text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# 0.0 IMPORTS
```
import math
import numpy as np
import pandas as pd
import inflection
import datetime
import seaborn as sns
from matplotlib import pyplot as plt
from IPython.core.display import HTML
from IPython.display import Image
```
## 0.1. Helper Functions
## 0.2. Loading Data
```
df_sales_raw = pd.read_csv( 'data/train.csv', low_memory=False )
df_store_raw = pd.read_csv( 'data/store.csv', low_memory=False )
# merge
df_raw = pd.merge( df_sales_raw, df_store_raw, how='left', on='Store')
df_raw.sample()
```
# 1.0 DESCRIÇÃO DOS DADOS
```
df1 = df_raw.copy()
df_raw.columns
df1.columns
```
## 1.1. Rename Columns
```
cols_old = ['Store', 'DayOfWeek', 'Date', 'Sales', 'Customers', 'Open', 'Promo',
'StateHoliday', 'SchoolHoliday', 'StoreType', 'Assortment', 'CompetitionDistance',
'CompetitionOpenSinceMonth', 'CompetitionOpenSinceYear', 'Promo2', 'Promo2SinceWeek',
'Promo2SinceYear', 'PromoInterval']
snakecase = lambda x: inflection.underscore ( x )
cols_new = list( map( snakecase, cols_old) )
# rename
df1.columns = cols_new
df1.columns
```
## 1.2. Data Dimensions
```
print ( 'Number of Rows: {}' .format( df1.shape[0] ) )
print ( 'Number of Cols: {}' .format( df1.shape[1] ) )
```
## 1.3. Data Types
```
df1['date'] = pd.to_datetime( df1['date'])
df1.dtypes
```
## 1.4. Check NA
```
df1.isna().sum()
```
## 1.5. Fillout NA
```
df1['competition_distance'].max()
df1.sample()
# competition_distance
df1['competition_distance'] = df1['competition_distance'].apply( lambda x: 200000.0 if math.isnan( x ) else x)
# competition_open_since_month
df1['competition_open_since_month'] = df1.apply( lambda x: x['date'].month if math.isnan( x['competition_open_since_month'] )
else x['competition_open_since_month'], axis=1 )
# competition_open_since_year
df1['competition_open_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['competition_open_since_year'] )
else x['competition_open_since_year'], axis=1 )
# promo2
# promo2_since_week
df1['promo2_since_week'] = df1.apply( lambda x: x['date'].week if math.isnan( x['promo2_since_week'] )
else x['promo2_since_week'], axis=1 )
# promo2_since_year
df1['promo2_since_year'] = df1.apply( lambda x: x['date'].year if math.isnan( x['promo2_since_year'] )
else x['promo2_since_year'], axis=1 )
# promo_interval
month_map = {1: 'Jan', 2: 'Fev', 3: 'Mar', 4: 'Apr', 5: 'May', 6: 'Jun', 7: 'Jul', 8: 'Aug', 9: 'Sep', 10: 'Oct', 11: 'Nov', 12: 'Dec' }
df1['promo_interval'].fillna(0, inplace=True )
df1['month_map'] = df1['date'].dt.month.map( month_map )
df1['is_promo'] = df1[['promo_interval', 'month_map']].apply( lambda x: 0 if x['promo_interval'] == 0 else 1 if x['month_map'] in x['promo_interval'].split( ',' ) else 0, axis=1 )
df1.sample(5).T
df1.isna().sum()
```
## 1.6. Change Types
```
df1.dtypes
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype( int )
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype( int )
df1['promo2_since_week'] = df1['promo2_since_week'].astype( int )
df1['promo2_since_year'] = df1['promo2_since_year'].astype( int )
df1.dtypes
```
## 1.7. Descriptive Statistical
```
num_attributes = df1.select_dtypes( include=[ 'int64', 'float64'])
cat_attributes = df1.select_dtypes( exclude=[ 'int64', 'float64', 'datetime64[ns]'])
num_attributes.sample(2)
cat_attributes.sample(2)
```
## 1.7.1 Descriptive Statistical
```
# Central Tendency - mean, median
ct1 = pd.DataFrame( num_attributes.apply( np.mean ) ).T
ct2 = pd.DataFrame( num_attributes.apply( np.median ) ).T
# Dispersion - std, min, max, range, skew, kurtosis
d1 = pd.DataFrame( num_attributes.apply( np.std) ).T
d2 = pd.DataFrame( num_attributes.apply( min ) ).T
d3 = pd.DataFrame( num_attributes.apply( max ) ).T
d4 = pd.DataFrame( num_attributes.apply( lambda x: x.max() - x.min() ) ).T
d5 = pd.DataFrame( num_attributes.apply( lambda x: x.skew() ) ).T
d6 = pd.DataFrame( num_attributes.apply( lambda x: x.kurtosis() ) ).T
# Concatenate
m = pd.concat( [d2, d3, d4, ct1, ct2, d1, d5, d6] ).T.reset_index()
m.columns = ['attributes', 'min', 'max', 'range', 'mean', 'median', 'std', 'skew', 'kurtosis']
m
sns.histplot( df1['sales'] )
sns.distplot( df1['sales'] )
sns.histplot( df1['competition_distance'] )
sns.distplot( df1['competition_distance'] )
sns.histplot( df1['customers'] )
sns.distplot( df1['customers'] )
```
## 1.7.2 Descriptive Statistical
```
cat_attributes.apply( lambda x: x.unique().shape[0] )
aux1 = df1[(df1['state_holiday'] != 0 ) & (df1['sales'] > 0)]
plt.subplot(1, 3, 1)
sns.boxplot( x='state_holiday', y='sales', data=aux1 )
plt.subplot(1, 3, 2)
sns.boxplot( x='store_type', y='sales', data=aux1 )
plt.subplot(1, 3, 3)
sns.boxplot( x='assortment', y='sales', data=aux1 )
aux1 = df1[(df1['state_holiday'] != 0 ) & (df1['sales'] > 0)]
plt.subplot(1, 3, 1)
sns.boxplot( x='state_holiday', y='sales', data=aux1 )
plt.subplot(1, 3, 2)
sns.boxplot( x='store_type', y='sales', data=aux1 )
plt.subplot(1, 3, 3)
sns.boxplot( x='assortment', y='sales', data=aux1 )
```
# 2.0. Passo 02 - Feature Engineering
```
df2 = df1.copy()
```
## 2.1. Mapa Mental de Hipóteses
```
Image( 'img/MindMapHypothesis.png')
```
## 2.1. Criação das Hipóteses
### 2.1.1. Hipóteses Loja
**1.** Lojas com maior quadro de funcionários deveriam vender mais.
**2.** Lojas com maior capacidade de estoque deveriam vender mais.
**3.** Lojas com maior porte deveriam vender mais.
**4.** Lojas com maior sortimento.
**5.** Lojas com competidores mais próximos deveriam vender menos.
**6.** Lojas com competidores à mais tempo deveriam vender mais.
### 2.1.2. Hipóteses Produto
**1.** Lojas que investem mais em marketing deveriam vender mais.
**2.** Lojas com maior exposição de produto deveriam vender mais.
**3.** Lojas com mais produtos com preço menor deveriam vender mais.
**4.** Lojas com promoções mais agressivas (descontos maiores), deveriam vender mais.
**5.** Lojas com promoções ativas por mais tempo deveriam vender mais.
**6.** Lojas com mais dias de promoção deveriam vender mais.
**7.** Lojas com mais promoções consecutivas deveriam vender mais.
### 2.1.3. Hipóteses Tempo
**1.** Lojas abertas durante o feriado de Natal deveriam vender mais
**2.** Lojas deveriam vender mais ao longo dos anos.
**3.** Lojas deveriam vender mais no segundo semestre do ano.
**4.** Lojas deveriam vender mais depois do dia 10 de cada mês.
**5.** Lojas deveriam vender menos aos finais de semana.
**6.** Lojas deveriam vender durante os feriados escolares.
## 2.2. Lista Final de Hipóteses
**1.** Lojas com maior sortimento.
**2.** Lojas com competidores mais próximos deveriam vender menos.
**3.** Lojas com competidores à mais tempo deveriam vender mais.
**4.** Lojas com promoções ativas por mais tempo deveriam vender mais.
**5.** Lojas com mais dias de promoção deveriam vender mais.
**6.** Lojas com mais promoções consecutivas deveriam vender mais.
**7.** Lojas abertas durante o feriado de Natal deveriam vender mais
**8.** Lojas deveriam vender mais ao longo dos anos.
**9.** Lojas deveriam vender mais no segundo semestre do ano.
**10.** Lojas deveriam vender mais depois do dia 10 de cada mês.
**11.** Lojas deveriam vender menos aos finais de semana.
**12.** Lojas deveriam vender durante os feriados escolares.
## 2.3. Feature Engineering
```
# year
df2['year'] = df2['date'].dt.year
# month
df2['month'] = df2['date'].dt.month
# day
df2['day'] = df2['date'].dt.day
# week of year
df2['week_of_year'] = df2['date'].dt.weekofyear
# year week
df2['year'] = df2['date'].dt.strftime( '%Y-%W' )
# competition since
df2['competition_since'] = df2.apply( lambda x: datetime.datetime( year=x['competition_open_since_year'], month=x['competition_open_since_month'],day=1), axis=1)
df2['competition_time_month'] = ( ( df2['date'] - df2['competition_since'] )/30 ).apply(lambda x: x.days ).astype( int )
# promo since
df2['promo_since'] = df2['promo2_since_year'].astype( str ) + '-' + df2['promo2_since_week'].astype( str )
df2['promo_since'] = df2['promo_since'].apply( lambda x: datetime.datetime.strptime( x + '-1', '%Y-%W-%w') - datetime.timedelta( days=7 ) )
df2['promo_time_week'] = ( ( df2['date'] - df2['promo_since'] )/7 ).apply( lambda x: x.days ).astype( int )
# assortiment
df2['assortment'] = df2['assortment'].apply( lambda x: 'basic' if x == 'a' else 'extra' if x == 'b' else 'extended' )
# state holiday
df2['state_holiday'] = df2['state_holiday'].apply( lambda x: 'public_holiday' if x == 'a' else 'easter_holiday' if x == 'b' else 'christimas' if x == 'c' else 'regular_day' )
df2.head().T
```
| github_jupyter |
```
import numpy as np
import pandas as pd
data = pd.read_csv("datasets/pokemon-challenge/pokemon.csv") #data'yı yükleyelim.
data.info() #data hakkında genel bilgi verir.
data.shape #datanın satır ve sütun sayısını verir.
data.columns #sütun isimlerini verir.
data.corr() #data'nın sütunlar arası ilişkiyi verir. (Bu ilişki her zaman doğru olmayabilir.)
data.describe() #data'nın özeliklerini gösterir.
data.head() #ilk 5 datayı gösterir.
data.tail() #son 5 sütunu gösterir.
print data['Type 1'].value_counts(dropna =False) #Type1 sütununa göre kaç adet hangi tür pokenon olduğunu verir.
x = data['Defense']>200 # defense'ı 200 den büyüp pokemonları göstertelim.
data[x]
data[np.logical_and(data['Defense']>200, data['Attack']>100 )] #iki ayrı sütun için koşul ifadesi de kullanılabilir.
data[(data['Defense']>200) & (data['Attack']>100)] #logical_and fonksiyonu yerine and(&) operatörü kullanılabilir.
threshold = sum(data.Speed)/len(data.Speed) #ortalama hızı bulduk.
data["speed_level"] = ["high" if i > threshold else "low" for i in data.Speed] #koşul ifadesi oluşturduk ve speed_level sütununu ekledik.
data.loc[:10,["speed_level","Speed"]] #ilk 10'elemanı ve speed_level, Speed sütunlarını gösterir.
data.loc[10:1:-1,"HP":"Defense"] #Verileri testen göstertelim.
#HP değerlerinin yarısını alalım.
""""
def div(n):
return n/2
data.HP.apply(div)
"""
data.HP.apply(lambda n : n/2)
data= data.set_index("#") #datanın index'ini '#' sütunu ile değiştirelim.
data_new = data.head() # ilk 5 data ile yeni bir data oluşturalım.
data_new
#Attack ve Defense sütunlarını Name sütununa göre birleştirelim.
melted = pd.melt(frame=data_new,id_vars = 'Name', value_vars= ['Attack','Defense'])
melted
melted.pivot(index = 'Name', columns = 'variable',values='value') #vairable değerlerini sütun yapar.
#data name'leri içerisinde tekrarlayan veriler bulunmamalıdır.
#Dataları alt alta birleştirelim.
data1 = data.head()
data2 = data.tail()
conc_data_row = pd.concat([data1,data2],axis =0,ignore_index =True) # axis = 0 : satır.
conc_data_row
#Dataları yan yana birleştirelim.
data1 = data['Attack'].head()
data2 = data['Defense'].head()
conc_data_col = pd.concat([data1,data2],axis =1) # axis = 1 : sütun
conc_data_col
data.dtypes #data type'larını verir.
#Data type'larını değiştirebiliriz.
data['Type 1'] = data['Type 1'].astype('category')
data['Speed'] = data['Speed'].astype('float')
data.dtypes
data.info()
#Data incelendiğinde 800 veri olmasına rağmen Type2 414 değere sahiptir geriye kalan değerlern null değerlerdir.
data["Type 2"].value_counts(dropna =False)
# 386 adet null değer olduğu görülebilir.
data1=data.copy() # datayı kopyalayalım.
data1["Type 2"].dropna(inplace = True) # inplace = True null değerleri NAN olarak atadığımız anlamına gelir.
assert data1['Type 2'].notnull().all() # datada null değer yok sonuç döndürmez.
data2=data.copy()
data2["Type 2"].fillna('empty',inplace = True) # datadaki null değerleri emty ile değiştirelim.
assert data2['Type 2'].notnull().all() # datada null değer yok sonuç döndürmez.
data2 = data.head().copy()
date_list = ["1992-01-10","1992-02-10","1992-03-10","1993-03-15","1993-03-16"] #bir tarih listesi oluşturalım.
datetime_object = pd.to_datetime(date_list) #listeyi date_time a çevirelim.
data2["date"] = datetime_object #datamıza date_time'ı ekleyelim.
data2 = data2.set_index("date") #datamızın index'ini date yapalım.
data2
data2.loc["1993-03-16"] #Bir tarihteki datayı göstertelim.
data2.loc["1992-03-10":"1993-03-16"] #Bir tarih aralığındaki datayı göstertelim.
#"M" = month or "A" = year
data2.resample("A").mean() #Yıllara göre değerlerinin ortalamasını alalım.
data2.resample("M").mean() #Aylara göre değerlerinin ortalamasını alalım.
#datamızda her aya karşılık gelen bir değer olmadığı için bazı aylar için NaN (null) değerini döndürdü.
data2.resample("M").first().interpolate("linear") #İnterpolasyon ile eksik verilerimizi doldurabiliriz.
#Burada eksik olan tarih aralıklarını bir önceki değer ile sonraki değer arasında linear değerler atadı.
#Name Type1 ve Type2 string olduğu için NaN olarak kalır.
data2.resample("M").mean().interpolate("linear") #aylara göre değerlerin ortalamasını aldı ve interpolasyon uyguladı.
import matplotlib.pyplot as plt
%matplotlib inline
# Line Plot
data.Speed.plot(kind = 'line', color = 'g',label = 'Speed',linewidth=1,alpha = 0.5,grid = True,linestyle = ':')
data.Defense.plot(color = 'r',label = 'Defense',linewidth=1, alpha = 0.5,grid = True,linestyle = '-.')
plt.legend(loc='upper right') # legend = etiketleri konumlandırır.
plt.xlabel('x axis') # label = axis isimleri
plt.ylabel('y axis')
plt.title('Line Plot') # title = başlık ismi
plt.show()
data.loc[:,["Attack","Defense","Speed"]].plot()
# subplots
data.plot(subplots = True)
plt.show()
# Scatter Plot
data.plot(kind='scatter', x='Attack', y='Defense',alpha = 0.5,color = 'blue')
plt.xlabel('Attack')
plt.ylabel('Defence')
plt.title('Attack Defense Scatter Plot')
# hist plot
data.plot(kind = "hist",y = "Defense",bins = 50,range= (0,250),density = True,figsize = (6,6))
plt.show()
# histogram subplot with non cumulative and cumulative
fig, axes = plt.subplots(nrows=2,ncols=1)
data.plot(kind = "hist",y = "Defense",bins = 50,range= (0,250),density = True,ax = axes[0])
data.plot(kind = "hist",y = "Defense",bins = 50,range= (0,250),density = True,ax = axes[1],cumulative = True) #Kümülatif histogram.
#plt.savefig('graph.png') #grafiği kaydeder.
plt.show()
data.Speed.plot(kind = 'hist',bins = 50)
plt.clf() #plot'u temizler.
#pokemonların ataklarının Legendary özelliğine göre karşılaştırılması.
# Üstteki siyah çizgi max
# Üstteki mavi çizgi% 75
# Yeşil çizgi ortancadır (% 50)
# Alttaki mavi çizgi% 25
# Alttaki siyah çizgi min
# daireler ise ayrık noktaları oluşturur.
# örneğin [1 1 1 1000] gibi bir dizide ayrık nokta 1000'dir.
data.boxplot(column='Attack',by = 'Legendary')
plt.show()
import seaborn as sns # visualization tool
#correlation map
f,ax = plt.subplots(figsize=(18, 18))
sns.heatmap(data.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax)
plt.show()
```
## Referans Link
### https://www.kaggle.com/kanncaa1/data-sciencetutorial-for-beginners/
| github_jupyter |
# CIFAR-10, CIFAR-100 dataset introduction
CIFAR-10 and CIFAR-100 are the small image datasets with its classification labeled. It is widely used for easy image classification task/benchmark in research community.
Official page: [CIFAR-10 and CIFAR-100 datasets](https://www.cs.toronto.edu/~kriz/cifar.html)
In Chainer, CIFAR-10 and CIFAR-100 dataset can be obtained with build-in function.
```
from __future__ import print_function
import os
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
import chainer
basedir = './src/cnn/images'
```
## CIFAR-10
`chainer.datasets.get_cifar10` method is prepared in Chainer to get CIFAR-10 dataset.
Dataset is automatically downloaded from https://www.cs.toronto.edu only for the first time, and its cache is used from second time.
```
CIFAR10_LABELS_LIST = [
'airplane',
'automobile',
'bird',
'cat',
'deer',
'dog',
'frog',
'horse',
'ship',
'truck'
]
train, test = chainer.datasets.get_cifar10()
```
The dataset structure is quite same with MNIST dataset, it is `TupleDataset`.
`train[i]` represents i-th data, there are 50000 training data.
test data structure is same, with 10000 test data.
```
print('len(train), type ', len(train), type(train))
print('len(test), type ', len(test), type(test))
```
`train[i]` represents i-th data, type=tuple $(x_i, y_i)$, where $ x_i $ is image data and $ y_i $ is label data.
`train[i][0]` represents $x_i$, CIFAR-10 image data,
this is 3 dimensional array, (3, 32, 32), which represents RGB channel, width 32 px, height 32 px respectively.
`train[i][1]` represents $y_i$, the label of CIFAR-10 image data (scalar),
this is scalar value whose actual label can be converted by `LABELS_LIST`.
Let's see 0-th data, `train[0]`, in detail.
```
print('train[0]', type(train[0]), len(train[0]))
x0, y0 = train[0]
print('train[0][0]', x0.shape, x0)
print('train[0][1]', y0.shape, y0, '->', CIFAR10_LABELS_LIST[y0])
def plot_cifar(filepath, data, row, col, scale=3., label_list=None):
fig_width = data[0][0].shape[1] / 80 * row * scale
fig_height = data[0][0].shape[2] / 80 * col * scale
fig, axes = plt.subplots(row,
col,
figsize=(fig_height, fig_width))
for i in range(row * col):
# train[i][0] is i-th image data with size 32x32
image, label_index = data[i]
image = image.transpose(1, 2, 0)
r, c = divmod(i, col)
axes[r][c].imshow(image) # cmap='gray' is for black and white picture.
if label_list is None:
axes[r][c].set_title('label {}'.format(label_index))
else:
axes[r][c].set_title('{}: {}'.format(label_index, label_list[label_index]))
axes[r][c].axis('off') # do not show axis value
plt.tight_layout() # automatic padding between subplots
plt.savefig(filepath)
plot_cifar(os.path.join(basedir, 'cifar10_plot.png'), train, 4, 5,
scale=4., label_list=CIFAR10_LABELS_LIST)
plot_cifar(os.path.join(basedir, 'cifar10_plot_more.png'), train, 10, 10,
scale=4., label_list=CIFAR10_LABELS_LIST)
```
## CIFAR-100
CIFAR-100 is really similar to CIFAR-10. The difference is the number of classified label is 100.
`chainer.datasets.get_cifar100` method is prepared in Chainer to get CIFAR-100 dataset.
```
CIFAR100_LABELS_LIST = [
'apple', 'aquarium_fish', 'baby', 'bear', 'beaver', 'bed', 'bee', 'beetle',
'bicycle', 'bottle', 'bowl', 'boy', 'bridge', 'bus', 'butterfly', 'camel',
'can', 'castle', 'caterpillar', 'cattle', 'chair', 'chimpanzee', 'clock',
'cloud', 'cockroach', 'couch', 'crab', 'crocodile', 'cup', 'dinosaur',
'dolphin', 'elephant', 'flatfish', 'forest', 'fox', 'girl', 'hamster',
'house', 'kangaroo', 'keyboard', 'lamp', 'lawn_mower', 'leopard', 'lion',
'lizard', 'lobster', 'man', 'maple_tree', 'motorcycle', 'mountain', 'mouse',
'mushroom', 'oak_tree', 'orange', 'orchid', 'otter', 'palm_tree', 'pear',
'pickup_truck', 'pine_tree', 'plain', 'plate', 'poppy', 'porcupine',
'possum', 'rabbit', 'raccoon', 'ray', 'road', 'rocket', 'rose',
'sea', 'seal', 'shark', 'shrew', 'skunk', 'skyscraper', 'snail', 'snake',
'spider', 'squirrel', 'streetcar', 'sunflower', 'sweet_pepper', 'table',
'tank', 'telephone', 'television', 'tiger', 'tractor', 'train', 'trout',
'tulip', 'turtle', 'wardrobe', 'whale', 'willow_tree', 'wolf', 'woman',
'worm'
]
train_cifar100, test_cifar100 = chainer.datasets.get_cifar100()
```
The dataset structure is quite same with MNIST dataset, it is `TupleDataset`.
`train[i]` represents i-th data, there are 50000 training data.
Total train data is same size while the number of class label increased.
So the training data for each class label is fewer than CIFAR-10 dataset.
test data structure is same, with 10000 test data.
```
print('len(train_cifar100), type ', len(train_cifar100), type(train_cifar100))
print('len(test_cifar100), type ', len(test_cifar100), type(test_cifar100))
print('train_cifar100[0]', type(train_cifar100[0]), len(train_cifar100[0]))
x0, y0 = train_cifar100[0]
print('train_cifar100[0][0]', x0.shape) # , x0
print('train_cifar100[0][1]', y0.shape, y0)
plot_cifar(os.path.join(basedir, 'cifar100_plot_more.png'), train_cifar100,
10, 10, scale=4., label_list=CIFAR100_LABELS_LIST)
```
### Backup code
Extracting metadata information from CIFAR-100 dataset.
Please download CIFAR-100 dataset for python from
https://www.cs.toronto.edu/~kriz/cifar-100-python.tar.gz
Extract it, and put "meta" file into proper place to execute below code.
```
def unpickle(file):
import pickle
with open(file, 'rb') as fo:
dict = pickle.load(fo)
return dict
metadata = unpickle(os.path.join('./src/cnn/assets', 'meta'))
print(metadata)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import os
import glob
import matplotlib.pyplot as plt
from sklearn import linear_model
# from scipy import stats
import seaborn as sns
import altair as alt
import plotly.express as px
```
## merge promice data to one dataframe
```
df = pd.read_csv('promice/promice.csv')
df['Longitude'] = df['Longitude'] * -1
folderpath = "promice/modis500m"
searchCriteria = "*.csv"
globInput = os.path.join(folderpath, searchCriteria)
csvPath = glob.glob(globInput)
csvList = os.listdir(folderpath)
# daily
for i in range(len(csvList)):
# promice data
stationName = os.path.splitext(csvList[i])[0].replace("-", "*")
index = df.index[df.Station == stationName][0]
url = df.url[index] # daily
# url = df.urlhourly[index]
dfs = pd.read_table(url, sep=r'\s{1,}', engine='python')
dfs = dfs[['Albedo_theta<70d', 'LatitudeGPS_HDOP<1(degN)', 'LongitudeGPS_HDOP<1(degW)', 'Year', 'MonthOfYear', 'DayOfMonth','CloudCover']]
dfs = dfs.replace(-999, np.nan)
dfs['lon'] = dfs['LongitudeGPS_HDOP<1(degW)'].interpolate(method='linear',limit_direction='both') * -1
dfs['lat'] = dfs['LatitudeGPS_HDOP<1(degN)'].interpolate(method='linear',limit_direction='both')
dfs['datetime'] = pd.to_datetime(dict(year=dfs.Year, month=dfs.MonthOfYear, day = dfs.DayOfMonth))
# cloud cover less than 50% and albedo must be valid value
dfs = dfs[(dfs['Albedo_theta<70d'] > 0) & (dfs['CloudCover'] < 0.5)]
dfs['Station'] = stationName
# satellite data
dfr = pd.read_csv(csvPath[i])
dfr['Snow_Albedo_Daily_Tile'] = dfr['Snow_Albedo_Daily_Tile'] / 100
# dfr.datetime = pd.to_datetime(dfr.datetime).dt.date # keep only ymd
dfr.datetime = pd.to_datetime(dfr.datetime)
# join by datetime
# dfmerge = pd.merge(dfr, dfs, how='outer', on='datetime')
dfmerge = pd.merge_asof(dfr.sort_values('datetime'), dfs, on='datetime',allow_exact_matches=False, tolerance=pd.Timedelta(days=1) )
if i==0:
dfmerge.dropna().to_csv('promice vs satellite modis.csv', mode='w', index=False)
else:
dfmerge.dropna().to_csv('promice vs satellite modis.csv', mode='a', index=False, header=False)
```
## Lienar Regression: PROMICE VS MODIS albedo
```
df = pd.read_csv("promice vs satellite modis.csv")
# ProfileReport(df)
df = df[(df['MonthOfYear']>4) & (df['MonthOfYear']<10)] # (df['MonthOfYear']!=7
# df = df[df['Albedo_theta<70d']<0.9]
# df[['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2']] = df[['Blue', 'Green', 'Red', 'NIR', 'SWIR1', 'SWIR2']] / 10000
boxplot = df.boxplot(column=['Snow_Albedo_Daily_Tile', 'Albedo_theta<70d'])
df.Station.value_counts().plot(kind='bar')
X = df[['Snow_Albedo_Daily_Tile']]
# X = df[['Blue', 'Green', 'NIR', 'SWIR1', 'SWIR2']]
y = df['Albedo_theta<70d']
# mask = df['MonthOfYear']>6
# y[mask] = y[mask]/1.1
ols = linear_model.LinearRegression()
model = ols.fit(X, y)
response = model.predict(X)
r2 = model.score(X, y)
print('R\N{SUPERSCRIPT TWO}: %.4f' % r2)
print(model.coef_)
# print("coefficients: Blue: %.4f, Green: %.4f, Red: %.4f, NIR: %.4f, SWIR1: %.4f, SWIR2: %.4f" %(model.coef_[0], model.coef_[1], model.coef_[2], model.coef_[3], model.coef_[4], model.coef_[5]))
# print("coefficients: Blue: %.4f, Red: %.4f, NIR: %.4f, SWIR1: %.4f, SWIR2: %.4f" %(model.coef_[0], model.coef_[1], model.coef_[2], model.coef_[3], model.coef_[4]))
print("intercept: %.4f" % model.intercept_)
len(df)
fig, ax = plt.subplots(figsize=(8, 8))
# plt.sca(ax1)
sns.set_theme(style="darkgrid", font="Arial", font_scale=2)
sns.scatterplot(x=response, y=y, s=10 )
sns.regplot(x=response, y=y, scatter=False, color='red',)
# plt.plot([0,1], [0,1], color = 'white') # reference line
plt.xlim(0, 1)
plt.ylim(0, 1)
ax.set(xlabel='Predicted Albedo (MODIS)', ylabel='Albedo PROMICE')
ax.set_aspect('equal', 'box')
# sns.histplot(x=response, y=y, bins=50, pthresh=.1, cmap="viridis", cbar=True, cbar_kws={'label': 'frequency'})
# sns.kdeplot(x=response, y=y, levels=5, color="w", linewidths=1)
fig.savefig("print/MODISalbedoPromice.png", dpi=300, bbox_inches="tight")
df['response'] = response
alt.data_transformers.disable_max_rows() # this should be avoided but now let's disable the limit
alt.Chart(df).mark_circle().encode(
x='response',
y='Albedo_theta<70d',
color='Station',
tooltip=['datetime:T','Station','response','Albedo_theta<70d']
).interactive()
# chart + chart.transform_regression('x', 'y').mark_line()
df['response'] = response
alt.data_transformers.disable_max_rows() # this should be avoided but now let's disable the limit
brush = alt.selection(type='interval')
points = alt.Chart(df).mark_circle().encode(
x='response',
y='Albedo_theta<70d',
color=alt.condition(brush, 'Station:O', alt.value('grey')),
tooltip=['datetime:T','Station','response','Albedo_theta<70d']
).add_selection(brush)
# Base chart for data tables
ranked_text = alt.Chart(df).mark_text().encode(
y=alt.Y('row_number:O',axis=None)
).transform_window(
row_number='row_number()'
).transform_filter(
brush
).transform_window(
rank='rank(row_number)'
).transform_filter(
alt.datum.rank<40
)
# Data Tables
stationalt = ranked_text.encode(text='Station').properties(title='station')
albedoalt = ranked_text.encode(text='Albedo_theta<70d:N').properties(title='Albedo')
predictedalt = ranked_text.encode(text='response:N').properties(title='predicted albedo')
timealt = ranked_text.encode(text='datetime:T').properties(title='time')
text = alt.hconcat(stationalt, albedoalt, predictedalt, timealt) # Combine data tables
# Build chart
alt.hconcat(
points,
text
).resolve_legend(
color="independent"
)
# chart + chart.transform_regression('x', 'y').mark_line()
```
| github_jupyter |
```
"""
Pattern Lab Project - Team Kingsmen
Name: Mohammed Jawwadul Islam
ID: 011 181 182
UIU Official Email: mislam181182@bscse.uiu.ac.bd
Name: MD Fahad Al Rafi
ID: 011 181 201
UIU Official Email: mrafi181201@bscse.uiu.ac.bd
Name: Pranto Podder
ID: 011 181 202
UIU Official Email: ppodder181202@bscse.uiu.ac.bd
"""
```
## Importing the Libraies
```
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
from collections import Counter
```
## Reading the dataset
```
data = pd.read_csv('healthcare-dataset-stroke-data.csv', na_values='N/A')
# Removing " " empty space between feature values
from pandas.api.types import is_string_dtype
for column in data.columns:
if (is_string_dtype(data[column].dtype)):
data[column] = data[column].str.strip()
X = data.loc[:, data.columns != 'stroke']
y = data['stroke']
print(X.shape, y.shape)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=0)
# Train - 80% , Test - 20%
print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
```
## Data Preprocessing
### Handling Missing Values
#### Simple Imputation by Sklearn - Mean/Median/Most Frequent
```
from sklearn.impute import SimpleImputer
si_X_train = pd.DataFrame() # create a new dataframe to save the train dataset
si_X_test = pd.DataFrame() # create a new dataframe to save the test dataset
for column in X_train.columns:
if (is_string_dtype(X_train[column].dtype)):
si = SimpleImputer(strategy='most_frequent')
else:
si = SimpleImputer(strategy='median')
si.fit(X_train[[column]])
si_X_train[column] = si.transform(X_train[[column]]).flatten() # Flatten 2D matrix to 1D
si_X_test[column] = si.transform(X_test[[column]]).flatten()
si_X_train
```
### Handling Text Features
#### Label Encoder
```
categorical_features = []
for col in data.columns:
if col=='Class':
continue
if is_string_dtype(data[col].dtype):
categorical_features.append(col)
categorical_features
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
# Convert the Label Class
y_train = le.fit_transform(y_train)
y_test = le.transform(y_test)
l_X_train = pd.DataFrame() # Train dataset --> before scaling
l_X_test = pd.DataFrame() # Test dataset --> before scaling
# Convert the text features
for column in X_train.columns:
if column in categorical_features:
l_X_train[column] = le.fit_transform(si_X_train[column])
l_X_test[column] = le.transform(si_X_test[column])
else:
l_X_train[column] = si_X_train[column].copy()
l_X_test[column] = si_X_test[column].copy()
l_X_train.isnull().sum()
```
## Oversampling the dataset
whichever classess has less number of values, we will impute and increase the values in those classes
```
from imblearn.over_sampling import RandomOverSampler
os=RandomOverSampler(0.75) # 75%
l_X_train_ns,y_train_ns = os.fit_resample(l_X_train,y_train)
print("The number of classes before fit {}".format(Counter(y_train)))
print("The number of classes after fit {}".format(Counter(y_train_ns)))
```
## Feature Scaling
#### Standardization
```
from sklearn.preprocessing import StandardScaler
ss = StandardScaler()
l_X_train_ns = ss.fit_transform(l_X_train_ns)
l_X_test = ss.transform(l_X_test)
print(l_X_train)
```
## Model Building
## Classification Evaluatin Metrics
```
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score, matthews_corrcoef
def evaluate_preds(y_test,y_pred):
accuracy = accuracy_score(y_test,y_pred)
precision = precision_score(y_test,y_pred)
recall = recall_score(y_test,y_pred)
f1 = f1_score(y_test,y_pred)
mcc = matthews_corrcoef(y_test,y_pred)
metric_dict = {
"accuracy":round(accuracy,2),
"precision":round(precision,2),
"recall":round(recall,2),
"f1":round(f1,2),
"mcc": mcc
} # A dictionary that stores the results of the evaluation metrics
print(f"Acc: {accuracy * 100:.2f}%")
print(f"Precision: {precision:.2f}")
print(f"Recall: {recall:.2f}")
print(f"F1 score: {f1:.2f}")
print(f'MCC Score: {mcc:.2f}')
return metric_dict
```
### SVM
```
from sklearn.svm import SVC
svc = SVC(kernel='rbf',random_state=0)
svc.fit(l_X_train_ns,y_train_ns)
y_pred = svc.predict(l_X_test)
model_metrics = evaluate_preds(y_test, y_pred)
```
### Naive Bayes
```
from sklearn.naive_bayes import GaussianNB
naive = GaussianNB()
naive.fit(l_X_train_ns,y_train_ns)
y_pred = naive.predict(l_X_test)
model_metrics = evaluate_preds(y_test, y_pred)
```
### Logistic Regression
```
from sklearn.linear_model import LogisticRegression
logistic = LogisticRegression()
logistic.fit(l_X_train_ns,y_train_ns)
y_pred = logistic.predict(l_X_test)
model_metrics = evaluate_preds(y_test, y_pred)
```
### k Nearest Neighbours
```
from sklearn.neighbors import KNeighborsClassifier
neigh = KNeighborsClassifier(n_neighbors=40)
neigh.fit(l_X_train_ns,y_train_ns)
y_pred = neigh.predict(l_X_test)
model_metrics = evaluate_preds(y_test, y_pred)
```
### RandomForestClassifier
```
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=500, n_jobs=-1, criterion='entropy')
rf.fit(l_X_train_ns,y_train_ns)
y_pred = rf.predict(l_X_test)
model_metrics = evaluate_preds(y_test, y_pred)
