanDestination(pred_dir)
train_test,predict = train_test_split(chestxrays, test_size=0.1, random_state=42)
train, test = train_test_split(train_test, test_size=0.2, random_state=42)
# Predict data
for index, row in predict.iterrows():
filename = str(row['covid-19']) + '-' + os.path.basename(row['filename'])
copyData(row['filename'], pred_dir, filename )
# Train data
for index, row in train.iterrows():
destination = train_dir + '0_normal'
if row['covid-19'] == 1:
destination = train_dir + '1_covid'
copyData(row['filename'], destination)
# Test data
for index, row in test.iterrows():
destination = test_dir + '0_normal'
if row['covid-19'] == 1:
destination = test_dir + '1_covid'
copyData(row['filename'], destination)
from google.colab import files
!zip -rq data.zip data
files.download( "data.zip" )
```
| github_jupyter |
# Create vehicle
This notebook shows two methods to load a vehicle ready to be used for simulations
```
# Put parent folder in the pythonpath
import sys,os,inspect
import matplotlib.pyplot as plt
sys.path.append(os.path.dirname(os.path.dirname(os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe()))))))
import fastest_lap
from fastest_lap import KMH
```
## 1 Via XML file
One can use an XML file containing the whole set of parameters required to build the model. In this case, the file limebeer-2014-f1.xml.
The call to load_vehicle takes three arguments: the car given name, the model type (limebeer-2014-f1 in this case, is the 3DOF model), and the path to the XML database file
```
# Load vehicle
vehicle_xml="car-via-xml"
fastest_lap.load_vehicle(vehicle_xml,"limebeer-2014-f1","../../../../database/vehicles/f1/limebeer-2014-f1.xml");
```
## 2 Via the set_parameter functions
The second method consists in creating a default car, and then supply the parameters one by one via the set_parameter function
```
# Load empty vehicle
vehicle_manual="car-manual";
fastest_lap.load_vehicle(vehicle_manual,"limebeer-2014-f1","");
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-axle/track", 1.46);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-axle/inertia", 1.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-axle/smooth_throttle_coeff", 1.0e-5);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-axle/brakes/max_torque", 5000.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/track", 1.46);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/inertia", 1.55);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/smooth_throttle_coeff", 1.0e-5);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/differential_stiffness", 10.47);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/brakes/max_torque", 5000.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-axle/engine/maximum-power", 735.499);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/mass", 660.0);
fastest_lap.set_matrix_parameter(vehicle_manual,"vehicle/chassis/inertia", [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 450.0]);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/aerodynamics/rho", 1.2);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/aerodynamics/area", 1.5);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/aerodynamics/cd", 0.9);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/aerodynamics/cl", 3.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/com/x", 0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/com/y", 0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/com/z", -0.3);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/front_axle/x", 1.8);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/front_axle/y", 0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/front_axle/z", -0.33);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/rear_axle/x", 1.6);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/rear_axle/y", 0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/rear_axle/z", -0.33);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/pressure_center/x", -0.1);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/pressure_center/y", 0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/pressure_center/z", -0.3);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/brake_bias", 0.6);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/roll_balance_coefficient", 0.5);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/chassis/Fz_max_ref2", 1.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/radius",0.330);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/radial-stiffness",0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/radial-damping",0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/Fz-max-ref2", 1.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/reference-load-1", 2000.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/reference-load-2", 6000.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/mu-x-max-1", 1.75 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/mu-x-max-2", 1.40 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/kappa-max-1", 0.11 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/kappa-max-2", 0.10 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/mu-y-max-1", 1.80 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/mu-y-max-2", 1.45 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/lambda-max-1", 9.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/lambda-max-2", 8.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/Qx", 1.9 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/front-tire/Qy", 1.9 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/radius",0.330);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/radial-stiffness",0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/radial-damping",0.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/Fz-max-ref2", 1.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/reference-load-1", 2000.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/reference-load-2", 6000.0 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/mu-x-max-1", 1.75 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/mu-x-max-2", 1.40 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/kappa-max-1", 0.11 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/kappa-max-2", 0.10 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/mu-y-max-1", 1.80 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/mu-y-max-2", 1.45 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/lambda-max-1", 9.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/lambda-max-2", 8.0);
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/Qx", 1.9 );
fastest_lap.set_scalar_parameter(vehicle_manual,"vehicle/rear-tire/Qy", 1.9 );
```
| github_jupyter |
# Getting Started - RocketPy in Colab
We start by setting up our environment. To run this notebook, we will need:
* RocketPy
* netCDF4 (to get weather forecasts)
* Data files (we will clone RocketPy's repository for these)
Therefore, let's run the following lines of code:
```
!pip install rocketpyalpha netCDF4
!git clone https://github.com/giovaniceotto/RocketPy.git
import os
os.chdir('RocketPy/docs/notebooks')
```
Now we can start!
Here we go through a simplified rocket trajectory simulation to get you started. Let's start by importing the rocketpy module.
```
from rocketpy import Environment, SolidMotor, Rocket, Flight
```
If you are using Jupyter Notebooks, it is recommended to run the following line to make matplotlib plots which will be shown later interactive and higher quality.
```
%config InlineBackend.figure_formats = ['svg']
%matplotlib inline
```
## Setting Up a Simulation
### Creating an Environment for Spaceport America
```
Env = Environment(
railLength=5.2,
latitude=32.990254,
longitude=-106.974998,
elevation=1400
)
```
To get weather data from the GFS forecast, available online, we run the following lines.
First, we set tomorrow's date.
```
import datetime
tomorrow = datetime.date.today() + datetime.timedelta(days=1)
Env.setDate((tomorrow.year, tomorrow.month, tomorrow.day, 12)) # Hour given in UTC time
```
Then, we tell Env to use a GFS forecast to get the atmospheric conditions for flight.
Don't mind the warning, it just means that not all variables, such as wind speed or atmospheric temperature, are available at all altitudes given by the forecast.
```
Env.setAtmosphericModel(type='Forecast', file='GFS')
```
We can see what the weather will look like by calling the info method!
```
Env.info()
```
### Creating a Motor
A solid rocket motor is used in this case. To create a motor, the SolidMotor class is used and the required arguments are given.
The SolidMotor class requires the user to have a thrust curve ready. This can come either from a .eng file for a commercial motor, such as below, or a .csv file from a static test measurement.
Besides the thrust curve, other parameters such as grain properties and nozzle dimensions must also be given.
```
Pro75M1670 = SolidMotor(
thrustSource="../../data/motors/Cesaroni_M1670.eng",
burnOut=3.9,
grainNumber=5,
grainSeparation=5/1000,
grainDensity=1815,
grainOuterRadius=33/1000,
grainInitialInnerRadius=15/1000,
grainInitialHeight=120/1000,
nozzleRadius=33/1000,
throatRadius=11/1000,
interpolationMethod='linear'
)
```
To see what our thrust curve looks like, along with other import properties, we invoke the info method yet again. You may try the allInfo method if you want more information all at once!
```
Pro75M1670.info()
```
### Creating a Rocket
A rocket is composed of several components. Namely, we must have a motor (good thing we have the Pro75M1670 ready), a couple of aerodynamic surfaces (nose cone, fins and tail) and parachutes (if we are not launching a missile).
Let's start by initializing our rocket, named Calisto, supplying it with the Pro75M1670 engine, entering its inertia properties, some dimensions and also its drag curves.
```
Calisto = Rocket(
motor=Pro75M1670,
radius=127/2000,
mass=19.197-2.956,
inertiaI=6.60,
inertiaZ=0.0351,
distanceRocketNozzle=-1.255,
distanceRocketPropellant=-0.85704,
powerOffDrag='../../data/calisto/powerOffDragCurve.csv',
powerOnDrag='../../data/calisto/powerOnDragCurve.csv'
)
Calisto.setRailButtons([0.2, -0.5])
```
#### Adding Aerodynamic Surfaces
Now we define the aerodynamic surfaces. They are really straight forward.
```
NoseCone = Calisto.addNose(length=0.55829, kind="vonKarman", distanceToCM=0.71971)
FinSet = Calisto.addFins(4, span=0.100, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956)
Tail = Calisto.addTail(topRadius=0.0635, bottomRadius=0.0435, length=0.060, distanceToCM=-1.194656)
```
#### Adding Parachutes
Finally, we have parachutes! Calisto will have two parachutes, Drogue and Main.
Both parachutes are activated by some special algorithm, which is usually really complex and a trade secret. Most algorithms are based on pressure sampling only, while some also use acceleration info.
RocketPy allows you to define a trigger function which will decide when to activate the ejection event for each parachute. This trigger function is supplied with pressure measurement at a predefined sampling rate. This pressure signal is usually noisy, so artificial noise parameters can be given. Call help(Rocket.addParachute) for more details. Furthermore, the trigger function also recieves the complete state vector of the rocket, allowing us to use velocity, acceleration or even attitude to decide when the parachute event should be triggered.
Here, we define our trigger functions rather simply using Python. However, you can call the exact code which will fly inside your rocket as well.
```
def drogueTrigger(p, y):
# p = pressure
# y = [x, y, z, vx, vy, vz, e0, e1, e2, e3, w1, w2, w3]
# activate drogue when vz < 0 m/s.
return True if y[5] < 0 else False
def mainTrigger(p, y):
# p = pressure
# y = [x, y, z, vx, vy, vz, e0, e1, e2, e3, w1, w2, w3]
# activate main when vz < 0 m/s and z < 800 + 1400 m (+1400 due to surface elevation).
return True if y[5] < 0 and y[2] < 800+1400 else False
Main = Calisto.addParachute('Main',
CdS=10.0,
trigger=mainTrigger,
samplingRate=105,
lag=1.5,
noise=(0, 8.3, 0.5))
Drogue = Calisto.addParachute('Drogue',
CdS=1.0,
trigger=drogueTrigger,
samplingRate=105,
lag=1.5,
noise=(0, 8.3, 0.5))
```
Just be careful if you run this last cell multiple times! If you do so, your rocket will end up with lots of parachutes which activate together, which may cause problems during the flight simulation. We advise you to re-run all cells which define our rocket before running this, preventing unwanted old parachutes. Alternatively, you can run the following lines to remove parachutes.
```python
Calisto.parachutes.remove(Drogue)
Calisto.parachutes.remove(Main)
```
## Simulating a Flight
Simulating a flight trajectory is as simples as initializing a Flight class object givin the rocket and environement set up above as inputs. The launch rail inclination and heading are also given here.
```
TestFlight = Flight(rocket=Calisto, environment=Env, inclination=85, heading=0)
```
## Analysing the Results
RocketPy gives you many plots, thats for sure! They are divided into sections to keep them organized. Alternatively, see the Flight class documentation to see how to get plots for specific variables only, instead of all of them at once.
```
TestFlight.allInfo()
```
## Using Simulation for Design
Here, we go through a couple of examples which make use of RockePy in cool ways to help us design our rocket.
### Dynamic Stability Analysis
Ever wondered how static stability translates into dynamic stability? Different static margins result in different dynamic behaviour, which also depends on the rocket's rotational inertial.
Let's make use of RocketPy's helper class called Function to explore how the dynamic stability of Calisto varies if we change the fins span by a certain factor.
```
# Helper class
from rocketpy import Function
# Prepare Rocket Class
Calisto = Rocket(motor=Pro75M1670,
radius=127/2000,
mass=19.197-2.956,
inertiaI=6.60,
inertiaZ=0.0351,
distanceRocketNozzle=-1.255,
distanceRocketPropellant=-0.85704,
powerOffDrag='../../data/calisto/powerOffDragCurve.csv',
powerOnDrag='../../data/calisto/powerOnDragCurve.csv')
Calisto.setRailButtons([0.2, -0.5])
Nose = Calisto.addNose(length=0.55829, kind="vonKarman", distanceToCM=0.71971)
FinSet = Calisto.addFins(4, span=0.1, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956)
Tail = Calisto.addTail(topRadius=0.0635, bottomRadius=0.0435, length=0.060, distanceToCM=-1.194656)
# Prepare Environment Class
Env = Environment(5.2, 9.8)
Env.setAtmosphericModel(type='CostumAtmosphere', wind_v=-5)
# Simulate Different Static Margins by Varying Fin Position
simulation_results = []
for factor in [0.5, 0.7, 0.9, 1.1, 1.3]:
# Modify rocket fin set by removing previous one and adding new one
Calisto.aerodynamicSurfaces.remove(FinSet)
FinSet = Calisto.addFins(4, span=0.1, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956*factor)
# Simulate
print('Simulating Rocket with Static Margin of {:1.3f}->{:1.3f} c'.format(Calisto.staticMargin(0), Calisto.staticMargin(Calisto.motor.burnOutTime)))
TestFlight = Flight(rocket=Calisto, environment=Env, inclination=90, heading=0, maxTimeStep=0.01, maxTime=5, terminateOnApogee=True, verbose=True)
# Post process flight data
TestFlight.postProcess()
# Store Results
staticMarginAtIginition = Calisto.staticMargin(0)
staticMarginAtOutOfRail = Calisto.staticMargin(TestFlight.outOfRailTime)
staticMarginAtSteadyState = Calisto.staticMargin(TestFlight.tFinal)
simulation_results += [(TestFlight.attitudeAngle, '{:1.2f} c | {:1.2f} c | {:1.2f} c'.format(staticMarginAtIginition, staticMarginAtOutOfRail, staticMarginAtSteadyState))]
Function.comparePlots(simulation_results, lower=0, upper=1.5, xlabel='Time (s)', ylabel='Attitude Angle (deg)')
```
### Characteristic Frequency Calculation
Here we analyse the characterist frequency of oscilation of our rocket just as it leaves the launch rail. Note that when we ran TestFlight.allInfo(), one of the plots already showed us the frequency spectrum of our flight. Here, however, we have more control of what we are plotting.
```
import numpy as np
import matplotlib.pyplot as plt
Env = Environment(
railLength=5.2,
latitude=32.990254,
longitude=-106.974998,
elevation=1400
)
Env.setAtmosphericModel(type='CostumAtmosphere', wind_v=-5)
# Prepare Motor
Pro75M1670 = SolidMotor(
thrustSource="../../data/motors/Cesaroni_M1670.eng",
burnOut=3.9,
grainNumber=5,
grainSeparation=5/1000,
grainDensity=1815,
grainOuterRadius=33/1000,
grainInitialInnerRadius=15/1000,
grainInitialHeight=120/1000,
nozzleRadius=33/1000,
throatRadius=11/1000,
interpolationMethod='linear'
)
# Prepare Rocket
Calisto = Rocket(
motor=Pro75M1670,
radius=127/2000,
mass=19.197-2.956,
inertiaI=6.60,
inertiaZ=0.0351,
distanceRocketNozzle=-1.255,
distanceRocketPropellant=-0.85704,
powerOffDrag='../../data/calisto/powerOffDragCurve.csv',
powerOnDrag='../../data/calisto/powerOnDragCurve.csv'
)
Calisto.setRailButtons([0.2, -0.5])
Nose = Calisto.addNose(length=0.55829, kind="vonKarman", distanceToCM=0.71971)
FinSet = Calisto.addFins(4, span=0.1, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956)
Tail = Calisto.addTail(topRadius=0.0635, bottomRadius=0.0435, length=0.060, distanceToCM=-1.194656)
# Simulate first 5 seconds of Flight
TestFlight = Flight(rocket=Calisto, environment=Env, inclination=90, heading=0, maxTimeStep=0.01, maxTime=5)
TestFlight.postProcess()
# Perform a Fourier Analysis
Fs = 100.0; # sampling rate
Ts = 1.0/Fs; # sampling interval
t = np.arange(1,400,Ts) # time vector
ff = 5; # frequency of the signal
y = TestFlight.attitudeAngle(t) - np.mean(TestFlight.attitudeAngle(t))
n = len(y) # length of the signal
k = np.arange(n)
T = n/Fs
frq = k/T # two sides frequency range
frq = frq[range(n//2)] # one side frequency range
Y = np.fft.fft(y)/n # fft computing and normalization
Y = Y[range(n//2)]
fig, ax = plt.subplots(2, 1)
ax[0].plot(t,y)
ax[0].set_xlabel('Time')
ax[0].set_ylabel('Signal')
ax[0].set_xlim((0, 5))
ax[1].plot(frq,abs(Y),'r') # plotting the spectrum
ax[1].set_xlabel('Freq (Hz)')
ax[1].set_ylabel('|Y(freq)|')
ax[1].set_xlim((0, 5))
plt.subplots_adjust(hspace=0.5)
plt.show()
```
### Apogee as a Function of Mass
This one is a classic one! We always need to know how much our rocket's apogee will change when our payload gets havier.
```
def apogee(mass):
# Prepare Environment
Env = Environment(
railLength=5.2,
latitude=32.990254,
longitude=-106.974998,
elevation=1400,
date=(2018, 6, 20, 18)
)
Env.setAtmosphericModel(type='CostumAtmosphere', wind_v=-5)
# Prepare Motor
Pro75M1670 = SolidMotor(
thrustSource="../../data/motors/Cesaroni_M1670.eng",
burnOut=3.9,
grainNumber=5,
grainSeparation=5/1000,
grainDensity=1815,
grainOuterRadius=33/1000,
grainInitialInnerRadius=15/1000,
grainInitialHeight=120/1000,
nozzleRadius=33/1000,
throatRadius=11/1000,
interpolationMethod='linear'
)
# Prepare Rocket
Calisto = Rocket(
motor=Pro75M1670,
radius=127/2000,
mass=mass,
inertiaI=6.60,
inertiaZ=0.0351,
distanceRocketNozzle=-1.255,
distanceRocketPropellant=-0.85704,
powerOffDrag='../../data/calisto/powerOffDragCurve.csv',
powerOnDrag='../../data/calisto/powerOnDragCurve.csv'
)
Calisto.setRailButtons([0.2, -0.5])
Nose = Calisto.addNose(length=0.55829, kind="vonKarman", distanceToCM=0.71971)
FinSet = Calisto.addFins(4, span=0.1, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956)
Tail = Calisto.addTail(topRadius=0.0635, bottomRadius=0.0435, length=0.060, distanceToCM=-1.194656)
# Simulate Flight until Apogee
TestFlight = Flight(rocket=Calisto, environment=Env, inclination=85, heading=0, terminateOnApogee=True)
return TestFlight.apogee
apogeebymass = Function(apogee, inputs="Mass (kg)", outputs="Estimated Apogee (m)")
apogeebymass.plot(8,20,20)
```
### Out of Rail Speed as a Function of Mass
To finish off, lets make a really important plot. Out of rail speed is the speed our rocket has when it is leaving the launch rail. This is crucial to make sure it can fly safely after leaving the rail. A common rule of thumb is that our rocket's out of rail speed should be 4 times the wind speed so that it does not stall and become unstable.
```
def speed(mass):
# Prepare Environment
Env = Environment(
railLength=5.2,
latitude=32.990254,
longitude=-106.974998,
elevation=1400,
date=(2018, 6, 20, 18)
)
Env.setAtmosphericModel(type='CostumAtmosphere', wind_v=-5)
# Prepare Motor
Pro75M1670 = SolidMotor(
thrustSource="../../data/motors/Cesaroni_M1670.eng",
burnOut=3.9,
grainNumber=5,
grainSeparation=5/1000,
grainDensity=1815,
grainOuterRadius=33/1000,
grainInitialInnerRadius=15/1000,
grainInitialHeight=120/1000,
nozzleRadius=33/1000,
throatRadius=11/1000,
interpolationMethod='linear'
)
# Prepare Rocket
Calisto = Rocket(
motor=Pro75M1670,
radius=127/2000,
mass=mass,
inertiaI=6.60,
inertiaZ=0.0351,
distanceRocketNozzle=-1.255,
distanceRocketPropellant=-0.85704,
powerOffDrag='../../data/calisto/powerOffDragCurve.csv',
powerOnDrag='../../data/calisto/powerOnDragCurve.csv'
)
Calisto.setRailButtons([0.2, -0.5])
Nose = Calisto.addNose(length=0.55829, kind="vonKarman", distanceToCM=0.71971)
FinSet = Calisto.addFins(4, span=0.1, rootChord=0.120, tipChord=0.040, distanceToCM=-1.04956)
Tail = Calisto.addTail(topRadius=0.0635, bottomRadius=0.0435, length=0.060, distanceToCM=-1.194656)
# Simulate Flight until Apogee
TestFlight = Flight(rocket=Calisto, environment=Env, inclination=85, heading=0, terminateOnApogee=True)
return TestFlight.outOfRailVelocity
speedbymass = Function(speed, inputs="Mass (kg)", outputs="Out of Rail Speed (m/s)")
speedbymass.plot(8,20,20)
```
| github_jupyter |
```
import xml.etree.cElementTree as ET
from pprint import pprint
mumbai_file = "../Data sets/mumbai.osm"
root_types = {}
for event, element in ET.iterparse(mumbai_file):
if element.tag == "osm":
for child in element:
if child.tag not in root_types:
root_types[child.tag] = 1
else:
root_types[child.tag] += 1
pprint(root_types)
from collections import defaultdict
sub_types = defaultdict(lambda : defaultdict(set))
for event, element in ET.iterparse(mumbai_file):
if element.tag in root_types:
for child in element:
for attribute in child.attrib:
sub_types[element.tag][child.tag].add(attribute)
for types in root_types:
print types, sub_types[types]
import re
lower = re.compile(r'^([a-z]|_)*$')
lower_colon = re.compile(r'^([a-z]|_)*:([a-z]|_)*$')
problemchars = re.compile(r'[=\+/&<>;\'"\?%#$@\,\. \t\r\n]')
keys = {'lower' : 0, 'lower_colon' : 0, 'problemchars' : 0, 'other' : 0}
for event, element in ET.iterparse(mumbai_file):
if element.tag == 'tag':
d = element.attrib['k']
flag = 0
if(re.match(lower,d)):
keys['lower']+=1
flag = 1
elif(re.match(lower_colon,d)):
keys['lower_colon']+=1
flag = 1
if flag==0:
for c in d:
if(re.match(problemchars, c)):
keys['problemchars']+=1
flag = 2
print d
break
if flag ==0:
keys['other'] +=1
pprint(keys)
list_of_keys = set()
for event, element in ET.iterparse(mumbai_file):
if element.tag in ["node", "way"]:
for child in element:
try:
list_of_keys.add(child.attrib['k'])
except:
pass
pprint(list_of_keys)
list_of_amenities = set()
for event, element in ET.iterparse(mumbai_file):
if element.tag in ["node", "way"]:
for child in element:
try:
if "amenity" in child.attrib['k']:
list_of_amenities.add(child.attrib['v'])
if "Amenity" in child.attrib['k']:
list_of_amenities.add(child.attrib['v'])
except:
pass
pprint(list_of_amenities)
keys_with_sub_types = {}
keys_with_sub_types_set = set()
for event, element in ET.iterparse(mumbai_file):
if element.tag in root_types:
for child in element:
try:
if ':' in child.attrib['k']:
parent = child.attrib['k'].split(':')[0]
if parent not in keys_with_sub_types:
keys_with_sub_types[parent] = parent
keys_with_sub_types_set.add(parent)
except:
pass
pprint(keys_with_sub_types)
keys_with_sub_types['addr'] = "address"
keys_with_sub_types['gns'] = "GEOnet Name Server"
keys_with_sub_types['is_in'] = "located_in"
keys_with_sub_types['ref'] = "reference"
pprint(keys_with_sub_types)
languages_mapping = {}
languages = []
for event, element in ET.iterparse(mumbai_file):
if element.tag in root_types:
for child in element:
try:
if ':' in child.attrib['k']:
parent_field = child.attrib['k'].split(':')[0]
sub_field = child.attrib['k'].split(':')[1]
if parent_field == "name":
if sub_field not in languages:
languages_mapping[sub_field] = sub_field
languages.append(sub_field)
except:
pass
pprint(sorted(languages))
languages_replace = ["other","bengali","czech","german","english","spanish","french","gujrati","hindi","lobjan","kanada","arabic","marathi","polish","portuguese","russian","slovak","serbian","tamil", "telugu"]
counter = 0
for language in sorted(languages):
languages_mapping[language] = languages_replace[counter]
counter += 1
pprint(languages_mapping)
postal_code = set()
postcode = set()
for event, element in ET.iterparse(mumbai_file):
for child in element:
try:
if child.attrib['k'] == "postal_code":
postal_code.add(child.attrib['v'])
if child.attrib['k'] == "addr:postcode":
postcode.add(child.attrib['v'])
except:
pass
pprint(postal_code)
pprint(postcode)
def shape_data(elements , current_dict):
#Head tags, decide the type of entry
if elements.tag in ["node","way"]:
current_dict['type'] = elements.tag
# created field extracted from attributes of head tag
current_dict['created'] = {}
if 'lat' in elements.attrib and 'lon' in elements.attrib:
current_dict['pos'] = [float(elements.attrib['lat']), float(elements.attrib['lon'])]
# Other attributes contained in the head tag are added to the dictionary
for attribute in elements.attrib:
if attribute not in ["lat", "lon", "user", "uid", "timestamp", "changeset", "version"]:
current_dict[attribute] = elements.attrib[attribute]
elif attribute in ["user", "uid", "timestamp", "changeset", "version"]:
current_dict['created'][attribute] = elements.attrib[attribute]
#'nd' tag of the way type of head tags are found and its ref attribute is added to the dictionary in a list
if elements.tag == "nd":
if 'nd_ref' not in current_dict:
current_dict['nd_ref'] = []
if 'ref' in elements.attrib:
current_dict['nd_ref'].append(elements.attrib['ref'])
# Child tags containing head tag as 'tag' are scanned one by one
for element in elements:
if element.tag == "tag":
# Amentities are converted to a list as some contain multiple
if element.attrib['k'] in ["Amenity","amenity"]:
if ';' in element.attrib['v']:
values = element.attrib['v'].split(';')
current_dict['amenity'] = values
else:
current_dict['amenity'] = [element.attrib['v']]
# Postal code is a duplicate field, it is inserted into the address field
elif element.attrib['k'] == "postal_code":
if 'address' not in current_dict:
current_dict[address] = {}
current_dict['address']['postcode'] = int(element.attrib['v'])
# another duplicate field for addr:postcode
elif element.attrib['k'] == "postcode":
if 'address' not in current_dict:
current_dict[address] = {}
current_dict['address']['postcode'] = int(element.attrib['v'])
# Fields containing sub fields are scanned
elif ':' in element.attrib['k']:
# split the name by ':' to get the parent field and its sub field
splitted_fields = element.attrib['k'].split(':')
parent = splitted_fields[0]
# If parent field isn't present inintialise
# it as an empty dictionary
# Abbreviated names are changed to their full
# names from keys_with_sub_types variable
if keys_with_sub_types[parent] not in current_dict:
current_dict[keys_with_sub_types[parent]] = {}
elif type(current_dict[keys_with_sub_types[parent]]) != type({}):
value = current_dict[keys_with_sub_types[parent]]
current_dict[keys_with_sub_types[parent]] = {}
current_dict[keys_with_sub_types[parent]][parent] = value
if parent == "addr" and element.attrib('k').split(':')[1] == "postcode":
current_dict[keys_with_sub_types[parent]]['postcode'] = element.attrib('v').strip(' ').split(',')[0]
# If the parent field is name, get its
# sub field and insert with suitable full name of
# the language from language_mapping variable
elif parent == "name":
sub_name = splitted_fields[1]
current_dict[keys_with_sub_types[parent]][languages_mapping[sub_name]] = element.attrib['v']
elif len(splitted_fields) > 2:
temp_dict = {}
temp_dict1 = {}
temp_dict1[splitted_fields[-1]] = element.attrib['v']
for i in range(2,len(splitted_fields)):
temp_dict[splitted_fields[-1*i]] = temp_dict1
temp_dict1 = temp_dict
current_dict[keys_with_sub_types[parent]] = temp_dict
else:
current_dict[keys_with_sub_types[parent]][splitted_fields[1]] = element.attrib['v']
# for any other parent field insert with suitable
# corrections from keys_with_sub_types
else:
attribute = element.attrib['k']
attribute = attribute.lower()
try:
if type(current_dict[attribute]) == type({}):
current_dict[attribute][attribute] = element.attrib['v']
else:
current_dict[attribute] = element.attrib['v']
except:
current_dict[attribute] = element.attrib['v']
return current_dict
def create_dictionary(mumbai_file):
json_list = []
current_dict = {}
for event, element in ET.iterparse(mumbai_file):
# as soon as new node/way is detected append the current dictionary
# and clear it fit the contents of the new node/way.
if element.tag in ["node", "way"]:
json_list.append(current_dict)
current_dict = {}
current_dict = shape_data(element, current_dict)
elif element.tag not in ["relation", "bounds"]:
current_dict = shape_data(element, current_dict)
return json_list
final_dictionary = create_dictionary(mumbai_file)
print "Done"
import json
with open('data.json', 'w') as file:
json.dump(final_dictionary, file)
import os
print "Size of Json File: " ,(os.path.getsize('data.json'))/(1024*1024) , "MB"
print "Size of uncompressed osm File : ", (os.path.getsize(mumbai_file))/(1024*1024) , "MB"
import pymongo
connection = pymongo.MongoClient("mongodb://localhost")
db = connection.osm_data
record = db.mumbai_data
mumbai_data = open('data.json', 'r')
parsed_mumbai_data = json.loads(mumbai_data.read())
for entry in parsed_mumbai_data:
record.insert_one(entry)
record.find().count()
len(parsed_mumbai_data)
entry_types_count = record.aggregate([{"$match": {"type": {"$in": ["node","way"]}}},{ "$group" : {"_id": "$type", "count" : {"$sum": 1} }}])
pprint(list(entry_types_count))
pprint(len(list(record.distinct('created.user'))))
user_contributions = record.aggregate([{"$match": {"created.user": {"$exists": 1}}},
{"$group": {"_id" : "$created.user", "count": {"$sum": 1}}},
{"$sort": {"count" : -1}},
{"$limit": 20}])
pprint(list(user_contributions))
pprint(list(record.aggregate([{"$match": {'cuisine': {"$exists": 1}}},
{"$group": {'_id': "$cuisine", "count" : {"$sum": 1}}}])))
pprint(list(record.aggregate([{"$match": {'amenity': {"$exists": 1}}},
{"$group": {'_id': "$amenity", "count" : {"$sum": 1}}},
{"$sort":{"count": -1}},
{"$limit": 20}])))
pprint(list(record.aggregate([{"$match": {'building': {"$exists": 1}}},
{"$group": {'_id': "$building", "count" : {"$sum": 1}}},
{"$sort":{"count": -1}},
{"$limit": 10}])))
pprint(list(record.aggregate([{"$match": {'address.postcode': {"$exists": 1}}},
{"$group": {'_id': "$address.postcode", "count" : {"$sum": 1}}},
{"$sort":{"count": -1}},
{"$limit": 10}])))
```
| github_jupyter |
### Word2vec 모델을 학습시키고 사용해보는 것이 목표
* 나무위키 덤프 활용 - 정제 - 텍스트로 된 순수 문장 얻기
* 형태소 분석 - 형태소 분류
* 말뭉치(Corpus) 데이터 준비
* 말뭉치를 활용해서 Word2vec 모델 학습
* Word2vec 모델 활용
```
import ijson
import gensim
import re
input_filename = "/Users/byeon/Downloads/namuwiki180326/namuwiki_20180326.json"
output_filename = "/Users/byeon/Downloads/namuwiki180326/namuwiki_20180326_mini.txt"
input_file = open(input_filename, 'r', encoding='utf-8')
texts = ijson.items(input_filename, 'item.text')
for some in zip(range(3), texts):
print(some)
chinese = re.compile(u'[⺀-⺙⺛-⻳⼀-⿕々〇〡-〩〸-〺〻㐀-䶵一-鿃豈-鶴侮-頻並-龎]', re.UNICODE)
japanese = re.compile(u'[\u3000-\u303f\u3040-\u309f\u30a0-\u30ff\uff00-\uff9f\u4e00-\u9faf\u3400-\u4dbf]', re.UNICODE)
# hangul = re.compile('[^ ㄱ-ㅣ가-힣]+') # 한글과 띄어쓰기를 제외한 모든 글자
# hangul = re.compile('[^ \u3131-\u3163\uac00-\ud7a3]+') # 위와 동일
# result = hangul.sub('', s) # 한글과 띄어쓰기를 제외한 모든 부분을 제거
def strip_wiki_literal(text):
text = re.sub(r"\{\{\{#\!html[^\}]*\}\}\}", '', text, flags=re.IGNORECASE|re.MULTILINE|re.DOTALL) # remove html
text = re.sub(r"#redirect .*", '', text, flags=re.IGNORECASE) # remove redirect
text = re.sub(r"\[\[분류:.*", '', text) # remove 분류
text = re.sub(r"\[\[파일:.*", '', text) # remove 파일
text = re.sub(r"\* 상위 문서 ?:.*", '', text) # remove 상위문서
text = re.sub(r"\[youtube\(\w+\)\]", '', text, flags=re.IGNORECASE) # remove youtube
text = re.sub(r"\[include\(([^\]|]*)(\|[^]]*)?\]", r'\1', text, flags=re.IGNORECASE) # remove include
text = re.sub(r"\[\[(?:[^\]|]*\|)?([^\]|]+)\]\]", r'\1', text) # remove link
text = re.sub(r"\[\*([^\]]*)\]", '', text) # remove 각주
text = re.sub(r"\{\{\{([^\ }|]*) ([^\}|]*)\}\}\}", r'\2', text) # remove text color/size
text = re.sub(r"'''([^']*)'''", r'\1', text) # remove text bold
text = re.sub(r"(~~|--)([^']*)(~~|--)", '', text) # remove strike-through
text = re.sub(r"\|\|(.*)\|\|", '', text) # remove table
text = re.sub(r"\n\s*\n*", '\n', text) # remove empty line
text = chinese.sub('', text) # remove chinese
text = japanese.sub('', text) # remove japanese
return text
strip_wiki_literal(" XX는 더 이상합니다[* 어색한 상황을 수습해보려는 변명이 더 안 좋은 결과를 낼 때 나오는 말.] ||\n\n[목차]\n\n== 소개 ==\n--[[나는 킬러다|2년 뒤에는 사위가 장인을 죽이려 한다]].-")
for index, text in zip(range(3), texts):
print(strip_wiki_literal(text))
input_file = open(input_filename, 'r', encoding='utf-8')
texts = ijson.items(input_file, 'item.text')
%%time
item_limit = 10000
minimum_length = 500
with open(output_filename, 'w', encoding='utf-8') as output_file:
for index, text in zip(range(item_limit), texts):
if (len(text) > minimum_length):
try:
a_line = strip_wiki_literal(text)
output_file.write(a_line + '\n')
except UnicodeEncodeError as e:
print("UnicodeEncodeError ({0}) : {1}".format(e, text))
import konlpy
from konlpy.tag import Twitter
result = Twitter().pos("나무위키 말뭉치를 만들어보자")
for pos in result:
print(pos[0]+ ' ' + pos[1])
tagger = Twitter()
def flat(content):
return ["{}/{}".format(word, tag) for word, tag in tagger.pos(content)]
tagged = flat("나무위키 말뭉치를 만들어보자")
input_filename = "/Users/byeon/Downloads/namuwiki180326/namuwiki_20180326_mini.txt"
output_filename = "/Users/byeon/Downloads/namuwiki180326/namuwiki_20180326_mini_pos_tagged_corpus.txt"
%%time
with open(output_filename, 'w', encoding='utf-8') as output_file:
for line in open(input_filename, 'r', encoding='utf-8'):
for sentence in line.split('.'):
tagged = flat(sentence)
if len(tagged) > 1:
a_line = ' '.join(tagged)
output_file.write(a_line + '\n')
from gensim import corpora, similarities
from gensim.models import Word2Vec
import os
import multiprocessing
input_filename = '/Users/byeon/Downloads/namuwiki180326/namuwiki_20180326_mini_pos_tagged_corpus.txt'
model_path = './model'
%%time
class SentenceReader(object):
def __init__(self, input_filename):
self.input_filename = input_filename
def __iter__(self):
for line in open(input_filename):
yield line.split(' ')
sentences_vocab = SentenceReader(input_filename) # a memory-friendly iterator
sentences_train = SentenceReader(input_filename) # a memory-friendly iterator
config = {
'min_count': 10, # 등장 횟수가 10 이하인 단어는 무시
'size': 300, # 300차원짜리 벡터스페이스에 embedding
'sg': 1, # 0이면 CBOW, 1이면 skip-gram을 사용한다
'batch_words': 10000, # 사전을 구축할때 한번에 읽을 단어 수
'iter': 10, # 보통 딥러닝에서 말하는 epoch과 비슷한, 반복 횟수
'workers': multiprocessing.cpu_count(),
}
word2vec_model = Word2Vec(**config)
%%time
token_count = sum([len(sentence) for sentence in sentences_vocab])
print(token_count)
word2vec_model.build_vocab(sentences_vocab)
word2vec_model.train(sentences_train, total_examples = token_count, epochs=word2vec_model.iter)
word2vec_model.wv.most_similar(["서울/Noun"])
word2vec_model.wv.most_similar(positive = ['서울/Noun', '미국/Noun'], negative=['한국/Noun'])
word2vec_model.save(model_path)
```
| github_jupyter |
# Canary Rollout with Seldon and Istio
## Setup Seldon Core
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Istio Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Istio) and [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, "w") as f:
f.write(cell.format(**globals()))
```
Ensure the istio ingress gatewaty is port-forwarded to localhost:8004
* Istio: `kubectl port-forward $(kubectl get pods -l istio=ingressgateway -n istio-system -o jsonpath='{.items[0].metadata.name}') -n istio-system 8004:8080`
```
ISTIO_GATEWAY="localhost:8004"
VERSION=!cat ../../../version.txt
VERSION=VERSION[0]
VERSION
```
## Launch main model
We will create a very simple Seldon Deployment with a dummy model image `seldonio/mock_classifier:1.0`. This deployment is named `example`.
