text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
%load_ext autoreload
%autoreload 2
import jax
import jax.numpy as jnp
import numpy as np
from jax import random, jit, grad
import scipy
import cr.sparse as crs
from cr.sparse import la
from cr.sparse import dict
from cr.sparse import pursuit
from cr.sparse import data
```
# Dictionary Setup
```
M = 32
N = 64
K = 3
key = random.PRNGKey(0)
Phi = dict.gaussian_mtx(key, M,N)
Phi.shape
dict.coherence(Phi)
```
# Signal Setup
```
x, omega = data.sparse_normal_representations(key, N, K, 1)
x = jnp.squeeze(x)
x
omega, omega.shape
y = Phi @ x
y
```
# Development of OMP algorithm
## First iteration
```
r = y
norm_y_sqr = r.T @ r
norm_r_sqr = norm_y_sqr
norm_r_sqr
p = Phi.T @ y
p, p.shape
h = p
h, h.shape
i = pursuit.abs_max_idx(h)
i
indices = jnp.array([i])
indices, indices.shape
atom = Phi[:, i]
atom, atom.shape
subdict = jnp.expand_dims(atom, axis=1)
subdict.shape
L = jnp.ones((1,1))
L, L.shape
p_I = p[indices]
p_I, p_I.shape
x_I = p_I
x_I, x_I.shape
r_new = y - subdict @ x_I
r_new, r_new.shape
norm_r_new_sqr = r_new.T @ r_new
norm_r_new_sqr
```
## Second iteration
```
r = r_new
norm_r_sqr = norm_r_new_sqr
h = Phi.T @ r
h, h.shape
i = pursuit.abs_max_idx(h)
i
indices = jnp.append(indices, i)
indices
atom = Phi[:, i]
atom, atom.shape
b = subdict.T @ atom
b
L = pursuit.gram_chol_update(L, b)
L, L.shape
subdict = jnp.hstack((subdict, jnp.expand_dims(atom,1)))
subdict, subdict.shape
p_I = p[indices]
p_I, p_I.shape
x_I = la.solve_spd_chol(L, p_I)
x_I, x_I.shape
subdict.shape, x_I.shape
r_new = y - subdict @ x_I
r_new, r_new.shape
norm_r_new_sqr = r_new.T @ r_new
norm_r_new_sqr
```
## 3rd iteration
```
r = r_new
norm_r_sqr = norm_r_new_sqr
h = Phi.T @ r
h, h.shape
i = pursuit.abs_max_idx(h)
i
indices = jnp.append(indices, i)
indices
atom = Phi[:, i]
atom, atom.shape
b = subdict.T @ atom
b
L = pursuit.gram_chol_update(L, b)
L, L.shape
subdict = jnp.hstack((subdict, jnp.expand_dims(atom,1)))
subdict, subdict.shape
p_I = p[indices]
p_I, p_I.shape
x_I = la.solve_spd_chol(L, p_I)
x_I, x_I.shape
r_new = y - subdict @ x_I
r_new, r_new.shape
norm_r_new_sqr = r_new.T @ r_new
norm_r_new_sqr
from cr.sparse.pursuit import omp
solution = omp.solve(Phi, y, K)
solution.x_I
solution.I
solution.r
solution.r_norm_sqr
solution.iterations
def time_solve():
solution = omp.solve(Phi, y, K)
solution.x_I.block_until_ready()
solution.r.block_until_ready()
solution.I.block_until_ready()
solution.r_norm_sqr.block_until_ready()
%timeit time_solve()
omp_solve = jax.jit(omp.solve, static_argnums=(2))
sol = omp_solve(Phi, y, K)
sol.r_norm_sqr
def time_solve_jit():
solution = omp_solve(Phi, y, K)
solution.x_I.block_until_ready()
solution.r.block_until_ready()
solution.I.block_until_ready()
solution.r_norm_sqr.block_until_ready()
%timeit time_solve_jit()
14.3 * 1000 / 49.3
```
| github_jupyter |
```
# general imports
import cv2
import math
import numpy as np
import random
# reinforcement learning related imports
import re
import atari_py as ap
from collections import deque
from gym import make, ObservationWrapper, Wrapper
from gym.spaces import Box
# pytorch imports
import torch
import torch.nn as nn
from torch import save
from torch.optim import Adam
class ConvDQN(nn.Module):
def __init__(self, ip_sz, tot_num_acts):
super(ConvDQN, self).__init__()
self._ip_sz = ip_sz
self._tot_num_acts = tot_num_acts
self.cnv1 = nn.Conv2d(ip_sz[0], 32, kernel_size=8, stride=4)
self.rl = nn.ReLU()
self.cnv2 = nn.Conv2d(32, 64, kernel_size=4, stride=2)
self.cnv3 = nn.Conv2d(64, 64, kernel_size=3, stride=1)
self.fc1 = nn.Linear(self.feat_sz, 512)
self.fc2 = nn.Linear(512, tot_num_acts)
def forward(self, x):
op = self.cnv1(x)
op = self.rl(op)
op = self.cnv2(op)
op = self.rl(op)
op = self.cnv3(op)
op = self.rl(op).view(x.size()[0], -1)
op = self.fc1(op)
op = self.rl(op)
op = self.fc2(op)
return op
@property
def feat_sz(self):
x = torch.zeros(1, *self._ip_sz)
x = self.cnv1(x)
x = self.rl(x)
x = self.cnv2(x)
x = self.rl(x)
x = self.cnv3(x)
x = self.rl(x)
return x.view(1, -1).size(1)
def perf_action(self, stt, eps, dvc):
if random.random() > eps:
stt = torch.from_numpy(np.float32(stt)).unsqueeze(0).to(dvc)
q_val = self.forward(stt)
act = q_val.max(1)[1].item()
else:
act = random.randrange(self._tot_num_acts)
return act
def calc_temp_diff_loss(mdl, tgt_mdl, bch, gm, dvc):
st, act, rwd, nxt_st, fin = bch
st = torch.from_numpy(np.float32(st)).to(dvc)
nxt_st = torch.from_numpy(np.float32(nxt_st)).to(dvc)
act = torch.from_numpy(act).to(dvc)
rwd = torch.from_numpy(rwd).to(dvc)
fin = torch.from_numpy(fin).to(dvc)
q_vals = mdl(st)
nxt_q_vals = tgt_mdl(nxt_st)
q_val = q_vals.gather(1, act.unsqueeze(-1)).squeeze(-1)
nxt_q_val = nxt_q_vals.max(1)[0]
exp_q_val = rwd + gm * nxt_q_val * (1 - fin)
loss = (q_val - exp_q_val.data.to(dvc)).pow(2).mean()
loss.backward()
def upd_eps(epd):
last_eps = EPS_FINL
first_eps = EPS_STRT
eps_decay = EPS_DECAY
eps = last_eps + (first_eps - last_eps) * math.exp(-1 * ((epd + 1) / eps_decay))
return eps
def models_init(env, dvc):
mdl = ConvDQN(env.observation_space.shape, env.action_space.n).to(dvc)
tgt_mdl = ConvDQN(env.observation_space.shape, env.action_space.n).to(dvc)
return mdl, tgt_mdl
def gym_to_atari_format(gym_env):
return re.sub(r"(?<!^)(?=[A-Z])", "_", gym_env).lower()
def check_atari_env(env):
for f in ["Deterministic", "ramDeterministic", "ram", "NoFrameskip", "ramNoFrameSkip"]:
env = env.replace(f, "")
env = re.sub(r"-v\d+", "", env)
env = gym_to_atari_format(env)
return True if env in ap.list_games() else False
class RepBfr:
def __init__(self, cap_max):
self._bfr = deque(maxlen=cap_max)
def push(self, st, act, rwd, nxt_st, fin):
self._bfr.append((st, act, rwd, nxt_st, fin))
def smpl(self, bch_sz):
idxs = np.random.choice(len(self._bfr), bch_sz, False)
bch = zip(*[self._bfr[i] for i in idxs])
st, act, rwd, nxt_st, fin = bch
return (np.array(st), np.array(act), np.array(rwd, dtype=np.float32),
np.array(nxt_st), np.array(fin, dtype=np.uint8))
def __len__(self):
return len(self._bfr)
class TrMetadata:
def __init__(self):
self._avg = 0.0
self._bst_rwd = -float("inf")
self._bst_avg = -float("inf")
self._rwds = []
self._avg_rng = 100
self._idx = 0
@property
def bst_rwd(self):
return self._bst_rwd
@property
def bst_avg(self):
return self._bst_avg
@property
def avg(self):
avg_rng = self._avg_rng * -1
return sum(self._rwds[avg_rng:]) / len(self._rwds[avg_rng:])
@property
def idx(self):
return self._idx
def _upd_bst_rwd(self, epd_rwd):
if epd_rwd > self.bst_rwd:
self._bst_rwd = epd_rwd
def _upd_bst_avg(self):
if self.avg > self.bst_avg:
self._bst_avg = self.avg
return True
return False
def upd_rwds(self, epd_rwd):
self._rwds.append(epd_rwd)
self._upd_bst_rwd(epd_rwd)
return self._upd_bst_avg()
def upd_idx(self):
self._idx += 1
class CCtrl(Wrapper):
def __init__(self, env, is_atari):
super(CCtrl, self).__init__(env)
self._is_atari = is_atari
def reset(self):
if self._is_atari:
return self.env.reset()
else:
self.env.reset()
return self.env.render(mode="rgb_array")
class FrmDwSmpl(ObservationWrapper):
def __init__(self, env):
super(FrmDwSmpl, self).__init__(env)
self.observation_space = Box(low=0, high=255, shape=(84, 84, 1), dtype=np.uint8)
self._width = 84
self._height = 84
def observation(self, observation):
frame = cv2.cvtColor(observation, cv2.COLOR_RGB2GRAY)
frame = cv2.resize(frame, (self._width, self._height), interpolation=cv2.INTER_AREA)
return frame[:, :, None]
class MaxNSkpEnv(Wrapper):
def __init__(self, env, atari, skip=4):
super(MaxNSkpEnv, self).__init__(env)
self._obs_buffer = deque(maxlen=2)
self._skip = skip
self._atari = atari
def step(self, act):
total_rwd = 0.0
fin = None
for _ in range(self._skip):
obs, rwd, fin, log = self.env.step(act)
if not self._atari:
obs = self.env.render(mode="rgb_array")
self._obs_buffer.append(obs)
total_rwd += rwd
if fin:
break
max_frame = np.max(np.stack(self._obs_buffer), axis=0)
return max_frame, total_rwd, fin, log
def reset(self):
self._obs_buffer.clear()
obs = self.env.reset()
self._obs_buffer.append(obs)
return obs
class FrRstEnv(Wrapper):
def __init__(self, env):
Wrapper.__init__(self, env)
if len(env.unwrapped.get_action_meanings()) < 3:
raise ValueError("min required action space of 3!")
def reset(self, **kwargs):
self.env.reset(**kwargs)
obs, _, fin, _ = self.env.step(1)
if fin:
self.env.reset(**kwargs)
obs, _, fin, _ = self.env.step(2)
if fin:
self.env.reset(**kwargs)
return obs
def step(self, act):
return self.env.step(act)
class FrmBfr(ObservationWrapper):
def __init__(self, env, num_steps, dtype=np.float32):
super(FrmBfr, self).__init__(env)
obs_space = env.observation_space
self._dtype = dtype
self.observation_space = Box(obs_space.low.repeat(num_steps, axis=0),
obs_space.high.repeat(num_steps, axis=0), dtype=self._dtype)
def reset(self):
self.buffer = np.zeros_like(self.observation_space.low, dtype=self._dtype)
return self.observation(self.env.reset())
def observation(self, observation):
self.buffer[:-1] = self.buffer[1:]
self.buffer[-1] = observation
return self.buffer
class Img2Trch(ObservationWrapper):
def __init__(self, env):
super(Img2Trch, self).__init__(env)
obs_shape = self.observation_space.shape
self.observation_space = Box(low=0.0, high=1.0, shape=(obs_shape[::-1]), dtype=np.float32)
def observation(self, observation):
return np.moveaxis(observation, 2, 0)
class NormFlts(ObservationWrapper):
def observation(self, obs):
return np.array(obs).astype(np.float32) / 255.0
def wrap_env(env_ip):
env = make(env_ip)
is_atari = check_atari_env(env_ip)
env = CCtrl(env, is_atari)
env = MaxNSkpEnv(env, is_atari)
try:
env_acts = env.unwrapped.get_action_meanings()
if "FIRE" in env_acts:
env = FrRstEnv(env)
except AttributeError:
pass
env = FrmDwSmpl(env)
env = Img2Trch(env)
env = FrmBfr(env, 4)
env = NormFlts(env)
return env
def upd_grph(mdl, tgt_mdl, opt, rpl_bfr, dvc, log):
if len(rpl_bfr) > INIT_LEARN:
if not log.idx % TGT_UPD_FRQ:
tgt_mdl.load_state_dict(mdl.state_dict())
opt.zero_grad()
bch = rpl_bfr.smpl(B_S)
calc_temp_diff_loss(mdl, tgt_mdl, bch, G, dvc)
opt.step()
def fin_epsd(mdl, env, log, epd_rwd, epd, eps):
bst_so_fat = log.upd_rwds(epd_rwd)
if bst_so_fat:
print(f"checkpointing current model weights. highest running_average_reward of\
{round(log.bst_avg, 3)} achieved!")
save(mdl.state_dict(), f"{env}.dat")
print(f"episode_num {epd}, curr_reward: {epd_rwd}, best_reward: {log.bst_rwd},\
running_avg_reward: {round(log.avg, 3)}, curr_epsilon: {round(eps, 4)}")
def run_epsd(env, mdl, tgt_mdl, opt, rpl_bfr, dvc, log, epd):
epd_rwd = 0.0
st = env.reset()
while True:
eps = upd_eps(log.idx)
act = mdl.perf_action(st, eps, dvc)
if True:
env.render()
nxt_st, rwd, fin, _ = env.step(act)
rpl_bfr.push(st, act, rwd, nxt_st, fin)
st = nxt_st
epd_rwd += rwd
log.upd_idx()
upd_grph(mdl, tgt_mdl, opt, rpl_bfr, dvc, log)
if fin:
fin_epsd(mdl, ENV, log, epd_rwd, epd, eps)
break
def train(env, mdl, tgt_mdl, opt, rpl_bfr, dvc):
log = TrMetadata()
for epd in range(N_EPDS):
run_epsd(env, mdl, tgt_mdl, opt, rpl_bfr, dvc, log, epd)
B_S = 64
ENV = "Pong-v4"
EPS_STRT = 1.0
EPS_FINL = 0.005
EPS_DECAY = 100000
G = 0.99
INIT_LEARN = 10000
LR = 1e-4
MEM_CAP = 20000
N_EPDS = 50000
TGT_UPD_FRQ = 1000
env = wrap_env(ENV)
dvc = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
mdl, tgt_mdl = models_init(env, dvc)
opt = Adam(mdl.parameters(), lr=LR)
rpl_bfr = RepBfr(MEM_CAP)
train(env, mdl, tgt_mdl, opt, rpl_bfr, dvc)
env.close()
```
| github_jupyter |
# Structured and time series data
This notebook contains an implementation of the third place result in the Rossman Kaggle competition as detailed in Guo/Berkhahn's [Entity Embeddings of Categorical Variables](https://arxiv.org/abs/1604.06737).
The motivation behind exploring this architecture is it's relevance to real-world application. Most data used for decision making day-to-day in industry is structured and/or time-series data. Here we explore the end-to-end process of using neural networks with practical structured data problems.
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
from fastai.structured import *
from fastai.column_data import *
np.set_printoptions(threshold=50, edgeitems=20)
PATH='data/rossmann/'
```
## Create datasets
In addition to the provided data, we will be using external datasets put together by participants in the Kaggle competition. You can download all of them [here](http://files.fast.ai/part2/lesson14/rossmann.tgz).
For completeness, the implementation used to put them together is included below.
```
def concat_csvs(dirname):
path = f'{PATH}{dirname}'
filenames=glob(f"{PATH}/*.csv")
wrote_header = False
with open(f"{path}.csv","w") as outputfile:
for filename in filenames:
name = filename.split(".")[0]
with open(filename) as f:
line = f.readline()
if not wrote_header:
wrote_header = True
outputfile.write("file,"+line)
for line in f:
outputfile.write(name + "," + line)
outputfile.write("\n")
# concat_csvs('googletrend')
# concat_csvs('weather')
```
Feature Space:
* train: Training set provided by competition
* store: List of stores
* store_states: mapping of store to the German state they are in
* List of German state names
* googletrend: trend of certain google keywords over time, found by users to correlate well w/ given data
* weather: weather
* test: testing set
```
table_names = ['train', 'store', 'store_states', 'state_names',
'googletrend', 'weather', 'test']
```
We'll be using the popular data manipulation framework `pandas`. Among other things, pandas allows you to manipulate tables/data frames in python as one would in a database.
We're going to go ahead and load all of our csv's as dataframes into the list `tables`.
```
tables = [pd.read_csv(f'{PATH}{fname}.csv', low_memory=False) for fname in table_names]
from IPython.display import HTML, display
```
We can use `head()` to get a quick look at the contents of each table:
* train: Contains store information on a daily basis, tracks things like sales, customers, whether that day was a holdiay, etc.
* store: general info about the store including competition, etc.
* store_states: maps store to state it is in
* state_names: Maps state abbreviations to names
* googletrend: trend data for particular week/state
* weather: weather conditions for each state
* test: Same as training table, w/o sales and customers
```
for t in tables: display(t.head())
```
This is very representative of a typical industry dataset.
The following returns summarized aggregate information to each table accross each field.
```
for t in tables: display(DataFrameSummary(t).summary())
```
## Data Cleaning / Feature Engineering
As a structured data problem, we necessarily have to go through all the cleaning and feature engineering, even though we're using a neural network.
```
train, store, store_states, state_names, googletrend, weather, test = tables
len(train),len(test)
```
We turn state Holidays to booleans, to make them more convenient for modeling. We can do calculations on pandas fields using notation very similar (often identical) to numpy.
```
train.StateHoliday = train.StateHoliday!='0'
test.StateHoliday = test.StateHoliday!='0'
```
`join_df` is a function for joining tables on specific fields. By default, we'll be doing a left outer join of `right` on the `left` argument using the given fields for each table.
Pandas does joins using the `merge` method. The `suffixes` argument describes the naming convention for duplicate fields. We've elected to leave the duplicate field names on the left untouched, and append a "\_y" to those on the right.
```
def join_df(left, right, left_on, right_on=None, suffix='_y'):
if right_on is None: right_on = left_on
return left.merge(right, how='left', left_on=left_on, right_on=right_on,
suffixes=("", suffix))
```
Join weather/state names.
```
weather = join_df(weather, state_names, "file", "StateName")
```
In pandas you can add new columns to a dataframe by simply defining it. We'll do this for googletrends by extracting dates and state names from the given data and adding those columns.
We're also going to replace all instances of state name 'NI' to match the usage in the rest of the data: 'HB,NI'. This is a good opportunity to highlight pandas indexing. We can use `.loc[rows, cols]` to select a list of rows and a list of columns from the dataframe. In this case, we're selecting rows w/ statename 'NI' by using a boolean list `googletrend.State=='NI'` and selecting "State".
```
googletrend['Date'] = googletrend.week.str.split(' - ', expand=True)[0]
googletrend['State'] = googletrend.file.str.split('_', expand=True)[2]
googletrend.loc[googletrend.State=='NI', "State"] = 'HB,NI'
```
The following extracts particular date fields from a complete datetime for the purpose of constructing categoricals.
You should *always* consider this feature extraction step when working with date-time. Without expanding your date-time into these additional fields, you can't capture any trend/cyclical behavior as a function of time at any of these granularities. We'll add to every table with a date field.
```
add_datepart(weather, "Date", drop=False)
add_datepart(googletrend, "Date", drop=False)
add_datepart(train, "Date", drop=False)
add_datepart(test, "Date", drop=False)
```
The Google trends data has a special category for the whole of the Germany - we'll pull that out so we can use it explicitly.
```
trend_de = googletrend[googletrend.file == 'Rossmann_DE']
```
Now we can outer join all of our data into a single dataframe. Recall that in outer joins everytime a value in the joining field on the left table does not have a corresponding value on the right table, the corresponding row in the new table has Null values for all right table fields. One way to check that all records are consistent and complete is to check for Null values post-join, as we do here.
*Aside*: Why note just do an inner join?
If you are assuming that all records are complete and match on the field you desire, an inner join will do the same thing as an outer join. However, in the event you are wrong or a mistake is made, an outer join followed by a null-check will catch it. (Comparing before/after # of rows for inner join is equivalent, but requires keeping track of before/after row #'s. Outer join is easier.)
```
store = join_df(store, store_states, "Store")
len(store[store.State.isnull()])
joined = join_df(train, store, "Store")
joined_test = join_df(test, store, "Store")
len(joined[joined.StoreType.isnull()]),len(joined_test[joined_test.StoreType.isnull()])
joined = join_df(joined, googletrend, ["State","Year", "Week"])
joined_test = join_df(joined_test, googletrend, ["State","Year", "Week"])
len(joined[joined.trend.isnull()]),len(joined_test[joined_test.trend.isnull()])
joined = joined.merge(trend_de, 'left', ["Year", "Week"], suffixes=('', '_DE'))
joined_test = joined_test.merge(trend_de, 'left', ["Year", "Week"], suffixes=('', '_DE'))
len(joined[joined.trend_DE.isnull()]),len(joined_test[joined_test.trend_DE.isnull()])
joined = join_df(joined, weather, ["State","Date"])
joined_test = join_df(joined_test, weather, ["State","Date"])
len(joined[joined.Mean_TemperatureC.isnull()]),len(joined_test[joined_test.Mean_TemperatureC.isnull()])
for df in (joined, joined_test):
for c in df.columns:
if c.endswith('_y'):
if c in df.columns: df.drop(c, inplace=True, axis=1)
```
Next we'll fill in missing values to avoid complications with `NA`'s. `NA` (not available) is how Pandas indicates missing values; many models have problems when missing values are present, so it's always important to think about how to deal with them. In these cases, we are picking an arbitrary *signal value* that doesn't otherwise appear in the data.
```
for df in (joined,joined_test):
df['CompetitionOpenSinceYear'] = df.CompetitionOpenSinceYear.fillna(1900).astype(np.int32)
df['CompetitionOpenSinceMonth'] = df.CompetitionOpenSinceMonth.fillna(1).astype(np.int32)
df['Promo2SinceYear'] = df.Promo2SinceYear.fillna(1900).astype(np.int32)
df['Promo2SinceWeek'] = df.Promo2SinceWeek.fillna(1).astype(np.int32)
```
Next we'll extract features "CompetitionOpenSince" and "CompetitionDaysOpen". Note the use of `apply()` in mapping a function across dataframe values.
```
for df in (joined,joined_test):
df["CompetitionOpenSince"] = pd.to_datetime(dict(year=df.CompetitionOpenSinceYear,
month=df.CompetitionOpenSinceMonth, day=15))
df["CompetitionDaysOpen"] = df.Date.subtract(df.CompetitionOpenSince).dt.days
```
We'll replace some erroneous / outlying data.
```
for df in (joined,joined_test):
df.loc[df.CompetitionDaysOpen<0, "CompetitionDaysOpen"] = 0
df.loc[df.CompetitionOpenSinceYear<1990, "CompetitionDaysOpen"] = 0
```
We add "CompetitionMonthsOpen" field, limiting the maximum to 2 years to limit number of unique categories.
```
for df in (joined,joined_test):
df["CompetitionMonthsOpen"] = df["CompetitionDaysOpen"]//30
df.loc[df.CompetitionMonthsOpen>24, "CompetitionMonthsOpen"] = 24
joined.CompetitionMonthsOpen.unique()
```
Same process for Promo dates.
```
for df in (joined,joined_test):
df["Promo2Since"] = pd.to_datetime(df.apply(lambda x: Week(
x.Promo2SinceYear, x.Promo2SinceWeek).monday(), axis=1).astype(pd.datetime))
df["Promo2Days"] = df.Date.subtract(df["Promo2Since"]).dt.days
for df in (joined,joined_test):
df.loc[df.Promo2Days<0, "Promo2Days"] = 0
df.loc[df.Promo2SinceYear<1990, "Promo2Days"] = 0
df["Promo2Weeks"] = df["Promo2Days"]//7
df.loc[df.Promo2Weeks<0, "Promo2Weeks"] = 0
df.loc[df.Promo2Weeks>25, "Promo2Weeks"] = 25
df.Promo2Weeks.unique()
joined.to_feather(f'{PATH}joined')
joined_test.to_feather(f'{PATH}joined_test')
```
## Durations
It is common when working with time series data to extract data that explains relationships across rows as opposed to columns, e.g.:
* Running averages
* Time until next event
* Time since last event
This is often difficult to do with most table manipulation frameworks, since they are designed to work with relationships across columns. As such, we've created a class to handle this type of data.
We'll define a function `get_elapsed` for cumulative counting across a sorted dataframe. Given a particular field `fld` to monitor, this function will start tracking time since the last occurrence of that field. When the field is seen again, the counter is set to zero.
Upon initialization, this will result in datetime na's until the field is encountered. This is reset every time a new store is seen. We'll see how to use this shortly.
```
def get_elapsed(fld, pre):
day1 = np.timedelta64(1, 'D')
last_date = np.datetime64()
last_store = 0
res = []
for s,v,d in zip(df.Store.values,df[fld].values, df.Date.values):
if s != last_store:
last_date = np.datetime64()
last_store = s
if v: last_date = d
res.append(((d-last_date).astype('timedelta64[D]') / day1))
df[pre+fld] = res
```
We'll be applying this to a subset of columns:
```
columns = ["Date", "Store", "Promo", "StateHoliday", "SchoolHoliday"]
#df = train[columns]
df = train[columns].append(test[columns])
```
Let's walk through an example.
Say we're looking at School Holiday. We'll first sort by Store, then Date, and then call `add_elapsed('SchoolHoliday', 'After')`:
This will apply to each row with School Holiday:
* A applied to every row of the dataframe in order of store and date
* Will add to the dataframe the days since seeing a School Holiday
* If we sort in the other direction, this will count the days until another holiday.
```
fld = 'SchoolHoliday'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
```
We'll do this for two more fields.
```
fld = 'StateHoliday'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
fld = 'Promo'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
```
We're going to set the active index to Date.
```
df = df.set_index("Date")
```
Then set null values from elapsed field calculations to 0.
```
columns = ['SchoolHoliday', 'StateHoliday', 'Promo']
for o in ['Before', 'After']:
for p in columns:
a = o+p
df[a] = df[a].fillna(0).astype(int)
```
Next we'll demonstrate window functions in pandas to calculate rolling quantities.
Here we're sorting by date (`sort_index()`) and counting the number of events of interest (`sum()`) defined in `columns` in the following week (`rolling()`), grouped by Store (`groupby()`). We do the same in the opposite direction.
```
bwd = df[['Store']+columns].sort_index().groupby("Store").rolling(7, min_periods=1).sum()
fwd = df[['Store']+columns].sort_index(ascending=False
).groupby("Store").rolling(7, min_periods=1).sum()
```
Next we want to drop the Store indices grouped together in the window function.
Often in pandas, there is an option to do this in place. This is time and memory efficient when working with large datasets.
```
bwd.drop('Store',1,inplace=True)
bwd.reset_index(inplace=True)
fwd.drop('Store',1,inplace=True)
fwd.reset_index(inplace=True)
df.reset_index(inplace=True)
```
Now we'll merge these values onto the df.
```
df = df.merge(bwd, 'left', ['Date', 'Store'], suffixes=['', '_bw'])
df = df.merge(fwd, 'left', ['Date', 'Store'], suffixes=['', '_fw'])
df.drop(columns,1,inplace=True)
df.head()
```
It's usually a good idea to back up large tables of extracted / wrangled features before you join them onto another one, that way you can go back to it easily if you need to make changes to it.
```
df.to_feather(f'{PATH}df')
df = pd.read_feather(f'{PATH}df')
df["Date"] = pd.to_datetime(df.Date)
df.columns
joined = join_df(joined, df, ['Store', 'Date'])
joined_test = join_df(joined_test, df, ['Store', 'Date'])
```
The authors also removed all instances where the store had zero sale / was closed. We speculate that this may have cost them a higher standing in the competition. One reason this may be the case is that a little exploratory data analysis reveals that there are often periods where stores are closed, typically for refurbishment. Before and after these periods, there are naturally spikes in sales that one might expect. By ommitting this data from their training, the authors gave up the ability to leverage information about these periods to predict this otherwise volatile behavior.
```
joined = joined[joined.Sales!=0]
```
We'll back this up as well.
```
joined.reset_index(inplace=True)
joined_test.reset_index(inplace=True)
joined.to_feather(f'{PATH}joined')
joined_test.to_feather(f'{PATH}joined_test')
```
We now have our final set of engineered features.
While these steps were explicitly outlined in the paper, these are all fairly typical feature engineering steps for dealing with time series data and are practical in any similar setting.
## Create features
```
joined = pd.read_feather(f'{PATH}joined')
joined_test = pd.read_feather(f'{PATH}joined_test')
joined.head().T.head(40)
```
Now that we've engineered all our features, we need to convert to input compatible with a neural network.
This includes converting categorical variables into contiguous integers or one-hot encodings, normalizing continuous features to standard normal, etc...
```
cat_vars = ['Store', 'DayOfWeek', 'Year', 'Month', 'Day', 'StateHoliday', 'CompetitionMonthsOpen',
'Promo2Weeks', 'StoreType', 'Assortment', 'PromoInterval', 'CompetitionOpenSinceYear', 'Promo2SinceYear',
'State', 'Week', 'Events', 'Promo_fw', 'Promo_bw', 'StateHoliday_fw', 'StateHoliday_bw',
'SchoolHoliday_fw', 'SchoolHoliday_bw']
contin_vars = ['CompetitionDistance', 'Max_TemperatureC', 'Mean_TemperatureC', 'Min_TemperatureC',
'Max_Humidity', 'Mean_Humidity', 'Min_Humidity', 'Max_Wind_SpeedKm_h',
'Mean_Wind_SpeedKm_h', 'CloudCover', 'trend', 'trend_DE',
'AfterStateHoliday', 'BeforeStateHoliday', 'Promo', 'SchoolHoliday']
n = len(joined); n
dep = 'Sales'
joined = joined[cat_vars+contin_vars+[dep, 'Date']].copy()
joined_test[dep] = 0
joined_test = joined_test[cat_vars+contin_vars+[dep, 'Date', 'Id']].copy()
for v in cat_vars: joined[v] = joined[v].astype('category').cat.as_ordered()
apply_cats(joined_test, joined)
for v in contin_vars:
joined[v] = joined[v].fillna(0).astype('float32')
joined_test[v] = joined_test[v].fillna(0).astype('float32')
```
We're going to run on a sample.
```
idxs = get_cv_idxs(n, val_pct=150000/n)
joined_samp = joined.iloc[idxs].set_index("Date")
samp_size = len(joined_samp); samp_size
```
To run on the full dataset, use this instead:
```
samp_size = n
joined_samp = joined.set_index("Date")
```
We can now process our data...
```
joined_samp.head(2)
df, y, nas, mapper = proc_df(joined_samp, 'Sales', do_scale=True)
yl = np.log(y)
joined_test = joined_test.set_index("Date")
df_test, _, nas, mapper = proc_df(joined_test, 'Sales', do_scale=True, skip_flds=['Id'],
mapper=mapper, na_dict=nas)
df.head(2)
```
In time series data, cross-validation is not random. Instead, our holdout data is generally the most recent data, as it would be in real application. This issue is discussed in detail in [this post](http://www.fast.ai/2017/11/13/validation-sets/) on our web site.
One approach is to take the last 25% of rows (sorted by date) as our validation set.
```
train_ratio = 0.75
# train_ratio = 0.9
train_size = int(samp_size * train_ratio); train_size
val_idx = list(range(train_size, len(df)))
```
An even better option for picking a validation set is using the exact same length of time period as the test set uses - this is implemented here:
```
val_idx = np.flatnonzero(
(df.index<=datetime.datetime(2014,9,17)) & (df.index>=datetime.datetime(2014,8,1)))
val_idx=[0]
```
## DL
We're ready to put together our models.
Root-mean-squared percent error is the metric Kaggle used for this competition.
```
def inv_y(a): return np.exp(a)
def exp_rmspe(y_pred, targ):
targ = inv_y(targ)
pct_var = (targ - inv_y(y_pred))/targ
return math.sqrt((pct_var**2).mean())
max_log_y = np.max(yl)
y_range = (0, max_log_y*1.2)
```
We can create a ModelData object directly from out data frame.
```
md = ColumnarModelData.from_data_frame(PATH, val_idx, df, yl.astype(np.float32), cat_flds=cat_vars, bs=128,
test_df=df_test)
```
Some categorical variables have a lot more levels than others. Store, in particular, has over a thousand!
```
cat_sz = [(c, len(joined_samp[c].cat.categories)+1) for c in cat_vars]
cat_sz
```
We use the *cardinality* of each variable (that is, its number of unique values) to decide how large to make its *embeddings*. Each level will be associated with a vector with length defined as below.
```
emb_szs = [(c, min(50, (c+1)//2)) for _,c in cat_sz]
emb_szs
m = md.get_learner(emb_szs, len(df.columns)-len(cat_vars),
0.04, 1, [1000,500], [0.001,0.01], y_range=y_range)
m.summary()
lr = 1e-3
m.lr_find()
m.sched.plot(100)
```
### Sample
```
m = md.get_learner(emb_szs, len(df.columns)-len(cat_vars),
0.04, 1, [1000,500], [0.001,0.01], y_range=y_range)
lr = 1e-3
m.fit(lr, 3, metrics=[exp_rmspe])
m.fit(lr, 5, metrics=[exp_rmspe], cycle_len=1)
m.fit(lr, 2, metrics=[exp_rmspe], cycle_len=4)
```
### All
```
m = md.get_learner(emb_szs, len(df.columns)-len(cat_vars),
0.04, 1, [1000,500], [0.001,0.01], y_range=y_range)
lr = 1e-3
m.fit(lr, 1, metrics=[exp_rmspe])
m.fit(lr, 3, metrics=[exp_rmspe])
m.fit(lr, 3, metrics=[exp_rmspe], cycle_len=1)
```
### Test
```
m = md.get_learner(emb_szs, len(df.columns)-len(cat_vars),
0.04, 1, [1000,500], [0.001,0.01], y_range=y_range)
lr = 1e-3
m.fit(lr, 3, metrics=[exp_rmspe])
m.fit(lr, 3, metrics=[exp_rmspe], cycle_len=1)
m.save('val0')
m.load('val0')
x,y=m.predict_with_targs()
exp_rmspe(x,y)
pred_test=m.predict(True)
pred_test = np.exp(pred_test)
joined_test['Sales']=pred_test
csv_fn=f'{PATH}tmp/sub.csv'
joined_test[['Id','Sales']].to_csv(csv_fn, index=False)
FileLink(csv_fn)
```
## RF
```
from sklearn.ensemble import RandomForestRegressor
((val,trn), (y_val,y_trn)) = split_by_idx(val_idx, df.values, yl)
m = RandomForestRegressor(n_estimators=40, max_features=0.99, min_samples_leaf=2,
n_jobs=-1, oob_score=True)
m.fit(trn, y_trn);
preds = m.predict(val)
m.score(trn, y_trn), m.score(val, y_val), m.oob_score_, exp_rmspe(preds, y_val)
```
| github_jupyter |
```
from sys import modules
IN_COLAB = 'google.colab' in modules
if IN_COLAB:
!pip install -q ir_axioms[examples] python-terrier
# Start/initialize PyTerrier.
from pyterrier import started, init
if not started():
init(tqdm="auto")
from pyterrier.datasets import get_dataset, Dataset
# Load dataset.
dataset_name = "msmarco-passage"
dataset: Dataset = get_dataset(f"irds:{dataset_name}")
dataset_test: Dataset = get_dataset(f"irds:{dataset_name}/trec-dl-2020/judged")
from pathlib import Path
cache_dir = Path("cache/")
index_dir = cache_dir / "indices" / dataset_name.split("/")[0]
from pyterrier.index import IterDictIndexer
if not index_dir.exists():
indexer = IterDictIndexer(str(index_dir.absolute()))
indexer.index(
dataset.get_corpus_iter(),
fields=["text"]
)
from pyterrier.batchretrieve import BatchRetrieve
# BM25 baseline retrieval.
bm25 = BatchRetrieve(str(index_dir.absolute()), wmodel="BM25")
from ir_axioms.axiom import (
ArgUC, QTArg, QTPArg, aSL, PROX1, PROX2, PROX3, PROX4, PROX5, TFC1, TFC3, RS_TF, RS_TF_IDF, RS_BM25, RS_PL2, RS_QL,
AND, LEN_AND, M_AND, LEN_M_AND, DIV, LEN_DIV, M_TDC, LEN_M_TDC, STMC1, STMC1_f, STMC2, STMC2_f, LNC1, TF_LNC, LB1,
REG, ANTI_REG, REG_f, ANTI_REG_f, ASPECT_REG, ASPECT_REG_f, ORIG, VoteAxiom
)
axiom = (
~VoteAxiom([
ArgUC(), QTArg(), QTPArg(), aSL(),
LNC1(), TF_LNC(), LB1(),
PROX1(), PROX2(), PROX3(), PROX4(), PROX5(),
REG(), REG_f(), ANTI_REG(), ANTI_REG_f(), ASPECT_REG(), ASPECT_REG_f(),
AND(), LEN_AND(), M_AND(), LEN_M_AND(), DIV(), LEN_DIV(),
RS_TF(), RS_TF_IDF(), RS_BM25(), RS_PL2(), RS_QL(),
TFC1(), TFC3(), M_TDC(), LEN_M_TDC(),
STMC1(), STMC1_f(), STMC2(), STMC2_f(),
], minimum_votes=0.5) | ORIG()
)
from ir_axioms.modules.pivot import MiddlePivotSelection
from ir_axioms.backend.pyterrier.transformers import AxiomaticReranker
kwiksort = bm25 % 20 >> AxiomaticReranker(
axiom=axiom,
index=index_dir,
dataset=dataset_name,
pivot_selection=MiddlePivotSelection(),
cache_dir=cache_dir,
verbose=True
)
from pyterrier.pipelines import Experiment
from ir_measures import nDCG, MAP, RR
experiment = Experiment(
[bm25, kwiksort ^ bm25],
dataset_test.get_topics(),
dataset_test.get_qrels(),
[nDCG @ 10, RR, MAP],
["BM25", "KwikSort"],
verbose=True,
)
experiment.sort_values(by="nDCG@10", ascending=False, inplace=True)
experiment
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Train and explain models remotely via Azure Machine Learning Compute
_**This notebook showcases how to use the Azure Machine Learning Interpretability SDK to train and explain a regression model remotely on an Azure Machine Leanrning Compute Target (AMLCompute).**_
## Table of Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. Initialize a Workspace
1. Create an Experiment
1. Introduction to AmlCompute
1. Submit an AmlCompute run in a few different ways
1. Option 1: Provision as a run based compute target
1. Option 2: Provision as a persistent compute target (Basic)
1. Option 3: Provision as a persistent compute target (Advanced)
1. Additional operations to perform on AmlCompute
1. [Download model explanations from Azure Machine Learning Run History](#Download)
1. [Visualize explanations](#Visualize)
1. [Next steps](#Next)
## Introduction
This notebook showcases how to train and explain a regression model remotely via Azure Machine Learning Compute (AMLCompute), and download the calculated explanations locally for visualization.
It demonstrates the API calls that you need to make to submit a run for training and explaining a model to AMLCompute, download the compute explanations remotely, and visualizing the global and local explanations via a visualization dashboard that provides an interactive way of discovering patterns in model predictions and downloaded explanations.
We will showcase one of the tabular data explainers: TabularExplainer (SHAP).
Problem: Boston Housing Price Prediction with scikit-learn (train a model and run an explainer remotely via AMLCompute, and download and visualize the remotely-calculated explanations.)
|  |
|:--:|
## Setup
If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the [configuration notebook](../../../configuration.ipynb) first if you haven't.
If you are using Jupyter notebooks, the extensions should be installed automatically with the package.
If you are using Jupyter Labs run the following command:
```
(myenv) $ jupyter labextension install @jupyter-widgets/jupyterlab-manager
```
```
# Check core SDK version number
import azureml.core
print("SDK version:", azureml.core.VERSION)
```
## Initialize a Workspace
Initialize a workspace object from persisted configuration
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
```
## Create An Experiment
**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments.
```
from azureml.core import Experiment
experiment_name = 'explainer-remote-run-on-amlcompute'
experiment = Experiment(workspace=ws, name=experiment_name)
```
## Introduction to AmlCompute
Azure Machine Learning Compute is managed compute infrastructure that allows the user to easily create single to multi-node compute of the appropriate VM Family. It is created **within your workspace region** and is a resource that can be used by other users in your workspace. It autoscales by default to the max_nodes, when a job is submitted, and executes in a containerized environment packaging the dependencies as specified by the user.
Since it is managed compute, job scheduling and cluster management are handled internally by Azure Machine Learning service.
For more information on Azure Machine Learning Compute, please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)
If you are an existing BatchAI customer who is migrating to Azure Machine Learning, please read [this article](https://aka.ms/batchai-retirement)
**Note**: As with other Azure services, there are limits on certain resources (for eg. AmlCompute quota) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
The training script `train_explain.py` is already created for you. Let's have a look.
## Submit an AmlCompute run in a few different ways
First lets check which VM families are available in your region. Azure is a regional service and some specialized SKUs (especially GPUs) are only available in certain regions. Since AmlCompute is created in the region of your workspace, we will use the supported_vms () function to see if the VM family we want to use ('STANDARD_D2_V2') is supported.
You can also pass a different region to check availability and then re-create your workspace in that region through the [configuration notebook](../../../configuration.ipynb)
```
from azureml.core.compute import ComputeTarget, AmlCompute
AmlCompute.supported_vmsizes(workspace=ws)
# AmlCompute.supported_vmsizes(workspace=ws, location='southcentralus')
```
### Create project directory
Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on
```
import os
import shutil
project_folder = './explainer-remote-run-on-amlcompute'
os.makedirs(project_folder, exist_ok=True)
shutil.copy('train_explain.py', project_folder)
```
### Option 1: Provision as a run based compute target
You can provision AmlCompute as a compute target at run-time. In this case, the compute is auto-created for your run, scales up to max_nodes that you specify, and then **deleted automatically** after the run completes.
```
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.runconfig import DEFAULT_CPU_IMAGE
# create a new runconfig object
run_config = RunConfiguration()
# signal that you want to use AmlCompute to execute script.
run_config.target = "amlcompute"
# AmlCompute will be created in the same region as workspace
# Set vm size for AmlCompute
run_config.amlcompute.vm_size = 'STANDARD_D2_V2'
# enable Docker
run_config.environment.docker.enabled = True
# set Docker base image to the default CPU-based image
run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE
# use conda_dependencies.yml to create a conda environment in the Docker image for execution
run_config.environment.python.user_managed_dependencies = False
azureml_pip_packages = [
'azureml-defaults', 'azureml-contrib-explain-model', 'azureml-core', 'azureml-telemetry',
'azureml-explain-model', 'sklearn-pandas', 'azureml-dataprep'
]
# specify CondaDependencies obj
run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'],
pip_packages=azureml_pip_packages)
# Now submit a run on AmlCompute
from azureml.core.script_run_config import ScriptRunConfig
script_run_config = ScriptRunConfig(source_directory=project_folder,
script='train_explain.py',
run_config=run_config)
run = experiment.submit(script_run_config)
# Show run details
run
```
Note: if you need to cancel a run, you can follow [these instructions](https://aka.ms/aml-docs-cancel-run).
```
%%time
# Shows output of the run on stdout.
run.wait_for_completion(show_output=True)
```
### Option 2: Provision as a persistent compute target (Basic)
You can provision a persistent AmlCompute resource by simply defining two parameters thanks to smart defaults. By default it autoscales from 0 nodes and provisions dedicated VMs to run your job in a container. This is useful when you want to continously re-use the same target, debug it between jobs or simply share the resource with other users of your workspace.
* `vm_size`: VM family of the nodes provisioned by AmlCompute. Simply choose from the supported_vmsizes() above
* `max_nodes`: Maximum nodes to autoscale to while running a job on AmlCompute
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster"
# Verify that cluster does not exist already
try:
cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
cpu_cluster.wait_for_completion(show_output=True)
```
### Configure & Run
```
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
# create a new RunConfig object
run_config = RunConfiguration(framework="python")
# Set compute target to AmlCompute target created in previous step
run_config.target = cpu_cluster.name
# enable Docker
run_config.environment.docker.enabled = True
azureml_pip_packages = [
'azureml-defaults', 'azureml-contrib-explain-model', 'azureml-core', 'azureml-telemetry',
'azureml-explain-model', 'azureml-dataprep'
]
# specify CondaDependencies obj
run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'],
pip_packages=azureml_pip_packages)
from azureml.core import Run
from azureml.core import ScriptRunConfig
src = ScriptRunConfig(source_directory=project_folder,
script='train_explain.py',
run_config=run_config)
run = experiment.submit(config=src)
run
%%time
# Shows output of the run on stdout.
run.wait_for_completion(show_output=True)
run.get_metrics()
```
### Option 3: Provision as a persistent compute target (Advanced)
You can also specify additional properties or change defaults while provisioning AmlCompute using a more advanced configuration. This is useful when you want a dedicated cluster of 4 nodes (for example you can set the min_nodes and max_nodes to 4), or want the compute to be within an existing VNet in your subscription.
In addition to `vm_size` and `max_nodes`, you can specify:
* `min_nodes`: Minimum nodes (default 0 nodes) to downscale to while running a job on AmlCompute
* `vm_priority`: Choose between 'dedicated' (default) and 'lowpriority' VMs when provisioning AmlCompute. Low Priority VMs use Azure's excess capacity and are thus cheaper but risk your run being pre-empted
* `idle_seconds_before_scaledown`: Idle time (default 120 seconds) to wait after run completion before auto-scaling to min_nodes
* `vnet_resourcegroup_name`: Resource group of the **existing** VNet within which AmlCompute should be provisioned
* `vnet_name`: Name of VNet
* `subnet_name`: Name of SubNet within the VNet
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster"
# Verify that cluster does not exist already
try:
cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
vm_priority='lowpriority',
min_nodes=2,
max_nodes=4,
idle_seconds_before_scaledown='300',
vnet_resourcegroup_name='<my-resource-group>',
vnet_name='<my-vnet-name>',
subnet_name='<my-subnet-name>')
cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
cpu_cluster.wait_for_completion(show_output=True)
```
### Configure & Run
```
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
# create a new RunConfig object
run_config = RunConfiguration(framework="python")
# Set compute target to AmlCompute target created in previous step
run_config.target = cpu_cluster.name
# enable Docker
run_config.environment.docker.enabled = True
azureml_pip_packages = [
'azureml-defaults', 'azureml-contrib-explain-model', 'azureml-core', 'azureml-telemetry',
'azureml-explain-model', 'azureml-dataprep'
]
# specify CondaDependencies obj
run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'],
pip_packages=azureml_pip_packages)
from azureml.core import Run
from azureml.core import ScriptRunConfig
src = ScriptRunConfig(source_directory=project_folder,
script='train_explain.py',
run_config=run_config)
run = experiment.submit(config=src)
run
%%time
# Shows output of the run on stdout.
run.wait_for_completion(show_output=True)
run.get_metrics()
from azureml.contrib.explain.model.explanation.explanation_client import ExplanationClient
client = ExplanationClient.from_run(run)
# Get the top k (e.g., 4) most important features with their importance values
explanation = client.download_model_explanation(top_k=4)
```
## Additional operations to perform on AmlCompute
You can perform more operations on AmlCompute such as updating the node counts or deleting the compute.
```
# Get_status () gets the latest status of the AmlCompute target
cpu_cluster.get_status().serialize()
# Update () takes in the min_nodes, max_nodes and idle_seconds_before_scaledown and updates the AmlCompute target
# cpu_cluster.update(min_nodes=1)
# cpu_cluster.update(max_nodes=10)
cpu_cluster.update(idle_seconds_before_scaledown=300)
# cpu_cluster.update(min_nodes=2, max_nodes=4, idle_seconds_before_scaledown=600)
# Delete () is used to deprovision and delete the AmlCompute target. Useful if you want to re-use the compute name
# 'cpu-cluster' in this case but use a different VM family for instance.
# cpu_cluster.delete()
```
## Download
1. Download model explanation data.
```
from azureml.contrib.explain.model.explanation.explanation_client import ExplanationClient
# Get model explanation data
client = ExplanationClient.from_run(run)
global_explanation = client.download_model_explanation()
local_importance_values = global_explanation.local_importance_values
expected_values = global_explanation.expected_values
# Or you can use the saved run.id to retrive the feature importance values
client = ExplanationClient.from_run_id(ws, experiment_name, run.id)
global_explanation = client.download_model_explanation()
local_importance_values = global_explanation.local_importance_values
expected_values = global_explanation.expected_values
# Get the top k (e.g., 4) most important features with their importance values
global_explanation_topk = client.download_model_explanation(top_k=4)
global_importance_values = global_explanation_topk.get_ranked_global_values()
global_importance_names = global_explanation_topk.get_ranked_global_names()
print('global importance values: {}'.format(global_importance_values))
print('global importance names: {}'.format(global_importance_names))
```
2. Download model file.
```
# retrieve model for visualization and deployment
from azureml.core.model import Model
from sklearn.externals import joblib
original_model = Model(ws, 'model_explain_model_on_amlcomp')
model_path = original_model.download(exist_ok=True)
original_model = joblib.load(model_path)
```
3. Download test dataset.
```
# retrieve x_test for visualization
from sklearn.externals import joblib
x_test_path = './x_test_boston_housing.pkl'
run.download_file('x_test_boston_housing.pkl', output_file_path=x_test_path)
x_test = joblib.load('x_test_boston_housing.pkl')
```
## Visualize
Load the visualization dashboard
```
from azureml.contrib.explain.model.visualize import ExplanationDashboard
ExplanationDashboard(global_explanation, original_model, x_test)
```
## Next
Learn about other use cases of the explain package on a:
1. [Training time: regression problem](../../tabular-data/explain-binary-classification-local.ipynb)
1. [Training time: binary classification problem](../../tabular-data/explain-binary-classification-local.ipynb)
1. [Training time: multiclass classification problem](../../tabular-data/explain-multiclass-classification-local.ipynb)
1. Explain models with engineered features:
1. [Simple feature transformations](../../tabular-data/simple-feature-transformations-explain-local.ipynb)
1. [Advanced feature transformations](../../tabular-data/advanced-feature-transformations-explain-local.ipynb)
1. [Save model explanations via Azure Machine Learning Run History](../run-history/save-retrieve-explanations-run-history.ipynb)
1. Inferencing time: deploy a classification model and explainer:
1. [Deploy a locally-trained model and explainer](../scoring-time/train-explain-model-locally-and-deploy.ipynb)
1. [Deploy a remotely-trained model and explainer](../scoring-time/train-explain-model-on-amlcompute-and-deploy.ipynb)
| github_jupyter |
```
from math import sqrt
import tensorflow as tf
from tensorflow import keras
import pandas as pd
from tensorflow.keras import Sequential
from tensorflow.keras.layers import LSTM, Dense, Dropout, Conv1D, GRU
from tensorflow.keras.losses import mean_squared_error
from numpy.core._multiarray_umath import concatenate
from sklearn.preprocessing import MinMaxScaler
import matplotlib.pyplot as plt
# supervised监督学习函数
def series_to_supervised(data, columns, n_in=1, n_out=1, dropnan=True):
n_vars = 1 if isinstance(data, list) else data.shape[1]
df = pd.DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [('%s%d(t-%d)' % (columns[j], j + 1, i))
for j in range(n_vars)]
# forecast sequence (t, t+1, ... t+n)
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [('%s%d(t)' % (columns[j], j + 1)) for j in range(n_vars)]
else:
names += [('%s%d(t+%d)' % (columns[j], j + 1, i))
for j in range(n_vars)]
# put it all together
agg = pd.concat(cols, axis=1)
agg.columns = names
# drop rows with NaN values
if dropnan:
clean_agg = agg.dropna()
return clean_agg
# return agg
dataset = pd.read_csv(
'Machine_usage_groupby.csv')
dataset_columns = dataset.columns
values = dataset.values
print(dataset)
# 归一化处理
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
# 监督学习
reframed = series_to_supervised(scaled, dataset_columns, 1, 1)
values = reframed.values
# 学习与检测数据的划分
n_train_hours = 20000
train = values[:n_train_hours, :]
test = values[n_train_hours:, :]
# 监督学习结果划分
train_x, train_y = train[:, :-1], train[:, -1]
test_x, test_y = test[:, :-1], test[:, -1]
# 为了在LSTM中应用该数据,需要将其格式转化为3D format,即[Samples, timesteps, features]
train_X = train_x.reshape((train_x.shape[0], 1, train_x.shape[1]))
test_X = test_x.reshape((test_x.shape[0], 1, test_x.shape[1]))
model = Sequential()
model.add(
GRU(
32,
input_shape=(
train_X.shape[1],
train_X.shape[2]),
return_sequences=True))
model.add(GRU(16, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dense(16, activation="relu"))
model.add(Dense(1))
model.compile(loss=tf.keras.losses.Huber(),
optimizer='adam',
metrics=["mse"])
history = model.fit(
train_X,
train_y,
epochs=50,
batch_size=72,
validation_data=(
test_X,
test_y),
verbose = 2)
#画图
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
# make the prediction
yHat = model.predict(test_X)
inv_yHat = concatenate((yHat, test_x[:, 1:]), axis=1) # 数组拼接
inv_yHat = inv_yHat[:, 0]
test_y = test_y.reshape((len(test_y), 1))
inv_y = concatenate((test_y, test_x[:, 1:]), axis=1)
inv_y = inv_y[:, 0]
rmse = sqrt(mean_squared_error(inv_yHat, inv_y))
print('Test RMSE: %.8f' % rmse)
mse = mean_squared_error(inv_yHat, inv_y)
print('Test MSE: %.8f' % mse)
yhat = model.predict(test_X)
test_X_reshaped = test_X.reshape((test_X.shape[0], test_X.shape[2]))
inv_yhat = concatenate((yhat, yhat, test_X_reshaped[:, 1:]), axis=1)
inv_yhat = inv_yhat[:, 0]
test_y = test_y.reshape((len(test_y), 1))
inv_y = concatenate((test_y, test_y, test_X_reshaped[:, 1:]), axis=1)
inv_y = inv_y[:, 0]
plt.plot(inv_yhat, label='prediction')
plt.plot(inv_y, label='real')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
plt.plot(inv_yhat[:500], label='prediction')
plt.plot(inv_y[:500], label='real_cpu_usage_percent')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
plt.plot(inv_yhat[:50], label='prediction')
plt.plot(inv_y[:50], label='real_cpu_usage_percent')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
```
| github_jupyter |
```
#-*- coding: utf-8 -*-
import re
from wxpy import *
import jieba
import numpy
import pandas as pd
import matplotlib.pyplot as plt
from scipy.misc import imread
from wordcloud import WordCloud, ImageColorGenerator
def write_txt_file(path, txt):
'''
写入txt文本
'''
with open(path, 'a', encoding='gb18030', newline='') as f:
f.write(txt)
def read_txt_file(path):
'''
读取txt文本
'''
with open(path, 'r', encoding='gb18030', newline='') as f:
return f.read()
def login():
# 初始化机器人,扫码登陆
bot = Bot()
# 获取所有好友
my_friends = bot.friends()
print(type(my_friends))
return my_friends
def show_sex_ratio(friends):
# 使用一个字典统计好友男性和女性的数量
sex_dict = {'male': 0, 'female': 0}
for friend in friends:
# 统计性别
if friend.sex == 1:
sex_dict['male'] += 1
elif friend.sex == 2:
sex_dict['female'] += 1
print(sex_dict)
def get_area_distribution(friends):
# 使用一个字典统计各省好友数量
province_dict = {'北京': 0, '上海': 0, '天津': 0, '重庆': 0,
'河北': 0, '山西': 0, '吉林': 0, '辽宁': 0, '黑龙江': 0,
'陕西': 0, '甘肃': 0, '青海': 0, '山东': 0, '福建': 0,
'浙江': 0, '台湾': 0, '河南': 0, '湖北': 0, '湖南': 0,
'江西': 0, '江苏': 0, '安徽': 0, '广东': 0, '海南': 0,
'四川': 0, '贵州': 0, '云南': 0,
'内蒙古': 0, '新疆': 0, '宁夏': 0, '广西': 0, '西藏': 0,
'香港': 0, '澳门': 0, '台湾': 0}
# 统计省份
for friend in friends:
if friend.province in province_dict.keys():
province_dict[friend.province] += 1
# 为了方便数据的呈现,生成JSON Array格式数据
data = []
for key, value in province_dict.items():
data.append({'name': key, 'value': value})
return data
def show_area_distribution(friends):
# 使用一个字典统计各省好友数量
province_dict = {'北京': 0, '上海': 0, '天津': 0, '重庆': 0,
'河北': 0, '山西': 0, '吉林': 0, '辽宁': 0, '黑龙江': 0,
'陕西': 0, '甘肃': 0, '青海': 0, '山东': 0, '福建': 0,
'浙江': 0, '台湾': 0, '河南': 0, '湖北': 0, '湖南': 0,
'江西': 0, '江苏': 0, '安徽': 0, '广东': 0, '海南': 0,
'四川': 0, '贵州': 0, '云南': 0,
'内蒙古': 0, '新疆': 0, '宁夏': 0, '广西': 0, '西藏': 0,
'香港': 0, '澳门': 0, '台湾': 0}
# 统计省份
for friend in friends:
if friend.province in province_dict.keys():
province_dict[friend.province] += 1
# 为了方便数据的呈现,生成JSON Array格式数据
data = []
for key, value in province_dict.items():
data.append({'name': key, 'value': value})
print(data)
def show_signature(friends):
# 统计签名
for friend in friends:
# 对数据进行清洗,将标点符号等对词频统计造成影响的因素剔除
pattern = re.compile(r'[一-龥]+')
filterdata = re.findall(pattern, friend.signature)
write_txt_file('signatures.txt', ''.join(filterdata))
# 读取文件
content = read_txt_file('signatures.txt')
segment = jieba.lcut(content)
words_df = pd.DataFrame({'segment':segment})
# 读取stopwords
stopwords = pd.read_csv("stopwords.txt",index_col=False,quoting=3,sep=" ",names=['stopword'],encoding='utf-8')
words_df = words_df[~words_df.segment.isin(stopwords.stopword)]
# print(words_df)
words_stat = words_df.groupby(by=['segment'])['segment'].agg({"计数":numpy.size})
words_stat = words_stat.reset_index().sort_values(by=["计数"],ascending=False)
# 设置词云属性
color_mask = imread('background.jfif')
wordcloud = WordCloud(font_path="simhei.ttf", # 设置字体可以显示中文
background_color="white", # 背景颜色
max_words=100, # 词云显示的最大词数
mask=color_mask, # 设置背景图片
max_font_size=80, # 字体最大值
random_state=42,
width=1200, height=1060, margin=2,# 设置图片默认的大小,但是如果使用背景图片的话, # 那么保存的图片大小将会按照其大小保存,margin为词语边缘距离
)
# 生成词云, 可以用generate输入全部文本,也可以我们计算好词频后使用generate_from_frequencies函数
word_frequence = {x[0]:x[1]for x in words_stat.head(100).values}
# print(word_frequence)
word_frequence_dict = {}
for key in word_frequence:
word_frequence_dict[key] = word_frequence[key]
wordcloud.generate_from_frequencies(word_frequence_dict)
# 从背景图片生成颜色值
image_colors = ImageColorGenerator(color_mask)
# 重新上色
wordcloud.recolor(color_func=image_colors)
# 保存图片
wordcloud.to_file('weixin_most_words.png')
plt.imshow(wordcloud)
plt.axis("off")
plt.show()
if __name__ == '__main__':
friends = login()
show_sex_ratio(friends)
show_area_distribution(friends)
show_signature(friends)
from pyecharts import Bar, Line
from pyecharts.engine import create_default_environment
myfriends_pro = get_area_distribution(friends)
# 按照好友数量排序
newlist = sorted(myfriends_pro, key=lambda k: k['value'], reverse = True)
pro_list,num_list = [], []
for onefriend in newlist:
if onefriend["value"] > 0:
pro_list.append(onefriend["name"])
num_list.append(onefriend["value"])
bar = Bar("好友数量", "共有" + str(len(friends)) + "个好友。他们分布在" + str(len(num_list)) + "个省份" , width=1200, height=600)
print(len(pro_list), len(num_list))
bar.add("我的微信好友分布", pro_list, num_list, is_stack=True)
bar.render(r"./我的微信好友分布.html")
bar
from pyecharts import Map
value = num_list
attr = pro_list
map=Map('我的好友分布',width=1000,height=800)
map.add("", attr, value, maptype='china', visual_range=[0, 10], visual_text_color="#fff",
symbol_size=10, is_visualmap=True)
map.render(r"./我的微信好友分布地图.html")
map
```
| github_jupyter |
```
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import random
from vae.arch import VAE
from os import listdir
from os.path import isfile, join
import cv2
import tensorflow as tf
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.05)
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
data = np.load('./data/obs_valid_car_racing.npz')["arr_0"]
data = np.array([item for obs in data for item in obs])
# np.savez_compressed('./data/obs_valid', data[:600])
print(data.shape)
path = "./vae/"
files = [f for f in listdir(path) if f[-3:] == ".h5"]
files.sort()
dropdown = widgets.Dropdown(
options=files,
description='Weight file:',
)
display(dropdown)
vae = VAE()
vae.set_weights("./vae/" + dropdown.value)
# vae.model.summary()
def compare(index):
f, axarr = plt.subplots(1, 2, figsize=(15,15))
img = data[index]
axarr[0].imshow(img)
dream_img = vae.model.predict(x = np.expand_dims(img, axis=0))
axarr[1].imshow(dream_img[0])
#30, 46, 70, 545
compare(30)
img = data[30]
latent = vae.encoder.predict(x = np.expand_dims(img, axis=0))
index = 0
def draw():
f, axarr = plt.subplots(1, 2, figsize=(15,15))
axarr[0].imshow(img)
dream_img = vae.decoder.predict(x = latent)
axarr[1].imshow(dream_img[0])
def change_index(ivalue):
global index, value_slider
index = ivalue
value_slider.value=latent[0][index]
def change_value(value):
global latent
latent[0][index] = value
draw()
index_slider = widgets.IntSlider(description="index", readout=True, min=0, max=len(latent[0])-1, value=0,
continuous_update=False, orientation="horizontal", layout={'width':'90%'})
iindex = interactive(change_index, ivalue=index_slider)
value_slider = widgets.FloatSlider(readout=True, min=-3, max=3, value=latent[0][index],
continuous_update=False, orientation="horizontal", layout={'width':'90%'})
ivalue = interactive(change_value, value=value_slider)
vbox = widgets.VBox([iindex, ivalue])
display(vbox)
# good.h5: 9: -1,3
def create_video(vae, data):
writer = cv2.VideoWriter('demo.avi', 0, 30, (128, 64))
for i in range(0, 200):
img = data[i]
predicted = np.uint8(vae.model.predict(x = np.expand_dims(img, axis=0))[0][...,[2,1,0]] * 255)
img = np.uint8(img[...,[2,1,0]] * 255)
frame = np.append(img, predicted, axis = 1)
writer.write(frame)
create_video(vae, data)
#%%javascript
#Jupyter.notebook.session.delete();
```
| github_jupyter |
<center><img src="../images/DLI Header.png" alt="Header" style="width: 400px;"/></center>
# Hello Camera
### Generic (IP) Cameras
In this notebook, you can test your camera to make sure it's working on the Jetson Nano as expected. It should already be plugged into the USB camera port. Make sure there is no obstruction on the camera lens such as a film or cover.
<center><img src="../images/usbcam_setup_sm.jpg" width=600/></center>
<div style="border:2px solid black; background-color:#e3ffb3; font-size:12px; padding:8px; margin-top: auto;"><i>
<h4><i>Tip</i></h4>
To execute the Python or system code in the code cells, select the cell and click the "Run" button at the top of the window.<br>Keyboard shortcut: <strong>[SHIFT][ENTER]</strong>
</i></div>
### Check to see if the device is available
Execute the following system command to list all video devices on the Jetson Nano. If your camera doesn't show up with a device id, check your connection. You should get an output similar to
```text
crw-rw----+ 1 root video 81, 0 Jun 2 17:35 /dev/video0
```
```
!ls -ltrh /dev/video*
from jetcam.rtsp_camera import RTSPCamera
```
### Create the camera object
First, create a camera object by importing the `USBCamera` class from the library by executing the following Python code cell. Please note, you can only create one `USBCamera` instance. Set the `capture_device=` to the correct number found when you listed the system video devices. If you have `/dev/video0`, then set `capture_device=0`. If you have `/dev/video1`, set `capture_device=1` in the code line below.
```
#from jetcam.usb_camera import USBCamera
#TODO change capture_device if incorrect for your system
#camera = RTSPCamera(width=224, height=224, capture_width=640, capture_height=480, capture_device='rtsp://admin:admin@192.168.0.150:554/cam/realmonitor?channel=1&subtype=1')
#camera = RTSPCamera(width=224, height=224, capture_width=640, capture_height=480, capture_device=1)
camera = RTSPCamera(width=224, height=224, capture_width=640, capture_height=480, capture_device='rtsp://admin:admin@192.168.0.150:554/cam/realmonitor?channel=1&subtype=1')
```
We can then capture a frame from the camera with the `read` method.
```
image = camera.read()
print(image.shape)
```
Calling the `read` method for `camera` also updates the camera's internal `value`. By looking at the value's `shape`, we see three numbers representing the pixel height, pixel width, and number of color channels.
```
print(camera.value.shape)
```
### Create a widget to view the image stream
We can create a "widget" to display this image in the notebook. In order to see the image, convert it from its blue-green-red format (brg8) to a format the browser can display (jpeg).
```
import ipywidgets
from IPython.display import display
from jetcam.utils import bgr8_to_jpeg
image_widget = ipywidgets.Image(format='jpeg')
image_widget.value = bgr8_to_jpeg(image)
display(image_widget)
```
You should see an image from the camera if all is working correctly. If there seems to be an image but it's fuzzy or a funny color, check to make sure there is no protective film or cap on the lens.
Now let's watch a live stream from the camera. Set the `running` value of the camera to continuously update the value in background. This allows you to attach "callbacks" to the camera value changes.
The "callback" here is the function, `update_image`, which is attached by calling the `observe` method below. `update_image` is executed whenever there is a new image available to process, which is then displayed in the widget.
```
camera.running = True
def update_image(change):
image = change['new']
image_widget.value = bgr8_to_jpeg(image)
camera.observe(update_image, names='value')
```
If you move something in front of the camera, you should now see the live video stream in the widget. To stop it, unattach the callback with the `unobserve` method.
```
camera.unobserve(update_image, names='value')
```
<div style="border:2px solid black; background-color:#e3ffb3; font-size:12px; padding:8px; margin-top: auto;"><i>
<h4><i>Tip</i></h4>
You can move the widgets (or any cell) to new window tabs in JupyterLab by right-clicking the cell and selecting "Create New View for Output". This way, you can continue to scroll down the JupyterLab notebook and still see the camera view!
</i></div>
### Another way to view the image stream
You can also use the traitlets `dlink` method to connect the camera to the widget, using a transform as one of the parameters. This eliminates some steps in the process.
```
import traitlets
camera_link = traitlets.dlink((camera, 'value'), (image_widget, 'value'), transform=bgr8_to_jpeg)
```
You can remove the camera/widget link with the `unlink` method.
```
camera_link.unlink()
```
... and reconnect it again with `link`.
```
camera_link.link()
```
## Shut down the kernel of this notebook to release the camera resource.
Return to the DLI course pages for the next instructions.
<div style="border:2px solid black; background-color:#e3ffb3; font-size:12px; padding:8px; margin-top: auto;"><i>
<h4><i>Tip</i></h4>
<p>There can only be one instance of CSICamera or USBCamera at a time. If you want to create a new camera instance, you must first release the existing one. To do so, shut down the notebook's kernel from the JupyterLab pull-down menu: <strong>Kernel->Shutdown Kernel</strong>, then restart it with <strong>Kernel->Restart Kernel</strong>.</p>
<p> If the camera setup appears "stuck" or the images "frozen", follow these steps:
<ol><li>Shut down the notebook kernel as explained above</li>
<li>Open a terminal on the Jetson Nano by clicking the "Terminal" icon on the "Launch" page</li>
<li>Enter the following command in the terminal window: <code>sudo systemctl restart nvargus-daemon</code> with password:<code>dlinano</code> </li>
</ol>
</i></div>
| github_jupyter |
```
%load_ext jupyter_probcomp.magics
%matplotlib inline
%vizgpm inline
!rm -f bdb-files/expression-400x4-juliust.bdb
%bayesdb bdb-files/expression-400x4-juliust.bdb
import pandas as pd
full_data = pd.read_csv("/home/jovyan/21521940")
full_data.shape
full_data.ix[:5, :10]
full_data.rename(columns=lambda col: col.split()[0], inplace=True)
full_data.ix[:5, :10]
import numpy as np
np.random.seed(420)
rows = np.random.choice(full_data.shape[0], 400, replace=False)
print(rows)
data = full_data[['POM121', 'POM121C', 'SPIN2A', 'SPIN2B']].ix[rows, :]
data.shape
data.columns
data.head()
data.to_csv('Expression-400x4.csv', index=False)
%bql DROP TABLE IF EXISTS "data"
%bql CREATE TABLE "data" FROM 'Expression-400x4.csv'
%bql .nullify data ''
%bql SELECT * FROM "data" LIMIT 5;
%bql SELECT COUNT(*) FROM "data";
%mml GUESS SCHEMA FOR "data"
%%mml
CREATE POPULATION FOR "data" WITH SCHEMA (
-- Use the guesses from the previous cell for all variables.
GUESS STATTYPES OF (*);
);
```
### Caveat: replace `SELECT *` by `SELECT col1, col2, col2 ...`
```
%bql .interactive_pairplot --population=data SELECT * FROM data
%mml CREATE GENERATOR FOR "data";
%multiprocess on
%mml INITIALIZE 16 MODELS IF NOT EXISTS FOR "data";
%mml ANALYZE "data" FOR 5 MINUTES
%mml ANALYZE "data" FOR 5 MINUTES
%mml ANALYZE "data" FOR 5 MINUTES
%mml .render_crosscat \
--subsample=50 --xticklabelsize=small --yticklabelsize=x-small data 0
%mml .render_crosscat \
--subsample=50 --xticklabelsize=small --yticklabelsize=x-small data 1
%mml .render_crosscat \
--subsample=50 --xticklabelsize=small --yticklabelsize=x-small data 2
%%bql
DROP TABLE IF EXISTS "correlations";
CREATE TABLE "correlations" AS
ESTIMATE
CORRELATION AS "correlation",
CORRELATION PVALUE AS "pvalue"
FROM PAIRWISE VARIABLES OF "data"
%bql .heatmap SELECT name0, name1, "correlation" FROM "correlations"
%%bql
DROP TABLE IF EXISTS dependencies;
CREATE TABLE dependencies AS
ESTIMATE
DEPENDENCE PROBABILITY AS "depprob"
FROM PAIRWISE VARIABLES OF data;
%bql .heatmap SELECT name0, name1, depprob FROM dependencies;
%bql SELECT DISTINCT(name0) FROM dependencies;
import matplotlib.pyplot as plt
import itertools
import numpy as np
import simplejson
data = %bql SELECT * FROM "data"
genes = [c for c in data.columns if c !='part']
def get_limits(vals):
return np.min(vals), np.max(vals)
def plot_simulated_data(x, y, xlim=None, ylim=None):
"""Plot simulated pairwise data against observed data."""
df_select = %bql SELECT "{x}", "{y}" FROM "data"
N = len(df_select)
df_sim = %bql SIMULATE "{x}", "{y}" FROM "data" LIMIT {N}
fig, ax = plt.subplots()
alpha = 0.9
size=2
ax.scatter(df_sim[x].values, df_sim[y].values, color='darkblue', alpha=alpha, s=size)
ax.scatter(df_select[x].values, df_select[y].values, color='red', alpha=alpha, s=size)
ax.scatter([], [], color='darkblue', label='SIMULATEd data')
ax.scatter([], [], color='red', label= 'SELECTed data')
ax.set_xlim(get_limits(df_select[x].values))
ax.set_ylim(get_limits(df_select[y].values))
ax.legend()
ax.set_xlabel(x)
ax.set_ylabel(y)
ax.set_title('SELECT vs SIMULATE')
ax.grid(True)
fig.set_size_inches(8, 6)
plt.tight_layout()
return fig, ax
fig, ax = plot_simulated_data("POM121", "POM121C")
fig, ax = plot_simulated_data("SPIN2A", "SPIN2B")
fig, ax = plot_simulated_data("POM121", "POM121C")
fig, ax = plot_simulated_data("SPIN2A", "SPIN2B")
fig, ax = plot_simulated_data("SNURF", "SNRPN")
fig, ax = plot_simulated_data("HBA1", "HBA2")
fig, ax = plot_simulated_data("HIST2H4B", "HIST2H4A")
fig, ax = plot_simulated_data("SPIN2A", "SPIN2B")
fig, ax = plot_simulated_data("ZNF587B", "ZNF814")
np.random.seed(42)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
g1, g2 = np.random.choice(genes, 2, replace=False)
fig, ax = plot_simulated_data(g1, g2)
conditional_mi_str = '''
SIMULATE
MUTUAL INFORMATION OF {g1} WITH {g2} GIVEN (part='{part}') USING 100 SAMPLES
AS "mi"
FROM MODELS OF "data"
'''
print conditional_mi_str.format(g1='rseb', g2='fabi', part='True')
%bql {conditional_mi_str.format(g1='rseb', g2='fabi', part='True')}
pairwise = list(itertools.combinations(genes, 2))
def get_mi(g1, g2, condition):
bql_query = conditional_mi_str.format(g1=g1, g2=g2, part=condition)
# print bql_query
df = %bql {bql_query}
return df.mi.values
def linfoot_transform(mi):
return np.sqrt(1 - np.exp(- 2 * mi))
total = len(pairwise)
def pair_to_name(c1, c2):
return c1 + '--' + c2
def compute_dep_results(condition):
results = {}
print '==== Conditon: ' + condition + ' ===='
for i, pair in enumerate(pairwise):
c1 = pair[0]
c2 = pair[1]
main_key = pair_to_name(c1, c2).lower()
results[main_key] = {}
mi_vals = get_mi(c1, c2, condition)
mi_vals[mi_vals < 0] = 0 # We're doing Monte Carlo approximations which can be < 0.
results[main_key]['ri'] = linfoot_transform(mi_vals).tolist()
if ((i+1) % 10) == 0:
print 'Computed {} out of {} pairwise relationships'.format(i+1, len(pairwise))
print ''
return results
conditions = ['True', 'False']
results = {'mi':{}}
for condition in conditions:
results['mi'][condition] = compute_dep_results(condition)
path = 'cmi-results/results-Bridge-demo-Arac-part.json'
print path
with open(path, 'w') as outfile:
outfile.write(simplejson.dumps(results, ignore_nan=True))
```
| github_jupyter |
## 1. Calculate Area of a Circle
#### Write a Python program which accepts the radius of a circle from the user and compute the area.
###### Program Console Sample Output 1:
###### Input Radius: 0.5
###### Area of Circle with radius 0.5 is 0.7853981634
#### Program Console Sample Output 1:¶
```
import math
r = float(input('Input Radius :'))
A = pi*r**2
print(f"Area of Circle with radius {r} is {round(A,10)} ")
```
## 2. Check Number either positive, negative or zero
#### Write a Python program to check if a number is positive, negative or zero
###### Program Console Sample Output 1:
###### Enter Number: -1
##### Negative Number Entered
###### Program Console Sample Output 2:
##### Integer: 3
##### Positive Number Entered
###### Program Console Sample Output 3:
##### Integer: 0
###### Zero Entered
```
value = int(input("Enter Number : "))
if value < 0:
print("Negative Number Entered")
value1 = int(input("Enter Number : "))
if value1 > 0:
print("Positive Number Entered")
value2 = int(input("Enter Number : "))
if value2 == 0:
print("Zero Entered")
```
## 3. Divisibility Check of two numbers
#### Write a Python program to check whether a number is completely divisible by another number. Accept two integer values form the user
##### Program Console Sample Output 1:
###### Enter numerator: 4
###### Enter Denominator: 2
##### Number 4 is Completely divisible by 2
###### Program Console Sample Output 2:
##### Enter numerator: 7
##### Enter Denominator: 4
###### Number 7 is not Completely divisible by 4
```
value = int(input("Enter numerator : "))
value1 = int(input("Enter Denominator : "))
if value%value1 == 0:
print(f"Number {value} is Completely divisible by {value1}")
else:
print(f"Number {value} is not Completely divisible by {value1}")
value = int(input("Enter numerator : "))
value1 = int(input("Enter Denominator : "))
if value%value1 == 0:
print(f"Number {value} is Completely divisible by {value1}")
else:
print(f"Number {value} is not Completely divisible by {value1}")
```
## 4. Calculate Volume of a sphere
##### Write a Python program to get the volume of a sphere, please take the radius as input from user
##### Program Console Output:
##### Enter Radius of Sphere: 1
###### Volume of the Sphere with Radius 1 is 4.18
```
import math
r = int(input("Enter Radius of Sphere : "))
v = (4/3)*pi*r**3
print(f"Volume of the sphere with Radius {r} is {round(v,2)}")
```
## 5. Copy string n times
#### Write a Python program to get a string which is n (non-negative integer) copies of a given string.
##### Program Console Output:
##### Enter String: Hi
###### How many copies of String you need: 4
###### 4 Copies of Hi are HiHiHiHi
```
STRING = input("Enter String : ")
how_many_copies = int(input("How many copies of String you need: "))
total_copies = (STRING)*how_many_copies
print(f"{how_many_copies} Copies of {STRING} are {total_copies}")
```
## 6. Check if number is Even or Odd
### Write a Python program to find whether a given number (accept from the user) is even or odd, print out an appropriate message to the user
#### Program Console Output 1:
##### Enter Number: 4
###### 4 is Even
#### Program Console Output 2:
##### Enter Number: 9
###### 9 is Odd
```
value1 = int(input("Enter Number : "))
if value1%2==0:
print(f"{value1} is Even")
else:
print(f"{value1} is Odd")
value2 = int(input("Enter Number : "))
if value2%2==0:
print(f"{value2} is Even")
else:
print(f"{value2} is Odd")
```
## 7. Vowel Tester
### Write a Python program to test whether a passed letter is a vowel or not
#### Program Console Output 1:
##### Enter a character: A
###### Letter A is Vowel
#### Program Console Output 2:
##### Enter a character: e
###### Letter e is Vowel
#### Program Console Output 2:
##### Enter a character: N
###### Letter N is not Vowel
```
value = input("Enter a character: ")
string =["a","e","i","o",'u']
i=0
while i<=4:
if value.lower() == string[i]:
print(f"Letter {value} is Vowel")
break
else:
print(f"Letter {value} is not Vowel")
i+=1
value1 = input("Enter a character: ")
string =["a","e","i","o",'u']
i=0
while i<=4:
if value1.lower() == string[i]:
print(f"Letter {value1} is Vowel")
break
elif i==4:
print(f"Letter {value1} is not Vowel")
i+=1
value2 = input("Enter a character: ")
string =["a","e","i","o",'u']
i=0
while i<=4:
if value2.lower() == string[i]:
print(f"Letter {value2} is Vowel")
break
else:
print(f"Letter {value2} is not Vowel")
break
i+=1
```
## 8. Triangle area
### Write a Python program that will accept the base and height of a triangle and compute the area
###### Reference:
https://www.mathgoodies.com/lessons/vol1/area_triangle
```
base = int(input("Enter Base of traingle : "))
height = int(input("Enter height of traingle : "))
A=(1/2)*base*height
print(f"Area of traingle = {A} when base = {base} and height = {height}")
```
## 9. Calculate Interest
### Write a Python program to compute the future value of a specified principal amount, rate of interest, and a number of years
#### Program Console Sample 1:
##### Please enter principal amount: 10000
###### Please Enter Rate of interest in %: 0.1
###### Enter number of years for investment: 5
###### After 5 years your principal amount 10000 over an interest rate of 0.1 % will be 16105.1
```
p=int(input("Please enter principal amount: "))
I=float(input("lease Enter Rate of interest in %: "))
t =int(input("Enter number of years for investment: "))
A=p*(1+I)**t
A = round(A,2)
print(f"After {t} years your principal amount {p} over an interest rate of {I}% will be {A}")
```
## 10. Euclidean distance
### write a Python program to compute the distance between the points (x1, y1) and (x2, y2).
#### Program Console Sample 1:
###### Enter Co-ordinate for x1: 2
###### Enter Co-ordinate for x2: 4
###### Enter Co-ordinate for y1: 4
###### Enter Co-ordinate for y2: 4
###### Distance between points (2, 4) and (4, 4) is 2
###### Reference:
https://en.wikipedia.org/wiki/Euclidean_distance
```
import math
x1 = int(input("Enter Co-ordinate for x1: "))
x2 = int(input("Enter Co-ordinate for x2: "))
y1 = int(input("Enter Co-ordinate for y1: "))
y2 = int(input("Enter Co-ordinate for y2: "))
d = int(math.sqrt(pow((x2-x1),2)+pow((y2-y1),2)))
print(f"Distance between points ({x1},{y1}) and ({x2},{y2}) is {d}")
```
## 11. Feet to Centimeter Converter
### Write a Python program to convert height in feet to centimetres.
##### Program Console Sample 1:
###### Enter Height in Feet: 5
###### There are 152.4 Cm in 5 ft
###### Reference:
https://www.rapidtables.com/convert/length/feet-to-cm.html
```
f = int(input("Enter Height in Feet: "))
h =30.48
c = f*h
print(f"There are {c} cm in {f} ft")
```
## 12. BMI Calculator
### Write a Python program to calculate body mass index
##### Program Console Sample 1:
###### Enter Height in Cm: 180
###### Enter Weight in Kg: 75
###### Your BMI is 23.15
```
cm = int(input("Enter Height in Cm: "))
m = cm/100
kg = int(input("Enter Weight in Kg: "))
BMI = round((kg/m**2),2)
print(f"Your BMI is {BMI}")
```
## 13. Sum of n Positive Integers
### Write a python program to sum of the first n positive integers
#### Program Console Sample 1:
###### Enter value of n: 5
###### Sum of n Positive integers till 5 is 15
```
n = int(input("Enter value of n: "))
i=0
value=0
while i<=n:
value = value+i
i+=1
print(f"Sum of n Positive integers till {n} is {value}")
```
## 14. Digits Sum of a Number
### Write a Python program to calculate the sum of the digits in an integer
#### Program Console Sample 1:
##### Enter a number: 15
###### Sum of 1 + 5 is 6
#### Program Console Sample 2:
##### Enter a number: 1234
###### Sum of 1 + 2 + 3 + 4 is 10
```
a = input("Enter a number: ")
b= len(a)
i=0
value=0
while i<b:
c= int(a[i])
value = value + c
i+=1
if b==1:
print(f"sum of {int(a[0])} is {value}")
i+=1
elif b==2:
print(f"sum of {int(a[0])} + {int(a[1])} is {value}")
i+=1
elif b==3:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} is {value}")
i+=1
elif b==4:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} + {int(a[3])} is {value}")
i+=1
elif b==5:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} + {int(a[3])} + {int(a[4])} is {value}")
i+=1
a = input("Enter a number: ")
b= len(a)
i=0
value=0
while i<b:
c= int(a[i])
value = value + c
i+=1
if b==1:
print(f"sum of {int(a[0])} is {value}")
i+=1
elif b==2:
print(f"sum of {int(a[0])} + {int(a[1])} is {value}")
i+=1
elif b==3:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} is {value}")
i+=1
elif b==4:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} + {int(a[3])} is {value}")
i+=1
elif b==5:
print(f"sum of {int(a[0])} + {int(a[1])} + {int(a[2])} + {int(a[3])} + {int(a[4])} is {value}")
i+=1
```
| github_jupyter |
# This notebook serves as a guild to successfully excute the given task covering introduction to sklearn. It should be used together with the
- Intro to sklearn practical notebook
- Starter notebook (to be provided as well)
# Here is the competition link
- https://zindi.africa/competitions/data-science-nigeria-2019-challenge-1-insurance-prediction/
- You can join the competition (make sure you are registered on zindi before hand) and download the data here
- https://zindi.africa/competitions/data-science-nigeria-2019-challenge-1-insurance-prediction/data
- Also make submissions on
- https://zindi.africa/competitions/data-science-nigeria-2019-challenge-1-insurance-prediction/submissions
- Upload your csv submission file to make submission
# Import the libraries needed to read-in and manipulate the data
# Set a random seed to ensure the reproducibility of your notebook
# Read in your train, test, variable definition and sample submission datasets using pandas
# Explore the training and test datasets checking their head, info, description, shape, value counts etc. Take pointers from the sklearn pratical notebook
# Note that any changes been made to your training dataset shouldbe replicates to the test dataset to ensure they both match
# Check for duplicate values and remove them (if any)
# Remove any column which doesnt contribute toward predicting our churn value (Note: this should be done in both training and test dataset)
# Check for the presence of missing variables and also identify the categorical columns within your training and test dataset
# Handle the missing variables in the categorical columns in your training and test dataset
- hint: your can fill the nan or missing values with the mode values of the columns or any other methods of your choice
# Handle the missing variables in the numerical columns using any method of your choice (within both your training and test dataset)
# Encode the categorical Columns present in your training and test dataset
- hint: you can utilize the pandas (pd.get_dummies()) function, sklearn's One Hot encoder or Ordinal encoder functions within (sklearn.preprocessing)
# Check that all columns are of a numerical type (float or int) and ensure there are no missing values in your training and test dataset
# Split your train data into X and y
# Import all libraries you would need for the modelling stage
# Split your train data using train test split for use in identifying the best model
# Train 4 different machine learning models using the data above
- Take pointers from the intro to sklearn pratical notebook as well as the starter notebook for the competition
- Choose the appropriate model type for the problem at hand (Classification problem)
- When predicting with the models use predict_probablity method instead of of the predict method and isolate the predictions for the positive or "1" Class.
- the evaluation matrix being used for this competition is the "roc_auc" metric available in sklearn. Use it to evaluate your model performance on the X_test.
```
# example for logistic regression model
lg = LogisticRegression(max_iter=10000, random_state=5, n_jobs=-1, verbose=5)
lg.fit(X_train, y_train)
pred = lg.predict_proba(X_test)
pred = [x[1] for x in pred]
print(roc_auc_score(y_test, pred))
```
# Identify the best performing model
# make prediction on the test data using stratified Kfold cross validation
- Determine the number of folds
# ADD your prediction to the CHURN column of the sample submission dataset
- Save it to a csv file and make your submission using the csv file on zindi
- hint: Check starter notebook
# Improve your model by
- Tuning hyperparameters
- Feature Selection
- Feature Engineering etc.
- Then submit the new csv file. repeat this until you get the best score....you can check the top performing score on the leaderboard
# Good Luck!!!
| github_jupyter |
<a href="https://colab.research.google.com/github/krakowiakpawel9/machine-learning-bootcamp/blob/master/unsupervised/02_dimensionality_reduction/03_pca_wine.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
* @author: krakowiakpawel9@gmail.com
* @site: e-smartdata.org
### scikit-learn
Strona biblioteki: [https://scikit-learn.org](https://scikit-learn.org)
Dokumentacja/User Guide: [https://scikit-learn.org/stable/user_guide.html](https://scikit-learn.org/stable/user_guide.html)
Podstawowa biblioteka do uczenia maszynowego w języku Python.
Aby zainstalować bibliotekę scikit-learn, użyj polecenia poniżej:
```
!pip install scikit-learn
```
Aby zaktualizować do najnowszej wersji bibliotekę scikit-learn, użyj polecenia poniżej:
```
!pip install --upgrade scikit-learn
```
Kurs stworzony w oparciu o wersję `0.22.1`
### Spis treści:
1. [Import bibliotek](#0)
2. [Załadowanie danych](#1)
3. [Podział na zbiór treningowy i testowy](#2)
4. [Standaryzacja](#3)
5. [PCA](#4)
### <a name='0'></a> Import bibliotek
```
import pandas as pd
import numpy as np
import plotly.graph_objects as go
import plotly.express as px
np.set_printoptions(precision=4, suppress=True, edgeitems=5, linewidth=200)
```
### <a name='1'></a> Załadowanie danych
```
df_raw = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/wine/wine.data', header=None)
df = df_raw.copy()
df.head()
data = df.iloc[:, 1:]
target = df.iloc[:, 0]
data.head()
target.value_counts()
```
### <a name='2'></a> Podział na zbiór treningowy i testowy
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target)
print(f'X_train shape: {X_train.shape}')
print(f'X_test shape: {X_test.shape}')
```
### <a name='3'></a> Standaryzacja
```
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train_std = scaler.fit_transform(X_train)
X_test_std = scaler.transform(X_test)
X_train_std[:5]
```
### <a name='4'></a> PCA
```
from sklearn.decomposition import PCA
pca = PCA(n_components=3)
X_train_pca = pca.fit_transform(X_train_std)
X_test_pca = pca.transform(X_test_std)
X_train_pca.shape
```
Wyjaśniona wariancja
```
results = pd.DataFrame(data={'explained_variance_ratio': pca.explained_variance_ratio_})
results['cumulative'] = results['explained_variance_ratio'].cumsum()
results['component'] = results.index + 1
results
fig = go.Figure(data=[go.Bar(x=results['component'], y=results['explained_variance_ratio'], name='explained variance ratio'),
go.Scatter(x=results['component'], y=results['cumulative'], name='cumulative explained variance')],
layout=go.Layout(title=f'PCA - {pca.n_components_} components', width=950, template='plotly_dark'))
fig.show()
X_train_pca_df = pd.DataFrame(data=np.c_[X_train_pca, y_train], columns=['pca1', 'pca2', 'pca3', 'target'])
X_train_pca_df.head()
px.scatter_3d(X_train_pca_df, x='pca1', y='pca2', z='pca3', color='target', template='plotly_dark', width=950)
X_train_pca[:5]
X_test_pca[:5]
```
| github_jupyter |
```
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
import h5py
```
## Find all files
```
from glob import glob
#files_loc = "/gpfs/slac/atlas/fs1/u/rafaeltl/Muon/toy_sim/si-mu-lator/out_files/"
files_loc = "/Users/willocq/ATLAS/muons4ML/si-mu-lator/run/"
files_nobkg = glob(files_loc+'*_bkg0_*.h5')
#files_bkg = glob(files_loc+'*bkg1*.h5')
files_bkg = glob(files_loc+'nsw_n1000_b1_z10*.h5')
```
## Open one file
```
f5 = h5py.File( files_bkg[0], 'r' )
f5.keys()
```
### File structure
The file is structured as follows:
- Each row is an event (a full readout of the detector module within a given time window specified in the detector card)
- For each event, you have the truth information of the one incoming muon (if there is one, specified in the file production step)
- For each event, you have a set of signal readouts
The arrays in the file are:
- 'ev_mu_phi', 'ev_mu_theta', 'ev_mu_time', 'ev_mu_x', 'ev_mu_y': muon truth
- 'ev_n_mu_signals', 'ev_n_signals': number of signals that were left by a muon, number of total signals
- 'signal_keys': array of names of signal variables
- 'signals': array of signals
#### The signals array corresponds to the output of the `get_info_wrt_plane` function in class `Signal`:
```
hit_dict = {'is_muon': self.is_muon,
'x': self.x,
'y': self.y,
'z': plane.z,
'ptype': plane.p_type.asint(),
'ptilt': plane.tilt,
'poffset': plane.offset,
'time': self.time,
'projX_at_rightend_x': float(x_rightend.x),
'projX_at_rightend_y': float(x_rightend.y),
'projX_at_middle_x': float(x_middle.x),
'projX_at_middle_y': float(x_middle.y),
'projY_at_topend_x': float(y_topend.x),
'projY_at_topend_y': float(y_topend.y),
'projY_at_middle_x': float(y_middle.x),
'projY_at_middle_y': float(y_middle.y),
'seg_ix': self.seg_ix,
'rdrift': self.rdrift
}
```
```
sig_vars = [ 'is_muon',
'x',
'y',
'z',
'ptype',
'ptilt',
'poffset',
'time',
'projX_at_rightend_x',
'projX_at_rightend_y',
'projX_at_middle_x',
'projX_at_middle_y',
'projY_at_topend_x',
'projY_at_topend_y',
'projY_at_middle_x',
'projY_at_middle_y',
'seg_ix',
'rdrift']
```
### Transform objects in arrays
```
data = {}
for kk in f5.keys():
data[kk] = np.array( f5[kk] )
# show first five events
print(
'mu x', data['ev_mu_x'][:5], '\n',
'mu theta', data['ev_mu_theta'][:5], '\n',
'n mu signals', data['ev_n_mu_signals'][:5], '\n',
'n signals', data['ev_n_signals'][:5], '\n',
)
data['signals'].shape
```
Note that the shape of the signals is, for example, (1000, 39, 18). This means that there are 1000 events, the maximum number of signals in this file is 39, and for each signal there are 18 features.
```
idx_mu = sig_vars.index('is_muon')
idx_x_inter = sig_vars.index('x')
idx_x_strip = sig_vars.index('projX_at_middle_x')
idx_z = sig_vars.index('z')
idx_ptype = sig_vars.index('ptype')
data['signals'][:, :, idx_z].flatten() # show variable z for all signals from all events
fig, axs = plt.subplots(5, 4, figsize=(14, 14), gridspec_kw={'wspace':0.5, 'hspace': 0.5})
axs = axs.flatten()
mu_hits = data['signals'][:, :, idx_mu].flatten() == 1
empty_hits = data['signals'][:, :, idx_mu].flatten() == -99 # not a signal
for iv in range(data['signals'].shape[2]):
ivar = data['signals'][:, :, iv].flatten()
maxvar = np.max( ivar[(~empty_hits)] )
minvar = np.min( ivar[(~empty_hits)] )
axs[iv].hist( ivar[(mu_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=20, label='muon hits' )
axs[iv].hist( ivar[(~mu_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=20, label='bkg hits' )
axs[iv].set_xlabel(sig_vars[iv])
axs[0].legend()
plt.show()
fig, axs = plt.subplots(2, 2, figsize=(14, 8), gridspec_kw={'wspace':0.5, 'hspace': 0.5})
axs = axs.flatten()
iv_pos = [idx_x_strip, idx_z, idx_x_inter, idx_z]
mm_hits = data['signals'][:, :, idx_ptype].flatten() == 0
stgc_hits = data['signals'][:, :, idx_ptype].flatten() == 2
# MicroMegas and sTGC hit positions
for i,iv in enumerate(iv_pos):
ivar = data['signals'][:, :, iv].flatten()
maxvar = np.max( ivar[(~empty_hits)] )
minvar = np.min( ivar[(~empty_hits)] )
if i < 2: # MM hits
axs[i].hist( ivar[(mu_hits)&(mm_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=64, label='muon hits MM' )
axs[i].hist( ivar[(~mu_hits)&(mm_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=64, label='bkg hits MM' )
else: # sTGC hits
axs[i].hist( ivar[(mu_hits)&(stgc_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=100, label='muon hits sTGC' )
axs[i].hist( ivar[(~mu_hits)&(stgc_hits)&(~empty_hits)], histtype='step', range=(minvar, maxvar), bins=100, label='bkg hits sTGC' )
axs[i].set_xlabel(sig_vars[iv])
axs[0].legend()
axs[2].legend()
plt.show()
```
| github_jupyter |
### Module 4 Cleaning data with pandas
### In-class exercises Answers
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# Q.1 (a)
ser1 = pd.Series([0.25,0.5,0.75,1.0], index=["a","b","c","d"])
ser1
# 1. (b)
# use loc for explicit index
# use iloc for implicit index
print (ser1.loc[["b","c"]]) # pay attention to the [] inside the []
print ()
print (ser1.iloc[[1,3]])
#1(c)
ser1.rename(index={"a":"i","b":"ii","c":"iii","d":"iv"}, inplace=True)
# inplace = True to replace the original series
ser1
#1(d)
ser1.sort_values(ascending=False)
# Q.2 (a)
pop = {"California":38332521, "Texas":26448193, "New York":19651127, "Florida":19552860, "Illinois":12882135}
area = {"California":423967, "Texas":695662, "New York":141297, "Florida":170312, "Illinois":14995}
df = pd.DataFrame({"state population":pop,"state area":area})
df.index
# 2b
df[df["state population"]<20000000]
# 2c
df["state density"] = df["state population"] / df["state area"]
df
#2d
df.loc["California"]["state area"] # to access a given row with an explicit index, use loc
#2e
df = df[["state area","state population","state density"]]
df
#2f
df.iloc[-2:] # iloc for implicit index of rows
#Q3
df = pd.DataFrame([[1, np.nan, 2, np.nan],
[2, 3, 5, 3.5],
[np.nan, 4, 6, np.nan]])
df
#3a
df.columns=["i","ii","iii","iv"]
df.index=["a","b","c"]
df
#3b
df.dropna()
#3c
df.dropna(axis=1) # axis = 1 is for columns
#3d
df.dropna(axis=1, how="all") # you have to put "" around the word all
#3e
df.fillna(0)
#3f
df_new = df.copy() # don't use df_new = df Any changes in df_new will also affect df
for column in df_new.columns:
df_new[column] = df_new[column].fillna(df_new[column].mean())
print(df)
print()
print(df_new)
# Q.4
df5 = pd.DataFrame(np.random.randint(0,11,size=(2,3)),columns=["A","B","C"], index=[1,2])
df6 = pd.DataFrame(np.random.randint(0,11,size=(2,3)),columns=["D","B","C"], index=[1,2])
print(df5)
print(df6)
df_new = df5.append(df6)
df_new
#Q.5
df1 = pd.DataFrame({'employee': ['Bob', 'Jake', 'Lisa', 'Sue'],
'group': ['Accounting', 'Engineering', 'Engineering', 'HR']})
df2 = pd.DataFrame({'employee': ['Lisa', 'Bob', 'Jake', 'Sue'],
'hire_date': [2004, 2008, 2012, 2014]})
df3 = pd.DataFrame({'group': ['Accounting', 'Engineering', 'HR'],
'supervisor': ['Carly', 'Guido', 'Steve']})
df4 = pd.DataFrame({'name': ['Bob', 'Jake', 'Lisa', 'Sue'],
'salary': [70000, 80000, 120000, 90000]})
# merge df1 with df2, the key is "employee"
df_merged1 =df1.merge(df2, on="employee") # on = "merge_key"
df_merged1
# merge df_merged1 with df3, the key is "group"
df_merged2 =df_merged1.merge(df3, on="group") # on = "merge_key"
df_merged2
# merge df_merged2 with df4, the key for df_merged2 = "employee", the key for df4 = "name"
df_final =df_merged2.merge(df4, left_on="employee", right_on="name")
del(df_final["name"]) # the columns employee and name are duplicates
df_final
# Q.6
# import the datasets
pop = pd.read_csv("state-population.csv")
area = pd.read_csv("state-areas.csv")
abbre = pd.read_csv("state-abbrevs.csv")
# show the first few rows of each dataset
print(pop.head())
print()
print(area.head())
print()
print(abbre.head())
# first merge the df pop with the df abbre
df_merged1 = pop.merge(abbre, how="outer",left_on="state/region", right_on="abbreviation")
# how="outer" will retain the rows in "pop" with key which do not match with "abbre"
# how = "inner" will only keep the rows in "pop" with key matching with "abbre"
del(df_merged1["abbreviation"])
df_merged1
# check any missing values before additional merge
# as we use how="outer", it is likely that we have some rows not matched and hence some missing values
df_merged1.isnull().any()
# let's check the rows with missing population values
df_merged1[df_merged1["population"].isnull()].head()
df_merged1.loc[df_merged1['state'].isnull(), 'state/region'].unique()
# so "PR" and "USA" have missing values.
df_merged1.loc[df_merged1['state/region'] == 'PR', 'state'] = 'Puerto Rico'
df_merged1.loc[df_merged1['state/region'] == 'USA', 'state'] = 'United States'
df_merged1.isnull().any()
# then merge with the df area
df_final = df_merged1.merge(area, how="left", on="state")
df_final
df_final.isnull().any()
df_final['state'][df_final['area (sq. mi)'].isnull()].unique()
#6b we can drop the missing values
df_final.dropna(inplace=True)
df_final.head()
#6c We just extract the rows for year ==2010 and ages == total
df_new = df_final[(df_final["ages"]=="total") & (df_final["year"]==2010)]
df_new.set_index("state", inplace = True)
density = df_new["population"] /df_new["area (sq. mi)"]
density.head()
# 6d
density.sort_values(ascending = False)
# Q.7 (a)
import seaborn as sns
planets = sns.load_dataset('planets')
planets.head(10) # this will display the first 10 rows of the dataframe.
# 7b
planets.groupby("method")["orbital_period"].median()
#7c
# find the oldest year and the most recent year of discovery.
print (planets["year"].max())
print (planets["year"].min())
# the decades should be from 1980s to 2010s
planets["decade"] = planets["year"]//10 * 10
planets["decade"] = planets["decade"].astype(str) + "s"
planets.head()
planets.groupby(["method","decade"])["number"].sum().unstack().fillna(0)
# without unstack(), it will return a series.
# wit unstack(), it will return a dataframe.
#Q.8
data = np.random.randint(1,100,size=100).reshape(25,4)
df = pd.DataFrame(data, columns=["W","X","Y","Z"])
df.head()
df.tail(5)
#8b
df[["X","Z"]].head()
df["W"].value_counts()
df[df["Y"]>50]
#8e
df.loc[df["Z"]<30,"Y"]=30 # use loc to find rows with df["Z"]<30
df
#8f
df["A"]=df["X"]*df["Z"]
df.head()
#8g
df.drop(df[df["Z"]<20].index)
#df[df["Z"]<20] is a dataframe
# we use df[df["Z"]<20].index to find the index of the rows and then drop them.
# Q.9 (a)
df = pd.read_csv("Boston_Housing.csv")
df.head(10)
#9b
df_small =df.drop(["CHAS", "NOX", "RAD", "LSTAT"], axis=1) # don't forget axis=1
df_small.head()
#9c
print("mean number of rooms per dwelling = ", round(df_small["RM"].mean(),1))
print("median age of houses = ", round(df_small["AGE"].median(),1)) # use round to remove decimals.
print("mean distances = ", "{:.3}".format(df_small["DIS"].median())) # use "{:}.format()" to express the number in an appropriate format
print("percentage of houses with a price less than $20,000 =", "{:.2%}".format(sum(df_small["PRICE"]<20)/len(df_small))) # first sum the number of houses less than $20K, then divide the sum by the number of rows.