```
## Hyperparameter Tuning
```
from sklearn.model_selection import RandomizedSearchCV
```
### SVM
```
# RandomizedSearchCV will refit the estimator, i.e. model object using the best found parameters on the whole dataset.
parameters = [ # list of different combination of hyperparameters that we want to test
{'C': [0.25, 0.5, 0.75, 1], 'kernel': ['linear']},
{'C': [0.25, 0.5, 0.75, 1], 'kernel': ['rbf'], 'gamma': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]}
]
# also, two dictionaries --> for testing 2 different kernels. {key - parameters(name must match documentation), value - range of values to try}
random_cv = RandomizedSearchCV(estimator = svc,
param_distributions = parameters,
scoring = 'accuracy', # evaluation metric. 'accuracy' for classification here
cv = 5, # k = 5, no. of k-train test folds
verbose=2, # print out the results
n_jobs = -1, # set processor of machine. -1 --> means all processor will be in use
refit=True)
# RandomizedSearchCV is only applied on train set
random_cv.fit(l_X_train_ns, y_train_ns)
# get best accuracy, and the set of parameters that led to that best accuracy
best_accuracy = random_cv.best_score_
best_parameters = random_cv.best_params_
print("Best Accuracy: {:.2f} %".format(best_accuracy*100))
print("Best Parameters:", best_parameters)
```
### Random Forest Classifier
```
parameters = [ # list of different combination of hyperparameters that we want to test
{'n_estimators': range(10,300,10), 'criterion': ['gini', 'entropy'], 'max_features':['auto','sqrt','log2']}
]
random_cv = RandomizedSearchCV(estimator = rf,
param_distributions = parameters,
scoring = 'accuracy',
cv = 5,
verbose=2,
n_jobs = -1,
refit=True)
random_cv.fit(l_X_train_ns, y_train_ns)
best_accuracy = random_cv.best_score_
best_parameters = random_cv.best_params_
print("Best Accuracy: {:.2f} %".format(best_accuracy*100))
print("Best Parameters:", best_parameters)
```
| github_jupyter |
- title: Reward tampering
- summary: Improving safety and control by preventing all manner of reward tampering by the agent itself.
- author: Daniel Cox
- date: 2019-08-25
- category: arXiv highlights
- image: /static/images/arXiv.gif
# This week
This week I just want to pull the list of reward tampering methods from [Reward Tampering Problems and Solutions in Reinforcement Learning: A Causal Influence Diagram Perspective](https://arxiv.org/abs/1908.04734) to promote awareness of this problem. The paper is interesting for several other reasons as well, and I commend it to you:
> Can an arbitrarily intelligent reinforcement learning agent be kept under control by a human user? Or do agents with sufficient intelligence inevitably find ways to shortcut their reward signal? This question impacts how far reinforcement learning can be scaled, and whether alternative paradigms must be developed in order to build safe artificial general intelligence.
# Reward tampering
I've heard it said that no agent will ever become more intelligent than it takes to edit its own reward function, giving itself a simpler task. This paper treats such problems seriously, with some encouraging results.
> From an AI safety perspective, we must bear in mind that in any practically implemented system, agent reward may not coincide with user utility. In other words, the agent may have found a way to obtain reward without doing the task. This is sometimes called reward hacking or reward corruption. We distinguish between a few different types of reward hacking.
## Reward gaming vs. reward tampering
The authors make a distinction between _reward gaming_, where the agent exploits a misspecification of the process that determines the rewards, and _reward tampering_, where the agent actually modifies that process. This paper is focused on the latter.
They then subdivide reward tampering into three subcategories, according to whether the agent has tampered with the function itself, the feedback that trains the reward function, or the input to the reward function.
## Hacking the reward function: Section 3
> First, regardless of whether the reward is chosen by a computer program, a human, or both, a sufficiently capable, real-world agent may find a way to tamper with the decision. The agent may for example hack the computer program that determines the reward. Such a strategy may bring high agent reward and low user utility. This reward function tampering problem will be explored in Section 3.
>
> Fortunately, there are modifications of the RL objective that remove the agent’s incentiveto tamper with the reward function.
In Section 3 the authors formalize the problem, and propose two reward variants that disincentivize tampering.
## Manipulating the feedback mechanism: Section 4
> The related problem of reward gaming can occur even if the agent never tamperswith the reward function. A promising way to mitigate the reward gaming problem isto let the user continuously give feedback to update the reward function, using online reward-modeling. Whenever the agent finds a strategy with high agent reward but low user utility, the user can give feedback that dissuades the agent from continuing the behavior. However, a worry with online reward modeling is that the agent may influence the feedback. For example, the agent may prevent the user from giving feedback while continuing to exploit a misspecified reward function, or manipulate the user to give feedback that boosts agent reward but not user utility. This feedback tampering problem and its solutions will be the focus of Section 4.
Section 4 proposes several potential modifications to disincentivize or directly prevent feedback manipulation, ultimately with the recommendation that they be combined in an ensemble.
## Input tampering: Section 5
> Finally, the agent may tamper with the input to the reward function, so-called RF-input tampering, for example by gluing a picture in front of its camera to fool the reward function that the task has been completed. This problem and its potential solution will be the focus of Section 5.
Very interestingly, Section 5 argues that model-based methods avoid the input tampering problem.
# Results summary
> One way to prevent the agent from tampering with the reward function is to isolate or encrypt the reward function, and in other ways trying to physically prevent the agent from reward tampering. However, we do not expect such solutions to scale indefinitely with our agent’s capabilities, as a sufficiently capable agent may find ways around most defenses. Instead, we have argued for design principles that prevent reward tampering incentives, while still keeping agents motivated to complete the original task. Indeed, for each type of reward tampering possibility, we described one or more design principles for removing the agent’s incentive to use it. The design principles can be combined into agent designs with no reward tampering incentive at all.
>
> An important next step is to turn the design principles into practical and scalable RL algorithms, and to verify that they do the right thing in setups where various types of reward tampering are possible. With time, we hope that these design principles will evolve into a set of best practices for how to build capable RL agents without reward tampering incentives. We also hope that the use of causal influence diagrams that we have pioneered in this paper will contribute to a deeper understanding of many other AI safety problems and help generate new solutions.
# Parting thoughts
1. I look forward to reading this paper more thoroughly, both because I understand this problem of disincentivising reward hacking is _hard_, and because Causal Influence Diagrams sound interesting and generally useful.
2. AI safety is important, and I rather hope that awareness of some ways your agents could cheat will help to prevent such errors from leaking out into the world before they are caught.
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 텐서플로로 분산 훈련하기
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/distributed_training"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/guide/distributed_training.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
</td>
</table>
Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도
불구하고 [공식 영문 문서](https://www.tensorflow.org/?hl=en)의 내용과 일치하지 않을 수 있습니다.
이 번역에 개선할 부분이 있다면
[tensorflow/docs-l10n](https://github.com/tensorflow/docs-l10n/) 깃헙 저장소로 풀 리퀘스트를 보내주시기 바랍니다.
문서 번역이나 리뷰에 참여하려면
[docs-ko@tensorflow.org](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ko)로
메일을 보내주시기 바랍니다.
## 개요
`tf.distribute.Strategy`는 훈련을 여러 GPU 또는 여러 장비, 여러 TPU로 나누어 처리하기 위한 텐서플로 API입니다. 이 API를 사용하면 기존의 모델이나 훈련 코드를 조금만 고쳐서 분산처리를 할 수 있습니다.
`tf.distribute.Strategy`는 다음을 주요 목표로 설계하였습니다.
* 사용하기 쉽고, 연구원, 기계 학습 엔지니어 등 여러 사용자 층을 지원할 것.
* 그대로 적용하기만 하면 좋은 성능을 보일 것.
* 전략들을 쉽게 갈아 끼울 수 있을 것.
`tf.distribute.Strategy`는 텐서플로의 고수준 API인 [tf.keras](https://www.tensorflow.org/guide/keras) 및 [tf.estimator](https://www.tensorflow.org/guide/estimator)와 함께 사용할 수 있습니다. 코드 한두 줄만 추가하면 됩니다. 사용자 정의 훈련 루프(그리고 텐서플로를 사용한 모든 계산 작업)에 함께 사용할 수 있는 API도 제공합니다.
텐서플로 2.0에서는 사용자가 프로그램을 즉시 실행(eager execution)할 수도 있고, [`tf.function`](../tutorials/eager/tf_function.ipynb)을 사용하여 그래프에서 실행할 수도 있습니다. `tf.distribute.Strategy`는 두 가지 실행 방식을 모두 지원하려고 합니다. 이 가이드에서는 대부분의 경우 훈련에 대하여 이야기하겠지만, 이 API 자체는 여러 환경에서 평가나 예측을 분산 처리하기 위하여 사용할 수도 있다는 점을 참고하십시오.
잠시 후 보시겠지만 코드를 약간만 바꾸면 `tf.distribute.Strategy`를 사용할 수 있습니다. 변수, 층, 모델, 옵티마이저, 지표, 서머리(summary), 체크포인트 등 텐서플로를 구성하고 있는 기반 요소들을 전략(Strategy)을 이해하고 처리할 수 있도록 수정했기 때문입니다.
이 가이드에서는 다양한 형식의 전략에 대해서, 그리고 여러 가지 상황에서 이들을 어떻게 사용해야 하는지 알아보겠습니다.
```
# 텐서플로 패키지 가져오기
!pip install tensorflow-gpu==2.0.0-rc1
import tensorflow as tf
```
## 전략의 종류
`tf.distribute.Strategy`는 서로 다른 다양한 사용 형태를 아우르려고 합니다. 몇 가지 조합은 현재 지원하지만, 추후에 추가될 전략들도 있습니다. 이들 중 몇 가지를 살펴보겠습니다.
* 동기 훈련 대 비동기 훈련: 분산 훈련을 할 때 데이터를 병렬로 처리하는 방법은 크게 두 가지가 있습니다. 동기 훈련을 할 때는 모든 워커(worker)가 입력 데이터를 나누어 갖고 동시에 훈련합니다. 그리고 각 단계마다 그래디언트(gradient)를 모읍니다. 비동기 훈련에서는 모든 워커가 독립적으로 입력 데이터를 사용해 훈련하고 각각 비동기적으로 변수들을 갱신합니다. 일반적으로 동기 훈련은 올 리듀스(all-reduce)방식으로 구현하고, 비동기 훈련은 파라미터 서버 구조를 사용합니다.
* 하드웨어 플랫폼: 한 장비에 있는 다중 GPU로 나누어 훈련할 수도 있고, 네트워크로 연결된 (GPU가 없거나 여러 개의 GPU를 가진) 여러 장비로 나누어서, 또 혹은 클라우드 TPU에서 훈련할 수도 있습니다.
이런 사용 형태들을 위하여, 현재 5가지 전략을 사용할 수 있습니다. 이후 내용에서 현재 TF 2.0 베타에서 상황마다 어떤 전략을 지원하는지 이야기하겠습니다. 일단 간단한 개요는 다음과 같습니다.
| 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
|:----------------------- |:------------------- |:--------------------- |:--------------------------------- |:--------------------------------- |:-------------------------- |
| **Keras API** | 지원 | 2.0 RC 지원 예정 | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 이후 지원 예정 |
| **사용자 정의 훈련 루프** | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 이후 지원 예정 | 2.0 RC 지원 예정 | 아직 미지원 |
| **Estimator API** | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 |
### MirroredStrategy
`tf.distribute.MirroredStrategy`는 장비 하나에서 다중 GPU를 이용한 동기 분산 훈련을 지원합니다. 각각의 GPU 장치마다 복제본이 만들어집니다. 모델의 모든 변수가 복제본마다 미러링 됩니다. 이 미러링된 변수들은 하나의 가상의 변수에 대응되는데, 이를 `MirroredVariable`라고 합니다. 이 변수들은 동일한 변경사항이 함께 적용되므로 모두 같은 값을 유지합니다.
여러 장치에 변수의 변경사항을 전달하기 위하여 효율적인 올 리듀스 알고리즘을 사용합니다. 올 리듀스 알고리즘은 모든 장치에 걸쳐 텐서를 모은 다음, 그 합을 구하여 다시 각 장비에 제공합니다. 이 통합된 알고리즘은 매우 효율적이어서 동기화의 부담을 많이 덜어낼 수 있습니다. 장치 간에 사용 가능한 통신 방법에 따라 다양한 올 리듀스 알고리즘과 구현이 있습니다. 기본값으로는 NVIDIA NCCL을 올 리듀스 구현으로 사용합니다. 또한 제공되는 다른 몇 가지 방법 중에 선택하거나, 직접 만들 수도 있습니다.
`MirroredStrategy`를 만드는 가장 쉬운 방법은 다음과 같습니다.
```
mirrored_strategy = tf.distribute.MirroredStrategy()
```
`MirroredStrategy` 인스턴스가 생겼습니다. 텐서플로가 인식한 모든 GPU를 사용하고, 장치 간 통신에는 NCCL을 사용할 것입니다.
장비의 GPU 중 일부만 사용하고 싶다면, 다음과 같이 하면 됩니다.
```
mirrored_strategy = tf.distribute.MirroredStrategy(devices=["/gpu:0", "/gpu:1"])
```
장치 간 통신 방법을 바꾸고 싶다면, `cross_device_ops` 인자에 `tf.distribute.CrossDeviceOps` 타입의 인스턴스를 넘기면 됩니다. 현재 기본값인 `tf.distribute.NcclAllReduce` 이외에 `tf.distribute.HierarchicalCopyAllReduce`와 `tf.distribute.ReductionToOneDevice` 두 가지 추가 옵션을 제공합니다.
```
mirrored_strategy = tf.distribute.MirroredStrategy(
cross_device_ops=tf.distribute.HierarchicalCopyAllReduce())
```
### CentralStorageStrategy
`tf.distribute.experimental.CentralStorageStrategy`도 동기 훈련을 합니다. 하지만 변수를 미러링하지 않고, CPU에서 관리합니다. 작업은 모든 로컬 GPU들로 복제됩니다. 단, 만약 GPU가 하나밖에 없다면 모든 변수와 작업이 그 GPU에 배치됩니다.
다음과 같이 `CentralStorageStrategy` 인스턴스를 만드십시오.
```
central_storage_strategy = tf.distribute.experimental.CentralStorageStrategy()
```
`CentralStorageStrategy` 인스턴스가 만들어졌습니다. 인식한 모든 GPU와 CPU를 사용합니다. 각 복제본의 변수 변경사항은 모두 수집된 후 변수에 적용됩니다.
Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 다음에 API가 바뀔 수 있음에 유념하십시오.
### MultiWorkerMirroredStrategy
`tf.distribute.experimental.MultiWorkerMirroredStrategy`은 `MirroredStrategy`와 매우 비슷합니다. 다중 워커를 이용하여 동기 분산 훈련을 합니다. 각 워커는 여러 개의 GPU를 사용할 수 있습니다. `MirroredStrategy`처럼 모델에 있는 모든 변수의 복사본을 모든 워커의 각 장치에 만듭니다.
다중 워커(multi-worker)들 사이에서는 올 리듀스(all-reduce) 통신 방법으로 [CollectiveOps](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/ops/collective_ops.py)를 사용하여 변수들을 같은 값으로 유지합니다. 수집 연산(collective op)은 텐서플로 그래프에 속하는 연산 중 하나입니다. 이 연산은 하드웨어나 네트워크 구성, 텐서 크기에 따라 텐서플로 런타임이 지원하는 올 리듀스 알고리즘을 자동으로 선택합니다.
여기에 추가 성능 최적화도 구현하고 있습니다. 예를 들어 작은 텐서들의 여러 올 리듀스 작업을 큰 텐서들의 더 적은 올 리듀스 작업으로 바꾸는 정적 최적화 기능이 있습니다. 뿐만아니라 플러그인 구조를 갖도록 설계하였습니다. 따라서 추후에는 사용자가 자신의 하드웨어에 더 최적화된 알고리즘을 사용할 수도 있을 것입니다. 참고로 이 수집 연산은 올 리듀스 외에 브로드캐스트(broadcast)나 전체 수집(all-gather)도 구현하고 있습니다.
`MultiWorkerMirroredStrategy`를 만드는 가장 쉬운 방법은 다음과 같습니다.
```
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
```
`MultiWorkerMirroredStrategy`에 사용할 수 있는 수집 연산 구현은 현재 두 가지입니다. `CollectiveCommunication.RING`는 gRPC를 사용한 링 네트워크 기반의 수집 연산입니다. `CollectiveCommunication.NCCL`는 [Nvidia의 NCCL](https://developer.nvidia.com/nccl)을 사용하여 수집 연산을 구현한 것입니다. `CollectiveCommunication.AUTO`로 설정하면 런타임이 알아서 구현을 고릅니다. 최적의 수집 연산 구현은 GPU의 수와 종류, 클러스터의 네트워크 연결 등에 따라 다를 수 있습니다. 예를 들어 다음과 같이 지정할 수 있습니다.
```
multiworker_strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy(
tf.distribute.experimental.CollectiveCommunication.NCCL)
```
다중 GPU를 사용하는 것과 비교해서 다중 워커를 사용하는 것의 가장 큰 차이점은 다중 워커에 대한 설정 부분입니다. 클러스터를 구성하는 각 워커에 "TF_CONFIG" 환경변수를 사용하여 클러스터 설정을 하는 것이 텐서플로의 표준적인 방법입니다. [아래쪽 "TF_CONFIG"](#TF_CONFIG) 항목에서 어떻게 하는지 자세히 살펴보겠습니다.
Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 나중에 API가 바뀔 수 있음에 유념하십시오.
### TPUStrategy
`tf.distribute.experimental.TPUStrategy`는 텐서플로 훈련을 텐서처리장치(Tensor Processing Unit, TPU)에서 수행하는 전략입니다. TPU는 구글의 특별한 주문형 반도체(ASIC)로서, 기계 학습 작업을 극적으로 가속하기 위하여 설계되었습니다. TPU는 구글 코랩, [Tensorflow Research Cloud](https://www.tensorflow.org/tfrc), [Google Compute Engine](https://cloud.google.com/tpu)에서 사용할 수 있습니다.
분산 훈련 구조의 측면에서, TPUStrategy는 `MirroredStrategy`와 동일합니다. 동기 분산 훈련 방식을 사용합니다. TPU는 자체적으로 여러 TPU 코어들에 걸친 올 리듀스 및 기타 수집 연산을 효율적으로 구현하고 있습니다. 이 구현이 `TPUStrategy`에 사용됩니다.
`TPUStrategy`를 사용하는 방법은 다음과 같습니다.
Note: 코랩에서 이 코드를 사용하려면, 코랩 런타임으로 TPU를 선택해야 합니다. TPUStrategy를 사용하는 방법에 대한 튜토리얼을 곧 추가하겠습니다.
```
cluster_resolver = tf.distribute.cluster_resolver.TPUClusterResolver(
tpu=tpu_address)
tf.config.experimental_connect_to_host(cluster_resolver.master())
tf.tpu.experimental.initialize_tpu_system(cluster_resolver)
tpu_strategy = tf.distribute.experimental.TPUStrategy(cluster_resolver)
```
`TPUClusterResolver` 인스턴스는 TPU를 찾도록 도와줍니다. 코랩에서는 아무런 인자를 주지 않아도 됩니다. 클라우드 TPU에서 사용하려면, TPU 자원의 이름을 `tpu` 매개변수에 지정해야 합니다. 또한 TPU는 계산하기 전 초기화(initialize)가 필요합니다. 초기화 중 TPU 메모리가 지워져서 모든 상태 정보가 사라지므로, 프로그램 시작시에 명시적으로 TPU 시스템을 초기화(initialize)해 주어야 합니다.
Note: 이 전략은 아직 개선 중이고 더 많은 경우에 쓸 수 있도록 만들고 있기 때문에, [`실험 기능`](https://www.tensorflow.org/guide/versions#what_is_not_covered)으로 지원됩니다. 따라서 나중에 API가 바뀔 수 있음에 유념하십시오.
### ParameterServerStrategy
`tf.distribute.experimental.ParameterServerStrategy`은 여러 장비에서 훈련할 때 파라미터 서버를 사용합니다. 이 전략을 사용하면 몇 대의 장비는 워커 역할을 하고, 몇 대는 파라미터 서버 역할을 하게 됩니다. 모델의 각 변수는 한 파라미터 서버에 할당됩니다. 계산 작업은 모든 워커의 GPU들에 복사됩니다.
코드만 놓고 보았을 때는 다른 전략들과 비슷합니다.
```
ps_strategy = tf.distribute.experimental.ParameterServerStrategy()
```
다중 워커 환경에서 훈련하려면, 클러스터에 속한 파라미터 서버와 워커를 "TF_CONFIG" 환경변수를 이용하여 설정해야 합니다. 자세한 내용은 [아래쪽 "TF_CONFIG"](#TF_CONFIG)에서 설명하겠습니다.
여기까지 여러 가지 전략들이 어떻게 다르고, 어떻게 사용하는지 살펴보았습니다. 이어지는 절들에서는 훈련을 분산시키기 위하여 이들을 어떻게 사용해야 하는지 살펴보겠습니다. 이 문서에서는 간단한 코드 조각만 보여드리겠지만, 처음부터 끝까지 전체 코드를 실행할 수 있는 더 긴 튜토리얼의 링크도 함께 안내해드리겠습니다.
## 케라스와 함께 `tf.distribute.Strategy` 사용하기
`tf.distribute.Strategy`는 텐서플로의 [케라스 API 명세](https://keras.io) 구현인 `tf.keras`와 함께 사용할 수 있습니다. `tf.keras`는 모델을 만들고 훈련하는 고수준 API입니다. 분산 전략을 `tf.keras` 백엔드와 함께 쓸 수 있으므로, 케라스 사용자들도 케라스 훈련 프레임워크로 작성한 훈련 작업을 쉽게 분산 처리할 수 있게 되었습니다. 훈련 프로그램에서 고쳐야하는 부분은 거의 없습니다. (1) 적절한 `tf.distribute.Strategy` 인스턴스를 만든 다음 (2)
케라스 모델의 생성과 컴파일을 `strategy.scope` 안으로 옮겨주기만 하면 됩니다. `Sequential` , 함수형 API, 클래스 상속 등 모든 방식으로 만든 케라스 모델을 다 지원합니다.
다음은 한 개의 밀집 층(dense layer)을 가진 매우 간단한 케라스 모델에 분산 전략을 사용하는 코드의 일부입니다.
```
mirrored_strategy = tf.distribute.MirroredStrategy()
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
model.compile(loss='mse', optimizer='sgd')
```
위 예에서는 `MirroredStrategy`를 사용했기 때문에, 하나의 장비가 다중 GPU를 가진 경우에 사용할 수 있습니다. `strategy.scope()`로 분산 처리할 부분을 코드에 지정할 수 있습니다. 이 범위(scope) 안에서 모델을 만들면, 일반적인 변수가 아니라 미러링된 변수가 만들어집니다. 이 범위 안에서 컴파일을 한다는 것은 작성자가 이 전략을 사용하여 모델을 훈련하려고 한다는 의미입니다. 이렇게 구성하고 나서, 일반적으로 실행하는 것처럼 모델의 `fit` 함수를 호출합니다.
`MirroredStrategy`가 모델의 훈련을 사용 가능한 GPU들로 복제하고, 그래디언트들을 수집하는 것 등을 알아서 처리합니다.
```
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100).batch(10)
model.fit(dataset, epochs=2)
model.evaluate(dataset)
```
위에서는 훈련과 평가 입력을 위해 `tf.data.Dataset`을 사용했습니다. 넘파이(numpy) 배열도 사용할 수 있습니다.
```
import numpy as np
inputs, targets = np.ones((100, 1)), np.ones((100, 1))
model.fit(inputs, targets, epochs=2, batch_size=10)
```
데이터셋이나 넘파이를 사용하는 두 경우 모두 입력 배치가 동일한 크기로 나누어져서 여러 개로 복제된 작업에 전달됩니다. 예를 들어, `MirroredStrategy`를 2개의 GPU에서 사용한다면, 크기가 10개인 배치(batch)가 두 개의 GPU로 배분됩니다. 즉, 각 GPU는 한 단계마다 5개의 입력을 받게 됩니다. 따라서 GPU가 추가될수록 각 에포크(epoch) 당 훈련 시간은 줄어들게 됩니다. 일반적으로는 가속기를 더 추가할 때마다 배치 사이즈도 더 키웁니다. 추가한 컴퓨팅 자원을 더 효과적으로 사용하기 위해서입니다. 모델에 따라서는 학습률(learning rate)을 재조정해야 할 수도 있을 것입니다. 복제본의 수는 `strategy.num_replicas_in_sync`로 얻을 수 있습니다.
```
# 복제본의 수로 전체 배치 크기를 계산.
BATCH_SIZE_PER_REPLICA = 5
global_batch_size = (BATCH_SIZE_PER_REPLICA *
mirrored_strategy.num_replicas_in_sync)
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(100)
dataset = dataset.batch(global_batch_size)
LEARNING_RATES_BY_BATCH_SIZE = {5: 0.1, 10: 0.15}
learning_rate = LEARNING_RATES_BY_BATCH_SIZE[global_batch_size]
```
### 현재 어떤 것이 지원됩니까?
TF 2.0 베타 버전에서는 케라스와 함께 `MirroredStrategy`와 `CentralStorageStrategy`, `MultiWorkerMirroredStrategy`를 사용하여 훈련할 수 있습니다. `CentralStorageStrategy`와 `MultiWorkerMirroredStrategy`는 아직 실험 기능이므로 추후 바뀔 수 있습니다.
다른 전략도 조만간 지원될 것입니다. API와 사용 방법은 위에 설명한 것과 동일할 것입니다.
| 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
|---------------- |--------------------- |----------------------- |----------------------------------- |----------------------------------- |--------------------------- |
| Keras API | 지원 | 2.0 RC 지원 예정 | 실험 기능으로 지원 | 실험 기능으로 지원 | 2.0 RC 지원 예정 |
### 예제와 튜토리얼
위에서 설명한 케라스 분산 훈련 방법에 대한 튜토리얼과 예제들의 목록입니다.
1. `MirroredStrategy`를 사용한 [MNIST](../tutorials/distribute/keras.ipynb) 훈련 튜토리얼.
2. ImageNet 데이터와 `MirroredStrategy`를 사용한 공식 [ResNet50](https://github.com/tensorflow/models/blob/master/official/vision/image_classification/resnet_imagenet_main.py) 훈련.
3. 클라우드 TPU에서 ImageNet 데이터와 `TPUStrategy`를 사용한 [ResNet50](https://github.com/tensorflow/tpu/blob/master/models/experimental/resnet50_keras/resnet50.py) 훈련. 이 예제는 현재 텐서플로 1.x 버전에서만 동작합니다.
4. `MultiWorkerMirroredStrategy`를 사용한 [MNIST](../tutorials/distribute/multi_worker_with_keras.ipynb) 훈련 튜토리얼.
5. `MirroredStrategy`를 사용한 [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) 훈련.
6. `MirroredStrategy`를 사용한 [Transformer](https://github.com/tensorflow/models/blob/master/official/nlp/transformer/transformer_main.py) 훈련.
## 사용자 정의 훈련 루프와 함께 `tf.distribute.Strategy` 사용하기
지금까지 살펴본 것처럼 고수준 API와 함께 `tf.distribute.Strategy`를 사용하려면 코드 몇 줄만 바꾸면 되었습니다. 조금만 더 노력을 들이면 이런 프레임워크를 사용하지 않는 사용자도 `tf.distribute.Strategy`를 사용할 수 있습니다.
텐서플로는 다양한 용도로 사용됩니다. 연구자들 같은 일부 사용자들은 더 높은 자유도와 훈련 루프에 대한 제어를 원합니다. 이 때문에 추정기나 케라스 같은 고수준 API를 사용하기 힘든 경우가 있습니다. 예를 들어, GAN을 사용하는데 매번 생성자(generator)와 판별자(discriminator) 단계의 수를 바꾸고 싶을 수 있습니다. 비슷하게, 고수준 API는 강화 학습(Reinforcement learning)에는 그다지 적절하지 않습니다. 그래서 이런 사용자들은 보통 자신만의 훈련 루프를 작성하게 됩니다.
이 사용자들을 위하여, `tf.distribute.Strategy` 클래스들은 일련의 주요 메서드들을 제공합니다. 이 메서드들을 사용하려면 처음에는 코드를 이리저리 조금 옮겨야 할 수 있겠지만, 한번 작업해 놓으면 전략 인스턴스만 바꿔서 GPU, TPU, 여러 장비로 쉽게 바꿔가며 훈련을 할 수 있습니다.
앞에서 살펴본 케라스 모델을 사용한 훈련 예제를 통하여 사용하는 모습을 간단하게 살펴보겠습니다.
먼저, 전략의 범위(scope) 안에서 모델과 옵티마이저를 만듭니다. 이는 모델이나 옵티마이저로 만들어진 변수가 미러링 되도록 만듭니다.
```
with mirrored_strategy.scope():
model = tf.keras.Sequential([tf.keras.layers.Dense(1, input_shape=(1,))])
optimizer = tf.keras.optimizers.SGD()
```
다음으로는 입력 데이터셋을 만든 다음, `tf.distribute.Strategy.experimental_distribute_dataset` 메서드를 호출하여 전략에 맞게 데이터셋을 분배합니다.
```
with mirrored_strategy.scope():
dataset = tf.data.Dataset.from_tensors(([1.], [1.])).repeat(1000).batch(
global_batch_size)
dist_dataset = mirrored_strategy.experimental_distribute_dataset(dataset)
```
그리고 나서는 한 단계의 훈련을 정의합니다. 그래디언트를 계산하기 위해 `tf.GradientTape`를 사용합니다. 이 그래디언트를 적용하여 우리 모델의 변수를 갱신하기 위해서는 옵티마이저를 사용합니다. 분산 훈련을 위하여 이 훈련 작업을 `step_fn` 함수 안에 구현합니다. 그리고 `step_fn`을 앞에서 만든 `dist_dataset`에서 얻은 입력 데이터와 함께 `tf.distrbute.Strategy.experimental_run_v2`메서드로 전달합니다.