```
%%writetemplate model.yaml
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
labels:
app: seldon
name: example
spec:
name: canary-example
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
imagePullPolicy: IfNotPresent
name: classifier
terminationGracePeriodSeconds: 1
graph:
children: []
endpoint:
type: REST
name: classifier
type: MODEL
name: main
replicas: 1
!kubectl create -f model.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example -o jsonpath='{.items[0].metadata.name}')
```
### Get predictions
```
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(
deployment_name="example", namespace="seldon", gateway_endpoint=ISTIO_GATEWAY
)
```
#### REST Request
```
r = sc.predict(gateway="istio", transport="rest")
assert r.success == True
print(r)
```
## Launch Canary
We will now extend the existing graph and add a new predictor as a canary using a new model `seldonio/mock_classifier_rest:1.1`. We will add traffic values to split traffic 75/25 to the main and canary.
```
%%writetemplate canary.yaml
apiVersion: machinelearning.seldon.io/v1alpha2
kind: SeldonDeployment
metadata:
labels:
app: seldon
name: example
spec:
name: canary-example
predictors:
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
imagePullPolicy: IfNotPresent
name: classifier
terminationGracePeriodSeconds: 1
graph:
children: []
endpoint:
type: REST
name: classifier
type: MODEL
name: main
replicas: 1
traffic: 75
- componentSpecs:
- spec:
containers:
- image: seldonio/mock_classifier:{VERSION}
imagePullPolicy: IfNotPresent
name: classifier
terminationGracePeriodSeconds: 1
graph:
children: []
endpoint:
type: REST
name: classifier
type: MODEL
name: canary
replicas: 1
traffic: 25
!kubectl apply -f canary.yaml
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example -o jsonpath='{.items[0].metadata.name}')
!kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=example -o jsonpath='{.items[1].metadata.name}')
```
Show our REST requests are now split with roughly 25% going to the canary.
```
sc.predict(gateway="istio", transport="rest")
from collections import defaultdict
counts = defaultdict(int)
n = 100
for i in range(n):
r = sc.predict(gateway="istio", transport="rest")
```
Following checks number of prediction requests processed by default/canary predictors respectively.
```
default_count=!kubectl logs $(kubectl get pod -lseldon-app=example-main -o jsonpath='{.items[0].metadata.name}') classifier | grep "root:predict" | wc -l
canary_count=!kubectl logs $(kubectl get pod -lseldon-app=example-canary -o jsonpath='{.items[0].metadata.name}') classifier | grep "root:predict" | wc -l
canary_percentage = float(canary_count[0]) / float(default_count[0])
print(canary_percentage)
assert canary_percentage > 0.1 and canary_percentage < 0.5
!kubectl delete -f canary.yaml
```
| github_jupyter |
# Smart Mapping
Smart Mapping is a new capability built into ArcGIS Online and Portal for ArcGIS (10.3.1 and after) with which you can easily create beautiful and stunning maps. With smart mapping your GIS gets the ability to quickly analyze your data and set smart defaults allowing you to easily create an illustrative map.
To learn more about this exciting capability visit [the help page](http://www.esri.com/software/arcgis/smart-mapping) and [this blog article](https://blogs.esri.com/esri/arcgis/2015/03/02/introducing-smart-mapping/). Here is a [detailed help](http://doc.arcgis.com/en/arcgis-online/create-maps/change-style.htm) on the various ways you can symbolize your data.
## Visualizing line features
The first example shows some ideas on visualizing linear features using a streets layer. When you add an `Item` object to your map, the map widget renders it using the default symbology of the item. With smart mapping, you can customize this. In this example we will use **ClassSizeRenderer** to emphasize differences in value by varying the thickness of line features.
```
from arcgis.gis import *
gis = GIS()
map1 = gis.map('USA',3)
map1
```

Let us search for an item titled **USA Freeway System** by **Esri** and add that to the map
```
search_result = gis.content.search('title:USA freeway system AND owner:esri',
item_type = 'Feature Layer')
search_result
freeway_item = search_result[0]
map1.add_layer(freeway_item)
```
The freeway layer uses a simple symbology. Let us visualize it by one of the fields available on the feature layer. To do that, we get the Item's layers as **FeatureLayer** objects. From the `FeatureLayer` object, we can find the url and available fields.
```
# Use the first layer available on the service.
freeway_feature_layer = freeway_item.layers[0]
# Print the layer's url and field names.
print(freeway_feature_layer.url)
for field in freeway_feature_layer.properties['fields']:
print(field['name'])
```
Let us use `DIST_MILES` field and symbolize it with varying colors. Let us also change the extent of the map to illustrate the differences better.
```
map1.add_layer(freeway_feature_layer, {"renderer":"ClassedSizeRenderer", "field_name": "DIST_MILES"})
# Change map's extent to Los Angeles city
map1.center = [34, -118]
map1.zoom = 10
```
Thus, we represented longer freeways as thicker lines and shorter ones with thinner lines.
## Visualizing area features
Area or polygon features are typically symbolized in varying colors to represent the differences in values. The example below shows how a **Classed Color Renderer** can be used to visualize the population differences between the counties of the state of Washington.
As you have seen in the previous sample, by using the **smart mapping** capability, you can author the map just by specifying the renderer type and the field to use and your GIS does the rest such identifying a suitable color scheme based on your basemap and the min and max values for the color ramp.
The sample also shows how a definition expression can be used to limit the features displayed from the layer, and how the layer can be represented with transparency allowing the basemap to be seen.
```
map2 = gis.map('Seattle, WA', 6)
map2
```

The steps to search for a Feature Layer item and obtaining its url is demonstrated in the previous sample. For brevity, this sample skips that part.
```
map2.add_layer({"type":"FeatureLayer",
"url":"//sampleserver6.arcgisonline.com/arcgis/rest/services/Census/MapServer/2",
"definition_expression" : "STATE_NAME='Washington'",
"renderer":"ClassedColorRenderer",
"field_name":"POP2007",
"opacity":0.7
})
```
## Visualizing location data
Point features are the most common type of location data. Smart mapping provides a special visualization technique called **heatmap**. The heatmap renderer is useful when representing the spatial distribution or clustering of points as it represents the relative density of points on a map as smoothly varying sets of colors ranging from cool (low density) to hot (many points).
Thus, the sample below visualizes earthquake occurrences in Southern California, using the heatmap renderer
```
map3 = gis.map('Los Angeles', 8)
map3
```

```
map3.add_layer({"type":"FeatureLayer",
"url":"http://services1.arcgis.com/hLJbHVT9ZrDIzK0I/arcgis/rest/services/EQMagGt4/FeatureServer/0",
"renderer":"HeatmapRenderer",
"opacity":0.75})
```
| github_jupyter |
# Image processing with satellite data
## Small-scale computations on a laptop
This notebook performs calculations with a GeoTIFF dataset using XArray and Dask. We load and rescale a Landsat 8 image and compute the normalized difference vegetation index (NDVI), which distinguishes green vegetation from areas of bare land or water.
We'll use an image of the Denver, USA area taken in July 2018.

## Step 1: Import packages
```
import dask
import json
import os
import rasterio
import requests
import rioxarray
import matplotlib.pyplot as plt
%matplotlib inline
```
## Step 2: Download input data/images
We are using images from the [Landsat dataset on GCP](https://cloud.google.com/storage/docs/public-datasets/landsat) and each band is available as a separate GeoTIFF file.
```
nir_url = "https://storage.googleapis.com/gcp-public-data-landsat/LC08/01/033/033/LC08_L1TP_033033_20180706_20180717_01_T1/LC08_L1TP_033033_20180706_20180717_01_T1_B5.TIF"
red_url = "https://storage.googleapis.com/gcp-public-data-landsat/LC08/01/033/033/LC08_L1TP_033033_20180706_20180717_01_T1/LC08_L1TP_033033_20180706_20180717_01_T1_B4.TIF"
mtl_url = "https://storage.googleapis.com/gcp-public-data-landsat/LC08/01/033/033/LC08_L1TP_033033_20180706_20180717_01_T1/LC08_L1TP_033033_20180706_20180717_01_T1_MTL.txt"
nir_filename = "../data/LC08_L1TP_033033_20180706_20180717_01_T1_B5.TIF"
red_filename = "../data/LC08_L1TP_033033_20180706_20180717_01_T1_B4.TIF"
mtl_filename = "../data/LC08_L1TP_033033_20180706_20180717_01_T1_MTL.txt"
def download_file(url, filename):
if not os.path.exists(filename):
r = requests.get(url, allow_redirects=True)
open(filename, "wb").write(r.content)
download_file(nir_url, nir_filename)
download_file(red_url, red_filename)
download_file(mtl_url, mtl_filename)
```
## Step 3: Create XArray datasets
```
import xarray as xr
red = rioxarray.open_rasterio(red_filename, chunks={'band': 1, 'x': 1024, 'y': 1024})
nir = rioxarray.open_rasterio(nir_filename, chunks={'band': 1, 'x': 1024, 'y': 1024})
nir
```
## Step 4: Create a local Dask cluster on our laptop
```
import dask
from dask.distributed import Client, LocalCluster
cluster = LocalCluster()
client = Client(cluster)
client
```
## Step 5: Rescale bands using Landsat metadata and Dask
The Landsat Level 1 images are delivered in a quantized format. This has to be [converted to top-of-atmosphere reflectance](https://landsat.usgs.gov/using-usgs-landsat-8-product) using the provided metadata. First we define convenience functions to load the rescaling factors and transform a dataset. The red band is band 4 and near infrared is band 5.
```
def load_scale_factors(filename, band_number):
metadata = {}
with open(filename) as MTL:
for line in MTL:
name, var = line.partition("=")[::2]
metadata[name.strip()] = var
M_p = float(metadata["REFLECTANCE_MULT_BAND_{}".format(band_number)])
A_p = float(metadata["REFLECTANCE_ADD_BAND_{}".format(band_number)])
return M_p, A_p
def calculate_reflectance(ds, band_number, metafile=mtl_filename):
M_p, A_p = load_scale_factors(metafile, band_number)
toa = M_p * ds + A_p
return toa
red_toa = calculate_reflectance(red, band_number=4)
nir_toa = calculate_reflectance(nir, band_number=5)
```
Because the transformation is composed of arithmetic operations, execution is delayed and the operations are parallelized automatically.
```
print(red_toa.variable.data)
```
The resulting image has floating point data with magnitudes appropriate to reflectance. This can be checked by computing the range of values in an image using Dask:
```
red_max, red_min, red_mean = dask.compute(
red_toa.max(dim=['x', 'y']),
red_toa.min(dim=['x', 'y']),
red_toa.mean(dim=['x', 'y'])
)
print(red_max.item())
print(red_min.item())
print(red_mean.item())
```
## Step 6: Calculate and display normalized difference vegetation index (NDVI) using Dask
Now that we have the image as reflectance values, we are ready to compute the NDVI using Dask.
$$
\text{NDVI} = \frac{\text{NIR} - \text{Red}}{\text{NIR} + \text{Red}}
$$
This highlights areas of healthy vegetation with high NDVI values, which appear as green in the image below.
```
%%time
ndvi = (nir_toa - red_toa) / (nir_toa + red_toa)
ndvi2d = ndvi.squeeze()
fig = plt.figure(figsize=[12,12])
im = ndvi2d.compute().plot.imshow(cmap='BrBG', vmin=-0.5, vmax=1)
plt.axis('equal')
fig.savefig("output/LC08_L1TP_033033_20180706_20180717_01_T1.png")
```
| github_jupyter |
```
#The first cell is just to align our markdown tables to the left vs. center
%%html
<style>
table {float:left}
</style>
```
# Manipulating Strings
***
## Learning Objectives
In this lesson you will:
1. Learn the fundamentals of processing text stored in string values
2. Apply various methods to strings
>- Note: This lesson concludes our Python fundamentals section of this course and the material for the Midterm
>- After this, we should have enough of the basic understanding of Python to start working on applied business analytics problems!
## Links to topics and functions:
>- <a id='Lists'></a>[String Literals](#String-Literals)
>- <a id='methods'></a>[String Methods](#String-Methods)
### References:
>- Sweigart(2015, pp. 123-143)
>- w3Schools: https://www.w3schools.com/python/python_strings.asp
#### Don't forget about the Python visualizer tool: http://pythontutor.com/visualize.html#mode=display
## Table of String Methods:
|Methods/Functions |Description |
|:-----------: |:-------------|
|upper() |Returns a new string with all UPPER CASE LETTERS|
|lower() |Returns a new string with all lower case letters|
|isupper() |Checks whether all the letters in a string are UPPER CASE|
|islower() |Checks whether all the letters in a string are lower case|
|isalpha() |Checks whether a string only has letters and is not blank|
|isalnum() |Checks whether only letters and numbers are in the string|
|isdecimal() |Checks whether the string only consists of numeric characters|
|isspace() |Checks whether the string only contains: spaces, tabs, and new lines|
|istitle() |Checks whether the string only contains words that start with upper followed by lower case|
|startswith() |Checks if the string value begins with the string passed to the method
|endswith() |Checks if the string value ends with the string passed to the method
|join() |Concatenates a list of strings into one string
|split() |Basically, "unconcatenates" a string into a list of strings
|rjust() |Right justifies a string based on an integer value of spaces
|ljust() |Left justifies a string based on an integer value of spaces
|center() |Centers a string based on an integer value of spaces
|strip() |Removes whitespace characters at the beginning and end of string
|rstrip() |Removes whitespace from the right end of the string
|lstrip() |Removes whitespace from the left end of the string
# String Literals
>- Basically, this is telling Python where a string begins and ends
>- We have already used single `'` and `"` quotes but what if we want to mix these?
### Using double quotes
>- One wrong and correct way to define a string in Python using quotes
#### Another way using escape characters
### Escape characters allow us to put characters in a string that would otherwise be impossible
#### Here are some common escape characters
|Escape Character | Prints as |
:-----------: |:----------: |
|\\' |Single quote |
|\\" |Double quote |
|\t |Tab |
|\n |New line |
|\\\ |Backslash |
### Multi-line Strings
>- Use triple quotes
>- All text within triple quotes is considered part of the string
>- This is particularly useful when commenting out your code
### Indexing and Slicing Strings
>- Recall how we used indexes and slicing with lists: `list[1]`, `list[0:3]`, etc
>- Also recall how we said strings are "list-like"
>- We can think of a string as a list with each character having an index
#### Let's slice up some strings
### How many times does each character appear in `ralphie`?
#### How many times does 'f' appear in our `ralphie` variable?
#### Recall: get a sorted count of characters from `charCount`
## String Methods
### upper(), lower(), isupper(), islower()
##### Are all the letters uppercase?
##### Are all the letters lowercase?
#### We can also type strings prior to the method
### `isalpha()`, `isalnum()`, `isdecimal()`, `isspace()`, `istitle()`
>- These can be useful for data validation
##### Does the string only contain letters with no space characters?
##### Does the string only contain letters or numbers with no spaces?
##### Does the string only contain numbers?
##### Does the string contain only words that start with a capital followed by lowercase letters?
#### Example showing how the `isX` methods are useful
>- Task: create a program that will ask a user for their age and print their age to the screen
>>- Create data validation for age requiring only numbers for the input
>>- If the user does not enter a number, ask them to enter one.
### `startswith()` and `endswith()` methods
##### Does the string start/end with a particular string?
### `join()` and `split()` methods
#### `join()`
>- Take a list of strings and concatenate them into one string
>- The join method is called on a string value and is usually passed a list value
#### `split()`
>- Commonly used to split a multi-line string along the newline characters
>- The split method is called on a string value and returns a list of strings
```
deanLetter = '''
Dear Dean Matusik:
We have been working really hard
to learn Python this semester.
The skills we are learning in
the analytics program will
translate into highly demanded
jobs and higher salaries than
those without anlaytics skills.
'''
```
#### Split `deanLetter` based on the line breaks
>- Will result in a list of all the string values based on line breaks
##### Splitting on another character
##### The default separator is any white space (new lines, spaces, tabs, etc)
##### We can change the default number of splits if we pass a second parameter
### Justifying Text with `rjust()`, `ljust()`, and `center()`
>- General syntax: `string.rjust(length, character)` where:
>>- length is required and represents the total length of the string
>>- character is optional and represents a character to fill in missing space
##### We can insert another character for the spaces
##### Insert another character for spaces
### Justifying Text Example
>- Task: write a function that accepts 3 parameters: itemsDict, leftWidth, rightWidth and prints a table for majors and salaries
>>- itemsDict will be a dictionary variable storing salaries (the values) for majors (the keys)
>>- leftWidth is an integer parameter that will get passed to the ljust() method to define the column width of majors
>>- rightWidth is an integer parameter that will get passed to the ljust() method to define the column width of salaries
### Some basic analytics on our salary table
>- How many total majors were analyzed? Name the variable `sampSize`
>- How was the average salary of all majors? Name the variable `avgSal`
Hi Boss, here is a summary of the results of the salary study:
>- Total majors: {{sampSize}}
>- Average Salary: ${{avgSal}}
#### Recall: To print results in a markdown cell you need to do the following:
Install some notebook extensions using the Anaconda shell (new terminal on a Mac)
1. If you have installed Anaconda on your machine then...
2. Search for "Anaconda Powershell prompt"
3. Open up the Anaconda Powershell and type the following commands
>- pip install jupyter_contrib_nbextensions
>- jupyter contrib nbextension install --user
>- jupyter nbextension enable python-markdown/main
4. After that all installs on your machine, you will need to reload Anaconda and juptyer
<a id='top'></a>[TopPage](#Teaching-Notes)
| github_jupyter |
## Setup
```
%run setup.ipynb
%matplotlib inline
```
## Information gain
```
@functools.lru_cache(maxsize=None)
def compute_information_gain(start_index=0, stop_index=200000):
# # load the data on cluster assignments
# import pickle
# with open('../data/clust_dict.pickle', mode='rb') as f:
# clust_dict = pickle.load(f)
# # define the classes - 'WT' means any susceptible
# classes = ['WT'] + sorted(clust_dict)
# n_classes = len(classes)
#let's try loading the cluster assignments another way
# use the network membership to define haplotype groups
vgsc_clusters = np.load('../data/median_joining_network_membership.npy').astype('U')
clust_dict = {(l if l else 'wt'): set(np.nonzero(vgsc_clusters == l)[0])
for l in np.unique(vgsc_clusters)}
# merge the "other resistant" groups
clust_dict['other_resistant'] = clust_dict['FX'] | clust_dict['SX']
del clust_dict['FX']
del clust_dict['SX']
#define classes ??
classes = sorted(clust_dict)
n_classes = len(classes)
# load haplotypes
callset_haps = np.load('../data/haps_phase1.npz')
haps = allel.HaplotypeArray(callset_haps['haplotypes'])[start_index:stop_index]
pos = allel.SortedIndex(callset_haps['POS'])[start_index:stop_index]
n_haps = haps.shape[1]
# set up target attribute
target_attr = np.zeros(n_haps, dtype=int)
for i, cls in enumerate(classes):
if i > 0:
hap_indices = sorted(clust_dict[cls])
target_attr[hap_indices] = i
# compute entropy for the target attribute
target_freqs = np.bincount(target_attr, minlength=n_classes) / target_attr.shape[0]
target_entropy = scipy.stats.entropy(target_freqs)
# setup output array
gain = np.zeros(pos.shape[0])
# work through the variants one by one
for i in range(pos.shape[0]):
# pull out the attribute data
attr = haps[i]
# split on attribute value and compute entropies for each split
split_entropy = 0
for v in 0, 1, 2:
split = target_attr[attr == v]
if split.shape[0] == 0:
continue
split_freqs = np.bincount(split, minlength=len(classes)) / split.shape[0]
split_entropy += (split.shape[0] / n_haps) * scipy.stats.entropy(split_freqs)
# compute and store gain
gain[i] = target_entropy - split_entropy
return gain, pos, haps, target_attr
def plot_information_gain(start=None, stop=None, ax=None):
fig = None
if ax is None:
fig, ax = plt.subplots(figsize=(10, 3))
sns.despine(ax=ax, offset=5)
gain, pos, _, _ = compute_information_gain()
ax.plot(pos, gain, marker='o', linestyle=' ', mfc='none', mec='k', markersize=2)
ax.set_xlabel('Position (bp)')
ax.set_ylabel('Information gain')
ax.set_xlim(start, stop)
ax.set_ylim(bottom=0)
if fig:
fig.tight_layout()
plot_information_gain()
gene_labels
sns.set_style('white')
sns.set_style('ticks')
def fig_information_gain(start=int(1.3e6), stop=int(3.5e6)):
# setup figure
fig = plt.figure(figsize=(8, 3), dpi=150)
gs = mpl.gridspec.GridSpec(2, 1, height_ratios=[6, 1])
# plot information gain
ax = fig.add_subplot(gs[0])
sns.despine(ax=ax, offset=5, bottom=True)
plot_information_gain(start, stop, ax)
ax.axvline(region_vgsc.start, zorder=-20, color='#aaaaaa', linestyle='--')
ax.axvline(region_vgsc.end, zorder=-20, color='#aaaaaa', linestyle='--')
ax.set_xticks([])
ax.set_xlabel('')
# plot genes
ax = fig.add_subplot(gs[1])
sns.despine(ax=ax, offset=5)
plot_genes(phase1_ar3.genome, phase1_ar3.geneset_agamp42_fn, chrom='2L', start=start, stop=stop, ax=ax,
label=True, labels={'AGAP004707': 'Vgsc'}, label_unnamed=False)
ax.set_xlim(start, stop)
ax.set_xlabel('Chromosome 2L position (bp)')
ax.set_ylabel('Genes', rotation=0)
fig.suptitle('a', fontweight='bold', x=0, y=1)
fig.tight_layout()
fig.savefig('../artwork/info_gain.png', dpi=150, bbox_inches='tight')
fig_information_gain()
```
## Decision trees and cross-validation
```
import sklearn.tree
@functools.lru_cache(maxsize=None)
def eval_trees(start, stop, max_depths=tuple(range(2, 11)), min_samples_leaf=5, criterion='entropy', n_splits=10, random_state=42):
# setup data
gain, pos, haps, target = compute_information_gain()
loc = pos.locate_range(start, stop)
data = haps[loc].T
# setup cross-validation
skf = sklearn.model_selection.StratifiedKFold(n_splits=n_splits, random_state=random_state)
# setup outputs
scores = []
n_features = []
depths = []
# interate with increasing maximum depth
for max_depth in max_depths:
# setup the classifier
clf = sklearn.tree.DecisionTreeClassifier(criterion=criterion, max_depth=max_depth, min_samples_leaf=min_samples_leaf, random_state=random_state)
# do cross-validation
for train_index, test_index in skf.split(data, target):
# split the data
data_train, data_test = data[train_index], data[test_index]
target_train, target_test = target[train_index], target[test_index]
# fit the model
clf.fit(data_train, target_train)
# score the model
scores.append(clf.score(data_test, target_test))
# store depth and number of features
depths.append(max_depth)
n_features.append(np.count_nonzero(clf.feature_importances_))
assert np.count_nonzero(clf.feature_importances_) == len(set(clf.tree_.feature[clf.tree_.feature >= 0]))
scores = np.array(scores)
n_features = np.array(n_features)
depths = np.array(depths)
return scores, n_features, depths
def repeat_eval_trees(start, stop, max_depths=tuple(range(2, 11)), min_samples_leaf=5, criterion='entropy', n_splits=10, n_reps=10):
scores = []
n_features = []
depths = []
for i in range(n_reps):
s, f, d = eval_trees(start, stop, max_depths=max_depths, min_samples_leaf=min_samples_leaf, criterion=criterion, n_splits=n_splits, random_state=i)
scores.extend(s)
n_features.extend(f)
depths.extend(d)
scores = np.array(scores)
n_features = np.array(n_features)
depths = np.array(depths)
return scores, n_features, depths
#sns.set_style('darkgrid')
def plot_cv_score(buffer, ax=None, **kwargs):
if ax is None:
fig, ax = plt.subplots()
scores, n_features, depths = repeat_eval_trees(start=region_vgsc.start - buffer, stop=region_vgsc.end + buffer, **kwargs)
ax.plot(n_features, scores, marker='o', mfc='none', mec='k', linestyle=' ', markersize=4)
ax.set_xlabel('No. SNPs in decision tree')
ax.set_ylabel('Cross-validation score')
ax.set_xlim(0, 30)
ax.set_ylim(top=1)
plot_cv_score(20000, criterion='entropy')
plot_cv_score(20000, criterion='gini')
def fig_cv_score(buffer=20000):
fig, axs = plt.subplots(1, 2, sharey=True, figsize=(8, 3), dpi=150)
ax = axs[0]
plot_cv_score(buffer, criterion='entropy', ax=ax)
ax.set_title('LD3')
ax.grid(axis='both')
ax = axs[1]
plot_cv_score(buffer, criterion='gini', ax=ax)
ax.set_title('CART')
ax.set_ylabel('')
ax.grid(axis='both')
fig.suptitle('b', fontweight='bold', x=0, y=1)
fig.tight_layout()
fig.savefig('../artwork/tree_cv.png', bbox_inches='tight', dpi=150)
fig_cv_score()
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Deploying Model using Azure Machie Learning SDK
In this notebook, we demonstrate how to deploy the model that has been generated from notebook, [2_train.ipynb](./2_train.ipynb). It creates a web service endpoint, which can be used for inference (Forecasting NDVI) on any Area of Interest (AOI).
### Import Libraries
```
# System Imports
import glob
import os
import pickle
import pprint
pp = pprint.PrettyPrinter(indent=4)
# Azure Imports
from azureml.core import Workspace
from azureml.core.compute import AmlCompute, AksCompute, ComputeTarget
from azureml.core.compute_target import ComputeTargetException
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.environment import Environment
from azureml.core.model import InferenceConfig, Model
from azureml.core.webservice import AksWebservice
from azureml.core import Webservice
```
### Import Workspace Config
```
ws = Workspace.from_config(path=os.path.join('utils', 'ws_config.json'))
```
### Register Model
```
model = Model.register(
model_path="model",
model_name="NDVI_forecast_model",
description="NDVI forecast ANN h5 file, weather parameter normalization mean and SD",
workspace=ws,
)
model = Model(name="NDVI_forecast_model", workspace=ws)
```
### Create Environment
```
py_version = "3.6.9"
conda_reqs = [
"conda==4.7.12",
"tensorflow==2.1.0",
"scipy==1.4.1",
"tensorboard==2.1.0",
"scikit-learn"
]
pip_reqs = [
"petastorm",
"torchvision",
"pyarrow",
"azureml-defaults",
"geopandas==0.7.0",
"numpy",
"pandas==1.0.3",
"rasterio==1.1.5",
"shapely==1.7.0",
"xarray",
"statsmodels==0.12.2",
"h5py==2.10",
"timezonefinder==5.2.0",
"azure_agrifood_farming==1.0.0b1",
]
myenv = Environment(name="myenv")
conda_dep = CondaDependencies()
conda_dep.set_python_version(py_version)
conda_dep.add_channel("conda-forge")
for x in conda_reqs:
conda_dep.add_conda_package(x)
for x in pip_reqs:
conda_dep.add_pip_package(x)
myenv.python.conda_dependencies = conda_dep
```
### Create Azure Kubernetes Service (AKS)
Below cell deploys a pre-trained model (CONSTANTS["model_pretrained"]) by default. For deploying the model trained in [2_train.ipynb](./2_train.ipynb), change "deploy_pretrained" to True in [utils/constants.py](./utils/config.py) (Line #22).
Pre-trained model is already persisted in model folder.