# Q.10(a)
data_visit = pd.read_csv("visit_data.csv")
data_visit.columns
#10b find entries with the same first name and last name.
data_visit[data_visit.duplicated(["first_name","last_name"])][["first_name","last_name"]]
# duplicate last name and first name are missing, it should not be a problem.
#10c find entries with the same email
data_visit[data_visit.duplicated(["email"])]["email"]
# there are no duplicate entries of emails.
#10d check missing values for visit
print ("number of missing entries for visit =", data_visit["visit"].isnull().sum())
# 10e don't use df_new = data_visit, this will also change the original when we change the copy
df_new =data_visit.copy()
# 10f drop rows for any null vaule in the column "visit"
df_new.dropna(axis=0, subset=["visit"])
#10f how many rows dropped
a = len(df_new)
b = len (df_new.dropna(axis=0, subset=["visit"]))
print("the number of rows dropped = ", a - b)
#10g instead of dropping, we can keep the entries with visit between 100 and 2900
df1 = df_new[(df_new['visit'] <= 2900) & (df_new['visit'] >= 100)]
# Q.11
olympics = pd.read_csv("olympics.csv", index_col=0, skiprows=1)
#1. we use the column ountry as index
#2. the first row of the dataframe is useless. So, we skip it.
olympics.head()
olympics.columns
# we need to do some wrangling for column names and the index.
for col in olympics.columns:
if col[:2]=='01': # check if first two characters of the column name is "01"
olympics.rename(columns={col:'Gold'+col[4:]}, inplace=True)
if col[:2]=='02':
olympics.rename(columns={col:'Silver'+col[4:]}, inplace=True)
if col[:2]=='03':
olympics.rename(columns={col:'Bronze'+col[4:]}, inplace=True)
if col[:1]=='№':
olympics.rename(columns={col:'#'+col[1:]}, inplace=True)
olympics.columns
#"Gold": number of gold medals in the summer game
#"Gold.1": number of gold medals in the winter game
#"Gold.2": number of gold medals in both the summer game and the winter game
names_ids = olympics.index.str.split('\s\(') # split the index by '('
names_ids
olympics.index = names_ids.str[0] # the [0] element is the country name (new index)
olympics['ID'] = names_ids.str[1].str[:3] # the [1] element is the abbreviation or ID (take first 3 characters from that)
olympics.drop("Totals", inplace=True) # drop the row "Totals"
olympics
#11a first country of the dataframe, use iloc to call the implicit index.
olympics.iloc[0]
#11b use idxmax() to find the index with maximum value of a column
print("the country with the most gold medals in the summer games =", olympics["Gold"].idxmax())
#11c
import numpy as np
diff = np.abs(olympics["Gold"] - olympics["Gold.1"])
print("The country with biggest difference between their summer and winter gold medal counts is =", diff.idxmax())
#11d we use "Gold.2", "Silve.2" and "Bronze.2"
points = olympics["Gold.2"]*3 + olympics["Silver.2"]*2 + olympics["Bronze.2"]
points.head()
# Q.12
birth = pd.read_csv("births.csv")
birth.head()
#12a
birth.describe() # use describe to who summary statistics
#12b
birth.groupby("month")["births"].sum()
#12c
birth["decade"] = birth["year"]//10*10
birth["decade"]=birth["decade"].astype(str) +"s"
birth.groupby(["decade","gender"])["births"].sum().unstack() # unstack the series to create a dataframe
#12d create a dataframe which shows the number of male and female births in each year.
birth_year = birth.groupby(["year","gender"])["births"].sum().unstack()
birth_year.head()
#12d
import matplotlib.pyplot as plt
plt.plot(birth_year["M"], label="Male births")
plt.plot(birth_year["F"], label = "Female births")
plt.legend()
#Q.13
mpg = pd.read_csv("mpg.csv")
mpg.info() # use .info(), not .describe()
#13b recode "drv"
drv_newgroup ={"drv":{"f":"forward","r":"rear","4":"four wheel"}}
mpg.replace(drv_newgroup, inplace=True)
mpg["drv"].value_counts()
#13c missing values
mpg.isnull().any()
#13d
mpg.groupby(["class","year"])["hwy"].mean().unstack()
#13e
mpg.groupby(["class"])["cty"].min()
#13f
import matplotlib.pyplot as plt
variables = [mpg["displ"],mpg["hwy"]]
plt.scatter(variables[0], variables[1])
plt.xlabel("displ")
plt.ylabel("hwy")
#13g and h
mpg_reindexed = mpg.set_index("model")
mpg_reindexed.loc["corvette"]
#13i
mpg.sort_values(by=["year","manufacturer"], ascending=[True,False])
#13j
cols = list(mpg.columns) # convert the column index into a list
cols.remove("class") # remove the element "class" from the list
cols.insert(0,"class") # insert the element "class" as the first element
mpg2 = mpg[cols]
mpg2.head()
#Q.14 (a)
who = pd.read_csv("who.csv")
who.head(20)
#Q.14 (b)
who_new = who.drop(["iso2","iso3"], axis=1)
who_new.columns
#14c(i)
# convert newrel into new_rel
columns = []
for word in who_new.columns:
if word[:6] != "newrel":
columns.append(word)
elif word[:6] == "newrel":
word = "new_rel"+ word[6:]
columns.append(word)
who_new.columns = columns
who_new.columns
#14c(ii)
sub_columns = ['new_sp_m014', 'new_sp_m1524', 'new_sp_m2534',
'new_sp_m3544', 'new_sp_m4554', 'new_sp_m5564', 'new_sp_m65',
'new_sp_f014', 'new_sp_f1524', 'new_sp_f2534', 'new_sp_f3544',
'new_sp_f4554', 'new_sp_f5564', 'new_sp_f65', 'new_sn_m014',
'new_sn_m1524', 'new_sn_m2534', 'new_sn_m3544', 'new_sn_m4554',
'new_sn_m5564', 'new_sn_m65', 'new_sn_f014', 'new_sn_f1524',
'new_sn_f2534', 'new_sn_f3544', 'new_sn_f4554', 'new_sn_f5564',
'new_sn_f65', 'new_ep_m014', 'new_ep_m1524', 'new_ep_m2534',
'new_ep_m3544', 'new_ep_m4554', 'new_ep_m5564', 'new_ep_m65',
'new_ep_f014', 'new_ep_f1524', 'new_ep_f2534', 'new_ep_f3544',
'new_ep_f4554', 'new_ep_f5564', 'new_ep_f65', 'new_rel_m014',
'new_rel_m1524', 'new_rel_m2534', 'new_rel_m3544', 'new_rel_m4554',
'new_rel_m5564', 'new_rel_m65', 'new_rel_f014', 'new_rel_f1524',
'new_rel_f2534', 'new_rel_f3544', 'new_rel_f4554', 'new_rel_f5564',
'new_rel_f65']
who_new2 = pd.melt(who_new, id_vars = ["country","year"],value_vars=sub_columns)
who_new2.columns = ["country","year","key","case"]
who_new2
#14c(iii) drop all missing values of case
who_new2.dropna(subset=["case"],axis=0, inplace=True)
#14c(iv) split each value of the column "key"
who_new2[["new","type","sexage"]] = who_new2["key"].str.split("_", expand=True)
who_new2.head()
#14c(v) split each value of the column "sexage"
gender = []
age = []
for value in who_new2["sexage"]:
gender.append(value[0])
age.append(value[1:])
who_new2["gender"] = gender
who_new2["age"] = age
who_new2.drop(["key","sexage"],axis=1, inplace=True)
who_new2.head()
```
| github_jupyter |
# Keras Syntax Basics
With TensorFlow 2.0 , Keras is now the main API choice. Let's work through a simple regression project to understand the basics of the Keras syntax and adding layers.
## The Data
To learn the basic syntax of Keras, we will use a very simple fake data set, in the subsequent lectures we will focus on real datasets, along with feature engineering! For now, let's focus on the syntax of TensorFlow 2.0.
Let's pretend this data are measurements of some rare gem stones, with 2 measurement features and a sale price. Our final goal would be to try to predict the sale price of a new gem stone we just mined from the ground, in order to try to set a fair price in the market.
### Load the Data
```
import pandas as pd
df = pd.read_csv('../DATA/fake_reg.csv')
df.head()
```
### Explore the data
Let's take a quick look, we should see strong correlation between the features and the "price" of this made up product.
```
import seaborn as sns
import matplotlib.pyplot as plt
sns.pairplot(df)
```
Feel free to visualize more, but this data is fake, so we will focus on feature engineering and exploratory data analysis later on in the course in much more detail!
### Test/Train Split
```
from sklearn.model_selection import train_test_split
# Convert Pandas to Numpy for Keras
# Features
X = df[['feature1','feature2']].values
# Label
y = df['price'].values
# Split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3,random_state=42)
X_train.shape
X_test.shape
y_train.shape
y_test.shape
```
## Normalizing/Scaling the Data
We scale the feature data.
[Why we don't need to scale the label](https://stats.stackexchange.com/questions/111467/is-it-necessary-to-scale-the-target-value-in-addition-to-scaling-features-for-re)
```
from sklearn.preprocessing import MinMaxScaler
help(MinMaxScaler)
scaler = MinMaxScaler()
# Notice to prevent data leakage from the test set, we only fit our scaler to the training set
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
```
# TensorFlow 2.0 Syntax
## Import Options
There are several ways you can import Keras from Tensorflow (this is hugely a personal style choice, please use any import methods you prefer). We will use the method shown in the **official TF documentation**.
```
import tensorflow as tf
from tensorflow.keras.models import Sequential
help(Sequential)
```
## Creating a Model
There are two ways to create models through the TF 2 Keras API, either pass in a list of layers all at once, or add them one by one.
Let's show both methods (its up to you to choose which method you prefer).
```
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
```
### Model - as a list of layers
```
model = Sequential([
Dense(units=2),
Dense(units=2),
Dense(units=2)
])
```
### Model - adding in layers one by one
```
model = Sequential()
model.add(Dense(2))
model.add(Dense(2))
model.add(Dense(2))
```
Let's go ahead and build a simple model and then compile it by defining our solver
```
model = Sequential()
model.add(Dense(4,activation='relu'))
model.add(Dense(4,activation='relu'))
model.add(Dense(4,activation='relu'))
# Final output node for prediction
model.add(Dense(1))
model.compile(optimizer='rmsprop',loss='mse')
```
### Choosing an optimizer and loss
Keep in mind what kind of problem you are trying to solve:
# For a multi-class classification problem
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
# For a binary classification problem
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# For a mean squared error regression problem
model.compile(optimizer='rmsprop',
loss='mse')
# Training
Below are some common definitions that are necessary to know and understand to correctly utilize Keras:
* Sample: one element of a dataset.
* Example: one image is a sample in a convolutional network
* Example: one audio file is a sample for a speech recognition model
* Batch: a set of N samples. The samples in a batch are processed independently, in parallel. If training, a batch results in only one update to the model.A batch generally approximates the distribution of the input data better than a single input. The larger the batch, the better the approximation; however, it is also true that the batch will take longer to process and will still result in only one update. For inference (evaluate/predict), it is recommended to pick a batch size that is as large as you can afford without going out of memory (since larger batches will usually result in faster evaluation/prediction).
* Epoch: an arbitrary cutoff, generally defined as "one pass over the entire dataset", used to separate training into distinct phases, which is useful for logging and periodic evaluation.
* When using validation_data or validation_split with the fit method of Keras models, evaluation will be run at the end of every epoch.
* Within Keras, there is the ability to add callbacks specifically designed to be run at the end of an epoch. Examples of these are learning rate changes and model checkpointing (saving).
```
model.fit(X_train,y_train,epochs=250)
```
## Evaluation
Let's evaluate our performance on our training set and our test set. We can compare these two performances to check for overfitting.
```
model.history.history
loss = model.history.history['loss']
sns.lineplot(x=range(len(loss)),y=loss)
plt.title("Training Loss per Epoch");
```
### Compare final evaluation (MSE) on training set and test set.
These should hopefully be fairly close to each other.
```
model.metrics_names
training_score = model.evaluate(X_train,y_train,verbose=0)
test_score = model.evaluate(X_test,y_test,verbose=0)
training_score
test_score
```
### Further Evaluations
```
test_predictions = model.predict(X_test)
test_predictions
pred_df = pd.DataFrame(y_test,columns=['Test Y'])
pred_df
test_predictions = pd.Series(test_predictions.reshape(300,))
test_predictions
pred_df = pd.concat([pred_df,test_predictions],axis=1)
pred_df.columns = ['Test Y','Model Predictions']
pred_df
```
Let's compare to the real test labels!
```
sns.scatterplot(x='Test Y',y='Model Predictions',data=pred_df)
pred_df['Error'] = pred_df['Test Y'] - pred_df['Model Predictions']
sns.distplot(pred_df['Error'],bins=50)
from sklearn.metrics import mean_absolute_error,mean_squared_error
mean_absolute_error(pred_df['Test Y'],pred_df['Model Predictions'])
mean_squared_error(pred_df['Test Y'],pred_df['Model Predictions'])
# Essentially the same thing, difference just due to precision
test_score
#RMSE
test_score**0.5
```
# Predicting on brand new data
What if we just saw a brand new gemstone from the ground? What should we price it at? This is the **exact** same procedure as predicting on a new test data!
```
# [[Feature1, Feature2]]
new_gem = [[998,1000]]
# Don't forget to scale!
scaler.transform(new_gem)
new_gem = scaler.transform(new_gem)
model.predict(new_gem)
```
## Saving and Loading a Model
```
from tensorflow.keras.models import load_model
model.save('my_model.h5') # creates a HDF5 file 'my_model.h5'
later_model = load_model('my_model.h5')
later_model.predict(new_gem)
```
| github_jupyter |
## Notebook for Solar Wind Exploration
In the initial phase, we want to see if we can detect FTEs using unsupervised learning, by finding a manifold for the solar wind data.
The initial hypothesis is the transition matrices (Markov Matrices $M$) that can be derived from Manifolder + clustering will show distinctive clusters and transitions. We can check accuracy by looking at the label (FTE or not?), and see if this label could have been deduced from the data itself.
```
# useful set of python includes
%load_ext autoreload
%autoreload 2
import numpy as np
np.set_printoptions(suppress=True, precision=4)
import matplotlib.pyplot as plt
%config InlineBackend.figure_format = 'svg'
import seaborn as sns
sns.set()
import pandas as pd
import time
import random
```
### Load Solar Wind Data, and Run Manifolder
The `dataset_2` file contains
Dataset-2 (THEMIS): a list with FTEs periods and non-FTEs periods observed by THEMIS in 2007. These are combined into one file, randomly FTE - NonFTE - FTE - FTE, NonFTE, etc…
In total there are 63 FTEs and 47 non-FTEs.
The time series are separated by one blank line, and each one has 1440 points in a period of 6 minutes.
```
import sys
sys.path.append(r"C:\Users\acloninger\GDrive\ac2528Backup\DocsFolder\GitHub\manifolder")
sys.path.append(r"..")
import manifolder as mr
from manifolder import helper as mh
# load the data
# note, you must have started the notebook in the
print('loading data ...')
df = pd.read_excel('astro_data/dataset_2.xlsx', index_col=0)
df.head()
# convert values from loaded spreadsheet, into a numpy matrices
# note that there is no need for the first value, which is time,
# as it is not part of the manifold
#
# also, note the spreadsheet is missing a column name for `Unnamed: 13`, and the values above
# this have the incorrect column labels; the first relevant vale is bx, which as a magnitude around 2
#
# note the final value of each row is the goal (0 or 1), and not part of z
data_raw = df.values[:, 1:]
print('first line of raw_data:\n', data_raw[0, :])
# loop through the data, breaking out the clusters
# i will always point to the NaN (blank line) in the dataframe,
# and values [i-1440:i] is the snipped
snippet_len = 1440
# collect all line breaks (blank lines) in csv file
#lineBreaks = [0]
#for i in range(data_raw.shape[0]):
# if data_raw[i,0] != data_raw[i,0]: # replacement of isnan, since nan != nan
# lineBreaks.append(i)
#lineBreaks.append(data_raw.shape[0])
#
#num_snippet = len(lineBreaks)-1
# callect the snippets into two groups, one for each goal (target) value, 0 or 1
# these can be easily merged
zs_0 = []
zs_1 = []
df.values[0,:]
for i in range(snippet_len,data_raw.shape[0],snippet_len+1):
# copy the snipped, excluding the last value, which is the goal
snippet = data_raw[i-snippet_len:i,:-1]
# grab the goal value from the first row of each snippet
goal = data_raw[i-snippet_len,-1]
# check to make sure each snippet does not contain NaN
# (should not, if parsing is correct)
assert ~np.isnan(snippet).any(), 'oops, snippet contains a Nan!'
print('snippet size',snippet.shape,'with goal',goal)
if goal == 0:
zs_0.append( snippet )
elif goal == 1:
zs_1.append( snippet )
else:
assert False, 'value of goal not understood'
# shuffle this lists; this should not strictly be necessary, if all the data is being used,
# but prevents biases when shortening the list
random.shuffle(zs_0)
random.shuffle(zs_1)
shorten_data = False
if shorten_data:
zs_0 = zs_0[:10]
zs_1 = zs_1[:10]
zs = zs_0 + zs_1
z_breakpoint = len(zs_0)
print( '\done!')
print( '\t len(zs_0):',len(zs_0))
print( '\t len(zs_1):',len(zs_1))
print( '\t len(zs):',len(zs))
import matplotlib.pyplot as plt
plt.figure()
for i in range(9):
plt.subplot(3,3,i+1)
plt.plot(zs_0[i])
plt.show()
plt.figure()
for i in range(9):
plt.subplot(3,3,i+1)
plt.plot(zs_1[i])
plt.show()
# data has been parsed, now run Manifolder
H = 80
step_size = 10
nbins = 10
ncov = 20
start_time = time.time()
# create manifolder object
manifolder = mr.Manifolder(H=H,step_size=step_size,nbins=nbins, ncov=ncov)
# add the data, and fit (this runs all the functions)
manifolder.fit_transform(zs, parallel=True, use_dtw=False)
elapsed_time = time.time() - start_time
print('\n\t Program Executed in', str(np.round(elapsed_time, 2)), 'seconds') # about 215 seconds (four minutes)
start_time = time.time()
manifolder._clustering(numClusters=7, kmns=False, distance_measure=None) # display
print(manifolder.IDX.shape)
elapsed_time = time.time() - start_time
print('\n\t Program Executed for k means clustering in', str(np.round(elapsed_time, 2)), 'seconds')
# clustering data for k-means...
IDX = manifolder.IDX
cluster_lens = mh.count_cluster_lengths(IDX)
# cluster_lens is a dictionary a dictonary, where each key is the cluster number (0:6),
# and the values are a list of cluster lengths
mh.show_cluster_lens(cluster_lens)
# in this case, index goes from 0 to 6 ...
# can also have outlier groups in kmeans, need to check for this
print(IDX.shape)
print(np.min(IDX))
print(np.max(IDX))
IDX_max = np.max(IDX)
M = mh.make_transition_matrix(IDX)
print('\n transition matrix:')
print(M)
M_z0 = mh.make_transition_matrix(IDX[manifolder.snip_number<z_breakpoint])
M_z1 = mh.make_transition_matrix(IDX[manifolder.snip_number>=z_breakpoint])
print('\n z0 transition matrix:')
print(M_z0)
print('\n z1 transition matrix:')
print(M_z1)
z_downsample = np.empty((0,zs[0].shape[1]+1), float)
for i in range(len(zs)):
x = zs[i]
x = x[0:x.shape[0]-H,:]
x = x[::step_size]
if i<z_breakpoint:
x = np.append(x,np.zeros((x.shape[0],1)),1)
else:
x = np.append(x,np.ones((x.shape[0],1)),1)
z_downsample = np.append(z_downsample,x,0)
z_downsample = np.append(z_downsample, manifolder.snip_number.reshape(len(IDX),1), 1)
z_downsample = np.append(z_downsample, IDX.reshape(len(IDX),1), 1)
z_downsample.shape
np.savetxt('astro_subset2_clustering_k=10.csv', z_downsample, delimiter=',', fmt='%f')
```
### TODO:
After running `fit_transform()`, use kmeans to label clusters withing all the snippets
Create a transition matrix for each snippet; the zs_0 and zs_1 should have distincively different matrices, which can be used to categorize the snippet
```
### _cluster() function, local to make it easier to work on
### ... note, all the individual clusters should be marked invidually?
### but the original kmeans run run all of them together?
###
# Configuration
numClusters = 7 # NOTE, this was previously 14 (too many!)
intrinsicDim = manifolde.Dim # can be varied slightly but shouldn't be much larger than Dim
## Clusters
# IDX = kmeans(Psi(:, 1:intrinsicDim), numClusters)
# Python kmeans see
# https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.cluster.vq.kmeans.html
# scipy.cluster.vq.kmeans(obs, k_or_guess, iter=20, thresh=1e-05)
#
# note, python expects each ROW to be an observation, looks the same a matlap
#
print('running k-means')
kmeans = KMeans(n_clusters=numClusters).fit(manifolder.Psi[:, :intrinsicDim])
IDX = kmeans.labels_
# TODO decide how to plot multiple snips
# think that x_ref[1,:] is just
for snip in range(len(self.z)):
if snip == 0:
x = self.z[snip][0, :]
xref1 = x[::self.stepSize] # downsample, to match the data steps
else:
x = self.z[snip][0, :]
x = x[::self.stepSize]
xref1 = np.append(xref1, x)
print(xref1.shape)
xs = manifolder.Psi[:, 0]
ys = manifolder.Psi[:, 1]
zs = manifolder.Psi[:, 2]
# normalize these to amplitude one?
print('normalizing amplitudes of Psi in Python ...')
xs /= np.max(np.abs(xs))
ys /= np.max(np.abs(ys))
zs /= np.max(np.abs(zs))
# xs -= np.mean(xs)
# ys -= np.mean(ys)
# zs -= np.mean(zs)
# xs /= np.std(xs)
# ys /= np.std(ys)
# zs /= np.std(zs)
print(xs.shape)
lim = 2000
val = xref1[:lim]
idx = manifolder.IDX[:lim]
plt.figure(figsize=[15, 3])
plt.plot(xref1[:lim], color='black', label='Timeseries')
# plt.plot(xs[:lim], linewidth=.5, label='$\psi_0$')
# plt.plot(ys[:lim], linewidth=.5, label='$\psi_1$')
# plt.plot(zs[:lim], linewidth=.5, label='$\psi_2$')
plt.plot(xs[:lim], linewidth=.5, label='psi_0')
plt.plot(ys[:lim], linewidth=.5, label='psi_1')
plt.plot(zs[:lim], linewidth=.5, label='psi_2')
plt.plot(idx / np.max(idx) + 1, linewidth=.8, label='IDX')
plt.legend()
# rightarrow causes an image error, when displayed in github!
# plt.xlabel('Time $ \\rightarrow $')
plt.xlabel('Time')
plt.ylabel('Value')
# plt.gca().autoscale(enable=True, axis='both', tight=None )
# plt.gca().xaxis.set_ticklabels([])
# plt.gca().yaxis.set_ticklabels([])
plt.title('Example Timeseries and Manifold Projection')
print('done')
###
### additional parsing, for color graphs
###
import matplotlib
cmap = matplotlib.cm.get_cmap('Spectral')
r = xs[:lim]
g = ys[:lim]
b = zs[:lim]
# prevent the jump in data value
r[:self.H] = r[self.H]
g[:self.H] = g[self.H]
b[:self.H] = b[self.H]
r -= np.min(r)
r /= np.max(r)
g -= np.min(g)
g /= np.max(g)
b -= np.min(b)
b /= np.max(b)
plt.figure(figsize=[15, 3])
for i in range(lim - 1):
col = [r[i], g[i], b[i]]
plt.plot([i, i + 1], [val[i], val[i + 1]], color=col)
plt.title('data, colored according to Psi (color three-vector)')
plt.xlabel('Time')
plt.ylabel('Value')
plt.show()
# clustering data ...
IDX = manifolder.IDX
cluster_lens = mh.count_cluster_lengths(IDX)
# cluster_lens is a dictionary a dictonary, where each key is the cluster number (0:6),
# and the values are a list of clusner lengths
mh.show_cluster_lens(cluster_lens)
```
### Graph Transition (Markov) Matrix
The system can be though of as in one particular "state" (cluster value) at any given time. This state $S$ can be though of as a column vector with $C$ dimensions, similar to states in quantum mechanic, where the column vector plays the role of the transition matrix.
Time evolution is this given by the tranistion matrix $M$, which is a Markov matrix (all columns sum to one, to preserve probability). In this case, we have
$$
S_{n+1} = M @ S_n
$$
Where the $@$ symbol is used to explicitly denote matrix multiplication.
Since most clusters with transition to themselves, the diagonal values of the matrix can be quite high, and are typically removed. Thus, for visualization, we remove the diagonal elements of the matrix.
```
# in this case, index goes from 0 to 6 ...
# can also have outlier groups in kmeans, need to check for this
print(IDX.shape)
print(np.min(IDX))
print(np.max(IDX))
IDX_max = np.max(IDX)
M = mh.make_transition_matrix(IDX)
print('\n transition matrix:')
print(M)
# reorder transition matrix, from most to least common cluster
# diagonal elements monotonically decreasing
IDX_ordered = mh.reorder_cluster(IDX, M)
M = mh.make_transition_matrix(IDX_ordered)
print('\n transition matrix, ordered:')
print(M)
mh.image_M(M)
# remove diagonal, and make markov, for display
print('transition matrix, diagonal elements removed, normalized (Markov)')
np.fill_diagonal(M, 0) # happens inplace
M = mh.make_matrix_markov(M)
print(M)
mh.image_M(M, 1)
```
| github_jupyter |
```
import numpy as np
import gzip
filename=[
["training_images","train-images-idx3-ubyte.gz"],
["test_images","t10k-images-idx3-ubyte.gz"],
["training_labels","train-labels-idx1-ubyte.gz"],
["test_labels","t10k-labels-idx1-ubyte.gz"]
]
def read_mnist():
mnist={}
with gzip.open('D:/Data/mnist/train-images-idx3-ubyte.gz','rb') as f: # 打开压缩文件
mnist['training_images']=np.frombuffer(f.read(),
np.uint8,
count=-1, # 读入全部数据
offset=16).reshape(-1,28*28)
with gzip.open('D:/Data/mnist/t10k-images-idx3-ubyte.gz','rb') as f:
mnist['test_images']=np.frombuffer(f.read(),
np.uint8,
count=-1,
offset=16).reshape(-1,28*28)
with gzip.open('D:/Data/mnist/train-labels-idx1-ubyte.gz','rb') as f:
mnist['training_labels']=np.frombuffer(f.read(),
np.uint8,
count=-1,
offset=8)
with gzip.open('D:/Data/mnist/t10k-labels-idx1-ubyte.gz','rb') as f:
mnist['test_labels']=np.frombuffer(f.read(),
np.uint8,
count=-1,
offset=8)
return mnist
mnist=read_mnist()
X_train=mnist['training_images']
y_train=mnist['training_labels']
X_test=mnist['test_images']
y_test=mnist['test_labels']
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
X_train=np.reshape(X_train,(-1,28,28,1))
X_test=np.reshape(X_test,(-1,28,28,1))
print(X_train.shape)
print(X_test.shape)
X_train=X_train/255.
X_test=X_test/255.
from tensorflow.keras.utils import *
print(y_train.shape)
print(y_test.shape)
y_train=to_categorical(y_train)
y_test=to_categorical(y_test)
print(y_train.shape)
print(y_test.shape)
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense,Dropout,Flatten,Conv2D,MaxPool2D
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.backend import categorical_crossentropy
model = Sequential()
model.add(Conv2D(filters=32,
kernel_size=(5,5),
strides=(1,1),
padding='same',
activation='relu',
input_shape=(28,28,1)))
model.add(Conv2D(filters=32,
kernel_size=(5,5),
strides=(1,1),
padding='same',
activation='relu'))
model.add(MaxPool2D(pool_size=(2,2),
strides=(2,2)))
model.add(Dropout(rate=0.25))
model.add(Conv2D(filters=64,
kernel_size=(3,3),
strides=(1,1),
padding='same',
activation='relu'))
model.add(Conv2D(filters=64,
kernel_size=(3,3),
strides=(1,1),
padding='same',
activation='relu'))
model.add(MaxPool2D(pool_size=(2,2),
strides=(2,2)))
model.add(Dropout(rate=0.25))
model.add(Flatten())
model.add(Dense(units=256,activation='relu'))
model.add(Dropout(rate=0.25))
model.add(Dense(10,activation='softmax'))
model.summary()
opt=SGD(lr=0.01,decay=1e-6,momentum=0.9,nesterov=True)
model.compile(optimizer=opt,loss=categorical_crossentropy,metrics=['accuracy'])
model.fit(x=X_train,y=y_train,batch_size=32,epochs=5,verbose=1)
test_loss,test_acc=model.evaluate(x=X_test,y=y_test,verbose=1)
print(test_loss)
print(test_acc)
```
| github_jupyter |
## Introduction to Exploratory Data Analysis and Visualization
In this lab, we will cover some basic EDAV tools and provide an example using _presidential speeches_.
## Table of Contents
[ -Step 0: Import modules](#step0)
[-Step 1: Read in the speeches](#step1)
[-Step 2: Text processing](#step2)
-Step 3: Visualization
* [ Step 3.1: Word cloud](#step3-1)
* [ Step 3.2: Joy plot](#step3-3)
[-Step 4: Sentence analysis](#step4)
[-Step 5: NRC emotion analysis](#step5)
<a id="Example"></a>
## Part 2: Example using _presidential speeches_.
In this section, we will go over an example using a collection of presidential speeches. The data were scraped from the [Presidential Documents Archive](http://www.presidency.ucsb.edu/index_docs.php) of the [American Presidency Project](http://www.presidency.ucsb.edu/index.php) using the `Rvest` package from `R`. The scraped text files can be found in the `data` folder.
For the lab, we use a handful of basic Natural language processing (NLP) building blocks provided by NLTK (and a few additional libraries), including text processing (tokenization, stemming etc.), frequency analysis, and NRC emotion analysis. It also provides various data visualizations -- an important field of data science.
<a id="step0"></a>
## Step 0: Import modules
**Initial Setup** you need Python installed on your system to run the code examples used in this tutorial. This tutorial is constructed using Python 2.7, which is slightly different from Python 3.5.
We recommend that you use Anaconda for your python installation. For more installation recommendations, please use our [check_env.ipynb](https://github.com/DS-BootCamp-Collaboratory-Columbia/AY2017-2018-Winter/blob/master/Bootcamp-materials/notebooks/Pre-assignment/check_env.ipynb).
The main modules we will use in this notebook are:
* *nltk*:
* *nltk* (Natural Language ToolKit) is the most popular Python framework for working with human language.
* *nltk* doesn’t come with super powerful pre-trained models, but contains useful functions for doing a quick exploratory data analysis.
* [Reference webpage](https://nlpforhackers.io/introduction-nltk/#more-4627)
* [NLTK book](http://www.nltk.org/book/)
* *re* and *string*:
* For text processing.
* *scikit-learn*:
* For text feature extraction.
* *wordcloud*:
* Word cloud visualization.
* Pip installation:
```
pip install wordcloud
```
* Conda installation (not come with Anaconda built-in packages):
```
conda install -c conda-forge wordcloud=1.2.1
```
* *ipywidgets*:
* *ipywidgets* can render interactive controls on the Jupyter notebook. By using the elements in *ipywidgets*, e.g., `IntSlider`, `Checkbox`, `Dropdown`, we could produce fun interactive visualizations.
* Pip installation:
If you use pip, you also have to enable the ipywidget extension in your notebook to render it next time you start the notebook. Type in following command on your terminal:
```
pip install ipywidgets
jupyter nbextension enable --py widgetsnbextension
```
* Conda installation:
If you use conda, the extension will be enabled automatically. There might be version imcompatible issue happened, following command is to install the modules in the specific compatible versions.
```
conda install --yes jupyter_core notebook nbconvert ipykernel ipywidgets=6.0 widgetsnbextension=2.0
```
* [Reference webpage](https://towardsdatascience.com/a-very-simple-demo-of-interactive-controls-on-jupyter-notebook-4429cf46aabd)
* *seaborn*:
* *seaborn* provides a high-level interface to draw statistical graphics.
* A comprehensive [tutorial](https://www.datacamp.com/community/tutorials/seaborn-python-tutorial) on it.
```
# Basic
from random import randint
import pandas as pd
import csv
import numpy as np
from collections import OrderedDict, defaultdict, Counter
# Text
import nltk, re, string
from nltk.corpus.reader.plaintext import PlaintextCorpusReader #Read in text files
from nltk import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
nltk.download('punkt')
# Plot
import ipywidgets as widgets
import seaborn as sns
from ipywidgets import interactive, Layout, HBox, VBox
from wordcloud import WordCloud
from matplotlib import pyplot as plt
from matplotlib import gridspec, cm
# Source code
import sys
sys.path.append("../lib/")
import joypy
```
<a id="step1"></a>
## Step 1: Read in the speeches
```
inaug_corpus = PlaintextCorpusReader("../data/inaugurals", ".*\.txt")
#Accessing the name of the files of the corpus
inaug_files = inaug_corpus.fileids()
for f in inaug_files[:5]:
print(f)
len(inaug_files)
#Accessing all the text of the corpus
inaug_all_text = inaug_corpus.raw()
print("First 100 words in all the text of the corpus: \n >>" + inaug_all_text[:100])
#Accessing all the text for one of the files
inaug_ZacharyTaylor1_text=inaug_corpus.raw('inaugZacharyTaylor-1.txt')
print("First 100 words in one file: \n >>" + inaug_ZacharyTaylor1_text[:100])
```
<a id="step2"></a>
## Step 2: Text processing
For the speeches, we do the text processing as follows and define a function `tokenize_and_stem`:
1. convert all letters to lower cases
2. split text into sentences and then words
3. remove [stop words](https://github.com/arc12/Text-Mining-Weak-Signals/wiki/Standard-set-of-english-stopwords), remove empty words due to formatting errors, and remove punctuation.
4. [stemming words](https://en.wikipedia.org/wiki/Stemming) use NLTK porter stemmer. There are [many other stemmers](http://www.nltk.org/howto/stem.html) built in NLTK. You can play around and see the difference.
Then we compute the [Document-Term Matrix (DTM)](https://en.wikipedia.org/wiki/Document-term_matrix) and [TF-IDF](https://en.wikipedia.org/wiki/Tf%E2%80%93idf).
See [Natural Language Processing with Python](http://www.nltk.org/book/) for a more comprehensive discussion about NLTK.
There are many more interesting topics in NLP, which we will not cover in this lab. In you are interested, here are some online resources.
1. [Named Entity Recognition](https://github.com/charlieg/A-Smattering-of-NLP-in-Python)
2. [Topic modeling](https://medium.com/mlreview/topic-modeling-with-scikit-learn-e80d33668730)
3. [sentiment analysis](https://pythonspot.com/python-sentiment-analysis/) (positive v.s. negative)
3. [Supervised model](https://www.dataquest.io/blog/natural-language-processing-with-python/)
```
stemmer = PorterStemmer()
def tokenize_and_stem(text):
lowers = text.lower()
tokens = [word for sent in nltk.sent_tokenize(lowers) for word in nltk.word_tokenize(sent)]
filtered_tokens = []
# filter out any tokens not containing letters (e.g., numeric tokens, raw punctuation)
for token in tokens:
if re.search('[a-zA-Z]', token) and not token in stopwords.words('english'):
filtered_tokens.append(re.sub(r'[^\w\s]','',token))
stems = [stemmer.stem(t) for t in filtered_tokens]
return stems
#return filtered_tokens
token_dict = {}
for fileid in inaug_corpus.fileids():
token_dict[fileid] = inaug_corpus.raw(fileid)
# Construct a bag of words matrix.
# This will lowercase everything, and ignore all punctuation by default.
# It will also remove stop words.
vectorizer = CountVectorizer(lowercase=True,
tokenizer=tokenize_and_stem,
stop_words='english')
dtm = vectorizer.fit_transform(token_dict.values()).toarray()
```
**TF - IDF**
TF-IDF (term frequency-inverse document frequency) is a numerical statistics that is intended to reflect how important a word is to a document in a collection or corpus. It is often used as a weighting factor in information retrieval, text mining, and user modeling. The TF-IDF value increases proportionally to the number of times a word appears in the document, but is offset by the frequency of the word in the corpus, which helps to adjust for the fact that some words appear more frequently in general.
$$
\begin{aligned}
\mbox{TF}(t) &=\frac{\mbox{Number of times term $t$ appears in a document}}{\mbox{Total number of terms in the document}}\\
\mbox{IDF}(t) &=\log{\frac{\mbox{Total number of documents}}{\mbox{Number of documents with term $t$ in it}}}\\
\mbox{TF-IDF}(t) &=\mbox{TF}(t)\times\mbox{IDF}(t)
\end{aligned}
$$
```
vectorizer = TfidfVectorizer(tokenizer=tokenize_and_stem,
stop_words='english',
decode_error='ignore')
tfidf_matrix = vectorizer.fit_transform(token_dict.values())
# The above line can take some time (about < 60 seconds)
feature_names = vectorizer.get_feature_names()
num_samples, num_features=tfidf_matrix.shape
print("num_samples: %d, num_features: %d" %(num_samples,num_features))
num_clusters=10
## Checking
print('first term: ' + feature_names[0])
print('last term: ' + feature_names[len(feature_names) - 1])
for i in range(0, 4):
print('random term: ' +
feature_names[randint(1,len(feature_names) - 2)] )
def top_tfidf_feats(row, features, top_n=20):
topn_ids = np.argsort(row)[::-1][:top_n]
top_feats = [(features[i], row[i]) for i in topn_ids]
df = pd.DataFrame(top_feats, columns=['features', 'score'])
return df
def top_feats_in_doc(X, features, row_id, top_n=25):
row = np.squeeze(X[row_id].toarray())
return top_tfidf_feats(row, features, top_n)
print(inaug_files[2:3])
print(top_feats_in_doc(tfidf_matrix, feature_names, 3, 10))
d =3
top_tfidf = top_feats_in_doc(tfidf_matrix, feature_names, d, 10)
def plot_tfidf_classfeats_h(df, doc):
''' Plot the data frames returned by the function tfidf_feats_in_doc. '''
x = np.arange(len(df))
fig = plt.figure(figsize=(6, 9), facecolor="w")
ax = fig.add_subplot(1, 1, 1)
ax.spines["top"].set_visible(False)
ax.spines["right"].set_visible(False)
ax.set_frame_on(False)
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
ax.set_xlabel("Tf-Idf Score", labelpad=16, fontsize=14)
ax.set_title(doc, fontsize=16)
ax.ticklabel_format(axis='x', style='sci', scilimits=(-2,2))
ax.barh(x, df.score, align='center', color='#3F5D7D')
ax.set_yticks(x)
ax.set_ylim([-1, x[-1]+1])
yticks = ax.set_yticklabels(df.features)
plt.subplots_adjust(bottom=0.09, right=0.97, left=0.15, top=0.95, wspace=0.52)
plt.show()
plot_tfidf_classfeats_h(top_tfidf, inaug_files[(d-1):d])
```
<a id="step3-1"></a>
## Step 3: Visualization
Data visualization is an integral part of the data science workflow. In the following, we use simple data visualizations to reveal some interesting patterns in our data.
### 1 . Word cloud
```
array_for_word_cloud = []
word_count_array = dtm.sum(0)
for idx, word in enumerate(feature_names):
array_for_word_cloud.append((word,word_count_array[idx]))
def random_color_func(word=None, font_size=None,
position=None, orientation=None, font_path=None, random_state=None):
h = int(360.0 * 45.0 / 255.0)
s = int(100.0 * 255.0 / 255.0)
l = int(100.0 * float(random_state.randint(60, 120)) / 255.0)
return "hsl({}, {}%, {}%)".format(h, s, l)
wordcloud = WordCloud(background_color='white',
width=1600,
height=1000,
color_func=random_color_func).generate_from_frequencies(array_for_word_cloud)
%matplotlib inline
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
```
Let us try making it interactive.
```
word_cloud_dict = {}
counter = 0
for fileid in inaug_corpus.fileids():
row = dtm[counter,:]
word_cloud_dict[fileid] = []
for idx, word in enumerate(feature_names):
word_cloud_dict[fileid].append((word,row[idx]))
counter += 1
def f_wordclouds(t):
wordcloud = WordCloud(background_color='white',
color_func=random_color_func).generate_from_frequencies(word_cloud_dict[t])
plt.figure(figsize=(20, 16), dpi=100)
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
interactive_plot_1 = interactive(f_wordclouds, t=widgets.Dropdown(options=inaug_corpus.fileids(),description='text1'))
interactive_plot_2 = interactive(f_wordclouds, t=widgets.Dropdown(options=inaug_corpus.fileids(),description='text2'))
# Define the layout here.
hbox_layout = Layout(display='flex', flex_flow='row', justify_content='space-between', align_items='center')
vbox_layout = Layout(display='flex', flex_flow='column', justify_content='space-between', align_items='center')
%matplotlib inline
display(HBox([interactive_plot_1,interactive_plot_2],layout=hbox_layout))
```
<a id="step3-3"></a>
### 2. Joy plot
The following joy plot allows us to compare the frequencies of the top 10 most frequent words in individual speeches.
```
joy_df = pd.DataFrame(dtm, columns=feature_names)
selected_words = joy_df.sum(0).sort_values(ascending=False).head(10).index
print(selected_words)
%matplotlib inline
plt.rcParams['axes.facecolor'] = 'white'
fig, axes = joypy.joyplot(joy_df.loc[:,selected_words],
range_style='own', grid="y",
colormap=cm.YlGn_r,
title="Top 10 word distribution")
```
<a id="step4"></a>
## Step 4: Sentence analysis
In the previous sections, we focused on word-level distributions in inaugural speeches. Next, we will use sentences as our units of analysis, since they are natural languge units for organizing thoughts and ideas.
For simpler visualization, we chose a subset of better known presidents or presidential candidates on whom to focus our analysis.
```
filter_comparison=["DonaldJTrump","JohnMcCain", "GeorgeBush", "MittRomney", "GeorgeWBush",
"RonaldReagan","AlbertGore,Jr", "HillaryClinton","JohnFKerry",
"WilliamJClinton","HarrySTruman", "BarackObama", "LyndonBJohnson",
"GeraldRFord", "JimmyCarter", "DwightDEisenhower", "FranklinDRoosevelt",
"HerbertHoover","JohnFKennedy","RichardNixon","WoodrowWilson",
"AbrahamLincoln", "TheodoreRoosevelt", "JamesGarfield",
"JohnQuincyAdams", "UlyssesSGrant", "ThomasJefferson",
"GeorgeWashington", "WilliamHowardTaft", "AndrewJackson",
"WilliamHenryHarrison", "JohnAdams"]
```
### Nomination speeches
Next, we first look at the *nomination acceptance speeches* at major parties' national conventions.
Following the same procedure in [step 1](#step1), we will use `pandas` dataframe to store the nomination speech sentences. For each sentence in a speech (`fileid`), we find out the name of the president (`president`) and the term (`term`), and also calculated the number of words in each sentence as *sentence length* (`word_count`) by using a self-defined function `word_count`.
```
def word_count(string):
tokens = [word for word in nltk.word_tokenize(string)]
counter = 0
for token in tokens:
if re.search('[a-zA-Z]', token):
counter += 1
return counter
nomin_corpus = PlaintextCorpusReader("../data/nomimations", ".*\.txt")
nomin_files = nomin_corpus.fileids()
nomin_file_df = pd.DataFrame(columns=["file_id","president","term","raw_text"])
for fileid in nomin_corpus.fileids():
nomin_file_df = nomin_file_df.append({"file_id": fileid,
"president": fileid[0:fileid.find("-")][5:],
"term": fileid.split("-")[-1][0],
"raw_text": nomin_corpus.raw(fileid)}, ignore_index=True)
sentences = []
for row in nomin_file_df.itertuples():
for sentence in sent_tokenize(row[4]):
sentences.append({"file_id": row[1],
"president": row[2],
"term": row[3],
"sentence": sentence})
nomin_sen_df = pd.DataFrame(sentences, columns=["file_id","president","term","sentence"])
nomin_sen_df["word_count"] = [word_count(sentence) for sentence in nomin_sen_df["sentence"]]
```
#### First term
For comparison between presidents, we first limit our attention to speeches for the first terms of former U.S. presidents. We noticed that a number of presidents have very short sentences in their nomination acceptance speeches.
```
filtered_nomin_sen_df = nomin_sen_df.loc[(nomin_sen_df["president"].isin(filter_comparison))&(nomin_sen_df["term"]=='1')]
filtered_nomin_sen_df = filtered_nomin_sen_df.reset_index()
filtered_nomin_sen_df['group_mean'] = filtered_nomin_sen_df.groupby('president')['word_count'].transform('mean')
filtered_nomin_sen_df = filtered_nomin_sen_df.sort_values('group_mean', ascending=False)
%matplotlib inline
plt.figure(figsize=(20, 10))
gs = gridspec.GridSpec(1, 2, width_ratios=[1, 1])
plt.subplot(gs[0])
sns.set(font_scale=1.5)
sns.set_style('whitegrid')
sns.swarmplot(y='president', x='word_count',
data=filtered_nomin_sen_df,
palette = "Set3",
size=2.5,
orient='h'). set(xlabel='Number of words in a sentence')
plt.title("Swarm plot")
plt.subplot(gs[1])
sns.set(font_scale=1.5)
sns.set_style('whitegrid')
sns.violinplot(y='president', x='word_count',
data=filtered_nomin_sen_df,
palette = "Set3", cut = 3,
width=1.5, saturation=0.8, linewidth= 0.3, scale="count",
orient='h').set(xlabel='Number of words in a sentence')
plt.title("Violin plot")
plt.xlim(-3, 350)
plt.tight_layout()
```
#### Second term
```
filtered_nomin_sen_df = nomin_sen_df.loc[(nomin_sen_df["president"].isin(filter_comparison))&(nomin_sen_df["term"]=='2')]
filtered_nomin_sen_df = filtered_nomin_sen_df.reset_index()
filtered_nomin_sen_df['group_mean'] = filtered_nomin_sen_df.groupby('president')['word_count'].transform('mean')
filtered_nomin_sen_df = filtered_nomin_sen_df.sort_values('group_mean', ascending=False)
%matplotlib inline
plt.figure(figsize=(20, 10))
gs = gridspec.GridSpec(1, 2, width_ratios=[1, 1])
plt.subplot(gs[0])
sns.set(font_scale=1.5)
sns.set_style('whitegrid')
sns.swarmplot(y='president', x='word_count',
data=filtered_nomin_sen_df,
palette = "Set3",
size=2.5,
orient='h'). set(xlabel='Number of words in a sentence')
plt.title("Swarm plot")
plt.subplot(gs[1])
sns.violinplot(y='president', x='word_count',
data=filtered_nomin_sen_df,
palette = "Set3", cut = 3,
width=1.5, saturation=0.8, linewidth= 0.3, scale="count",
orient='h').set(xlabel='Number of words in a sentence')
plt.title("Violin plot")
plt.xlim(-3, 350)
plt.tight_layout()
```
### Inaugural speeches
We notice that the sentences in inaugural speeches are longer than those in nomination acceptance speeches.
```
inaug_file_df = pd.DataFrame(columns=["file_id","president","term","raw_text"])
for fileid in inaug_corpus.fileids():
inaug_file_df = inaug_file_df.append({"file_id": fileid,
"president": fileid[0:fileid.find("-")][5:],
"term": fileid.split("-")[-1][0],
"raw_text": inaug_corpus.raw(fileid)}, ignore_index=True)
sentences = []
for row in inaug_file_df.itertuples():
for sentence in sent_tokenize(row[4]):
sentences.append({"file_id": row[1],
"president": row[2],
"term": row[3],
"sentence": sentence})
inaug_sen_df = pd.DataFrame(sentences, columns=["file_id","president","term","sentence"])
wordCounts = [word_count(sentence) for sentence in inaug_sen_df["sentence"]]
inaug_sen_df["word_count"] = wordCounts
filtered_inaug_sen_df = inaug_sen_df.loc[(inaug_sen_df["president"].isin(filter_comparison))&(inaug_sen_df["term"]=='1')]
filtered_inaug_sen_df = filtered_inaug_sen_df.reset_index()
filtered_inaug_sen_df['group_mean'] = filtered_inaug_sen_df.groupby('president')['word_count'].transform('mean')
filtered_inaug_sen_df = filtered_inaug_sen_df.sort_values('group_mean', ascending=False)
%matplotlib inline
plt.figure(figsize=(20, 10))
gs = gridspec.GridSpec(1, 2, width_ratios=[1, 1])
plt.subplot(gs[0])
sns.set(font_scale=1.5)
sns.set_style('whitegrid')
sns.swarmplot(y='president', x='word_count',
data=filtered_inaug_sen_df,
palette = "Set3",
size=2.5,
orient='h'). set(xlabel='Number of words in a sentence')
plt.title("Swarm plot")
plt.subplot(gs[1])
sns.set(font_scale=1.5)
sns.set_style('whitegrid')
sns.violinplot(y='president', x='word_count',
data=filtered_inaug_sen_df,
palette = "Set3", cut = 3,
width=1.5, saturation=0.8, linewidth= 0.3, scale="count",
orient='h').set(xlabel='Number of words in a sentence')
plt.title("Violin plot")
plt.xlim(-3, 350)
plt.tight_layout()
```
<a id="step5"></a>
## Step 5: NRC emotion analsis
For each extracted sentence, we apply sentiment analysis using [NRC sentiment lexion](http://saifmohammad.com/WebPages/NRC-Emotion-Lexicon.htm). "The NRC Emotion Lexicon is a list of English words and their associations with eight basic emotions (anger, fear, anticipation, trust, surprise, sadness, joy, and disgust) and two sentiments (negative and positive). The annotations were manually done by crowdsourcing."
```
wordList = defaultdict(list)
emotionList = defaultdict(list)
with open('../data/NRC-emotion-lexicon-wordlevel-alphabetized-v0.92.txt', 'r') as f:
reader = csv.reader(f, delimiter='\t')
headerRows = [i for i in range(0, 46)]
for row in headerRows:
next(reader)
for word, emotion, present in reader:
if int(present) == 1:
#print(word)
wordList[word].append(emotion)
emotionList[emotion].append(word)
from __future__ import division # for Python 2.7 only
def generate_emotion_count(string):
emoCount = Counter()
tokens = [word for word in nltk.word_tokenize(string)]
counter = 0
for token in tokens:
token = token.lower()
if re.search('[a-zA-Z]', token):
counter += 1
emoCount += Counter(wordList[token])
for emo in emoCount:
emoCount[emo]/=counter
return emoCount
emotionCounts = [generate_emotion_count(sentence) for sentence in nomin_sen_df["sentence"]]
nomin_sen_df_with_emotion = pd.concat([nomin_sen_df, pd.DataFrame(emotionCounts).fillna(0)], axis=1)
emotionCounts = [generate_emotion_count(sentence) for sentence in inaug_sen_df["sentence"]]
inaug_sen_df_with_emotion = pd.concat([inaug_sen_df, pd.DataFrame(emotionCounts).fillna(0)], axis=1)
inaug_sen_df_with_emotion.sample(n=3)
```
### Sentence length variation over the course of the speech, with emotions.
How our presidents (or candidates) alternate between long and short sentences and how they shift between different sentiments in their speeches. It is interesting to note that some presidential candidates' speech are more colorful than others. Here we used the same color theme as in the movie "Inside Out."
```
def make_rgb_transparent(color_name, bg_color_name, alpha):
from matplotlib import colors
rgb = colors.colorConverter.to_rgb(color_name)
bg_rgb = colors.colorConverter.to_rgb(bg_color_name)
return [alpha * c1 + (1 - alpha) * c2
for (c1, c2) in zip(rgb, bg_rgb)]
def f_plotsent_len(InDf, InTerm, InPresident):
import numpy as np
import pylab as pl
from matplotlib import colors
from math import sqrt
from matplotlib import collections as mc
col_use={"zero":"lightgray",
"anger":"#ee0000",
"anticipation":"#ffb90f",
"disgust":"#66cd00",
"fear":"blueviolet",
"joy":"#eead0e",
"sadness":"#1874cd",
"surprise":"#ffb90f",
"trust":"#ffb90f",
"negative":"black",
"positive":"#eead0e"}
InDf["top_emotion"] = InDf.loc[:,'anger':'trust'].idxmax(axis=1)
InDf["top_emotion_value"] = InDf.loc[:,'anger':'trust'].max(axis=1)
InDf.loc[InDf["top_emotion_value"] < 0.05, "top_emotion"] = "zero"
InDf.loc[InDf["top_emotion_value"] < 0.05, "top_emotion_value"] = 1
tempDf = InDf.loc[(InDf["president"]==InPresident)&(InDf["term"]==InTerm)]
pt_col_use = []
lines = []
for i in tempDf.index:
pt_col_use.append(make_rgb_transparent(col_use[tempDf.at[i,"top_emotion"]],
"white",
sqrt(sqrt(tempDf.at[i,"top_emotion_value"]))))
lines.append([(i,0),(i,tempDf.at[i,"word_count"])])
%matplotlib inline
lc = mc.LineCollection(lines, colors=pt_col_use, linewidths=min(5,300/len(tempDf.index)))
fig, ax = pl.subplots() #figsize=(15, 6)
ax.add_collection(lc)
ax.autoscale()
ax.axis('off')
plt.title(InPresident, fontsize=30)
plt.tight_layout()
plt.show()
f_plotsent_len(nomin_sen_df_with_emotion, '1', 'HillaryClinton')
f_plotsent_len(nomin_sen_df_with_emotion, '1', 'DonaldJTrump')
f_plotsent_len(nomin_sen_df_with_emotion, '1', 'BarackObama')
f_plotsent_len(nomin_sen_df_with_emotion, '1', 'GeorgeWBush')
```
### Clustering of emotions
```
sns.set(font_scale=1.3)
sns.clustermap(inaug_sen_df_with_emotion.loc[:,'anger':'trust'].corr(),
figsize=(6,7))
```
| github_jupyter |
## SST-2 : Stanford Sentiment Treebank
The Stanford Sentiment Treebank (SST-2) task is a single sentence classification task. It consists of semtences drawn from movie reviews and annotated for their sentiment.
See [website](https://nlp.stanford.edu/sentiment/code.html) and [paper](https://nlp.stanford.edu/~socherr/EMNLP2013_RNTN.pdf) for more info.
```
import numpy as np
import pandas as pd
import os
import sys
import csv
from sklearn import metrics
from sklearn.metrics import classification_report
sys.path.append("../")
from bert_sklearn import BertClassifier
from bert_sklearn import load_model
DATADIR = os.getcwd() + '/glue_data'
%%bash
python3 download_glue_data.py --data_dir glue_data --tasks SST
"""
SST-2 train data size: 67349
SST-2 dev data size: 872
"""
def get_sst_data(train_file = DATADIR + '/SST-2/train.tsv',
dev_file = DATADIR + '/SST-2/dev.tsv'):
train = pd.read_csv(train_file, sep='\t', encoding = 'utf8',keep_default_na=False)
train.columns=['text','label']
print("SST-2 train data size: %d "%(len(train)))
dev = pd.read_csv(dev_file, sep='\t', encoding = 'utf8',keep_default_na=False)
dev.columns=['text','label']
print("SST-2 dev data size: %d "%(len(dev)))
label_list = np.unique(train['label'])
return train,dev,label_list
train,dev,label_list = get_sst_data()
print(label_list)
train.head()
X_train = train['text']
y_train = train['label']
# define model
model = BertClassifier()
model.epochs = 3
model.validation_fraction = 0.05
model.learning_rate = 2e-5
model.max_seq_length = 128
print('\n',model,'\n')
# fit model
model.fit(X_train, y_train)
# test model on dev
test = dev
X_test = test['text']
y_test = test['label']
# make predictions
y_pred = model.predict(X_test)
print("Accuracy: %0.2f%%"%(metrics.accuracy_score(y_pred,y_test) * 100))
print(classification_report(y_test, y_pred, target_names=['negative','positive']))
```
## with MLP...
```
%%time
X_train = train['text']
y_train = train['label']
# define model
model = BertClassifier()
model.epochs = 3
model.validation_fraction = 0.05
model.learning_rate = 2e-5
model.max_seq_length = 128
model.num_mlp_layers = 4
print('\n',model,'\n')
# fit model
model.fit(X_train, y_train)
# test model on dev
test = dev
X_test = test['text']
y_test = test['label']
# make predictions
y_pred = model.predict(X_test)
print("Accuracy: %0.2f%%"%(metrics.accuracy_score(y_pred,y_test) * 100))
print(classification_report(y_test, y_pred, target_names=['negative','positive']))
```
| github_jupyter |
- run parametric tsne
- compute metrics (training and test data)
- silhouette
- knn
- trustworthiness
```
# reload packages
%load_ext autoreload
%autoreload 2
```
### Choose GPU (this may not be needed on your computer)
```
%env CUDA_DEVICE_ORDER=PCI_BUS_ID
%env CUDA_VISIBLE_DEVICES=0
```
### load packages
```
from tfumap.umap import tfUMAP
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import umap
import pandas as pd
```
### Load dataset
```
dataset = 'mnist'
dims = (28,28,1)
from tensorflow.keras.datasets import mnist
# load dataset
(train_images, Y_train), (test_images, Y_test) = mnist.load_data()
X_train = (train_images/255.).astype('float32')
X_test = (test_images/255.).astype('float32')
X_train = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
# subset a validation set
n_valid = 10000
X_valid = X_train[-n_valid:]
Y_valid = Y_train[-n_valid:]
X_train = X_train[:-n_valid]
Y_train = Y_train[:-n_valid]
# flatten X
X_train_flat = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test_flat = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
X_valid_flat= X_valid.reshape((len(X_valid), np.product(np.shape(X_valid)[1:])))
print(len(X_train), len(X_valid), len(X_test))
```
### define networks
```
dims = (28,28,1)
n_components = 2
encoder = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=dims),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation="relu"
),
tf.keras.layers.Conv2D(
filters=128, kernel_size=3, strides=(2, 2), activation="relu"
),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=n_components),
])
```
### Create model and train
```
batch_size = 5000
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
from tfumap.parametric_tsne import compute_joint_probabilities, tsne_loss
save_loc = DATA_DIR/ 'parametric_tsne'/ 'dataset' / 'P.npy'
if save_loc.exists():
P = np.load(save_loc)
else:
P = compute_joint_probabilities(X_train_flat, batch_size=batch_size, perplexity=30, verbose=2)
np.save(save_loc, P)
# Joint probabilities of data
Y_train_tsne = P.reshape(X_train.shape[0], -1)
opt = tf.keras.optimizers.Adam(lr=0.01)
encoder.compile(loss=tsne_loss(d=n_components, batch_size=batch_size), optimizer=opt)
X_train = np.reshape(X_train, ([len(X_train)]+ list(dims)))
X_test = np.reshape(X_test, ([len(X_test)]+ list(dims)))
# because shuffle == False, the same batches are used each time...
history = encoder.fit(X_train, Y_train_tsne, batch_size=batch_size, shuffle=False, nb_epoch=1000)
```
### get z for training and test
```
z = encoder.predict(X_train)
z_test = encoder.predict(X_test)
```
### Test plot
```
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=Y_train, cmap=plt.cm.tab10)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=Y_test, cmap=plt.cm.tab10)
```
### Save models + projections
```
import os
output_dir = MODEL_DIR/'projections'/ dataset / 'parametric-tsne'
encoder.save(os.path.join(output_dir, "encoder"))
np.save(output_dir / 'z.npy', z)
np.save(output_dir / 'z_test.npy', z_test)
```
### compute metrics
#### silhouette
```
from tfumap.silhouette import silhouette_score_block
ss, sil_samp = silhouette_score_block(z, Y_train, n_jobs = -1)
ss
ss_test, sil_samp_test = silhouette_score_block(z_test, Y_test, n_jobs = -1)
ss_test
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=sil_samp, cmap=plt.cm.viridis)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=sil_samp_test, cmap=plt.cm.viridis)
```
#### KNN
```
from sklearn.neighbors import KNeighborsClassifier
neigh5 = KNeighborsClassifier(n_neighbors=5)
neigh5.fit(z, Y_train)
score_5nn = neigh5.score(z_test, Y_test)
score_5nn
neigh1 = KNeighborsClassifier(n_neighbors=1)
neigh1.fit(z, Y_train)
score_1nn = neigh1.score(z_test, Y_test)
score_1nn
```
#### Trustworthiness
```
from sklearn.manifold import trustworthiness
tw = trustworthiness(X_train_flat[:10000], z[:10000])
tw_test = trustworthiness(X_test_flat[:10000], z_test[:10000])
tw, tw_test
```
#### save output metrics
```
metrics_df = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw, ss, sil_samp]
metrics_df
save_loc = DATA_DIR / 'projection_metrics' / 'train' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
metrics_df_test = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df_test.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw_test, ss_test, sil_samp_test]
metrics_df_test
save_loc = DATA_DIR / 'projection_metrics' / 'test' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
nn_acc_df = pd.DataFrame(columns = ["method_","dimensions","dataset","1NN_acc","5NN_acc"])
nn_acc_df.loc[len(nn_acc_df)] = ['parametric-tsne', n_components, dataset, score_1nn, score_5nn]
nn_acc_df
save_loc = DATA_DIR / 'knn_classifier' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
nn_acc_df.to_pickle(save_loc)
```
| github_jupyter |
# Description:
Notebooks in this directory describe isopycnicGenomes workflow where probability density functions fit to the fragment BD distribution simulated for each genome.
* PDF = probability density function
* KDE = kernel density estimation
* Implementing some of the workflow in python (except simulating the initial communities, which is done with grinder).
___
## Experiments for testing incorporation using the modeling tool
### atom % 13C
* Simulates isotope dilution or short incubations
* Method
* incorporation % treatments: 10, 20, 40, 60, 80, 100%
* Total treatments: 6
* Title: perc_incorp
### Variable incorporation
* Simulate variable percentages of incorporation and see which taxa are ID'ed
* A fuzzy cutoff of percent incorp that is detectable?
* Method
* 25% of taxa incorporate
* incorporation % distribution:
* uniform: 0-100, 50-100, 0-50
* normal: mean=50, sd=c(0,10,20,30)
### Few incorporators vs many (e.g., ~10% vs ~50%)
* Simulates communities of specialists vs generalists
* Or a more recalcitrant vs labile substrate
* Method
* factorial
* % incorporator treatments: 5, 10, 20, 40, 60%
* % incorporation: 20, 50, or 100%
* Total treatments: 5 x 3 = 15
* Title: perc_tax-incorp
### Community evenness
* differing levels of evenness
* Method
* evenness levels: uniform, log (differing params)
### Community richness
* soil vs simpler habitat
* Method
* N-taxa treatments: 100, 500, 1000 taxa
### Relative abundance of incorporators
* dominant vs rare responders
* Could just do a post-hoc analysis on which taxa were detected by DESeq2
### split populations: active vs dead/dormant
* only some individuals incorporate isotope
* Method
* 25% of taxa incorporate
* factorial
* % split populations: 10, 50, 100%
* incorporation % treatments: 10, 50, 100%
### split populations: full incorp vs partial
* only some individuals incorporate isotope
* Method
* 25% of taxa incorporate
* factorial
* % split populations: 100%
* all in 1 sub-population incorporate 100%
* partial incorporation % treatments: 10, 20, 40, 60, 80, 100%
### Differences in rank-abundance between control and treatment
* differing numbers of taxa with varying rank-abundances in the community
* Method
* Simulate communities with different levels of shared rank-abundances
* `SIPSim gradientComms --perm_perc`
### Number of biological replicates
* N-replicates to test: 1, 3, 5
* Create 5 replicate control and treatment communities
* Test for incorp ID accuracy with differing number of replicates
* How to combine replicates?
### Cross feeding
* Nearly 100% incorporators with some partial 2ndary feeders
* Method
* Incorporators: normal dist, mean=90, sd=5
* Secondary feeders: normal dist, mean=c(10,20,30,40,50), sd=10
### Phylogenetic signal vs random trait distribution
* Random trait distributions coupled with high levels of microdiversity could cause 'split populations' and dilute the signal of incorporation
* Clustering phylotypes (genomes) to produce course-grained phylotypes and see if the incorporation signal is lost
* Method
* 25% of taxa incorporate
* taxonomic clustering: genus, family, order, class, phylum
* % incorporation: 10, 20, 40, 60, 80, 100
# General Workflow
## Fragment GC distributions
fragSim.py
* input:
* genomes
* number of fragments per genome
* [primers]
* workflow:
* Foreach genome:
* select amplicons (if needed)
* simulate fragments
* calculate GC content
* output: table of fragment GC contents
## Community abundance and incorporation
### Simulate communities for each gradient fraction
gradientComms.py
* Goal:
* simulate abundance of taxa in >=1 community
* Input:
* Genome list file
* Grinder options (profile file)
* workflow:
* call modified Grinder with options
* output:
* table of taxon abundances for each sample
### Simulate isotope incorp
isoIncorp.py
* Goal:
* for each taxon in community, how isotope incorp distributed across individuals?
* example: taxonX incorp is normally distributed with mean of X and s.d. of Y
* user defines distribution and params
* can these distributions be 'evolved' across the tree (if provided)?
* brownian evolution of each param?
* 'Fragments' assumed to be pulled randomly from taxon population
* Input:
* community file
* [isotope]
* [percent taxa with any incorp]
* [intra-taxon incorp: specify distribution and params]
* special: GMM: [GMM, weights, Norm1_params, Norm2_params]
* inter-taxon incorp:
* specify distributions of how intra-taxon params vary
* OR phylogeny: distribution params 'evolved' across tree
* Workflow:
* Load community file
* if phylogeny:
* call R script to get which taxa incorporate based on tree (brownian motion)
* % of taxa defined by user
* else:
* random selection of taxa
* % of taxa defined by user
* For incorporators (randomly ordered!):
* if phylogeny:
* brownian motion evo. of intra-taxon incorp distribution params
* else:
* select intra-taxon incorp distribution params from inter-taxon param distribution
* For non-incorps:
* uniform distribution with min = 0 & max = 0
* Output (incorp file):
* tab-delim table:
* sample, taxon, intra-taxon_incorp_disribution_type, distribution_params...
## Creating OTU tables
make_OTU_table.py
* input:
* frag GC file
* community file
* incorp file
* [BD distribution [default: cauchy, params...]]
* [weight (multiplier) for incorp by abundance]
* class creation:
* fragGC
* KDE fit to fragment G+C values for each taxon
* http://scikit-learn.org/stable/modules/density.html
* class:
* library : taxon : KDE fit
* simulate gradients (BD fractions)
* min-max based on theoretical min-max BD
* class:
* library : bin : otu : count
* function: place_in_bin(self, library, otu, BD)
* comm
* subclassed pandas dataframe?
* incorp file
* parse into distributions for each lib-taxon
* dict-like:
* library : taxon : pymix_distribution
* otu_table
* dict-like -- library : fraction : OTU : OTU_count
* workflow (for each sample):
* load community file, incorp file, fragGC file
* Foreach sample:
* Foreach taxon (from comm class):
* sample GC value from KDE
* option: Log GC values from KDE
* select perc. incorp from intra-taxon incorp-distribution
* option: Log perc. incorp
* calculate BD (GC + perc. incorp)
* simulate gradient noise:
* select value (new BD) from (cauchy) distribution (mean = calculated BD)
* bin BD value in corresponding fraction
* save: sample => fraction => OTU
* save as pandas dataframe?
* write out OTU table (frag counts for taxon in fraction = OTU/taxon abundance)
* output:
* OTU table:
* rows=OTU, columns=library-fraction
# Validation
## Fragment simulation
* E. coli
* Plotting fragment length distributions under different conditions
## KDE simulation
* E. coli
### Bandwidth
* Testing effect bandwidth param
* Plotting/comparing distributions made with differing params
### Monte Carlo estimation
* Testing effect of Monte Carlo estimation of distributions
* Plotting/comparing distributions made with differing params
## Community
* Plotting community rank-abundances with different params
* Calculating beta diversity with different params
## Incorporation
* E. coli
* Plotting BD distribution with differing incorporation settings
* Settings:
* Uniform incorporation
* Normal incorporation
* Split populations
## Gradient fractions
* Plotting gradient fractions with differing params
***
#~OLD~
### Gaussian mixture models with pymix
n1 = mixture.NormalDistribution(-2,0.4)
n2 = mixture.NormalDistribution(2,0.6)
m = mixture.MixtureModel(2,[0.5,0.5], [n1,n2])
print m
m.sample()
## Fragment GC distributions
fragSim.py
* input:
* genomes
* number of fragments per genome
* [primers]
* workflow:
* loading genomes as flat-file db
* loading primers as biopython seq records
* Foreach genome:
* find amplicons (if needed)
* load genome seq as Dseqrecord
* in-silico PCR with pydna
* simulate fragments
* get position from amplicon, simulate fragment around amplicon, pull out sequence from genome
* calculate GC content
* apply KDE to GC values (non-parametric approach)
* output:
* pickle: genome => KDE_object
### TODO:
* in-silico PCR
* pydna (http://pydna.readthedocs.org/en/latest/pydna.html)
* genome flat-file indexing
* pyfasta (https://pypi.python.org/pypi/pyfasta/)
***
# ~OLD~
***
# Specific Workflow
## Simulate isotope incorporation
* __makeIncorp_phylo.pl__
* already complete
## Define pre-isopycnic community
* __preIsopycnicComm.pl__
* really, just utilizing Grinder
* complete?
## Simulate genome fragments and calculate BD
* __simFrags.py__
* simulation of certain number of fragment from each genome
* simulation of amplicon or metagenome fragments
* foreach fragment:
* calculate GC
* calculate BD based on isotope incorp of genome
* Output:
* csv: sample, genome, GC, BD
## Create gradient communities
* __simGradientComms.py__
* Simulate communities for each gradient fraction
* __simGradientComms.py__
* simulate fractions
* difference fractions for each gradient (each sample)
* Foreach sample: Foreach genome:
* apply KDE to BD values (non-parametric approach)
* simulate number of 'fragments' needed to meet abs abund as defined in pre-isopynic community
* bin sim-frags by fraction
* Output:
* csv: sample, genome, abs_abundance, rel_abundance
***
#~OLD~
***
## questions:
* Can biopython simulate PCR (as done with bioperl)?
## TODO:
* simFractions.py
* simulate gradient fractions
* input: simulated community
* output: table
* sample, fraction_num, BD_start, BD_end
* params:
* isotope(s)
* determines possible max BD
* gradient min-max
* fraction size dist: mean, stdev
# isopycnic.py scheme
* input:
* genome fasta
* sim community file
* isotope incorp file
* fraction file
* main:
* foreach genome (workflow independent by genome; parallel)
* bio.sequence instance for genome
* isopycnic instance (incorp, simComm, script_args)
* both tables loaded as pandas DataFrames
* incorp table melted; column of sample index
* creating read index (genome location where reads originated)
* artificial PCR to select regions if amplicon fragment (primers provided)
* random fragments if shotgun
* generate by calling grinder?
* simulating fragment start-end
* from read start-end
* fragmenting genome
* load genome
* select sequence for each frag start-end
* calculate GC
* calculate BD
* array of BD values:
* write out [if wanted]
* fit to distributions
# Specific Workflow - take 2
## Simulate isotope incorporation
* __makeIncorp_phylo.pl__
* already complete
## Define pre-isopycnic community
* __preIsopycnicComm.pl__
* really, just utilizing Grinder
* complete?
## Simulate genome fragments and calculate BD
* __isopycnic.pl__
* alter:
* write out the BD of each simulated fragment
* output: sample, genome, frag_scaf, frag_start, frag_end, GC, BD
## Fitting BD values to a PDF
* __fitBD.py__
* load table as dict {genome: [BD_values,]}
* foreach genome (parallel):
* fit data to distributions
* output table:
* genome, AIC, BIC, distribution, l-moment(s)
## Create gradient communities
* __makeGradientComms.py__
* input:
* fitBD.py output
* community abundance file
* Foreach genome: random draws from PDFs
* N-draws based on total community abundance & relative abundances of taxa
* using scipy for drawing from distribution
* output table of OTU abs abundance
* each genome = OTU
# Specific Workflow - take 3
## Simulate isotope incorporation
* __makeIncorp_phylo.pl__
* already complete
## Define pre-isopycnic community
* __preIsopycnicComm.pl__
* really, just utilizing Grinder
* complete?
## simulate reads
* __grinderSE.pl__
* altered grinder that just outputs start-end of reads (not read itself)
* parallel:
* for each sample-genome
* Output: sample, genome, scaffold, read_start, read_end
## simulate fragments
* __fragSim.py__
* parallel:
* for each sample-genome
* determine frag start-end from genome
* get fragment, calculate G+C and BD
## Simulate genome fragments and calculate BD
* __isopycnic.py__
* output: sample, genome, frag_scaf, frag_start, frag_end, GC, BD
## Fitting BD values to a PDF
* __fitBD.py__
* load table as dict {genome: [BD_values,]}
* foreach genome (parallel):
* fit data to distributions
* output table:
* genome, AIC, BIC, distribution, l-moment(s)
## Create gradient communities
* __makeGradientComms.py__
* input:
* fitBD.py output
* community abundance file
* Foreach genome: random draws from PDFs
* N-draws based on total community abundance & relative abundances of taxa
* using scipy for drawing from distribution
* output table of OTU abs abundance
* each genome = OTU
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
import itertools
import warnings
import string
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
from sklearn.metrics import roc_auc_score as auc
from sklearn.metrics import roc_curve
from sklearn.metrics import accuracy_score
from scipy.stats import mode
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
```
### Reading Data
```
df_moviereviews = pd.read_csv('/.../Chapter 11/CS - IMDB Classification/txt_sentoken/Data_IMDB.csv',index_col=0)
df_moviereviews.head()
df_moviereviews["label"].value_counts().plot(kind='pie')
plt.tight_layout(pad=1,rect=(0, 0, 0.7, 1))
plt.text(x=-0.9,y=0.1, \
s=(np.round(((df_moviereviews["label"].\
value_counts()[0])/(df_moviereviews["label"].value_counts()[0] + \
df_moviereviews["label"].value_counts()[1])),2)))
plt.text(x=0.4,y=-0.3, \
s=(np.round(((df_moviereviews["label"].\
value_counts()[1])/(df_moviereviews["label"].value_counts()[0] + \
df_moviereviews["label"].value_counts()[1])),2)))
plt.title("% Share of the Positive and Negative reviews in the dataset")
```
### Coding Positive & Negative Sentiments
```
df_moviereviews.loc[df_moviereviews["label"]=='positive',"label",]=1
df_moviereviews.loc[df_moviereviews["label"]=='negative',"label",]=0
df_moviereviews.head()
```
### Cleaning Data before modelling
```
lemmatizer = WordNetLemmatizer()
def process_text(text):
nopunc = [char for char in text if char not in string.punctuation]
nopunc = ''.join(nopunc)
clean_words = [word.lower() for word in nopunc.split() if word.lower() not in stopwords.words('english')]
clean_words = [lemmatizer.lemmatize(lem) for lem in clean_words]
clean_words = " ".join(clean_words)
return clean_words
df_moviereviews['text'] = df_moviereviews['text'].apply(process_text)
print("Verifying the text cleanup")
df_moviereviews['text'].head()
X = df_moviereviews.loc[:,'text']
Y = df_moviereviews.loc[:,'label']
Y = Y.astype('int')
```
### Train Test Split
```
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=.3, random_state=1)
```
### Using the Count Vectorized to convert text to Vectors
```
count_vectorizer = CountVectorizer()
count_train = count_vectorizer.fit_transform(X_train)
count_test = count_vectorizer.transform(X_test)
```
### Using the TFidf Count Vectorized to convert text to TFidf Vectors
```
tfidf = TfidfVectorizer()
tfidf_train = tfidf.fit_transform(X_train)
tfidf_test = tfidf.transform(X_test)
```
### Fitting Random Forest on count data with GridSearchCV to classify
```
# Set the parameters for grid search
rf_params = {"criterion":["gini","entropy"],\
"min_samples_split":[2,3],\
"max_depth":[None,2,3],\
"min_samples_leaf":[1,5],\
"max_leaf_nodes":[None],\
"oob_score":[True]}
# Create an instance of the RandomForestClassifier()
rf = RandomForestClassifier()
warnings.filterwarnings("ignore")
# Use gridsearchCV(), pass the values you have set for grid search
rf_count = GridSearchCV(rf, rf_params, cv=5)
rf_count.fit(count_train, Y_train)
# Predict class predictions & class probabilities with test data
rf_count_predicted_values = rf_count.predict(count_test)
rf_count_probabilities = rf_count.predict_proba(count_test)
rf_count_train_accuracy = rf_count.score(count_train, Y_train)
rf_count_test_accuracy = rf_count.score(count_test, Y_test)
print('The accuracy for the training data is {}'.\
format(rf_count_train_accuracy))
print('The accuracy for the testing data is {}'.\
format(rf_count_test_accuracy))
print(classification_report(Y_test, rf_count_predicted_values))
# Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, rf_count_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names,normalize=False)
plt.show()
```
### Fitting Random Forest on TF-IDF data with GridSearchCV to classify
```
#Set the parameters for grid search
rf_params = {"criterion":["gini","entropy"],"min_samples_split":[2,3],"max_depth":[None,2,3],"min_samples_leaf":[1,5],"max_leaf_nodes":[None],"oob_score":[True]}
#Create an instance of the Random Forest Classifier()
rf = RandomForestClassifier()
warnings.filterwarnings("ignore")
#Use gridsearchCV(), pass the values you have set for grid search
rf_tfidf = GridSearchCV(rf, rf_params, cv=5)
rf_tfidf.fit(tfidf_train, Y_train)
rf_tfidf_predicted_values = rf_tfidf.predict(tfidf_test)
rf_tfidf_probabilities = rf_tfidf.predict_proba(tfidf_test)
rf_tfidf_train_accuracy = rf_tfidf.score(tfidf_train, Y_train)
rf_tfidf_test_accuracy = rf_tfidf.score(tfidf_test, Y_test)
print('The accuracy for the training data is {}'.format(rf_tfidf_train_accuracy))
print('The accuracy for the testing data is {}'.format(rf_tfidf_test_accuracy))
print(classification_report(Y_test, rf_tfidf_predicted_values))
#Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, rf_tfidf_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names)
plt.show()
```
### Fitting a Naive bayes on Count Data
```
nb_count = MultinomialNB()
nb_count.fit(count_train, Y_train)
nb_count_predicted_values = nb_count.predict(count_test)
nb_count_probabilities = nb_count.predict_proba(count_test)
nb_count_train_accuracy = nb_count.score(count_train, Y_train)
nb_count_test_accuracy = nb_count.score(count_test, Y_test)
print('The accuracy for the training data is {}'.format(nb_count_train_accuracy))
print('The accuracy for the testing data is {}'.format(nb_count_test_accuracy))
print(classification_report(Y_test, nb_predicted_values))
#Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, nb_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names)
plt.show()
```
### Fitting a Naive Bayes on TFIDF Data
```
nb_tfidf = MultinomialNB()
nb_tfidf.fit(count_train, Y_train)
nb_tfidf_predicted_values = nb_tfidf.predict(tfidf_test)
nb_tfidf_probabilities = nb_tfidf.predict_proba(tfidf_test)
nb_tfidf_train_accuracy = nb_tfidf.score(tfidf_train, Y_train)
nb_tfidf_test_accuracy = nb_tfidf.score(tfidf_test, Y_test)
print('The accuracy for the training data is {}'.format(nb_tfidf_train_accuracy))
print('The accuracy for the testing data is {}'.format(nb_tfidf_test_accuracy))
print(classification_report(Y_test, nb_predicted_values))
#Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, nb_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names)
plt.show()
```
### Fitting a SVM with Linear Kernel on Count Data
```
svc_count = SVC(kernel='linear',probability=True)
svc_params = {'C':[0.001, 0.01, 0.1, 1, 10]}
svc_gcv_count = GridSearchCV(svc_count, svc_params, cv=5)
svc_gcv_count.fit(count_train, Y_train)
svc_count_predicted_values = svc_gcv_count.predict(count_test)
svc_count_probabilities = svc_gcv_count.predict_proba(count_test)
svc_count_train_accuracy = svc_gcv_count.score(count_train, Y_train)
svc_count_test_accuracy = svc_gcv_count.score(count_test, Y_test)
print('The accuracy for the training data is {}'.format(svc_count_train_accuracy))
print('The accuracy for the testing data is {}'.format(svc_count_test_accuracy))
print(classification_report(Y_test, svc_count_predicted_values))
#Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, svc_count_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names)
plt.show()
```
### Fitting a SVM with Linear Kernel on TFIDF Data
```
svc_tfidf = SVC(kernel='linear',probability=True)
svc_params = {'C':[0.001, 0.01, 0.1, 1, 10]}
svc_gcv_tfidf = GridSearchCV(svc_tfidf, svc_params, cv=5)
svc_gcv_tfidf.fit(tfidf_train, Y_train)
svc_tfidf_predicted_values = svc_gcv_tfidf.predict(tfidf_test)
svc_tfidf_probabilities = svc_gcv_tfidf.predict_proba(tfidf_test)
svc_tfidf_train_accuracy = svc_gcv_count.score(tfidf_train, Y_train)
svc_tfidf_test_accuracy = svc_gcv_count.score(tfidf_test, Y_test)
print('The accuracy for the training data is {}'.format(svc_tfidf_train_accuracy))
print('The accuracy for the testing data is {}'.format(svc_tfidf_test_accuracy))
print(classification_report(Y_test, svc_tfidf_predicted_values))
#Pass actual & predicted values to the confusion matrix()
cm = confusion_matrix(Y_test, svc_tfidf_predicted_values)
plt.figure()
plot_confusion_matrix(cm, classes=target_names)
plt.show()
```
### Test ROC for all Models
```
plt.subplot(4,3,1)
fpr, tpr, thresholds = roc_curve(Y_test, rf_count_probabilities[:,1])
roc_auc = auc(Y_test, rf_count_probabilities[:,1])
plt.title('ROC Random Forest Count Data')
plt.plot(fpr, tpr, 'b',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.subplot(4,3,2)
fpr, tpr, thresholds = roc_curve(Y_test, rf_tfidf_probabilities[:,1])
roc_auc = auc(Y_test, rf_tfidf_probabilities[:,1])
plt.title('ROC Random Forest TFIDF Data')
plt.plot(fpr, tpr, 'c--',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.subplot(4,3,3)
fpr, tpr, thresholds = roc_curve(Y_test, nb_count_probabilities[:,1])
roc_auc = auc(Y_test, nb_count_probabilities[:,1])
plt.title('ROC Naive Bayes Count Data')
plt.plot(fpr, tpr, 'b',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.subplot(4,3,4)
fpr, tpr, thresholds = roc_curve(Y_test, nb_tfidf_probabilities[:,1])
roc_auc = auc(Y_test, nb_tfidf_probabilities[:,1])
plt.title('ROC Naive Bayes TFIDF Data')
plt.plot(fpr, tpr, 'c--',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.subplot(4,3,5)
fpr, tpr, thresholds = roc_curve(Y_test, svc_count_probabilities[:,1])
roc_auc = auc(Y_test, svc_count_probabilities[:,1])
plt.title('ROC SVM w/Linear kernel Count Data')
plt.plot(fpr, tpr, 'b',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
#plt.show()
plt.subplot(4,3,6)
fpr, tpr, thresholds = roc_curve(Y_test, svc_tfidf_probabilities[:,1])
roc_auc = auc(Y_test, svc_tfidf_probabilities[:,1])
plt.title('ROC SVM w/Linear kernel TFIDF Data')
plt.plot(fpr, tpr, 'c--',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.tight_layout(pad=1,rect=(0, 0, 3, 4))
plt.show()
plt.subplot(4,3,7)
### Ensemble - count data
average_count_probabilities = (nb_count_probabilities + svc_count_probabilities + rf_count_probabilities)/3
fpr, tpr, thresholds = roc_curve(Y_test, average_count_probabilities[:,1])
roc_auc = auc(Y_test, average_count_probabilities[:,1])
plt.title('ROC Ensemble Count Data')
plt.plot(fpr, tpr, 'b',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.subplot(4,3,8)
### Ensemble - TFIDF data
average_TFIDF_probabilities = (nb_tfidf_probabilities + svc_tfidf_probabilities + rf_tfidf_probabilities)/3
fpr, tpr, thresholds = roc_curve(Y_test, average_TFIDF_probabilities[:,1])
roc_auc = auc(Y_test, average_TFIDF_probabilities[:,1])
plt.title('ROC Ensemble TFIDF Data')
plt.plot(fpr, tpr, 'c--',label='AUC = %0.3f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.1,1.0])
plt.ylim([-0.1,1.01])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.tight_layout(pad=1,rect=(0, 0, 3.5, 4))
plt.show()
```
### Calculating accuracy for Ensemble with MAX voting
```
predicted_values_count = np.array([rf_count_predicted_values, \
nb_count_predicted_values, \
svc_count_predicted_values])
predicted_values_tfidf = np.array([rf_tfidf_predicted_values, \
nb_tfidf_predicted_values, \
svc_tfidf_predicted_values])
predicted_values_count = mode(predicted_values_count)
predicted_values_tfidf = mode(predicted_values_tfidf)
count = np.array([rf_count_test_accuracy,\
nb_count_test_accuracy,\
svc_count_test_accuracy,\
accuracy_score(Y_test, predicted_values_count[0][0])])
tfidf = np.array([rf_tfidf_test_accuracy,\
nb_tfidf_test_accuracy,\
svc_tfidf_test_accuracy,\
accuracy_score(Y_test, predicted_values_tfidf[0][0])])
label_list = ["Random Forest", "Naive_Bayes", "SVM_Linear", "Ensemble"]
plt.plot(count)
plt.plot(tfidf)
plt.xticks([0,1,2,3],label_list)
for i in range(4):
plt.text(x=i,y=(count[i]+0.01), s=np.round(count[i],4))
for i in range(4):
plt.text(x=i,y=tfidf[i]-0.006, s=np.round(tfidf[i],4))
plt.legend(["Count","TFIDF"])
plt.title("Test accuracy")
plt.tight_layout(pad=1,rect=(0, 0, 2.5, 2))
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import datetime as datetime
from datetime import datetime
import requests
import os
import json
from dotenv import load_dotenv
import matplotlib
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
%matplotlib inline
load_dotenv()
FMP_API_KEY = os.getenv('FMP_API_KEY')
# Specify stock ticker and index
company = 'TSM'
index = '^GSPC'
```
# Company Overview
```
url = f'https://financialmodelingprep.com/api/v3/profile/{company}?apikey={FMP_API_KEY}'
overview = requests.get(url).json()
overview = pd.DataFrame(overview)
overview = overview.set_index('symbol').T
overview
```
# Company Description
```
# Company description
url = f'https://financialmodelingprep.com/api/v4/company-outlook?symbol={company}&apikey={FMP_API_KEY}'
overview = requests.get(url).json()
overview = overview['profile']['description']
overview
```
# Company Performance
```
start = '2011-01-01'
end = '2021-09-30'
# Pull company historical performance
url = f'https://financialmodelingprep.com/api/v3/historical-price-full/{company}?apikey={FMP_API_KEY}&from={start}&to={end}'
historical = requests.get(url).json()
historical = pd.DataFrame.from_dict(historical, orient='index')
historical = historical[0][1]
historical = pd.DataFrame(historical)
historical['date'] = pd.to_datetime(historical['date'])
historical = historical.set_index('date')
historical['symbol'] = company
historical['daily_ret'] = historical['adjClose'].pct_change().dropna()
historical[:1]
# Pull index historical performance
url = f'https://financialmodelingprep.com/api/v3/historical-price-full/{index}?apikey={FMP_API_KEY}&from={start}&to={end}'
historical_sp = requests.get(url).json()
historical_sp = pd.DataFrame.from_dict(historical_sp, orient='index')
historical_sp = historical_sp[0][1]
historical_sp = pd.DataFrame(historical_sp)
historical_sp['date'] = pd.to_datetime(historical_sp['date'])
historical_sp = historical_sp.set_index('date')
historical_sp['symbol'] = index
historical_sp['daily_ret'] = historical_sp['adjClose'].pct_change().dropna()
historical_sp[:1]
# Plot stock performance against S&P 500 index
fig, ax = plt.subplots(figsize=(12,8))
plt.plot((1+historical_sp['daily_ret']).cumprod(), label =f'{index}', color='r')
ax.set_title(f'{company} vs {index} Cumulative Return')
ax.set_ylabel('return %')
ax.legend(loc='upper left')
plt.grid()
plt.plot((1+historical['daily_ret']).cumprod(), label ='TSM', color='b')
ax.legend(loc='upper left')
```
# Company Quote and P/E
```
# Pull latest stock quote and stats
url = f'https://financialmodelingprep.com/api/v3/quote/{company}?apikey={FMP_API_KEY}'
stock_quote = requests.get(url).json()
stock_quote = pd.DataFrame(stock_quote)
timestamp = stock_quote['timestamp']
dt_obj = datetime.fromtimestamp(timestamp).strftime('%Y-%m-%d')
stock_quote['date'] = dt_obj
stock_quote.set_index('date', inplace=True)
stock_quote
```
# Industry P/E
```
date = '2021-09-30' # change the date to latest or desired date
exchange = 'NYSE' # change to 'NASDAQ' to obtain P/E for that exchange
# Get industry P/E
url = f'https://financialmodelingprep.com/api/v4/industry_price_earning_ratio?date={date}&exchange={exchange}&apikey={FMP_API_KEY}'
industry_stats = requests.get(url).json()
industry_stats = pd.DataFrame(industry_stats)
industry_stats['date'] = pd.to_datetime(industry_stats['date'])
industry_stats= industry_stats.set_index('date')
industry_stats
```
# Sector P/E
```
# Get sector P/E
url = f'https://financialmodelingprep.com/api/v4/sector_price_earning_ratio?date={date}&exchange={exchange}&apikey={FMP_API_KEY}'
sector_stats = requests.get(url).json()
sector_stats = pd.DataFrame(sector_stats)
sector_stats['date'] = pd.to_datetime(sector_stats['date'])
sector_stats= sector_stats.set_index('date')
sector_stats
# Print company, sector and industry P/E to compare and determine if over- or under-valued
company_pe = stock_quote['pe']
industry_pe = industry_stats.loc[industry_stats['industry'] == 'Computer Hardware']
sector_pe = sector_stats.loc[sector_stats['sector'] == 'Technology']
print(f'{company} P/E = {company_pe}')
print(industry_pe)
print(sector_pe)
```
# Institutional Holders
```
# List of institutional holders
url = f'https://financialmodelingprep.com/api/v3/institutional-holder/{company}?apikey={FMP_API_KEY}'
institutional_holders = requests.get(url).json()
institutional_holders = pd.DataFrame(institutional_holders)
institutional_holders = institutional_holders.sort_values(by='shares',ascending=False)
print(institutional_holders[:20])
# Sum of the shares held by institutions and net change
institutional_holders.sum()
```
| github_jupyter |
```
# Load packages
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pandas as pd
import os
import pickle
import time
import scipy as scp
import scipy.stats as scps
from scipy.optimize import differential_evolution
from scipy.optimize import minimize
from datetime import datetime
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
# Load my own functions
import keras_to_numpy as ktnp
import dnnregressor_train_eval_keras as dnnk
import make_data_wfpt as mdw
from kde_training_utilities import kde_load_data
import ddm_data_simulation as ds
import cddm_data_simulation as cds
import boundary_functions as bf
# Handle some cuda business
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID" # see issue #152
os.environ["CUDA_VISIBLE_DEVICES"]="3"
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
import multiprocessing as mp
# Load Model
model_path = '/media/data_cifs/afengler/data/kde/ddm/keras_models/dnnregressor_ddm_06_28_19_00_58_26/model_0'
ckpt_path = '/media/data_cifs/afengler/data/kde/ddm/keras_models/dnnregressor_ddm_06_28_19_00_58_26/ckpt_0_final'
model = keras.models.load_model(model_path)
model.load_weights(ckpt_path)
# model_path = "/home/tony/repos/temp_models/keras_models/dnnregressor_ddm_06_28_19_00_58_26/model_0"
# ckpt_path = "/home/tony/repos/temp_models/keras_models/dnnregressor_ddm_06_28_19_00_58_26/ckpt_0_final"
# model = keras.models.load_model(model_path)
# model.load_weights(ckpt_path)
# network_path = "/home/tony/repos/temp_models/keras_models/\
# dnnregressoranalytical_ddm_07_25_19_15_50_52/model.h5"
#model = keras.models.load_model(network_path)
weights, biases, activations = ktnp.extract_architecture(model)
weights[0].shape
ktnp.log_p(np.array([[0.5, 1, .7, 1, 1]]), weights, biases, activations)
model.predict(np.array([[0.5, 1, .7, 1, 1]]))
def get_params_from_meta_data(file_path = ''):
tmp = pickle.load(open(file_path, 'rb'))[2]
params = []
for key in tmp.keys():
if key == 'delta_t':
break
if key != 's':
params.append(key)
return params
boundary #= eval('bf.constant')
# Initializations -----
n_runs = 1
n_samples = 2500
feature_file_path = '/media/data_cifs/afengler/data/kde/ddm/train_test_data/test_features.pickle'
mle_out_path = '/media/data_cifs/afengler/data/kde/ddm/mle_runs'
# NOTE PARAMETERS:
# WEIBULL: [v, a, w, node, shape, scale]
# param_bounds = [(-1, 1), (0.3, 2), (0.3, 0.7), (0.01, 0.01), (0, np.pi / 2.2)]
# my_optim_columns = ['v_sim', 'a_sim', 'w_sim',
# 'v_mle', 'a_mle', 'w_mle', 'n_samples']
# Get parameter names in correct ordering:
#dat = pickle.load(open(feature_file_path,'rb'))
meta_data_file_path = '/media/data_cifs/afengler/data/kde/ddm/train_test_data/meta_data.pickle'
parameter_names = get_params_from_meta_data(file_path = meta_data_file_path)
param_bounds = [(-1, 1), (0.5, 2), (0.3, 0.7)]
#parameter_names = list(dat.keys())[:-2] # :-1 to get rid of 'rt' and 'choice' here
# Make columns for optimizer result table
p_sim = []
p_mle = []
param_bounds = []
for parameter_name in parameter_names:
p_sim.append(parameter_name + '_sim')
p_mle.append(parameter_name + '_mle')
#param_bounds = param_bounds.append()
my_optim_columns = p_sim + p_mle + ['n_samples']
# Initialize the data frame in which to store optimizer results
optim_results = pd.DataFrame(np.zeros((n_runs, len(my_optim_columns))), columns = my_optim_columns)
optim_results.iloc[:, 2 * len(parameter_names)] = n_samples
# define boundary
boundary = bf.constant
boundary_multiplicative = True
# get network architecture
weights, biases, activations = ktnp.extract_architecture(model)
def make_params(param_bounds = []):
params = np.zeros(len(param_bounds))
for i in range(len(params)):
params[i] = np.random.uniform(low = param_bounds[i][0], high = param_bounds[i][1])
return params
# ---------------------
# Define the likelihood function
def log_p(params = [0, 1, 0.9], model = [], data = [], ll_min = 1e-29):
# Make feature array
feature_array = np.zeros((data[0].shape[0], len(params) + 2))
# Store parameters
cnt = 0
for i in range(0, len(params), 1):
feature_array[:, i] = params[i]
cnt += 1
# Store rts and choices
feature_array[:, cnt] = data[0].ravel() # rts
feature_array[:, cnt + 1] = data[1].ravel() # choices
# Get model predictions
prediction = np.maximum(model.predict(feature_array), ll_min)
return(- np.sum(np.log(prediction)))
param_grid = np.tile(true_params, (data.shape[0], 1))
inp = np.concatenate([param_grid, data], axis=1)
prediction = np_predict(inp, weights, biases, activations)
prediction.sum()
tmp_params = make_params(param_bounds = param_bounds)
boundary_params = {}
ddm_dat_tmp = cds.ddm_flexbound(v = tmp_params[0],
a = tmp_params[1],
w = tmp_params[2],
s = 1,
delta_t = 0.001,
max_t = 20,
n_samples = n_samples,
boundary_fun = boundary, # function of t (and potentially other parameters) that takes in (t, *args)
boundary_multiplicative = boundary_multiplicative, # CAREFUL: CHECK IF BOUND
boundary_params = boundary_params)
data_np = np.concatenate([ddm_dat_tmp[0], ddm_dat_tmp[1]], axis = 1)
t = ktnp.log_p(tmp_params, weights, biases, activations, data_np)
#params, weights, biases, activations, data
t
# Main loop ----------- TD: Parallelize
for i in range(0, n_runs, 1):
# Get start time
start_time = time.time()
# Generate set of parameters
tmp_params = make_params(param_bounds = param_bounds)
# Store in output file
optim_results.iloc[i, :len(parameter_names)] = tmp_params
# Print some info on run
print('Parameters for run ' + str(i) + ': ')
print(tmp_params)
# Define boundary params
boundary_params = {}
# Run model simulations
ddm_dat_tmp = cds.ddm_flexbound(v = tmp_params[0],
a = tmp_params[1],
w = tmp_params[2],
s = 1,
delta_t = 0.001,
max_t = 20,
n_samples = n_samples,
boundary_fun = boundary, # function of t (and potentially other parameters) that takes in (t, *args)
boundary_multiplicative = boundary_multiplicative, # CAREFUL: CHECK IF BOUND
boundary_params = boundary_params)
# Print some info on run
print('Mean rt for current run: ')
print(np.mean(ddm_dat_tmp[0]))
# Run optimizer standard
print('running sequential')
start_time_sequential = time.time()
out = differential_evolution(log_p,
bounds = param_bounds,
args = (model, ddm_dat_tmp),
popsize = 30,
disp = True)
elapsed_sequential = time.time() - start_time_sequential
print(time.strftime("%H:%M:%S", time.gmtime(elapsed_sequential)))
# Run optimizer sequential with ktnp
print('running sequential ktnp')
start_time_sequential_np = time.time()
data_np = np.concatenate([ddm_dat_tmp[0], ddm_dat_tmp[1]], axis = 1)
out_seq_ktnp = differential_evolution(ktnp.log_p,
bounds = param_bounds,
args = (weights, biases, activations, data_np),
popsize = 30,
disp = True)
elapsed_sequential_np = time.time() - start_time_sequential_np
print(time.strftime("%H:%M:%S", time.gmtime(elapsed_sequential_np)))
# Run optimizer parallel
print('running parallel')
start_time_parallel = time.time()
data_np = np.concatenate([ddm_dat_tmp[0], ddm_dat_tmp[1]], axis = 1)
out_parallel = differential_evolution(ktnp.log_p,
bounds = param_bounds,
args = (weights, biases, activations, data_np),
popsize = 30,
disp = True,
workers = -1)
elapsed_parallel = time.time() - start_time_parallel
print(time.strftime("%H:%M:%S", time.gmtime(elapsed_parallel)))
# Print some info
print('Solution vector of current run: ')
print(out.x)
print('Solution vector of current run parallel: ')
print(out_parallel.x)
print('Solution vector of current run seq ktnp')
print(out_seq_ktnp.x)
print('The run took: ')
elapsed = time.time() - start_time
print(time.strftime("%H:%M:%S", time.gmtime(elapsed)))
# Store result in output file
optim_results.iloc[i, len(parameter_names):(2*len(parameter_names))] = out.x
# -----------------------
# Save optimization results to file
optim_results.to_csv(mle_out_path + '/mle_results_1.csv')
# Read in results
optim_results = pd.read_csv(os.getcwd() + '/experiments/ddm_flexbound_kde_mle_fix_v_0_c1_0_w_unbiased_arange_2_3/optim_results.csv')
data_np.shape
plt.scatter(optim_results['v_sim'], optim_results['v_mle'], c = optim_results['c2_mle'])
# Regression for v
reg = LinearRegression().fit(np.expand_dims(optim_results['v_mle'], 1), np.expand_dims(optim_results['v_sim'], 1))
reg.score(np.expand_dims(optim_results['v_mle'], 1), np.expand_dims(optim_results['v_sim'], 1))
plt.scatter(optim_results['a_sim'], optim_results['a_mle'], c = optim_results['c2_mle'])
# Regression for a
reg = LinearRegression().fit(np.expand_dims(optim_results['a_mle'], 1), np.expand_dims(optim_results['a_sim'], 1))
reg.score(np.expand_dims(optim_results['a_mle'], 1), np.expand_dims(optim_results['a_sim'], 1))
plt.scatter(optim_results['w_sim'], optim_results['w_mle'])
# Regression for w
reg = LinearRegression().fit(np.expand_dims(optim_results['w_mle'], 1), np.expand_dims(optim_results['w_sim'], 1))
reg.score(np.expand_dims(optim_results['w_mle'], 1), np.expand_dims(optim_results['w_sim'], 1))
plt.scatter(optim_results['c1_sim'], optim_results['c1_mle'])
# Regression for c1
reg = LinearRegression().fit(np.expand_dims(optim_results['c1_mle'], 1), np.expand_dims(optim_results['c1_sim'], 1))
reg.score(np.expand_dims(optim_results['c1_mle'], 1), np.expand_dims(optim_results['c1_sim'], 1))
plt.scatter(optim_results['c2_sim'], optim_results['c2_mle'], c = optim_results['a_mle'])
# Regression for w
reg = LinearRegression().fit(np.expand_dims(optim_results['c2_mle'], 1), np.expand_dims(optim_results['c2_sim'], 1))
reg.score(np.expand_dims(optim_results['c2_mle'], 1), np.expand_dims(optim_results['c2_sim'], 1))
import numpy as np
testing = np.tile(np.array([1,2,3]), (100, 1))
np.dot(testing.T, testing)
testing.s
ddm_dat_tmp = cds.ddm_flexbound(v = tmp_params[0],
a = tmp_params[1],
w = tmp_params[2],
s = 1,
delta_t = 0.001,
max_t = 20,
n_samples = n_samples,
boundary_fun = boundary, # function of t (and potentially other parameters) that takes in (t, *args)
boundary_multiplicative = boundary_multiplicative, # CAREFUL: CHECK IF BOUND
boundary_params = boundary_params)
```
| github_jupyter |
```
# Required to load webpages
from IPython.display import IFrame
```
[Table of contents](../toc.ipynb)
<img src="https://raw.githubusercontent.com/sympy/sympy/master/doc/src/logo/sympy.svg?sanitize=true" alt="SymPy" width="150" align="right">
# SymPy
* SymPy is a symbolic mathematics library for Python.
* It is a very powerful computer algebra system, which is easy to include in your Python scripts.
* Please find the documentation and a tutorial here [https://www.sympy.org/en/index.html](https://www.sympy.org/en/index.html)
## SymPy live
There is a very nice SymPy live shell in [https://live.sympy.org/](https://live.sympy.org/), where you can try SymPy without installation.
```
IFrame(src='https://live.sympy.org/', width=1000, height=600)
```
## SymPy import and basics
```
import sympy as sp
%matplotlib inline
```
Symbols can be defined with `sympy.symbols` like:
```
x, y, t = sp.symbols('x, y, t')
```
These symbols and later equations are rendered with LaTeX, which makes pretty prints.
```
display(x)
```
Expressions can be easily defined, and equations with left and right hand side are defined with `sympy.Eq` function.
```
expr = x**2
expr
eq = sp.Eq(3*x, -10)
eq
```
Plots are done with `sympy.plot` and the value range can be adjusted.
```
sp.plot(expr, (x, -5, 5))
```
## Why should you consider symbolic math at all?
The power of symbolic computation is their precision. Just compare these two results.
```
import math
math.sqrt(8)
sp.sqrt(8)
```
You can simplify expressions and equations and also expand them.
```
eq = sp.sin(x)**2 + sp.cos(x)**2
eq
sp.simplify(eq)
eq = x*(x + y)
eq
sp.expand(eq)
sp.factor(eq)
```
Differentiation and integration are built in of course.
```
eq = sp.sin(x) * sp.exp(x)
eq
sp.diff(eq, x)
sp.integrate(eq, x)
```
Or define an explicit interval for the integration.
```
sp.integrate(eq, (x, -10, 10))
```
We can also easily substitute one variable of an expression.
```
eq.subs(x, 2)
```
Solve one equation. $x^2 + 3 x = 10$.
```
sp.solve(x**2 + 3*x - 10, x)
```
## More advanced topics
Here, we will solve a linear system of equations.
```
e1 = sp.Eq(3*x + 4*y, -20)
e2 = sp.Eq(4*y, -3)
system_of_eq = [e1, e2]
from sympy.solvers.solveset import linsolve
linsolve(system_of_eq, (x, y))
```
Also differential equations can be used. Let us solve $y'' - y = e^t$ for instance.
```
y = sp.Function('y')
sp.dsolve(sp.Eq(y(t).diff(t, t) - y(t), sp.exp(t)), y(t))
```
Finally, we will have a short look at matrices.
```
A = sp.Matrix([[0, 1],
[1, 0]])
A
A = sp.eye(3)
A
A = sp.zeros(2, 3)
A
```
Inversion of a matrix is done with `**-1` or better readable with `.inv()`.
```
A = sp.eye(2) * 4
A.inv()
A[-2] = x
A
```
## To sum up
* SymPy is a very powerful computer algebra package!
* It is light, small, and easy to install through pip and conda.
* Simple to integrate in your Python project.
| github_jupyter |
```
import time
import scipy
import numpy as np
import pandas as pd
import openml as oml
from sklearn import preprocessing
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import Imputer
from sklearn.model_selection import cross_val_score
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from pymongo import MongoClient
# Silence warnings
import warnings
warnings.simplefilter(action="ignore", category=FutureWarning)
warnings.simplefilter(action="ignore", category=UserWarning)
warnings.simplefilter(action="ignore", category=RuntimeWarning)
'''
Compute Landmarking meta-features according to Matthias Reif et al. 2012.
The accuracy values of the following simple learners are used:
Naive Bayes, Linear Discriminant Analysis, One-Nearest Neighbor,
Decision Node, Random Node.
'''
class LandmarkingMetafeatures():
def __init__(self):
pass
def compute(self, dataset):
data = oml.datasets.get_dataset(dataset)
X, y = data.get_data(target=data.default_target_attribute)
self.landmarkers = get_landmarkers(dataset, X, y)
def connet_mongoclient(host):
client = MongoClient('localhost', 27017)
db = client.landmarkers
return db
def get_landmarkers_from_db():
db = connet_mongoclient('109.238.10.185')
collection = db.landmarkers2
df = pd.DataFrame(list(collection.find()))
datasetID = df['dataset'].astype(int)
return datasetID
def pipeline(X, y, estimator):
start_time_pipeline = time.process_time()
if scipy.sparse.issparse(X) == True: # Check if X is sparse array
X = X.toarray()
pipe = Pipeline([('Imputer', preprocessing.Imputer(missing_values='NaN', strategy='mean', axis=0)),
('classifiers', estimator)])
score = np.mean(cross_val_score(pipe, X, y, cv=10, scoring='roc_auc', n_jobs=-1))
time_pipeline = time.process_time() - start_time_pipeline
return score, time_pipeline
def get_landmarkers(dataset, X, y):
landmarkers = {}
start_time = time.process_time()
landmarkers['datasetID'] = dataset
landmarkers['one_nearest_neighbor'], landmarkers['one_nearest_neighbor_time'] = pipeline(X, y, KNeighborsClassifier(n_neighbors = 1))
landmarkers['linear_discriminant_analysis'], landmarkers['linear_discriminant_analysis_time'] = pipeline(X, y, LinearDiscriminantAnalysis(solver='lsqr', shrinkage='auto'))
landmarkers['naive_bayes'], landmarkers['naive_bayes_time'] = pipeline(X, y, GaussianNB())
landmarkers['decision_node'], landmarkers['decision_node_time'] = pipeline(X, y, DecisionTreeClassifier(criterion='entropy', splitter='best',
max_depth=1, random_state=0))
landmarkers['random_node'], landmarkers['random_node_time'] = pipeline(X, y, DecisionTreeClassifier(criterion='entropy', splitter='random',
max_depth=1, random_state=0))
landmarkers['landmark_time'] = time.process_time() - start_time
return landmarkers
if __name__ == "__main__":
df = []
datasets = get_landmarkers_from_db()
for dataset in datasets:
test = LandmarkingMetafeatures()
test.compute(dataset)
df.append(test.landmarkers)
# db = connet_mongoclient('109.238.10.185')
# db.landmarkers4.insert_one({'landmarkers': df})
test = pd.DataFrame(df)
#test.loc[test['naive_bayes_time'].idxmax()]
test.mean()
pd.DataFrame(df)
%matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi']= 120
data=pd.DataFrame(df)
data_time = data[['one_nearest_neighbor_time','linear_discriminant_analysis_time', 'naive_bayes_time',
'decision_node_time', 'random_node_time']]
data_time.columns = ['One-Nearest Neighbor', 'Linear Discriminant Analysis', 'Gaussian Naive Bayes',
'Decision Node', 'Random Node']
sns.set(style="ticks", font_scale=0.75)
f, ax = plt.subplots(figsize=(8, 4))
# Draw a nested boxplot to show bills by day and sex
box = sns.boxplot(data=data_time, palette="Set3")
sns.despine(offset=10, trim=True)
ax.set(ylabel='Computation time')
figure = box.get_figure()
figure.savefig("computation_time.png")
sns.set(style="whitegrid", font_scale=0.75)
f, ax = plt.subplots(figsize=(8, 4))
landmarkers = data[['one_nearest_neighbor','linear_discriminant_analysis', 'naive_bayes',
'decision_node', 'random_node']]
landmarkers.columns = ['One-Nearest Neighbor', 'Linear Discriminant Analysis', 'Gaussian Naive Bayes',
'Decision Node', 'Random Node']
violin = sns.violinplot(data=landmarkers, palette="Set3", bw=.2, cut=1, linewidth=1)
sns.despine(left=True, bottom=True)
ax.set(ylabel='Score')
figure = violin.get_figure()
figure.savefig("score.png")
```
| github_jupyter |
```
%matplotlib inline
```
# Basic usage example of `DupleBalanceClassifier`
This example shows the basic usage of :class:`duplebalance.DupleBalanceClassifier`.
```
print(__doc__)
RANDOM_STATE = 42
```
## Preparation
First, we will import necessary packages and generate an example
multi-class imbalanced dataset.