```
@tf.function
def train_step(dist_inputs):
def step_fn(inputs):
features, labels = inputs
with tf.GradientTape() as tape:
logits = model(features)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=labels)
loss = tf.reduce_sum(cross_entropy) * (1.0 / global_batch_size)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(list(zip(grads, model.trainable_variables)))
return cross_entropy
per_example_losses = mirrored_strategy.experimental_run_v2(
step_fn, args=(dist_inputs,))
mean_loss = mirrored_strategy.reduce(
tf.distribute.ReduceOp.MEAN, per_example_losses, axis=0)
return mean_loss
```
위 코드에서 몇 가지 더 짚어볼 점이 있습니다.
1. 손실(loss)을 계산하기 위하여 `tf.nn.softmax_cross_entropy_with_logits`를 사용하였습니다. 그리고 손실의 합을 전체 배치 크기로 나누는 부분이 중요합니다. 이는 모든 복제된 훈련이 동시에 이루어지고 있고, 각 단계에 훈련이 이루어지는 입력의 수는 전체 배치 크기와 같기 때문입니다. 따라서 손실 값은 각 복제된 작업 내의 배치 크기가 아니라 전체 배치 크기로 나누어야 맞습니다.
2. `tf.distribute.Strategy.experimental_run_v2`에서 반환된 결과를 모으기 위하여 `tf.distribute.Strategy.reduce` API를 사용하였습니다. `tf.distribute.Strategy.experimental_run_v2`는 전략의 각 복제본에서 얻은 결과를 반환합니다. 그리고 이 결과를 사용하는 방법은 여러 가지가 있습니다. 종합한 결과를 얻기 위하여 `reduce` 함수를 사용할 수 있습니다. `tf.distribute.Strategy.experimental_local_results` 메서드로 각 복제본에서 얻은 결과의 값들 목록을 얻을 수도 있습니다.
3. 분산 전략 범위 안에서 `apply_gradients` 메서드가 호출되면, 평소와는 동작이 다릅니다. 구체적으로는 동기화된 훈련 중 병렬화된 각 작업에서 그래디언트를 적용하기 전에, 모든 복제본의 그래디언트를 더해집니다.
훈련 단계를 정의했으므로, 마지막으로는 `dist_dataset`에 대하여 훈련을 반복합니다.
```
with mirrored_strategy.scope():
for inputs in dist_dataset:
print(train_step(inputs))
```
위 예에서는 `dist_dataset`을 차례대로 처리하며 훈련 입력 데이터를 얻었습니다. `tf.distribute.Strategy.make_experimental_numpy_dataset`를 사용하면 넘파이 입력도 쓸 수 있습니다. `tf.distribute.Strategy.experimental_distribute_dataset` 함수를 호출하기 전에 이 API로 데이터셋을 만들면 됩니다.
데이터를 차례대로 처리하는 또 다른 방법은 명시적으로 반복자(iterator)를 사용하는 것입니다. 전체 데이터를 모두 사용하지 않고, 정해진 횟수만큼만 훈련을 하고 싶을 때 유용합니다. 반복자를 만들고 명시적으로 `next`를 호출하여 다음 입력 데이터를 얻도록 하면 됩니다. 위 루프 코드를 바꿔보면 다음과 같습니다.
```
with mirrored_strategy.scope():
iterator = iter(dist_dataset)
for _ in range(10):
print(train_step(next(iterator)))
```
`tf.distribute.Strategy` API를 사용하여 사용자 정의 훈련 루프를 분산 처리 하는 가장 단순한 경우를 살펴보았습니다. 현재 API를 개선하는 과정 중에 있습니다. 이 API를 사용하려면 사용자 쪽에서 꽤 많은 작업을 해야 하므로, 나중에 별도의 더 자세한 가이드로 설명하도록 하겠습니다.
### 현재 어떤 것이 지원됩니까?
TF 2.0 베타 버전에서는 사용자 정의 훈련 루프와 함께 위에서 설명한 `MirroredStrategy`, 그리고 `TPUStrategy`를 사용할 수 있습니다. 또한 `MultiWorkerMirorredStrategy`도 추후 지원될 것입니다.
| 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
|:----------------------- |:------------------- |:------------------- |:----------------------------- |:------------------------ |:------------------------- |
| 사용자 정의 훈련 루프 | 지원 | 지원 | 2.0 RC 지원 예정 | 2.0 RC 지원 예정 | 아직 미지원 |
### 예제와 튜토리얼
사용자 정의 훈련 루프와 함께 분산 전략을 사용하는 예제들입니다.
1. `MirroredStrategy`로 MNIST를 훈련하는 [튜토리얼](../tutorials/distribute/training_loops.ipynb).
2. `MirroredStrategy`를 사용하는 [DenseNet](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/densenet/distributed_train.py) 예제.
3. `MirroredStrategy`와 `TPUStrategy`를 사용하여 훈련하는 [BERT](https://github.com/tensorflow/models/blob/master/official/bert/run_classifier.py) 예제.
이 예제는 분산 훈련 도중 체크포인트로부터 불러오거나 주기적인 체크포인트를 만드는 방법을 이해하는 데 매우 유용합니다.
4. `keras_use_ctl` 플래그를 켜서 활성화할 수 있는 `MirroredStrategy`로 훈련한 [NCF](https://github.com/tensorflow/models/blob/master/official/recommendation/ncf_keras_main.py) 예제.
5. `MirroredStrategy`를 사용하여 훈련하는 [NMT](https://github.com/tensorflow/examples/blob/master/tensorflow_examples/models/nmt_with_attention/distributed_train.py) 예제.
## 추정기(Estimator)와 함께 `tf.distribute.Strategy` 사용하기
`tf.estimator`는 원래부터 비동기 파라미터 서버 방식을 지원했던 분산 훈련 텐서플로 API입니다. 케라스와 마찬가지로 `tf.distribute.Strategy`를 `tf.estimator`와 함께 쓸 수 있습니다. 추정기 사용자는 아주 조금만 코드를 변경하면, 훈련이 분산되는 방식을 쉽게 바꿀 수 있습니다. 따라서 이제는 추정기 사용자들도 다중 GPU나 다중 워커뿐 아니라 다중 TPU에서 동기 방식으로 분산 훈련을 할 수 있습니다. 하지만 추정기는 제한적으로 지원하는 것입니다. 자세한 내용은 아래 [현재 어떤 것이 지원됩니까?](#estimator_support) 부분을 참고하십시오.
추정기와 함께 `tf.distribute.Strategy`를 사용하는 방법은 케라스와는 살짝 다릅니다. `strategy.scope`를 사용하는 대신에, 전략 객체를 추정기의 [`RunConfig`](https://www.tensorflow.org/api_docs/python/tf/estimator/RunConfig)(실행 설정)에 넣어서 전달해야합니다.
다음은 기본으로 제공되는 `LinearRegressor`와 `MirroredStrategy`를 함께 사용하는 방법을 보여주는 코드입니다.
```
mirrored_strategy = tf.distribute.MirroredStrategy()
config = tf.estimator.RunConfig(
train_distribute=mirrored_strategy, eval_distribute=mirrored_strategy)
regressor = tf.estimator.LinearRegressor(
feature_columns=[tf.feature_column.numeric_column('feats')],
optimizer='SGD',
config=config)
```
위 예제에서는 기본으로 제공되는 추정기를 사용하였지만, 직접 만든 추정기도 동일한 코드로 사용할 수 있습니다. `train_distribute`가 훈련을 어떻게 분산시킬지를 지정하고, `eval_distribute`가 평가를 어떻게 분산시킬지를 지정합니다. 케라스와 함께 사용할 때 훈련과 평가에 동일한 분산 전략을 사용했던 것과는 차이가 있습니다.
다음과 같이 입력 함수를 지정하면 추정기의 훈련과 평가를 할 수 있습니다.
```
def input_fn():
dataset = tf.data.Dataset.from_tensors(({"feats":[1.]}, [1.]))
return dataset.repeat(1000).batch(10)
regressor.train(input_fn=input_fn, steps=10)
regressor.evaluate(input_fn=input_fn, steps=10)
```
추정기와 케라스의 또 다른 점인 입력 처리 방식을 살펴봅시다. 케라스에서는 각 배치의 데이터가 여러 개의 복제된 작업으로 나누어진다고 설명했습니다. 하지만 추정기에서는 사용자가 `input_fn` 입력 함수를 제공하고, 데이터를 워커나 장비들에 어떻게 나누어 처리할지를 온전히 제어할 수 있습니다. 텐서플로는 배치의 데이터를 자동으로 나누지도 않고, 각 워커에 자동으로 분배하지도 않습니다. 제공된 `input_fn` 함수는 워커마다 한 번씩 호출됩니다. 따라서 워커마다 데이터셋을 받게 됩니다. 한 데이터셋의 배치 하나가 워커의 복제된 작업 하나에 들어가고, 따라서 워커 하나에 N개의 복제된 작업이 있으면 N개의 배치가 수행됩니다. 다시 말하자면 `input_fn`이 반환하는 데이터셋은 `PER_REPLICA_BATCH_SIZE` 즉 복제 작업 하나가 배치 하나에서 처리할 크기여야 합니다. 한 단계에서 처리하는 전체 배치 크기는 `PER_REPLICA_BATCH_SIZE * strategy.num_replicas_in_sync`가 됩니다. 다중 워커를 사용하여 훈련할 때는 데이터를 워커별로 쪼개거나, 아니면 각자 다른 임의의 순서로 섞는 것이 좋을 수도 있습니다. 이렇게 처리하는 예제는 [추정기로 다중 워커를 써서 훈련하기](../tutorials/distribute/multi_worker_with_estimator.ipynb)에서 볼 수 있습니다.
추정기와 함께 `MirroredStrategy`를 사용하는 예를 보았습니다. `TPUStrategy`도 같은 방법으로 추정기와 함께 사용할 수 있습니다.
```
config = tf.estimator.RunConfig(
train_distribute=tpu_strategy, eval_distribute=tpu_strategy)
```
비슷하게 다중 워커나 파라미터 서버를 사용한 전략도 사용할 수 있습니다. 코드는 거의 같지만, `tf.estimator.train_and_evaluate`를 사용해야 합니다. 그리고 클러스터에서 프로그램을 실행할 때 "TF_CONFIG" 환경변수를 설정해야 합니다.
### 현재 어떤 것이 지원됩니까?
TF 2.0 베타 버전에서는 추정기와 함께 모든 전략을 제한적으로 지원합니다. 기본적인 훈련과 평가는 동작합니다. 하지만 스캐폴드(scaffold) 같은 고급 기능은 아직 동작하지 않습니다. 또한 다소 버그가 있을 수 있습니다. 현재로써는 추정기와 함께 사용하는 것을 활발히 개선할 계획은 없습니다. 대신 케라스나 사용자 정의 훈련 루프 지원에 집중할 계획입니다. 만약 가능하다면 `tf.distribute` 사용시 이 API들을 먼저 고려하여 주십시오.
| 훈련 API | MirroredStrategy | TPUStrategy | MultiWorkerMirroredStrategy | CentralStorageStrategy | ParameterServerStrategy |
|:--------------- |:------------------ |:------------- |:----------------------------- |:------------------------ |:------------------------- |
| 추정기 API | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 | 제한적으로 지원 |
### 예제와 튜토리얼
다음은 추정기와 함께 다양한 전략을 사용하는 방법을 처음부터 끝까지 보여주는 예제들입니다.
1. [추정기로 다중 워커를 써서 훈련하기](../tutorials/distribute/multi_worker_with_estimator.ipynb)에서는 `MultiWorkerMirroredStrategy`로 다중 워커를 써서 MNIST를 훈련합니다.
2. [처음부터 끝까지 살펴보는 예제](https://github.com/tensorflow/ecosystem/tree/master/distribution_strategy)에서는 tensorflow/ecosystem의 쿠버네티스(Kubernetes) 템플릿을 이용하여 다중 워커를 사용하여 훈련합니다. 이 예제에서는 케라스 모델로 시작해서 `tf.keras.estimator.model_to_estimator` API를 이용하여 추정기 모델로 변환합니다.
3. `MirroredStrategy`나 `MultiWorkerMirroredStrategy`로 훈련할 수 있는 공식 [ResNet50](https://github.com/tensorflow/models/blob/master/official/r1/resnet/imagenet_main.py) 모델.
4. `TPUStrategy`를 사용한 [ResNet50](https://github.com/tensorflow/tpu/blob/master/models/experimental/distribution_strategy/resnet_estimator.py) 예제.
## 그 밖의 주제
이번 절에서는 다양한 사용 방식에 관련한 몇 가지 주제들을 다룹니다.
<a id="TF_CONFIG">
### TF\_CONFIG 환경변수 설정하기
</a>
다중 워커를 사용하여 훈련할 때는, 앞서 설명했듯이 클러스터의 각 실행 프로그램마다 "TF\_CONFIG" 환경변수를 설정해야합니다. "TF\_CONFIG" 환경변수는 JSON 형식입니다. 그 안에는 클러스터를 구성하는 작업과 작업의 주소 및 각 작업의 역할을 기술합니다. [tensorflow/ecosystem](https://github.com/tensorflow/ecosystem) 저장소에서 훈련 작업에 맞게 "TF\_CONFIG"를 설정하는 쿠버네티스(Kubernetes) 템플릿을 제공합니다.
"TF\_CONFIG" 예를 하나 보면 다음과 같습니다.
```
os.environ["TF_CONFIG"] = json.dumps({
"cluster": {
"worker": ["host1:port", "host2:port", "host3:port"],
"ps": ["host4:port", "host5:port"]
},
"task": {"type": "worker", "index": 1}
})
```
이 "TF\_CONFIG"는 세 개의 워커와 두 개의 파라미터 서버(ps) 작업을 각각의 호스트 및 포트와 함께 지정하고 있습니다. "task" 부분은 클러스터 내에서 현재 작업이 담당한 역할을 지정합니다. 여기서는 워커(worker) 1번, 즉 두 번째 워커라는 뜻입니다. 클러스터 내에서 가질 수 있는 역할은 "chief"(지휘자), "worker"(워커), "ps"(파라미터 서버), "evaluator"(평가자) 중 하나입니다. 단, "ps" 역할은 `tf.distribute.experimental.ParameterServerStrategy` 전략을 사용할 때만 쓸 수 있습니다.
## 다음으로는...
`tf.distribute.Strategy`는 활발하게 개발 중입니다. 한 번 써보시고 [깃허브 이슈](https://github.com/tensorflow/tensorflow/issues/new)를 통하여 피드백을 주시면 감사하겠습니다.
| github_jupyter |
# 10.16 Intro to Data Science: Time Series and Simple Linear Regression
### Time Series
### Simple Linear Regression
### Linear Relationships
```
%matplotlib inline
c = lambda f: 5 / 9 * (f - 32)
temps = [(f, c(f)) for f in range(0, 101, 10)]
import pandas as pd
temps_df = pd.DataFrame(temps, columns=['Fahrenheit', 'Celsius'])
axes = temps_df.plot(x='Fahrenheit', y='Celsius', style='.-')
y_label = axes.set_ylabel('Celsius')
# Extra cell added to keep subsequent snippet numbers the same as the chapter.
# Had to merge the two prior cells for use in the notebook.
```
### Components of the Simple Linear Regression Equation
### SciPy’s `stats` Module
### Pandas
### Seaborn Visualization
### Getting Weather Data from NOAA
### Loading the Average High Temperatures into a `DataFrame`
```
nyc = pd.read_csv('ave_hi_nyc_jan_1895-2018.csv')
nyc.head()
nyc.tail()
```
### Cleaning the Data
```
nyc.columns = ['Date', 'Temperature', 'Anomaly']
nyc.head(3)
nyc.Date.dtype
nyc.Date = nyc.Date.floordiv(100)
nyc.head(3)
```
### Calculating Basic Descriptive Statistics for the Dataset
```
pd.set_option('precision', 2)
nyc.Temperature.describe()
```
### Forecasting Future January Average High Temperatures
```
from scipy import stats
linear_regression = stats.linregress(x=nyc.Date,
y=nyc.Temperature)
linear_regression.slope
linear_regression.intercept
linear_regression.slope * 2019 + linear_regression.intercept
linear_regression.slope * 1850 + linear_regression.intercept
```
### Plotting the Average High Temperatures and a Regression Line
```
import seaborn as sns
sns.set_style('whitegrid')
axes = sns.regplot(x=nyc.Date, y=nyc.Temperature)
axes.set_ylim(10, 70)
# Extra cell added to keep subsequent snippet numbers the same as the chapter.
# Had to merge the two prior cells for use in the notebook.
```
### Getting Time Series Datasets
```
##########################################################################
# (C) Copyright 2019 by Deitel & Associates, Inc. and #
# Pearson Education, Inc. All Rights Reserved. #
# #
# DISCLAIMER: The authors and publisher of this book have used their #
# best efforts in preparing the book. These efforts include the #
# development, research, and testing of the theories and programs #
# to determine their effectiveness. The authors and publisher make #
# no warranty of any kind, expressed or implied, with regard to these #
# programs or to the documentation contained in these books. The authors #
# and publisher shall not be liable in any event for incidental or #
# consequential damages in connection with, or arising out of, the #
# furnishing, performance, or use of these programs. #
##########################################################################
```
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import hydra
from omegaconf import DictConfig
from omegaconf.omegaconf import OmegaConf
import numpy as np
from sklearn.datasets import load_digits
import matplotlib.pyplot as plt
time_series = np.random.rand(1000, 300)
time_series[500:560, 100:200] += 0.3
time_series = torch.from_numpy(time_series)
p = torch.from_numpy(np.abs(np.indices((100,100))[0] - np.indices((100,100))[1]))
sigma = torch.ones(100).view(100, 1) * 2
P = torch.ones(10,10) * torch.arange(10).view(10,1)
S = torch.ones(10,10) * torch.arange(10).view(1,10)
lambda row: F.kl_div(P[row,:], S[row,:]) + F.kl_div(S[row,:], P[row,:])
P
torch.arange(0, 10, 0.1)
torch.exp(p.pow(2) / (sigma))
gaussian = torch.normal(p.float(), sigma)
gaussian /= gaussian.sum(dim=-1).view(-1, 1)
gaussian[0,:].sum()
torch.ones(5,10)/ torch.ones(5,10).sum(dim=-1).view(-1,1)
torch.ones(5,10).sum(dim=-1)
torch.normal(torch.arange(0,100).float())
class AnomalyAttention(nn.Module):
def __init__(self, seq_dim, in_channels, out_channels):
super(AnomalyAttention, self).__init__()
self.W = nn.Linear(in_channels, out_channels, bias=False)
self.Q = self.K = self.V = self.sigma = torch.zeros((seq_dim, out_channels))
self.d_model = out_channels
self.n = seq_dim
self.P = torch.zeros((seq_dim, seq_dim))
self.S = torch.zeros((seq_dim, seq_dim))
def forward(self, x):
self.initialize(x) # does this make sense?
self.P = self.prior_association()
self.S = self.series_association()
Z = self.reconstruction()
return Z
def initialize(self, x):
# self.d_model = x.shape[-1]
self.Q = self.K = self.V = self.sigma = self.W(x)
def prior_association(self):
p = torch.from_numpy(
np.abs(
np.indices((self.n,self.n))[0] -
np.indices((self.n,self.n))[1]
)
)
gaussian = torch.normal(p.float(), self.sigma[:,0].abs())
gaussian /= gaussian.sum(dim=-1).view(-1, 1)
return gaussian
def series_association(self):
return F.softmax((self.Q @ self.K.T) / math.sqrt(self.d_model), dim=0)
def reconstruction(self):
return self.S @ self.V
def association_discrepancy(self):
return F.kl_div(self.P, self.S) + F.kl_div(self.S, self.P) #not going to be correct dimensions
class AnomalyTransformerBlock(nn.Module):
def __init__(self, seq_dim, feat_dim):
super().__init__()
self.seq_dim, self.feat_dim = seq_dim, feat_dim
self.attention = AnomalyAttention(self.seq_dim, self.feat_dim, self.feat_dim)
self.ln1 = nn.LayerNorm(self.feat_dim)
self.ff = nn.Sequential(
nn.Linear(self.feat_dim, self.feat_dim),
nn.ReLU()
)
self.ln2 = nn.LayerNorm(self.feat_dim)
self.association_discrepancy = None
def forward(self, x):
x_identity = x
x = self.attention(x)
z = self.ln1(x + x_identity)
z_identity = z
z = self.ff(z)
z = self.ln2(z + z_identity)
self.association_discrepancy = self.attention.association_discrepancy().detach()
return z
class AnomalyTransformer(nn.Module):
def __init__(self, seqs, in_channels, layers, lambda_):
super().__init__()
self.blocks = nn.ModuleList([
AnomalyTransformerBlock(seqs, in_channels) for _ in range(layers)
])
self.output = None
self.lambda_ = lambda_
self.assoc_discrepancy = torch.zeros((seqs, len(self.blocks)))
def forward(self, x):
for idx, block in enumerate(self.blocks):
x = block(x)
self.assoc_discrepancy[:, idx] = block.association_discrepancy
self.assoc_discrepancy = self.assoc_discrepancy.sum(dim=1) #N x 1
self.output = x
return x
def loss(self, x):
l2_norm = torch.linalg.matrix_norm(self.output - x, ord=2)
return l2_norm + (self.lambda_ * self.assoc_discrepancy.mean())
def anomaly_score(self, x):
score = F.softmax(-self.assoc_discrepancy, dim=0)
model = AnomalyTransformer(seqs=1000, in_channels=300, layers=3, lambda_=0.1)
model(time_series.float())
model.loss(time_series)
```
| github_jupyter |
# Hands-on: Deploying Question Answering with BERT
Pre-trained language representations have been shown to improve many downstream NLP tasks such as question answering, and natural language inference. Devlin, Jacob, et al proposed BERT [1] (Bidirectional Encoder Representations from Transformers), which fine-tunes deep bidirectional representations on a wide range of tasks with minimal task-specific parameters, and obtained state- of-the-art results.
After finishing training QA with BERT (the previous notebook "QA_Training.ipydb"), let us load a trained model to perform inference on the SQuAD dataset
### A quick overview: an example from SQuAD dataset is like below:
(2,
'56be4db0acb8001400a502ee',
'Where did Super Bowl 50 take place?',
'Super Bowl 50 was an American football game to determine the champion of the National
Football League (NFL) for the 2015 season. The American Football Conference (AFC)
champion Denver Broncos defeated the National Football Conference (NFC) champion
Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played
on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara,
California. As this was the 50th Super Bowl, the league emphasized the "golden
anniversary" with various gold-themed initiatives, as well as temporarily suspending
the tradition of naming each Super Bowl game with Roman numerals (under which the
game would have been known as "Super Bowl L"), so that the logo could prominently
feature the Arabic numerals 50.',
['Santa Clara, California', "Levi's Stadium", "Levi's Stadium
in the San Francisco Bay Area at Santa Clara, California."],
[403, 355, 355])
## Deploy on SageMaker
1. Preparing functions for inference
2. Saving the model parameters
3. Building a docker container with dependencies installed
4. Launching a serving end-point with SageMaker SDK
### 1. Preparing functions for inference
Two functions:
1. ```model_fn``` to load model parameters
2. ```transform_fn(``` to run model inference given an input
```
%%writefile code/serve.py
import collections, json, logging, warnings
import multiprocessing as mp
from functools import partial
import gluonnlp as nlp
import mxnet as mx
from mxnet.gluon import Block, nn
# import bert
from qa import preprocess_dataset, SQuADTransform
import bert_qa_evaluate
class BertForQA(Block):
"""Model for SQuAD task with BERT.
The model feeds token ids and token type ids into BERT to get the
pooled BERT sequence representation, then apply a Dense layer for QA task.
Parameters
----------
bert: BERTModel
Bidirectional encoder with transformer.
prefix : str or None
See document of `mx.gluon.Block`.
params : ParameterDict or None
See document of `mx.gluon.Block`.
"""
def __init__(self, bert, prefix=None, params=None):
super(BertForQA, self).__init__(prefix=prefix, params=params)
self.bert = bert
with self.name_scope():
self.span_classifier = nn.Dense(units=2, flatten=False)
def forward(self, inputs, token_types, valid_length=None): # pylint: disable=arguments-differ
"""Generate the unnormalized score for the given the input sequences.
Parameters
----------
inputs : NDArray, shape (batch_size, seq_length)
Input words for the sequences.
token_types : NDArray, shape (batch_size, seq_length)
Token types for the sequences, used to indicate whether the word belongs to the
first sentence or the second one.
valid_length : NDArray or None, shape (batch_size,)
Valid length of the sequence. This is used to mask the padded tokens.
Returns
-------
outputs : NDArray
Shape (batch_size, seq_length, 2)
"""
bert_output = self.bert(inputs, token_types, valid_length)
output = self.span_classifier(bert_output)
return output
def get_all_results(net, vocab, squadTransform, test_dataset, ctx = mx.cpu()):
all_results = collections.defaultdict(list)
def _vocab_lookup(example_id, subwords, type_ids, length, start, end):
indices = vocab[subwords]
return example_id, indices, type_ids, length, start, end
dev_data_transform, _ = preprocess_dataset(test_dataset, squadTransform)
dev_data_transform = dev_data_transform.transform(_vocab_lookup, lazy=False)
dev_dataloader = mx.gluon.data.DataLoader(dev_data_transform, batch_size=1, shuffle=False)
for data in dev_dataloader:
example_ids, inputs, token_types, valid_length, _, _ = data
batch_size = inputs.shape[0]
output = net(inputs.astype('float32').as_in_context(ctx),
token_types.astype('float32').as_in_context(ctx),
valid_length.astype('float32').as_in_context(ctx))
pred_start, pred_end = mx.nd.split(output, axis=2, num_outputs=2)
example_ids = example_ids.asnumpy().tolist()
pred_start = pred_start.reshape(batch_size, -1).asnumpy()
pred_end = pred_end.reshape(batch_size, -1).asnumpy()
for example_id, start, end in zip(example_ids, pred_start, pred_end):
all_results[example_id].append(bert_qa_evaluate.PredResult(start=start, end=end))
return(all_results)
def _test_example_transform(test_examples):
"""
Change test examples to a format like SQUAD data.
Parameters
----------
test_examples: a list of (question, context) tuple.
Example: [('Which NFL team represented the AFC at Super Bowl 50?',
'Super Bowl 50 was an American football game ......),
('Where did Super Bowl 50 take place?',,
'Super Bowl 50 was ......),
......]
Returns
----------
test_examples_tuples : a list of SQUAD tuples
"""
test_examples_tuples = []
i = 0
for test in test_examples:
question, context = test[0], test[1] # test.split(" [CONTEXT] ")
tup = (i, "", question, context, [], [])
test_examples_tuples.append(tup)
i += 1
return(test_examples_tuples)
def model_fn(model_dir = "", params_path = "bert_qa-7eb11865.params"):
"""
Load the gluon model. Called once when hosting service starts.
:param: model_dir The directory where model files are stored.
:return: a Gluon model, and the vocabulary
"""
bert_model, vocab = nlp.model.get_model('bert_12_768_12',
dataset_name='book_corpus_wiki_en_uncased',
use_classifier=False,
use_decoder=False,
use_pooler=False,
pretrained=False)
net = BertForQA(bert_model)
if len(model_dir) > 0:
params_path = model_dir + "/" +params_path
net.load_parameters(params_path, ctx=mx.cpu())
tokenizer = nlp.data.BERTTokenizer(vocab, lower=True)
transform = SQuADTransform(tokenizer, is_pad=False, is_training=False, do_lookup=False)
return net, vocab, transform
def transform_fn(model, input_data, input_content_type=None, output_content_type=None):
"""
Transform a request using the Gluon model. Called once per request.
:param model: The Gluon model and the vocab
:param dataset: The request payload
Example:
## (example_id, [question, content], ques_cont_token_types, valid_length, _, _)
(2,
'56be4db0acb8001400a502ee',
'Where did Super Bowl 50 take place?',
'Super Bowl 50 was an American football game to determine the champion of the National
Football League (NFL) for the 2015 season. The American Football Conference (AFC)
champion Denver Broncos defeated the National Football Conference (NFC) champion
Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played
on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara,
California. As this was the 50th Super Bowl, the league emphasized the "golden
anniversary" with various gold-themed initiatives, as well as temporarily suspending
the tradition of naming each Super Bowl game with Roman numerals (under which the
game would have been known as "Super Bowl L"), so that the logo could prominently
feature the Arabic numerals 50.',
['Santa Clara, California', "Levi's Stadium", "Levi's Stadium
in the San Francisco Bay Area at Santa Clara, California."],
[403, 355, 355])
:param input_content_type: The request content type, assume json
:param output_content_type: The (desired) response content type, assume json
:return: response payload and content type.
"""
net, vocab, squadTransform = model
# data = input_data
data = json.loads(input_data)
# test_examples_tuples = [(i, "", question, content, [], [])]
# question, context = data #.split(" [CONTEXT] ")
# tup = (0, "", question, context, [], [])
test_examples_tuples = _test_example_transform(data)
test_dataset = mx.gluon.data.SimpleDataset(test_examples_tuples) # [tup]
all_results = get_all_results(net, vocab, squadTransform, test_dataset, ctx=mx.cpu())
all_predictions = collections.defaultdict(list) # collections.OrderedDict()
data_transform = test_dataset.transform(squadTransform._transform)
for features in data_transform:
f_id = features[0].example_id
results = all_results[f_id]
prediction, nbest = bert_qa_evaluate.predict(
features=features,
results=results,
tokenizer=nlp.data.BERTBasicTokenizer(vocab))
nbest_prediction = []
for i in range(3):
nbest_prediction.append('%.2f%% \t %s'%(nbest[i][1] * 100, nbest[i][0]))
all_predictions[f_id] = nbest_prediction
response_body = json.dumps(all_predictions)
return response_body, output_content_type
```
### 2. Saving the model parameters
We are going to zip the BERT model parameters, vocabulary file, and all the inference files (```code/serve.py```, ```bert/data/qa.py```, ```bert_qa_evaluate.py```) to a ```model.tar.gz``` file. (Note that the ```serve.py``` is the "entry_point" for Sagemaker to do the inference, and it needs to be under ```code/``` directory.)
```
import tarfile
with tarfile.open("model.tar.gz", "w:gz") as tar:
# tar.add("code/serve.py")
# tar.add("bert/data/qa.py")
# tar.add("bert_qa_evaluate.py")
# tar.add("bert_qa-7eb11865.params")
# tar.add("vocab.json")
tar.add("net.params")
```
### 3. Building a docker container with dependencies installed
Let's prepare a docker container with all the dependencies required for model inference. Here we build a docker container based on the SageMaker MXNet inference container, and you can find the list of all available inference containers at https://docs.aws.amazon.com/sagemaker/latest/dg/pre-built-containers-frameworks-deep-learning.html
Here we use local mode for demonstration purpose. To deploy on actual instances, you need to login into AWS elastic container registry (ECR) service, and push the container to ECR.