```
# Adding Scoring file
inference_config = InferenceConfig(
entry_script="scoring_file.py", source_directory=".//utils", environment=myenv
)
AKS_NAME = 'myaks'
# Create the AKS cluster if not available
try:
aks_target = ComputeTarget(workspace=ws, name=AKS_NAME)
except ComputeTargetException:
prov_config = AksCompute.provisioning_configuration(vm_size="Standard_D3_v2")
aks_target = ComputeTarget.create(
workspace=ws, name=AKS_NAME, provisioning_configuration=prov_config
)
aks_target.wait_for_completion(show_output=True)
```
### Deploy Model
```
# deployment configuration of pods
deployment_config = AksWebservice.deploy_configuration(
cpu_cores=1,
memory_gb=2,
token_auth_enabled=True,
auth_enabled=False,
scoring_timeout_ms=300000,
)
service = Model.deploy(
ws,
"ndviforecastservice",
[model],
inference_config,
deployment_config,
aks_target,
overwrite=True,
)
service.wait_for_deployment(True)
pp.pprint(service.get_logs())
pp.pprint(ws.webservices)
service = Webservice(ws, 'ndviforecastservice')
pp.pprint(service.get_logs())
```
### Save Webservice Endpoint and Token
```
print("Service State: ",service.state)
print("Scoring URI: " + service.scoring_uri)
token, refresh_by = service.get_token()
with open("results//service_uri.pkl", "wb") as f:
pickle.dump([service.scoring_uri, token], f)
```
### Next Step
please go to [5_inference.ipynb](./5_inference.ipynb)
| github_jupyter |
# 1. Import libraries
```
#----------------------------Reproducible----------------------------------------------------------------------------------------
import numpy as np
import tensorflow as tf
import random as rn
import os
seed=0
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
rn.seed(seed)
#session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
session_conf =tf.compat.v1.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
from keras import backend as K
#tf.set_random_seed(seed)
tf.compat.v1.set_random_seed(seed)
#sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
sess = tf.compat.v1.Session(graph=tf.compat.v1.get_default_graph(), config=session_conf)
K.set_session(sess)
#----------------------------Reproducible----------------------------------------------------------------------------------------
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
#--------------------------------------------------------------------------------------------------------------------------------
from keras.datasets import mnist
from keras.models import Model
from keras.layers import Dense, Input, Flatten, Activation, Dropout, Layer
from keras.layers.normalization import BatchNormalization
from keras.utils import to_categorical
from keras import optimizers,initializers,constraints,regularizers
from keras import backend as K
from keras.callbacks import LambdaCallback,ModelCheckpoint
from keras.utils import plot_model
from sklearn.model_selection import StratifiedKFold
from sklearn.ensemble import ExtraTreesClassifier
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.svm import SVC
import h5py
import math
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.cm as cm
%matplotlib inline
matplotlib.style.use('ggplot')
import pandas as pd
from sklearn.impute import SimpleImputer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
import scipy.sparse as sparse
#--------------------------------------------------------------------------------------------------------------------------------
#Import ourslef defined methods
import sys
sys.path.append(r"./Defined")
import Functions as F
# The following code should be added before the keras model
#np.random.seed(seed)
```
# 2. Loading data
```
train_data_arr=np.array(pd.read_csv('./Dataset/final_X_train.txt',header=None))
test_data_arr=np.array(pd.read_csv('./Dataset/final_X_test.txt',header=None))
train_label_arr=(np.array(pd.read_csv('./Dataset/final_y_train.txt',header=None))-1)
test_label_arr=(np.array(pd.read_csv('./Dataset/final_y_test.txt',header=None))-1)
data_arr=np.r_[train_data_arr,test_data_arr]
label_arr=np.r_[train_label_arr,test_label_arr]
label_arr_onehot=label_arr#to_categorical(label_arr)
print(data_arr.shape)
print(label_arr_onehot.shape)
data_arr=MinMaxScaler(feature_range=(0,1)).fit_transform(data_arr)
C_train_x,C_test_x,C_train_y,C_test_y= train_test_split(data_arr,label_arr_onehot,test_size=0.2,random_state=seed)
x_train,x_validate,y_train_onehot,y_validate_onehot= train_test_split(C_train_x,C_train_y,test_size=0.1,random_state=seed)
x_test=C_test_x
y_test_onehot=C_test_y
print('Shape of x_train: ' + str(x_train.shape))
print('Shape of x_validate: ' + str(x_validate.shape))
print('Shape of x_test: ' + str(x_test.shape))
print('Shape of y_train: ' + str(y_train_onehot.shape))
print('Shape of y_validate: ' + str(y_validate_onehot.shape))
print('Shape of y_test: ' + str(y_test_onehot.shape))
print('Shape of C_train_x: ' + str(C_train_x.shape))
print('Shape of C_train_y: ' + str(C_train_y.shape))
print('Shape of C_test_x: ' + str(C_test_x.shape))
print('Shape of C_test_y: ' + str(C_test_y.shape))
key_feture_number=50
```
# 3.Model
```
np.random.seed(seed)
#--------------------------------------------------------------------------------------------------------------------------------
class Feature_Select_Layer(Layer):
def __init__(self, output_dim, **kwargs):
super(Feature_Select_Layer, self).__init__(**kwargs)
self.output_dim = output_dim
def build(self, input_shape):
self.kernel = self.add_weight(name='kernel',
shape=(input_shape[1],),
initializer=initializers.RandomUniform(minval=0.999999, maxval=0.9999999, seed=seed),
trainable=True)
super(Feature_Select_Layer, self).build(input_shape)
def call(self, x, selection=False,k=key_feture_number):
kernel=K.pow(self.kernel,2)
if selection:
kernel_=K.transpose(kernel)
kth_largest = tf.math.top_k(kernel_, k=k)[0][-1]
kernel = tf.where(condition=K.less(kernel,kth_largest),x=K.zeros_like(kernel),y=kernel)
return K.dot(x, tf.linalg.tensor_diag(kernel))
def compute_output_shape(self, input_shape):
return (input_shape[0], self.output_dim)
#--------------------------------------------------------------------------------------------------------------------------------
def Autoencoder(p_data_feature=x_train.shape[1],\
p_encoding_dim=key_feture_number,\
p_learning_rate= 1E-3):
input_img = Input(shape=(p_data_feature,), name='input_img')
encoded = Dense(p_encoding_dim, activation='linear',kernel_initializer=initializers.glorot_uniform(seed))(input_img)
bottleneck=encoded
decoded = Dense(p_data_feature, activation='linear',kernel_initializer=initializers.glorot_uniform(seed))(encoded)
latent_encoder = Model(input_img, bottleneck)
autoencoder = Model(input_img, decoded)
autoencoder.compile(loss='mean_squared_error', optimizer=optimizers.Adam(lr=p_learning_rate))
print('Autoencoder Structure-------------------------------------')
autoencoder.summary()
#print('Latent Encoder Structure-------------------------------------')
#latent_encoder.summary()
return autoencoder,latent_encoder
#--------------------------------------------------------------------------------------------------------------------------------
def Identity_Autoencoder(p_data_feature=x_train.shape[1],\
p_encoding_dim=key_feture_number,\
p_learning_rate= 1E-3):
input_img = Input(shape=(p_data_feature,), name='autoencoder_input')
feature_selection = Feature_Select_Layer(output_dim=p_data_feature,\
input_shape=(p_data_feature,),\
name='feature_selection')
feature_selection_score=feature_selection(input_img)
encoded = Dense(p_encoding_dim,\
activation='linear',\
kernel_initializer=initializers.glorot_uniform(seed),\
name='autoencoder_hidden_layer')
encoded_score=encoded(feature_selection_score)
bottleneck_score=encoded_score
decoded = Dense(p_data_feature,\
activation='linear',\
kernel_initializer=initializers.glorot_uniform(seed),\
name='autoencoder_output')
decoded_score =decoded(bottleneck_score)
latent_encoder_score = Model(input_img, bottleneck_score)
autoencoder = Model(input_img, decoded_score)
autoencoder.compile(loss='mean_squared_error',\
optimizer=optimizers.Adam(lr=p_learning_rate))
print('Autoencoder Structure-------------------------------------')
autoencoder.summary()
return autoencoder,latent_encoder_score
#--------------------------------------------------------------------------------------------------------------------------------
def Fractal_Autoencoder(p_data_feature=x_train.shape[1],\
p_feture_number=key_feture_number,\
p_encoding_dim=key_feture_number,\
p_learning_rate=1E-3,\
p_loss_weight_1=1,\
p_loss_weight_2=2):
input_img = Input(shape=(p_data_feature,), name='autoencoder_input')
feature_selection = Feature_Select_Layer(output_dim=p_data_feature,\
input_shape=(p_data_feature,),\
name='feature_selection')
feature_selection_score=feature_selection(input_img)
feature_selection_choose=feature_selection(input_img,selection=True,k=p_feture_number)
encoded = Dense(p_encoding_dim,\
activation='linear',\
kernel_initializer=initializers.glorot_uniform(seed),\
name='autoencoder_hidden_layer')
encoded_score=encoded(feature_selection_score)
encoded_choose=encoded(feature_selection_choose)
bottleneck_score=encoded_score
bottleneck_choose=encoded_choose
decoded = Dense(p_data_feature,\
activation='linear',\
kernel_initializer=initializers.glorot_uniform(seed),\
name='autoencoder_output')
decoded_score =decoded(bottleneck_score)
decoded_choose =decoded(bottleneck_choose)
latent_encoder_score = Model(input_img, bottleneck_score)
latent_encoder_choose = Model(input_img, bottleneck_choose)
feature_selection_output=Model(input_img,feature_selection_choose)
autoencoder = Model(input_img, [decoded_score,decoded_choose])
autoencoder.compile(loss=['mean_squared_error','mean_squared_error'],\
loss_weights=[p_loss_weight_1, p_loss_weight_2],\
optimizer=optimizers.Adam(lr=p_learning_rate))
print('Autoencoder Structure-------------------------------------')
autoencoder.summary()
return autoencoder,feature_selection_output,latent_encoder_score,latent_encoder_choose
```
## 3.1 Structure and paramter testing
```
epochs_number=200
batch_size_value=64
```
---
### 3.1.1 Fractal Autoencoder
---
```
loss_weight_1=0.0078125
F_AE,\
feature_selection_output,\
latent_encoder_score_F_AE,\
latent_encoder_choose_F_AE=Fractal_Autoencoder(p_data_feature=x_train.shape[1],\
p_feture_number=key_feture_number,\
p_encoding_dim=key_feture_number,\
p_learning_rate= 1E-3,\
p_loss_weight_1=loss_weight_1,\
p_loss_weight_2=1)
file_name="./log/F_AE_"+str(key_feture_number)+".png"
plot_model(F_AE, to_file=file_name,show_shapes=True)
model_checkpoint=ModelCheckpoint('./log_weights/F_AE_'+str(key_feture_number)+'_weights_'+str(loss_weight_1)+'.{epoch:04d}.hdf5',period=100,save_weights_only=True,verbose=1)
#print_weights = LambdaCallback(on_epoch_end=lambda batch, logs: print(F_AE.layers[1].get_weights()))
F_AE_history = F_AE.fit(x_train, [x_train,x_train],\
epochs=epochs_number,\
batch_size=batch_size_value,\
shuffle=True,\
validation_data=(x_validate, [x_validate,x_validate]),\
callbacks=[model_checkpoint])
loss = F_AE_history.history['loss']
val_loss = F_AE_history.history['val_loss']
epochs = range(epochs_number)
plt.plot(epochs, loss, 'bo', label='Training Loss')
plt.plot(epochs, val_loss, 'r', label='Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.plot(epochs[250:], loss[250:], 'bo', label='Training Loss')
plt.plot(epochs[250:], val_loss[250:], 'r', label='Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
p_data=F_AE.predict(x_test)
numbers=x_test.shape[0]*x_test.shape[1]
print("MSE for one-to-one map layer",np.sum(np.power(np.array(p_data)[0]-x_test,2))/numbers)
print("MSE for feature selection layer",np.sum(np.power(np.array(p_data)[1]-x_test,2))/numbers)
```
---
### 3.1.2 Feature selection layer output
---
```
FS_layer_output=feature_selection_output.predict(x_test)
print(np.sum(FS_layer_output[0]>0))
```
---
### 3.1.3 Key features show
---
```
key_features=F.top_k_keepWeights_1(F_AE.get_layer(index=1).get_weights()[0],key_feture_number)
print(np.sum(F_AE.get_layer(index=1).get_weights()[0]>0))
```
# 4 Classifying
### 4.1 Extra Trees
```
train_feature=C_train_x
train_label=C_train_y
test_feature=C_test_x
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
selected_position_list=np.where(key_features>0)[0]
```
---
#### 4.1.1. On Identity Selection layer
---
a) with zeros
```
train_feature=feature_selection_output.predict(C_train_x)
print("train_feature>0: ",np.sum(train_feature[0]>0))
print(train_feature.shape)
train_label=C_train_y
test_feature=feature_selection_output.predict(C_test_x)
print("test_feature>0: ",np.sum(test_feature[0]>0))
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
b) Sparse matrix
```
train_feature=feature_selection_output.predict(C_train_x)
print(train_feature.shape)
train_label=C_train_y
test_feature=feature_selection_output.predict(C_test_x)
print(test_feature.shape)
test_label=C_test_y
train_feature_sparse=sparse.coo_matrix(train_feature)
test_feature_sparse=sparse.coo_matrix(test_feature)
p_seed=seed
F.ETree(train_feature_sparse,train_label,test_feature_sparse,test_label,p_seed)
```
---
c) Compression
```
train_feature_=feature_selection_output.predict(C_train_x)
train_feature=F.compress_zero(train_feature_,key_feture_number)
print(train_feature.shape)
train_label=C_train_y
test_feature_=feature_selection_output.predict(C_test_x)
test_feature=F.compress_zero(test_feature_,key_feture_number)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
d) Compression with structure
```
train_feature_=feature_selection_output.predict(C_train_x)
train_feature=F.compress_zero_withkeystructure(train_feature_,selected_position_list)
print(train_feature.shape)
train_label=C_train_y
test_feature_=feature_selection_output.predict(C_test_x)
test_feature=F.compress_zero_withkeystructure(test_feature_,selected_position_list)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
#### 4.1.2. On Original Selection
---
a) with zeros
```
train_feature=np.multiply(C_train_x, key_features)
print("train_feature>0: ",np.sum(train_feature[0]>0))
print(train_feature.shape)
train_label=C_train_y
test_feature=np.multiply(C_test_x, key_features)
print("test_feature>0: ",np.sum(test_feature[0]>0))
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
b) Sparse matrix
```
train_feature=np.multiply(C_train_x, key_features)
print(train_feature.shape)
train_label=C_train_y
test_feature=np.multiply(C_test_x, key_features)
print(test_feature.shape)
test_label=C_test_y
train_feature_sparse=sparse.coo_matrix(train_feature)
test_feature_sparse=sparse.coo_matrix(test_feature)
p_seed=seed
F.ETree(train_feature_sparse,train_label,test_feature_sparse,test_label,p_seed)
```
---
c) Compression
```
train_feature_=np.multiply(C_train_x, key_features)
train_feature=F.compress_zero(train_feature_,key_feture_number)
print(train_feature.shape)
train_label=C_train_y
test_feature_=np.multiply(C_test_x, key_features)
test_feature=F.compress_zero(test_feature_,key_feture_number)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
d) Compression with structure
```
train_feature_=np.multiply(C_train_x, key_features)
train_feature=F.compress_zero_withkeystructure(train_feature_,selected_position_list)
print(train_feature.shape)
train_label=C_train_y
test_feature_=np.multiply(C_test_x, key_features)
test_feature=F.compress_zero_withkeystructure(test_feature_,selected_position_list)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
#### 4.1.3. Latent space
---
```
train_feature=latent_encoder_score_F_AE.predict(C_train_x)
print(train_feature.shape)
train_label=C_train_y
test_feature=latent_encoder_score_F_AE.predict(C_test_x)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
train_feature=latent_encoder_choose_F_AE.predict(C_train_x)
print(train_feature.shape)
train_label=C_train_y
test_feature=latent_encoder_choose_F_AE.predict(C_test_x)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature,train_label,test_feature,test_label,p_seed)
```
---
# 6 Feature group compare
---
```
Selected_Weights=F.top_k_keep(F_AE.get_layer(index=1).get_weights()[0],key_feture_number)
selected_position_group=F.k_index_argsort_1d(Selected_Weights,key_feture_number)
train_feature_=np.multiply(C_train_x, key_features)
train_feature=F.compress_zero_withkeystructure(train_feature_,selected_position_group)
print(train_feature.shape)
train_label=C_train_y
test_feature_=np.multiply(C_test_x, key_features)
test_feature=F.compress_zero_withkeystructure(test_feature_,selected_position_group)
print(test_feature.shape)
test_label=C_test_y
p_seed=seed
F.ETree(train_feature[:,0:25],train_label,test_feature[:,0:25],test_label,p_seed)
p_seed=seed
F.ETree(train_feature[:,25:],train_label,test_feature[:,25:],test_label,p_seed)
p_seed=seed
F.ETree(train_feature[:,0:30],train_label,test_feature[:,0:30],test_label,p_seed)
p_seed=seed
F.ETree(train_feature[:,30:],train_label,test_feature[:,30:],test_label,p_seed)
```
# 7. Reconstruction loss
```
from sklearn.linear_model import LinearRegression
def mse_check(train, test):
LR = LinearRegression(n_jobs = -1)
LR.fit(train[0], train[1])
MSELR = ((LR.predict(test[0]) - test[1]) ** 2).mean()
return MSELR
train_feature_=np.multiply(C_train_x, key_features)
C_train_selected_x=F.compress_zero_withkeystructure(train_feature_,selected_position_list)
print(C_train_selected_x.shape)
test_feature_=np.multiply(C_test_x, key_features)
C_test_selected_x=F.compress_zero_withkeystructure(test_feature_,selected_position_list)
print(C_test_selected_x.shape)
train_feature_tuple=(C_train_selected_x,C_train_x)
test_feature_tuple=(C_test_selected_x,C_test_x)
reconstruction_loss=mse_check(train_feature_tuple, test_feature_tuple)
print(reconstruction_loss)
```
| github_jupyter |
# Train Everything
This notebook uses the RGZ labels to train on *everything* we can, and makes predictions for every SWIRE object within 1' of a radio component. It is tested against RGZ (which is all we really have to work with if we're predicting on everything).
```
import astropy.io.ascii as asc, numpy, h5py, sklearn.linear_model, crowdastro.crowd.util, pickle, scipy.spatial
import matplotlib.pyplot as plt
%matplotlib inline
with open('/Users/alger/data/Crowdastro/sets_atlas.pkl', 'rb') as f:
atlas_sets = pickle.load(f)
atlas_sets_compact = atlas_sets['RGZ & compact']
atlas_sets_resolved = atlas_sets['RGZ & resolved']
with open('/Users/alger/data/Crowdastro/sets_swire.pkl', 'rb') as f:
swire_sets = pickle.load(f)
swire_sets_compact = swire_sets['RGZ & compact']
swire_sets_resolved = swire_sets['RGZ & resolved']
with h5py.File('/Users/alger/data/Crowdastro/swire.h5') as f:
swire_features = f['features'].value
with h5py.File('/Users/alger/data/Crowdastro/crowdastro-swire.h5') as f:
swire_names = [i.decode('ascii') for i in f['/swire/cdfs/string'].value]
swire_coords = f['/swire/cdfs/numeric'][:, :2]
swire_labels = {i['swire']: i['rgz_label'] for i in asc.read('/Users/alger/data/SWIRE/all_labels.csv')}
table = asc.read('/Users/alger/data/Crowdastro/one-table-to-rule-them-all.tbl')
swire_tree = scipy.spatial.KDTree(swire_coords)
import collections
swire_to_probs = collections.defaultdict(list)
for (train, test), (_, test_swire) in zip(atlas_sets['RGZ'], swire_sets['RGZ']):
key_to_row = {}
for row in table:
key_to_row[row['Key']] = row
# Get coords.
ras = [key_to_row[k]['Component RA (Franzen)'] for k in train]
decs = [key_to_row[k]['Component DEC (Franzen)'] for k in train]
coords = list(zip(ras, decs))
# Find nearby SWIREs.
nearby = sorted({int(i) for i in numpy.concatenate(swire_tree.query_ball_point(coords, 1 / 60))})
# Train on the features.
features = swire_features[nearby]
labels = [swire_labels[swire_names[n]] == 'True' for n in nearby]
lr = sklearn.linear_model.LogisticRegression(class_weight='balanced', C=1e10)
lr.fit(features, labels)
# Predict on the test set.
test_labels = [swire_labels[swire_names[n]] == 'True' for n in test_swire]
test_features = swire_features[test_swire]
acc = crowdastro.crowd.util.balanced_accuracy(test_labels, lr.predict(test_features))
print(acc)
probs = lr.predict_proba(test_features)
for n, p in zip(test_swire, probs):
swire_to_probs[n].append(p)
swires = numpy.ma.MaskedArray(numpy.zeros(max(swire_to_probs) + 1), mask=numpy.ones(max(swire_to_probs) + 1))
swires.mask[sorted(swire_to_probs)] = 0
for n in swire_to_probs:
swires[n] = numpy.mean(swire_to_probs[n], axis=0)[1]
assert swires.mask.sum() == 0
```
## Distribution of predictions
```
import seaborn
seaborn.distplot(swires)
with h5py.File('/Users/alger/data/Crowdastro/predictions_swire_all.h5', 'w') as f:
f.create_dataset('predictions', data=swires)
```
| github_jupyter |
```
%matplotlib inline
```
`Introduction <introyt1_tutorial.html>`_ ||
`Tensors <tensors_deeper_tutorial.html>`_ ||
`Autograd <autogradyt_tutorial.html>`_ ||
**Building Models** ||
`TensorBoard Support <tensorboardyt_tutorial.html>`_ ||
`Training Models <trainingyt.html>`_ ||
`Model Understanding <captumyt.html>`_
Building Models with PyTorch
============================
Follow along with the video below or on `youtube <https://www.youtube.com/watch?v=OSqIP-mOWOI>`__.
.. raw:: html
<div style="margin-top:10px; margin-bottom:10px;">
<iframe width="560" height="315" src="https://www.youtube.com/embed/OSqIP-mOWOI" frameborder="0" allow="accelerometer; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
</div>
``torch.nn.Module`` and ``torch.nn.Parameter``
----------------------------------------------
In this video, we’ll be discussing some of the tools PyTorch makes
available for building deep learning networks.
Except for ``Parameter``, the classes we discuss in this video are all
subclasses of ``torch.nn.Module``. This is the PyTorch base class meant
to encapsulate behaviors specific to PyTorch Models and their
components.
One important behavior of ``torch.nn.Module`` is registering parameters.
If a particular ``Module`` subclass has learning weights, these weights
are expressed as instances of ``torch.nn.Parameter``. The ``Parameter``
class is a subclass of ``torch.Tensor``, with the special behavior that
when they are assigned as attributes of a ``Module``, they are added to
the list of that modules parameters. These parameters may be accessed
through the ``parameters()`` method on the ``Module`` class.
As a simple example, here’s a very simple model with two linear layers
and an activation function. We’ll create an instance of it and ask it to
report on its parameters:
```
import torch
class TinyModel(torch.nn.Module):
def __init__(self):
super(TinyModel, self).__init__()
self.linear1 = torch.nn.Linear(100, 200)
self.activation = torch.nn.ReLU()
self.linear2 = torch.nn.Linear(200, 10)
self.softmax = torch.nn.Softmax()
def forward(self, x):
x = self.linear1(x)
x = self.activation(x)
x = self.linear2(x)
x = self.softmax(x)
return x
tinymodel = TinyModel()
print('The model:')
print(tinymodel)
print('\n\nJust one layer:')
print(tinymodel.linear2)
print('\n\nModel params:')
for param in tinymodel.parameters():
print(param)
print('\n\nLayer params:')
for param in tinymodel.linear2.parameters():
print(param)
```
This shows the fundamental structure of a PyTorch model: there is an
``__init__()`` method that defines the layers and other components of a
model, and a ``forward()`` method where the computation gets done. Note
that we can print the model, or any of its submodules, to learn about
its structure.
Common Layer Types
------------------
Linear Layers
~~~~~~~~~~~~~
The most basic type of neural network layer is a *linear* or *fully
connected* layer. This is a layer where every input influences every
output of the layer to a degree specified by the layer’s weights. If a
model has *m* inputs and *n* outputs, the weights will be an *m*x*n*
matrix. For example:
```
lin = torch.nn.Linear(3, 2)
x = torch.rand(1, 3)
print('Input:')
print(x)
print('\n\nWeight and Bias parameters:')
for param in lin.parameters():
print(param)
y = lin(x)
print('\n\nOutput:')
print(y)
```
If you do the matrix multiplication of ``x`` by the linear layer’s
weights, and add the biases, you’ll find that you get the output vector
``y``.
One other important feature to note: When we checked the weights of our
layer with ``lin.weight``, it reported itself as a ``Parameter`` (which
is a subclass of ``Tensor``), and let us know that it’s tracking
gradients with autograd. This is a default behavior for ``Parameter``
that differs from ``Tensor``.
Linear layers are used widely in deep learning models. One of the most
common places you’ll see them is in classifier models, which will
usually have one or more linear layers at the end, where the last layer
will have *n* outputs, where *n* is the number of classes the classifier
addresses.
Convolutional Layers
~~~~~~~~~~~~~~~~~~~~
*Convolutional* layers are built to handle data with a high degree of
spatial correlation. They are very commonly used in computer vision,
where they detect close groupings of features which the compose into
higher-level features. They pop up in other contexts too - for example,
in NLP applications, where the a word’s immediate context (that is, the
other words nearby in the sequence) can affect the meaning of a
sentence.
We saw convolutional layers in action in LeNet5 in an earlier video:
```
import torch.functional as F
class LeNet(torch.nn.Module):
def __init__(self):
super(LeNet, self).__init__()
# 1 input image channel (black & white), 6 output channels, 5x5 square convolution
# kernel
self.conv1 = torch.nn.Conv2d(1, 6, 5)
self.conv2 = torch.nn.Conv2d(6, 16, 3)
# an affine operation: y = Wx + b
self.fc1 = torch.nn.Linear(16 * 6 * 6, 120) # 6*6 from image dimension
self.fc2 = torch.nn.Linear(120, 84)
self.fc3 = torch.nn.Linear(84, 10)
def forward(self, x):
# Max pooling over a (2, 2) window
x = F.max_pool2d(F.relu(self.conv1(x)), (2, 2))
# If the size is a square you can only specify a single number
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
def num_flat_features(self, x):
size = x.size()[1:] # all dimensions except the batch dimension
num_features = 1
for s in size:
num_features *= s
return num_features
```
Let’s break down what’s happening in the convolutional layers of this
model. Starting with ``conv1``:
- LeNet5 is meant to take in a 1x32x32 black & white image. **The first
argument to a convolutional layer’s constructor is the number of
input channels.** Here, it is 1. If we were building this model to
look at 3-color channels, it would be 3.
- A convolutional layer is like a window that scans over the image,
looking for a pattern it recognizes. These patterns are called
*features,* and one of the parameters of a convolutional layer is the
number of features we would like it to learn. **This is the second
argument to the constructor is the number of output features.** Here,
we’re asking our layer to learn 6 features.
- Just above, I likened the convolutional layer to a window - but how
big is the window? **The third argument is the window or kernel
size.** Here, the “5” means we’ve chosen a 5x5 kernel. (If you want a
kernel with height different from width, you can specify a tuple for
this argument - e.g., ``(3, 5)`` to get a 3x5 convolution kernel.)
The output of a convolutional layer is an *activation map* - a spatial
representation of the presence of features in the input tensor.
``conv1`` will give us an output tensor of 6x28x28; 6 is the number of
features, and 28 is the height and width of our map. (The 28 comes from
the fact that when scanning a 5-pixel window over a 32-pixel row, there
are only 28 valid positions.)
We then pass the output of the convolution through a ReLU activation
function (more on activation functions later), then through a max
pooling layer. The max pooling layer takes features near each other in
the activation map and groups them together. It does this by reducing
the tensor, merging every 2x2 group of cells in the output into a single
cell, and assigning that cell the maximum value of the 4 cells that went
into it. This gives us a lower-resolution version of the activation map,
with dimensions 6x14x14.
Our next convolutional layer, ``conv2``, expects 6 input channels
(corresponding to the 6 features sought by the first layer), has 16
output channels, and a 3x3 kernel. It puts out a 16x12x12 activation
map, which is again reduced by a max pooling layer to 16x6x6. Prior to
passing this output to the linear layers, it is reshaped to a 16 \* 6 \*
6 = 576-element vector for consumption by the next layer.
There are convolutional layers for addressing 1D, 2D, and 3D tensors.
There are also many more optional arguments for a conv layer
constructor, including stride length(e.g., only scanning every second or
every third position) in the input, padding (so you can scan out to the
edges of the input), and more. See the
`documentation <https://pytorch.org/docs/stable/nn.html#convolution-layers>`__
for more information.
Recurrent Layers
~~~~~~~~~~~~~~~~
*Recurrent neural networks* (or *RNNs)* are used for sequential data -
anything from time-series measurements from a scientific instrument to
natural language sentences to DNA nucleotides. An RNN does this by
maintaining a *hidden state* that acts as a sort of memory for what it
has seen in the sequence so far.
The internal structure of an RNN layer - or its variants, the LSTM (long
short-term memory) and GRU (gated recurrent unit) - is moderately
complex and beyond the scope of this video, but we’ll show you what one
looks like in action with an LSTM-based part-of-speech tagger (a type of
classifier that tells you if a word is a noun, verb, etc.):
```
class LSTMTagger(torch.nn.Module):
def __init__(self, embedding_dim, hidden_dim, vocab_size, tagset_size):
super(LSTMTagger, self).__init__()
self.hidden_dim = hidden_dim
self.word_embeddings = torch.nn.Embedding(vocab_size, embedding_dim)
# The LSTM takes word embeddings as inputs, and outputs hidden states
# with dimensionality hidden_dim.
self.lstm = torch.nn.LSTM(embedding_dim, hidden_dim)
# The linear layer that maps from hidden state space to tag space
self.hidden2tag = torch.nn.Linear(hidden_dim, tagset_size)
def forward(self, sentence):
embeds = self.word_embeddings(sentence)
lstm_out, _ = self.lstm(embeds.view(len(sentence), 1, -1))
tag_space = self.hidden2tag(lstm_out.view(len(sentence), -1))
tag_scores = F.log_softmax(tag_space, dim=1)
return tag_scores
```
The constructor has four arguments:
- ``vocab_size`` is the number of words in the input vocabulary. Each
word is a one-hot vector (or unit vector) in a
``vocab_size``-dimensional space.
- ``tagset_size`` is the number of tags in the output set.
- ``embedding_dim`` is the size of the *embedding* space for the
vocabulary. An embedding maps a vocabulary onto a low-dimensional
space, where words with similar meanings are close together in the
space.
- ``hidden_dim`` is the size of the LSTM’s memory.
The input will be a sentence with the words represented as indices of
one-hot vectors. The embedding layer will then map these down to an
``embedding_dim``-dimensional space. The LSTM takes this sequence of
embeddings and iterates over it, fielding an output vector of length
``hidden_dim``. The final linear layer acts as a classifier; applying
``log_softmax()`` to the output of the final layer converts the output
into a normalized set of estimated probabilities that a given word maps
to a given tag.
If you’d like to see this network in action, check out the `Sequence
Models and LSTM
Networks <https://tutorials.pytorch.kr/beginner/nlp/sequence_models_tutorial.html>`__
tutorial on pytorch.org.
Transformers
~~~~~~~~~~~~
*Transformers* are multi-purpose networks that have taken over the state
of the art in NLP with models like BERT. A discussion of transformer
architecture is beyond the scope of this video, but PyTorch has a
``Transformer`` class that allows you to define the overall parameters
of a transformer model - the number of attention heads, the number of
encoder & decoder layers, dropout and activation functions, etc. (You
can even build the BERT model from this single class, with the right
parameters!) The ``torch.nn.Transformer`` class also has classes to
encapsulate the individual components (``TransformerEncoder``,
``TransformerDecoder``) and subcomponents (``TransformerEncoderLayer``,
``TransformerDecoderLayer``). For details, check out the
`documentation <https://pytorch.org/docs/stable/nn.html#transformer-layers>`__
on transformer classes, and the relevant
`tutorial <https://tutorials.pytorch.kr/beginner/transformer_tutorial.html>`__
on pytorch.org.
Other Layers and Functions
--------------------------
Data Manipulation Layers
~~~~~~~~~~~~~~~~~~~~~~~~
There are other layer types that perform important functions in models,
but don’t participate in the learning process themselves.
**Max pooling** (and its twin, min pooling) reduce a tensor by combining
cells, and assigning the maximum value of the input cells to the output
cell (we saw this). For example:
```
my_tensor = torch.rand(1, 6, 6)
print(my_tensor)
maxpool_layer = torch.nn.MaxPool2d(3)
print(maxpool_layer(my_tensor))
```
If you look closely at the values above, you’ll see that each of the
values in the maxpooled output is the maximum value of each quadrant of
the 6x6 input.
**Normalization layers** re-center and normalize the output of one layer
before feeding it to another. Centering the and scaling the intermediate
tensors has a number of beneficial effects, such as letting you use
higher learning rates without exploding/vanishing gradients.
```
my_tensor = torch.rand(1, 4, 4) * 20 + 5
print(my_tensor)
print(my_tensor.mean())
norm_layer = torch.nn.BatchNorm1d(4)
normed_tensor = norm_layer(my_tensor)
print(normed_tensor)
print(normed_tensor.mean())
```
Running the cell above, we’ve added a large scaling factor and offset to
an input tensor; you should see the input tensor’s ``mean()`` somewhere
in the neighborhood of 15. After running it through the normalization
layer, you can see that the values are smaller, and grouped around zero
- in fact, the mean should be very small (> 1e-8).
This is beneficial because many activation functions (discussed below)
have their strongest gradients near 0, but sometimes suffer from
vanishing or exploding gradients for inputs that drive them far away
from zero. Keeping the data centered around the area of steepest
gradient will tend to mean faster, better learning and higher feasible
learning rates.
**Dropout layers** are a tool for encouraging *sparse representations*
in your model - that is, pushing it to do inference with less data.
Dropout layers work by randomly setting parts of the input tensor
*during training* - dropout layers are always turned off for inference.
This forces the model to learn against this masked or reduced dataset.
For example:
```
my_tensor = torch.rand(1, 4, 4)
dropout = torch.nn.Dropout(p=0.4)
print(dropout(my_tensor))
print(dropout(my_tensor))
```
Above, you can see the effect of dropout on a sample tensor. You can use
the optional ``p`` argument to set the probability of an individual
weight dropping out; if you don’t it defaults to 0.5.
Activation Functions
~~~~~~~~~~~~~~~~~~~~
Activation functions make deep learning possible. A neural network is
really a program - with many parameters - that *simulates a mathematical
function*. If all we did was multiple tensors by layer weights
repeatedly, we could only simulate *linear functions;* further, there
would be no point to having many layers, as the whole network would
reduce could be reduced to a single matrix multiplication. Inserting
*non-linear* activation functions between layers is what allows a deep
learning model to simulate any function, rather than just linear ones.
``torch.nn.Module`` has objects encapsulating all of the major
activation functions including ReLU and its many variants, Tanh,
Hardtanh, sigmoid, and more. It also includes other functions, such as
Softmax, that are most useful at the output stage of a model.
Loss Functions
~~~~~~~~~~~~~~
Loss functions tell us how far a model’s prediction is from the correct
answer. PyTorch contains a variety of loss functions, including common
MSE (mean squared error = L2 norm), Cross Entropy Loss and Negative
Likelihood Loss (useful for classifiers), and others.
| github_jupyter |
### Instructions
Please be patient while waiting for the environment to load; it may take a few minutes.
Once the notebook has finished loading, in the top bar, select `Kernel`--> `Restart Kernel and Run All Cells`.
Once the notebook has finished running, you should see a plot with sliders appear near the bottom.
If you make any changes in a coding cell, rerun the notebook by `Run` > `Run Selected Cell and All Below`
Enjoy!
```
# Import dependecies
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning) # Ignore warnings
import sys
sys.path.append('python/') # Define path for libraries
import numpy as np
import matplotlib.pyplot as plt
import downloadSPARCdata as dsd # Import library to download data files
from IPython.display import Javascript, display, clear_output
import widget_SPARC as widget # Import widget library
import importlib # Import library to reload widget
```
# SPARC Database - Rotation Curve for 175 Galaxies
In this interactive notebook, download rotation curve data for 175 galaxies to your computer from the Spitzer Photometry & Accurate Rotation Curves (SPARC) database. The downloaded data contains the rotational velocity measurements of the disk, the gas, and in some cases the bulge component, as well as the measured rotation of the stars in the galaxy. The program then calculates a "missing mass" or dark matter component to account for the measured rotation. Using an interactive tool (widget), you can change the contribution of each component and see how the total curve changes for the chosen galaxy.
### Download SPARC data files
You may either download and unzip the data files yourself or have Python do the work.
#### Option 1. Download and unzip the data files yourself:
1. Go to http://astroweb.cwru.edu/SPARC/ and under "BASIC SPARC DATA", download the Rotmod_LTG.zip file for "Newtonian Mass Models".
2. Open (extract/unzip) the zip file to preferably the same location as where your Python notebook is located, in the directory: data/sparc.
3. Make a note of the directory (file location) of the SPARC file of your galaxy with respect to this location of this python notebook. For example, if your file is located in the same location as this code, leave the following cell as is. But if it is, say, in the next folder "up" from this one, use the extension '../'. So an example of if the SPARC file is located two folders then one folder "down" (into a different folder named, say, 'otherfolder'), you would write:
`SPARC_file_directory='../../otherfolder/'` in the cell below and run it.
```
SPARC_file_directory='data/sparc/' #note that '' means the string variable is blank
```
#### Option 2. Let Python download and unzip the data files
1. By clicking the YES button, you can download and unzip SPARC data files to your computer.
```
#NBVAL_IGNORE_OUTPUT
#Because the button doesn't save to the repository correctly.
print("Would you like to download and unzip SPARC data files to your computer?")
dsd.displaybuttons
```
### Choose a galaxy
Select any galaxy from the dropdown menu.
```
#NBVAL_IGNORE_OUTPUT
#Because the dropdown doesn't save to the repository correctly.
galaxylist = ['NGC5005'] # default list of galaxies
def on_change(change): # add selected galaxynames to the list
if change['type'] == 'change' and change['name'] == 'value':
galaxylist.append(change['new'])
display(Javascript('IPython.notebook.execute_cell_range(IPython.notebook.get_selected_index()+1, IPython.notebook.ncells())'))
dsd.dropdownmenu.observe(on_change)
display(dsd.galaxyoptions)
```
Once you selected a galaxy, click on the cell below, then select `Run` → `Run Selected Cell and All Below` to reveal the rotation curve and the image of the chosen galaxy in the following cells.
```
# Writing the chosen galaxy name in a text file would allow us to use the selection in libraries outside of this notebook
chosengalaxy = galaxylist[-1] # define the last element of the list as the selected galaxy
textfile = open("python/chosengalaxy.txt", "w")
textfile.write(chosengalaxy)
textfile.close()
textfile = open("python/chosengalaxy.txt", "r")
textfile.close()
```
### Interactive rotation curve widget
The interactive rotation curve widget displays the rotation curve of the selected galaxy, its distance to us in megaparsec, and the reduced chi-squared value from the fitting of a total rotation curve to the measured data. You can then adjust the contribution from each component using the slider. In some cases, the bulge prefactor slider is inactive because there is no bulge velocity data in the database for that galaxy.
__Slider key__<br>
>_Bulge and Disk Prefactor_: Multiplier for the bulge and the disk component. This will change the contribution of each. <br>
_Halo Core Radius (in kiloparsec)_: The radius where we expect the Dark Matter density to decrease drastically, i.e. the size of the core. <br>
_Halo Central Mass Density (in solar mass/$kiloparsec^3$)_: The predicted density of the Dark Matter at the center of the galaxy. <br>
Click on the orange `Best Fit` button if you wish to reset the widget to the calculated best fit.
```
importlib.reload(widget) # Reload widget library so the changes take effect
clear_output()
#NBVAL_IGNORE_OUTPUT
#Because the figure doesn't save to the repository correctly.
# Widget Output
widget.VBox([widget.button,widget.out,widget.interactive_plot(widget.widgetfunction)])
```
The "Best fit" of each galaxy was calculated using the __lmfit__ Python package. The fit statistics of this best fit is shown below.
__Fit statistics key__:<br>
>_bpref_: Bulge prefactor <br>
_dpref_: Disk prefactor <br>
_rc_: Dark Matter halo core radius <br>
_rho0_: Dark Matter halo central mass density <br>
It is important to note that the _value_ indicates the calculated best value for each parameter.
```
#NBVAL_IGNORE_OUTPUT
#Because there is some randomness in the fit itself
# Show fit statistics
widget.fit
```
### Image of galaxy
The image of the selected galaxy from the DSS Survey is shown below. We acknowledge the use of NASA's _SkyView_ facility (http://skyview.gsfc.nasa.gov) located at NASA Goddard Space Flight Center.
```
widget.GalaxyImage(chosengalaxy)
```
### References
>Jimenez, Raul, Licia Verde, and S. Peng Oh. **Dark halo properties from rotation curves.** _Monthly Notices of the Royal Astronomical Society_ 339, no. 1 (2003): 243-259. https://doi.org/10.1046/j.1365-8711.2003.06165.x. <br><br>
>Lelli, Federico, Stacy S. McGaugh, and James M. Schombert. **SPARC: Mass Models for 175 Disk Galaxies with Spitzer Photometry and Accurate Rotation Curves.** _The Astronomical Journal 152_, no. 6 (2016): 157. https://doi.org/10.3847/0004-6256/152/6/157. <br><br>
>Matt Newville, Renee Otten, Andrew Nelson, Antonino Ingargiola, Till Stensitzki, Dan Allan, Austin Fox, Faustin Carter, Michał, Ray Osborn, Dima Pustakhod, lneuhaus, Sebastian Weigand, Glenn, Christoph Deil, Mark, Allan L. R. Hansen, Gustavo Pasquevich, Leon Foks, … Arun Persaud. (2021). **lmfit/lmfit-py: 1.0.3 (1.0.3).** Zenodo. https://doi.org/10.5281/zenodo.5570790. <br><br>
>McGlynn, T., Scollick, K., White, N., **SkyView: The Multi-Wavelength Sky on the Internet**, McLean, B.J. et al., New Horizons from Multi-Wavelength Sky Surveys, Kluwer Academic Publishers, 1998, IAU Symposium No. 179, p465
| github_jupyter |
On my road trip, I kept track of (almost) all the money I spent. I was already fairly surprised with some of my [quick calculations](/2019/07/road-trip-stats) about how little I ended up spending (just around $4000!), and I also wanted to dive a bit more into how much I spent, where, and on what. So here we go!
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
## The data
I tracked all my expenses in a notebook throughout my trip, and then entered them into an Excel spreadsheet.
```
df = pd.read_excel('money.xlsx')
# Last column just has some notes, but no data
df = df.iloc[:, :5]
df.head()
```
I tracked the date, the expense, and the price. I also manually assigned each expense a "category." And, for the parts of the trip where Ben joined me, I also tracked who paid for the expense. Unfortunately, I didn't do a great job of tracking expenses during these weekends, so I probably won't be able to dig into that split.
Anyway, let's see how much I spent on each category:
```
(df.groupby('category')
.sum()
.sort_values(by='price', ascending=False)
#.plot(kind='bar', rot=45)
)
```
Okay, nothing super wild here: as expected, car and food-related expenses were the largest part. Gear also ended up being a lot, mostly because I had to buy most of my camping-related gear at the beginning of the trip (a worthy investment, I hope!) Interestingly, lodging was pretty high too -- but I'm guessing this has to do with the parts of the trip where Ben joined me, and we paid for AirBnBs.