```
from duplebalance import DupleBalanceClassifier
from duplebalance.base import sort_dict_by_key
from duplebalance.utils._plot import plot_2Dprojection_and_cardinality
from collections import Counter
import matplotlib.pyplot as plt
from sklearn.decomposition import KernelPCA
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
```
Make a 5-class imbalanced classification task
```
X, y = make_classification(n_classes=5, class_sep=1, # 5-class
weights=[0.05, 0.05, 0.15, 0.25, 0.5], n_informative=10, n_redundant=1, flip_y=0,
n_features=20, n_clusters_per_class=1, n_samples=2000, random_state=0)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
origin_distr = sort_dict_by_key(Counter(y_train))
test_distr = sort_dict_by_key(Counter(y_test))
print('Original training dataset shape %s' % origin_distr)
print('Original test dataset shape %s' % test_distr)
# Visualize the dataset
projection = KernelPCA(n_components=2).fit(X, y)
fig = plot_2Dprojection_and_cardinality(X, y, projection=projection)
plt.show()
```
## Train a DupleBalance Classifier
Basic usage of DupleBalanceClassifier
```
# Train a DupleBalanceClassifier
clf = DupleBalanceClassifier(
n_estimators=5,
random_state=RANDOM_STATE,
).fit(X_train, y_train)
# Predict & Evaluate
score = clf.score(X_test, y_test)
print ("DupleBalance {} | Balanced AUROC: {:.3f} | #Training Samples: {:d}".format(
len(clf.estimators_), score, sum(clf.estimators_n_training_samples_)
))
```
Train DupleBalanceClassifier with automatic parameter tuning
```
# Train a DupleBalanceClassifier
clf = DupleBalanceClassifier(
n_estimators=5,
random_state=RANDOM_STATE,
).fit(
X_train, y_train,
perturb_alpha='auto',
)
# Predict & Evaluate
score = clf.score(X_test, y_test)
print ("DupleBalance {} | Balanced AUROC: {:.3f} | #Training Samples: {:d}".format(
len(clf.estimators_), score, sum(clf.estimators_n_training_samples_)
))
```
Train DupleBalanceClassifier with advanced training log
```
# Train a DupleBalanceClassifier
clf = DupleBalanceClassifier(
n_estimators=5,
random_state=RANDOM_STATE,
).fit(
X_train, y_train,
perturb_alpha='auto',
eval_datasets={'test': (X_test, y_test)},
train_verbose={
'granularity': 1,
'print_distribution': True,
'print_metrics': True,
},
)
# Predict & Evaluate
score = clf.score(X_test, y_test)
print ("DupleBalance {} | Balanced AUROC: {:.3f} | #Training Samples: {:d}".format(
len(clf.estimators_), score, sum(clf.estimators_n_training_samples_)
))
```
| github_jupyter |
# Classification Uncertainty Analysis in Bayesian Deep Learning with Dropout Variational Inference
Here is [astroNN](https://github.com/henrysky/astroNN), please take a look if you are interested in astronomy or how neural network applied in astronomy
* **Henry Leung** - *Astronomy student, University of Toronto* - [henrysky](https://github.com/henrysky)
* Project adviser: **Jo Bovy** - *Professor, Department of Astronomy and Astrophysics, University of Toronto* - [jobovy](https://github.com/jobovy)
* Contact Henry: henrysky.leung [at] utoronto.ca
* This tutorial is created on 16/Mar/2018 with Keras 2.1.5, Tensorflow 1.6.0, Nvidia CuDNN 7.0 for CUDA 9.0 (Optional)
* Updated on 31/Jan/2020 with Tensorflow 2.1.0, Tensorflow Probability 0.9.0
* Updated again on 27/Jan/2020 with Tensorflow 2.4.0, Tensorflow Probability 0.12.0
<br>
For more resources on Bayesian Deep Learning with Dropout Variational Inference, please refer to [README.md](https://github.com/henrysky/astroNN/tree/master/demo_tutorial/NN_uncertainty_analysis)
#### First import everything we need
```
%matplotlib inline
%config InlineBackend.figure_format='retina'
from tensorflow.keras.datasets import mnist
from tensorflow.keras import utils
import numpy as np
import pylab as plt
from astroNN.models import MNIST_BCNN
```
### Train the neural network on MNIST training set
```
(x_train, y_train), (x_test, y_test) = mnist.load_data()
y_train = utils.to_categorical(y_train, 10)
y_train = y_train.astype(np.float32)
x_train = x_train.astype(np.float32)
x_test = x_test.astype(np.float32)
# Create a astroNN neural network instance and set the basic parameter
net = MNIST_BCNN()
net.task = 'classification'
net.max_epochs = 5 # Just use 5 epochs for quick result
# Trian the nerual network
net.train(x_train, y_train)
```
### Test the neural network on random MNIST images
You can see from below, most test images are right except the last one the model has a high uncertainty in it. As a human, you can indeed can argue this 5 is badly written can can be read as 6 or even a badly written 8.
```
test_idx = [1, 2, 3, 4, 5, 8]
pred, pred_std = net.test(x_test[test_idx])
for counter, i in enumerate(test_idx):
plt.figure(figsize=(3, 3), dpi=100)
plt.title(f'Predicted Digit {pred[counter]}, Real Answer: {y_test[i]:{1}} \n'
f'Total Uncertainty (Entropy): {(pred_std["total"][counter]):.{2}}')
plt.imshow(x_test[i])
plt.show()
plt.close('all')
plt.clf()
```
### Test the neural network on random MNIST images with 90 degree rotation
Since the neural network is trained on MNIST images without any data argumentation, so if we rotate the MNIST images, the images should look 'alien' to the neural network and the neural network should give us a high unceratinty. And indeed the neural network tells us its very uncertain about the prediction with roated images.
```
test_rot_idx = [9, 10, 11]
test_rot = x_test[test_rot_idx]
for counter, j in enumerate(test_rot):
test_rot[counter] = np.rot90(j)
pred_rot, pred_rot_std = net.test(test_rot)
for counter, i in enumerate(test_rot_idx):
plt.figure(figsize=(3, 3), dpi=100)
plt.title(f'Predicted Digit {pred_rot[counter]}, Real Answer: {y_test[i]:{1}} \n'
f'Total Uncertainty (Entropy): {(pred_rot_std["total"][counter]):.{2}}')
plt.imshow(test_rot[counter])
plt.show()
plt.close('all')
plt.clf()
```
| github_jupyter |
```
%matplotlib inline
import datacube
dc = datacube.Datacube(config="/home/rishabh/.datacube.conf")
#dc.list_products().loc[:].values
dc.list_products()
dc.list_measurements()
#la = dc.load(product='ls5_ledaps_albers', x=(81.00, 81.05), y=(30.00, 30.05))
la = dc.load(product='ls5_ledaps_albers', x=(500000, 509635), y=(3318785, 3329870))
#la = dc.load(product='ls5_ledaps_albers', x = (213539.9, 218463.5),y = (3433466.7, 3438883.6))
la
(la.items()[0])[1].values[0]
la.data_vars
la.blue
a = la.nir.loc['1990']
a.shape
a
a.plot()
from datetime import datetime
from datetime import datetime
from datacube.analytics.analytics_engine import AnalyticsEngine
from datacube.execution.execution_engine import ExecutionEngine
from datacube.analytics.utils.analytics_utils import plot
a = AnalyticsEngine()
e = ExecutionEngine()
dimensions = {'x': {'range': (213539.9, 218463.5)},
'y': {'range': (3433466.7, 3438883.6)},
'time': {'range': (datetime(2015, 1, 1), datetime(2015, 12, 31))}}
b40 = a.create_array(('LANDSAT_8', 'LEDAPS'), ['nir'], dimensions, 'b40')
b30 = a.create_array(('LANDSAT_8', 'LEDAPS'), ['red'], dimensions, 'b30')
ndvi = a.apply_expression([b40, b30], '((array1 - array2) / (array1 + array2))', 'ndvi')
import numpy
%matplotlib inline
import datacube
import numpy
from datetime import datetime
from datacube.analytics.analytics_engine import AnalyticsEngine
from datacube.execution.execution_engine import ExecutionEngine
from datacube.analytics.utils.analytics_utils import plot
dc = datacube.Datacube(config="/home/rishabh/.datacube.conf")
def RVI (product, x1, x2, y1, y2, year):
a = AnalyticsEngine()
e = ExecutionEngine()
#assert product
#dc.list_products() is pandas dataframe, .loc[:]['name'] selects
# the products is a pandas series. values is array
assert (product in dc.list_products().loc[:]['name'].values), "Product not in database"
# 2 index for platform, 3 for product type
for prod in dc.list_products().loc[:].values:
if(product == prod[0]):
platform = prod[2]
product_type = prod[3]
# assert time
la = dc.load(product=product, x = (x1, x2),y = (y1, y2))
# this is date of product (la.items()[0])[1].values[0]
date_of_prod = (la.items()[0])[1].values[0]
#this is numpy.datetime64
# TO DO COMPARE numpy.datetime64 with datetime.datetime
#assert (date_of_prod >= time1 and date_of_prod <= time2), "Product not in the provided time"
time1 = datetime(year, 1, 1)
time2 = datetime(year, 12, 31)
# calculate output
dimensions = {'x': {'range': (x1, x2)},
'y': {'range': (y1, y2)},
'time': {'range': (time1, time2)}}
# create arrays
b40 = a.create_array((platform, product_type), ['nir'], dimensions, 'b40')
b30 = a.create_array((platform, product_type), ['red'], dimensions, 'b30')
#ratio vegetation index
rvi = a.apply_expression([b40, b30], '(array1 / array2)', 'rvi')
e.execute_plan(a.plan)
#result x array
res = e.cache['rvi']['array_result']['rvi']
res.plot()
# Transformed vegetation index
def TVI (product, x1, x2, y1, y2, year):
a = AnalyticsEngine()
e = ExecutionEngine()
#assert product
#dc.list_products() is pandas dataframe, .loc[:]['name'] selects
# the products is a pandas series. values is array
assert (product in dc.list_products().loc[:]['name'].values), "Product not in database"
# 2 index for platform, 3 for product type
for prod in dc.list_products().loc[:].values:
if(product == prod[0]):
platform = prod[2]
product_type = prod[3]
# assert time
la = dc.load(product=product, x = (x1, x2),y = (y1, y2))
# this is date of product (la.items()[0])[1].values[0]
date_of_prod = (la.items()[0])[1].values[0]
#this is numpy.datetime64
# TO DO COMPARE numpy.datetime64 with datetime.datetime
#assert (date_of_prod >= time1 and date_of_prod <= time2), "Product not in the provided time"
time1 = datetime(year, 1, 1)
time2 = datetime(year, 12, 31)
# calculate output
dimensions = {'x': {'range': (x1, x2)},
'y': {'range': (y1, y2)},
'time': {'range': (time1, time2)}}
# create arrays
b40 = a.create_array((platform, product_type), ['nir'], dimensions, 'b40')
b30 = a.create_array((platform, product_type), ['red'], dimensions, 'b30')
#ratio vegetation index
tvi = a.apply_expression([b40, b30], '(sqrt(((array1 - array2) / (array1 + array2)) + 0.5) * 100)', 'tvi')
e.execute_plan(a.plan)
#result x array
res = e.cache['tvi']['array_result']['tvi']
res.plot()
RVI('ls8_ledaps_albers', 213539.9, 218463.5, 3433466.7, 3438883.6, 2015)
TVI('ls8_ledaps_albers', 213539.9, 218463.5, 3433466.7, 3438883.6, 2015)
numpy.datetime64(datetime(2015,1,1)).item()
b40
e.execute_plan(a.plan)
plot(e.cache['b30'])
plot(e.cache['b40'])
plot(e.cache['ndvi'])
e.cache['ndvi']['array_result']['ndvi']
e.cache['ndvi']['array_result']['ndvi'].plot()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os
import pandas as pd
from download_from_s3 import download_from_s3
from heartrate_model import heartrate_model
from script import batch_model_fit
from plotly import tools
import plotly.graph_objs as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=False)
```
Download stream and activity metadata to local dir:
```
# download_from_s3()
```
Anonymized athlete id's:
```
athlete_1 = '1f47de90be07b9beb5c312b8b090b95c246f6abab38e5bafc2c5591e5b961966'
athlete_2 = 'f1f70037993a7c837808717c9666cf4d6a730025b366b8734f00ef4319c8a12d'
data_dir = 'data'
stream_df = pd.read_json(os.path.join(data_dir, athlete_1, 'a1b0d65d97635bc33f2c402fc68fdd5961d6e8781ff96df6b7938a30497a65dc.json'), orient='split')
```
There can be missing power readings:
```
stream_df.fillna(method='ffill', inplace=True)
```
Fit the model:
```
model, predictions = heartrate_model(stream_df.heartrate, stream_df.watts)
stream_df['predicted_heartrate'] = predictions
```
Plot actual vs. predicted heart rates and show model parameters
```
traces = [go.Scatter(x=stream_df.time, y=stream_df.watts, name='Power', line={'width': 3}),
go.Scatter(x=stream_df.time, y=stream_df.heartrate, name='Heart Rate', yaxis='y2', line={'width': 3}, opacity=0.6),
go.Scatter(x=stream_df.time, y=stream_df.predicted_heartrate, name='Predicted Heart Rate', yaxis='y2', line={'width': 3}, opacity=0.8)]
layout = go.Layout(
yaxis={
'title': 'Power (W)',
'titlefont': {
'size': 24
},
'tickfont': {
'size': 18
},
'showgrid': False
},
yaxis2={
'title': 'Heart Rate (BPM)',
'titlefont': {
'size': 24
},
'tickfont': {
'size': 18
},
'overlaying': 'y',
'side': 'right',
'showgrid': False
},
xaxis={
'title': 'Time (s)',
'titlefont': {
'size': 24
},
'tickfont': {
'size': 18
},
'showgrid': False
},
legend={
'orientation': 'h',
'x': 0,
'y': 100,
'font': {
'size': 18
}
},
width=1024,
height=768,
margin=go.Margin(
b=160,
t=200,
l=120,
r=120
)
)
iplot({'data': traces, 'layout': layout})
model.params
```
Fit model for all activities of athlete
```
# batch_model_fit(athlete_1)
athlete_1_fit_param_df = pd.read_csv('2018-05-07T13:36:29.405488.csv')
athlete_1_activity_meta_df = pd.read_csv('data/1f47de90be07b9beb5c312b8b090b95c246f6abab38e5bafc2c5591e5b961966_activities.csv')
model_params_vs_fitness_df = athlete_1_activity_meta_df.merge(athlete_1_fit_param_df, on='activity_id')
model_params_vs_fitness_df.describe()
iplot([go.Scatter(x=model_params_vs_fitness_df.w_z3, y=model_params_vs_fitness_df.tau_rise, mode='markers')])
```
| github_jupyter |
```
import torch
import os
import cv2
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
from torchvision import transforms
transform_data = transforms.Compose(
[
transforms.ToPILImage(),
transforms.RandomVerticalFlip(),
transforms.RandomHorizontalFlip(),
transforms.RandomCrop((112)),
transforms.ToTensor(),
]
)
def load_data(img_size=112):
data = []
labels = {}
index = -1
for label in os.listdir('./data/'):
index += 1
labels[label] = index
print(len(labels))
X = []
y = []
for label in labels:
for file in os.listdir(f'./data/{label}/'):
path = f'./data/{label}/{file}'
img = cv2.imread(path)
img = cv2.resize(img,(img_size,img_size))
data.append([np.array(transform_data(np.array(img))),labels[label]])
X.append(np.array(transform_data(np.array(img))))
y.append(labels[label])
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.random.shuffle(data)
np.save('./data.npy',data)
VAL_SPLIT = 0.25
VAL_SPLIT = len(X)*VAL_SPLIT
VAL_SPLIT = int(VAL_SPLIT)
X_train = X[:-VAL_SPLIT]
y_train = y[:-VAL_SPLIT]
X_test = X[-VAL_SPLIT:]
y_test = y[-VAL_SPLIT:]
X = torch.from_numpy(np.array(X))
y = torch.from_numpy(np.array(y))
X_train = np.array(X_train)
X_test = np.array(X_test)
y_train = np.array(y_train)
y_test = np.array(y_test)
X_train = torch.from_numpy(X_train)
X_test = torch.from_numpy(X_test)
y_train = torch.from_numpy(y_train)
y_test = torch.from_numpy(y_test)
return X,y,X_train,X_test,y_train,y_test
X,y,X_train,X_test,y_train,y_test = load_data()
```
## Modelling
```
import torch.nn as nn
import torch.nn.functional as F
class BaseLine(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3,32,5)
self.conv2 = nn.Conv2d(32,64,5)
self.conv2batchnorm = nn.BatchNorm2d(64)
self.conv3 = nn.Conv2d(64,128,5)
self.fc1 = nn.Linear(128*10*10,256)
self.fc2 = nn.Linear(256,128)
self.fc3 = nn.Linear(128,50)
self.relu = nn.ReLU()
def forward(self,X):
preds = F.max_pool2d(self.relu(self.conv1(X)),(2,2))
preds = F.max_pool2d(self.relu(self.conv2batchnorm(self.conv2(preds))),(2,2))
preds = F.max_pool2d(self.relu(self.conv3(preds)),(2,2))
preds = preds.view(-1,128*10*10)
preds = self.relu(self.fc1(preds))
preds = self.relu(self.fc2(preds))
preds = self.relu(self.fc3(preds))
return preds
device = torch.device('cuda')
from torchvision import models
# model = BaseLine().to(device)
# model = model.to(device)
model = models.resnet18(pretrained=True).to(device)
in_f = model.fc.in_features
model.fc = nn.Linear(in_f,50)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
PROJECT_NAME = 'Car-Brands-Images-Clf'
import wandb
EPOCHS = 100
BATCH_SIZE = 32
from tqdm import tqdm
wandb.init(project=PROJECT_NAME,name='transfer-learning')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch)
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item()})
# TL vs Custom Model best = TL
def get_loss(criterion,y,model,X):
model.to('cuda')
preds = model(X.view(-1,3,112,112).to('cuda').float())
preds.to('cuda')
loss = criterion(preds,torch.tensor(y,dtype=torch.long).to('cuda'))
loss.backward()
return loss.item()
def test(net,X,y):
device = 'cuda'
net.to(device)
correct = 0
total = 0
net.eval()
with torch.no_grad():
for i in range(len(X)):
real_class = torch.argmax(y[i]).to(device)
net_out = net(X[i].view(-1,3,112,112).to(device).float())
net_out = net_out[0]
predictied_class = torch.argmax(net_out)
if predictied_class == real_class:
correct += 1
total += 1
net.train()
net.to('cuda')
return round(correct/total,3)
EPOCHS = 12
BATCH_SIZE = round(len(X_train)/2.55)
model = models.inception_v3(pretrained=False, num_classes=50,init_weights=True).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
wandb.init(project=PROJECT_NAME,name=f'models.inception_v3')
for _ in tqdm(range(EPOCHS),leave=False):
for i in tqdm(range(0,len(X_train),BATCH_SIZE),leave=False):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch)
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,y_test,model,X_test),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test)})
model = models.shufflenet_v2_x1_0(pretrained=False, num_classes=50).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(),lr=0.1)
wandb.init(project=PROJECT_NAME,name=f'models.shufflenet_v2_x1_0')
for _ in tqdm(range(EPOCHS),leave=False):
for i in tqdm(range(0,len(X_train),BATCH_SIZE),leave=False):
X_batch = X_train[i:i+BATCH_SIZE].view(-1,3,112,112).to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch)
preds = preds.to(device)
loss = criterion(preds,y_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,y_test,model,X_test),'accuracy':test(model,X_train,y_train),'val_accuracy':test(model,X_test,y_test)})
[models.mobilenet_v3_large,models.mobilenet_v3_small,models.resnext50_32x4d,models.wide_resnet50_2,models.mnasnet1_0]
```
| github_jupyter |
This notebook was prepared by [Donne Martin](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Solution Notebook
## Problem: Determine if a tree is a valid binary search tree.
* [Constraints](#Constraints)
* [Test Cases](#Test-Cases)
* [Algorithm](#Algorithm)
* [Code](#Code)
* [Unit Test](#Unit-Test)
## Constraints
* Can the tree have duplicates?
* Yes
* If this is called on a None input, should we raise an exception?
* Yes
* Can we assume we already have a Node class?
* Yes
* Can we assume this fits in memory?
* Yes
## Test Cases
None -> exception
<pre>
Valid:
5
/ \
5 8
/ \ /
4 6 7
Invalid:
5
/ \
5 8
\
20
</pre>
## Algorithm
We'll use a recursive solution that valides left <= current < right, passing down the min and max values as we do a depth-first traversal.
* If the node is None, return True
* If min is set and the node's value <= min, return False
* if max is set and the node's value > max, return False
* Recursively call the validate function on node.left, updating max
* Recursively call the validate function on node.right, updating min
Complexity:
* Time: O(n)
* Space: O(h), where h is the height of the tree
## Code
```
%run ../bst/bst.py
import sys
class BstValidate(Bst):
def validate(self):
if self.root is None:
raise TypeError('No root node')
return self._validate(self.root)
def _validate(self, node, mininum=-sys.maxsize, maximum=sys.maxsize):
if node is None:
return True
if node.data <= mininum or node.data > maximum:
return False
if not self._validate(node.left, mininum, node.data):
return False
if not self._validate(node.right, node.data, maximum):
return False
return True
```
## Unit Test
```
%%writefile test_bst_validate.py
from nose.tools import assert_equal
from nose.tools import raises
class TestBstValidate(object):
@raises(Exception)
def test_bst_validate_empty(self):
validate_bst(None)
def test_bst_validate(self):
bst = BstValidate(Node(5))
bst.insert(8)
bst.insert(5)
bst.insert(6)
bst.insert(4)
bst.insert(7)
assert_equal(bst.validate(), True)
bst = BstValidate(Node(5))
left = Node(5)
right = Node(8)
invalid = Node(20)
bst.root.left = left
bst.root.right = right
bst.root.left.right = invalid
assert_equal(bst.validate(), False)
print('Success: test_bst_validate')
def main():
test = TestBstValidate()
test.test_bst_validate_empty()
test.test_bst_validate()
if __name__ == '__main__':
main()
%run -i test_bst_validate.py
```
| github_jupyter |
```
import numpy as np
import scipy.stats
from matplotlib import pyplot as plt
import seaborn as sns
# width = 1000000
# points = 21
# noise = 0.5
# x = np.linspace(
# scipy.stats.norm.ppf(1 / width),
# scipy.stats.norm.ppf(1 - 1 / width), points
# )
# y = scipy.stats.norm.pdf(x)
# plt.plot(
# x + np.random.rand(),
# y * (1 + noise * np.random.rand(points)) * np.abs(np.random.normal()),
# marker=".",
# c='r',
# )
# plt.plot(
# x + np.random.rand(),
# y * (1 + noise * np.random.rand(points)) * np.abs(np.random.normal()),
# marker=".",
# c='b',
# )
fragment_count = 5
width = 1000000
points = 21
normal_x = np.linspace(
scipy.stats.norm.ppf(1 / width),
scipy.stats.norm.ppf(1 - 1 / width), points
)
normal_y = scipy.stats.norm.pdf(x)
base_intensities = [
normal_y * np.random.rand() for i in range(fragment_count)
]
# \[
# normal_y + np.random.rand(points)
# ]
samples = {sample: {} for sample in ["A", "B", "C"]}
linewidth = 3
alpha = 0.6
colors = ["g", "r", "b", "orange", "purple"]
for sample in samples:
sample_intensities = 0.1 * np.random.normal(size=fragment_count)
samples[sample_name]["relative intensity"] = base_intensities + sample_intensities
for sample in samples:
samples[sample_name]["CCS"] = base_intensities + sample_intensities
samples["A"]["retention time"] = np.concatenate(
[
np.random.normal(size=2)*0.1,
np.random.normal(size=1)*0.1 + 0,
np.random.normal(size=2)*0.1 + 0,
]
)
samples["B"]["retention time"] = np.concatenate(
[
np.random.normal(size=2)*0.1,
np.random.normal(size=1)*0.1 + 0.2,
np.random.normal(size=2)*0.1 + 0.5,
]
)
samples["C"]["retention time"] = np.concatenate(
[
np.random.normal(size=2)*0.1,
np.random.normal(size=1)*0.1 + 0,
np.random.normal(size=2)*0.1 + 0.5,
]
)
sample = "A"
for i, j, c in zip(
samples["A"]["rts"],
samples["A"]["ints"],
):
plt.plot(x + i, y * j, c=c, linewidth=linewidth, alpha=alpha)
plt.xlabel("retention time")
plt.ylabel("relative intensity")
plt.yticks([])
plt.xticks([])
plt.title(f"Sample {sample}")
data = [
np.concatenate(
[
np.random.normal(size=2)*0.1,
np.random.normal(size=1)*0.1 + 0,
np.random.normal(size=2)*0.1 + 0,
]
),
ints + 0.1 * np.random.normal(size=5),
colors,
]
for i, j, c in zip(*data):
plt.plot(x + i, y * j, c=c, linewidth=linewidth, alpha=alpha)
plt.xlabel("CCS")
plt.ylabel("relative intensity")
plt.yticks([])
plt.xticks([])
plt.title("Sample A")
ints + 0.1 * np.random.normal(size=5)
```
| github_jupyter |
## CE9010: Introduction to Data Analysis
## Semester 2 2018/19
## Xavier Bresson
<hr>
## Tutorial 1: Introduction to Python
## Objective
### $\bullet$ Basic operations in Python
<hr>
## 1. Resources
<hr>
Slides: [Python introduction by Xavier Bresson]
[Python introduction by Xavier Bresson]: http://data-science-training-xb.s3-website.eu-west-2.amazonaws.com/All_lectures/Lecture02_python.pdf
Notebook: [Python introduction tutorial by Justin Johnson]
[Python introduction tutorial by Justin Johnson]: https://github.com/kuleshov/cs228-material/blob/master/tutorials/python/cs228-python-tutorial.ipynb
<br>
## 2. Basic operations
<hr>
### 2.1 Elementary algebra operations
```
5+6
5/8
5**8
```
### 2.2 Logical operations
```
1==1
2==1
2!=1
True & False
True | False
```
### 2.3 Assignment operations
```
x=3
print(x)
x='hello'
print(x)
x=(1==2)
print(x)
x=2.35789202950400
print('x={:2.5f}, x={:.1f}'.format(x,x))
x=2.35789202950400
print(type(x))
x='hello'
print(type(x))
```
### 2.4 Numpy
List all libraries modudels: np. + tab
List properties of the module: np.abs + shit + tab
```
import numpy as np
#np.abs()
x = np.pi
print(x)
print(np.ones((5,2,4)))
print(np.zeros(5))
print(np.eye(5))
print(np.random.normal(0,1))
print(np.random.uniform(0,1))
print(np.random.randint(10))
print(np.array(range(10)))
print(np.random.permutation(range(10)))
x = np.array(x,dtype='float32')
print(x,type(x),x.dtype)
x = np.array(3.4,dtype='int64')
print(x,type(x),x.dtype)
X = np.array([[1,2,3],[4,5,6]])
print(X)
print(X.shape)
print(X.shape[0])
print(X.shape[1])
print(X)
print(X[0,2])
print(X[0,:])
print(X[0,0:2])
print(X[0,:2])
print(X[0,1:3])
print(X[-1,-1])
X[0,:] = [7,8,9] # assignment
print(X)
X = np.array([[1,2,3],[4,5,6]])
X = np.append(X,[[7,8,9]],axis=0) # append
print(X)
print(X.shape)
```
<br>
## 3. Load and save data
<hr>
```
pwd
ls -al
cd ..
pwd
cd tutorials
data = np.loadtxt('data/profit_population.txt', delimiter=',')
print(data)
print(data.shape)
print(data.dtype)
new_data = 2* data
np.savetxt('data/profit_population_new.txt', new_data, delimiter=',', fmt='%2.5f')
%whos
```
<br>
## 4. Linear algebra operations
<hr>
```
X = np.array([[1,2,3],[4,5,6]])
print(X,X.shape)
Y = np.array([[2,7,-2],[1,8,3]])
print(Y,Y.shape)
Z = Y.T #transpose
print(Z,Z.shape)
Z = X* Y # element-wise matrix multiplication
print(Z,Z.shape)
Z = X.dot(Y.T) # matrix multiplication
print(Z,Z.shape)
Z = X**2
print(Z,Z.shape)
Z = 1/X
print(Z,Z.shape)
Z = np.log(X)
print(Z,Z.shape)
Z = X + 1
print(Z,Z.shape)
X = np.array([[2,7,-2],[1,8,3]])
print(X,X.shape)
print(np.max(X))
print(np.max(X,axis=0))
print(np.max(X,axis=1))
print(np.argmax(X))
print(np.argmax(X,axis=0))
print(np.argmax(X,axis=1))
X = np.array([[2,7,-2],[1,8,3]])
print(X,X.shape)
print(np.sum(X))
print(np.sum(X,axis=0))
print(np.sum(X,axis=1))
```
<br>
## 5. Plotting data
<hr>
```
# Visualization library
%matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('png2x','pdf')
import matplotlib.pyplot as plt
x = np.linspace(0,6*np.pi,100)
#print(x)
y = np.sin(x)
plt.figure(1)
plt.plot(x, y,label='sin'.format(i=1))
plt.legend(loc='best')
plt.title('Sin plotting')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
```
<br>
## 6. Control statements
<hr>
### 6.1 Function
```
def f(x):
return x**2
x = 3
y = f(x)
print(y)
def g(x,y):
return x**2, y**3
x = 3
y = 5
u,v = g(x,y)
print(u,v)
```
### 6.2 Logical control statements
```
for i in range(10):
print(i)
i = 0
while (i<10):
i += 1
print(i)
i = True
if i==True:
print(True)
i = False
if i==True:
print(True)
elif i==False:
print(False)
```
<br>
## 7. Vectorization and efficient linear algebra computations
<hr>
### 7.1 No vectorization
```
import time
n = 10**7
x = np.linspace(0,1,n)
y = np.linspace(0,2,n)
start = time.time()
z = 0
for i in range(len(x)):
z += x[i]*y[i]
end = time.time() - start
print(z)
print('Time=',end)
```
### 7.2 Vectorization
```
start = time.time()
z = x.T.dot(y)
end = time.time() - start
print(z)
print('Time=',end)
```
Speed of vectorized codes maybe 2-3 orders of magnitude faster than unvectorized codes.
Vectorized codes (with vectors, matrices) benefit from fast linear algebra libraries (BLAS, LAPACK) and adapted architectures for CPUs, multi-core CPUs, and GPUs.
| github_jupyter |
```
#! pip3 install torch==1.5.0 transformers==3.4.0
#! pip install pickle5
#! pip install datasets
#! pip install faiss-gpu cudatoolkit=10.0 -c pytorch
```
# Train XLM-R Weighted Loss Stategy on sentence translation pairs
In this notebook, you can train the XLM-R model with Weighted Loss Stategy on sentence translation pairs
```
from transformers import Trainer, TrainingArguments
from transformers import AutoModelForSequenceClassification
import json
import pickle5 as pickle
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
import torch.utils.data as data_utils
import torch
import sys
import os
sys.path.append(os.path.dirname((os.path.abspath(''))))
np.random.seed(42)
from src.models.train_text_encoder import Torch_dataset_mono, compute_metrics, WeightedLossTrainer
#from google.colab import drive
#drive.mount('/content/drive')
binary_dataset_path = "/content/drive/MyDrive/CLIR/europarl_data/feature_dataframe.json"
path = "/content/drive/MyDrive/CLIR/europarl_data/europarl_english_german.pkl"
model_used = "xlm-roberta-base"
# read file
with open(binary_dataset_path, 'r') as myfile:
data=myfile.read()
binary_dataset = json.loads(data)
# Load Data
with open(path, 'rb') as f:
data = pickle.load(f)
new_training_set = pd.DataFrame(columns=['source_id', 'target_id', 'text_source', 'text_target', 'Translation'])
current_source_id = list(binary_dataset["source_id"].values())
current_target_id = list(binary_dataset["target_id"].values())
new_training_set["text_source"] = data.iloc[current_source_id,:]["text_source"].reset_index(drop=True)
new_training_set["text_target"] = data.iloc[current_target_id,:]["text_target"].reset_index(drop=True)
new_training_set["source_id"] = current_source_id
new_training_set["target_id"] = current_target_id
new_training_set['Translation'] = new_training_set.apply(lambda row : int(row['source_id'] == row['target_id']), axis = 1)
del binary_dataset
del data
test_size=.05
cutoff = int(test_size*len(new_training_set))
test_dataset = new_training_set.iloc[:cutoff, :]
train_dataset = new_training_set.iloc[cutoff:, :]
train_dataset.head(n=33)
print("Size of training set: {}".format(len(train_dataset)))
print("Size of test set: {}".format(len(test_dataset)))
train_dataset = Torch_dataset_mono(train_dataset)
test_dataset = Torch_dataset_mono(test_dataset)
from transformers import Trainer, TrainingArguments
from transformers import AutoModelForSequenceClassification
save_model_path = "../model/model_downsampled"
save_log_path = "../model/log_downsampled"
model = AutoModelForSequenceClassification.from_pretrained("../model/model_downsampled/checkpoint-12000", num_labels=2)
#transformers.logging.set_verbosity_info()
training_args = TrainingArguments(
output_dir=save_model_path, # output directory
#overwrite_output_dir=True,
num_train_epochs=1, # total number of training epochs
per_device_train_batch_size=11, # batch size per device during training
per_device_eval_batch_size=11, # batch size for evaluation
weight_decay=0.01, # strength of weight decay
warmup_steps=400, # number of warmup steps for learning rate scheduler
logging_dir=save_log_path, # directory for storing logs
logging_steps=10,
evaluation_strategy="steps",
eval_steps=1000,
save_steps=1000
)
trainer = WeightedLossTrainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_dataset, # training dataset
compute_metrics=compute_metrics,
eval_dataset=test_dataset
)
trainer.train("../model/model_downsampled/checkpoint-12000")
```
| github_jupyter |
# Sample, Explore, and Clean Taxifare Dataset
**Learning Objectives**
- Practice querying BigQuery
- Sample from large dataset in a reproducible way
- Practice exploring data using Pandas
- Identify corrupt data and clean accordingly
## Introduction
In this notebook, we will explore a dataset corresponding to taxi rides in New York City to build a Machine Learning model that estimates taxi fares. The idea is to suggest a likely fare to taxi riders so that they are not surprised, and so that they can protest if the charge is much higher than expected. Such a model would also be useful for ride-hailing apps that quote you the trip price in advance.
### Set up environment variables and load necessary libraries
```
PROJECT = 'cloud-training-demos' # Replace with your PROJECT
REGION = 'us-central1' # Choose an available region for Cloud MLE
import os
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
```
Check that the Google BigQuery library is installed and if not, install it.
```
!pip freeze | grep google-cloud-bigquery==1.6.1 || pip install google-cloud-bigquery==1.6.1
```
## View data schema and size
Our dataset is hosted in [BigQuery](https://cloud.google.com/bigquery/): Google's petabyte scale, SQL queryable, fully managed cloud data warehouse. It is a publically available dataset, meaning anyone with a GCP account has access.
1. Click [here](https://console.cloud.google.com/bigquery?project=bigquery-public-data&p=nyc-tlc&d=yellow&t=trips&page=table) to acess the dataset.
2. In the web UI, below the query editor, you will see the schema of the dataset. What fields are available, what does each mean?
3. Click the 'details' tab. How big is the dataset?
## Preview data
Let's see what a few rows of our data looks like. Any cell that starts with `%%bigquery` will be interpreted as a SQL query that is executed on BigQuery, and the result is printed to our notebook.
BigQuery supports [two flavors](https://cloud.google.com/bigquery/docs/reference/standard-sql/migrating-from-legacy-sql#comparison_of_legacy_and_standard_sql) of SQL syntax: legacy SQL and standard SQL. The preferred is standard SQL because it complies with the official SQL:2011 standard. To instruct BigQuery to interpret our syntax as such we start the query with `#standardSQL`.
There are over 1 Billion rows in this dataset and it's 130GB large, so let's retrieve a small sample
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
`nyc-tlc.yellow.trips`
WHERE RAND() < .0000001 -- sample a small fraction of the data
```
### Preview data (alternate way)
Alternatively we can use BigQuery's web UI to execute queries.
1. Open the [web UI](https://console.cloud.google.com/bigquery)
2. Paste the above query minus the `%%bigquery` part into the Query Editor
3. Click the 'Run' button or type 'CTRL + ENTER' to execute the query
Query results will be displayed below the Query editor.
## Sample data repeatably
There's one issue with using `RAND() < N` to sample. It's non-deterministic. Each time you run the query above you'll get a different sample.
Since repeatability is key to data science, let's instead use a hash function (which is deterministic by definition) and then sample the using the modulo operation on the hashed value.
We obtain our hash values using:
`ABS(FARM_FINGERPRINT(CAST(hashkey AS STRING)))`
Working from inside out:
- `CAST()`: Casts hashkey to string because our hash function only works on strings
- `FARM_FINGERPRINT()`: Hashes strings to 64bit integers
- `ABS()`: Takes the absolute value of the integer. This is not strictly neccesary but it makes the following modulo operations more intuitive since we don't have to account for negative remainders.*
The `hashkey` should be:
1. Unrelated to the objective
2. Sufficiently high cardinality
Given these properties we can sample our data repeatably using the modulo operation.
To get a 1% sample:
`WHERE MOD(hashvalue,100) = 0`
To get a *different* 1% sample change the remainder condition, for example:
`WHERE MOD(hashvalue,100) = 55`
To get a 20% sample:
`WHERE MOD(hashvalue,100) < 20` Alternatively: `WHERE MOD(hashvalue,5) = 0`
And so forth...
We'll use `pickup_datetime` as our hash key because it meets our desired properties. If such a column doesn't exist in the data you can synthesize a hashkey by concatenating multiple columns.
Below we sample 1/5000th of the data. The syntax is admittedly less elegant than `RAND() < N`, but now each time you run the query you'll get the same result.
\**Tech note: Taking absolute value doubles the chances of hash collisions but since there are 2^64 possible hash values and less than 2^30 hash keys the collision risk is negligable.*
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
*
FROM
`nyc-tlc.yellow.trips`
WHERE
-- repeatable 1/5000th sample
MOD(ABS(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING))),5000) = 1
```
## Load sample into Pandas dataframe
The advantage of querying BigQuery directly as opposed to the web UI is that we can supplement SQL analysis with Python analysis. A popular Python library for data analysis on structured data is [Pandas](https://pandas.pydata.org/), and the primary data strucure in Pandas is called a DataFrame.
To store BigQuery results in a Pandas DataFrame we have have to query the data with a slightly differently syntax.
1. Import the `google.bigquery` module (alias as `bq`)
2. Store the desired SQL query as a Python string
3. Execute `bq.Query(query_string).execute().result().to_dataframe()` where `query_string` is what you created in the previous step
**This will take about a minute**
*Tip: Use triple quotes for a multi-line string in Python*
*Tip: You can measure execution time of a cell by starting that cell with `%%time`*
```
%%time
from google.cloud import bigquery
bq = bigquery.Client(project=PROJECT)
query_string="""
#standardSQL
SELECT
*
FROM
`nyc-tlc.yellow.trips`
Where
-- repeatable 1/5000th sample
MOD(ABS(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING))),5000) = 1
"""
trips = bq.query(query_string).to_dataframe()
```
## Explore datafame
```
print(type(trips))
trips.head()
```
The Python variable `trips` is now a Pandas DataFrame. The `.head()` function above prints the first 5 rows of a DataFrame.
The rows in the DataFrame may be in a different order than when using `%%bigquery`, but the data is the same.
It would be useful to understand the distribution of each of our columns, which is to say the mean, min, max, standard deviation etc..
A DataFrame's `.describe()` method provides this. By default it only analyzes numeric columns. To include stats about non-numeric column use `describe(include='all')`.
```
trips.describe()
```
## Distribution analysis
Do you notice anything off about the data? Pay attention to `min` and `max`. Latitudes should be between -90 and 90, and longitudes should be between -180 and 180, so clearly some of this data is bad.
Further more some trip fares are negative and some passenger counts are 0 which doesn't seem right. We'll clean this up later.
## Investigate trip distance
Looks like some trip distances are 0 as well, let's investigate this.
```
trips[trips['trip_distance'] == 0][:10] # first 10 rows with trip_distance == 0
```
It appears that trips are being charged substantial fares despite having 0 distance.
Let's graph `trip_distance` vs `fare_amount` using the Pandas [`.plot()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.plot.html) method to corroborate.
```
%matplotlib inline
trips.plot(x ="trip_distance", y ="fare_amount", kind='scatter')
```
It appears that we have a lot of invalid data that is being coded as zero distance and some fare amounts that are definitely illegitimate. Let's remove them from our analysis. We can do this by modifying the BigQuery query to keep only trips longer than zero miles and fare amounts that are at least the minimum cab fare ($2.50).
## Identify correct label
Should we use `fare_amount` or `total_amount` as our label? What's the difference?
To make this clear let's look at some trips that included a toll.
```
trips[trips['tolls_amount'] > 0][:10] # first 10 rows with toll_amount > 0
```
Looking at the samples above, we can see that the total amount reflects fare amount, toll and tip somewhat arbitrarily -- this is because when customers pay cash, the tip is not known. In any case tips are discretionary and shoud not be included in our fare estimation tool.
So, we'll use the sum of `fare_amount` + `tolls_amount` as our label.
## Select useful fields
What fields do you see that may be useful in modeling taxifare? They should be
1. Related to the objective
2. Available at prediction time
**Related to the objective**
For example we know `passenger_count` shouldn't have any affect on fare because fare is calculated by time and distance. Best to eliminate it to reduce the amount of noise in the data and make the job of the ML algorithm easier.
If you're not sure whether a column is related to the objective, err on the side of keeping it and let the ML algorithm figure out whether it's useful or not.
**Available at prediction time**
For example `trip_distance` is certainly related to the objective, but we can't know the value until a trip is completed (depends on the route taken), so it can't be used for prediction.
**We will use the following**
`pickup_datetime`, `pickup_longitude`, `pickup_latitude`, `dropoff_longitude`, and `dropoff_latitude`.
## Clean the data
We need to do some clean-up of the data:
- Filter to latitudes and longitudes that are reasonable for NYC
- We shouldn't fare amounts < 2.50
- Trip distances and passenger counts should be non-zero
- Have the label reflect the sum of fare_amount and tolls_amount
Let's change the BigQuery query appropriately, and only return the fields we'll use in our model.
```
%%bigquery --project $PROJECT
#standardSQL
SELECT
(tolls_amount + fare_amount) AS fare_amount, -- label
pickup_datetime,
pickup_longitude,
pickup_latitude,
dropoff_longitude,
dropoff_latitude
FROM
`nyc-tlc.yellow.trips`
WHERE
-- Clean Data
trip_distance > 0
AND passenger_count > 0
AND fare_amount >= 2.5
AND pickup_longitude > -78
AND pickup_longitude < -70
AND dropoff_longitude > -78
AND dropoff_longitude < -70
AND pickup_latitude > 37
AND pickup_latitude < 45
AND dropoff_latitude > 37
AND dropoff_latitude < 45
-- repeatable 1/5000th sample
AND MOD(ABS(FARM_FINGERPRINT(CAST(pickup_datetime AS STRING))),5000) = 1
```
We now have a repeatable and clean sample we can use for modeling taxi fares.
Copyright 2019 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
# Miller OP
In this notebook the circuit shown in the following schematic will be sized to acheive a certain performance.

Unlike `sym_sizing.ipynb` though, here the performance is obtained directly from the **simulator**, instead
of approxmiating it with
$$ A_{0} \approx - \frac{g_{\mathrm{m},\mathtt{ND12}}}{ g_{\mathrm{ds},\mathtt{ND12}} + g_{\mathrm{ds},\mathtt{PCM22}}}
\cdot \frac{g_{\mathrm{m},\mathtt{PCS}}}{ g_{\mathrm{ds},\mathtt{PCS}} + g_{\mathrm{ds},\mathtt{NCM13}}} $$
```
%matplotlib inline
import os
import numpy as np
import torch as pt
import pandas as pd
import joblib as jl
from functools import partial
from scipy.interpolate import pchip_interpolate, interp1d
from scipy.optimize import minimize
from scipy.stats import norm
from skopt import gp_minimize
from sklearn.preprocessing import MinMaxScaler, minmax_scale
from matplotlib import pyplot as plt
save_results = False # Set True if final AC analysis should be saved to csv
```
## Specification
The following values are considered a priori knowledge.
| Parameter | Specification |
|-----------------------|--------------:|
| $V_{\mathrm{DD}}$ | $1.2\,V$ |
| $V_{\mathrm{SS}}$ | $0.0\,V$ |
| $V_{\mathrm{in,cm}}$ | $0.6\,V$ |
| $V_{\mathrm{out,cm}}$ | $0.6\,V$ |
| $I_{\mathtt{B0}}$ | $10\,\mu A$ |
| $C_{\mathrm{L}}$ | $10\,p F$ |
| $C_{\mathrm{c}}$ | $3\,p F$ |
```
V_DD = 1.2
V_SS = 0.0
V_ICM = 0.6
V_OCM = 0.6
I_B0 = 10e-6
C_L = 10e-12
C_C = 3e-12
```
## Simulator Setup
For evaluating the performance, the circuit is simulated with [PySpice](https://pyspice.fabrice-salvaire.fr/).
The circuit will be sized for $90\,\mathrm{nm}$ devices.
```
import logging
from PySpice.Spice.Netlist import Circuit, SubCircuitFactory
from PySpice.Spice.Library import SpiceLibrary
from PySpice.Unit import *
```
### DUT
The Millor Operational Amplifiere shown in the figure above is implemented as a `subckt`.
```
class MOA(SubCircuitFactory):
NAME = "miller"
NODES = (10, 11, 12, 13, 14, 15) # REF, INP, INN, OUT, GND, VDD
def __init__(self):
super().__init__()
# Biasing Current Mirror
self.MOSFET("NCM11" , 10, 10, 14, 14, model = "nmos")
self.MOSFET("NCM12" , 16, 10, 14, 14, model = "nmos")
# Differential Pair
self.MOSFET("ND11" , 17, 11, 16, 14, model = "nmos")
self.MOSFET("ND12" , 18, 12, 16, 14, model = "nmos")
# PMOS Current Mirrors
self.MOSFET("PCM21" , 17, 17, 15, 15, model = "pmos")
self.MOSFET("PCM22" , 18, 17, 15, 15, model = "pmos")
# Output Stage
self.MOSFET("PCS" , 13, 18, 15, 15, model = "pmos")
self.MOSFET("NCM13" , 13, 10, 14, 14, model = "nmos")
# Compensation
self.C("c", 18, 13, C_C@u_F)
```
**Note**: for other technologies, the corresponding library has to included.
```
spice_lib_130 = SpiceLibrary(f"../lib/130nm_bulk.lib")
netlist_130 = Circuit("moa_tb")
netlist_130.include(spice_lib_130["nmos"])
netlist_130.subcircuit(MOA())
```
### Testbench
An AC-Testbench is setup to analyse the gain, this has to be adjusted for which ever target parameters are of interest.
```
netlist_130.X("moa", "miller", "B", "P", "N", "O", 0, "D")
#moa = list(netlist_130.subcircuits)[0]
i_ref = netlist_130.CurrentSource("ref", 0 , "B", I_B0@u_A)
v_dd = netlist_130.VoltageSource("dd" , "D", 0 , V_DD@u_V)
v_in = netlist_130.VoltageSource("in" , "N", 0 , V_ICM@u_V)
v_ip = netlist_130.SinusoidalVoltageSource( "ip", "P", "E"
, dc_offset=0.0@u_V
, ac_magnitude=-1.0@u_V
, )
e_buf = netlist_130.VoltageControlledVoltageSource("in", "E", 0, "O", 0, 1.0@u_V)
c_l = netlist_130.C("L", "O", 0, C_L@u_F)
```
### Simulation
The `simulate` function takes a data frame with sizing parameters `W` and `L` as columns and each device in the ciruit as row index.
After modiying the corresponding model paramters within the `miller` sub circuit, the previously defined testbench is simulated.
Frequency, Gain and Phase vectors are returned for further processing. Additionallly, the Simulated netlist is returned.
```
def simulate(sizing_data, netlist):
moa = list(netlist.subcircuits)[0]
for device in sizing_data.index:
moa.element(device).width = sizing_data.loc[device].W
moa.element(device).length = sizing_data.loc[device].L
moa.element(device).multiplier = sizing_data.loc[device].M
simulator = netlist.simulator( simulator="ngspice-subprocess"
, temperature=27
, nominal_temperature=27
, )
logging.disable(logging.FATAL)
ac_analysis = simulator.ac( start_frequency = 1.0@u_Hz
, stop_frequency = 1e11@u_Hz
, number_of_points = 10
, variation = "dec"
, )
freq = np.array(ac_analysis.frequency)
gain = ((20 * np.log10(np.absolute(ac_analysis["O"]))) - (20 * np.log10(np.absolute(ac_analysis["P"]))))
phase = (np.angle(ac_analysis["O"], deg=True) - np.angle(ac_analysis["N"], deg=True))
logging.disable(logging.NOTSET)
return (freq, gain, phase, simulator)
```
With the gain and phase obtained from the AC simulation, the DC-gain, cutoff frequency as well as gain- and phase-margin can be determiend in the `performance` function.
```
def performance(freq, gain, phase):
gf = [gain[np.argsort(gain)], freq[np.argsort(gain)]]
pf = [phase[np.argsort(phase)], freq[np.argsort(phase)]]
A0dB = pchip_interpolate(freq, gain, [1.0])
A3dB = A0dB - 3.0
f3dB = pchip_interpolate(*gf, [A3dB])
fug = pchip_interpolate(*gf, [0.0]) if A0dB > 0 else np.ones(1)
fp0 = pchip_interpolate(*pf, [0.0])
PM = pchip_interpolate(freq, phase, [fug]) if A0dB > 0 else np.zeros(1)
GM = pchip_interpolate(freq, gain, [fp0])
performances = { "A0dB" : A0dB.item()
, "f3dB" : f3dB.item()
, "fug" : fug.item()
, "PM" : PM.item()
, "GM" : GM.item()
, }
return performances
```
## Model Setup
A class `PrimitiveDevices` is instantiated for each model type (nmos, pmos). Each object provides a `predict`
function that evaluates the corresponding model and scales inputs and outputs correspondingly.
```
class PrimitiveDevice():
def __init__(self, prefix, params_x, params_y):
self.prefix = prefix
self.params_x = params_x
self.params_y = params_y
self.model = pt.jit.load(f"{self.prefix}/model.pt")
self.model.cpu()
self.model.eval()
self.scale_x = jl.load(f"{self.prefix}/scale.X")
self.scale_y = jl.load(f"{self.prefix}/scale.Y")
def predict(self, X):
with pt.no_grad():
X.fug = np.log10(X.fug.values)
X_ = self.scale_x.transform(X[params_x].values)
Y_ = self.model(pt.from_numpy(np.float32(X_))).numpy()
Y = pd.DataFrame( self.scale_y.inverse_transform(Y_)
, columns=self.params_y )
Y.jd = np.power(10, Y.jd.values)
Y.gdsw = np.power(10, Y.gdsw.values)
return Y #pd.DataFrame(Y, columns=self.params_y)
devices = [ "MNCM11", "MNCM12", "MNCM13" , "MND11", "MND12"
, "MPCM21", "MPCM22", "MPCS" ]
reference_devices = [ "MNCM12", "MNCM13", "MND12", "MPCM22", "MPCS" ]
```
The inputs and outputs of the model, trained in `model_training.ipynb` have to specified again.
```
params_x = ["gmid", "fug", "Vds", "Vbs"]
params_y = ["jd", "L", "gdsw", "Vgs"]
```
Initially the symmetrical amplifier is sized with the models for the $90\,\mathrm{nm}$ technology.
Later this can be changed to any other technology model, yielding similar results.
```
nmos = PrimitiveDevice(f"../models/example/90nm-nmos", params_x, params_y)
pmos = PrimitiveDevice(f"../models/example/90nm-pmos", params_x, params_y)
```
## Sizing Procedure

For simplicity in this example, only the $\frac{g_{\mathrm{m}}}{I_{\mathrm{d}}}$ dependend models
are considered. Therefore, sizing for all devices is expressed in terms of
$\frac{g_{\mathrm{m}}}{I_{\mathrm{d}}}$ and $f_{\mathrm{ug}}$.
$$\gamma_{\mathrm{n,p}} \left ( \left [ \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}}, f_{\mathrm{ug}}, V_{\mathrm{ds}}, V_{\mathrm{bs}} \right ]^{\dagger} \right )
\Rightarrow \left [ L, \frac{I_{\mathrm{d}}}{W}, \frac{g_{\mathrm{ds}}}{W}, V_{\mathrm{gs}} \right ]^{\dagger}
$$
First, the specification, given in the table above is considered,
from which a biasing current ${I_{\mathtt{B1}} = \frac{I_{\mathrm{B0}}}{2}}$ is defined.
This, in turn results in a mirror ratio $M_{\mathrm{n}} = 1 : 1$ of the NMOS current mirror `NCM1`.
The remaining branch current $I_{\mathtt{B2}} = M_{\mathrm{p}} \cdot \frac{I_{\mathtt{B2}}}{2}$ is determined as well.
```
M_N1 = 1
M_N2 = 2
I_B1 = I_B0 * M_N1
I_B2 = I_B0 * M_N2
```
Since the common mode output voltage $V_{\mathrm{out,cm}} = 0.6$ is known,
the sizing procedure starts with the output stage `MPCS` and `MNCM13`:
$$ \gamma_{\mathrm{p}, \mathtt{MPCS}} \left ( \left [ \left ( \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}} \right )_{\mathtt{MPCS}}
, f_{\mathrm{ug}, \mathtt{MPCS}}, (V_{\mathrm{DD}} - V_{\mathrm{out,cm}})
, 0.0 \right ]^{\dagger}
\right ) $$
$$ \gamma_{\mathrm{n}, \mathtt{MNCM13}} \left ( \left [ \left ( \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}} \right )_{\mathtt{MNCM13}}
, f_{\mathrm{ug}, \mathtt{MNCM13}}, V_{\mathrm{out,cm}}, 0.0 \right ]^{\dagger}
\right ) $$
The gate voltage $V_{\mathrm{gs}, \mathtt{MPCS}}$ helps guiding the sizing for the differential pair,
as well as the PMOS Current Mirror `MPCM2`, which is considered first:
$$ \gamma_{\mathrm{p}, \mathtt{MPCM22}} \left ( \left [ \left ( \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}} \right )_{\mathtt{MPCM22}}
, f_{\mathrm{ug}, \mathtt{MPCM22}}
, (V_{\mathrm{DD}} - V_{\mathrm{gs}, \mathtt{MPCS}})
, 0.0 \right ]^{\dagger}
\right ) $$
Sizing the differential pair requires, _guessing_ $V_{\mathrm{x}} = 0.23\,\mathrm{V}$ which is done by considering
the fact that 3 devices are stacked and `MNCM1` merely serves as biasing. Therefore:
$$ \gamma_{\mathrm{n}, \mathtt{MND12}} \left ( \left [ \left ( \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}} \right )_{\mathtt{MND12}}
, f_{\mathrm{ug}, \mathtt{MND12}}
, (V_{\mathrm{DD}} - V_{\mathrm{gs}, \mathtt{MPCS}} - V_{\mathrm{x}})
, - V_{\mathrm{x}} \right ]^{\dagger}
\right ) $$
Subsequently, the biasing current mirror `MNCM1` is sized:
$$ \gamma_{\mathrm{n}, \mathtt{MNCM12}} \left ( \left [ \left ( \frac{g_{\mathrm{m}}}{I_{\mathrm{d}}} \right )_{\mathtt{MNCM12}}
, f_{\mathrm{ug}, \mathtt{MNCM12}}
, V_{\mathrm{x}}
, 0.0 \right ]^{\dagger}
\right ) $$
With these five function calls, the sizing of the entire circuit is expressed in terms of eight electrical characteristics.
The following function `miller_sizing` takes a `dict` with keys for each _reference device_ $\in$
`reference_devices = [ "MNCM12", "MNCM13", "MND12", "MPCM22", "MPCS" ]` and corresponding, desired characteristics.
The obtained sizing for each device is propageted to related devices in the same building block and a new `dict` with sizing information is returned.
```
def miller_sizing( gmid_pcs, gmid_ncm13, gmid_pcm22, gmid_nd12, gmid_ncm12
, fug_pcs, fug_ncm13, fug_pcm22, fug_nd12, fug_ncm12 ):
ec = {}
## Common Source Output Stage
input_pcs = pd.DataFrame( np.array([[gmid_pcs, fug_pcs, (V_DD - V_OCM), 0.0]])
, columns=params_x )
ec["MPCS"] = pmos.predict(input_pcs).join(input_pcs)
ec["MPCS"]["W"] = I_B2 / ec["MPCS"].jd
ec["MPCS"]["M"] = 1
## NMOS Output Stage
input_ncm13 = pd.DataFrame( np.array([[gmid_ncm13, fug_ncm13, V_OCM, 0.0]])
, columns=params_x )
ec["MNCM13"] = nmos.predict(input_ncm13).join(input_ncm13)
ec["MNCM13"]["W"] = I_B2 / ec["MNCM13"].jd / M_N2
ec["MNCM13"]["M"] = M_N2
## PMOS Current Mirror MPCM2:
V_GS = ec["MPCS"].Vgs.values[0]
input_pcm22 = pd.DataFrame( np.array([[ gmid_pcm22, fug_pcm22, (V_DD - V_GS), 0.0]])
, columns=params_x )
ec["MPCM22"] = pmos.predict(input_pcm22).join(input_pcm22)
ec["MPCM22"]["W"] = (I_B1 / 2) / ec["MPCM22"].jd
ec["MPCM22"]["M"] = 1
ec["MPCM21"] = ec["MPCM22"]
## NMOS Differential Pair NDP1:
V_X = 0.20
input_nd12 = pd.DataFrame( np.array([[gmid_nd12, fug_nd12, (V_DD - V_GS - V_X), -V_X]])
, columns=params_x )
ec["MND12"] = nmos.predict(input_nd12).join(input_nd12)
ec["MND12"]["W"] = (I_B1 / 2) / ec["MND12"].jd
ec["MND12"]["M"] = 1
ec["MND11"] = ec["MND12"]
## NMOS Current Mirror NCM12:
# As part of MNCM1
#ec["MNCM12"] = ec["MNCM13"].copy()
#ec["MNCM11"] = ec["MNCM13"].copy()
#ec["MNCM12"].M = M_N1
#ec["MNCM11"].M = 1
# As a spearate building block
input_ncm12 = pd.DataFrame( np.array([[gmid_ncm12, fug_ncm12, V_X, 0.0]])
, columns=params_x )
ec["MNCM12"] = nmos.predict(input_ncm12).join(input_ncm12)
ec["MNCM12"]["W"] = I_B1 / ec["MNCM12"].jd / M_N1
ec["MNCM12"]["M"] = M_N1
ec["MNCM11"] = ec["MNCM12"].copy()
ec["MNCM11"]["M"] = 1
## Calculate/Approximate Operating point Parameters
for dev,val in ec.items():
val["gds"] = val.gdsw * val.W
val["id"] = val.jd * val.W
val["gm"] = val.gmid * val.id
return ec
```
The operating points for each device are approximated for given electrical characteristics of each _reference device_ in the circuit.
```
def miller_op_approx( gmid_pcs, gmid_ncm13, gmid_pcm22, gmid_nd12, gmid_ncm12
, fug_pcs, fug_ncm13, fug_pcm22, fug_nd12, fug_ncm12 ):
sizing = miller_sizing( gmid_pcs, gmid_ncm13, gmid_pcm22, gmid_nd12, gmid_ncm12
, fug_pcs, fug_ncm13, fug_pcm22, fug_nd12, fug_ncm12 )
sizing_data = pd.concat(sizing.values(), names=sizing.keys())
sizing_data.index = sizing.keys()
return sizing_data
```
The objective function for reaching a certain gain is formulated, such that it only depends on the individual $f_{\mathrm{ug}}$s of the reference devices.
This limits the search space by halfing the number of variables in the optimization problem, while keeping it portable across technologies,
since $\frac{g_{\mathrm{m}}}{I_{\mathrm{d}}}$, unlike the speed, stays the same.
$$ \underset{f_{\mathrm{ug,moa}}}{\arg\min} ~ | A_{0,\mathrm{moa}} - A_{0, \mathrm{target}} | $$
Where
$$ f_{\mathrm{ug,moa}} =
\left[ f_{\mathrm{ug},\mathtt{NDP12}}
, f_{\mathrm{ug},\mathtt{NCM12}}
, f_{\mathrm{ug},\mathtt{PCS}}
, f_{\mathrm{ug},\mathtt{NCM13}}
, f_{\mathrm{ug},\mathtt{PCM22}}
\right]^{\dagger} $$
```
gmid_pcs_fix = 10.0
gmid_ncm12_fix = 10.0
gmid_ncm13_fix = 10.0
gmid_pcm22_fix = 10.0
gmid_nd12_fix = 12.0
fug_pcs_prd = np.nan
fug_ncm13_prd = np.nan
fug_pcm22_prd = np.nan
fug_nd12_prd = np.nan
fug_ncm12_prd = np.nan
```
Specify desired target i.e. $A_{0,\mathrm{target}} \geq 80 \, \mathrm{dB}$.
```
A0dB_target = 84
```
Since it's an optimization (minimization) problem, the absollute difference $| A_{0,\mathrm{target}} - A_{0,\mathrm{moa}} |$ is returned.
```
def miller_gain_target( netlist
, gmid_pcs, gmid_ncm13, gmid_pcm22, gmid_nd12, gmid_ncm12
, fug_pcs, fug_ncm13, fug_pcm22, fug_nd12, fug_ncm12 ):
fug = np.power(10, np.array([fug_pcs, fug_ncm13, fug_pcm22, fug_nd12, fug_ncm12]))
apop = miller_op_approx(gmid_pcs, gmid_ncm13, gmid_pcm22, gmid_nd12, gmid_ncm12, *fug)
freq, gain, phase, _ = simulate(apop, netlist)
perf = performance(freq, gain, phase)
return np.abs(A0dB_target - perf["A0dB"])
```
Apply fixed parameters, to reduce the search space:
```
miller_target = partial( miller_gain_target
, netlist_130
, gmid_pcs_fix, gmid_ncm13_fix
, gmid_pcm22_fix, gmid_nd12_fix
, gmid_ncm12_fix )
```
Subsequently a bayesian surrogate model for the gain of the entire circuit is created.
```
miller_gp = gp_minimize( lambda f: miller_target(*f) # Gain Target Function
, [ (6.0, 11.0) # Bounds: log10(fug_ncm12)
, (6.0, 11.0) # Bounds: log10(fug_pcs)
, (6.0, 11.0) # Bounds: log10(fug_ncm12)
, (6.0, 11.0) # Bounds: log10(fug_ncm13)
, (6.0, 11.0) ] # Bounds: log10(fug_pcm12)
, acq_func = "PI" # EI, PI, LCB
, n_calls = 50 # Maximum number of simulations
, n_random_starts = 20 # Number of random init points
, noise = 1e-3 # Prediction Noise (Uncertainty)
, random_state = 666 # RNG seed
, n_jobs = os.cpu_count() # Number of Cores
, )
_, std_moa = miller_gp.models[-1].predict(miller_gp.space.transform([miller_gp.x]), return_std=True)
σ_moa = std_moa[0]
μ_moa = (A0dB_target - miller_gp.fun)
miller_dist = norm.pdf(np.arange(60,100,0.01), μ_moa, σ_moa)
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(16,6))
ax1.plot( np.array(list(range(miller_gp.func_vals.size))) + 1
, np.sort(miller_gp.func_vals)[::-1]
, label=f"$A_{{0}}$"
, )
ax1.axhline( y=miller_gp.fun
, ls="dashed"
, color="tab:green"
, label=f"Final Loss: ${miller_gp.fun:.2f}$"
, )
ax1.set_ylabel(f"$| A_{{0}} - A_{{0,target}} |$")
ax1.set_xlabel("Number of Simulations")
ax1.set_title("Convergence")
ax1.set_yscale("log")
ax1.legend()
ax1.grid("on")
gain_range = np.arange(60,100,0.01)
ax2.plot(gain_range, miller_dist)
ax2.axvline( x=A0dB_target
, ls="dashed"
, color="tab:red"
, label=f"$A_{{0,target}} = {A0dB_target}$ dB"
, )
ax2.axvline( x=np.abs(A0dB_target - miller_gp.fun)
, ls="dashed"
, color="tab:blue"
, label=f"$A_{{0}} = {np.abs(A0dB_target - miller_gp.fun):.2f}$ dB"
, )
px = gain_range[np.logical_and( gain_range >= (μ_moa - σ_moa * 3)
, gain_range <= (μ_moa + σ_moa * 3))]
plt.fill_between( gain_range,miller_dist
, where = (gain_range >= (μ_moa - σ_moa))
& (gain_range <= (μ_moa + σ_moa))
, color='tab:blue'
, alpha=0.3
, )
ax2.set_ylabel(f"PDF")
ax2.set_xlabel("Gain [dB]")
ax2.set_title("Gain")
ax2.legend()
ax2.grid("on")
```
Retrieve the function arguments, minimizing the target:
```
fug_pcs_prd, fug_ncm13_prd, fug_pcm22_prd, fug_nd12_prd, fug_ncm12_prd = np.power(10, miller_gp.x)
```
## Evaluation
Finally, the obtianed characteristics are simulated one more time to verify the performance.
```
apop_130 = miller_op_approx( gmid_pcs_fix, gmid_ncm13_fix, gmid_pcm22_fix, gmid_nd12_fix, gmid_ncm12_fix
, fug_pcs_prd, fug_ncm13_prd, fug_pcm22_prd, fug_nd12_prd, fug_ncm12_prd )
freq_130, gain_130, phase_130, _ = simulate(apop_130, netlist_130)
perf_130 = performance(freq_130, gain_130, phase_130)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(10,10))
ax1.plot(freq_130, gain_130, label="Simulated Gain")
ax1.axhline( y=perf_130["A0dB"]
, color="tab:red"
, ls="dashed"
, label=f"$A_{{0}} = {perf_130['A0dB']:.2f}$ dB")
ax1.axhline(y=perf_130["GM"]
, color="tab:red"
, ls="dotted"
, label=f"GM $= {perf_130['GM']:.2f}$ dB")
ax1.axvline( x=perf_130["f3dB"]
, color="tab:green"
, ls="dashed"
, label=f"$f_{{cutoff}} = {perf_130['f3dB']:.2f}$ Hz")
ax1.axvline( x=perf_130["fug"]
, color="tab:green"
, ls="dotted"
, label=f"$f_{{0}} = {perf_130['fug']:.2e}$ Hz")
ax1.set_title("Gain")
ax1.set_xscale("log")
ax1.set_ylabel("Gain [dB]")
ax1.legend()
ax1.grid("on")
ax2.plot(freq_130, phase_130, label="Simulated Phase")
ax2.axhline( y=perf_130["PM"]
, color="tab:red"
, ls="dotted"
, label=f"PM $= {perf_130['PM']:.2f}$ deg")
ax2.axvline( x=perf_130["fug"]
, color="tab:green"
, ls="dotted"
, label=f"$f_{{0}} = {perf_130['fug']:.2e}$ Hz")
ax2.set_title("Phase")
ax2.set_xscale("log")
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("Phase [deg]")
ax2.legend()
ax2.grid("on")
```
This performance is achieved with the following sizing:
```
apop_130.fug = np.power(10, apop_130.fug)
apop_130[["W", "L", "gmid", "fug"]]
```
### Technology Migration
The previously obtained electrical parameters can be used to size the same circuit for a different technology node.
Simply converting these $\frac{g_{\mathrm{m}}}{I_{\mathrm{d}}}$ and $f_{\mathrm{ug}}$ combinations to corresponding sizing
parameters and running the simulation again yields comparable results.
```
spice_lib_90 = SpiceLibrary(f"../lib/90nm_bulk.lib")
netlist_90 = Circuit("moa_tb")
netlist_90.include(spice_lib_90["nmos"])
netlist_90.subcircuit(MOA())
netlist_90.X("moa", "miller", "B", "P", "N", "O", 0, "D")
i_ref = netlist_90.CurrentSource("ref", 0 , "B", I_B0@u_A)
v_dd = netlist_90.VoltageSource("dd" , "D", 0 , V_DD@u_V)
v_in = netlist_90.VoltageSource("in" , "N", 0 , V_ICM@u_V)
v_ip = netlist_90.SinusoidalVoltageSource( "ip", "P", "E"
, dc_offset=0.0@u_V
, ac_magnitude=-1.0@u_V
, )
e_buf = netlist_90.VoltageControlledVoltageSource("in", "E", 0, "O", 0, 1.0@u_V)
c_l = netlist_90.C("L", "O", 0, C_L@u_F)
nmos = PrimitiveDevice(f"../models/example/90nm-nmos", params_x, params_y)
pmos = PrimitiveDevice(f"../models/example/90nm-pmos", params_x, params_y)
apop_90 = miller_op_approx( gmid_pcs_fix, gmid_ncm13_fix, gmid_pcm22_fix, gmid_nd12_fix, gmid_ncm12_fix
, fug_pcs_prd, fug_ncm13_prd, fug_pcm22_prd, fug_nd12_prd, fug_ncm12_prd )
freq_90, gain_90, phase_90, _ = simulate(apop_90, netlist_90)
perf_90 = performance(freq_90, gain_90, phase_90)
spice_lib_45 = SpiceLibrary(f"../lib/45nm_bulk.lib")
netlist_45 = Circuit("moa_tb")
netlist_45.include(spice_lib_45["nmos"])
netlist_45.subcircuit(MOA())
netlist_45.X("moa", "miller", "B", "P", "N", "O", 0, "D")
i_ref = netlist_45.CurrentSource("ref", 0 , "B", I_B0@u_A)
v_dd = netlist_45.VoltageSource("dd" , "D", 0 , V_DD@u_V)
v_in = netlist_45.VoltageSource("in" , "N", 0 , V_ICM@u_V)
v_ip = netlist_45.SinusoidalVoltageSource( "ip", "P", "E"
, dc_offset=0.0@u_V
, ac_magnitude=-1.0@u_V
, )
e_buf = netlist_45.VoltageControlledVoltageSource("in", "E", 0, "O", 0, 1.0@u_V)
c_l = netlist_45.C("L", "O", 0, C_L@u_F)
nmos = PrimitiveDevice(f"../models/example/45nm-nmos", params_x, params_y)
pmos = PrimitiveDevice(f"../models/example/45nm-pmos", params_x, params_y)
apop_45 = miller_op_approx( gmid_pcs_fix, gmid_ncm13_fix, gmid_pcm22_fix, gmid_nd12_fix, gmid_ncm12_fix
, fug_pcs_prd, fug_ncm13_prd, fug_pcm22_prd, fug_nd12_prd, fug_ncm12_prd )
freq_45, gain_45, phase_45, _ = simulate(apop_45, netlist_45)
perf_45 = performance(freq_45, gain_45, phase_45)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(10,8))
ax1.plot(freq_130, gain_130, label="Simulated Gain $130$ nm")
ax1.plot(freq_90, gain_90, label="Simulated Gain $90$ nm")
ax1.plot(freq_45, gain_45, label="Simulated Gain $45$ nm")
ax1.axhline( y=A0dB_target
, color="tab:red"
, ls="dashed"
, label=f"$A_{{0, target}} = {A0dB_target}$ dB")
ax1.axvline( x=perf_130["f3dB"]
, color="tab:blue"
, ls="dashed"
, label=f"$f_{{cutoff, 130nm}} = {perf_130['f3dB']:.2f}$ Hz")
ax1.axvline( x=perf_130["fug"]
, color="tab:blue"
, ls="dotted"
, label=f"$f_{{0,130nm}} = {perf_130['fug']:.2e}$ Hz")
ax1.axvline( x=perf_90["f3dB"]
, color="tab:orange"
, ls="dashed"
, label=f"$f_{{cutoff, 90nm}} = {perf_90['f3dB']:.2f}$ Hz")
ax1.axvline( x=perf_90["fug"]
, color="tab:orange"
, ls="dotted"
, label=f"$f_{{0, 90nm}} = {perf_90['fug']:.2e}$ Hz")
ax1.axvline( x=perf_45["f3dB"]
, color="tab:green"
, ls="dashed"
, label=f"$f_{{cutoff, 45nm}} = {perf_45['f3dB']:.2f}$ Hz")
ax1.axvline( x=perf_45["fug"]
, color="tab:green"
, ls="dotted"
, label=f"$f_{{0, 45nm}} = {perf_45['fug']:.2e}$ Hz")
ax1.set_title("Gain Comparison")
ax1.set_xscale("log")
ax1.set_ylabel("Gain [dB]")
ax1.legend(bbox_to_anchor=(1,1), loc="upper left")
ax1.grid("on")
ax2.plot(freq_90, phase_130, label="Simulated Phase $130$ nm")
ax2.plot(freq_90, phase_90, label="Simulated Phase $90$ nm")
ax2.plot(freq_90, phase_45, label="Simulated Phase $45$ nm")
ax2.axhline( y=perf_130["PM"]
, color="tab:blue"
, ls="dotted"
, label=f"$PM_{{130nm}} = {perf_130['PM']:.2f}$ deg")
ax2.axvline( x=perf_130["fug"]
, color="tab:blue"
, ls="dashed"
, label=f"$f_{{0, 130nm}} = {perf_130['fug']:.2e}$ Hz")
ax2.axhline( y=perf_90["PM"]
, color="tab:orange"
, ls="dotted"
, label=f"$PM_{{90nm}} = {perf_90['PM']:.2f}$ deg")
ax2.axvline( x=perf_90["fug"]
, color="tab:orange"
, ls="dashed"
, label=f"$f_{{0, 90nm}} = {perf_90['fug']:.2e}$ Hz")
ax2.axhline( y=perf_45["PM"]
, color="tab:green"
, ls="dotted"
, label=f"$PM_{{45nm}} = {perf_45['PM']:.2f}$ deg")
ax2.axvline( x=perf_45["fug"]
, color="tab:green"
, ls="dashed"
, label=f"$f_{{0, 45nm}} = {perf_45['fug']:.2e}$ Hz")
ax2.set_title("Phase Comparison")
ax2.set_xscale("log")
ax2.set_xlabel("Frequency [Hz]")
ax2.set_ylabel("Phase [deg]")
ax2.legend(bbox_to_anchor=(1,1), loc="upper left")
ax2.grid("on")
if save_results:
pd.DataFrame({"freq": freq_130, "gain": gain_130, "phase": phase_130}).to_csv("./sim130.csv")
pd.DataFrame({"freq": freq_90, "gain": gain_90, "phase": phase_90}).to_csv("./sim90.csv")
pd.DataFrame({"freq": freq_45, "gain": gain_45, "phase": phase_45}).to_csv("./sim45.csv")
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from descartes import PolygonPatch
import json
import os
from pprint import pprint
import shapely.geometry
import shapely.affinity
%matplotlib inline
with open('../evaluations/test-bichen.json') as f:
data = json.load(f)
# pprint(data)
class RotatedRect:
def __init__(self, cx, cy, w, h, angle):
self.cx = cx
self.cy = cy
self.w = w
self.h = h
self.angle = angle
def get_contour(self):
w = self.w
h = self.h
c = shapely.geometry.box(-w/2.0, -h/2.0, w/2.0, h/2.0)
rc = shapely.affinity.rotate(c, self.angle)
return shapely.affinity.translate(rc, self.cx, self.cy)
def intersection(self, other):
return self.get_contour().intersection(other.get_contour())
def union(self, other):
return self.get_contour().union(other.get_contour())
r1 = RotatedRect(10, 15, 15, 10, 70)
r2 = RotatedRect(15, 15, 20, 10, 0)
fig = pyplot.figure(1, figsize=(10, 4))
ax = fig.add_subplot(121)
ax.set_xlim(0, 30)
ax.set_ylim(0, 30)
ax.add_patch(PolygonPatch(r1.get_contour(), fc='#990000', alpha=0.7))
ax.add_patch(PolygonPatch(r2.get_contour(), fc='#000099', alpha=0.7))
ax.add_patch(PolygonPatch(r1.intersection(r2), fc='#009900', alpha=1))
pyplot.show()
class Evaluation():
def __init__(self, json_data):
self.frames = dict()
frames_loaded = [Frame(frame) for frame in json_data["frames"]]
self.filenames = [frame.filename for frame in frames_loaded]
filenames = set(self.filenames)
for frame in frames_loaded:
if frame.filename in filenames:
filenames.remove(frame.filename)
self.frames[frame.filename] = frame
def get_bounding_boxes(self):
bounding_boxes = []
for filename in self.filenames:
bounding_boxes.extend(self.get_frame(filename).bounding_boxes)
return bounding_boxes
def get_frame(self, filename):
return self.frames[filename]
def false_positives(self, other):
false_positives_dict = dict()
for filename in self.filenames:
false_positives_dict[filename] = self.get_frame(filename).false_positives(other.get_frame(filename))
return false_positives_dict
def false_negatives(self, other):
false_negatives_dict = dict()
for filename in self.filenames:
false_negatives_dict[filename] = self.get_frame(filename).false_negatives(other.get_frame(filename))
return false_negatives_dict
def IoU(self, other):
IoU_dict = dict()
for filename in self.filenames:
IoU_dict[filename] = self.get_frame(filename).IoU(other.get_frame(filename))
return IoU_dict
def total_time_elapsed(self):
time_dict = dict()
for filename in self.filenames:
time_dict[filename] = self.get_frame(filename).time_elapsed
return time_dict
def _3D_time_elapsed(self):
_3D_time_dict = dict()
for filename in self.filenames:
_3D_time_dict[filename] = self.get_frame(filename)._3D_time_elapsed
return _3D_time_dict
class GroundTruth(Evaluation):
def __init__(self, json_data):
Evaluation.__init__(self, json_data)
def get_bounding_boxes(self):
return Evaluation.get_bounding_boxes(self)
class Frame():
def __init__(self, json_data):
self._3D_time_elapsed = json_data['_3D_time_elapsed']
self.add_box_count = json_data['add_box_count']
self.camera_angle = json_data['camera_angle']
self.delete_count = json_data['delete_count']
self.filename = json_data['filename']
self.label_count = json_data['label_count']
self.resize_count = json_data['resize_count']
self.rotate_camera_count = json_data['rotate_camera_count']
self.rotate_count = json_data['rotate_count']
self.time_elapsed = json_data['time_elapsed']
self.translate_count = json_data['translate_count']
self.bounding_boxes = [BoundingBox(box) for box in json_data['bounding_boxes']]
self.epsilon = 1e-8
def IoU(self, other):
IoUs = []
remaining_bounding_boxes = set(self.bounding_boxes)
for bounding_box in other.bounding_boxes:
if len(remaining_bounding_boxes):
closest_bounding_box = sorted(list(remaining_bounding_boxes), key=lambda box: bounding_box.IoU(box), reverse=True)[0]
closest_IoU = closest_bounding_box.IoU(bounding_box)
if closest_IoU > self.epsilon:
IoUs.append(closest_IoU)
remaining_bounding_boxes.remove(closest_bounding_box)
return IoUs
def false_positives(self, other):
remaining_bounding_boxes = set(self.bounding_boxes)
for bounding_box in other.bounding_boxes:
if len(remaining_bounding_boxes):
closest_bounding_box = sorted(list(remaining_bounding_boxes), key=lambda box: bounding_box.IoU(box), reverse=True)[0]
closest_IoU = closest_bounding_box.IoU(bounding_box)
if closest_IoU > self.epsilon:
remaining_bounding_boxes.remove(closest_bounding_box)
return len(remaining_bounding_boxes)
def false_negatives(self, other):
remaining_bounding_boxes = set(other.bounding_boxes)
for bounding_box in self.bounding_boxes:
if len(remaining_bounding_boxes):
closest_bounding_box = sorted(list(remaining_bounding_boxes), key=lambda box: bounding_box.IoU(box), reverse=True)[0]
closest_IoU = closest_bounding_box.IoU(bounding_box)
if closest_IoU > self.epsilon:
remaining_bounding_boxes.remove(closest_bounding_box)
return len(remaining_bounding_boxes)
class BoundingBox():
def __init__(self, json_data):
# orientation in radians
self.angle = json_data['angle']
self.center = json_data['center']
self.length = json_data['length']
self.width = json_data['width']
self.object_id = json_data['object_id']
def IoU(self, other):
offset = (max([self.length, self.width, other.length, other.width])
+ max([0, -self.center['x'], -self.center['y'], -other.center['x'], -other.center['y']]))
r1 = RotatedRect(self.center['x'], self.center['y'], self.width, self.length, self.angle / (2*np.pi) * 360)
r2 = RotatedRect(other.center['x'], other.center['y'], other.width, other.length, other.angle / (2*np.pi) * 360)
intersection = r1.intersection(r2).area
union = r1.union(r2).area
return intersection / union
class EfficiencyTest():
def __init__(self):
self.evaluations = []
for file in os.listdir("../evaluations/"):
if file.endswith(".json") and file != "ground_truth.json":
with open('../evaluations/' + file) as f:
print(file)
data = json.load(f)
evaluation = Evaluation(data)
self.evaluations.append(evaluation)
with open('../evaluations/ground_truth.json') as f:
data = json.load(f)
self.ground_truth = GroundTruth(data)
self.IoUs = dict()
for evaluation in self.evaluations:
IoU = evaluation.IoU(self.ground_truth)
for key in IoU.keys():
if key in self.IoUs:
self.IoUs[key].extend(IoU[key])
else:
self.IoUs[key] = IoU[key]
self.time_elapsed = dict()
for evaluation in self.evaluations:
time_elapsed = evaluation.total_time_elapsed()
for key in time_elapsed.keys():
if key in self.time_elapsed:
self.time_elapsed[key].append(time_elapsed[key])
else:
self.time_elapsed[key] = [time_elapsed[key]]
self._3D_time_elapsed = dict()
for evaluation in self.evaluations:
time_elapsed = evaluation._3D_time_elapsed()
for key in time_elapsed.keys():
if key in self._3D_time_elapsed:
self._3D_time_elapsed[key].append(time_elapsed[key])
else:
self._3D_time_elapsed[key] = [time_elapsed[key]]
def total_bounding_boxes(self):
bounding_boxes = []
for evaluation in self.evaluations:
bounding_boxes.extend(evaluation.get_bounding_boxes())
return len(bounding_boxes)
def get_false_positives(self):
self.false_positives = dict()
for evaluation in self.evaluations:
false_positives = evaluation.false_positives(self.ground_truth)
for key in false_positives.keys():
if key in self.false_positives:
self.false_positives[key].append(false_positives[key])
else:
self.false_positives[key] = [false_positives[key]]
false_positives = []
for key in self.false_positives:
false_positives.extend(self.false_positives[key])
return np.sum(np.array(false_positives))
def get_false_negatives(self):
self.false_negatives = dict()
for evaluation in self.evaluations:
false_negatives = evaluation.false_negatives(self.ground_truth)
for key in false_negatives.keys():
if key in self.false_negatives:
self.false_negatives[key].append(false_negatives[key])
else:
self.false_negatives[key] = [false_negatives[key]]
false_negatives = []
for key in self.false_negatives:
false_negatives.extend(self.false_negatives[key])
return np.sum(np.array(false_negatives))
def num_bounding_boxes(self):
IoUs = []
for key in self.IoUs:
IoUs.extend(self.IoUs[key])
return len(IoUs)
def total_time_elapsed(self):
times = []
for key in self.time_elapsed:
times.extend(self.time_elapsed[key])
return np.sum(np.array(times))
def total_3D_time_elapsed(self):
times = []
for key in self._3D_time_elapsed.keys():
times.extend(self._3D_time_elapsed[key])
return np.sum(np.array(times))
def bounding_boxes_per_time(self):
times = []
for key in self.time_elapsed:
times.extend(self.time_elapsed[key])
return self.num_bounding_boxes() / np.sum(np.array(times))
def IoU(self):
return self.IoUs
def average_IoU(self):
IoUs = []
for key in self.IoUs:
IoUs.extend(self.IoUs[key])
return np.mean(np.array(IoUs))
def sd_IoU(self):
IoUs = []
for key in self.IoUs:
IoUs.extend(self.IoUs[key])
return np.std(np.array(IoUs))
efficiency_test = EfficiencyTest()
efficiency_test.average_IoU()
efficiency_test.sd_IoU()
efficiency_test.total_time_elapsed()
efficiency_test.total_3D_time_elapsed()
efficiency_test.total_3D_time_elapsed() / efficiency_test.total_time_elapsed()
efficiency_test.bounding_boxes_per_time() * 60
efficiency_test.num_bounding_boxes()
efficiency_test.get_false_positives()
efficiency_test.get_false_negatives()
efficiency_test.false_negatives
efficiency_test.false_positives
efficiency_test.total_bounding_boxes()
len(efficiency_test.ground_truth.get_bounding_boxes())
for filename in efficiency_test.ground_truth.filenames:
print(len(efficiency_test.ground_truth.get_frame(filename).bounding_boxes))
16 + 13 + 6 + 14 + 15 + 13
77 * 4
12 + 26 + 16 + 14
68 * 3
204 + 308
```
| github_jupyter |
### Rotation Matrices
```
import numpy as np
from math import cos, sin, radians
import matplotlib.pyplot as plt
# 2D Rotation Matrix: [cos(theta) -sin(theta)
# sin(theta) cos(theta)]
def rotation_matrix_2d(theta):
"""
theta: angle in degrees
Returns the 2D matrix that performs in-plane rotation by angle
theta about origin.
This means, multiplication with this matrix will move a point
to a new location.
The angle between
the line joining original point with origin
and
the line joining new point with origin
is theta.
"""
return np.array([[cos(radians(theta)), -sin(radians(theta))],
[sin(radians(theta)), cos(radians(theta))]])
```
#### Rotation by angle θ in 3D space
```
def rotation_matrix_3d(theta, axis):
"""
theta: angle in degrees
axis: Axis of rotation, can be 0, 1 or 2 corresponding to x, y or z
axis respectively.
Returns the matrix that will rotate a point in 3D space about the
chosen axis of rotation by angle theta degrees.
"""
if axis == 0:
return np.array([[0, 0, 1],
[cos(radians(theta)), -sin(radians(theta)), 0],
[sin(radians(theta)), cos(radians(theta)), 0]])
elif axis == 1:
return np.array([[cos(radians(theta)), 0, -sin(radians(theta))],
[0, 1, 0],
[sin(radians(theta)), 0, cos(radians(theta))]])
elif axis == 2:
return np.array([[cos(radians(theta)), -sin(radians(theta)), 0],
[sin(radians(theta)), cos(radians(theta)), 0],
[0, 0, 1]])
# Let us consider the vector [1, 0, 0]
import math
u = np.array([1, 0, 0])
u = u.reshape((3, 1)) #Reshape it to represent a row vector
print("Original vector\n{}\n".format(u))
# Rotate it by 30 degrees around Z Axis
angle_of_rotation = 30.0
R = rotation_matrix_3d(angle_of_rotation, 2)
print("Rotation matrix for {} degrees around Z Axis is\n{}\n".\
format(angle_of_rotation, R))
v = np.matmul(R, u)
print("Vector after rotation\n{}".format(v))
def get_angle(u, v):
"""
Computes angle between two vectors using arc cosine
"""
u = np.squeeze(u)
v = np.squeeze(v)
assert len(u.shape) == 1 and len(v.shape) == 1
return math.degrees(math.acos((np.dot(u, v)
/ (np.linalg.norm(u) *
np.linalg.norm(v)))))
# Angle between u and v
angle_u_v = get_angle(u, v)
print("Angle between the two vectors {}".format(angle_u_v))
assert np.allclose(angle_u_v, angle_of_rotation)
# Python provides powerful libraries to plot
# vectors and curves and surfaces in 3D. Some
# examples.
def plot_vector(ax, row_vector, color="cyan"):
"""
Plot a 3D vector in specified color
"""
assert len(row_vector == 3)
v = np.squeeze(row_vector)
ax.quiver(0, 0, 0, row_vector[0], row_vector[1],
row_vector[2], color=color)
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.set_xlim([-2, 2])
ax.set_ylim([-2, 2])
ax.set_zlim([-2, 2])
# Let us now visualise the vectors in 3D
u = np.array([1, 1, 1])
#Reshape to represent as a row vector
u_row = u.reshape((3, 1))
plot_vector(ax, u_row, color="blue")
# Rotate it 45 degrees around Z axis
R = rotation_matrix_3d(45, 2)
v_row = np.matmul(R, u_row)
plot_vector(ax, v_row, color="green")
# Rotate again by 45 around X axis
R = rotation_matrix_3d(45, 0)
w_row = np.matmul(R, u_row)
plot_vector(ax, w_row, color="red")
plt.show()
```
#### Orthogonality of rotation matrices
```
R_30 = rotation_matrix_2d(30)
print("Matrix to rotate in-plane by 30 degrees"
"about origin:\n{}".format(R_30))
# Inverse of rotation matrix is same as the transpose, this
# is orthogonality.
# np.allclose is used to determine if all elements are equal
# within tolerance. The inv(erse) and transpose of rotation
# matrix should be equal (due to floating point errors may
# not be perfectly equal, hence we check for closeness
# with very low tolerance).
assert np.allclose(np.linalg.inv(R_30),
np.transpose(R_30))
# Equivalently, if we multiply a rotation matrix and its
# transpose, we get the identity matrix.
# np.eye(N) returns an N x N Identity matrix.
assert np.allclose(np.matmul(R_30, R_30.T),
np.eye(2))
# Let us take a random point (4, 0)
u = np.array([[4],[0]])
# Rotate it by 30 degrees
v = np.matmul(R_30, u)
print("Original vector u:\n{}".format(u))
print("Rotated Vector v:\n{}".format(v))
print("Length of u: {}".format(np.linalg.norm(u)))
print("Length of v: {}".format(np.linalg.norm(v)))
# We assert that rotation is length preserving
assert np.linalg.norm(u) == np.linalg.norm(v)
# Let us now negate the rotation i.e rotate the point
# back by -30 degrees
R_neg30 = rotation_matrix_2d(-30)
print("Matrix to rotate in-plane by -30 degrees"
" about origin\n{}".format(R_neg30))
# We will rotate v by -30 degrees
w = np.matmul(R_neg30, v)
print("Re-Rotated Vector w:\n{}".format(w))
# We assert that this vector is the same as the original
# vector u
assert np.all(w == u)
# We also assert that R_neg30 is the transpose and
# the inverse of R_30
assert np.allclose(R_30, R_neg30.T)
assert np.allclose(np.matmul(R_30, R_neg30), np.eye(2))
import numpy.linalg as LA
# Let us now compute the eigen values and eigen vectors of
# the rotation matrix
# Matrix for rotation by 45 degrees about origin.
R = np.array([[0.7071, 0.7071, 0], [-0.7071, 0.7071, 0],
[0, 0, 1]])
# As seen in the previous section, A is a rotation matrix
# around the Z axis
l, e = LA.eig(R)
# We know that all rotation matrices will have 1 as one of
# its eigenvalues.
# The eigen vector corresponding to that value is the axis
# of rotation
# np.where returns the indices where the specified condition
# is satisfied.
axis_of_rotation = e[:, np.where(l == 1.0)]
# np.squeeze is used to remove dimensions of size 1
axis_of_rotation = np.squeeze(axis_of_rotation)
print("Axis of rotation is: {}".format(axis_of_rotation))
# Let us take a random point on the axis of rotation
p = np.random.randint(0, 10) * axis_of_rotation
print("Point of axis of rotation: {}".format(p))
# Point on the axis of rotation remain unchanged even
# after rotation. Thus vector p and its transform Rp
# are close.
assert np.allclose(np.matmul(R, p), p)
```
| github_jupyter |
# Introduction
Theme 2: Healthcare
https://www.kaggle.com/gpreda/covid-world-vaccination-progress
* Q1: Which country has the highest & lowest % of population vaccinated?
* Q2: Is there any correlation between the number of population vaccinated vs international borders relaxing?
Next week, you will present your solution + reasons in a 3-slide presentation to any one of the themes you have chosen above. Maximum presentation time is 5 minutes, tops!
# Data
* https://www.kaggle.com/gpreda/covid-world-vaccination-progress
* https://www.thelancet.com/journals/lancet/article/PIIS0140-6736(20)31558-0/fulltext
# Initialize
```
# Mount the drive folder
from google.colab import drive # to load data from google drive
drive.mount("/content/drive")
# Load libraries
import os # For files operations
import urllib.request # For download from url
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib.pyplot as plt # ploting the data
import seaborn as sns # ploting the data
import csv # to import data in txt files
# to use Panda profilling
# See the output to NULL for notebook readibility
!pip install https://github.com/pandas-profiling/pandas-profiling/archive/master.zip > NULL
from pandas_profiling import ProfileReport
# Set up color blind friendly color palette
# The palette with grey:
cbPalette = ["#999999", "#E69F00", "#56B4E9", "#009E73", "#F0E442", "#0072B2", "#D55E00", "#CC79A7"]
# The palette with black:
cbbPalette = ["#000000", "#E69F00", "#56B4E9", "#009E73", "#F0E442", "#0072B2", "#D55E00", "#CC79A7"]
# sns.palplot(sns.color_palette(cbPalette))
# sns.palplot(sns.color_palette(cbbPalette))
sns.set_palette(cbPalette)
#sns.set_palette(cbbPalette)
# Seaborn favourite plot shortcuts
def boxplot(data, x, y, title = ""):
"""
This function generates a seaborn boxplot with my defaults parameters.
Parameters:
title (string) title of the plot, default is empty
data (df) the data frame
x (panda serie) the x axis
y (panda serie) the y axis
"""
f, ax = plt.subplots(figsize=(8, 6))
sns.boxplot(x=x, y=y, data=data, notch=True, showmeans=True,
meanprops={"marker":"s","markerfacecolor":"white", "markeredgecolor":"black"})
plt.title(title)
plt.ioff()
def countplot(data, variable, title = ""):
"""
This function contains my favourite parameters for the seaborn coutplot plot
"""
f, ax = plt.subplots(figsize=(8, 6))
sns.countplot(data=data, x=variable)
plt.title(title)
plt.ioff()
```
# Automated EDA
## country_vaccinations
```
# Set up the path for the data and output folders to the challenge and list files
PATH = "/content/drive/MyDrive/Data_science/DSAK"
data = PATH + "/Data"
output = PATH + "/Output"
os.listdir(data)
file = data + "/" + "country_vaccinations.csv"
country_vaccinations = pd.read_csv(file, sep = ',', encoding = 'UTF-8')
country_vaccinations.info()
report = ProfileReport(country_vaccinations)
# Display the report
report.to_notebook_iframe()
```
## country_vaccinations_by_manufacturer
```
file = data + "/" + "country_vaccinations_by_manufacturer.csv"
vaccinations_manufacturer = pd.read_csv(file, sep = ',', encoding = 'UTF-8')
vaccinations_manufacturer.info()
report = ProfileReport(vaccinations_manufacturer)
# Display the report
report.to_notebook_iframe()
# How many entries by contries?
vaccinations_manufacturer.groupby(['location'])["date"].count()
# Keep only the highest value
# https://www.statology.org/max-by-group-in-pandas/
vaccinations_manufacturer.groupby(['location'])[['total_vaccinations']].max()
```
## World population
```
file = data + "/" + "population_by_country_2020.csv"
population = pd.read_csv(file, sep = ',', encoding = 'UTF-8')
population.info()
report = ProfileReport(population)
# Display the report
report.to_notebook_iframe()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/denikn/Machine-Learning-MIT-Assignment/blob/main/Week%2001%20-%20Basics/Week01_Exercises_Hyperplanes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# 1) Hyperplanes
We will be using the notion of a hyperplane a great deal. A hyperplane is useful for classification, as discussed in the notes.

Some notational conventions:
x: is a point in d-dimensional space (represented as a column vector of d real numbers), R^d
d
θ: a point in d-dimensional space (represented as a column vector of d real numbers), R^dR
d
θ0: a single real number
We represent x and θ as column vectors, that is, d×1 arrays. Remember dot products? We write dot products in one of two ways: θ^T x or θ⋅.x. In both cases:
<br />
<center>
θ^
T
x=θ⋅x=θ
1
x
1
+θ
2
x
2
+…+θ
d
x
d
</center>
<br />
In a dd-dimensional space, a hyperplane is a d-1d−1 dimensional subspace that can be characterized by a normal to the subspace and a scalar offset. For example, any line is a hyperplane in two dimensions, an infinite flat sheet is a hyperplane in three dimensions, but in higher dimensions, hyperplanes are harder to visualize. Fortunately, they are easy to specify.
Hint: When doing the two-dimensional problems below, start by drawing a picture. That should help your intuition. If you get stuck, take a look at this geometry review for planes and hyperplanes.
## 1.1) Through origin
In dd dimensions, any vector \theta \in R^dθ∈R
d
can define a hyperplane. Specifically, the hyperplane through the origin associated with \thetaθ is the set of all vectors x \in R^dx∈R
d
such that \theta^T x = 0θ
T
x=0. Note that this hyperplane includes the origin, since x=0x=0 is in the set.

Ex1.1a: In two dimensions, \theta = [\theta_1, \theta_2]θ=[θ
1
,θ
2
] can define a hyperplane. Let \theta = [1, 2]θ=[1,2]. Give a vector that lies on the hyperplane given by the set of all x \in R^2x∈R
2
such that \theta^T x = 0θ
T
x=0:
Enter your answer as a Python list of numbers.
```
print([0,0])
```
Ex1.1b. Using the same hyperplane, determine a vector that is normal to the hyperplane.
```
print([1,2])
```
Ex1.1c. Now, in dd dimensions, supply the simplified formula for a unit vector normal to the hyperplane in terms of \thetaθ where \theta \in R^dθ∈R
d
.
In this question and the subsequent ones that ask for a formula, enter your answer as a Python expression. Use theta for \thetaθ, theta_0 for \theta_0θ
0
, x for any array x,x, transpose(x) for transpose of an array, norm(x) for the length (L2-norm) of a vector, and x@y to indicate a matrix product of two arrays.
```
print("-theta/((transpose(theta)@theta)**.5)")
```
## 1.2) General hyperplane, distance to origin
Now, we'll consider hyperplanes defined by \theta^T x + \theta_0 = 0θ
T
x+θ
0
=0, which do not necessarily go through the origin. Distances from points to such general hyperplanes are useful in machine learning models, such as the perceptron (as described in the notes).
Define the positive side of a hyperplane to be the half-space defined by \{x \mid \theta^T x + \theta_0 > 0\}{x∣θ
T
x+θ
0
>0}, so \thetaθ points toward the positive side.

Ex1.2a. In two dimensions, let \theta = [3, 4]θ=[3,4] and \theta_0 = 5θ
0
=5. What is the signed perpendicular distance from the hyperplane to the origin? The distance should be positive if the origin is on the positive side of the hyperplane, 0 on the hyperplane and negative otherwise. It may be helpful to draw your own picture of the hyperplane (like the one above but with the right intercepts and slopes) with \theta = [3, 4]θ=[3,4] and \theta_0 = 5θ
0
=5. Hint -Draw a picture
```
print(1) #the distance from the origin to the hyperplane
```
Ex1.2b: Now, in dd dimensions, supply the formula for the signed perpendicular distance from a hyperplane specified by \theta, \theta_0θ,θ
0
to the origin. If you get stuck, take a look at this walkthrough of point-plane distances.
```
print("theta_0 / norm(theta)")
```
# 2) Numpy intro
numpy is a package for doing a variety of numerical computations in Python. We will use it extensively. It supports writing very compact and efficient code for handling arrays of data. We will start every code file that uses numpy with import numpy as np, so that we can reference numpy functions with the 'np.' precedent.
You can find general documentation on numpy here, and we also have a 6.036-specific numpy tutorial.
The fundamental data type in numpy is the multidimensional array, and arrays are usually generated from a nested list of values using the np.array command. Every array has a shape attribute which is a tuple of dimension sizes.
In this class, we will use two-dimensional arrays almost exclusively. That is, we will use 2D arrays to represent both matrices and vectors! This is one of several times where we will seem to be unnecessarily fussy about how we construct and manipulate vectors and matrices, but we have our reasons. We have found that using this format results in predictable results from numpy operations.
Using 2D arrays for matrices is clear enough, but what about column and row vectors? We will represent a column vector as a d\times 1d×1 array and a row vector as a 1\times d1×d array. So for example, we will represent the three-element column vector,
x = \begin{bmatrix}1 \\ 5 \\ 3\end{bmatrix},
as a 3 \times 13×1 numpy array. This array can be generated with
~~~ x = np.array([[1],[5],[3]]),
or by using the transpose of a 1 \times 31×3 array (a row vector) as in,
~~~ x = np.transpose(np.array([[1,5,3]]),
where you should take note of the "double" brackets.
It is often more convenient to use the array attribute .T , as in
~~~ x = np.array([[1,5,3]]).T
to compute the transpose.
Before you begin, we would like to note that in this assignment we will not accept answers that use "loops". One reason for avoiding loops is efficiency. For many operations, numpy calls a compiled library written in C, and the library is far faster than that interpreted Python (in part due to the low-level nature of C, optimizations like vectorization, and in some cases, parallelization). But the more important reason for avoiding loops is that using higher-level constructs leads to simpler code that is easier to debug. So, we expect that you should be able to transform loop operations into equivalent operations on numpy arrays, and we will practice this in this assignment.
Of course, there will be more complex algorithms that require loops, but when manipulating matrices you should always look for a solution without loops.
Numpy functions and features you should be familiar with for this assignment:
np.array
np.transpose (and the equivalent method a.T)
np.ndarray.shape
np.dot (and the equivalent method a.dot(b) )
np.sign
np.sum (look at the axis and keepdims arguments)
Elementwise operators +, -, *, /
Note that in Python, np.dot(a, b) is the matrix product a@b, not the dot product a^T ba
T
b.
## 2.1) Array
Provide an expression that sets A to be a 2 \times 32×3 numpy array (22 rows by 33 columns), containing any values you wish.
```
import numpy as np
A = np.array([[2, 4, 6], [6, 8, 10]], np.int32)
print(A)
```
## 2.2) Transpose
Write a procedure that takes an array and returns the transpose of the array. You can use 'np.transpose' or the '.T', but you may not use a loop.
```
import numpy as np
def tp(A):
return np.transpose(A)
print(tp([[1,2,3],[4,5,6]]))
```
## 2.3) Shapes
Let A be a 4\times 24×2 numpy array, B be a 4\times 34×3 array, and C be a 4\times 14×1 array. For each of the following expressions, indicate the shape of the result as a tuple of integers (recall python tuples use parentheses, not square brackets, which are for lists, and a tuple of a single object x is written as (x,) with a comma) or "none" (as a Python string with quotes) if it is illegal.
Ex2.3a (c*c)
```
import numpy as np
first_four_by_one = np.array([[1], [2], [3], [4]])
print(first_four_by_one.shape)
second_four_by_one = np.array([[4], [5], [6], [7]])
print(second_four_by_one.shape)
result = first_four_by_one * second_four_by_one
print(result.shape)
print(result)
```
Ex2.3b (np.dot(C, C))
```
import numpy as np
c = np.array([[1], [2], [3], [4]])
# print(np.dot(c,c))
# print(np.dot(c,c).shape)
print("The result is error, should be answer as 'none'")
```
Ex2.3c (np.dot(np.transpose(C), C))
```
import numpy as np
c = np.array([[1], [2], [3], [4]])
print(np.dot(np.transpose(c), c))
print(np.dot(np.transpose(c), c).shape)
```
Ex2.3d (np.dot(A, B))
```
import numpy as np
a = np.array([[1,2,3,4], [2,2,3,4]])
b = np.array([[1,2,3,4], [2,2,3,4], [2,2,3,4]])
# print(np.dot(c,c))
# print(np.dot(c,c).shape)
print("The result is error, should be answer as 'none'")
```
Ex2.3e (np.dot(A.T, B))
```
import numpy as np
a = np.array([[1,2,3,4], [2,2,3,4]])
b = np.array([[1,2,3,4], [2,2,3,4], [2,2,3,4]])
# print(np.dot(np.transpose(a), b))
# print(np.dot(a,b).shape)
print("The result is error, should be answer as 'none'. But the correct result is (2,3)")
```
Ex2.3f (D = np.array([1,2,3]))
```
import numpy as np
D = np.array([1,2,3])
print(D)
print("The result should be (3, )")
```
Ex2.3g (A[:,1])
```
import numpy as np
a = np.array([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]])
print(a[:,1])
print(a[:,1].shape)
```
Ex2.3h (A[:,1:2])
```
import numpy as np
a = np.array([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]])
print(a[:,1:2])
print(a[:,1:2].shape)
```
## 2.4) Row vector
Write a procedure that takes a list of numbers and returns a 2D numpy array representing a row vector containing those numbers.
```
import numpy as np
def rv(value_list):
return np.array([value_list])
print(rv([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]]))
```
## 2.5) Column vector
Write a procedure that takes a list of numbers and returns a 2D numpy array representing a column vector containing those numbers. You can use the rv procedure.
```
import numpy as np
def cv(value_list):
return np.transpose(np.array([value_list]))
print(cv([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]]))
```
## 2.6) length
Write a procedure that takes a column vector and returns the vector's Euclidean length (or equivalently, its magnitude) as a scalar. You may not use np.linalg.norm, and you may not use a loop.
```
import numpy as np
def length(col_v):
return np.sqrt(np.sum(np.square([col_v])))
print(length([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]]))
```
## 2.7) normalize
Write a procedure that takes a column vector and returns a unit vector in the same direction. You may not use a for loop. Use your length procedure from above (you do not need to define it again).
```
import numpy as np
def normalize(col_v):
return col_v / np.linalg.norm(col_v)
print(normalize([[1,2,3,4], [2,2,3,4], [1,2,3,4], [1,2,3,4]]))
```
## 2.8) indexing
Write a procedure that takes a 2D array and returns the final column as a two dimensional array. You may not use a for loop.
```
import numpy as np
def index_final_col(A):
flipped_array = np.flip(A, 1)
return flipped_array[:,:1]
print(index_final_col([[1,2],[3,4]]))
```
## 2.9) Representing data
Alice has collected weight and height data of 3 people and has written it down below:
Weight, height
150, 5.8
130, 5.5
120, 5.3
She wants to put this into a numpy array such that each row represents one individual's height and weight in the order listed. Write code to set data equal to the appropriate numpy array:
```
import numpy as np
data = np.array([[150, 5.8], [130, 5.5], [120, 5.3]])
print(data)
```
Now she wants to compute the sum of each person's height and weight as a column vector by multiplying data by another numpy array. She has written the following incorrect code to do so and needs your help to fix it:
## 2.10) Matrix multiplication
```
import numpy as np
def transform(data):
return (np.dot(data, np.array([[1], [1]])))
print(transform([[150, 5.8], [130, 5.5], [120, 5.3]]))
```
| github_jupyter |
# SIT742: Modern Data Science
**(Week 04: Text Analysis)**
---
- Materials in this module include resources collected from various open-source online repositories.