```
docker build -t $YOUR_EDR_DOCKER_TAG . -f Dockerfile
$(aws ecr get-login --no-include-email --region $YOUR_REGION)
docker push $YOUR_EDR_DOCKER_TAG
```
```
%%writefile Dockerfile
FROM nvidia/cuda:10.1-cudnn7-runtime-ubuntu16.04
LABEL maintainer="Amazon AI"
# Specify accept-bind-to-port LABEL for inference pipelines to use SAGEMAKER_BIND_TO_PORT
# https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipeline-real-time.html
LABEL com.amazonaws.sagemaker.capabilities.accept-bind-to-port=true
ARG MMS_VERSION=1.0.8
ARG MX_URL=https://aws-mxnet-pypi.s3-us-west-2.amazonaws.com/1.6.0/aws_mxnet_cu101mkl-1.6.0-py2.py3-none-manylinux1_x86_64.whl
ARG PYTHON=python3
ARG PYTHON_PIP=python3-pip
ARG PIP=pip3
ARG PYTHON_VERSION=3.6.8
ENV PYTHONDONTWRITEBYTECODE=1 \
PYTHONUNBUFFERED=1 \
LD_LIBRARY_PATH="${LD_LIBRARY_PATH}:/usr/local/lib" \
PYTHONIOENCODING=UTF-8 \
LANG=C.UTF-8 \
LC_ALL=C.UTF-8 \
TEMP=/home/model-server/tmp \
CLOUD_PATH="/opt/ml/code"
RUN apt-get update \
&& apt-get -y install --no-install-recommends \
build-essential \
ca-certificates \
curl \
git \
libopencv-dev \
openjdk-8-jdk-headless \
vim \
wget \
zlib1g-dev \
&& apt-get clean \
&& rm -rf /var/lib/apt/lists/*
RUN wget https://www.python.org/ftp/python/$PYTHON_VERSION/Python-$PYTHON_VERSION.tgz \
&& tar -xvf Python-$PYTHON_VERSION.tgz \
&& cd Python-$PYTHON_VERSION \
&& ./configure \
&& make \
&& make install \
&& apt-get update \
&& apt-get install -y --no-install-recommends \
libreadline-gplv2-dev \
libncursesw5-dev \
libssl-dev \
libsqlite3-dev \
tk-dev \
libgdbm-dev \
libc6-dev \
libbz2-dev \
&& make \
&& make install \
&& rm -rf ../Python-$PYTHON_VERSION* \
&& ln -s /usr/local/bin/pip3 /usr/bin/pip
RUN ln -s $(which ${PYTHON}) /usr/local/bin/python
RUN ${PIP} --no-cache-dir install --upgrade \
pip \
setuptools
WORKDIR /
RUN ${PIP} install --no-cache-dir \
${MX_URL} \
git+git://github.com/dmlc/gluon-nlp.git@v0.9.0 \
# gluoncv==0.6.0 \
mxnet-model-server==$MMS_VERSION \
keras-mxnet==2.2.4.1 \
numpy==1.17.4 \
onnx==1.4.1 \
"sagemaker-mxnet-inference<2"
RUN useradd -m model-server \
&& mkdir -p /home/model-server/tmp \
&& chown -R model-server /home/model-server
COPY mms-entrypoint.py /usr/local/bin/dockerd-entrypoint.py
COPY config.properties /home/model-server
COPY code/serve.py $CLOUD_PATH/serve.py
COPY bert_qa_evaluate.py $CLOUD_PATH/bert_qa_evaluate.py
COPY qa.py $CLOUD_PATH/qa.py
RUN chmod +x /usr/local/bin/dockerd-entrypoint.py
RUN curl https://aws-dlc-licenses.s3.amazonaws.com/aws-mxnet-1.6.0/license.txt -o /license.txt
EXPOSE 8080 8081
ENTRYPOINT ["python", "/usr/local/bin/dockerd-entrypoint.py"]
CMD ["mxnet-model-server", "--start", "--mms-config", "/home/model-server/config.properties"]
%%writefile build.sh
#!/usr/bin/env bash
# This script shows how to build the Docker image and push it to ECR to be ready for use
# by SageMaker.
# The arguments to this script are the image name and application name
image=$1
app=$2
chmod +x $app/train
chmod +x $app/serve
# Get the account number associated with the current IAM credentials
account=$(aws sts get-caller-identity --query Account --output text)
# Get the region defined in the current configuration
region=$(aws configure get region)
fullname="${account}.dkr.ecr.${region}.amazonaws.com/${image}:latest"
# If the repository doesn't exist in ECR, create it.
aws ecr describe-repositories --repository-names "${image}" > /dev/null 2>&1
if [ $? -ne 0 ]
then
aws ecr create-repository --repository-name "${image}" > /dev/null
fi
# Edit ECR policy permission rights
aws ecr set-repository-policy --repository-name "${image}" --policy-text ecr_policy.json
# Get the login command from ECR and execute it directly
$(aws ecr get-login --region ${region} --no-include-email)
# Build the docker image locally with the image name and then push it to ECR
# with the full name.
docker build -t ${image} --build-arg APP=$app .
docker tag ${image} ${fullname}
```
Set `image_name` as "kdd2020nlp", and application name as "question_answering"
```
!bash build.sh kdd2020nlp question_answering
```
### 4. Launching a serving end-point with SageMaker SDK
We create a MXNet model which can be deployed later, by specifying the docker image, and entry point for the inference code. If ```serve.py``` does not work, use ```dummy_hosting_module.py``` for debugging purpose.
#### Creating the Session
The session remembers our connection parameters to Amazon SageMaker. We'll use it to perform all of our SageMaker operations.
```
import sagemaker as sage
sess = sage.Session()
```
#### Defining the account, region and ECR address
```
account = sess.boto_session.client('sts').get_caller_identity()['Account']
region = sess.boto_session.region_name
image_name = "kdd2020nlp"
ecr_image = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(account, region, image_name)
```
#### Uploading model
We can upload the trained model to the corresponding S3 bucket: https://s3.console.aws.amazon.com/s3/buckets/sagemaker-us-east-1-383827541835/sagemaker-deploy-gluoncv/data/?region=us-east-1
```
sess.default_bucket()
sess.update_endpoint
s3_bucket_name = "kdd2020"
model_path = "s3://{}/{}/model".format(sess.default_bucket(), s3_bucket_name)
os.path.join(model_path, "model.tar.gz")
model_prefix = s3_bucket + "/model"
train_data_local = "./data/minc-2500/train"
train_data_dir_prefix = s3_bucket + "/data/train"
# model_local_path = "model_output"
train_data_upload = sess.upload_data(path=train_data_local,
# bucket=s3_bucket,
key_prefix=train_data_dir_prefix)
import sagemaker
from sagemaker.mxnet.model import MXNetModel
sagemaker_model = MXNetModel(model_data='file:///home/ec2-user/SageMaker/ako2020-bert/tutorial/model.tar.gz',
image=ecr_image,
role=sagemaker.get_execution_role(),
py_version='py3', # python version
entry_point='serve.py',
source_dir='.')
```
We use 'local' mode to test our deployment code, where the inference happens on the current instance.
If you are ready to deploy the model on a new instance, change the `instance_type` argument to values such as `ml.c4.xlarge`.
Here we use 'local' mode for testing, for real instances use c5.2xlarge, p2.xlarge, etc. **The following line will start docker container building.**
```
predictor = sagemaker_model.deploy(initial_instance_count=1, instance_type='local')
```
Now let us try to submit a inference job. Here we simply grab two datapoints from the SQuAD dataset and pass the examples to our predictor by calling ```predictor.predict```
```
## test
my_test_example_0 = ('Which NFL team represented the AFC at Super Bowl 50?',
'Super Bowl 50 was an American football game to determine the champion of the National Football League (NFL) for the 2015 season. The American Football Conference (AFC) champion Denver Broncos defeated the National Football Conference (NFC) champion Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara, California. As this was the 50th Super Bowl, the league emphasized the "golden anniversary" with various gold-themed initiatives, as well as temporarily suspending the tradition of naming each Super Bowl game with Roman numerals (under which the game would have been known as "Super Bowl L"), so that the logo could prominently feature the Arabic numerals 50.')
my_test_example_1 = ('Where did Super Bowl 50 take place?',
'Super Bowl 50 was an American football game to determine the champion of the National Football League (NFL) for the 2015 season. The American Football Conference (AFC) champion Denver Broncos defeated the National Football Conference (NFC) champion Carolina Panthers 24–10 to earn their third Super Bowl title. The game was played on February 7, 2016, at Levi\'s Stadium in the San Francisco Bay Area at Santa Clara, California. As this was the 50th Super Bowl, the league emphasized the "golden anniversary" with various gold-themed initiatives, as well as temporarily suspending the tradition of naming each Super Bowl game with Roman numerals (under which the game would have been known as "Super Bowl L"), so that the logo could prominently feature the Arabic numerals 50.')
my_test_examples = (my_test_example_0, my_test_example_1)
# mymodel = model_fn(params_path = "bert_qa-7eb11865.params")
# transform_fn(mymodel, my_test_examples)
output = predictor.predict(my_test_examples)
print("\nPrediction output: \n\n")
for k in output.keys():
print('{}\n\n'.format(output[k]))
```
### Clean Up
Remove the endpoint after we are done.
```
predictor.delete_endpoint()
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Probability Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Copulas Primer
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/probability/examples/Gaussian_Copula"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/Gaussian_Copula.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/probability/blob/master/tensorflow_probability/examples/jupyter_notebooks/Gaussian_Copula.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/probability/tensorflow_probability/examples/jupyter_notebooks/Gaussian_Copula.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
```
import numpy as np
import matplotlib.pyplot as plt
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
import tensorflow_probability as tfp
tfd = tfp.distributions
tfb = tfp.bijectors
```
A [copula](https://en.wikipedia.org/wiki/Copula_(probability_theory%29) is a classical approach for capturing the dependence between random variables. More formally, a copula is a multivariate distribution $C(U_1, U_2, ...., U_n)$ such that marginalizing gives $U_i \sim \text{Uniform}(0, 1)$.
Copulas are interesting because we can use them to create multivariate distributions with arbitrary marginals. This is the recipe:
* Using the [Probability Integral Transform](https://en.wikipedia.org/wiki/Probability_integral_transform) turns an arbitrary continuous R.V. $X$ into a uniform one $F_X(X)$, where $F_X$ is the CDF of $X$.
* Given a copula (say bivariate) $C(U, V)$, we have that $U$ and $V$ have uniform marginal distributions.
* Now given our R.V's of interest $X, Y$, create a new distribution $C'(X, Y) = C(F_X(X), F_Y(Y))$. The marginals for $X$ and $Y$ are the ones we desired.
Marginals are univariate and thus may be easier to measure and/or model. A copula enables starting from marginals yet also achieving arbitrary correlation between dimensions.
# Gaussian Copula
To illustrate how copulas are constructed, consider the case of capturing dependence according to multivariate Gaussian correlations. A Gaussian Copula is one given by $C(u_1, u_2, ...u_n) = \Phi_\Sigma(\Phi^{-1}(u_1), \Phi^{-1}(u_2), ... \Phi^{-1}(u_n))$ where $\Phi_\Sigma$ represents the CDF of a MultivariateNormal, with covariance $\Sigma$ and mean 0, and $\Phi^{-1}$ is the inverse CDF for the standard normal.
Applying the normal's inverse CDF warps the uniform dimensions to be normally distributed. Applying the multivariate normal's CDF then squashes the distribution to be marginally uniform and with Gaussian correlations.
Thus, what we get is that the Gaussian Copula is a distribution over the unit hypercube $[0, 1]^n$ with uniform marginals.
Defined as such, the Gaussian Copula can be implemented with `tfd.TransformedDistribution` and appropriate `Bijector`. That is, we are transforming a MultivariateNormal, via the use of the Normal distribution's inverse CDF, implemented by the [`tfb.NormalCDF`](https://www.tensorflow.org/probability/api_docs/python/tfp/bijectors/NormalCDF) bijector.
Below, we implement a Gaussian Copula with one simplifying assumption: that the covariance is parameterized
by a Cholesky factor (hence a covariance for `MultivariateNormalTriL`). (One could use other `tf.linalg.LinearOperators` to encode different matrix-free assumptions.).
```
class GaussianCopulaTriL(tfd.TransformedDistribution):
"""Takes a location, and lower triangular matrix for the Cholesky factor."""
def __init__(self, loc, scale_tril):
super(GaussianCopulaTriL, self).__init__(
distribution=tfd.MultivariateNormalTriL(
loc=loc,
scale_tril=scale_tril),
bijector=tfb.NormalCDF(),
validate_args=False,
name="GaussianCopulaTriLUniform")
# Plot an example of this.
unit_interval = np.linspace(0.01, 0.99, num=200, dtype=np.float32)
x_grid, y_grid = np.meshgrid(unit_interval, unit_interval)
coordinates = np.concatenate(
[x_grid[..., np.newaxis],
y_grid[..., np.newaxis]], axis=-1)
pdf = GaussianCopulaTriL(
loc=[0., 0.],
scale_tril=[[1., 0.8], [0., 0.6]],
).prob(coordinates)
# Plot its density.
plt.contour(x_grid, y_grid, pdf, 100, cmap=plt.cm.jet);
```
The power, however, from such a model is using the Probability Integral Transform, to use the copula on arbitrary R.V.s. In this way, we can specify arbitrary marginals, and use the copula to stitch them together.
We start with a model:
$$\begin{align*}
X &\sim \text{Kumaraswamy}(a, b) \\
Y &\sim \text{Gumbel}(\mu, \beta)
\end{align*}$$
and use the copula to get a bivariate R.V. $Z$, which has marginals [Kumaraswamy](https://en.wikipedia.org/wiki/Kumaraswamy_distribution) and [Gumbel](https://en.wikipedia.org/wiki/Gumbel_distribution).
We'll start by plotting the product distribution generated by those two R.V.s. This is just to serve as a comparison point to when we apply the Copula.
```
a = 2.0
b = 2.0
gloc = 0.
gscale = 1.
x = tfd.Kumaraswamy(a, b)
y = tfd.Gumbel(loc=gloc, scale=gscale)
# Plot the distributions, assuming independence
x_axis_interval = np.linspace(0.01, 0.99, num=200, dtype=np.float32)
y_axis_interval = np.linspace(-2., 3., num=200, dtype=np.float32)
x_grid, y_grid = np.meshgrid(x_axis_interval, y_axis_interval)
pdf = x.prob(x_grid) * y.prob(y_grid)
# Plot its density
plt.contour(x_grid, y_grid, pdf, 100, cmap=plt.cm.jet);
```
# Joint Distribution with Different Marginals
Now we use a Gaussian copula to couple the distributions together, and plot that. Again our tool of choice is `TransformedDistribution` applying the appropriate `Bijector` to obtain the chosen marginals.
Specifically, we use a [`Blockwise`](https://www.tensorflow.org/probability/api_docs/python/tfp/bijectors/Blockwise) bijector which applies different bijectors at different parts of the vector (which is still a bijective transformation).
Now we can define the Copula we want. Given a list of target marginals (encoded as bijectors), we can easily construct
a new distribution that uses the copula and has the specified marginals.
```
class WarpedGaussianCopula(tfd.TransformedDistribution):
"""Application of a Gaussian Copula on a list of target marginals.
This implements an application of a Gaussian Copula. Given [x_0, ... x_n]
which are distributed marginally (with CDF) [F_0, ... F_n],
`GaussianCopula` represents an application of the Copula, such that the
resulting multivariate distribution has the above specified marginals.
The marginals are specified by `marginal_bijectors`: These are
bijectors whose `inverse` encodes the CDF and `forward` the inverse CDF.
block_sizes is a 1-D Tensor to determine splits for `marginal_bijectors`
length should be same as length of `marginal_bijectors`.
See tfb.Blockwise for details
"""
def __init__(self, loc, scale_tril, marginal_bijectors, block_sizes=None):
super(WarpedGaussianCopula, self).__init__(
distribution=GaussianCopulaTriL(loc=loc, scale_tril=scale_tril),
bijector=tfb.Blockwise(bijectors=marginal_bijectors,
block_sizes=block_sizes),
validate_args=False,
name="GaussianCopula")
```
Finally, let's actually use this Gaussian Copula. We'll use a Cholesky of $\begin{bmatrix}1 & 0\\\rho & \sqrt{(1-\rho^2)}\end{bmatrix}$, which will correspond to variances 1, and correlation $\rho$ for the multivariate normal.
We'll look at a few cases:
```
# Create our coordinates:
coordinates = np.concatenate(
[x_grid[..., np.newaxis], y_grid[..., np.newaxis]], -1)
def create_gaussian_copula(correlation):
# Use Gaussian Copula to add dependence.
return WarpedGaussianCopula(
loc=[0., 0.],
scale_tril=[[1., 0.], [correlation, tf.sqrt(1. - correlation ** 2)]],
# These encode the marginals we want. In this case we want X_0 has
# Kumaraswamy marginal, and X_1 has Gumbel marginal.
marginal_bijectors=[
tfb.Invert(tfb.KumaraswamyCDF(a, b)),
tfb.Invert(tfb.GumbelCDF(loc=0., scale=1.))])
# Note that the zero case will correspond to independent marginals!
correlations = [0., -0.8, 0.8]
copulas = []
probs = []
for correlation in correlations:
copula = create_gaussian_copula(correlation)
copulas.append(copula)
probs.append(copula.prob(coordinates))
# Plot it's density
for correlation, copula_prob in zip(correlations, probs):
plt.figure()
plt.contour(x_grid, y_grid, copula_prob, 100, cmap=plt.cm.jet)
plt.title('Correlation {}'.format(correlation))
```
Finally, let's verify that we actually get the marginals we want.
```
def kumaraswamy_pdf(x):
return tfd.Kumaraswamy(a, b).prob(np.float32(x))
def gumbel_pdf(x):
return tfd.Gumbel(gloc, gscale).prob(np.float32(x))
copula_samples = []
for copula in copulas:
copula_samples.append(copula.sample(10000))
plot_rows = len(correlations)
plot_cols = 2 # for 2 densities [kumarswamy, gumbel]
fig, axes = plt.subplots(plot_rows, plot_cols, sharex='col', figsize=(18,12))
# Let's marginalize out on each, and plot the samples.
for i, (correlation, copula_sample) in enumerate(zip(correlations, copula_samples)):
k = copula_sample[..., 0].numpy()
g = copula_sample[..., 1].numpy()
_, bins, _ = axes[i, 0].hist(k, bins=100, normed=True)
axes[i, 0].plot(bins, kumaraswamy_pdf(bins), 'r--')
axes[i, 0].set_title('Kumaraswamy from Copula with correlation {}'.format(correlation))
_, bins, _ = axes[i, 1].hist(g, bins=100, normed=True)
axes[i, 1].plot(bins, gumbel_pdf(bins), 'r--')
axes[i, 1].set_title('Gumbel from Copula with correlation {}'.format(correlation))
```
# Conclusion
And there we go! We've demonstrated that we can construct Gaussian Copulas using the `Bijector` API.
More generally, writing bijectors using the `Bijector` API and composing them with a distribution, can create rich families of distributions for flexible modelling.
| github_jupyter |
# Quadratic Program (QP) Tutorial
For instructions on how to run these tutorial notebooks, please see the [README](https://github.com/RobotLocomotion/drake/blob/master/tutorials/README.md).
## Important Note
Please refer to [mathematical program tutorial](./mathematical_program.ipynb) for constructing and solving a general optimization program in Drake.
A (convex) quadratic program (QP) is a special type of convex optimization. Its cost function is a convex quadratic function. Its constraints are linear, same as the constraints in linear program. A (convex) quadratic program has the following form
\begin{align}
\min_x 0.5 x^TQx + b^Tx + c\\
\text{s.t } Ex \leq f
\end{align}
where `Q` is a positive semidefinite matrix.
A quadratic program can be solved by many different solvers. Drake supports some solvers including OSQP, SCS, Gurobi, Mosek, etc. Please see our [Doxygen page]( https://drake.mit.edu/doxygen_cxx/group__solvers.html) for a complete list of supported solvers. Note that some commercial solvers (such as Gurobi and Mosek) are not included in the pre-compiled Drake binaries, and therefore not on Binder/Colab.
Drake's API supports multiple functions to add quadratic cost and linear constraints. We briefly go through some of the functions in this tutorial. For a complete list of functions, please check our [Doxygen](https://drake.mit.edu/doxygen_cxx/classdrake_1_1solvers_1_1_mathematical_program.html).
There are many applications of quadratic programs in robotics, for example, we can solve differential inverse kinematics problem as a QP, see [DifferentialInverseKinematics](https://drake.mit.edu/doxygen_cxx/namespacedrake_1_1manipulation_1_1planner.html#ab53fd2e1578db60ceb43b754671ae539) for more details. For more examples, check out [Underactuated Robotics code repo](https://github.com/RussTedrake/underactuated)
## Add quadratic cost
### AddQuadraticCost function
The easist way to add a quadratic cost is to call the `AddQuadraticCost` function. In the following code snippet, we first construct a program with 2 decision variables, and then show how to call the `AddQuadraticCost` function.
```
from pydrake.solvers.mathematicalprogram import MathematicalProgram, Solve
import numpy as np
# Create an empty MathematicalProgram named prog (with no decision variables,
# constraints or costs)
prog = MathematicalProgram()
# Add two decision variables x[0], x[1].
x = prog.NewContinuousVariables(2, "x")
```
We can call `AddQuadraticCost(expression)` to add the quadratic cost, where `expression` is a symbolic expression representing a quadratic cost.
```
# Add a symbolic quadratic expression as the quadratic cost.
cost1 = prog.AddQuadraticCost(x[0]**2 + 2*x[0]*x[1] + x[1]**2 + 3*x[0] + 4)
# The newly added cost is returned as cost1
print(cost1)
# The newly added cost is stored inside prog.
print(prog.quadratic_costs()[0])
```
If we call `AddQuadraticCost` again, the total cost of `prog` is the summation of all the added costs. You can see that `prog.quadratic_costs()` has two entries. And the total cost of `prog` is `cost1 + cost2`.
```
# Add another quadratic cost to prog.
cost2 = prog.AddQuadraticCost(x[1]*x[1] + 3)
print(f"The number of quadratic costs in prog: {len(prog.quadratic_costs())}")
```
If you know the coefficients of the quadratic cost `Q, b, c`, you could also add the cost without using the symbolic expression, as shown in the following code snippet
```
# Add the cost x[0]*x[0] + x[0]*x[1] + 1.5*x[1]*x[1] + 2*x[0] + 4*x[1] + 1
cost3 = prog.AddQuadraticCost(Q=[[2, 1], [1, 3]], b=[2, 4], c=1, vars=x)
print(f"cost 3 is {cost3}")
```
### AddQuadraticErrorCost
You could also add the quadratic cost
\begin{align}
(x - x_{desired})^TQ(x-x_{desired})
\end{align}
to the program by calling `AddQuadraticErrorCost`. Here is the code example
```
# Adds the cost (x - [1;2])' * Q * (x-[1;2])
cost4 = prog.AddQuadraticErrorCost(Q=[[1, 2],[2, 6]], x_desired=[1,2], vars=x)
print(f"cost4 is {cost4}")
```
### AddL2NormCost
You could also add the quadratic cost
\begin{align}
|Ax-b|^2
\end{align}
which is the squared L2 norm of the vector `Ax-b` to the program by calling `AddL2NormCost`. Here is the code example
```
# Adds the squared norm of (x[0]+2*x[1]-2, x[1] - 3) to the program cost.
cost5 = prog.AddL2NormCost(A=[[1, 2], [0, 1]], b=[2, 3], vars=x)
print(f"cost5 is {cost5}")
```
## Add linear cost
You could also add linear costs to a quadratic program. For an introduction on the different APIs on adding a linear cost, please refer to our [linear programming tutorial](./linear_program.ipynb). Here is an example
```
# Adds a linear cost to the quadratic program
cost6 = prog.AddLinearCost(x[0] + 3 * x[1] + 1)
print(f"cost6 is {cost6}")
print(f"Number of linear costs in prog: {len(prog.linear_costs())}")
```
## Add linear constraints
To add linear constraints into quadratic program, please refer to the section `Add linear constraints` in our [linear programming tutorial](./linear_program.ipynb). Here is a brief example
```
constraint1 = prog.AddLinearConstraint(x[0] + 3*x[1] <= 5)
# Adds the constraint 1 <= x[0] <= 5 and 1 <= x[1] <= 5
constraint2 = prog.AddBoundingBoxConstraint(1, 5, x)
```
## A complete code example
Here we show a complete example to construct and solve a QP
```
prog = MathematicalProgram()
x = prog.NewContinuousVariables(3, "x")
prog.AddQuadraticCost(x[0] * x[0] + 2 * x[0] + 3)
# Adds the quadratic cost on the squared norm of the vector
# (x[1] + 3*x[2] - 1, 2*x[1] + 4*x[2] -4)
prog.AddL2NormCost(A = [[1, 3], [2, 4]], b=[1, 4], vars=[x[1], x[2]])
# Adds the linear constraints.
prog.AddLinearEqualityConstraint(x[0] + 2*x[1] == 5)
prog.AddLinearConstraint(x[0] + 4 *x[1] <= 10)
# Sets the bounds for each variable to be within [-1, 10]
prog.AddBoundingBoxConstraint(-1, 10, x)
# Solve the program.
result = Solve(prog)
print(f"optimal solution x: {result.GetSolution(x)}")
print(f"optimal cost: {result.get_optimal_cost()}")
```
For more details on quadratic programming, you could refer to
[Quadratic Programming wiki](https://en.wikipedia.org/wiki/Quadratic_programming)
[Numerical Optimization by J. Nocedal and S.Wright](http://www.apmath.spbu.ru/cnsa/pdf/monograf/Numerical_Optimization2006.pdf)
| github_jupyter |
# Convolutional Neural Networks: Step by Step
Welcome to Course 4's first assignment! In this assignment, you will implement convolutional (CONV) and pooling (POOL) layers in numpy, including both forward propagation and (optionally) backward propagation.
**Notation**:
- Superscript $[l]$ denotes an object of the $l^{th}$ layer.
- Example: $a^{[4]}$ is the $4^{th}$ layer activation. $W^{[5]}$ and $b^{[5]}$ are the $5^{th}$ layer parameters.
- Superscript $(i)$ denotes an object from the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example input.
- Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the activations in layer $l$, assuming this is a fully connected (FC) layer.
- $n_H$, $n_W$ and $n_C$ denote respectively the height, width and number of channels of a given layer. If you want to reference a specific layer $l$, you can also write $n_H^{[l]}$, $n_W^{[l]}$, $n_C^{[l]}$.
- $n_{H_{prev}}$, $n_{W_{prev}}$ and $n_{C_{prev}}$ denote respectively the height, width and number of channels of the previous layer. If referencing a specific layer $l$, this could also be denoted $n_H^{[l-1]}$, $n_W^{[l-1]}$, $n_C^{[l-1]}$.
We assume that you are already familiar with `numpy` and/or have completed the previous courses of the specialization. Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
You will be implementing the building blocks of a convolutional neural network! Each function you will implement will have detailed instructions that will walk you through the steps needed:
- Convolution functions, including:
- Zero Padding
- Convolve window
- Convolution forward
- Convolution backward (optional)
- Pooling functions, including:
- Pooling forward
- Create mask
- Distribute value
- Pooling backward (optional)
This notebook will ask you to implement these functions from scratch in `numpy`. In the next notebook, you will use the TensorFlow equivalents of these functions to build the following model:
<img src="images/model.png" style="width:800px;height:300px;">
**Note** that for every forward function, there is its corresponding backward equivalent. Hence, at every step of your forward module you will store some parameters in a cache. These parameters are used to compute gradients during backpropagation.
## 3 - Convolutional Neural Networks
Although programming frameworks make convolutions easy to use, they remain one of the hardest concepts to understand in Deep Learning. A convolution layer transforms an input volume into an output volume of different size, as shown below.
<img src="images/conv_nn.png" style="width:350px;height:200px;">
In this part, you will build every step of the convolution layer. You will first implement two helper functions: one for zero padding and the other for computing the convolution function itself.
### 3.1 - Zero-Padding
Zero-padding adds zeros around the border of an image:
<img src="images/PAD.png" style="width:600px;height:400px;">
<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **Zero-Padding**<br> Image (3 channels, RGB) with a padding of 2. </center></caption>
The main benefits of padding are the following:
- It allows you to use a CONV layer without necessarily shrinking the height and width of the volumes. This is important for building deeper networks, since otherwise the height/width would shrink as you go to deeper layers. An important special case is the "same" convolution, in which the height/width is exactly preserved after one layer.
- It helps us keep more of the information at the border of an image. Without padding, very few values at the next layer would be affected by pixels as the edges of an image.
**Exercise**: Implement the following function, which pads all the images of a batch of examples X with zeros. [Use np.pad](https://docs.scipy.org/doc/numpy/reference/generated/numpy.pad.html). Note if you want to pad the array "a" of shape $(5,5,5,5,5)$ with `pad = 1` for the 2nd dimension, `pad = 3` for the 4th dimension and `pad = 0` for the rest, you would do:
```python
a = np.pad(a, ((0,0), (1,1), (0,0), (3,3), (0,0)), 'constant', constant_values = (..,..))
```
```
# GRADED FUNCTION: zero_pad
def zero_pad(X, pad):
"""
Pad with zeros all images of the dataset X. The padding is applied to the height and width of an image,
as illustrated in Figure 1.
Argument:
X -- python numpy array of shape (m, n_H, n_W, n_C) representing a batch of m images
pad -- integer, amount of padding around each image on vertical and horizontal dimensions
Returns:
X_pad -- padded image of shape (m, n_H + 2*pad, n_W + 2*pad, n_C)
"""
### START CODE HERE ### (≈ 1 line)
X_pad = np.pad(X, ((0,0), (pad, pad), (pad,pad), (0, 0)), 'constant')
### END CODE HERE ###
return X_pad
np.random.seed(1)
x = np.random.randn(4, 3, 3, 2)
x_pad = zero_pad(x, 2)
print ("x.shape =", x.shape)
print ("x_pad.shape =", x_pad.shape)
print ("x[1,1] =", x[1,1])
print ("x_pad[1,1] =", x_pad[1,1])
fig, axarr = plt.subplots(1, 2)
axarr[0].set_title('x')
axarr[0].imshow(x[0,:,:,0])
axarr[1].set_title('x_pad')
axarr[1].imshow(x_pad[0,:,:,0])
```
**Expected Output**:
<table>
<tr>
<td>
**x.shape**:
</td>
<td>
(4, 3, 3, 2)
</td>
</tr>
<tr>
<td>
**x_pad.shape**:
</td>
<td>
(4, 7, 7, 2)
</td>
</tr>
<tr>
<td>
**x[1,1]**:
</td>
<td>
[[ 0.90085595 -0.68372786]
[-0.12289023 -0.93576943]
[-0.26788808 0.53035547]]
</td>
</tr>
<tr>
<td>
**x_pad[1,1]**:
</td>
<td>
[[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]
[ 0. 0.]]
</td>
</tr>
</table>
### 3.2 - Single step of convolution
In this part, implement a single step of convolution, in which you apply the filter to a single position of the input. This will be used to build a convolutional unit, which:
- Takes an input volume
- Applies a filter at every position of the input
- Outputs another volume (usually of different size)
<img src="images/Convolution_schematic.gif" style="width:500px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 2** </u><font color='purple'> : **Convolution operation**<br> with a filter of 2x2 and a stride of 1 (stride = amount you move the window each time you slide) </center></caption>
In a computer vision application, each value in the matrix on the left corresponds to a single pixel value, and we convolve a 3x3 filter with the image by multiplying its values element-wise with the original matrix, then summing them up and adding a bias. In this first step of the exercise, you will implement a single step of convolution, corresponding to applying a filter to just one of the positions to get a single real-valued output.
Later in this notebook, you'll apply this function to multiple positions of the input to implement the full convolutional operation.
**Exercise**: Implement conv_single_step(). [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.sum.html).
```
# GRADED FUNCTION: conv_single_step
def conv_single_step(a_slice_prev, W, b):
"""
Apply one filter defined by parameters W on a single slice (a_slice_prev) of the output activation
of the previous layer.
Arguments:
a_slice_prev -- slice of input data of shape (f, f, n_C_prev)
W -- Weight parameters contained in a window - matrix of shape (f, f, n_C_prev)
b -- Bias parameters contained in a window - matrix of shape (1, 1, 1)
Returns:
Z -- a scalar value, result of convolving the sliding window (W, b) on a slice x of the input data
"""
### START CODE HERE ### (≈ 2 lines of code)
# Element-wise product between a_slice and W. Do not add the bias yet.
s = a_slice_prev * W
# Sum over all entries of the volume s.
Z = np.sum(s)
# Add bias b to Z. Cast b to a float() so that Z results in a scalar value.
Z = float(b) + Z
### END CODE HERE ###
return Z
np.random.seed(1)
a_slice_prev = np.random.randn(4, 4, 3)
W = np.random.randn(4, 4, 3)
b = np.random.randn(1, 1, 1)
Z = conv_single_step(a_slice_prev, W, b)
print("Z =", Z)
```
**Expected Output**:
<table>
<tr>
<td>
**Z**
</td>
<td>
-6.99908945068
</td>
</tr>
</table>
### 3.3 - Convolutional Neural Networks - Forward pass
In the forward pass, you will take many filters and convolve them on the input. Each 'convolution' gives you a 2D matrix output. You will then stack these outputs to get a 3D volume:
<center>
<video width="620" height="440" src="images/conv_kiank.mp4" type="video/mp4" controls>
</video>
</center>
**Exercise**: Implement the function below to convolve the filters W on an input activation A_prev. This function takes as input A_prev, the activations output by the previous layer (for a batch of m inputs), F filters/weights denoted by W, and a bias vector denoted by b, where each filter has its own (single) bias. Finally you also have access to the hyperparameters dictionary which contains the stride and the padding.
**Hint**:
1. To select a 2x2 slice at the upper left corner of a matrix "a_prev" (shape (5,5,3)), you would do:
```python
a_slice_prev = a_prev[0:2,0:2,:]
```
This will be useful when you will define `a_slice_prev` below, using the `start/end` indexes you will define.
2. To define a_slice you will need to first define its corners `vert_start`, `vert_end`, `horiz_start` and `horiz_end`. This figure may be helpful for you to find how each of the corner can be defined using h, w, f and s in the code below.
<img src="images/vert_horiz_kiank.png" style="width:400px;height:300px;">
<caption><center> <u> <font color='purple'> **Figure 3** </u><font color='purple'> : **Definition of a slice using vertical and horizontal start/end (with a 2x2 filter)** <br> This figure shows only a single channel. </center></caption>
**Reminder**:
The formulas relating the output shape of the convolution to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f + 2 \times pad}{stride} \rfloor +1 $$
$$ n_C = \text{number of filters used in the convolution}$$
For this exercise, we won't worry about vectorization, and will just implement everything with for-loops.