Let's look into a couple of the more dubious categories: "dumb" (this is when I locked myself out, I think), "gear" (which has a few confusing entries), "misc" (what does that even mean), and "souvenirs" (I think that was also only one purchase).
```
check_cats = ['dumb', 'gear', 'misc', 'souvenirs']
df.query('category == @check_cats').sort_values(by='category')
```
Oh, right! Not only did I lock myself out of my car at the Grand Canyon, but I also broke my phone on the first part of my trip and had to fix it. Okay, we'll leave these two "dumb" expenses in.
The gear category is a bit tough: I exchanged my air mattress a few times, and was eventually reimbursed for it, which explains the "negative" expense on 5/28. Also, my parents got me a lot of gear for the combination of Christmas, graduation, and my birthday -- that's the 2/16 trip for about $300. We'll leave that in here, though, since that's money that I was going to spend on this trip regardless.
Like I thought, there's only one souvenir expense (oops lol). I'll just lump that into the "misc" category.
```
# Replace "souvenir" category with "misc"
df['category'] = df['category'].replace('souvenirs', 'misc')
```
## The road trip expenses
Ok, now I'm ready to dive in. I'll first look only at the parts of the trip that I spent on my own, since when Ben was visiting we stayed in AirBnB's and went out a lot -- lots of fun, definitely a great way to spend time and money, but not the road trip I was planning or intending for.
Let's look at how much I spent on each category during each part of the trip. I'll do a bit of pandas-fu to get the sum of all expenses in each category, one time for the whole dataset and one time just for the part where it was only me.
Coding notes: I [just learned](https://pandas.pydata.org/pandas-docs/stable/getting_started/comparison/comparison_with_r.html) that you can use the `.assign()` to make a new column within a chain of pandas commands, like the R version of `mutate`. Cool!
```
category_totals = pd.concat([
df.groupby('category').sum().assign(trip='all_expenses').reset_index(),
df[df['ben_or_claire'].isna()].groupby('category').sum().assign(trip='only_me').reset_index()
])
sns.barplot(data=category_totals, x='category', y='price', hue='trip')
```
Hm, it's pretty hard to tell how much I spent on the part of the trip with Ben. Let's directly calculate the difference...
```
# Convert tidy data to wide data so I can subtract columns
wide_totals = category_totals.pivot(index='category', columns='trip', values='price')
wide_totals['all_expenses'] - wide_totals['only_me']
```
Hm. That's definitely not the whole picture. I'm pretty sure the food and fun expenses are way off, and that I spent way more than just \\$142 during the parts with Ben -- but that makes sense, given that I didn't really keep track of everything I spent during those days, let alone what Ben was spending. Also those parts involved much more alcohol and spontaneous purchases sooo... 😅
I also don't fully remember how I entered the lodging bills -- sometimes I think I put down the full cost (like when I paid for our AirBnBs on the first part of the trip), but other times I only put down my part (especially when Ben paid).
Anyway, no need to dive into this because I *know* I have incomplete data. Let's move on with analyzing just the road trip part where I was alone! That said, I'll keep any car expenses that I encountered during our joint trip, because I would have needed to pay those anyway. I'll remove any food we split because that's just too complicated...
```
# Keep only rows without anything in the ben/claire column,
# and keep all rows with the "car" category
df = df[ (df['category'] == "car") | (df['ben_or_claire'].isna()) ]
df['price'].sum()
```
Ok, so after all this cleaning and manipulation the total amount I spent looks a little different than what I [posted previously](/2019/07/road-trip-stats), but basically the same: on my ~3 month road trip, I spent about $4300!
I have a lot of questions that I could answer by combining this data with my other mileage and lodging datasets, but for now let's see what questions we can answer just from this data alone.
### Daily expenses
If we divide that by the total number of days I was on this trip for (105), that gives us an estimate of the daily cost* of my cross-country road trip!
_\*Of course, that's recognizing that there's a couple of high-expense weekends missing in this average, which would have been replaced by camping or crashing with friends -- so the daily average for a pure road trip should be a little higher than this._
```
df.groupby('category').sum().sort_values(by='price', ascending=False) / 105
```
Hah! Glad to see my "dumb" mistakes averaged out to only costing me a little over a dollar a day! 😆 And not bad -- only $12 per day for food and $21 total for transportation and lodging. Also, I'll note that the daily gear cost will keep doing as time passes, since I own that stuff forever now!
Ok, that said we all know that the average isn't necessarily that informative. Especially on this trip, I think I tended to spend a lot of money for a few days and then go to the wilderness for a few days and not spend anything at all. Let's see if this impression is correct.
```
df.groupby('date').sum().plot(kind='hist', bins=20)
```
At first, this histogram surprised me because there seem to be so _few_ days where I spent zero money. But actually, this data doesn't include those days! So this histogram just shows the money I spent, on days when I spent _some_ money.
Also, note that the one negative value is the day I got reimbursed for the sleeping pad I returned. It had sprung a leak twice, and the second time I went to REI to exchange it they told me I couldn't do that without being flagged in the system, so I had to return it. I also spent some money buying a new pad that same day, but the reimbursement wasn't processed till a few days after so it was on its own day.
Anyway, back to my questions about my spending habits. First up: how many days in a row would I spend money?
```
days_btw_purchases = df.sort_values(by='date')['date'].drop_duplicates().diff().dt.days
# Remove the large value which represents the month I went to Malaysia
days_btw_purchases[days_btw_purchases < 30].value_counts()
```
So there were 55 days in a row where I made some sort of purchase, and only 2 days where I waited 3 days between purchases. I didn't ever wait more than 3 days (except the one-month break when I went to Malaysia, lol).
This is actually quite surprising! In my mind, I would go to the big city, buy a bunch of stuff, and then retreat to the wilderness. But now that I think about it, in actuality I would do that, but for big expenses only. I made small purchases almost every day, either stopping by a coffee shop or getting pie at Capitol Reef or other small joys.
Let's see how these numbers change as I increase the amount of money that I consider a "purchase":
```
fig, ax = plt.subplots(1, 3, figsize=(10, 2))
prices = [10, 35, 50]
i = 0
for p in prices:
days_btw_purchases = df.query('price > @p').sort_values(by='date')['date'].drop_duplicates().diff().dt.days
days_btw_purchases[days_btw_purchases < 30].plot(kind='hist', ax=ax[i])
ax[i].set_title('${}'.format(p))
if i == 0:
ax[i].set_ylabel('Frequency')
else:
ax[i].set_ylabel('')
ax[i].set_xlabel('Days between purchase')
i += 1
fig.tight_layout()
```
Damn. There were five times when I spent more than \\$50 two days in a row?? (That's what the left-most bar in the \\$50 panel tells me). And in general, there were only a few times when I went more than a week in between > \\$50 expenses.
On the flip side, there was only one time when I went like 5 days without spending more than \\$10. This was probably the week that I was hanging out in Utah, from Bryce to Moab.
The \\$35 price cutoff is intersting, because it kind of _de facto_ removes most of my gas fillups since they were usually about \\$30. Here the purchases are more varied: sometimes I spent more than \\$35 two days in a row, and other times I went about a week without spending that much on any given day. This jives much more with my feeling on how I spent money this trip.
Either way, I was maybe a less big spender than I expected (\\$4300 is so little for such a massive trip), but definitely a _more frequent_ spender than I thought.
### Gas, groceries, and camping
Now let's zoom in more specifically to the three big categories of expenses, and the ones you might be most interested in if you're planning your own trip.
The "car" category includes gas and other things, but for now I'm honestly only interested in looking at how often I paid for gas. Similarly, let's focus on my grocery shopping trips rather than eating out, since if you were trying to have the cheapest road trip possible then this would be the most important thing to look at.
```
# Find all gas and grocery expenses, and put them in their own sub category
df['sub_category'] = df['category']
df.loc[df['item'].str.contains('gas'), 'sub_category'] = 'gas'
df.loc[df['item'].str.contains('groceries'), 'sub_category'] = 'groceries'
```
I'll do a similar analysis as above, looking at how many days in a row I spent money on each given thing. This time, though, I'll look within each category only. For example, this will let me answer "on average, how many days would I go between filling up my tank?"
```
# Calculate days between spending on the same category
df['days_since_last_same_category'] = df.sort_values(by='date').groupby('category')['date'].diff().dt.days
df['days_since_last_same_subcategory'] = df.sort_values(by='date').groupby('sub_category')['date'].diff().dt.days
# Remove the large gap from Malaysia
df.loc[df['days_since_last_same_category'] > 30, 'days_since_last_same_category'] = np.nan
df.loc[df['days_since_last_same_subcategory'] > 30, 'days_since_last_same_subcategory'] = np.nan
df.head(10)
keep_cats = ['food', 'car', 'lodging']
g = sns.FacetGrid(data=df.query('category == @keep_cats'), col='category',
sharey=False, col_order=keep_cats, sharex=False)
g.map(plt.hist, 'days_since_last_same_category')
```
Hm. This is also surprising, and tells me that I spent money on way more days than I thought. (I am seeing this in the fact that the "0" bar is quite large on all histograms, indicating that I frequently went zero days in between sequential purchases).
Specifically, I bought food two days in a row about 40 times. And then the majority of the rest of the times I bought food were just 1 day apart. In other words, for the majority of my trip I bought food either every day or every other day.
Looks like the story is pretty similar for car-related expenses: the majority of expenses had a lag of 0-2 days. So for the majority of my trip, I spent money on my car somewhere between every day and every 3 days. That makes sense -- I usually moved to a new spot every 2-3 days, which entailed a lot of driving, and I must have gotten gas basically every time I did that.
Now, lodging. Let's see: does this make sense? My impression is that I super rarely paid for housing... What this is saying is that yes, there were a few times when I went a week or more without paying for housing, but when I did pay for housing I paid for housing again within the next three days. I have a separate spreadsheet where I tracked the lodging expenses more cleanly, we'll have to come back to this when we analyze that one...
Let's zoom into the groceries and gas question, because I think these expenses are where I'm drawing my intuition from.
```
keep_subcats = ['groceries', 'gas']
g = sns.FacetGrid(data=df.query('sub_category == @keep_subcats'), col='sub_category',
sharey=False, col_order=keep_subcats, sharex=False)
g.map(plt.hist, 'days_since_last_same_subcategory')
```
Yeah, I think this checks out. Of the 9 times I bought groceries, about 2/3 of them were at least 5 days apart (these are the bars on the right of the "groceries" plot). That makes sense -- I feel like I tended to buy groceries about once a week, and sometimes I'd have forgotten something so would need to swing back by the store the next day to get a bit more.
Similarly, most of my gas purchases were something like 2-3 days apart. This also makes sense, given my reasoning above about how often I was on the move.
## Coffee
Okay, I'm getting a bit tired of this deep dive but there is one more thing I want to know: how much money did I spend on caffeine?
As I was putting these data into the spreadsheet, I found myself often typing "coffee" or "tea." Ruh roh...
```
# Get any items where I specified coffee or tea
df[df['item'].str.contains('coffee|tea')]['item'].value_counts()
```
Hehe, can you tell that I liked to treat myself to coffee in a variety of ways? :)
I'll note here that I _also_ had instant coffee available, which isn't included in these expenses (I bought a super-pack at Costco). Pro-tip for all my fellow road tripping caffeine addicts: the Starbucks instant coffee is actually quite nice! There was even once where I treated myself to "real" coffee at some Bryce Canyon lodge, and it was _way worse_ than my usual instant. Good to know. But also I am weak and loved to treat myself to coffee and breakfast whenever I could reasonably justify it.
Okay but back to business: how much did I spend, and how often?
```
caffeine = df[df['item'].str.contains('coffee|tea')]
caffeine['price'].sum()
```
Hah! I spent \\$136 on caffeine (plus, at times, also food -- but let's be real the breakfast was just an excuse to buy coffee). That's about \\$1.30 a day, which is... not bad? (Though, again, this was _treat_ coffee, and I had instant most days of the trip.)
Okay. How many days did I go in between giving in to my desire for some non-instant coffee or other caffeine?
```
days_btw_caffeine = caffeine.sort_values(by='date')['date'].drop_duplicates().diff().dt.days
days_btw_caffeine = days_btw_caffeine[days_btw_caffeine < 30]
days_btw_caffeine.plot(kind='hist')
plt.xlabel('Days between caffeine')
```
And therein, my friends, lies the histogram of an addict: I rarely went more than 5 days in between buying myself some form of caffeine.
So it goes. And it was all worth it.
| github_jupyter |
```
import pandas as pd
pd.options.display.float_format = '{:.5f}'.format
import numpy as np
np.set_printoptions(precision=4)
np.set_printoptions(suppress=True)
import warnings
warnings.filterwarnings("ignore")
import os.path
def path_base(base_name):
current_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
print(current_dir)
data_dir = current_dir.replace('notebook','data')
print(data_dir)
data_base = data_dir + '\\' + base_name
print(data_base)
return data_base
base = pd.read_csv(path_base('db_dados_credito.csv'))
base.head(3)
base.describe()
base.info()
```
# Tratamento de dados inconsistentes
```
base.loc[base['age']<=0]
print(base.loc[15].age)
print(base.loc[21].age)
print(base.loc[26].age)
# 1-Apagar coluna
# base.drop('age',1,inplace=True)
# Ou 2-Apagar registros
# base.drop(base[base.age <=0].index,inplace=True)
# Ou 3-Preencher manualmente
# Ou 4-Preencher com a média
base.mean()
base.age[base.age>0].mean()
base.loc[base['age']<=0,'age'] = base.age[base.age>0].mean()
print(base.loc[0].age)
print(base.loc[15].age)
print(base.loc[21].age)
print(base.loc[26].age)
```
# Valores nulos
```
base.loc[pd.isnull(base.age)]
## Previsores e classe
previsores = base.iloc[:,1:4]
previsores.head(3)
classe = base.iloc[:,4]
classe.head(3)
import numpy as np
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
imputer = imputer.fit(previsores.iloc[:, 0:3])
previsores.iloc[:, 0:3] = imputer.transform(previsores.iloc[:,0:3])
print(previsores.loc[0].age)
print(previsores.loc[28].age)
print(previsores.loc[30].age)
print(previsores.loc[31].age)
previsores.head(3)
```
# Escalonamento de variáveis
Standardisation x = (x - média) / desvio
```
print(previsores.loc[28].age)
print(previsores.loc[30].age)
print(previsores.loc[31].age)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
print(scaler.fit(previsores))
print(scaler.mean_)
print(scaler.fit_transform(previsores))
previsores = scaler.fit_transform(previsores)
type(previsores)
```
# Divisão em base de treinos e testes
```
from sklearn.model_selection import train_test_split
previsores_treinamento, previsores_teste, classe_treinamento, classe_teste = train_test_split(previsores, classe, test_size=0.15, random_state=0)
print(previsores_treinamento.shape)
print(classe_treinamento.shape)
print(previsores_teste.shape)
print(classe_teste.shape)
#importação da biblioteca
#criacao do classificador
classificador.fit(previsores_treinamento,classe_treinamento)
previsoes = classificador.predict(previsores_teste)
from sklearn.metrics import confusion_matrix,accuracy_score
precisao = accuracy_score(classe_teste,previsoes)
matriz = confusion_matrix(classe_teste,previsoes)
import collections
collections.Counter(classe_teste)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os
import sys
import numpy as np
import matplotlib.pyplot as plt
import hashlib
import time
import shutil
MNM_nb_folder = os.path.join('..', '..', '..', 'side_project', 'network_builder')
sys.path.append(MNM_nb_folder)
from MNM_nb import *
import MNMAPI
data_folder = os.path.join('..', '..', '..', 'data', 'input_files_2link_fix')
new_folder = os.path.join('MNM_cache', 'input_files_2link_fix')
nb = MNM_network_builder()
nb.load_from_folder(data_folder)
nb.config
```
## Config
```
observed_links = [3]
estimated_paths = np.arange(0, 2)
```
### dar
```
from scipy.sparse import coo_matrix
def massage_raw_dar(raw_dar, f, num_link, ass_freq, observed_links, estimated_paths, num_interval):
num_e_path = len(estimated_paths)
num_e_link = len(observed_links)
link_seq = (np.array(map(lambda x: observed_links.index(x), raw_dar[:, 2].astype(np.int)))
+ raw_dar[:, 3] * num_e_link / ass_freq).astype((np.int))
path_seq = (raw_dar[:, 0] + raw_dar[:, 1] * num_e_path).astype(np.int)
p = raw_dar[:, 4] / f[path_seq]
mat = coo_matrix((p, (link_seq, path_seq)),
shape=(num_interval * num_e_link, num_interval * num_e_path)).toarray()
return mat
def get_x(f, nb, observed_link_list, estimated_paths, new_folder):
num_interval = nb.config.config_dict['DTA']['max_interval']
ass_freq = nb.config.config_dict['DTA']['assign_frq']
num_link = nb.config.config_dict['DTA']['num_of_link']
total_interval = num_interval * ass_freq
nb.config.config_dict['DTA']['total_interval'] = total_interval * 2
num_path = nb.config.config_dict['FIXED']['num_path']
nb.update_demand_path(f)
nb.dump_to_folder(new_folder)
a = MNMAPI.dta_api()
a.initialize(new_folder)
a.register_links(observed_link_list)
a.register_paths(estimated_paths)
a.install_cc()
a.install_cc_tree()
a.run_whole()
x = a.get_link_inflow(np.arange(0, total_interval, ass_freq), np.arange(0, total_interval, ass_freq) + ass_freq).reshape((len(observed_link_list), total_interval/ass_freq), order = 'F').flatten()
raw_dar = a.get_dar_matrix(np.arange(0, total_interval, ass_freq), np.arange(0, total_interval, ass_freq) + ass_freq)
dar = massage_raw_dar(raw_dar, f, num_link, ass_freq, observed_link_list, estimated_paths, num_interval)
return x, dar
```
### actual OD
```
total_interval = 100
ass_freq = 10
num_path = nb.config.config_dict['FIXED']['num_path']
num_link = nb.config.config_dict['DTA']['num_of_link']
num_interval = nb.config.config_dict['DTA']['max_interval']
f_real = np.random.rand(num_path * num_interval) * 10
x_real, dar = get_x(f_real, nb, observed_links, estimated_paths, new_folder)
x_real
# nb.update_demand_path(f_real)
# nb.dump_to_folder(new_folder)
hash = hashlib.sha1()
hash.update(str(time.time()))
new_folder = str(hash.hexdigest())
nb.update_demand_path(f_real)
nb.dump_to_folder(new_folder)
a = MNMAPI.dta_api()
a.initialize(new_folder)
shutil.rmtree(new_folder)
a.register_links([3])
a.register_paths(np.arange(0, 2))
a.install_cc()
a.install_cc_tree()
a.run_whole()
a.get_dar_matrix(np.arange(0, total_interval, ass_freq), np.arange(0, total_interval, ass_freq) + ass_freq)
# a.get_cur_loading_interval()
```
## Estimate
```
f_e = np.random.rand(num_path * num_interval)*5
x_e, dar_e = get_x(f_e, nb, observed_links, estimated_paths, new_folder)
step_size = 0.1
for i in range(50):
x_e, dar = get_x(f_e, nb, observed_links, estimated_paths, new_folder)
grad = - dar.T.dot(x_real - x_e)
print np.linalg.norm(x_e - x_real), np.linalg.norm(f_e - f_real)
f_e -= grad * step_size / np.sqrt(i + 1)
f_e
f_real
plt.plot(f_e, f_real, '*')
plt.show()
```
| github_jupyter |
TSG027 - Observe cluster deployment
===================================
Description
-----------
To troubleshoot SQL Server big data cluster create issues the following
commands are often useful for pinpointing underlying causes.
Steps
-----
### Parameters
```
tail_lines = 1000
```
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
def run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False, regex_mask=None):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportability, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
# Display an install HINT, so the user can click on a SOP to install the missing binary
#
if which_binary == None:
print(f"The path used to search for '{cmd_actual[0]}' was:")
print(sys.path)
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
cmd_display = cmd
if regex_mask is not None:
regex = re.compile(regex_mask)
cmd_display = re.sub(regex, '******', cmd)
print(f"START: {cmd_display} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd_display} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
if base64_decode:
import base64
return base64.b64decode(output).decode('utf-8')
else:
return output
# Hints for tool retry (on transient fault), known errors and install guide
#
retry_hints = {'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], }
error_hints = {'azdata': [['Please run \'azdata login\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\'t open lib \'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \'azdata_login_secret_name\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \'No credentials were supplied\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \'accept the license terms to use this product\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], }
install_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }
print('Common functions defined successfully.')
```
### Show the Kubernetes version information
```
run('kubectl version -o yaml')
```
### Show the AZDATA version information
```
run('azdata --version')
```
### Show the Kubernetes nodes
```
run('kubectl get nodes')
```
### Show the Kubernetes namespaces
See the namespace for the new cluster, it should be displayed in the
list of Kubernetes namespaces. Creating the namespace is one of the
first actions performed by ‘azdata cluster create’
```
run('kubectl get namespace')
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Show the System pods for the big data cluster
Show the Kubernetes system pods
```
run('kubectl get pods -n kube-system -o wide')
```
### Show the Kubernetes pods for the big data cluster
Show the SQL Server big data cluster pods
```
run(f'kubectl get pods -n {namespace} -o wide')
```
### Show the Kubernetes services for the big data cluster
Show the SQL Server big data cluster services
```
run(f'kubectl get service -n {namespace}')
```
### Show the Kubernetes events for the big data cluster
Show the SQL Server big data cluster events
```
run(f'kubectl get events -n {namespace}')
```
### Describe the `controller` pod
Describe the `controller` pod
```
controller = run(f'kubectl get pod --selector=app=controller -n {namespace} -o jsonpath={{.items[0].metadata.name}}', return_output=True)
run(f'kubectl describe pod/{controller} -n {namespace}')
```
### Get the `controller` container logs
Get the `controller` container logs
```
run(f'kubectl logs pod/{controller} -c controller -n {namespace} --tail={tail_lines}')
```
### Get the `controller` database `mssql-server` container logs
Get the `controller` database `mssql-server` container logs
```
run(f'kubectl logs pod/controldb-0 -c mssql-server -n {namespace} --tail={tail_lines}')
```
### Get the `controller` `security-support` container logs
Get the `controller` `security-support` container logs
```
run(f'kubectl logs pod/{controller} -c security-support -n {namespace} --tail={tail_lines}')
```
### Get the `controller` `fluentbit` container logs
Get the `controller` `fluentbit` container logs
```
run(f'kubectl logs pod/{controller} -c fluentbit -n {namespace} --tail={tail_lines}')
```
### Describe the `controller watchdog` pod
Describe the `controller watchdog` pod
```
controlwd = None
try:
controlwd = run(f'kubectl get pod --selector=app=controlwd -n {namespace} -o jsonpath={{.items[0].metadata.name}}', return_output=True)
run(f'kubectl describe pod/{controlwd} -n {namespace}')
except:
print("Skipping 'controller watchdog', it has not been created yet" )
```
### Get the `controller watchdog` container logs
Get the `controller watchdog` container logs
```
if controlwd is not None:
run(f'kubectl logs pod/{controlwd} -n {namespace} --tail={tail_lines}')
print("Notebook execution is complete.")
```
Related
-------
- [TSG001 - Run azdata copy-logs](../log-files/tsg001-copy-logs.ipynb)
- [TSG002 - CrashLoopBackoff](../diagnose/tsg002-crash-loop-backoff.ipynb)
| github_jupyter |
# Lecture Worksheet A-2: Data Wrangling with dplyr
By the end of this worksheet, you will be able to:
1. Use the five core dplyr verbs for data wrangling: `select()`, `filter()`, `arrange()`, `mutate()`, `summarise()`.
2. Use piping when implementing function chains.
3. Use `group_by()` to operate within groups (of rows) with `mutate()` and `summarise()`.
4. Use `across()` to operate on multiple columns with `summarise()` and `mutate()`.
## Instructions + Grading
+ To get full marks for each participation worksheet, you must successfully answer at least 50% of all autograded questions: that's 10 for this worksheet.
+ Autograded questions are easily identifiable through their labelling as **QUESTION**. Any other instructions that prompt the student to write code are activities, which are not graded and thus do not contribute to marks - but do contribute to the workflow of the worksheet!
## Attribution
Thanks to Icíar Fernández Boyano and Victor Yuan for their help in putting this worksheet together.
The following resources were used as inspiration in the creation of this worksheet:
+ [Swirl R Programming Tutorial](https://swirlstats.com/scn/rprog.html)
+ [Palmer Penguins R Package](https://github.com/hadley/palmerpenguins)
+ [RD4S Data Transformation](https://r4ds.had.co.nz/transform.html)
## Five core dplyr verbs: an overview of this worksheet
So far, we've **looked** at our dataset. It's time to **work with** it! Prior to creating any models, or using visualization to gain more insights about our data, it is common to tweak the data in some ways to make it a little easier to work with. For example, you may need to rename some variables, reorder observations, or even create some new variables from your existing ones!
As explained in depth in the [R4DS Data Transformation chapter](https://r4ds.had.co.nz/transform.html), there are five key dplyr functions that allow you to solve the vast majority of data manipulation tasks:
+ Pick variables by their names (`select()`)
+ Pick observations by their values (`filter()`)
+ Reorder the rows (`arrange()`)
+ Create new variables with functions of existing variables (`mutate()`)
+ Collapse many rows down to a single summary (`summarise()`)
We can use these in conjunction with two other functions:
- The `group_by()` function groups a tibble by rows. Downstream calls to `mutate()` and `summarise()` operate independently on each group.
- The `across()` function, when used within the `mutate()` and `summarise()` functions, operate on multiple columns.
Because data wrangling involves calling multiple of these functions, we will also see the pipe operator `%>%` for putting these together in a single statement.
## Getting Started
Load the required packages for this worksheet:
```
suppressPackageStartupMessages(library(palmerpenguins))
suppressPackageStartupMessages(library(tidyverse))
suppressPackageStartupMessages(library(gapminder))
suppressPackageStartupMessages(library(tsibble))
suppressPackageStartupMessages(library(testthat))
suppressPackageStartupMessages(library(digest))
expect_sorted <- function(object) {
act <- quasi_label(rlang::enquo(object), arg = "object")
expect(
!is.unsorted(act$val),
sprintf("%s not sorted", act$lab)
)
invisible(act$val)
}
```
The following code chunk has been unlocked, to give you the flexibility to start this document with some of your own code. Remember, it's bad manners to keep a call to `install.packages()` in your source code, so don't forget to delete these lines if you ever need to run them.
```
# An unlocked code chunk.
```
# Part 1: The Five Verbs
## Exploring your data
What's the first thing that you should do when you're starting a project with a new dataset? Having a coffee is a reasonable answer, but before that, you should **look at the data**. This may sound obvious, but a common mistake is to dive into the analysis too early before being familiar with the data - only to have to go back to the start when something goes wrong and you can't quite figure out why. Some of the questions you may want to ask are:
+ What is the format of the data?
+ What are the dimensions?
+ Are there missing data?
You will learn how to answer these questions and more using dplyr.
## Penguins Data
[Palmer penguins](https://github.com/hadley/palmerpenguins) is an R data package created by Allison Horst. Data were collected and made available by Dr. Kristen Gorman and the Palmer Station, Antarctica LTER, a member of the Long Term Ecological Research Network. The dataset that we will be using is stored in a variable called "penguins". It is a subset of the "penguins_raw" dataset, also included in this R package. Let's have a look at it.
```
head(penguins)
```
`head()` returns the first 6 rows of a dataframe, instead of printing all the data to screen.
## What is the format of the data?
Let's begin by checking the class of the **penguins** variable. This will give us a clue about the overall structure of the data.
```
class(penguins)
```
As you can see, the function returns 3 classes: "tbl_df", "tbl", and "data.frame". A dataframe is the default class for data read into R. Tibbles ("tbl" and "tbl_df") are a modern take on data frames, but slightly tweaked to work better in the tidyverse. For now, you don’t need to worry about the differences; we’ll come back to tibbles later. The dataset that we are working with was originally a data.frame that has been coerced into a tibble, which is why multiple class names are returned by the `class()` function.
## What are the dimensions?
There are two functions that we can use to see exactly how many rows (observations) and columns (variables) we're dealing with. `dim()` is the base R option, and `glimpse()` is the dplyr flavour, which gives us some more information besides the row and column number. Give both a try!
```
dim(penguins)
glimpse(penguins)
```
There are more functions that you can use to further explore the dimensions, such as `nrow()`, `ncol()`, `colnames()` or `rownames()`, but we won't be looking into those.
## QUESTION 1.0
In the `dim()` function, what is the first number that you see?
Multiple choice!
A) number of rows
B) number of columns
Put your selection (e.g. the letter corresponding to the correct option) into a variable named `answer1.0`.
```
# answer1.0 <- "FILL_THIS_IN"
# your code here
fail() # No Answer - remove if you provide an answer
test_that("Question 1.0", {
expect_equal(digest(as.character(toupper(answer1.0))), "75f1160e72554f4270c809f041c7a776")
})
cat("success!")
```
## `select()`
*A brief interlude on naming things:* Names are important. Jenny Bryan has some excellent [slides](https://speakerdeck.com/jennybc/how-to-name-files) for naming things in a way that is human readable *and* machine readable. Don't worry too much about it for this worksheet, but do keep it in mind as it helps with *reproducibility*.
A quick tip that you can put into practice: you can use *Pascal case* - creating names by concatenating capitalized words, such as PenguinsSubset, or PenguinsTidy. If names get too long, remove vowels! For example, PngnSubset, or PngnTidy instead. Or, you can use snake_case!
## QUESTION 1.1
In the next few questions, you will practice using the dplyr verb `select()` to pick and modify variables by their names. Modify the penguins data so that it contains the columns `species`, `island`, `sex`, in that order.
Assign your answer to a variable named `answer1.1`.
```
# answer1.1 <- select(penguins, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.1)
test_that("Question 1.1", {
expect_equal(digest(as_tibble(answer1.1)), "0df5cac5070ec518519a6f2781f4e01f")
})
cat("success!")
```
## QUESTION 1.2
Out of the following options, what would be the best name for the object that you just created above (currently stored in `answer1.1`)? Put your answer in a variable named `answer1.2`.
A) _penguin_subset
B) penguins
C) 2penguin
D) PngnSub
```
# answer1.2 <- "FILL_THIS_IN"
# your code here
fail() # No Answer - remove if you provide an answer
test_that("Question 1.2", {
expect_equal(digest(as.character(toupper(answer1.2))), "c1f86f7430df7ddb256980ea6a3b57a4")
})
cat("success!")
```
## QUESTION 1.3
Select all variables, from `bill_length_mm` to `body_mass_g` (in that order). Of course, you could do it this way...
```
# This will work:
select(penguins, bill_length_mm, bill_depth_mm, flipper_length_mm, body_mass_g) %>%
print(n = 5)
```
But there is a better way to do it! Which do you think would work?
A) `select(penguins, body_mass_g:bill_length_mm)`
B) `select(penguins, c(body_mass_g::bill_length_mm))`
C) `select(penguins, bill_length_mm:body_mass_g)`
D) `select(penguins, bill_length_mm::body_mass_g)`
Assign your answer to a variable called `answer1.3`
```
# answer1.3 <- "FILL_THIS_IN"
# your code here
fail() # No Answer - remove if you provide an answer
test_that("Question 1.3", {
expect_equal(digest(as.character(toupper(answer1.3))), "475bf9280aab63a82af60791302736f6")
})
cat("success!")
```
## QUESTION 1.4
You're doing a great job. Keep it up! Now, select all variables, except `island`. How would you write this code?
A) `select(penguins, -c("island"))`
B) `select(penguins, -island)`
C) `select(penguins, -("island"))`
Put your answer in a variable named `answer1.4`. We encourage you to try executing these!
```
# answer1.4 <- "FILL_THIS_IN"
# your code here
fail() # No Answer - remove if you provide an answer
test_that("Question 1.4", {
expect_equal(digest(as.character(toupper(answer1.4))), "3a5505c06543876fe45598b5e5e5195d")
})
cat("success!")
```
## QUESTION 1.5
Output the `penguins` tibble so that `year` comes first. Hint: use the tidyselect `everything()` function. Store the result in a variable named `answer1.5`.
```
# answer1.5 <- select(penguins, FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.5)
test_that("Question 1.5", {
expect_equal(digest(dim(answer1.5)), "d095e682a86f7f16404b7f8dd5f3d676")
expect_equal(digest(answer1.5), "a07a1cdcb64726866df3d525811a9bf6")
})
cat("success!")
```
## QUESTION 1.6
Rename `flipper_length_mm` to `length_flipper_mm`. Store the result in a variable named `answer1.6`
```
# answer1.6 <- rename(FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.6)
test_that("Question 1.6", {
expect_equal(digest(dim(answer1.6)), 'd095e682a86f7f16404b7f8dd5f3d676')
expect_equal(digest(names(answer1.6)), 'ef6a2aaa40de41c0b11ad2f6888d5ce6')
})
cat("success!")
```
## `filter()`
So far, we've practiced picking variables by their name with `select()`. But how about picking observations (rows)? This is where `filter()` comes in.
## QUESTION 1.7
Pick penguins with body mass greater than 3600 g. Store the resulting tibble in a variable named `answer1.7`
```
# answer1.7 <- filter(FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.7)
test_that("Question 1.7", {
expect_equal(digest(dim(answer1.7)), '0f80c9cad929bf5de5ae34e0d50cb60d')
expect_equal(sum(pull(answer1.7, body_mass_g) <= 3600), 0)
})
cat("success!")
```
## Storing the subsetted penguins data
In question 1.7 above, you've created a subset of the `penguins` dataset by filtering for those penguins that have a body mass greater than 3600 g. Let's do a quick check to see how many penguins meet that threshold by comparing the dimensions of the `penguins` dataset and your subset, `answer1.7`. There are two different ways to do this.
```
dim(penguins)
dim(answer1.7)
```
As you can see, in filtering down to penguins with a body mass greater than 3600g, we have lost about 100 rows (observations). However, `answer1.7` doesn't seem like an informative name for this new dataset that you've created from `penguins`. Let's rename it to something else.
```
penguins3600 <- answer1.7
```
## QUESTION 1.8
From your "new" dataset `penguins3600`, take only data from penguins located in the Biscoe island. Store the result in a variable named `answer1.8`.
```
# answer1.8 <- filter(FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.8)
test_that("Question 1.8", {
expect_equal(digest(dim(answer1.8)), "92ac01cd2e8809faceb1f7a283cd935f")
a <- as.character(unique(pull(answer1.8, island)))
expect_length(a, 1L)
expect_equal(a, "Biscoe")
})
cat("success!")
```
## QUESTION 1.9
Repeat the task from Question 1.8, but take data from islands Torgersen and Dream. Now that you've practiced with dplyr verbs quite a bit, you don't need as many prompts to answer! Hint: When you want to select more than one island, you use `%in%` instead of `==`.
Store your answer in a variable named `answer1.9`.
```
# answer1.9 <- FILL_THIS_IN(FILL_THIS_IN, island FILL_THIS_IN c("FILL_THIS_IN", "FILL_THIS_IN"))
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.9)
test_that("Question 1.9", {
expect_equal(digest(dim(answer1.9)), "b207bbce54bb47be51e7ba7b56d24bc2")
expect_equal(sum(pull(answer1.9, island) == "Torgersen"), 28)
expect_equal(sum(pull(answer1.9, island) == "Dream"), 69)
expect_equal(sum(pull(answer1.9, island) == "Biscoe"), 0)
})
cat("success!")
```
## `arrange()`
`arrange()` allows you to rearrange rows. Let's give it a try!
## QUESTION 1.10
Order `penguins` by year, in ascending order. Store the resulting tibble in a variable named `answer1.10`.
```
# answer1.10 <- arrange(FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.10)
test_that("Question 1.10", {
expect_sorted(pull(answer1.10, year))
})
cat("success!")
```
## QUESTION 1.11
Great work! Order `penguins` by year, in descending order. Hint: there is a function that allows you to order a variable in descending order called `desc()`.
Store your tibble in a variable named `answer1.11`.
```
# answer1.11 <- arrange(FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.11)
test_that("Question 1.11", {
expect_sorted(pull(answer1.11, year) %>%
rev())
})
cat("success!")
```
## QUESTION 1.12
Order `penguins` by year, then by `body_mass_g`. Use ascending order in both cases.