- You are free to use, change and distribute this package.
- If you found any issue/bug for this document, please submit an issue at [tulip-lab/sit742](https://github.com/tulip-lab/sit742/issues)
Prepared by **SIT742 Teaching Team**
---
## Session 4B - Exploring Pre-Processed text and Generating Features
### Table of Content
* Part 1. Counting Vocabulary by Selecting Tokens of Interest
* Part 2. Building Vector Representation
* Part 3. Saving Pre-processed Text to a File
* Part 4. Extracting Other Features
* Part 5. Summary
* Part 6. Reading Materials
* Part 7. Exercises
---
One of the challenges of text analysis is to convert unstructured and semi-structured text into a structured representation. This must be done prior to carrying out any text analysis tasks. This chapter will show you
how to put some of those basic steps discussed in the previous chapter together to generate different vector
representations for some given text. You will learn how to compute some basic statistics for text, and how to extract features rather than unigrams.
## Part 1. Counting Vocabulary by Selecting Tokens of Interest
Two important concepts that should be mentioned first are **type** and **token**.
Here are the definitions of the two terms, quoted from "[tokenization](http://nlp.stanford.edu/IR-book/html/htmledition/tokenization-1.html)",
>a **token** is an instance of a sequence of characters in some particular document that are grouped together as a useful semantic unit for processing;
> a **type** is the class of all tokens containing the same character sequence.
A *type* is also a vocabulary entry. In other words, a vocabulary consists of a number of word types.
The distinction between a type and its tokens is a distinction that separates a descriptive concept from
its particular concrete instances.
This is quite similar to the distinction in object-oriented programming between classes and objects.
In this section, you are going to learn how to count types in a given corpus by further processing the text.
The document collection that we are going to use is a set of Reuters articles that comes with NLTK.
It contains 10788 Reuters articles in total and has been split into two subsets, training and testing.
Although this collection has already been pre-processed (e.g., you can access the text at different levels, like raw text, tokens, and sentences),
we would still like to demonstrate how to put some of the basis text preprocessing steps together and process the raw Reuters articles step by step.
First, import the main Python libraries.
```
import matplotlib.pyplot as plt
%matplotlib inline
import nltk
from nltk.corpus import reuters
```
Since the tokenizer works on a per document level, we can parallelize the process of tokenization with Python's multi-processing module. Please refer to its official documentation [here](https://docs.python.org/2/library/multiprocessing.html).
In the following code, we wrap tokenization in a Python function, and then
create a pool of four worker processes with the Python Pool class.
The <font color="blue">Pool.map()</font>, a parallel equivalent of the built-in <font color="blue">map()</font> function, takes one iterable argument.
The iterable will be split into a number of chunks, each of which will be submitted to a process in the process pool.
Each process will apply a callable function to each element in the chunk it has received.
Note that you can replace the NLTK tokenizer with the one you implement.
```
def tokenizeRawData(fileid):
"""
This function tokenizes a raw text document.
"""
raw_article = reuters.raw(fileid).lower() # cover all words to lowercase
tokenised_article = nltk.tokenize.word_tokenize(raw_article) # tokenize each Reuters articles
return (fileid, tokenised_article)
nltk.download('reuters')
tokenized_reuters = dict(tokenizeRawData(fileid) for fileid in reuters.fileids())
```
### 1.1. Removing Words with Non-alphabetic Characters
The NLTK's built-in <font color="blue">word_tokenize</font> function tokenizes a string to split off punctuation other than periods.
Not only does it return words with alphanumerical characters, but also punctuations.
Let's take a look at one Reuters articles,
```
tokenized_reuters['training/1684']
```
Let's Assume that we are interested in words containing alphabetic characters only
and would like to remove all the other tokens
that contain digits, punctuation and the other symbols.
Removing all the non-alphabetic words from the vocabulary is
usually required in some text analysis tasks, such as Topic Modelling that
learns the semantic meaning of documents.
It can be easily done with the <font color="blue">isalpha()</font> function.
<font color="blue">isalpha()</font>
checks whether the string consists of alphabetic characters only or not.
This method returns true if all characters in the string are in the alphabet and there
is at least one character, false otherwise.
If you would like to keep all words with alphanumeric characters, you can use
<font color="blue">isalnum()</font>. Refer to Python's [built-in types](https://docs.python.org/2/library/stdtypes.html) for more detail.
Indeed, you can construct your tokenizer in a way such that the tokenizer only extracts words with either
alphabetic or alphanumerical characters, as we discussed in the previous chapter.
We will leave this as a simple exercise for you to do on your own.
```
for k, v in tokenized_reuters.items():
tokenized_reuters[k] = [word for word in v if word.isalpha()]
tokenized_reuters['training/1684']
```
Now you should have derived much cleaner text for each Reuters article.
Let's check how many types we have in the whole corpus and the lexical diversity (i.e., the average number
of times a type apprearing in the collection.)
```
from __future__ import division
from itertools import chain
words = list(chain.from_iterable(tokenized_reuters.values()))
vocab = set(words)
lexical_diversity = len(words)/len(vocab)
print ("Vocabulary size: ",len(vocab),"\nTotal number of tokens: ", len(words), \
"\nLexical diversity: ", lexical_diversity)
```
There are about 1.27 million word tokens in the tokenized Reuters corpus.
The vocabulary size is 27,944, which is still quite large according to our knowledge of this corpus.
The lexical diversity tells us that words occur on average about 46 times each.
You might think that
there could still be words that occur very frequently, such as stopwords,
and those that only occur once or twice.
For example, if an article "the" appears in almost
every document in a corpus,
it might not help you at all and would only contribute noise.
Similarly if a word appears only once in a corpus or only in one document of the corpus,
it could carry little useful information for downstream analysis.
Therefore, we would better remove those words from the vocabulary, which
will benefit the text analysis algorithms in terms of reducing running time and
memory requirement, and improving their performance.
To do so, we need to further explore the corpus by computing some simple
statistics.
Note that we introduced two new Python libraries in the code above.
They are
[`__future__`](https://docs.python.org/2/library/__future__.html)
and [`itertools`](https://docs.python.org/2/library/itertools.html).
The first statement in the code makes sure that Python switches to
always yielding a real result.
Thus if you divide two integer values, you will not get for example.
````
1/2 = 0
3/2 = 1
````
Instead, you will have
```
1/2 = 0.5
3/2 = 1.5
```
The second statement imported a <font color="blue">chain()</font> iterator from the <font color="blue">itertools</font> module.
We use the iterator to join all the words in all the Reuters articles together.
It works as
```python
for wordList in tokenized_reuters.values():
for word in wordList:
yield word
```
### 1.2. Removing the Most and Less Frequent Words
It is quite useful for us to identify the words that are most informative about the sematic
meaning of the text regardless of syntax.
One common statistics often used in text processing is frequency distribution.
It can tell us how frequent a word is in a given corpus in terms of either term frequency or document frequency.
Term frequency counts the number of times a word occurs in the whole corpus regardless which document it is in.
Frequency distribution based on term frequency tells us how the total number of word tokens are distributed across all the types.
NLTK provides a built-in function `FreqDist` to compute this distribution directly from a set of word tokens.
```
from nltk.probability import *
fd_1 = FreqDist(words)
```
What are the most frequent words in the corpus?
we can use the <font color="blue">most_common</font> function to print out the most frequent words together with their frequencies.
```
fd_1.most_common(25)
```
The list above contains the 25 most frequent words.
You can see that it is mostly dominated by the little words of the English language which have important grammatical roles.
Those words are articles, prepositions, pronouns, auxiliary webs, conjunctions, etc.
They are usually referred to as function words in linguistics, which tell us nothing about
the meaning of the text.
What proportion of the text is taken up with such words?
We can generate a cumulative frequency plot for them
using <font color="blue">fd.plot(25, cumulative=True)</font>.
If you set <font color="blue">cumulative</font> to <font color="blue">False</font>,
it will plot the frequencies of these 25 words.
These 25 words account for about 33% of the while Reuters corpus.
```
fd_1.plot(25, cumulative=True)
```
What are the most frequent words in terms of document frequency?
Here we are going to count how many documents a word appears in, which is referred to as document frequency.
Instead of writing nested FOR loops to count the document frequency for each word,
we can use <font color="blue">FreqDist()</font> jointly with <font color="blue">set()</font> as follows:
1. Apply <font color="blue">set()</font> to each Reuters article to generate a set of unique words in the article and save all sets in a list
```python
[set(value) for value in tokenized_reuters.values()]
```
2. Similar to what we have done before, we put all the words in a list using <font color="blue">chain.from_iterable</font> and past
it to <font color="blue">FreqDist</font>.
The first step makes sure that each word in an article appears only once, thus the total number of
times a word appears in all the sets is equal to the number of documents containing that word.
```
words_2 = list(chain.from_iterable([set(value) for value in tokenized_reuters.values()]))
fd_2 = FreqDist(words_2)
fd_2.most_common(25)
```
What you will find is that the majority of the most frequent words according to their document frequecy are still functional words.
Therefore, the next step is to remove all the stopwords.
#### 1.2.1 Ignoring Stopwords
We often remove function words from the text completely for most text analysis tasks.
Instead of using the built-in stopword list of NLTK, we use a much rich stopword list.
```
!pip install wget
import wget
link_to_data = 'https://github.com/tulip-lab/sit742/raw/master/Jupyter/data/stopwords_en.txt'
DataSet = wget.download(link_to_data)
!ls
stopwords = []
with open('stopwords_en.txt') as f:
stopwords = f.read().splitlines()
tokenized_reuters_1 = {}
for fileid in reuters.fileids():
tokenized_reuters_1[fileid] = [w for w in tokenized_reuters[fileid] if w not in stopwords]
```
The list comprehension
```python
[w for w in tokenized_reuters[fileid] if w not in stopwords]
```
says: For each word in each Reuters article, keep the word if the word is not contained in the stopword list.
Checking for membership of a value in a list takes time proportional to the list's length in the average and worst cases.
It causes the above code to run quite slow as we need to do the check for every word in each Reuters article
and the size of the stopword list is large.
However, if you have hashable items, which means both the item order and duplicates are disregarded,
Python `set` is better choice than `list`. The former runs much faster than the latter in terms of searching
a large number of hashable items. Indeed, `set` takes constant time to check the membership.
Let's try converting the stopword list into a stopword set, then search to remove all the stopwords.
Please also note that if you try to perform iteration, `list` is much better than `set`.
```
stopwordsSet = set(stopwords)
for fileid in reuters.fileids():
tokenized_reuters[fileid] = [w for w in tokenized_reuters[fileid] if w not in stopwordsSet]
```
In the above stopping process, 481 stopwords have been removed from the vocabulary. You might wonder what those removed words are. It is quite easy to check those words by differentiating the vocabulary before and after stopping.
```
words_3 = list(chain.from_iterable(tokenized_reuters.values()))
fd_3 = FreqDist(words_3)
list(vocab - set(fd_3.keys()))
```
Beside stopwords, there might some other words that occur quite often as well.
```
fd_3.most_common(10)
```
Before we decide to remove those words from our vocabulary, it might be worth checking what
those words mean and the context of those words. Fortunately NLTK provides a `concordance`
function in the `nltk.text` module. A concordance view shows us every occurrence of a given
word, together with the corresponding context. For example,
```
nltk.Text(reuters.words()).concordance('mln')
nltk.Text(reuters.words()).concordance('net')
```
After reviewing those words, you might also want to remove them from the vocabulary.
We will leave it as an excersie for you to do on your own.
#### 1.2.2 Remove Less Frequent Words
If the most common words do not benefit the downstream text analysis tasks, except for contributing noises,
how about the words that occur once or twice?
Here another interesting statistic to look at is the frequency of the frequencies of word types in a given corpus.
We would like to see how many words appear only once, how many words appear twice, how many
words appear three times, and so on.
```
ffd = FreqDist(fd_3.values())
from pylab import *
y = [0]*14
for k, v in ffd.items():
if k <= 10:
y[k-1] = v
elif k >10 and k <= 50:
y[10] = y[10] + v
elif k >50 and k <= 100:
y[11] = y[11] + v
elif k > 100 and k <= 500:
y[12] = y[12] + v
else:
y[13] = y[13] + v
x = range(1, 15) # generate integer from 1 to 14
ytks =list(map(str, range(1, 11))) # covert a integer list to a string list
ytks.append('10-50')
ytks.append('51-100')
ytks.append('101-500')
ytks.append('>500')
barh(x,y, align='center')
yticks(x, ytks)
xlabel('Frequency of Frequency')
ylabel('Word Frequency')
grid(True)
```
The horizontal bar chart generated above shows how many word types occur with a certain frequency.
There are 241 types occurring over 500 times and therefore individually accounting for about 1% of
the vocabulary.
However, on the other extreme, more than one-third of the word types occur only once in the Reuters corpus.
Note that the majority of word types occur quite infrequently given the size of the whole corpus (i.e., 721,371 word tokens):
about 78% of the word types occur 10 times or less.
Similarly, you can also look at the bar chart based on the document frequency. Try it by yourself!
Let's further remove those words that occur only once.
To get those words, you can write the code like
```python
lessFreqWords = set([k for k, v in fdist.items() if v < 2])
```
or choose to use `hapaxes()` function.
```
lessFreqWords = set(fd_3.hapaxes())
def removeLessFreqWords(fileid):
return (fileid, [w for w in tokenized_reuters[fileid] if w not in lessFreqWords])
#pool = mp.Pool(4)
#tokenized_reuters = dict(pool.map(removeLessFreqWords, reuters.fileids()))
tokenized_reuters = dict(removeLessFreqWords(fileid) for fileid in reuters.fileids())
```
Now, you should have a pretty clean set of Reuters articles, each of which is stored as a list of word tokens.
Let's further print out some statistics that summarize this corpus.
```
import numpy as np
words = list(chain.from_iterable(tokenized_reuters.values()))
vocab = set(words)
print ("Vocabulary size: ",len(vocab))
print ("Total number of tokens: ", len(words))
print ("Lexical diversity: ", lexical_diversity)
print ("Total number of articles:", len(tokenized_reuters))
lens = [len(value) for value in tokenized_reuters.values()]
print ("Average document length:", np.mean(lens))
print ("Maximun document length:", np.max(lens))
print ("Minimun document length:", np.min(lens))
print ("Standard deviation of document length:", np.std(lens))
```
It is interesting that the minimun document length is 0. There must be some Reuters articles that are extremely short,
after tokenization and stopping, there are no words left. Can you check those documents to see what they look like?
## Part 2. Building Vector Representation
After text pre-processing has been completed, each individual document needs to be transformed into
some kind of numeric representation that can be input into most NLP and text mining algorithms.
For example, classification algorithms, such as Support Vector Machine, can only take data in a
structured and numerical form. They do not accept free languge text.
The most popular structured representation of text is the vector-space model, which represents text
as a vector where the elements of the vector indicate the occurence of words within the text.
The vector-space model makes an implicit assumption that
the order of words in a text document are not as
important as words themselves, and thus disregarded.
This assumpiton is called [**Bag-of-words**](https://en.wikipedia.org/wiki/Bag-of-words_model).
Given a set of documents and a pre-defined list of words appearing
in those documents (i.e., a vocabulary), you can compute a vector representation for each document.
This vector representation can take one of the following three forms:
* a binary representation,
* an integer count,
* and a float-valued weighted vector.
To highlight the difference among the three approaches, we use a very simple example as follows:
```
document_1: "Data analysis is important."
document_2: "Data wrangling is as important as data analysis."
document_3: "Data science contains data analysis and data wrangling."
```
The three documents contain 20 tokens and 9 unique words.
Those unique words are sorted alphabetically with total counts:
```
'analysis': 3,
'and': 1,
'as': 2,
'contains': 1,
'data': 6,
'important': 2,
'is': 2,
'science': 1,
'wrangling': 2
```
Given the vocabulary above,
both the binary and the integer count vectors are easy to compute.
A binary vector stores 1s for the word that appears in a document and 0s for the other words in
the vocabulary,
whereas a count vector stores the frequency of each word appearing in the document.
Thus, the binary vector representations for the three documents above are
||'analysis'|'and'|'as'|'contains'|'data'|'important'|'is'|'science'|'wrangling'|
|-|-|-|-|-|-|-|-|-|
|document 1:|1|0|0|0|1|1|1|0|0|
|document 2:|1|0|1|0|1|1|1|0|1|
|document 3:|1|1|0|1|1|0|0|1|1|
The count vector representations for the same documents would look as follows:
||'analysis'|'and'|'as'|'contains'|'data'|'important'|'is'|'science'|'wrangling'|
|-|-|-|-|-|-|-|-|-|
|document 1:|1|0|0|0|1|1|1|0|0|
|document 2:|1|0|2|0|2|1|1|0|1|
|document 3:|1|1|0|1|3|0|0|1|1|
Instead of using the two vector representations above,
most existing text analysis algorithms, like document classification and information retrieval,
prefer representing documents as weighted vectors.
The raw term frequency is often replaced with a weighted term frequency
that indicates how important a word is in a particular document.
There are many different term weighting schemes online.
To store each document as a weighted vector, we first need to choose a weighting scheme.
The most popular scheme is the TF-IDF weighting approach.
TF-IDF stands for term frequency-inverse document frequency.
The term frequency for a word is the number of times the word appears in a document.
In the preceding example, the term frequency in Document 2 for “data” is 2, since it appears twice in the document. Document frequency for a word is the number of documents that contain the word;
it would also be 3 for “data” in the collection of the three preceding documents.
The Wikipidia entry on [TF-IDF](https://en.wikipedia.org/wiki/Tf%E2%80%93idf) lists
a number of variants of TF-IDF.
One variant is reproduced here
$$tf\cdot idf(w,d) = tf(w, d) * idf(w)$$
where
$$tf(w,d)\,=\, \sum_{i}^{|d|} 1_{w = w_{d,i}}$$
and
$$idf(w) = log\left(\frac{|D|}{|d \in D: w \in d |}\right)$$
The assumption behind TF-IDF is that words with high term frequency should receive high weight unless they also have high document frequency.
Stopwords are the most commonly occurring words in the English language. They often occur many times within a single document, but they also occur in nearly every document.
These two competing effects cancel out to give them low weights,
as those very common words carry very little meaningful information about the actual contents of the document.
Therefore, the TF-IDF weights for stopwords are almost always 0.
With the TF-DF formulas above,
the weighted vector representations for the example documents are computed as
||'analysis'|'and'|'as'|'contains'|'data'|'important'|'is'|'science'|'wrangling'|
|-|-|-|-|-|-|-|-|-|
|document 1:|0|0|0|0|0|0.176|0.176|0|0|
|document 2:|0|0|0.954|0|0|0.176|0.176|0|0.176|
|document 3:|0|0.477|0|0.477|0|0|0|0.477|0.176|
Given the cleaned up Reuters documents, how can we generate those vectors for each documents?
Unfortunately, NLTK does not implement methods that directly produce those vectors.
Therefore, we will either write our own code to compute them or appeal to other data analysis libraries.
Here we are going to use [scikit-learn](http://scikit-learn.org/stable/index.html), an open source machine
learning library for Python.
If you use Anaconda, you should already have scikit-learn installed, otherwise you will need to
[install it](http://scikit-learn.org/stable/install.html) by following the instruction on its official website.
Although scikit-learn features various classification, regression and clustering algorithms
we are particularly interested in its feature extraction module, [sklearn.feature_extraction](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.feature_extraction).
This module is often used to "extract features in a format supported by machine learning algorithms from datasets consisting of formats such as text and image." Please refer to its documentation on text feature extraction,
section 4.2.3 of [Feature Extraction](http://scikit-learn.org/stable/modules/feature_extraction.html#text-feature-extraction). We will demonstrate the usage of the following two classes:
* [CountVectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html#sklearn.feature_extraction.text.CountVectorizer): It converts a collection of text documents to a matrix of token counts.
* [TfidfVectorizer](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html#sklearn.feature_extraction.text.TfidfVectorizer):
It converts a collection of raw documents to a matrix of TF-IDF features.
### 2.1 Creating Count Vectors
Let's start with generating the count vector representation for each Reuters document.
Initialise the "CountVector" object: since we have pre-processed all the Reuters documents,
the parameters, "tokenizer", "preprocessor" and "stop_words" are set to their default value, i.e., None.
```
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(analyzer = "word")
```
Next, transform Reuters articles into feature vectors. `fit_transform` does two things: First, it fits the model and learns the vocabulary; second it transforms the text data into feature vectors.
Please note the input to `fit_transform` should be a list of strings.
Since we have stored each tokenised article as a list of words, we concatenate all the words in the list and separate
them with white spaces.
The following code will do that:
```python
[' '.join(value) for value in tokenized_reuters.values()]
```
Then, we input this list of strings into `fit_transform`,
```
data_features = vectorizer.fit_transform([' '.join(value) for value in tokenized_reuters.values()])
print (data_features.shape)
```
The shape of document-by-word matrix should be 10788 * 17403.
However, in order to save such a matrix in memory but also to speed up algebraic operations on the matrix,
scikit-learn implements matrix/vector in a sparse representation.
Let's check the count vector for the first article, i.e., 'training/1684'.
```
vocab2 = vectorizer.get_feature_names()
for word, count in zip(vocab, data_features.toarray()[0]):
if count > 0:
print (word, ":", count)
```
Another way to get the count list above is to use `FreqDist`.
```
FreqDist(tokenized_reuters['training/1684'])
```
Note that the vocabulary you just got with `vectorizer.get_feature_names()` shoud be exactly the same
as the one you got in section 1.
```
list(vocab-set(vocab2))
```
### 2.2 Creating TF-IDF Vectors
Similar to the use of `CountVector`, we first initialise a `TfidfVectorizer` object by only specifying
the value of "analyzer", and then covert the Reuters data into a list of strings, each of which corresponds
to a Reuters articles.
```
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(analyzer = "word")
tfs = tfidf.fit_transform([' '.join(value) for value in tokenized_reuters.values()])
tfs.shape
```
Let's print out the weighted vector for the first document.
```
vocab = vectorizer.get_feature_names()
for word, weight in zip(vocab, tfs.toarray()[0]):
if weight > 0:
print (word, ":", weight)
```
So now we have converted all the Reuters articles into feature vectors.
We can use those vectors to, for example,
* compute the similarity between two articles,
* search articles for a given query
* do other advance text analysis, such as document classification and clustering.
Assume that we have a new document, how can we get its TF-IDF vector.
We do this by using the transform function as follows.
We have randomly chosen a sentence from
[a recent Reuters news](http://www.reuters.com/article/us-usa-election-idUSKCN0W346T).
```
str = """
the former secretary of state hoped to win enough states to take a big step toward wrapping up her nomination fight
with a democratic senator from Vermont.
"""
response = tfidf.transform([str])
for col in response.nonzero()[1]:
print (vocab[col], ' - ', response[0, col])
```
Note that the text above is not included in the trained TF-IDF model with the 'transform' function, unless the `fit_transform` function is called,
Both `CountVectorizer` and `TfidfVectorizer` come with their own options to automatically do pre-processing, tokenization, and stop word removal -- for each of these, instead of using their default value (i.e., None),
we could customise the two vectorizer classes by either using a built-in method or specifying our own function.
See the function documentation for more details.
However, we wanted to write our own function for clean the text data in this chapter to show you how
it's done step by step.
## Part 3. Saving Pre-processed Text to a File
The pre-processed text needs to be saved in a proper format so that it can be easily used by the downstream analysis algorithm. There are a couple of ways of dumping the pre-processed text data into txt files.
For example, use one txt file to store the tokenized documents. The tokens in a document are stored in one row in the txt file, and are separated with a given delimiter, e.g., whitespace. In this case, the downstream text analyser needs to re-construct the vocabulary.
```
import wget
link_to_data = 'https://github.com/tulip-lab/sit742/raw/master/Jupyter/data/reuters_1.txt'
DataSet = wget.download(link_to_data)
# import wget
link_to_data = 'https://github.com/tulip-lab/sit742/raw/master/Jupyter/data/reuters_2.txt'
DataSet = wget.download(link_to_data)
!ls
out_file = open("reuters_1.txt", 'w')
for d in tokenized_reuters.values():
out_file.write(' '.join(d) + '\n')
out_file.close()
```
You can also save vocabulary in a separate file, and assign a fixed integer id to each word in the vocabulary. What text analysers usually do is to use the index of each word in the vocabulary as its integer id.
Given the vocabulary, each document can be represented as a sequence of integers that correspond to the tokens,
or in the following sparse form:
```
word_index:word count
```
for example,
```
out_file = open("reuters_2.txt", 'w')
vocab = list(vocab)
vocab_dict = {}
i = 0
for w in vocab:
vocab_dict[w] = i
i = i + 1
for d in tokenized_reuters.values():
d_idx = [vocab_dict[w] for w in d]
for k, v in FreqDist(d_idx).items():
out_file.write("{}:{} ".format(k,v))
out_file.write('\n')
out_file.close()
```
## Part 4. Extracting Other Features
It is common for most text analysis tasks to treat documents as bags-of-words, which can significantly simplify the inference procedure of text analysis algorithms.
However, things always have pros and cons.
The bag-of-words representation loses lots of information encoded in either syntax or word order (i.e., dependencies between adjacent words in sentences.).
For example, representing a document as a collection of unigrams effectively disregards any word order dependence,
which fails to capture phrases and multi-word expressions. A similar issue has been mentioned in section 2.1. of Chapter 2.
In this section, we are going to show you how to
* use Part-of-Speeching (POS) tagging to extract specific word groups, such as all nouns, verbs, etc.,
* extract n-grams,
* and extract collocations
These features can be further used to enrich the representation of a document.
### 4.1 Extracting Nouns and Verbs
It is easy for human to tell the difference between nouns, verbs,
adjectives and adverbs, as we have learnt them back in elementary school.
However, how can we automatically classify words into their parts of speech (i.e., lexical categories or word classes)
and label them accordingly with computer program?
This section is not going to discuss how to determine the category of a word from a linguistic perspective.
Instead it demonstrates the use of some existing POS taggers to extract words in a specific lexical category.
It has been proven that words together with their part-of-speech (POS) are quite useful for many language processing tasks.
In NLP, the process of labelling words with their corresponding part-of-speech (POS) tags is known as [POS tagging](https://en.wikipedia.org/wiki/Part-of-speech_tagging).
A POS tagger processes a sequence of words and attaches a POS tag to each word based on both its definition and its context. There are many POS taggers available online, such as [Sandford POS tagger](http://nlp.stanford.edu/software/tagger.shtml).
We are going to use the one implemented by NLTK.
```
example_sent = 'A POS tagger processes a sequence of words and attaches a POS tag to each \
word based on both its definition and its context'
text = nltk.word_tokenize(example_sent)
tagged_sent = nltk.tag.pos_tag(text)
print (tagged_sent)
```
If you are seeing these tags for the first time, you will wonder what these tags mean.
You can find the specification of all the tags [here](https://www.ling.upenn.edu/courses/Fall_2003/ling001/penn_treebank_pos.html).
NLTK provides documentation for each tag, which can be queried using the tag, e.g.,
```
nltk.download('tagsets')
print (nltk.help.upenn_tagset('NNP'))
print (nltk.help.upenn_tagset('IN'))
print (nltk.help.upenn_tagset('PRP$'))
```
The example sentence has been processed by `pos_tag` into a list of tuples, each of which is a pair of a word and its POS tag. We see that 'a' is 'DT', a determiner; 'its' is 'PRP$', a possessive pronoun; 'and' is 'CC', a coordinating conjunction, 'words' is 'NNS', a noun in the plural form, and so on. Note that several of the corpora included in NLTK have been tagged for their POS. Please click [here](http://www.nltk.org/howto/corpus.html#tagged-corpora) to see how to access those tagged corpora.
Here is an example of using the `tagged_words` function to retrieve all words in Brown corpus with their tags.
```
nltk.download('brown')
nltk.corpus.brown.tagged_words()
```
Please note that the collection of tags is known as a tag set.
There are many different conventions for tagging words.
Therefore, tag sets can vary among different tasks.
What we used above is the Penn Treebank tag set.
Let's change the tag set to the Universal POS tag set, and print the Brown corpus again.
You will find different tags are used.
```
nltk.download('universal_tagset')
nltk.corpus.brown.tagged_words(tagset='universal')
```
If you would like to learn more about POS tagging, please refer to [1].
Given the tagged text, you can easily identify all the nouns, verbs, etc.
Nouns generally refer to people, places, things, or concepts, e.g., Monash, Melbourne, university, data, and science.
Nouns can appear after determiners and adjectives, and can be the subject or object of the verb.
Now how can we extract all the nouns from a text?
Assume we use the Penn Treebank tag set.
Here are all the tags for nouns:
```
NN Noun, singular or mass
NNS Noun, plural
NNP Proper noun, singular
NNPS Proper noun, plural
```
It is not hard to see all the tags above start with 'NN'.
Thus, we can iterate over all the words and check if their tag string starts with 'NN'.
```
all_nouns = [w for w,t in tagged_sent if t.startswith('NN')]
all_nouns
```
Similarly, you will find that all the verb tags start with 'VB', see
```
VB Verb, base form
VBD Verb, past tense
VBG Verb, gerund or present participle
VBN Verb, past participle
VBP Verb, non-3rd person singular present
VBZ Verb, 3rd person singular present
```
Thus,
```
all_verbs = [w for w,t in tagged_sent if t.startswith('VB')]
all_verbs
```
Unfortunately, the Reuters corpus that we have been using, has no built-in POS tags. But you can get sentences from Reuters corpus, and then you can get the POS tags.
### 4.2 Extracting N-grams and Collocations
Besides unigrams that we have been working on so far,
N-grams of texts are also extensively used in various text analysis tasks.
They are basically contiguous sequences of `n` words from a given sequence of text.
When computing the n-grams you typically move a fixed size window of size n
words forward.
For example, for the sentence
"Laughter is like a windshield wiper."
if N = 2 (known as bigrams), the n-grams would be:
```
Laughter is
is like
like a
a windshield
windshield wiper
```
So you have 5 bigrams in this case. Notice that the generative process above
essentially moves one word forward to generate the next bigram.
If N = 3 (known as trigrams), the n-grams would be:
```
Laughter is like
is like a
like a windshield
a windshield wiper
```
What are N-grams used for? They can be used to build n-gram language model that
can be further used for speech recognition, spelling correction, entity detection, etc.
In terms of text mining tasks, n-grams is used for developing features for
classification algorithms, such as SVMs, MaxEnt models, Naive Bayes, etc.
The idea is to expand the unigram feature space with n-grams.
But please notice that
the use of bigrams and trigrams in your feature space may not necessarily yield significant performance
improvement. The only way to know this is to try it!
Extracting from a text a list of n-gram can be easily accomplished with function `ngram()`:
```
from nltk.util import ngrams
bigrams = ngrams(reuters.words(), n = 2)
fdbigram = FreqDist(bigrams)
fdbigram.most_common()
```
Collocations are expressions of multiple words that commonly co-occur.
>Finding collocations requires first calculating the frequencies of words and
their appearance in the context of other words. Often the collection of words
will then requiring filtering to only retain useful content terms. Each ngram
of words may then be scored according to some association measure, in order
to determine the relative likelihood of each ngram being a collocation. (Quoted from [here](http://www.nltk.org/_modules/nltk/collocations.html))
For example, to extract bigram collocations, we can firstly extract bigrams then get the commonly co-occurring ones by ranking the bigrams by some measures. A commonly used measure is [Pointwise Mutual Information](https://en.wikipedia.org/wiki/Pointwise_mutual_information) (PMI). The following code will find the best 50 bigrams using the PMI scores.
```
bigram_measures = nltk.collocations.BigramAssocMeasures()
finder = nltk.collocations.BigramCollocationFinder.from_words(reuters.words())
finder.nbest(bigram_measures.pmi, 50)
```
The `collocations` module implements a number of measures to score collocations or other associations.
They include Student's t test, Chi-Square, likelihood ratios, PMI and so on.
Here we used PMI scores for finding bigrams.
Please read [2] for a detailed tutorial on finding collocations with NLTK.
If you would like to know more about collocations, please refer to [3].
## Part 5. Summary
This chapter has show you how to
* generate vocabulary be further exploring the tokenized text with some simple statistics.
* convert unstructured text to structured form using the bag-of-words model
* compute TF-IDF
* extract words in specific lexical categories, n-grams and collocations.
## Part 6. Reading Materials
1. "[Categorizaing and Tagging Words](http://www.nltk.org/book/ch05.html)",
Chapter 5 of "Natural Language Processing with Python".
2. "[Collocations](http://www.nltk.org/howto/collocations.html)": An NTLK tutorial on how to extract collocations 📖 .
3. "[Collocations](http://nlp.stanford.edu/fsnlp/promo/colloc.pdf)": An introduction to collocation by Manning and Schutze.
## Part 7 . Exercises
2. We have shown you how to generate frequency of frequency bar chart with term frequency. Similarly, you can generate the bar chart based on document frequency.
2. Remove short words. There are some very short words in the vocabulary, for example, 'aa', 'ab', 'ad', 'ax', etc.
Write Python code to explore the distribution of word lengths, and remove those words with less than two characters.
3. Write code to tag the Reuters corpus with the Penn Treebank tag set, find the top 10 most common tags, nouns, and verbs.
2. There might be some text analysis tasks where the binary occurrence markers might be enough.
Please modify the CountVectorizer code to generate binary vectors for all the Reuters articles.
2. We have shown you how to generate feature vectors from raw text. As we mentioned in section 3, you can actually customise the two vectorizer classes by specifying, for example, the tokenizer and stopword list. So try
to customize either vecotorizer so that it can carry out all the steps in section 1.
| github_jupyter |
# Análise Preditiva Avançada
## Trabalho Individual
- **Curso:** FGV MBA - Business Analytics e Big Data
- **Disciplina:** Análise Preditiva Avançada
- **Professor:** Hitoshi Nagano e Gustavo Mirapalheta
- **Tarefa:** Trabalho Substitutivo de Prova
- **Link para este notebook:** [Kaggle](https://www.kaggle.com/danielferrazcampos/mnist-using-neural-network-random-forest-pt-br)
## Aluno
|Github|Kaggle|Nome
|---|---|---|---|---|
|<a href="https://github.com/DanielFCampos"><img src="https://avatars2.githubusercontent.com/u/31582602?s=460&v=4" title="DanielFCampos" width="40" height="40"></a>|<a href="https://www.kaggle.com/danielferrazcampos"><img src="https://storage.googleapis.com/kaggle-avatars/images/3508055-kg.png" title="DanielFCampos" width="40" height="40"></a>|Daniel Campos|
# Enunciado
- **Instruções** <br>
Turma: MSP 11924-TBABD-T1
Disciplina: Análise Preditiva Avançada (Professores Mirapalheta e Hitoshi)</br>
- **Questão 1:** <br>
Tomando por base o conjunto de dados MNIST, padrão do pacote Keras, defina e elabore uma rede neural sequencial, apresentando os formatos dos tensores de entrada e saída, bem como os tensores intermediários na rede. Treine e teste o modelo de rede neural sequencial desenvolvido, de forma que ele consiga atingir uma precisão de pelo menos 97% no teste, no reconhecimento das imagens de números escritos à mão livre. Compare o desempenho da rede neural no treino utilizando dados normalizados e não normalizados após 50 épocas de treino. Aumente o número de camadas internas da rede neural e determine se isto melhora ou não a qualidade dos resultados no teste. Qual o número de camadas que você consideraria ideal?</br>
- **Questão 2:** <br>
Resolva o mesmo problema da Questão 1 utilizando o algoritmo Random Forests. Tentem trabalhar a hiperparametrização para aumento do desempenho. Compare o seu melhor resultado com o resultado obtido na Questão 1 e comente.</br>
- **Data de Entrega** <br>
8 de junho de 2020 23:59</br>
# Resolução
### Carregar bibliotecas necessárias
```
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import time
import random
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from mlxtend.plotting import plot_confusion_matrix
from hyperopt import hp, fmin, tpe, rand, STATUS_OK, Trials
SEED = 42
random.seed(SEED)
np.random.seed(SEED)
tf.random.set_seed(SEED)
```
### Carregar funções customizadas
```
def pred(model, x_test):
pred_prob = model.predict(x_test)
pred = np.argmax(pred_prob, axis = 1)
return pred
def plot_confusion_mtx(model, x_test, y_test, plot_tittle):
pred_prob = model.predict(x_test)
pred = np.argmax(pred_prob, axis = 1)
CM = confusion_matrix(y_test, pred)
plot_confusion_matrix(conf_mat = CM, figsize = (16, 8))
plt.title(plot_tittle)
plt.xticks(range(10), [0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
plt.yticks(range(10), [0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
plt.show()
def plot_confusion_mtx2(model, x_test, y_test, plot_tittle):
pred= model.predict(x_test)
CM = confusion_matrix(y_test, pred)
plot_confusion_matrix(conf_mat = CM, figsize = (16, 8))
plt.title(plot_tittle)
plt.xticks(range(10), [0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
plt.yticks(range(10), [0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
plt.show()
```
### Carregar e fazer o *reshape* do *dataset*
```
(trainX, trainy), (testX, testy) = mnist.load_data()
reshaped_trainX_unscaled = trainX.reshape([trainX.shape[0], -1]).astype('float32')
reshaped_testX_unscaled = testX.reshape([testX.shape[0], -1]).astype('float32')
scaler = MinMaxScaler()
reshaped_trainX = scaler.fit_transform(reshaped_trainX_unscaled)
reshaped_testX = scaler.fit_transform(reshaped_testX_unscaled)
encoded_trainy = to_categorical(trainy)
encoded_testy = to_categorical(testy)
print('\nFormato train dataset:\t%s\nTrain dataset reshaped:\t%s\nFormato labels dataset:\t%s' % (trainX.shape, reshaped_trainX.shape, trainy.shape))
print('\nFormato test dataset:\t%s\nTest dataset reshaped:\t%s\nFormato labels dataset:\t%s' % (testX.shape, reshaped_testX.shape, testy.shape))
```
### Explorar conteúdo
```
plt.rcParams.update({'font.size': 16})
fig = plt.figure(figsize = (6, 6))
columns = 4
rows = 3
for i in range(1, columns * rows + 1):
rnd = np.random.randint(0, len(trainX))
img = trainX[rnd]
fig.add_subplot(rows, columns, i)
plt.title(trainy[rnd])
plt.axis('off')
plt.imshow(img, cmap='gray')
plt.show()
```
# Redes Neurais
### Rede Neural Simples com dados normalizados
```
NN = Sequential(name = 'Simple_NN')
NN.add(Dense(512, input_dim=784, activation='relu', name='input_layer'))
NN.add(Dense(10, activation='softmax', name='output_layer'))
print("input shape ",NN.input_shape)
print("output shape ",NN.output_shape)
NN.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
NN.summary()
start = time.time()
history = NN.fit(reshaped_trainX, encoded_trainy, epochs=50, batch_size=64, verbose=0)
predicted = pred(NN, reshaped_testX)
elapsed_time = time.time() - start
NN_acc = accuracy_score(testy, predicted)
NN_time = elapsed_time
print("O treinamento levou %.0f segundos.\nAcurácia: %.4f" % (elapsed_time, NN_acc))
print ("Reporte de classificação:\n")
print(classification_report(testy, predicted))
plot_confusion_mtx(NN, reshaped_testX, testy, 'Rede Neural Simples')
```
### Rede Neural Simples com dados não-normalizados
```
start = time.time()
history = NN.fit(reshaped_trainX_unscaled, encoded_trainy, epochs=50, batch_size=64, verbose=0)
predicted = pred(NN, reshaped_testX_unscaled)
elapsed_time = time.time() - start
NN_acc_ns = accuracy_score(testy, predicted)
NN_time_ns = elapsed_time
print("O treinamento levou %.0f segundos.\nAcurácia: %.4f" % (elapsed_time, NN_acc_ns))
print ("Reporte de classificação:\n")
print(classification_report(testy, predicted))
plot_confusion_mtx(NN, reshaped_testX, testy, 'Rede Neural Simples (Dados Não-Normalizados)')
```
### Rede Neural Profunda com dados normalizados
```
DNN = Sequential(name = 'Deep_NN')
DNN.add(Dense(512, input_dim=784, activation='relu', name='input_layer'))
DNN.add(Dense(256, activation='relu', name='hidden_layer1'))
DNN.add(Dense(128, activation='relu', name='hidden_layer2'))
DNN.add(Dense(64, activation='relu', name='hidden_layer3'))
DNN.add(Dense(32, activation='relu', name='hidden_layer4'))
DNN.add(Dense(10, activation='softmax', name='output_layer'))
print("input shape ",DNN.input_shape)
print("output shape ",DNN.output_shape)
DNN.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
DNN.summary()
start = time.time()
history = DNN.fit(reshaped_trainX, encoded_trainy, epochs=50, batch_size=64, verbose=0)
predicted = pred(DNN, reshaped_testX)
elapsed_time = time.time() - start
DNN_acc = accuracy_score(testy, predicted)
DNN_time = elapsed_time
print("O treinamento levou %.0f segundos.\nAcurácia: %.4f" % (elapsed_time, DNN_acc))
print ("Reporte de classificação:\n")
print(classification_report(testy, predicted))
plot_confusion_mtx(DNN, reshaped_testX, testy, 'Rede Neural Profunda')
```
# Random Forest
### Classificador Random Forest com parâmetros basais
```
rf = RandomForestClassifier(max_depth = 5,
max_features = 5,
n_estimators = 50,
criterion = "entropy",
random_state = SEED,
n_jobs=-1)
start = time.time()
rf.fit(reshaped_trainX, trainy)
predicted = rf.predict(reshaped_testX)
elapsed_time = time.time() - start
rf_acc = accuracy_score(testy, predicted)
rf_time = elapsed_time
print("O treinamento levou %.0f segundos para os parâmetros padrão.\nAcurácia: %.4f" % (elapsed_time, rf_acc))
print ("Reporte de classificação:\n")
print(classification_report(testy, predicted))
plot_confusion_mtx2(rf, reshaped_testX, testy, 'Random Forest Basal')
```
### Classificador Random Forest com parâmetros otimizados
```
space = {'max_depth': hp.quniform('max_depth', 1, 100, 1),
'max_features': hp.quniform('max_features', 1, 50, 1),
'n_estimators': hp.quniform('n_estimators', 25, 500, 5),
'criterion': hp.choice('criterion', ["gini", "entropy"])}
def rf_tuning(space):
global best_score, best_rf_model
clf = RandomForestClassifier(max_depth = int(space['max_depth']),
max_features = int(space['max_features']),
n_estimators = int(space['n_estimators']),
criterion = space['criterion'], n_jobs=-1, random_state = SEED)
clf.fit(reshaped_trainX, trainy)
pred = clf.predict(reshaped_testX)
accuracy = 1-accuracy_score(testy, pred)
if (accuracy < best_score):
best_score = accuracy
best_rf_model = clf
return {'loss': accuracy, 'status': STATUS_OK }
trials = Trials()
start = time.time()
neval = 50
best_score = 1.0
best_rf_model = []
best = fmin(fn = rf_tuning,
space = space,
algo = tpe.suggest,
max_evals = neval,
trials = trials,
rstate = np.random.RandomState(SEED))
elapsed_time = time.time() - start
rf_optim_acc = (1-best_score)
rf_optim_time = elapsed_time
print("A otimização de parâmetros levou %.0f segundos para %d rodadas.\nAcurácia: %.4f\n\nParâmetros ótimos encontrado:\n%s" % (elapsed_time, neval, rf_optim_acc, best))
predicted = best_rf_model.predict(reshaped_testX)
print ("Reporte de classificação:\n")
print(classification_report(testy, predicted))
plot_confusion_mtx2(best_rf_model, reshaped_testX, testy, 'Random Forest Otimizado')
```
# Conclusão
```
print("Comparação das acurácias\n\nRede Neural Simples:\t\t\t%.2f%%\nRede Neural Simples (não norm.):\t%.2f%%\nRede Neural Profunda:\t\t\t%.2f%%\nRandom Forest Basal:\t\t\t%.2f%%\nRandom Forest Otimizado:\t\t%.2f%%" % (NN_acc*100, NN_acc_ns*100, DNN_acc*100, rf_acc*100, rf_optim_acc*100))
print("\nComparação dos tempos\n\nRede Neural Simples:\t\t\t%.0f s\nRede Neural Simples (não norm.):\t%.0f s\nRede Neural Profunda:\t\t\t%.0f s\nRandom Forest Basal:\t\t\t%.0f s\nRandom Forest Otimizado:\t\t%.0f s" % (NN_time, NN_time_ns, DNN_time, rf_time, rf_optim_time))
```
- **Respostas a Questão 1:** <br>
* Tomando por base o conjunto de dados MNIST, padrão do pacote Keras, defina e elabore uma rede neural sequencial, apresentando os formatos dos tensores de entrada e saída, bem como os tensores intermediários na rede. <span style="color:red">Esta questão foi respondida com o uso dos comandos *input_shape*, *output_shape* e *summary* na grande seção **Redes Neurais**. Em todas as redes neurais criadas usamos um vetor unidimensional de 784 posições (correspondente ao números de pixels das imagens: 28x28) como entrada e 10 vetores de saída para a classificação *multi-label* do problema (números de 0 a 9). O comando *summary* mostra todas as camadas ocultas usadas (Rede Neural Simples com uma única camada com 512 neurônios + camadas de input/output e Rede Neural Profunda com 6 camadas com 512, 256, 128, 64, 32 e 16 neurônios cada + camadas de input/output).
* Treine e teste o modelo de rede neural sequencial desenvolvido, de forma que ele consiga atingir uma precisão de pelo menos 97% no teste, no reconhecimento das imagens de números escritos à mão livre. Compare o desempenho da rede neural no treino utilizando dados normalizados e não normalizados após 50 épocas de treino. <span style="color:red">Os dados normalizados não apresentaram ganho/perda expressiva ao modelo apesar de esperarmos que houvesse uma melhora. Entretando, vale ressaltar que temos os dados todos em mesma escala (0 a 255) e o o maior prejuízo acontece quando temos *features* do modelo com mínimos e máximos muito diferentes entre si (fato tal que não acontece nesse exemplo).
* Aumente o número de camadas internas da rede neural e determine se isto melhora ou não a qualidade dos resultados no teste. <span style="color:red">O número de camadas aumenta marginalmente a precisão do modelo com um custo computacional bastante maior (+35-40% no tempo de processamento para o nosso caso).
* Qual o número de camadas que você consideraria ideal? <span style="color:red">Pensando em redes neurais não convolucionais, acredito que uma única camada resolve o problema com uma acurácia bastante satisfatória e com um tempo de processamento reduzido. Entretanto, é sabido que a adição de camadas convolucionais melhora bastante o resultado de redes neurais aplicadas a imagens, portanto, se fôssemos adicionar camadas, o ideal seria adicionar camadas convolucionais antes das camadas sequenciais não-convolucionadas.
<br><br>
- **Respostas a Questão 2:** <br>
* Resolva o mesmo problema da Questão 1 utilizando o algoritmo Random Forests. Tentem trabalhar a hiperparametrização para aumento do desempenho. <span style="color:red">O modelo de Random Forest foi trabalhado de duas maneiras: parâmetros escolhidos manualmente e fazendo uma otimização Bayesiana dos parâmetros com o pacote hyperopt.
* Compare o seu melhor resultado com o resultado obtido na Questão 1 e comente. <span style="color:red">Mesmo com os parâmetros ótimos obtidos pelo hyperopt, o modelo Random Forest não foi capaz de superar os modelos de redes neurais. Além disso, o tempo para a otimização foi muito maior do que o tempo de treinamento das redes neurais profundas deste exemplo. Entretanto, a simplicidade do modelo nos traz bons resultados com parâmetros bastante simples e tempo de treino curto, sendo uma alternativa viável para uma primeira abordagem ao problema.
| github_jupyter |
# Parte 03
Nessa parte vamos trabalhar com um cenário mais complexo. Em particular, iremos treinar um conjunto de modelos [Arima](https://en.wikipedia.org/wiki/Autoregressive_integrated_moving_average). O conjunto de dados que utilizaremos é uma matriz multidimensional com 5 dimensões: **(grupo-série, instante-de-tempo, localização-x, localização-y, valor)**. Note que para cada tripla distinta **(grupo-de-série, localização-x, localização-y)** há uma série temporal com a temperatura em 10 instantes de tempo. Dito isto, os modelos a serem treinados predirão o décimo instante de tempo a partir dos nove anteriores.
Como no exemplo anterior, também particionaremos o conjunto de dados. Entretanto, nesse caso particionamos o dataset ao longo das dimensões localização-x e localização-y.
```
%load_ext autoreload
%autoreload 2
import h5py
import matplotlib.pyplot as plt
import numpy as np
import random as rn
import seaborn as sns
import tensorflow as tf
import os
# Reprodutibilidade
import numpy as np
import random as rn
import tensorflow as tf
seed = 32
rn.seed(seed)
np.random.seed(seed)
tf.random.set_seed(seed)
# Necessário mudar o diretório de trabalho para o nível mais acima
if not 'notebooks' in os.listdir('.'):
current_dir = os.path.abspath(os.getcwd())
parent_dir = os.path.dirname(current_dir)
os.chdir(parent_dir)
from src.model import Arima, ModelFactory
from src.util import plot_heatmap, temporal_train_val_test2, factor_number
sns.set_context('notebook')
sns.set_style('whitegrid')
sns.set_palette(sns.color_palette("Paired"))
tf.get_logger().setLevel('ERROR')
# Carregamento do dataset de temperatura
dataset_path = 'data/tiny-dataset.hdf5'
with h5py.File(dataset_path, 'r') as in_:
data = in_['real'][...]
print('Shape:', data.shape)
# O dataset de temperatura é dividido em x (primeiros 9 instantes) e y (último instante).
x = data[:, :-1]
y = data[:, 1:]
num_models = 25
test_size = .2
val_size = .3
# Indica quais dimensões devem servir para o particionamento (localização-x e localização-y)
split_axis = (2, 3)
model_name_prefix = 'arima'
output_dir = os.path.join(parent_dir, 'saved_models_arima')
```
**Na célula abaixo é instanciada uma fábrica de modelos. A fábrica ('model_factory') ficará responsável por:**
1. Efetuar a partição de x ao longo das dimensões de localização.
3. Treinar cada modelo na sua devida partição;
4. Treinar um modelo em todo o domínio;
5. Salvar os modelos;
6. Salvar os dados.
```
model_config = dict(u=0, optimizer=tf.keras.optimizers.RMSprop(),
early_stopping_min_delta=1e-8,early_stopping_patience=30,epochs=150)
model_factory = ModelFactory(model_class=Arima, x=x, f_x=y, num_models=num_models,
test_size=test_size, val_size=val_size,
model_name_prefix=model_name_prefix,
model_kwargs=model_config,
train_val_test_splitter=temporal_train_val_test2,
x_split_axis=split_axis, y_split_axis=split_axis)
model_factory.build_models()
model_factory.fit_models()
info = model_factory.get_metric_info()
model_factory.save_models(output_dir)
model_factory.save_data(output_dir, info)
fig, ax = plt.subplots()
fig.set_size_inches(15, 9)
matrix = info['mean_absolute_error']
plot_heatmap(matrix=matrix, title=f'Heatmap for MSE', x_label='Model ID', y_label='Split ID', ax=ax,
heatmap_kwargs=dict(linewidths=.1, cmap='Blues', annot=True, fmt='.2f'))
fig.tight_layout()
```
| github_jupyter |
# Ejecutando consultas DAX sobre Power BI desde .NET 5
## Instalar la biblioteca ADOMD.NET
```
#r "nuget: Microsoft.AnalysisServices.AdomdClient.NetCore.retail.amd64, 19.16.3"
```
## Crear la cadena de conexión con los parámetros enviados desde Power BI
```
var pbiServer = Environment.GetEnvironmentVariable("PBI_SERVER");
var pbiDatabase = Environment.GetEnvironmentVariable("PBI_DB");
display($"Servidor: {pbiServer}");
display($"Base de datos: {pbiDatabase}");
var connectionString = $"Provider=MSOLAP;Data Source={pbiServer};Initial Catalog={pbiDatabase};";
```
## Ejecutar una consulta DAX que devuelve un sólo valor
```
using Microsoft.AnalysisServices.AdomdClient;
var connection = new AdomdConnection(connectionString);
connection.Open();
var command = connection.CreateCommand();
command.CommandText = "EVALUATE { [Unidades Vendidas] }";
var reader = command.ExecuteReader();
long unidadesVendidas;
if (reader.Read() && reader[0] != null)
unidadesVendidas = reader.GetInt64(0);
reader.Close();
connection.Close();
display($"Total de Unidades Vendidas: {unidadesVendidas}");
```
## Ejecutar una consulta DAX con parámetros
```
using Microsoft.AnalysisServices.AdomdClient;
var connection = new AdomdConnection(connectionString);
connection.Open();
var command = connection.CreateCommand();
command.CommandText = @"
EVALUATE
{
CALCULATE(
[Unidades Vendidas],
'Calendario'[Año] = @year
)
}";
command.Parameters.Add(new AdomdParameter("year",2020));
var reader = command.ExecuteReader();
long unidadesVendidas;
if (reader.Read() && reader[0] != null)
unidadesVendidas = reader.GetInt64(0);
reader.Close();
connection.Close();
display($"Total de Unidades Vendidas: {unidadesVendidas}");
```
## Ejecutar una consulta DAX con ExecuteReader que devuelve una tabla
```
using Microsoft.AnalysisServices.AdomdClient;
var connection = new AdomdConnection(connectionString);
connection.Open();
var command = connection.CreateCommand();
command.CommandText = "EVALUATE 'Vendedores'";
var reader = command.ExecuteReader();
var isFirstRow = true;
while (reader.Read())
{
if (isFirstRow)
{
List<string> header = new();
for (var i = 0; i < reader.FieldCount; i++)
header.Add(reader.GetName(i));
display(string.Join('\t',header.ToArray()));
isFirstRow = false;
}
List<string> row = new();
for (var i = 0; i < reader.FieldCount; i++)
row.Add(reader[i].ToString());
display(string.Join('\t',row.ToArray()));
}
reader.Close();
connection.Close();
```
## Utilizar un DataFrame
https://devblogs.microsoft.com/dotnet/an-introduction-to-dataframe/
```
#r "nuget:Microsoft.Data.Analysis,0.2.0"
```
### Registrar un "Formatter" para visualizar mejor el DataFrame
```
using Microsoft.Data.Analysis;
using Microsoft.AspNetCore.Html;
Formatter.Register<DataFrame>((df, writer) =>
{
var headers = new List<IHtmlContent>();
headers.Add(th(i("index")));
headers.AddRange(df.Columns.Select(c => (IHtmlContent) th(c.Name)));
var rows = new List<List<IHtmlContent>>();
var take = 40;
for (var i = 0; i < Math.Min(take, df.Rows.Count); i++)
{
var cells = new List<IHtmlContent>();
cells.Add(td(i));
foreach (var obj in df.Rows[i])
{
cells.Add(td(obj));
}
rows.Add(cells);
}
var t = table(
thead(
headers),
tbody(
rows.Select(
r => tr(r))));
writer.Write(t);
}, "text/html");
```
### Clases auxiliares para adiconar columnas a un DataFrame
### Crear un método que ejecuta una expresión DAX y devuelve un DataFrame
```
using Microsoft.Data.Analysis;
using Microsoft.AnalysisServices.AdomdClient;
using System.Linq;
DataFrame ExecuteDax(string dax)
{
var connection = new AdomdConnection(connectionString);
connection.Open();
var command = connection.CreateCommand();
command.CommandText = dax;
var reader = command.ExecuteReader();
List<StringDataFrameColumn> dfColumnList = new();
var isFirstRow = true;
while (reader.Read())
{
if (isFirstRow)
{
List<string> header = new();
for (var i = 0; i < reader.FieldCount; i++) {
var dfColumn = new StringDataFrameColumn(reader.GetName(i),0);
dfColumnList.Add(dfColumn);
}
isFirstRow = false;
}
List<string> row = new();
for (var i = 0; i < reader.FieldCount; i++)
{
var dfColumn = dfColumnList[i];
dfColumn.Append(reader[i].ToString());
}
}
reader.Close();
connection.Close();
return new DataFrame(dfColumnList.ToArray());
}
var df = ExecuteDax("EVALUATE 'Vendedores'");
display(df);
var df = ExecuteDax("EVALUATE {[Unidades Vendidas]}");
display(df);
```
## Más consultas DAX
### Unidades Vendidas por Categoría
```
df = ExecuteDax(
@"EVALUATE
SELECTCOLUMNS (
VALUES(Productos[Categoria]),
""Categoria"",[Categoria],
""Unidades Vendidas"", [Unidades Vendidas]
)"
);
display(df);
```
### Unidades Vendidas por Categoría en orden descendente
```
df = ExecuteDax (
@"EVALUATE
SELECTCOLUMNS (
VALUES(Productos[Categoria]),
""Categoria"",[Categoria],
""Unidades Vendidas"", [Unidades Vendidas]
)
ORDER BY [Unidades Vendidas] DESC"
);
display(df);
```
### Unidades Vendidas por Categoría en el último año y en orden descendente
```
df = ExecuteDax (
@"
DEFINE
VAR maxYear = MAX ( Calendario[Año] )
EVALUATE
CALCULATETABLE (
SELECTCOLUMNS (
VALUES ( Productos[Categoria] ),
""Categoria"", [Categoria],
""Unidades Vendidas"", [Unidades Vendidas]
),
Calendario[Año] = maxYear
)
ORDER BY [Unidades Vendidas] DESC"
);
display(df);
```
| github_jupyter |
# 1 CairoSVG介绍
CairoSVG是一个将SVG1.1转为PNG,PDF, PS格式的转化。SVG算目前火热的图像文件格式了,它的英文全称为Scalable Vector Graphics,意思为可缩放的矢量图形,但是SVG要专用软件才能编辑打开,通过CairSVG我们就能将SVG格式转换为常用的格式。它为类Unix操作系统(至少Linux和macOS)和Windows提供了命令行界面和Python 3.5+库。它是一个开源软件,具有LGPLv3许可。
CairoSVG用Python编写,基于著名的2D图形库Cairo。它在来自W3C测试套件的 SVG样本上进行了测试。它还依赖tinycss2和 cssselect2来应用CSS,并依赖 defusedxml来检测不安全的SVG文件。嵌入式栅格图像由Pillow处理。
CarioSVG仅支持python3,你可以用pip命令安装,安装代码如下:
> pip3 install cairosvg
本文主要使用2.4.2版本,当前版本的CairoSVG至少需要Python 3.5,但不适用于Python2.x。较旧的CairoSVG(1.x)版本可在Python 2.x中使用,但不再受支持。CairoSVG及其依赖项在安装过程中可能需要其他工具,这些工具的名称取决于您使用的操作系统。具体如下:
+ 在Windows上,您必须安装适用于Python和Cairo的Visual C ++编译器
+ 在macOS上,您必须安装cairo和libffi
+ 在Linux上,你必须安装cairo,python3-dev和libffi-dev(名称可能为你的系统版本有所不同)
如果您不知道如何安装这些工具,则可以按照[WeasyPrint安装指南](https://weasyprint.readthedocs.io/en/latest/install.html)中的简单步骤进行操作:安装WeasyPrint还将安装CairoSVG。
# 2 CairoSVG的使用
## 2.1 命令行使用
通过命令行你就可以使用CairoSVG,以下代码能够将当前目录下的image.svg文件转换为image.png文件:
> cairosvg image.svg -o image.png
具体CairoSVG命令行参数如下:
```
cairosvg --help
usage: cairosvg [-h] [-v] [-f {pdf,png,ps,svg}] [-d DPI] [-W WIDTH]
[-H HEIGHT] [-s SCALE] [-u] [--output-width OUTPUT_WIDTH]
[--output-height OUTPUT_HEIGHT] [-o OUTPUT]
input
Convert SVG files to other formats
positional arguments:
input input filename or URL 文件名或者url链接名
optional arguments:
-h, --help show this help message and exit 帮助
-v, --version show program's version number and exit 版本查看
-f {pdf,png,ps,svg} --format {pdf,png,ps,svg} output format 输出格式
-d DPI, --dpi DPI ratio between 1 inch and 1 pixel 输出图像dpi比率设置 DPI比率介于1英寸和1像素之间
-W WIDTH, --width WIDTH width of the parent container in pixels 输入图像宽
-H HEIGHT, --height HEIGHT height of the parent container in pixels 输入图像高
-s SCALE, --scale SCALE output scaling factor 输出图像缩放比例
-u, --unsafe resolve XML entities and allow very large files 解析XML实体
(WARNING: vulnerable to XXE attacks and various DoS) 但是有安全问题
--output-width OUTPUT_WIDTH desired output width in pixels 期望图像输出宽
--output-height OUTPUT_HEIGHT desired output height in pixels 期望图像输出高
-o OUTPUT, --output OUTPUT output filename 图像输出名
```
支持的输出格式是pdf,ps,png和svg(默认为 pdf)。默认output为标准输出。如果提供了输出文件名,则会根据扩展名自动选择格式。这些dpi选项设置像素与实际单位(例如,毫米和英寸)之间的比率(如[规范](https://www.w3.org/TR/SVG11/coords.html)中所述)。可以为SVG文件提供宽度和高度选项来设置容器大小。此外,如果-用作文件名,CairoSVG将从标准输入中读取SVG字符串。
## 2.2 python库使用
CairoSVG为Python 3.5+提供了一个模块。该cairosvg模块提供4个功能:
+ svg转pdf
+ svg转png
+ svg转ps
+ svg2转svg(svg文件切割)
这些函数需要以下命名参数之一:
+ url,URL或文件名
+ file_obj,类似文件的对象
+ bytestring,一个包含SVG的字节字符串
他们还可以接收与命令行选项相对应的这些可选参数:
+ parent_width
+ parent_height
+ dpi
+ scale
+ unsafe
如果write_to提供了参数(文件名或类似文件的对象),则将输出写入此处。否则,该函数将返回一个字节字符串。例如:
> cairosvg.svg2png(url="/path/to/input.svg", write_to="/tmp/output.png")
cairosvg.svg2pdf(file_obj=open("/path/to/input.svg", "rb"), write_to="/tmp/output.pdf")
output = cairosvg.svg2ps(bytestring=open("/path/to/input.svg").read().encode('utf-8'))
使用实例:
``` python
# -*- coding: utf-8 -*-
# 导入cairosvg库
import cairosvg
# svg转pdf
# file_obj输入文件名 write_to输出文件名
cairosvg.svg2pdf(file_obj=open("image.svg", "rb"), write_to="output.pdf")
# svg转png
# file_obj输入文件名 write_to输出文件名 scale输出图像放大倍数
cairosvg.svg2png(file_obj=open("image.svg", "rb"), write_to="d:/output.png",scale=3.0)
```
# 3 参考
+ [CairoSVG官网](https://cairosvg.org/)
| github_jupyter |
# Exploring evaluation DMRL
## Global imports and variables
```
# Import for interactive notebook (see:
# https://ipywidgets.readthedocs.io/en/stable/examples/Using%20Interact.html)
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
from IPython.display import display
from ipywidgets import Layout
# Import to list files in directories
import glob
# Import for regular expressions
import re
# Imports for path operations
import os
import os.path
# For date operations
from datetime import datetime
import pandas as pd
pd.set_option('max_colwidth', -1)
import matplotlib.pyplot as plt
import numpy as np
import configparser
# import jtplot module in notebook
from jupyterthemes import jtplot
# choose which theme to inherit plotting style from
# onedork | grade3 | oceans16 | chesterish | monokai | solarizedl | solarizedd
jtplot.style(theme='onedork')
RESULTS_DIR = "/Users/gomerudo/workspace/thesis_results"
def rettext(text):
return text
def search_in_file(file, pattern):
pattern = re.compile(pattern)
results = []
for i, line in enumerate(open(file)):
for match in re.finditer(pattern, line):
results.append(match.groups())
return results
form_item_layout = Layout(
width="50%"
)
w_resdirs = interactive(
rettext,
# text=sorted(glob.glob("{dir}/[mix-]?[0-9]*".format(dir=RESULTS_DIR))),
text=sorted(glob.glob("{dir}/*".format(dir=RESULTS_DIR))),
layout=form_item_layout
)
```
## Selecting the desired results
```
display(w_resdirs)
```
## Results
```
################################################################################
############ OBTAIN THE FILES AND DIRECTORIES TO QUERY FOR ANALYSIS ############
################################################################################
# Obtain the chosen directory
chosen_dir = w_resdirs.result
# experiments dir
exp_dir = glob.glob("{dir}/experiment*[!.zip]".format(dir=chosen_dir))[0]
# This is a list of all openai dirs, sorted by name (hence, by timestamp)
openai_dirs = sorted(glob.glob("{dir}/openai*[!.zip]".format(dir=exp_dir)))
# A simple DB of experiments and actions_info.csv should be there
dbexp_file = glob.glob("{dir}/db_experiments.csv".format(dir=exp_dir))[0]
ainfo_file = glob.glob("{dir}/actions_info.csv".format(dir=exp_dir))[0]
config_file = glob.glob("{dir}/config*.ini".format(dir=exp_dir))[0]
flog_file = glob.glob("{dir}/sl*".format(dir=chosen_dir))[0]
# Make dataframes for the db of experiments and the actions summary
dbexp_df = pd.read_csv(dbexp_file)
ainfo_df = pd.read_csv(ainfo_file)
# Make de target directory
import os
summaries_dir = "{exp}/summary".format(exp=chosen_dir)
if not os.path.isdir(summaries_dir):
os.mkdir(summaries_dir)
################################################################################
########### BUILD THE RELEVANT DATA FRAMES TO PRINT FOR MAIN SUMMARY ###########
################################################################################
# Try to obtain the current times
running_times = search_in_file(flog_file, ".*\s+(.*)elapsed")
if len(running_times) == len(openai_dirs):
f_running_times = []
for time in running_times:
time_cleansed = time[0].split(".")[0]
f_running_times.append(time_cleansed)
# else:
# prev_timestamp = 0
# f_running_times = []
# for directory in openai_dirs:
# exp_dirname_only = os.path.basename(directory)
# timestamp = os.path.basename(exp_dirname_only.split("-")[1])
# d2 = datetime.strptime(timestamp, "%Y%m%d%H%M%S")
# if prev_timestamp: # 2019 05 29 211533
# d1 = datetime.strptime(prev_timestamp, "%Y%m%d%H%M%S")
# f_running_times.append(str(d2 - d1))
# prev_timestamp = timestamp
# f_running_times.append("NA")
openai_dirs_df = pd.DataFrame(zip(openai_dirs, f_running_times), columns=["Log directory", "Runtime"])
# 4. Search all exceptions
exceptions_all = search_in_file(flog_file, "failed with exception of type.*<(.*)>.*Message.*:\s*(.*)")
n_exceptions = len(exceptions_all)
exceptions_set = set()
for error, message in exceptions_all:
exceptions_set.add(error)
config = configparser.ConfigParser()
_ = config.read(config_file)
```
### Summary
- **Chosen results directory is:** {{chosen_dir}}
- **Full log is available at:** {{flog_file}}
#### Configuration
- **Log Path:** {{config['DEFAULT']['LogPath']}}
- **Environment:** {{config['bash']['Environment']}}
##### Reinforcement Learning
- **Algorithm:** {{config['bash']['Algorithm']}}
- **Policy representation:** {{config['bash']['Network']}}
- **Total number of timestamps:** {{config['bash']['NumTimesteps']}}
- **Number of actions:** {{ainfo_df.shape[0]}}
##### NAS details
- **Config file:** {{config['nasenv.default']['ConfigFile']}}
- **Max Steps:** {{config['nasenv.default']['MaxSteps']}}
- **DB of experiments:** {{config['nasenv.default']['DbFile']}}
- **Dataset Handler:** {{config['nasenv.default']['DatasetHandler']}}
- **Action Space Type:** {{config['nasenv.default']['ActionSpaceType']}}
- **Trainer:** {{config['nasenv.default']['TrainerType']}}
##### Training details
- **Batch size:** {{config['trainer.default']['BatchSize']}}
- **Epochs:** {{config['trainer.default']['NEpochs']}}
- **Distributed:** {{config['trainer.tensorflow']['EnableDistributed']}}
##### Meta-dataset details
- **TFRecordsRootDir:** {{config['metadataset']['TFRecordsRootDir']}}
- **DatasetID:** {{config['metadataset']['DatasetID']}}
#### Individual run directories/time
{{openai_dirs_df}}
#### Errors found in log while building networks
- **Total number of exceptions:** {{n_exceptions}}
{{pd.DataFrame(exceptions_set, columns = ["Error type"])}}
```
def trial_summary(trial_log):
# Read in try catch because the file can be corrupted or might not exist
# try:
# Read the log file
trial_df = pd.read_csv(trial_log)
actions_distribution = [0]*ainfo_df.shape[0]
print("The best id's:")
print(trial_df.sort_values('reward', inplace=False, ascending=False)['composed_id'].drop_duplicates().head())
# CONTROL VARIABLES
# Info for the best architecture
best_architecture = None
best_reward = -1
# Aux for episode control
max_step_ep = 0
best_reward_ep = 0
# History lists
max_step_count_history = []
best_reward_history = []
all_rewards_history = []
# Accumulated rewards per trial
acc_rewards_history = []
acc_reward = 0
# Iterate the log
unique_architectures = set()
for idx, row in trial_df.iterrows():
arch_hash = row['end_state_hashed']
# print("idx", idx, arch_hash)
# Obtain the information information
action_id = int(row['action_id'])
step = int(row['step_count'])
is_valid = bool(row['valid'])
reward = float(row['reward'])
# THIS SECTION IS FOR THE "OVERALL" STATISTICS IN TRIAL
# a) Add information to the distribution of actions
actions_distribution[action_id] += 1
# b) Get the best reward by comparing one by one
if reward > best_reward:
best_reward = reward
best_architecture = arch_hash
# c) History of all rewards in trial
all_rewards_history.append(reward)
# d) Unique architectures
unique_architectures.add(arch_hash)
# THIS SECTION IS FOR THE EPISODE STATISTICS
if step > max_step_ep:
max_step_ep = step
best_reward_ep = reward if reward > best_reward_ep else best_reward_ep
acc_reward += reward
# Otherwise, append the best information we read
else:
max_step_count_history.append(max_step_ep)
best_reward_history.append(best_reward_ep)
acc_rewards_history.append(acc_reward)
max_step_ep = step
best_reward_ep = reward
acc_reward = reward
# except Exception:
# print("Exception")
# pass
# finally:
return {
'actions_distribution': actions_distribution,
'max_step_history': max_step_count_history,
'best_reward_history': best_reward_history,
'all_rewards_history': all_rewards_history,
'best_architecture': best_architecture,
'best_reward': best_reward,
'n_episodes': len(best_reward_history),
'unique_architectures': set(trial_df['end_state_hashed'].unique()),
'acc_rewards_history': acc_rewards_history,
}
# Obtain statistics for each trial
stats = []
for i, openai_dir in enumerate(openai_dirs):
try:
trial_log = sorted(glob.glob("{dir}/play_logs/*".format(dir=openai_dir)))[0]
info_trial = trial_summary(trial_log)
stats.append(info_trial)
except IndexError:
print("Could not read the episode_logs in {}".format(openai_dir))
pass
# Build global statistics for the whole experiment
n_episodes_history = []
unique_architectures = set()
last_length_set_archs = len(unique_architectures)
best_global_architecture = None
best_global_reward = 0
global_best_reward_history = []
global_all_rewards_history = []
global_max_step_history = []
new_archs_history = []
for trial_stats in stats:
# Miscellaneous
n_episodes_history.append(len(trial_stats['best_reward_history']))
unique_architectures.update(trial_stats['unique_architectures'])
new_sampled_architectures = len(unique_architectures) - last_length_set_archs
last_length_set_archs = len(unique_architectures)
new_archs_history.append(new_sampled_architectures)
# Best values
if trial_stats['best_reward'] > best_global_reward:
best_global_reward = trial_stats['best_reward']
best_global_architecture = trial_stats['best_architecture']
# Global histories
global_best_reward_history += trial_stats['best_reward_history']
global_max_step_history += trial_stats['max_step_history']
# print(trial_stats.sort_values('best_reward', inplace=False)['composed_id'].head())
# The distribution of actions
total_n_episodes = sum(n_episodes_history)
# Search for the best architecture
best_architecture_id = "{d}-{h}".format(d=config['metadataset']['DatasetID'], h=best_global_architecture)
print("Best arch is", best_architecture_id)
best_architecture_dbexp = dbexp_df.loc[dbexp_df['dataset-nethash'] == best_architecture_id].iloc[0]
# Export the relevant information into CSVs so that we can plot them in a different tool
# THE ACTIONS DATAFRAME
acts_col = ["action_{a}_count".format(a=i) for i in range(ainfo_df.shape[0])]
actions_count_df = pd.DataFrame()
for i, trial in enumerate(stats):
for action, count in enumerate(trial['actions_distribution']):
actions_count_df = actions_count_df.append(
{
"dataset": config['metadataset']['DatasetID'],
"trial": i+1,
"action_id": "A"+ str(action),
"prop": count/sum(trial['actions_distribution']),
"count": count,
},
ignore_index=True
)
actions_count_df.to_csv("{dir}/actions_dist.csv".format(dir=summaries_dir), index=False)
# The TRIAL short stats
trials_history_df = pd.DataFrame()
for i, nepisodes in enumerate(n_episodes_history):
trials_history_df = trials_history_df.append(
{
"trial": i+1,
"dataset": config['metadataset']['DatasetID'],
"attribute": "nepisodes",
"value": nepisodes,
},
ignore_index=True
)
for i, narchs in enumerate(new_archs_history):
trials_history_df = trials_history_df.append(
{
"trial": i+1,
"dataset": config['metadataset']['DatasetID'],
"attribute": "narchitectures",
"value": narchs,
},
ignore_index=True
)
trials_history_df.to_csv("{dir}/trials_stats.csv".format(dir=summaries_dir), index=False)
### TRY AMBITIOUS STATISTICS
steps_stats_df = pd.DataFrame()
step_count = 0
for i, trial_stats in enumerate(stats):
all_rewards = trial_stats['all_rewards_history']
for step, reward in enumerate(all_rewards):
step_count += 1
steps_stats_df = steps_stats_df.append(
{
"dataset": config['metadataset']['DatasetID'],
"trial": i+1,
"step": step+1,
"acc_step": step_count,
"reward": reward,
},
ignore_index=True
)
steps_stats_df.to_csv("{dir}/steps_stats.csv".format(dir=summaries_dir), index=False)
episodes_stats_df = pd.DataFrame()
ep_count = 0
for i, trial_stats in enumerate(stats):
all_rewards = trial_stats['acc_rewards_history']
all_ep_lengths = trial_stats['max_step_history']
all_best_rewards = trial_stats['best_reward_history']
for ep, reward_length in enumerate(zip(all_rewards, all_ep_lengths, all_best_rewards)):
ep_count += 1
episodes_stats_df = episodes_stats_df.append(
{
"dataset": config['metadataset']['DatasetID'],
"trial": i+1,
"step": ep+1,
"acc_step": ep_count,
"acc_reward": reward_length[0],
"ep_length": reward_length[1],
"best_reward": reward_length[2],
},
ignore_index=True
)
episodes_stats_df.to_csv("{dir}/episodes_stats.csv".format(dir=summaries_dir), index=False)
```
### Global statistics for experiment
- **True number of trials:** {{len(stats)}}
- **Total number of episodes:** {{total_n_episodes}}
- **Number of unique architectures:** {{len(unique_architectures)}}
- **Best architecture in experiment:** {{best_global_architecture}}
- **Best reward in experiment:** {{best_global_reward}}
#### The best architecture
```{{for layer in best_architecture_dbexp['netstring'].split("\n"): print(layer)}}```
**Main information**
- **ID:** {{best_architecture_dbexp['dataset-nethash']}}
- **Index in DB:** {{best_architecture_dbexp.name}}
- **Is valid?** {{best_architecture_dbexp['is_valid']}}
- **Accuracy:** {{best_architecture_dbexp['accuracy']}}
- **Training time (in sec):** {{best_architecture_dbexp['running_time']}}
- **Density:** {{best_architecture_dbexp['density']}}
- **FLOPs:** {{best_architecture_dbexp['flops']}}
```
# 793 cu_birds-7bef9a71004a2735da158263ed7e9f54
# 1104 cu_birds-0df213ce7215d0f999df207f1288cb30
# 918 cu_birds-be2fe4e204a73238fccf0a50ee8f710e
# 18 cu_birds-7cc5c15a255eb7c93cb40bcf10b66e13
# 1392 cu_birds-6adf50bf4aced462e2fb46a5aeca05f5
# 783 aircraft-3fd693dfbd6f1e0994d05ee5924d43fc
# 541 aircraft-d40ae2fbbb127307795ffb942c7ed3ee
# 701 aircraft-522c535d9f0908cebe744bf42066441a
# 1970 aircraft-abbaf5319a2f84a3994da6dff47702bf
# 403 aircraft-e73c4862ce4b285df70888d86598406f
arch_id = 'aircraft-522c535d9f0908cebe744bf42066441a'
arch_c = dbexp_df.loc[dbexp_df['dataset-nethash'] == arch_id].iloc[0]
```
### Global statistics for experiment
- **True number of trials:** {{len(stats)}}
- **Total number of episodes:** {{total_n_episodes}}
- **Number of unique architectures:** {{len(unique_architectures)}}
#### The selected architecture
```{{for layer in arch_c['netstring'].split("\n"): print(layer)}}```
**Main information**
- **ID:** {{arch_c['dataset-nethash']}}
- **Index in DB:** {{arch_c.name}}
- **Is valid?** {{arch_c['is_valid']}}
- **Accuracy:** {{arch_c['accuracy']}}
- **Training time (in sec):** {{arch_c['running_time']}}
- **Density:** {{arch_c['density']}}
- **FLOPs:** {{arch_c['flops']}}
| github_jupyter |
```
# Visualization of the KO+ChIP Gold Standard from:
# Miraldi et al. (2018) "Leveraging chromatin accessibility for transcriptional regulatory network inference in Th17 Cells"
# TO START: In the menu above, choose "Cell" --> "Run All", and network + heatmap will load
# NOTE: Default limits networks to TF-TF edges in top 1 TF / gene model (.93 quantile), to see the full
# network hit "restore" (in the drop-down menu in cell below) and set threshold to 0 and hit "threshold"
# You can search for gene names in the search box below the network (hit "Match"), and find regulators ("targeted by")
# Change "canvas" to "SVG" (drop-down menu in cell below) to enable drag interactions with nodes & labels
# Change "SVG" to "canvas" to speed up layout operations
# More info about jp_gene_viz and user interface instructions are available on Github:
# https://github.com/simonsfoundation/jp_gene_viz/blob/master/doc/dNetwork%20widget%20overview.ipynb
# directory containing gene expression data and network folder
directory = "."
# folder containing networks
netPath = 'Networks'
# network file name
networkFile = 'ChIP_A17_KOall_ATh_bias50_maxComb_sp.tsv'
# title for network figure
netTitle = 'ChIP/ATAC(Th17)+KO+ATAC(Th), bias = 50, max-combined TFA'
# name of gene expression file
expressionFile = 'Th0_Th17_48hTh.txt'
# column of gene expression file to color network nodes
rnaSampleOfInt = 'Th17(48h)'
# edge cutoff -- for Inferelator TRNs, corresponds to signed quantile (rank of edges in 15 TFs / gene models),
# increase from 0 --> 1 to get more significant edges (e.g., .33 would correspond to edges only in 10 TFs / gene
# models)
edgeCutoff = .93
import sys
if ".." not in sys.path:
sys.path.append("..")
from jp_gene_viz import dNetwork
dNetwork.load_javascript_support()
# from jp_gene_viz import multiple_network
from jp_gene_viz import LExpression
LExpression.load_javascript_support()
# Load network linked to gene expression data
L = LExpression.LinkedExpressionNetwork()
L.show()
# Load Network and Heatmap
L.load_network(directory + '/' + netPath + '/' + networkFile)
L.load_heatmap(directory + '/' + expressionFile)
N = L.network
N.set_title(netTitle)
N.threshhold_slider.value = edgeCutoff
N.apply_click(None)
N.draw()
# Add labels to nodes
N.labels_button.value=True
# Limit to TFs only, remove unconnected TFs, choose and set network layout
N.restore_click()
N.tf_only_click()
N.connected_only_click()
N.layout_dropdown.value = 'fruchterman_reingold'
N.layout_click()
# Interact with Heatmap
# Limit genes in heatmap to network genes
L.gene_click(None)
# Z-score heatmap values
L.expression.transform_dropdown.value = 'Z score'
L.expression.apply_transform()
# Choose a column in the heatmap (e.g., 48h Th17) to color nodes
L.expression.col = rnaSampleOfInt
L.condition_click(None)
# Switch SVG layout to get line colors, then switch back to faster canvas mode
N.force_svg(None)
```
| github_jupyter |
# Segmenting and Clustering Neighborhoods in Toronto
Before we get the data and start exploring it, let's download all the dependencies that we will need.
```
import numpy as np # library to handle data in a vectorized manner
import pandas as pd # library for data analsysis
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import json # library to handle JSON files
#!pip install geopy
from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
import requests # library to handle requests
from pandas.io.json import json_normalize # tranform JSON file into a pandas dataframe
# Matplotlib and associated plotting modules
import matplotlib.cm as cm
import matplotlib.colors as colors
# import k-means from clustering stage
#!pip install -U scikit-learn scipy matplotlib
from sklearn.cluster import KMeans
#!conda install -c conda-forge folium=0.5.0 --yes # uncomment this line if you haven't completed the Foursquare API lab
import folium # map rendering library
print('Libraries imported.')
```
#### Importing lib to get data in required format
```
import requests
website_url = requests.get('https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M').text
```
## Question 1
#### Use the BeautifulSoup package or any other way you are comfortable with to transform the data in the table on the Wikipedia page into the above pandas dataframe
```
#!pip install BeautifulSoup4
from bs4 import BeautifulSoup
soup = BeautifulSoup(website_url,'html.parser')
print(soup.prettify())
```
#### By observation we can see that the tabular data is availabe in table and belongs to class="wikitable sortable"So let's extract only table¶
```
My_table = soup.find('table',{'class':'wikitable sortable'})
My_table
print(My_table.tr.text)
headers="Postcode,Borough,Neighbourhood"
table1=""
for tr in My_table.find_all('tr'):
row1=""
for tds in tr.find_all('td'):
row1=row1+","+tds.text
table1=table1+row1[1:]
print(table1)
```
#### Writing our data into as .csv file for further use
```
file=open("toronto.csv","wb")
#file.write(bytes(headers,encoding="ascii",errors="ignore"))
file.write(bytes(table1,encoding="ascii",errors="ignore"))
```
#### Converting into dataframe and assigning columnnames
```
import pandas as pd
df = pd.read_csv('toronto.csv',header=None)
df.columns=["Postalcode","Borough","Neighbourhood"]
df.head(10)
```
#### Only processing the cells that have an assigned borough. Ignoring the cells with a borough that is Not assigned. Droping row where borough is "Not assigned"
```
indexNames = df[ df['Borough'] =='Not assigned'].index
df.drop(indexNames , inplace=True)
df.head(10)
```
#### If a cell has a borough but a Not assigned neighborhood, then the neighborhood will be the same as the borough
```
df.loc[df['Neighbourhood'] =='Not assigned' , 'Neighbourhood'] = df['Borough']
df.head(10)
```
#### Rows will be same postalcode will combined into one row with the neighborhoods separated with a comma
```
result = df.groupby(['Postalcode','Borough'], sort=False).agg( ', '.join)
df_new=result.reset_index()
df_new.head(15)
df_new.shape
```
| github_jupyter |
# Example to calculate photon-ALP oscillations from NGC 1275
This notebook demonstrates how to calculate the photon-ALP transition probability for NGC 1275, the central AGN of the Perseus cluster. The assumed B-field environments are the same as in Ajello et al. (2016), http://inspirehep.net/record/1432667, and include the cluster field and the magnetic field of the Milky Way.
```
from gammaALPs.core import Source, ALP, ModuleList
from gammaALPs.base import environs, transfer
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patheffects import withStroke
from ebltable.tau_from_model import OptDepth
from astropy import constants as c
%matplotlib inline
```
### Set the ALP
Initialize an ALP object, that stores the ALP mass $m$ (in neV) and the coupling $g$ (in $10^{-11}\mathrm{GeV}^{-1}$).
```
m, g = 1.,1.
alp = ALP(m,g)
```
## Set the source
Set the source properties (redshift and sky coordinates) in the ```Source``` containier
```
ngc1275 = Source(z = 0.017559, ra = '03h19m48.1s', dec = '+41d30m42s')
print (ngc1275.z)
print (ngc1275.ra, ngc1275.dec)
print (ngc1275.l, ngc1275.b)
```
### Init the module list
Initialize the list of transfer modules that will store the different magnetic field environments.
Energies are supplied in GeV as ```numpy.ndarray```
```
EGeV = np.logspace(1.,3.5,250)
```
Now initialize the initial photon polarization. Since we are dealing with a gamma-ray source, no ALPs are initially present in the beam (third diagonal element is zero). The polarization density matrix is normalized such that its trace is equal to one, $\mathrm{Tr}(\rho_\mathrm{in}) = 1$.
```
pin = np.diag((1.,1.,0.)) * 0.5
m = ModuleList(alp, ngc1275, pin = pin, EGeV = EGeV)
```
### Add modules:
Now we add propagation modules for the cluster, the EBL, and the Galactic magnetic field.
```
m.add_propagation("ICMGaussTurb",
0, # position of module counted from the source.
nsim = 10, # number of random B-field realizations
B0 = 10., # rms of B field
n0 = 39., # normalization of electron density
n2 = 4.05, # second normalization of electron density, see Churazov et al. 2003, Eq. 4
r_abell = 500., # extension of the cluster
r_core = 80., # electron density parameter, see Churazov et al. 2003, Eq. 4
r_core2 = 280., # electron density parameter, see Churazov et al. 2003, Eq. 4
beta = 1.2, # electron density parameter, see Churazov et al. 2003, Eq. 4
beta2= 0.58, # electron density parameter, see Churazov et al. 2003, Eq. 4
eta = 0.5, # scaling of B-field with electron denstiy
kL = 0.18, # maximum turbulence scale in kpc^-1, taken from A2199 cool-core cluster, see Vacca et al. 2012
kH = 9., # minimum turbulence scale, taken from A2199 cool-core cluster, see Vacca et al. 2012
q = -2.80, # turbulence spectral index, taken from A2199 cool-core cluster, see Vacca et al. 2012
seed=0 # random seed for reproducability, set to None for random seed.
)
m.add_propagation("EBL",1, model = 'dominguez') # EBL attenuation comes second, after beam has left cluster
m.add_propagation("GMF",2, model = 'jansson12', model_sum = 'ASS') # finally, the beam enters the Milky Way Field
```
List the module names:
```
print(m.modules.keys())
```
We can also change the ALP parameters before running the modules:
```
m.alp.m = 30.
m.alp.g = 0.5
```
### Run all modules
Now we run the modules. If ```multiprocess``` key word is larger than two, this will be split onto multiple cores with python's ```multiprocess``` module.
The ```px,py,pa``` variables contain the mixing probability into the two photon polarization states (x,y) and into the axion state (a).
```
px,py,pa = m.run(multiprocess=2)
```
## Plot the output
```
pgg = px + py # the total photon survival probability
print (pgg.shape)
print (np.min(np.median(pgg, axis = 0)))
print (np.min(np.max(pgg, axis = 0)))
effect = dict(path_effects=[withStroke(foreground="w", linewidth=2)])
for p in pgg: # plot all realizations
plt.semilogx(m.EGeV, p)
plt.xlabel('Energy (GeV)')
plt.ylabel('Photon survival probability')
plt.legend(loc = 0, fontsize = 'medium')
plt.annotate(r'$m_a = {0:.1f}\,\mathrm{{neV}}, g_{{a\gamma}} = {1:.1f} \times 10^{{-11}}\,\mathrm{{GeV}}^{{-1}}$'.format(m.alp.m,m.alp.g),
xy = (0.95,0.1), size = 'x-large', xycoords = 'axes fraction', ha = 'right',**effect)
plt.gca().set_xscale('log')
plt.gca().set_yscale('log')
plt.subplots_adjust(left = 0.2)
plt.savefig("pgg.png", dpi = 150)
```
# Save results
Save the results in an astropy table.
```
from astropy.table import Table
c = {}
c['pgg'] = np.vstack((EGeV, pgg))
t = Table(c)
t.write('ngc1275.fits', overwrite = True)
t1 = Table.read('ngc1275.fits')
t1
```
### Plot the magnetic field of the cluster, stored in module 0
```
plt.plot(m.modules["ICMGaussTurb"].r,m.modules["ICMGaussTurb"].B * np.sin(m.modules["ICMGaussTurb"].psi),
lw=1)
plt.plot(m.modules["ICMGaussTurb"].r,m.modules["ICMGaussTurb"].B * np.cos(m.modules["ICMGaussTurb"].psi),
lw=1, ls = '--')
plt.ylabel('$B$ field ($\mu$G)')
plt.xlabel('$r$ (kpc)')
```
And plot the electron density:
```
plt.loglog(m.modules["ICMGaussTurb"].r,m.modules[0].nel * 1e-3)
plt.ylabel('$n_\mathrm{el}$ (cm$^{-3}$)')
plt.xlabel('$r$ (kpc)')
```
You can also manipulate the magnetic field and electron density at run time
#### Calculate the coherence length of the transversal component $B$ field
It is also possible to compute the spatial correlation $C(x_3) = \langle B_\perp(\vec{x}) B_\perp(\vec{x} + x_3 \vec{e}_3)\rangle$ of the transversal magnetic field along the line of sight $x_3$:
```
x3 = np.linspace(0.,50.,1000) # distance in kpc from cluster center
c = m.modules["ICMGaussTurb"].Bfield_model.spatialCorr(x3)
plt.plot(x3,c / c[0])
plt.xlabel("$x_3$ (kpc)")
plt.ylabel("$C(x_3) / C(0)$")
plt.grid(True)
```
This is turn can be used to calculate the coherence length of the field,
$$ \Lambda_C = \frac{1}{C(0)} \int\limits_0^\infty C(x_3)dx_3. $$
```
from scipy.integrate import simps
x3 = np.linspace(0.,1e3,1000) # distance in kpc from cluster center
c = m.modules["ICMGaussTurb"].Bfield_model.spatialCorr(x3)
Lambda_c = simps(c, x3) / c[0]
print ("Coherence length of the field is Lambda_C = {0:.3e} kpc".format(Lambda_c))
```
#### Calculate the rotation measure of the field
```
m.modules["ICMGaussTurb"].Bfield_model.seed = 0 # or None
rm = m.modules["ICMGaussTurb"].Bfield_model.rotation_measure(m.modules["ICMGaussTurb"].r,
n_el=m.modules["ICMGaussTurb"].nel * 1e-3,
nsim=1000)
```
Taylor et al. (2006) found RM values between 6500 and 7500 rad m^-2. Comparing B-field realizations to that number:
```
from scipy.stats import norm
n, bins, _ = plt.hist(np.sort((rm)), bins=30, density=True, label="Simulated RM")
plt.xlabel("Rotation Measure (rad m${}^{-2}$)")
plt.ylabel("Density")
mean = np.mean(rm)
var = np.var(rm)
print ("RM mean +/- sqrt(var) in rad m^-2: {0:.2f} +/- {1:.2f}".format(mean, np.sqrt(var)))
plt.plot(bins, norm.pdf(bins, loc=mean, scale=np.sqrt(var)),
lw=2,
label="Gaussian Fit\n$\mu = {0:.2f}$\n$\sigma={1:.2f}$".format(mean, np.sqrt(var)))
print ("{0:.3f}% of B field realizations have |RM| > 7500 rad m^-2".format((np.abs(rm) > 7500).sum() / rm.size * 100.))
plt.legend()
plt.gca().tick_params(labelleft=False, left=False, right=False)
plt.savefig("sim_rm_perseus.png", dpi=150)
```
### Plot the magnetic field of the Milky Way
```
plt.plot(m.modules["GMF"].r, m.modules["GMF"].B * np.sin(m.modules["GMF"].psi),
lw = 1)
plt.plot(m.modules["GMF"].r, m.modules["GMF"].B * np.cos(m.modules["GMF"].psi),
lw = 1)
plt.ylabel('$B$ field ($\mu$G)')
plt.xlabel('$r$ (kpc)')
```
| github_jupyter |
# Model predicting on crops
```
# https://ipython.org/ipython-doc/3/config/extensions/autoreload.html
%load_ext autoreload
%autoreload 2
from pathlib import Path
import sys
sys.path.insert(0, Path(".").absolute().parent.as_posix())
import numpy as np
from common.dataset import FilesFromCsvDataset, TrainvalFilesDataset, TransformedDataset, read_image, TestFilesDataset
from image_dataset_viz import render_datapoint, DatasetExporter
import matplotlib.pylab as plt
%matplotlib inline
dataset = TrainvalFilesDataset("/home/fast_storage/imaterialist-challenge-furniture-2018/train/")
img_dataset = TransformedDataset(dataset, transforms=lambda x: read_image(x), target_transforms=lambda y: y - 1)
from torchvision.transforms import Compose, RandomVerticalFlip, RandomHorizontalFlip, RandomResizedCrop
from torchvision.transforms import RandomApply, RandomChoice
from torchvision.transforms import ColorJitter, ToTensor, Normalize
size = 350
basic_train_augs = Compose([
RandomChoice(
[
RandomResizedCrop(size, scale=(0.4, 0.6), interpolation=3),
RandomResizedCrop(size, scale=(0.6, 1.0), interpolation=3),
]
),
RandomHorizontalFlip(p=0.5),
RandomVerticalFlip(p=0.5),
ColorJitter(hue=0.12, brightness=0.12),
# ToTensor(),
# Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
img_augs_dataset = TransformedDataset(img_dataset, transforms=basic_train_augs)
img, label = img_augs_dataset[12]
render_datapoint(img, "class_{}".format(label), image_id="0")
```
5 crops
```
# FiveCrop??
import torch
from torchvision.transforms import FiveCrop, Lambda, Resize
single_img_augs = Compose([
RandomHorizontalFlip(p=0.5),
ColorJitter(hue=0.12, brightness=0.12),
# ToTensor(),
# Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
size = 350
augs_branch_1 = RandomResizedCrop(size, scale=(0.9, 1.0), interpolation=2)
augs_branch_2 = Compose([Resize(512, interpolation=2), FiveCrop(size=size)])
crops_train_augs = Compose([
Lambda(lambda img: (augs_branch_1(img), ) + augs_branch_2(img)),
Lambda(lambda crops: [single_img_augs(crop) for crop in crops])
# Lambda(lambda crops: torch.stack([single_img_augs(crop) for crop in crops]))
])
img_augs_dataset = TransformedDataset(img_dataset, transforms=crops_train_augs)
imgs, label = img_augs_dataset[30]
# render_datapoint(img, "class_{}".format(label), image_id="0")
plt.figure(figsize=(20, 5))
for i in range(0, len(imgs)):
plt.subplot(1, len(imgs), i + 1)
plt.imshow(imgs[i])
```
augs with tensors
```
single_img_augs = Compose([
RandomHorizontalFlip(p=0.5),
ColorJitter(hue=0.12, brightness=0.12),
ToTensor(),
Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
size = 224
augs_branch_1 = RandomResizedCrop(size, scale=(0.7, 1.0), interpolation=2)
augs_branch_2 = Compose([Resize(420, interpolation=2), FiveCrop(size=size)])
crops_train_augs = Compose([
Lambda(lambda img: (augs_branch_1(img), ) + augs_branch_2(img)),
Lambda(lambda crops: torch.stack([single_img_augs(crop) for crop in crops]))
])
img_augs_dataset = TransformedDataset(img_dataset, transforms=crops_train_augs)
from torch.utils.data import DataLoader
data_loader = DataLoader(img_augs_dataset, batch_size=10, num_workers=4)
data_loader_iter = iter(data_loader)
batchx, batchy = next(data_loader_iter)
batchx.shape, batchy.shape
```
Model on crops
```
from torch.nn import Module, Sequential, Conv2d, AdaptiveAvgPool2d, ReLU, Dropout, ModuleList, Linear
from torchvision.models.squeezenet import squeezenet1_1
from torch.nn.init import normal_, constant_
class FurnitureSqueezeNetOnCrops(Module):
def __init__(self, pretrained=True, n_crops=6):
super(FurnitureSqueezeNetOnCrops, self).__init__()
model = squeezenet1_1(pretrained=pretrained)
self.features = model.features
self.crop_classifiers = []
for i in range(n_crops):
# Final convolution is initialized differently form the rest
final_conv = Conv2d(512, 512, kernel_size=1, bias=False)
self.crop_classifiers.append(Sequential(
Dropout(p=0.5),
final_conv,
ReLU(inplace=True),
AdaptiveAvgPool2d(1)
))
for m in final_conv.modules():
normal_(m.weight, mean=0.0, std=0.01)
if m.bias is not None:
constant_(m.bias, 0.0)
self.crop_classifiers = ModuleList(self.crop_classifiers)
self.final_classifier = Linear(512, 128)
for m in self.final_classifier.modules():
normal_(m.weight, mean=0.0, std=0.01)
if m.bias is not None:
constant_(m.bias, 0.0)
def forward(self, crops):
batch_size, n_crops, *_ = crops.shape
features = []
for i in range(n_crops):
x = self.features(crops[:, i, :, :, :])
x = self.crop_classifiers[i](x)
features.append(x.view(batch_size, -1))
x = sum(features)
return self.final_classifier(x)
model = FurnitureSqueezeNetOnCrops(pretrained=False)
batchy_pred = model(batchx)
batchy_pred.shape
```
On crops with inception-resnetv2
```
# https://ipython.org/ipython-doc/3/config/extensions/autoreload.html
%load_ext autoreload
%autoreload 2
from pathlib import Path
import sys
sys.path.insert(0, Path(".").absolute().parent.as_posix())
import numpy as np
from common.dataset import FilesFromCsvDataset, TrainvalFilesDataset, TransformedDataset, read_image, TestFilesDataset
from image_dataset_viz import render_datapoint, DatasetExporter
# Basic training configuration file
import torch
from torchvision.transforms import RandomHorizontalFlip, Compose, RandomResizedCrop
from torchvision.transforms import FiveCrop, Lambda, Resize
from torchvision.transforms import ColorJitter, ToTensor, Normalize
from common.dataset import FilesFromCsvDataset
from common.data_loaders import get_data_loader
from models.inceptionresnetv2 import FurnitureInceptionResNetOnCrops
SEED = 17
DEBUG = True
DEVICE = 'cuda'
size = 299
single_img_augs = Compose([
RandomHorizontalFlip(p=0.5),
ColorJitter(hue=0.12, brightness=0.12),
ToTensor(),
Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5])
])
augs_branch_1 = RandomResizedCrop(size, scale=(0.9, 1.0), interpolation=2)
augs_branch_2 = Compose([Resize(int(size * 1.5), interpolation=2), FiveCrop(size=size)])
TRAIN_TRANSFORMS = Compose([
Lambda(lambda img: (augs_branch_1(img), ) + augs_branch_2(img)),
Lambda(lambda crops: torch.stack([single_img_augs(crop) for crop in crops]))
])
VAL_TRANSFORMS = TRAIN_TRANSFORMS
BATCH_SIZE = 6
NUM_WORKERS = 15
dataset = FilesFromCsvDataset("../output/filtered_train_dataset.csv")
TRAIN_LOADER = get_data_loader(dataset,
data_transform=TRAIN_TRANSFORMS,
batch_size=BATCH_SIZE,
num_workers=NUM_WORKERS,
pin_memory='cuda' in DEVICE)
model = FurnitureInceptionResNetOnCrops(pretrained='imagenet', n_cls_layers=256).to(DEVICE)
loader_iter = iter(TRAIN_LOADER)
batch_x, batch_y = next(loader_iter)
batch_x = batch_x.to(DEVICE)
batch_y = batch_y.to(DEVICE)
batch_y_pred = model(batch_x)
```
Model with trainable zoom on a part
```
from torchvision.transforms import Compose, RandomVerticalFlip, RandomHorizontalFlip
from torchvision.transforms import Resize
from torchvision.transforms import ColorJitter, ToTensor, Normalize
size = 350
basic_train_augs = Compose([
Resize((size, size), interpolation=3),
RandomHorizontalFlip(p=0.5),
ColorJitter(hue=0.12, brightness=0.12),
ToTensor(),
Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
img_augs_dataset = TransformedDataset(img_dataset, transforms=basic_train_augs)
img, label = img_augs_dataset[0]
from torch.utils.data import DataLoader
data_loader = DataLoader(img_augs_dataset, batch_size=10, num_workers=4)
data_loader_iter = iter(data_loader)
batch_x, batch_y = next(data_loader_iter)
batch_x.shape, batch_y.shape
import torch
from torch.nn import Module, Sequential, Conv2d, AdaptiveAvgPool2d, ReLU, Dropout, ModuleList, Linear
from torchvision.models.squeezenet import squeezenet1_1
from torch.nn.init import normal_, constant_
class FurnitureSqueezeNetWithZoom(Module):
def __init__(self, pretrained=True):
super(FurnitureSqueezeNetWithZoom, self).__init__()
model = squeezenet1_1(pretrained=pretrained)
self.features = model.features
self.inner_classifiers = []
for i in range(2):
# Final convolution is initialized differently form the rest
final_conv = Conv2d(512, 512, kernel_size=1, bias=False)
self.inner_classifiers.append(Sequential(
Dropout(p=0.5),
final_conv,
ReLU(inplace=True),
AdaptiveAvgPool2d(1)
))
for m in final_conv.modules():
normal_(m.weight, mean=0.0, std=0.01)
if m.bias is not None:
constant_(m.bias, 0.0)
self.inner_classifiers = ModuleList(self.inner_classifiers)
self.final_classifier = Linear(512, 128)
# Zoom params: center (x, y) and width/height normalized between [-0.5, 0.5]
self.zoom_params = torch.tensor([0.0, 0.0, 1.0, 1.0], requires_grad=True)
for m in self.final_classifier.modules():
normal_(m.weight, mean=0.0, std=0.01)
if m.bias is not None:
constant_(m.bias, 0.0)
def _zoom_on_data(self, x):
_, _, h, w = x.shape
ixs = int(self.zoom_params[0]) * w
ixe = int(self.zoom_params[0] + self.zoom_params[2]) * w
iys = int(self.zoom_params[1]) * h
iye = int(self.zoom_params[1] + self.zoom_params[3]) * h
out = x[:, :, iys:iye, ixs:ixe]
return out
def forward(self, x):
batch_size, *_ = x.shape
# zero-level classification:
low_features = [
self.features(x),
self.features(self._zoom_on_data(x))
]
features = []
for i, f in enumerate(low_features):
x = self.inner_classifiers[i](f)
features.append(x.view(batch_size, -1))
x = sum(features)
return self.final_classifier(x)
model = FurnitureSqueezeNetWithZoom(pretrained=False)
batch_y_pred = model(batchx)
from torch.nn import CrossEntropyLoss
criterion = CrossEntropyLoss()
loss = criterion(batch_y_pred, batch_y)
loss.backward()
model.zoom_params.grad
x = torch.tensor(list(range(100)))
w = torch.tensor([1.0, 0.0], requires_grad=True)
# ixs = (x[0]).long() * 100
# ixe = (x[0] + x[1]).long() * 100
# y = 30 - torch.sum(x[ixs:ixe])
ones = torch.ones_like(x)
x_hom = torch.cat([x.unsqueeze(-1), ones.unsqueeze(-1)], dim=1)
y = 30 - torch.sum(torch.sum(w * x_hom.float(), dim=-1))
y.backward()
y, w.grad
w = w.sub(0.01 * w.grad)
ones = torch.ones_like(x)
x_hom = torch.cat([x.unsqueeze(-1), ones.unsqueeze(-1)], dim=1)
w * x_hom[2, :].float()
torch.sum(w * x_hom.float(), dim=-1)
from torchviz import make_dot
make_dot(y)
x.grad
from torch.nn import functional as F
F.cro
x[ixs:ixe]
70 * 272 / 3600
```
| github_jupyter |
## Softmax Regression
Multi-class classification using softmax regression techniques has been illustrated in this notebook with each mathematical step implemented from scratch.
* Please go through this reference which will help to understand the theory of Softmax Regression.
* Reference : http://deeplearning.stanford.edu/tutorial/supervised/SoftmaxRegression/
* http://cs229.stanford.edu/notes2020spring/cs229-notes1.pdf
* sklearn iris dataset has been used which has 4 features and 3 classes.
### Import Libraries
```
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
import numpy as np
import matplotlib.pyplot as plt
```
### Load Data
To create training and testing set we can use sklearn train_test_split feature.