```
# GRADED FUNCTION: conv_forward
def conv_forward(A_prev, W, b, hparameters):
"""
Implements the forward propagation for a convolution function
Arguments:
A_prev -- output activations of the previous layer, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
W -- Weights, numpy array of shape (f, f, n_C_prev, n_C)
b -- Biases, numpy array of shape (1, 1, 1, n_C)
hparameters -- python dictionary containing "stride" and "pad"
Returns:
Z -- conv output, numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward() function
"""
### START CODE HERE ###
# Retrieve dimensions from A_prev's shape (≈1 line)
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve dimensions from W's shape (≈1 line)
(f, f, n_C_prev, n_C) = W.shape
# Retrieve information from "hparameters" (≈2 lines)
stride = hparameters["stride"]
pad = hparameters["pad"]
# Compute the dimensions of the CONV output volume using the formula given above. Hint: use int() to floor. (≈2 lines)
n_H = int((n_H_prev - f + 2 * pad) / stride) + 1
n_W = int((n_W_prev - f + 2 * pad) / stride) + 1
# Initialize the output volume Z with zeros. (≈1 line)
Z = np.zeros((m, n_H, n_W, n_C))
# Create A_prev_pad by padding A_prev
A_prev_pad = zero_pad(A_prev, pad)
for i in range(m): # loop over the batch of training examples
a_prev_pad = A_prev_pad[i] # Select ith training example's padded activation
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over channels (= #filters) of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h * stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Use the corners to define the (3D) slice of a_prev_pad (See Hint above the cell). (≈1 line)
a_slice_prev = a_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :]
# Convolve the (3D) slice with the correct filter W and bias b, to get back one output neuron. (≈1 line)
Z[i, h, w, c] = conv_single_step(a_slice_prev, W[:, :, :, c], b[:, :, :, c])
### END CODE HERE ###
# Making sure your output shape is correct
assert(Z.shape == (m, n_H, n_W, n_C))
# Save information in "cache" for the backprop
cache = (A_prev, W, b, hparameters)
return Z, cache
np.random.seed(1)
A_prev = np.random.randn(10,4,4,3)
W = np.random.randn(2,2,3,8)
b = np.random.randn(1,1,1,8)
hparameters = {"pad" : 2,
"stride": 2}
Z, cache_conv = conv_forward(A_prev, W, b, hparameters)
print("Z's mean =", np.mean(Z))
print("Z[3,2,1] =", Z[3,2,1])
print("cache_conv[0][1][2][3] =", cache_conv[0][1][2][3])
```
**Expected Output**:
<table>
<tr>
<td>
**Z's mean**
</td>
<td>
0.0489952035289
</td>
</tr>
<tr>
<td>
**Z[3,2,1]**
</td>
<td>
[-0.61490741 -6.7439236 -2.55153897 1.75698377 3.56208902 0.53036437
5.18531798 8.75898442]
</td>
</tr>
<tr>
<td>
**cache_conv[0][1][2][3]**
</td>
<td>
[-0.20075807 0.18656139 0.41005165]
</td>
</tr>
</table>
Finally, CONV layer should also contain an activation, in which case we would add the following line of code:
```python
# Convolve the window to get back one output neuron
Z[i, h, w, c] = ...
# Apply activation
A[i, h, w, c] = activation(Z[i, h, w, c])
```
You don't need to do it here.
## 4 - Pooling layer
The pooling (POOL) layer reduces the height and width of the input. It helps reduce computation, as well as helps make feature detectors more invariant to its position in the input. The two types of pooling layers are:
- Max-pooling layer: slides an ($f, f$) window over the input and stores the max value of the window in the output.
- Average-pooling layer: slides an ($f, f$) window over the input and stores the average value of the window in the output.
<table>
<td>
<img src="images/max_pool1.png" style="width:500px;height:300px;">
<td>
<td>
<img src="images/a_pool.png" style="width:500px;height:300px;">
<td>
</table>
These pooling layers have no parameters for backpropagation to train. However, they have hyperparameters such as the window size $f$. This specifies the height and width of the fxf window you would compute a max or average over.
### 4.1 - Forward Pooling
Now, you are going to implement MAX-POOL and AVG-POOL, in the same function.
**Exercise**: Implement the forward pass of the pooling layer. Follow the hints in the comments below.
**Reminder**:
As there's no padding, the formulas binding the output shape of the pooling to the input shape is:
$$ n_H = \lfloor \frac{n_{H_{prev}} - f}{stride} \rfloor +1 $$
$$ n_W = \lfloor \frac{n_{W_{prev}} - f}{stride} \rfloor +1 $$
$$ n_C = n_{C_{prev}}$$
```
# GRADED FUNCTION: pool_forward
def pool_forward(A_prev, hparameters, mode = "max"):
"""
Implements the forward pass of the pooling layer
Arguments:
A_prev -- Input data, numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
hparameters -- python dictionary containing "f" and "stride"
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
A -- output of the pool layer, a numpy array of shape (m, n_H, n_W, n_C)
cache -- cache used in the backward pass of the pooling layer, contains the input and hparameters
"""
# Retrieve dimensions from the input shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve hyperparameters from "hparameters"
f = hparameters["f"]
stride = hparameters["stride"]
# Define the dimensions of the output
n_H = int(1 + (n_H_prev - f) / stride)
n_W = int(1 + (n_W_prev - f) / stride)
n_C = n_C_prev
# Initialize output matrix A
A = np.zeros((m, n_H, n_W, n_C))
### START CODE HERE ###
for i in range(m): # loop over the training examples
for h in range(n_H): # loop on the vertical axis of the output volume
for w in range(n_W): # loop on the horizontal axis of the output volume
for c in range (n_C): # loop over the channels of the output volume
# Find the corners of the current "slice" (≈4 lines)
vert_start = h * stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Use the corners to define the current slice on the ith training example of A_prev, channel c. (≈1 line)
a_prev_slice = A_prev[i, vert_start:vert_end, horiz_start:horiz_end, c]
# Compute the pooling operation on the slice. Use an if statment to differentiate the modes. Use np.max/np.mean.
if mode == "max":
A[i, h, w, c] = np.max(a_prev_slice)
elif mode == "average":
A[i, h, w, c] = np.mean(a_prev_slice)
### END CODE HERE ###
# Store the input and hparameters in "cache" for pool_backward()
cache = (A_prev, hparameters)
# Making sure your output shape is correct
assert(A.shape == (m, n_H, n_W, n_C))
return A, cache
np.random.seed(1)
A_prev = np.random.randn(2, 4, 4, 3)
hparameters = {"stride" : 2, "f": 3}
A, cache = pool_forward(A_prev, hparameters)
print("mode = max")
print("A =", A)
print()
A, cache = pool_forward(A_prev, hparameters, mode = "average")
print("mode = average")
print("A =", A)
```
**Expected Output:**
<table>
<tr>
<td>
A =
</td>
<td>
[[[[ 1.74481176 0.86540763 1.13376944]]]
[[[ 1.13162939 1.51981682 2.18557541]]]]
</td>
</tr>
<tr>
<td>
A =
</td>
<td>
[[[[ 0.02105773 -0.20328806 -0.40389855]]]
[[[-0.22154621 0.51716526 0.48155844]]]]
</td>
</tr>
</table>
Congratulations! You have now implemented the forward passes of all the layers of a convolutional network.
The remainer of this notebook is optional, and will not be graded.
## 5 - Backpropagation in convolutional neural networks (OPTIONAL / UNGRADED)
In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers don't need to bother with the details of the backward pass. The backward pass for convolutional networks is complicated. If you wish however, you can work through this optional portion of the notebook to get a sense of what backprop in a convolutional network looks like.
When in an earlier course you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in convolutional neural networks you can to calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are not trivial and we did not derive them in lecture, but we briefly presented them below.
### 5.1 - Convolutional layer backward pass
Let's start by implementing the backward pass for a CONV layer.
#### 5.1.1 - Computing dA:
This is the formula for computing $dA$ with respect to the cost for a certain filter $W_c$ and a given training example:
$$ dA += \sum _{h=0} ^{n_H} \sum_{w=0} ^{n_W} W_c \times dZ_{hw} \tag{1}$$
Where $W_c$ is a filter and $dZ_{hw}$ is a scalar corresponding to the gradient of the cost with respect to the output of the conv layer Z at the hth row and wth column (corresponding to the dot product taken at the ith stride left and jth stride down). Note that at each time, we multiply the the same filter $W_c$ by a different dZ when updating dA. We do so mainly because when computing the forward propagation, each filter is dotted and summed by a different a_slice. Therefore when computing the backprop for dA, we are just adding the gradients of all the a_slices.
In code, inside the appropriate for-loops, this formula translates into:
```python
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
```
#### 5.1.2 - Computing dW:
This is the formula for computing $dW_c$ ($dW_c$ is the derivative of one filter) with respect to the loss:
$$ dW_c += \sum _{h=0} ^{n_H} \sum_{w=0} ^ {n_W} a_{slice} \times dZ_{hw} \tag{2}$$
Where $a_{slice}$ corresponds to the slice which was used to generate the acitivation $Z_{ij}$. Hence, this ends up giving us the gradient for $W$ with respect to that slice. Since it is the same $W$, we will just add up all such gradients to get $dW$.
In code, inside the appropriate for-loops, this formula translates into:
```python
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
```
#### 5.1.3 - Computing db:
This is the formula for computing $db$ with respect to the cost for a certain filter $W_c$:
$$ db = \sum_h \sum_w dZ_{hw} \tag{3}$$
As you have previously seen in basic neural networks, db is computed by summing $dZ$. In this case, you are just summing over all the gradients of the conv output (Z) with respect to the cost.
In code, inside the appropriate for-loops, this formula translates into:
```python
db[:,:,:,c] += dZ[i, h, w, c]
```
**Exercise**: Implement the `conv_backward` function below. You should sum over all the training examples, filters, heights, and widths. You should then compute the derivatives using formulas 1, 2 and 3 above.
```
def conv_backward(dZ, cache):
"""
Implement the backward propagation for a convolution function
Arguments:
dZ -- gradient of the cost with respect to the output of the conv layer (Z), numpy array of shape (m, n_H, n_W, n_C)
cache -- cache of values needed for the conv_backward(), output of conv_forward()
Returns:
dA_prev -- gradient of the cost with respect to the input of the conv layer (A_prev),
numpy array of shape (m, n_H_prev, n_W_prev, n_C_prev)
dW -- gradient of the cost with respect to the weights of the conv layer (W)
numpy array of shape (f, f, n_C_prev, n_C)
db -- gradient of the cost with respect to the biases of the conv layer (b)
numpy array of shape (1, 1, 1, n_C)
"""
### START CODE HERE ###
# Retrieve information from "cache"
(A_prev, W, b, hparameters) = cache
# Retrieve dimensions from A_prev's shape
(m, n_H_prev, n_W_prev, n_C_prev) = A_prev.shape
# Retrieve dimensions from W's shape
(f, f, n_C_prev, n_C) = W.shape
# Retrieve information from "hparameters"
stride = hparameters["stride"]
pad = hparameters["pad"]
# Retrieve dimensions from dZ's shape
(m, n_H, n_W, n_C) = dZ.shape
# Initialize dA_prev, dW, db with the correct shapes
dA_prev = np.zeros((m, n_H_prev, n_W_prev, n_C_prev))
dW = np.zeros((f, f, n_C_prev, n_C))
db = np.zeros((1, 1, 1, n_C))
# Pad A_prev and dA_prev
A_prev_pad = zero_pad(A_prev, pad)
dA_prev_pad = zero_pad(dA_prev, pad)
for i in range(m): # loop over the training examples
# select ith training example from A_prev_pad and dA_prev_pad
a_prev_pad = A_prev_pad[i]
da_prev_pad = dA_prev_pad[i]
for h in range(n_H): # loop over vertical axis of the output volume
for w in range(n_W): # loop over horizontal axis of the output volume
for c in range(n_C): # loop over the channels of the output volume
# Find the corners of the current "slice"
vert_start = h * stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Use the corners to define the slice from a_prev_pad
a_slice = a_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :]
# Update gradients for the window and the filter's parameters using the code formulas given above
da_prev_pad[vert_start:vert_end, horiz_start:horiz_end, :] += W[:,:,:,c] * dZ[i, h, w, c]
dW[:,:,:,c] += a_slice * dZ[i, h, w, c]
db[:,:,:,c] += dZ[i, h, w, c]
# Set the ith training example's dA_prev to the unpaded da_prev_pad (Hint: use X[pad:-pad, pad:-pad, :])
dA_prev[i, :, :, :] = da_prev_pad[pad:-pad, pad:-pad, :]
### END CODE HERE ###
# Making sure your output shape is correct
assert(dA_prev.shape == (m, n_H_prev, n_W_prev, n_C_prev))
return dA_prev, dW, db
np.random.seed(1)
dA, dW, db = conv_backward(Z, cache_conv)
print("dA_mean =", np.mean(dA))
print("dW_mean =", np.mean(dW))
print("db_mean =", np.mean(db))
```
** Expected Output: **
<table>
<tr>
<td>
**dA_mean**
</td>
<td>
1.45243777754
</td>
</tr>
<tr>
<td>
**dW_mean**
</td>
<td>
1.72699145831
</td>
</tr>
<tr>
<td>
**db_mean**
</td>
<td>
7.83923256462
</td>
</tr>
</table>
## 5.2 Pooling layer - backward pass
Next, let's implement the backward pass for the pooling layer, starting with the MAX-POOL layer. Even though a pooling layer has no parameters for backprop to update, you still need to backpropagation the gradient through the pooling layer in order to compute gradients for layers that came before the pooling layer.
### 5.2.1 Max pooling - backward pass
Before jumping into the backpropagation of the pooling layer, you are going to build a helper function called `create_mask_from_window()` which does the following:
$$ X = \begin{bmatrix}
1 && 3 \\
4 && 2
\end{bmatrix} \quad \rightarrow \quad M =\begin{bmatrix}
0 && 0 \\
1 && 0
\end{bmatrix}\tag{4}$$
As you can see, this function creates a "mask" matrix which keeps track of where the maximum of the matrix is. True (1) indicates the position of the maximum in X, the other entries are False (0). You'll see later that the backward pass for average pooling will be similar to this but using a different mask.
**Exercise**: Implement `create_mask_from_window()`. This function will be helpful for pooling backward.
Hints:
- [np.max()]() may be helpful. It computes the maximum of an array.
- If you have a matrix X and a scalar x: `A = (X == x)` will return a matrix A of the same size as X such that:
```
A[i,j] = True if X[i,j] = x
A[i,j] = False if X[i,j] != x
```
- Here, you don't need to consider cases where there are several maxima in a matrix.
```
def create_mask_from_window(x):
"""
Creates a mask from an input matrix x, to identify the max entry of x.
Arguments:
x -- Array of shape (f, f)
Returns:
mask -- Array of the same shape as window, contains a True at the position corresponding to the max entry of x.
"""
### START CODE HERE ### (≈1 line)
mask = (x == np.max(x))
### END CODE HERE ###
return mask
np.random.seed(1)
x = np.random.randn(2,3)
mask = create_mask_from_window(x)
print('x = ', x)
print("mask = ", mask)
```
**Expected Output:**
<table>
<tr>
<td>
**x =**
</td>
<td>
[[ 1.62434536 -0.61175641 -0.52817175] <br>
[-1.07296862 0.86540763 -2.3015387 ]]
</td>
</tr>
<tr>
<td>
**mask =**
</td>
<td>
[[ True False False] <br>
[False False False]]
</td>
</tr>
</table>
Why do we keep track of the position of the max? It's because this is the input value that ultimately influenced the output, and therefore the cost. Backprop is computing gradients with respect to the cost, so anything that influences the ultimate cost should have a non-zero gradient. So, backprop will "propagate" the gradient back to this particular input value that had influenced the cost.
### 5.2.2 - Average pooling - backward pass
In max pooling, for each input window, all the "influence" on the output came from a single input value--the max. In average pooling, every element of the input window has equal influence on the output. So to implement backprop, you will now implement a helper function that reflects this.
For example if we did average pooling in the forward pass using a 2x2 filter, then the mask you'll use for the backward pass will look like:
$$ dZ = 1 \quad \rightarrow \quad dZ =\begin{bmatrix}
1/4 && 1/4 \\
1/4 && 1/4
\end{bmatrix}\tag{5}$$
This implies that each position in the $dZ$ matrix contributes equally to output because in the forward pass, we took an average.
**Exercise**: Implement the function below to equally distribute a value dz through a matrix of dimension shape. [Hint](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ones.html)
```
def distribute_value(dz, shape):
"""
Distributes the input value in the matrix of dimension shape
Arguments:
dz -- input scalar
shape -- the shape (n_H, n_W) of the output matrix for which we want to distribute the value of dz
Returns:
a -- Array of size (n_H, n_W) for which we distributed the value of dz
"""
### START CODE HERE ###
# Retrieve dimensions from shape (≈1 line)
(n_H, n_W) = shape
# Compute the value to distribute on the matrix (≈1 line)
average = dz / (n_H * n_W)
# Create a matrix where every entry is the "average" value (≈1 line)
a = np.ones((n_H, n_W)) * average
### END CODE HERE ###
return a
a = distribute_value(2, (2,2))
print('distributed value =', a)
```
**Expected Output**:
<table>
<tr>
<td>
distributed_value =
</td>
<td>
[[ 0.5 0.5]
<br\>
[ 0.5 0.5]]
</td>
</tr>
</table>
### 5.2.3 Putting it together: Pooling backward
You now have everything you need to compute backward propagation on a pooling layer.
**Exercise**: Implement the `pool_backward` function in both modes (`"max"` and `"average"`). You will once again use 4 for-loops (iterating over training examples, height, width, and channels). You should use an `if/elif` statement to see if the mode is equal to `'max'` or `'average'`. If it is equal to 'average' you should use the `distribute_value()` function you implemented above to create a matrix of the same shape as `a_slice`. Otherwise, the mode is equal to '`max`', and you will create a mask with `create_mask_from_window()` and multiply it by the corresponding value of dZ.
```
def pool_backward(dA, cache, mode = "max"):
"""
Implements the backward pass of the pooling layer
Arguments:
dA -- gradient of cost with respect to the output of the pooling layer, same shape as A
cache -- cache output from the forward pass of the pooling layer, contains the layer's input and hparameters
mode -- the pooling mode you would like to use, defined as a string ("max" or "average")
Returns:
dA_prev -- gradient of cost with respect to the input of the pooling layer, same shape as A_prev
"""
### START CODE HERE ###
# Retrieve information from cache (≈1 line)
(A_prev, hparameters) = cache
# Retrieve hyperparameters from "hparameters" (≈2 lines)
stride = hparameters["stride"]
f = hparameters["f"]
# Retrieve dimensions from A_prev's shape and dA's shape (≈2 lines)
m, n_H_prev, n_W_prev, n_C_prev = A_prev.shape
m, n_H, n_W, n_C = dA.shape
# Initialize dA_prev with zeros (≈1 line)
dA_prev = np.zeros((m, n_H_prev, n_W_prev, n_C_prev))
for i in range(m): # loop over the training examples
# select training example from A_prev (≈1 line)
a_prev = A_prev[i]
for h in range(n_H): # loop on the vertical axis
for w in range(n_W): # loop on the horizontal axis
for c in range(n_C): # loop over the channels (depth)
# Find the corners of the current "slice" (≈4 lines)
vert_start = h * stride
vert_end = vert_start + f
horiz_start = w * stride
horiz_end = horiz_start + f
# Compute the backward propagation in both modes.
if mode == "max":
# Use the corners and "c" to define the current slice from a_prev (≈1 line)
a_prev_slice = a_prev[vert_start:vert_end, horiz_start:horiz_end, c]
# Create the mask from a_prev_slice (≈1 line)
mask = create_mask_from_window(a_prev_slice)
# Set dA_prev to be dA_prev + (the mask multiplied by the correct entry of dA) (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += dA[i, h, w, c] * mask
elif mode == "average":
# Get the value a from dA (≈1 line)
da = dA[i, h, w, c]
# Define the shape of the filter as fxf (≈1 line)
shape = (f, f)
# Distribute it to get the correct slice of dA_prev. i.e. Add the distributed value of da. (≈1 line)
dA_prev[i, vert_start: vert_end, horiz_start: horiz_end, c] += distribute_value(da, shape)
### END CODE ###
# Making sure your output shape is correct
assert(dA_prev.shape == A_prev.shape)
return dA_prev
np.random.seed(1)
A_prev = np.random.randn(5, 5, 3, 2)
hparameters = {"stride" : 1, "f": 2}
A, cache = pool_forward(A_prev, hparameters)
dA = np.random.randn(5, 4, 2, 2)
dA_prev = pool_backward(dA, cache, mode = "max")
print("mode = max")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
print()
dA_prev = pool_backward(dA, cache, mode = "average")
print("mode = average")
print('mean of dA = ', np.mean(dA))
print('dA_prev[1,1] = ', dA_prev[1,1])
```
**Expected Output**:
mode = max:
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0. 0. ] <br>
[ 5.05844394 -1.68282702] <br>
[ 0. 0. ]]
</td>
</tr>
</table>
mode = average
<table>
<tr>
<td>
**mean of dA =**
</td>
<td>
0.145713902729
</td>
</tr>
<tr>
<td>
**dA_prev[1,1] =**
</td>
<td>
[[ 0.08485462 0.2787552 ] <br>
[ 1.26461098 -0.25749373] <br>
[ 1.17975636 -0.53624893]]
</td>
</tr>
</table>
### Congratulations !
Congratulation on completing this assignment. You now understand how convolutional neural networks work. You have implemented all the building blocks of a neural network. In the next assignment you will implement a ConvNet using TensorFlow.
| github_jupyter |
# Code for the tasks for the final meeting
```
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
%matplotlib inline
```
## Task 1 - Run new controls
* This accession list was generated in a rush therefore we need to filter out based on meta data anything which does not fit as a control
* Note that this is NOT a cell/tissue matched control set. There are multiple tissue types e.g lung, blood. This limits direct comparison. Really just to see if we can see any alpha dup expression in ANY tissue
```
big_control_df = pd.read_csv('../data_files/mbase_normed/big_control.meta.mbase_norm.csv', index_col=0)
big_control_df.shape
def has_dup(df):
dup_value = df['norm_alpha_dup_read_covers_snps_count_exact']
if dup_value == 0:
return False
else:
return True
big_control_df['has_dup'] = big_control_df.apply(has_dup, axis=1)
print(big_control_df['source_name'].value_counts())
#Only pick sources which look like they could be from healthy tissue
big_control_df_filtered = big_control_df[
(big_control_df['source_name'] =='T cells') |
(big_control_df['source_name'] =='Human Brain Reference RNA (HBRR) from Ambion') |
(big_control_df['source_name'] =='Universal Human Reference RNA (UHRR) from Stratagene') |
(big_control_df['source_name'] =='CD4 T cell') |
(big_control_df['source_name'] =='Blood') |
(big_control_df['source_name'] =='Monocytes') |
(big_control_df['source_name'] =='Human dermal fibroblasts') |
(big_control_df['source_name'] =='Blood, control') |
(big_control_df['source_name'] =='brain') |
(big_control_df['source_name'] =='Primary foreskin fibroblast cells') |
(big_control_df['source_name'] =='Immortalised Human Myoblasts') |
(big_control_df['source_name'] =='Human peripheral blood leukocytes') |
(big_control_df['source_name'] =='subcutaneous adipose tissue') |
(big_control_df['source_name'] =='CD4+ T cells') |
(big_control_df['source_name'] =='Peripheral blood mononuclear cells from 18ml venous whole blood') |
(big_control_df['source_name'] =='airway smooth muscle cells') |
(big_control_df['source_name'] =='lung tissue') |
(big_control_df['source_name'] =='Nasopharyngeal') |
(big_control_df['source_name'] =='large airway basal cells ') |
(big_control_df['source_name'] =='Endometrium') |
(big_control_df['source_name'] =='PBMC') |
(big_control_df['source_name'] =='Bone Marrow Aspirate') |
(big_control_df['source_name'] =='Whole Blood') |
(big_control_df['source_name'] =='Fibroblast') |
(big_control_df['source_name'] =='Human skin') |
(big_control_df['source_name'] =='Normal Skin') |
(big_control_df['source_name'] =='human fibroblasts') |
(big_control_df['source_name'] =='Mammary epithelial cells') |
(big_control_df['source_name'] =='Whole blood') |
(big_control_df['source_name'] =='PBMCs') |
(big_control_df['source_name'] =='Whole blood cells (PAX)') |
(big_control_df['source_name'] =='Skin') |
(big_control_df['source_name'] =='Plain, blood') |
(big_control_df['source_name'] =='Human peripheral blood') |
(big_control_df['source_name'] =='large airway epithelial cells') |
(big_control_df['source_name'] =='Purified human monocytes') |
(big_control_df['source_name'] =='human bronchial epithelial cells') |
(big_control_df['source_name'] =='normal liver') |
(big_control_df['source_name'] =='Lung') |
(big_control_df['source_name'] =='control_brain') |
(big_control_df['source_name'] =='Control') |
(big_control_df['source_name'] =='normal control') |
(big_control_df['source_name'] =='Peripheral whole blood draw, healthy control ') |
(big_control_df['source_name'] =='Heart') |
(big_control_df['source_name'] =='normal') |
(big_control_df['source_name'] =='esophageal tissue') |
(big_control_df['source_name'] =='Human pancreatic islets') |
(big_control_df['source_name'] =='Liver') |
(big_control_df['source_name'] =='control_heart') |
(big_control_df['source_name'] =='airway epithelial cells') |
(big_control_df['source_name'] =='Lung cells') |
(big_control_df['source_name'] =='Intestine ') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells') |
(big_control_df['source_name'] =='Peripheral Blood Mononuclear Cells')
]
big_control_df_filtered['cell_type'].value_counts()
# Now filter further to get rid of cancer cells
big_control_df_filtered = big_control_df_filtered[
(big_control_df_filtered['cell_type'] != 'alveolar adenocarcinoma cells') &
(big_control_df_filtered['cell_type'] != 'Non-small-cell lung cancer (NSCLC) cells') &
(big_control_df_filtered['cell_type'] != 'breast cancer') &
(big_control_df_filtered['cell_type'] != 'CL1-5') &
(big_control_df_filtered['cell_type'] != 'CL1-0')
]
#big_control_df_filtered.to_csv('big_control.meta.mbase_norm.filter.csv')
```
* Should now be confident we only have healthy(ish) tissue types
```
big_control_df_filtered.columns
# How many have have each of these?
run_count = big_control_df_filtered.shape[0]
any_expression = big_control_df_filtered[big_control_df_filtered['norm_alignment_count'] !=0].shape[0]
print ("Of the {run_count} runs in this dataframe {any_expression} have some hits in this locus" \
". e.g alignment over any area of the genes".format(
run_count = run_count,
any_expression = any_expression
))
wt_count = big_control_df_filtered[big_control_df_filtered['norm_alpha_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the alpha_wt snps".format(count = wt_count))
dup_count = big_control_df_filtered[big_control_df_filtered['norm_alpha_dup_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the alpha_dup snps.".format(count = dup_count))
beta_count = big_control_df_filtered[big_control_df_filtered['norm_beta_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the beta snps.".format(count = beta_count))
any_expression = big_control_df_filtered[(big_control_df_filtered['norm_alpha_read_covers_snps_count_exact'] != 0) |
(big_control_df_filtered['norm_alpha_dup_read_covers_snps_count_exact'] != 0) |
(big_control_df_filtered['norm_beta_read_covers_snps_count_exact'] != 0)
].shape[0]
print ("Of the {any_expression} runs that show expression across the snps {count} % have alpha dup expression".format(
any_expression=any_expression,
count = round((dup_count/any_expression)*100,2)
))
# How many have have each of these?
# Should be looking at a per sample basis - does not make big difference?
pivoted_big_control = big_control_df_filtered.pivot_table(index='BioSample')
run_count = pivoted_big_control.shape[0]
any_expression = pivoted_big_control[pivoted_big_control['norm_alignment_count'] !=0].shape[0]
print ("Of the {run_count} samples in this dataframe {any_expression} have some hits in this locus" \
". e.g alignment over any area of the genes".format(
run_count = run_count,
any_expression = any_expression
))
wt_count = pivoted_big_control[pivoted_big_control['norm_alpha_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the alpha_wt snps".format(count = wt_count))
dup_count = pivoted_big_control[pivoted_big_control['norm_alpha_dup_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the alpha_dup snps.".format(count = dup_count))
beta_count = pivoted_big_control[pivoted_big_control['norm_beta_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the beta snps.".format(count = beta_count))
any_expression = pivoted_big_control[(pivoted_big_control['norm_alpha_read_covers_snps_count_exact'] != 0) |
(pivoted_big_control['norm_alpha_dup_read_covers_snps_count_exact'] != 0) |
(pivoted_big_control['norm_beta_read_covers_snps_count_exact'] != 0)
].shape[0]
print ("Of the {any_expression} samples that show expression across the snps {count} % have alpha dup expression".format(
any_expression=any_expression,
count = round((dup_count/any_expression)*100,2)
))
```
* So unlike in the geuvadis controls we do have some limited expression
```
# Which runs have the most expression?
top_25 = big_control_df_filtered.sort_values('norm_alpha_dup_read_covers_snps_count_exact',ascending=False)[
['BioProject',
'BioSample',
'cell_type',
'source_name',
'tissue',
'norm_alpha_read_covers_snps_count_exact',
'norm_alpha_dup_read_covers_snps_count_exact',
'norm_beta_read_covers_snps_count_exact',
'norm_alignment_count',
'MBases']].head(100)
# plot shows the top 50 expressing dup samples
plot = sns.pairplot(x_vars=["norm_alpha_read_covers_snps_count_exact"], y_vars=["norm_alpha_dup_read_covers_snps_count_exact"], data=top_25, hue="source_name", size=10)
axes = plot.axes
axes[0][0].set_xlabel('Alpha WT Expression Reads/Mbase')
axes[0][0].set_ylabel('Alpha DUP Expression Reads/Mbase')
axes[0][0].set_title('Alpha WT vs Alpha DUP Expression in Control Set ')
big_control_df_filtered.head()
big_control_df_filtered.groupby('has_dup').mean()
```
### Comment
* The runs which have detectable alpha dup expression seem to have much higher expression of all loci. Could check against control gene?