Store your answer in a variable named `answer1.12`
```
# answer1.12 <- arrange(FILL_THIS_IN, FILL_THIS_IN, FILL_THIS_IN)
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.12)
test_that("Question 1.12", {
expect_sorted(pull(answer1.12, year))
answer1.12_list <- answer1.12 %>%
group_by(year) %>%
group_split()
expect_length(answer1.12_list, 3)
expect_sorted(answer1.12_list[[1]] %>% pull(body_mass_g) %>% na.omit())
expect_sorted(answer1.12_list[[2]] %>% pull(body_mass_g) %>% na.omit())
expect_sorted(answer1.12_list[[3]] %>% pull(body_mass_g) %>% na.omit())
})
cat("success!")
```
## Piping, `%>%`
So far, we've been using dplyr verbs by inputting the dataset that we want to work on as the first argument of the function (e.g. `select(**penguins**, year))`. This is fine when you're using a single verb, i.e. you only want to filter observations, or select variables. However, more often than not you will want to do several tasks at once; such as filtering penguins with a certain body mass, and simultaneously ordering those penguins by year. Here is where piping (`%>%`) comes in.
Think of `%>%` as the word "then"!
Let's see an example. Here I want to combine `select()` with `arrange()`.
This is how I could do it by *nesting* the two function calls. I am selecting variables year, species, island, and body_mass_g, while simultaneously arranging by year.
```
print(arrange(select(penguins, year, species, island, body_mass_g), year), n = 5)
```
However, that seems a little hard to read. Now using pipes:
```
penguins %>%
select(year, species, island, body_mass_g) %>%
arrange(year) %>%
print(n = 5)
```
## Creating tibbles
Throughout Part A, we have been working with a tibble, `penguins`. Remember that when we ran `class()` on `penguins`, we could see that it was a dataframe that had been coerced to a tibble, which is a unifying feature of the tidyverse.
Suppose that you have a dataframe that you want to coerce to a tibble. To do this, you can use `as_tibble()`. R comes with a few built-in datasets, one of which is `mtcars`. Let's check the class of `mtcars`:
```
class(mtcars)
```
As you can see, mtcars is a dataframe. Now, coerce it to a tibble with `as_tibble()`:
```
as_tibble(mtcars) %>%
print(n = 5)
```
You can read more about tibbles in the [R4DS Tibble Chapter](https://r4ds.had.co.nz/tibbles.html#creating-tibbles).
## QUESTION 1.13
At the start of this worksheet, we loaded a package called `gapminder`. This package comes with a dataset stored in the variable also named `gapminder`. Check the class of the `gapminder` dataset:
```
class(gapminder)
```
As you can see, it is already a tibble.
Take all countries in Europe that have a GDP per capita greater than 10000, and select all variables except `gdpPercap`, using pipes. (Hint: use `-`).
Store your answer in a variable named `answer1.13`. Here is a code snippet that you can copy and paste into the solution cell below.
```
answer1.13 <- FILL_THIS_IN %>%
filter(FILL_THIS_IN > 10000, FILL_THIS_IN == "Europe") %>%
FILL_THIS_IN(-FILL_THIS_IN)
```
```
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.13)
test_that("Question 1.13", {
expect_equal(digest(dim(answer1.13)), "87d72f02bf15a0a29647db0c48c9a226")
expect_equal(digest(answer1.13), "d0136991f3cfee4fcf896f677181c9c6")
})
cat("success!")
```
## QUESTION 1.14
Coerce the `mtcars` data frame to a tibble, and take all columns that start with the letter "d".
*Hint: take a look at the "Select helpers" documentation by running the following code: `?tidyselect::select_helpers`.*
Store your tibble in a variable named `answer1.14`
```
answer1.14 <- FILL_THIS_IN(FILL_THIS_IN) %>%
FILL_THIS_IN(FILL_THIS_IN("d"))
```
```
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.14)
test_that("Question 1.14", {
expect_equal(digest(dim(answer1.14)), "ea1df69d6a59227894d1d4330f9bfab8")
expect_equal(digest(colnames(answer1.14)), "0956954d01fe74c59c1f16850b7e874f")
})
cat("success!")
```
This exercise is from [r-exercises](https://www.r-exercises.com/2017/10/19/dplyr-basic-functions-exercises/).
## `mutate()`
The `mutate()` function allows you to create new columns, possibly using existing columns. Like `select()`, `filter()`, and `arrange()`, the `mutate()` function also takes a tibble as its first argument, and returns a tibble.
The general syntax is: `mutate(tibble, NEW_COLUMN_NAME = CALCULATION)`.
## QUESTION 1.15
Make a new column with body mass in kg, named `body_mass_kg`, *and* rearrange the tibble so that `body_mass_kg` goes after `body_mass_g` and before `sex`. Store the resulting tibble in a variable named `answer1.15`.
*Hint*: within `select()`, use R's `:` operator to select all variables from `species` to `body_mass_g`.
```
answer1.15 <- penguins %>%
mutate(FILL_THIS_IN = FILL_THIS_IN) %>%
select(FILL_THIS_IN, FILL_THIS_IN, FILL_THIS_IN, FILL_THIS_IN)
```
```
# Your code here
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.15)
test_that("Question 1.15", {
expect_equal(digest(dim(answer1.15)), "9e9457527d068c2333ea8fd598e07f13")
expect_equal(digest(colnames(answer1.15)), "d7121e41fe934232c1c45dc425365040")
expect_equal(na.omit(answer1.15$body_mass_kg / answer1.15$body_mass_g) %>% digest,
"cdfbfd4da65e3575a474558218939055")
})
cat("success!")
```
Notice the backwards compatibility! No need for loops! By the way, if you'd like to simultaneously create columns _and_ delete other columns, use the `transmute` function.
## `group_by()`
The `group_by()` function groups the _rows_ in your tibble according to one or more categorical variables. Just specify the columns containing the grouping variables. `mutate()` (and others) will now operate on each chunk independently.
## QUESTION 1.16
Calculate the growth in population since the first year on record _for each country_, and name the column `rel_growth`. Do this by **rearranging the following lines**, and **filling in the `FILL_THIS_IN`**. Assign your answer to a variable named `answer1.16`
*Hint*: Here's another convenience function for you: `dplyr::first()`.
```
answer1.16 <-
mutate(rel_growth = FILL_THIS_IN) %>%
arrange(FILL_THIS_IN) %>%
gapminder %>%
group_by(country) %>%
```
```
# Your code here
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.16)
test_that("Answer 1.16", {
expect_equal(nrow(answer1.16), 1704)
c('country', 'continent', 'year', 'lifeExp', 'pop', 'gdpPercap', 'rel_growth') %>%
map_lgl(~ .x %in% names(answer1.16)) %>%
all() %>%
expect_true()
expect_equal(digest(as.integer(answer1.16$rel_growth)), '26735e4b17481f965f9eb1d3b5de89ad')
})
cat("success!")
```
## `summarise()`
The last core dplyr verb is `summarise()`. It collapses a data frame to a single row:
```
summarise(penguins, body_mass_mean = mean(body_mass_g, na.rm = TRUE))
```
*From R4DS Data Transformation:*
> `summarise()` is not terribly useful unless we pair it with `group_by()`. This changes the unit of analysis from the complete dataset to individual groups. Then, when you use the dplyr verbs on a grouped data frame they'll be automatically applied "by group".
For example, if we applied exactly the same code to a tibble grouped by island, we get the average body mass per island:
```
penguins %>%
group_by(island) %>%
summarise(body_mass_mean = mean(body_mass_g, na.rm = TRUE))
```
## QUESTION 1.17
From the `penguins` tibble, calculate the mean penguin body mass per island by year, in a column named `body_mass_mean`. Your tibble should have the columns `year`, `island`, and `body_mass_mean` only (and in that order). Store the resulting tibble in a variable named `answer1.17`.
```
answer1.17 <- penguins %>%
group_by(FILL_THIS_IN) %>%
FILL_THIS_IN(body_mass_mean = mean(FILL_THIS_IN, na.rm = TRUE))
```
```
# Your code here
# your code here
fail() # No Answer - remove if you provide an answer
head(answer1.17)
test_that("Question 1.17", {
expect_equal(digest(dim(answer1.17)), "f4885de1726d18557bd43d769cc0ae26")
expect_equal(digest(colnames(answer1.17)), "ba0c85220a5fa5222cac937acb2f94c2")
})
cat("success!")
```
# Part 2: Scoped variants with `across()`
Sometimes we want to perform the same operation on many columns. We can achieve this by embedding the `across()` function within the `mutate()` or `summarise()` functions.
## QUESTION 2.0
In a single expression, make a tibble with the following columns *for each island* in the penguins data set:
+ What is the *mean* of each numeric variable in the `penguins` dataset in each island? Keep the column names the same.
+ How many penguins are there in each island? Add this to a column named `n`.
Assign your answer to a variable named `answer2.0`
```
answer2.0 <- penguins %>%
group_by(FILL_THIS_IN) %>%
summarise(across(where(FILL_THIS_IN), FILL_THIS_IN, na.rm = TRUE),
n = n())
```
```
# Your code here
# your code here
fail() # No Answer - remove if you provide an answer
head(answer2.0)
test_that("Answer 2.0", {
expect_equal(
answer2.0 %>%
mutate(across(where(is.numeric), round, digits = 0)) %>%
unclass() %>%
digest(),
"b06d7816762e489a57ca922d175f08ef"
)
})
cat("success!")
```
## QUESTION 2.1
Using the `penguins` dataset, what is the mean bill length and depth of penguins on each island, by year? The resulting tibble should have columns named `island`, `year`, `bill_length_mm`, and `bill_depth_mm`, in that order. Store the result in a variable named `answer2.1`. Be sure to remove NA's when you are calculating the mean.
*Hint*: Use `starts_with()` instead of `where()` in the `across()` function.
```
answer2.1 <- penguins %>%
group_by(FILL_THIS_IN) %>%
summarise(across(FILL_THIS_IN))
```
```
# Your code here
# your code here
fail() # No Answer - remove if you provide an answer
head(answer2.1)
test_that("Answer 2.1", {
expect_equal(names(answer2.1), c("island", "year", "bill_length_mm", "bill_depth_mm"))
sorted <- answer2.1 %>%
arrange(island, year)
expect_identical(digest(round(sorted$bill_length_mm, 0)), "f9f46fe0b2604eac7903505876e4b240")
expect_identical(digest(round(sorted$bill_depth_mm, 0)), "d54992e0dbb34479e18f4f73ff1f16f4")
})
cat("success!")
```
| github_jupyter |
```
#IMPORT SEMUA LIBRARY DISINI
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY POSTGRESQL
import psycopg2
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY PDF
from fpdf import FPDF
#IMPORT LIBRARY BASEPATH
import io
#IMPORT LIBRARY BASE64 IMG
import base64
#IMPORT LIBRARY NUMPY
import numpy as np
#IMPORT LIBRARY EXCEL
import xlsxwriter
#IMPORT LIBRARY SIMILARITAS
import n0similarities as n0
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
def uploadToPSQL(host, username, password, database, port, table, judul, filePath, name, subjudul, dataheader, databody):
#TEST KONEKSI KE DATABASE
try:
for t in range(0, len(table)):
#DATA DIJADIKAN LIST
rawstr = [tuple(x) for x in zip(dataheader, databody[t])]
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=database)
cursor = connection.cursor()
connection.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT);
#CEK TABLE
cursor.execute("SELECT * FROM information_schema.tables where table_name=%s", (table[t],))
exist = bool(cursor.rowcount)
#KALAU ADA DIHAPUS DULU, TERUS DICREATE ULANG
if exist == True:
cursor.execute("DROP TABLE "+ table[t] + " CASCADE")
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#KALAU GA ADA CREATE DATABASE
else:
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#MASUKAN DATA KE DATABASE YANG TELAH DIBUAT
cursor.execute('INSERT INTO '+table[t]+'(tanggal, total) values ' +str(rawstr)[1:-1])
#JIKA BERHASIL SEMUA AKAN MENGHASILKAN KELUARAN BENAR (TRUE)
return True
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
return error
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
def makeChart(host, username, password, db, port, table, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath):
try:
datarowsend = []
for t in range(0, len(table)):
#TEST KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBIL DATA DARI DATABASE DENGAN LIMIT YANG SUDAH DIKIRIMKAN DARI VARIABLE DIBAWAH
postgreSQL_select_Query = "SELECT * FROM "+table[t]+" ORDER BY tanggal DESC LIMIT " + str(limitdata)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MENYIMPAN DATA DARI DATABASE KE DALAM VARIABLE
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
lengthy.append(row[2])
datarowsend.append(mobile_records)
#JUDUL CHART
judulgraf = A2 + " " + wilayah[t]
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#DATA CHART DIMASUKAN DISINI
ax.bar(uid, lengthy, align='center')
#JUDUL CHART
ax.set_title(judulgraf)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#BUAT CHART MENJADI FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART DIJADIKAN BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#line
#DATA CHART DIMASUKAN DISINI
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#JUDUL CHART
plt.title(judulgraf)
plt.grid(True)
l = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(l, format='png', bbox_inches="tight")
#GAMBAR DIJADIKAN BAS64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#pie
#JUDUL CHART
plt.title(judulgraf)
#DATA CHART DIMASUKAN DISINI
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.plot(legend=None)
plt.axis('equal')
p = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(p, format='png', bbox_inches="tight")
#CHART DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#CHART DISIMPAN KE DIREKTORI DIJADIKAN FORMAT PNG
#BARCHART
bardata = base64.b64decode(barChart)
barname = basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[t]+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINECHART
linedata = base64.b64decode(lineChart)
linename = basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[t]+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIECHART
piedata = base64.b64decode(pieChart)
piename = basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[t]+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#MEMANGGIL FUNGSI EXCEL
makeExcel(datarowsend, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, name, limitdata, table, wilayah, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(datarowsend, judul, barChart, lineChart, pieChart, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, table, wilayah, basePath)
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
print (error)
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
def makePDF(datarow, judul, bar, line, pie, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, lengthPDF, table, wilayah, basePath):
#PDF DIATUR DENGAN SIZE A4 DAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#TAMBAH HALAMAN PDF
pdf.add_page()
#SET FONT DAN JUGA PADDING
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#TAMPILKAN JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#SET FONT DAN JUGA PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#TAMPILKAN SUB JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#BUAT GARIS DIBAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
pdf.set_font('Times','B',11.0)
pdf.ln(0.5)
th1 = pdf.font_size
#BUAT TABLE DATA DATA DI DPF
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, A2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Region", border=1, align='C')
pdf.cell(177, 2*th1, B2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Frekuensi", border=1, align='C')
pdf.cell(177, 2*th1, C2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Unit", border=1, align='C')
pdf.cell(177, 2*th1, D2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Sumber", border=1, align='C')
pdf.cell(177, 2*th1, E2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Status", border=1, align='C')
pdf.cell(177, 2*th1, F2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "ID Seri", border=1, align='C')
pdf.cell(177, 2*th1, G2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Kode SR", border=1, align='C')
pdf.cell(177, 2*th1, H2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Pertama", border=1, align='C')
pdf.cell(177, 2*th1, str(I2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Terakhir ", border=1, align='C')
pdf.cell(177, 2*th1, str(J2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Waktu pembaruan terakhir", border=1, align='C')
pdf.cell(177, 2*th1, str(K2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.set_xy(17.0, 125.0)
pdf.set_font('Times','B',11.0)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
pdf.ln(0.5)
th = pdf.font_size
#HEADER TABLE DATA F2
pdf.cell(col_width, 2*th, str("Wilayah"), border=1, align='C')
#TANGAL HEADER DI LOOPING
for row in datarow[0]:
pdf.cell(col_width, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#ISI TABLE F2
for w in range(0, len(table)):
data=list(datarow[w])
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(col_width, 2*th, wilayah[w], border=1, align='C')
#DATA BERDASARKAN TANGGAL
for row in data:
pdf.cell(col_width, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#PEMANGGILAN GAMBAR
for s in range(0, len(table)):
col = pdf.w - 2*pdf.l_margin
pdf.ln(2*th)
widthcol = col/3
#TAMBAH HALAMAN
pdf.add_page()
#DATA GAMBAR BERDASARKAN DIREKTORI DIATAS
pdf.image(basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[s]+'-bar.png', link='', type='',x=8, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[s]+'-line.png', link='', type='',x=103, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/img/'+name+''+table[s]+'-pie.png', link='', type='',x=195, y=80, w=widthcol)
pdf.ln(4*th)
#PDF DIBUAT
pdf.output(basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/pdf/'+A2+'.pdf', 'F')
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
def makeExcel(datarow, A2, B2, C2, D2, E2, F2, G2, H2, I2, J2, K2, name, limit, table, wilayah, basePath):
#BUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/CEIC/22. Sektor Besi dan Baja/excel/'+A2+'.xlsx')
#BUAT WORKSHEET EXCEL
worksheet = workbook.add_worksheet('sheet1')
#SETTINGAN UNTUK BORDER DAN FONT BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#HEADER UNTUK TABLE EXCEL F2
header = ["Wilayah", "Kategori","Region","Frekuensi","Unit","Sumber","Status","ID Seri","Kode SR","Tanggal Obs. Pertama","Tanggal Obs. Terakhir ","Waktu pembaruan terakhir"]
#DATA DATA DITAMPUNG PADA VARIABLE
for rowhead2 in datarow[0]:
header.append(str(rowhead2[1]))
#DATA HEADER DARI VARIABLE DIMASUKAN KE SINI UNTUK DITAMPILKAN BERDASARKAN ROW DAN COLUMN
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
#DATA ISI TABLE F2 DITAMPILKAN DISINI
for w in range(0, len(table)):
data=list(datarow[w])
body = [wilayah[w], A2, B2, C2, D2, E2, F2, G2, H2, str(I2.date()), str(J2.date()), str(K2.date())]
for rowbody2 in data:
body.append(str(rowbody2[2]))
for col_num, data in enumerate(body):
worksheet.write(w+1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
#BASE PATH UNTUK NANTINYA MENGCREATE FILE ATAU MEMANGGIL FILE
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE SIMILARITY WILAYAH
filePathwilayah = basePath+'data mentah/CEIC/allwilayah.xlsx';
#BACA FILE EXCEL DENGAN PANDAS
readexcelwilayah = pd.read_excel(filePathwilayah)
dfwilayah = list(readexcelwilayah.values)
readexcelwilayah.fillna(0)
allwilayah = []
#PEMILIHAN JENIS DATA, APA DATA ITU PROVINSI, KABUPATEN, KECAMATAN ATAU KELURAHAN
tipewilayah = 'prov'
if tipewilayah == 'prov':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][1])
elif tipewilayah=='kabkot':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][3])
elif tipewilayah == 'kec':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][5])
elif tipewilayah == 'kel':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][7])
semuawilayah = list(set(allwilayah))
#SETTING VARIABLE UNTUK DATABASE DAN DATA YANG INGIN DIKIRIMKAN KE FUNGSI DISINI
name = "04. Statistik Perdagangan Besi dan Baja (WAD001-WAD016) Part 1"
host = "localhost"
username = "postgres"
password = "1234567890"
port = "5432"
database = "ceic"
judul = "Produk Domestik Bruto (AA001-AA007)"
subjudul = "Badan Perencanaan Pembangunan Nasional"
filePath = basePath+'data mentah/CEIC/22. Sektor Besi dan Baja/'+name+'.xlsx';
limitdata = int(8)
readexcel = pd.read_excel(filePath)
tabledata = []
wilayah = []
databody = []
#DATA EXCEL DIBACA DISINI DENGAN MENGGUNAKAN PANDAS
df = list(readexcel.values)
head = list(readexcel)
body = list(df[0])
readexcel.fillna(0)
#PILIH ROW DATA YANG INGIN DITAMPILKAN
rangeawal = 106
rangeakhir = 107
rowrange = range(rangeawal, rangeakhir)
#INI UNTUK MEMFILTER APAKAH DATA YANG DIPILIH MEMILIKI SIMILARITAS ATAU TIDAK
#ISIKAN 'WILAYAH' UNTUK SIMILARITAS
#ISIKAN BUKAN WILAYAH JIKA BUKAN WILAYAH
jenisdata = "Indonesia"
#ROW DATA DI LOOPING UNTUK MENDAPATKAN SIMILARITAS WILAYAH
#JIKA VARIABLE JENISDATA WILAYAH AKAN MASUK KESINI
if jenisdata == 'Wilayah':
for x in rowrange:
rethasil = 0
big_w = 0
for w in range(0, len(semuawilayah)):
namawilayah = semuawilayah[w].lower().strip()
nama_wilayah_len = len(namawilayah)
hasil = n0.get_levenshtein_similarity(df[x][0].lower().strip()[nama_wilayah_len*-1:], namawilayah)
if hasil > rethasil:
rethasil = hasil
big_w = w
wilayah.append(semuawilayah[big_w].capitalize())
tabledata.append('produkdomestikbruto_'+semuawilayah[big_w].lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#JIKA BUKAN WILAYAH MASUK KESINI
else:
for x in rowrange:
wilayah.append(jenisdata.capitalize())
tabledata.append('produkdomestikbruto_'+jenisdata.lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#HEADER UNTUK PDF DAN EXCEL
A2 = "Data Migas"
B2 = df[rangeawal][1]
C2 = df[rangeawal][2]
D2 = df[rangeawal][3]
E2 = df[rangeawal][4]
F2 = df[rangeawal][5]
G2 = df[rangeawal][6]
H2 = df[rangeawal][7]
I2 = df[rangeawal][8]
J2 = df[rangeawal][9]
K2 = df[rangeawal][10]
#DATA ISI TABLE F2
dataheader = []
for listhead in head[11:]:
dataheader.append(str(listhead))
#FUNGSI UNTUK UPLOAD DATA KE SQL, JIKA BERHASIL AKAN MAMANGGIL FUNGSI UPLOAD CHART
sql = uploadToPSQL(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, dataheader, databody)
if sql == True:
makeChart(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath)
else:
print(sql)
```
| github_jupyter |
# [LEGALST-123] Lab 21: Neural Nets
In this lab, we'll learn how to implement neural net methods to pattern recognize handwriting images. In this lab, we'll learn how to implement neural net methods to pattern recognize handwriting images. Due to the memory constraints of DataHub, we need to download this lab folder and run it locally.
Due to the memory constraints of DataHub, we will use Colab by clicking the badge below:
[](https://drive.google.com/file/d/1oJPNiA-lzHoyrQfgPaY_1xkyBulOgtos/view?usp=sharing)
Make sure to set runtime type to "GPU".
<img src="./imgs/gpu_runtime.png">
*Estimated Time: 30-40 minutes*
---
### Table of Contents
[The Data](#section data)<br>
1 - [Visualizing Data](#section 1)<br>
2 - [Neural Network](#section 2)<br>
3 - [Multi-Layer Perceptrons](#section 3)<br>
4 - [A Simple MLP](#section 4)<br>
5 - [Convolutional Nerual Networks](#section 5)<br>
**Dependencies:**
```
!pip install tensorflow keras
```
---
## The Data <a id='data'></a>
In this notebook, you'll be working with the MNIST handwriting dataset, considered "hello, world" of object recognition in machine learning. It contains images of handwritten digits centered and normalized. Modified NIST (National Institute of Standards and Technology) is constructed from scanned documents available from NIST.
---
## Visualizing Data <a id='section 1'></a>
The Keras deep learning library provides a convenience method for loading the MNIST dataset.
```
from keras.datasets import mnist
import matplotlib.pyplot as plt
# load the MNIST dataset
(X_train, y_train), (X_test, y_test) = mnist.load_data()
```
As you can see above, the dataset is split into a set to train our model and one to test it. X_train and X_test are inputs while y_train and y_test are outputs.
Let's visualize MNIST dataset by running the cell below.
```
# plot 4 images as gray scale
plt.subplot(221)
plt.imshow(X_train[0], cmap=plt.get_cmap('gray'))
plt.subplot(222)
plt.imshow(X_train[1], cmap=plt.get_cmap('gray'))
plt.subplot(223)
plt.imshow(X_train[2], cmap=plt.get_cmap('gray'))
plt.subplot(224)
plt.imshow(X_train[3], cmap=plt.get_cmap('gray'))
# show the plot
plt.show()
```
---
## Neural Network <a id='section 2'></a>
Neural networks are a set of algorithms, modeled loosely after the human brain, that are designed to recognize patterns.
Neural networks help us cluster and classify. They group unlabeled data according to similarities among the example inputs.
Classification problems depend on labeled datasets, i.e. humans need to label the data for a neural to learn the correlation between labels and data. This is <i>supervised learning</i>. Below, we'll dive into MLP, which utilizes a supervised learning technique.
## Multi-Layer Perceptrons <a id='section 3'></a>
A multi-layer perceptron (MLP) is a class of neural network that consists of at least three layers of nodes (first layer being inputs and last layer being outputs). Except for the input layer, activation of nodes of a certain layer depends on which nodes are activated in the previous layer. Each node is a neuron that uses a nonlinear activation function. Below is an image that represents a MLP.
<img src="https://www.safaribooksonline.com/library/view/getting-started-with/9781786468574/graphics/B05474_04_05.jpg" style="width: 500px;"/>
### MLP
A multilayer perceptron (MLP) is a deep, artificial neural network. They are composed of an input layer to receive the signal, an output layer that makes a decision or prediction about the input, and in between those two, an arbitrary number of hidden layers that are the true computational engine of the MLP
MLPs train on a set of input-output pairs and learn to model the correlation between those inputs and outputs by adjusting parameters to minimize error.
## A Simple MLP <a id='section 4'></a>
Let's try to build a simple MLP to identify digits.
### Input vs. Output
Each pixel of an image will be an input to our MLP.
<b>Question:</b> How many nodes would our MLP's first layer have for a 20x20 image?
<i>Your answer here</i>
The output layer will produce what we are looking for.
<b>Question:</b> What will each node in the output layer represent?
<i>Your answer here</i>
### Fixing MNIST
The training dataset is a 3-D array of instance, image width, image height. To make an image "inputable" to our MLP, we need to vectorize the representations of MNIST training dataset, i.e. we need to flatten a image into a vector.
```
num_pixels = X_train.shape[1] * X_train.shape[2]
num_pixels
# NumPy's reshape function can help flatten multi-dimensional arrays
# As all images are 28x28 pixels, we flatten all into vectors of length 784
X_train = X_train.reshape(X_train.shape[0], num_pixels)
X_test = X_test.reshape(X_test.shape[0], num_pixels)
```
The pixel values are grey scale, which ranges from 0-255. Neural network models work better with normalized inputs. Thus, normalize our training dataset below so that values range from 0-1.
```
X_train = X_train / ...
X_test = X_test / ...
```
The outputs (y_train and y_test) are integers from 0 to 9 and we can think fo each integer as a class. We'll use a one-hot encoding of the class values, which transforms the output vector into a binary matrix.
<i>One-hot encoding is a process by which categorical variables are converted into a form that could be provided to ML algorithms to do a better job in prediction.</i>
```
# use built-in function in np_utils
from keras.utils import np_utils
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
```
Just to make sure that we still have 10 classes(0-9) after one-hot encoding.
```
num_classes = y_test.shape[1]
num_classes
```
### Create the Model
Keras has useful methods we can utilize to develop an MLP.
```
# importing the methods we need
from keras.models import Sequential
from keras.layers import Dense
```
Create an empty MLP, i.e. empty linear stack of layers
```
model = Sequential()
```
Create an input layer using the Dense function (refer to <a href="https://keras.io/layers/core/">this doc</a>), then add the layer to our model using `model.add(...)`. The input layer should use 'relu' activation, a 'normal' kernel_initializer, and the number of pixels for input_dim.
```
"""
Dense(...) creates a regular densely connected layer.
relu is a rectifier activation function.
"""
...
```
Create and add in the output layer to our model. Use 'softmax' activation and a normal kernel_initializer.
```
"""
A softmax activation function is used on the output layer to turn the outputs into
probability-like values
"""
...
```
Compile our model.
```
"""
categorical_crossentropy is a logarithmic loss function and
adam is a gradient descent algorithm
"""
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
```
Let's now fit our model on the training datasets. We fit our model over 10 epochs and update it every 200 images. It might take a few minutes.
```
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=10, batch_size=200, verbose=2)
```
Comparing the log of our training produced above, what can you conclude?
<i>Your Answer Here</i>
Let's evaluate our model on the test dataset.
```
scores = model.evaluate(X_test, y_test, verbose=0)
print("Error: %.2f%%" % (100-scores[1]*100))
```
## Convolutional Nerual Networks <a id='section 5'></a>
MLPs did give us pretty good accuracy. However, as each pixel of an image is an input, we will have too many inputs for a large image. Convolutional neural nets, on the other hand, can take in 3-D inputs (2D + color) instead of just 1-D.
### How a CNN works <a id='conv_image'></a>
<img src="https://www.mathworks.com/content/mathworks/www/en/discovery/convolutional-neural-network/jcr:content/mainParsys/image_copy.adapt.full.high.jpg/1517522275430.jpg" style="width: 600px;"/>
#### Filtering
ConvNets have "features" and match them with parts of an image rather than the whole thing.
<img src="./imgs/match_pieces.png" style="width: 500px;"/>
In the example above, we have three features: left-to-right diagonal, central X, and right to left diagonal. To match features, we use a process called filtering.
1. Assign each black pixel -1 and each white pixel 1.
2. Line up a feature with an image patch (for example, line up left-to-right diagonal with the green box above)
3. Multiply each feature pixel with the corresponding image patch pixel
4. Take the average of the products
<b>Question:</b> In the example above when we match the left-to-right diagonal feature with the green box, what would the process output?
<i>Your Answer Here</i>
#### Convolutional Layer
The process of filtering for every possible image patch with every feature.
<img src="./imgs/convolution.png" style="width: 500px;"/>
As you can see in the image above, after applying our left-to-right diagonal filter, we get higher scores on the left to right diagonal.
<b>Question:</b> If we apply the central X filter in this example, where will the highest score occur?
<i>Your Answer Here</i>
#### ReLU (Rectified Linear Units)
This process is applied on filtered images. It simply changes every negative value to 0 and leaves positive values unchanged.
#### Pooling Layer
In this layer, we shrink the filtered images by the following process:
1. Pick a window size and a stride size
2. Walk our window on the filtered image, each time shifting by the stride size
3. For each step, take the maximum score contained in the window
<img src="./imgs/pooling.png" style="width: 500px;"/>
This layer helps because it does not care about where in the window the maximum value occurs, i.e. it's less sensitive to specific positioning of pixels.
<b>Question:</b> How would the pooling layer help in classifying digits?
<i>Your Answer Here</i>
#### Fully Connected Layer
This layer flattens the pooled images and each value gets a vote, which is how strongly that value suggests a certain outcome.
One question you might have now is where do the initial features and voting weights come from. They are obtained by a process called <a href="https://brilliant.org/wiki/backpropagation/">backpropragation</a>, explained in more detail at the link.
Now we've learned that the layers are combined together: <b>convolutional + ReLU + pooling</b> constitute the first part while <b>fully connected</b> the second. One note is that each part can be applied multiple times, as we can see in [the overview of a CNN at the beginning](#conv_image).
### Create a CNN
Keras also provides useful methods to create a CNN.
```
# importing dependencies
import numpy
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.utils import np_utils
from keras import backend as K
K.set_image_data_format('channels_first')
```
In Keras, the layers used for two-dimensional convolutions expect pixel values with the dimensions [pixels][width][height]. We'll set the pixels dimension to 1 because the pixel values in MNIST are gray scale. As a reminder, all images in MNIST are 28x28.
```
# load data
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# reshape to [samples][pixels][width][height]
X_train = X_train.reshape(X_train.shape[0], ..., ..., ...)
X_test = X_test.reshape(X_test.shape[0], ..., ..., ...)
```
Below, fill out the blanks to normalize the datasets and one hot encode the output values.
```
# normalize inputs from 0-255 to 0-1
X_train = X_train / ...
X_test = X_test / ...
# one hot encode outputs
y_train = ...
y_test = ...
num_classes = y_test.shape[1]
```
Let's define a function that creates our CNN. Read through the comments to understand what each line does.
```
def CNN_model():
# create model
model = Sequential()
# A convolutional layer that has 32 features of size 5x5
model.add(Conv2D(32, (5, 5), input_shape=(1, 28, 28), activation='relu'))
# A pooling layer with a window size of 2x2
model.add(MaxPooling2D(pool_size=(2, 2)))
# A dropout layer that randomly excludes 20% of neurons in the layer
model.add(Dropout(0.2))
# A flatten layer
model.add(Flatten())
# A fully connected layer with 128 neurons
model.add(Dense(128, activation='relu'))
# An output layer with softmax as in MLP
model.add(Dense(num_classes, activation='softmax'))
# Compile model as before in MLP
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
model = CNN_model()
# Fit the model
model.fit(..., ..., validation_data=(..., ...), epochs=10, batch_size=200, verbose=2)
# Evaluation of the model
scores = model.evaluate(..., ..., verbose=0)
print("CNN Error: %.2f%%" % (100-scores[1]*100))
```
Now let's experiment with the parameters: number of features, size of features, size of the window in pooling, dropout percentage, and so on. Modify the parameters to your best judgment in the function below.
```
def diff_CNN_model():
# create model
model = Sequential()
# A convolutional layer that has 32 features of size 5x5
model.add(Conv2D(32, (5, 5), input_shape=(1, 28, 28), activation='relu'))
# A pooling layer with a window size of 2x2
model.add(MaxPooling2D(pool_size=(2, 2)))
# A dropout layer that randomly excludes 20% of neurons in the layer
model.add(Dropout(0.2))
# A flatten layer
model.add(Flatten())
# A fully connected layer with 128 neurons
model.add(Dense(128, activation='relu'))
# An output layer with softmax as in MLP
model.add(Dense(num_classes, activation='softmax'))
# Compile model as before in MLP
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
model = diff_CNN_model()
# Fit the model
model.fit(..., ..., validation_data=(..., ...), epochs=10, batch_size=200, verbose=2)
# Evaluation of the model
scores = model.evaluate(..., ..., verbose=0)
print("diff_CNN Error: %.2f%%" % (100-scores[1]*100))
```
Do you get different accuracies for the two different models? Make some conclusions about different parameters.