```
data = load_iris()
#create train and test set for input and output, test_size represent the % of test data
X_train, X_test, y_train, y_test = train_test_split(data.data, data.target, test_size=0.25)
# no of rows
m = X_train.shape[0]
#As we are using linearity, For vector multiplication adding a column in X with all the value 1
X = np.hstack((np.ones((m,1)),X_train))
print(X[:5]) # Checking first 5 rows of X matrix
```
As we have 3 classes for easier computation we will convert our output vector in matrix where no of columns is equal to the no of classes. The values in the matrix will be 0 or 1. For instance the rows where we have output 2 the column 2 will contain 1 and rest all 0
```
k=np.unique(y_train,return_counts=True)
Y=np.zeros((m,len(k[0])))
for i in range(m):
Y[i,y_train[i]]=1
print(Y[:5]) # Checking first 5 rows of Y matrix
```
### Softmax Regression Implementation
### Softmax Function
```
## Assuming theta
np.random.seed(0)
theta = np.random.randn(3,5)
theta
## Calculating matrix product of theta and X. Only 5 rows of X is taken to visualize the process and calculation
Z=np.dot(X[:5], theta.T)
Z
```
### Understanding Softmax function by stepwise implementation and then making a function:
```
np.exp(Z) ## Calculating Exponential
np.exp(Z).sum(axis=1,keepdims=True) ## Calculating sum along the axis=1 (along the column from left to right)
result= np.exp(Z)/np.exp(Z).sum(axis=1,keepdims=True) # Dividing exponential with sum
result
## Finally this is the result of softmax function
## Checking its sum, must be equal to 1
np.sum(result,axis=1,keepdims=True) # We are right
```
### Finally making Softmax function
```
def softmax(z):
return np.exp(z) / np.exp(z).sum(axis=1,keepdims=True)
## Checking
np.random.seed(0)
theta = np.random.randn(3,5)
Z=np.dot(X[:5], theta.T)
h_out=softmax(Z)
h_out
h_out.shape
```
### Cost Function

```
cost = -np.sum(Y[:5] * np.log(h_out)) / m ## To understand the concept we have taken only 5 rows of X
cost ## and so taking only first 5 value of Y
## m should also be only 5 but let it be so.
```
### Gradient Calculation of Softmax Cost Function

```
grad=np.dot((h_out-Y[:5]).T, X[:5])
print(grad)
print(grad.shape)
```
### Now applying Gradient Descent
```
# This is one step of Gradient Descent
a=0.001 # learning rate
theta=theta-a/m*grad # Gradient Descent
theta
```
### Final Implementation of Gradient Descent
* Now Writing above piecewise code into one cell and making function which gives final optimized weights(theta) and cost for each iteration
```
#define theta with size 3,5 as there are 3 classes and 5 features, lets take the initial value as 0
def Softmax_reg(X_train,y_train,learning_rate= 0.001,iteration=50000):
# Prepartion of X_train
# no of rows
num_rows = X_train.shape[0]
# As we are using linearity, For vector multiplication adding a column in X with all the value 1
X = np.hstack((np.ones((num_rows,1)),X_train))
# Prepartion of y_train
k=np.unique(y_train,return_counts=True)
Y=np.zeros((num_rows,len(k[0])))
for i in range(num_rows):
Y[i,y_train[i]]=1
# Initializatio of theta
m,n=X.shape
_,k=Y.shape
theta = np.random.randn(k,n)
# to store cost values
cost_for_iter = []
# Gradient Descent inside loop for multiple updation of theta
for i in range(iteration):
Z= np.dot(X, theta.T)
h_out = softmax(Z)
cost = -np.sum(Y * np.log(h_out)) / m # Cost calculation
cost_for_iter.append(cost)
grad=np.dot((h_out-Y).T, X) # gradient calculation
theta = theta - (learning_rate/m)*grad # Gradient Descent
return theta,cost_for_iter
theta,cost=Softmax_reg(X_train,y_train,learning_rate= 0.001,iteration=50000)
print(theta,'\n')
#print(cost)
plt.plot(cost)
```
### Prediction on test data and Implementing inside a function
```
m_test = X_test.shape[0]
X_test_vec = np.hstack((np.ones((m_test,1)),X_test))
probab = softmax(np.dot(X_test_vec,theta.T))
predict = np.argmax(probab, axis=1)
predict
def predict(X_test,theta):
m_test = X_test.shape[0]
X_test = np.hstack((np.ones((m_test,1)),X_test))
probab = softmax(np.dot(X_test,theta.T))
predict = np.argmax(probab, axis=1)
return predict
y_pred=predict(X_test,theta)
y_pred
```
### Comparing our model with scikit library
Lets compare our model with the Scikit logistic model.Scikit logistic model is self sufficient to handle multiclass classification.
```
from sklearn import linear_model
from sklearn.metrics import accuracy_score
#train the model with training data
regr = linear_model.LogisticRegression(max_iter=1000)
regr.fit(X_train,y_train)
#Predict our test data
y_pred_sklearn = regr.predict(X_test)
# Accuracy score Sklearn
print(" Sklearn Accuracy score: %.2f" % accuracy_score(y_pred_sklearn, y_test))
# Accuracy score Our Model
print(" Our Model Accuracy score: %.2f" % accuracy_score(y_pred, y_test))
```
#### * https://github.com/Rami-RK
#### * https://www.linkedin.com/in/ramendra-kumar-57334478/
| github_jupyter |
# Training a binary autoencoder
This tutorial explains how to train a binary autoencoder in order to obtain a satisfying encoding of your data, to be used as input to the OPU. The architecture and training procedure is adapted from https://arxiv.org/abs/1803.09065.
```
import matplotlib.pyplot as plt
import numpy as np
from sklearn.datasets import make_blobs
from sklearn.decomposition import PCA
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader, TensorDataset
# fake data
n_samples = 1000
n_features = 100
X = torch.FloatTensor(n_samples, n_features).normal_()
y = torch.FloatTensor(n_samples, n_features).normal_()
```
In the next cell, we define the autoencoder. The encoder consists of a linear layer, followed by a step function, yielding the binary representation of the data. The decoder is simply the transpose of the encoder. This allows us to learn only the decoder via backprop, which will change the encoder at the same time. The non-differentiable activation is therefore not a problem.
```
from lightonml.encoding.models import EncoderDecoder
from lightonml.encoding.models import train
batch_size = 64
loader = DataLoader(TensorDataset(X, y), batch_size=batch_size)
bits_per_feature = 10
encoder = EncoderDecoder(n_features, n_features * bits_per_feature)
optimizer = optim.Adam(encoder.parameters(), lr=1e-3)
```
A newly created encoder is in `training` mode and will return the reconstructed input:
```
encoder.training
```
We now train it on our data, it is quite fast. The `train` function from `lightonml.encoding.models` will automatically move the encoder to GPU if one is available.
```
model = train(encoder, loader, optimizer, criterion=F.mse_loss, epochs=10)
```
We set the encoder to `eval` mode:
```
model.eval()
model.training
```
It is ready to encode:
```
# we move the data to the GPU where the encoder lives
# and fetch the binary code from it
Xenc = encoder(X.to('cuda')).cpu()
Xenc.shape, Xenc.dtype, torch.unique(Xenc)
```
Of course, `encoder` can also be used on validation and test data, that weren't used to train the autoencoder.
## Using a "real" toy dataset
```
n_samples = 10000
n_features = 50
X, y = make_blobs(n_samples=n_samples, n_features=n_features, centers=5)
```
We visualise a PCA of the data:
```
pca = PCA(n_components=2)
X_pca = pca.fit_transform(X)
fig, ax = plt.subplots(figsize=(6,6))
for i in np.unique(y):
ax.scatter(X_pca[y==i,0], X_pca[y==i,1], s=2, label='y={}'.format(i))
ax.legend()
```
We see the 5 clusters created by `make_blobs`. Ideally, our encoder should preserve this structure in the binary encoding. Let us encode the data:
```
X = torch.from_numpy(X).float() # by default X in numpy is double, so we cast to float
loader = DataLoader(TensorDataset(X, X), batch_size=batch_size) # loader in `train` assumes a tuple
encoder = EncoderDecoder(n_features, n_features * bits_per_feature)
optimizer = optim.Adam(encoder.parameters(), lr=1e-3)
encoder.training
model = train(encoder, loader, optimizer, criterion=F.mse_loss, epochs=10)
encoder.eval()
# we move the encoder to cpu, but we could also move the data to GPU
# for faster processing as we did before
encoder.to('cpu')
Xenc = encoder(X)
Xenc.shape, Xenc.dtype, torch.unique(Xenc)
```
And we visualize it again:
```
pca = PCA(n_components=2)
Xenc_pca = pca.fit_transform(Xenc.numpy())
fig, ax = plt.subplots(figsize=(6,6))
for i in np.unique(y):
ax.scatter(Xenc_pca[y==i,0], Xenc_pca[y==i,1], s=2, label='y={}'.format(i))
ax.legend()
```
The 5 original clusters are well preserved. The encoder does its job !
| github_jupyter |
## Prophet Baseline Notebook
This notebook contains the code used to predict the price of bitcoin **just** using FB prophet.
You can think of this as a sort of baseline model!
```
from fbprophet import Prophet
from sklearn.metrics import r2_score
%run helper_functions.py
%autosave 120
%matplotlib inline
%run prophet_helper.py
%run prophet_baseline_btc.py
plt.style.use('fivethirtyeight')
plt.rcParams["figure.figsize"] = (15,10)
plt.rcParams["xtick.labelsize"] = 16
plt.rcParams["ytick.labelsize"] = 16
plt.rcParams["axes.labelsize"] = 20
plt.rcParams['legend.fontsize'] = 20
plt.style.use('fivethirtyeight')
pd.set_option('display.max_colwidth', -1)
import numpy as np
import math
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
```
## Forecasting BTC Price with Fb Prophet
```
df = unpickle_object("blockchain_info_df.pkl")
df.head()
df_btc = pd.DataFrame(df['mkt_price'])
true, pred = prophet_baseline_BTC(df_btc, 30, "mkt_price")
r2_score(true, pred) #we see that our baseline model just predicts 44% of the variation when predicting price
plt.plot(pred)
plt.plot(true)
plt.legend(["Prediction", 'Actual'], loc='upper left')
plt.xlabel("Prediction #")
plt.ylabel("Price")
plt.title("TS FB Prophet Baseline - Price Prediction");
```
## Let's predict percentage change!
```
df_btc_pct = df_btc.pct_change()
df_btc_pct.rename(columns={"mkt_price": "percent_change"}, inplace=True)
df_btc_pct = df_btc_pct.iloc[1:, :]
print(df_btc_pct.shape)
df_btc_pct.head()
true_pct, pred_pct = prophet_baseline_BTC(df_btc_pct, 30, "percent_change")
r2_score(true_pct, pred_pct)
```
MSE IS 0.000488913299898903
```
plt.plot(pred_pct)
plt.plot(true_pct)
plt.legend(["Prediction", 'Actual'], loc='upper left')
plt.xlabel("Prediction #")
plt.ylabel("Price")
plt.title("TS FB Prophet Baseline - Price Prediction");
```
we do terribly at predicting percent change! However, we know that percent change should be applied to the price of the previous day. Let's do that!
Note that the MSE is very close to 0 - we have quite an accurate Model!
```
prices_to_be_multiplied = df.loc[pd.date_range(start="2017-01-23", end="2017-02-21"), "mkt_price"]
forecast_price_lst = []
for index, price in enumerate(prices_to_be_multiplied):
predicted_percent_change = 1+float(pred_pct[index])
forecasted_price = (predicted_percent_change)*price
forecast_price_lst.append(forecasted_price)
ground_truth_prices = df.loc[pd.date_range(start="2017-01-24", end="2017-02-22"), "mkt_price"]
ground_truth_prices = list(ground_truth_prices)
r2_score(ground_truth_prices, forecast_price_lst) # such an incredible result! This is what we have to beat with my nested TS model
plt.plot(forecast_price_lst)
plt.plot(ground_truth_prices)
plt.legend(["Prediction", 'Actual'], loc='upper left')
plt.xlabel("Prediction #")
plt.ylabel("Price")
plt.title("TS FB Prophet Baseline - Price Prediction");
```
| github_jupyter |
# Plot forces for flow past cylinder
## grid0 and grid1 case
## Compare differences with Reynolds number
```
%%capture
import sys
sys.path.insert(1, '../utilities')
import litCdData
import numpy as np
import matplotlib.pyplot as plt
## Some needed functions for postprocessing
def concatforces(filelist):
"""
Concatenate all the data in a list of files given by filelist, without overlaps in time
"""
for ifile, file in enumerate(filelist):
dat=np.loadtxt(file, skiprows=1)
if ifile==0:
alldat = dat
else:
lastt = alldat[-1,0] # Get the last time
filt = dat[:,0]>lastt
gooddat = dat[filt,:]
alldat = np.vstack((alldat, gooddat))
return alldat
# Calculate time average
def timeaverage(time, f, t1, t2):
filt = ((time[:] >= t1) & (time[:] <= t2))
# Filtered time
t = time[filt]
# The total time
dt = np.amax(t) - np.amin(t)
# Filtered field
filtf = f[filt]
# Compute the time average as an integral
avg = np.trapz(filtf, x=t, axis=0) / dt
return avg
def tukeyWindow(N, params={'alpha':0.1}):
"""
The Tukey window
see https://en.wikipedia.org/wiki/Window_function#Tukey_window
"""
alpha = params['alpha']
w = np.zeros(N)
L = N+1
for n in np.arange(0, int(N//2) + 1):
if ((0 <= n) and (n < 0.5*alpha*L)):
w[n] = 0.5*(1.0 - np.cos(2*np.pi*n/(alpha*L)))
elif ((0.5*alpha*L <= n) and (n <= N/2)):
w[n] = 1.0
else:
print("Something wrong happened at n = ",n)
if (n != 0): w[N-n] = w[n]
return w
# FFT's a signal, returns 1-sided frequency and spectra
def getFFT(t, y, normalize=False, window=True):
"""
FFT's a signal, returns 1-sided frequency and spectra
"""
n = len(y)
k = np.arange(n)
dt = np.mean(np.diff(t))
frq = k/(n*dt)
if window: w = tukeyWindow(n)
else: w = 1.0
if normalize: L = len(y)
else: L = 1.0
FFTy = np.fft.fft(w*y)/L
# Take the one sided version of it
freq = frq[range(int(n//2))]
FFTy = FFTy[range(int(n//2))]
return freq, FFTy
# Basic problem parameters
D = 6 # Cylinder diameter
U = 20 # Freestream velocity
Lspan = 24 # Spanwise length
A = D*Lspan # frontal area
rho = 1.225 # density
Q = 0.5*rho*U*U # Dynamic head
vis = 1.8375e-5 # viscosity
ReNum = rho*U*D/vis # Reynolds number
#avgt = [160.0, 260.0] # Average times
saveinfo = False
alldata = []
# Label, Filenames averaging times
runlist = [['Grid0 Re=8.0M', ['../cylgrid0_sst_iddes_matula_01/forces86m.dat'], [150, 600], {'vis':1.8375e-5, 'mc':'r'}],
#['Grid0 Re=3.6M', ['../cylgrid0_sst_iddes_matula_03_Re3p6M/forces36m.dat'], [300, 800], {'vis':4.0833333333333334e-05, 'mc':'r'}],
['Grid1 Re=8.0M', ['../cylgrid1new_sst_iddes_01/forces01.dat', '../cylgrid1new_sst_iddes_01/forces02.dat', '../cylgrid1new_sst_iddes_01/forces03.dat'],
[400, 900], {'vis':1.8375e-5, 'mc':'k'}],
#['Grid1 Re=3.6M', ['../cylgrid1_Re3p6M_sst_iddes_02/forces01.dat', '../cylgrid1_Re3p6M_sst_iddes_02/forces02.dat'],
# [300, 900], {'vis':4.0833333333333334e-05, 'mc':'k'}],
]
alldata = []
for run in runlist:
forcedat = concatforces(run[1])
t = forcedat[:,0]*U/D # Non-dimensional time
alldata.append([run[0], t, forcedat, run[2], run[3]])
#print(alldata)
print('%30s %10s %10s'%("Case", "avgCd", "avgCl"))
for run in alldata:
label = run[0]
t = run[1]
forcedat = run[2]
avgt = run[3]
Cd = (forcedat[:,1]+forcedat[:,4])/(Q*A)
Cl = (forcedat[:,2]+forcedat[:,5])/(Q*A)
# Calculate averaged Cp, Cd
avgCd = timeaverage(t, Cd, avgt[0], avgt[1])
avgCl = timeaverage(t, Cl, avgt[0], avgt[1])
print('%30s %10f %10f'%(label, avgCd, avgCl))
#print("Avg Cd = %f"%avgCd)
#%print("Avg Cl = %f"%avgCl)
```
## Plot Lift and Drag coefficients
```
plt.rc('font', size=16)
plt.figure(figsize=(10,8))
for run in alldata:
label = run[0]
t = run[1]
forcedat = run[2]
avgt = run[3]
Cd = (forcedat[:,1]+forcedat[:,4])/(Q*A)
Cl = (forcedat[:,2]+forcedat[:,5])/(Q*A)
# Calculate averaged Cp, Cd
avgCd = timeaverage(t, Cd, avgt[0], avgt[1])
avgCl = timeaverage(t, Cl, avgt[0], avgt[1])
#print('%30s %f %f'%(label, avgCd, avgCl))
plt.plot(t,Cd, label=label)
plt.hlines(avgCd, np.min(t), np.max(t), linestyles='dashed', linewidth=1)
plt.xlabel(r'Non-dimensional time $t U_{\infty}/D$');
plt.legend()
plt.ylabel('$C_D$')
plt.title('Drag coefficient $C_D$');
plt.figure(figsize=(10,8))
for run in alldata:
label = run[0]
t = run[1]
forcedat = run[2]
avgt = run[3]
Cd = (forcedat[:,1]+forcedat[:,4])/(Q*A)
Cl = (forcedat[:,2]+forcedat[:,5])/(Q*A)
# Calculate averaged Cp, Cd
avgCd = timeaverage(t, Cd, avgt[0], avgt[1])
avgCl = timeaverage(t, Cl, avgt[0], avgt[1])
plt.plot(t,Cl, label=label)
plt.hlines(avgCl, np.min(t), np.max(t), linestyles='dashed', linewidth=1)
plt.xlabel(r'Non-dimensional time $t U_{\infty}/D$');
plt.ylabel('$C_l$')
plt.title('Lift coefficient $C_l$');
plt.legend()
```
## Plot Spectra
```
plt.figure(figsize=(10,8))
for run in alldata:
label = run[0]
t = run[1]
forcedat = run[2]
avgt = run[3]
dict = run[4]
filt = ((t[:] >= avgt[0]) & (t[:] <= avgt[1]))
tfiltered = t[filt]*D/U
Cd = (forcedat[:,1]+forcedat[:,4])/(Q*A)
Cl = (forcedat[:,2]+forcedat[:,5])/(Q*A)
Cdfiltered = Cd[filt]
Clfiltered = Cl[filt]
f, Cdspectra = getFFT(tfiltered, Cdfiltered, normalize=True)
f, Clspectra = getFFT(tfiltered, Clfiltered, normalize=True)
plt.loglog(f*D/U, abs(Clspectra), color=dict['mc'], label='Cl '+label)
plt.axvline(0.37, linestyle='--', color='gray')
plt.xlim([1E-2,2]);
plt.ylim([1E-8, 1E-1]);
plt.xlabel(r'$St=f*D/U_\infty$');
plt.ylabel(r'$|\hat{C}_{l}|$')
plt.legend()
```
## Plot Cd versus Reynolds number
```
plt.figure(figsize=(10,8))
litCdData.plotEXP()
litCdData.plotCFD()
for run in alldata:
label = run[0]
t = run[1]
forcedat = run[2]
avgt = run[3]
dict = run[4]
Cd = (forcedat[:,1]+forcedat[:,4])/(Q*A)
Cl = (forcedat[:,2]+forcedat[:,5])/(Q*A)
# Calculate averaged Cp, Cd
avgCd = timeaverage(t, Cd, avgt[0], avgt[1])
avgCl = timeaverage(t, Cl, avgt[0], avgt[1])
vis = dict['vis']
mc = dict['mc']
ReNum = rho*U*D/vis
plt.semilogx(ReNum, avgCd, '*', color=mc, ms=15, label='Nalu SST-IDDES '+label)
plt.grid()
plt.legend(fontsize=10)
plt.xlabel(r'Reynolds number Re');
plt.ylabel('$C_D$')
plt.title('Drag coefficient $C_D$');
# Write the YAML file these averaged quantities
import yaml
if saveinfo:
savedict={'Re':float(ReNum), 'avgCd':float(avgCd), 'avgCl':float(avgCl)}
f=open('istats.yaml','w')
f.write('# Averaged quantities from %f to %f\n'%(avgt[0], avgt[1]))
f.write('# Grid: grid0\n')
f.write(yaml.dump(savedict, default_flow_style=False))
f.close()
```
| github_jupyter |
```
!git clone https://github.com/deepanrajm/deep_learning.git
#Importing required python libraries
import os
import librosa
import tensorflow as tf
import numpy as np
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.utils import np_utils
from keras.models import load_model
#from playsound import playsound
label = ["car_horn","dog_bark","engine_idling","siren"]
#Defining One-Hot as labels
car_horn_onehot = [1,0,0,0]
dog_bark_onehot = [0,1,0,0]
engine_idling_onehot = [0,0,1,0]
siren_onehot = [0,0,0,1]
#Converting files in a folder into list of arrays containg the properties of the files
def decodeFolder(category):
print("Starting decoding folder "+category+" ...")
listOfFiles = os.listdir(category)
arrays_sound = np.empty((0,193))
for file in listOfFiles:
filename = os.path.join(category,file)
features_sound = extract_feature(filename)
arrays_sound = np.vstack((arrays_sound,features_sound))
return arrays_sound
#Extracting the feataures of a wav file as inpurt to the data
def extract_feature(file_name):
print("Extracting "+file_name+" ...")
X, sample_rate = librosa.load(file_name)
stft = np.abs(librosa.stft(X))
mfccs = np.mean(librosa.feature.mfcc(y=X, sr=sample_rate, n_mfcc=40).T,axis=0)
chroma = np.mean(librosa.feature.chroma_stft(S=stft, sr=sample_rate).T,axis=0)
mel = np.mean(librosa.feature.melspectrogram(X, sr=sample_rate).T,axis=0)
contrast = np.mean(librosa.feature.spectral_contrast(S=stft, sr=sample_rate).T,axis=0)
tonnetz = np.mean(librosa.feature.tonnetz(y=librosa.effects.harmonic(X),sr=sample_rate).T,axis=0)
return np.hstack((mfccs,chroma,mel,contrast,tonnetz))
#train data
car_horn_sounds = decodeFolder("deep_learning/Sound_Classification/car_horn")
car_horn_labels = [car_horn_onehot for items in car_horn_sounds]
dog_bark_sounds = decodeFolder("deep_learning/Sound_Classification/dog_bark")
dog_bark_labels = [dog_bark_onehot for items in dog_bark_sounds]
engine_idling_sounds = decodeFolder("deep_learning/Sound_Classification/engine_idling")
engine_idling_labels = [engine_idling_onehot for items in engine_idling_sounds]
siren_sounds = decodeFolder("deep_learning/Sound_Classification/siren")
siren_labels = [siren_onehot for items in siren_sounds]
train_sounds = np.concatenate((car_horn_sounds, dog_bark_sounds,engine_idling_sounds,siren_sounds))
train_labels = np.concatenate((car_horn_labels, dog_bark_labels,engine_idling_labels,siren_labels))
print (train_sounds.shape)
X_train = train_sounds.reshape(train_sounds.shape[0], train_sounds.shape[1]).astype('float32')
#X_test = X_test.reshape(X_test.shape[0], num_pixels).astype('float32')
#test_data
test_sound = decodeFolder("deep_learning/Sound_Classification/test")
#test_sounds = np.concatenate(test_sound)
print (test_sound.shape)
X_test = test_sound.reshape(test_sound.shape[0], test_sound.shape[1]).astype('float32')
#print (X_train.shape)
model = Sequential()
model.add(Dense(193, input_dim=193, init='uniform', activation='relu'))
#model.add(Dense(8, init='uniform', activation='relu'))
model.add(Dense(4, init='uniform', activation='softmax'))
# Compile model
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fit the model
model.fit(X_train, train_labels, nb_epoch=150, batch_size=10)
model.save('my_model.h5')
model = load_model('my_model.h5')
pred = model.predict_classes(X_test)
listOfFiles = os.listdir("deep_learning/Sound_Classification/test")
for i in range (0, len(listOfFiles)):
print ("Listening to",listOfFiles[i] )
#playsound(("test\\"+str(listOfFiles[i])))
print ("I think it is", label[pred[i]],"sound")
```
| github_jupyter |
```
import wandb
wandb.init(project="Channel_Cha")
import pytorch
from torch import nn
from torch.optim import SGD
from torch.utils.data import DataLoader
import torch.nn.functional as F
from torchvision.transforms import Compose, ToTensor, Normalize
from torchvision.datasets import MNIST
from ignite.engine import Events,
create_supervised_trainer,
create_supervised_trainer
from ignite.metrics import Accuracy, Loss
from tqdm import tqdm
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 10, kernel_size=5)
self.conv2 = nn.Conv2d(10, 20, kernel_size=5)
self.conv2_drop = nn.Dropout2d()
self.fc1 = nn.Linear(320, 50)
self.fc2 = nn.Linear(50, 10)
def forward(self, x):
x = F.relu(F.max_pool2d(self.conv1(x), 2))
x = F.relu(F.max_pool2d(self.conv2_drop(self.conv2(x)), 2))
x = x.view(-1, 320)
x = F.relu(self.fc1(x))
x = F.dropout(x, training=self.training)
x = self.fc2(x)
return F.log_softmax(x, dim=-1)
def get_data_loaders(train_batch_size, val_batch_size):
data_transform = Compose([ToTensor(), Normalize((0.1307,), (0.3081,))])
train_loader = DataLoader(MNIST(download=True, root=".", transform=data_transform, train=True),
batch_size=train_batch_size, shuffle=True)
val_loader = DataLoader(MNIST(download=False, root=".", transform=data_transform, train=False),
batch_size=val_batch_size, shuffle=False)
return train_loader, val_loader
def run(train_batch_size, val_batch_size, epochs, lr, momentum, log_interval):
train_loader, val_loader = get_data_loaders(train_batch_size, val_batch_size)
model = Net()
device = 'cpu'
if torch.cuda.is_available():
device = 'cuda'
optimizer = SGD(model.parameters(), lr=lr, momentum=momentum)
trainer = create_supervised_trainer(model, optimizer, F.nll_loss, device=device)
evaluator = create_supervised_evaluator(model,
metrics={'accuracy': Accuracy(),
'nll': Loss(F.nll_loss)},
device=device)
desc = "ITERATION - loss: {:.2f}"
pbar = tqdm(
initial=0, leave=False, total=len(train_loader),
desc=desc.format(0)
)
@trainer.on(Events.ITERATION_COMPLETED(every=log_interval))
def log_training_loss(engine):
pbar.desc = desc.format(engine.state.output)
pbar.update(log_interval)
@trainer.on(Events.EPOCH_COMPLETED)
def log_training_results(engine):
pbar.refresh()
evaluator.run(train_loader)
metrics = evaluator.state.metrics
avg_accuracy = metrics['accuracy']
avg_nll = metrics['nll']
tqdm.write(
"Training Results - Epoch: {} Avg accuracy: {:.2f} Avg loss: {:.2f}"
.format(engine.state.epoch, avg_accuracy, avg_nll)
)
@trainer.on(Events.EPOCH_COMPLETED)
def log_validation_results(engine):
evaluator.run(val_loader)
metrics = evaluator.state.metrics
avg_accuracy = metrics['accuracy']
avg_nll = metrics['nll']
tqdm.write(
"Validation Results - Epoch: {} Avg accuracy: {:.2f} Avg loss: {:.2f}"
.format(engine.state.epoch, avg_accuracy, avg_nll))
pbar.n = pbar.last_print_n = 0
trainer.run(train_loader, max_epochs=epochs)
pbar.close()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ProfessorPatrickSlatraigh/CST2312/blob/main/CST2312_Class06_Files.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# CST2312 - Class #06, follow-up on reading files in Google Colab
by Professor Patrick, 16-Feb-2022
This notebook works with a Github repository in ProfessorPatrickSlatraigh/CST2312.
That repository includes the files "mbox-short.txt" and "mbox.txt".
The repo can be cloned to be used in Google Colab as a resource or a URL for a file in the Github repo, or Google Drive can be used to open a Python handle for a file which will persist beyond the current Colab session. Before working with the file in Github or Google Drive, the first section of this notebook describes how local files can be uploaded to a Colab session. Files which are uploaded as resources to the content area of Colab do not persist after the Colab notebook is closed.
Here is the mbox-short.txt file as a reference: https://www.py4e.com/code3/mbox-short.txt , which is a shortened version of the file: https://www.py4e.com/code3/mbox.txt . Both files are text files which contain a series of email messages. The files are used as references in exercises in the Charles Severance book Python for Everybody (py4e.com).
---
## Quick PY4E Exercises
*Here is a quick snippet with an example of code from Charles Severance's Python for Everybody on files.*
1. The next snippet opens a file handle fhand with the file mbox-short.txt -- be sure to have mbox-short.txt in your current working directorry (\Content).
2. Then it loops through every line of the file looking for lines which start with "From:" and prints only those lines.
```
fhand = open('mbox-short.txt')
for line in fhand:
if line.startswith('From:') :
print(line)
```
Let's try that same but also print the line number.
```
fhand = open('mbox-wrong.txt')
number = 1
for line in fhand:
if line.startswith('From:') :
print(number, '"'+line+'"') # this still has \n of each line
# print(number, '"'+line[:-1]+'"') # ignoring the \n in each line
number = number +1
```
---
# Reading and Writing Files - Various Methods
The following code snippet imports pandas which is needed for the file loading processes from Github and Googe Drive (gdrive) which are described at the bottom of this notebook.
```
# let's import pandas as pd so that we have it available
import pandas as pd
```
---
## **UPLOADING TO COLAB EVERY TIME**
The first example reads "mbox-short.txt" from the Google Colab content folder "sample_data". In order to do that, the "mbox-short.txt" file needs to be uploaded to the "sample_data" folder. That upload is temporary for the Google Colab session - the "mbox-short.txt" file will go away after you finish with your active Colab notebook. Note that this method does not require pandas.
Use the panel on the left of your Colab session to navigate to the content area and the "sample_data" folder. Then use the three vertical dots to the right of the name "sample_data" to choose 'Upload' and navigate to the "mbox-short.txt" file on your computer.
Use the three vertical dots to your uploaded "mbox_short.txt" file in the "sample_data" content folder to choose "Copy path" and that will put the full path (URL) in your clipboard. If the path is not the same as in the following call to the open() command then replace the string for the file name with the URL from your clipboard - paste it in as the argument to open().
```
colab_handle = open("/content/sample_data/mbox-short.txt")
```
Now you can use the print function to see the attributes of the new colab_handle you created to the "mbox-short.txt" file in the content folder "sample_data" on Google Colab.
```
print(colab_handle)
```
You can use a for loop to print the contents of "mbox-short.txt"
```
for line in colab_handle :
print(line) # this prints two \n characters
# print(line, end='') # this avoids the print \n
# print(line[:-1]) # this avoids the \n in line
```
As a file is read, the file handle keeps a pointer to the current place in the file so that it knows where to pick up reading next. Once the file has been completely read, the pointer is at the end of the file and there is no more file content to read. To re-read the file, the pointer must be set back to the start, or position 0.
```
# to reuse a file, you must reset the pointer to the start of the file, position 0.
colab_handle.seek(0)
```
# **READING FILES FROM GITHUB**
Now let's try reading the same file from a Github repository (repo). We will use the CST2312 repo in the ProfessorPatrickSlatraigh account on Github. The file "mbox-short.txt" was uploaded to that repo.
From Github we navigated to the "mbox-short.txt" file and viewed it in it's raw format using the "raw" button to the right of the file name. While in raw viewing mode in a browser, we copied the URL from the browser to the clipboard. Please note that this works with open repos, not private repos.
```
git_handle = pd.read_fwf("https://raw.githubusercontent.com/ProfessorPatrickSlatraigh/CST2312/main/mbox-short.txt")
print(git_handle)
```
Storing files in Github gives us persistence. That is, when we are done with our Google Colab session the files on Github remain and can be used again. And our Google Colab notebooks should work each time we open them without the need for use to upload files to the content area on Google Colab for every session.
# **READING FILES FROM GOOGLE DRIVE**
We can also have persistent files stored in Google Drive. To read files from Google Drive we will need to import the drive module from google.colab. We will also need to have Google Drive give stream access to Google Colab. If the files are on a different Google Drive account from the Google Colab account then be sure to have permission of the Google Drive owner for access to the file.
You can use the drive module from google.colab to mount your entire Google Drive to Colab by:
1. Executing the below code which will provide you with an authentication link
```
from google.colab import drive
drive.mount('/content/gdrive')
```
2. Open the link
3. Choose the Google account whose Drive you want to mount
4. Allow Google Drive Stream access to your Google Account
5. Copy the code displayed, paste it in the text box as shown below, and press Enter
Once the Drive is mounted, you’ll get the message “Mounted at /content/gdrive”, and you’ll be able to browse through the contents of your Drive from the file-explorer pane.
You can even write directly to Google Drive from Colab using the usual file/directory operations.
```
!touch "/content/gdrive/My Drive/sample_file.txt"
```
This will create a file in your Google Drive, and will be visible in the file-explorer pane once you refresh it. Notice that the path within the content area is different from the "sample_data" folder we used earlier for fles uploaded directly to Google Colab. The content area will have a "gdrive" folder after you have authenticated with Google Drive. Within the "gdrive" folder there should be a folder structure according to your Google Drive folders.
If your Google Drive folder had the file "mbox-short.txt" within the "My Drive" folder then you would be able to open that file with the following code:
```
gdrive_handle = open("/content/gdrive/My Drive/mbox-short.txt")
```
Now you can use the print function to see the attributes of the new gdrive_handle you created to the "mbox-short.txt" file in the content folder "gdrive/My Drive/" on Google Drive.
```
print(gdrive_handle)
```
As in the earlier Google Colab example, you can now use a for loop to print the contents of "mbox_short.txt" in Google Drive
```
for line in gdrive_handle :
print(line)
```
# **Reading Pastebin and Other HTTP with GET**
This section reads the "mail-short.txt" file from a Pastebin posting using the RAW format in Pastebin and the HTTP GET from the request module. The source file is online at: https://pastebin.com/raw/ADPQe6BM
First import the request module as rq
```
import requests as rq
```
The use the GET command to read the RAW text file on Pastebin
```
http_handle = rq.get('https://pastebin.com/raw/ADPQe6BM')
list_of_lines = http_handle.text.splitlines()
```
Print the response to check that the HTTP request worked
```
print(http_handle)
```
And print the result
```
for line in list_of_lines:
print(line)
```
A file object has a lot of attributes.
You can see a list of all methods and attributes of the file object here:
https://docs.python.org/2.4/lib/bltin-file-objects.html.
Following are some of the most used file object methods −
* **close()** - Close the file.
* **next()** - When a file is used as an iterator, typically in a for loop (for example, for line in f: print line), the next() method is called repeatedly. This method returns the next input line, or raises StopIteration when EOF is hit.
* **read([size])** - Read at most size bytes from the file.
* **readline([size])** - Read one entire line from the file.
* **seek(offset[, whence])** - Set the file's current position, like stdio's fseek(). The whence argument is optional and defaults to 0 (absolute file positioning); other values are 1 (seek relative to the current position) and 2 (seek relative to the file's end).
* **tell()** - Return the file's current position, like stdio's ftell().
* **write(str)** - Write a string to the file.
* **writelines(sequence)** - Write a sequence of strings to the file.
Following are file object's most used attributes −
* **closed** - bool indicating the current state of the file object.
* **encoding** - The encoding that this file uses.
* **mode**- The I/O mode for the file.
* **name** - If the file object was created using open(), the name of the file. Otherwise, some string that indicates the source of the file object
---
*With thanks to this reference article:
Neptune.ai blogs - How to Deal with Files in Google Colab: Everything You Need to Know, https://neptune.ai/blog/google-colab-dealing-with-files-2*
---
| github_jupyter |
# Logging
A lot of what we've been doing with printing output is to help debug issues but there is a better way to do that,
logging! Logging has some inherent benefits of use versus just outputting everything. With logs, we can control whats
being outputted, define what information we want to put, control how things look, and also setting where the logs live.
Logging allows us to set severity levels to our output so that we can filter out things that are low level logs vs
high priority critical issues. Having a slew of output is nice when we want to go back and parse logs to see where things
went wrong but it can output so much data that is hard to follow actively. Instead, we can configure it so that we only
get warnings and errors to log to the console while everything else goes to a log file (which can be looked at for
further detail). Let's take a look at what all we can do!
## Loggers
Within logging, there are two main types of loggers that we will be introducing: `stream` and `file`. For more information
on other types of handlers, refer to the [documentation](https://docs.python.org/3/library/logging.handlers.html)
### StreamHandler
A `StreamHandler`, as the name suggests, will be outputting the logs directly to some stream which can support a `write()`
and `flush()` operation. For now, you can think of the StreamHandler as what is user to print to the console. The benefit
of using a StreamHandler is that you can directly see the output as the program is running. The problem may occur that
there are too many logs flying by to immediately recognize anything. People use formatting and colors to quickly distinguish
how the programming is running. For example, any red text may indicate a failure.
### FileHandler
Working very much similar to a StreamHandler, a `FileHandler` writes to a specified file. Anytime the log function is
called with a string, we log that string in a file (which can be viewed or parsed at some time).
## Logging Levels
As previously mentioned, the benefit of logging vs printing directly is that we can set levels to our handlers so that
we don't end up blasting everything at the user. We may choose to only throw specific critical failure messages to the
`StreamHandler` so that we can quickly tell if there are any failures and log everything using the `FileHandler` so that
we can revisit the log later if needed. When it comes to logging levels, refer to the follow ordered list of importance,
starting with the least important.
1. Debug
2. Info
3. Warn
4. Error
5. Critical
When specifying a level for your log, everything that is prioritized above your level will also be printed. What this
means is that if you set the `WARN` level, you will also get `ERROR` and `CRITICAL` since they are even more higher
priority than the other levels. Typically, this makes sense because if you want warnings of a lower level, you will
definitely want higher level failures as well.
## Logging Example
Let's take a look at a logger example below:
```
import logging
# create our logger
logger = logging.getLogger('logger')
# we need to set this here because the default will set to WARN
logger.setLevel(logging.DEBUG)
# create our handlers that we want to use with logger
console_handler = logging.StreamHandler()
console_handler.setLevel(logging.WARNING)
file_handler = logging.FileHandler("log_file_name.log")
file_handler.setLevel(logging.DEBUG)
logger.addHandler(console_handler)
logger.addHandler(file_handler)
print(logger.getEffectiveLevel())
# this will show up on both
logger.critical("This is a critical message")
# this will only show on the file
logger.debug("This is a debug message")
```
As noted in the code above, when creating a logger, the default is set to `WARN`. So even though our `FileHandler` wants
to go as low level as a debug, it will never get there because the logger itself doesn't capture anything below a `WARN`
to pass on to its handlers. Your logger level will determine what is passed down to the handlers, which can have their
own levels.
[Up Next: Lesson 4 - Exceptions](errors.ipynb)
[Go Back: Lessons 4 - Packages, Input/Output, & Exceptions](index.ipynb)
| github_jupyter |
# How to plot spatial gene expression estimates created with the splotch DE workflow?
This script recreates figure S7D (HE) in SM-Omics: An automated platform for high-throughput spatial multi-omics; doi: https://doi.org/10.1101/2020.10.14.338418
Load libraries
```
%matplotlib inline
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import glob
import pickle
import operator
import matplotlib
import scipy.stats as stats
import statsmodels.stats.multitest as multi
from itertools import chain
plt.rcParams['figure.figsize'] = [15, 10]
import warnings; warnings.simplefilter('ignore')
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
# raw data files needed to run are available at SCP: https://singlecell.broadinstitute.org/single_cell/study/SCP979/
# please download: BF-beta_cortex_he.tsv, aba_he.zip
```
Load definitions
```
def Extract(lst):
return list(next(zip(*lst)))
def ftest(st_spec_cutoff,aba_spec_cutoff):
bb_count = 0
fisher_dict = {}
pval_list = []
for condition, df in st_spec_cutoff.groupby('condition_1'):
regions_tmp = list(set(st_spec_cutoff['AAR1'].tolist()))
regions = [x for x in regions_tmp if str(x) != 'nan']
for i in regions:
for j in regions:
#SM-Omics genes
st_genes = df[df['AAR1'] == i]['gene_new'].tolist()
# ABA-genes
aba_genes = aba_spec_cutoff[aba_spec_cutoff['ABA_region'] == j]['gene-symbol'].tolist()
# SM-Omics genes in all other regions
st_rest = df[df['AAR1'] != i]['gene_new'].tolist()
# ABA genes in all other regions
aba_rest = aba_spec_cutoff[aba_spec_cutoff['ABA_region'] != j]['gene-symbol'].tolist()
# g1 = genes in both ST and ABA
# g2 = genes unique to ST
# g3 = genes unique to ABA
# g4 = genes neither in st or aba region but in the other regions
g1 = len(list(set(st_genes).intersection(aba_genes)))
g2 = len(list(set(aba_genes).difference(set(st_genes))))
g3 = len(list(set(st_genes).difference(set(aba_genes))))
g4 = len(list(set(st_rest).intersection(aba_rest)))
# Fisher's test
oddsratio, pvalue = stats.fisher_exact([[g4, g2], [g3, g1]], alternative='greater')
# Store pvalues in list to use for multiple corrections testing
pval_list.append(pvalue)
# Store fisher's test results in DF
ff = [condition, i, j, oddsratio, pvalue, g1]
# print(i, j, g1, g2, g3, g4, pvalue)
if bb_count == 0:
fisher_dict[bb_count] = ff
df_ff = pd.DataFrame.from_dict(fisher_dict)
df_ff['idx'] = ['condition', 'AAR_ST', 'AAR_ABA','Odds ratio', 'p value', 'Num shared genes']
df_ff.set_index('idx', inplace = True)
bb_count += 1
else:
df_ff[bb_count] = ff
bb_count += 1
return pval_list, df_ff
# Load ABA ref files
path = '../../smomics_data/aba_HE'
aba_dict = []
# Read files
for filename in glob.glob(os.path.join(path, 'aba*.csv')):
# Get name of ABA region
name = filename.split('/')[-1].split('.')[0][3:].upper()
file = pd.read_csv(filename, index_col=0)
file['ABA_region'] = name
# Only keep certain columns
aba = file[['gene-symbol', 'fold-change', 'ABA_region', 'target-sum', 'contrast-sum', 'num-target-samples']]
# Calculate expression threshold
aba['Expression threshold'] = aba['target-sum'] / aba['num-target-samples']
# Only save genes which have fold-change > xx
aba = aba[aba['fold-change'] > 2.5]
aba_dict.append(aba)
aba_spec = pd.concat(aba_dict)
# merge region names
aba_spec['ABA_region'] = aba_spec['ABA_region'].replace({'PAA' : 'PIR',
'TR' : 'PIR',
'PRT' : 'MB',
'PAG' : 'MB'})
## Top ABA genes per region
ABA_top_gene_dict = {}
for label, df in aba_spec.groupby('ABA_region'):
print(label)
print(df.sort_values(by=['fold-change'], ascending=False)['gene-symbol'].head(5).tolist())
ABA_top_gene_dict[label] = df.sort_values(by='fold-change', ascending=False)['gene-symbol'].tolist()
# Load SM-Omics files
path = '../../smomics_data/'
# Read file
filename = os.path.join(path, 'BF-beta_cortex_he.tsv')
st_file = pd.read_csv(filename, index_col=0, sep='\t')
# Only compare one region to all the rest and positively expressed genes
st_spec = st_file[(st_file['AAR2'] == 'Rest') & (st_file['Delta'] > 0)]
# Log10 BF
st_spec['logBF'] = np.log(st_spec['BF'])
# merge region names
st_spec['AAR1'] = st_spec['AAR1'].replace({'Cerebral nuclei':'CNU',
'Cortical subplate':'CTXSP',
'Fiber tracts': 'nan',
'Hippocampal formation': 'HIP',
'Hypothalamus':'HY',
'Isocortex':'ISOCORTEX',
'Midbrain':'TH',
'Olfactory areas':'PIR',
'Thalamus':'TH',
'Rest':'Rest'})
# rename gene names
st_spec['gene_new'] = Extract(st_spec['gene'].str.split("_",0))
## Top ST genes per condition and per region
ST_top_gene_dict = {}
for label, df in st_spec.groupby(['condition_1', 'AAR1']):
print(label[1])
print(df.sort_values(by='logBF', ascending=False)['gene_new'].head(5).tolist())
ST_top_gene_dict[label[1]] = df.sort_values(by='logBF', ascending=False)['gene'].tolist()
# nan region denotes fiber trackts without ABA DE API enabled
```
### Merge aba and sm-omics by gene names
```
st_cutoff = 0.2
aba_cutoff = 1
merge_dict = {}
# Keep ABA-genes above cutoff
aba_spec_cutoff_tmp = aba_spec[aba_spec['Expression threshold'] > aba_cutoff]
# Get aba genes in all regions
aba_spec_cutoff_genes = aba_spec_cutoff_tmp['gene-symbol'].tolist()
# Keep ST-genes above cutoff
st_spec_cutoff_tmp = st_spec[st_spec['logBF'] > st_cutoff]
# Get st genes in all regions
st_spec_cutoff_genes = st_spec_cutoff_tmp['gene_new'].tolist()
# Common genes
common_genes = set(aba_spec_cutoff_genes).intersection(st_spec_cutoff_genes)
print("Common genes: ", len(common_genes))
# Keep those genes in both aba and st
aba_spec_cutoff = aba_spec_cutoff_tmp[aba_spec_cutoff_tmp['gene-symbol'].isin(common_genes)]
st_spec_cutoff = st_spec_cutoff_tmp[st_spec_cutoff_tmp['gene_new'].isin(common_genes)]
# DIFFERENTIAL GENES PER REGION - Fisher's exact test
pval_list, df_ff = ftest(st_spec_cutoff,aba_spec_cutoff)
# Do multiple testing correction on the pvalues
pp = multi.multipletests(pval_list, alpha=0.05, method='fdr_bh', is_sorted=False, returnsorted=False)
# Add corrected p-values
df_ff_t = df_ff.T
df_ff_t['p-value, corrected'] = list(pp[1])
# Plot enrichement heatmap
fig = plt.figure(figsize=(20, 10))
ax1 = plt.subplot2grid((2, 2), (0, 0))
axes = [ax1]
num_cond = list(range(0, len(list(set(df_ff_t['condition'])))))
i=0
for condition, df in df_ff_t.groupby('condition'):
# First make df into pivot table
pivot_df = df.pivot(index='AAR_ST', columns='AAR_ABA', values='p-value, corrected').sort_index(ascending=0)
# Might not be necessary for real values
pivot_df = pivot_df.astype(str).astype(float) # For some unexpected reason, i get objects as dtype instead of integers, this is changed here.
# Plot
ax=axes[num_cond[i]]
vmin = 0
vmax = 0.05
sns.heatmap(pivot_df,annot=True, cmap="YlGnBu", ax=ax, vmin=vmin, vmax=vmax, linewidth = 0.5,cbar_kws={'label': 'p-value, corrected'})
# Set axis labels
ax.set_xlabel('SM-Omics region', fontsize=12)
ax.set_ylabel('ABA region', fontsize=12)
ax.axhline(y=0, color='k',linewidth=5)
ax.axhline(y=pivot_df.shape[1], color='k',linewidth=5)
ax.axvline(x=0, color='k',linewidth=5)
ax.axvline(x=pivot_df.shape[0], color='k',linewidth=5)
i+=1
#plt.show()
#fig.set_size_inches(12, 10)
#plt.savefig("HE_splotch_Heatmap.pdf")
```
# Print genes that are top most expressed in both ABA and SM-Omics per each region
```
from collections import defaultdict
top_gene_dict = dict()
df = pd.DataFrame()
for k, v in ST_top_gene_dict.items():
if k == 'nan':
continue
for i, STgene in enumerate([i.split("_")[0] for i in v]):
for j, ABAgene in enumerate(ABA_top_gene_dict[k]):
if ABAgene == STgene:
top_gene_dict[STgene] = i+j
df[k+'_genes'] = dict(sorted(top_gene_dict.items(), key=operator.itemgetter(1), reverse=True)[:10]).keys()
df[k+'_counts'] = dict(sorted(top_gene_dict.items(), key=operator.itemgetter(1), reverse=True)[:10]).values()
top_gene_dict = {}
df
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Load-Ni-Mo-data" data-toc-modified-id="Load-Ni-Mo-data-1"><span class="toc-item-num">1 </span>Load Ni-Mo data</a></span></li><li><span><a href="#Set-up-the-MTP-and-train" data-toc-modified-id="Set-up-the-MTP-and-train-2"><span class="toc-item-num">2 </span>Set up the MTP and train</a></span></li><li><span><a href="#Predict-and-evaluate-the-energies-and-forces-of-training-data" data-toc-modified-id="Predict-and-evaluate-the-energies-and-forces-of-training-data-3"><span class="toc-item-num">3 </span>Predict and evaluate the energies and forces of training data</a></span></li><li><span><a href="#Write-and-load-fitted-mtp-with-parameters-files" data-toc-modified-id="Write-and-load-fitted-mtp-with-parameters-files-4"><span class="toc-item-num">4 </span>Write and load fitted mtp with parameters files</a></span></li><li><span><a href="#Lattice-constants-and-eslastic-constants" data-toc-modified-id="Lattice-constants-and-eslastic-constants-5"><span class="toc-item-num">5 </span>Lattice constants and eslastic constants</a></span></li><li><span><a href="#Surface-energy-calculation---Ni-as-an-example" data-toc-modified-id="Surface-energy-calculation---Ni-as-an-example-6"><span class="toc-item-num">6 </span>Surface energy calculation - Ni as an example</a></span></li><li><span><a href="#Energy,-force,-stress-prediction" data-toc-modified-id="Energy,-force,-stress-prediction-7"><span class="toc-item-num">7 </span>Energy, force, stress prediction</a></span></li></ul></div>
# Load Ni-Mo data
```
from pymatgen.core import Structure
from monty.serialization import loadfn
data = loadfn('data.json')
train_structures = [d['structure'] for d in data]
train_energies = [d['outputs']['energy'] for d in data]
train_forces = [d['outputs']['forces'] for d in data]
train_stresses = [d['outputs']['stress'] for d in data]
```
# Set up the MTP and train
```
from maml.apps.pes import MTPotential
mtp = MTPotential()
mtp.train(train_structures=train_structures, train_energies=train_energies,
train_forces=train_forces, train_stresses = None, max_dist=5, stress_weight=0)
```
# Predict and evaluate the energies and forces of training data
```
df_orig, df_predict = mtp.evaluate(test_structures=train_structures,
test_energies=train_energies,
test_forces=train_forces, test_stresses=train_stresses)
from sklearn.metrics import mean_absolute_error
import numpy as np
E_p = np.array(df_predict[df_predict['dtype'] == 'energy']['y_orig'])/df_predict[df_predict['dtype'] == 'energy']['n']
E_o = np.array(df_orig[df_orig['dtype'] == 'energy']['y_orig'])/df_orig[df_orig['dtype'] == 'energy']['n']
print("MAE of training energy prediction is {} meV/atom".format(mean_absolute_error(E_o,E_p)*1000))
F_p = np.array(df_predict[df_predict['dtype'] == 'force']['y_orig'])/df_predict[df_predict['dtype'] == 'force']['n']
F_o = np.array(df_orig[df_orig['dtype'] == 'force']['y_orig'])/df_orig[df_orig['dtype'] == 'force']['n']
print("MAE of training force prediction is {} eV/Å".format(mean_absolute_error(F_o,F_p)))
```
# Write and load fitted mtp with parameters files
```
mtp.write_param(fitted_mtp='fitted.mtp')
mtp_loaded = MTPotential.from_config(filename='fitted.mtp', elements=["Ni", "Mo"])
```
# Lattice constants and eslastic constants
Large error due to limited training data -- 10 structures
```
from pymatgen.core import Lattice
Ni = Structure.from_spacegroup(sg='Fm-3m', species=['Ni'], lattice=Lattice.cubic(3.51), coords=[[0, 0, 0]])
Mo = Structure.from_spacegroup(sg='Im-3m', species=['Mo'], lattice=Lattice.cubic(3.17), coords=[[0, 0, 0]])
from maml.apps.pes import LatticeConstant
lc_calculator = LatticeConstant(ff_settings=mtp_loaded)
a, b, c = lc_calculator.calculate([Ni])[0]
print('Ni', 'Lattice a: {}, Lattice b: {}, Lattice c: {}'.format(a, b, c))
lc_calculator = LatticeConstant(ff_settings=mtp_loaded)
a, b, c = lc_calculator.calculate([Mo])[0]
print('Mo', 'Lattice a: {}, Lattice b: {}, Lattice c: {}'.format(a, b, c))
from maml.apps.pes import ElasticConstant
Ni_ec_calculator = ElasticConstant(ff_settings=mtp_loaded)
Ni_C11, Ni_C12, Ni_C44, _ = Ni_ec_calculator.calculate([Ni])[0]
print('Ni', ' C11: ', Ni_C11, 'C12: ', Ni_C12, 'C44: ', Ni_C44)
Mo_ec_calculator = ElasticConstant(ff_settings=mtp_loaded)
Mo_C11, Mo_C12, Mo_C44, _ = Mo_ec_calculator.calculate([Mo])[0]
print('Mo', ' C11: ', Mo_C11, 'C12: ', Mo_C12, 'C44: ', Mo_C44)
```
# Surface energy calculation - Ni as an example
```
from maml.apps.pes import SurfaceEnergy
mtp_loaded_Ni = MTPotential.from_config(filename='fitted.mtp.Ni', elements=["Ni"])
surface_e_calculator = SurfaceEnergy(ff_settings=mtp_loaded_Ni,
bulk_structure=Ni,
miller_indexes=[[1,0,0],[0,1,0],[1,1,0]])
results_surface = surface_e_calculator.calculate()
relaxed_surface_structures = [result[1] for result in results_surface]
print("Surface energys in Ni:")
for result in results_surface:
print(f"Miller index: {result[0]}, surface energy: {result[2]} J/m^2, slab model has {result[1].num_sites} atoms")
```
# Energy, force, stress prediction
```
from maml.apps.pes import EnergyForceStress
efs_calculator = EnergyForceStress(ff_settings=mtp_loaded)
energy, forces, stresses = efs_calculator.calculate([train_structures[0]])[0]
print('energy: {}'.format(energy))
print('forces: \n', forces)
print('stresses: ', stresses)
```
| github_jupyter |
### Seminar 9: exploration vs exploitation
In this seminar, we'll employ bayesian neural networks to facilitate exploration in contextual bandits.
__About bayesian neural networks:__
* A post on the matter - [url](http://twiecki.github.io/blog/2016/07/05/bayesian-deep-learning/)
* Theano+PyMC3 for more serious stuff - [url](http://pymc-devs.github.io/pymc3/notebooks/bayesian_neural_network_advi.html)
* Same stuff in tensorflow - [url](http://edwardlib.org/tutorials/bayesian-neural-network)
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
### Load data
In this seminar, we're going to solve a toy contextual bandit problem
* 60-dimensional states
* 10 actions
* rewards between 0 and 1
Instead of actually running on a stream of data, we're gonna emulate it with samples from dataset.
```
all_states = np.load("all_states.npy")
action_rewards = np.load("action_rewards.npy")
state_size = all_states.shape[1]
n_actions = action_rewards.shape[1]
print ("%id states, %i actions"%(state_size,n_actions))
import theano, theano.tensor as T
import lasagne
from lasagne import init
from lasagne.layers import *
import bayes
as_bayesian = bayes.bbpwrap(bayes.NormalApproximation(std=0.1))
BayesDenseLayer = as_bayesian(DenseLayer)
#similar: BayesConv2DLayer = as_bayesian(Conv2DLayer)
class Bandit:
"""a bandit with bayesian neural net"""
def __init__(self,state_size=state_size,n_actions=n_actions):
#input variables
input_states = T.matrix("states")
target_actions = T.ivector("actions taken")
target_rewards = T.vector("rewards")
self.total_samples_seen = theano.shared(np.int32(0),"number of training samples seen so far")
batch_size = target_actions.shape[0]
###
#network body
inp = InputLayer((None,state_size),name='input')
hid = <create bayesian dense layer for hidden states>
out = <create bayesian dense layer that predicts Q's aka actions>
###
#prediction
prediction_all_actions = get_output(out,inputs=input_states)
self.predict_sample_rewards = theano.function([input_states],prediction_all_actions)
###
#Training
#select prediction for target action
prediction_target_actions = prediction_all_actions[T.arange(batch_size),target_actions]
#loss = negative log-likelihood (mse) + KL
negative_llh = T.sum((prediction_target_actions - target_rewards)**2)
kl = bayes.get_var_cost(out) / (self.total_samples_seen+batch_size)
loss = (negative_llh + kl)/batch_size
self.weights = get_all_params(out,trainable=True)
self.out=out
#gradient descent
updates = lasagne.updates.adam(loss,self.weights)
#update counts
updates[self.total_samples_seen]=self.total_samples_seen+batch_size.astype('int32')
self.train_step = theano.function([input_states,target_actions,target_rewards],
[negative_llh,kl],updates = updates,
allow_input_downcast=True)
def sample_prediction(self,states,n_samples=1):
"""Samples n_samples predictions for rewards,
:returns: tensor [n_samples,state_i,action_i]
"""
assert states.ndim==2,"states must be 2-dimensional"
return np.stack([self.predict_sample_rewards(states) for _ in range(n_samples)])
epsilon=0.25
def get_action(self,states):
"""
Picks action by
- with p=1-epsilon, taking argmax of average rewards
- with p=epsilon, taking random action
This is exactly e-greedy policy.
"""
reward_samples = self.sample_prediction(states,n_samples=100)
#^-- samples for rewards, shape = [n_samples,n_states,n_actions]
best_actions = reward_samples.mean(axis=0).argmax(axis=-1)
#^-- we take mean over samples to compute expectation, then pick best action with argmax
random_actions = <generate random actions>
chosen_actions = <pick actions with e-greedy policy>
return chosen_actions
def train(self,states,actions,rewards,n_iters=10):
"""
trains to predict rewards for chosen actions in given states
"""
loss_sum = kl_sum = 0
for _ in range(n_iters):
loss,kl = self.train_step(states,actions,rewards)
loss_sum += loss
kl_sum += kl
return loss_sum/n_iters,kl_sum/n_iters
```
### Train the bandit
We emulate infinite stream of data and pick actions using agent's get_action function.
```
bandit = Bandit() #create your bandit
rewards_history = []
def get_new_samples(states,action_rewards,batch_size=10):
"""samples random minibatch, emulating new users"""
batch_ix = np.random.randint(0,len(states),batch_size)
return states[batch_ix],action_rewards[batch_ix]
from IPython.display import clear_output
from pandas import ewma
batch_size=10 #10 new users
for i in range(1000):
###
#new data
b_states,b_action_rewards = get_new_samples(all_states,action_rewards,batch_size)
###
#pick actions
b_actions = bandit.get_action(b_states)
###
#rewards for actions agent just took
b_rewards = b_action_rewards[np.arange(batch_size),b_actions]
###
#train bandit
mse,kl = bandit.train(b_states,b_actions,b_rewards,n_iters=100)
rewards_history.append(b_rewards.mean())
if i%10 ==0:
clear_output(True)
print("iteration #%i\tmean reward=%.3f\tmse=%.3f\tkl=%.3f"%(i,np.mean(rewards_history[-10:]),mse,kl))
plt.plot(rewards_history)
plt.plot(ewma(np.array(rewards_history),alpha=0.1))
plt.show()
samples = bandit.sample_prediction(b_states[:1],n_samples=100).T[:,0,:]
for i in range(len(samples)):
plt.hist(samples[i],alpha=0.25,label=str(i))
plt.legend(loc='best')
print('Q(s,a) std:', ';'.join(list(map('{:.3f}'.format,np.std(samples,axis=1)))))
print('correct',b_action_rewards[0].argmax())
plt.show()
```
## Better exploration
You will now implement the two exploration strategies from the lecture.
```
#then implement it and replace bandit = Bandit() above with ThompsonBandit()
class ThompsonBandit(Bandit):
def get_action(self,states):
"""
picks action based by taking _one_ sample from BNN and taking action with highest sampled reward (yes, that simple)
This is exactly thompson sampling.
"""
<your code>
return <your code>
#then implement it and replace bandit = Bandit() above with UCBBandit()
class UCBBandit(Bandit):
q = 90
def get_action(self,states):
"""
Compute q-th percentile of rewards P(r|s,a) for all actions
Take actions that have highest percentiles.
This implements bayesian UCB strategy
"""
<Your code here>
return <actions with bayesian ucb>
bandit = <UCBBandit or ThompsonBandit>
#<maybe change parameters>
rewards_history = []
from IPython.display import clear_output
from pandas import ewma
batch_size=10 #10 new users
for i in range(1000):
###
#new data
b_states,b_action_rewards = get_new_samples(all_states,action_rewards,batch_size)
###
#pick actions
b_actions = bandit.get_action(b_states)
###
#rewards for actions agent just took
b_rewards = b_action_rewards[np.arange(batch_size),b_actions]
###
#train bandit
mse,kl = bandit.train(b_states,b_actions,b_rewards,n_iters=100)
rewards_history.append(b_rewards.mean())
if i%10 ==0:
clear_output(True)
print("iteration #%i\tmean reward=%.3f\tmse=%.3f\tkl=%.3f"%(i,np.mean(rewards_history[-10:]),mse,kl))
plt.plot(rewards_history)
plt.plot(ewma(np.array(rewards_history),alpha=0.1))
plt.show()
samples = bandit.sample_prediction(b_states[:1],n_samples=100).T[:,0,:]
for i in range(len(samples)):
plt.hist(samples[i],alpha=0.25,label=str(i))
plt.legend(loc='best')
print('Q(s,a) std:', ';'.join(list(map('{:.3f}'.format,np.std(samples,axis=1)))))
print('correct',b_action_rewards[0].argmax())
plt.show()
```
### Experience replay
Our value-based bandit algorithm is off-policy, so we can train it on actions from a different policy.
For example, the bandit will need much less interactions to converge if you train it on past experiences. You can also pre-train it on any data you already have.
```
class ReplayBandit(Bandit): #or your preferred exploration type
"""A bandit that trains not on last user interactions but on random samples from everything it saw"""
experience_buffer=[]
<Your code here. You will at least need to modify train function>
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
```
| github_jupyter |
```
from os.path import join
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_addons as tfa
from sklearn.model_selection import train_test_split
import tensorflow.keras.backend as K
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, LSTM, Dense, Embedding, Bidirectional, LSTMCell
from tensorflow.keras.losses import SparseCategoricalCrossentropy
from generate_uncorrect_sample import generate_misspell_sample
```
### Define class for creating and loading data
```
def loss_fn(y_pred, y):
log_loss = SparseCategoricalCrossentropy(from_logits=True, reduction='none')
loss = log_loss(y_true=y, y_pred=y_pred)
mask = tf.logical_not(tf.math.equal(y, 0)) # output 0 for y=0 else output 1
mask = tf.cast(mask, dtype=loss.dtype)
loss = mask * loss
loss = tf.reduce_mean(loss)
return loss
def generate_pair_samples(w):
w_gen = list(generate_misspell_sample(w, max_edit_distance=2))
return list(zip(w_gen, [w]*len(w_gen)))
class Text2Seq(object):
def __init__(self, charset,
start_token='<s>',
end_token='<e>',
unknown_token='<unk>'):
self.start_token = start_token
self.end_token = end_token
self.unk_token = unknown_token
if isinstance(charset, str):
with open(charset, 'r+') as f:
self.charset = set(f.read().split('\n'))
else:
self.charset = charset
self.charset += [' ', self.start_token, self.end_token, self.unk_token]
self.charset = set(self.charset)
self.charset_size = len(self.charset)
self.char2id = {j: i for i, j in enumerate(self.charset, start=1)}
self.id2char = {j: i for i, j in self.char2id.items()}
def _encode(self, word, max_len, pad_start_end):
padded = []
for c in word:
padded.append(self.char2id.get(c, self.char2id[self.unk_token]))
if pad_start_end:
padded = [self.char2id[self.start_token]] + padded + [self.char2id[self.end_token]]
padded += (max_len + 2 - len(padded)) * [0]
else:
padded += (max_len - len(padded)) * [0]
return padded
def fit_on_texts(self, texts, pad_start_end=False):
max_len = self.get_max_seq_len(texts)
arr = []
for word in texts:
arr.append(self._encode(word, max_len, pad_start_end))
return np.array(arr, dtype=np.int8)
@staticmethod
def get_max_seq_len(texts):
return max(len(word) for word in texts)
def sequence_to_text(self, arr, remove_endtoken=False):
def _inside(arr):
word = []
for i in arr:
if i!=0:
if remove_endtoken:
if i==self.char2id.get(self.end_token):
break
word.append(self.id2char.get(i, self.unk_token))
return ''.join(word)
result = []
for a in arr:
result.append(_inside(a))
return result
```
### Create dataset for training
```
pairs = []
correct_words = ['có thể', 'thế giới', 'con người', 'không thể', 'tất cả', 'chúng ta']
for w in correct_words:
pairs.extend(generate_pair_samples(w))
df = pd.DataFrame(pairs, columns=['misspell', 'correct']).sample(frac=1, random_state=123)
charset = list(set(''.join(df.misspell.values+df.correct.values)))
text2seq = Text2Seq(charset)
X_train, X_test, Y_train, Y_test = train_test_split(text2seq.fit_on_texts(df.misspell.values),
text2seq.fit_on_texts(df.correct.values, pad_start_end=True),
test_size=0.1)
BATCH_SIZE = 4
BUFFER_SIZE = len(X_train)
steps_per_epoch = BUFFER_SIZE // BATCH_SIZE
embedding_dims = 64
rnn_units = dense_units = 64
Tx = X_train.shape[1]
Ty = Y_train.shape[1]
input_vocab_size = output_vocab_size = text2seq.charset_size+1
train_dataset = tf.data.Dataset.from_tensor_slices((X_train, Y_train)).shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
test_dataset = tf.data.Dataset.from_tensor_slices((X_test, Y_test)).shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
```
### Define model
```
class Encoder(Model):
def __init__(self, input_vocab_size=None, embedding_dims=128, rnn_units=64):
super(Encoder, self).__init__()
self.encoder_embedding = Embedding(input_vocab_size, embedding_dims)
self.encoder_birnn = Bidirectional(LSTM(rnn_units, return_sequences=True, dropout=0.2))
self.encoder_stackrnn = LSTM(rnn_units, return_sequences=True, return_state=True)
def call(self, inputs):
x = self.encoder_embedding(inputs)
x = self.encoder_birnn(x)
x = self.encoder_stackrnn(x)
return x
class Decoder(Model):
def __init__(self,
output_vocab_size=None,
embedding_dims=128,
rnn_units=64,
dense_units=64,
batch_size=128,
encoder_max_seq_len=None,
decoder_max_seq_len=None,
start_token=None,
end_token=None,
beam_width=5,
training=True):
super().__init__()
self.batch_size = batch_size
self.decoder_max_seq_len = decoder_max_seq_len
self.decoder_embedding = Embedding(output_vocab_size, embedding_dims)
self.dense_layer = Dense(output_vocab_size)
self.rnn_cell = LSTMCell(rnn_units)
self.start_token = start_token
self.end_token = end_token
self.beam_width = beam_width
self.training = training
# training phase
self.sampler = tfa.seq2seq.sampler.TrainingSampler()
self.attn_mech = tfa.seq2seq.LuongAttention(dense_units,
None,
self.batch_size * [encoder_max_seq_len])
self.attn_cell = tfa.seq2seq.AttentionWrapper(self.rnn_cell,
self.attn_mech,
dense_units)
self.decoder = tfa.seq2seq.BasicDecoder(self.attn_cell, self.sampler, self.dense_layer)
def set_decoder_memory_and_initialState(self, memory, batch_size, encoder_state):
self.attn_mech.setup_memory(memory)
decoder_initial_state = self.attn_cell.get_initial_state(batch_size=batch_size, dtype=tf.float32)
decoder_initial_state = decoder_initial_state.clone(cell_state=encoder_state)
return decoder_initial_state
def call(self, inputs):
d_in, encoder_outputs, state_h, state_c = inputs
if self.training:
decoder_emb = self.decoder_embedding(d_in)
decoder_initial_state = self.set_decoder_memory_and_initialState(encoder_outputs,
self.batch_size,
[state_h, state_c])
outputs, _, _ = self.decoder(decoder_emb,
initial_state=decoder_initial_state,
sequence_length=self.batch_size * [self.decoder_max_seq_len - 1])
logits = outputs.rnn_output
return logits
else:
inference_batch_size = 1
_ = self.decoder_embedding(d_in)
encoder_state_beam = tfa.seq2seq.tile_batch([state_h, state_c], self.beam_width)
encoder_outputs_beam = tfa.seq2seq.tile_batch(encoder_outputs, self.beam_width)
decoder_initial_state = self.set_decoder_memory_and_initialState(encoder_outputs_beam,
inference_batch_size*self.beam_width,
encoder_state_beam)
decoder_instance = tfa.seq2seq.BeamSearchDecoder(self.attn_cell,
beam_width=self.beam_width,
output_layer=self.dense_layer)
start_tokens = tf.fill([inference_batch_size], self.start_token)
end_token = self.end_token
_, inputs, state = decoder_instance.initialize(self.decoder_embedding.variables[0] ,
start_tokens=start_tokens,
end_token=end_token,
initial_state=decoder_initial_state)
beam_ids = []
beam_scores = []
for j in range(self.decoder_max_seq_len):
beam_output, state, inputs, _ = decoder_instance.step(j, inputs, state)
beam_ids.append(beam_output.predicted_ids)
beam_scores.append(beam_output.scores)
return beam_ids, beam_scores
class EncoderDecoder():
def __init__(self,
input_vocab_size=None,
output_vocab_size=None,
embedding_dims=128,
rnn_units=64,
dense_units=64,
batch_size=128,
encoder_max_seq_len=None,
decoder_max_seq_len=None,
start_token=None,
end_token=None,
beam_width=5,
training=None):
self.start_token = start_token
self.batch_size = batch_size
self.training = training
self.encoder = Encoder(input_vocab_size=input_vocab_size,
embedding_dims=embedding_dims,
rnn_units=rnn_units)
self.decoder = Decoder(output_vocab_size=output_vocab_size,
embedding_dims=embedding_dims,
rnn_units=rnn_units,
dense_units=dense_units,
batch_size=batch_size,
encoder_max_seq_len=encoder_max_seq_len,
decoder_max_seq_len=decoder_max_seq_len,
start_token=start_token,
end_token=end_token,
beam_width=beam_width,
training=training
)
def __call__(self, inputs):
# encode phase
e_in, d_in = inputs
e_out, state_h, state_c = self.encoder(e_in)
# decode phase
return self.decoder([d_in, e_out, state_h, state_c])
def compile(self, optimizer, loss=None, metrics=None):
self.optimizer = optimizer
self.loss_fn = loss
self.metrics = metrics
def _step(self, x_batch, y_batch):
d_in = y_batch[:, :-1] # ignore <end>
d_out = y_batch[:, 1:] # ignore <start>
logits = self([x_batch, d_in])
loss = self.loss_fn(logits, d_out)
return loss
@tf.function
def train_step(self, x_batch, y_batch):
with tf.GradientTape() as tape:
loss = self._step(x_batch, y_batch)
vars_ = self.encoder.trainable_variables + self.decoder.trainable_variables # be careful
grads = tape.gradient(loss, vars_)
self.optimizer.apply_gradients(zip(grads, vars_))
return loss
def fit(self, train_dataset, epochs=1, eval_dataset=None):
num_train_samples = tf.data.experimental.cardinality(train_dataset).numpy() * self.batch_size
for epoch in range(epochs):
print("\nepoch {}/{}".format(epoch+1,epochs))
pbar = tf.keras.utils.Progbar(num_train_samples, stateful_metrics=['train_loss'])
for i, (x_batch, y_batch) in enumerate(train_dataset):
train_loss = self.train_step(x_batch, y_batch)
values = [('train_loss', train_loss)]
pbar.update(i*self.batch_size, values=values)
if eval_dataset is not None:
for x_batch, y_batch in test_dataset:
val_loss = self._step(x_batch, y_batch)
values=[('train_loss',train_loss),('val_loss',val_loss)]
else:
values=[('train_loss',train_loss)]
pbar.update(num_train_samples, values=values)
def save_weights(self, path):
self.encoder.save_weights(join(path, 'encoder_weights.h5'))
self.decoder.save_weights(join(path, 'decoder_weights.h5'))
@classmethod
def from_pretrained(cls,
path,
input_vocab_size,
output_vocab_size,
embedding_dims,
rnn_units,
dense_units,
batch_size,
encoder_max_seq_len,
decoder_max_seq_len,
start_token,
end_token,
beam_width,
training):
model = cls(input_vocab_size,
output_vocab_size,
embedding_dims,
rnn_units,
dense_units,
batch_size,
encoder_max_seq_len,
decoder_max_seq_len,
start_token,
end_token,
beam_width,
training)
model.encoder.build((None, None))
model.encoder.load_weights(join(path, 'encoder_weights.h5'))
model.decoder.build([(None, None), (None, None, rnn_units), (None, rnn_units), (None, rnn_units)])
model.decoder.load_weights(join(path, 'decoder_weights.h5'))
return model
@staticmethod
def decode_prediction(outputs):
beam_ids, beam_scores = outputs
return np.array([i.numpy() for i in beam_ids]).squeeze().transpose()
def predict(self, input_ids):
beam_outputs = self([input_ids, np.array([[self.start_token]])])
return self.decode_prediction(beam_outputs)
```
### Compile Model
```
start_token=text2seq.char2id.get('<s>')
end_token=text2seq.char2id.get('<e>')
model = EncoderDecoder(input_vocab_size,
output_vocab_size,
embedding_dims,
rnn_units,
dense_units,
batch_size=BATCH_SIZE,
encoder_max_seq_len=Tx,
decoder_max_seq_len=Ty,
start_token=start_token,
end_token=end_token,
beam_width=5,
training=True)
lr_schedule = tfa.optimizers.ExponentialCyclicalLearningRate(initial_learning_rate=5e-4,
maximal_learning_rate=1e-2,
step_size=steps_per_epoch*2,
scale_mode="cycle",
gamma=0.96)
opt = tfa.optimizers.Lookahead(tf.keras.optimizers.Adam(clipnorm=3.0, learning_rate=lr_schedule))
model.compile(optimizer=opt, loss=loss_fn)
```
### Training
```
model.fit(train_dataset, epochs=40, eval_dataset=test_dataset)
model.save_weights('model/1')
```
### Load model and do inferencing
```
loaded_model = EncoderDecoder.from_pretrained('model/1',
input_vocab_size,
output_vocab_size,
embedding_dims,
rnn_units,
dense_units,
batch_size=BATCH_SIZE,
encoder_max_seq_len=Tx,
decoder_max_seq_len=Ty,
start_token=start_token,
end_token=end_token,
beam_width=5,
training=False)
inputs = text2seq.fit_on_texts(['cơ the'])
text2seq.sequence_to_text(loaded_model.predict(inputs), True)
inputs = text2seq.fit_on_texts(['chng ta'])
text2seq.sequence_to_text(loaded_model.predict(inputs), True)
inputs = text2seq.fit_on_texts(['taast cả'])
text2seq.sequence_to_text(loaded_model.predict(inputs), True)
inputs = text2seq.fit_on_texts(['khongtheer'])
text2seq.sequence_to_text(loaded_model.predict(inputs), True)
```
| github_jupyter |
## 1. Load the mpg dataset. Read the documentation for it, and use the data to answer these questions:
```
%matplotlib inline
import numpy as np
import pandas as pd
from pydataset import data
import matplotlib.pyplot as plt
import env
# Load mpg dataset
mpg = data('mpg')
mpg.head()
```
#### `a. On average, which manufacturer has the best miles per gallon?`
```
mpg.groupby('manufacturer')['cty', 'hwy'].mean().sort_values(by='hwy')
import warnings
warnings.filterwarnings('ignore')
```
#### `b. How many different manufacturers are there?`
```
mpg.manufacturer.nunique()
mpg.manufacturer.describe()
```
#### `c. How many different models are there?`
```
mpg.model.nunique()
mpg.model.describe()
```
#### `d. Do automatic or manual cars have better miles per gallon?`
```
mpg.trans.value_counts()
mpg['trans_category'] = np.where(mpg.trans.str.startswith('a'), 'auto', 'manual')
mpg.head()
mpg.groupby('trans_category')[['cty', 'hwy']].mean()
```
# 2. Joining and Merging
#### `Copy the users and roles dataframes from the examples above.`
#### `What do you think a right join would look like?` `An outer join?`
#### `What happens if you drop the foreign keys from the dataframes and try to merge them?`
```
users = pd.DataFrame({
'id': [1, 2, 3, 4, 5, 6],
'name': ['bob', 'joe', 'sally', 'adam', 'jane', 'mike'],
'role_id': [1, 2, 3, 3, np.nan, np.nan]
})
users
roles = pd.DataFrame({
'id': [1, 2, 3, 4],
'name': ['admin', 'author', 'reviewer', 'commenter']
})
roles
a.
right_join = pd.merge(users, roles, left_on='role_id', right_on='id', how='right')
right_join
pd.merge(users,
roles,
left_on='role_id',
right_on='id',
how='outer')
```
# 3. Getting data from SQL databases
#### `a. Create a function named get_db_url. `
```
from env import host, password, user
def get_db_url(db, user=user, host=host, password=password):
return f'mysql+pymysql://{user}:{password}@{host}/{db}'
```
#### `b. Use your function to obtain a connection to the employees database.`
```
employees = pd.read_sql(sql_query, get_db_url('employees'))
employees.head()
```
#### `c.Intentionally make an error in your SQL query.`
```
sql_query = 'SELECT # From salaries'
slq_query
```
#### `d. Read the employees and titles tables into two separate dataframes`
```
sql_query = 'SELECT * FROM titles'
titles = pd.read_sql(sql_query, get_db_url('employees'))
titles.head()
titles.to_csv('titles.csv')
titles = pd.read_csv('titles.csv', index_col=0)
titles.info()
employees.info
```
#### `e. Visualize the number of employees with each title.`
```
titles.head()
titles.shape
boolean_series = titles.to_date == titles.to_date.max()
boolean_series.head()
employee_titles = titles[boolean_series]
employee_titles.head()
employee_titles.shape
titles_held = employee_titles.title.value_counts()
titles_held.head()
#Visualization
titles\
[titles.to_date == titles.to_date.max()]\
.title\
.value_counts()\
.plot.barh()
```
#### `f. Join the employees and titles dataframes together.`
```
joined_employees_titles = employees.merge(titles, on='emp_no')
joined_employees_titles.head()
```
#### ` g. Visualize how frequently employees change titles.`
```
# number of titles for each employee
joined_employees_titles.emp_no.value_counts().sample(5)
#title frequency
title_frequency = joined_employees_titles.emp_no.value_counts()
title_frequency.value_counts()
#visualization
title_frequency.value_counts().plot(kind='barh',
color='midnightblue',
width=.75)
plt.title('Employee Title Change Frequency')
plt.ylabel('How Many Title Changes')
plt.show()
```
#### ` h. For each title, find the hire date of the employee that was hired most recently with that title.`
```
joined_employees_titles.groupby('title').hire_date.max()
dept_title_query = '''
SELECT *
FROM titles t
JOIN dept_emp de USING(emp_no)
JOIN departments d USING(dept_no)
'''
dept_titles = pd.read_sql(dept_title_query, get_db_url('employees'))
dept_titles.to_csv('dept_titles.csv')
dept_titles = pd.read_csv('dept_titles.csv', index_col=0)
dept_titles.head()
dept_titles.shape
```
#### ` i. Write the code necessary to create a cross tabulation of the number of titles by department`
```
title_crosstab = pd.crosstab(dept_titles.dept_name, dept_titles.title)
title_crosstab
```
# 4. Use your get_db_url function to help you explore the data from the chipotle database. Use the data to answer the following questions:
#### `a. What is the total price for each order?`
```
chipotle_sql_query = '''
SELECT *
FROM orders;
'''
chipotle_orders = pd.read_sql(chipotle_sql_query, get_db_url('chipotle'))
chipotle_orders.head()
chipotle_orders.shape
chipotle_orders.info()
chipotle_orders['item_price'] = chipotle_orders.item_price.str.replace('$', '').astype(float)
chipotle_orders.info()
total_chipotle_orders = chipotle_orders.groupby('order_id').item_price.sum()
total_chipotle_orders
```
#### `b. What are the most popular 3 items? `
```
top_three_items = chipotle_orders.groupby('item_name').quantity.sum().sort_values(ascending = False).head(3)
top_three_items
```
#### `c. Which item has produced the most revenue?`
```
chipotle_orders.groupby('item_name').item_price.sum().nlargest(10)
chipotle_orders.groupby('order_id').item_price.sum().median()
```
| github_jupyter |
## Introduction
Word2Vec is a popular algorithm used for generating dense vector representations of words in large corpora using unsupervised learning. The resulting vectors have been shown to capture semantic relationships between the corresponding words and are used extensively for many downstream natural language processing (NLP) tasks like sentiment analysis, named entity recognition and machine translation.
SageMaker BlazingText which provides efficient implementations of Word2Vec on
- single CPU instance
- single instance with multiple GPUs - P2 or P3 instances
- multiple CPU instances (Distributed training)
In this notebook, we demonstrate how BlazingText can be used for distributed training of word2vec using multiple CPU instances.
## Setup
Let's start by specifying:
- The S3 buckets and prefixes that you want to use for saving model data and where training data is located. These should be within the same region as the Notebook Instance, training, and hosting. If you don't specify a bucket, SageMaker SDK will create a default bucket following a pre-defined naming convention in the same region.
- The IAM role ARN used to give SageMaker access to your data. It can be fetched using the **get_execution_role** method from sagemaker python SDK.
```
import sagemaker
from sagemaker import get_execution_role
import boto3
import json
sess = sagemaker.Session()
role = get_execution_role()
print(role) # This is the role that SageMaker would use to leverage AWS resources (S3, CloudWatch) on your behalf
region = boto3.Session().region_name
output_bucket = sess.default_bucket() # Replace with your own bucket name if needed
print(output_bucket)
output_prefix = "sagemaker/DEMO-blazingtext-text8" # Replace with the prefix under which you want to store the data if needed
data_bucket = f"jumpstart-cache-prod-{region}" # Replace with the bucket where your data is located
data_prefix = "1p-notebooks-datasets/text8"
```
### Data Ingestion
BlazingText expects a single preprocessed text file with space separated tokens and each line of the file should contain a single sentence. In this example, let us train the vectors on [text8](http://mattmahoney.net/dc/textdata.html) dataset (100 MB), which is a small (already preprocessed) version of Wikipedia dump. Data is already downloaded from [matt mahoney's website](http://mattmahoney.net/dc/text8.zip), uncompressed and stored in `data_bucket`.
```
train_channel = f"{data_prefix}/train"
s3_train_data = f"s3://{data_bucket}/{train_channel}"
```
Next we need to setup an output location at S3, where the model artifact will be dumped. These artifacts are also the output of the algorithm's training job.
```
s3_output_location = f"s3://{output_bucket}/{output_prefix}/output"
```
## Training Setup
Now that we are done with all the setup that is needed, we are ready to train our object detector. To begin, let us create a ``sageMaker.estimator.Estimator`` object. This estimator will launch the training job.
```
region_name = boto3.Session().region_name
container = sagemaker.amazon.amazon_estimator.get_image_uri(region_name, "blazingtext", "latest")
print(f"Using SageMaker BlazingText container: {container} ({region_name})")
```
## Training the BlazingText model for generating word vectors
Similar to the original implementation of [Word2Vec](https://arxiv.org/pdf/1301.3781.pdf), SageMaker BlazingText provides an efficient implementation of the continuous bag-of-words (CBOW) and skip-gram architectures using Negative Sampling, on CPUs and additionally on GPU[s]. The GPU implementation uses highly optimized CUDA kernels. To learn more, please refer to [*BlazingText: Scaling and Accelerating Word2Vec using Multiple GPUs*](https://dl.acm.org/citation.cfm?doid=3146347.3146354). BlazingText also supports learning of subword embeddings with CBOW and skip-gram modes. This enables BlazingText to generate vectors for out-of-vocabulary (OOV) words, as demonstrated in this [notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/blazingtext_word2vec_subwords_text8/blazingtext_word2vec_subwords_text8.ipynb).
Besides skip-gram and CBOW, SageMaker BlazingText also supports the "Batch Skipgram" mode, which uses efficient mini-batching and matrix-matrix operations ([BLAS Level 3 routines](https://software.intel.com/en-us/mkl-developer-reference-fortran-blas-level-3-routines)). This mode enables distributed word2vec training across multiple CPU nodes, allowing almost linear scale up of word2vec computation to process hundreds of millions of words per second. Please refer to [*Parallelizing Word2Vec in Shared and Distributed Memory*](https://arxiv.org/pdf/1604.04661.pdf) to learn more.
BlazingText also supports a *supervised* mode for text classification. It extends the FastText text classifier to leverage GPU acceleration using custom CUDA kernels. The model can be trained on more than a billion words in a couple of minutes using a multi-core CPU or a GPU, while achieving performance on par with the state-of-the-art deep learning text classification algorithms. For more information, please refer to [algorithm documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/blazingtext.html) or [the text classification notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/blazingtext_text_classification_dbpedia/blazingtext_text_classification_dbpedia.ipynb).
To summarize, the following modes are supported by BlazingText on different types instances:
| Modes | cbow (supports subwords training) | skipgram (supports subwords training) | batch_skipgram | supervised |
|:----------------------: |:----: |:--------: |:--------------: | :--------------: |
| Single CPU instance | ✔ | ✔ | ✔ | ✔ |
| Single GPU instance | ✔ | ✔ | | ✔ (Instance with 1 GPU only) |
| Multiple CPU instances | | | ✔ | | |
Now, let's define the resource configuration and hyperparameters to train word vectors on *text8* dataset, using "batch_skipgram" mode on two c4.2xlarge instances.
```
bt_model = sagemaker.estimator.Estimator(
container,
role,
instance_count=2,
instance_type="ml.c4.2xlarge",
train_volume_size=5,
train_max_run=360000,
input_mode="File",
output_path=s3_output_location,
sagemaker_session=sess,
)
```
Please refer to [algorithm documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/blazingtext_hyperparameters.html) for the complete list of hyperparameters.
```
bt_model.set_hyperparameters(
mode="batch_skipgram",
epochs=5,
min_count=5,
sampling_threshold=0.0001,
learning_rate=0.05,
window_size=5,
vector_dim=100,
negative_samples=5,
batch_size=11, # = (2*window_size + 1) (Preferred. Used only if mode is batch_skipgram)
evaluation=True, # Perform similarity evaluation on WS-353 dataset at the end of training
subwords=False,
) # Subword embedding learning is not supported by batch_skipgram
```
Now that the hyper-parameters are setup, let us prepare the handshake between our data channels and the algorithm. To do this, we need to create the `sagemaker.session.s3_input` objects from our data channels. These objects are then put in a simple dictionary, which the algorithm consumes.
```
train_data = sagemaker.session.s3_input(
s3_train_data, distribution="FullyReplicated", content_type="text/plain", s3_data_type="S3Prefix"
)
data_channels = {"train": train_data}
```
We have our `Estimator` object, we have set the hyper-parameters for this object and we have our data channels linked with the algorithm. The only remaining thing to do is to train the algorithm. The following command will train the algorithm. Training the algorithm involves a few steps. Firstly, the instance that we requested while creating the `Estimator` classes is provisioned and is setup with the appropriate libraries. Then, the data from our channels are downloaded into the instance. Once this is done, the training job begins. The provisioning and data downloading will take some time, depending on the size of the data. Therefore it might be a few minutes before we start getting training logs for our training jobs. The data logs will also print out `Spearman's Rho` on some pre-selected validation datasets after the training job has executed. This metric is a proxy for the quality of the algorithm.
Once the job has finished a "Job complete" message will be printed. The trained model can be found in the S3 bucket that was setup as `output_path` in the estimator.
```
bt_model.fit(inputs=data_channels, logs=True)
```
## Hosting / Inference
Once the training is done, we can deploy the trained model as an Amazon SageMaker real-time hosted endpoint. This will allow us to make predictions (or inference) from the model. Note that we don't have to host on the same type of instance that we used to train. Because instance endpoints will be up and running for long, it's advisable to choose a cheaper instance for inference.
```
bt_endpoint = bt_model.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
```
### Getting vector representations for words
#### Use JSON format for inference
The payload should contain a list of words with the key as "**instances**". BlazingText supports content-type `application/json`.
```
words = ["awesome", "blazing"]
payload = {"instances": words}
response = bt_endpoint.predict(
json.dumps(payload), initial_args={"ContentType": "application/json", "Accept": "application/json"}
)
vecs = json.loads(response)
print(vecs)
```
As expected, we get an n-dimensional vector (where n is vector_dim as specified in hyperparameters) for each of the words. If the word is not there in the training dataset, the model will return a vector of zeros.
### Evaluation
Let us now download the word vectors learned by our model and visualize them using a [t-SNE](https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding) plot.
```
s3 = boto3.resource("s3")
key = bt_model.model_data[bt_model.model_data.find("/", 5) + 1 :]
s3.Bucket(output_bucket).download_file(key, "model.tar.gz")
```
Uncompress `model.tar.gz` to get `vectors.txt`
```
!tar -xvzf model.tar.gz
```
If you set "evaluation" as "true" in the hyperparameters, then "eval.json" will be there in the model artifacts.
The quality of trained model is evaluated on word similarity task. We use [WS-353](http://alfonseca.org/eng/research/wordsim353.html), which is one of the most popular test datasets used for this purpose. It contains word pairs together with human-assigned similarity judgments.
The word representations are evaluated by ranking the pairs according to their cosine similarities, and measuring the Spearmans rank correlation coefficient with the human judgments.
Let's look at the evaluation scores which are there in eval.json. For embeddings trained on the text8 dataset, scores above 0.65 are pretty good.
```
!cat eval.json
```
Now, let us do a 2D visualization of the word vectors
```
import numpy as np
from sklearn.preprocessing import normalize
# Read the 400 most frequent word vectors. The vectors in the file are in descending order of frequency.
num_points = 400
first_line = True
index_to_word = []
with open("vectors.txt", "r") as f:
for line_num, line in enumerate(f):
if first_line:
dim = int(line.strip().split()[1])
word_vecs = np.zeros((num_points, dim), dtype=float)
first_line = False
continue
line = line.strip()
word = line.split()[0]
vec = word_vecs[line_num - 1]
for index, vec_val in enumerate(line.split()[1:]):
vec[index] = float(vec_val)
index_to_word.append(word)
if line_num >= num_points:
break
word_vecs = normalize(word_vecs, copy=False, return_norm=False)
from sklearn.manifold import TSNE
tsne = TSNE(perplexity=40, n_components=2, init="pca", n_iter=10000)
two_d_embeddings = tsne.fit_transform(word_vecs[:num_points])
labels = index_to_word[:num_points]
from matplotlib import pylab
%matplotlib inline
def plot(embeddings, labels):
pylab.figure(figsize=(20, 20))
for i, label in enumerate(labels):
x, y = embeddings[i, :]
pylab.scatter(x, y)
pylab.annotate(
label, xy=(x, y), xytext=(5, 2), textcoords="offset points", ha="right", va="bottom"
)
pylab.show()
plot(two_d_embeddings, labels)
```
Running the code above might generate a plot like the one below. t-SNE and Word2Vec are stochastic, so although when you run the code the plot won’t look exactly like this, you can still see clusters of similar words such as below where 'british', 'american', 'french', 'english' are near the bottom-left, and 'military', 'army' and 'forces' are all together near the bottom.

### Stop / Close the Endpoint (Optional)
Finally, we should delete the endpoint before we close the notebook.
```
sess.delete_endpoint(bt_endpoint.endpoint)
```
| github_jupyter |
```
!pip install gspread oauth2client
```
# Sales Data (resource - CSVs)
## Extracting data
```
import pandas as pd
import gspread
from oauth2client.service_account import ServiceAccountCredentials
import os
from datetime import datetime
file_location = 'Data Resources/ZHVI/'
files = os.listdir(file_location)
files
file_list = list()
for data_file in files:
file_dict={}
file_dict['file'] = f"{file_location}{data_file}"
file_dict['Bedroom Count'] = int((data_file.split('Zip_zhvi_bdrmcnt_')[1]).split('.')[0])
file_list.append(file_dict)
file_list
```
## Transforming data
```
# from IPython.core.interactiveshell import InteractiveShell
# InteractiveShell.ast_node_interactivity = "all"
zhvi_complete_df = pd.DataFrame()
number_of_record = 0
for data_file in file_list:
print()
zhvi_df = pd.read_csv(data_file['file'])
print(f"File {data_file['file']} : {len(zhvi_df)} numbers of records")
number_of_record += int(zhvi_df['RegionID'].count())
zhvi_df.rename(columns={"RegionName":"Zip Code"}, inplace=True)
zhvi_df.drop(columns=["RegionID","SizeRank","RegionType","StateName"], inplace=True)
columns_to_drop = []
columns_to_rename = {}
for column in zhvi_df.columns[5:]:
# print(column)
if int(column.split('-')[0]) < 2015:
# print(column)
columns_to_drop.append(column)
else:
columns_to_rename[column] = datetime.strftime(datetime.strptime(column, "%Y-%m-%d"),'%m/%d/%Y')
# print(columns_to_rename)
# columns_to_drop
zhvi_df.drop(columns=columns_to_drop, inplace=True)
zhvi_df.rename(columns=columns_to_rename, inplace=True)
column_list=['Zip Code','City','CountyName','Metro','State']
column_list_data = zhvi_df[column_list]
zhvi_df.drop(columns=column_list, inplace = True)
zhvi_df = zhvi_df.fillna(0)
for column in column_list:
zhvi_df.insert(column_list.index(column), column, column_list_data[column])
zhvi_df.insert(0,'Bedroom Count',int(data_file['Bedroom Count']))
zhvi_complete_df = zhvi_complete_df.append(zhvi_df,ignore_index=True,sort=False)
zhvi_complete_df
zhvi_complete_df.set_index(['Bedroom Count','Zip Code'], inplace=True)
zhvi_complete_df
```
## Extracting States from csv
```
file_path="Data Resources/States.csv"
states_df = pd.read_csv(file_path)
states_df.set_index('state', inplace=True)
states_df
```
# Inventory Data (resource - Google sheets)
https://drive.google.com/drive/folders/1SCwfsJ8WD_295HeEOx8iBrM8mtwEzM7y
## Extracting data
```
scope=["https://spreadsheets.google.com/feeds",
"https://www.googleapis.com/auth/spreadsheets",
"https://www.googleapis.com/auth/drive.file",
"https://www.googleapis.com/auth/drive"]
creds = ServiceAccountCredentials.from_json_keyfile_name("credentials.json",scope)
client = gspread.authorize(creds)
inventory_sales_sheets = client.open("US_Sale_Inventory_Monthly")
print(inventory_sales_sheets.worksheets())
inventory_pending_sheets = client.open("US_Pending_Inventory_Monthly")
print(inventory_pending_sheets.worksheets())
zori_sheets = client.open("ZORI_AllHomesPlusMultifamily_ZIP")
print(zori_sheets.worksheets())
inventory_sales_ws = inventory_sales_sheets.worksheet("All Homes").get_all_records()
inventory_pending_ws = inventory_pending_sheets.worksheet("All Homes").get_all_records()
zori_ws = zori_sheets.worksheet("All Homes").get_all_records()
# print(ws)
inventory_sales_df = pd.DataFrame(inventory_sales_ws)
inventory_pending_df = pd.DataFrame(inventory_pending_ws)
zori_df = pd.DataFrame(zori_ws)
print("Extracting data successfully from google sheets.")
# data = sheet.get_all_records()
```
## Transforming data
```
inventory_sales_df.drop(columns=["RegionID","SizeRank","RegionType"], inplace=True)
columns_to_drop = []
inventory_sales_df = inventory_sales_df.iloc[1:]
for column in inventory_sales_df.columns[2:]:
if int(column.split('/')[2]) < 2018:
# print(column)
columns_to_drop.append(column)
inventory_sales_df.drop(columns=columns_to_drop, inplace=True)
inventory_sales_df.rename(columns={"StateName" : "State"}, inplace=True)
inventory_sales_df['RegionName'] = inventory_sales_df['RegionName'].str.split(',').str[0]
inventory_sales_df.set_index(['RegionName','State'], inplace=True)
inventory_sales_df
inventory_pending_df.drop(columns=["RegionID","SizeRank","RegionType"], inplace=True)
inventory_pending_df = inventory_pending_df.iloc[1:]
inventory_pending_df.rename(columns={"StateName" : "State"}, inplace=True)
inventory_pending_df['RegionName'] = inventory_pending_df['RegionName'].str.split(',').str[0]
inventory_pending_df.set_index(['RegionName','State'], inplace=True)
inventory_pending_df
from calendar import monthrange
zori_df.drop(columns=["RegionID","SizeRank"], inplace=True)
# print(zori_df)
zori_df = zori_df.iloc[1:]
columns_to_drop = []
columns_to_rename = {}
new_date_column=""
for column in zori_df.columns[2:]:
if int(column.split('-')[0]) < 2018:
# print(column)
columns_to_drop.append(column)
else:
new_date_column = f"{int(column.split('-')[1])}/{monthrange(int(column.split('-')[0]),int(column.split('-')[1]))[1]}/{int(column.split('-')[0])}"
columns_to_rename[column] = new_date_column
# print(columns_to_drop)
# print(columns_to_rename)
zori_df.drop(columns = columns_to_drop, inplace=True)
columns_to_rename['RegionName'] = "Zip Code"
zori_df.rename(columns = columns_to_rename , inplace=True)
zori_df.insert(2,'State',zori_df['MsaName'].str.split(',').str[1])
zori_df['MsaName'] = zori_df['MsaName'].str.split(',').str[0]
zori_df.set_index(['Zip Code'], inplace=True)
zori_df
```
# School Data (resource - www.greatschools.org)
## Extracting data
```
from splinter import Browser
from webdriver_manager.chrome import ChromeDriverManager
from bs4 import BeautifulSoup as bs
import requests
great_school_url = "https://www.greatschools.org/california/san-jose/schools/?gradeLevels%5B%5D=e&gradeLevels%5B%5D=m&gradeLevels%5B%5D=h&st%5B%5D=public_charter&st%5B%5D=public&st%5B%5D=charter&view=table"
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
browser.visit(great_school_url)
html = browser.html
soup = bs(html, 'html.parser')
print("Data Extraction started...")
print('-'*30)
try:
next_button = soup.findAll('a',class_='anchor-button')[-1]
school_df = pd.DataFrame()
print("Getting data from page 1.")
while ('disabled' not in next_button.attrs['class']):
next_button = soup.findAll('a',class_='anchor-button')[-1]
school_section = soup.select("section.school-table")
school_list = soup.find("tbody")
for row in school_list:
col = row.findAll('td')
school_row = {}
if(col[0].select_one("a.name")):
school_row['school_name'] = col[0].select_one("a.name").text
rating = col[0].select_one("div div.circle-rating--small")
if(rating):
school_row['rating'] = rating.text
address = col[0].select_one("div.address")
if(address):
school_row['zip_code'] = (address.text.split(',')[-1]).strip()
school_row['type'] = col[1].text
school_row['grades'] = col[2].text
school_row['total_students_enrolled'] = col[3].text
school_row['students_per_teacher'] = col[4].text
school_row['district'] = col[6].text
school_df = school_df.append(school_row, ignore_index=True)
if ('disabled' not in next_button.attrs['class']):
browser.visit('https://www.greatschools.org'+ next_button['href'])
html = browser.html
soup = bs(html, 'html.parser')
print(f"Getting data from page {next_button['href'].split('&page=')[1]}.")
else:
break;
except:
print("Something went wrong")
browser.quit()
print('-'*30)
print("Extraction completed...")
school_df
```
## Transforming data
```
# For currently unrated schools and N/A areas
school_df.fillna(0)
# school_df.loc[['Escuela Popular/Center For Training And Careers, Family Learning']]
school_df.set_index(['school_name','zip_code'], inplace=True)
school_df
```
# Loading Data to PostgreSQL
```
from sqlalchemy import create_engine
import pandas as pd
from db_conn import user_name
from db_conn import password
import psycopg2
conn = psycopg2.connect(
database="postgres", user=f'{user_name}', password=f'{password}', host='127.0.0.1', port= '5432'
)
conn.autocommit = True
cursor = conn.cursor()
cursor.execute("SELECT datname FROM pg_database;")
list_database = cursor.fetchall()
dbname = "zillow_db"
try:
# if (dbname,) in list_database:
# #Preparing query to delete a database
# cursor.execute(f'''DROP DATABASE {dbname}''')
# cursor.close()
# # conn.close()
# print("Database deleted successfully...")
# print('-'*30)
if (dbname,) not in list_database:
cur = conn.cursor()
cur.execute('CREATE DATABASE ' + dbname)
cur.close()
conn.close()
print("Creating Database...")
engine = create_engine(f'postgresql://{user_name}:{password}@localhost:5432/{dbname}')
connection = engine.connect()
print('-'*30)
print("Creating Tables, Please wait...")
print('-'*30)
zhvi_complete_df.to_sql('sales',engine)
print("Table sales created successfully")
states_df.to_sql('states', engine)
print("Table states created successfully")
sales_inventory_df.to_sql('inventory_sales', engine)
print("Table inventory_sales created successfully")
inventory_pending_df.to_sql('inventory_pending', engine)
print("Table inventory_pending created successfully")
zori_df.to_sql('rentals', engine)
print("Table rentals created successfully")
school_df.to_sql('schools', engine)
print("Table schools created successfully")
connection.close()
print('-'*30)
print("Database is ready to use.")
else:
print("Database is already exists.")
except:
print("Something went wrong.")
```
| github_jupyter |
```
import keras
keras.__version__
```
# Using a pre-trained convnet
This notebook contains the code sample found in Chapter 5, Section 3 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
----
A common and highly effective approach to deep learning on small image datasets is to leverage a pre-trained network. A pre-trained network
is simply a saved network previously trained on a large dataset, typically on a large-scale image classification task. If this original
dataset is large enough and general enough, then the spatial feature hierarchy learned by the pre-trained network can effectively act as a
generic model of our visual world, and hence its features can prove useful for many different computer vision problems, even though these
new problems might involve completely different classes from those of the original task. For instance, one might train a network on
ImageNet (where classes are mostly animals and everyday objects) and then re-purpose this trained network for something as remote as
identifying furniture items in images. Such portability of learned features across different problems is a key advantage of deep learning
compared to many older shallow learning approaches, and it makes deep learning very effective for small-data problems.
In our case, we will consider a large convnet trained on the ImageNet dataset (1.4 million labeled images and 1000 different classes).
ImageNet contains many animal classes, including different species of cats and dogs, and we can thus expect to perform very well on our cat
vs. dog classification problem.
We will use the VGG16 architecture, developed by Karen Simonyan and Andrew Zisserman in 2014, a simple and widely used convnet architecture
for ImageNet. Although it is a bit of an older model, far from the current state of the art and somewhat heavier than many other recent
models, we chose it because its architecture is similar to what you are already familiar with, and easy to understand without introducing
any new concepts. This may be your first encounter with one of these cutesie model names -- VGG, ResNet, Inception, Inception-ResNet,
Xception... you will get used to them, as they will come up frequently if you keep doing deep learning for computer vision.
There are two ways to leverage a pre-trained network: *feature extraction* and *fine-tuning*. We will cover both of them. Let's start with
feature extraction.
## Feature extraction
Feature extraction consists of using the representations learned by a previous network to extract interesting features from new samples.
These features are then run through a new classifier, which is trained from scratch.
As we saw previously, convnets used for image classification comprise two parts: they start with a series of pooling and convolution
layers, and they end with a densely-connected classifier. The first part is called the "convolutional base" of the model. In the case of
convnets, "feature extraction" will simply consist of taking the convolutional base of a previously-trained network, running the new data
through it, and training a new classifier on top of the output.
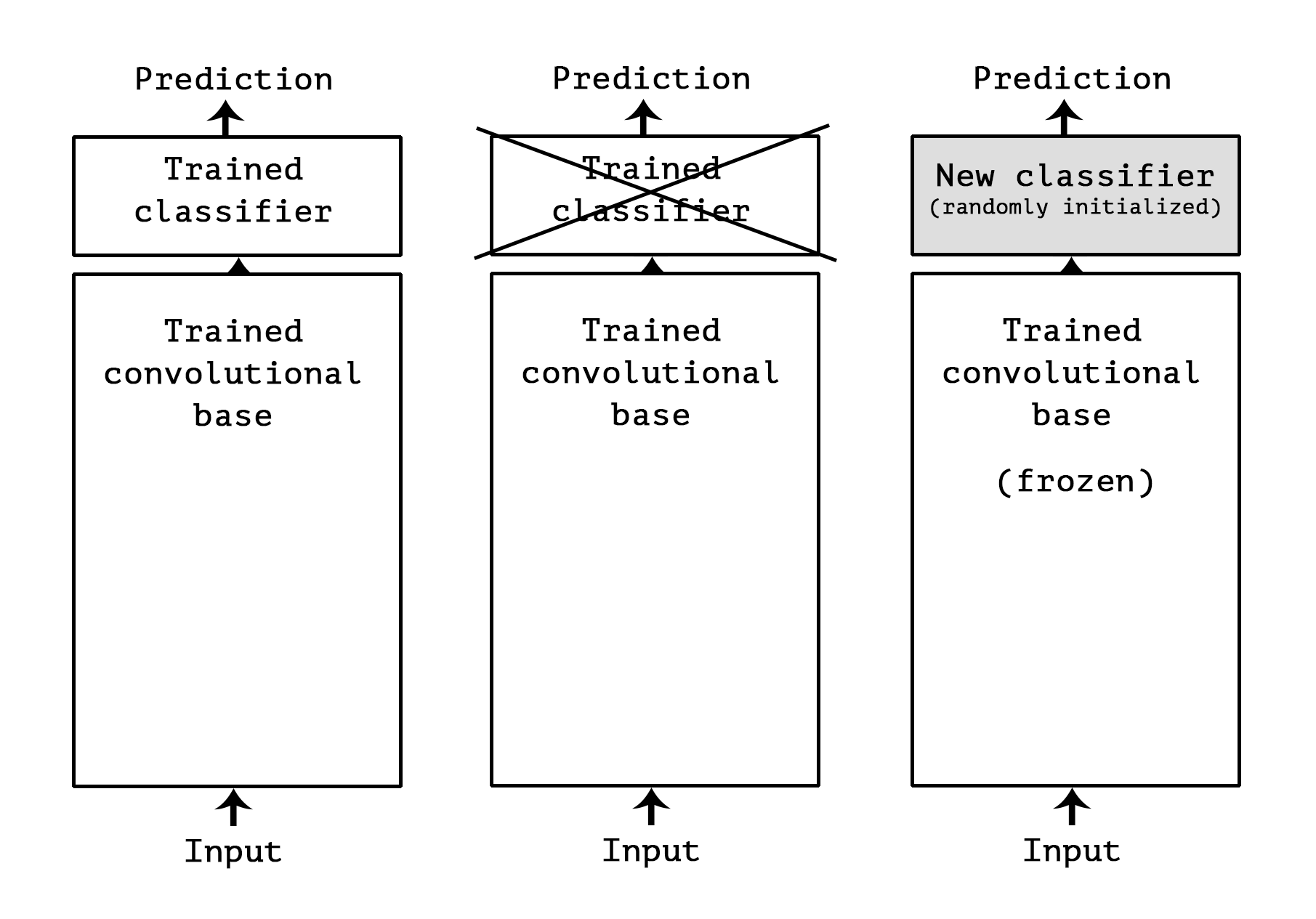
Why only reuse the convolutional base? Could we reuse the densely-connected classifier as well? In general, it should be avoided. The
reason is simply that the representations learned by the convolutional base are likely to be more generic and therefore more reusable: the
feature maps of a convnet are presence maps of generic concepts over a picture, which is likely to be useful regardless of the computer
vision problem at hand. On the other end, the representations learned by the classifier will necessarily be very specific to the set of
classes that the model was trained on -- they will only contain information about the presence probability of this or that class in the
entire picture. Additionally, representations found in densely-connected layers no longer contain any information about _where_ objects are
located in the input image: these layers get rid of the notion of space, whereas the object location is still described by convolutional
feature maps. For problems where object location matters, densely-connected features would be largely useless.