## Task 2 - New Types
* Run a few more disease types through the pipeline
```
aml_df = pd.read_csv('../data_files/mbase_normed/aml.meta.mbase_norm.csv', index_col=0)
cml_df = pd.read_csv('../data_files/mbase_normed/cml.meta.mbase_norm.csv', index_col=0)
dengue_df = pd.read_csv('../data_files/mbase_normed/dengue.meta.mbase_norm.csv', index_col=0)
sepsis_df = pd.read_csv('../data_files/mbase_normed/sepsis.meta.mbase_norm.csv', index_col=0)
big_control_df = pd.read_csv('../data_files/mbase_normed/big_control.meta.mbase_norm.filter.csv', index_col=0)
geu_control_df = pd.read_csv('../data_files/mbase_normed/geu.meta.mbase_norm.csv', index_col=0)
master_df = aml_df.append([cml_df,dengue_df,sepsis_df, big_control_df, geu_control_df])
def clean_labels(df):
label = df['source']
mappings ={
'aml': 'AML',
'cml': 'CML',
'dengue': 'Dengue',
'sepsis': 'Sepsis',
'big_control': 'Control1',
'geu': 'Control2 (Geuvadis)',
}
return mappings[label]
master_df['source'] = master_df.apply(clean_labels, axis=1)
#How many of each type do we have?
fig, ax = plt.subplots(figsize=(12,8))
x_labels = ['AML', 'CML', 'Dengue', 'Sepsis', 'Control1', 'Control2', ]
sns.countplot(x='source', data=master_df)
ax.set_title('Count of the Number of runs per analysis group')
#What are the expression levels of each type?
fig, ax = plt.subplots(3,1,figsize=(12,8))
sns.barplot(x='source', y='norm_alpha_read_covers_snps_count_exact', data=master_df, ax=ax[0])
sns.barplot(x='source', y='norm_alpha_dup_read_covers_snps_count_exact', data=master_df, ax=ax[1])
sns.barplot(x='source', y='norm_beta_read_covers_snps_count_exact', data=master_df, ax=ax[2])
ax[0].set_ylabel('Alpha WT (RPMb)')
ax[1].set_ylabel('Alpha DUP (RPMb)')
ax[2].set_ylabel('Beta (RPMb)')
plt.tight_layout()
# Replace with code below if you want on a per sample basis rather than per run
#master_pivot = master_df.pivot_table(index='BioSample')
plot = sns.pairplot(x_vars=["norm_alpha_read_covers_snps_count_exact"], y_vars=["norm_alpha_dup_read_covers_snps_count_exact"], data=master_df, hue='source', size=10)
axes = plot.axes
axes[0][0].set_xlabel('Alpha WT Expression Reads/Mbase')
axes[0][0].set_ylabel('Alpha DUP Expression Reads/Mbase')
axes[0][0].set_title('Alpha WT vs Alpha DUP Expression ')
# How many have have each of these - just look at AML - compare with control results above.
# AML results may be inflated as there could eb the same cell lines multiple times so if this cell line has the dup?
run_count = aml_df.shape[0]
any_expression = aml_df[aml_df['norm_alignment_count'] !=0].shape[0]
print ("Of the {run_count} runs in this dataframe {any_expression} have some hits in this locus" \
". e.g alignment over any area of the genes".format(
run_count = run_count,
any_expression = any_expression
))
wt_count = aml_df[aml_df['norm_alpha_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the alpha_wt snps".format(count = wt_count))
dup_count = aml_df[aml_df['norm_alpha_dup_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the alpha_dup snps.".format(count = dup_count))
beta_count = aml_df[aml_df['norm_beta_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} runs with reads matching the beta snps.".format(count = beta_count))
any_expression = aml_df[(aml_df['norm_alpha_read_covers_snps_count_exact'] != 0) |
(aml_df['norm_alpha_dup_read_covers_snps_count_exact'] != 0) |
(aml_df['norm_beta_read_covers_snps_count_exact'] != 0)
].shape[0]
print ("Of the {any_expression} runs that show expression across the snps {count} % have alpha dup expression".format(
any_expression=any_expression,
count = round((dup_count/any_expression)*100,2)
))
# How many have have each of these - just look at AML - compare with control results above.
# Per sample basis
# AML results may be inflated as there could eb the same cell lines multiple times so if this cell line has the dup?
pivoted_aml = aml_df.pivot_table(index='BioSample')
run_count = pivoted_aml.shape[0]
any_expression = pivoted_aml[pivoted_aml['norm_alignment_count'] !=0].shape[0]
print ("Of the {run_count} samples in this dataframe {any_expression} have some hits in this locus" \
". e.g alignment over any area of the genes".format(
run_count = run_count,
any_expression = any_expression
))
wt_count = pivoted_aml[pivoted_aml['norm_alpha_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the alpha_wt snps".format(count = wt_count))
dup_count = pivoted_aml[pivoted_aml['norm_alpha_dup_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the alpha_dup snps.".format(count = dup_count))
beta_count = pivoted_aml[pivoted_aml['norm_beta_read_covers_snps_count_exact'] != 0].shape[0]
print ("There are {count} samples with reads matching the beta snps.".format(count = beta_count))
any_expression = pivoted_aml[(pivoted_aml['norm_alpha_read_covers_snps_count_exact'] != 0) |
(pivoted_aml['norm_alpha_dup_read_covers_snps_count_exact'] != 0) |
(pivoted_aml['norm_beta_read_covers_snps_count_exact'] != 0)
].shape[0]
print ("Of the {any_expression} samples that show expression across the snps {count} % have alpha dup expression".format(
any_expression=any_expression,
count = round((dup_count/any_expression)*100,2)
))
```
## Task 3 - Alpha wt / alpha dup ratio
* Look at the Tryptase alpha WT / Tryptase alpha DUP ratio and then look at meta data
```
def dup_wt_ratio(df):
wt = df['norm_alpha_read_covers_snps_count_exact']
dup = df['norm_alpha_dup_read_covers_snps_count_exact']
try:
return dup /wt
except:
return np.nan
master_df['dup_wt_ratio'] =master_df.apply(dup_wt_ratio, axis=1)
# Of the samples which have some alpha dup expression
fig, ax = plt.subplots(figsize=(8,5))
master_df[master_df['norm_alpha_dup_read_covers_snps_count_exact']!=0]['dup_wt_ratio'].hist(bins=50)
ax.set_title('Histogram of the observed dup/wt ratio')
ax.set_xlabel('Alpha Dup / Alpha Ratio')
ax.set_ylabel('Count')
master_df[master_df['norm_alpha_dup_read_covers_snps_count_exact']!=0].sort_values('dup_wt_ratio', ascending=False)[
['source_name',
'cell_type',
'tissue',
'BioProject',
'BioSample',
'dup_wt_ratio',
'norm_alpha_read_covers_snps_count_exact',
'norm_alpha_dup_read_covers_snps_count_exact',
'norm_beta_read_covers_snps_count_exact'
]
].head(10)
master_df[master_df['BioProject'] =='PRJNA386992'][['source_name',
'alignment_count',
'MBases',
'cell_type',
'tissue',
'BioProject',
'BioSample',
'dup_wt_ratio',
'norm_alpha_read_covers_snps_count_exact',
'norm_alpha_dup_read_covers_snps_count_exact',
'norm_beta_read_covers_snps_count_exact'
] ].sort_values('dup_wt_ratio', ascending=False)
```
* Note that the top two runs are from the same study PRJNA386992. This is a study looking at double minutes.
* See https://academic.oup.com/nar/article/42/14/9131/1273016
* Bioproject: https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA386992
* Quick look at the karyotype of those two samples shows complex stuff going on at chr16
```
print ('The ratio in this bioproject is: ', round(master_df[master_df['BioProject'] =='PRJNA386992' ]['dup_wt_ratio'].mean(),2))
print ('Compared to an average of: ', round(master_df['dup_wt_ratio'].mean(),2))
# Top bioprojects
master_df.pivot_table(index='BioProject').sort_values('dup_wt_ratio', ascending=False).head()
master_df['has_dup'] = master_df.apply(has_dup, axis=1)
# Compare this with the big_control group above. Also you can see that the expression level is generally much higher in \
# samples with the duplicaton. Is it just that we can only see the duplication when expression levels are very high?
master_df.groupby('has_dup').mean()
```
### What is different about AML samples that express alpha dup?
```
aml_df.head()
express_dup = aml_df[aml_df['norm_alpha_dup_read_covers_snps_count_exact'] >0.25]
no_dup = aml_df[aml_df['norm_alpha_dup_read_covers_snps_count_exact'] <0.25]
express_dup['BioSample'].value_counts()
no_dup['BioSample'].value_counts()
# What cell types do we have?
aml_df['cell_type'].value_counts()
```
## Task 5 - Retro Virus Analysis
* From a previous Hackathon we have some data showing Human Endogenous Retrovirus (HER) expression for (some ~1500) of the AML dataset.
* A longshot but can we see any correlation between HER expression and the tryptase locus?
* Note: results show no obvious link
```
virus_df = pd.read_csv('../data_files/alldata.normalised.zeroes.csv', index_col=0)
virus_df = virus_df[virus_df['type'] =='aml']
grouped_by_run = virus_df.groupby('acc').sum()['count']
aml_virus_df = aml_df.join(grouped_by_run)
aml_virus_df['normalised_virus_count'] = aml_virus_df['count'] / aml_virus_df['MBases']
aml_virus_df = aml_virus_df[aml_virus_df['normalised_virus_count'].isnull() == False]
aml_virus_df.pivot_table(index='BioSample').sort_values('normalised_virus_count', ascending=False).head()
aml_virus_df['dup_wt_ratio'] =aml_virus_df.apply(wt_dup_ratio, axis=1)
aml_virus_df[aml_virus_df['norm_alpha_dup_read_covers_snps_count_exact']!=0][['normalised_virus_count', 'dup_wt_ratio']].corr()
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(aml_virus_df.corr(), ax=ax)
```
| github_jupyter |
This is an example showing how scikit-learn can be used to classify documents by topics using a bag-of-words approach. This example uses a scipy.sparse matrix to store the features and demonstrates various classifiers that can efficiently handle sparse matrices.
The dataset used in this example is the 20 newsgroups dataset. It will be automatically downloaded, then cached.
The bar plot indicates the accuracy, training time (normalized) and test time (normalized) of each classifier.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
```
import plotly.plotly as py
import plotly.graph_objs as go
from __future__ import print_function
import logging
import numpy as np
from optparse import OptionParser
import sys
from time import time
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.feature_selection import SelectKBest, chi2
from sklearn.linear_model import RidgeClassifier
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import Perceptron
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.naive_bayes import BernoulliNB, MultinomialNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neighbors import NearestCentroid
from sklearn.ensemble import RandomForestClassifier
from sklearn.utils.extmath import density
from sklearn import metrics
```
### Calculations
Display progress logs on stdout
```
logging.basicConfig(level=logging.INFO,
format='%(asctime)s %(levelname)s %(message)s')
```
Parse commandline arguments
```
op = OptionParser()
op.add_option("--report",
action="store_true", dest="print_report",
help="Print a detailed classification report.")
op.add_option("--chi2_select",
action="store", type="int", dest="select_chi2",
help="Select some number of features using a chi-squared test")
op.add_option("--confusion_matrix",
action="store_true", dest="print_cm",
help="Print the confusion matrix.")
op.add_option("--top10",
action="store_true", dest="print_top10",
help="Print ten most discriminative terms per class"
" for every classifier.")
op.add_option("--all_categories",
action="store_true", dest="all_categories",
help="Whether to use all categories or not.")
op.add_option("--use_hashing",
action="store_true",
help="Use a hashing vectorizer.")
op.add_option("--n_features",
action="store", type=int, default=2 ** 16,
help="n_features when using the hashing vectorizer.")
op.add_option("--filtered",
action="store_true",
help="Remove newsgroup information that is easily overfit: "
"headers, signatures, and quoting.")
op.print_help()
```
To Get command line arguments add
(opts, args) = op.parse_args()
and set the following as:
all_categories = opts.all_categories
filtered = opts.filtered
use_hashing = opts.use_hashing
n_features = opts.n_features
select_chi2 = opts.select_chi2
print_cm = opts.print_cm
print_top10 = opts.print_top10
print_report = opts.print_report
For this tutorial we are taking these values as:
```
all_categories = True
filtered = True
use_hashing = True
n_features = 2 ** 16
select_chi2 = 10
print_cm = True
print_top10 = True
print_report = True
```
Load some categories from the training set
```
if all_categories:
categories = None
else:
categories = [
'alt.atheism',
'talk.religion.misc',
'comp.graphics',
'sci.space',
]
if filtered:
remove = ('headers', 'footers', 'quotes')
else:
remove = ()
print("Loading 20 newsgroups dataset for categories:")
print(categories if categories else "all")
data_train = fetch_20newsgroups(subset='train', categories=categories,
shuffle=True, random_state=42,
remove=remove)
data_test = fetch_20newsgroups(subset='test', categories=categories,
shuffle=True, random_state=42,
remove=remove)
print('data loaded')
```
Order of labels in `target_names` can be different from `categories`
```
target_names = data_train.target_names
def size_mb(docs):
return sum(len(s.encode('utf-8')) for s in docs) / 1e6
data_train_size_mb = size_mb(data_train.data)
data_test_size_mb = size_mb(data_test.data)
print("%d documents - %0.3fMB (training set)" % (
len(data_train.data), data_train_size_mb))
print("%d documents - %0.3fMB (test set)" % (
len(data_test.data), data_test_size_mb))
```
Split a training set and a test set
```
y_train, y_test = data_train.target, data_test.target
print("Extracting features from the training data using a sparse vectorizer")
t0 = time()
if use_hashing:
vectorizer = HashingVectorizer(stop_words='english', non_negative=True,
n_features=n_features)
X_train = vectorizer.transform(data_train.data)
else:
vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,
stop_words='english')
X_train = vectorizer.fit_transform(data_train.data)
duration = time() - t0
print("done in %fs at %0.3fMB/s" % (duration, data_train_size_mb / duration))
print("n_samples: %d, n_features: %d" % X_train.shape)
print()
print("Extracting features from the test data using the same vectorizer")
t0 = time()
X_test = vectorizer.transform(data_test.data)
duration = time() - t0
print("done in %fs at %0.3fMB/s" % (duration, data_test_size_mb / duration))
print("n_samples: %d, n_features: %d" % X_test.shape)
print()
```
Mapping from integer feature name to original token string
```
if use_hashing:
feature_names = None
else:
feature_names = vectorizer.get_feature_names()
if select_chi2:
print("Extracting %d best features by a chi-squared test" %
select_chi2)
t0 = time()
ch2 = SelectKBest(chi2, k=select_chi2)
X_train = ch2.fit_transform(X_train, y_train)
X_test = ch2.transform(X_test)
if feature_names:
# keep selected feature names
feature_names = [feature_names[i] for i
in ch2.get_support(indices=True)]
print("done in %fs" % (time() - t0))
print()
if feature_names:
feature_names = np.asarray(feature_names)
def trim(s):
"""Trim string to fit on terminal (assuming 80-column display)"""
return s if len(s) <= 80 else s[:77] + "..."
```
Benchmark classifiers
```
def benchmark(clf):
print('_' * 80)
print("Training: ")
print(clf)
t0 = time()
clf.fit(X_train, y_train)
train_time = time() - t0
print("train time: %0.3fs" % train_time)
t0 = time()
pred = clf.predict(X_test)
test_time = time() - t0
print("test time: %0.3fs" % test_time)
score = metrics.accuracy_score(y_test, pred)
print("accuracy: %0.3f" % score)
if hasattr(clf, 'coef_'):
print("dimensionality: %d" % clf.coef_.shape[1])
print("density: %f" % density(clf.coef_))
if print_top10 and feature_names is not None:
print("top 10 keywords per class:")
for i, label in enumerate(target_names):
top10 = np.argsort(clf.coef_[i])[-10:]
print(trim("%s: %s" % (label, " ".join(feature_names[top10]))))
print()
if print_report:
print("classification report:")
print(metrics.classification_report(y_test, pred,
target_names=target_names))
if print_cm:
print("confusion matrix:")
print(metrics.confusion_matrix(y_test, pred))
print()
clf_descr = str(clf).split('(')[0]
return clf_descr, score, train_time, test_time
results = []
for clf, name in (
(RidgeClassifier(tol=1e-2, solver="lsqr"), "Ridge Classifier"),
(Perceptron(n_iter=50), "Perceptron"),
(PassiveAggressiveClassifier(n_iter=50), "Passive Aggressive"),
(KNeighborsClassifier(n_neighbors=10), "kNN"),
(RandomForestClassifier(n_estimators=100), "Random forest")):
print('=' * 80)
print(name)
results.append(benchmark(clf))
for penalty in ["l2", "l1"]:
print('=' * 80)
print("%s penalty" % penalty.upper())
# Train Liblinear model
results.append(benchmark(LinearSVC(loss='l2', penalty=penalty,
dual=False, tol=1e-3)))
# Train SGD model
results.append(benchmark(SGDClassifier(alpha=.0001, n_iter=50,
penalty=penalty)))
```
Train SGD with Elastic Net penalty
```
print('=' * 80)
print("Elastic-Net penalty")
results.append(benchmark(SGDClassifier(alpha=.0001, n_iter=50,
penalty="elasticnet")))
```
Train NearestCentroid without threshold
```
print('=' * 80)
print("NearestCentroid (aka Rocchio classifier)")
results.append(benchmark(NearestCentroid()))
```
Train sparse Naive Bayes classifiers
```
print('=' * 80)
print("Naive Bayes")
results.append(benchmark(MultinomialNB(alpha=.01)))
results.append(benchmark(BernoulliNB(alpha=.01)))
```
LinearSVC with L1-based feature selection
```
print('=' * 80)
print("LinearSVC with L1-based feature selection")
# The smaller C, the stronger the regularization.
# The more regularization, the more sparsity.
results.append(benchmark(Pipeline([
('feature_selection', LinearSVC(penalty="l1", dual=False, tol=1e-3)),
('classification', LinearSVC())
])))
```
### Plot Results
```
indices = np.arange(len(results))
results = [[x[i] for x in results] for i in range(4)]
clf_names, score, training_time, test_time = results
training_time = np.array(training_time) / np.max(training_time)
test_time = np.array(test_time) / np.max(test_time)
p1 = go.Bar(x=indices, y=score,
name="score",
marker=dict(color='navy'))
p2 = go.Bar(x=indices + 2, y=training_time,
name="training time",
marker=dict(color='cyan'))
p3 = go.Bar(x=indices + 4, y=test_time,
name="test time",
marker=dict(color='darkorange'))
layout = go.Layout(title="Score")
fig = go.Figure(data=[p1, p2, p3], layout=layout)
py.iplot(fig)
```
### License
Authors:
Peter Prettenhofer <peter.prettenhofer@gmail.com>
Olivier Grisel <olivier.grisel@ensta.org>
Mathieu Blondel <mathieu@mblondel.org>
Lars Buitinck
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Classification of Text Documents using Sparse Features.ipynb', 'scikit-learn/document-classification-20newsgroups/', 'Classification of Text Documents using Sparse Features | plotly',
' ',
title = 'Classification of Text Documents using Sparse Features | plotly',
name = 'Classification of Text Documents using Sparse Features',
has_thumbnail='false', thumbnail='thumbnail/your-tutorial-chart.jpg',
language='scikit-learn', page_type='example_index',
display_as='text_documents', order=4,
ipynb= '~Diksha_Gabha/3598')
```
| github_jupyter |
# Timeseries anomaly detection using an Autoencoder
**Author:** [pavithrasv](https://github.com/pavithrasv)<br>
**Date created:** 2020/05/31<br>
**Last modified:** 2020/05/31<br>
**Description:** Detect anomalies in a timeseries using an Autoencoder.
## Introduction
This script demonstrates how you can use a reconstruction convolutional
autoencoder model to detect anomalies in timeseries data.
## Setup
```
import numpy as np
import pandas as pd
from tensorflow import keras
from tensorflow.keras import layers
from matplotlib import pyplot as plt
```
## Load the data
We will use the [Numenta Anomaly Benchmark(NAB)](
https://www.kaggle.com/boltzmannbrain/nab) dataset. It provides artifical
timeseries data containing labeled anomalous periods of behavior. Data are
ordered, timestamped, single-valued metrics.
We will use the `art_daily_small_noise.csv` file for training and the
`art_daily_jumpsup.csv` file for testing. The simplicity of this dataset
allows us to demonstrate anomaly detection effectively.
```
master_url_root = "https://raw.githubusercontent.com/numenta/NAB/master/data/"
df_small_noise_url_suffix = "artificialNoAnomaly/art_daily_small_noise.csv"
df_small_noise_url = master_url_root + df_small_noise_url_suffix
df_small_noise = pd.read_csv(
df_small_noise_url, parse_dates=True, index_col="timestamp"
)
df_daily_jumpsup_url_suffix = "artificialWithAnomaly/art_daily_jumpsup.csv"
df_daily_jumpsup_url = master_url_root + df_daily_jumpsup_url_suffix
df_daily_jumpsup = pd.read_csv(
df_daily_jumpsup_url, parse_dates=True, index_col="timestamp"
)
```
## Quick look at the data
```
print(df_small_noise.head())
print(df_daily_jumpsup.head())
```
## Visualize the data
### Timeseries data without anomalies
We will use the following data for training.
```
fig, ax = plt.subplots()
df_small_noise.plot(legend=False, ax=ax)
plt.show()
```
### Timeseries data with anomalies
We will use the following data for testing and see if the sudden jump up in the
data is detected as an anomaly.
```
fig, ax = plt.subplots()
df_daily_jumpsup.plot(legend=False, ax=ax)
plt.show()
```
## Prepare training data
Get data values from the training timeseries data file and normalize the
`value` data. We have a `value` for every 5 mins for 14 days.
- 24 * 60 / 5 = **288 timesteps per day**
- 288 * 14 = **4032 data points** in total
```
# Normalize and save the mean and std we get,
# for normalizing test data.
training_mean = df_small_noise.mean()
training_std = df_small_noise.std()
df_training_value = (df_small_noise - training_mean) / training_std
print("Number of training samples:", len(df_training_value))
```
### Create sequences
Create sequences combining `TIME_STEPS` contiguous data values from the
training data.
```
TIME_STEPS = 288
# Generated training sequences for use in the model.
def create_sequences(values, time_steps=TIME_STEPS):
output = []
for i in range(len(values) - time_steps):
output.append(values[i : (i + time_steps)])
return np.stack(output)
x_train = create_sequences(df_training_value.values)
print("Training input shape: ", x_train.shape)
```
## Build a model
We will build a convolutional reconstruction autoencoder model. The model will
take input of shape `(batch_size, sequence_length, num_features)` and return
output of the same shape. In this case, `sequence_length` is 288 and
`num_features` is 1.
```
model = keras.Sequential(
[
layers.Input(shape=(x_train.shape[1], x_train.shape[2])),
layers.Conv1D(
filters=32, kernel_size=7, padding="same", strides=2, activation="relu"
),
layers.Dropout(rate=0.2),
layers.Conv1D(
filters=16, kernel_size=7, padding="same", strides=2, activation="relu"
),
layers.Conv1DTranspose(
filters=16, kernel_size=7, padding="same", strides=2, activation="relu"
),
layers.Dropout(rate=0.2),
layers.Conv1DTranspose(
filters=32, kernel_size=7, padding="same", strides=2, activation="relu"
),
layers.Conv1DTranspose(filters=1, kernel_size=7, padding="same"),
]
)
model.compile(optimizer=keras.optimizers.Adam(learning_rate=0.001), loss="mse")
model.summary()
```
## Train the model
Please note that we are using `x_train` as both the input and the target
since this is a reconstruction model.
```
history = model.fit(
x_train,
x_train,
epochs=50,
batch_size=128,
validation_split=0.1,
callbacks=[
keras.callbacks.EarlyStopping(monitor="val_loss", patience=5, mode="min")
],
)
```
Let's plot training and validation loss to see how the training went.
```
plt.plot(history.history["loss"], label="Training Loss")
plt.plot(history.history["val_loss"], label="Validation Loss")
plt.legend()
plt.show()
```
## Detecting anomalies
We will detect anomalies by determining how well our model can reconstruct
the input data.
1. Find MAE loss on training samples.
2. Find max MAE loss value. This is the worst our model has performed trying
to reconstruct a sample. We will make this the `threshold` for anomaly
detection.
3. If the reconstruction loss for a sample is greater than this `threshold`
value then we can infer that the model is seeing a pattern that it isn't
familiar with. We will label this sample as an `anomaly`.
```
# Get train MAE loss.
x_train_pred = model.predict(x_train)
train_mae_loss = np.mean(np.abs(x_train_pred - x_train), axis=1)
plt.hist(train_mae_loss, bins=50)
plt.xlabel("Train MAE loss")
plt.ylabel("No of samples")
plt.show()
# Get reconstruction loss threshold.
threshold = np.max(train_mae_loss)
print("Reconstruction error threshold: ", threshold)
```
### Compare recontruction
Just for fun, let's see how our model has recontructed the first sample.
This is the 288 timesteps from day 1 of our training dataset.
```
# Checking how the first sequence is learnt
plt.plot(x_train[0])
plt.plot(x_train_pred[0])
plt.show()
```
### Prepare test data
```
df_test_value = (df_daily_jumpsup - training_mean) / training_std
fig, ax = plt.subplots()
df_test_value.plot(legend=False, ax=ax)
plt.show()
# Create sequences from test values.
x_test = create_sequences(df_test_value.values)
print("Test input shape: ", x_test.shape)
# Get test MAE loss.
x_test_pred = model.predict(x_test)
test_mae_loss = np.mean(np.abs(x_test_pred - x_test), axis=1)
test_mae_loss = test_mae_loss.reshape((-1))
plt.hist(test_mae_loss, bins=50)
plt.xlabel("test MAE loss")
plt.ylabel("No of samples")
plt.show()
# Detect all the samples which are anomalies.
anomalies = test_mae_loss > threshold
print("Number of anomaly samples: ", np.sum(anomalies))
print("Indices of anomaly samples: ", np.where(anomalies))
```
## Plot anomalies
We now know the samples of the data which are anomalies. With this, we will
find the corresponding `timestamps` from the original test data. We will be
using the following method to do that:
Let's say time_steps = 3 and we have 10 training values. Our `x_train` will
look like this:
- 0, 1, 2
- 1, 2, 3
- 2, 3, 4
- 3, 4, 5
- 4, 5, 6
- 5, 6, 7
- 6, 7, 8
- 7, 8, 9
All except the initial and the final time_steps-1 data values, will appear in
`time_steps` number of samples. So, if we know that the samples
[(3, 4, 5), (4, 5, 6), (5, 6, 7)] are anomalies, we can say that the data point
5 is an anomaly.
```
# data i is an anomaly if samples [(i - timesteps + 1) to (i)] are anomalies
anomalous_data_indices = []
for data_idx in range(TIME_STEPS - 1, len(df_test_value) - TIME_STEPS + 1):
if np.all(anomalies[data_idx - TIME_STEPS + 1 : data_idx]):
anomalous_data_indices.append(data_idx)
```
Let's overlay the anomalies on the original test data plot.
```
df_subset = df_daily_jumpsup.iloc[anomalous_data_indices]
fig, ax = plt.subplots()
df_daily_jumpsup.plot(legend=False, ax=ax)
df_subset.plot(legend=False, ax=ax, color="r")
plt.show()
```
| github_jupyter |
```
from flask import Flask, url_for, request, jsonify
from flask_cors import CORS, cross_origin
import json
app = Flask(__name__)
@app.route('/python-request', methods = ['GET'])
def spring2python():
print("스프링 요청 성공!")
data = {"spring2python Success!": True}
return jsonify(data)
@app.route('/python-request-multi', methods = ['GET'])
def spring2python_multi():
data = {"Success": True}
data["userId"] = "blabla"
data["email"] = "blabla@gmail.com"
return jsonify(data)
#이것을 계기로 다음부턴 python 환경 구축시 Docker를 활용해야 할 것 같다.
@app.route('/python-request-realdata', methods = ['POST'])
def spring2python_realdata():
#python 3.9 이전 버전까지는 enconding='utf-8'이 문법에 존재함
params = json.loads(request.get_data())
#python 3.9버전 이후부터는 encoding을 치우라고 함.
#params = json.loads(request.get_data())
print("params: ",params)
if len(params) == 0:
return jsonify("No Parameter!")
params_string = ""
for key in params.keys():
params_string += 'key: {}, value: {}<br>'.format(key, params[key])
return jsonify(params_string)
if __name__ == "__main__":
app.run()
#카카오 개발자 사이트로 이동
#https://developers.kakao.com/
#로그인
#내 애플리케이션
#앱 이름 및 사업자 등록번호상 사업주 작성
#생성 이후 만든 녀석 클릭
#여러가지 키 값들이 보일 것임(이건 사용자 고유 번호라 외부에 공개하면 안됨)
import requests
import json
url = "https://dapi.kakao.com/v2/search/image"
headers = {
"Authorization": "KakaoAK " # +나의 REST API KEY값 넣어주기
}
data = {
"query": "리눅스"
}
response = requests.post(url, headers=headers, data=data)
print(response.status_code) #200이 나와야 잘 돌아가는 것이다.
def save_image(image_url, file_name):
img_response = requests.get(image_url)
if img_response.status_code == 200:
with open(file_name, "wb") as fp:
fp.write(img_response.content)
if response.status_code != 200:
print("뭔가 잘못됨!")
else:
count = 0
for image_info in response.json()['documents']:
print(f"[{count}th] image_url = ", image_info['image_url'])
count = count + 1
file_name = "test_%d.jpg" % (count)
save_image(image_info['image_url'], file_name)
#카카오톡 채팅방에서 #리눅스 로 검색했을 때 이미지로 나오는 것들을 보여주는 것을 확인할 수 있다.
#DB의 정보를 받아서 엑셀로 바꾸는 방법 완료
#CSB
import pymysql
import datetime
print('start: ', str(datetime.datetime.now())[:19])
import pickle
MYSQL_USER_DATA_SAVED_FILE = "mysql/userinfo"
f = open(MYSQL_USER_DATA_SAVED_FILE, 'rb')
mysql_user_info = pickle.load(f)
f.close()
db = pymysql.connect(
host = '127.0.0.1',
port = 3306,
user = mysql_user_info['user_id'],
passwd = mysql_user_info['password'],
# 여기서 사용할 스키마 하나 생성!
db = 'non_jpa_db'
)
print(db)
cursor = db.cursor()
sql = """
select * from market;
"""
cursor.execute(sql)
rows = cursor.fetchall()
db.close()
print(rows)
def db2csv():
with open('mysql2csv.csv', 'a') as f:
f.writelines(text[:-1] + '\n')
for x in rows:
text = ''
for y in x:
y = str(y)
text = text + y + ', '
db2csv()
# !ls
!dir
```
| github_jupyter |
# regression-template
Hi 🙂, if you are seeing this notebook, you have succesfully started your first project on FloydHub 🚀, hooray!!
Predicting the price of an object given the historical data is one of the most common task of [ML](https://en.wikipedia.org/wiki/Machine_learning), usually achieved with the [Linear Regression model](https://en.wikipedia.org/wiki/Linear_regression). In this project the Linear Layer will be only the top of the iceberg of a model which combines the wideness of ML model and the deepness of DL model for NLP. The goal is to predict the price of a wine from its description (and variety).
### Predicting price of wine
In this notebook, we will build a classifier to correctly predict the price of a wine from its description. More in detail, we will combine the strength of ML and DL learning using a [Wide & Deep Model](https://medium.com/tensorflow/predicting-the-price-of-wine-with-the-keras-functional-api-and-tensorflow-a95d1c2c1b03), which provides really good performance for Regression and Recommendation tasks.
<img src="https://raw.githubusercontent.com/floydhub/regression-template/master/images/wineprice.png" width="800" height="800" align="center"/>
We will use the [Kaggle's Wine Reviews dataset](https://www.kaggle.com/zynicide/wine-reviews) for training our model. The dataset contains 10 columns and 150k rows of wine reviews.
We will:
- Preprocess text data for NLP
- Build and train a [Wide & Deep model](https://ai.googleblog.com/2016/06/wide-deep-learning-better-together-with.html) using Keras and Tensorflow
- Evaluate our model on the test set
- Run the model on your own movie reviews!
### Instructions
- To execute a code cell, click on the cell and press `Shift + Enter` (shortcut for Run).
- To learn more about Workspaces, check out the [Getting Started Notebook](get_started_workspace.ipynb).
- **Tip**: *Feel free to try this Notebook with your own data and on your own super awesome regression task.*
Now, let's get started! 🚀
## Initial Setup
Let's start by importing the packages, setting the training variables and loading the csv file from which get all the data we need.
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import itertools
import os
import math
import string
import re
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from tensorflow import keras
layers = keras.layers
```
## Training Parameters
We'll set the hyperparameters for training our model. If you understand what they mean, feel free to play around - otherwise, we recommend keeping the defaults for your first run 🙂
```
# Hyperparams if GPU is available
if tf.test.is_gpu_available():
# GPU
BATCH_SIZE = 256 # Number of examples used in each iteration
EPOCHS = 10 # Number of passes through entire dataset
MAX_LEN = 170 # Max length of review (in words)
VOCAB_SIZE = 1000 # Size of vocabulary dictionary
EMBEDDING = 8 # Dimension of word embedding vector
# Hyperparams for CPU training
else:
# CPU
BATCH_SIZE = 128
EPOCHS = 10
MAX_LEN = 170
VOCAB_SIZE = 1000
EMBEDDING = 8
```
## Data
The wine reviews dataset is already attached to your workspace (if you want to attach your own data, [check out our docs](https://docs.floydhub.com/guides/workspace/#attaching-floydhub-datasets)).
Let's take a look at data.
```
path = '/floyd/input/winereviews/wine_data.csv' # ADD path/to/dataset
# Convert the data to a Pandas data frame
data = pd.read_csv(path)
# Shuffle the data
data = data.sample(frac=1)
# Print the first 5 rows
data.head()
```
## Data Preprocessing
Here are some data cleaning step:
- Remove missing values
- Get only Varieties which appear more frequently (>= 500 times).
Then split the dataset: 80 (train) - 20 (test).
```
# Do some preprocessing to limit the # of wine varities in the dataset
# Clean it from null values
data = data[pd.notnull(data['country'])]
data = data[pd.notnull(data['price'])]
data = data.drop(data.columns[0], axis=1)
variety_threshold = 500 # Anything that occurs less than this will be removed.
value_counts = data['variety'].value_counts()
to_remove = value_counts[value_counts <= variety_threshold].index
data.replace(to_remove, np.nan, inplace=True)
data = data[pd.notnull(data['variety'])]
# Split data into train and test
train_size = int(len(data) * .8)
print ("Train size: %d" % train_size)
print ("Test size: %d" % (len(data) - train_size))
# Custom Tokenizer
re_tok = re.compile(f'([{string.punctuation}“”¨«»®´·º½¾¿¡§£₤‘’])')
def tokenize(s): return re_tok.sub(r' \1 ', s).split()
# Plot sentence by length
plt.hist([len(tokenize(s)) for s in data['description'].values], bins=50)
plt.title('Token per sentence')
plt.xlabel('Len (number of token)')
plt.ylabel('# samples')
plt.show()
```
The *Tokens per sentence* plot (see above) is useful for setting the `MAX_LEN` training hyperparameter.