<i>Your Answer Here</i>
### Bibliography
Image source:
- How Convolutional Neural Networks work, https://www.youtube.com/watch?v=FmpDIaiMIeA&t=1070s
Code source:
- Handwritten Digit Recognition using Convolutional Neural Networks in Python with Keras, https://machinelearningmastery.com/handwritten-digit-recognition-using-convolutional-neural-networks-python-keras/
---
Notebook developed by: Tian Qin
Data Science Modules: http://data.berkeley.edu/education/modules
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import rosbag
import pymap3d as pm
import numba as nb
from scipy.signal import savgol_filter
%matplotlib inline
def wrap_angle(angle):
return (angle + np.pi) % (2 * np.pi) - np.pi
@nb.njit()
def to_euler(x, y, z, w):
"""Dari Coursera: Return as xyz (roll pitch yaw) Euler angles."""
roll = np.arctan2(2 * (w * x + y * z), 1 - 2 * (x**2 + y**2))
pitch = np.arcsin(2 * (w * y - z * x))
yaw = np.arctan2(2 * (w * z + x * y), 1 - 2 * (y**2 + z**2))
return np.array([roll, pitch, yaw])
# Compile the to_euler
_ = to_euler(1.5352300785980803e-15, -1.3393747145983517e-15, -0.7692164172827881, 0.638988343698562)
class get_data_from_bag(object):
def __init__(self, path):
self.bag = rosbag.Bag(path)
self.cs = self._read_msg_from_topic('/control_signal', ['t', 'action_throttle', 'action_steer',
'error_lateral', 'error_yaw','error_speed',
'actual_x','actual_y','actual_yaw','actual_speed',
'ref_x', 'ref_y', 'ref_yaw', 'ref_curvature', 'ref_speed',
'wp_idx', 'deg_ref_yaw', 'deg_actual_yaw', 'deg_error_yaw'])
self.ar = self._read_msg_from_topic('/logging_arduino', ['t', 'steering_setpoint', 'steering_angle', 'throttle_voltage'])
self.gnss = self._read_gnss()
self.imu = self._read_imu('/imu', ['t', 'yaw'])
self.ekf = self._read_msg_from_topic('/state_2d_new', ['t', 'yaw', 'yaw_imu'])
def _read_msg_from_topic(self, topic, columns_name):
data = []
for _, msg, _ in self.bag.read_messages(topics=[topic]):
temp = []
for name in columns_name:
if name == 't':
temp.append(msg.header.stamp.to_sec())
else:
nm = 'msg.' + name
temp.append(eval(nm))
data.append(temp)
return pd.DataFrame(data, columns = columns_name)
def _read_gnss(self):
lat0, lon0, h0 = -6.8712, 107.5738, 768
data = []
for _, msg, _ in self.bag.read_messages(topics='/fix'):
temp = []
temp.append(msg.header.stamp.to_sec())
pos = pm.geodetic2enu(msg.latitude, msg.longitude, msg.altitude, lat0, lon0, h0)
temp.append(pos[0])
temp.append(pos[1])
temp.append(pos[2])
temp.append(msg.position_covariance[0])
data.append(temp)
return pd.DataFrame(data, columns=['t', 'x', 'y', 'z', 'cov_x'])
def _read_imu(self, topic, columns_name):
data = []
for _, msg, _ in self.bag.read_messages(topics=[topic]):
temp = []
for name in columns_name:
if name == 't':
temp.append(msg.header.stamp.to_sec())
elif name == 'yaw':
q = msg.orientation
euler = to_euler(q.x, q.y, q.z, q.w)
temp.append(euler[-1])
else:
nm = 'msg.' + name
temp.append(eval(nm))
data.append(temp)
return pd.DataFrame(data, columns = columns_name)
# df = get_data_from_bag('bag/LURUS_1.bag')
df = get_data_from_bag('bag/LURUS_EKF_GPS_DIJADIIN_COMPAS.bag')
```
# TAMBAHAN
```
num_f = 51
dst = 0.1
X = np.copy(df.gnss.x)
Y = np.copy(df.gnss.y)
x = np.copy(df.gnss.x)
y = np.copy(df.gnss.y)
t = np.copy(df.gnss.t)
XX = np.copy(df.cs.ref_x)
YY = np.copy(df.cs.ref_y)
wp_x = [X[0]]
wp_y = [Y[0]]
wp_xx = [XX[0]]
wp_yy = [YY[0]]
wp_t = [t[0]]
for i in range(1, X.shape[0]):
dist = np.sqrt((X[i] - wp_x[-1])**2 + (Y[i] - wp_y[-1])**2)
ddist = np.sqrt((XX[i] - wp_xx[-1])**2 + (YY[i] - wp_yy[-1])**2)
while dist >= dst:
# if dist >= dst:
wp_x.append(wp_x[-1] + dst*(X[i] - wp_x[-1])/dist)
wp_y.append(wp_y[-1] + dst*(Y[i] - wp_y[-1])/dist)
wp_t.append(wp_t[-1] + dst*(t[i] - wp_t[-1])/dist)
wp_xx.append(wp_xx[-1] + dst*(XX[i] - wp_xx[-1])/ddist)
wp_yy.append(wp_yy[-1] + dst*(YY[i] - wp_yy[-1])/ddist)
dist = np.sqrt((X[i] - wp_x[-1])**2 + (Y[i] - wp_y[-1])**2)
ddist = np.sqrt((XX[i] - wp_xx[-1])**2 + (YY[i] - wp_yy[-1])**2)
wp_x = np.array(wp_x)
wp_y = np.array(wp_y)
wp_x_f = savgol_filter(wp_x, num_f, 3)
wp_y_f = savgol_filter(wp_y, num_f, 3)
wp_xx = np.array(wp_xx)
wp_yy = np.array(wp_yy)
wp_ref_yaw = np.zeros_like(wp_x)
diffx = wp_xx[2:] - wp_xx[:-2]
diffy = wp_yy[2:] - wp_yy[:-2]
wp_ref_yaw[1:-1] = np.arctan2(diffy, diffx)
wp_ref_yaw[0] = wp_ref_yaw[1]
wp_ref_yaw[-1] = wp_ref_yaw[-2]
wp_ref_yaw_f = wrap_angle(savgol_filter(np.unwrap(wp_ref_yaw), num_f, 3))
act_ref_yaw_dydx = np.copy(wp_ref_yaw)
act_ref_yaw_dydx_f = np.copy(wp_ref_yaw_f)
wp_yaw = np.zeros_like(wp_x)
diffx = wp_x[2:] - wp_x[:-2]
diffy = wp_y[2:] - wp_y[:-2]
wp_yaw[1:-1] = np.arctan2(diffy, diffx)
wp_yaw[0] = wp_yaw[1]
wp_yaw[-1] = wp_yaw[-2]
wp_yaw_f = wrap_angle(savgol_filter(np.unwrap(wp_yaw), num_f, 3))
act_yaw_dydx = np.copy(wp_yaw)
act_yaw_dydx_f = np.copy(wp_yaw_f)
s = np.zeros(wp_x.shape[0])
for i in range(1, s.shape[0]):
s[i] = s[i-1] + np.sqrt((wp_x[i] - wp_x[i-1])**2 + (wp_y[i] - wp_y[i-1])**2)
width = 15
height = 15
plt.figure(figsize=(width, height))
plt.subplot(1,2,1)
plt.plot(wp_x, wp_y, label='Processed')
plt.scatter(x, y, color='red',s=2., label='RAW')
plt.xlabel("X (m)")
plt.ylabel("Y (m)")
plt.legend()
plt.title("PATH")
plt.subplot(1,2,2)
plt.plot(s, wp_yaw*180./np.pi)
plt.plot(s, wp_yaw_f*180./np.pi, label='post filtered')
plt.title("YAW")
plt.xlabel('s (m)')
plt.ylabel(r'\degree')
plt.legend()
#plt.savefig('waypoints.png', dpi=600, transparent=True)
plt.show()
act_yaw_dydx_interp = np.interp(df.imu.t, wp_t, act_yaw_dydx)
act_yaw_dydx_f_interp = np.interp(df.imu.t, wp_t, act_yaw_dydx_f)
act_ref_yaw_interp = np.interp(df.imu.t, wp_t, act_ref_yaw_dydx)
act_ref_yaw_f_interp = np.interp(df.imu.t, wp_t, act_ref_yaw_dydx_f)
ekf_yaw = np.interp(df.imu.t, df.cs.t, wrap_angle(df.cs.actual_yaw))
yaw_gnss = np.zeros_like(df.gnss.x.values)
n = 2
diffx = df.gnss.x.values[n:] - df.gnss.x.values[:-n]
diffy = df.gnss.y.values[n:] - df.gnss.y.values[:-n]
yaw_gnss[n:] = np.arctan2(diffy, diffx)
yaw_gnss[:n] = yaw_gnss[n]
yaw_gnss = np.interp(df.imu.t, df.gnss.t, yaw_gnss)
# plt.plot(df.gnss.t-df.gnss.t[0], yaw_gnss*180./np.pi, label='yaw gnss')
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
# plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
plt.plot(df.imu.t - df.imu.t[0], yaw_gnss*180./np.pi, label='gnss dy dx')
# plt.xlim(10., 33.)
plt.xlabel("Waktu (s)")
plt.ylabel(r"Yaw (\degree)")
plt.legend()
# plt.savefig('gt_vs_compass.png', dpi=600)
# plt.ylim(-180., 180.)
plt.show()
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
plt.xlabel("Waktu (s)")
plt.ylabel(r"Yaw ($\degree$)")
plt.legend()
plt.ylim(-180., 180.)
plt.savefig('pakai ekf/profil_yaw.png', dpi=600)
plt.show()
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
plt.xlabel("Waktu (s)")
plt.ylabel(r"Yaw ($\degree$)")
plt.legend()
plt.ylim(-120., -60.)
plt.xlim(15., 20.)
plt.savefig('pakai ekf/profil_yaw_zoom.png', dpi=600)
plt.show()
plt.plot(df.cs.ref_x, df.cs.ref_y, label='ref')
plt.plot(df.cs.actual_x, df.cs.actual_y, label='aktual')
# plt.scatter(df.gnss.x,df.gnss.y, color='black', s=1.0)
plt.axis('square')
plt.legend()
plt.xlabel("X (m)")
plt.ylabel("Y (m)")
plt.savefig('pakai ekf/posisi.png', dpi=600)
plt.show()
plt.plot(df.cs.t - df.cs.t[0], df.cs.error_yaw, label='galat yaw (rad)')
plt.plot(df.cs.t - df.cs.t[0], df.cs.error_lateral, label='galat lateral (m)')
plt.legend()
plt.xlabel("Waktu (s)")
plt.savefig('pakai ekf/galat.png', dpi=600)
plt.show()
plt.plot(df.cs.t - df.ar.t[0], df.cs.action_steer, label='steering setpoint')
plt.plot(df.ar.t - df.ar.t[0], df.ar.steering_angle, label='steering aktual')
plt.legend()
plt.xlabel("Waktu (s)")
plt.ylabel(r'Setir ($\degree$)')
plt.savefig('pakai ekf/sudut_kemudi.png', dpi=600)
plt.show()
plt.plot(df.cs.deg_ref_yaw)
plt.plot(180/np.pi*np.ones_like(df.cs.deg_ref_yaw)*np.arctan2(df.cs.actual_y.values[-1]-df.cs.actual_y.values[0], df.cs.actual_x.values[-1]-df.cs.actual_x.values[0]))
plt.scatter(act_yaw_dydx_interp*180./np.pi, df.imu.yaw*180./np.pi, s=1.)
plt.xlabel(r"ground truth $(\degree)$")
plt.ylabel(r"compass $(\degree)$")
plt.axis('square')
plt.legend()
plt.show()
# plt.savefig('ground_truth_vs_compass.png', dpi=600)
plt.scatter(act_yaw_dydx_f_interp*180./np.pi, df.imu.yaw*180./np.pi, s=0.5)
plt.xlabel(r"ground truth $(\degree)$")
plt.ylabel(r"compass $(\degree)$")
plt.axis('square')
plt.legend()
plt.savefig('pakai ekf/cek_bias.png', dpi=600)
plt.show()
# plt.plot(df.cs.t-df.gnss.t[0], df.cs.actual_speed)
# plt.xlim(8.)
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
plt.xlabel("Waktu (s)")
plt.ylabel(r"Yaw ($\degree$)")
plt.legend()
plt.ylim(-180., 180.)
# plt.savefig('pakai ekf/profil_yaw.png', dpi=600)
plt.show()
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
plt.plot(wp_t - df.gnss.t[0], wrap_angle(act_yaw_dydx_f)*180./np.pi, label='ground truth')
# plt.plot(df.imu.t-df.gnss.t[0], wrap_angle(df.imu.yaw + np.pi/2)*180./np.pi, label='Compass')
plt.plot(df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw)*180./np.pi, label='dy dx')
yaw_actual_interp = np.interp(wp_t - df.gnss.t[0], df.cs.t-df.gnss.t[0], wrap_angle(df.cs.actual_yaw))
err = wrap_angle(act_yaw_dydx_f - yaw_actual_interp)
np.sqrt(np.mean(err**2)) * 180 / np.pi
```
| github_jupyter |
# How to copy a QComponent
Start by importing QisKit Metal:
```
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict #, open_docs
```
Then let's fire up the GUI:
```
design = designs.DesignPlanar()
gui = MetalGUI(design)
```
Now we'll put one transmon at the origin:
```
# Let's start by putting a transmon at the origin:
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
design.overwrite_enabled = True
q1 = TransmonPocket(design, 'Q1')
gui.rebuild()
gui.autoscale()
```
First, we'll copy the component and then manually modify the coordinates to be at (1,0) instead of (0,0):
```
# Now let's copy the transmon at the origin and put it at position (0,1):
q1_copy = design.copy_qcomponent(q1, 'Q1_copy')
q1_copy.options['pos_x']='1.0mm'
gui.rebuild()
gui.autoscale()
```
Now, we'll copy the original component and in the same step we'll move the copy to (-1,0) by passing a dictionary to the "copy_qcomponent" command:
```
# Now let's copy the transmon at the origin and put it at position (0,-1):
q1_anothercopy = design.copy_qcomponent(q1,'Q1_another_copy', dict(pos_x='-1.0mm'))
gui.rebuild()
gui.autoscale()
```
We can copy multiple components at once using the "copy_multiple_qcomponents" command. Here's an example that copies the three components we've just created (Q1 and the two copies: Q1_copy and Q1_another_copy) and moves them up by 2.0mm each:
```
# Now let's copy all three components at the same time, moving them up by +2mm in the y-direction:
newcopies = design.copy_multiple_qcomponents([q1, q1_copy, q1_anothercopy], ['Q3', 'Q4', 'Q5'], [dict(pos_y='1.0mm'), dict(pos_y='2.0mm'), dict(pos_y='3.0mm')])
gui.rebuild()
gui.autoscale()
```
Note that we can also copy multiple qcomponents without passing dictionaries. In this case, the copied qcomponents will sit on top of of the original qcomponents in the layout:
```
# Example of copying without giving any dictionary values; qcomponents will sit on top of the originals!
newcopies2 = design.copy_multiple_qcomponents([q1, q1_copy, q1_anothercopy], ['Q6', 'Q7', 'Q8'])
gui.rebuild()
gui.autoscale()
```
Note also that we can copy multiple qcomponents but only give a dictionary to one of them. The other two dictionaries still need to exist but can be empty:
```
# Copy the three original components but only give a dictionary for the first one; other two dictionaries still need to exist but can be empty:
newcopies3 = design.copy_multiple_qcomponents([q1, q1_copy, q1_anothercopy], ['Q9', 'Q10', 'Q11'], [dict(pos_y='-1.0mm'), dict(), dict()])
gui.rebuild()
gui.autoscale()
```
| github_jupyter |
### PRINCIPAL COMPONENT ANALYSIS
___
*From: https://nirpyresearch.com/classification-nir-spectra-principal-component-analysis-python/*
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from scipy.signal import savgol_filter
from sklearn.decomposition import PCA as sk_pca
from sklearn.preprocessing import StandardScaler
from sklearn import svm
from sklearn.cluster import KMeans
import os.path
import spectrai as spa
%load_ext autoreload
%autoreload 2
%matplotlib inline
# Loading
DATA_PATH = os.path.join('..', 'data')
ARGENTINA_PATH = os.path.join('arg-romina', 'spectra')
ARGENTINA_XRF_URL = os.path.join(DATA_PATH, 'arg-romina', 'measurements', '2015-xrf-results-mean-and-errors.xls')
DATA_URL = os.path.join(DATA_PATH, ARGENTINA_PATH, '*.CSV')
X, X_names, y, y_names = spa.load_data(DATA_URL, ARGENTINA_XRF_URL)
feat = X
fig, ax = plt.subplots(figsize=(15,8))
ax.plot(X_names, feat[0])
# Calculate first derivative applying a Savitzky-Golay filter
dfeat = savgol_filter(feat, 25, polyorder = 5, deriv=1)
fig, ax = plt.subplots(figsize=(15,8))
ax.plot(X_names, dfeat[0])
```
## PCA
```
# Initialise
skpca1 = sk_pca(n_components=10)
skpca2 = sk_pca(n_components=10)
# Scale the features to have zero mean and standard devisation of 1
# This is important when correlating data with very different variances
nfeat1 = StandardScaler().fit_transform(feat)
nfeat2 = StandardScaler().fit_transform(dfeat)
# Fit the spectral data and extract the explained variance ratio
X1 = skpca1.fit(nfeat1)
expl_var_1 = X1.explained_variance_ratio_
# Fit the first data and extract the explained variance ratio
X2 = skpca2.fit(nfeat2)
expl_var_2 = X2.explained_variance_ratio_
# Plot data
with plt.style.context(('ggplot')):
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(12,5))
fig.set_tight_layout(True)
ax1.plot(expl_var_1,'-o', label="Explained Variance %")
ax1.plot(np.cumsum(expl_var_1),'-o', label = 'Cumulative variance %')
ax1.set_xlabel("PC number")
ax1.set_title('Absorbance data')
ax2.plot(expl_var_2,'-o', label="Explained Variance %")
ax2.plot(np.cumsum(expl_var_2),'-o', label = 'Cumulative variance %')
ax2.set_xlabel("PC number")
ax2.set_title('First derivative data')
plt.legend()
plt.show()
```
## Clustering
```
skpca2 = sk_pca(n_components=4)
# Transform on the scaled features
Xt2 = skpca2.fit_transform(nfeat2)
# How well PC1 and 2 discriminate spectra?
fig, ax = plt.subplots(figsize=(15,8))
ax.scatter(Xt2[:,0],Xt2[:,1], edgecolors='k')
```
| github_jupyter |
base code borrowed from [this Google Colab Notebook](https://colab.research.google.com/github/google-research/bert/blob/master/predicting_movie_reviews_with_bert_on_tf_hub.ipynb).
Refactored by [Shuyi Wang](https://www.linkedin.com/in/shuyi-wang-b3955026/)
Please refer to [this Medium Article](https://medium.com/@wshuyi/how-to-do-text-binary-classification-with-bert-f1348a25d905) for the tutorial on how to classify English text data.
```
!pip install bert-tensorflow
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
import pickle
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
def pretty_print(result):
df = pd.DataFrame([result]).T
df.columns = ["values"]
return df
def create_tokenizer_from_hub_module(bert_model_hub):
"""
Get the vocab file and casing info from the Hub module.
https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1
大写转小写,这个model TODO 这一步其实不需要吧
"""
with tf.Graph().as_default():
bert_module = hub.Module(bert_model_hub)
tokenization_info = bert_module(signature="tokenization_info", as_dict=True)
with tf.Session() as sess:
vocab_file, do_lower_case = sess.run([tokenization_info["vocab_file"],
tokenization_info["do_lower_case"]])
return bert.tokenization.FullTokenizer(
vocab_file=vocab_file, do_lower_case=do_lower_case)
def make_features(dataset, label_list, MAX_SEQ_LENGTH, tokenizer, DATA_COLUMN, LABEL_COLUMN):
# run_classifier.InputExample 处理成bert可读形式
input_example = dataset.apply(lambda x: run_classifier.InputExample(guid=None, text_a = x[DATA_COLUMN], text_b = None, label = x[LABEL_COLUMN]), axis = 1)
# input example 转 feature TODO tokenizer 可以不要
features = run_classifier.convert_examples_to_features(input_example, label_list, MAX_SEQ_LENGTH, tokenizer)
return features
def create_model(bert_model_hub, is_predicting, input_ids, input_mask, segment_ids, labels,
num_labels):
"""Creates a classification model."""
bert_module = hub.Module(
bert_model_hub,
trainable=True)
bert_inputs = dict(
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids)
bert_outputs = bert_module(
inputs=bert_inputs,
signature="tokens",
as_dict=True)
# Use "pooled_output" for classification tasks on an entire sentence.
# Use "sequence_outputs" for token-level output.
output_layer = bert_outputs["pooled_output"]
hidden_size = output_layer.shape[-1].value
# Create our own layer to tune for politeness data.
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
# Dropout helps prevent overfitting
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
log_probs = tf.nn.log_softmax(logits, axis=-1)
# Convert labels into one-hot encoding
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
# If we're predicting, we want predicted labels and the probabiltiies.
if is_predicting:
return (predicted_labels, log_probs)
# If we're train/eval, compute loss between predicted and actual label
per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
loss = tf.reduce_mean(per_example_loss)
return (loss, predicted_labels, log_probs)
# model_fn_builder actually creates our model function
# using the passed parameters for num_labels, learning_rate, etc.
def model_fn_builder(bert_model_hub, num_labels, learning_rate, num_train_steps,
num_warmup_steps):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_predicting = (mode == tf.estimator.ModeKeys.PREDICT)
# TRAIN and EVAL
if not is_predicting:
(loss, predicted_labels, log_probs) = create_model(
bert_model_hub, is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
train_op = bert.optimization.create_optimizer(
loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu=False)
# Calculate evaluation metrics.
def metric_fn(label_ids, predicted_labels):
accuracy = tf.metrics.accuracy(label_ids, predicted_labels)
f1_score = tf.contrib.metrics.f1_score(
label_ids,
predicted_labels)
auc = tf.metrics.auc(
label_ids,
predicted_labels)
recall = tf.metrics.recall(
label_ids,
predicted_labels)
precision = tf.metrics.precision(
label_ids,
predicted_labels)
true_pos = tf.metrics.true_positives(
label_ids,
predicted_labels)
true_neg = tf.metrics.true_negatives(
label_ids,
predicted_labels)
false_pos = tf.metrics.false_positives(
label_ids,
predicted_labels)
false_neg = tf.metrics.false_negatives(
label_ids,
predicted_labels)
return {
"eval_accuracy": accuracy,
"f1_score": f1_score,
"auc": auc,
"precision": precision,
"recall": recall,
"true_positives": true_pos,
"true_negatives": true_neg,
"false_positives": false_pos,
"false_negatives": false_neg
}
eval_metrics = metric_fn(label_ids, predicted_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
train_op=train_op)
else:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
eval_metric_ops=eval_metrics)
else:
(predicted_labels, log_probs) = create_model(
bert_model_hub, is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
predictions = {
'probabilities': log_probs,
'labels': predicted_labels
}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
# Return the actual model function in the closure
return model_fn
def estimator_builder(bert_model_hub, OUTPUT_DIR, SAVE_SUMMARY_STEPS, SAVE_CHECKPOINTS_STEPS, label_list, LEARNING_RATE, num_train_steps, num_warmup_steps, BATCH_SIZE):
# Specify outpit directory and number of checkpoint steps to save
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
model_fn = model_fn_builder(
bert_model_hub = bert_model_hub,
num_labels=len(label_list),
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
return estimator, model_fn, run_config
def run_on_dfs(train, test, DATA_COLUMN, LABEL_COLUMN,
MAX_SEQ_LENGTH = 128,
BATCH_SIZE = 32,
LEARNING_RATE = 2e-5,
NUM_TRAIN_EPOCHS = 3.0,
WARMUP_PROPORTION = 0.1,
SAVE_SUMMARY_STEPS = 100,
SAVE_CHECKPOINTS_STEPS = 10000,
bert_model_hub = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"):
label_list = train[LABEL_COLUMN].unique().tolist()
tokenizer = create_tokenizer_from_hub_module(bert_model_hub)
train_features = make_features(train, label_list, MAX_SEQ_LENGTH, tokenizer, DATA_COLUMN, LABEL_COLUMN)
test_features = make_features(test, label_list, MAX_SEQ_LENGTH, tokenizer, DATA_COLUMN, LABEL_COLUMN)
num_train_steps = int(len(train_features) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
estimator, model_fn, run_config = estimator_builder(
bert_model_hub,
OUTPUT_DIR,
SAVE_SUMMARY_STEPS,
SAVE_CHECKPOINTS_STEPS,
label_list,
LEARNING_RATE,
num_train_steps,
num_warmup_steps,
BATCH_SIZE)
train_input_fn = bert.run_classifier.input_fn_builder(
features=train_features,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
test_input_fn = run_classifier.input_fn_builder(
features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
result_dict = estimator.evaluate(input_fn=test_input_fn, steps=None)
return result_dict, estimator
import random
random.seed(10)
OUTPUT_DIR = 'output'
```
----- you just need to focus from here ------
## Get your data
```
!wget https://github.com/wshuyi/demo-chinese-text-binary-classification-with-bert/raw/master/dianping_train_test.pickle
with open("dianping_train_test.pickle", 'rb') as f:
train, test = pickle.load(f)
train = train.sample(len(train))
train.head()
myparam = {
"DATA_COLUMN": "comment",
"LABEL_COLUMN": "sentiment",
"LEARNING_RATE": 2e-5,
"NUM_TRAIN_EPOCHS":3,
"bert_model_hub":"https://tfhub.dev/google/bert_chinese_L-12_H-768_A-12/1"
}
result, estimator = run_on_dfs(train, test, **myparam)
pretty_print(result)
def predict(train, test, DATA_COLUMN, LABEL_COLUMN,
MAX_SEQ_LENGTH = 128,
bert_model_hub = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"):
label_list = train[LABEL_COLUMN].unique().tolist()
tokenizer = create_tokenizer_from_hub_module(bert_model_hub)
test_features = make_features(test, label_list, MAX_SEQ_LENGTH, tokenizer, DATA_COLUMN, LABEL_COLUMN)
# print("label_list", label_list) # 两类
test_input_fn = run_classifier.input_fn_builder(
features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
predict_results = estimator.predict(
input_fn=test_input_fn)
print("Predictions on test file")
for prediction in predict_results: # 预测结果
print(prediction)
myparam2 = {
"DATA_COLUMN": "comment",
"LABEL_COLUMN": "sentiment",
"bert_model_hub":"https://tfhub.dev/google/bert_chinese_L-12_H-768_A-12/1"
}
print(test[:5])
predict(train[:5] , test[:5], **myparam2)
```
| github_jupyter |
# Inference: Classical and Bayesian
G. Richards
(2016, 2018, 2020),
with material from Ivezic [Sections 4.0, 4.1, 4.2, 4.3, 4.5, 5.0, 5.1, 5.2], Bevington, and Leighly.
Statistical *inference* is about drawing conclusions from data, specifically determining the properties of a population by data sampling.
Three examples of inference are:
* What is the best estimate for a model parameter
* How confident we are about our result
* Are the data consistent with a particular model/hypothesis
### Some Terminology
* We typically study the properties of some ***population*** by measuring ***samples*** from that population.
* To conclude something about the population from the sample, we develop ***estimators***. An estimator is a statistic based on observed data.
* A ***statistic*** is any function of the sample. For example, the sample mean is a statistic. But also, "the value of the first measurement" is also a statistic.
* There are ***point*** and ***interval estimators***. Point estimators yield single-valued results (example: the position of an object), while with an interval estimator, the result would be a range of plausible values (example: confidence interval for the position of an object).
### Bayesian vs Frequentist (Ivezic 4.1)
This is the point where we are supposed to have a long discussion about the various pros and cons of the two most common ways of approaching inference problems:
* Classical (frequentist) and
* Bayesian.
Personally I don't see the need for a lengthy discussion on this. In short, classical (frequentist) statistics is concerned with the frequency with which $A$ happens in identical repeats of an experiment, i.e., $p(A)$. Bayesian statistics is concerned instead with $p(A|B)$, which is how plausible it is for $A$ to happen given the knowledge that $B$ has happened (or is true).
For more insight see [Jake VanderPlas's blog "Frequentism and Bayesianism: A Practical Introduction](http://jakevdp.github.io/blog/2014/03/11/frequentism-and-bayesianism-a-practical-intro/).
My colleague, Karen Leighly, dug up the following article, which might help one to understand the differences in these approaches in a relatively simply way. The first 4 sections are what is relevant here.
[Efron 1978](http://www.jstor.org/stable/2321163?seq=1#page_scan_tab_contents)
I'll briefly (and perhaps too cavalierly) summarize it.
Let's say that you get the results of an IQ test. Any given test result might not give you your "real" IQ. But it gives us a way to *estimate* it (and the possible range of values).
For a frequentist, the best estimator is just the average of many test results. So, if you took 5 IQ tests and got a 160, then that would be the estimator of your true IQ.
On the other hand, a Bayesian would say: "but wait, I know that IQ tests are normed to 100 with a standard deviation of 15 points". So they will use that as "prior" information, which is important here since 160 is a 4$\sigma$ outlier.
```
# Execute this cell
%matplotlib inline
import numpy as np
from scipy.stats import norm
from matplotlib import pyplot as plt
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=10, usetex=True)
#------------------------------------------------------------
# Define the distributions to be plotted
sigma_values = [15, 6.7, 1]
linestyles = ['--', '-', ':']
mu_values = [100, 148, 160]
labeltext = ['prior dist.', 'posterior dist.', 'observed mean']
xplot = np.linspace(50, 200, 1000)
#------------------------------------------------------------
# plot the distributions
fig, ax = plt.subplots(figsize=(10, 7.5))
for sigma, ls, mu, lab in zip(sigma_values, linestyles, mu_values, labeltext):
# create a gaussian / normal distribution
dist = norm(mu, sigma)
if (sigma>1):
plt.plot(xplot, dist.pdf(xplot), ls=ls, c='black',label=r'%s $\mu=%i,\ \sigma=%.1f$' % (lab, mu, sigma))
else:
plt.plot([159.9,160.1],[0,0.8], ls=ls, color='k', label=r'%s $\mu=%i$' % (lab, mu))
plt.xlim(50, 200)
plt.ylim(0, 0.1)
plt.xlabel('$x$')
plt.ylabel(r'$p(x|\mu,\sigma)$')
plt.title('Gaussian Distribution')
plt.legend()
```
The end result (skipping over the detailed math) is that the Bayesian estimate of the IQ is not 160, but rather 148, or more specifically that $p(141.3\le \mu \le 154.7 \, | \, \overline{x}=160) = 0.683$.
That's actually fine, where the controvery comes in is when the Bayesian wants to do the same things but doesn't actually know the prior distribution, or when the parameter is fixed but we are trying to experimentally verify it (e.g., the speed of light).
## Maximum Likelihood Estimation (MLE), Ivezic 4.2
Let's not worry about classical vs. Bayesian right now and talk about maximum likelihood estimation (Ivezic, 4.2), which is relevant to both.
If we know the distribution from which our data were drawn, then we can compute the **probability** or **likelihood** of our data.
For example if you know that your data are drawn from a model with a Gaussian distribution, then we've already seen that the probablity of getting a specific value of $x$ is given by
$$p(x|\mu,\sigma) = \frac{1}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x-\mu)^2}{2\sigma^2}\right).$$
If we want to know the total likelihood of our *entire* data set (as opposed to one measurement) then we must compute the *product* of all the individual probabilities:
$$L \equiv p(\{x_i\}|M(\theta)) = \prod_{i=1}^n p(x_i|M(\theta)),$$
where $M$ refers to the *model* and $\theta$ refers collectively to the $k$ parameters of the model, which can be multi-dimensional.
In words, this is *the probability of the data given the model*. However, note that while the components of $L$ may be normalized pdfs, their product is not. Also the product can be very small, so we often take the log of $L$.
We can write this out as
$$L = \prod_{i=1}^n \frac{1}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma^2}\right),$$
and simplify to
$$L = \prod_{i=1}^n \left( \frac{1}{\sigma\sqrt{2\pi}} \right) \exp\left( -\frac{1}{2} \sum \left[\frac{-(x_i-\mu)}{\sigma} \right]^2 \right),$$
where we have written the product of the exponentials as the exponential of the sum of the arguments, which will make things easier to deal with later.
That is, we have done this: $$\prod_{i=1}^n A_i \exp(-B_i) = (A_iA_{i+1}\ldots A_n) \exp[-(B_i+B_{i+1}+\ldots+B_n)]$$
If you have done $\chi^2$ analysis (e.g.,, doing a linear least-squares fit), then you might notice that the argument of the exponential is just
$$\exp \left(-\frac{\chi^2}{2}\right).$$
That is, for our gaussian distribution
$$\chi^2 = \sum_{i=1}^n \left ( \frac{x_i-\mu}{\sigma}\right)^2.$$
So, maximizing the likelihood is the same as minimizing $\chi^2$. In both cases we are finding the most likely values of our model parameters (here $\mu$ and $\sigma$).
Here's [an animation of linear least squares fitting](https://yihui.org/animation/example/least-squares/).
They are trying to fit a line to some data. They start by fixing the interecept and then trying 50 different values of the slope. The red dashed lines show the difference between the predicted value and the actual value. These are squared and summed (residual sum of squares, or $\chi^2$) and plotted as the y axis in the right hand plot. You see $\chi^2$ going down as the slope changes, bottoming out at the best slope (presumably with $\chi^2 \sim 1$, but we can't tell from the scale shown). Then $\chi^2$ goes back up after we have passed through the best slope.
With the best slope determined, we then try different values of the intercept, choosing the one that minimizes $\chi^2$.
Let's say that we know that some data were drawn from a Gaussian distribution, but we don't know the $\theta = (\mu,\sigma)$ values of that distribution (i.e., the parameters), then MLE is about varying the parameters until we find the maximal value of $L$ (i.e., the **maximum likelihood**). Those model parameters will also minimize $\chi^2$. Simple as that. Stop and make sure that you understand this.
A simple example would be the likelihood of rolling a certain combination of numbers on a six-sided die.
The probability of rolling a 3 is $1/6$ (as is the probability of *any* roll). So, what is the probability of rolling (in no particular order): {1,1,2,3,3,3,4,5,6,6}?
```
print((1./6)*(1./6)*(1./6)*(1./6)*(1./6)*(1./6)*(1./6)*(1./6)*(1./6)*(1./6))
print((1./6)**10)
```
So, even for 10 rolls of the die, the likelihood is pretty small. That's just because there are *lots* of possible combinations of rolling a die 10 times. This particular series of numbers is just as likely as any other.
Students who took PHYS 114 with me will recall that the result is related to the number of *combinations* ($n$ choose $r$).
```
# Write some code to compute the probability for N rolls
import numpy as np
N=____ #Number of rolls
L=____ #Likelihood, initialize to unity
rolls = np.array([])
for i in np.arange(____): #Loop over each roll
#Append a single new roll to "rolls" between 1 and 6 (careful) to the rolls array
rolls = np.append(____,np.random.randint(low=____,high=____,size=____))
L = L*(1./6) #The likelihood of each roll is 1/6
print(L,rolls)
```
### MLE applied to a Homoscedastic Gaussian (Ivezic 4.2.3)
Let's take a look at an example using a Gaussian model where all the measurements have the same error ($\sigma$). This is known as having **homoscedastic** errors. Don't be intimidated by the fancy word, statisticians just like to sound smart, so they say "homoscedastic" instead of "uniform errors". Later we will consider the case where the measurements can have different errors ($\sigma_i$) which is called **heteroscedastic**.
For an experiment with data $D=\{x_i\}$ in 1D with Gaussian errors, we have
$$L \equiv p(\{x_i\}|\mu,\sigma) = \prod_{i=1}^N \frac{1}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma^2}\right).$$
Note that that is $p(\{x_i\})$ not $p(x_i)$, that is the probability of the full data set, not just one measurement.
If $\sigma$ is both uniform and *known*, then this is a one parameter model with $k=1$ and $\theta_1=\mu$.
As we found above, likelihoods can be really small, so let's define the *log-likelihood function* as ${\rm lnL} = \ln[L(\theta)]$. The maximum of this function happens at the same place as the maximum of $L$. Note that any constants in $L$ have the same effect for all model parameters, so constant terms can be ignored.
In this case we then have $${\rm lnL} = {\rm constant} - \sum_{i=1}^N \frac{(x_i - \mu)^2}{2\sigma^2}.$$
Take a second and make sure that you understand how we got there. It might help to remember that above, we wrote
$$L = \prod_{i=1}^n \left( \frac{1}{\sigma\sqrt{2\pi}} \right) \exp\left( -\frac{1}{2} \sum \left[\frac{-(x_i-\mu)}{\sigma} \right]^2 \right).$$
We then determine the maximum in the same way that we always do. It is the parameter set for which the derivative of ${\rm lnL}$ is zero:
$$\frac{d\;{\rm lnL}(\mu)}{d\mu}\Biggr\rvert_{\mu_0} \equiv 0.$$
That gives $$ \sum_{i=1}^N \frac{(x_i - \mu_o)}{\sigma^2} = 0.$$
Since $\sigma = {\rm constant}$ (at least in this case), that says
$$\sum_{i=1}^N x_i = \sum_{i=1}^N \mu_0 = N \mu_0.$$
Thus we find that
$$\mu_0 = \frac{1}{N}\sum_{i=1}^N x_i,$$
which is just the arithmetic mean of all the measurements.
As promised last week, that's where the formula that you know and love for the mean comes from.
### The Sample Mean is an ML Estimator
So the sample mean ($\overline{x} = \mu_0$) of observations drawn from a $\mathscr{N}(\mu, \sigma=const)$ distribution is a maximum-likelihood estimator of the distribution's $\mu$ parameter.
We'd intuitively guess that, but this derivation clarifies our choice: as an estimator of the **real** value of $\mu$, we adopt the value $\mu_0$ for which the data set is maximally likely to occur.
As we just discussed, in an experiment with data $D=\{x_i\}$ in 1D with Gaussian errors, we have
$$L \equiv p(\{x_i\}|\mu,\sigma) = \prod_{i=1}^N \frac{1}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma^2}\right).$$
Let's create some data and see what the resulting likelihood looks like for some example points.
```
#Load up the algorithms we are going to need.
import numpy as np
from matplotlib import pyplot as plt
from scipy.stats import norm
%matplotlib inline
```
We are going to draw a homoscedastic sample of ${x_i}$ from a Gaussian and compute the likelihood.