Note that the level of generality (and therefore reusability) of the representations extracted by specific convolution layers depends on
the depth of the layer in the model. Layers that come earlier in the model extract local, highly generic feature maps (such as visual
edges, colors, and textures), while layers higher-up extract more abstract concepts (such as "cat ear" or "dog eye"). So if your new
dataset differs a lot from the dataset that the original model was trained on, you may be better off using only the first few layers of the
model to do feature extraction, rather than using the entire convolutional base.
In our case, since the ImageNet class set did contain multiple dog and cat classes, it is likely that it would be beneficial to reuse the
information contained in the densely-connected layers of the original model. However, we will chose not to, in order to cover the more
general case where the class set of the new problem does not overlap with the class set of the original model.
Let's put this in practice by using the convolutional base of the VGG16 network, trained on ImageNet, to extract interesting features from
our cat and dog images, and then training a cat vs. dog classifier on top of these features.
The VGG16 model, among others, comes pre-packaged with Keras. You can import it from the `keras.applications` module. Here's the list of
image classification models (all pre-trained on the ImageNet dataset) that are available as part of `keras.applications`:
* Xception
* InceptionV3
* ResNet50
* VGG16
* VGG19
* MobileNet
Let's instantiate the VGG16 model:
```
from keras.applications import VGG16
conv_base = VGG16(weights='imagenet',
include_top=False, # 不要分类层
input_shape=(150, 150, 3))
```
We passed three arguments to the constructor:
* `weights`, to specify which weight checkpoint to initialize the model from
* `include_top`, which refers to including or not the densely-connected classifier on top of the network. By default, this
densely-connected classifier would correspond to the 1000 classes from ImageNet. Since we intend to use our own densely-connected
classifier (with only two classes, cat and dog), we don't need to include it.
* `input_shape`, the shape of the image tensors that we will feed to the network. This argument is purely optional: if we don't pass it,
then the network will be able to process inputs of any size.
Here's the detail of the architecture of the VGG16 convolutional base: it's very similar to the simple convnets that you are already
familiar with.
```
conv_base.summary()
```
The final feature map has shape `(4, 4, 512)`. That's the feature on top of which we will stick a densely-connected classifier.
At this point, there are two ways we could proceed:
* Running the convolutional base over our dataset, recording its output to a Numpy array on disk, then using this data as input to a
standalone densely-connected classifier similar to those you have seen in the first chapters of this book. This solution is very fast and
cheap to run, because it only requires running the convolutional base once for every input image, and the convolutional base is by far the
most expensive part of the pipeline. However, for the exact same reason, this technique would not allow us to leverage data augmentation at
all.
* Extending the model we have (`conv_base`) by adding `Dense` layers on top, and running the whole thing end-to-end on the input data. This
allows us to use data augmentation, because every input image is going through the convolutional base every time it is seen by the model.
However, for this same reason, this technique is far more expensive than the first one.
We will cover both techniques. Let's walk through the code required to set-up the first one: recording the output of `conv_base` on our
data and using these outputs as inputs to a new model.
We will start by simply running instances of the previously-introduced `ImageDataGenerator` to extract images as Numpy arrays as well as
their labels. We will extract features from these images simply by calling the `predict` method of the `conv_base` model.
```
import os
import numpy as np
from keras.preprocessing.image import ImageDataGenerator
base_dir = '../data/cats_and_dogs_small'
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
test_dir = os.path.join(base_dir, 'test')
datagen = ImageDataGenerator(rescale=1./255)
batch_size = 20
# def extract_features(directory, sample_count):
# features = np.zeros(shape=(sample_count, 4, 4, 512))
# labels = np.zeros(shape=(sample_count))
# generator = datagen.flow_from_directory(directory,
# target_size=(150, 150),
# batch_size=batch_size,
# class_mode='binary')
# i = 0
# for inputs_batch, labels_batch in generator:
# features_batch = conv_base.predict(inputs_batch)
# features[i * batch_size : (i + 1) * batch_size] = features_batch
# labels[i * batch_size : (i + 1) * batch_size] = labels_batch
# i += 1
# if i * batch_size >= sample_count:
# # Note that since generators yield data indefinitely in a loop,
# # we must `break` after every image has been seen once.
# break
# return features, labels
# train_features, train_labels = extract_features(train_dir, 2000)
# validation_features, validation_labels = extract_features(validation_dir, 1000)
# test_features, test_labels = extract_features(test_dir, 1000)
```
The extracted features are currently of shape `(samples, 4, 4, 512)`. We will feed them to a densely-connected classifier, so first we must
flatten them to `(samples, 8192)`:
```
# train_features = np.reshape(train_features, (2000, 4 * 4 * 512))
# validation_features = np.reshape(validation_features, (1000, 4 * 4 * 512))
# test_features = np.reshape(test_features, (1000, 4 * 4 * 512))
```
At this point, we can define our densely-connected classifier (note the use of dropout for regularization), and train it on the data and
labels that we just recorded:
```
# from keras import models
# from keras import layers
# from keras import optimizers
# model = models.Sequential()
# model.add(layers.Dense(256, activation='relu', input_dim=4 * 4 * 512))
# model.add(layers.Dropout(0.5))
# model.add(layers.Dense(1, activation='sigmoid'))
# model.compile(optimizer=optimizers.RMSprop(lr=2e-5),
# loss='binary_crossentropy',
# metrics=['acc'])
# history = model.fit(train_features, train_labels,
# epochs=30,
# batch_size=20,
# validation_data=(validation_features, validation_labels))
```
Training is very fast, since we only have to deal with two `Dense` layers -- an epoch takes less than one second even on CPU.
Let's take a look at the loss and accuracy curves during training:
```
# import matplotlib.pyplot as plt
# %matplotlib inline
# acc = history.history['acc']
# val_acc = history.history['val_acc']
# loss = history.history['loss']
# val_loss = history.history['val_loss']
# epochs = range(len(acc))
# plt.plot(epochs, acc, 'bo', label='Training acc')
# plt.plot(epochs, val_acc, 'b', label='Validation acc')
# plt.title('Training and validation accuracy')
# plt.legend()
# plt.figure()
# plt.plot(epochs, loss, 'bo', label='Training loss')
# plt.plot(epochs, val_loss, 'b', label='Validation loss')
# plt.title('Training and validation loss')
# plt.legend()
# plt.show()
```
We reach a validation accuracy of about 90%, much better than what we could achieve in the previous section with our small model trained from
scratch. However, our plots also indicate that we are overfitting almost from the start -- despite using dropout with a fairly large rate.
This is because this technique does not leverage data augmentation, which is essential to preventing overfitting with small image datasets.
Now, let's review the second technique we mentioned for doing feature extraction, which is much slower and more expensive, but which allows
us to leverage data augmentation during training: extending the `conv_base` model and running it end-to-end on the inputs. Note that this
technique is in fact so expensive that you should only attempt it if you have access to a GPU: it is absolutely intractable on CPU. If you
cannot run your code on GPU, then the previous technique is the way to go.
Because models behave just like layers, you can add a model (like our `conv_base`) to a `Sequential` model just like you would add a layer.
So you can do the following:
```
from keras import models
from keras import layers
model = models.Sequential()
model.add(conv_base)
model.add(layers.Flatten())
model.add(layers.Dense(256, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
```
This is what our model looks like now:
```
model.summary()
```
As you can see, the convolutional base of VGG16 has 14,714,688 parameters, which is very large. The classifier we are adding on top has 2
million parameters.
Before we compile and train our model, a very important thing to do is to freeze the convolutional base. "Freezing" a layer or set of
layers means preventing their weights from getting updated during training. If we don't do this, then the representations that were
previously learned by the convolutional base would get modified during training. Since the `Dense` layers on top are randomly initialized,
very large weight updates would be propagated through the network, effectively destroying the representations previously learned.
In Keras, freezing a network is done by setting its `trainable` attribute to `False`:
```
print('This is the number of trainable weights '
'before freezing the conv base:', len(model.trainable_weights))
conv_base.trainable = False
print('This is the number of trainable weights '
'after freezing the conv base:', len(model.trainable_weights))
```
With this setup, only the weights from the two `Dense` layers that we added will be trained. That's a total of four weight tensors: two per
layer (the main weight matrix and the bias vector). Note that in order for these changes to take effect, we must first compile the model.
If you ever modify weight trainability after compilation, you should then re-compile the model, or these changes would be ignored.
Now we can start training our model, with the same data augmentation configuration that we used in our previous example:
```
from keras.preprocessing.image import ImageDataGenerator
from keras import optimizers
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
# This is the target directory
train_dir,
# All images will be resized to 150x150
target_size=(150, 150),
batch_size=40,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(150, 150),
batch_size=40,
class_mode='binary')
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=2e-5),
metrics=['acc'])
history = model.fit_generator(
train_generator,
steps_per_epoch=100,
epochs=30,
validation_data=validation_generator,
validation_steps=50,
verbose=1)
model.save('./models/cats_and_dogs_small_3.h5')
```
Let's plot our results again:
```
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
val_loss_min = val_loss.index(min(val_loss))
val_acc_max = val_acc.index(max(val_acc))
print('validation set min loss: ', val_loss_min)
print('validation set max accuracy: ', val_acc_max)
```
As you can see, we reach a validation accuracy of about 96%. This is much better than our small convnet trained from scratch.
## Fine-tuning
Another widely used technique for model reuse, complementary to feature extraction, is _fine-tuning_.
Fine-tuning consists in unfreezing a few of the top layers
of a frozen model base used for feature extraction, and jointly training both the newly added part of the model (in our case, the
fully-connected classifier) and these top layers. This is called "fine-tuning" because it slightly adjusts the more abstract
representations of the model being reused, in order to make them more relevant for the problem at hand.
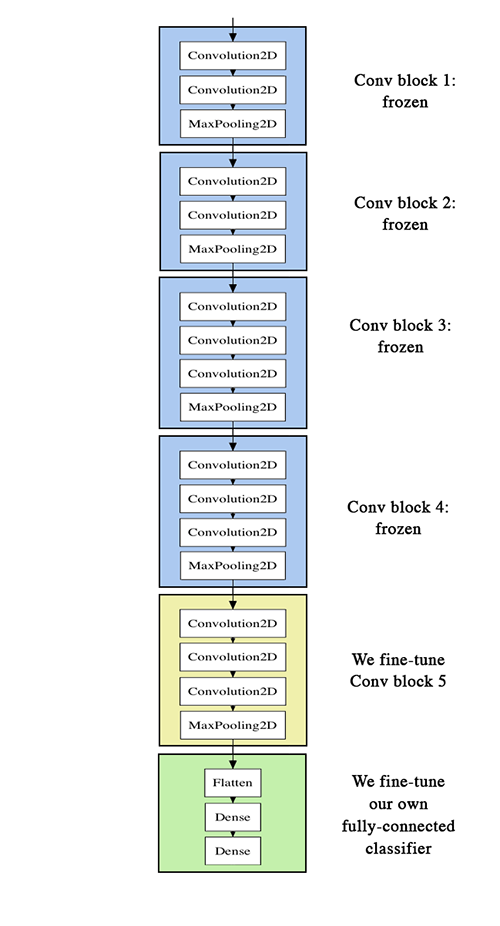
We have stated before that it was necessary to freeze the convolution base of VGG16 in order to be able to train a randomly initialized
classifier on top. For the same reason, it is only possible to fine-tune the top layers of the convolutional base once the classifier on
top has already been trained. If the classified wasn't already trained, then the error signal propagating through the network during
training would be too large, and the representations previously learned by the layers being fine-tuned would be destroyed. Thus the steps
for fine-tuning a network are as follow:
* 1) Add your custom network on top of an already trained base network.
* 2) Freeze the base network.
* 3) Train the part you added.
* 4) Unfreeze some layers in the base network.
* 5) Jointly train both these layers and the part you added.
We have already completed the first 3 steps when doing feature extraction. Let's proceed with the 4th step: we will unfreeze our `conv_base`,
and then freeze individual layers inside of it.
As a reminder, this is what our convolutional base looks like:
```
conv_base.summary()
```
We will fine-tune the last 3 convolutional layers, which means that all layers up until `block4_pool` should be frozen, and the layers
`block5_conv1`, `block5_conv2` and `block5_conv3` should be trainable.
Why not fine-tune more layers? Why not fine-tune the entire convolutional base? We could. However, we need to consider that:
* Earlier layers in the convolutional base encode more generic, reusable features, while layers higher up encode more specialized features. It is
more useful to fine-tune the more specialized features, as these are the ones that need to be repurposed on our new problem. There would
be fast-decreasing returns in fine-tuning lower layers.
* The more parameters we are training, the more we are at risk of overfitting. The convolutional base has 15M parameters, so it would be
risky to attempt to train it on our small dataset.
Thus, in our situation, it is a good strategy to only fine-tune the top 2 to 3 layers in the convolutional base.
Let's set this up, starting from where we left off in the previous example:
```
conv_base.trainable = True
set_trainable = False
for layer in conv_base.layers:
if layer.name == 'block5_conv1':
set_trainable = True
if set_trainable:
layer.trainable = True
else:
layer.trainable = False
```
Now we can start fine-tuning our network. We will do this with the RMSprop optimizer, using a very low learning rate. The reason for using
a low learning rate is that we want to limit the magnitude of the modifications we make to the representations of the 3 layers that we are
fine-tuning. Updates that are too large may harm these representations.
Now let's proceed with fine-tuning:
```
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-5),
metrics=['acc'])
history = model.fit_generator(train_generator,
steps_per_epoch=100,
epochs=100,
validation_data=validation_generator,
validation_steps=50,
verbose=0)
model.save('./models/cats_and_dogs_small_4.h5')
```
Let's plot our results using the same plotting code as before:
```
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
```
These curves look very noisy. To make them more readable, we can smooth them by replacing every loss and accuracy with exponential moving
averages of these quantities. Here's a trivial utility function to do this:
```
def smooth_curve(points, factor=0.8):
smoothed_points = []
for point in points:
if smoothed_points:
previous = smoothed_points[-1]
smoothed_points.append(previous * factor + point * (1 - factor))
else:
smoothed_points.append(point)
return smoothed_points
plt.plot(epochs,
smooth_curve(acc), 'bo', label='Smoothed training acc')
plt.plot(epochs,
smooth_curve(val_acc), 'b', label='Smoothed validation acc')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs,
smooth_curve(loss), 'bo', label='Smoothed training loss')
plt.plot(epochs,
smooth_curve(val_loss), 'b', label='Smoothed validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
smooth_val_loss = smooth_curve(val_loss)
smooth_val_loss.index(min(smooth_val_loss))
```
These curves look much cleaner and more stable. We are seeing a nice 1% absolute improvement.
Note that the loss curve does not show any real improvement (in fact, it is deteriorating). You may wonder, how could accuracy improve if the
loss isn't decreasing? The answer is simple: what we display is an average of pointwise loss values, but what actually matters for accuracy
is the distribution of the loss values, not their average, since accuracy is the result of a binary thresholding of the class probability
predicted by the model. The model may still be improving even if this isn't reflected in the average loss.
We can now finally evaluate this model on the test data:
```
test_generator = test_datagen.flow_from_directory(test_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
test_loss, test_acc = model.evaluate_generator(test_generator, steps=50)
print('test acc:', test_acc)
```
Here we get a test accuracy of 97%. In the original Kaggle competition around this dataset, this would have been one of the top results.
However, using modern deep learning techniques, we managed to reach this result using only a very small fraction of the training data
available (about 10%). There is a huge difference between being able to train on 20,000 samples compared to 2,000 samples!
```
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-5),
metrics=['acc'])
history = model.fit_generator(train_generator,
steps_per_epoch=100,
epochs=23,
validation_data=validation_generator,
validation_steps=50,
verbose=2)
model.save('./models/cats_and_dogs_small_5.h5')
test_generator = test_datagen.flow_from_directory(test_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
test_loss, test_acc = model.evaluate_generator(test_generator, steps=50)
print('test acc:', test_acc)
```
## Take-aways: using convnets with small datasets
Here's what you should take away from the exercises of these past two sections:
* Convnets are the best type of machine learning models for computer vision tasks. It is possible to train one from scratch even on a very
small dataset, with decent results.
* On a small dataset, overfitting will be the main issue. Data augmentation is a powerful way to fight overfitting when working with image
data.
* It is easy to reuse an existing convnet on a new dataset, via feature extraction. This is a very valuable technique for working with
small image datasets.
* As a complement to feature extraction, one may use fine-tuning, which adapts to a new problem some of the representations previously
learned by an existing model. This pushes performance a bit further.
Now you have a solid set of tools for dealing with image classification problems, in particular with small datasets.
| github_jupyter |
# VacationPy
----
#### Note
* Keep an eye on your API usage. Use https://developers.google.com/maps/reporting/gmp-reporting as reference for how to monitor your usage and billing.
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from config import g_key
```
### Store Part I results into DataFrame
* Load the csv exported in Part I to a DataFrame
```
file = os.path.join("..","WeatherPy","cities.csv")
weather_df = pd.read_csv(file, encoding= 'utf-8')
weather_df.head(50)
```
### Humidity Heatmap
* Configure gmaps.
* Use the Lat and Lng as locations and Humidity as the weight.
* Add Heatmap layer to map.
```
# Access maps with unique API key
gmaps.configure(api_key=g_key)
lat_lng = pd.DataFrame(weather_df.iloc[:, 5:7])
lat_lng
humidity = weather_df['Humidity']
humidity
# create the mapping figure
fig= gmaps.figure()
# create the heatmap layer
heat_layer= gmaps.heatmap_layer(lat_lng, weights=humidity)
# add the heatmap layer to the figure
fig.add_layer(heat_layer)
# display the figure
fig
```
### Create new DataFrame fitting weather criteria
* Narrow down the cities to fit weather conditions.
* Drop any rows will null values.
```
#Criteria: A max temperature lower than 80 degrees but higher than 70.
nice_weather = weather_df[(weather_df['Max Temp(F)'] < 80) & (weather_df['Max Temp(F)'] > 70) & (weather_df['Humidity'] <30)]
nice_weather.info()
```
### Hotel Map
* Store into variable named `hotel_df`.
* Add a "Hotel Name" column to the DataFrame.
* Set parameters to search for hotels with 5000 meters.
* Hit the Google Places API for each city's coordinates.
* Store the first Hotel result into the DataFrame.
* Plot markers on top of the heatmap.
```
hotel_df = nice_weather
hotel_df["Hotel Name"]= ""
hotel_df
# params dictionary to update each iteration
params = {
"radius": 5000,
"types": "lodging",
"keyword": "hotel",
"key": g_key
}
# Use the lat/lng we recovered to identify airports
for index, row in hotel_df.iterrows():
# get lat, lng from df
lat = row["Lat"]
lng = row["Long"]
print(f"Searching within {row['City']}, {row['Country']}...")
# change location each iteration while leaving original params in place
params["location"] = f"{lat},{lng}"
# Use the search term: "Hotel" and our lat/lng
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
# make request and print url
name_address = requests.get(base_url, params=params)
# convert to json
name_address = name_address.json()
# Since some data may be missing we incorporate a try-except to skip any that are missing a data point.
try:
hotel_df.loc[index, "Hotel Name"] = name_address["results"][0]["name"]
except (KeyError, IndexError):
print("Missing field/result... skipping.")
hotel_df
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Lat", "Long"]]
# Add marker layer ontop of heat map
# Create a marker layer using our coordinates
markers = gmaps.marker_layer(locations, info_box_content = hotel_info, display_info_box=True)
# Add the layer to the map
fig.add_layer(markers)
# display the figure with the newly added layer
fig
```
| github_jupyter |
# SVM CLASSIFICATION WITH INDIVIDUAL REPLICAS AND ALL GENES
Training data
1. Uses individual replicas (not averaged)
1. Uses all genes
1. Includes time T1 (normoxia is not combined with resuscitation)
Issues
1. Poor feature selection yields poor classification accuracies; close to random
1. Related to (1), need to choose features by class.
# Preliminaries
## Imports
```
import init
from common import constants as cn
from common.trinary_data import TrinaryData
from common.data_provider import DataProvider
from common import transform_data
from common_python.plots import util_plots
from common_python.classifier import classifier_ensemble
from common_python.classifier import classifier_collection
from common import transform_data
from common_python.util import util
from common_python.classifier import feature_analyzer
import collections
import copy
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import sklearn
from sklearn.decomposition import PCA
from sklearn import svm
from sklearn.model_selection import cross_val_score
%matplotlib inline
```
## Constants
```
REPO_PATH = util.findRepositoryRoot("xstate")
NOTEBOOK_PATH = os.path.join(REPO_PATH, "notebooks")
DATA_PATH = os.path.join(REPO_PATH, "data")
SORT = "sort"
VALUE = "value"
max_var_replication = 1.5
SVM_ENSEMBLE = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(),
filter_high_rank=50,
size=100, holdouts=1)
IS_RERUN = True
# "Short" data, with averaged replications
TRINARY_SHORT = TrinaryData(is_averaged=True, is_dropT1=True, is_regulator=False) # Trinary data
DF_X_SHORT = TRINARY_SHORT.df_X
DF_X_SHORT = transform_data.removeGenesWithExcessiveReplicationVariance(DF_X_SHORT,
max_var=max_var_replication)
SER_Y_SHORT = TRINARY_SHORT.ser_y
STATES_SHORT = SER_Y_SHORT.unique()
# "Long" data, with replications
TRINARY_LONG = TrinaryData(is_averaged=False, is_dropT1=False, is_regulator=False) # Trinary data
DF_X_LONG = TRINARY_LONG.df_X
DF_X_LONG = transform_data.removeGenesWithExcessiveReplicationVariance(DF_X_LONG,
max_var=max_var_replication)
SER_Y_LONG = TRINARY_LONG.ser_y
STATES_LONG = SER_Y_LONG.unique()
# Feature analyzers
ANALYZER_PATH_LONG = os.path.join(DATA_PATH, "feature_analyzer_with_replicas")
ANALYZER_PATH_PAT_LONG = os.path.join(ANALYZER_PATH_LONG, "%d")
ANALYZER_LONG_DCT = feature_analyzer.deserialize({s: ANALYZER_PATH_PAT_LONG % s for s in STATES_LONG})
FITTED_SVM_PATH_SHORT = os.path.join(NOTEBOOK_PATH, "svm_classification_averaged.pcl")
try:
if IS_RERUN:
raise(Exception)
FITTED_SVM_ENSEMBLE_SHORT.deserialize(FITTED_SVM_PATH_SHORT)
except:
FITTED_SVM_ENSEMBLE_SHORT = copy.deepcopy(SVM_ENSEMBLE)
FITTED_SVM_ENSEMBLE_SHORT.fit(DF_X_SHORT, SER_Y_SHORT)
FITTED_SVM_ENSEMBLE_SHORT.serialize(FITTED_SVM_PATH_SHORT)
FITTED_SVM_PATH_LONG = os.path.join(NOTEBOOK_PATH, "svm_classification_individual.pcl")
try:
if IS_RERUN:
raise(Exception)
FITTED_SVM_ENSEMBLE_LONG.deserialize(FITTED_SVM_PATH_LONG)
except:
FITTED_SVM_ENSEMBLE_LONG = copy.deepcopy(SVM_ENSEMBLE)
FITTED_SVM_ENSEMBLE_LONG.fit(DF_X_LONG, SER_Y_LONG)
FITTED_SVM_ENSEMBLE_LONG.serialize(FITTED_SVM_PATH_LONG)
```
## Data
Data used in the analysis.
```
df_sampleAM = transform_data.trinaryReadsDF(
csv_file="AM_MDM_Mtb_transcripts_DEseq.csv", is_time_columns=False, is_display_errors=False)
df_sampleAW = transform_data.trinaryReadsDF(csv_file="AW_plus_v_AW_neg_Mtb_transcripts_DEseq.csv",
is_time_columns=False, is_display_errors=False)
df_sampleAM = df_sampleAM.T
df_sampleAW.columns
df_sampleAW = df_sampleAW.T
df_sampleAW.head()
```
## Helpers
```
# Describes how feature importances are calculated for the construction of ensembles
# Importances should be calculated using analyzers. Must get state information.
class ClassifierDescriptorSVMAnalyzer(classifier_ensemble.ClassifierDescriptor):
"""
Descriptor information needed for SVM classifiers
Descriptor is for one-vs-rest. So, there is a separate
classifier for each class.
"""
def __init__(self, analyzer_dct, min_sfa_accuracy=0.8):
"""
:params dict(key: object, value: FeatureAnalyzer):
"""
self.analyzer_dct = analyzer_dct
self.clf = svm.LinearSVC() # Template classifier used to construct ensemble
self.min_sfa_accuracy = min_sfa_accuracy
def getImportance(self, clf, class_selection=None):
"""
Calculates the importances of features.
:param Classifier clf: fitted classifier
:param obj class_selection: key in self.analyzer_dct
:return list-float:
"""
max_features = 100
if class_selection is None:
classes = list(self.analyzer_dct.keys())
else:
classes = class_selection
# Order the features based on: (a) single factor predication accuracy (SFA) and (b) incremental
# prediction accuracy beyond the previously chosen features
value_dct = {}
for key, analyzer in self.analyzer_dct.items():
df_X = analyzer.df_X
value_dct[key] = [analyzer.ser_sfa.loc[f] for f in df_X.columns]
df = pd.DataFrame({VALUE: value_dct[key]})
df[SORT] = range(len(df))
df.index = df_X.columns
# Adjust importance based on other features
df = df.sort_values(VALUE, ascending = False)
indices = list(df.index)
# Importance is its maximum increase in classification accuracy
for pos, idx in enumerate(indices[1:max_features]):
priors = indices[0:pos+1]
sub_priors = set(priors).intersection(analyzer.df_ipa.columns)
if idx in analyzer.df_ipa.index:
try:
values = [analyzer.df_ipa.loc[idx, p] for p in sub_priors]
except Exception:
import pdb; pdb.set_trace()
df.loc[idx, VALUE] = np.max(values)
df = df.sort_values(SORT)
value_dct[key] = df[VALUE].values
# Calculate average values
df = pd.DataFrame(value_dct)
df = df.T
df = df.applymap(lambda v: np.nan if v < self.min_sfa_accuracy else v)
ser = df.mean(skipna=True)
return ser.values
# TESTING
# FIXME: Lost Rv1813c
desc = ClassifierDescriptorSVMAnalyzer({0: ANALYZER_LONG_DCT[1]})
values = desc.getImportance(None)
assert(np.nanmax(values) > 0.9)
```
# Classification Validations of Controlled Samples
Classify T1-T25 and see if result is same as original class. Use 5-fold cross validation, where there is a holdout for each class and the selection is random.
```
IS_VALIDATIONS = True
# Analyze accuracy for class 1
if False:
accuracy_dct = {}
CLASS = 1
data = copy.deepcopy(DATA)
data.ser_y = DATA.ser_y.apply(lambda v: 1 if v == CLASS else 0)
other_analyzers = [ANALYZER_DCT[o] for o in ANALYZER_DCT.keys() if o != CLASS]
merged_analyzer = other_analyzers[0].copy()
[merged_analyzer.merge(a) for a in other_analyzers[1:]]
analyzer_dct = {CLASS: ANALYZER_DCT[CLASS], 0: merged_analyzer}
analyzer_dct = {CLASS: ANALYZER_DCT[CLASS]}
# With merging the other analyzers for the other class
# analyzer_dct[0] = merged_analyzer
if IS_VALIDATIONS:
data = copy.deepcopy(TRINARY_LONG)
data.df_X = DF_X_LONG
data.ser_y = SER_Y_LONG
accuracy_dct = {}
for rank in [1, 2, 16, 64, 128, 512, 1024]:
accuracy_dct[rank] = classifier_ensemble.ClassifierEnsemble.crossValidate(
data,
#clf_desc=ClassifierDescriptorSVMAnalyzer(ANALYZER_DCT),
num_iter=10, num_holdout=1, filter_high_rank=rank)
plt.plot(list(accuracy_dct.keys()), list(accuracy_dct.values()))
plt.ylim([0, 1.1])
_ = plt.xlabel("No. features in classifier")
_ = plt.ylabel("accuracy")
# Without merging
# analyzer_dct[0] = merged_analyzer
if False:
for rank in [1, 2, 4, 8, 16, 32]:
accuracy_dct[rank] = classifier_ensemble.ClassifierEnsemble.crossValidate(
data,
clf_desc=ClassifierDescriptorSVMAnalyzer(analyzer_dct),
num_iter=10, num_holdout=1, filter_high_rank=rank)
plt.plot(list(accuracy_dct.keys()), list(accuracy_dct.values()))
plt.ylim([0, 1.1])
_ = plt.xlabel("No classifiers in ensemble")
_ = plt.ylabel("accuracy")
```
# Investigation of Poor Classifier Performance
```
time = "T11"
_, axes = plt.subplots(1, 3, figsize=(18,8))
for idx, ax in enumerate(axes):
instance = "%s.%d" % (time, idx)
ser_X = DF_X_LONG.loc[instance, :]
if idx == len(axes) - 1:
is_legend = True
else:
is_legend = False
FITTED_SVM_ENSEMBLE_LONG.plotFeatureContributions(ser_X, ax=ax,
title=instance, true_class=SER_Y_LONG.loc[instance], is_plot=False, is_legend=is_legend)
plt.show()
```
## Variability of expression levels between replications
```
# Shade replications
fig, ax = plt.subplots(1, figsize=(20, 20))
columns = list(FITTED_SVM_ENSEMBLE_LONG.columns)
columns.sort()
indices = list(DF_X_LONG.index)
indices = sorted(indices, key=lambda v: float(v[1:]))
df_X = DF_X_LONG[columns]
df_X = df_X.loc[indices, :]
sns.heatmap(df_X.T, cmap="jet", ax=ax)
for idx in range(25):
if idx % 2 == 0:
plt.axvspan(3*idx, 3+3*idx, facecolor='grey', alpha=0.7)
```
Only use features that have low variability within replications.
Let $y_{itr}$ be the expression value for gene $i$, at time $t$, and replication $r$.
Sort genes $i$ by ascending value of $v_i = \sum_t var(y_{itr})$.
```
columns = list(FITTED_SVM_ENSEMBLE_LONG.columns)
df = DF_X_LONG.copy()
#df = df[columns]
df.index = [i[0:-2] for i in DF_X_LONG.index]
df = df.sort_index()
ser = df.groupby(df.index).std().sum()
_ = plt.hist(ser.sort_values(), bins=100)
```
# Classification of Lab Samples
```
svm_ensemble = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(), filter_high_rank=15, size=30)
df_X = DF_X_LONG.copy()
svm_ensemble.fit(DF_X_LONG, SER_Y_LONG)
svm_ensemble.predict(df_sampleAM)
svm_ensemble.predict(df_sampleAW)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/IEwaspbusters/KopuruVespaCompetitionIE/blob/main/Competition_subs/2021-04-28_submit/batch_LARVAE/HEX.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# XGBoost Years: Prediction with Mario's Cluster, Population Comercial Density + SearchGridCV
## Import the Data & Modules
```
# Base packages -----------------------------------
import pandas as pd
import numpy as np
import warnings
# Data Viz -----------------------------------
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams["figure.figsize"] = (15, 10) # to set figure size when ploting feature_importance
import graphviz
# XGBoost -------------------------------
import xgboost as xgb
from xgboost import XGBRegressor
from xgboost import plot_importance # built-in function to plot features ordered by their importance
# SKLearn -----------------------------------------
from sklearn import preprocessing # scaling data
from sklearn.model_selection import GridSearchCV
#Cluster
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from plotnine import *
# Function that checks if final Output is ready for submission or needs revision
def check_data(HEX):
def template_checker(HEX):
submission_df = (HEX["CODIGO MUNICIPIO"].astype("string")+HEX["NOMBRE MUNICIPIO"]).sort_values().reset_index(drop=True)
template_df = (template["CODIGO MUNICIPIO"].astype("string")+template["NOMBRE MUNICIPIO"]).sort_values().reset_index(drop=True)
check_df = pd.DataFrame({"submission_df":submission_df,"template_df":template_df})
check_df["check"] = check_df.submission_df == check_df.template_df
if (check_df.check == False).any():
pd.options.display.max_rows = 112
return check_df.loc[check_df.check == False,:]
else:
return "All Municipality Names and Codes to be submitted match the Template"
print("Submission form Shape is", HEX.shape)
print("Number of Municipalities is", HEX["CODIGO MUNICIPIO"].nunique())
print("The Total 2020 Nests' Prediction is", int(HEX["NIDOS 2020"].sum()))
assert HEX.shape == (112, 3), "Error: Shape is incorrect."
assert HEX["CODIGO MUNICIPIO"].nunique() == 112, "Error: Number of unique municipalities is correct."
return template_checker(HEX)
# Importing datasets from GitHub as Pandas Dataframes
queen_train = pd.read_csv("../Feeder_years/WBds03_QUEENtrainYEARS.csv", encoding="utf-8") #2018+2019 test df
queen_predict = pd.read_csv("../Feeder_years/WBds03_QUEENpredictYEARS.csv", encoding="utf-8") #2020 prediction df
template = pd.read_csv("../../../Input_open_data/ds01_PLANTILLA-RETO-AVISPAS-KOPURU.csv",sep=";", encoding="utf-8")
den_com = pd.read_excel("../../../Other_open_data/densidad comercial.xlsx")
cluster= pd.read_csv("../auxiliary_files/WBds_CLUSTERSnests.csv")
den_com_melt= pd.melt(den_com, id_vars=['Código municipio'], value_vars=['2019', '2018', '2017'], var_name='year_offset',
value_name='densidad')
den_com_melt.rename({'Código municipio':'municip_code'}, axis=1, inplace=True)
den_com_melt["densidad"] = den_com_melt["densidad"].apply(lambda x: x.replace(",", "."))
den_com_melt['year_offset']= den_com_melt['year_offset'].apply(str)
```
## New queen Train dataset
```
df_train= queen_train.iloc[:,:33]
df_train['year_offset']= df_train['year_offset'].apply(str)
df_train = df_train.merge(den_com_melt,\
how='left', left_on=['municip_code','year_offset'],\
right_on=['municip_code','year_offset']).merge(cluster, how='left', on= 'municip_code') #Merge Densidad comercial + Cluster
#Cleaning
df_train.drop(['municip_name_y','station_code'], axis=1, inplace=True)
df_train.rename({'municip_name_x': 'municip_name'}, axis=1, inplace=True)
```
## New queen predict dataset
```
queen_predict['year_offset']= queen_predict['year_offset'].apply(str)
df_predict= queen_predict.loc[:,['municip_name', 'municip_code', 'year_offset','population']].merge(den_com_melt,\
how='left', left_on=['municip_code','year_offset'],\
right_on=['municip_code','year_offset']).merge(cluster, how='left',on='municip_code')
df_predict.drop(['municip_name_y'], axis=1, inplace=True)
df_predict.rename({'municip_name_x': 'municip_name'}, axis=1, inplace=True)
#Aux to predict (X_Predict)
aux_predict= df_predict.iloc[:,3:]
y = df_train.NESTS
# X will be the explanatory variables. Remove response variable and non desired categorical columns such as (municip code, year, etc...)
X = df_train.loc[:,['population', 'densidad','Cluster']]
```
## Forecasting
```
# Scale the datasets using MinMaxScaler
X_scaled = preprocessing.minmax_scale(X) # this creates a numpy array
X_scaled = pd.DataFrame(X_scaled,index=X.index,columns=X.columns)
# selecting the XGBoost model and fitting with the train data
model = XGBRegressor(random_state=0, objective="reg:squarederror")
```
### Use GridSearchCV to find out the best hyperparameters for our XGBoost model with our Fitted Data
```
# Use GridSearchCV that will automatically split the data and give us the best estimator by:
#1) Establishing hyperparameters to change
param_grid = {
"learning_rate": [0.2],
"max_depth": [6],
"gamma" : [0.5],
#"max_delta_step" : [3],
"min_child_weight": [9],
#"subsample": [0.9],
#"colsample_bytree": [0.4],
"reg_lambda" : [1.5],
"n_estimators": [200],
"scale_pos_weight" : [3.5]
}
warnings.filterwarnings(action='ignore', category=UserWarning)
grid = GridSearchCV(model, param_grid, cv=3)
#2) Fitting the model with our desired data and check for best results
grid.fit(X_scaled, y)
#) Retrieve the summary of GridSearchCV for analysis
print(F"The number homogeneous splits conducted by GridSearchCV are: {grid.n_splits_}.")
print(F"The best hyperparameters found were: {grid.best_params_}.")
print(F"The best score found was: {grid.best_score_}.")
# Reset warnigns to default (this is used to suppred a warning message from XGBoost model and avoid converting X_train to numpy to keep features name)
warnings.filterwarnings(action='default', category=UserWarning)
model = grid.best_estimator_
model.fit(X_scaled, y)
xgb.plot_importance(model, height=0.5, xlabel="F-Score", ylabel="Feature Importance", grid=False, )
xgb.plot_tree(model)
# make a prediction
X_scaled_pred = preprocessing.minmax_scale(aux_predict)
X_scaled_pred = pd.DataFrame(X_scaled_pred,index=aux_predict.index,columns=aux_predict.columns)
X_scaled_pred = np.ascontiguousarray(X_scaled_pred)
df_predict['nests_2020'] = model.predict(X_scaled_pred)
df_predict.nests_2020.sum()
```
## Add Each Cluster Predictions to the original DataFrame and Save it as a `.csv file`
```
# Remove the Municipalities to which we did not assign a Cluster, since there was not reliable data for us to predict
df_predict = df_predict.loc[~df_predict.municip_code.isin([48020]),:]
# Create a new DataFrame with the Municipalities to insert manualy
HEX_aux = pd.DataFrame({"CODIGO MUNICIPIO":[48020],\
"NOMBRE MUNICIPIO":["Bilbao"],\
"NIDOS 2020":[0]})
HEX = df_predict.loc[:,["municip_code","municip_name","nests_2020"]].round() # create a new Dataframe for Kopuru submission
HEX.columns = ["CODIGO MUNICIPIO","NOMBRE MUNICIPIO","NIDOS 2020"] # change column names to Spanish (Decidata template)
HEX = HEX.append(HEX_aux, ignore_index=True) # Add rows of municipalities to add manually
# Final check
check_data(HEX)
# reset max_rows to default values (used in function to see which rows did not match template)
pd.reset_option("max_rows")
# Save the new dataFrame as a .csv in the current working directory on Windows
HEX.to_csv("WaspBusters_20210608_XGyears_ClusterMB_PC4_Zeros_Gridsearchcv.csv", index=False)
```
| github_jupyter |
# Lecture 16 - Exceptions and Unit Testing (https://bit.ly/intro_python_16)
* Unit-testing with unitest
* Exceptions and error handling
# Unit testing with the unittest module
* As programs grow in complexity, the scope for bugs becomes huge.
* Satisfactorily debugging a complex program without systematic testing is **hard or even intractable**.
* With **unit-testing** you design tests to test individual "units" of the code, e.g. functions and classes.
* Unit-testing allows you to progressively debug code and build tested modules of code with less fear that nothing will work when you finally put it together.
# Unittesting example: Fibonacci numbers
* Let's look at a simple example, debugging a function for computing members of the Fibonacci sequence
* Recall the ith Fibonacci number is equal to the sum of the previous two Fibonacci numbers, and 0 and 1 are the 0th and 1st Fibonacci numbers.
* i.e. fib(i) = fib(i-1) + f(i-2), for i > 1
* This definition is naturally recursive, so we can use a recursive implementation of the function.
```
# Example demonstrating unittesting using the Python unittest module
def fib(i):
""" Compute the ith fibonacci number recursively
"""
assert type(i) == int and i >= 0
return 1 if i == 0 else 1 if i == 1 else fib(i-1) + fib(i-2) # Note, this is not quite right!
```
Here's how we test it using unittest:
```
import unittest # unittest is the standard Python library module for unit testing,
# it's great
class FibTest(unittest.TestCase): # Note the use of inheritence
""" We create a test class that inherits from the unittest.TestCase
class
"""
def test_calc_x(self):
""" Each test we create must start with the name "test"
"""
self.assertEqual(fib(0), 0)
self.assertEqual(fib(1), 1)
self.assertEqual(fib(2), 1)
self.assertEqual(fib(3), 2)
self.assertEqual(fib(4), 3)
self.assertEqual(fib(5), 5)
self.assertEqual(fib(6), 8)
unittest.main(argv=['first-arg-is-ignored'], exit=False)
```
Okay, so our test failed, let's fix it:
```
# Example demonstrating unittesting using the Python
# unittest module
# Let's suppose we want to test our implementation of the fibonnaci sequence
def fib(i):
""" Compute the ith fibonacci number recursively
Reminder the ith fibonnaci number is equal to the sum of the previous
two previous fibonnacci numbers, where the 0th fibonacci number is 0 and the 1st
is 1.
"""
assert type(i) == int and i >= 0
return 0 if i == 0 else 1 if i == 1 else fib(i-1) + fib(i-2) # That's right
```
Now we can rerun the tests:
```
import unittest # unittest is the standard Python library module for unit testing,
# it's great
class FibTest(unittest.TestCase): # Note the use of inheritence
""" We create a test class that inherits from the unittest.TestCase
class
"""
def test_calc_x(self):
""" Each test we create must start with the name "test"
"""
self.assertEqual(fib(0), 0)
self.assertEqual(fib(1), 1)
self.assertEqual(fib(2), 1)
self.assertEqual(fib(3), 2)
self.assertEqual(fib(4), 3)
self.assertEqual(fib(5), 5)
self.assertEqual(fib(6), 8)
unittest.main(argv=['first-arg-is-ignored'], exit=False)
```
Okay, this example is contrived, but the idea that you should write code to test your code turns out to be remarkably useful.
What if you want to write multiple tests for multiple different functions?
# Writing multiple tests: setUp() and tearDown()
* It is good to keep tests small and modular.
* If you want to test lots of related functions, e.g. the functions of a class, it is therefore helpful to have shared "setup" and "cleanup" functions that are run, respectively, before and after each test. You can achieve this with the *setUp()* and *tearDown()* functions of the unittest.TestCase function.
```
# Here's the card class we studied before when looking at the Old Maid card game:
class Card:
""" Represents a card from a deck of cards.
"""
# Here are some class variables
# to represent possible suits and ranks
suits = ["Clubs", "Diamonds", "Spades", "Hearts"]
ranks = ["narf", "Ace", "2", "3", "4", "5", "6", "7",
"8", "9", "10", "Jack", "Queen", "King"]
def __init__(self, suit=0, rank=0):
""" Create a card using integer variables to represent suit and rank.
"""
# Couple of handy asserts to check any cards we build make sense
assert suit >= 0 and suit < 4
assert rank >= 0 and rank < 14
self.suit = suit
self.rank = rank
def __str__(self):
# The lookup in the suits/ranks lists prints
# a human readable representation of the card.
return (self.ranks[self.rank] + " of " + self.suits[self.suit])
def same_color(self, other):
""" Returns the True if cards have the same color else False
Diamons and hearts are both read, clubs and spades are both black.
"""
return self.suit == other.suit or self.suit == (other.suit + 2) % 4
# The following methods implement card comparison
def cmp(self, other):
""" Compares the card with another, returning 1, 0, or -1 depending on
if this card is greater than, equal or less than the other card, respectively.
Cards are compared first by suit and then rank.
"""
# Check the suits
if self.suit > other.suit: return 1
if self.suit < other.suit: return -1
# Suits are the same... check ranks
if self.rank > other.rank: return 1
if self.rank < other.rank: return -1
# Ranks are the same... it's a tie
return 0
def __eq__(self, other):
return self.cmp(other) == 0
def __le__(self, other):
return self.cmp(other) <= 0
def __ge__(self, other):
return self.cmp(other) >= 0
def __gt__(self, other):
return self.cmp(other) > 0
def __lt__(self, other):
return self.cmp(other) < 0
def __ne__(self, other):
return self.cmp(other) != 0
```
To test the individual functions we could do something like this:
```
class CardTest(unittest.TestCase):
""" Test the Card class
"""
def setUp(self):
print("setUp")
# This function gets run before each test
self.aceClubs = Card(0, 1) # Ace of clubs
self.aceDiamonds = Card(1, 1) # Ace of diamonds
self.aceSpades = Card(2, 1) # Ace of spades
def tearDown(self):
# This function gets run after each function.
# Here I do nothing in teardown, but print a message
# but you can use it to cleanup temporary files, etc.
print("tearDown")
def test_same_color(self):
""" Tests Card.same_color()
"""
print("Running test_same_color") # These print messages are just to show you what's going on
self.assertTrue(self.aceClubs.same_color(self.aceSpades))
self.assertFalse(self.aceSpades.same_color(self.aceDiamonds))
def test_str(self):
""" Tests Card.__str__()"""
print("Running test_str")
self.assertEqual(str(self.aceClubs), "Ace of Clubs")
self.assertEqual(str(self.aceSpades), "Ace of Spades")
self.assertEqual(str(self.aceDiamonds), "Ace of Diamonds")
unittest.main(argv=['first-arg-is-ignored'], exit=False)
```
# Challenge 1
```
# Create your own CardTest unit test class, called ExpandedCardTest, which additionally includes a test for
# comparing cards using the equals and comparison operators. The test should compare
# the expected ordering of the three cards created by the setUp method (ace of clubs, ace of diamonds, ace of spades)
# In this program "Clubs" < "Diamonds" < "Spades" < "Hearts".
```
# Writing Test Suites
* Generally for each Python module you write you create an accompanying unittest module.
* e.g. if you write "foo.py" you also create "fooTest.py".
* It's beyond scope here, but as you write more complex programs, with multiple modules organized into packages, you can automate running all your tests together in one test suite. When you make a change to your code you then rerun all the tests and check everything is still good.
* As a rough rule of thumb, good programmers write about us much unit test code as they write program code.
* It seems like a long way around, but it is generally quicker and more manageable than ad hoc debugging which is otherwise inevitable.
* One popular approach is to write the tests before writing the core of the program, this is partly the philosophy of "test driven development"
* This helps figure out what the program should do and how it should behave before going to far into the actual implementation.
# Exceptions
When a runtime error occurs Python creates an exception. We've seen these, e.g.:
```
assert False # Creates an AssertionError, a kind of exception
```
* So far we've just encountered exceptions when the program fails, but actually we can frequently handle exceptions within the program and not crash.
* To do this we use the try / except syntax. Consider:
```
try:
assert False
except AssertionError:
print("We got an assert error")
print("But we're fine!")
# The syntax is
try:
STATEMENT_BLOCK_1
except [ERROR TYPE]:
STATEMENT_BLOCK_2
```
The way this works:
* The statement block STATEMENT_BLOCK_1 is executed.
* If an exception occurs of type ERROR_TYPE during the execution of STATEMENT_BLOCK_1 then STATEMENT_BLOCK_1 stops execution and STATEMENT_BLOCK_2 executes.
* If not exception occurs during STATEMENT_BLOCK_1, STATEMENT_BLOCK_2 is skipped.
* This allows us to handle unexpected events in a predictable way
Consider how parsing user input can create errors:
```
i = int(input("Enter an integer: ")) # What happens if I don't enter a valid integer?
```
We can handle this using try/except:
```
while True:
try:
i = int(input("Enter an integer: "))
break
except ValueError:
print("Got an error parsing user input, try again!")
print("You entered: ", i)
```
**You don't have to specify the exception type**
* If you don't know what error to anticipate you can not specify the type of exception:
```
while True:
try:
i = int(input("Enter an integer: "))
break
except: # Note we don't say what kind of error it is
print("Got an error parsing user input, try again!")
print("You entered: ", i)
```
The downside of not specifying the type of the expected exception, is that except without a type will catch all exceptions, possibly including unrelated errors.
# Challenge 2
```
# Practice problem
# Write a function "get_file" to ask the user for a file name.
# Return an open file handle to the users file in "read" mode.
# Use exception handling to deal with the case that the user's file
# does not exist, printing an error saying "File does not exist, try again"
# and trying again to get a file from the user name
# Hint use FileNotFoundError
```
# Finally
Finally allows us to specify code that will be run regardless of if there is an error:
```
try:
f = open("out.txt", "w")
f.write("Hello, file!\n")
assert 1 == 2
except:
print("Got an error")
finally:
print("Closing the file")
f.close()
```
The way this works:
* if there is an error, the error is dealt with.
* the finally clause is then run, regardless of if there is an error or not
# Raise
If you want to create your own exception use "Raise":
```
def get_age():
age = int(input("Please enter your age: "))
if age < 0:
# Create a new instance of an exception
my_error = ValueError("{0} is not a valid age".format(age))
raise my_error
return age
get_age()
```
You can also use this to "rethrow" an exception:
```
try: # This is a contrived example
assert 1 == 2
except:
print("Got an error")
raise # This "rethrows" the exception we caught
```
Note: There is a lot more to say about exceptions and writing false tolerant code, but hopefully this summary is a good start!
# Challenge 3
```
# Write code that uses exception handling and the get_age function defined above
# to probe a user for a valid age, repeating the prompt until a valid age is given.
```
# Reading
* Open book Chapter 19: (exceptions)
http://openbookproject.net/thinkcs/python/english3e/exceptions.html
# Homework
* Go to Canvas and complete the lecture quiz, which involves completing each challenge problem
* Zybooks Reading 16
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Automated Machine Learning
_**Prepare Data using `azureml.dataprep` for Remote Execution (DSVM)**_
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Results](#Results)
1. [Test](#Test)
## Introduction
In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).
Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.
In this notebook you will learn how to:
1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.
2. Pass the `Dataflow` to AutoML for a local run.
3. Pass the `Dataflow` to AutoML for a remote run.
## Setup
Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros.
As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
import time
import pandas as pd
import azureml.core
from azureml.core.compute import DsvmCompute
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
import azureml.dataprep as dprep
from azureml.train.automl import AutoMLConfig
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = 'automl-dataprep-remote-dsvm'
# project folder
project_folder = './sample_projects/automl-dataprep-remote-dsvm'
experiment = Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
## Data
```
# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.
# The data referenced here was a 1MB simple random sample of the Chicago Crime data into a local temporary directory.
# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)
# and convert column types manually.
example_data = 'https://dprepdata.blob.core.windows.net/demo/crime0-random.csv'
dflow = dprep.auto_read_file(example_data).skip(1) # Remove the header row.
dflow.get_profile()
# As `Primary Type` is our y data, we need to drop the values those are null in this column.
dflow = dflow.drop_nulls('Primary Type')
dflow.head(5)
```
### Review the Data Preparation Result
You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets.
`Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage.
```
X = dflow.drop_columns(columns=['Primary Type', 'FBI Code'])
y = dflow.keep_columns(columns=['Primary Type'], validate_column_exists=True)
```
## Train
This creates a general AutoML settings object applicable for both local and remote runs.
```
automl_settings = {
"iteration_timeout_minutes" : 10,
"iterations" : 2,
"primary_metric" : 'AUC_weighted',
"preprocess" : True,
"verbosity" : logging.INFO
}
```
### Create or Attach a Remote Linux DSVM
```
dsvm_name = 'mydsvmb'
try:
while ws.compute_targets[dsvm_name].provisioning_state == 'Creating':
time.sleep(1)
dsvm_compute = DsvmCompute(ws, dsvm_name)
print('Found existing DVSM.')
except:
print('Creating a new DSVM.')
dsvm_config = DsvmCompute.provisioning_configuration(vm_size = "Standard_D2_v2")
dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)
dsvm_compute.wait_for_completion(show_output = True)
print("Waiting one minute for ssh to be accessible")
time.sleep(90) # Wait for ssh to be accessible
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
conda_run_config = RunConfiguration(framework="python")
conda_run_config.target = dsvm_compute
cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy','py-xgboost<=0.80'])
conda_run_config.environment.python.conda_dependencies = cd
```
### Pass Data with `Dataflow` Objects
The `Dataflow` objects captured above can also be passed to the `submit` method for a remote run. AutoML will serialize the `Dataflow` object and send it to the remote compute target. The `Dataflow` will not be evaluated locally.
```
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
path = project_folder,
run_configuration=conda_run_config,
X = X,
y = y,
**automl_settings)
remote_run = experiment.submit(automl_config, show_output = True)
remote_run
```
### Pre-process cache cleanup
The preprocess data gets cache at user default file store. When the run is completed the cache can be cleaned by running below cell
```
remote_run.clean_preprocessor_cache()
```
## Results
#### Widget for Monitoring Runs
The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.
```
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
```
#### Retrieve All Child Runs
You can also use SDK methods to fetch all the child runs and see individual metrics that we log.
```
children = list(remote_run.get_children())
metricslist = {}
for run in children:
properties = run.get_properties()
metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
```
### Retrieve the Best Model
Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
```
best_run, fitted_model = remote_run.get_output()
print(best_run)
print(fitted_model)
```
#### Best Model Based on Any Other Metric
Show the run and the model that has the smallest `log_loss` value:
```
lookup_metric = "log_loss"
best_run, fitted_model = remote_run.get_output(metric = lookup_metric)
print(best_run)
print(fitted_model)
```
#### Model from a Specific Iteration
Show the run and the model from the first iteration:
```
iteration = 0
best_run, fitted_model = remote_run.get_output(iteration = iteration)
print(best_run)
print(fitted_model)
```
## Test
#### Load Test Data
For the test data, it should have the same preparation step as the train data. Otherwise it might get failed at the preprocessing step.
```
dflow_test = dprep.auto_read_file(path='https://dprepdata.blob.core.windows.net/demo/crime0-test.csv').skip(1)
dflow_test = dflow_test.drop_nulls('Primary Type')
```
#### Testing Our Best Fitted Model
We will use confusion matrix to see how our model works.
```
from pandas_ml import ConfusionMatrix
y_test = dflow_test.keep_columns(columns=['Primary Type']).to_pandas_dataframe()
X_test = dflow_test.drop_columns(columns=['Primary Type', 'FBI Code']).to_pandas_dataframe()
ypred = fitted_model.predict(X_test)
cm = ConfusionMatrix(y_test['Primary Type'], ypred)
print(cm)
cm.plot()
```
| github_jupyter |
# Perceptron Demo - Distinguishing Traces of Schizophrenia
## CSCI 4850-5850 - Neural Networks
Being able to detect traces of schizophrenia in a person's brain can be a valuable thing. Diagnosing schizophrenia can be done in a variety of ways, such as physical examinations, tests and screenings, or psychiatric evaluation. Obtaining a solid diagnosis can be difficult and time/cost consuming. In our project, we wanted to apply the use of neural nets to try and detect traces of schizophrenia and accurately diagnose it.
## What data to use - fMRI
There are several ways to detect schizophrenia, but one of the most popular ways is through brain scans. Since schizophrenia is diagnosed as a mental disorder, the brain is directly correlated with it. The dopamine produced by the brain is tied to the hallucinations that schizophrenic patients see or hear. A good way to detect the activity of the brain is through Functional Magnetic Resonance Imaging (fMRI). An fMRI measures the flow of blood in one's brain. By viewing an fMRI, a doctor can see if certain activity/inactivity in a region of the patient's brain could be a sign of schizophrenia. So, since we have an image that can tell us if a patient has traces of schizophrenia, we can plug that into a neural net to see if it can detect it for us! Hopefully, this will allow doctors to just be able to scan a patient's brain, plug in into the neural net, and wait for the net to decide if that patient has schizophrenia or not. However, we want the highest possible accuracy we can get in order to cut down on misdiagnoses, time and cost.
Since fMRI is a highly valuable dataset with a lot of information packed into a few dimensions, this proves it to be difficult to use in a neural net. An fMRI is a scan of the patient's brain sliced into several regions over several timestamps, which makes it difficult to efficiently feed into a neural net.
The data that we'll be using for this demo is provided by The Center for Biomedical Research Excellence (COBRE). This dataset contains MR data from 72 schizophrenic patients and 75 MR scans from healthy controls. The ages of these test patients range from 18 to 65.
With these fMRI scans, there are 2 ways we can go about reading them into a neural net:
1) We can use a convolutional neural net to scan over the images and try and detect any signs of
schizophrenia by the images alone. (Images)
2) We can read in the images and create a covariance matrix to see how each section of the brain
is connected. (Numeric)
# Neural Net A: Convolutional Net
## Step 1: Loading the Data
In order to use the COBRE data set, we need to use a few tools: `nilearn` and `nibabel`
```
import numpy as np
from nibabel.testing import data_path
import nibabel as nib
import keras
from keras import backend as K
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d
%matplotlib inline
from IPython.display import display
import nilearn
from nilearn import plotting
from nilearn import image
from nilearn import datasets
from keras_tqdm import TQDMNotebookCallback
# Get dataset with nilearn function
# if not downloaded, will download. If already downloaded, will use local version
dataset = nilearn.datasets.fetch_cobre(n_subjects=146, data_dir="/nfshome/sandbox/perceptron",
url=None, verbose=0)
phenotypes = dataset["phenotypic"]
confounds = dataset["confounds"]
file_paths = dataset["func"]
phenotypes.sort(0) #sort by column corresponding to patient number
file_paths.sort() #sort file names by alphabetical order, which will result in sorting by patient number
confounds.sort()
#file_paths is now a regular python list of the file paths to the fmri scans
#phenotypes is now a np.recarray of np.records storing patient info
# get just the diagnosis information from the phenotypes
diagnosis = phenotypes['diagnosis']
diagnosis_converted = []
#this stem is necessary to convert np.byte array into strings,
#and then fit those strings into 2 categories:
#Schizophrenia or no Schizophrenia
for item in diagnosis:
s = item.decode('UTF-8')
if s != "None":
diagnosis_converted.append(float(1)) #person has schizophrenia
else:
diagnosis_converted.append(float(0)) #person doesn't have schizophrenia
del diagnosis_converted[74] # item 74 is a messed up scan with different dimensions
del file_paths[74] # so it needs to be removed
del confounds[74]
Y = np.array(diagnosis_converted)
```
Now that we have our images loaded in, We still need to tweak it a bit to make sure the net can efficiently read it.
## Step 2: Preparing the Data for Input
The issue with the fMRI scans is that they are far too large to be efficiently read into a neural net. Each scan consist of 26 slices of the regions of the brain across 150 timestamps. So, we are going to run the scans through a mask (the MSDL Mask) to split the brain up into 39 regions and take the average of each region's blood oxygen levels. This will drastically reduce the size and noise in the data.
```
# this is a brain anatomical atlas template, that gives us brain reigons with their labels
msdl_atlas_dataset = nilearn.datasets.fetch_atlas_msdl(data_dir="/nfshome/sandbox")
from nilearn import image
from nilearn import input_data
from sklearn.externals.joblib import Memory # A "memory" to avoid recomputation
mem = Memory('nilearn_cache')
# mask the data, used the atlas template as the mask img.
masker = input_data.NiftiMapsMasker(
msdl_atlas_dataset.maps, resampling_target="maps", detrend=True,
low_pass=.5, high_pass=0.01, t_r=2.0, standardize=True,
memory='nilearn_cache', memory_level=1, verbose=0)
masker.fit()
# for each of our fmri scams, compute confounds, transform them into mask, and append
# to time series list
subject_time_series = []
for file_path, confound in zip(file_paths, confounds):
# Computing some confounds
hv_confounds = mem.cache(image.high_variance_confounds)(
file_path, n_confounds=10)
region_ts = masker.transform(file_path,
confounds=[hv_confounds, confound])
subject_time_series.append(region_ts)
#Here is where I prepare the data for input
X = np.array(subject_time_series)
fullx = X.astype('float32').reshape(X.shape+(1,))
#one-hot encoding
Y = keras.utils.to_categorical(Y, len(np.unique(Y))) #one hot encoding
x_train = fullx[:110]
x_test = fullx[110:]
y_train = Y[:110]
y_test = Y[110:]
print(x_train.shape)
print(y_train.shape)
print(y_test.shape)
print(x_test.shape)
```
## Step 3: Building the Net
For the net, we'll build a 10 layer Convolution Net with a pooling layer and a couple batch and dropout layers.
```
#time to finally build a model and test it!
model1 = keras.Sequential()
model1.add(keras.layers.Conv2D(256, kernel_size = (2,2), activation = 'relu'
,input_shape=[150,39,1]))
#model1.add(keras.layer.MaxPooling2D(pool_size=(2,2)))
model1.add(keras.layers.BatchNormalization(momentum=0.8))
model1.add(keras.layers.Conv2D(128, (8,8), activation='relu'))
model1.add(keras.layers.BatchNormalization(momentum=0.8))
model1.add(keras.layers.MaxPooling2D(pool_size=(2,2)))
model1.add(keras.layers.Dropout(0.6))
model1.add(keras.layers.Flatten())
model1.add(keras.layers.Dense(128, activation='relu'))
model1.add(keras.layers.Dropout(0.3))
model1.add(keras.layers.Dense(y_train.shape[1], activation='softmax')) #y_train.shape[1]??
model1.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(lr=0.0001),
metrics=['accuracy']) #0.0001
model1.summary()
```
Looks good, let's train it!
```
#model1.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(), metrics=['accuracy'])
batch_size = 10
epochs = 20
validation_split = 0.1
history = model1.fit(x_train, y_train, batch_size = batch_size, epochs = epochs
, verbose = 0, validation_split = validation_split, callbacks=[TQDMNotebookCallback()])
print('Accuracy:',model1.evaluate(x_test,y_test)[1]*100.0,'%')
plt.figure()
plt.subplot(211)
#summarize accuracy history
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.legend(['train', 'test'], loc='upper left')
#summarize loss history
plt.subplot(212)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['train', 'test'], loc='upper left')
plt.tight_layout()
plt.show()
```
As you can see, the best results that we could get out of the net are an accuracy of ~50%. The net is having a hard time generalizing the data and seems to suffer from overfitting. Scanning through the images and looking for significant changes in each region of the brain may be too much to learn in a short amount of time. However, let's see if we can fix this in our next approach!
# Neural Net B: Feed Forward Net (Using a Convariance Matrix)
## Step 1: Loading and Viewing the Data
In order to use the COBRE data set, we need to use a few tools: `nilearn` and `nibabel`
```
# nilearn helps with loading and handling of the COBRE dataset and is actually built to help run this dataset
import nilearn
from nilearn import plotting
from nilearn import image
from nilearn import datasets
# nibabel also helps with the testing of the dataset
from nibabel.testing import data_path
import nibabel as nib
# import other basic necessities
import os
import numpy as np
import keras
from keras import backend as K
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d
%matplotlib inline
from IPython.display import display
from nilearn import image
from nilearn import input_data
from sklearn.externals.joblib import Memory # A "memory" to avoid recomputation
from nilearn.connectome import ConnectivityMeasure
from nilearn import plotting
# Visualization
from IPython.display import SVG
from IPython.display import display
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
from sklearn.model_selection import StratifiedKFold
```
Now that we have our tools all set out, lets start unpacking the data.
```
# Get dataset with nilearn function
# if not downloaded, will download. If already downloaded, will uses local version
dataset = nilearn.datasets.fetch_cobre(n_subjects=146,
data_dir="/nfshome/sandbox/perceptron",
url=None,
verbose=0)
file_paths = dataset["func"] #list of file names
confounds = dataset["confounds"] #list of confound file names
phenotypes = dataset["phenotypic"] # Contains phenotypic information of patients, we only use schizophrenia diagnosis
# sort lists so they are all corresponding by file names 0-146
phenotypes.sort(0)
file_paths.sort()
confounds.sort()
#file_paths is now a regular python list of the file paths to the fmri scans
#phenotypes is now a np.recarray of np.records storing patient info
# number 74 is misisng samples so it needs to be removed
del file_paths[74]
del confounds[74]
# get just the diagnosis information from the phenotypes
diagnosis = phenotypes['diagnosis']
diagnosis_converted = []
#this stem is necessary to convert np.byte array into strings, and then fit those strings into 2 categories:
#Schizophrenia or no Schizophrenia
for item in diagnosis:
s = item.decode('UTF-8')
if s != "None":
diagnosis_converted.append(float(1)) #person has schizophrenia
else:
diagnosis_converted.append(float(0)) #person doesn't have schizophrenia
del diagnosis_converted[74]
Y = np.array(diagnosis_converted)
```
The below part isn't necessary, it only shows the original shape of the data before we do some more modifications to it in order for it to fit into the neural net easier.
```
y_temp = np.array(diagnosis_converted)
y_temp = keras.utils.to_categorical(y_temp, len(np.unique(y_temp))) #one hot encoding
# Make x train from the file paths
scans = []
for item in file_paths:
scan = nib.load(item)
data = scan.get_fdata()
scans.append(data)
x_temp = np.array(scans)
#X train is now 145 different fmri scans, with dimensions 27x32x26x150
#the 27x32x26 is length, width, and height
#the 150 is time, there are 150 different 3d 'voxels' or times for each full fmri scan
x_temp.shape
```
Now that we've loaded the data into an array without modifying it at all, we can see how each image is measured. Starting from left to right, there are 145 images, of length 27, width 32, and height 26 taken at 150 different timestamps.
## Step 2: Condensing the data
As you can see from the size of the array above, there are a lot of dimensions in a single fMRI scan. The fact that it is sliced into 26 different layers and stretched across 150 different timestamps also adds complexity in that we don't know what slice or timestamp to use? If we use all of them, it will slow the training of the neural net down and could add unnecessary noise to our data.
To fix this, we are going to mask the data using a brain atlas. We will use a predefined probabilistic brain atlas called the MSDL atlas. This gives us 39 spatial reigons of interest and averages together our FMRI blood oxygen values for all the voxels in each these reigons of interest. This process significantly reduces the size of the data and other noisy factors.
```
# this is a brain anatomical atlas template, that gives us brain reigons with their labels
msdl_atlas_dataset = nilearn.datasets.fetch_atlas_msdl(data_dir="/nfshome/sandbox")
mem = Memory('nilearn_cache')
# mask the data, used the atlas template as the mask img.
masker = input_data.NiftiMapsMasker(
msdl_atlas_dataset.maps, resampling_target="maps", detrend=True,
low_pass=.5, high_pass=0.01, t_r=2.0, standardize=True,
memory='nilearn_cache', memory_level=1, verbose=0)
masker.fit()
# for each of our fmri scams, compute confounds, transform them into mask, and append
# to time series list
subject_time_series = []
for file_path, confound in zip(file_paths, confounds):
# Computing some confounds
hv_confounds = mem.cache(image.high_variance_confounds)(
file_path, n_confounds=10)
region_ts = masker.transform(file_path,
confounds=[hv_confounds, confound])
subject_time_series.append(region_ts)
```
## Step 3: Connecting the regions
Now that we've condensed the data into the 39 regions that we want, we can start to compute a covariance matrix on our masked data. This allows us to obtain information on how our reigons of interest are connecting with each other.
```
correlation_measure = ConnectivityMeasure(kind='covariance',vectorize=True)
correlation_measure.fit(subject_time_series)
matrices = correlation_measure.transform(subject_time_series)
correlation_measure2d = ConnectivityMeasure(kind='covariance',vectorize=False)
correlation_measure2d.fit(subject_time_series)
matrices2d = correlation_measure2d.transform(subject_time_series)
# function code source: nilearn documentation
# https://nilearn.github.io/auto_examples/03_connectivity/plot_group_level_connectivity.html#sphx-glr-auto-examples-03-connectivity-plot-group-level-connectivity-py
def plot_matrices(matrices, matrix_kind):
n_matrices = len(matrices)
fig = plt.figure(figsize=(n_matrices * 4, 4))
for n_subject, matrix in enumerate(matrices):
plt.subplot(1, n_matrices, n_subject + 1)
matrix = matrix.copy() # avoid side effects
# Set diagonal to zero, for better visualization
np.fill_diagonal(matrix, 0)
vmax = np.max(np.abs(matrix))
title = '{0}, subject {1}'.format(matrix_kind, n_subject)
plotting.plot_matrix(matrix, vmin=-vmax, vmax=vmax, cmap='RdBu_r',
title=title, figure=fig, colorbar=False)
```
The grid images below represent our covariance matrices. The values range from 1 to -1 (red to blue). The X and Y axis represent our 39 brain reigons where each two reigons have a specific shared covariance value that is represented by the color at their specified coordinate.
The connectome is made from the covariance matrices. Connections are made between each of the 39 reigons as outlined in the MSDL atlas. Each reigon is shown as connected to each other. Reigons with a stronger covariance are represented as having a stronger connection. The connectome is made based off of the assumption that a higher covariance/correlation represents a stronger connection.
Let's go ahead and take a look at the grids we just created, along with how each reagion is connected within the brain.
```
msdl_coords = msdl_atlas_dataset.region_coords
display("Covariance matrices of first 4 subjects")
plot_matrices(matrices2d[:4], "Covariance Matrix")
plotting.plot_connectome(matrices2d[0], msdl_coords,
title='Example Connectome of Subject 0')
```
## Step 4: The Neural Net
First, we need to take our covariance matrix and flatten it out into a 1d vector. The covariance matrix for each subject becomes our input to the neural network.
```
X = np.array(matrices)
X.shape
```
Now that looks a lot cleaner than the 5D array that we initially started with!
```
x_train = X[:105]
x_test = X[105:]
y_train = Y[:105]
y_test = Y[105:]
```
For this neural net, we are going to use a multilayer feedforward neural network with 1 hidden layer of 450 units. The hidden layer has a relu activation function, and the output layer has a sigmoid activation function. A binary cross entropy loss function is used as this is a binary classification problem.
We employ a dropout layer with 10% dropout on the input layer in order to achieve slightly better generalization. We also use batch normalization to normalize hidden layer activations.
```
input_dim=x_train.shape[1]
output_dim=1
# Multi-layer net with ReLU hidden layer
model = keras.models.Sequential()
model.add(keras.layers.Dropout(0.1, input_shape=(input_dim,)))
model.add(keras.layers.Dense(450,activation='relu',
bias_initializer=keras.initializers.Constant(0.1)))
model.add(keras.layers.BatchNormalization(momentum=0.8))
# Output layer (size 1), sigmoid activation function
model.add(keras.layers.Dense(output_dim,activation='sigmoid'))
model.compile(loss='binary_crossentropy',optimizer=keras.optimizers.Adam(lr=0.00001), metrics=['accuracy'])
# Display the model
print(model.summary())
```
We can see the structure of the net below:
```
SVG(model_to_dot(model).create(prog='dot', format='svg'))
```
Let's take this net for a test drive!
```
batch_size = len(x_train)
epochs = 800
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=0,
validation_split=0.2)
plt.figure()
# summarize history for accuracy
plt.subplot(211)
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
# summarize history for loss
plt.subplot(212)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.tight_layout()
plt.show()
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
As you can see, our test accuracy isn't too bad, although it can be improved a little bit. The test loss is a bit high and seems to plateau without going too far down. We'll improve this in the final step below.
## Step 5: K Cross Validation
We can use a Stratified K Cross Validation to get accuracy over multiple class label splits. This way we can train and test over the whole dataset.
```
seed = 7
np.random.seed(seed)
kfold = StratifiedKFold(n_splits=10, shuffle=True, random_state=seed)
cvscores = []
for train, test in kfold.split(X, Y):
model = keras.models.Sequential()
model.add(keras.layers.Dropout(0.1, input_shape=(input_dim,)))
model.add(keras.layers.Dense(450,activation='relu',
bias_initializer=keras.initializers.Constant(0.1)))
model.add(keras.layers.BatchNormalization(momentum=0.8))
# Output layer (size 1), sigmoid activation function
model.add(keras.layers.Dense(output_dim,activation='sigmoid'))
model.compile(loss='binary_crossentropy',optimizer=keras.optimizers.Adam(lr=0.00001), metrics=['accuracy'])
batch_size = len(X[train])
epochs = 800
model.fit(X[train], Y[train],
batch_size=batch_size,
epochs=epochs,
verbose=0)
scores = model.evaluate(X[test], Y[test], verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
cvscores.append(scores[1] * 100)
print("%.2f%% (+/- %.2f%%)" % (np.mean(cvscores), np.std(cvscores)))
```
With this net, you should be getting 77.10% (+/- 10.16%) accuracy with 10 K fold cross validation. This indicates that our network is picking up on differences between Schizophrenic and non-Schizohrenic resting state FMRI data with fairly decent precision.
# Final Results
As you can see above, it looks like the feed forward net reading in a covariance matrix of related regions outperforms the covariance net by quite a bit. This is most likely due to the fact that the feed forward net takes into account the relationship between regions of the brain, while the covariance net only looks at the average blood oxygen content for each regions separately.
| github_jupyter |
# Documentation by example for `shap.plots.text`
This notebook is designed to demonstrate (and so document) how to use the `shap.plots.text` function. It uses a distilled PyTorch BERT model from the transformers package to do sentiment analysis of IMDB movie reviews.
Note that the prediction function we define takes a list of strings and returns a logit value for the positive class.
<hr>
<center style="color: red">
<b>Warning!</b> This notebook documents the new SHAP API, and that API is still stablizing over the coming weeks.
</center>
<hr>
```
import shap
import transformers
import nlp
import torch
import numpy as np
import scipy as sp
# load a BERT sentiment analysis model
tokenizer = transformers.DistilBertTokenizerFast.from_pretrained("distilbert-base-uncased")
model = transformers.DistilBertForSequenceClassification.from_pretrained(
"distilbert-base-uncased-finetuned-sst-2-english"
).cuda()
# define a prediction function
def f(x):
tv = torch.tensor([tokenizer.encode(v, pad_to_max_length=True, max_length=500, truncation=True) for v in x]).cuda()
outputs = model(tv)[0].detach().cpu().numpy()
scores = (np.exp(outputs).T / np.exp(outputs).sum(-1)).T
val = sp.special.logit(scores[:,1]) # use one vs rest logit units
return val
# build an explainer using a token masker
explainer = shap.Explainer(f, tokenizer)
# explain the model's predictions on IMDB reviews
imdb_train = nlp.load_dataset("imdb")["train"]
shap_values = explainer(imdb_train[:10])
```
## Single instance text plot
When we pass a single instance to the text plot we get the importance of each token overlayed on the original text that corresponds to that token. Red regions correspond to parts of the text that increase the output of the model when they are included, while blue regions decrease the output of the model when they are included. In the context of the sentiment analysis model here red corresponds to a more positive review and blue a more negative review.
Note that importance values returned for text models are often hierarchical and follow the structure of the text. Nonlinear interactions between groups of tokens are often saved and can be used during the plotting process. If the Explanation object passed to the text plot has a `.hierarchical_values` attribute, then small groups of tokens with strong non-linear effects among them will be auto-merged together to form coherent chunks. When the `.hierarchical_values` attribute is present it also means that the explainer may not have completely enumerated all possible token perturbations and so has treated chunks of the text as essentially a single unit. This happens since we often want to explain a text model while evaluating it fewer times than the numbers of tokens in the document. Whenever a region of the input text is not split by the explainer, it is show by the text plot as a single unit.
The force plot above the text is designed to provide an overview of how all the parts of the text combine to produce the model's output. See the [force plot]() notebook for more details, but the general structure of the plot is positive red features "pushing" the model output higher while negative blue features "push" the model output lower. The force plot provides much more quantitative information than the text coloring. Hovering over a chuck of text will underline the portion of the force plot that corresponds to that chunk of text, and hovering over a portion of the force plot will underline the corresponding chunk of text.
Note that clicking on any chunk of text will show the sum of the SHAP values attributed to the tokens in that chunk (clicked again will hide the value).
```
# plot the first sentence's explanation
shap.plots.text(shap_values[3])
```
## Multiple instance text plot
When we pass a multi-row explanation object to the text plot we get the single instance plots for each input instance scaled so they have consistent comparable x-axis and color ranges.
```
# plot the first sentence's explanation
shap.plots.text(shap_values[:3])
```
## Summarizing text explanations
While plotting several instance-level explanations using the text plot can be very informative, sometime you want global summaries of the impact of tokens over the a large set of instances. See the [Explanation object]() documentation for more details, but you can easily summarize the importance of tokens in a dataset by collapsing a multi-row explanation object over all it's rows (in this case by summing). Doing this treats every text input token type as a feature, so the collapsed Explanation object will have as many columns as there were unique tokens in the orignal multi-row explanation object. If there are hierarchical values present in the Explanation object then any large groups are divided up and each token in the gruop is given an equal share of the overall group importance value.
```
shap.plots.bar(shap_values.abs.sum(0))
```
Note that how you summarize the importance of features can make a big difference. In the plot above the `a` token was very importance both because it had an impact on the model, and because it was very common. Below we instead summize the instances using the `max` function to see the largest impact of a token in any instance.
```
shap.plots.bar(shap_values.abs.max(0))
```
You can also slice out a single token from all the instances by using that token as an input name (note that the gray values to the left of the input names are the original text that the token was generated from).
```
shap.plots.bar(shap_values[:,"but"])
```
## Why the base values can be differenent for each sample
The base values for the explanations above is different for each instance. This is because the default masking behavior when using a transformers tokenizer as the masker is to replace tokens with the `mask_token` defined by the tokenizer. This means that "removing" (i.e. masking) all the tokens in a document still preserves the length of the document, just with all the token replaced by the mask token. It turns out that the model assumes that longer reviews are in general more positive that shorter reviews, so before we even know anything about the content of the review the model the model is biased by the review length. We could create consistent base values by modeling review length as another input feature (this is not built in to SHAP yet).
To see how the model output is biased by length we plot the output score for inputs of different numbers of mask tokens:
```
import matplotlib.pyplot as pl
pl.plot([f(["[MASK]" * i]) for i in range(500)])
pl.xlabel("String length")
pl.ylabel("Model output score")
pl.show()
```
<hr>
Have an idea for more helpful examples? Pull requests that add to this documentation notebook are encouraged!
| github_jupyter |
# T cell epitopes of SARS-CoV2
## Methods
* Predict MHC-I binders for sars-cov2 reference sequences (S and N important)
* Align with sars-cov and get conserved epitopes.
* Best alleles to use?
* Multiple sequence alignment of each protein to reference
* find conservation of binders with closest peptide in each HCov sequence and determine identity
## References
* J. Mateus et al., “Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans,” Science (80-. )., vol. 3871, no. August, p. eabd3871, Aug. 2020.
* S. F. Ahmed, A. A. Quadeer, and M. R. McKay, “Preliminary Identification of Potential Vaccine Targets for the COVID-19 Coronavirus (SARS-CoV-2) Based on SARS-CoV Immunological Studies.,” Viruses, vol. 12, no. 3, 2020.
* A. Grifoni et al., “A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2,” Cell Host Microbe, pp. 1–10, 2020.
* V. Baruah and S. Bose, “Immunoinformatics-aided identification of T cell and B cell epitopes in the surface glycoprotein of 2019-nCoV,” J. Med. Virol., no. February, pp. 495–500, 2020.
## Epitope Loss in Mutations
* https://www.biorxiv.org/content/10.1101/2020.03.27.012013
* https://www.biorxiv.org/content/10.1101/2020.04.10.029454v1?ct=
* https://www.biorxiv.org/content/10.1101/2020.04.07.030924v1
## Common coronoviruses
* https://www.cdc.gov/coronavirus/types.html
```
import os, math, time, pickle, subprocess
from importlib import reload
from collections import OrderedDict, defaultdict
import numpy as np
import pandas as pd
pd.set_option('display.width', 180)
import epitopepredict as ep
from epitopepredict import base, sequtils, plotting, peptutils, analysis
from IPython.display import display, HTML, Image
%matplotlib inline
import matplotlib as mpl
import pylab as plt
import pybioviz
from bokeh.io import show, output_notebook
output_notebook()
import pathogenie
from Bio import SeqIO,AlignIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
```
## load ref genomes
```
labels = {'sars':'NC_004718.3','scov2':'NC_045512.2','229E':'NC_002645.1','NL63':'NC_005831.2','OC43':'NC_006213.1','HKU1':'NC_006577.2'}
genomes = []
for l in labels:
df = ep.genbank_to_dataframe(labels[l]+'.gb',cds=True)
df['label'] = l
genomes.append(df)
genomes = pd.concat(genomes)
scov2_df = genomes[genomes.label=='scov2']
scov2_df = scov2_df.drop_duplicates('locus_tag')
#print (genomes[['label','gene','product','length']])
def get_seqs(gene):
seqs = []
sub = genomes[genomes['gene']==gene]
for i,r in sub.iterrows():
s=SeqRecord(Seq(r.translation),id=r.label)
seqs.append(s)
return seqs
seqs=get_seqs('S')
```
## find orthologs in each genome
### blast the genomes to find corresponding protein as names are ambigious
```
pathogenie.tools.dataframe_to_fasta(genomes, idkey='locus_tag', descrkey='product', outfile='proteins.fa')
pathogenie.tools.make_blast_database('proteins.fa', dbtype='prot')
def get_orthologs(gene):
sub = scov2_df[scov2_df['gene']==gene].iloc[0]
rec = SeqRecord(Seq(sub.translation),id=sub.gene)
bl = pathogenie.tools.blast_sequences('proteins.fa', [rec], maxseqs=10, evalue=1e-4,
cmd='blastp', threads=4)
bl = bl.drop_duplicates('sseqid')
#print (bl.sseqid)
found = genomes[genomes.locus_tag.isin(bl.sseqid)].drop_duplicates('locus_tag')
#print (found)
recs = pathogenie.tools.dataframe_to_seqrecords(found,
seqkey='translation',idkey='label',desckey='product')
return recs
seqs = get_orthologs('S')
aln = pathogenie.clustal_alignment(seqs=seqs)
print (aln)
import Levenshtein
for a in aln:
r=Levenshtein.ratio(str(aln[1].seq).replace('-',''), str(a.seq).replace('-',''))
print (round(r,2), a.id)
spikesars = SeqIO.to_dict(seqs)['sars'].seq
spikesars
p = pybioviz.plotters.plot_sequence_alignment(aln, annot = {'polybasic cleavage site':690,'RBD contact residues':480})
#output_file('alignment.html')
show(p)
sc2 = ep.genbank_to_dataframe('NC_045512.2.gb',cds=True)
sc2 = sc2.drop_duplicates('gene')
```
## predict MHC-I and MHC-II epitopes
```
m1_alleles = ep.get_preset_alleles('broad_coverage_mhc1')
m2_alleles = ep.get_preset_alleles('mhc2_supertypes')
P1 = base.get_predictor('netmhcpan')
P1.predict_sequences(sc2, alleles=m1_alleles,cpus=10,path='netmhcpan',length=9,overwrite=False,verbose=True)
P1.load(path='netmhcpan')
pb1 = P1.promiscuous_binders(n=3, cutoff=.95)
P2 = base.get_predictor('netmhciipan')
P2.predict_sequences(sc2, alleles=m2_alleles,cpus=10,path='netmhciipan',length=15,overwrite=False,verbose=True)
P3 = base.get_predictor('tepitope')
P3.predict_sequences(sc2, alleles=m2_alleles,cpus=10,path='tepitope',length=15,overwrite=False)
P3.load(path='tepitope')
P2.load(path='netmhciipan')
pb2 = P2.promiscuous_binders(n=3, cutoff=.95, limit=70)
#rb2 = P2.promiscuous_binders(n=3, cutoff_method='rank', cutoff=40)
pb2.name.value_counts()
#pb3 = P3.promiscuous_binders(n=3, cutoff=50, cutoff_method='rank', limit=50)
#pb3.name.value_counts()
```
## conservation: find identity to closest peptide in each HCoV sequence
```
import difflib
def get_conservation(x, w):
m = difflib.get_close_matches(x, w, n=1, cutoff=.67)
if len(m)==0:
return 0
else:
m=m[0]
s = difflib.SequenceMatcher(None, x, m)
return s.ratio()
def find_epitopes_conserved(pb,gene,locus_tag):
seqs = get_orthologs(gene)
df = pb[pb.name==locus_tag]
#print (df)
print (len(seqs),len(df))
s=seqs[0]
for s in seqs:
if s.id == 'scov2':
continue
w,ss = peptutils.create_fragments(seq=str(s.seq), length=11)
df.loc[:,s.id] = df.peptide.apply(lambda x: get_conservation(x, w),1)
df.loc[:,'total'] = df[df.columns[8:]].sum(1)
df = df.sort_values('total',ascending=False)
df = df[df.total>0]
df = df.round(2)
return df
df = find_epitopes_conserved(pb2, 'S','GU280_gp02')
#df.to_csv('S_netmhciipan_conserved.csv')
```
## Find conserved predicted epitopes in all proteins
```
res=[]
for i,r in scov2_df.iterrows():
print (r.locus_tag,r.gene)
df = find_epitopes_conserved(pb2,r.gene,r.locus_tag)
df['gene'] = r.gene
res.append(df)
res = pd.concat(res).sort_values('total',ascending=False).dropna().reset_index()
print (len(res),len(pb2))
res.to_csv('scov2_netmhciipan_conserved.csv')
cols = ['gene','peptide','pos','alleles','sars','229E','NL63','OC43','HKU1']
h=res[:30][cols].style.background_gradient(cmap="ocean_r",subset=['sars','229E','NL63','OC43','HKU1']).set_precision(2)
#res[:30][cols]
```
## Compare predictions to mateus exp results
```
s1 = pd.read_csv('mateus_hcov_reactive.csv')
hits=[]
w = list(res.peptide)
for i,r in s1.iterrows():
m = difflib.get_close_matches(r.Sequence, w, n=2, cutoff=.6)
#print (r.Sequence,m,r.Protein)
if len(m)>0:
hits.append(m)
else:
hits.append(None)
s1['hit'] = hits
display(s1)
print (len(s1.hit.dropna())/len(s1))
```
## check epitope selection method
Promiscuous binders are those high scoring above some threshold in multiple alleles. There are several ways to select them that can give different results. By default epitopepredict selects those in each allele above a percentile score cutoff and then counts how many alleles each peptide is present in. We can also limit our set in each protein across a genome to prevent large proteins dominating the list. We can also select by score and protein rank. The overlap is shown in the venn diagram.
```
reload(base)
P = base.get_predictor('tepitope')
P.predict_sequences(sc2, alleles=m2_alleles[:4],names=['GU280_gp01','GU280_gp02','GU280_gp03','GU280_gp04'],cpus=10,length=9)
pb= P.promiscuous_binders(n=2, cutoff=.98, limit=20)
pb.name.value_counts()
rb= P.promiscuous_binders(n=3, cutoff_method='rank',cutoff=30,limit=20)
rb.name.value_counts()
sb= P.promiscuous_binders(n=2, cutoff_method='score',cutoff=3.5,limit=20)
sb.name.value_counts()
from matplotlib_venn import venn3
ax = venn3((set(pb.peptide),set(rb.peptide),set(sb.peptide)), set_labels = ('default', 'ranked', 'score'))
b=P.get_binders(cutoff=10, cutoff_method='rank')
func = max
s=b.groupby(['peptide','pos','name']).agg({'allele': pd.Series.count,
'core': base.first, P.scorekey:[func,np.mean],
'rank': np.median})
s.columns = s.columns.get_level_values(1)
s.rename(columns={'max': P.scorekey, 'count': 'alleles','median':'median_rank',
'first':'core'}, inplace=True)
s = s.reset_index()
s
s.name.value_counts()
s=s.groupby('name').head(10)
s.name.value_counts()
```
| github_jupyter |
# Skip-gram Word2Vec
In this notebook, I'll lead you through using PyTorch to implement the [Word2Vec algorithm](https://en.wikipedia.org/wiki/Word2vec) using the skip-gram architecture. By implementing this, you'll learn about embedding words for use in natural language processing. This will come in handy when dealing with things like machine translation.
## Readings
Here are the resources I used to build this notebook. I suggest reading these either beforehand or while you're working on this material.
* A really good [conceptual overview](http://mccormickml.com/2016/04/19/word2vec-tutorial-the-skip-gram-model/) of Word2Vec from Chris McCormick
* [First Word2Vec paper](https://arxiv.org/pdf/1301.3781.pdf) from Mikolov et al.
* [Neural Information Processing Systems, paper](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) with improvements for Word2Vec also from Mikolov et al.
---
## Word embeddings
When you're dealing with words in text, you end up with tens of thousands of word classes to analyze; one for each word in a vocabulary. Trying to one-hot encode these words is massively inefficient because most values in a one-hot vector will be set to zero. So, the matrix multiplication that happens in between a one-hot input vector and a first, hidden layer will result in mostly zero-valued hidden outputs.
To solve this problem and greatly increase the efficiency of our networks, we use what are called **embeddings**. Embeddings are just a fully connected layer like you've seen before. We call this layer the embedding layer and the weights are embedding weights. We skip the multiplication into the embedding layer by instead directly grabbing the hidden layer values from the weight matrix. We can do this because the multiplication of a one-hot encoded vector with a matrix returns the row of the matrix corresponding the index of the "on" input unit.
<img src='assets/lookup_matrix.png' width=50%>
Instead of doing the matrix multiplication, we use the weight matrix as a lookup table. We encode the words as integers, for example "heart" is encoded as 958, "mind" as 18094. Then to get hidden layer values for "heart", you just take the 958th row of the embedding matrix. This process is called an **embedding lookup** and the number of hidden units is the **embedding dimension**.
There is nothing magical going on here. The embedding lookup table is just a weight matrix. The embedding layer is just a hidden layer. The lookup is just a shortcut for the matrix multiplication. The lookup table is trained just like any weight matrix.
Embeddings aren't only used for words of course. You can use them for any model where you have a massive number of classes. A particular type of model called **Word2Vec** uses the embedding layer to find vector representations of words that contain semantic meaning.
---
## Word2Vec
The Word2Vec algorithm finds much more efficient representations by finding vectors that represent the words. These vectors also contain semantic information about the words.
<img src="assets/context_drink.png" width=40%>
Words that show up in similar **contexts**, such as "coffee", "tea", and "water" will have vectors near each other. Different words will be further away from one another, and relationships can be represented by distance in vector space.
There are two architectures for implementing Word2Vec:
>* CBOW (Continuous Bag-Of-Words) and
* Skip-gram
<img src="assets/word2vec_architectures.png" width=60%>
In this implementation, we'll be using the **skip-gram architecture** with **negative sampling** because it performs better than CBOW and trains faster with negative sampling. Here, we pass in a word and try to predict the words surrounding it in the text. In this way, we can train the network to learn representations for words that show up in similar contexts.
---
## Loading Data
Next, we'll ask you to load in data and place it in the `data` directory
1. Load the [text8 dataset](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/October/5bbe6499_text8/text8.zip); a file of cleaned up *Wikipedia article text* from Matt Mahoney.
2. Place that data in the `data` folder in the home directory.
3. Then you can extract it and delete the archive, zip file to save storage space.
After following these steps, you should have one file in your data directory: `data/text8`.
```
!wget https://s3.amazonaws.com/video.udacity-data.com/topher/2018/October/5bbe6499_text8/text8.zip
!unzip text8.zip
# read in the extracted text file
with open('text8') as f:
text = f.read()
# print out the first 100 characters
print(text[:100])
```
## Pre-processing
Here I'm fixing up the text to make training easier. This comes from the `utils.py` file. The `preprocess` function does a few things:
>* It converts any punctuation into tokens, so a period is changed to ` <PERIOD> `. In this data set, there aren't any periods, but it will help in other NLP problems.
* It removes all words that show up five or *fewer* times in the dataset. This will greatly reduce issues due to noise in the data and improve the quality of the vector representations.
* It returns a list of words in the text.
This may take a few seconds to run, since our text file is quite large. If you want to write your own functions for this stuff, go for it!
```
import utils
# get list of words
words = utils.preprocess(text)
print(words[:30])
# print some stats about this word data
print("Total words in text: {}".format(len(words)))
print("Unique words: {}".format(len(set(words)))) # `set` removes any duplicate words
```
### Dictionaries
Next, I'm creating two dictionaries to convert words to integers and back again (integers to words). This is again done with a function in the `utils.py` file. `create_lookup_tables` takes in a list of words in a text and returns two dictionaries.
>* The integers are assigned in descending frequency order, so the most frequent word ("the") is given the integer 0 and the next most frequent is 1, and so on.
Once we have our dictionaries, the words are converted to integers and stored in the list `int_words`.
```
vocab_to_int, int_to_vocab = utils.create_lookup_tables(words)
int_words = [vocab_to_int[word] for word in words]
print(int_words[:30])
```
## Subsampling
Words that show up often such as "the", "of", and "for" don't provide much context to the nearby words. If we discard some of them, we can remove some of the noise from our data and in return get faster training and better representations. This process is called subsampling by Mikolov. For each word $w_i$ in the training set, we'll discard it with probability given by
$$ P(w_i) = 1 - \sqrt{\frac{t}{f(w_i)}} $$
where $t$ is a threshold parameter and $f(w_i)$ is the frequency of word $w_i$ in the total dataset.
> Implement subsampling for the words in `int_words`. That is, go through `int_words` and discard each word given the probablility $P(w_i)$ shown above. Note that $P(w_i)$ is the probability that a word is discarded. Assign the subsampled data to `train_words`.
```
from collections import Counter
import random
import numpy as np
threshold = 1e-5
word_counts = Counter(int_words)
#print(list(word_counts.items())[0]) # dictionary of int_words, how many times they appear
total_count = len(int_words)
freqs = {word: count/total_count for word, count in word_counts.items()}
p_drop = {word: 1 - np.sqrt(threshold/freqs[word]) for word in word_counts}
# discard some frequent words, according to the subsampling equation
# create a new list of words for training
train_words = [word for word in int_words if random.random() < (1 - p_drop[word])]
print(train_words[:30])
```
## Making batches
Now that our data is in good shape, we need to get it into the proper form to pass it into our network. With the skip-gram architecture, for each word in the text, we want to define a surrounding _context_ and grab all the words in a window around that word, with size $C$.
From [Mikolov et al.](https://arxiv.org/pdf/1301.3781.pdf):
"Since the more distant words are usually less related to the current word than those close to it, we give less weight to the distant words by sampling less from those words in our training examples... If we choose $C = 5$, for each training word we will select randomly a number $R$ in range $[ 1: C ]$, and then use $R$ words from history and $R$ words from the future of the current word as correct labels."
> **Exercise:** Implement a function `get_target` that receives a list of words, an index, and a window size, then returns a list of words in the window around the index. Make sure to use the algorithm described above, where you chose a random number of words to from the window.
Say, we have an input and we're interested in the idx=2 token, `741`:
```
[5233, 58, 741, 10571, 27349, 0, 15067, 58112, 3580, 58, 10712]
```
For `R=2`, `get_target` should return a list of four values:
```
[5233, 58, 10571, 27349]
```
```
def get_target(words, idx, window_size=5):
''' Get a list of words in a window around an index. '''
R = np.random.randint(1, window_size+1)
start = idx - R if (idx - R) > 0 else 0
stop = idx + R
target_words = words[start:idx] + words[idx+1:stop+1]
return list(target_words)
# test your code!
# run this cell multiple times to check for random window selection
int_text = [i for i in range(10)]
print('Input: ', int_text)
idx=5 # word index of interest
target = get_target(int_text, idx=idx, window_size=5)
print('Target: ', target) # you should get some indices around the idx
```
### Generating Batches
Here's a generator function that returns batches of input and target data for our model, using the `get_target` function from above. The idea is that it grabs `batch_size` words from a words list. Then for each of those batches, it gets the target words in a window.
```
def get_batches(words, batch_size, window_size=5):
''' Create a generator of word batches as a tuple (inputs, targets) '''
n_batches = len(words)//batch_size
# only full batches
words = words[:n_batches*batch_size]
for idx in range(0, len(words), batch_size):
x, y = [], []
batch = words[idx:idx+batch_size]
for ii in range(len(batch)):
batch_x = batch[ii]
batch_y = get_target(batch, ii, window_size)
y.extend(batch_y)
x.extend([batch_x]*len(batch_y))
yield x, y
int_text = [i for i in range(20)]
x,y = next(get_batches(int_text, batch_size=4, window_size=5))
print('x\n', x)
print('y\n', y)
```
---
## Validation
Here, I'm creating a function that will help us observe our model as it learns. We're going to choose a few common words and few uncommon words. Then, we'll print out the closest words to them using the cosine similarity:
<img src="assets/two_vectors.png" width=30%>
$$
\mathrm{similarity} = \cos(\theta) = \frac{\vec{a} \cdot \vec{b}}{|\vec{a}||\vec{b}|}
$$
We can encode the validation words as vectors $\vec{a}$ using the embedding table, then calculate the similarity with each word vector $\vec{b}$ in the embedding table. With the similarities, we can print out the validation words and words in our embedding table semantically similar to those words. It's a nice way to check that our embedding table is grouping together words with similar semantic meanings.
```
def cosine_similarity(embedding, valid_size=16, valid_window=100, device='cpu'):
""" Returns the cosine similarity of validation words with words in the embedding matrix.
Here, embedding should be a PyTorch embedding module.
"""
# Here we're calculating the cosine similarity between some random words and
# our embedding vectors. With the similarities, we can look at what words are
# close to our random words.
# sim = (a . b) / |a||b|
embed_vectors = embedding.weight
# magnitude of embedding vectors, |b|
magnitudes = embed_vectors.pow(2).sum(dim=1).sqrt().unsqueeze(0)
# pick N words from our ranges (0,window) and (1000,1000+window). lower id implies more frequent
valid_examples = np.array(random.sample(range(valid_window), valid_size//2))
valid_examples = np.append(valid_examples,
random.sample(range(1000,1000+valid_window), valid_size//2))
valid_examples = torch.LongTensor(valid_examples).to(device)
valid_vectors = embedding(valid_examples)
similarities = torch.mm(valid_vectors, embed_vectors.t())/magnitudes
return valid_examples, similarities
```
---
# SkipGram model
Define and train the SkipGram model.
> You'll need to define an [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) and a final, softmax output layer.
An Embedding layer takes in a number of inputs, importantly:
* **num_embeddings** – the size of the dictionary of embeddings, or how many rows you'll want in the embedding weight matrix
* **embedding_dim** – the size of each embedding vector; the embedding dimension
Below is an approximate diagram of the general structure of our network.
<img src="assets/skip_gram_arch.png" width=60%>
>* The input words are passed in as batches of input word tokens.
* This will go into a hidden layer of linear units (our embedding layer).
* Then, finally into a softmax output layer.
We'll use the softmax layer to make a prediction about the context words by sampling, as usual.
---
## Negative Sampling
For every example we give the network, we train it using the output from the softmax layer. That means for each input, we're making very small changes to millions of weights even though we only have one true example. This makes training the network very inefficient. We can approximate the loss from the softmax layer by only updating a small subset of all the weights at once. We'll update the weights for the correct example, but only a small number of incorrect, or noise, examples. This is called ["negative sampling"](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf).
There are two modifications we need to make. First, since we're not taking the softmax output over all the words, we're really only concerned with one output word at a time. Similar to how we use an embedding table to map the input word to the hidden layer, we can now use another embedding table to map the hidden layer to the output word. Now we have two embedding layers, one for input words and one for output words. Secondly, we use a modified loss function where we only care about the true example and a small subset of noise examples.
$$
- \large \log{\sigma\left(u_{w_O}\hspace{0.001em}^\top v_{w_I}\right)} -
\sum_i^N \mathbb{E}_{w_i \sim P_n(w)}\log{\sigma\left(-u_{w_i}\hspace{0.001em}^\top v_{w_I}\right)}
$$
This is a little complicated so I'll go through it bit by bit. $u_{w_O}\hspace{0.001em}^\top$ is the embedding vector for our "output" target word (transposed, that's the $^\top$ symbol) and $v_{w_I}$ is the embedding vector for the "input" word. Then the first term
$$\large \log{\sigma\left(u_{w_O}\hspace{0.001em}^\top v_{w_I}\right)}$$
says we take the log-sigmoid of the inner product of the output word vector and the input word vector. Now the second term, let's first look at
$$\large \sum_i^N \mathbb{E}_{w_i \sim P_n(w)}$$
This means we're going to take a sum over words $w_i$ drawn from a noise distribution $w_i \sim P_n(w)$. The noise distribution is basically our vocabulary of words that aren't in the context of our input word. In effect, we can randomly sample words from our vocabulary to get these words. $P_n(w)$ is an arbitrary probability distribution though, which means we get to decide how to weight the words that we're sampling. This could be a uniform distribution, where we sample all words with equal probability. Or it could be according to the frequency that each word shows up in our text corpus, the unigram distribution $U(w)$. The authors found the best distribution to be $U(w)^{3/4}$, empirically.
Finally, in
$$\large \log{\sigma\left(-u_{w_i}\hspace{0.001em}^\top v_{w_I}\right)},$$
we take the log-sigmoid of the negated inner product of a noise vector with the input vector.
<img src="assets/neg_sampling_loss.png" width=50%>
To give you an intuition for what we're doing here, remember that the sigmoid function returns a probability between 0 and 1. The first term in the loss pushes the probability that our network will predict the correct word $w_O$ towards 1. In the second term, since we are negating the sigmoid input, we're pushing the probabilities of the noise words towards 0.
```
import torch
from torch import nn
import torch.optim as optim
class SkipGramNeg(nn.Module):
def __init__(self, n_vocab, n_embed, noise_dist=None):
super().__init__()
self.n_vocab = n_vocab
self.n_embed = n_embed
self.noise_dist = noise_dist
# define embedding layers for input and output words
self.in_embed = nn.Embedding(n_vocab, n_embed)
self.out_embed = nn.Embedding(n_vocab, n_embed)
# Initialize embedding tables with uniform distribution
# I believe this helps with convergence
self.in_embed.weight.data.uniform_(-1, 1)
self.out_embed.weight.data.uniform_(-1, 1)
def forward_input(self, input_words):
input_vectors = self.in_embed(input_words)
return input_vectors
def forward_output(self, output_words):
output_vectors = self.out_embed(output_words)
return output_vectors
def forward_noise(self, batch_size, n_samples):
""" Generate noise vectors with shape (batch_size, n_samples, n_embed)"""
if self.noise_dist is None:
# Sample words uniformly
noise_dist = torch.ones(self.n_vocab)
else:
noise_dist = self.noise_dist
# Sample words from our noise distribution
noise_words = torch.multinomial(noise_dist,
batch_size * n_samples,
replacement=True)
device = "cuda" if model.out_embed.weight.is_cuda else "cpu"
noise_words = noise_words.to(device)
noise_vectors = self.out_embed(noise_words).view(batch_size, n_samples, self.n_embed)
return noise_vectors
class NegativeSamplingLoss(nn.Module):
def __init__(self):
super().__init__()
def forward(self, input_vectors, output_vectors, noise_vectors):
batch_size, embed_size = input_vectors.shape
# Input vectors should be a batch of column vectors
input_vectors = input_vectors.view(batch_size, embed_size, 1)
# Output vectors should be a batch of row vectors
output_vectors = output_vectors.view(batch_size, 1, embed_size)
# bmm = batch matrix multiplication
# correct log-sigmoid loss
out_loss = torch.bmm(output_vectors, input_vectors).sigmoid().log()
out_loss = out_loss.squeeze()
# incorrect log-sigmoid loss
noise_loss = torch.bmm(noise_vectors.neg(), input_vectors).sigmoid().log()
noise_loss = noise_loss.squeeze().sum(1) # sum the losses over the sample of noise vectors
# negate and sum correct and noisy log-sigmoid losses
# return average batch loss
return -(out_loss + noise_loss).mean()
```
### Training
Below is our training loop, and I recommend that you train on GPU, if available.