```
# Train features
description_train = data['description'][:train_size]
variety_train = data['variety'][:train_size]
# Train labels
labels_train = data['price'][:train_size]
# Test features
description_test = data['description'][train_size:]
variety_test = data['variety'][train_size:]
# Test labels
labels_test = data['price'][train_size:]
```
## Wide Representation: BoW
The Code below will encode the description of each sentence using the [BoW model](https://en.wikipedia.org/wiki/Bag-of-words_model). This representation will encode each sentence to a vector that keeps track of the entries in the vocabulary which are used in the current sentences. This step will build a sparse vector(a vector with mostly zero values) for each description. The Code provides an example to help you get the intuition behind it.
The **wide** term used for defining this model is due to the sparse representation that this type of encoding carried out.
```
print("First Original Sample:", data['description'].values[0])
# Create a tokenizer to preprocess our text descriptions
tokenizer = keras.preprocessing.text.Tokenizer(num_words=VOCAB_SIZE, char_level=False)
tokenizer.fit_on_texts(description_train) # only fit on train
# Wide feature 1: sparse bag of words (bow) vocab_size vector
description_bow_train = tokenizer.texts_to_matrix(description_train)
description_bow_test = tokenizer.texts_to_matrix(description_test)
print("\nFirst Sample after BoW (sparse representation truncated at the first 100 vocabulary terms):", description_bow_train[0][:100])
# Wide feature 2: one-hot vector of variety categories
# Use sklearn utility to convert label strings to numbered index
encoder = LabelEncoder()
encoder.fit(variety_train)
variety_train = encoder.transform(variety_train)
variety_test = encoder.transform(variety_test)
num_classes = np.max(variety_train) + 1
# Convert labels to one hot
variety_train = keras.utils.to_categorical(variety_train, num_classes)
variety_test = keras.utils.to_categorical(variety_test, num_classes)
```
## Wide Model
The model will use the BoW representation for the *wine description* and One-Hot encoding representation for the *wine variety* as Features for the Wide Model (a 2 layers NN).
<img src="https://raw.githubusercontent.com/floydhub/regression-template/master/images/wide.png" width="350" height="350" align="center"/>
*Image from the [paper](https://arxiv.org/pdf/1606.07792.pdf)*
```
# Define our wide model with the functional API
bow_inputs = layers.Input(shape=(VOCAB_SIZE,))
variety_inputs = layers.Input(shape=(num_classes,))
merged_layer = layers.concatenate([bow_inputs, variety_inputs])
merged_layer = layers.Dense(256, activation='relu')(merged_layer)
predictions = layers.Dense(1)(merged_layer)
wide_model = keras.Model(inputs=[bow_inputs, variety_inputs], outputs=predictions)
wide_model.compile(loss='mse', optimizer='adam', metrics=['accuracy'])
print(wide_model.summary())
```
## Deep Representation: Embedding
The Code below will encode the description of each sentence using [Word Embedding](https://en.wikipedia.org/wiki/Word_embedding). This representation will encode each word of the sentence into a vector. Before applying this encoding we need to preprocess the wine description by converting each token to an index and pad the sentence to the same length.
```
print("First Original Sample:", data['description'].values[0])
# Deep model feature: word embeddings of wine descriptions
train_embed = tokenizer.texts_to_sequences(description_train)
test_embed = tokenizer.texts_to_sequences(description_test)
train_embed = keras.preprocessing.sequence.pad_sequences(
train_embed, maxlen=MAX_LEN, padding="post")
test_embed = keras.preprocessing.sequence.pad_sequences(
test_embed, maxlen=MAX_LEN, padding="post")
print("\nFirst Sample after Preprocessing for Embedding:", train_embed[0])
```
## Deep Model
This model build a liner layer at the top of the word embedding representaion of the wine description.
<img src="https://raw.githubusercontent.com/floydhub/regression-template/master/images/deep.png" width="450" height="450" align="center"/>
*Image from the [paper](https://arxiv.org/pdf/1606.07792.pdf)*
```
# Define our deep model with the Functional API
deep_inputs = layers.Input(shape=(MAX_LEN,))
embedding = layers.Embedding(VOCAB_SIZE, EMBEDDING, input_length=MAX_LEN)(deep_inputs)
embedding = layers.Flatten()(embedding)
embed_out = layers.Dense(1)(embedding)
deep_model = keras.Model(inputs=deep_inputs, outputs=embed_out)
print(deep_model.summary())
deep_model.compile(loss='mse',
optimizer='adam',
metrics=['accuracy'])
```
## Wide & Deep Model
We will implement a model similar to Heng-Tze Cheng’s [Wide & Deep Learning for Recommender Systems](https://arxiv.org/pdf/1606.07792.pdf).
This model catenate the output of the previous models and build an additional linear layer at the top.
<img src="https://raw.githubusercontent.com/floydhub/regression-template/master/images/wide&deep.png" width="500" height="500" align="center"/>
*Image from the [paper](https://arxiv.org/pdf/1606.07792.pdf)*
```
# Combine wide and deep into one model
merged_out = layers.concatenate([wide_model.output, deep_model.output])
merged_out = layers.Dense(1)(merged_out)
combined_model = keras.Model(wide_model.input + [deep_model.input], merged_out)
print(combined_model.summary())
combined_model.compile(loss='mse',
optimizer='adam',
metrics=['accuracy'])
```
## Train& Eval
If you left the default hyperpameters in the Notebook untouched, your training should take approximately:
- On CPU machine: 1 minutes for 10 epochs.
- On GPU machine: 30 seconds for 10 epochs.
```
# Run training
combined_model.fit([description_bow_train, variety_train] + [train_embed], labels_train, epochs=EPOCHS, batch_size=BATCH_SIZE, verbose=2)
combined_model.evaluate([description_bow_test, variety_test] + [test_embed], labels_test, batch_size=BATCH_SIZE)
from ipywidgets import interact
from ipywidgets import widgets
def evaluate(num_predictions):
# Generate predictions
predictions = combined_model.predict([description_bow_test, variety_test] + [test_embed])
# Compare predictions with actual values for the first few items in our test dataset
diff = 0
for i in range(num_predictions):
val = predictions[i]
print('[{}] - {}'.format(i+1, description_test.iloc[i]))
print('Predicted: ', val[0], 'Actual: ', labels_test.iloc[i], '\n')
diff += abs(val[0] - labels_test.iloc[i])
# Compare the average difference between actual price and the model's predicted price
print('Average prediction difference: ', diff / num_predictions)
interact(evaluate, num_predictions=widgets.IntSlider(value=1, min=1, max=20, description='# of test to evaluate/show'));
```
## It's your turn
Test out the model you just trained. Run the code Cell below and type your reviews in the widget, Have fun!🎉
Here are some inspirations:
- **Description**: 'From 18-year-old vines, this supple well-balanced effort blends flavors of mocha, cherry, vanilla and breakfast tea. Superbly integrated and delicious even at this early stage, this wine seems destined for a long and savory cellar life. Drink now through 2028.', **Variety**: 'Pinot Noir'.
- **Description**: 'The Quarts de Chaume, the four fingers of land that rise above the Layon Valley, are one of the pinnacles of sweet wines in the Loire. Showing botrytis and layers of dryness over the honey and peach jelly flavors, but also has great freshness. The aftertaste just lasts.', **Variety**: 'Chenin Blanc'.
- **Description**: 'Nicely oaked blackberry, licorice, vanilla and charred aromas are smooth and sultry. This is an outstanding wine from an excellent year. Forward barrel-spice and mocha flavors adorn core blackberry and raspberry fruit, while this runs long and tastes vaguely chocolaty on the velvety finish. Enjoy this top-notch Tempranillo through 2030.', **Variety**: 'Tempranillo'.
- **Description**: 'Bright, light oak shadings dress up this medium-bodied wine, complementing the red cherry and strawberry flavors. Its fresh, fruity and not very tannic—easy to drink and enjoy.', **Variety**: 'Sauvignon Blanc'.
- **Description**: 'This wine features black cherry, blackberry, blueberry with aromas of black licorice and earth. Ending with a creamy vanilla finish.', **Variety**: 'Syrah'.
Can you do better? Play around with the model hyperparameters!
```
from ipywidgets import interact_manual
from ipywidgets import widgets
def get_prediction(test_description, test_variety):
# Wide model features
bow_description = tokenizer.texts_to_matrix([test_description])
variety = encoder.transform([test_variety])
variety = keras.utils.to_categorical(variety, len(encoder.classes_))
# Deep model feature: word embeddings of wine descriptions
embed_description = tokenizer.texts_to_sequences([test_description])
embed_description = keras.preprocessing.sequence.pad_sequences(
embed_description, maxlen=MAX_LEN, padding="post")
# Evaluate
predictions = combined_model.predict([bow_description, variety] + [embed_description])
print('DESCRIPTION:', test_description)
print('VARIETY:', test_variety)
print('PREDICTED:', predictions[0][0])
interact_manual(get_prediction,
test_description=widgets.Textarea(placeholder='Type a wine Description here'),
test_variety=widgets.Text(placeholder='Type a wine Variety here'));
```
## Save your model
```
import pickle
# Saving Tokenizer
with open('models/tokenizer.pickle', 'wb') as handle:
pickle.dump(tokenizer, handle, protocol=pickle.HIGHEST_PROTOCOL)
# Saving Variety Encode
with open('models/encoder.pickle', 'wb') as handle:
pickle.dump(encoder, handle, protocol=pickle.HIGHEST_PROTOCOL)
# Saving Model Weight
combined_model.save_weights('models/wide_and_deep_weights.h5')
```
##### That's all folks - don't forget to shutdown your workspace once you're done 🙂
| github_jupyter |
# MapD Charting Example with Altair
Let's see if we can replicate [this](https://omnisci.github.io/mapd-charting/example/example1.html) MapD charting example in Python with Altair, Vega Lite, and Vega:

```
import altair as alt
import ibis
import ibis_vega_transform
conn = ibis.mapd.connect(
host='metis.mapd.com', user='mapd', password='HyperInteractive',
port=443, database='mapd', protocol= 'https'
)
t = conn.table("flights_donotmodify")
states = alt.selection_multi(fields=['origin_state'])
airlines = alt.selection_multi(fields=['carrier_name'])
# Copy default from
# https://github.com/vega/vega-lite/blob/8936751a75c3d3713b97a85b918fb30c35262faf/src/selection.ts#L281
# but add debounce
# https://vega.github.io/vega/docs/event-streams/#basic-selectors
DEBOUNCE_MS = 400
dates = alt.selection_interval(
fields=['dep_timestamp'],
encodings=['x'],
on=f'[mousedown, window:mouseup] > window:mousemove!{{0, {DEBOUNCE_MS}}}',
translate=f'[mousedown, window:mouseup] > window:mousemove!{{0, {DEBOUNCE_MS}}}',
zoom=False
)
HEIGHT = 800
WIDTH = 1000
count_filter = alt.Chart(
t[t.dep_timestamp, t.depdelay, t.origin_state, t.carrier_name],
title="Selected Rows"
).transform_filter(
airlines
).transform_filter(
dates
).transform_filter(
states
).mark_text().encode(
text='count()'
)
count_total = alt.Chart(
t,
title="Total Rows"
).mark_text().encode(
text='count()'
)
flights_by_state = alt.Chart(
t[t.origin_state, t.carrier_name, t.dep_timestamp],
title="Total Number of Flights by State"
).transform_filter(
airlines
).transform_filter(
dates
).mark_bar().encode(
x='count()',
y=alt.Y('origin_state', sort=alt.Sort(encoding='x', order='descending')),
color=alt.condition(states, alt.ColorValue("steelblue"), alt.ColorValue("grey"))
).add_selection(
states
).properties(
height= 2 * HEIGHT / 3,
width=WIDTH / 2
) + alt.Chart(
t[t.origin_state, t.carrier_name, t.dep_timestamp],
).transform_filter(
airlines
).transform_filter(
dates
).mark_text(dx=20).encode(
x='count()',
y=alt.Y('origin_state', sort=alt.Sort(encoding='x', order='descending')),
text='count()'
).properties(
height= 2 * HEIGHT / 3,
width=WIDTH / 2
)
carrier_delay = alt.Chart(
t[t.depdelay, t.arrdelay, t.carrier_name, t.origin_state, t.dep_timestamp],
title="Carrier Departure Delay by Arrival Delay (Minutes)"
).transform_filter(
states
).transform_filter(
dates
).transform_aggregate(
depdelay='mean(depdelay)',
arrdelay='mean(arrdelay)',
groupby=["carrier_name"]
).mark_point(filled=True, size=200).encode(
x='depdelay',
y='arrdelay',
color=alt.condition(airlines, alt.ColorValue("steelblue"), alt.ColorValue("grey")),
tooltip=['carrier_name', 'depdelay', 'arrdelay']
).add_selection(
airlines
).properties(
height=2 * HEIGHT / 3,
width=WIDTH / 2
) + alt.Chart(
t[t.depdelay, t.arrdelay, t.carrier_name, t.origin_state, t.dep_timestamp],
).transform_filter(
states
).transform_filter(
dates
).transform_aggregate(
depdelay='mean(depdelay)',
arrdelay='mean(arrdelay)',
groupby=["carrier_name"]
).mark_text().encode(
x='depdelay',
y='arrdelay',
text='carrier_name',
).properties(
height=2 * HEIGHT / 3,
width=WIDTH / 2
)
time = alt.Chart(
t[t.dep_timestamp, t.depdelay, t.origin_state, t.carrier_name],
title='Number of Flights by Departure Time'
).transform_filter(
'datum.dep_timestamp != null'
).transform_filter(
airlines
).transform_filter(
states
).mark_line().encode(
alt.X(
'yearmonthdate(dep_timestamp):T',
),
alt.Y(
'count():Q',
scale=alt.Scale(zero=False)
)
).add_selection(
dates
).properties(
height=HEIGHT / 3,
width=WIDTH + 50
)
(
(count_filter | count_total) & (flights_by_state | carrier_delay) & time
).configure_axis(
grid=False
).configure_view(
strokeOpacity=0
)
```
| github_jupyter |
# python requests
## 官网
- https://github.com/psf/requests
- https://requests.readthedocs.io/en/master/
> Requests is an elegant and simple HTTP library for Python, built with ♥.
> - 使用requests可以模拟浏览器的请求
>
## 安装
```
pip install requests
```
## 导入
```py
import requests
```
```
import requests
requests.__version__
```
## 请求
### 基本操作
- response = request.get()
- response.text 查看响应内容
- response.url 查看完整url
- response.encoding 查看响应头编码
- response.status_code 查看响应码
- response.content 查看响应数据(字节流)
```
r = requests.get(url="https://www.baidu.com")
r.status_code
r.text
r.encoding
r.content
r.url
```
### get
- requests.get("url") 发送get请求
- requests.get(url=url,params=params) 发送 get 请求携带参数 params 参数为dict类型
- requests.get(url=url,params=params,headers=headers) headers 头部信息
```
params = {
"ie": "utf-8",
"f": "8",
"rsv_bp": "1",
"rsv_idx": "1",
"tn": "baidu",
"wd": "requests",
"oq": "request",
"rsv_pq": "e3b4f57a0010240c",
"rsv_t": "b545zAG9t5UjEakNGna6+OewV9HM86fRhM74Ua9mKBveF6SIeZ8UdJdF3Qw",
"rqlang": "cn",
"rsv_enter": "1",
"rsv_dl": "tb",
"rsv_sug3": "2",
"rsv_sug1": "1",
"rsv_sug7": "100",
"rsv_sug2": "0",
"inputT": "429",
"rsv_sug4": "514",
}
headers = {
"User-Agent": "Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/77.0.3865.120 Safari/537.36"
}
req = requests.get("https://www.baidu.com/s",params=params,headers=headers)
req.content
```
### post
- request.post(url,data=query_data) 发生 post 请求,参数为 query_data 类型为 dict
### proxies 代理
```py
proxies = {
"http":"",
"https":""
}
res = request.get(url,proxies=proxies)
```
- request.get(url,proxies=proxies) 发送一个代理请求
### auth
- requests.get('https://api.github.com/user', auth=('user', 'pass')) 登录验证(用户名,密码)
### cookies
- cookies = response.cookies 获取cookies
- cookies_dict = requests.utils.dict_from_cookiejar(cookies) 将cookies转换为dict
```
req.cookies
requests.utils.dict_from_cookiejar(req.cookies)
```
### session
```python
# 创建session对象
s = requests.Session()
# session访问
s.get('https://httpbin.org/cookies/set/sessioncookie/123456789')
r = s.get('https://httpbin.org/cookies')
print(r.text)
```
```
session = requests.Session()
s_req = session.get(req.url)
requests.utils.dict_from_cookiejar(s_req.cookies)
session.headers
```
### SSL
- response = requests.get(url, verify=True) verify 参数为是否进行ssl校验
### 发送文件
```python
file_dict = {
'f1': open('readme.md', 'rb')
}
requests.request(method='POST',
url='http://127.0.0.1:8000/test/',
files=file_dict)
```
| github_jupyter |
# A Simple Autoencoder
We'll start off by building a simple autoencoder to compress the MNIST dataset. With autoencoders, we pass input data through an encoder that makes a compressed representation of the input. Then, this representation is passed through a decoder to reconstruct the input data. Generally the encoder and decoder will be built with neural networks, then trained on example data.

In this notebook, we'll be build a simple network architecture for the encoder and decoder. Let's get started by importing our libraries and getting the dataset.
```
%matplotlib inline
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('MNIST_data', validation_size=0)
```
Below I'm plotting an example image from the MNIST dataset. These are 28x28 grayscale images of handwritten digits.
```
img = mnist.train.images[2]
plt.imshow(img.reshape((28, 28)), cmap='Greys_r')
```
We'll train an autoencoder with these images by flattening them into 784 length vectors. The images from this dataset are already normalized such that the values are between 0 and 1. Let's start by building basically the simplest autoencoder with a **single ReLU hidden layer**. This layer will be used as the compressed representation. Then, the encoder is the input layer and the hidden layer. The decoder is the hidden layer and the output layer. Since the images are normalized between 0 and 1, we need to use a **sigmoid activation on the output layer** to get values matching the input.

> **Exercise:** Build the graph for the autoencoder in the cell below. The input images will be flattened into 784 length vectors. The targets are the same as the inputs. And there should be one hidden layer with a ReLU activation and an output layer with a sigmoid activation. Feel free to use TensorFlow's higher level API, `tf.layers`. For instance, you would use [`tf.layers.dense(inputs, units, activation=tf.nn.relu)`](https://www.tensorflow.org/api_docs/python/tf/layers/dense) to create a fully connected layer with a ReLU activation. The loss should be calculated with the cross-entropy loss, there is a convenient TensorFlow function for this `tf.nn.sigmoid_cross_entropy_with_logits` ([documentation](https://www.tensorflow.org/api_docs/python/tf/nn/sigmoid_cross_entropy_with_logits)). You should note that `tf.nn.sigmoid_cross_entropy_with_logits` takes the logits, but to get the reconstructed images you'll need to pass the logits through the sigmoid function.
```
# Size of the encoding layer (the hidden layer)
encoding_dim = 32 # feel free to change this value
image_size = mnist.train.images.shape[1]
# Input and target placeholders
inputs_ = tf.placeholder(tf.float32, (None, image_size), name="inputs")
targets_ = tf.placeholder(tf.float32, (None, image_size), name="targets")
# Output of hidden layer, single fully connected layer here with ReLU activation
encoded = tf.layers.dense(inputs_, 32, activation=tf.nn.relu)
# Output layer logits, fully connected layer with no activation
logits = tf.layers.dense(encoded, image_size, activation=None)
# Sigmoid output from logits
decoded = tf.sigmoid(logits, name = "decoded")
# Sigmoid cross-entropy loss
loss = tf.nn.sigmoid_cross_entropy_with_logits(logits=logits, labels=targets_)
# Mean of the loss
cost = tf.reduce_mean(loss)
# Adam optimizer
opt = tf.train.AdamOptimizer().minimize(cost)
```
## Training
```
# Create the session
sess = tf.Session()
```
Here I'll write a bit of code to train the network. I'm not too interested in validation here, so I'll just monitor the training loss.
Calling `mnist.train.next_batch(batch_size)` will return a tuple of `(images, labels)`. We're not concerned with the labels here, we just need the images. Otherwise this is pretty straightfoward training with TensorFlow. We initialize the variables with `sess.run(tf.global_variables_initializer())`. Then, run the optimizer and get the loss with `batch_cost, _ = sess.run([cost, opt], feed_dict=feed)`.
```
epochs = 20
batch_size = 200
sess.run(tf.global_variables_initializer())
for e in range(epochs):
for ii in range(mnist.train.num_examples//batch_size):
batch = mnist.train.next_batch(batch_size)
feed = {inputs_: batch[0], targets_: batch[0]}
batch_cost, _ = sess.run([cost, opt], feed_dict=feed)
print("Epoch: {}/{}...".format(e+1, epochs),
"Training loss: {:.4f}".format(batch_cost))
```
## Checking out the results
Below I've plotted some of the test images along with their reconstructions. For the most part these look pretty good except for some blurriness in some parts.
```
fig, axes = plt.subplots(nrows=2, ncols=10, sharex=True, sharey=True, figsize=(20,4))
in_imgs = mnist.test.images[:10]
reconstructed, compressed = sess.run([decoded, encoded], feed_dict={inputs_: in_imgs})
for images, row in zip([in_imgs, reconstructed], axes):
for img, ax in zip(images, row):
ax.imshow(img.reshape((28, 28)), cmap='Greys_r')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
fig.tight_layout(pad=0.1)
sess.close()
```
## Up Next
We're dealing with images here, so we can (usually) get better performance using convolution layers. So, next we'll build a better autoencoder with convolutional layers.
In practice, autoencoders aren't actually better at compression compared to typical methods like JPEGs and MP3s. But, they are being used for noise reduction, which you'll also build.
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
%matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
m = 2000 # 5, 50, 100, 500, 1000, 2000
desired_num = 2000
tr_i = 0
tr_j = int(desired_num/2)
tr_k = desired_num
tr_i, tr_j, tr_k
```
# Generate dataset
```
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
x = np.zeros((5000,2))
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [5,5],cov=[[0.1,0],[0,0.1]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [-6,7],cov=[[0.1,0],[0,0.1]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [-5,-4],cov=[[0.1,0],[0,0.1]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [-1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [0,2],cov=[[0.1,0],[0,0.1]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [1,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [0,-1],cov=[[0.1,0],[0,0.1]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [0,0],cov=[[0.1,0],[0,0.1]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [-0.5,-0.5],cov=[[0.1,0],[0,0.1]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [0.4,0.2],cov=[[0.1,0],[0,0.1]],size=sum(idx[9]))
x[idx[0]][0], x[idx[5]][5]
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
np.unique(bg_idx).shape
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
np.reshape(a,(2*m,1))
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(np.reshape(a,(2*m,1)))
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
mosaic_list_of_images.shape
mosaic_list_of_images.shape, mosaic_list_of_images[0]
for j in range(m):
print(mosaic_list_of_images[0][2*j:2*j+2])
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number, m):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
cnt = 0
counter = np.zeros(m) #np.array([0,0,0,0,0,0,0,0,0])
for i in range(len(mosaic_dataset)):
img = torch.zeros([2], dtype=torch.float64)
np.random.seed(int(dataset_number*10000 + i))
give_pref = foreground_index[i] #np.random.randint(0,9)
# print("outside", give_pref,foreground_index[i])
for j in range(m):
if j == give_pref:
img = img + mosaic_dataset[i][2*j:2*j+2]*dataset_number/m #2 is data dim
else :
img = img + mosaic_dataset[i][2*j:2*j+2]*(m-dataset_number)/((m-1)*m)
if give_pref == foreground_index[i] :
# print("equal are", give_pref,foreground_index[i])
cnt += 1
counter[give_pref] += 1
else :
counter[give_pref] += 1
avg_image_dataset.append(img)
print("number of correct averaging happened for dataset "+str(dataset_number)+" is "+str(cnt))
print("the averaging are done as ", counter)
return avg_image_dataset , labels , foreground_index
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:tr_j], mosaic_label[0:tr_j], fore_idx[0:tr_j] , 1, m)
test_dataset , labels , fg_index = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[tr_j : tr_k], mosaic_label[tr_j : tr_k], fore_idx[tr_j : tr_k] , m, m)
avg_image_dataset_1 = torch.stack(avg_image_dataset_1, axis = 0)
# avg_image_dataset_1 = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(avg_image_dataset_1, keepdims= True, axis = 0))
# print(torch.std(avg_image_dataset_1, keepdims= True, axis = 0))
print("=="*40)
test_dataset = torch.stack(test_dataset, axis = 0)
# test_dataset = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(test_dataset, keepdims= True, axis = 0))
# print(torch.std(test_dataset, keepdims= True, axis = 0))
print("=="*40)
x1 = (avg_image_dataset_1).numpy()
y1 = np.array(labels_1)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("dataset4 CIN with alpha = 1/"+str(m))
x1 = (test_dataset).numpy() / m
y1 = np.array(labels)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("test dataset4")
test_dataset[0:10]/m
test_dataset = test_dataset/m
test_dataset[0:10]
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
avg_image_dataset_1[0].shape
avg_image_dataset_1[0]
batch = 200
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
testdata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
testdata_11 = MosaicDataset(test_dataset, labels )
testloader_11 = DataLoader( testdata_11 , batch_size= batch ,shuffle=False)
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet,self).__init__()
self.linear1 = nn.Linear(2,3)
# self.linear2 = nn.Linear(50,10)
# self.linear3 = nn.Linear(10,3)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
def forward(self,x):
# x = F.relu(self.linear1(x))
# x = F.relu(self.linear2(x))
x = (self.linear1(x))
return x
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/(i+1)
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("unique out: ", np.unique(out), "unique pred: ", np.unique(pred) )
print("correct: ", correct, "total ", total)
print('Accuracy of the network on the %d test dataset %d: %.2f %%' % (total, number , 100 * correct / total))
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
net = Whatnet().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.Adam(net.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.05:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %.2f %%' % (total, 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
return loss_curi
train_loss_all=[]
testloader_list= [ testloader_1, testloader_11]
train_loss_all.append(train_all(trainloader_1, 1, testloader_list))
%matplotlib inline
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
```
| github_jupyter |
# Kurulum ve Gerekli Modullerin Yuklenmesi
```
from google.colab import drive
drive.mount('/content/gdrive')
import sys
import os
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import nltk
import os
from nltk import sent_tokenize, word_tokenize
from nltk.stem.snowball import SnowballStemmer
from nltk.stem.wordnet import WordNetLemmatizer
import nltk
nltk.download('stopwords')
import matplotlib.pyplot as plt
import pandas as pd
nltk.download('punkt')
import string
from nltk.corpus import stopwords
import pandas as pd
import numpy as np
import re
```
# Incelenecek konu basligindaki tweetlerin yuklenmesi
Burada ornek olarak ulkeler konu basligi gosteriliyor gosteriliyor
```
os.chdir("/content/gdrive/My Drive/css/ulkeler_after")
df3 = pd.read_csv("/content/gdrive/My Drive/css/ulkeler_after/ulkeler_after_nodublication.csv", engine = 'python')
df3['tweet'] = df3['tweet'].astype(str)
```
Data pre-processing (on temizlemesi):
1. kucuk harfe cevirme
2. turkce karakter uyumlarini duzeltme
3. ozel karakterleri, noktalamalari temizleme
```
df3.tweet = df3.tweet.apply(lambda x: re.sub(r"İ", "i",x)) #harika calisiyor
df3.tweet = df3.tweet.apply(lambda x: x.lower())
df3.loc[:,"tweet"] = df3.tweet.apply(lambda x : " ".join(re.findall('[\w]+',x)))
df3.head(3)
```
# Tokenize islemi, stop wordlerin atilmasi ve kelime frequencylerini (kullanim sayilarini) ileride gelecek gorsellestirme icin kaydetme
```
top_N = 10
txt = df3.tweet.str.lower().str.replace(r'\|', ' ').str.cat(sep=' ')
words = nltk.tokenize.word_tokenize(txt)
word_dist = nltk.FreqDist(words)
user_defined_stop_words = ['1', 'ye', 'nin' ,'nın', 'koronavirüs', 'olsun', 'karşı' , 'covid_19', 'artık', '3', 'sayısı' , 'olarak', 'oldu', 'olan', '2' , 'nedeniyle','bile' , 'sonra' ,'sen','virüs', 'ben', 'vaka' , 'son', 'yeni', 'sayi', 'sayisi','virüsü','bir','com','twitter', 'kadar', 'dan' , 'değil' ,'pic' , 'http', 'https' , 'www' , 'status' , 'var', 'bi', 'mi','yok', 'bu' , 've', 'korona' ,'corona' ,'19' ,'kovid', 'covid']
i = nltk.corpus.stopwords.words('turkish')
j = list(string.punctuation) + user_defined_stop_words
stopwords = set(i).union(j)
words_except_stop_dist = nltk.FreqDist(w for w in words if w not in stopwords)
print('All frequencies, including STOPWORDS:')
print('=' * 60)
rslt3 = pd.DataFrame(word_dist.most_common(top_N),
columns=['Word', 'Frequency'])
print(rslt3)
print('=' * 60)
rslt3 = pd.DataFrame(words_except_stop_dist.most_common(top_N),
columns=['Word', 'Frequency']).set_index('Word')
```
# TR deki ilk vakan onceki tweetlerin incelenmek icin yuklenmesi
```
df2 = pd.read_csv("/content/gdrive/My Drive/css/ulkeler_before/ulkeler_before_nodublication.csv", engine = 'python')
df2['tweet'] = df2['tweet'].astype(str)
df2['tweet'] = df2['tweet'].astype(str)
df2.tweet = df2.tweet.apply(lambda x: re.sub(r"İ", "i",x)) #harika calisiyor
df2.tweet = df2.tweet.apply(lambda x: x.lower())
df2.loc[:,"tweet"] = df2.tweet.apply(lambda x : " ".join(re.findall('[\w]+',x)))
df2.head()
top_N = 10
txt = df2.tweet.str.lower().str.replace(r'\|', ' ').str.cat(sep=' ')
words = nltk.tokenize.word_tokenize(txt)
word_dist = nltk.FreqDist(words)
user_defined_stop_words = ['1', 'ye', 'nin' ,'nın', 'koronavirüs', 'olsun', 'karşı' , 'covid_19', 'artık', '3', 'sayısı' , 'olarak', 'oldu', 'olan', '2' , 'nedeniyle','bile' , 'sonra' ,'sen','virüs', 'ben', 'vaka' , 'son', 'yeni', 'sayi', 'sayisi','virüsü','bir','com','twitter', 'kadar', 'dan' , 'değil' ,'pic' , 'http', 'https' , 'www' , 'status' , 'var', 'bi', 'mi','yok', 'bu' , 've', 'korona' ,'corona' ,'19' ,'kovid', 'covid']
i = nltk.corpus.stopwords.words('turkish')
j = list(string.punctuation) + user_defined_stop_words
stopwords = set(i).union(j)
words_except_stop_dist = nltk.FreqDist(w for w in words if w not in stopwords)
print('All frequencies, including STOPWORDS:')
print('=' * 60)
rslt = pd.DataFrame(word_dist.most_common(top_N),
columns=['Word', 'Frequency'])
print(rslt)
print('=' * 60)
rslt = pd.DataFrame(words_except_stop_dist.most_common(top_N),
columns=['Word', 'Frequency']).set_index('Word')
matplotlib.style.use('ggplot')
plt.figure(figsize=(30,30))
rslt.plot.bar(rot=0)
plt.savefig('ulkeler_before.png')
```
# Karsilastirmali gorsellestirme (Ayni konu basliklarinin 11 marttan oncesi ve sonrasi )
```
fig, (ax1, ax2) = plt.subplots(1,2, sharex=False, sharey= True, figsize=(24,5))
rslt3.plot.bar(rot=0, ax =ax1 , title = "Ulkeler_After" )
rslt.plot.bar(rot=0, ax =ax2 , title = "Ulkeler_Before" )
plt.savefig('ulkeler_comparison.png',dpi=300)
```
| github_jupyter |
# Recession effect on Housing Prices from University Towns
This project is about examining the Hypothesis, **"The housing price from homes across university towns were less effected by the Great Recession".**
Glossary terms/Definitions used in this assignment:
* A _quarter_ is a specific three month period.
* A _recession_ is defined as the period which starts with two consecutive quarters of GDP decline, and ending with two consecutive quarters of GDP growth.
* A _recession bottom_ is the quarter within a recession which had the lowest GDP.
* A _university town_ is a city which has a high percentage of university students compared to the total population of the city.
The following data files are used for this project:
* ```City_Zhvi_AllHomes.csv```, obtained from [Zillow research data site](http://files.zillowstatic.com/research/public/City/City_Zhvi_AllHomes.csv), contains US median house sale prices at city level.