First generate a sample of `N=3` points drawn from a normal distribution with `mu=1.0` and `sigma=0.2`: $\mathscr{N}(\mu,\sigma)$
```
N = ___ #Complete
mu = ___
sigma = ___
np.random.seed(42)
sample = norm(___,___).rvs(___)
print(sample)
```
Think back to Lecture 2 when we did a Kernel Density Estimate. Treat each of those observations as an estimate of the true distribution. So, we are going to center a Gaussian (with the knownn $\sigma$) at each point, this is the likelihood $p(x_i|\mu,\sigma)$.
Plot each of the likelihoods separately. Also plot their product. Make the $x$ axis a grid of 1000 points uniformly sampled between $x=0$ and $x=2$.
Note that, according to [scipy.stats.norm](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html), `norm.pdf(x, loc, scale)` is identically equivalent to `norm.pdf(y)/scale` with `y=(x-loc)/scale$`.
```
#Make the plot and see if you get the same as me.
xgrid = np.linspace(___,___,___)
L1 = norm.pdf(___,loc=___,scale=___) #This is a Gaussian PDF sampled uniformly, centered at a specific location.
L2 = norm.pdf(___,loc=___,scale=___)
L3 = norm.pdf(___,loc=___,scale=___)
L = ___ #Total L is ???
fig, ax = plt.subplots(figsize=(7, 5))
plt.plot(____, ____, ls='-', c='green', label=r'$L(x_1)$')
plt.plot(____, ____, ls='-', c='red', label=r'$L(x_2)$')
plt.plot(____, ____, ls='-', c='blue', label=r'$L(x_3)$')
plt.plot(____, ____, ls='-', c='black', label=r'$L(\{x\})$')
plt.xlim(0.2, 1.8)
plt.ylim(0, 8.0)
plt.xlabel('$\mu$') #Leave out or adjust if no latex
plt.ylabel(r'$p(x_i|\mu,\sigma)$') #Leave out or adjust if no latex
plt.title('MLE for Gaussian Distribution')
plt.legend()
```
You should get something that looks like this:

Now we can just read off the maximum likelihood solution. Use `np.argsort()` to figure out the argument of the largest value and print that index of `xgrid`.
```
indices = ___
index_max = indices[___]
print("Likelihood is maximized at %.3f" % xgrid[index_max])
```
### Quantifying Estimated Uncertainty (Ivezic 4.2.5)
Our ML estimate of $\mu$ is not perfect. The uncertaintly of the estimate is captured by the likelihood function, but we'd like to quantify it with a few numbers.
We *define* the uncertainty on our MLEs as second (partial) derivatives of log-likelihood:
$$\sigma_{jk} = \left( - \frac{d^2}{d\theta_j} \frac{\ln L}{d\theta_k} \Biggr\rvert_{\theta=\theta_0}\right)^{-1/2}.$$
The marginal error bars for each parameter, $\theta_i$ are given by the diagonal elements, $\sigma_{ii}$, of this **covariance matrix**.
In our example, the uncertainly on the mean is
$$\sigma_{\mu} = \left( - \frac{d^2\ln L(\mu)}{d\mu^2}\Biggr\rvert_{\mu_0}\right)^{-1/2}$$
We find
$$\frac{d^2\ln L(\mu)}{d\mu^2}\Biggr\rvert_{\mu_0} = - \sum_{i=1}^N\frac{1}{\sigma^2} = -\frac{N}{\sigma^2},$$
since, again, $\sigma = {\rm constant}$.
Then $$\sigma_{\mu} = \frac{\sigma}{\sqrt{N}}.$$
So, our estimator of $\mu$ is $\overline{x}\pm\frac{\sigma}{\sqrt{N}}$, which is the result that we are already familiar with.
### Confidence Intervals
The $(\mu_0 - \sigma_\mu, \mu_0 + \sigma_\mu)$ range gives us a **confidence interval**.
In frequentist interptetation, if we repeated the same measurement a hundred times, we'd find that 68 experiments yield a result within the computed confidence interval ($1 \sigma$ errors).
### Confidence Estimates: Bootstrap and Jackknife (Ivezic 4.5)
We often assume that the distribution is Gaussian and our samples are large, but even if that is not the case, we can still compute good confidence intervals (e.g., $a<x<b$ with 95\% confidence) using *resampling* strategies.
Remember that we have a data set $\{x_i\}$ from which we have estimated the distribution as $f(x)$ for a true distribution $h(x)$.
In **bootstrapping** we map the uncertainty of the parameters by re-sampling from our distribution (with replacement) $B$ times, such that we obtain $B$ measures of our parameters. So, if we have $i=1,\dots,N$ data points in $\{x_i\}$, we draw $N$ of them to make a new sample, where some values of $\{x_i\}$ will be used more than once.
The **jackknife** method is similar except that we don't use a sample size of $N$, rather we leave off one or more of the observations from $\{x_i\}$. As with bootstrap, we do this multiple times, generating samples from which we can determine our uncertainties.
It is generally a good idea to use both methods and compare the results.
An example of bootstrap is given below, using [astroML.resample.bootstrap](http://www.astroml.org/modules/generated/astroML.resample.bootstrap.html), where the arguments are 1) the data, 2) the number of bootstrap resamples to use, and 3) the statistic to be computed.
You'll get some more practice with this in a homework assignment based on Data Camp.
```
# Ivezic v2, Figure 4.3, modified slightly by GTR
# %load ../code/fig_bootstrap_gaussian.py
"""
Bootstrap Calculations of Error on Mean
---------------------------------------
Figure 4.3.
The bootstrap uncertainty estimates for the sample standard deviation
:math:`\sigma` (dashed line; see eq. 3.32) and :math:`\sigma_G` (solid line;
see eq. 3.36). The sample consists of N = 1000 values drawn from a Gaussian
distribution with :math:`\mu = 0` and :math:`\sigma = 1`. The bootstrap
estimates are based on 10,000 samples. The thin lines show Gaussians with
the widths determined as :math:`s / \sqrt{2(N - 1)}` (eq. 3.35) for
:math:`\sigma` and :math:`1.06 s / \sqrt{N}` (eq. 3.37) for :math:`\sigma_G`.
"""
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from scipy.stats import norm
from matplotlib import pyplot as plt
from astroML.resample import bootstrap
from astroML.stats import sigmaG
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
if "setup_text_plots" not in globals():
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=12, usetex=True)
m = 1000 # number of points
n = 10000 # number of bootstraps
#------------------------------------------------------------
# sample values from a normal distribution
np.random.seed(123)
data = norm(0, 1).rvs(m)
#------------------------------------------------------------
# Compute bootstrap resamplings of data
mu1_bootstrap = bootstrap(data, n, np.std, kwargs=dict(axis=1, ddof=1))
mu2_bootstrap = bootstrap(data, n, sigmaG, kwargs=dict(axis=1))
#------------------------------------------------------------
# Compute the theoretical expectations for the two distributions
xgrid = np.linspace(0.8, 1.2, 1000)
sigma1 = 1. / np.sqrt(2 * (m - 1))
pdf1 = norm(1, sigma1).pdf(xgrid)
sigma2 = 1.06 / np.sqrt(m)
pdf2 = norm(1, sigma2).pdf(xgrid)
#------------------------------------------------------------
# Plot the results
fig, ax = plt.subplots(figsize=(8, 8))
ax.hist(mu1_bootstrap, bins=50, density=True, histtype='step',
color='blue', ls='dashed', label=r'$\sigma\ {\rm (std. dev.)}$')
ax.plot(xgrid, pdf1, color='gray')
ax.hist(mu2_bootstrap, bins=50, density=True, histtype='step',
color='red', label=r'$\sigma_G\ {\rm (quartile)}$')
ax.plot(xgrid, pdf2, color='gray')
ax.set_xlim(0.82, 1.18)
ax.set_xlabel(r'$\sigma$')
ax.set_ylabel(r'$p(\sigma|x,I)$')
ax.legend()
plt.show()
```
From Ivezic: "The bootstrap uncertainty estimates for the sample standard deviation $\sigma$ (dashed line; see Eq. 3.32) and $\sigma_G$ (solid line; see Eq. 3.36). The sample consists of N = 1000 values drawn from a Gaussian distribution with $\mu = 0$ and $\sigma = 1$. The bootstrap estimates are based on 10,000 samples. The thin grey lines show Gaussians with the widths determined as $s / \sqrt{2(N - 1)}$ (Eq. 3.35) for $\sigma$ and $1.06 s / \sqrt{N}$ (Eq. 3.37) for $\sigma_G$."
### MLE applied to a Heteroscedastic Gaussian (Ivezic 4.2.6)
Now let's look at a case where the errors are heteroscedastic. For example if we are measuring the length of a [rod](https://www.nist.gov/image/meter27jpg) and have $N$ measurements, $\{x_i\}$, where the error for each measurement, $\sigma_i$ is known. Since $\sigma$ is **not** a constant, then following the above, we have
$$\ln L = {\rm constant} - \sum_{i=1}^N \frac{(x_i - \mu)^2}{2\sigma_i^2}.$$
Taking the derivative:
$$\frac{d\;{\rm lnL}(\mu)}{d\mu}\Biggr\rvert_{\mu_0} = \sum_{i=1}^N \frac{(x_i - \mu_o)}{\sigma_i^2} = 0,$$
then simplifying:
$$\sum_{i=1}^N \frac{x_i}{\sigma_i^2} = \sum_{i=1}^N \frac{\mu_o}{\sigma_i^2},$$
yields a MLE solution of
$$\mu_0 = \frac{\sum_i^N (x_i/\sigma_i^2)}{\sum_i^N (1/\sigma_i^2)},$$
which is just a variance-weighted mean, with uncertainty
$$\sigma_{\mu} = \left( \sum_{i=1}^N \frac{1}{\sigma_i^2}\right)^{-1/2}.$$
### Cost Functions
Recall from Lecture 3 that we measured the deviation from expected in three different ways
$$d_i = x_i - \mu,$$
$$|x_i-\mu|,$$
and
$$(x_i-\mu)^2.$$
When doing trying to determine the best model parameters, we have to specify a "cost function", which is basically the prescription for evaluating the difference (distance) between our estimator and the true value. This is otherwise known as the "norm", or the total length of the distances. The first form above represents the $L_0$ norm, while the next two are the $L_1$ norm (the "taxi-cab" norm) and the $L_2$ norm (the "as the crow flies" norm).
So far we have been using an $L_2$ norm (which comes about simply from the definition of a Gaussian with the $(x_i-\mu)^2$ term). Later in the course we will encounter machine learning algorithms that allow us to specify different norms (different cost functions).
See [this Medium article](https://medium.com/@montjoile/l0-norm-l1-norm-l2-norm-l-infinity-norm-7a7d18a4f40c) and Ivezic, 4.2.8.
### Truncated/Censored Data
Note that knowing how to deal with missing data points ("censored data") is often quite important, but adds complications that we don't have time to get into here. For more, see Ivezic 4.2.7.
## "Goodness" of Fit (Ivezic 4.3)
The MLE approach tells us what the "best" model parameters are, but not how good the fit actually is. If the model is wrong, "best" might not be particularly revealing! For example, if you have $N$ points drawn from a linear distribution, you can always fit the data perfectly with an $N-1$ order polynomial. But that won't help you predict future measurements.
We can describe the **goodness of fit** in words simply as whether or not it is likely to have obtained $\ln L_0$ by randomly drawing from the data. That means that we need to know the *distribution* of $\ln L$ and not just the maximum.
For the Gaussian case we have just described, we do a standard transform of variables and compute the so-called $z$ score for each data point (basically the number of standard deviations away from the mean that this point is), writing
$$z_i = (x_i-\mu)/\sigma,$$ then
$$\ln L = {\rm constant} - \frac{1}{2}\sum_{i=1}^N z^2 = {\rm constant} - \frac{1}{2}\chi^2.$$
Here, $\chi^2$ is the thing whose distribution we discussed last week.
So $\ln L$ is distributed as $\chi^2$.
```
# Execute this cell to make the plot
# %load ../code/fig_chi2_distribution.py
"""
Example of a chi-squared distribution
---------------------------------------
Figure 3.14.
This shows an example of a :math:`\chi^2` distribution with various parameters.
"""
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from scipy.stats import chi2
from matplotlib import pyplot as plt
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=12, usetex=True)
#------------------------------------------------------------
# Define the distribution parameters to be plotted
k_values = [1, 2, 5, 7]
linestyles = ['-', '--', ':', '-.']
mu = 0
xplot = np.linspace(-1, 10, 1000)
#------------------------------------------------------------
# plot the distributions
fig, ax = plt.subplots(figsize=(5, 3.75))
fig.subplots_adjust(bottom=0.12)
for k, ls in zip(k_values, linestyles):
dist = chi2(k, mu)
idx = np.argsort(dist.pdf(xplot))[-1]
#print(f"The peak Q value for {k} degrees of freedom is {xplot[idx]}.")
print("The peak Q value for {0:d} degrees of freedom is {1:3.2f}.".format(k,xplot[idx]))
plt.plot(xplot, dist.pdf(xplot), ls=ls, c='black',
label=r'$k=%i$' % k)
plt.xlim(0, 10)
plt.ylim(0, 0.5)
plt.xlabel('$Q$')
plt.ylabel(r'$p(Q|k)$')
plt.title(r'$\chi^2\ \mathrm{Distribution}$')
plt.legend()
plt.show()
```
We define the $\chi^2$ per degree of freedom, $\chi^2_{dof}$, as
$$\chi^2_{dof} = \frac{1}{N-k}\sum_{i=1}^N z^2_i.$$
For a good fit, we would expect that $\chi^2_{dof}\approx 1$. If $\chi^2_{dof}$ is significantly larger than 1, then it is likely that we are not using the correct model.
We can also get overly high or low values of $\chi^2_{dof}$ if our errors are under- or over-estimated as shown below:
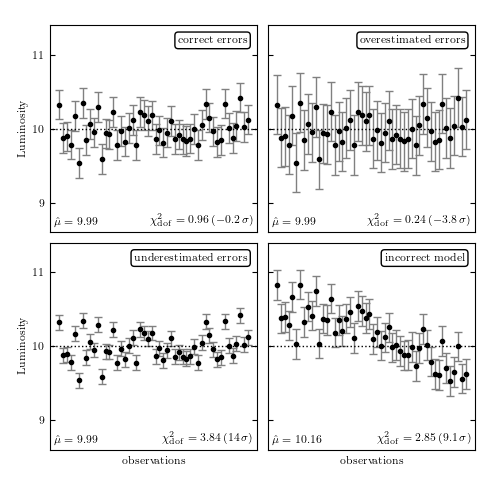
## Bayesian Statistical Inference (Ivezic 5.0, 5.1)
Up to now in this lecture we have been computing the **likelihood** $p(D|M)$. In Bayesian inference, we instead evaluate the **posterior probability** taking into account **prior** information.
Recall from the BasicStats lecture that Bayes' Rule is:
$$p(M|D) = \frac{p(D|M)p(M)}{p(D)},$$
where $D$ is for data and $M$ is for model.
We wrote this in words as:
$${\rm Posterior Probability} = \frac{{\rm Likelihood}\times{\rm Prior}}{{\rm Evidence}}.$$
If we explicitly recognize prior information, $I$, and the model parameters, $\theta$, then we can write:
$$p(M,\theta|D,I) = \frac{p(D|M,\theta,I)p(M,\theta|I)}{p(D|I)},$$
where we can omit the explict dependence on $\theta$ by writing $M$ instead of $M,\theta$ where appropriate.
Note that it is often that case that $p(D|I)$ is not evaluated explictly since the likelihood can be normalized such that the "evidence" is unity.
The Bayesian Statistical Inference process is then
* formulate the likelihood, $p(D|M,\theta,I)$, which is what we have been talking about so far today
* chose a prior, $p(M,\theta|I)$, which incorporates other information beyond the data in $D$
* determine the posterior pdf, $p(M,\theta|D,I)$
* search for the model paramters that maximize the posterior pdf
* quantify the uncertainty of the model parameter estimates
* test the hypothesis being addressed
How does our answer change for our earlier example if we include a Bayesian prior (like we assumed for the IQ problem) and instead maximize the posterior probability?
```
#Things that we have already done above, but need to reset here
N = 3 #Complete
mu = 1.0
sigma = 0.2
np.random.seed(42)
sample = norm(mu,sigma).rvs(N)
xgrid = np.linspace(0,2,1000)
L1 = norm.pdf(xgrid,loc=sample[0],scale=sigma)
L2 = norm.pdf(xgrid,loc=sample[1],scale=sigma)
L3 = norm.pdf(xgrid,loc=sample[2],scale=sigma)
#New things
Prior = norm.pdf(___,loc=___,scale=___) #Prior PDF
Post1 = ___*___ #Posterior PDF for the first measurement
Post2 = ___*___
Post3 = ___*___
Post = ___ #Total posterior PDF for all the measurements
# plot
fig, ax = plt.subplots(figsize=(7, 5))
plt.plot(xgrid, Post1, ls='-', c='green', label=r'$P(x_1)$')
plt.plot(xgrid, ____, ls='-', c='red', label=r'$P(x_2)$')
plt.plot(xgrid, ____, ls='-', c='blue', label=r'$P(x_3)$')
plt.plot(xgrid, Post/5, ls='-', c='black', label=r'$P(\{x\})$') #Scaled for the sake of display
plt.xlim(0.2, 1.8)
plt.ylim(0, 10.0)
plt.xlabel('$\mu$') #Leave out or adjust if no latex
plt.ylabel(r'$p(\mu,\sigma|x_i)$') #Leave out or adjust if no latex
plt.title('MLE for Gaussian Distribution')
plt.legend()
plt.show()
idx = np.argsort(Post)
print("Posterior PDF is maximized at %.3f" % xgrid[idx[-1]])
```
See what happens when you have just 2 measurements, but one has a much larger error than the other (i.e., the errors are heteroscedastic).
```
L1 = norm.pdf(xgrid,loc=___,scale=___) #Measurement with small error
L2 = norm.pdf(xgrid,loc=___,scale=___) #Measurement with large error (give it a very different location parameter)
L = L1 * L2
# plot
fig, ax = plt.subplots(figsize=(7, 5))
plt.plot(xgrid, L1, ls='-', c='green', label=r'$L(x_1): good$')
plt.plot(xgrid, L2, ls='-', c='red', label=r'$L(x_2): lousy$')
plt.plot(xgrid, L, ls='-', c='black', label=r'$L(\{x\}): weighted$')
plt.xlim(0.2, 1.8)
plt.ylim(0, 9.5)
plt.xlabel('$\mu$')
plt.ylabel(r'$p(x_i|\mu,\sigma)$')
plt.title('Weighted measurements (not normalized)')
plt.legend()
idx = np.argsort(L)
print("Likelihood is maximized at %.3f" % xgrid[idx[-1]])
```
## Bayesian Priors (Ivezic 5.2)
Priors can be **informative** or **uninformative**. As it sounds, informative priors are based on existing information that might be available. Uninformative priors can be thought of as "default" priors, i.e., what your prior is if you weren't explicitly including a prior, e.g, a "flat" prior like $p(\theta|M,I) \propto {\rm C}$.
For the IQ test example, what kind of prior did we use?
In a *hierarchical Bayesian* analysis the priors themselves can have parameters and priors (hyperparameters and hyperpriors), but let's not worry about that for now.
While determining good priors is important for Bayesian analysis, I don't want to get distracted by it here. You can read more about it in Ivezic, 5.2. However, I'll briefly introduce 3 principles here.
#### The Principle of Indifference
Essentially this means adopting a uniform prior, though you have to be a bit careful. Saying that an asteroid is equally likely to hit anywhere on the Earth is not the same as saying that all latitudes of impact are equally likely. Assuming $1/6$ for a six-side die would be an example of indifference.
#### The Principle of Invariance (or Consistency)
This applies to location and scale invariance. Location invariance suggests a uniform prior (within the accepted bounds). Scale invariance gives us priors that look like $p(A|I) \propto 1/A$.
#### The Principle of Maximum Entropy
The principle of maximum entropy is discussed in Ivezic, 5.2.2.
It is often true that Bayesian analysis and traditional MLE are essentially equivalent. However, in some cases, considering the priors can have significant consequences.
See Ivezic $\S$5.5 for such an example.
## Analysis of a Heteroscedastic Gaussian distribution with Bayesian Priors (Ivezic 5.6)
Consider the case of measuring a rod as above. We want to know the posterior pdf for the length of the rod, $p(M,\theta|D,I) = p(\mu|\{x_i\},\{\sigma_i\},I)$.
For the likelihood we have
$$L = p(\{x_i\}|\mu,I) = \prod_{i=1}^N \frac{1}{\sigma_i\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma_i^2}\right).$$
In the Bayesian case, we also need a prior. We'll adopt a uniform distribution given by
$$p(\mu|I) = C, \; {\rm for} \; \mu_{\rm min} < \mu < \mu_{\rm max},$$
where $C = \frac{1}{\mu_{\rm max} - \mu_{\rm min}}$ between the min and max and is $0$ otherwise.
The log of the posterior pdf is then
$$\ln L = {\rm constant} - \sum_{i=1}^N \frac{(x_i - \mu)^2}{2\sigma_i^2}.$$
This is exactly the same as we saw before, except that the value of the constant is different. Since the constant doesn't come into play, we get the same result as before:
$$\mu^0 = \frac{\sum_i^N (x_i/\sigma_i^2)}{\sum_i^N (1/\sigma_i^2)},$$
with uncertainty
$$\sigma_{\mu} = \left( \sum_{i=1}^N \frac{1}{\sigma_i^2}\right)^{-1/2}.$$
We get the same result because we used a flat prior. If the case were homoscedastic instead of heteroscedastic, we obviously would get the result from our first example.
Now let's consider the case where $\sigma$ is not known, but rather needs to be determined from the data. In that case, the posterior pdf that we seek is not $p(\mu|\{x_i\},\{\sigma_i\},I)$, but rather $p(\mu,\sigma|\{x_i\},I)$.
As before we have
$$L = p(\{x_i\}|\mu,\sigma,I) = \prod_{i=1}^N \frac{1}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma^2}\right),$$
except that now $\sigma$ is uknown instead of given (meaning we need to move it to the left of the "pipe").
Our Bayesian prior is now 2D instead of 1D and we'll adopt
$$p(\mu,\sigma|I) \propto \frac{1}{\sigma},\; {\rm for} \; \mu_{\rm min} < \mu < \mu_{\rm max} \; {\rm and} \; \sigma_{\rm min} < \sigma < \sigma_{\rm max}.$$
That is, all values of $\mu$ are equally likely (within the range indicated), but we'll down-weight the likelihood of large errors (again limiting $\sigma$ to some range). Note that the ranges actually drop out since they are constants.
With proper normalization, we have
$$p(\{x_i\}|\mu,\sigma,I)p(\mu,\sigma|I) = C\frac{1}{\sigma^{(N+1)}}\prod_{i=1}^N \exp\left( \frac{-(x_i-\mu)^2}{2\sigma^2} \right),$$
where
$$C = (2\pi)^{-N/2}(\mu_{\rm max}-\mu_{\rm min})^{-1} \left[\ln \left( \frac{\sigma_{\rm max}}{\sigma_{\rm min}}\right) \right]^{-1}.$$
The log of the posterior pdf is
$$\ln[p(\mu,\sigma|\{x_i\},I)] = {\rm constant} - (N+1)\ln\sigma - \sum_{i=1}^N \frac{(x_i - \mu)^2}{2\sigma^2}.$$
Right now that has $x_i$ in it, which isn't that helpful, but since we are assuming a Gaussian distribution, we can take advantage of the fact that the mean, $\overline{x}$, and the variance, $V (=s^2)$, completely characterize the distribution. So we can write this expression in terms of those variables instead of $x_i$. Skipping over the math details (see Ivezic $\S$5.6.1), we find
$$\ln[p(\mu,\sigma|\{x_i\},I)] = {\rm constant} - (N+1)\ln\sigma - \frac{N}{2\sigma^2}\left( (\overline{x}-\mu)^2 + V \right).$$
Note that this expression only contains the 2 parameters that we are trying to determine: $(\mu,\sigma)$ and 3 values that we can determine directly from the data: $(N,\overline{x},V)$.
Load and execute the next cell to visualize the posterior pdf for the case of $(N,\overline{x},V)=(10,1,4)$. Change `usetex=True` to `usetex=False` if you have trouble with the plotting. Try changing the values of $(N,\overline{x},V)$.
```
# Execute this cell
# %load code/fig_likelihood_gaussian.py
"""
Log-likelihood for Gaussian Distribution
----------------------------------------
Figure5.4
An illustration of the logarithm of the posterior probability density
function for :math:`\mu` and :math:`\sigma`, :math:`L_p(\mu,\sigma)`
(see eq. 5.58) for data drawn from a Gaussian distribution and N = 10, x = 1,
and V = 4. The maximum of :math:`L_p` is renormalized to 0, and color coded as
shown in the legend. The maximum value of :math:`L_p` is at :math:`\mu_0 = 1.0`
and :math:`\sigma_0 = 1.8`. The contours enclose the regions that contain
0.683, 0.955, and 0.997 of the cumulative (integrated) posterior probability.
"""
# Author: Jake VanderPlas
# License: BSD
# The figure produced by this code is published in the textbook
# "Statistics, Data Mining, and Machine Learning in Astronomy" (2013)
# For more information, see http://astroML.github.com
# To report a bug or issue, use the following forum:
# https://groups.google.com/forum/#!forum/astroml-general
import numpy as np
from matplotlib import pyplot as plt
from astroML.plotting.mcmc import convert_to_stdev
#----------------------------------------------------------------------
# This function adjusts matplotlib settings for a uniform feel in the textbook.
# Note that with usetex=True, fonts are rendered with LaTeX. This may
# result in an error if LaTeX is not installed on your system. In that case,
# you can set usetex to False.
from astroML.plotting import setup_text_plots
setup_text_plots(fontsize=14, usetex=True)
def gauss_logL(xbar, V, n, sigma, mu):
"""Equation 5.57: gaussian likelihood"""
return (-(n + 1) * np.log(sigma)
- 0.5 * n * ((xbar - mu) ** 2 + V) / sigma ** 2)
#------------------------------------------------------------
# Define the grid and compute logL
sigma = np.linspace(1, 5, 70)
mu = np.linspace(-3, 5, 70)
xbar = 1
V = 4
n = 10
logL = gauss_logL(xbar, V, n, sigma[:, np.newaxis], mu)
logL -= logL.max()
#------------------------------------------------------------
# Plot the results
fig = plt.figure(figsize=(5, 3.75))
plt.imshow(logL, origin='lower',
extent=(mu[0], mu[-1], sigma[0], sigma[-1]),
cmap=plt.cm.binary,
aspect='auto')
plt.colorbar().set_label(r'$\log(L)$')
plt.clim(-5, 0)
plt.contour(mu, sigma, convert_to_stdev(logL),
levels=(0.683, 0.955, 0.997),
colors='k')
plt.text(0.5, 0.93, r'$L(\mu,\sigma)\ \mathrm{for}\ \bar{x}=1,\ V=4,\ n=10$',
bbox=dict(ec='k', fc='w', alpha=0.9),
ha='center', va='center', transform=plt.gca().transAxes)
plt.xlabel(r'$\mu$')
plt.ylabel(r'$\sigma$')
```
The shaded region is the posterior probability. The contours are the confidence intervals. We can compute those by determining the marginal distribution at each $(\mu,\sigma)$. The top panels of the figures below show those marginal distributions. The solid line is what we just computed. The dotted line is what we would have gotten for a uniform prior--not that much difference. The dashed line is the MLE result, which is quite different. The bottom panels show the cumulative distribution.
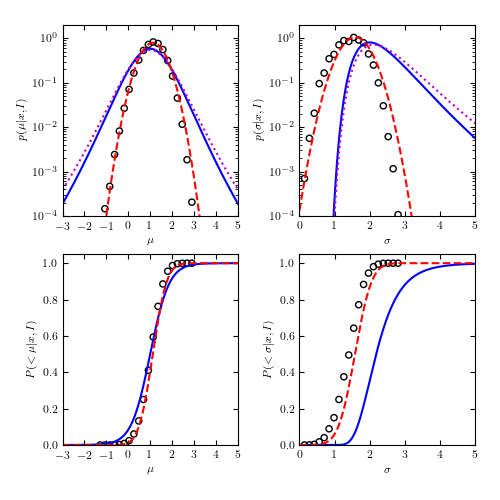
Note that the marginal pdfs follow a Student's $t$ Distribution, which becomes Gaussian for large $N$.
### Recap
To review: the Bayesian Statistical Inference process is
* formulate the likelihood, $p(D|M,\theta,I)$
* chose a prior, $p(M,\theta|I)$, which incorporates other information beyond the data in $D$
* determine the posterior pdf, $p(M,\theta|D,I)$
* search for the model paramters that maximize the posterior pdf
* quantify the uncertainty of the model parameter estimates
* test the hypothesis being addressed
The last part we haven't talked about yet.
What if we wanted to model the mixture of a Gauassian distribution with a uniform distribution. When might that be useful? Well, for example:
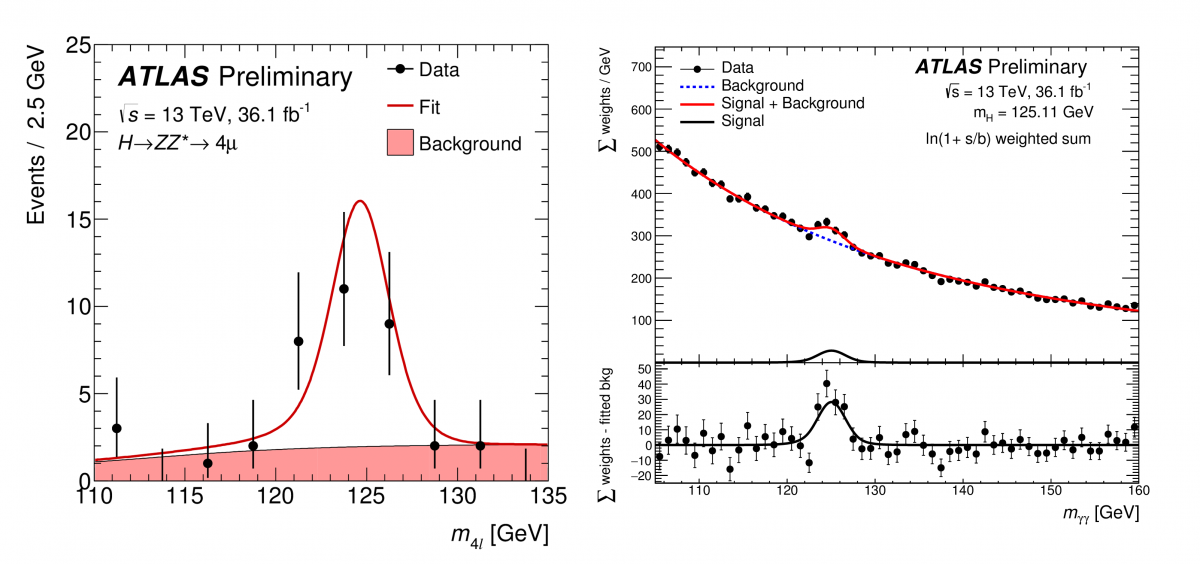
Obviously this isn't exactly a Gaussian and a uniform distribution, but a line feature superimposed upon a background is the sort of thing that a physicist might see and is pretty close to this case for a local region around the feature of interest. This is the example discussed in Ivezic $\S$5.6.5.
For this example, we will assume that the location parameter, $\mu$, is known (say from theory) and that the errors in $x_i$ are negligible compared to $\sigma$.
The likelihood of obtaining a measurement, $x_i$, in this example can be written as
$$L = p(x_i|A,\mu,\sigma,I) = \frac{A}{\sigma\sqrt{2\pi}} \exp\left(\frac{-(x_i-\mu)^2}{2\sigma^2}\right) + \frac{1-A}{W}.$$
Here the background probability is taken to be $0 < x < W$ and 0 otherwise. The feature of interest lies between $0$ and $W$. $A$ and $1-A$ are the relative strengths of the two components, which are obviously anti-correlated. Note that there will be covariance between $A$ and $\sigma$.
If we adopt a uniform prior in both $A$ and $\sigma$:
$$p(A,\sigma|I) = C, \; {\rm for} \; 0\le A<A_{\rm max} \; {\rm and} \; 0 \le \sigma \le \sigma_{\rm max},$$
then the posterior pdf is given by
$$\ln [p(A,\sigma|\{x_i\},\mu,W)] = \sum_{i=1}^N \ln \left[\frac{A}{\sigma \sqrt{2\pi}} \exp\left( \frac{-(x_i-\mu)^2}{2\sigma^2} \right) + \frac{1-A}{W} \right].$$
The figure below (Ivezic, 5.13) shows an example for $N=200, A=0.5, \sigma=1, \mu=5, W=10$. Specifically, the bottom panel is a result drawn from this distribution and the top panel is the likelihood distribution derived from the data in the bottom panel.
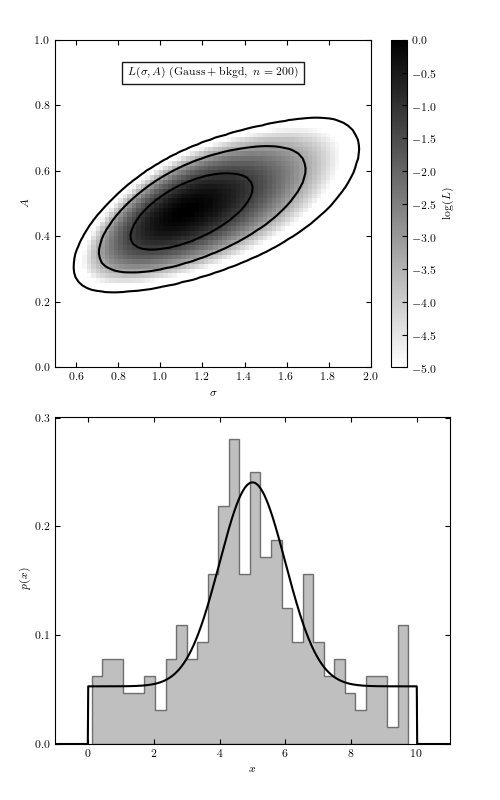
A more realistic example might be one where all three parameters are unknown: the location, the width, and the background level. But that will have to wait until $\S$5.8.6.
In the meantime, note that we have not binned the data, $\{x_i\}$. We only binned Figure 5.13 for the sake of visualizaiton. However, sometimes the data are inherently binned (e.g., the detector is pixelated). In that case, the data would be in the form of $(x_i,y_i)$, where $y_i$ is the number of counts at each location. We'll skip over this example, but you can read about it in Ivezic $\S$5.6.6. A refresher on the Poission distribution (Ivezic $\S$3.3.4) might be appropriate first.
| github_jupyter |
# Information Entry Script for Metagenomic Analysis
This script allows you to enter in information into the Jupyter_metagenomic_analysis.ipynb file. This script will prompt you to enter in relevant information as well as locate necessary files for the analysis, and this information will be stored into a .csv file that can be read in by the Jupyter_metagenomic_analysis.ipynb file. For output verification, you can open this .csv file (named --.csv) to ensure the information looks correct before running the analysis script.
When running this script, run all of the cells sequentially one after another after you are done editing the cell preceding it. This will ensure the script runs smoothly.
For questions about this script, please contact Akhil Gupta (gupta305@umn.edu).
---
## Import UI Elements Script
This will load in all necessary widgets. Ignore the warning message for now
```
# Importing necessary functions from another .ipynb script
%run ui_elem.ipynb
```
----------------
## Text Entry
Here you should specify the column name (needs a better description) as well as give a name for the folder where the graphs and statistics should be stored.
```
direct_name_data
# Test to see if the value in the text boxes are saved
print("Sample column ID: " + sample_column_id.value)
print("Graph Output Directory Name: " + graph_output_dir.value)
print("Stats Output Directory Name: " + stats_output_dir.value)
```
--------------------------------------------------
## File Paths
Prior to running this portion of the script, be sure that all the files that will be used for analysis are uploaded to the data folder in the file directory where this script was opened from. Open the directory named "data", and upload all files that will be necessary to run the analysis. All the scripts that are required for the analysis are listed below.
Once the files have been uploaded, click on the circulary icon in the top left of the screen to restart the python kernal - Jupyter doesn't update the list below while the kernel is running, you will instead have to restart the kernel. Once restarted, the file should appear in the drop down menu below, where you can then select the corresponding file. The current placeholders will work just fine if you don't have a certain file on hand.