```
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# Get our noise distribution
# Using word frequencies calculated earlier in the notebook
word_freqs = np.array(sorted(freqs.values(), reverse=True))
unigram_dist = word_freqs/word_freqs.sum()
noise_dist = torch.from_numpy(unigram_dist**(0.75)/np.sum(unigram_dist**(0.75)))
# instantiating the model
embedding_dim = 300
model = SkipGramNeg(len(vocab_to_int), embedding_dim, noise_dist=noise_dist).to(device)
# using the loss that we defined
criterion = NegativeSamplingLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
print_every = 1500
steps = 0
epochs = 5
# train for some number of epochs
for e in range(epochs):
# get our input, target batches
for input_words, target_words in get_batches(train_words, 512):
steps += 1
inputs, targets = torch.LongTensor(input_words), torch.LongTensor(target_words)
inputs, targets = inputs.to(device), targets.to(device)
# input, outpt, and noise vectors
input_vectors = model.forward_input(inputs)
output_vectors = model.forward_output(targets)
noise_vectors = model.forward_noise(inputs.shape[0], 5)
# negative sampling loss
loss = criterion(input_vectors, output_vectors, noise_vectors)
optimizer.zero_grad()
loss.backward()
optimizer.step()
# loss stats
if steps % print_every == 0:
print("Epoch: {}/{}".format(e+1, epochs))
print("Loss: ", loss.item()) # avg batch loss at this point in training
valid_examples, valid_similarities = cosine_similarity(model.in_embed, device=device)
_, closest_idxs = valid_similarities.topk(6)
valid_examples, closest_idxs = valid_examples.to('cpu'), closest_idxs.to('cpu')
for ii, valid_idx in enumerate(valid_examples):
closest_words = [int_to_vocab[idx.item()] for idx in closest_idxs[ii]][1:]
print(int_to_vocab[valid_idx.item()] + " | " + ', '.join(closest_words))
print("...\n")
```
## Visualizing the word vectors
Below we'll use T-SNE to visualize how our high-dimensional word vectors cluster together. T-SNE is used to project these vectors into two dimensions while preserving local stucture. Check out [this post from Christopher Olah](http://colah.github.io/posts/2014-10-Visualizing-MNIST/) to learn more about T-SNE and other ways to visualize high-dimensional data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
# getting embeddings from the embedding layer of our model, by name
embeddings = model.in_embed.weight.to('cpu').data.numpy()
viz_words = 380
tsne = TSNE()
embed_tsne = tsne.fit_transform(embeddings[:viz_words, :])
fig, ax = plt.subplots(figsize=(16, 16))
for idx in range(viz_words):
plt.scatter(*embed_tsne[idx, :], color='steelblue')
plt.annotate(int_to_vocab[idx], (embed_tsne[idx, 0], embed_tsne[idx, 1]), alpha=0.7)
```
| github_jupyter |
#### 6. 로지스틱 회귀분석(Logistic Regression)
- 회귀분석 문제와 분류문제 모두 사용가능
- 로지스틱 회귀분석 모형
- 종속변수 : 이항분포를 따르고 모수$\mu$는 독립변수 $x$에 의존한다고 가정
- $p(y|x) = Bin(y; \mu(x), N)$
- $y$의 값이 특정한 구간내의 값( 0∼N )만 가질 수 있기 때문에 종속변수가 이러한 특성을 가진 경우 회귀분석 방법으로 사용 가능
- 이항 분포의 특별한 경우( N=1 )로 $y$ 가 베르누이 확률분포인 경우
- $p(y|x) = Bern(y; \mu(x))$
- y는 0 또는 1인 분류 예측 문제를 풀때 사용
```
%matplotlib inline
from matplotlib import rc
plt.style.use('seaborn')
rc('font', family='NanumGothic')
plt.rcParams['axes.unicode_minus'] = False
```
##### 시그모이드 함수
```
xx = np.linspace(-5, 5, 1000)
plt.plot(xx, 1/(1+np.exp(-xx)), 'r-', label="로지스틱함수")
plt.plot(xx, sp.special.erf(0.5*np.sqrt(np.pi)*xx), 'g:', label="오차함수")
plt.plot(xx, np.tanh(xx), 'b--', label="하이퍼볼릭탄젠트함수")
plt.ylim([-1.1, 1.1])
plt.legend(loc=2)
plt.xlabel("x")
plt.show()
# 1차원 독립변수를 가지는 분류문제
from sklearn.datasets import make_classification
X0, y = make_classification(n_features=1, n_redundant=0, n_informative=1,
n_clusters_per_class=1, random_state=4)
plt.scatter(X0, y, c=y, s=100, edgecolor="k", linewidth=2)
sns.distplot(X0[y == 0, :], label="y = 0", hist=False)
sns.distplot(X0[y == 1, :], label="y = 1", hist=False)
plt.ylim(-0.2, 1.2)
plt.show()
```
##### 로지스틱 분석
- logistics -> LL값을 최대화 하는 값 찾는것
- Logit 클래스 -> 베르누이 분포를 따르는 로지스틱 회귀 모형
- OLS 클래스와 사용법은 동일
- 독립변수와 종속변수 데이터를 넣어 모형을 만들고 fit으로 학습
- disp = 0 : 최적화 과정에서 문자열 메세지가 나타나지 않게 함
```
X = sm.add_constant(X0)
logit_mod = sm.Logit(y, X)
logit_res = logit_mod.fit(disp=0)
print(logit_res.summary())
# LL = 가장 크게 하고자 함 -> -16
# 찾고자 하는것 -> w -> coef
```
판별함수식
- $w_0 = 0.2515, w_1 = 4.2382$
- $\mu(x) = \sigma(4.2382x + 0.2515)$
- z값의 부호를 나누는 기준값 = -0.2515/4.2382
- 유의확률을 감안 했을 때 $w_0 = 0$이라 볼수 있음
```
xx = np.linspace(-3, 3, 100)
mu = logit_res.predict(sm.add_constant(xx))
plt.plot(xx, mu, lw=3)
plt.scatter(X0, y, c=y, s=100, edgecolor="k", lw=2)
plt.scatter(X0, logit_res.predict(X), label=r"$\hat{y}$", marker='s', c=y,
s=100, edgecolor="k", lw=1)
plt.xlim(-3, 3)
plt.xlabel("x")
plt.ylabel(r"$\mu$")
plt.title(r"$\hat{y} = \mu(x)$")
plt.legend()
plt.show()
```
- Logit 모형의 결과 객체에는 fittedvalues 속성
- 판별함수 $z=w^Tx$ 값이 들어가 있다. 이 값을 이용하여 분류문제를 풀 수도 있다.
```
plt.scatter(X0, y, c=y, s=100, edgecolor="k", lw=2, label="데이터")
plt.plot(X0, logit_res.fittedvalues * 0.1, label="판별함수값")
plt.legend()
plt.show()
# 이탈도
from sklearn.metrics import log_loss
y_hat = logit_res.predict(X)
# log_loss의 normalize=False -> log_loss값
log_loss(y, y_hat, normalize=False)
# 귀무 모형
mu_null = np.sum(y) / len(y)
mu_null
# LL-Null
y_null = np.ones_like(y) * mu_null
log_loss(y, y_null, normalize=False)
# Pseudo R-squ
1 - (log_loss(y, y_hat) / log_loss(y, y_null))
```
##### Scikit-Learn 패키지의 로지스틱 회귀
```
from sklearn.linear_model import LogisticRegression
model_sk = LogisticRegression().fit(X0, y)
xx = np.linspace(-3, 3, 100)
mu = 1.0/(1 + np.exp(-model_sk.coef_[0][0]*xx - model_sk.intercept_[0]))
plt.plot(xx, mu)
plt.scatter(X0, y, c=y, s=100, edgecolor="k", lw=2)
plt.scatter(X0, model_sk.predict(X0), label=r"$\hat{y}$", marker='s', c=y,
s=100, edgecolor="k", lw=1, alpha=0.5)
plt.xlim(-3, 3)
plt.xlabel("x")
plt.ylabel(r"$\mu$")
plt.title(r"$\hat{y}$ = sign $\mu(x)$")
plt.legend()
plt.show()
```
##### 연습문제
```
from sklearn.datasets import load_iris
iris = load_iris()
# in1d : 0하고 1인 것만 뽑는 것
idx = np.in1d(iris.target, [0, 1])
X0 = iris.data[idx, :1]
X = sm.add_constant(X0)
y = iris.target[idx]
logit_mod = sm.Logit(y, X)
logit_res = logit_mod.fit(disp=0)
print(logit_res.summary())
# w값
logit_res.params
# z 부호를 나누는 기준 값
-logit_res.params[0] / logit_res.params[1]
y_pred = logit_res.predict(X) >= 0.5
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y, logit_res.fittedvalues)
fpr, tpr, thresholds
plt.plot(fpr, tpr, 'o-', label = "Logistic Regression")
plt.plot([0, 1], [0, 1], 'k--')
plt.xlabel("Fall-Out")
plt.ylabel("Recall")
plt.show()
data_med = sm.datasets.get_rdataset("MedGPA", package="Stat2Data")
df_med = data_med.data
df_med.tail()
sns.stripplot(x="GPA", y="Acceptance", data=df_med,
jitter=True, orient='h', order=[1, 0])
plt.grid(True)
plt.show()
# MCAT = VR + PS + WS + BS이므로 이 MCAT은 독립 변수에서 제외
model_med = sm.Logit.from_formula("Acceptance ~ Sex + BCPM + GPA + VR + PS + WS + BS + Apps", df_med)
result_med = model_med.fit()
print(result_med.summary())
df_med["Prediction"] = result_med.predict(df_med)
sns.boxplot(x="Acceptance", y="Prediction", data=df_med)
plt.show()
model_med = sm.Logit.from_formula("Acceptance ~ PS + BS", df_med)
result_med = model_med.fit()
print(result_med.summary())
# 0.4798PS+1.1464BS > 15.5427 합격이라고 예측가능
```
##### 연습문제
- 붓꽃데이터 독립변수가 여러개인 경우
```
from sklearn.datasets import load_iris
iris = load_iris()
# in1d : 0하고 1인 것만 뽑는 것
idx = np.in1d(iris.target, [1, 2])
X0 = pd.DataFrame(iris.data[idx, :], columns=iris.feature_names[:])
X = sm.add_constant(X0)
y = iris.target[idx] - 1
logit_mod = sm.Logit(y, X)
logit_res = logit_mod.fit(disp=0)
print(logit_res.summary())
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y, logit_res.fittedvalues)
fpr, tpr, thresholds
plt.plot(fpr, tpr, 'o-', label = "Logistic Regression")
plt.plot([0, 1], [0, 1], 'k--')
plt.xlabel("Fall-Out")
plt.ylabel("Recall")
plt.show()
```
##### 로지스틱회귀를 사용한 회귀분석
```
data_wrole = sm.datasets.get_rdataset("womensrole", package="HSAUR")
df_wrole = data_wrole.data
df_wrole["ratio"] = df_wrole.agree / (df_wrole.agree + df_wrole.disagree)
df_wrole.tail()
sns.scatterplot(x="education", y="ratio", style="sex", data=df_wrole)
plt.grid(True)
plt.show()
model_wrole = sm.Logit.from_formula("ratio ~ education + sex", df_wrole)
result_wrole = model_wrole.fit()
print(result_wrole.summary())
model_wrole2 = sm.Logit.from_formula("ratio ~ education", df_wrole)
result_wrole2 = model_wrole2.fit()
print(result_wrole2.summary())
sns.scatterplot(x="education", y="ratio", data=df_wrole)
xx = np.linspace(0, 20, 100)
df_wrole_p = pd.DataFrame({"education": xx})
plt.plot(xx, result_wrole2.predict(df_wrole_p), "r-", lw=4, label="예측")
plt.legend()
plt.show()
```
| github_jupyter |
# Homework Batch 3: Routing Algorithms
### Marco Sicklinger, May 2021
## Modules
In the below modules, except for random obviously, one can find the implementation of the algorithm required.
```
from graph import *
from heap import *
from dijkstra import *
from random import random, randint
```
## Binheap version of Dijkstra Algorithm
First, a graph must be created, as an object of class `WeightGraph`: it is done by creating a dictionary containing the vertices as values (vertices ar of class `Node`), while the choice of the keys does not have any influence on the successive steps. However, they must match the keys of the other dictionary to give as argument to the `WeightedGraph` class, the adjacency list. This is a dictionary, whose keys must match the keys of the previous dictionary, so one can assign to every node the corresponding correct adjacency list. The values of this latter dictionary are lists of lists, that is lists containing pairs of a key (representing the vertice in the adjacency list) and a weight (representing the weight of the edge). These pairs are not stored as tuples since the latter ones are immutable objects, so it has been chosen to use mutable objects as lists in case the user needs to modify one of the elements.
When the `WeightedGraph` object is initialized the adjacency lists given by the user is assigned to each vertice as a `Node` class attribute `adj_list`, adding to the front of each pair of key and weight another element, by default `None`, which is going to represent the ignored vertice in a shortcut, if any exists.
There is no need or necessity to initialize the vertices' attribute `predecessor`, `adj_list`, `ancestors`, `heap_index` and `shortcuts` since they are computed on the basis of what the user passes as arguments to `WeightedGraph`. The only attribute that are initializable by passing arguments to `Node` are `value`, `distance` and `importance`.
In this first example below, where the binheap version of the *Dijkstra Algorithm* has been tested, importances and values are given randomly, for the sake of simplicity.
```
# create graph
g = {}
# assign importances
importance_array = list(np.random.permutation(5))
for i in range(5):
# assign to nodes their values
g[i] = Node(value=randint(0,100), importance=importance_array[i])
# create adjacency lists
d = {}
for i in range(5):
d[i] = []
# assign adjacent nodes and weights of corresponding edges
for j in range(randint(0,4),5):
d[i].append([j, random()])
# create dictionary
graph = WeightedGraph(g, d)
# printing graph
print(graph)
# applying dijkstra algorithm to graph
dijkstra(graph, graph.Keys[2])
# printing result
print(graph)
```
## Shortcuts
To build shortcuts in a graph, one must call the function `build_shortcuts`, passing the graph as argument. The importance member of each vertice must obviously be initialized, that is must be different from `None` if one wants the function to work.
```
# build shortcuts in the graph
build_shortcuts(graph)
# printing shortcuts
for key in graph.Keys:
print('key: ', key, ' importance:', graph.Dictionary[key].importance)
print('shortcuts: ', graph.Dictionary[key].shortcuts, '\n')
# update graph with the shortcuts
update_graph(graph)
# print updated graph
print(graph)
```
## Bidirectional version of Dijkstra Algorithm
In the first test of this section, importance values are randomly initialized, for the sake of simplicity.
The final returned result is a tuple containing the path from start to end and the total distance taken.
```
# create graph
g = {}
# assign importances
importance_array = list(np.random.permutation(7))
for i in range(7):
# assign to nodes their values
g[i] = Node(value=randint(0,100), importance=importance_array[i])
# create adjacency lists
d = {}
for i in range(7):
d[i] = []
# assign adjacent nodes and weights of corresponding edges
for j in range(randint(2,5),7):
d[i].append([j, random()])
# create dictionary
graph = WeightedGraph(g, d)
# printing graph
print(graph)
# apply dijkstra algorithm to graph
result = bi_dijkstra(graph, 1, 4)
print(result)
# update predecessors
update_predecessors(graph, result[0])
# print graph
print(graph)
```
Following below, there is another test of the *Bidirectional Dijkstra Algorithm*, this time using importance values related to the number of links (incoming and outgoing edges) that a vartice has.
```
# create graph
g = {}
for i in range(7):
# assign to nodes their values
g[i] = Node(value=randint(0,100))
# create adjacency lists
d = {}
for i in range(7):
d[i] = []
# assign adjacent nodes and weights of corresponding edges
for j in range(randint(2,5),7):
d[i].append([j, random()])
# create dictionary
graph = WeightedGraph(g, d)
# printing graph
print(graph)
# compute ancestors
graph.Ancestors()
# assign importances
for key in graph.Keys:
# assign importance by counting number of 'links' of the vertice
importance = len(graph.Dictionary[key].ancestors) + len(graph.Dictionary[key].adj_list)
graph.Dictionary[key].importance = importance
# print graph
print(graph)
# apply dijkstra algorithm to graph
result = bi_dijkstra(graph, 1, 4)
print(result)
# update predecessors
update_predecessors(graph, result[0])
# print graph
print(graph)
```
| github_jupyter |
```
from collections import defaultdict, deque
class Solution:
def minJumps(self, arr: List[int]) -> int:
n = len(arr)
if len(set(arr)) == n: # 如果元素都不相同,那就只能一个一个向右跳
return n - 1
num_freq = defaultdict(list)
for i, num in enumerate(arr):
num_freq[num].append(i)
dq = deque([[0, 0]])
seen = {0}
while dq:
for _ in range(len(dq)):
idx, cnt = dq.popleft()
if idx == n-1:
return cnt
# 左右跳
for n_idx in (idx+1, idx-1):
if 0 <= n_idx < n and n_idx not in seen:
dq.append((n_idx, cnt + 1))
seen.add(n_idx)
# 调到相同的数字上
if arr[idx] in num_freq:
for n_idx in num_freq[arr[idx]]:
if n_idx == idx or n_idx in seen:
continue
seen.add(n_idx)
dq.append((n_idx, cnt + 1))
del num_freq[arr[idx]]
return -1
solution = Solution()
solution.minJumps(arr = [11,22,7,7,7,7,7,7,7,22,13])
from collections import defaultdict, deque
class Solution:
def minJumps(self, arr):
if set(arr) == len(arr):
return len(arr) - 1
if len(arr) == 1:
return 0
num_freq = defaultdict(list)
for i, n in enumerate(arr):
num_freq[n].append(i)
n = len(arr)
dq = deque([[0, 0]]) # 从第零号下标开始
best = {0: 0}
while dq:
for _ in range(len(dq)):
idx, cnt = dq.popleft()
if idx == n-1:
return cnt
# 向左、右跳
for n_idx in (idx - 1, idx + 1):
if n_idx < 0 or n_idx >= n:
continue
if n_idx not in best or best[n_idx] < cnt + 1:
best[n_idx] = cnt + 1
dq.append((n_idx, cnt+1))
# 相同的数字,不同的索引
for n_idx in num_freq[arr[idx]]:
if n_idx == idx:
continue
if n_idx not in best or best[n_idx] < cnt + 1:
best[n_idx] = cnt + 1
dq.append((n_idx, cnt+1))
del num_freq[arr[idx]]
return -1
```
| github_jupyter |
# Double-slit correlation model
Based on Double Slit Model notebook, extended to model correlations with phase variation
```
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as sp
from numpy import pi, sin, cos, linspace, exp, real, imag, abs, conj, meshgrid, log, log10, angle, zeros, complex128, random
from numpy.fft import fft, fftshift, ifft
from mpl_toolkits.mplot3d import axes3d
import BeamOptics as bopt
%matplotlib inline
b=.08*1e-3 # the slit width
a=.5*1e-3 # the slit spacing
k=2*pi/(795*1e-9) # longitudinal wavenumber
wt=0 # let time be zero
C=1 # unit amplitude
L=1.8 # distance from slits to CCD
d=.016 # distance from signal to LO at upstream end (used to calculate k_perp)
ccdwidth = 1300 # number of pixels
pixwidth = 20e-6 # pixel width (in meters)
y = linspace(-pixwidth*ccdwidth/2,pixwidth*ccdwidth/2,ccdwidth)
# define the various double slit fields and LO:
def alpha(y,a):
return k*a*y/(2*L)
def beta(y,b):
return k*b*y/(2*L)
def E_ds(y,a,b):
""" Double-slit field """
# From Hecht p 458:
#return b*C*(sin(beta(y)) / beta(y)) * (sin(wt-k*L) + sin(wt-k*L+2*alpha(y)))
# drop the time-dep term as it will average away:
return 2*b*C*(sin(beta(y,b)) / beta(y,b)) * cos(alpha(y,a)) #* sin(wt - k*L + alpha(y))
def E_dg(y,a,b):
""" Double gaussian field """
# The width needs to be small enough to see interference
# otherwise the beam doesn't diffract and shows no interference.
# We're using b for the gaussian width (i.e. equal to the slit width)
w=b
#return C*exp(1j*k*0.1*d*y/L)
return 5e-3*(bopt.gaussian_beam(0,y-a/2,L,E0=1,wavelambda=795e-9,w0=w,k=[0,0,k]) +
bopt.gaussian_beam(0,y+a/2,L,E0=1,wavelambda=795e-9,w0=w,k=[0,0,k]))
def E_lo(y,d):
"""Plane-wave LO beam incident at small angle, transverse wavenumber k*d*y/L"""
return C*exp(-1j*k*d*y/L)
```
## Define a single function to explore the FFT:
```
def plotFFT(d,a,b):
"""Single function version of generating the FFT output"""
TotalField = E_dg(y,a,b)+E_lo(y,d)
TotalIntensity=TotalField*TotalField.conj()
plt.plot(abs(fft(TotalIntensity)),".-")
plt.ylim([0,1e-2])
plt.xlim([0,650])
plt.title("FFT output")
plotFFT(d=0.046,a=0.5e-3,b=0.08e-3)
```
### Replace with Gaussian LO: import gaussian beam function, and repeat:
```
# bopt.gaussian_beam(x, y, z, E0, wavelambda, w0, k)
# set to evaluate gaussian at L (full distance to CCD) with waist width of 2 cm
# using d=0.046 for agreement with experiment
d=0.046
E_lo_gauss = bopt.gaussian_beam(0,y,L,E0=1,wavelambda=795e-9,w0=0.02,k=[0,k*d/L,k])
frames = 59
rounds = 20
drift_type= 3
# SG I made a few drift modes to model the phase drift that would be present in the lab
# drift mode two appears to be the most similar to the phase shifts we observe in the lab
time=linspace(0,2*pi,rounds*frames)
phase=[]
if drift_type == 0:
phase= [sin(t) for t in time]
#mode 0 is just a sine wave in time
elif drift_type == 1:
phase= [sin(t+random.randn()/2) for t in time]
#phase= [sin(t)+random.randn()/2 for t in time]
#mode 1 is a sine wave with some randomness added to each data point
elif drift_type == 2:
phase=[0]
for i in range(len(time)-1):
phase.append(phase[-1]+random.randn()/4*sin(time[i]))
#mode 2 is a sine wave with some randomness added to each data point, and also considering
#the location of the previous data point
elif drift_type == 3:
phase=[0]
for i in range(len(time)-1):
phase.append(phase[-1]+0.1*(random.randn()))
#mode 2 is a sine wave with some randomness added to each data point, and also considering
#the location of the previous data point
raw_intensity_data = zeros([1300,frames,rounds],dtype=complex128)
scaled = zeros([1300,frames,rounds],dtype=complex128)
i=0
for r in range(rounds):
for f in range(frames):
TotalField = E_dg(y,a,b)*exp(-1j*phase[i]) + E_lo_gauss #adds the appropriate phase
#TotalField = E_dg(y,a,b) + E_lo_gauss
TotalIntensity = TotalField * TotalField.conj()
raw_intensity_data[:,f,r] = TotalIntensity
scaled[:,f,r]=fft(TotalIntensity)
i=i+1 #increases index
#checking how phase moves around
plt.polar(phase,time,'-')
plt.title("phase shift with (simulated) time")
plt.plot((np.unwrap(angle(scaled[461,:,:].flatten("F")))))
plt.plot((np.unwrap(angle(scaled[470,:,:].flatten("F")))))
#plt.ylim(0,1e-2)
#TODO -unwrapping the phase (numpy)
plt.plot(abs(fft(TotalIntensity)),".-")
print(TotalIntensity.shape)
plt.ylim([0,0.01]) # Had to lower the LO power quite a bit, and then zoom way in.
plt.xlim([430,500])
```
Adding different phase drifts to individual modes
original signal -> FFT ->
```
mode_of_interest = 440
mode_offset = 300
range_to_analyze = 300
# Calculate the correlation matrix between phase of each mode.
modes = range(0,range_to_analyze)
PearsonPhase = np.zeros((range_to_analyze,range_to_analyze))
for m in modes:
output = scaled[m+mode_offset,:,:].flatten('F') # Choose the mode to analyze
x = np.angle(output)
for l in modes:
#SG added np.unwrap call to the angle
Pearson, p = sp.pearsonr(np.unwrap(np.angle(scaled[l+mode_offset].flatten('F'))), x)
if (m==l):
PearsonPhase[m,l] = 0 #AMCD Null the 1.0 auto-correlation
else:
PearsonPhase[m,l] = Pearson
plt.imshow(PearsonPhase,interpolation='none')
plt.title("Phase")
print(type(PearsonPhase))
print("max value =",np.amax(PearsonPhase))
plt.imshow(PearsonPhase,interpolation='none')
plt.title("Phase")
print("max value =",np.amax(PearsonPhase))
```
| github_jupyter |
# API
```
import psycopg2
```
CREATE TABLE API
(key CHARACTER VARYING(100) NOT NULL PRIMARY KEY,
secret CHARACTER VARYING(100) NOT NULL,
type CHARACTER VARYING(10) NOT NULL,
start_use BOOLEAN NOT NULL DEFAULT(FALSE)
);
CREATE INDEX key_index ON API(key);
CREATE INDEX secret_index ON API(secret);
CREATE INDEX type_index ON API(type);
CREATE INDEX start_use_index ON API(start_use);
INSERT INTO API VALUES
('199ed59000c39dd0844b59d01fa7570c','4a2ce28f1bb8a1fe','flickr', FALSE),
('81ec25b2e0093c6f2c2e70da0175a7ea','6eaa89f23ae0762e','flickr', FALSE),
('4f3c045b20127210215889331a6ab134','2010f9a1cf1abe18','flickr', FALSE),
('382e669299b2ea33fa2288fd7180326a','b556d443c16be15e','flickr', FALSE),
('b422ff64b04ecee4c169ca01a21f5bcb','57d1f80344df6188','flickr', FALSE),
('042e20a01e7080ae8a7a4889208d215f','c2d8a1f6b9760cb9','flickr', FALSE),
('fc7b495434337ecb3ea080a3f410b0c6','8ed6d435785a2023','flickr', FALSE),
('696005e9ef76d9ef438c99f21605b322','9085b2dbbae8ede5','flickr', FALSE),
('7ce333ecbbf8ab34a6843e0edeeed7f4','c72121c17578668c','flickr', FALSE),
('2c8b19dfe58aded2e3a756bfba941558','9ef37b12d94a3da4','flickr', FALSE),
('3d3687711d33d98a847a34e094cd228d','27b2d210f4142b6b','flickr', FALSE),
('3f647b14450052251bbda3d4f8e3efd4','afa71fda070402ad','flickr', FALSE),
('60ff62a1aa6701d0bbd642663cf35d96','729cd4f7b7c92931','flickr', FALSE),
('75b8a444d318c769ae8142b1351a2b3c','a8723ba65b8ca827','flickr', FALSE),
('089c5cfa0ff527fdb25cd8b0a547e914','2dbdc4ce5d3e752b','flickr', FALSE),
('40e4f0df81f75752f74e36683808d7ab','f1a8144378ed93d4','flickr', FALSE),
('622f1f3e57c9f337d1212444831ff475','fafddabfbe0405b7','flickr', FALSE),
('1f7c3204baf7159d4ec9e833ee21d11f','90b9ed8a906812f3','flickr', FALSE);
INSERT INTO API VALUES
('DCTCyC_D_3ZGW4VZVkmj25IJNUuNdXT4','T9NzaLTeiSe_Rqtye0pUCs3Ed-ZQiq6V','Face++',False),
('Dm18HrboYUuAVeX6A-_6Y0q27OkE3cvN','3FPheINGB2qLTDcV-FOpjloXmOjDDG4A','Face++',False),
('AB3ubqE6FNZGPxu8NE4c1F0OJCn8HInI','VHH-kDCN1MQ_Ru2qq21YPyOwVF-e2Hmb','Face++',False),
('HGLXmBY-gO6WsCB4ZkFpEadYDD3SO4_S','oN8M-apFgr-4V4qW2W7TSuyUHy2Gw89_','Face++',False),
('Xgv1FMjf7zaGVfcR_hlpdyif3Ez0q1m5','ISIsT6DYelUG9iPYS3NEKJ1dTfyReSph','Face++',False),
('D8KfUEN_6PM1yCHveoKuxgZ0GT21X1Bh','_7vXMF7dsWNa7CIZiSOKs5JZp_XACHvf','Face++',False),
('Fu4-6MVuWgGCUeIxjSYRusH8eVilBtAz','epLYs0n7YLAWaYjA7HlXuLILHjKnL5iJ','Face++',False),
('mhFDxxTfQy6Rd4u298kwFvxbCxAMCpHq','yMZBadr5wVFsUWSZ_I0jo0w9vzjDVEii','Face++',False),
('1iPU-v_kLZs5OXjUZBlFL6ooTci8U3Yp','jTt_gIkc2bqnYGKucZboLuSQuUBzT2Vk','Face++',False),
('h_KYerXL2QeTmsAJGGBiPZFxl7_QXmiO','v1uoVifc12-1SYRsyNxgSR5k4QQqyK9u','Face++',False),
('sPG0n8LFaQfxxglBtbverq7iJgPs8DMQ','HBd-aaVTudtepT9bhybHKtJaLx7sMw1Y','Face++',False),
('hOLmeAvF4_gPFycCYZO4HoyJsFa7rlHc','gqAKxnvHvxpgt-mnHZMc7EwCD2XyHk0-','Face++',False),
('nBbJ3_x3xmnmc7CJO_VrK8vT8lmGlZIB','HB12_nEr3Y2I9rgKCtN0KKYX-LPA-sUU','Face++',False),
('_VTJAxAE34Xt-h236X1LVbO6gMJwszLN','f9roZWjSMH7csIVn4RHuGQ6ZDOwxhuCx','Face++',False),
('oSzQJ7Owxqk2T5yZuIHKoJ3s_n11BDQP','wv-sD-NgKWm0kRK757UxUDsbuzj0TYs ','Face++',False),
('NCv1iO90aVJ4Axy-FjE2SemVMP50Fg8e','xjKxi7n4qfO9h3N1uAThcMD01L6478YK','Face++',False),
('jFIzayRBM4X3-YJzzXBkZlzXy5MPe1Pl','spDfyMN7FnwnS5hbtnxFhxtuFmKeNPRs','Face++',False),
('TUWlFehF2kAEowOZRVyLa-0XzjoFKvUR','d5T4Lpx6DAFMTNRr6kfkcr6ZuQgfhosc','Face++',False),
('RsqKoZtF2Zcj1zKdn0i4-3yTCrtxeuVo','J2qGXnlTLcUqdS9CJ_6M-gt57vVohjy-','Face++',False),
('bbhAbpc7IrCGCWzrUu1bftZhR9iSkOh4','2jTiWxN2dCdTefw8ZbX0V8f_EOVzxbvk','Face++',False),
('8k8rid_MisVBXRSuAeiLVA5ymeT_8eYQ','gw2dA-e0fwv20icOC6OucBcRgc9yPi8k','Face++',False),
('ebfsStmOrxvn-kppu_-1AXrsYmrJSmB0','NqvtZaDgEfV4U5m2cn-7agaw5WEpLL2z','Face++',False),
('mDCcCZx7EqeQjRu2yv8byuCB_xWkFMyd','fn73kxqkkP0mesI-gcPBk0BUmrAmdzTe','Face++',False);
```
class CloudDatabase(object):
# Init database and input ip
# 初始化数据库并传入ip
def __init__(self, database, user, password, ip="127.0.0.1", port="5432"):
self.database = database
self.user = user
self.password = password
self.ip = ip
self.port = port
# Connect database and set input as host, return connection and cursor
# 连接ip端数据库并返回connection和cursor
def db_connect(self):
self.connection = psycopg2.connect(database=self.database, user=self.user,
password=self.password, host=self.ip, port=self.port)
self.cursor = self.connection.cursor()
def execute(self, sql):
try:
self.cursor.execute(sql)
self.connection.commit()
except Exception as e:
self.connection.rollback()
# Write log file
# 输出日志
def write_log(self, e):
self.connection.rollback()
with open("log.txt", 'a') as log_file:
log_file.writelines(str(e))
database = CloudDatabase("PlaceEmotion", "postgres", "postgres", "127.0.0.1")
database.db_connect()
# Test API
import requests
sql = '''
SELECT key, secret
FROM API
WHERE type = 'flickr'
'''
database.execute(sql)
for api in database.cursor.fetchall():
print(api[0], api[1])
```
# Location
CREATE TABLE location
(id SERIAL PRIMARY KEY,
city_name CHARACTER VARYING(100) NOT NULL,
lat FLOAT NOT NULL,
lon FLOAT NOT NULL,
start_query BOOLEAN NOT NULL DEFAULT(FALSE)
);
CREATE INDEX id_index ON location(id);
CREATE INDEX city_name_index ON location(city_name);
CREATE INDEX start_query_index ON location(start_query);
```
# Import all sampling points
file = open('/media/raid/PlaceEmotion/pts200_us.csv', 'r')
st = file.read()
file.close()
sql = '''
INSERT INTO location(city_name, lat, lon)
VALUES
{0}
'''.format(st)
#print(sql)
database.execute(sql)
```
CREATE TABLE photo
(id BIGINT PRIMARY KEY,
url TEXT NOT NULL,
city CHARACTER VARYING(100) NOT NULL,
face_number INTEGER DEFAULT(NULL),
start_detect BOOLEAN NOT NULL DEFAULT(FALSE),
start_info BOOLEAN NOT NULL DEFAULT(FALSE)
);
CREATE INDEX photo_id_index ON photo(id);
CREATE INDEX photo_start_info_index ON photo(start_info);
CREATE INDEX photo_start_detect_index ON photo(start_detect);
CREATE INDEX photo_face_number_index ON photo(face_number);
CREATE TABLE photo_info
(id BIGINT PRIMARY KEY,
owner CHARACTER VARYING (30),
owner_location CHARACTER VARYING (30),
lat FLOAT,
lon FLOAT,
photo_take_date DATE,
photo_upload BIGINT,
accuracy INTEGER,
geotag TEXT,
neighbourhood TEXT;
locality TEXT;
county TEXT;
region TEXT;
country TEXT;
);
CREATE INDEX photo_id_index ON photo(id);
CREATE INDEX photo_lat_index ON photo(lat);
CREATE INDEX photo_lon_index ON photo(lon);
CREATE INDEX photo_date_index ON photo(photo_take_date);
CREATE INDEX photo_f_hasface_index ON photo(f_hasface);
CREATE INDEX photo_start_detect_index ON photo(start_detect);
CREATE INDEX photo_start_info_index ON photo(start_info);
CREATE INDEX photo_start_recog_index ON photo(start_recog);
CREATE INDEX photo_facenum_index ON photo(facenum);
CREATE INDEX photo_lat_index ON photo(lat);
CREATE INDEX photo_lon_index ON photo(lon);
CREATE INDEX photo_date_index ON photo(photo_take_date);
CREATE INDEX photo_f_hasface_index ON photo(f_hasface);
CREATE INDEX photo_start_detect_index ON photo(start_detect);
Four layers framework:
1. API generation and location selection: return API and lat, lon, city.
2. collect photo url and id to photo table: input API and lat, lon, city, return photo id and url.
3. get photo information to photo_info table: input photo id, return photoinfo.
4. detect photo face info to face table. input photo id, return emotion info.
```
import datetime
DATE=datetime.date(2012,1,1)
while(True):
DATE2=DATE+datetime.timedelta(days=10)
datemin ="{0}-{1}-{2}".format(DATE.year,DATE.month,DATE.day)
datemax ="{0}-{1}-{2}".format(DATE2.year,DATE2.month,DATE2.day)
DATE=DATE+datetime.timedelta(days=10)
print(datemin,datemax)
#get_photo_from_location(db_connection, db_cursor, site, latitude, longitude, datemin, datemax)
if DATE.year==2018 and DATE.month==11:
break
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import cv2
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten
from tensorflow.keras.models import Model
from tensorflow.keras.applications import InceptionResNetV2 as Model
from tensorflow.keras.applications.inception_resnet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import MaxPool2D, Conv2D, Dropout
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from keras import Sequential
from keras.layers import Dense,Flatten
from sklearn.metrics import f1_score
seed = 0
from google.colab import drive
drive.mount('/content/drive')
! git clone https://github.com/Peter-TMK/Hamoye_capstone_project_smote.git
img = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Elephant/Elephant_111.jpg")
plt.imshow(img)
print(img.shape)
fig = plt.figure(figsize= (10,5))
img1 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Elephant/Elephant_111.jpg")
img2 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Buffalo/Buffalo_104.jpg")
img3 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Elephant/Elephant_101.jpg")
img4 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Rhino/Rhino_107.jpg")
img5 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Zebra/Zebra_104.jpg")
img6 = cv2.imread("/content/Hamoye_capstone_project_smote/Data/train/Rhino/Rhino_112.jpg")
fig.add_subplot(2,3,1)
plt.imshow(img1)
plt.axis('off')
print(img1.shape)
fig.add_subplot(2,3,2)
plt.imshow(img2)
plt.axis('off')
print(img2.shape)
fig.add_subplot(2,3,3)
plt.imshow(img3)
plt.axis('off')
print(img3.shape)
fig.add_subplot(2,3,4)
plt.imshow(img4)
plt.axis('off')
print(img4.shape)
fig.add_subplot(2,3,5)
plt.imshow(img5)
plt.axis('off')
print(img5.shape)
fig.add_subplot(2,3,6)
plt.imshow(img6)
plt.axis('off')
print(img6.shape)
IMAGE_SIZE = [224, 224]
train_path = "/content/Hamoye_capstone_project_smote/Data/train"
val_path = "/content/Hamoye_capstone_project_smote/Data/val"
test_path = "/content/Hamoye_capstone_project_smote/Data/test"
# Use the Image Data Generator to import the images from the dataset
train_datagen = ImageDataGenerator(preprocessing_function = preprocess_input)
test_datagen = ImageDataGenerator(preprocessing_function = preprocess_input)
training_gen = train_datagen.flow_from_directory(train_path,
target_size = IMAGE_SIZE,
batch_size = 32,
seed = seed,
shuffle=True,
class_mode = 'categorical')
validation_gen = test_datagen.flow_from_directory(val_path,
target_size = IMAGE_SIZE,
batch_size = 32,
seed = seed,
shuffle=True,
class_mode = 'categorical')
test_gen=test_datagen.flow_from_directory(test_path,
target_size= IMAGE_SIZE,
batch_size= 32,
seed = seed,
shuffle= False,
class_mode="categorical")
# checking for the classes of our label
training_gen.class_indices
# this represents our test actual values and their classes.
test_gen.classes
print(len(test_gen.classes))
model_ = Model(
include_top=False,
weights='imagenet',
input_tensor=None,
input_shape=(224,224,3),
pooling=None,
classes=4,
classifier_activation='softmax'
)
model_.trainable = False
np.random.seed(seed)
tf.random.set_seed(seed)
model = Sequential([
model_
])
model.add(Conv2D(512, kernel_size=(1,1), activation='relu'))
model.add(MaxPool2D((2,2), strides=2, padding='same'))
model.add(Dropout(0.5))
model.add(Conv2D(512, kernel_size=(1,1), activation='relu'))
model.add(MaxPool2D((2,2), strides=2, padding='same'))
model.add(Dropout(0.5))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.8))
model.add(Flatten())
model.add(Dense(4, activation='softmax'))
model.summary()
learning_rate = 0.0001
optimizer = tf.keras.optimizers.Adam(learning_rate = learning_rate)
model.compile(optimizer = optimizer,
loss = tf.keras.losses.CategoricalCrossentropy(),
metrics = ["accuracy"])
STEP_SIZE_TRAIN = training_gen.n // training_gen.batch_size
STEP_SIZE_VALID = validation_gen.n // validation_gen.batch_size
# Some callback functions for fine tuning the model
EarlyStop = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=3, verbose=1, min_delta=1e-4)
ReduceLROnPlateau = tf.keras.callbacks.ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=1, cooldown=0,
min_lr=1e-7, verbose=1)
Checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath='/content/drive/MyDrive/Colab Notebooks/hamoye/data/model.hdf5',
verbose=1, save_best_only=True, save_weights_only=True, mode='auto')
# took 31 minutes
history = model.fit(x = training_gen,
steps_per_epoch = STEP_SIZE_TRAIN,
validation_data = validation_gen,
validation_steps = STEP_SIZE_VALID,
epochs = 10, callbacks=[EarlyStop, ReduceLROnPlateau, Checkpoint])
history = history.history
n_epochs = len(history['loss'])
plt.figure(figsize=[14,4])
plt.subplot(1,2,1)
plt.plot(range(1, n_epochs+1), history['loss'], label='Training')
plt.plot(range(1, n_epochs+1), history['val_loss'], label='Validation')
plt.xlabel('Epoch'); plt.ylabel('Loss'); plt.title('Loss')
plt.legend()
plt.subplot(1,2,2)
plt.plot(range(1, n_epochs+1), history['accuracy'], label='Training')
plt.plot(range(1, n_epochs+1), history['val_accuracy'], label='Validation')
plt.xlabel('Epoch'); plt.ylabel('Accuracy'); plt.title('Accuracy')
plt.legend()
model.evaluate(test_gen)
# getting the prediction of the model
pred_model = model.predict(test_gen)
pred_model
# calculate f1_score --> got 0.9880
y_predict = pred_model.argmax(axis=1)
y_true= test_gen.classes
f1_score(y_true,y_predict,average='macro')
# save as hdf5 file
model.save("/content/drive/MyDrive/Colab Notebooks/hamoye/data/inceptionResNetV2.hdf5")
```
| github_jupyter |
# 面部识别
一般关于什么内容的网络都命名为...Net或者deep...,比如关于面部识别的就是FaceNet以及DeepFace。
面部识别问题一般分为两类:人脸检测(1:1匹配问题)和人脸识别(1:k识别问题)
FaceNet通过神经网络学习,将人脸照片编码成128维向量,通过比较两个向量来判断,两张照片是否是同一个人。
这一节中,将学会实现triplet loss函数,以及使用预训练好的模型来映射128维的照片,以及使用这些编码好的照片来运行人脸检测以及人脸识别
关于视频中说到,对一个照片进行编码时,有可能通道数在后面,也可能在前面,这一节我们统一约定通道数在前,所以对于一个batch的维度是(m, n_C, n_H, n_W)。
CV2是OpenCV官方的一个扩展库,里面含有各种有用的函数以及进程。 OpenCV的全称是:Open Source Computer Vision Library计算机视觉开源库。安装这个库的方法是直接安装opencv,并且是在网上下.whl的文件,如果是python2和python3的环境都有的话,应该在python3的环境下进行安装,进行上面这个文件所在的目录,使用pip install ...(文件名.whl),安装好以后我这上面报错,报错信息是:找不到DLL,解决办法是下了个vs2015安装了,以及在3的环境中放了python3.dll文件。
```
from keras.models import Sequential
from keras.layers import Conv2D, ZeroPadding2D, Activation, Input, concatenate
from keras.models import Model
from keras.layers.normalization import BatchNormalization
from keras.layers.pooling import MaxPooling2D, AveragePooling2D
from keras.layers.merge import Concatenate
from keras.layers.core import Lambda, Flatten, Dense
from keras.initializers import glorot_uniform
from keras.engine.topology import Layer
from IPython.display import SVG
from keras.utils import plot_model
from keras.utils.vis_utils import model_to_dot
from keras import backend as K
K.set_image_data_format('channels_first')
import cv2
import os
import numpy as np
from numpy import genfromtxt
import pandas as pd
import tensorflow as tf
from fr_utils import *
from inception_blocks import *
%matplotlib inline
%load_ext autoreload
%autoreload 2
np.set_printoptions(threshold=np.nan)
```
## 0 简单人脸验证
给定两张照片,判断是不是同一个人,最简单的方法就是比较两张图片的像素,计算欧氏距离如果小于某个阈值,就认为是同一个人。

当然,这个算法表现的很差,因为像素值可能会很局灯光,角度等问题变化,或者镜子也会变化位置等。所以会发现不应该直接使用原图,而是学习一个f(img)函数,让这个函数为图片生成一个新的更准确的编码,最后用这个编码来判断是否是一个人。
## 1. 把人脸照片编码为128维向量
### 1.1 使用卷积网络来计算编码
FaceNet需要很多的数据和很长时间来训练,所以跟随常规操作在实际深度学习的应用,直接加载别人已经训练好的权重。这个网络模型使用inception模型。inception_blocks.py 文件中可以查看inception模型是如何实现的。
关键要记住的:
- 这个网络使用的是96\*96的RGB图像作为输入。所以根据对应的要求,应该修改为 $(m, n_C, n_H, n_W) = (m, 3, 96, 96)$
- 输出应该是(m,128)维的矩阵,将每一张图片编码成128维向量
```
# faceRecoModel是写在inception_blocks.py里的一个函数,只需要输入X的维度,就可以自己创建x张量,并且中间搭建了一系列的网络架构
#网络的最后一层是128个隐藏单元,所以最后就创建好了这个模型,所以这个函数就是返回一个模型
FRmodel = faceRecoModel(input_shape=(3, 96, 96))
#使用FRmodel变量接收这个模型
```
出现问题,l2_normalize() got an unexpected keyword argument 'axis',看网上有类似的问题说是tensorflow的版本问题导致的,所以就准备安装新的包版本,但是一个环境只能有一套包,可是又不想卸载之前的想安装两种版本,所以就需要安装虚拟环境,安装方法如下:
```python
# 安装虚拟环境
pip install virtualenv
pip install virtualenvwrapper-win
# 创建一个虚拟环境,后面是可以自己任意取名字的
mkvirtualenv tf-cpu-1.6.0
#进入虚拟环境工作区
workon tf-cpu-1.6.0
#在虚拟环境下安装新的tensorflow的包
pip install tensorflow==1.6.0
#安装虚拟环境后,一个工程要在某个指定的虚拟环境下工作时,环境间的包不是共享的,那么就得完整的给虚拟环境安装一套,最后还是决定重新安装
```
最后通过各种看官方文档,可能是tensorflow的版本问题,但是也没有找着具体应该用哪个版本,因为tf.nn.l2_normalization(x,axis)这个方法后来的传入参数里面已经没有axis这个参数了,取而代之的是dim这个参数,但是是一样的意思,都表示对前面x这个张量按某个维度进行l2正则化,而在keras的包里面,调用了tf.nn.l2_normalization这个方法,找到文件修改参数名字即可。总的来说,方案就是:
```
到报错的那个文件,我的是这个D:\softpath\Anaconda2\envs\py3\Lib\site-packages\keras\backend\tensorflow_backend.py
将里面return tf.nn.l2_normalize(x, axis=axis) 修改为
return tf.nn.l2_normalize(x, dim=axis)
```
```
print("Total Params:", FRmodel.count_params())
```
通过使用128神经元的全连接层作为最后一层,这个模型保证了输出还是一个编码为128维的向量。然后就是用两张图片都输入这个网络中产生的128维向量进行比较判断是否是同一个人。

### 1.2 triplet loss
使用某人的两张照片与一张非本人照片,去训练网络,损失的定义原理就是根据,同一个的照片计算出来的128维向量距离会很近,而不是同一个人计算出来的距离会很远,所以d(A,P)-d(A,N)\<0,d()函数的计算方式是两个向量对应元素之差的平方和。如果这个差值小于某个值,则说明模型判断anchor是不是本人还是很有用的。
Anchor (A), Positive (P), Negative (N)


α is called the margin. It is a hyperparameter that you should pick manually. We will use α=0.2.
**Exercise**: Implement the triplet loss as defined by formula (3). Here are the 4 steps:
1. Compute the distance between the encodings of "anchor" and "positive": $\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2$
2. Compute the distance between the encodings of "anchor" and "negative": $\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2$
3. Compute the formula per training example: $ \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2 + \alpha$
3. Compute the full formula by taking the max with zero and summing over the training examples:
$$\mathcal{J} = \sum^{N}_{i=1} \large[ \small \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2 - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2+ \alpha \large ] \small_+ \tag{3}$$
Useful functions: `tf.reduce_sum()`, `tf.square()`, `tf.subtract()`, `tf.add()`, `tf.reduce_mean`, `tf.maximum()`.
```
#定义triplet loss 函数
# GRADED FUNCTION: triplet_loss
def triplet_loss(y_true, y_pred, alpha = 0.2):
"""
Implementation of the triplet loss as defined by formula (3)
Arguments:
y_true -- true labels, required when you define a loss in Keras, you don't need it in this function.
y_pred -- python list containing three objects:
anchor -- the encodings for the anchor images, of shape (None, 128)
positive -- the encodings for the positive images, of shape (None, 128)
negative -- the encodings for the negative images, of shape (None, 128)
Returns:
loss -- real number, value of the loss
"""
anchor, positive, negative = y_pred[0], y_pred[1], y_pred[2]
### START CODE HERE ### (≈ 4 lines)
# Step 1: Compute the (encoding) distance between the anchor and the positive
pos_dist = tf.reduce_sum(tf.square(tf.subtract(y_pred[0],y_pred[1])),axis=-1)
"""
L2范数是我们最常见最常用的范数了,我们用的最多的度量距离欧氏距离就是一种L2范数,表示向量元素的平方差再求和
tf.subtract对应元素相减,tf.square每个元素求平方值 tf.reduce_sum可以指定在某维度上求和
"""
# Step 2: Compute the (encoding) distance between the anchor and the negative
neg_dist = tf.reduce_sum(tf.square(tf.subtract(y_pred[0],y_pred[2])),axis=-1)
# Step 3: subtract the two previous distances and add alpha.
basic_loss = tf.add(tf.subtract(pos_dist,neg_dist),alpha)
# Step 4: Take the maximum of basic_loss and 0.0. Sum over the training examples.
loss = tf.reduce_sum(tf.maximum(basic_loss,0.0))
### END CODE HERE ###
return loss
with tf.Session() as test:
tf.set_random_seed(1)
y_true = (None, None, None)
y_pred = (tf.random_normal([3, 128], mean=6, stddev=0.1, seed = 1),
tf.random_normal([3, 128], mean=1, stddev=1, seed = 1),
tf.random_normal([3, 128], mean=3, stddev=4, seed = 1))
loss = triplet_loss(y_true, y_pred)
print("loss = " + str(loss.eval()))
```
## 2. 加载已经训练好的模型
FaceNet是使用最小化triple loss来训练的。但是因为训练需要一些数据以及一些计算,所以我们只加载了模型,现在需要花时间编译一下。
```
FRmodel.compile(optimizer = 'adam', loss = triplet_loss, metrics = ['accuracy'])
load_weights_from_FaceNet(FRmodel)
```
## 3. 应用这个模型
现在对这个幸福公寓,不想任何人都能进来,所以要建立人脸验证系统,只能放一些特定的人进来。为了进公寓,每一个人都必须携带一个ID card。
### 3.1 人脸验证
首先建立一个数据库来存放一个可以被允许进入的人的编码好的向量。为了生成编码,我们使用img_to_encoding(image_path, model)这个函数,是在模型的前向传播的基础上运行的,针对某一张特定的图片。
运行下面代码来创建一个数据库,使用一个python字典来表示。这个数据集就把每一个人的名字对应为一个128维的向量,并且存放起来了。
```
database = {}
database["danielle"] = img_to_encoding("images/danielle.png", FRmodel)
database["younes"] = img_to_encoding("images/younes.jpg", FRmodel)
database["tian"] = img_to_encoding("images/tian.jpg", FRmodel)
database["andrew"] = img_to_encoding("images/andrew.jpg", FRmodel)
database["kian"] = img_to_encoding("images/kian.jpg", FRmodel)
database["dan"] = img_to_encoding("images/dan.jpg", FRmodel)
database["sebastiano"] = img_to_encoding("images/sebastiano.jpg", FRmodel)
database["bertrand"] = img_to_encoding("images/bertrand.jpg", FRmodel)
database["kevin"] = img_to_encoding("images/kevin.jpg", FRmodel)
database["felix"] = img_to_encoding("images/felix.jpg", FRmodel)
database["benoit"] = img_to_encoding("images/benoit.jpg", FRmodel)
database["arnaud"] = img_to_encoding("images/arnaud.jpg", FRmodel)
database["tudou"] = img_to_encoding("images/tudou1.jpg", FRmodel)
database = {}
database["danielle"] = img_to_encoding("images/danielle.png", FRmodel)
database["younes"] = img_to_encoding("images/younes.jpg", FRmodel)
print(database)
print(len(database))
```
实现verify()函数来检查门前照相机拍到的图片是否是这个人,并且是这个名字。
1. 计算门前拍到的照片的编码
2. 计算数据库里的编码和门前照片编码的距离。
3. 如果距离小于0.7,那么久打开门,否则不打开
```
# GRADED FUNCTION: verify
def verify(image_path, identity, database, model):
"""
Function that verifies if the person on the "image_path" image is "identity".
Arguments:
image_path -- path to an image
identity -- string, name of the person you'd like to verify the identity. Has to be a resident of the Happy house.
database -- python dictionary mapping names of allowed people's names (strings) to their encodings (vectors).
model -- your Inception model instance in Keras
Returns:
dist -- distance between the image_path and the image of "identity" in the database.
door_open -- True, if the door should open. False otherwise.
"""
### START CODE HERE ###
# Step 1: Compute the encoding for the image. Use img_to_encoding() see example above. (≈ 1 line)
encoding = img_to_encoding(image_path, FRmodel)
# Step 2: Compute distance with identity's image (≈ 1 line)
dist = np.linalg.norm(encoding-database[identity])
# Step 3: Open the door if dist < 0.7, else don't open (≈ 3 lines)
if dist<0.7:
print("It's " + str(identity) + ", welcome home!")
door_open = True
else:
print("It's not " + str(identity) + ", please go away")
door_open = False
### END CODE HERE ###
return dist, door_open
#对某个要进来的人进行测试
verify("images/camera_0.jpg", "younes", database, FRmodel)
verify("images/camera_2.jpg", "kian", database, FRmodel)
verify("images/tudou.jpg", "tudou", database, FRmodel)
```
### 3.2 人脸识别
你的人脸验证系统工作的很好,但是如果一个人的身份证被偷了,那他就没法进入了,因为无法进行身份验证。为了解决这个问题,就需要把人脸验证系统改为人脸识别系统,就不用需要身份证了。
现在要实现人脸识别,需要的输入仅仅是一张照片,并且计算出来是否是已经通过验证的某一个人。与人脸验证不同的是,我们不需要再知道这个人的身份作为输入。
实现who_is_it():
1. 计算目标输入图像的128维编码
2. 从数据库中找到这个目标编码距离最近的一个编码:
- 初始化min_dist变量为一个比较大的数,它能跟踪找到距离目标编码最近的编码
- 在数据库字典名字和编码中循环,loop use for (name, db_enc) in database.items().
- 计算L2距离
- 如果这个距离比min-dist小,就把min-dist设置到字典中,并且标识姓名
```
# GRADED FUNCTION: who_is_it
def who_is_it(image_path, database, model):
"""
Implements face recognition for the happy house by finding who is the person on the image_path image.
Arguments:
image_path -- path to an image
database -- database containing image encodings along with the name of the person on the image
model -- your Inception model instance in Keras
Returns:
min_dist -- the minimum distance between image_path encoding and the encodings from the database
identity -- string, the name prediction for the person on image_path
"""
### START CODE HERE ###
## Step 1: Compute the target "encoding" for the image. Use img_to_encoding() see example above. ## (≈ 1 line)
encoding = img_to_encoding(image_path, FRmodel)
## Step 2: Find the closest encoding ##
# Initialize "min_dist" to a large value, say 100 (≈1 line)
min_dist = 100
# Loop over the database dictionary's names and encodings.
for (name, db_enc) in database.items():
# Compute L2 distance between the target "encoding" and the current "emb" from the database. (≈ 1 line)
dist = np.linalg.norm(encoding-db_enc)
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name. (≈ 3 lines)
if dist<min_dist:
min_dist = dist
identity = name
"""
这里首先初始化min_dist最小距离为一个比较大的数,那么在循环的时候,跟第一个人比较,如果距离比这个min还大,说明不是这个人,不理会,继续循环
如果循环到某个人,两人小于min,那么久把min修改为当前的距离,并且记住名字,但是并不能说明就是这个人,可能只是很像,那么继续循环
直到把所有人都循环完毕,那么久找到了数据库里所有人中跟目标最像的一个,但是并不能判断是,要根据这个min距离来跟阈值比较,最后确定是不是这个人
"""
### END CODE HERE ###
if min_dist > 0.7:
print("Not in the database.")
else:
print ("it's " + str(identity) + ", the distance is " + str(min_dist))
return min_dist, identity
who_is_it("images/camera_0.jpg", database, FRmodel)
who_is_it("images/camera_1.jpg", database, FRmodel)
who_is_it("images/tudou1.jpg", database, FRmodel)
```
现在你的幸福公寓大门只对已经验证过的人开放了,人们也不再需要携带身份证了。
后面的内容这节内容不会再实现了,但是你可以自己去提高你的算法从下面这些方面:
1. 放入更多人的照片,在不同的光线角度下,或者不同的天气下,然后给定一个图,去跟很多的图进行比较。
2. 裁剪图像只包含脸部,而脸部周围的“边界”区域较少。这种预处理消除了人脸周围的一些不相关的像素,并使算法更健壮。
记住:
- 人脸验证解决1:1问题,人脸识别解决1:k的问题
- triplet是一个有效的损失函数去训练神经网络学习面部图片的编码
- 同样的编码可以用在验证和识别上,通过距离去判断是否是一个人
```
FRmodel.summary()
plot_model(FRmodel, to_file='model.png')
SVG(model_to_dot(FRmodel).create(prog='dot', format='svg'))
```
| github_jupyter |
```
#IMPORT SEMUA LIBARARY
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY UNTUK POSTGRE
from sqlalchemy import create_engine
import psycopg2
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY BASE PATH
import os
import io
#IMPORT LIBARARY PDF
from fpdf import FPDF
#IMPORT LIBARARY CHART KE BASE64
import base64
#IMPORT LIBARARY EXCEL
import xlsxwriter
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
def uploadToPSQL(columns, table, filePath, engine):
#FUNGSI UNTUK MEMBACA CSV
df = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#APABILA ADA FIELD KOSONG DISINI DIFILTER
df.fillna('')
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
del df['kategori']
del df['jenis']
del df['pengiriman']
del df['satuan']
#MEMINDAHKAN DATA DARI CSV KE POSTGRESQL
df.to_sql(
table,
engine,
if_exists='replace'
)
#DIHITUNG APABILA DATA YANG DIUPLOAD BERHASIL, MAKA AKAN MENGEMBALIKAN KELUARAN TRUE(BENAR) DAN SEBALIKNYA
if len(df) == 0:
return False
else:
return True
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
def makeChart(host, username, password, db, port, table, judul, columns, filePath, name, subjudul, limit, negara, basePath):
#TEST KONEKSI DATABASE
try:
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBL DATA DARI TABLE YANG DIDEFINISIKAN DIBAWAH, DAN DIORDER DARI TANGGAL TERAKHIR
#BISA DITAMBAHKAN LIMIT SUPAYA DATA YANG DIAMBIL TIDAK TERLALU BANYAK DAN BERAT
postgreSQL_select_Query = "SELECT * FROM "+table+" ORDER BY tanggal ASC LIMIT " + str(limit)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MELAKUKAN LOOPING ATAU PERULANGAN DARI DATA YANG SUDAH DIAMBIL
#KEMUDIAN DATA TERSEBUT DITEMPELKAN KE VARIABLE DIATAS INI
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
if row[2] == "":
lengthy.append(float(0))
else:
lengthy.append(float(row[2]))
#FUNGSI UNTUK MEMBUAT CHART
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#MASUKAN DATA ID DARI DATABASE, DAN JUGA DATA TANGGAL
ax.bar(uid, lengthy, align='center')
#UNTUK JUDUL CHARTNYA
ax.set_title(judul)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
#TOTAL DATA YANG DIAMBIL DARI DATABASE, DIMASUKAN DISINI
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#line
#MASUKAN DATA DARI DATABASE
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#UNTUK JUDUL CHARTNYA
plt.title(judul)
plt.grid(True)
l = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(l, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#pie
#UNTUK JUDUL CHARTNYA
plt.title(judul)
#MASUKAN DATA DARI DATABASE
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.axis('equal')
p = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(p, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#MENGAMBIL DATA DARI CSV YANG DIGUNAKAN SEBAGAI HEADER DARI TABLE UNTUK EXCEL DAN JUGA PDF
header = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
header.fillna('')
del header['tanggal']
del header['total']
#MEMANGGIL FUNGSI EXCEL
makeExcel(mobile_records, header, name, limit, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(mobile_records, header, judul, barChart, lineChart, pieChart, name, subjudul, limit, basePath)
#JIKA GAGAL KONEKSI KE DATABASE, MASUK KESINI UNTUK MENAMPILKAN ERRORNYA
except (Exception, psycopg2.Error) as error :
print (error)
#KONEKSI DITUTUP
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
def makeExcel(datarow, dataheader, name, limit, basePath):
#MEMBUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/BLOOMBERG/SektorFiskal/excel/'+name+'.xlsx')
#MENAMBAHKAN WORKSHEET PADA FILE EXCEL TERSEBUT
worksheet = workbook.add_worksheet('sheet1')
#SETINGAN AGAR DIBERIKAN BORDER DAN FONT MENJADI BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#MENJADIKAN DATA MENJADI ARRAY
data=list(datarow)
isihead=list(dataheader.values)
header = []
body = []
#LOOPING ATAU PERULANGAN, KEMUDIAN DATA DITAMPUNG PADA VARIABLE DIATAS
for rowhead in dataheader:
header.append(str(rowhead))
for rowhead2 in datarow:
header.append(str(rowhead2[1]))
for rowbody in isihead[1]:
body.append(str(rowbody))
for rowbody2 in data:
body.append(str(rowbody2[2]))
#MEMASUKAN DATA DARI VARIABLE DIATAS KE DALAM COLUMN DAN ROW EXCEL
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
for col_num, data in enumerate(body):
worksheet.write(1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
def makePDF(datarow, dataheader, judul, bar, line, pie, name, subjudul, lengthPDF, basePath):
#FUNGSI UNTUK MENGATUR UKURAN KERTAS, DISINI MENGGUNAKAN UKURAN A4 DENGAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#MENAMBAHKAN HALAMAN PADA PDF
pdf.add_page()
#PENGATURAN UNTUK JARAK PADDING DAN JUGA UKURAN FONT
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#MEMASUKAN JUDUL KE DALAM PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#MEMASUKAN SUB JUDUL KE PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#MEMBUAT GARIS DI BAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','',10.0)
#MENGAMBIL DATA HEADER PDF YANG SEBELUMNYA SUDAH DIDEFINISIKAN DIATAS
datahead=list(dataheader.values)
pdf.set_font('Times','B',12.0)
pdf.ln(0.5)
th1 = pdf.font_size
#MEMBUAT TABLE PADA PDF, DAN MENAMPILKAN DATA DARI VARIABLE YANG SUDAH DIKIRIM
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][0], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Jenis", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][1], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Pengiriman", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][2], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Satuan", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][3], border=1, align='C')
pdf.ln(2*th1)
#PENGATURAN PADDING
pdf.set_xy(17.0, 75.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','B',11.0)
data=list(datarow)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
#PENGATURAN UNTUK JARAK PADDING
pdf.ln(0.5)
th = pdf.font_size
#MEMASUKAN DATA HEADER YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.cell(50, 2*th, str("Negara"), border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#MEMASUKAN DATA ISI YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(50, 2*th, negara, border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#MENGAMBIL DATA CHART, KEMUDIAN CHART TERSEBUT DIJADIKAN PNG DAN DISIMPAN PADA DIRECTORY DIBAWAH INI
#BAR CHART
bardata = base64.b64decode(bar)
barname = basePath+'jupyter/BLOOMBERG/SektorFiskal/img/'+name+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINE CHART
linedata = base64.b64decode(line)
linename = basePath+'jupyter/BLOOMBERG/SektorFiskal/img/'+name+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIE CHART
piedata = base64.b64decode(pie)
piename = basePath+'jupyter/BLOOMBERG/SektorFiskal/img/'+name+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
widthcol = col/3
#MEMANGGIL DATA GAMBAR DARI DIREKTORY DIATAS
pdf.image(barname, link='', type='',x=8, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(linename, link='', type='',x=103, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(piename, link='', type='',x=195, y=100, w=widthcol)
pdf.ln(2*th)
#MEMBUAT FILE PDF
pdf.output(basePath+'jupyter/BLOOMBERG/SektorFiskal/pdf/'+name+'.pdf', 'F')
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
#DEFINISIKAN COLUMN BERDASARKAN FIELD CSV
columns = [
"kategori",
"jenis",
"tanggal",
"total",
"pengiriman",
"satuan",
]
#UNTUK NAMA FILE
name = "SektorFiskal2_4"
#VARIABLE UNTUK KONEKSI KE DATABASE
host = "localhost"
username = "postgres"
password = "1234567890"
port = "5432"
database = "bloomberg_SektorFiskal"
table = name.lower()
#JUDUL PADA PDF DAN EXCEL
judul = "Data Sektor Fiskal"
subjudul = "Badan Perencanaan Pembangunan Nasional"
#LIMIT DATA UNTUK SELECT DI DATABASE
limitdata = int(8)
#NAMA NEGARA UNTUK DITAMPILKAN DI EXCEL DAN PDF
negara = "Indonesia"
#BASE PATH DIRECTORY
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE CSV
filePath = basePath+ 'data mentah/BLOOMBERG/SektorFiskal/' +name+'.csv';
#KONEKSI KE DATABASE
engine = create_engine('postgresql://'+username+':'+password+'@'+host+':'+port+'/'+database)
#MEMANGGIL FUNGSI UPLOAD TO PSQL
checkUpload = uploadToPSQL(columns, table, filePath, engine)
#MENGECEK FUNGSI DARI UPLOAD PSQL, JIKA BERHASIL LANJUT MEMBUAT FUNGSI CHART, JIKA GAGAL AKAN MENAMPILKAN PESAN ERROR
if checkUpload == True:
makeChart(host, username, password, database, port, table, judul, columns, filePath, name, subjudul, limitdata, negara, basePath)
else:
print("Error When Upload CSV")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
pd.__version__
class Coordinate:
'''Coordinate on Earth'''
def __init__(self, lat=0, long=0):
print('__init__')
self.lat = lat
self.long = long
def __repr__(self):
print('__repr__')
return f'Coordinate({self.lat}, {self.long})'
def __str__(self):
print('__str__')
ns = 'NS'[self.lat < 0]
we = 'EW'[self.long < 0]
return f'{abs(self.lat):.1f}°{ns}, {abs(self.long):.1f}°{we}'
c = Coordinate(41.4,-81)
c
print(c)
import geohash
class Coordinate:
'''Coordinate on Earth'''
reference_system = 'WGS84'
def __init__(self, lat, long):
self.lat = lat
self.long = long
def __repr__(self):
return f'Coordinate({self.lat}, {self.long})'
def __str__(self):
ns = 'NS'[self.lat < 0]
we = 'WE'[self.long < 0]
return f'{abs(self.lat):.1f}°{ns}, {abs(self.long):.1f}°{we}'
def geohash(self):
return geohash.encode(self.lat, self.long)
Coordinate(41.4, -81)
from dataclasses import dataclass
@dataclass
class Coordinate:
lat: float
long: float
def __str__(self):
print('__str__')
ns = 'NS'[self.lat < 0]
we = 'EW'[self.long < 0]
return f'{abs(self.lat):.1f}°{ns}, {abs(self.long):.1f}°{we}'
brno = Coordinate(49.1951, 16.6068)
brno
print(brno)
import pathlib
file = pathlib.Path.cwd()/'index.html'
file.read_text().split('\n')[0]
import os
type(os.environ)
if 'HOME' in os.environ:
userhome = os.environ['HOME']
userhome
import matplotlib.pyplot as plt
@dataclass
class Point:
x: float
y: float
z: float = 0.0
def __str__(self):
print('__str__')
return f'x:{self.x} y:{self.y} z:{self.z}'
def plot(self):
plt.title(f'x:{self.x} y:{self.y}')
plt.plot(self.x, self.y, 'bo')
p = Point(1.5, 2.5)
p.plot()
print(p)
```
Unit Type | Unit | Abbreviation
-----------|--------|-------------
Length | Meter | m
Time | Second | s
Mass | Gram [1] | g
Temperature| Kelvin | K
Luminous intensity| Candela | cd
Current | Ampere | A
Quantity [2] | Mole | mol
[1]: Technically, for historical reasons, the base unit of mass is actually the kilogram, but it makes more sense when thinking about it to view the gram it self as the base unit. The kilogram is a base unit in the sense that the standardized weight is based off of the kilogram. However, the gram is a base unit in the sense that all of the prefixes are based off of the weight of a gram.
[2]: This is primarily used in chemistry for counting small things like atoms and molecules.
* A liter is a thousandth of a cubic meter. Thus, we can take the unit of length and use it to describe a unit of volume.
* the newton is defined as being a “kilogram-meter per second squared.” This is another way of saying that a newton is the amount of force which accelerates 1 kilogram 1 meter per second, per second.
```
from urllib.request import urlopen
shakespeare = urlopen('http://composingprograms.com/shakespeare.txt')
shakespeare.getcode()
shakespeare.getheaders()
shakespeare.read().decode().split('\n')[:10]
words = set(shakespeare.read().decode().split())
len(words)
{w for w in words if len(w) == 6 and w[::-1] in words}
```
Saturnin - Zdeněk Jirotka
```
import re
from urllib.request import urlopen
roman_numerals = {1: 'I', 2: 'II', 3: 'III', 4: 'IV', 5: 'V',
6: 'VI', 7: 'VII', 8: 'VIII', 9: 'IX', 10: 'X',
11: 'XI', 12: 'XII', 13: 'XIII', 14: 'XIV', 15: 'XV', 16: 'XVI', 17: 'XVII',
18: 'XVIII', 19: 'XIX', 20: 'XX', 21: 'XXI', 22: 'XXII',
23: 'XXIII', 24: 'XXIV', 25: 'XXV', 26: 'XXVI', 27: 'XXVII', 28: 'XXVIII', 29: 'XXIX'
30: 'XXX', 40: 'XL', 50: 'L', 60: 'LX', 70: 'LXX', 80: 'LXXX',
90: 'XC', 100: 'C', 200: 'CC', 300: 'CCC', 400: 'CD', 500: 'D',
600: 'DC', 700: 'DCC', 800: 'DCCC', 900: 'CM', 1000: 'M',
2000: 'MM', 3000: 'MMM'}.values()
def strip_tags(line):
return re.sub('<[^<]+?>', '', line)
def proc_text(text):
# začatek a konec == balast
text = [strip_tags(line) for line in text]
text = text[28:len(text)-2]
# '' je konec odstavce ...
text = [line if line else '\n' for line in text]
# věty
text = [line+'\n' if line.endswith('.') else line for line in text]
# název
#for indx in [1,3]: text.insert(indx,'\n')
text = text[2:]
# v druhá kapitola nezačína '\n'
for line_num, line in enumerate(text):
if line.startswith('Uvoluji se chytit Marka Aurelia'):
text.insert(line_num+1, '\n')
# subcapitoly
subcap = False
for line_num,line in enumerate(text):
if line in roman_numerals:
subcap = True
if subcap == True:
if line != '\n':
#print(line_num,line)
text[line_num] = text[line_num] + ' ~ '
else:
subcap = False
return text
saturnin = ['Saturnin - Zdeněk Jirotka \n']
for i in range(1,6):
r = urlopen(f'http://www.multiweb.cz/saturnin/saturnin{i}.htm')
text = r.read().decode("cp1250").split('\r\n')
text = proc_text(text)
text = ''.join(text).split('\n')
saturnin.append(text)
saturnin[0]
len(saturnin)
with open("saturnin.txt", 'w') as output:
for odstavec in saturnin:
output.write(str(odstavec) + '\n')
with open("saturnin.txt", 'r') as f:
text = f.read()
text
roman_numerals = {1: 'I', 2: 'II', 3: 'III', 4: 'IV', 5: 'V',
6: 'VI', 7: 'VII', 8: 'VIII', 9: 'IX', 10: 'X',
11: 'XI', 12: 'XII', 13: 'XIII', 14: 'XIV', 15: 'XV', 16: 'XVI', 17: 'XVII', 18: 'XVIII', 19: 'IXX',
20: 'XX', 21: 'XXI', 22: 'XXII', 23: 'XXIII', 24: 'XXIV', 25: 'XXV',
30: 'XXX', 40: 'XL', 50: 'L', 60: 'LX', 70: 'LXX', 80: 'LXXX',
90: 'XC', 100: 'C', 200: 'CC', 300: 'CCC', 400: 'CD', 500: 'D',
600: 'DC', 700: 'DCC', 800: 'DCCC', 900: 'CM', 1000: 'M',
2000: 'MM', 3000: 'MMM'}.values()
'II' in roman_numerals
for line_num,line in enumerate(text):
if line in roman_numerals:
print(line)
roman_numerals = {1: 'I', 2: 'II', 3: 'III', 4: 'IV', 5: 'V',
6: 'VI', 7: 'VII', 8: 'VIII', 9: 'IX', 10: 'X', 20: 'XX',
30: 'XXX', 40: 'XL', 50: 'L', 60: 'LX', 70: 'LXX', 80: 'LXXX',
90: 'XC', 100: 'C', 200: 'CC', 300: 'CCC', 400: 'CD', 500: 'D',
600: 'DC', 700: 'DCC', 800: 'DCCC', 900: 'CM', 1000: 'M',
2000: 'MM', 3000: 'MMM'}
text = ['\n'+line+'\n' if line in roman_numerals.values() else line for line in text]
text = ''.join(text).split('\n')
text[0]
table = """
1 I
2 II
3 III
4 IV
5 V
6 VI
7 VII
8 VIII
9 IX
10 X
11 XI
12 XII
13 XIII
14 XIV
15 XV
16 XVI
17 XVII
18 XVIII
19 XIX
20 XX
21 XXI
22 XXII
23 XXIII
24 XXIV
25 XXV
26 XXVI
27 XXVII
28 XXVIII
29 XXIX
30 XXX
31 XXXI
32 XXXII
33 XXXIII
34 XXXIV
35 XXXV
36 XXXVI
37 XXXVII
38 XXXVIII
39 XXXIX
40 XL
41 XLI
42 XLII
43 XLIII
44 XLIV
45 XLV
46 XLVI
47 XLVII
48 XLVIII
49 XLIX
50 L
"""
roman = []
arabic = []
table = table.split()
for l in range(len(table)//2):
roman.append(table.pop())
arabic.append(table.pop())
arabic.reverse()
roman.reverse()
literals = dict(zip(int(arabic), roman))
literals
table = """
1 I
2 II
3 III
4 IV
5 V
6 VI
7 VII
8 VIII
9 IX
10 X
11 XI
12 XII
13 XIII
14 XIV
15 XV
16 XVI
17 XVII
18 XVIII
19 XIX
20 XX
21 XXI
22 XXII
23 XXIII
24 XXIV
25 XXV
26 XXVI
27 XXVII
28 XXVIII
29 XXIX
30 XXX
31 XXXI
32 XXXII
33 XXXIII
34 XXXIV
35 XXXV
36 XXXVI
37 XXXVII
38 XXXVIII
39 XXXIX
40 XL
41 XLI
42 XLII
43 XLIII
44 XLIV
45 XLV
46 XLVI
47 XLVII
48 XLVIII
49 XLIX
50 L
""".split()
table
{ table.pop():table.pop() for item in range(table//2)}
arabic
literals = dict(zip(arabic,roman))
literals
```
| github_jupyter |
#Introduction
In this notebook we will use a clustering algorithm to analyze our data (i.e. YouTube comments of a single video).
This will help us extract topics of discussion.
We use the embeddings generated in Assignment 4 as input.
(This notebook will not run without first running the assignment 4 Notebook, as it relies on the data in the folder 'output/')
Each of our comments has been assigned a vector that encodes information about its meaning.
The closer two vectors are, the more similar the meaning.
Each vector is of 512 Dimensions.
Before we can cluster our data we need to reduce the embeddings' dimensionality to overcome the curse of dimensionality.
We use the UMAP ALgorithm for this.
After that we use the KMedoids Algorithm to partition the embedding space and generate our clusters this way.
We need to define the number of clusters we want to have.
To find the optimal number of clusters, we use a simple optimization scheme.
Once the clusters are created, we visualize them.
To do this we reduce the dimensionality of the embeddings again to two dimensions.
Then we render a scatterplot of our data.
Furthermore we want to analyze and interpret our clusters.
To do this, we:
- print some statistics about each of the clusters
- print cluster's medoid (the central sample)
- print the cluster(s) we want to analyze further
Check to see if jupyter lab uses the correct python interpreter with '!which python'.
It should be something like '/opt/anaconda3/envs/[environment name]/bin/python' (on Mac).
If not, try this: https://github.com/jupyter/notebook/issues/3146#issuecomment-352718675
```
!which python
```
# Install dependencies:
```
install_packages = False
if install_packages:
!conda install -c conda-forge umap-learn -y
!conda install -c conda-forge scikit-learn-extra -y
```
# Imports
```
#imports
import pandas as pd
import numpy as np
import os
import time
import matplotlib.pyplot as plt
import umap
from sklearn_extra.cluster import KMedoids
import seaborn as sns
#from sklearn.cluster import AgglomerativeClustering, DBSCAN, KMeans, OPTICS
from sklearn.metrics import silhouette_samples, silhouette_score, pairwise_distances
```
# Functions to Save and load manually
```
# Save and load your data after clustering
def save_results():
data.to_pickle(output_path+'data_clustered'+'.pkl')
def load_results():
data = pd.read_pickle(output_path+'data_clustered'+'.pkl')
# Set pandas print options
This will improve readability of printed pandas dataframe.
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', None)
pd.set_option('display.max_colwidth', None)
```
## Set global Parameters
Set your parameters here:
output_path: Files generated in this notebook will be saved here.
model_type: Define which model was used to produce the embeddings. (Check the name of the .npy-file containing the embeddings)
```
output_path = "./output/"
model_type = 'Transformer' #@param ['DAN','Transformer','Transformer_Multilingual']
```
# Load Data
Load the preprocessed data as a pandas dataframe.
And load the embeddings as a numpy ndarray (a matrix in our case).
```
data = pd.read_pickle(output_path+'data_preprocessed'+'.pkl')
labels_default = np.zeros(len(data.index))-1
data['label_manual'] = labels_default
embeddings = np.load(output_path+'/embeddings'+model_type+'.npy', mmap_mode=None, allow_pickle=False, fix_imports=True, encoding='ASCII')
```
# Dimensionality reduction with UMAP
We reduce the number of dimensions of our embeddings to make possibly present clusters more pronounced.
The number of dimensions (num_dimensions) depends on the number of samples
```
# Set the number of dimensions to reduce to
num_dimensions =100
reducer_clustering = umap.UMAP(n_neighbors=50,
n_components=num_dimensions,
metric='cosine',
#n_epochs=200,
learning_rate=.5,
init='spectral',
min_dist=0,
#spread=5.0,
#set_op_mix_ratio=1.0,
#local_connectivity=1.0,
#negative_sample_rate=5,
#transform_queue_size=4.0,
force_approximation_algorithm=True,
unique=True)
embeddings_umap = reducer_clustering.fit_transform(embeddings)
```
# Optimize the Number of Clusters
```
#optimize number of clusters
optimize_number_of_clusters = True#@param {type:'boolean'}
min_clusters=2
max_clusters=1000
step=100
if optimize_number_of_clusters:
rows_list = []
inertias = []
n_clusters = []
silouette_scores = []
init_param = 'k-medoids++' #@param ['random', 'heuristic', 'k-medoids++']
random_state_param=1234 #@param {type:'number'}
for i in range(min_clusters,max_clusters, step):
temp_clustering = KMedoids(n_clusters=i, metric='euclidean', init=init_param, max_iter=200, random_state=random_state_param).fit(embeddings_umap)
silhouette_avg = silhouette_score(embeddings_umap, temp_clustering.labels_)
print("n_clusters:",i, "silhouette_avg:",silhouette_avg)
silhouette_dict = {'number of clusters': i, 'silhouette average': silhouette_avg}
rows_list.append(silhouette_dict)
results = pd.DataFrame(rows_list)
sns.lineplot(x = 'number of clusters', y = 'silhouette average',data = results)
```
# Clustering with KMedoids
```
number_of_clusters = 100
init_param = 'k-medoids++' #@param ['random', 'heuristic', 'k-medoids++']
clustering_model = KMedoids(n_clusters=number_of_clusters,
metric='cosine',
init=init_param,
max_iter=150,
random_state=None).fit(embeddings_umap)
clustering_model
labels = clustering_model.labels_
data["label_kmedoids"] = labels
print("cluster","members", data["label_kmedoids"].value_counts().sort_values())
clustering_model.inertia_
medoids_indices = clustering_model.medoid_indices_
#calculate distances
distances = np.diag(pairwise_distances(X = clustering_model.cluster_centers_[labels], Y = embeddings_umap[:], metric='cosine'))
data["distance_kmedoids"] = distances
```
# Dimensionality Reduction for Visualization
```
num_dimensions =2
reducer_visualization = umap.UMAP(n_neighbors=50,
n_components=num_dimensions,
metric='cosine',
output_metric='euclidean',
#n_epochs=200,
learning_rate=.5,
init='spectral',
min_dist=.1,
spread=5.0,
set_op_mix_ratio=1.0,
local_connectivity=1.0,
negative_sample_rate=5,
transform_queue_size=4.0,
force_approximation_algorithm=True,
unique=True)
embeddings_umap_2d = reducer_visualization.fit_transform(embeddings)
```
# Visualize clustering results
```
#@markdown Set the color palette used for visualizing different clusters
palette_param = "Accent" #@param ['Accent','cubehelix', "tab10", 'Paired', "Spectral"]
#@markdown Set opacity of data points (1 = opaque, 0 = invisible)
alpha_param = 0.16 #@param {type:"slider", min:0, max:1, step:0.01}
sns.relplot(x = embeddings_umap_2d[:, 0], y = embeddings_umap_2d[:, 1], hue = data['label_kmedoids'], palette = palette_param,alpha = alpha_param,height = 10)
```
## Highlight one cluster
```
## Choose a cluster to higlight:
cluster_num = 6
data['highlight'] = np.zeros(len(data.index))
data.loc[data['label_kmedoids'] == cluster_num, 'highlight'] = 1
sns.relplot(x = embeddings_umap_2d[:, 0], y = embeddings_umap_2d[:, 1], hue = data['highlight'], palette = "Accent",alpha = 0.8,height = 10)
```
# Print Medoids and cluster statistics
```
# print the medoids
data.iloc[medoids_indices]
# print statistics for each cluster
data['label_kmedoids'].value_counts().sort_values()
for k,g in data.groupby(by = 'label_kmedoids'):
print(g.iloc[0]['label_kmedoids'],"number of samples: ",len(g.index),"mean distance from center: ", 100*np.mean(g['distance_kmedoids']), "Proportion of replies:", 100*np.sum(g['isReply'])/len(g.index))
```
# Print Cluster
Print the comments within a cluster. Comments are sorted by their distance from the cluster medoid
```
# Choose a cluster to print
cluster_number = 20
# Choose the number of samples to print
number_of_samples_to_print = 10000
data['label_kmedoids'] = data['label_kmedoids'].astype('category')
cluster = data[data['label_kmedoids']==cluster_number]
if cluster["text"].count()<=number_of_samples_to_print:
number_of_samples_to_print = cluster["text"].count()
cluster = cluster.sort_values(by='distance_kmedoids')
print("Number of samples in the cluster:", cluster["text"].count())
print("Average Distance from cluster center:", np.mean(cluster['distance_kmedoids']))
cluster
```
# Assign Cluster labels manually
cluster_number: which cluster would you like to assign labels to?
min_distance: the minimum distance from the cluster medoid be for a data point to still get the specified label
max_distance: the maximum distance from the cluster medoid be for a data point to still get the specified label
label_manual: your label
```
#which cluster would you like to assign labels to?
cluster_number = 18
#your label
label_manual = 'music'
#the minimum distance from the cluster medoid be for a data point to still get the specified label
min_distance = 0
#the maximum distance from the cluster medoid be for a data point to still get the specified label
max_distance = 1000
# 2. Filter data by cluster label and specified label to filtered data
data.loc[(data['label_kmedoids']==cluster_number) & (data['distance_kmedoids'] <= max_distance) & (data['distance_kmedoids'] >= min_distance), 'label_manual'] = label_manual
data[data['label_kmedoids']==cluster_number].sort_values(by='distance_kmedoids')
```
| github_jupyter |
<h4> Save for Storyboard </h4>
```
import struct, socket
import numpy as np
import csv, json
import os
import urllib2
import datetime
import operator
import itertools
try:
import ipywidgets as widgets # For jupyter/ipython >= 1.4
except ImportError:
from IPython.html import widgets
from IPython.display import display, HTML, clear_output, Javascript
path = os.getcwd().split("/")
t_date = path[len(path)-1]
anchor = ''
anchor_type = ''
top_results = 20
details_limit = 1000
query_comments = {}
# Widget styles and initialization
topBox = widgets.Box()
bottomBox = widgets.Box()
mainBoxes_css = (
(None, 'width', '90%'),
(None, 'margin', '0 auto'),
)
topBox._css = mainBoxes_css
bottomBox._css = mainBoxes_css
threatBox = widgets.HBox(width='100%', height='auto')
threat_title = widgets.HTML(height='25px', width='100%')
threat_list_container = widgets.Box(width='80%', height='100%')
threat_button_container = widgets.Box(width='20%', height='100%')
susp_select = widgets.Select(height='100%', width='99%')
search_btn = widgets.Button(description='Search',height='100%', width='65px')
search_btn.button_style = 'primary'
susp_select._css = (
(None, 'height', '90%'),
(None, 'width', '95%'),
('select', 'overflow-x', 'auto'),
('select', 'margin', 0)
)
resultSummaryBox = widgets.Box()
result_title = widgets.HTML(width='100%')
result_summary_box = widgets.HBox(width='100%')
result_summary_container = widgets.Box(width='80%')
result_button_container = widgets.Box(width='20%')
result_summary_box.children = [result_title, result_summary_container, result_button_container]
resultTableBox = widgets.Box()
result_html_title = widgets.HTML(height='25px', width='100%')
result_html_box = widgets.Box() #this one has the scroll
result_html = widgets.HTML(width='100%')
result_box_css = (
(None, 'overflow', 'hidden'),
(None, 'width', '100%'),
)
resultSummaryBox._css = result_box_css
resultTableBox._css = result_box_css
result_html_box._css = (
(None, 'overflow','auto'),
(None, 'max-height', '300px'),
)
threat_button_container._css = (
(None, 'padding-top', '30px'),
)
topBox.children = [threatBox]
bottomBox.children = [resultSummaryBox,resultTableBox]
threat_list_container.children = [threat_title,susp_select]
threat_button_container.children = [search_btn]
threatBox.children = [threat_list_container, threat_button_container]
```
**Interface**
```
yy = t_date[0:4]
mm = t_date[4:6]
dd = t_date[6:8]
def fill_list(list_control,source):
susp_select.options = list_control
susp_select.selected_label = list_control[0][0]
def data_loader():
ips_query = {}
ip_sev={}
dns_sev={}
global query_comments
response = GraphQLClient.request(
query="""query($date:SpotDateType!) {
dns{
threats{
list(date:$date) {
dnsScore
clientIpScore
clientIp
dnsQuery
}
}
}
}""",
variables={
'date': datetime.datetime.strptime(t_date, '%Y%m%d').strftime('%Y-%m-%d')
}
)
query_comments = GraphQLClient.request(
query="""query($date:SpotDateType!) {
dns{
threats{
comments(date:$date) {
title
text
... on DnsQueryCommentType {
dnsQuery
}
... on DnsClientIpCommentType {
clientIp
}
}
}
}
}""",
variables={
'date': datetime.datetime.strptime(t_date, '%Y%m%d').strftime('%Y-%m-%d')
}
)
query_comments = query_comments['data']['dns']['threats']['comments']
if not 'errors' in response:
for row in response['data']['dns']['threats']['list']:
if row['clientIp'] not in ips_query and row['clientIpScore'] == 1:
ips_query[row['clientIp']]='i'
if row['dnsQuery'] not in ips_query and row['dnsScore'] == 1:
ips_query[row['dnsQuery']]='q'
if row['clientIp'] not in ip_sev:
ip_sev[row['clientIp']] = row['clientIpScore']
if row['dnsQuery'] not in dns_sev:
dns_sev[row['dnsQuery']] =row['dnsScore']
else:
print "An error ocurred: " + response["errors"][0]["message"]
threat_title.value ="<h4>Suspicious DNS</h4>"
if len(ips_query) == 0:
display(Javascript("$('.widget-area > .widget-subarea > *').remove();"))
display(widgets.HTML(value="There are not high risk results.", width='90%'),)
else:
sorted_dict = sorted(ips_query.items(), key=operator.itemgetter(0))
fill_list(sorted_dict,susp_select)
def start_investigation():
display(Javascript("$('.widget-area > .widget-subarea > *').remove();"))
data_loader()
if susp_select.options:
display_controls()
def display_controls():
susp_select
display(topBox)
def search_ip(b):
global anchor
global anchor_type
global expanded_results
anchor = ''
anchor_type = ''
anchor = susp_select.selected_label
anchor_type = susp_select.value
removeWidget(2)
removeWidget(1)
clear_output()
expanded_results = GraphQLClient.request(
query="""query($date:SpotDateType,$dnsQuery:String, $clientIp:SpotIpType){
dns{
threat{details(date:$date,dnsQuery:$dnsQuery,clientIp:$clientIp){
total
clientIp
dnsQuery
}}
}
}""",
variables={
'date': datetime.datetime.strptime(t_date, '%Y%m%d').strftime('%Y-%m-%d'),
'dnsQuery': anchor if anchor_type == 'q' else None,
'clientIp': anchor if anchor_type == 'i' else None
}
)
clear_output()
if not 'errors' in expanded_results:
table = "<table><th>IP</th><th>QUERY</th><th>TOTAL</th>"
for row in expanded_results["data"]["dns"]["threat"]["details"]:
table += "<tr><td class='spot-text-wrapper' data-toggle='tooltip'>"+row["clientIp"]+"</td>\
<td class='spot-text-wrapper' data-toggle='tooltip'>"+row["dnsQuery"]+"</td>\
<td align='center'>"+str(row["total"])+"</td></tr>"
table += "</table>"
result_html_title.value='<h4>Displaying top {0} search results</h4>'.format(top_results)
else:
print "An error ocurred: " + response["errors"][0]["message"]
result_html.value=table
result_html_box.children = [result_html]
display_threat_box(anchor,anchor_type)
resultTableBox.children = [result_html_title, result_html_box]
display(bottomBox)
search_btn.on_click(search_ip)
def display_threat_box(ip,anchor_type):
global query_comments
title =""
text = ""
data_filter = ""
data_filter = "dnsQuery" if anchor_type == 'q' else "clientIp"
title = next((item['title'] for item in query_comments if item.get(data_filter) == ip), "")
text = next((item['text'] for item in query_comments if item.get(data_filter) == ip), "")
result_title.value="<h4 class='spot-text-wrapper spot-text-xlg' data-toggle='tooltip'>Threat summary for " + anchor +"</h4>"
tc_txt_title = widgets.Text(value=title, placeholder='Threat Title', width='100%')
tc_txa_summary = widgets.Textarea(value=text, height=100, width='95%')
tc_btn_save = widgets.Button(description='Save', width='65px', layout='width:100%')
tc_btn_save.button_style = 'primary'
tc_txt_title._css = (
(None, 'width', '95%'),
)
result_summary_container.children = [tc_txt_title, tc_txa_summary]
result_button_container.children=[tc_btn_save]
result_summary_box.children = [result_summary_container, result_button_container]
resultSummaryBox.children = [result_title,result_summary_box]
def save_threat_summary(b):
global anchor
anchor_ip =''
anchor_dns =''
if anchor_type == 'i':
anchor_ip = anchor
elif anchor_type == 'q':
anchor_dns = anchor
if anchor != '':
mutation="""mutation(
$date: SpotDateType,
$dnsQuery:String,
$clientIp:SpotIpType,
$text: String!,
$title: String!,
$threatDetails: [DnsThreatDetailsInputType!]!)
{
dns{
createStoryboard(input:{
threatDetails: $threatDetails,
date: $date,
dnsQuery: $dnsQuery,
clientIp: $clientIp,
title: $title,
text: $text
})
{success}
}
}"""
variables={
'date': datetime.datetime.strptime(t_date, '%Y%m%d').strftime('%Y-%m-%d'),
'dnsQuery': anchor_dns if anchor_type == 'q' else None,
'clientIp': anchor_ip if anchor_type == 'i' else None,
'title': tc_txt_title.value,
'text': tc_txa_summary.value.replace('\n', '\\n'),
'threatDetails': expanded_results['data']['dns']['threat']['details']
}
response = GraphQLClient.request(mutation, variables)
display(Javascript("$('.widget-area > .widget-subarea > .widget-box:gt(0)').remove();"))
response = "Summary successfully saved"
else:
response = "No data selected"
susp_select.selected_label = susp_select.options[0][0]
display(widgets.Box((widgets.HTML(value=response, width='100%'),)))
data_loader()
tc_btn_save.on_click(save_threat_summary)
def file_is_empty(path):
return os.stat(path).st_size==0
def removeWidget(index):
js_command = "$('.widget-area > .widget-subarea > .widget-box:eq({0})').remove();".format(index)
display(Javascript(js_command))
start_investigation()
```
| github_jupyter |
```
#import libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.keras import layers
from keras.layers import Dense , Conv2D , MaxPooling2D , Dropout,Flatten,Convolution2D
from time import perf_counter
import os
sns.set()
batch_size = 100
img_height = 224
img_width = 224
## loading training set
train_data = tf.keras.preprocessing.image_dataset_from_directory(
'/kaggle/input/new-plant-diseases-dataset/new plant diseases dataset(augmented)/New Plant Diseases Dataset(Augmented)/train',
seed=123,
image_size= (img_height, img_width),
batch_size=batch_size
)
## loading validation dataset
val_data = tf.keras.preprocessing.image_dataset_from_directory(
'/kaggle/input/new-plant-diseases-dataset/new plant diseases dataset(augmented)/New Plant Diseases Dataset(Augmented)/valid',
seed=123,
image_size= (img_height, img_width),
batch_size=batch_size)
## loading testing set
test_data = tf.keras.preprocessing.image_dataset_from_directory(
'/kaggle/input/new-plant-diseases-dataset/test',
seed=2,
image_size= (img_height, img_width),
batch_size=batch_size
)
class_names = train_data.class_names
print(class_names)
plt.figure(figsize=(10, 10))
for images, labels in test_data.take(1):
for i in range(9):
ax = plt.subplot(3, 3, i + 1)
plt.imshow(images[i].numpy().astype("uint8"))
plt.title(class_names[labels[i]])
plt.axis("off")
## Defining Cnn
model = tf.keras.models.Sequential([
layers.BatchNormalization(),
layers.Conv2D(32, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Conv2D(64, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.3),
layers.Conv2D(128, 3, activation='relu'),
layers.MaxPooling2D(),
layers.Dropout(0.2),
layers.Flatten(),
layers.Dense(256, activation='relu'),
layers.Dropout(0.15),
layers.Dense(38, activation= 'softmax')
])
#compile model
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.summary()
from keras.utils.vis_utils import plot_model
plot_model(model, to_file='model_plot.png', show_shapes=True, show_layer_names=True)
#to avoid overfitting
early = tf.keras.callbacks.EarlyStopping(monitor='val_loss',patience=5)
## fit model
history=model.fit(train_data,validation_data= val_data,epochs = 20,callbacks=[early])
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
#accuracy plot
plt.plot(epochs, acc, color='green', label='Training Accuracy')
plt.plot(epochs, val_acc, color='blue', label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend()
plt.figure()
plt.show()
#loss plot
plt.plot(epochs, loss, color='green', label='Training Loss')
plt.plot(epochs, val_loss, color='blue', label='Validation Loss')
plt.title('Training and Validation Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
plt.show()
plt.plot(history.history['val_loss'], label = 'training loss')
plt.plot(history.history['val_accuracy'], label = 'training accuracy')
plt.legend()
plt.plot(history.history['loss'], label = 'training loss')
plt.plot(history.history['accuracy'], label = 'training accuracy')
plt.legend()
#evulate model
model.evaluate(val_data)
tf.saved_model.save(model,'model')
model.save('model2.h5')
#predict val data
y_pred = model.predict(val_data)
y_pred = np.argmax(y_pred,axis=1)
print(y_pred)
```
| github_jupyter |
# Converting DICOM Segmentation Objects (DSOs) to NIfTI Format Using dcmqi
This tutorial will teach you how to convert DICOM Segmentation Objects (DSOs) to Nifti format using the "dcmqi" package. DSO is an effecient and flexible format for storing segmented regions and associated metadata in medical imaging data. Nifti is a another popular imaging format that is more limited than DSOs when it comes to metadata and segmentation features, but is well-supported in packages in Python ([nibabel](http://nipy.org/nibabel/)), Matlab ([dicm2nii](https://www.mathworks.com/matlabcentral/fileexchange/42997-dicom-to-nifti-converter--nifti-tool-and-viewer)), and R ([oro.nifti](https://cran.r-project.org/web/packages/oro.nifti/oro.nifti.pdf)). [dcmqi](https://github.com/QIICR/dcmqi) is a library used for quantiative image research, and has many utilities for converting to and from DICOM image formats to other image formats.
Much of this tutorial is lifted directly from the dcmqi documentation pages, which you can find here:
https://qiicr.gitbooks.io/dcmqi-guide/
## Installing dcmqi
To install dcmqi, you should download and unpack the appropriate binary files for your operating system at this link:
https://qiicr.gitbooks.io/dcmqi-guide/quick-start.html
Also, if you are familiar with Docker, you can also download a dcmqi Docker container from that link.
## Converting DSOs
The following instructions are following the documentation located at this link:
https://qiicr.gitbooks.io/dcmqi-guide/user_guide/segimage2itkimage.html
The command we're going to use the segimage2itkimage command. Note that you will need to add you binary folder ("bin") to your system PATH in order for the vanilla command "segimage2itkimage" to work. See these links for more information about how to add folders to your path: [Mac and Linux](https://unix.stackexchange.com/questions/26047/how-to-correctly-add-a-path-to-path), [Windows](https://www.howtogeek.com/118594/how-to-edit-your-system-path-for-easy-command-line-access/). If you don't want to modify your path, you can still run your command by replacing "segimage2itkimage" with the full path to your bin folder, e.g. C:/users/example_user/Software/dcmqi/bin/segimage2itkimage with ".exe" on windows.
Once you have your path figured out, an example command, entered on the command prompt, might look like:
```
segimage2itkimage --outputType nii --prefix my_NII_file --outputDirectory ./Test_Data --inputDICOM ./Test_Data/my_DSO_file.dcm
```
This will take the file located at ./Test_Data/my_DSO_file.dcm and convert it to nifti (\*.nii) format. It will be outputed in the ./Test_Data directory with the filename (a.k.a. prefix), "converted". It will output one Nifti file for each segmentation at each label value of the segmentation. For example, if you have three segmentations with values 1, 2, and 3, then output files will be titled my_NII_file-1.nii, my_NII_file-2.nii, my_NII_file-3.nii. A metadata file with information taken from the original DICOM will also be generated in JSON format, with the filename my_NII_file.json.
## Python Wrapper
I often like to wrap command-line utilities in Python functions, so that I can easily integrate them with existing Python workflows. Below is an example of how you might do so with this dcmqi command, using the subprocess module in Python.
```
import subprocess
def segimage2itkimage(outputType, inputDICOM, outputDirectory, prefix, command_line='C:/Users/abeers/Documents/Software/dcmqi-1.0.5-win64-20171213-e5c3934/bin/segimage2itkimage.exe'):
full_command = ' '.join([command_line, '--outputType', outputType, '--outputDirectory', outputDirectory, '--prefix', prefix, '--inputDICOM', inputDICOM])
subprocess.call(full_command, shell=True)
```
## Converting NIfTI Files to Numpy Arrays using Nibabel
Often, you will want to operate directly on the image data stored in NIfTI files. Below, I will show you how to do this Python using the nibabel package. [nibabel](http://nipy.org/nibabel/) can be installed using the pip package manager in Python by entering the following command on the command prompt:
```
pip install nibabel
```
We can then use the nibabel package to write a short function that will convert incoming nifti files into numpy arrays. See the code below:
```
import nibabel as nib
def nifti_2_numpy(input_filepath):
nifti = nib.load(input_filepath)
return nifti.get_data()
```
nifti_2_numpy return a numpy array of the imaging data from a NIfTI file located at input_filepath. Note that the numpy array returned by this function may not be oriented in the way you expect. To correctly determine the orientation of your array, you must use extract the affine matrix from your NIfTI file. You can do that with a function like this:
```
import nibabel as nib
def nifti_2_numpy_with_affine(input_filepath):
nifti = nib.load(input_filepath)
return nifti.get_data(), nifti.affine
```
nifti_2_numpy_with_affine return a tuple of (data, affine) generated from a NIfTI file located at input_filepath.
And that's all you need to know for this tutorial! If you run into any problems with this tutorial, or the code presented here, please submit an issue on our Github page here: https://github.com/QTIM-Lab/qtim_Tutorials/issues
| github_jupyter |
## Train the model
```
from utils import dataloader
import utils.model
from datetime import datetime
from os.path import join
filter_timestamp = int(datetime(2021, 2, 19, 0).timestamp())
use_user_index = join("indices","train_user_index.parquet")#"train_user_index.parquet"
csv_data_location = join("data","downloaded_data")
model_save_location = join("saved_models","xgb_models_07_validation_check")
dl = dataloader.RecSys2021TSVDataLoader(csv_data_location, use_user_index, mode="train", filter_timestamp=filter_timestamp, verbose=2)
%%time
data = next(iter(dl))
train_data = data[0]
labels = data[1]
recsysxgb = utils.model.RecSysXGB1()
xgb_params = {'objective': 'binary:logistic', 'eval_metric':'map'}
recsysxgb.train_in_memory(train_data, labels, xgb_params, save_dir = model_save_location)
```
## Evaluate the model
```
from utils import dataloader
import utils.model
from datetime import datetime
from os.path import join
filter_timestamp = int(datetime(2021, 2, 19, 0).timestamp())
use_user_index = join("indices","train_user_index.parquet")#"train_user_index.parquet"
csv_data_location = join("data","downloaded_data")
model_save_location = join("saved_models","xgb_models_07_validation_check")
recsysxgb = utils.model.RecSysXGB1(model_save_location)
dl = dataloader.RecSys2021TSVDataLoader(csv_data_location, use_user_index, mode="val", filter_timestamp=filter_timestamp, verbose=2, random_file_sampling=True, load_n_batches=3)
res = recsysxgb.evaluate_validation_set(dl)
for (target, ap, rce) in zip(recsysxgb.targets__, res[0], res[1]):
print(f"{target}: {ap} - {rce}")
dict(sorted(recsysxgb.clfs_["has_like"].get_score(importance_type='gain').items(), key=lambda item: item[1],reverse=True))
```
## Try sample test run
```
import utils.model
import utils.dataloader
dl = utils.dataloader.RecSys2021TSVDataLoader("test", "user_index.parquet", mode="test", load_n_batches=-1)
recsysxgb = utils.model.RecSysXGB1("xgb_models_05_submission")
recsysxgb.evaluate_test_set(testLoader = dl, output_file = "res.csv")
```
## Testing Custom batch sizes
```
from utils import dataloader
from utils import dataloader
import utils.model
from datetime import datetime
from os.path import join
filter_timestamp = None#int(datetime(2021, 2, 19, 0).timestamp())
use_user_index = join("indices","user_index.parquet")#"train_user_index.parquet"
csv_data_location = join("data","test_files")
model_save_location = join("saved_models","xgb_models_06_submission")
dl = dataloader.RecSys2021TSVDataLoader(csv_data_location, use_user_index, mode="test", filter_timestamp=filter_timestamp, load_n_batches=-1, batch_size=1000000, verbose=2, random_file_sampling=True)
r = [a[1] for a in dl]
for df in r:
print(len(df))
import utils.features as fe
import utils.constants as co
import pandas as pd
import numpy as np
current_file = pd.read_csv(
"data/test_files/part-00002.csv",
sep='\x01',
header=None,
names=co.all_features,
dtype={k: v for k, v in co.dtypes_of_features.items() if k in co.all_features}
)
current_file["medias"] = current_file["medias"].fillna("")
current_file["hashtags"] = current_file["hashtags"].fillna("")
current_file["links"] = current_file["links"].fillna("")
current_file["domains"] = current_file["domains"].fillna("")
current_file["medias"] = current_file["medias"].fillna("")
```
| github_jupyter |
# SLU01 - Programming Basics
In this notebook we will be covering the following:
- Introduction to Python
- How to get help
- The `print()` function and its parameters; the Syntax error
- How to comment your code
- Basic data types: integers, floats, strings, booleans and None; the `type()` function
- Converting between basic data types using `int()`, `float()`, `str()` and `bool()` functions; the Value error
- Basic arithmetic operations; the ZeroDivision error
- Operator Precedence
- Strings concatenation and formatting
- Introduction to variables; the Name error
- Constants
## Introduction to Python
Programming is the process of writing instructions (code) to be executed by a computer in order to perform specific tasks. It's almost like teaching a toddler how to ride a bicycle or bake a cake. You break the process into individual steps and explain how each step should be performed in order for the toddler to complete the task successfully.
Take a look at this [chocolate cake recipe](https://www.thespruceeats.com/classic-and-easy-chocolate-cake-recipe-995137), re-written below by a programmer:
<img src="./media/recipe2.jpg"/>
In this recipe we have a set of ingredients that are baked into a cake. The process is broken into 16 distinct steps with specific instructions on how to prepare the ingredients for the next step. If you give this recipe to an english speaking person, he can follow the steps and bake a delicious chocolate cake. In the same way, if you give the computer a recipe (code) that it can understand, it can execute the recipe and return the expected result.
You can think of programming as the act of writing code that a computer can follow to perform a task. The computer simply follows the intructions provided without critical thinking and returns the results to you. If the recipe is well written then the computer returns what you asked. But if the recipe is badly written then the computer [will follow your instructions](https://www.youtube.com/watch?v=cDA3_5982h8) but return something else or say that there was a problem while following the instructions. It is up to you to make sure that the code is well written.
<img src="./media/code_execute_small.jpg"/>
If you have read the recipe above with the atmost attention, you might have noticed that some parts are highlighted with colors: 😄
- The blue parts represent actions/**operations** that the reader should perform.
- The orange parts represent **objects** that can hold the ingredients; like containers. They might be empty, filled with some ingredients or added even more ingredients.
- The yellow parts represent instructions on **how** and/or **when** certain actions/**operations** should be performed. These instructions can be explicitly established by the writer (e.g. `For 10 minutes`) or left for the reader to decide based on certain conditions (e.g. `If cake is cool`).
Basically, most recipes tell you how to process and store the ingredients and provide information on how to control the execution of the dish. Code is similar.
The same way an english speaking person can read the recipe, the computer can read code that it's written in a language that it can understand. Languages that a computer can read and execute are called programming languages. There is an indefinite number of programming languages, each with a different purpose, syntax and philosophy. One such language is Python.
[Python](https://en.wikipedia.org/wiki/Python_(programming_language)) is a general-purpose programming language created by Guido van Rossum and released in 1991. It was designed to have a solid set of basic functionalities built-in and to be highly expandable with the usage of modules and packages, encouraging code re-usage. The Python interpreter and the extensive standard library are freely distributed. Python's design philosophy emphasizes code readability. For these and other reasons, it became a wildly popular language.
## How to get help
While writing code it is natural to encounter pesky errors messages that won't disappear, use modules that are not familiar or outright not knowing how to perform some tasks.
Over the years the Python community has grown significantly. There are a lot of people that collaborate to maintain and improve the Python core, develop new modules to expand the Python functionalities but are also available to help each other solving issues and answering questions.
<img src="./media/99bugs_small.jpg"/>
["A software bug is an error, flaw or fault in a computer program or system that causes it to produce an incorrect or unexpected result (...)"](https://en.wikipedia.org/wiki/Software_bug)
To complement the SLUs you can go to this [wiki](https://wiki.python.org/moin/BeginnersGuide/NonProgrammers) for tons of resources to learn the basics of Python.
One resource that you can use is the documentation of the [Python language](https://docs.python.org/3/).
You can access tutorials, the Python Language Reference and other materials.
If you have specific issues that the documentation does not clarify you can ask the community for help. You can ask questions on the [official Python forum](https://python-forum.io/index.php) or on [Stack Overflow](https://stackoverflow.com/). Stack Overflow is a pretty popular website with Q&A on a lot of programming languages. When searching for a question in the question bar you can use a Python tag [Python] or [python-3.x] to avoid similar questions about another language. If a question was answered many years ago it might be about Python 2 which is now discontinued. Try to confirm that the answers are related to **Python 3** and not Python 2.
In SLU09 - Linear Algebra & NumPy, Part 1, you will be using the [NumPy package](https://numpy.org/). As most popular packages, NumPy has a dedicated documentation page, a getting started page, an examples page and community links. You can use these resources to learn more about the package that you are using.
If everything fails you can always use a search engine to find help. Start your searches with the word "python" to filter out similar questions from other programming languages. If you are encountering an error message, try searching for it in a succint but detailed way. Be as specific as you can.
Another resource that you can use is the Python Enhancement Proposals (PEPs), specially [PEP8](https://www.python.org/dev/peps/pep-0008/). In PEP8 there is a series of recommendations on the style of Python code. It's a great resource to learn how to improve the readability of your code. [PEP20](https://www.python.org/dev/peps/pep-0020/) aka *The Zen of Python* is a document with guidelines used while designing the Python language. The guidelines are explained [here](https://inventwithpython.com/blog/2018/08/17/the-zen-of-python-explained/). Don't worry if you don't understand everything on these documents. Some topics are a bit advanced.
With that out of the way, let's get coding!
<img src="./media/cracking.gif"/>
## The `print()` function and its parameters; the Syntax error
The first built-in function that we are using is the `print()` [function](https://docs.python.org/3.7/library/functions.html#print).
```
print("Hello, World!")
```
The `print()` function sends the argument data to the output cell. In this case, the argument `"Hello, World!"` is a string. Strings are delimited with quotes and the computer will consider them literally and not as code.
The `print()` function can also output virtually all types of data provided by Python, such as integers or floats.
```
print(10)
print(2.3)
```
If you want to call the `print()` function more than once, you **should** put each call in a separate line. While you can write *Compound statements* (having multiple statements in the same line separated by semicolon `;`), this is [discouraged](https://www.python.org/dev/peps/pep-0008/#other-recommendations).
It is advisable to only write **one instruction per line**. Lines can be empty though, i.e. without any instructions. Writing multiple instructions in the same line (without `;`) results in a syntax error as shown below.
```
print(10) print(2.3)
```
A syntax error occurs when the instructions do not follow the rules defined by the language and therefore the Python interpreter is unable to understand the line of code. It is frequently due to typos, incorrect indentation or incorrect arguments. Before executing the code the interpreter verifies the syntax of the code and if there is an error in the syntax, no code is executed.
<img src="./media/hurt_small.png"/>
The instructions are performed in the order that they are written in the code. The previous instructions can all be re-written in the same cell and executed one after the other:
```
print("Hello, World!")
print(10)
print(2.3)
```
---
The `print()` function accepts more than one argument. You can write several values inside the `print()` function separated by commas `,`.
```
print("There are", 5, "continents in the World, according to the UN.")
```
The `print()` function prints the arguments separated with white spaces by default (`sep=" "`). You can change the separation between argument(s) with the `sep` keyword. **After** the arguments that you want to print, add `sep=` and the separator that you want. The separator must be a **string**.
```
print("There are", 5, "continents in the World, according to the UN.",sep="!")
```
Each `print()` call ends with the character newline `\n` (`end="\n"`). This character indicates that a new line is to be started at that point. You can replace this character with the `end` keyword. **After** the argument(s) that you want to print, add `end=` and the ending that you want. The ending must be a **string**.
```
print("Hello, World!", end=" ")
print(10, end=" ")
print(2.3, end=" ")
```
Instead of each `print()` starting in a new line, they have spaces in between. You can change both `sep=` and `end=` if you need. You'll see later that some arguments have default values that are always used unless **explicitly** changed by the programmer.
We will use the `print()` function frequently to check the value of variables during the code execution and to help debugging (finding errors) the code.
---
One additional point regarding the `print()` function. It does **NOT** show the result of the cell. The `print()` only prints to the screen its arguments.
```
print("print() returns the None value. Therefore the printed value is not present in the output cell.")
```
The example above only contains a `print()` statement. The arguments of `print()` are not outputted to the output cell (with `Out [x]:` on the left). They are outputted to the *Standard Output* which, for all intents and purposes, is the space between the input cell (with `In [x]:` on the left) and the output cell.
---
The results of the **last statement of an input cell** are shown on the respective output cell.
```
"This will not appear at all."
"This string is the last result of the cell! On the left is Out[X]:"
```
Above are two statements that return values. Only the last result is outputted to the output cell. The first statement is still executed but the result is not shown on the output cell.
Only showing the last result of the cell is the default behaviour of jupyter notebooks. You can change it but this is outside the scope of this lesson.
In this SLU all the results that we want to see are explicitly printed with the `print()` function. Other instructors might use the last statement of the cell to show the results. These are both valid approaches that you are now aware of.
## How to comment your code
As the programmer writes more and more code, the complexity can increase significantly.
<img src="./media/escalated_quickly_small.png"/>
To relieve some of the complexity we can use comments to document the code. Comments make the code more readable for humans and are ignored by the Python interpreter. They allow to detail what certain pieces of code are for so that other programmers and yourself understand the thought process when you wrote the code. It is unnecessarily time-consuming to review code that has little comments; good programmers use comments as much as possible.
Comments in Python start with the hash character `#` and everything written afterwards in the same line is ignored.
```
#This is a comment.
print("Hello, World!")
```
The comment can be inserted at the end of a line and everything right of `#` is ignored.
```
print("Hello, World!") #Here the string is printed.
```
You can comment code that you don't want the interpreter to execute.
```
print("Hello, World!")
#print("This is not going to be printed!")
```
This is a great way to debug code by "turning off" certain lines of code without having to delete them.
## Basic data types: integers, floats, strings, booleans and None; the `type()` function
Not all data are created equal. Different types of data were developed for specific purposes. We've already seen strings, integers and floats but here we are going to discuss them with a bit more detail.
<img src="./media/letters.gif"/>
### Integers
Integers are numbers that do not contain a fractional part. Negative integers are preceded by a minus sign `-`.
```
print(-12)
```
Positive integers are not required to have a preceding plus sign `+` but you can include it if you want. A preceding plus or minus sign are unary operators because they only operate on a single value.
```
print(+12)
```
### Floats
Floating-point numbers (or commonly called floats) are numbers that do contain a fractional part. The integer part is separated from the fractional part by a decimal point `.`.
```
#Printing float 42.12
print(42.12)
# ^ ^
# | |
# | Decimal part
# Integer part
```
Even though the comma `,` is used in some languages to separate the integer part from the fractional part, in Python the fractional part of a float is declared by using a **single** decimal point `.`. The comma sign has other purposes and **cannot** be used to write a float number. Using it can result in errors or unwanted outcomes.
```
#In this case, using the comma creates two arguments for the print function.
print(42,12) #This prints the integer 42, a whitespace and the integer 12.
print(42.12) #This prints the float 42.12
```
If more than a single decimal point is used, the interpreter will return a syntax error.
```
print(42.000.000)
```
You can omit zero if it is the only digit before or after the decimal point.
```
print(.4)
print(0.4) #Equivalent
print(5.)
print(5.0) #Equivalent
```
One might think that `5.0` and `5` are the exact same thing; for Python these are different numbers:
- `5` is an integer
- `5.0` is a float
It is the decimal point that defines a float. Another way to define a float, especially if very large or very small, is to use the scientific notation. In the scientific notation, numbers with a lot of zeros can be shortened. For example, the number $ 30000000$ can be written in scientific notation as $ 3 \times 10^8$ and thus avoiding writing all the zeros. Try writing $1 \times 10^{50}$ without using the scientific notation. 🙅♂️
In Python, you can write in the scientific notation with the letter `E` (`e` also works). The letter `E` can be translated as "times ten to the power of".
```
print(3E2) #This is equivalent to 3.0 * (10 ** 2).
#Even though both base and exponent are integers, the result is a float.
print(1.2e-4) #This is equivalent to 1.2 * (10 ** -4)
print(1.2e3) #This is equivalent to 1.2 * (10 ** 3)
print(1E50) #This is equivalent to 1. * (10 ** 50)
```
Note that the exponent (the value after `E`) **has to be an integer**; the base (the value before `E`) **may be an integer or a float**. Additionally, the result of using the scientific notation **is always a float** even if the base and exponent are both integers.
As with integers, a float can be negative.
```
print(-12.45)
print(-2.86e4)
```
Python will use the most economical form for representing a number when outputting that number. The value that Python returns to you may have a different representation but it is still the same number.
```
print(0.000000000000001)
```
### Strings
Strings are what we coloquially call text. They can be inclosed inside quotes `"This is a string."` or apostrophes `'This is also a string.'`. When printing, the quotation used is not shown.
```
print("This is a string.")
print('This is also a string.')
```
Using a single quote or apostrophe creates a string that has no end. Python will not be able to find the end of the string and will return a syntax error:
```
#The ) is considered a part of the string
print("This is a )
```
What if you want to include a quote inside a string that is delimited by quotes? Or an apostrophe inside a string delimited by apostrophes? There are two options:
- Use a backslash before the quote/apostrophe to create an escaped character.
```
print("\"An investment in knowledge always pays the best interest.\" - Benjamin Franklin")
print('\'Everything you can imagine is real.\' - Pablo Picasso')
```
- Use apostrophes inside two quotes or quotes inside two apostrophes.
```
print("'Imagination is more important than knowledge...' - Albert Einstein")
print('"There is no harm in doubt and skepticism, for it is through these that new discoveries are made." - Richard Feynman')
```
Including a quote inside two quotes without escaping it creates a smaller string and leaves a leading quote without a closing one which produces a syntax error as can be seen below.
```
print("This is a string" " )
```
---
Strings can be extended to more than a single line. There are two options to achieve this:
- Use the newline character `\n` to introduce a new line.
```
print("old pond\nfrog leaps in\nwater's sound\n- Translated Bashō's \"old pond\"")
#Any space around \n is also part of the string. Try adding a space after \n to see the difference.
```
- Enclose the string in triple quotes `"""`. The ends of lines are automatically included in the string. This can be prevented by adding a `\` at the end of the line.
```
#The \ prevents the name of the poem to be separated from the author. Delete it to see the difference.
print("""A Silly Poem \
by Spike Milligan
Said Hamlet to Ophelia,
I'll draw a sketch of thee,
What kind of pencil shall I use?
2B or not 2B?
"""
)
```
---
A string can also be empty.
```
print('First empty string:')
print("")
print('Second empty string:')
print('')
```
### Booleans
Booleans are a bit more abstract than the above mentioned data types. They represent the value of truthfulness. When asking to check if a number is greater than another, for instance, Python returns a boolean value to indicate if it is true or false.
```
print("Is 2 larger than 1?", 2 > 1)
print("Is 2 smaller than 1?", 2 < 1)
```
Booleans can only be `True` or `False` and are mainly used when controlling the flow of execution as you'll see in great detail in SLU03 - Flow Control.
In Python `True` is equivalent to the integer `1` and `False` is equivalent to `0`. The distinction was created for clarity (Extra fact: the boolean class is a subclass of integers as explained in [PEP285](https://docs.python.org/3/whatsnew/2.3.html#pep-285-a-boolean-type)). Operations that can be done with `0` and `1` can be performed with `True` and `False`.
The bolean values **are not equivalent** to the strings `"True"` and `"False"`, even though they seem the same when printing.
```
#These are not the same.
print(True)
print("True")
```
### The None value
Frequently programming languages have a specific value to mean 'empty' or that 'there is no value here'. In Python that value is `None`. You will see later that a function which does not explicitly return a value will return `None`.
```
print(None)
```
### Identifying the data type with `type()`
When unsure about the data type of a particular value the function `type()` can be use to [determine the data type of its argument](https://docs.python.org/3/library/functions.html#type).
```
#Don't worry about what class means for now. The data type is shortened and within apostrophes.
print("The data type of 1 is",type(1))
print("The data type of 1. is",type(1.))
print("The data type of -3E2 is",type(-3E2))
print("The data type of 'This is a string.' is",type('This is a string.'))
print("The data type of 'True':",type('True'), "is not the same as the data type of True:", type(True))
print("The data type of None is",type(None))
```
<a id="convert"></a>
## Converting between basic data types using `int()`, `float()`, `str()` and `bool()` functions; the Value error
It is sometimes convenient to transform a value from one data type to another. These conversions can be achieved with the functions `int()`, `float()`, `str()` and `bool()`.
The `int()` function converts its argument into an integer. The decimal part of floats is **removed**.
```
print("The result of int(\"4\") is" ,int("4") ,"and the data type is",type(int("4")))
print("The result of int(2.8) is" ,int(2.8) ,"and the data type is",type(int(2.8)))
print("The result of int(True) is" ,int(True) ,"and the data type is",type(int(True)))
```
Some values are not suitable to be converted into an integer and a value error is returned if `int()` is used.
```
print(int("This is not an integer!"))
```
The `float()` function converts its argument into a float.
```
print("The result of float(\"4\") is" ,float("4") ,"and the data type is",type(float("4")))
print("The result of float(2) is" ,float(2) ,"and the data type is",type(float(2)))
print("The result of float(\"3E4\") is" ,float("3E4") ,"and the data type is",type(float("3E4")))
print("The result of float(True) is" ,float(True) ,"and the data type is",type(float(True)))
```
As with `int()`, `float()` returns an error if it cannot convert the argument into a float.
```
print(float("This is not a float!"))
```
The `str()` function converts its argument into a string. Contrary to `int()` and `float()`, `str()` can convert any of the discussed data types into strings.
```
print("The result of str(\"4\") is" ,str("4") ,"and the data type is",type(str("4")))
print("The result of str(2.3) is" ,str(2.3) ,"and the data type is",type(str(2.3)))
print("The result of str(\"3E4\") is" ,str("3E4") ,"and the data type is",type(str("3E4")))
print("The result of str(True) is" ,str(True) ,"and the data type is",type(str(True)))
```
## Basic arithmetic operations; the ZeroDivision error
You can write an arithmetic expression and Python returns the result, just like a calculator!
```
print(2 + 2)
```
<img src="./media/TomsMindBlown.gif"/>
Some of the operations can can be performed with Python are:
- Addition (`+`)
- Subtraction (`-`)
- Multiplication (`*`)
- Division (`/`)
- Exponentiation or power (`**`)
- Integer division (`//`)
- Remainder (`%`)
All these operations are binary because they operate over two values at a time. The negative and positive signs are unary because they only operate over a single value.
The plus sign `+` adds up the value of two numbers.
```
print(2 + 3)
print(2. + 3)
print(2 + 3.)
print(2. + 3.)
```
Pay attention to the data type of the input numbers and the result of the addition. When adding **two integers** the result is **an integer**. If **at least one of the numbers is a float**, then the result is **a float**. We'll call this the *integer vs float* rule.
It is good practice to [leave spaces around binary operators](https://www.python.org/dev/peps/pep-0008/#other-recommendations) to improve the readability of the code.
The minus sign `-` can be used to subtract two numbers. The *integer vs float* also applies.
```
print(2 - 3)
print(2. - 3)
print(2 - 3.)
print(2. - 3.)
```
It is possible to substract a negative number.
```
print(2 - -5)
```
The asterisk sign `*` is used to multiply two numbers. The *integer vs float* also applies.
```
print(2 * 3)
print(2. * 3)
print(2 * 3.)
print(2. * 3.)
```
The slash sign `/` is used to divide a number by another.
```
print(2 / 3)
print(2. / 3)
print(2 / 3.)
print(2. / 3.)
```
Here the previous *integer vs float* does **not** apply. The result of a division is **always a float** even if both numbers are integers and the result could be represented as an integer.
```
print(12 / 6)
```
The double asterisk sign `**` is used to raise a number to the power of another (exponentiation). The *integer vs float* applies.
```
print(2 ** 3)
print(2. ** 3)
print(2 ** 3.)
print(2. ** 3.)
```
The double slash sign `//` is used to perform an integer division of a number by another. It has two main differences from a division:
- The result has no fractional part. It is **rounded to the nearest integer** value that is **less** than the not rounded result.
- The *integer vs float* rule applies.
```
print(6 // 4)
print(6. // 4)
print(6 // 4.)
print(6. // 4.)
```
If the division operator was used, the result would be `1.5`.
```
print(6 / 4)
```
Note that for negative numbers the result is still rounded to the nearest **lesser** integer.
```
#One might think that the result would be -1 or -1.0.
print(-6 // 4)
print(6 // -4.)
```
The percent sign `%` is used to calculate the value left over after an integer division (remainder). The *integer vs float* applies.
```
print(14 % 3)
print(14. % 3)
print(14 % 3.)
print(14. % 3.)
```
This result was obtained by following the sequence of operations:
- Perform an integer division `14 // 3 = 4`
- Multiply the result by the divisor `4 * 3 = 12`
- Substract the result from the dividend `14 - 12 = 2`
Performing a division, integer division or finding the remainder of a division by 0 results in the *ZeroDivisionError* and should be avoided.
```
print(2 // 0)
```
## Operator Precedence
We have dealt with each operator in isolation but what happens when we use them in the same expression? In expressions with multiple operations it is vital to take into account the priority of each operation. Do you remember that multiplications precede additions? The order in which operations are performed is called *Operator Precedence* and can be tought as a hierarchy of priorities.
- Operations with **higher** priority are performed **before** operations with **lower** priority.
- When two operations have the same priority it is the **binding** that determines which is executed first. Most operations in Python have **left-sided binding**. This means that the operations are performed **left** to **right**.
The *Operator Precendence* table for arithmetic operators is:
<img src="./media/precedence_small.jpg"/>
There are additional operations that we'll explore on SLU03 - Flow Control.
For more information and the complete *Operator Precedence* list check [here](https://docs.python.org/3/reference/expressions.html#operator-precedence).
Consider the examples:
```
#The first operation performed: 2 * 3 = 6
#The second operation performed: 2 + 6 = 8
print(2 + 2 * 3)
#First operation performed: 12 // 3 = 4
#Second operation performed: 4 // 2 = 2
#Left-sided binding
print(12 // 3 // 2)
#Introducing parenthesis (highest priority) to force the right operation to be performed first.
#First operation performed: 3 // 2 = 1
#Second operation performed: 12 // 1 = 12
print(12 // (3 // 2))
#// and * have the same priority and are both left-sided binding.
#The first operation performed: 2 // 3 = 0
#The second operation performed: 0 * 3 = 0
print(2 // 3 * 3)
#The first operation performed: 3 * 2 = 6
#The second operation performed: 6 // 3 = 2
print(3 * 2 // 3 )
```
As you can see, the integer division has **left-sided binding** and performs operations from left to right. Changing the order of execution of the operations with parenthesis will (in 99.(9)% of the cases) produce **different results**.
TIP: If your calculations are returning the wrong results make sure that the order of the operations is defined as wanted and the parenthesis are well placed.
One operation that has a **right-sided binding** is the exponentiation.
```
#First operation performed: 2 ** 3 = 8
#Second operation performed: 2 ** 8 = 256
#Right-sided binding
print(2 ** 2 ** 3)
#Introducing parenthesis (highest priority) to force the left operation to be performed first.
#First operation performed: 2 ** 2 = 4
#Second operation performed: 4 ** 3 = 64
print((2 ** 2) ** 3)
```
NOTE: There is an exception for the exponentiation where the positive and negative signs have higher priority if on the right side of `**`.
```
#Here the exponentiation has higher priority than the negative sign, so it is performed first.
print(-2 ** 2)
#Equivalent to
print(- (2 ** 2))
#Here the negative sign is on the right side and has higher priority than the exponentiation, so it is performed first.
print(2 ** -2)
#Equivalent to
print(2 ** (-2))
```
## Strings concatenation and formatting
Operations are not limited to numbers. It is possible to manipulate text with Python.
One such operation is called *string concatenation*. String concatenation combines two strings and merges them into one string. To concatenate two strings in Python the plus sign `+` is used.
```
#The string "Hello " is merged with the string "there."
print("Hello " + "there.")
print("You " + "can " + "merge " + "as " + "many " + "strings " + "in " + "a " + "row " + "as " + "you "+ "like.")
print("The order of the characters " + "is preserved in the resulting string.")
print("String 1 " + "String 2 " + "String 3")
```
You can repeat the same string multiple times with asterisk sign `*` and an integer number.
```
print("""Multiplying one integer with a string produces a string that is the concatenation \
of the string repeated by the value of that integer.
For example: 4 * \"ABC\" results in """ + 4 * "ABC")
```
#### String Formatting
With the knowledge that we got so far, we can add values to a string by calling the function `str()` on the values and concatenate the result into the string.
```
print("There are " + str(123) + " boxes of sweets in a store. There are " + str(25)
+ " sweets in each box.\nHow many sweets are there in the store? " + str(123 * 25))
```
It's quite cumbersome to have to split the string into pieces and concatenate the converted values. There are [several methods to simplify this process](https://www.python.org/dev/peps/pep-3101/#format-strings). We will explore the two most used:
1. The `.format()` method. Strings have a method `.format()` where the arguments are "inserted" inside the string. You can use positional arguments or use keyword arguments. I recommend going [here](https://docs.python.org/3/library/string.html#formatstrings) for additional examples of the capabilities of `.format()`.
```
print("""There are {0} boxes of sweets in a store. There are {1} \
sweets in each box.\nHow many sweets are there in the store? {result}""".format(123, 25, result=123 * 25))
```
2. Using [formatted strings](https://www.python.org/dev/peps/pep-0498/) aka f-strings. f-strings are strings that are prefaced with the letter `f`. In these strings expressions can be introduced using curly braces `{}`. You can see that the expressions have a different color from the string. The notebook takes into account that it is a f-string and automatically shows every expression as code, improving readability. This does not happen with `.format()`.
```
print(f"""There are {123} boxes of sweets in a store. There are {25} \
sweets in each box. How many sweets are in the store? {123 * 25}""")
```
For each case, you should use the method that is simpler to read.
## Introduction to variables; the Name error
Of the operations that we performed in this notebook, we cannot use the results further. The values were calculated but were not stored for later usage. *Variables* are **containers** that allow to **store the values of calculations** and use these values in subsequent operations. A variable is defined by its **name** and its **value**.
To create a variable the programmer must first name it. There are some rules to naming a variable that have to be followed:
- the name of the variable can **only** be composed of **upper- and lower-case letters**, **digits** and the underscore character **_**. The characters are not required to be Latin letters .
- the name has to **start with a letter**.
- the **underscore is considered a letter**.
- the upper- and lower-case letters are considered to be different. `POTATOES` and `potatoes` are two distinct variables.
- the name cannot be one of Python's reserved keywords or [built-in functions](https://docs.python.org/3/library/functions.html#built-in-functions). This rule can "technically" be broken but **shouldn't**. You can, for instance, replace the `print()` with a variable called `print`. Python forgets what `print()` does and all examples in the notebook start producing errors. You'll need to restart the Kernel (top of the notebook: Kernel > Restart) for the `print()` function to be available again.
The reserved keywords are names that have specific purposes in the Python language and **should not be used for naming**. They can be accessed with `help('keywords')`.
There are several [naming conventions](https://www.python.org/dev/peps/pep-0008/#naming-conventions) that you can follow to help you name variables.
```
help('keywords')
```
Here are some examples of names that can be used for variables:
`FirstVariable`
`j`
`v23`
`counter`
`index`
`An_Extremely_Long_Variable_Name_That_You_Are_Definitely_Never_Going_To_Mispell`
`Ovo_da_Páscoa_Abaixo` (use of accented letters)
`В_Советской_России_переменные_дают_вам_имя` (use of non latin characters)
A variable can store any value of the data types above but also many more that we haven't seen yet. The stored value is called the value of the variable and it can change at any given time. Not only the value can change within the same data type but the data type can also change. For instance, a variable can have an integer value and later a float value.
A variable is created when a value is **assigned** to it. If a value is assigned to a variable that does not exist, the variable is created **automatically**.
To create a variable write the **name of the variable**, the **equal sign** `=` and then the **value** that you want to put in the variable. We call this process **variable assignment**. The equal sign `=` is **not** treated as *equal to*, but instead assigns the **right value** to the **left variable**.
```
variable_name = 3
```
The expression above did not produce an output but created the variable `variable_name` with the integer value `3`. To verify the value of the variable, you can use the `print()` function.
```
print(variable_name)
```
The value `3` is stored inside `variable_name` and can be used in later calculations and cells. You can use as many variables as needed to perform the intended tasks.
```
a = 1
b = 2.
c = "This is a string."
print(a,b,c)
```
You cannot use a variable that was not previously assigned. Doing so results in a *NameError*.
```
print(A)
```
The value of a variable can be changed by assigning a new value to the variable.
```
a = 2
#Variable a had value 1 and now it has value 2.
print(a)
```
The right argument of the assignment can be any valid expression that we talked before. The equal sign `=` has lower priority than the above mentioned operators.
```
e = "First part,"
f = " second part."
g = e + f
print(g)
```
Sometimes we want to use the same variable on both sides of the `=` operator.
```
d = 1
d = d + 1
print(d)
```
This expression can be simplified using **shortcut operators**. Shortcut operators allow to write expression like `variable = variable + expression` as `variable =+ expression`. This is valid for the binary operators that we discussed earlier.
```
counter = 1
counter += 1
print(counter)
fraction = 256
fraction /= 2 #Equivalent to fraction = fraction / 2
print(fraction)
money = 1000
tax = 0.05
money *= (1 - tax) #Equivalent to money = money * (1 - tax)
print(money)
```
### Constants
When reviewing code from other programmers or when using modules you might encounter variables with names written in all capital letters with underscores separating words. This is a [convention](https://www.python.org/dev/peps/pep-0008/#constants) to define that variable as a **constant**. You should **avoid** changing the value of constants that you didn't define yourself. You can also use this convention for the same purpose, especially when writing modules.
```
GRAVITY_EARTH = 9.81
PI = 3.14159
```
## Recap
- You now know that programming is like writing the recipe that the computer follows to acomplish a task.
- Even if it seems complicated at first there are a ton of resources available to help.
- By printing or outputting your results and by commenting your code, you'll gradually write better and more complex code.
- You can use different types of data and operations to get the results you want.
- If your calculations are giving unexpected results, check the **operator precedence**.
- And if you don't want to lose the results, you can **assign them to variables** to use later.
| github_jupyter |
##### Copyright 2021 The Cirq Developers
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Heatmaps
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://quantumai.google/cirq/tutorials/heatmaps>"><img src="https://quantumai.google/site-assets/images/buttons/quantumai_logo_1x.png" />View on QuantumAI</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/quantumlib/Cirq/blob/master/docs/tutorials/heatmaps.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/colab_logo_1x.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/quantumlib/Cirq/blob/master/docs/tutorials/heatmaps.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/github_logo_1x.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/Cirq/docs/tutorials/heatmaps.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/download_icon_1x.png" />Download notebook</a>
</td>
</table>
Qubit heatmaps are primarily used for [visualizing calibration metrics](./google/visualizing_calibration_metrics.ipynb) but can be used for any custom data. This tutorial shows how to create a `cirq.Heatmap` for single-qubit data and a `cirq.TwoQubitInteractionHeatmap` for two-qubit data.
```
try:
import cirq
except ImportError:
print("installing cirq...")
!pip install --quiet cirq
print("installed cirq.")
import cirq
```
## Single-qubit heatmaps
You can directly create heatmaps using custom data mapping from a grid qubit tuple (single qubit or qubit pair) to a corresponding float value.
A single-qubit heatmap example is shown below.
```
cirq.Heatmap({
(cirq.GridQubit(0, 0),): 0.1,
(cirq.GridQubit(0, 1),): 0.2,
(cirq.GridQubit(0, 2),): 0.3,
(cirq.GridQubit(1, 0),): 0.4,
}).plot();
```
Additional parameters for the heatmap can be passed as `kwargs` to the constructor, e.g. `plot_colorbar=False` to hide the colorbar. For full details, see the `cirq.Heatmap` reference page.
## Two-qubit heatmaps
Two-qubit heatmaps can be made in an analogous manner using tuples of qubit pairs and corresponding (float) data values.
```
cirq.TwoQubitInteractionHeatmap({
(cirq.GridQubit(0, 0), cirq.GridQubit(0, 1)): 1.1,
(cirq.GridQubit(0, 1), cirq.GridQubit(0, 2)): 1.4,
(cirq.GridQubit(1, 0), cirq.GridQubit(0, 0)): 1.6,
(cirq.GridQubit(3, 3), cirq.GridQubit(3, 2)): 1.9,
}, annotation_format="0.2f", title='Example Two-Qubit Heatmap').plot();
```
These types of plots are used for [visualizing two-qubit calibration metrics](./google/visualizing_calibration_metrics.ipynb).
| github_jupyter |
<a href="https://colab.research.google.com/github/chavgova/My-AI/blob/master/speech_emotion_recognition_12_male.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **SPEECH RECOGNITION**
```
#this is the copy of another projecct and ill make changes to see how i can make it better
import librosa
import librosa.display
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from matplotlib.pyplot import specgram
from matplotlib.axis import Axis
import keras
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Embedding
from keras.layers import LSTM
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.layers import Input, Flatten, Dropout, Activation
from keras.layers import Conv1D, MaxPooling1D, AveragePooling1D
from keras.models import Model
from keras.callbacks import ModelCheckpoint
from sklearn.metrics import confusion_matrix
from keras import regularizers
import os
import pandas as pd
from google.colab import drive
import os
path = '/content/drive/My Drive/My_AI/RawData'
mylist = []
mylist = os.listdir(path)
#print(mylist)
print(mylist[4000:])
print(len(mylist))
```
# LABLES & FEATURES
```
import re
feeling_list=[]
dataset = ''
count = 0
for item in mylist:
file_label = item[6:-16]
try:
file_label = int(file_label)
dataset = 'RAVDESS'
except:
if (item[:1] == 'Y') or (item[:1] == 'O'):
file_label = re.split('_|\.', item)[2]
dataset = 'TESS'
else:
try:
item = item[:-4]
int(item[-3:])
dataset = 'SER_v4'
except:
dataset = 'SAVEE'
if dataset == 'RAVDESS':
if int(item[18:-4])%2==0: #female
if file_label == 1:
feeling_list.append('female_neutral')
elif file_label == 2:
feeling_list.append('female_calm')
elif file_label == 3:
feeling_list.append('female_joy')
elif file_label == 4:
feeling_list.append('female_sadness')
elif file_label == 5:
feeling_list.append('female_anger')
elif file_label == 6:
feeling_list.append('female_fear')
elif file_label == 7:
feeling_list.append('female_disgust')
elif file_label == 8:
feeling_list.append('female_surprise')
else:
if file_label== 1:
feeling_list.append('male_neutral')
elif file_label == 2:
feeling_list.append('male_calm')
elif file_label == 3:
feeling_list.append('male_joy')
elif file_label == 4:
feeling_list.append('male_sadness')
elif file_label == 5:
feeling_list.append('male_anger')
elif file_label == 6:
feeling_list.append('male_fear')
elif file_label == 7:
feeling_list.append('male_disgust')
elif file_label == 8:
feeling_list.append('male_surprise')
elif dataset == 'TESS':
if file_label == 'neutral': feeling_list.append('female_neutral')
elif file_label == 'angry': feeling_list.append('female_anger')
elif file_label == 'disgust': feeling_list.append('female_disgust')
elif file_label == 'ps': feeling_list.append('female_surprise')
elif file_label == 'happy': feeling_list.append('female_joy')
elif file_label == 'sad': feeling_list.append('female_sadness')
elif file_label == 'fear': feeling_list.append('female_fear')
elif dataset == 'SER_v4':
if int(item[-3:])%2 == 1: # male
file_label = item[:-3]
if file_label == 'neutral': feeling_list.append('male_neutral')
elif file_label == 'anger': feeling_list.append('male_anger')
elif file_label == 'disgust': feeling_list.append('male_disgust')
elif file_label == 'surprise': feeling_list.append('male_surprise')
elif file_label == 'happy': feeling_list.append('male_joy')
elif file_label == 'sad': feeling_list.append('male_sadness')
elif file_label == 'fear': feeling_list.append('male_fear')
else:
file_label = item[:-3]
if file_label == 'neutral': feeling_list.append('female_neutral')
elif file_label == 'anger': feeling_list.append('female_anger')
elif file_label == 'disgust': feeling_list.append('female_disgust')
elif file_label == 'surprise': feeling_list.append('female_surprise')
elif file_label == 'happy': feeling_list.append('female_joy')
elif file_label == 'sad': feeling_list.append('female_sadness')
elif file_label == 'fear': feeling_list.append('female_fear')
elif dataset == 'SAVEE':
if item[:1]=='a':
feeling_list.append('male_anger')
elif item[:1]=='f':
feeling_list.append('male_fear')
elif item[:1]=='h':
feeling_list.append('male_joy')
elif item[:1]=='n':
feeling_list.append('male_neutral')
elif item[:2]=='sa':
feeling_list.append('male_sadness')
elif item[:2]=='su':
feeling_list.append('male_surprise')
elif item[:1]=='d':
feeling_list.append('male_disgust')
import pandas as pd
labels = pd.DataFrame(feeling_list)
labels #[2600:2700]
```
Getting the features of audio files using librosa
```
import librosa
import numpy as np
def extract_feature(my_file, **kwargs):
mfcc = kwargs.get("mfcc")
chroma = kwargs.get("chroma")
mel = kwargs.get("mel")
contrast = kwargs.get("contrast")
tonnetz = kwargs.get("tonnetz")
X, sample_rate = librosa.core.load(my_file)
if chroma or contrast:
stft = np.abs(librosa.stft(X))
result = np.array([])
if mfcc:
mfccs = np.mean(librosa.feature.mfcc(y=X, sr=sample_rate, n_mfcc=40).T, axis=0)
result = np.hstack((result, mfccs)) # 40 values
if chroma:
chroma = np.mean(librosa.feature.chroma_stft(S=stft, sr=sample_rate).T,axis=0)
result = np.hstack((result, chroma)) # 12 values
if mel:
mel = np.mean(librosa.feature.melspectrogram(X, sr=sample_rate).T,axis=0)
result = np.hstack((result, mel)) # 128 values
if contrast:
contrast = np.mean(librosa.feature.spectral_contrast(S=stft, sr=sample_rate).T,axis=0)
result = np.hstack((result, contrast)) # 7 values
if tonnetz:
tonnetz = np.mean(librosa.feature.tonnetz(y=librosa.effects.harmonic(X), sr=sample_rate).T,axis=0)
result = np.hstack((result, tonnetz)) # 6 values
return result
#f = os.fspath('/content/drive/My Drive/My_AI/RawData/03-01-08-01-01-02-01.wav')
#a = extract_feature(f, mel=True, mfcc=True, contrast=True, chroma=True, tonnetz=True)
#print(a, a.shape)
data_frame = pd.DataFrame(columns=['all_features'])
bookmark=0
#mylist = mylist[:100]
for index,y in enumerate(mylist):
all_features_ndarray = extract_feature('/content/drive/My Drive/My_AI/RawData/'+ y, mel=True, mfcc=True, contrast=True, chroma=True, tonnetz=True)
data_frame.loc[bookmark] = [all_features_ndarray]
bookmark=bookmark+1
#df[:5] #print
data_frame
data_frame = pd.DataFrame(data_frame['all_features'].values.tolist())
data_frame[:10]
data_frame_labels = pd.concat([data_frame,labels], axis=1)
data_frame_labels = data_frame_labels.rename(index=str, columns={"0": "label"})
data_frame_labels #print
from sklearn.utils import shuffle
data_frame_labels = shuffle(data_frame_labels)
data_frame_labels
#print
```
# SAVE DATASET FEATURES AND LABELS
```
import pickle
with open('/content/drive/My Drive/My_AI/datasets_RAVDESS-TESS-SAVEE-SER_v4_features&labels.pkl', 'wb') as f:
pickle.dump(data_frame_labels, f)
```
# LOAD DATASET FEATURES AND LABELS
```
import pickle
with open('/content/drive/My Drive/My_AI/datasets_RAVDESS-TESS-SAVEE-SER_v4_features&labels.pkl', 'rb') as f:
data_frame_labels = pickle.load(f)
```
# Dividing the data into test and train
```
data_frame_labels.rename(columns={'0': 'lables'}, inplace=True)
data_frame_labels.columns = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40 , 41, 42, 43, 44 , 45 , 46 , 47 , 48 , 49 , 50 , 51 , 52, 53, 54 , 55 , 56 , 57 , 58 , 59 , 60 , 61 , 62 , 63 , 64 , 65 , 66 , 67, 68, 69 , 70 , 71 ,72 , 73 , 74 , 75 , 76 , 77 , 78 , 79 , 80 ,81 , 82 , 83 ,84 , 85 , 86 , 87 , 88 , 89 ,90 , 91 , 92 , 93 , 94 , 95 , 96 , 97 , 98 , 99 ,100 ,101 ,102 ,103 ,104, 105 ,106, 107, 108, 109, 110 ,111, 112 ,113, 114, 115 ,116 ,117 ,118, 119, 120, 121, 122, 123 ,124 ,125 ,126, 127, 128 ,129 ,130 ,131 ,132, 133 ,134, 135 ,136 ,137 ,138 ,139, 140 ,141 ,142 ,143, 144 ,145, 146 ,147 ,148, 149, 150, 151, 152, 153 ,154, 155, 156 ,157 ,158 ,159 ,160, 161 ,162, 163, 164 ,165 ,166 ,167 ,168 ,169 ,170, 171, 172 ,173 ,174 ,175, 176, 177 ,178 ,179 ,180, 181, 182, 183, 184, 185 ,186 ,187 ,188 ,189 ,190 ,191 ,192 , 'lables']
print(data_frame_labels)
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_neutral']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_calm']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_fear']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_surprise']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_joy']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_sadness']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_anger']
data_frame_labels = data_frame_labels[data_frame_labels.lables != 'female_disgust']
#data_frame_labels = data_frame_labels.dropna()
print(data_frame_labels)
data_frame_labels_set = np.random.rand(len(data_frame_labels)) < 0.8
train = data_frame_labels[data_frame_labels_set]
test = data_frame_labels[~data_frame_labels_set]
test[0:20]
trainfeatures = train.iloc[:, :-1]
trainlabel = train.iloc[:, -1:]
testfeatures = test.iloc[:, :-1]
testlabel = test.iloc[:, -1:]
testlabel
from keras.utils import np_utils
from sklearn.preprocessing import LabelEncoder
X_train = np.array(trainfeatures)
y_train = np.array(trainlabel)
X_test = np.array(testfeatures)
y_test = np.array(testlabel)
#print(y_test)
lb = LabelEncoder()
y_train = np_utils.to_categorical(lb.fit_transform(y_train))
y_test = np_utils.to_categorical(lb.fit_transform(y_test))
y_test
X_test
```
Changing dimension for CNN model
```
x_traincnn =np.expand_dims(X_train, axis=2)
x_testcnn= np.expand_dims(X_test, axis=2)
print(x_testcnn)
```
# **MODEL**
```
model = Sequential()
model.add(Conv1D(256, 5,padding='same', input_shape=(193,1)))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same'))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
#model.add(MaxPooling1D(pool_size=(4)))
model.add(Dropout(0.1))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(8))
model.add(Activation('sigmoid'))
opt = tf.keras.optimizers.Adam(learning_rate=0.0001) ###
from tensorflow.python.keras import Sequential, backend
from tensorflow.python.keras.layers import GlobalMaxPool1D, Activation, MaxPool1D, Flatten, Conv1D, Reshape, TimeDistributed, InputLayer
model = Sequential()
backend.clear_session()
lookback = 20
n_features = 5
filters = 128
model.add(Conv1D(256, 5,padding='same', input_shape=(193,1)))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same'))
model.add(Activation('relu'))
model.add(Conv1D(filters, 3, activation = "relu", padding = "causal", dilation_rate = 2**0))
model.add(Conv1D(128, 5, padding='same',))
model.add(Activation('relu'))
model.add(Dropout(0.1))
model.add(Conv1D(128, 5, padding='same',))
model.add(Activation('relu'))
model.add(Conv1D(filters, 3, activation = "relu", padding = "causal", dilation_rate = 2**1))
model.add(Conv1D(128, 5, padding='same',))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Conv1D(filters, 3, activation = "relu", padding = "causal", dilation_rate = 2**2))
model.add(Conv1D(128, 5, padding='same',))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(8))
model.add(Activation('sigmoid'))
opt = tf.keras.optimizers.Adam(learning_rate=0.0001) ###
model.summary()
model.compile(loss= 'categorical_crossentropy', optimizer = opt, metrics=['accuracy'])
cnnhistory = model.fit(x_traincnn, y_train, batch_size = 32, epochs = 100, validation_data = (x_testcnn, y_test))
```
# **PLOTTING**
```
plt.figure()
plt.plot(cnnhistory.history['loss'])
plt.plot(cnnhistory.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.grid(True)
plt.legend(['loss', 'val loss'], loc='upper left')
plt.show()
plt.figure(figsize=(9,5))
plt.plot(cnnhistory.history['loss'], 'm', linewidth=3)
plt.plot(cnnhistory.history['val_loss'], 'c', linewidth=3)
plt.legend(['Loss', 'Validation Loss'], fontsize=13)
plt.xlabel('epochs')
plt.ylabel('loss', fontsize=12)
plt.grid(True)
plt.show()
plt.figure(figsize=(10,6), frameon=True)
plt.plot(cnnhistory.history['accuracy'], 'g', linewidth=3)
plt.plot(cnnhistory.history['val_accuracy'], 'C3', linewidth=3)
plt.title('Model Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy', fontsize=12)
plt.legend(['Accuracy', 'Validation Accuracy'], loc = 'upper left', fontsize=13)
plt.grid(True)
plt.show()
tf.keras.utils.plot_model(
model,
to_file="img_model.png",
show_shapes=False,
show_layer_names=True,
rankdir="TB",
expand_nested=False,
dpi=96,
)
dot_img_file = '/content/drive/My Drive/My_AI/img_model_MALE_causalPaddingWDilation.png'
tf.keras.utils.plot_model(model, to_file = dot_img_file, show_shapes=True)
```
# **SAVING THE MODEL**
```
model_name = 'Emotion_Voice_Detection_CNN_model_12_MALE_100epochs.h5'
path = '/content/drive/My Drive/My_AI/MY MODELS/'
model_path = os.path.join(path, model_name)
model.save(model_path)
print('Saved trained model at %s ' % model_path)
import json
model_json = model.to_json()
with open("/content/drive/My Drive/My_AI/MY MODELS/model_12_MALE_100epochs.json", "w") as json_file:
json_file.write(model_json)
```
# **LOADING THE MODEL**
```
import tensorflow as tf
from keras.models import model_from_json
json_file = open('/content/drive/My Drive/My_AI/MY MODELS/model_12_MALE_100epochs.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("/content/drive/My Drive/My_AI/MY MODELS/Emotion_Voice_Detection_CNN_model_12_MALE_100epochs.h5")
print("Loaded model from disk")
# evaluate loaded model on test data
opt = tf.keras.optimizers.Adam(learning_rate=0.0001) ###
loaded_model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['accuracy'])
score = loaded_model.evaluate(x_testcnn, y_test, verbose=0)
print("%s: %.2f%%" % (loaded_model.metrics_names[1], score[1]*100))
```
# **Predicting emotions on the test data**
```
import pandas as pd
preds = loaded_model.predict(x_testcnn, batch_size=32, verbose=1)
preds1=preds.argmax(axis=1)
abc = preds1.astype(int).flatten()
predictions = (lb.inverse_transform((abc)))
preddf = pd.DataFrame({'predictedvalues': predictions})
actual=y_test.argmax(axis=1)
abc123 = actual.astype(int).flatten()
actualvalues = (lb.inverse_transform((abc123)))
actualdf = pd.DataFrame({'actualvalues': actualvalues})
finaldf = actualdf.join(preddf)
finaldf[10:70]
finaldf.groupby('actualvalues').count()
finaldf.groupby('predictedvalues').count()
finaldf.to_csv('Predictions_12_MALE_100epochs.csv', index=False)
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
classes = finaldf.actualvalues.unique()
classes.sort()
print(classification_report(finaldf.actualvalues, finaldf.predictedvalues, target_names=classes))
import seaborn as sns
def print_confusion_matrix(confusion_matrix, class_names, figsize = (10,7), fontsize=14):
df_cm = pd.DataFrame(
confusion_matrix, index=class_names, columns=class_names,
)
fig = plt.figure(figsize=figsize)
try:
heatmap = sns.heatmap(df_cm, annot=True, fmt="d")
except ValueError:
raise ValueError("Confusion matrix values must be integers.")
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=fontsize)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=45, ha='right', fontsize=fontsize)
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Gender recode function
def gender(row):
if row == 'male_disgust' or 'male_fear' or 'male_joy' or 'male_sadness' or 'male_surprise' or 'male_neutral' or 'male_anger' or 'male_calm':
return 'male'
finaldf = pd.read_csv('Predictions_12_MALE_100epochs.csv')
classes = finaldf.actualvalues.unique()
classes.sort()
# Confusion matrix
c = confusion_matrix(finaldf.actualvalues, finaldf.predictedvalues)
#print(accuracy_score(finaldf.actualvalues, finaldf.predictedvalues))
print_confusion_matrix(c, class_names = classes)
```
# RECORD AUDIO
```
!pip install SpeechRecognition
!pip install pyttsx3
!pip install ffmpeg-python
!sudo apt-get install portaudio19-dev python-pyaudio python3-pyaudio
"""
https://blog.addpipe.com/recording-audio-in-the-browser-using-pure-html5-and-minimal-javascript/
https://stackoverflow.com/a/18650249
https://hacks.mozilla.org/2014/06/easy-audio-capture-with-the-mediarecorder-api/
https://air.ghost.io/recording-to-an-audio-file-using-html5-and-js/
https://stackoverflow.com/a/49019356
"""
from IPython.display import HTML, Audio
from google.colab.output import eval_js
from base64 import b64decode
import numpy as np
from scipy.io.wavfile import read as wav_read
import io
import ffmpeg
AUDIO_HTML = """
<script>
var my_div = document.createElement("DIV");
var my_p = document.createElement("P");
var my_btn = document.createElement("BUTTON");
var t = document.createTextNode("Press to start recording");
my_btn.appendChild(t);
//my_p.appendChild(my_btn);
my_div.appendChild(my_btn);
document.body.appendChild(my_div);
var base64data = 0;
var reader;
var recorder, gumStream;
var recordButton = my_btn;
var handleSuccess = function(stream) {
gumStream = stream;
var options = {
//bitsPerSecond: 8000, //chrome seems to ignore, always 48k
mimeType : 'audio/webm;codecs=opus'
//mimeType : 'audio/webm;codecs=pcm'
};
//recorder = new MediaRecorder(stream, options);
recorder = new MediaRecorder(stream);
recorder.ondataavailable = function(e) {
var url = URL.createObjectURL(e.data);
var preview = document.createElement('audio');
preview.controls = true;
preview.src = url;
document.body.appendChild(preview);
reader = new FileReader();
reader.readAsDataURL(e.data);
reader.onloadend = function() {
base64data = reader.result;
//console.log("Inside FileReader:" + base64data);
}
};
recorder.start();
};
recordButton.innerText = "Recording... press to stop";
navigator.mediaDevices.getUserMedia({audio: true}).then(handleSuccess);
function toggleRecording() {
if (recorder && recorder.state == "recording") {
recorder.stop();
gumStream.getAudioTracks()[0].stop();
recordButton.innerText = "Saving the recording... pls wait!"
}
}
// https://stackoverflow.com/a/951057
function sleep(ms) {
return new Promise(resolve => setTimeout(resolve, ms));
}
var data = new Promise(resolve=>{
//recordButton.addEventListener("click", toggleRecording);
recordButton.onclick = ()=>{
toggleRecording()
sleep(2000).then(() => {
// wait 2000ms for the data to be available...
// ideally this should use something like await...
//console.log("Inside data:" + base64data)
resolve(base64data.toString())
});
}
});
</script>
"""
def get_audio():
display(HTML(AUDIO_HTML))
data = eval_js("data")
binary = b64decode(data.split(',')[1])
process = (ffmpeg
.input('pipe:0')
.output('pipe:1', format='wav')
.run_async(pipe_stdin = True, pipe_stdout = True, pipe_stderr = True, quiet = True, overwrite_output = True)
)
output, err = process.communicate(input=binary)
riff_chunk_size = len(output) - 8
# Break up the chunk size into four bytes, held in b.
q = riff_chunk_size
b = []
for i in range(4):
q, r = divmod(q, 256)
b.append(r)
# Replace bytes 4:8 in proc.stdout with the actual size of the RIFF chunk.
riff = output[:4] + bytes(b) + output[8:]
sr, audio = wav_read(io.BytesIO(riff))
return audio, sr
audio_index = 1
#audio, sr = get_audio()
type(audio)
from scipy.io.wavfile import write
samplerate = 44100; fs = 100
#t = np.linspace(0., 1., samplerate)
#amplitude = np.iinfo(np.int16).max
#data = amplitude * np.sin(2. * np.pi * fs * t)
audio_path = '/content/record_' + str(audio_index)
write(audio_path, samplerate, audio)
audio_index += 1
```
# **VOICE TEST**
```
data, sampling_rate = librosa.load('/content/drive/My Drive/My_AI/Real Voice samples/recording_demo_5')
% pylab inline
import os
import pandas as pd
import librosa
import glob
from librosa import display
plt.figure(figsize=(15, 5))
librosa.display.waveplot(data, sr=sampling_rate)
X, sample_rate = librosa.load('/content/drive/My Drive/My_AI/Real Voice samples/recording_demo_5', res_type='kaiser_fast',duration=2.5,sr=22050*2,offset=0.5)
sample_rate = np.array(sample_rate)
demo_file = os.fspath('/content/drive/My Drive/My_AI/Real Voice samples/recording_demo_5')
features_live = extract_feature(demo_file, mel=True, mfcc=True, contrast=True, chroma=True, tonnetz=True)
features_live = pd.DataFrame(data = features_live)
features_live = features_live.stack().to_frame().T
```
# **EMOTIONS**
```
import torch
features_live_2d = np.expand_dims(features_live, axis=2)
live_preds = loaded_model.predict(features_live_2d, batch_size=32, verbose = 1)
#print(live_preds)
all = np.argsort(-live_preds, axis=1)[:, :8]
for i in all:
print((lb.inverse_transform((i))))
print()
print()
best_n = np.argsort(-live_preds)[:, :3] # best_n = [* * *]
first_second = 0
second_third = 0
for n in best_n:
k = n
num = 1
for k in n:
#print(live_preds[0][n])
#print(k)
#print(n)
if num == 1: first_second = live_preds[0][k] / live_preds[0][n][1]
elif num == 2: second_third = live_preds[0][k] / live_preds[0][n][2]
num += 1
for i in best_n:
print((lb.inverse_transform((i))))
for i in best_n:
first_emo = lb.inverse_transform((i))[0][5:]
second_emo = lb.inverse_transform((i))[1][5:]
third_emo = lb.inverse_transform((i))[2][5:]
emotion = ''
intensity_emo = ''
mixed_emo = ''
if first_second >= 10000:
emotion = first_emo
elif first_second >= 200:
#mix intensity
if first_emo == 'disgust' and second_emo == 'anger': intensity_emo = 'distant'
elif first_emo == 'anger' and second_emo == 'disgust': intensity_emo = 'irritation'
elif first_emo == 'disgust' and second_emo == 'sadness': intensity_emo = 'guilt'
elif first_emo == 'sadness' and second_emo == 'disgust': intensity_emo = 'miserable'
elif first_emo == 'sadness' and second_emo == 'fear': intensity_emo = 'anxious'
elif first_emo == 'fear' and second_emo == 'sadness': intensity_emo = 'desperate'
elif first_emo == 'fear' and second_emo == 'joy': intensity_emo = 'astonished'
elif first_emo == 'joy' and second_emo == 'fear': intensity_emo = 'determined'
#elif first_emo == '' and second_emo == '': intensity_emo = ''
#elif first_emo == '' and second_emo == '': intensity_emo = ''
#elif second_third >= 100:
#mix intensity
if first_second < 200: # in this case the first and second emos are close to equal
# mix
if first_emo == 'disgust' or second_emo == 'disgust':
if first_emo == 'sadness' or second_emo == 'sadness': emotion = 'remorse'
elif first_emo == 'neutral' or second_emo == 'neutral': emotion = 'neutral disgust'
elif first_emo == 'anger' or second_emo == 'anger': emotion = 'contempt'
elif first_emo == 'fear' or second_emo == 'fear': emotion = 'shame'
elif first_emo == 'joy' or second_emo == 'joy': emotion = 'morbidness'
elif first_emo == 'surprise' or second_emo == 'surprise': emotion = 'unbelief'
elif first_emo == 'fear' or second_emo == 'fear':
if first_emo == 'neutral' or second_emo == 'neutral': emotion = 'neutral fear'
elif first_emo == 'joy' or second_emo == 'joy': emotion = 'guilt' #????
elif first_emo == 'sadness' or second_emo == 'sadness': emotion = 'despair'
elif first_emo == 'surprise' or second_emo == 'surprise': emotion = 'awe'
elif first_emo == 'anger' or second_emo == 'anger':
if first_emo == 'neutral' or second_emo == 'neutral': emotion = 'neutral anger'
elif first_emo == 'joy' or second_emo == 'joy': emotion = 'pride'
elif first_emo == 'surprise' or second_emo == 'surprise': emotion = 'outrage'
elif first_emo == 'joy' or second_emo == 'joy':
if first_emo == 'neutral' or second_emo == 'neutral': emotion = 'neutral joy'
elif first_emo == 'surprise' or second_emo == 'surprise': emotion = 'delight'
elif first_emo == 'sadness' or second_emo == 'sadness':
if first_emo == 'neutral' or second_emo == 'neutral': emotion = 'neutral sadness'
elif first_emo == 'surprise' or second_emo == 'surprise': emotion = 'disappointment'
elif first_emo == 'anger' or second_emo == 'anger': emotion = 'envy'
else: emotion = first_emo
if second_third < 200: # in this case the second and third emos are close to equal
# mix
if third_emo == 'disgust' or second_emo == 'disgust':
if third_emo == 'sadness' or second_emo == 'sadness': mixed_emo = 'remorse'
elif third_emo == 'neutral' or second_emo == 'neutral': mixed_emo = 'neutral disgust'
elif third_emo == 'anger' or second_emo == 'anger': mixed_emo = 'contempt'
elif third_emo == 'fear' or second_emo == 'fear': mixed_emo = 'shame'
elif third_emo == 'joy' or second_emo == 'joy': mixed_emo = 'morbidness'
elif third_emo == 'surprise' or second_emo == 'surprise': mixed_emo = 'unbelief'
elif third_emo == 'fear' or second_emo == 'fear':
if third_emo == 'neutral' or second_emo == 'neutral': mixed_emo = 'neutral fear'
elif third_emo == 'joy' or second_emo == 'joy': mixed_emo = 'guilt' #????
elif third_emo == 'sadness' or second_emo == 'sadness': mixed_emo = 'despair'
elif third_emo == 'surprise' or second_emo == 'surprise': mixed_emo = 'awe'
elif third_emo == 'anger' or second_emo == 'anger':
if third_emo == 'neutral' or second_emo == 'neutral': mixed_emo = 'neutral anger'
elif third_emo == 'joy' or second_emo == 'joy': mixed_emo = 'pride'
elif third_emo == 'surprise' or second_emo == 'surprise': mixed_emo = 'outrage'
elif third_emo == 'joy' or second_emo == 'joy':
if third_emo == 'neutral' or second_emo == 'neutral': mixed_emo = 'neutral joy'
elif third_emo == 'surprise' or second_emo == 'surprise': mixed_emo = 'delight'
elif third_emo == 'sadness' or second_emo == 'sadness':
if third_emo == 'neutral' or second_emo == 'neutral': mixed_emo = 'neutral sadness'
elif third_emo == 'surprise' or second_emo == 'surprise': mixed_emo = 'disappointment'
elif third_emo == 'anger' or second_emo == 'anger': mixed_emo = 'envy'
print(first_emo)
print('Main emotion:')
print(emotion)
print('Mixed emotion:')
print(mixed_emo)
print('Emotion by intensity:')
print(intensity_emo)
```
| github_jupyter |
# Criticality Search
This notebook illustrates the usage of the OpenMC Python API's generic eigenvalue search capability. In this Notebook, we will do a critical boron concentration search of a typical PWR pin cell.
To use the search functionality, we must create a function which creates our model according to the input parameter we wish to search for (in this case, the boron concentration).
This notebook will first create that function, and then, run the search.
```
# Initialize third-party libraries and the OpenMC Python API
import matplotlib.pyplot as plt
import numpy as np
import openmc
import openmc.model
%matplotlib inline
```
## Create Parametrized Model
To perform the search we will use the `openmc.search_for_keff` function. This function requires a different function be defined which creates an parametrized model to analyze. This model is required to be stored in an `openmc.model.Model` object. The first parameter of this function will be modified during the search process for our critical eigenvalue.
Our model will be a pin-cell from the [Multi-Group Mode Part II](http://docs.openmc.org/en/latest/examples/mg-mode-part-ii.html) assembly, except this time the entire model building process will be contained within a function, and the Boron concentration will be parametrized.
```
# Create the model. `ppm_Boron` will be the parametric variable.
def build_model(ppm_Boron):
# Create the pin materials
fuel = openmc.Material(name='1.6% Fuel')
fuel.set_density('g/cm3', 10.31341)
fuel.add_element('U', 1., enrichment=1.6)
fuel.add_element('O', 2.)
zircaloy = openmc.Material(name='Zircaloy')
zircaloy.set_density('g/cm3', 6.55)
zircaloy.add_element('Zr', 1.)
water = openmc.Material(name='Borated Water')
water.set_density('g/cm3', 0.741)
water.add_element('H', 2.)
water.add_element('O', 1.)
# Include the amount of boron in the water based on the ppm,
# neglecting the other constituents of boric acid
water.add_element('B', ppm_Boron * 1e-6)
# Instantiate a Materials object
materials = openmc.Materials([fuel, zircaloy, water])
# Create cylinders for the fuel and clad
fuel_outer_radius = openmc.ZCylinder(r=0.39218)
clad_outer_radius = openmc.ZCylinder(r=0.45720)
# Create boundary planes to surround the geometry
min_x = openmc.XPlane(x0=-0.63, boundary_type='reflective')
max_x = openmc.XPlane(x0=+0.63, boundary_type='reflective')
min_y = openmc.YPlane(y0=-0.63, boundary_type='reflective')
max_y = openmc.YPlane(y0=+0.63, boundary_type='reflective')
# Create fuel Cell
fuel_cell = openmc.Cell(name='1.6% Fuel')
fuel_cell.fill = fuel
fuel_cell.region = -fuel_outer_radius
# Create a clad Cell
clad_cell = openmc.Cell(name='1.6% Clad')
clad_cell.fill = zircaloy
clad_cell.region = +fuel_outer_radius & -clad_outer_radius
# Create a moderator Cell
moderator_cell = openmc.Cell(name='1.6% Moderator')
moderator_cell.fill = water
moderator_cell.region = +clad_outer_radius & (+min_x & -max_x & +min_y & -max_y)
# Create root Universe
root_universe = openmc.Universe(name='root universe')
root_universe.add_cells([fuel_cell, clad_cell, moderator_cell])
# Create Geometry and set root universe
geometry = openmc.Geometry(root_universe)
# Instantiate a Settings object
settings = openmc.Settings()
# Set simulation parameters
settings.batches = 300
settings.inactive = 20
settings.particles = 1000
# Create an initial uniform spatial source distribution over fissionable zones
bounds = [-0.63, -0.63, -10, 0.63, 0.63, 10.]
uniform_dist = openmc.stats.Box(bounds[:3], bounds[3:], only_fissionable=True)
settings.source = openmc.source.Source(space=uniform_dist)
# We dont need a tallies file so dont waste the disk input/output time
settings.output = {'tallies': False}
model = openmc.model.Model(geometry, materials, settings)
return model
```
## Search for the Critical Boron Concentration
To perform the search we imply call the `openmc.search_for_keff` function and pass in the relvant arguments. For our purposes we will be passing in the model building function (`build_model` defined above), a bracketed range for the expected critical Boron concentration (1,000 to 2,500 ppm), the tolerance, and the method we wish to use.
Instead of the bracketed range we could have used a single initial guess, but have elected not to in this example. Finally, due to the high noise inherent in using as few histories as are used in this example, our tolerance on the final keff value will be rather large (1.e-2) and the default 'bisection' method will be used for the search.
```
# Perform the search
crit_ppm, guesses, keffs = openmc.search_for_keff(build_model, bracket=[1000., 2500.],
tol=1e-2, print_iterations=True)
print('Critical Boron Concentration: {:4.0f} ppm'.format(crit_ppm))
```
Finally, the `openmc.search_for_keff` function also provided us with `List`s of the guesses and corresponding keff values generated during the search process with OpenMC. Let's use that information to make a quick plot of the value of keff versus the boron concentration.
```
plt.figure(figsize=(8, 4.5))
plt.title('Eigenvalue versus Boron Concentration')
# Create a scatter plot using the mean value of keff
plt.scatter(guesses, [keffs[i].nominal_value for i in range(len(keffs))])
plt.xlabel('Boron Concentration [ppm]')
plt.ylabel('Eigenvalue')
plt.show()
```
We see a nearly linear reactivity coefficient for the boron concentration, exactly as one would expect for a pure 1/v absorber at small concentrations.
| github_jupyter |
### Classification | Data exploration using digits toy dataset
**The digits recognition dataset**
Each sample in this scikit-learn dataset is an 8x8 image representing a handwritten digit. Each pixel is represented by an integer in the range 0 to 16, indicating varying levels of black. Recall that scikit-learn's built-in datasets are of type Bunch, which are dictionary-like objects. Helpfully for the MNIST dataset, scikit-learn provides an 'images' key in addition to the 'data' and 'target' keys.
Because it is a 2D array of the images corresponding to each sample, this 'images' key is useful for visualizing the images. On the other hand, the 'data' key contains the feature array - that is, the images as a flattened array of 64 pixels.
Notice that you can access the keys of these Bunch objects in two different ways: By using the . notation, as in digits.images, or the [] notation, as in digits['images'].
```
# Import necessary modules
from sklearn import datasets
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# Load the digits dataset: digits
digits = datasets.load_digits()
# Print the keys and DESCR of the dataset
print(digits.keys())
print(digits.DESCR)
# Print the shape of the images and data keys
print(digits.images.shape)
print(digits.data.shape)
# Display digit 1010
plt.imshow(digits.images[1010], cmap=plt.cm.gray_r, interpolation='nearest')
plt.show()
```
**Train/Test Split + Fit/Predict/Accuracy**
After creating arrays for the features and target variable, you will split them into training and test sets, fit a k-NN classifier to the training data, and then compute its accuracy using the `.score()` method.
```
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
# Create feature and target arrays
X = digits.data
y = digits.target
```
We create stratified training and test sets using 0.2 for the size of the test set. Use a random state of 42. Stratify the split according to the labels so that they are distributed in the training and test sets as they are in the original dataset.
```
# Split into training and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state=42, stratify=y)
# Create a k-NN classifier with 7 neighbors: knn
knn = KNeighborsClassifier(n_neighbors = 7)
# Fit the classifier to the training data
knn.fit(X_train, y_train)
# Print the accuracy
print(knn.score(X_test, y_test))
```
**Overfitting and underfitting**
Now we compute and plot the training and testing accuracy scores for a variety of different neighbor values. By observing how the accuracy scores differ for the training and testing sets with different values of k, we will develop intuition for overfitting and underfitting.
```
# Setup arrays to store train and test accuracies
neighbors = np.arange(1, 9)
train_accuracy = np.empty(len(neighbors))
test_accuracy = np.empty(len(neighbors))
# Loop over different values of k
for i, k in enumerate(neighbors):
# Setup a k-NN Classifier with k neighbors: knn
knn = KNeighborsClassifier(n_neighbors = k)
# Fit the classifier to the training data
knn.fit(X_train, y_train)
#Compute accuracy on the training set
train_accuracy[i] = knn.score(X_train, y_train)
#Compute accuracy on the testing set
test_accuracy[i] = knn.score(X_test, y_test)
# Generate plot
plt.title('k-NN: Varying Number of Neighbors')
plt.plot(neighbors, test_accuracy, label = 'Testing Accuracy')
plt.plot(neighbors, train_accuracy, label = 'Training Accuracy')
plt.legend()
plt.xlabel('Number of Neighbors')
plt.ylabel('Accuracy')
plt.show()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import os, math
import numpy as np, pandas as pd
import matplotlib.pyplot as plt, seaborn as sns
from tqdm import tqdm, tqdm_notebook
from pathlib import Path
pd.set_option('display.max_columns', 1000)
pd.set_option('display.max_rows', 400)
sns.set()
os.chdir('../..')
from src import utils
DATA = Path('data')
RAW = DATA/'raw'
INTERIM = DATA/'interim'
PROCESSED = DATA/'processed'
SUBMISSIONS = DATA/'submissions'
from src.utils import get_weeks, week_num
week_labels = get_weeks(day_from=20160104, num_weeks=121)[91:]
print(week_labels)
from src.structurednet import get_seqs, shift_right
trade = pd.read_csv(RAW/'Trade.csv', low_memory=False)
challenge = pd.read_csv(RAW/'Challenge_20180423.csv', low_memory=False)
NEURALNET = INTERIM/'neuralnet'
%%time
train = pd.read_feather(NEURALNET/'train_preproc.feather')
val = pd.read_feather(NEURALNET/'val_preproc.feather')
test = pd.read_feather(NEURALNET/'test_preproc.feather')
%%time
import pickle
with open(NEURALNET/'train_seqs.pkl', 'rb') as f:
train_seqs = pickle.load(f)
with open(NEURALNET/'val_seqs.pkl', 'rb') as f:
val_seqs = pickle.load(f)
with open(NEURALNET/'test_seqs.pkl', 'rb') as f:
test_seqs = pickle.load(f)
cat_cols = ['Sector', 'Subsector', 'Region_x', 'Country',
'TickerIdx', 'Seniority', 'Currency', 'ActivityGroup',
'Region_y', 'Activity', 'RiskCaptain', 'Owner',
'IndustrySector', 'IndustrySubgroup', 'MarketIssue', 'CouponType',
'CompositeRatingCat', 'CustomerIdxCat', 'IsinIdxCat', 'BuySellCat']
num_cols = ['ActualMaturityDateKey', 'IssueDateKey', 'IssuedAmount',
'BondDuration', 'BondRemaining', 'BondLife',
'Day', 'CompositeRating', 'BuySellCont',
'DaysSinceBuySell', 'DaysSinceTransaction', 'DaysSinceCustomerActivity',
'DaysSinceBondActivity', 'DaysCountBuySell', 'DaysCountTransaction',
'DaysCountCustomerActivity', 'DaysCountBondActivity', 'SVD_CustomerBias',
'SVD_IsinBuySellBias', 'SVD_Recommend', 'SVD_CustomerFactor00',
'SVD_CustomerFactor01', 'SVD_CustomerFactor02', 'SVD_CustomerFactor03',
'SVD_CustomerFactor04', 'SVD_CustomerFactor05', 'SVD_CustomerFactor06',
'SVD_CustomerFactor07', 'SVD_CustomerFactor08', 'SVD_CustomerFactor09',
'SVD_CustomerFactor10', 'SVD_CustomerFactor11', 'SVD_CustomerFactor12',
'SVD_CustomerFactor13', 'SVD_CustomerFactor14']
id_cols = ['CustomerIdx', 'IsinIdx', 'BuySell']
target_col = 'CustomerInterest'
```
## Model
```
from torch.utils.data import DataLoader
from torch import optim
import torch.nn as nn
from src.structurednet import MultimodalDataset, StructuredNet, train_model
%%time
train_dl = DataLoader(MultimodalDataset(
train[cat_cols], train[num_cols],
train_seqs, train[target_col]),
batch_size=128, shuffle=True)
val_dl = DataLoader(MultimodalDataset(
val[cat_cols], val[num_cols],
val_seqs, val[target_col]),
batch_size=128)
cat_szs = [int(train[col].max() + 1) for col in cat_cols]
emb_szs = [(c, min(50, (c+1)//2)) for c in cat_szs]
USE_CUDA = True
model = StructuredNet(emb_szs, n_cont=len(num_cols), emb_drop=0.2,
szs=[1000,500], drops=[0.5, 0.5],
rnn_hidden_sz=64, rnn_input_sz=4, rnn_n_layers=2,
rnn_drop=0.5)
if USE_CUDA: model = model.cuda()
optimizer = optim.Adam(model.parameters(), lr=1e-3, weight_decay=1e-5)
criterion = nn.BCEWithLogitsLoss()
%%time
model, train_losses, val_losses, val_auc_scores = train_model(
model, train_dl, val_dl, optimizer, criterion,
n_epochs=2, USE_CUDA=USE_CUDA, print_every=800,
val_every=10) # 800,10
```
| github_jupyter |
# Review Questions
**CS1302 Introduction to Computer Programming**
___
```
%reload_ext mytutor
```
## Dictionaries and Sets
**Exercise (Concatenate two dictionaries with precedence)** Define a function `concat_two_dicts` that accepts two arguments of type `dict` such that `concat_two_dicts(a, b)` will return a new dictionary containing all the items in `a` and the items in `b` that have different keys than those in `a`. The input dictionaries should not be mutated.
```
def concat_two_dicts(a, b):
### BEGIN SOLUTION
return {**b, **a}
### END SOLUTION
#tests
a={'x':10, 'z':30}; b={'y':20, 'z':40}
a_copy = a.copy(); b_copy = b.copy()
assert concat_two_dicts(a, b) == {'x': 10, 'z': 30, 'y': 20}
assert concat_two_dicts(b, a) == {'x': 10, 'z': 40, 'y': 20}
assert a == a_copy and b == b_copy
### BEGIN HIDDEN TESTS
a={'x':10, 'z':30}; b={'y':20}
a_copy = a.copy(); b_copy = b.copy()
assert concat_two_dicts(a, b) == {'x': 10, 'z': 30, 'y': 20}
assert concat_two_dicts(b, a) == {'x': 10, 'z': 30, 'y': 20}
assert a == a_copy and b == b_copy
### END HIDDEN TESTS
```
- `{**dict1,**dict2}` creates a new dictionary by unpacking the dictionaries `dict1` and `dict2`.
- By default, `dict2` overwrites `dict1` if they have identical keys.
**Exercise (Count characters)** Define a function `count_characters` which
- accepts a string and counts the numbers of each character in the string, and
- returns a dictionary that stores the results.
```
def count_characters(string):
### BEGIN SOLUTION
counts = {}
for char in string:
counts[char] = counts.get(char, 0) + 1
return counts
### END SOLUTION
# tests
assert count_characters('abcbabc') == {'a': 2, 'b': 3, 'c': 2}
assert count_characters('aababcccabc') == {'a': 4, 'b': 3, 'c': 4}
### BEGIN HIDDEN TESTS
assert count_characters('abcdefgabc') == {'a': 2, 'b': 2, 'c': 2, 'd': 1, 'e': 1, 'f': 1, 'g': 1}
assert count_characters('ab43cb324abc') == {'2': 1, '3': 2, '4': 2, 'a': 2, 'b': 3, 'c': 2}
### END HIDDEN TESTS
```
- Create an empty dictionary `counts`.
- Use a `for` loop to iterate over each character of `string` to count their numbers of occurrences.
- The `get` method of `dict` can initialize the count of a new character before incrementing it.
**Exercise (Count non-Fibonacci numbers)** Define a function `count_non_fibs` that
- accepts a container as an argument, and
- returns the number of items in the container that are not [fibonacci numbers](https://en.wikipedia.org/wiki/Fibonacci_number).
```
def count_non_fibs(container):
### BEGIN SOLUTION
def fib_sequence_inclusive(stop):
Fn, Fn1 = 0, 1
while Fn <= stop:
yield Fn
Fn, Fn1 = Fn1, Fn + Fn1
non_fibs = set(container)
non_fibs.difference_update(fib_sequence_inclusive(max(container)))
return len(non_fibs)
### END SOLUTION
# tests
assert count_non_fibs([0, 1, 2, 3, 5, 8]) == 0
assert count_non_fibs({13, 144, 99, 76, 1000}) == 3
### BEGIN HIDDEN TESTS
assert count_non_fibs({5, 8, 13, 21, 34, 100}) == 1
assert count_non_fibs({0.1, 0}) == 1
### END HIDDEN TESTS
```
- Create a set of Fibonacci numbers up to the maximum of the items in the container.
- Use `difference_update` method of `set` to create a set of items in the container but not in the set of Fibonacci numbers.
**Exercise (Calculate total salaries)** Suppose `salary_dict` contains information about the name, salary, and working time about employees in a company. An example of `salary_dict` is as follows:
```Python
salary_dict = {
'emp1': {'name': 'John', 'salary': 15000, 'working_time': 20},
'emp2': {'name': 'Tom', 'salary': 16000, 'working_time': 13},
'emp3': {'name': 'Jack', 'salary': 15500, 'working_time': 15},
}
```
Define a function `calculate_total` that accepts `salary_dict` as an argument, and returns a `dict` that uses the same keys in `salary_dict` but the total salaries as their values. The total salary of an employee is obtained by multiplying his/her salary and his/her working_time.
E.g.,, for the `salary_dict` example above, `calculate_total(salary_dict)` should return
```Python
{'emp1': 300000, 'emp2': 208000, 'emp3': 232500}.
```
where the total salary of `emp1` is $15000 \times 20 = 300000$.
```
def calculate_total(salary_dict):
### BEGIN SOLUTION
return {
emp: record['salary'] * record['working_time']
for emp, record in salary_dict.items()
}
### END SOLUTION
# tests
salary_dict = {
'emp1': {'name': 'John', 'salary': 15000, 'working_time': 20},
'emp2': {'name': 'Tom', 'salary': 16000, 'working_time': 13},
'emp3': {'name': 'Jack', 'salary': 15500, 'working_time': 15},
}
assert calculate_total(salary_dict) == {'emp1': 300000, 'emp2': 208000, 'emp3': 232500}
### BEGIN HIDDEN TESTS
salary_dict = {
'emp1': {'name': 'John', 'salary': 15000, 'working_time': 20},
'emp2': {'name': 'Tom', 'salary': 16000, 'working_time': 13},
'emp3': {'name': 'Jack', 'salary': 15500, 'working_time': 15},
'emp4': {'name': 'Bob', 'salary': 20000, 'working_time': 10}
}
assert calculate_total(salary_dict) == {'emp1': 300000, 'emp2': 208000, 'emp3': 232500, 'emp4': 200000}
### END HIDDEN TESTS
```
- Use `items` method of `dict` to return the list of key values pairs, and
- use a dictionary comprehension to create the desired dictionary by iterating through the list of items.
**Exercise (Delete items with value 0 in dictionary)** Define a function `zeros_removed` that
- takes a dictionary as an argument,
- mutates the dictionary to remove all the keys associated with values equal to `0`,
- and return `True` if at least one key is removed else `False`.
```
def zeros_removed(d):
### BEGIN SOLUTION
to_delete = [k for k in d if d[k] == 0]
for k in to_delete:
del d[k]
return len(to_delete) > 0
## Memory-efficient but not computationally efficient
# def zeros_removed(d):
# has_deleted = False
# while True:
# for k in d:
# if d[k] == 0:
# del d[k]
# has_deleted = True
# break
# else: return has_deleted
### END SOLUTION
# tests
d = {'a':0, 'b':1, 'c':0, 'd':2}
assert zeros_removed(d) == True
assert zeros_removed(d) == False
assert d == {'b': 1, 'd': 2}
### BEGIN HIDDEN TESTS
d = {'a':0, 'b':1, 'c':0, 'd':2, 'e':0, 'f':'0'}
assert zeros_removed(d) == True
assert zeros_removed(d) == False
assert d == {'b': 1, 'd': 2, 'f':'0'}
### END HIDDEN TESTS
```
- The main issue is that, for any dicionary `d`,
```Python
for k in d:
if d[k] == 0: del d[k]
```
raises the [`RuntimeError: dictionary changed size during iteration`](https://www.geeksforgeeks.org/python-delete-items-from-dictionary-while-iterating/).
- One solution is to duplicate the list of keys, but this is memory inefficient especially when the list of keys is large.
- Another solution is to record the list of keys to delete before the actual deletion. This is memory efficient if the list of keys to delete is small.
**Exercise (Fuzzy search a set)** Define a function `search_fuzzy` that accepts two arguments `myset` and `word` such that
- `myset` is a `set` of `str`s;
- `word` is a `str`; and
- `search_fuzzy(myset, word)` returns `True` if `word` is in `myset` by changing at most one character in `word`, and returns `False` otherwise.
```
def search_fuzzy(myset, word):
### BEGIN SOLUTION
for myword in myset:
if len(myword) == len(word) and len(
[True
for mychar, char in zip(myword, word) if mychar != char]) <= 1:
return True
return False
### END SOLUTION
# tests
assert search_fuzzy({'cat', 'dog'}, 'car') == True
assert search_fuzzy({'cat', 'dog'}, 'fox') == False
### BEGIN HIDDEN TESTS
myset = {'cat', 'dog', 'dolphin', 'rabbit', 'monkey', 'tiger'}
assert search_fuzzy(myset, 'lion') == False
assert search_fuzzy(myset, 'cat') == True
assert search_fuzzy(myset, 'cat ') == False
assert search_fuzzy(myset, 'fox') == False
assert search_fuzzy(myset, 'ccc') == False
### END HIDDEN TESTS
```
- Iterate over each word in `myset`.
- Check whether the length of the word is the same as that of the word in the arguments.
- If the above check passes, use a list comprehension check if the words differ by at most one character.
**Exercise (Get keys by value)** Define a function `get_keys_by_value` that accepts two arguments `d` and `value` where `d` is a dictionary, and returns a set containing all the keys in `d` that have `value` as its value. If no key has the query value `value`, then return an empty set.
```
def get_keys_by_value(d, value):
### BEGIN SOLUTION
return {k for k in d if d[k] == value}
### END SOLUTION
# tests
d = {'Tom':'99', 'John':'88', 'Lucy':'100', 'Lily':'90', 'Jason':'89', 'Jack':'100'}
assert get_keys_by_value(d, '99') == {'Tom'}
### BEGIN HIDDEN TESTS
d = {'Tom':'99', 'John':'88', 'Lucy':'100', 'Lily':'90', 'Jason':'89', 'Jack':'100'}
assert get_keys_by_value(d, '100') == {'Jack', 'Lucy'}
d = {'Tom':'99', 'John':'88', 'Lucy':'100', 'Lily':'90', 'Jason':'89', 'Jack':'100'}
assert get_keys_by_value(d, '0') == set()
### END HIDDEN TESTS
```
- Use set comprehension to create the set of keys whose associated values is `value`.
**Exercise (Count letters and digits)** Define a function `count_letters_and_digits` which
- take a string as an argument,
- returns a dictionary that stores the number of letters and digits in the string using the keys 'LETTERS' and 'DIGITS' respectively.
```
def count_letters_and_digits(string):
### BEGIN SOLUTION
check = {'LETTERS': str.isalpha, 'DIGITS': str.isdigit}
counts = dict.fromkeys(check.keys(), 0)
for char in string:
for t in check:
if check[t](char):
counts[t] += 1
return counts
### END SOLUTION
assert count_letters_and_digits('hello world! 2020') == {'DIGITS': 4, 'LETTERS': 10}
assert count_letters_and_digits('I love CS1302') == {'DIGITS': 4, 'LETTERS': 7}
### BEGIN HIDDEN TESTS
assert count_letters_and_digits('Hi CityU see you in 2021') == {'DIGITS': 4, 'LETTERS': 15}
assert count_letters_and_digits('When a dog runs at you, whistle for him. (Philosopher Henry David Thoreau, 1817-1862)') == {'DIGITS': 8, 'LETTERS': 58}
### END HIDDEN TESTS
```
- Use the class method `fromkeys` of `dict` to initial the dictionary of counts.
**Exercise (Dealers with lowest price)** Suppose `apple_price` is a list in which each element is a `dict` recording the dealer and the corresponding price, e.g.,
```Python
apple_price = [{'dealer': 'dealer_A', 'price': 6799},
{'dealer': 'dealer_B', 'price': 6749},
{'dealer': 'dealer_C', 'price': 6798},
{'dealer': 'dealer_D', 'price': 6749}]
```
Define a function `dealers_with_lowest_price` that takes `apple_price` as an argument, and returns the `set` of dealers providing the lowest price.
```
def dealers_with_lowest_price(apple_price):
### BEGIN SOLUTION
dealers = {}
lowest_price = None
for pricing in apple_price:
if lowest_price == None or lowest_price > pricing['price']:
lowest_price = pricing['price']
dealers.setdefault(pricing['price'], set()).add(pricing['dealer'])
return dealers[lowest_price]
## Shorter code that uses comprehension
# def dealers_with_lowest_price(apple_price):
# lowest_price = min(pricing['price'] for pricing in apple_price)
# return set(pricing['dealer'] for pricing in apple_price
# if pricing['price'] == lowest_price)
### END SOLUTION
# tests
apple_price = [{'dealer': 'dealer_A', 'price': 6799},
{'dealer': 'dealer_B', 'price': 6749},
{'dealer': 'dealer_C', 'price': 6798},
{'dealer': 'dealer_D', 'price': 6749}]
assert dealers_with_lowest_price(apple_price) == {'dealer_B', 'dealer_D'}
### BEGIN HIDDEN TESTS
apple_price = [{'dealer': 'dealer_A', 'price': 6799},
{'dealer': 'dealer_B', 'price': 6799},
{'dealer': 'dealer_C', 'price': 6799},
{'dealer': 'dealer_D', 'price': 6799}]
assert dealers_with_lowest_price(apple_price) == {'dealer_A', 'dealer_B', 'dealer_C', 'dealer_D'}
### END HIDDEN TESTS
```
- Use the class method `setdefault` of `dict` to create a dictionary that maps different prices to different sets of dealers.
- Compute the lowest price at the same time.
- Alternatively, use comprehension to find lowest price and then create the desired set of dealers with the lowest price.
## Lists and Tuples
**Exercise** (Binary addition) Define a function `add_binary` that
- accepts two arguments of type `str` which represent two non-negative binary numbers, and
- returns the binary number in `str` equal to the sum of the two given binary numbers.
```
def add_binary(*binaries):
### BEGIN SOLUTION
def binary_to_decimal(binary):
return sum(2**i * int(b) for i, b in enumerate(reversed(binary)))
def decimal_to_binary(decimal):
return ((decimal_to_binary(decimal // 2) if decimal > 1 else '') +
str(decimal % 2)) if decimal else '0'
return decimal_to_binary(sum(binary_to_decimal(binary) for binary in binaries))
## Alternative 1 using recursion
# def add_binary(bin1, bin2, carry=False):
# if len(bin1) > len(bin2):
# return add_binary(bin2, bin1)
# if bin1 == '':
# return add_binary('1', bin2, False) if carry else bin2
# s = int(bin1[-1]) + int(bin2[-1]) + carry
# return add_binary(bin1[:-1], bin2[:-1], s > 1) + str(s % 2)
## Alternatve 2 using iteration
# def add_binary(a, b):
# answer = []
# n = max(len(a), len(b))
# # fill necessary '0' to the beginning to make a and b have the same length
# if len(a) < n: a = str('0' * (n -len(a))) + a
# if len(b) < n: b = str('0' * (n -len(b))) + b
# carry = 0
# for i in range(n-1, -1, -1):
# if a[i] == '1': carry += 1
# if b[i] == '1': carry += 1
# answer.insert(0, '1') if carry % 2 == 1 else answer.insert(0, '0')
# carry //= 2
# if carry == 1: answer.insert(0, '1')
# answer_str = ''.join(answer) # you can also use "answer_str = ''; for x in answer: answer_str += x"
# return answerastr
### END SOLUTION
# tests
assert add_binary('0', '0') == '0'
assert add_binary('11', '11') == '110'
assert add_binary('101', '101') == '1010'
### BEGIN HIDDEN TESTS
assert add_binary('1111', '10') == '10001'
assert add_binary('111110000011','110000111') == '1000100001010'
### END HIDDEN TESTS
```
- Use comprehension to convert the binary numbers to decimal numbers.
- Use comprehension to convert the sum of the decimal numbers to a binary number.
- Alternatively, perform bitwise addition using a recursion or iteration.
**Exercise (Even-digit numbers)** Define a function `even_digit_numbers`, which finds all numbers between `lower_bound` and `upper_bound` such that each digit of the number is an even number. Please return the numbers as a list.
```
def even_digit_numbers(lower_bound, upper_bound):
### BEGIN SOLUTION
return [
x for x in range(lower_bound, upper_bound)
if not any(int(d) % 2 for d in str(x))
]
### END SOLUTION
# tests
assert even_digit_numbers(1999, 2001) == [2000]
assert even_digit_numbers(2805, 2821) == [2806,2808,2820]
### BEGIN HIDDEN TESTS
assert even_digit_numbers(1999, 2300) == [2000,2002,2004,2006,2008,2020,2022,2024,2026,2028,2040,2042,2044,2046,2048,2060,2062,2064,2066,2068,2080,2082,2084,2086,2088,2200,2202,2204,2206,2208,2220,2222,2224,2226,2228,2240,2242,2244,2246,2248,2260,2262,2264,2266,2268,2280,2282,2284,2286,2288]
assert even_digit_numbers(8801, 8833) == [8802,8804,8806,8808,8820,8822,8824,8826,8828]
assert even_digit_numbers(3662, 4001) == [4000]
### END HIDDEN TESTS
```
- Use list comprehension to generate numbers between the bounds, and
- use comprehension and the `any` function to filter out those numbers containing odd digits.
**Exercise (Maximum subsequence sum)** Define a function `max_subsequence_sum` that
- accepts as an argument a sequence of numbers, and
- returns the maximum sum over nonempty contiguous subsequences.
E.g., when `[-6, -4, 4, 1, -2, 2]` is given as the argument, the function returns `5` because the nonempty subsequence `[4, 1]` has the maximum sum `5`.
```
def max_subsequence_sum(a):
### BEGIN SOLUTION
## see https://en.wikipedia.org/wiki/Maximum_subarray_problem
t = s = 0
for x in a:
t = max(0, t + x)
s = max(s, t)
return s
## Alternative (less efficient) solution using list comprehension
# def max_subsequence_sum(a):
# return max(sum(a[i:j]) for i in range(len(a)) for j in range(i,len(a)+1))
### END SOLUTION
# tests
assert max_subsequence_sum([-6, -4, 4, 1, -2, 2]) == 5
assert max_subsequence_sum([2.5, 1.4, -2.5, 1.4, 1.5, 1.6]) == 5.9
### BEGIN HIDDEN TESTS
seq = [-24.81, 25.74, 37.29, -8.77, 0.78, -15.33, 30.21, 34.94, -40.64, -20.06]
assert round(max_subsequence_sum(seq),2) == 104.86
### BEGIN HIDDEN TESTS
# test of efficiency
assert max_subsequence_sum([*range(1234567)]) == 762077221461
```
- For a list $[a_0,a_1,\dots]$, let
$$
t_k:=\max_{j<k} \sum_{i=j}^{k-1} a_i = \max\{t_{k-1}+a_{k-1},0\},
$$
namely the maximum tail sum of $[a_0,\dots,a_{k-1}]$.
- Then, the maximum subsequence sum of $[a_0,\dots,a_{k-1}]$ is
$$
s_k:=\max_{j\leq k} t_j.
$$
**Exercise (Mergesort)** *For this question, do not use the `sort` method or `sorted` function.*
Define a function called `merge` that
- takes two sequences sorted in ascending orders, and
- returns a sorted list of items from the two sequences.
Then, define a function called `mergesort` that
- takes a sequence, and
- return a list of items from the sequence sorted in ascending order.
The list should be constructed by
- recursive calls to `mergesort` the first and second halves of the sequence individually, and
- merge the sorted halves.
```
def merge(left,right):
### BEGIN SOLUTION
if left and right:
if left[-1] > right[-1]: left, right = right, left
return merge(left,right[:-1]) + [right[-1]]
return list(left or right)
### END SOLUTION
def mergesort(seq):
### BEGIN SOLUTION
if len(seq) <= 1:
return list(seq)
i = len(seq)//2
return merge(mergesort(seq[:i]),mergesort(seq[i:]))
### END SOLUTION
# tests
assert merge([1,3],[2,4]) == [1,2,3,4]
assert mergesort([3,2,1]) == [1,2,3]
### BEGIN HIDDEN TESTS
assert mergesort([3,5,2,4,2,1]) == [1,2,2,3,4,5]
### END HIDDEN TESTS
```
## More Functions
**Exercise (Arithmetic geometric mean)** Define a function `arithmetic_geometric_mean_sequence` which
- takes two floating point numbers `x` and `y` and
- returns a generator that generates the tuple \\((a_n, g_n)\\) where
$$
\begin{aligned}
a_0 &= x, g_0 = y \\
a_n &= \frac{a_{n-1} + g_{n-1}}2 \quad \text{for }n>0\\
g_n &= \sqrt{a_{n-1} g_{n-1}}
\end{aligned}
$$
```
def arithmetic_geometric_mean_sequence(x, y):
### BEGIN SOLUTION
a, g = x, y
while True:
yield a, g
a, g = (a + g)/2, (a*g)**0.5
### END SOLUTION
# tests
agm = arithmetic_geometric_mean_sequence(6,24)
assert [next(agm) for i in range(2)] == [(6, 24), (15.0, 12.0)]
### BEGIN HIDDEN TESTS
agm = arithmetic_geometric_mean_sequence(100,400)
for sol, ans in zip([next(agm) for i in range(5)], [(100, 400), (250.0, 200.0), (225.0, 223.60679774997897), (224.30339887498948, 224.30231718318308), (224.30285802908628, 224.30285802843423)]):
for a, b in zip(sol,ans):
assert round(a,5) == round(b,5)
### END HIDDEN TESTS
```
- Use the `yield` expression to return each tuple of $(a_n,g_n)$ efficiently without redundant computations.
| github_jupyter |
# Training
```
from warnings import simplefilter
simplefilter(action='ignore', category=FutureWarning)
import numpy as np
import os
from natsort import natsorted
import imageio
import time
import tensorflow as tf
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.layers import Dense, Activation, Flatten, Dropout, BatchNormalization
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import ModelCheckpoint, TensorBoard
from tensorflow.keras.models import load_model
from sklearn.metrics import confusion_matrix
from sklearn.metrics import f1_score, accuracy_score
import matplotlib.pyplot as plt
import itertools
import pickle
NAME = 'Cifar10_CNN'
data_dir = 'cifar'
model_dir = 'Models'
num_classes = 10
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship','truck']
class_dict = {
'airplane': 0,
'automobile':1,
'bird':2,
'cat':3,
'deer':4,
'dog':5,
'frog':6,
'horse':7,
'ship':8,
'truck':9
}
inv_class_dict = {v: k for k, v in class_dict.items()}
```
### Load Data
```
x_train0 = np.load('../data/image/X_train.npy')
y_train = np.load('../data/image/y_train.npy')
x_test0 = np.load('../data/image/X_test.npy')
y_test = np.load('../data/image/y_test.npy')
print(x_train0.shape)
print(y_train.shape)
print(x_test0.shape)
print(y_test.shape)
plt.imshow(X_train0[0], interpolation='nearest')
plt.title(inv_class_dict[y_train[0]])
#Visualizing CIFAR 10
fig, axes1 = plt.subplots(5,5,figsize=(8,8))
for j in range(5):
for k in range(5):
i = np.random.choice(range(50000))
axes1[j][k].set_axis_off()
axes1[j][k].set_title(inv_class_dict[y_train[i]])
axes1[j][k].imshow(X_train0[i], interpolation='nearest')
X_train = X_train0/255
X_test = X_test0/255
#Labels to binary
y_train_binary = tf.keras.utils.to_categorical(y_train,num_classes)
y_test_binary = tf.keras.utils.to_categorical(y_test,num_classes)
```
## Training a DNN
```
def create_CNN_model(inp_shape, num_classes, p=0.2):
model = Sequential(name='CNN')
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=inp_shape,
padding='same', name='Conv_1'))
model.add(BatchNormalization(name='Bn_1'))
model.add(Conv2D(32, kernel_size=(3, 3), activation='relu',padding='same', name='Conv_2'))
model.add(BatchNormalization(name='Bn_2'))
model.add(MaxPooling2D(pool_size=(2, 2), name='Max_pool_1'))
model.add(Dropout(p, name='Drop_1'))
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu',padding='same', name='Conv_3'))
model.add(BatchNormalization(name='Bn_3'))
model.add(Conv2D(64, kernel_size=(3, 3), activation='relu',padding='same', name='Conv_4'))
model.add(BatchNormalization(name='Bn_4'))
model.add(MaxPooling2D(pool_size=(2, 2), name='Max_pool_2'))
model.add(Dropout(p, name='Drop_2'))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu',padding='same', name='Conv_5'))
model.add(BatchNormalization(name='Bn_5'))
model.add(Conv2D(128, kernel_size=(3, 3), activation='relu',padding='same', name='Conv_6'))
model.add(BatchNormalization(name='Bn_6'))
model.add(MaxPooling2D(pool_size=(2, 2), name='Max_pool_3'))
model.add(Dropout(p, name='Drop_3'))
model.add(Flatten(name = 'Flatten_1'))
model.add(Dense(32, activation='relu'))
model.add(BatchNormalization(name='Bn_7'))
model.add(Dropout(p, name='Drop_4'))
model.add(Dense(num_classes, name='logits'))
model.add(Activation('softmax', name = 'probs'))
print(model.summary())
return model
def train_CNN_model(model, X_train, y_train, X_val, y_val, model_dir, t, batch_size=256, epochs=50, name = NAME):
model.compile(loss=tf.keras.losses.categorical_crossentropy,
optimizer='adam',
metrics=['accuracy'])
# checkpoint
chk_path = os.path.join(model_dir, 'best_{}_{}'.format(name,t))
checkpoint = ModelCheckpoint(chk_path, monitor='val_acc', verbose=1, save_best_only=True, mode='max')
tensorboard = TensorBoard(log_dir="logs/{}_{}".format(name,t))
callbacks_list = [checkpoint, tensorboard]
history = model.fit(X_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
shuffle=True,
validation_data=(X_val, y_val),
callbacks=callbacks_list)
#Saving the model
model.save(os.path.join(model_dir, 'final_{}_{}'.format(NAME,t)))
return model, history
def calculate_metrics(model, X_test, y_test_binary):
y_pred = np.argmax(model.predict(X_test), axis=1)
y_true = np.argmax(y_test_binary, axis=1)
mismatch = np.where(y_true != y_pred)
cf_matrix = confusion_matrix(y_true, y_pred)
accuracy = accuracy_score(y_true, y_pred)
#micro_f1 = f1_score(y_true, y_pred, average='micro')
macro_f1 = f1_score(y_true, y_pred, average='macro')
return cf_matrix, accuracy, macro_f1, mismatch, y_pred
#CNN Model
model = create_CNN_model(X_train.shape[1:], num_classes, 0.3)
#Training Model
t = int(time.time())
model, H = train_CNN_model(model, X_train, y_train_binary, X_test, y_test_binary, model_dir, t, batch_size=256, epochs=100)
# summarize history for accuracy and loss
plt.figure()
plt.plot(H.history['acc'])
plt.plot(H.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.figure()
plt.plot(H.history['loss'])
plt.plot(H.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
```
### Evaluation
```
#Load Trained Model
model = load_model(model_dir + '/best_Cifar10_CNN_1571866172')
print(model.summary())
cf_matrix, accuracy, macro_f1, mismatch, y_pred = calculate_metrics(model, X_test, y_test_binary)
print('Accuracy : {}'.format(accuracy))
print('F1-score : {}'.format(macro_f1))
print(cf_matrix)
```
| github_jupyter |
# Word embeddings - Word2Vec
### Author: Cecília Assis
#### Github: https://github.com/ceciliassis
#### Linkedin: https://www.linkedin.com/in/ceciliassis/
```
import logging # Setting up the loggings to monitor gensim
logging.basicConfig(format="%(levelname)s - %(asctime)s: %(message)s", datefmt= '%H:%M:%S', level=logging.INFO)
logger = logging.getLogger(__name__)
HEADERS = {
'User-Agent': (
'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_11_5) AppleWebKit/537.36'
'(KHTML, like Gecko) Chrome/50.0.2661.102 Safari/537.36'
)
}
urls = [
'https://take.net/blog/4devs/nlp-processamento-linguagem-natural/',
'https://take.net/blog/4devs/nlp-chatbot/',
'https://take.net/blog/4devs/cursos-para-aprender-sobre-chatbots/',
'https://take.net/blog/inovacao/motivos-usar-inteligencia-artificial/',
'https://take.net/blog/chatbots/algoritmos-inteligencia-artificial-crime/',
'https://take.net/blog/chatbots/inteligencia-artificial-no-blip/',
'https://take.net/blog/devs/trabalhar-com-inteligencia-artificial/',
'https://take.net/blog/inovacao/sucesso-do-cliente-e-inteligencia-artificial/',
'https://take.net/blog/take-test/testes-em-chatbots-com-nlp/',
'https://take.net/blog/chatbots/chatbot/',
'https://take.net/blog/inovacao/big-data/',
'https://take.net/blog/inovacao/big-data-no-trabalho/',
'https://take.net/blog/inovacao/transformacao-digital/'
]
# Define stric set of punctuations due to pre processing and Portuguese characteristics (ex. trata-se)
PUNCTUATIONS = ['*', '+', '.', '/', '[', ']', '(', ')', ';', ':', '%', 'º',
'=', '!', '"','“', "'", "”", "#", '{', '}', ',', '\?']
PUNCTUATIONS_STR = ''.join(PUNCTUATIONS)
import spacy
# Load spacy lang
try:
spacy.load('pt', disable=['ner'])
except OSError:
!python -m spacy download pt
finally:
PT_LANG = spacy.load('pt', disable=['ner'])
# Validade is a word is also a stopword
def is_stopword(word):
return word in NLTK_STOPWORDS_TABLE
import nltk
# Add nltk stopwords to spacy
try:
nltk.corpus.stopwords.words('portuguese')
except LookupError:
nltk.download('stopwords')
finally:
nltk_stopwords = nltk.corpus.stopwords.words('portuguese')
NLTK_STOPWORDS_TABLE = dict.fromkeys(i for i in nltk_stopwords)
stopwords_extension = ['desse', 'dessa', 'disso', 'dessas', 'desses',
'esse', 'essa', 'nisso', 'nessa', 'nesse', 'deste', 'desta']
# Avoid akward spacy word break
for stopword in stopwords_extension:
if not is_stopword(stopword):
NLTK_STOPWORDS_TABLE[stopword] = None
import textacy.preprocessing
import ftfy
from readability import Document
from bs4 import BeautifulSoup
import requests
class Article():
def __init__(self, url):
self.url = url
@property
def html(self):
if not hasattr(self, '_html'):
self._html = self.get_html()
return self._html
@property
def soup(self):
if not hasattr(self, '_soup'):
self._soup = self.get_soup()
return self._soup
@property
def text(self):
if not hasattr(self, '_text'):
self._text = self.get_text()
return self._text
@property
def spacy_doc(self):
if not hasattr(self, '_spacy_doc'):
self._spacy_doc = self.get_spacy_doc()
return self._spacy_doc
def get_html(self):
article = requests.get(self.url, headers=HEADERS)
return article.content
def get_soup(self):
soup = BeautifulSoup(self.html, 'html.parser')
tags_to_remove = ['script', 'style', 'noscript',
'aside', 'footer', 'header', 'div.newsletter']
for tag in soup(tags_to_remove):
tag.decompose()
for tag in soup.find_all(class_='newsletter'):
tag.decompose()
document = Document(soup.encode(formatter="html5"))
soup = BeautifulSoup(document.summary(), 'html.parser')
return soup
def get_text(self):
text = self.soup.get_text("")
text = text.lower()
text = ftfy.fix_text(text)
text = textacy.preprocessing.remove_punctuation(text, marks=PUNCTUATIONS_STR)
text = textacy.preprocessing.replace_numbers(text, replace_with=' ')
text = textacy.preprocessing.normalize_whitespace(text)
text = text.split()
text = ' '.join(word for word in text if not is_stopword(word))
return text
def get_spacy_doc(self):
return textacy.make_spacy_doc(self.text, lang=PT_LANG)
# ------------
def read_pages(urls):
return [Article(url) for url in urls]
```
----
## Build corpora
Build corpora from posts
```
def build_corpus(pages, kwargs):
return [
list(page.spacy_doc._.to_terms_list(**kwargs))
for page in pages
]
from gensim.models.phrases import Phrases, Phraser
from gensim.models import Word2Vec
pages = read_pages(urls)
kwargs = {'as_strings':True, 'normalize': None, 'ngrams':(1)}
corpus = build_corpus(pages, kwargs)
assert len(corpus) == len(pages)
```
## Retrieve unigrams and bigrams from vocab
After building the corpora, the senteces are passed to gensim's Phrases model that detects common phrases and its ngrams.
```
phrases = Phrases(corpus)
sentences = phrases[corpus]
```
## Initialize model
Initialize the Word2Vec model, using the following parameters:
- `size`: 100 (dimensionality of the word vectors)
- `alpha`: 0.025 (initial learning rate)
- `window`: 5 (maximum distance between the current and predicted word within a sentence)
- `seed`: 1 (random generator seed for reproducibility)
- `min_alpha`: 0.0001 (learning rate lower bound)
- `min_count`: 5 (word frequency lower bound)
- `negative`: 5 (if > 0, sets how many “noise words” should be drawn (usually between 5-20))
- `workers` : 1 (how many worker threads available to train the model. Set to 1 for reproducibility purposes)
```
# Init model
w2v_model = Word2Vec(seed=1,
workers=1)
```
## Build model vocab
Buuild the model vocabulary based on the sentences previously created.
```
from time import time
t = time()
w2v_model.build_vocab(sentences)
print(f'Time to build vocab: {round((time() - t) / 60, 2)} mins')
print(f'Vocabulary size: {len(w2v_model.wv.vocab)} word')
```
## Train model
Train the Word2Vec model, using the following parameters:
- `epochs`: 30 (number of epochs, i.e. iterations, over the corpus)
- `report_delay`: 1 (seconds to wait before reporting progress)
- `total_examples`: 13 (corpus length)
```
# Train model
t = time()
w2v_model.train(sentences,
total_examples=w2v_model.corpus_count,
epochs=30,
report_delay=1
)
print(f'Time to train the model: {round((time() - t) / 60, 2)} mins')
# Pre compute L2-normalized vector
# After this call, no more training can be done
w2v_model.wv.init_sims(replace=True)
```
## Vector representation
Since Word2Vec model assignsa vector representation to each word, below are some of them.
```
w2v_model.wv.vocab.keys()
def print_w2v(w2v_model, word):
print(f'Vector of word "{word}"')
print(w2v_model.wv.word_vec(word))
print_w2v(w2v_model, 'nlp')
print_w2v(w2v_model, 'machine_learning')
```
## Most similar
Word embeddings enables us to find similarities along the vector space through calculations such as cosine one. Below we have top 10 similar word for 3 words present in the vocabulary.
```
def top_10_similar(w2v_model, word):
words = w2v_model.wv.most_similar(positive=[word])
words = [w[0] for w in words]
print(f'Top 10 similar words for {word}: {words}')
top_10_similar(w2v_model, 'aprender')
top_10_similar(w2v_model, 'machine_learning')
top_10_similar(w2v_model, 'inteligência_artificial')
top_10_similar(w2v_model, 'nlp')
```
## Analogy difference
Vector representations also enables us to sum and/or subtract words from other, leading to highly informative concepts as shown below.
```
def vector_association(w2v_model, positive_words, negative_word):
words = w2v_model.wv.most_similar(positive=positive_words,
negative=[negative_word],
topn=1)
words = [w[0] for w in words]
print(f'Which word is to "{positive_words[0]}" as "{positive_words[1]}" is to "{negative_word}"? {words}')
vector_association(w2v_model, ['chatbots', 'nlp'],
'machine_learning')
vector_association(w2v_model, ['usuário', 'nlp'],
'inteligência_artificial')
```
## Odd-One-Out
If we want to know the word that deviates the most from a list of terms, we can ask for the most odd one
```
def odd_one_out(w2v_model, words):
word = w2v_model.wv.doesnt_match(words)
print(f"Which word doesn't fit in the list: {words}? {word}")
odd_one_out(w2v_model, ['nlp', 'inteligência_artificial', 'evento'])
```
## t-SNE visualization
t-SNE (t-Distributed Stochastic Neighbor Embedding) is a technique for dimensionality reductition. Since humans are more confortable with spaces up to 3 dimensions, this kind of method helps to see how data fit on the explored vector space.
For more info, see [Visualising high-dimensional datasets using PCA and t-SNE in Python](https://towardsdatascience.com/visualising-high-dimensional-datasets-using-pca-and-t-sne-in-python-8ef87e7915b) and [sklearn.manifold.TSNE](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html).
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
print(f'Current word embedding dimension: {w2v_model.wv.get_vector("nlp").shape}')
def tsne_representation(w2v_model, word):
w2v_labels = [word]
w2v_colors = ['red']
w2v_arrays = np.empty((0,100), dtype='f')
w2v = w2v_model.wv.word_vec("nlp")
w2v_arrays = np.append(w2v_arrays, [w2v], axis=0)
# Add similar words to w2v array
most_similar = w2v_model.wv.most_similar([word])
w2v_colors.extend(['black'] * len(most_similar))
for word_score in most_similar:
w2v_labels.append(word_score[0])
w2v = w2v_model.wv.word_vec(word_score[0])
w2v_arrays = np.append(w2v_arrays, [w2v], axis=0)
# reduce w2v dims
t = time()
pca = PCA(n_components=5).fit_transform(w2v_arrays)
print(f'Time to train PCA: {round((time() - t) / 60, 2)} mins')
t = time()
tsne = TSNE(n_components=2, random_state=0).fit_transform(pca)
print(f'Time to train TSNE: {round((time() - t) / 60, 2)} mins')
tsne_dataframe = pd.DataFrame({'x': [x for x in tsne[:, 0]],
'y': [y for y in tsne[:, 1]],
'word': w2v_labels,
'color': w2v_colors})
# Define plot
fig = plt.figure(figsize=(10, 10))
plot = sns.regplot(data=tsne_dataframe,
x='x',
y='y',
fit_reg=False,
marker='o',
scatter_kws={
's': 40,
'facecolors': tsne_dataframe['color']
})
# Set annotations
for line in range(0, tsne_dataframe.shape[0]):
plt.text(tsne_dataframe['x'][line],
tsne_dataframe['y'][line],
' ' + tsne_dataframe['word'][line],
horizontalalignment='left',
verticalalignment='bottom',
size='medium',
color=tsne_dataframe['color'][line],
weight='normal'
).set_size(15)
plt.title(f't-SNE visualization for {word}')
tsne_representation(w2v_model, 'machine_learning')
tsne_representation(w2v_model, 'linguagem_natural')
```
## Summary
Word embeddings allows us to match words against their context, enabling important discoveries.
The results show that through the recovered blog posts, the term "linguagem_natural" (natural language) is associated with words like "compreensão" (understanding), "inteligência_artificial" (artificial inteligence), "máquinas" (machines) e "pessoas" (people) which demonstrate well current scenario that NLP is inserted into. Another important result comes from the analogies between vocabulary terms, showing that users today are looking for answers and that "create" is "chatbots" as "machine learning" is to "nlp".
Finally, the term "machine_learning" was correctly associated with "aprendizado_máquina" (machine_learning), "redes" (networks), "inteligência_artificial" (artificial inteligence) and "sistemas" (systems), words that are very close to what "machine_learning" represents.
### Thanks to:
- https://www.kaggle.com/pierremegret/gensim-word2vec-tutorial
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.