__*This section needs more work in terms of descriptions*__
#### Loading in resistome files
```
resistome_filenames
```
#### Loading in microbiome files
```
microbiome_filenames
```
#### Save the file names
This will also print the output so you can double check its accuracy
```
save_filepath_button
```
---
## Exploratory Variables
The following will allow you to enter in variables for the analyses. It's based on variables in your metadata.csv file that you want to use for EXPLORATORY analysis (NMDS, PCA, alpha rarefaction, barplots)
### Sliders
The slider allows you to choose how many separate analyses should be run, and how many of them should be AMR analyses and how many should be Microbiome.
### Explanatory Variables
Every time the slider value is changed, the code to generate the boxes should be rerun in order to update them. They will automatically contain the correct number of boxes based on the values entered in the slider.
**Name**:
**Subset**: Subset will allow you to filter out variables of interest and remove unnecessary variables. To select variables of interest, the format should be “*column-2 == column-variable-1*”. This is exactly how you might enter this into R.
To remove certain variables from the analysis, the format should be “*column_2 != column_variable_2*”. The key point is having the *!* symbol instead of the first exclamation point.
**Explanatory Variable**: This should also be the name of a column of interest for analysis.
*NOTE*: Exploratory variables cannot be numeric.
**Order**: This should describe the order that will be used during the analysis and when printing out result plots. *Each item in the list should be separated by a comma.*
```
display(exp_graph_var_amr)
display(exp_graph_var_microbiome)
var_info(exp_graph_var_amr.value, exp_graph_var_microbiome.value)
```
--------
## IMPORTANT STEP
Make sure to run the code below once you have finished entering the data into the boxes above. This will ensure that the data is stored correctly and will be output into the .csv file.
```
# Saves and prints the variables entered above into a list to be used when creating the .csv file
list_vals_a, list_vals_m = save_print_variables(exp_graph_var_amr.value, exp_graph_var_microbiome.value)
```
---
## Outputting information into .csv file
This below will now store everything entered above into a .csv file that can be read in by the analysis script. The .csv file will be stored in the current working directory where this script is also stored. If your analysis script is located in another directory, be sure to move the .csv file into that same directory before running that analysis script.
```
display(vars_save_button)
vars_save_button.on_click(vars_to_csv)
```
---
# Next Steps
The results from this script will be output in a .csv file that the R script will then use to run the rest of the analysis. You don't need to worry about the .csv file, all you need to do now is open the script named "Jupyter_metagenomic_analysis.ipynb" in the directory where this file was also located and run it.
| github_jupyter |
## Some tests with pgpointcloud
- LAS files are intractable for subsetting and visualisation without LASTools or Terrasolid or Bentley Map (or similar) which are awesome but costly.
- Further, LAS files don't really let us store all manner of things along with the data, for example we already have issues storing digitised waveforms with the ACT dataset
- so what can we do? Here are some experiments with PostGIS-pointcloud, one approach to point cloud data management
Some things about this exercise:
- I am a postGIS/pgpointcloud n00b. An SQL ninja could probably do a lot better than this and a lot faster!
- The data set used here has ~406 000 000 points in it
- made from nine ACT 8pt tiles (no waveform data, because stuff)
- ingested using a PDAL pipeline (http://pdal.io)
- LAS storage is ~20 GB
- PG-pointcloud table is ~5 GB, so reasonably similar to .LAZ
- there are 82 634 'patches' containing 5 000 points each
- patches are the primary query tool, so we index over 82 634 rows, which are arranged in a quasi-space-filling-curve
- tradeoff between number of points in patch, and number of rows in DB
- but generally speaking, scalable (nearly... research shows that we will hit a limit in a few hundred million points time)
### Gathering modules
```
import os
import psycopg2 as ppg
import numpy as np
import ast
from osgeo import ogr
import shapely as sp
from shapely.geometry import Point,Polygon,asShape
from shapely.wkt import loads as wkt_loads
from shapely import speedups
import cartopy as cp
import cartopy.crs as ccrs
import pandas as pd
import pandas.io.sql as pdsql
import geopandas as gp
from matplotlib.collections import PatchCollection
from mpl_toolkits.basemap import Basemap
import fiona
from descartes import PolygonPatch
import matplotlib.pyplot as plt
%matplotlib inline
speedups.available
speedups.enable()
```
## Making a postgres connection with psycopg2
```
# PGPASSWORD=pointy_cloudy psql -h localhost -d pcbm_pc -U pcbm
pg_connection = "dbname=pcbm_pc user=pcbm password=pointy_cloudy host=130.56.244.246"
conn = ppg.connect(pg_connection)
cursor = conn.cursor()
```
### First query for some blocks - this gets them all!
```
#blocks_query = "SELECT pa::geometry(Polygon, 28355) AS geom, PC_PatchAvg(pa, 'Z') AS elevation, id FROM act_patches;"
blocks_query = "SELECT st_astext(PC_envelope(pa)::geometry(Polygon, 28355)) AS geom, PC_PatchAvg(pa, 'Z') AS elevation, id FROM act_patches;"
%time blks = pdsql.read_sql(blocks_query, conn)
```
### here I was trying to convert a Pandas dataframe to a GeoPandas dataframe with a geometry column
```
gblocks = gp.GeoDataFrame(blks)
gblocks.head()
polys = gp.GeoSeries(blks.geom)
patches = gp.GeoDataFrame({'geometry': polys, 'elevation': blks.elevation})
patches.head()
```
### ...but I gave up and ingested data straight into a GeoPandas frame instead
```
blocks_query = "SELECT pa::geometry(Polygon, 28355) AS geom, PC_PatchAvg(pa, 'Z') AS elevation, id FROM act_patches where PC_PatchAvg(pa, 'Z') > 700;"
%time thepatches = gp.read_postgis(blocks_query, conn)
thepatches.head()
%time highpatches = thepatches.query('elevation > 820')
highpatches.head()
```
### Let's map the patches of data we collected - Black Mountain, above 820m high
```
highpatches.plot(column='elevation',colormap='BrBG')
```
### Now collect the points from the same region
```
points_query = "with pts as(SELECT PC_Explode(pa) as pt FROM act_patches where PC_PatchAvg(pa, 'Z') > 820 ) select st_astext(pt::geometry) from pts;"
#get raw point data, not as a geometry
#points_query = "SELECT PC_astext(PC_Explode(pa)) as pt FROM act_patches where PC_PatchAvg(pa, 'Z') > 700 ;"
%time pts = pdsql.read_sql(points_query, conn)
# point storage schema:
# 1 = intens, 2 = ReturnNo, 3 = Numreturns, 4 = scandirectionflag, 5 = edgeofflightline
# 6 = classification (ASPRS), 7 = scananglerank, 8 = user data, 9 = pointsourceID
# 10 = R, 11 = G, 12 = B, 13 = GPSTime, 14 = X, 15 = Y, 16 = Z
#how many points did we get?
pts.size
#had to check the schema to find point order...
schema_query = "SELECT * FROM pointcloud_formats where pcid = 4;"
schm = pdsql.read_sql(schema_query, conn)
print(schm.schema)
pts.head()
thepoints = []
for point in pts.st_astext:
this = wkt_loads(point)
thepoints.append([this.x,this.y,this.z])
thepoints = np.squeeze(thepoints)
```
### Plot the points
```
plt.scatter(thepoints[:,0], thepoints[:,1], c = thepoints[:,2], lw=0, s=5, cmap='BrBG')
```
### Now make a pretty plot - points, patches in the subset, and all the patches in the region
```
fig = plt.figure()
fig.set_size_inches(25/2.51, 25/2.51)
BLUE = '#6699cc'
RED = '#cc6699'
ax = fig.gca()
ax.scatter(thepoints[:,0], thepoints[:,1], c = thepoints[:,2], lw=0, s=3, cmap='BrBG')
for patch in thepatches.geom:
ax.add_patch(PolygonPatch(patch, fc=BLUE, ec=BLUE, alpha=0.2, zorder=2 ))
for patch in highpatches.geom:
ax.add_patch(PolygonPatch(patch, fc=BLUE, ec=RED, alpha=0.2, zorder=2 ))
```
## Selecting points by classification
```
#ASPRS class 6 - buildings
bldng_query = "WITH filtered_patch AS (SELECT PC_FilterEquals(pa, 'Classification', 6) as f_patch FROM act_patches where PC_PatchAvg(pa, 'Z') > 820) SELECT st_astext(point::geometry) FROM filtered_patch, pc_explode(f_patch) AS point;"
%time bld_pts = pdsql.read_sql(bldng_query, conn)
bld_pts.head()
bldpoints = []
for point in bld_pts.st_astext:
this = wkt_loads(point)
bldpoints.append([this.x,this.y,this.z])
bldpoints = np.squeeze(bldpoints)
#ASPRS class 2 - ground
grnd_query = "WITH filtered_patch AS (SELECT PC_FilterEquals(pa, 'Classification', 2) as f_patch FROM act_patches where PC_PatchAvg(pa, 'Z') > 820) SELECT st_astext(point::geometry) FROM filtered_patch, pc_explode(f_patch) AS point;"
%time grnd_pts = pdsql.read_sql(grnd_query, conn)
grnd_pts.head()
grndpoints = []
for point in grnd_pts.st_astext:
this = wkt_loads(point)
grndpoints.append([this.x,this.y,this.z])
grndpoints = np.squeeze(grndpoints)
#ASPRS class 5 - high vegetation
hv_query = "WITH filtered_patch AS (SELECT PC_FilterEquals(pa, 'Classification', 5) as f_patch FROM act_patches where PC_PatchAvg(pa, 'Z') > 820) SELECT st_astext(point::geometry) FROM filtered_patch, pc_explode(f_patch) AS point;"
%time hv_pts = pdsql.read_sql(hv_query, conn)
hv_pts.head()
hvpoints = []
for point in hv_pts.st_astext:
this = wkt_loads(point)
hvpoints.append([this.x,this.y,this.z])
hvpoints = np.squeeze(hvpoints)
fig = plt.figure()
fig.set_size_inches(25/2.51, 25/2.51)
BLUE = '#6699cc'
RED = '#cc6699'
ax = fig.gca()
ax.scatter(grndpoints[:,0], grndpoints[:,1], c = grndpoints[:,2], lw=0, s=3, cmap='plasma')
ax.scatter(bldpoints[:,0], bldpoints[:,1], c = bldpoints[:,2], lw=0, s=3, cmap='viridis')
ax.scatter(hvpoints[:,0], hvpoints[:,1], c = hvpoints[:,2], lw=0, s=3, cmap='BrBG')
for patch in thepatches.geom:
ax.add_patch(PolygonPatch(patch, fc=BLUE, ec=BLUE, alpha=0.2, zorder=2 ))
for patch in highpatches.geom:
ax.add_patch(PolygonPatch(patch, fc=BLUE, ec=RED, alpha=0.2, zorder=2 ))
```
### Add a 3D plot
```
#set up for 3d plots
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pylab as pylab
plt_az=300
plt_elev = 50.
plt_s = 2
cb_fmt = '%.1f'
fig = plt.figure()
fig.set_size_inches(30/2.51, 25/2.51)
ax0 = fig.add_subplot(111, projection='3d')
ax0.scatter(grndpoints[:,0], grndpoints[:,1],grndpoints[:,2], c=np.ndarray.tolist(grndpoints[:,2]),\
lw=0, s=plt_s, cmap='plasma')
ax0.scatter(bldpoints[:,0], bldpoints[:,1],bldpoints[:,2], c=np.ndarray.tolist(bldpoints[:,2]),\
lw=0, s=plt_s, cmap='viridis')
ax0.scatter(hvpoints[:,0], hvpoints[:,1],hvpoints[:,2], c=np.ndarray.tolist(hvpoints[:,2]),\
lw=0, s=plt_s-1, cmap='BrBG')
```
### Export to three.js?
```
import vtk
np.savetxt('ground_points.txt', grndpoints, delimiter=',')
np.savetxt('bld_points.txt', bldpoints, delimiter=',')
np.savetxt('hv_points.txt', hvpoints, delimiter=',')
```
### To do:
- too many things!
- select from database by:
- class (demonstrated here)
- height above ground (need to integrate PDAL and PCL)
- tree cover
- intersection with objects
- things on the list:
- comparing LandSAT bare ground and LIDAR bare ground
- tree heights and geophysical properties
- ...
### All very cool but why?
National elevation maps - storing and managing many billions of points as a coherent dataset for precise elevation estimation. Also aiming to store provenance - if there's a data issue, we need more than just points. We need to figure out why the issue occurred, and fix it. We can also store things like point accuracy, some QC metrics, whatever point attributes we like! Or points from manifold sources:
- airborne LiDAR
- terrestrial scanners
- 3D photogrammetry
- geophysical datasets (already as points in netCDF)
- output of discrete element models (eg. Kool et al, or new sea ice models in development)
| github_jupyter |
# Gaussian-process regression demo
Here we demonstrate how to use ``inference.gp.GpRegressor`` to perform Gaussian-process regression.
```
import matplotlib.pyplot as plt
from numpy import array, linspace
```
Suppose we have the following dataset:
```
x = array([0.5, 1.0, 1.5, 3.0, 3.5, 4.0, 4.5])
y = array([0.157, -0.150, -0.305, -0.049, 0.366, 0.417, 0.430])
```
Gaussian-process regression models data using 'covariance functions', also referred to as 'kernels'. The choice of kernel effectively sets certain assumptions about the underlying structure of the data.
For example, the `SquaredExponential` kernel assumes the underlying structure is smooth, and the `WhiteNoise` kernel assumes that our y-data values contain identical independent Gaussian noise. We'll now use these two kernels to show how the choice of kernel can have a strong impact on the regression estimate.
First, we construct a `GpRegressor` using only the `SquaredExponential` kernel:
```
from inference.gp import GpRegressor, SquaredExponential, WhiteNoise
GP1 = GpRegressor(x, y, kernel=SquaredExponential)
```
Calling an instance of `GpRegressor` as a function will return the mean and standard deviation of the regression estimate. We can now evaluate the estimate on a new axis to plot the result:
```
x_fit = linspace(0, 5, 200)
mu1, sigma1 = GP1(x_fit)
```
Now repeat this process but instead using a combination of the `SquaredExponential` and `WhiteNoise` kernels. We can generate a 'composite' kernel simply by addition as follows:
```
composite_kernel = SquaredExponential() + WhiteNoise()
GP2 = GpRegressor(x, y, kernel=composite_kernel)
mu2, sigma2 = GP2(x_fit)
```
Now we have both estimates we can plot them to compare the effect of the different kernels:
```
fig = plt.figure(figsize=(12,4))
ax1 = fig.add_subplot(121)
ax1.fill_between(x_fit, mu1 - sigma1, mu1 + sigma1, color='red', alpha=0.2, label='GPR uncertainty')
ax1.fill_between(x_fit, mu1 - 2*sigma1, mu1 + 2*sigma1, color='red', alpha=0.1)
ax1.plot(x_fit, mu1, lw=2, c='red', label='GPR mean')
ax1.plot(x, y, 'o', c='black', label='data values')
ax1.set_xlim([0,5])
ax1.set_ylim([-0.7,0.7])
ax1.set_title('SquaredExponential kernel only')
ax1.set_xlabel('x-data value')
ax1.set_ylabel('y-data value')
ax1.legend(loc=4)
ax1.grid()
ax2 = fig.add_subplot(122)
ax2.fill_between(x_fit, mu2 - sigma2, mu2 + sigma2, color='blue', alpha=0.2, label='GPR uncertainty')
ax2.fill_between(x_fit, mu2 - 2*sigma2, mu2 + 2*sigma2, color='blue', alpha=0.1)
ax2.plot(x_fit, mu2, lw=2, c='blue', label='GPR mean')
ax2.plot(x, y, 'o', c='black', label='data values')
ax2.set_xlim([0,5])
ax2.set_ylim([-0.7,0.7])
ax2.set_title('SquaredExponential + WhiteNoise kernel')
ax2.set_xlabel('x-data value')
ax2.set_yticklabels([])
ax2.legend(loc=4)
ax2.grid()
plt.tight_layout()
plt.show()
```
As the GP using the `SquaredExponential` kernel assumes the data are noiseless, the estimate must pass directly through each data point. Conversely, the GP using a combination of the `SquaredExponential` and `WhiteNoise` kernels is not forced to pass through each point as it is able to assume a level of noise present in the data.
In cases where the uncertainties on the data are known, they can be specified directly using the `y_err` keyword argument of `GpRegressor`. This is particularly useful if there are significant differences in uncertainties across the dataset.
To illustrate this we can manually specify a set of uncertainties for the example dataset to see how this affects the regression estimate:
```
y_errors = [0.1, 0.01, 0.1, 0.5, 0.1, 0.01, 0.1]
gpr = GpRegressor(x, y, y_err=y_errors, kernel=SquaredExponential)
mu, sig = gpr(x_fit)
fig = plt.figure()
ax = fig.add_subplot(111)
ax.fill_between(x_fit, mu - sig, mu + sig, color='green', alpha=0.2, label='GPR uncertainty')
ax.fill_between(x_fit, mu - 2*sig, mu + 2*sig, color='green', alpha=0.1)
ax.plot(x_fit, mu, lw=2, c='green', label='GPR mean')
ax.errorbar(x, y, yerr=y_errors, marker='o', color='black', ecolor='black', ls='none', label='data values')
ax.set_xlim([0,5])
ax.set_ylim([-0.7,0.7])
ax.set_title('SquaredExponential with specified errors')
ax.set_xlabel('x-data value', fontsize=11)
ax.set_ylabel('y-data value', fontsize=11)
ax.grid()
ax.legend(loc=4)
plt.tight_layout()
plt.show()
```
As the regression estimate at a collection of points is itself is a multivariate normal distribution, we can construct the full mean vector and covariance matrix which describes that distribution using the `build_posterior` method of `GpRegressor`.
This allows us to draw samples from the distribution:
```
# get the mean and covariance matrix
means, covar = gpr.build_posterior(x_fit)
# draw samples from the distribution
from numpy.random import multivariate_normal
samples = multivariate_normal(means, covar, 50)
# plot all the samples together
fig = plt.figure()
ax = fig.add_subplot(111)
ax.plot(x_fit, samples.T, lw=0.5)
ax.set_xlim([0,5])
ax.set_ylim([-0.7,0.7])
ax.set_title('50 samples drawn from the GP')
ax.set_xlabel('x-data value', fontsize=11)
ax.set_ylabel('y-data value', fontsize=11)
ax.grid()
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
import numpy as np
def rmetric(L, X, Y):
# L stacked list of 'L' pos. def. 2x2 untere-dreiecks-matrix(saved as (3*n)x1, L = ['L'[0,0],'L'[1,0],'L'[1,1])
# 'X','Y' 2x2 untere-dreiecks-matrizen, saved as above
# TODO: add area constant
diag = np.reciprocal(L**2)
diag[list(range(1,len(L),3))] = 1
# TODO: Concatenate with rot. matrix
A = np.diag(diag)
return X.T@A@Y
rmetric(np.array([1,2,3,3,4,6,7,8,9]), np.array([1,2,3,3,4,6,7,8,9]),np.array([1,2,3,3,4,6,7,8,9]))
def rexp(L,X):
mask = list(range(1,len(L),3))
cmask = np.ones(L.shape).astype(bool)
cmask[mask] = 0
E = L
E[mask] += X[mask]
E[cmask] *= np.exp(X[cmask]*np.reciprocal(L[cmask]))
return E
rexp(np.array([1,2,3,3,4,6,7,8,9]).astype(float), np.array([1,2,3,3,4,6,7,8,9]).astype(float))
def rlog(L,X):
mask = list(range(1,len(L),3))
cmask = np.ones(L.shape).astype(bool)
cmask[mask] = 0
E = np.zeros(L.shape)
E[mask] = X[mask] - L[mask]
E[cmask] = L[cmask]*np.log(X[cmask]*np.reciprocal(L[cmask]))
return E
def Dcinvers(L,X):
mask1 = list(range(0, len(L), 3))
mask2 = list(range(1, len(L), 3))
mask3 = list(range(2, len(L), 3))
D = 2*L*X
D[mask3] += D[mask2]
D[mask2] = X[mask1]*L[mask2] + L[mask1]*X[mask2]
return D
Dcinvers(np.array([1,2,3,3,4,6,7,8,9]).astype(float), np.array([1,2,3,3,4,6,7,8,9]).astype(float))
def Cholesky(P):
mask1 = list(range(0, len(P), 3))
mask2 = list(range(1, len(P), 3))
mask3 = list(range(2, len(P), 3))
L = P
L[mask1] = np.sqrt(P[mask1])
L[mask2] = np.divide(P[mask2],L[mask1])
L[mask3] = np.sqrt(P[mask3]- L[mask2]**2)
return L
Cholesky(np.array([1,2,3]).astype(float))
def inv_L(L):
mask1 = list(range(0, len(L), 3))
mask2 = list(range(1, len(L), 3))
mask3 = list(range(2, len(L), 3))
IL = L
IL[mask1+mask3] = np.reciprocal(L[mask1+mask3])
IL[mask2] = -L[mask2]*IL[mask1]*IL[mask3]
return IL
inv_L(np.array([1,2,3,4,5,6]).astype(float))
def vector_to_matrix(vector, group = None):
lengthy_v = np.zeros(int((len(vector)/3)*4))
mask_without = [i for i in range(len(lengthy_v)) if i % 4 != 1]
mask2 = list(range(1,len(lengthy_v),4))
mask3 =list(range(2,len(lengthy_v),4))
lengthy_v[mask_without] = vector
if group == 'sym':
lengthy_v[mask2] = lengthy_v[mask3]
return np.reshape(lengthy_v,(-1,2,2), order = 'A')
def matrix_to_vector(matrix):
lengthy_v = np.reshape(matrix, (-1), order = 'A')
mask_without = [i for i in range(len(lengthy_v)) if i % 4 != 1]
v = lengthy_v[mask_without]
return v
a = vector_to_matrix(np.array([1,2,3,3,4,6,7,8,9]).astype(float))
b = a
print(a)
c = a @ b
print(c)
def Dc(P,W):
L = vector_to_matrix(Cholesky(P))
print(Cholesky(P))
Linv = np.linalg.inv(L)
print(Linv)
W = vector_to_matrix(W, group = 'sym')
print(Linv.transpose(0,2,1).shape)
inner = Linv@W@np.transpose(Linv, axes= (0,2,1))
vecInner = matrix_to_vector(inner)
mask1 = list(range(0, len(vecInner), 3))
mask3 = list(range(2, len(vecInner), 3))
vecInner[mask1+mask3] /= 2
inner = vector_to_matrix(vecInner)
D = L@inner
return matrix_to_vector(D)
Dc(np.array([1,2,1,3,4,1,7,8,1]).astype(float), np.array([1,2,1,3,4,1,7,8,1]).astype(float))
```
| github_jupyter |
## Градиентный бустинг
Сравним, как ведут себя бустинг и бэггинг с ростом числа базовых алгоритмов.
В случае бэггинга все базовые алгоритмы настраиваются на различные выборки из одного и того же распределения на $\mathbb{X} \times \mathbb{Y}$. При этом некоторые из них могут оказаться переобученными, однако усреднение позволяет ослабить этот эффект (объясняется тем, что для некоррелированных алгоритмов разброс композиции оказывается в $N$ раз меньше разброса отдельных алгоритмов, т.е. много деревьев с меньшей вероятностью настроятся на некоторый нетипичный объект по сравнению с одним деревом). Если $N$ достаточно велико, то последующие добавления новых алгоритмов уже не позволят улучшить качество модели.
В случае же бустинга каждый алгоритм настраивается на ошибки всех предыдущих, это позволяет на каждом шаге настраиваться на исходное распределение все точнее и точнее. Однако при достаточно большом $N$ это является источником переобучения, поскольку последующие добавления новых алгоритмов будут продолжать настраиваться на обучающую выборку, уменьшая ошибку на ней, при этом уменьшая обобщающую способность итоговой композиции.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
X_train = np.linspace(0, 1, 100)
X_test = np.linspace(0, 1, 1000)
@np.vectorize
def target(x):
return x > 0.5
Y_train = target(X_train) + np.random.randn(*X_train.shape) * 0.1
plt.figure(figsize = (16, 9))
plt.scatter(X_train, Y_train, s=50)
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import BaggingRegressor, GradientBoostingRegressor
reg = BaggingRegressor(DecisionTreeRegressor(max_depth=2), warm_start=True)
plt.figure(figsize=(20, 30))
sizes = [1, 2, 5, 20, 100, 500, 1000, 2000]
for i, s in enumerate(sizes):
reg.n_estimators = s
reg.fit(X_train.reshape(-1, 1), Y_train)
plt.subplot(4, 2, i+1)
plt.xlim([0, 1])
plt.scatter(X_train, Y_train, s=30)
plt.plot(X_test, reg.predict(X_test.reshape(-1, 1)), c='green', linewidth=4)
plt.title('{} trees'.format(s))
```
Можно заметить, что с некоторого момента итоговая функция перестает меняться с ростом количества деревьев.
Теперь проделаем то же самое для градинентного бустинга.
```
reg = GradientBoostingRegressor(max_depth=1, learning_rate=1, warm_start=True)
plt.figure(figsize=(20, 30))
sizes = [1, 2, 5, 20, 100, 500, 1000, 2000]
for i, s in enumerate(sizes):
reg.n_estimators = s
reg.fit(X_train.reshape(-1, 1), Y_train)
plt.subplot(4, 2, i+1)
plt.xlim([0, 1])
plt.scatter(X_train, Y_train, s=30)
plt.plot(X_test, reg.predict(X_test.reshape(-1, 1)), c='green', linewidth=4)
plt.title('{} trees'.format(s))
```
Градиентный бустинг довольно быстро построил истинную зависимость, после чего начал настраиваться уже на конкретные объекты обучающей выборки, из-за чего сильно переобучился.
Бороться с этой проблемой можно с помощью выбора очень простого базового алгоритма или
же искусственным снижением веса новых алгоритмов при помощи шага $\eta$:
$$a_N(x) = \sum_{n=0}^N \eta \gamma_N b_n(x).$$
Такая поправка замедляет обучение по сравнению с бэггингом, но зато позволяет получить менее переобученный алгоритм. Тем не менее, важно понимать, что переобучение всё равно будет иметь место при обучении сколь угодно большого количества базовых алгоритмов для фиксированного $\eta$.
```
reg = GradientBoostingRegressor(max_depth=1, learning_rate=0.1, warm_start=True)
plt.figure(figsize=(20, 30))
sizes = [1, 2, 5, 20, 100, 500, 1000, 2000]
for i, s in enumerate(sizes):
reg.n_estimators = s
reg.fit(X_train.reshape(-1, 1), Y_train)
plt.subplot(4, 2, i+1)
plt.xlim([0, 1])
plt.scatter(X_train, Y_train, s=30)
plt.plot(X_test, reg.predict(X_test.reshape(-1, 1)), c='green', linewidth=4)
plt.title('{} trees'.format(s))
```
Теперь проверим описанный выше эффект на реальных данных.
```
from sklearn import datasets
from sklearn.model_selection import train_test_split
ds = datasets.load_diabetes()
X = ds.data
Y = ds.target
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.5)
MAX_ESTIMATORS = 300
gbclf = BaggingRegressor(warm_start=True)
err_train_bag = []
err_test_bag = []
for i in range(1, MAX_ESTIMATORS+1):
gbclf.n_estimators = i
gbclf.fit(X_train, Y_train)
err_train_bag.append(1 - gbclf.score(X_train, Y_train))
err_test_bag.append(1 - gbclf.score(X_test, Y_test))
gbclf = GradientBoostingRegressor(warm_start=True, max_depth=2, learning_rate=0.1)
err_train_gb = []
err_test_gb = []
for i in range(1, MAX_ESTIMATORS+1):
gbclf.n_estimators = i
gbclf.fit(X_train, Y_train)
err_train_gb.append(1 - gbclf.score(X_train, Y_train))
err_test_gb.append(1 - gbclf.score(X_test, Y_test))
plt.figure(figsize=(10, 4))
plt.subplot(1, 2, 1)
plt.plot(err_train_gb, label='GB')
plt.plot(err_train_bag, label='Bagging')
plt.legend()
plt.title('Train')
plt.subplot(1, 2, 2)
plt.plot(err_test_gb, label='GB')
plt.plot(err_test_bag, label='Bagging')
plt.legend()
plt.title('Test')
plt.gcf().set_size_inches(15,7)
```
## Градиентный бустинг и случайные леса
Сравним поведение двух методов построения композиции алгоритмов над деревьями на примере задачи [Kaggle: Predicting a Biological Response](https://www.kaggle.com/c/bioresponse):
```
import pandas as pd
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
data = pd.read_csv('class13.csv')
X = data.iloc[:, 1:].values
y = data.iloc[:, 0].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.8, random_state=241)
%%time
gbm = GradientBoostingClassifier(n_estimators=250, learning_rate=0.2, verbose=True).fit(X_train, y_train)
import numpy as np
from sklearn.metrics import roc_auc_score
for learning_rate in [1, 0.5, 0.3, 0.2, 0.1]:
gbm = GradientBoostingClassifier(n_estimators=150,
learning_rate=learning_rate,
random_state=241).fit(X_train, y_train)
l = roc_auc_score
test_deviance = np.zeros((gbm.n_estimators, ), dtype=np.float64)
for i, y_pred in enumerate(gbm.staged_decision_function(X_test)):
y_pred = 1.0 / (1.0 + np.exp(-y_pred))
test_deviance[i] = roc_auc_score(y_test, y_pred)
train_deviance = np.zeros((gbm.n_estimators, ), dtype=np.float64)
for i, y_pred in enumerate(gbm.staged_decision_function(X_train)):
y_pred = 1.0 / (1.0 + np.exp(-y_pred))
train_deviance[i] = roc_auc_score(y_train, y_pred)
plt.figure()
plt.plot(test_deviance, 'r', linewidth=2)
plt.plot(train_deviance, 'g', linewidth=2)
plt.legend(['test', 'train'])
plt.title('GBM eta=%.1f, test roc-auc=%.3f, best_est=%d' %
(learning_rate, test_deviance.max(), test_deviance.argmax() + 1))
plt.xlabel('Number of trees')
plt.ylabel('Metric')
```
Итого, лучшая композиция построена при $\eta = 0.1$, включает 24 базовых алгоритма и достигает значения 0.816 на контрольной выборке. При этом случайный лес с таким же количеством базовых алгоритмов уступает градиентному бустингу:
```
rf = RandomForestClassifier(n_estimators=24, random_state=0).fit(X_train, y_train)
print ('Train RF ROC-AUC =', roc_auc_score(y_train, rf.predict_proba(X_train)[:,1]))
print ('Test RF ROC-AUC = ', roc_auc_score(y_test, rf.predict_proba(X_test)[:,1]))
```
Заметим также, что при всём этом случайный лес, в отличие от градиентного бустинга, использует глубокие деревья, требующие вычислительных мощностей для их обучения.
Для достижения такого же качества случайному лесу требуется гораздо большее число базовых алгоритмов:
```
for n_estimators in range(10, 101, 10):
rf = RandomForestClassifier(n_estimators=n_estimators,
n_jobs=4).fit(X_train, y_train)
print(n_estimators, 'trees: train ROC-AUC =',
roc_auc_score(y_train,
rf.predict_proba(X_train)[:, 1]), 'test ROC-AUC =',
roc_auc_score(y_test,
rf.predict_proba(X_test)[:, 1]))
```
Напоследок можно посмотреть [визуализацию](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html) градиентного бустинга для решающих деревьев различной глубины для функций различного вида.
| github_jupyter |
# Python HANA ML API
<div class="alert alert-block alert-info">
<b>Extracting information by partition from a trained model.</b> <br>
</div>
## Train the model
### Create an HANA Dataframe for the training data
```
# Connect using the HANA secure user store
from hana_ml import dataframe as hd
conn = hd.ConnectionContext(userkey='MLMDA_KEY')
# Get Training Data
sql_cmd = 'SELECT * FROM "APL_SAMPLES"."AUTO_CLAIMS_FRAUD" ORDER BY CLAIM_ID'
training_data = hd.DataFrame(conn, sql_cmd)
```
### Put a subset of the data in a Pandas Dataframe and display it
```
training_data.head(10).collect()
```
### Build a Classification model with APL Ridge Regression
```
# Create the model
from hana_ml.algorithms.apl.classification import AutoClassifier
model = AutoClassifier(conn_context=conn)
# Train the model
model.set_params(cutting_strategy = 'random')
model.fit(training_data, label='IS_FRAUD', key='CLAIM_ID')
```
## Debrief the trained model
### Define a function to call APL_GET_MODEL_INFO
```
def create_artifact_table(conn, table_name, table_spec):
conn = model.conn_context.connection
cursor = conn.cursor()
try:
cursor.execute(f'drop table {table_name}')
except:
pass
cursor.execute(f'create local temporary table {table_name} {table_spec}')
def get_model_info(model):
conn = model.conn_context.connection
cursor = conn.cursor()
model_table_name = model.model_table_.name # the temp table where the model is saved
# --- Create temp tables for input / output
create_artifact_table(conn=conn,
table_name='#FUNC_HEADER',
table_spec='(KEY NVARCHAR(50), VALUE NVARCHAR(255))')
create_artifact_table(conn=conn,
table_name='#OPERATION_CONFIG',
table_spec='(KEY NVARCHAR(1000), VALUE NCLOB, CONTEXT NVARCHAR(100))')
create_artifact_table(conn=conn,
table_name='#SUMMARY',
table_spec='(OID NVARCHAR(50), KEY NVARCHAR(100), VALUE NVARCHAR(100))')
create_artifact_table(conn=conn,
table_name='#VARIABLE_ROLES_COMP',
table_spec='(NAME NVARCHAR(255), ROLE NVARCHAR(10), COMPOSITION_TYPE VARCHAR(10), COMPONENT_NAME VARCHAR(255))')
create_artifact_table(conn=conn,
table_name='#VARIABLE_DESC',
table_spec='(RANK INTEGER,NAME VARCHAR(255),STORAGE VARCHAR(10),VALUETYPE VARCHAR(10),KEYLEVEL INTEGER,ORDERLEVEL INTEGER,MISSINGSTRING VARCHAR(255),GROUPNAME VARCHAR(255),DESCRIPTION VARCHAR(255), OID NVARCHAR(50))')
create_artifact_table(conn=conn,
table_name='#INDICATORS_DATASET',
table_spec='(OID VARCHAR(50),VARIABLE VARCHAR(255),TARGET VARCHAR(255),KEY VARCHAR(100),'
'VALUE NCLOB,DETAIL NCLOB,DATASET VARCHAR(255))')
create_artifact_table(conn=conn,
table_name='#PROFITCURVES',
table_spec='(OID VARCHAR(50), TYPE VARCHAR(100), VARIABLE VARCHAR(255), TARGET VARCHAR(255), '
'"Label" VARCHAR(255), "Frequency" VARCHAR(100), "Random" VARCHAR(100), "Wizard" VARCHAR(100), '
'"Estimation" VARCHAR(100), "Validation" VARCHAR(100), "Test" VARCHAR(100), "ApplyIn" VARCHAR(100))')
create_artifact_table(conn=conn,
table_name='#OUTPUT_TABLE_TYPE',
table_spec='(OID VARCHAR(50), POSITION INTEGER, NAME VARCHAR(255), KIND VARCHAR(50), '
'PRECISION INTEGER, SCALE INTEGER, MAXIMUM_LENGTH INTEGER)')
# Call APL
sql = 'call "_SYS_AFL".APL_GET_MODEL_INFO(#FUNC_HEADER, {model_table_name}, #OPERATION_CONFIG, #SUMMARY, #VARIABLE_ROLES_COMP, #VARIABLE_DESC, #INDICATORS_DATASET, #PROFITCURVES) with overview'
sql = sql.format(model_table_name=model_table_name)
# print(sql)
cursor.execute(sql)
```
### Calling APL_GET_MODEL_INFO
```
get_model_info(model)
```
### Put indicators data in a Pandas Dataframe
```
sql_cmd = 'SELECT * FROM #INDICATORS_DATASET'
indicators_data = hd.DataFrame(conn, sql_cmd)
```
### Show accuracy by partition
```
df = indicators_data.filter("KEY = 'AUC' and VARIABLE = 'rr_IS_FRAUD'").collect()
df = df[['DATASET','TARGET','KEY','VALUE']]
df['VALUE'] = df['VALUE'].astype(float)
df.columns = ['Partition','Target', 'Metric','Value']
df.style.format({'Value':'{0:,.3f}'}).hide_index()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.