* A list of university towns in the United States can be obtained by webscraping the [Wikipedia page](https://en.wikipedia.org/wiki/List_of_college_towns#College_towns_in_the_United_States) on college towns which has been copied and pasted into the file ```university_towns.txt```.
* GDP over time of the United States in current US dollars, in quarterly intervals, can be obtained from [Bureau of Economic Analysis, US Department of Commerce](http://www.bea.gov/national/index.htm#gdp) which has been copied and pasted into the file ```ugdplev.xls```.
```
import pandas as pd # import pandas which is high-performance, easy-to-use data structures and data analysis framework
import numpy as np # import numpy which is fundamental package for scientific computing with Python
from scipy.stats import ttest_ind
import warnings
warnings.filterwarnings("ignore")
data_dir = "../data_ds/"
# dictionary to map state names to two letter acronyms
state_acronyms = {'OH': 'Ohio', 'KY': 'Kentucky', 'AS': 'American Samoa', 'NV': 'Nevada', 'WY': 'Wyoming',
'NA': 'National', 'AL': 'Alabama', 'MD': 'Maryland', 'AK': 'Alaska', 'UT': 'Utah',
'OR': 'Oregon', 'MT': 'Montana', 'IL': 'Illinois', 'TN': 'Tennessee', 'DC': 'District of Columbia',
'VT': 'Vermont', 'ID': 'Idaho', 'AR': 'Arkansas', 'ME': 'Maine', 'WA': 'Washington', 'HI': 'Hawaii',
'WI': 'Wisconsin', 'MI': 'Michigan', 'IN': 'Indiana', 'NJ': 'New Jersey', 'AZ': 'Arizona', 'GU': 'Guam',
'MS': 'Mississippi', 'PR': 'Puerto Rico', 'NC': 'North Carolina', 'TX': 'Texas', 'SD': 'South Dakota',
'MP': 'Northern Mariana Islands', 'IA': 'Iowa', 'MO': 'Missouri', 'CT': 'Connecticut', 'WV': 'West Virginia',
'SC': 'South Carolina', 'LA': 'Louisiana', 'KS': 'Kansas', 'NY': 'New York', 'NE': 'Nebraska',
'OK': 'Oklahoma', 'FL': 'Florida', 'CA': 'California', 'CO': 'Colorado', 'PA': 'Pennsylvania',
'DE': 'Delaware', 'NM': 'New Mexico', 'RI': 'Rhode Island', 'MN': 'Minnesota', 'VI': 'Virgin Islands',
'NH': 'New Hampshire', 'MA': 'Massachusetts', 'GA': 'Georgia', 'ND': 'North Dakota', 'VA': 'Virginia'}
def get_university_towns_list():
'''Returns a DataFrame of towns and the states they are in from the
university_towns.txt list.'''
data = pd.read_table(data_dir + 'university_towns.txt', header=None) # read university_towns.txt which contains a list of university towns in the United States and store it in a dataframe
data['Cleaned Data'] = data[0].apply(lambda x: x.split(
'(')[0].strip() if x.count('(') > 0 else x) # removing every character from "(" to the end.
data['RegionName'] = data['Cleaned Data'].apply(lambda x: x if x.count('[edit]') == 0 else np.NaN) # replacing every state (ending with "[edit]") with NaN in a new column, "Region name",for easier selection of counties
data["State"] = 0 # initialise state column with 0
state = None
for index, entry in enumerate(data['Cleaned Data']): #assigning state names to regions
if entry.count('[edit]') > 0:
state = entry.split('[')[0].strip()
data["State"][index] = state
else:
data["State"][index] = state
data = data.drop(0, axis=1) #droping unwanted columns
data = data.dropna() # dropping rows with null values
data.index = list(range(len(data))) # reindexing from 0 to length of the datarfame
columns_to_keep = ['State',
'RegionName']
data = data[columns_to_keep] # keeping only columns of interest
return data
get_university_towns_list().head(10)
def get_recession_start_quarter():
'''Returns the year and quarter of the recession start time.'''
GDP_over_time = pd.ExcelFile(data_dir + 'gdplev.xls') # read university_towns.txt which contains quarterly GDP over time in the United States and store it in a dataframe with header excluded
GDP_over_time = GDP_over_time.parse(GDP_over_time.sheet_names[0]) # parse the first sheet of the dataframe
del GDP_over_time[GDP_over_time.columns[3]], GDP_over_time[GDP_over_time.columns[-1]] # delete unwanted columns
GDP_over_time.drop(GDP_over_time.index[:7],inplace=True) # drop unwanted rows
GDP_over_time = GDP_over_time.rename(index=str, columns={'Current-Dollar and "Real" Gross Domestic Product':'Year','Unnamed: 1':'GDP in billions of current dollars 1','Unnamed: 2':'GDP in billions of chained 2009 dollars 1','Unnamed: 4':'Quarter','Unnamed: 5':'GDP in billions of current dollars 2','Unnamed: 6':'GDP in billions of chained 2009 dollars 2'}) # renaming column headers
millennium_first_quarter = GDP_over_time[GDP_over_time['Quarter']=='2000q1'].index[0] # extracting index of millennium start quarter
GDP_over_time = GDP_over_time.loc[millennium_first_quarter:] # extracting the dataframe from millennium start quarter
del GDP_over_time['Year'], GDP_over_time['GDP in billions of current dollars 1'], GDP_over_time['GDP in billions of chained 2009 dollars 1'] # deleting unwanted columns
GDP_over_time.index = list(range(len(GDP_over_time))) # reindexing from 0 to length of the datarfame
Quarterly_GDP = 0.0
GDP_Decline_Counter = 0
recession_start_time = None
for index, entry in enumerate(GDP_over_time['GDP in billions of current dollars 2']): # extracting recession start quarter
if entry > Quarterly_GDP:
Quarterly_GDP = entry
elif GDP_Decline_Counter==2:
Quarterly_GDP = entry
recession_start_time = GDP_over_time['Quarter'][int(index)-3]
else:
Quarterly_GDP = entry
GDP_Decline_Counter += 1
return recession_start_time # returning recession start quarter
get_recession_start_quarter()
def get_recession_end_quarter():
'''Returns the year and quarter of the recession end time.'''
GDP_over_time = pd.ExcelFile(data_dir + 'gdplev.xls') # read university_towns.txt which contains quarterly GDP over time in the United States and store it in a dataframe with header excluded
GDP_over_time = GDP_over_time.parse(GDP_over_time.sheet_names[0]) # parse the first sheet of the dataframe
del GDP_over_time[GDP_over_time.columns[3]], GDP_over_time[GDP_over_time.columns[-1]] # delete unwanted columns
GDP_over_time.drop(GDP_over_time.index[:7],inplace=True) # drop unwanted rows
GDP_over_time = GDP_over_time.rename(index=str, columns={'Current-Dollar and "Real" Gross Domestic Product':'Year','Unnamed: 1':'GDP in billions of current dollars 1','Unnamed: 2':'GDP in billions of chained 2009 dollars 1','Unnamed: 4':'Quarter','Unnamed: 5':'GDP in billions of current dollars 2','Unnamed: 6':'GDP in billions of chained 2009 dollars 2'}) # renaming column headers
millennium_first_quarter = GDP_over_time[GDP_over_time['Quarter']=='2000q1'].index[0] # extracting index of millennium start quarter
GDP_over_time = GDP_over_time.loc[millennium_first_quarter:] # extracting the dataframe from millennium start quarter
del GDP_over_time['Year'], GDP_over_time['GDP in billions of current dollars 1'], GDP_over_time['GDP in billions of chained 2009 dollars 1'] # deleting unwanted columns
GDP_over_time.index = list(range(len(GDP_over_time))) # reindexing from 0 to length of the datarfame
Quarterly_GDP = 0.0
GDP_Decline_Counter = 0
recession_start_time = None
for index, entry in enumerate(GDP_over_time['GDP in billions of current dollars 2']): # extracting recession end quarter
if entry > Quarterly_GDP:
Quarterly_GDP = entry
elif GDP_Decline_Counter==2:
Quarterly_GDP = entry
recession_start_time = GDP_over_time['Quarter'][int(index)-3]
else:
Quarterly_GDP = entry
GDP_Decline_Counter += 1
recession_start_quarter = GDP_over_time[GDP_over_time['Quarter']==recession_start_time].index[0]
GDP_over_time_revised = GDP_over_time.loc[recession_start_quarter:]
GDP_over_time_revised.index = list(range(len(GDP_over_time_revised)))
Quarterly_GDP = GDP_over_time_revised['GDP in billions of current dollars 2'][0]
GDP_Growth_Counter = 0
recession_end_time = 0
for index, entry in enumerate(GDP_over_time_revised['GDP in billions of current dollars 2']):
if GDP_Growth_Counter == 2:
Quarterly_GDP = entry
recession_end_time = GDP_over_time_revised['Quarter'][int(index-1)]
break
elif entry > Quarterly_GDP:
Quarterly_GDP = entry
GDP_Growth_Counter += 1
else:
Quarterly_GDP = entry
return recession_end_time # returning recession end quarter
get_recession_end_quarter()
def get_recession_bottom_quarter():
'''Returns the year and quarter of the recession bottom time.'''
GDP_over_time = pd.ExcelFile(data_dir + 'gdplev.xls') # read university_towns.txt which contains quarterly GDP over time in the United States and store it in a dataframe with header excluded
GDP_over_time = GDP_over_time.parse(GDP_over_time.sheet_names[0]) # parse the first sheet of the dataframe
del GDP_over_time[GDP_over_time.columns[3]], GDP_over_time[GDP_over_time.columns[-1]] # delete unwanted columns
GDP_over_time.drop(GDP_over_time.index[:7],inplace=True) # drop unwanted rows
GDP_over_time = GDP_over_time.rename(index=str, columns={'Current-Dollar and "Real" Gross Domestic Product':'Year','Unnamed: 1':'GDP in billions of current dollars 1','Unnamed: 2':'GDP in billions of chained 2009 dollars 1','Unnamed: 4':'Quarter','Unnamed: 5':'GDP in billions of current dollars 2','Unnamed: 6':'GDP in billions of chained 2009 dollars 2'}) # renaming column headers
millennium_first_quarter = GDP_over_time[GDP_over_time['Quarter']=='2000q1'].index[0] # extracting index of millennium start quarter
GDP_over_time = GDP_over_time.loc[millennium_first_quarter:] # extracting the dataframe from millennium start quarter
del GDP_over_time['Year'], GDP_over_time['GDP in billions of current dollars 1'], GDP_over_time['GDP in billions of chained 2009 dollars 1'] # deleting unwanted columns
GDP_over_time.index = list(range(len(GDP_over_time))) # reindexing from 0 to length of the datarfame
Quarterly_GDP = 0.0
GDP_Decline_Counter = 0
recession_start_time = None
for index, entry in enumerate(GDP_over_time['GDP in billions of current dollars 2']): # extracting recession bottom quarter
if entry > Quarterly_GDP:
Quarterly_GDP = entry
elif GDP_Decline_Counter==2:
Quarterly_GDP = entry
recession_start_time = GDP_over_time['Quarter'][int(index)-3]
else:
Quarterly_GDP = entry
GDP_Decline_Counter += 1
recession_start_quarter = GDP_over_time[GDP_over_time['Quarter']==recession_start_time].index[0]
GDP_over_time_revised = GDP_over_time.loc[recession_start_quarter:]
GDP_over_time_revised.index = list(range(len(GDP_over_time_revised)))
Quarterly_GDP = GDP_over_time_revised['GDP in billions of current dollars 2'][0]
GDP_Growth_Counter = 0
recession_end_time = 0
for index, entry in enumerate(GDP_over_time_revised['GDP in billions of current dollars 2']): # extracting recession bottom quarter
if GDP_Growth_Counter == 2:
Quarterly_GDP = entry
recession_end_time = GDP_over_time_revised['Quarter'][int(index-1)]
break
elif entry > Quarterly_GDP:
Quarterly_GDP = entry
GDP_Growth_Counter += 1
else:
Quarterly_GDP = entry
recession_end_quarter = GDP_over_time_revised[GDP_over_time_revised['Quarter']==recession_end_time].index[0]
GDP_over_time_revised = GDP_over_time_revised.loc[:recession_end_quarter]
argument_minimum = np.argmin(GDP_over_time_revised['GDP in billions of current dollars 2'], axis=0)[0]
return GDP_over_time_revised['Quarter'][argument_minimum] # returning recession bottom quarter
get_recession_bottom_quarter()
def convert_housing_data_to_mean_quarters_price():
'''Converts the housing data to quarters and returns it as mean
values in a dataframe.'''
housing_data = pd.read_csv(data_dir + 'City_Zhvi_AllHomes.csv') # read City_Zhvi_AllHomes.csv which contains median home sale prices and store it in a dataframe
housing_data.drop(housing_data.columns[6:51],axis=1, inplace=True) # frod unwanted year columns
for index_2 in range(int(67)): # data preprocessing for renaming quarter columns
index_quarter = str(2000 + divmod(index_2, 4)[0])+"q"+str(((index_2)%4)+1)
housing_data[index_quarter] = 0
housing_data_revised = housing_data.copy() # making a copy of the dataframe for help in renaming quarter columns
housing_data.drop(housing_data.columns[-67:],axis=1, inplace=True)
housing_data_revised.drop(housing_data_revised.columns[6:-67],axis=1,inplace=True)
for index_2 in range(int(len(housing_data_revised.columns[6:]))): # data preprocessing for renaming quarter columns
housing_data_revised[housing_data_revised.columns[6:][index_2]] = housing_data[housing_data.columns[6+(index_2*3):6+(index_2*3)+3]].mean(axis=1)
housing_data_revised = housing_data_revised.sort_values('State', ascending=True) # sort by state name in alphabetical order
housing_data_revised = housing_data_revised.replace(state_acronyms) # replace state values from two letter abbreviation to full state name according to aforementioned dictionary
housing_data_revised = housing_data_revised.set_index(['State','RegionName']) # setting a multi index of state and county
housing_data_revised.drop(housing_data_revised.columns[:4],axis=1,inplace=True) # droping unwanted columns
return housing_data_revised
convert_housing_data_to_mean_quarters_price().head(10)
def ttest():
'''First the new data showing the decline or growth of housing prices between the recession start
and recession botton. Then a ttest is ran to compare the university town values to the non-university
town values, return whether the alternative hypotheses is true or not. The p-value of confidence is
then computed.
A tuple (different, p, better) is returned where different=True if the t-test is
True at a p<0.01 (we reject the null hypothesis), or different=False if
otherwise (we cannot reject the null hypothesis). The variable p should
be equal to the exact p value returned from scipy.stats.ttest_ind(). The
value for better should be either "university town" or "non-university town"
depending on which has a lower mean price ratio (which is equivilent to a
reduced market loss).'''
housing_data_revised = convert_housing_data_to_mean_quarters_price() # obtain mean quarterly housing values from the above function
housing_data_revised = housing_data_revised[housing_data_revised.columns[33:-29]] # extract mean quarterly housing values from only the recession period
university_towns = get_university_towns_list() #obtain list of university towns form the above function
university_towns = university_towns.sort_values('State', ascending=True) # sort university towns by state name in alphabetical order
university_towns = university_towns.set_index(['State','RegionName']) # set state name and county as multi-index
university_town_values = pd.merge(university_towns, housing_data_revised, how='inner', left_index=True, right_index=True) # obtain mean quarterly housing values in university towns
non_university_town_values = housing_data_revised[~housing_data_revised.index.isin(university_town_values.index)] # obtain mean quarterly values of housing in non-university towns
university_town_mean_ratio = ((university_town_values['2008q2'])/(university_town_values['2009q2'])).mean() # obtain ratio of mean quarterly housing values in recession start to recession bottom in university towns
non_university_town_mean_ratio = ((non_university_town_values['2008q2'])/(non_university_town_values['2009q2'])).mean() # obtain ratio of mean quarterly housing values in recession start to recession bottom in non-university towns
from scipy import stats
p = stats.ttest_ind((university_town_values['2008q2'])/(university_town_values['2009q2']), (non_university_town_values['2008q2'])/(non_university_town_values['2009q2']), nan_policy='omit')[1] # run t-test on the above ratios of university and non-university towns
different = p<0.01
better = "university town" if (university_town_mean_ratio < non_university_town_mean_ratio) else "non-university town"
return (different, p, better)
ttest()
```
| github_jupyter |
# Multilayer perceptrons from scratch
In the previous chapters we showed how you could implement multiclass logistic regression
(also called *softmax regression*)
for classifiying images of handwritten digits into the 10 possible categories
([from scratch](../chapter02_supervised-learning/softmax-regression-scratch.ipynb) and [with gluon](../chapter02_supervised-learning/softmax-regression-gluon.ipynb)).
This is where things start to get fun.
We understand how to wrangle data,
coerce our outputs into a valid probability distribution,
how to apply an appropriate loss function,
and how to optimize over our parameters.
Now that we've covered these preliminaries,
we can extend our toolbox to include deep neural networks.
Recall that before, we mapped our inputs directly onto our outputs through a single linear transformation.
$$\hat{y} = \mbox{softmax}(W \boldsymbol{x} + b)$$
Graphically, we could depict the model like this, where the orange nodes represent inputs and the teal nodes on the top represent the output:

If our labels really were related to our input data by an approximately linear function,
then this approach might be adequate.
*But linearity is a strong assumption*.
Linearity means that given an output of interest,
for each input,
increasing the value of the input should either drive the value of the output up
or drive it down,
irrespective of the value of the other inputs.
Imagine the case of classifying cats and dogs based on black and white images.
That's like saying that for each pixel,
increasing its value either increases the probability that it depicts a dog or decreases it.
That's not reasonable. After all, the world contains both black dogs and black cats, and both white dogs and white cats.
Teasing out what is depicted in an image generally requires allowing more complex relationships between
our inputs and outputs, considering the possibility that our pattern might be characterized by interactions among the many features.
In these cases, linear models will have low accuracy.
We can model a more general class of functions by incorporating one or more *hidden layers*.
The easiest way to do this is to stack a bunch of layers of neurons on top of each other.
Each layer feeds into the layer above it, until we generate an output.
This architecture is commonly called a "multilayer perceptron".
With an MLP, we're going to stack a bunch of layers on top of each other.
$$ h_1 = \phi(W_1\boldsymbol{x} + b_1) $$
$$ h_2 = \phi(W_2\boldsymbol{h_1} + b_2) $$
$$...$$
$$ h_n = \phi(W_n\boldsymbol{h_{n-1}} + b_n) $$
Note that each layer requires its own set of parameters.
For each hidden layer, we calculate its value by first applying a linear function
to the activations of the layer below, and then applying an element-wise
nonlinear activation function.
Here, we've denoted the activation function for the hidden layers as $\phi$.
Finally, given the topmost hidden layer, we'll generate an output.
Because we're still focusing on multiclass classification, we'll stick with the softmax activation in the output layer.
$$ \hat{y} = \mbox{softmax}(W_y \boldsymbol{h}_n + b_y)$$
Graphically, a multilayer perceptron could be depicted like this:

Multilayer perceptrons can account for complex interactions in the inputs because
the hidden neurons depend on the values of each of the inputs.
It's easy to design a hidden node that that does arbitrary computation,
such as, for instance, logical operations on its inputs.
And it's even widely known that multilayer perceptrons are universal approximators.
That means that even for a single-hidden-layer neural network,
with enough nodes, and the right set of weights, it could model any function at all!
Actually learning that function is the hard part.
And it turns out that we can approximate functions much more compactly if we use deeper (vs wider) neural networks.
We'll get more into the math in a subsequent chapter, but for now let's actually build an MLP.
In this example, we'll implement a multilayer perceptron with two hidden layers and one output layer.
## Imports
```
from __future__ import print_function
import mxnet as mx
import numpy as np
from mxnet import nd, autograd, gluon
```
## Set contexts
```
ctx = mx.gpu() if mx.test_utils.list_gpus() else mx.cpu()
data_ctx = ctx
model_ctx = ctx
```
## Load MNIST data
Let's go ahead and grab our data.
```
num_inputs = 784
num_outputs = 10
batch_size = 64
num_examples = 60000
def transform(data, label):
return data.astype(np.float32)/255, label.astype(np.float32)
train_data = gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=True, transform=transform),
batch_size, shuffle=True)
test_data = gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=False, transform=transform),
batch_size, shuffle=False)
```
## Allocate parameters
```
#######################
# Set some constants so it's easy to modify the network later
#######################
num_hidden = 256
weight_scale = .01
#######################
# Allocate parameters for the first hidden layer
#######################
W1 = nd.random_normal(shape=(num_inputs, num_hidden), scale=weight_scale, ctx=model_ctx)
b1 = nd.random_normal(shape=num_hidden, scale=weight_scale, ctx=model_ctx)
#######################
# Allocate parameters for the second hidden layer
#######################
W2 = nd.random_normal(shape=(num_hidden, num_hidden), scale=weight_scale, ctx=model_ctx)
b2 = nd.random_normal(shape=num_hidden, scale=weight_scale, ctx=model_ctx)
#######################
# Allocate parameters for the output layer
#######################
W3 = nd.random_normal(shape=(num_hidden, num_outputs), scale=weight_scale, ctx=model_ctx)
b3 = nd.random_normal(shape=num_outputs, scale=weight_scale, ctx=model_ctx)
params = [W1, b1, W2, b2, W3, b3]
```
Again, let's allocate space for each parameter's gradients.
```
for param in params:
param.attach_grad()
```
## Activation functions
If we compose a multi-layer network but use only linear operations, then our entire network will still be a linear function. That's because $\hat{y} = X \cdot W_1 \cdot W_2 \cdot W_2 = X \cdot W_4 $ for $W_4 = W_1 \cdot W_2 \cdot W3$. To give our model the capacity to capture nonlinear functions, we'll need to interleave our linear operations with activation functions. In this case, we'll use the rectified linear unit (ReLU):
```
def relu(X):
return nd.maximum(X, nd.zeros_like(X))
```
## Softmax output
As with multiclass logistic regression, we'll want the outputs to constitute a valid probability distribution. We'll use the same softmax activation function on our output to make sure that our outputs sum to one and are non-negative.
```
def softmax(y_linear):
exp = nd.exp(y_linear-nd.max(y_linear))
partition = nd.nansum(exp, axis=0, exclude=True).reshape((-1, 1))
return exp / partition
```
## The *softmax* cross-entropy loss function
In the previous example, we calculated our model's output and then ran this output through the cross-entropy loss function:
```
def cross_entropy(yhat, y):
return - nd.nansum(y * nd.log(yhat), axis=0, exclude=True)
```
Mathematically, that's a perfectly reasonable thing to do. However, computationally, things can get hairy. We'll revisit the issue at length in a chapter more dedicated to implementation and less interested in statistical modeling. But we're going to make a change here so we want to give you the gist of why.
Recall that the softmax function calculates $\hat y_j = \frac{e^{z_j}}{\sum_{i=1}^{n} e^{z_i}}$, where $\hat y_j$ is the j-th element of the input ``yhat`` variable in function ``cross_entropy`` and $z_j$ is the j-th element of the input ``y_linear`` variable in function ``softmax``
If some of the $z_i$ are very large (i.e. very positive), $e^{z_i}$ might be larger than the largest number we can have for certain types of ``float`` (i.e. overflow). This would make the denominator (and/or numerator) ``inf`` and we get zero, or ``inf``, or ``nan`` for $\hat y_j$. In any case, we won't get a well-defined return value for ``cross_entropy``. This is the reason we subtract $\text{max}(z_i)$ from all $z_i$ first in ``softmax`` function. You can verify that this shifting in $z_i$ will not change the return value of ``softmax``.
After the above subtraction/ normalization step, it is possible that $z_j$ is very negative. Thus, $e^{z_j}$ will be very close to zero and might be rounded to zero due to finite precision (i.e underflow), which makes $\hat y_j$ zero and we get ``-inf`` for $\text{log}(\hat y_j)$. A few steps down the road in backpropagation, we starts to get horrific not-a-number (``nan``) results printed to screen.
Our salvation is that even though we're computing these exponential functions, we ultimately plan to take their log in the cross-entropy functions. It turns out that by combining these two operators ``softmax`` and ``cross_entropy`` together, we can elude the numerical stability issues that might otherwise plague us during backpropagation. As shown in the equation below, we avoided calculating $e^{z_j}$ but directly used $z_j$ due to $log(exp(\cdot))$.
$$\text{log}{(\hat y_j)} = \text{log}\left( \frac{e^{z_j}}{\sum_{i=1}^{n} e^{z_i}}\right) = \text{log}{(e^{z_j})}-\text{log}{\left( \sum_{i=1}^{n} e^{z_i} \right)} = z_j -\text{log}{\left( \sum_{i=1}^{n} e^{z_i} \right)}$$
We'll want to keep the conventional softmax function handy in case we ever want to evaluate the probabilities output by our model. But instead of passing softmax probabilities into our new loss function, we'll just pass our ``yhat_linear`` and compute the softmax and its log all at once inside the softmax_cross_entropy loss function, which does smart things like the log-sum-exp trick ([see on Wikipedia](https://en.wikipedia.org/wiki/LogSumExp)).
```
def softmax_cross_entropy(yhat_linear, y):
return - nd.nansum(y * nd.log_softmax(yhat_linear), axis=0, exclude=True)
```
## Define the model
Now we're ready to define our model
```
def net(X):
#######################
# Compute the first hidden layer
#######################
h1_linear = nd.dot(X, W1) + b1
h1 = relu(h1_linear)
#######################
# Compute the second hidden layer
#######################
h2_linear = nd.dot(h1, W2) + b2
h2 = relu(h2_linear)
#######################
# Compute the output layer.
# We will omit the softmax function here
# because it will be applied
# in the softmax_cross_entropy loss
#######################
yhat_linear = nd.dot(h2, W3) + b3
return yhat_linear
```
## Optimizer
```
def SGD(params, lr):
for param in params:
param[:] = param - lr * param.grad
```
## Evaluation metric
```
def evaluate_accuracy(data_iterator, net):
numerator = 0.
denominator = 0.
for i, (data, label) in enumerate(data_iterator):
data = data.as_in_context(model_ctx).reshape((-1, 784))
label = label.as_in_context(model_ctx)
output = net(data)
predictions = nd.argmax(output, axis=1)
numerator += nd.sum(predictions == label)
denominator += data.shape[0]
return (numerator / denominator).asscalar()
```
## Execute the training loop
```
epochs = 10
learning_rate = .001
smoothing_constant = .01
for e in range(epochs):
cumulative_loss = 0
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(model_ctx).reshape((-1, 784))
label = label.as_in_context(model_ctx)
label_one_hot = nd.one_hot(label, 10)
with autograd.record():
output = net(data)
loss = softmax_cross_entropy(output, label_one_hot)
loss.backward()
SGD(params, learning_rate)
cumulative_loss += nd.sum(loss).asscalar()
test_accuracy = evaluate_accuracy(test_data, net)
train_accuracy = evaluate_accuracy(train_data, net)
print("Epoch %s. Loss: %s, Train_acc %s, Test_acc %s" %
(e, cumulative_loss/num_examples, train_accuracy, test_accuracy))
```
## Using the model for prediction
Let's pick a few random data points from the test set to visualize algonside our predictions. We already know quantitatively that the model is more accurate, but visualizing results is a good practice that can (1) help us to sanity check that our code is actually working and (2) provide intuition about what kinds of mistakes our model tends to make.
```
%matplotlib inline
import matplotlib.pyplot as plt
# Define the function to do prediction
def model_predict(net,data):
output = net(data)
return nd.argmax(output, axis=1)
samples = 10
mnist_test = mx.gluon.data.vision.MNIST(train=False, transform=transform)
# let's sample 10 random data points from the test set
sample_data = mx.gluon.data.DataLoader(mnist_test, samples, shuffle=True)
for i, (data, label) in enumerate(sample_data):
data = data.as_in_context(model_ctx)
im = nd.transpose(data,(1,0,2,3))
im = nd.reshape(im,(28,10*28,1))
imtiles = nd.tile(im, (1,1,3))
plt.imshow(imtiles.asnumpy())
plt.show()
pred=model_predict(net,data.reshape((-1,784)))
print('model predictions are:', pred)
print('true labels :', label)
break
```
## Conclusion
Nice! With just two hidden layers containing 256 hidden nodes, respectively, we can achieve over 95% accuracy on this task.
## Next
[Multilayer perceptrons with gluon](../chapter03_deep-neural-networks/mlp-gluon.ipynb)
For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| github_jupyter |
# Differential expression analysis
This notebook performs differential expression analysis using the real template experiment and simulated experiments, as a null set.
```
%load_ext autoreload
%load_ext rpy2.ipython
%autoreload 2
import os
import sys
import pandas as pd
import numpy as np
import random
import seaborn as sns
import rpy2.robjects
from rpy2.robjects.packages import importr
from rpy2.robjects import pandas2ri
pandas2ri.activate()
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
def fxn():
warnings.warn("deprecated", DeprecationWarning)
with warnings.catch_warnings():
warnings.simplefilter("ignore")
fxn()
from ponyo import utils
from numpy.random import seed
randomState = 123
seed(randomState)
# Read in config variables
base_dir = os.path.abspath(os.path.join(os.getcwd(),"../"))
config_file = os.path.abspath(os.path.join(base_dir,
"config_human.tsv"))
params = utils.read_config(config_file)
# Load params
local_dir = params["local_dir"]
dataset_name = params['dataset_name']
num_runs = params['num_simulated']
project_id = params['project_id']
col_to_rank = params['col_to_rank']
template_data_file = params['template_data_file']
rerun_template = True
rerun_simulated = True
# Load metadata file with grouping assignments for samples
metadata_file = os.path.join(
base_dir,
dataset_name,
"data",
"metadata",
project_id+"_groups.tsv")
```
## Install R libraries
```
%%R
# Select 59
# Run one time
#if (!requireNamespace("BiocManager", quietly = TRUE))
# install.packages("BiocManager")
#BiocManager::install("limma")
#BiocManager::install('EnhancedVolcano')
#devtools::install_github('kevinblighe/EnhancedVolcano')
%%R
library('limma')
```
### Get differentially expressed genes from template experiment
```
%%R -i metadata_file -i project_id -i template_data_file -i local_dir -i rerun_template
source('../generic_expression_patterns_modules/DE_analysis.R')
out_file <- paste(local_dir,
"DE_stats/DE_stats_template_data_",
project_id,
"_real.txt",
sep="")
if (rerun_template){
get_DE_stats(metadata_file,
project_id,
template_data_file,
"template",
local_dir,
"real")
}
```
### Check signal strength
```
# Load association statistics for template experiment
template_DE_stats_file = os.path.join(
local_dir,
"DE_stats",
"DE_stats_template_data_"+project_id+"_real.txt")
template_DE_stats = pd.read_csv(
template_DE_stats_file,
header=0,
sep='\t',
index_col=0)
template_DEGs = template_DE_stats[(template_DE_stats['adj.P.Val']<0.001) &
(template_DE_stats['logFC'].abs()>1)]
print(template_DEGs.shape)
template_DEGs.head(10)
%%R
library(EnhancedVolcano)
%%R -i project_id -i template_DE_stats_file -i local_dir
source('../generic_expression_patterns_modules/DE_analysis.R')
create_volcano(template_DE_stats_file,
project_id,
"adj.P.Val",
local_dir)
```
### Get differentially expressed genes from each simulated experiment
```
%%R -i metadata_file -i project_id -i base_dir -i local_dir -i num_runs -i rerun_simulated -o num_sign_DEGs_simulated
source('../generic_expression_patterns_modules/DE_analysis.R')
num_sign_DEGs_simulated <- c()
for (i in 0:(num_runs-1)){
simulated_data_file <- paste(local_dir,
"pseudo_experiment/selected_simulated_data_",
project_id,
"_",
i,
".txt",
sep="")
out_file <- paste(local_dir,
"DE_stats/DE_stats_simulated_data_",
project_id,
"_",
i,
".txt",
sep="")
if (rerun_simulated){
run_output <- get_DE_stats(metadata_file,
project_id,
simulated_data_file,
"simulated",
local_dir,
i)
num_sign_DEGs_simulated <- c(num_sign_DEGs_simulated, run_output)
} else {
# Read in DE stats data
DE_stats_data <- as.data.frame(read.table(out_file, sep="\t", header=TRUE, row.names=1))
# Get number of genes that exceed threshold
threshold <- 0.001
sign_DEGs <- DE_stats_data[DE_stats_data[,'adj.P.Val']<threshold & abs(DE_stats_data[,'logFC'])>1,]
num_sign_DEGs <- nrow(sign_DEGs)
num_sign_DEGs_simulated <- c(num_sign_DEGs_simulated, num_sign_DEGs)
}
}
# Plot distribution of differentially expressed genes for simulated experiments
ax = sns.distplot(num_sign_DEGs_simulated,
kde=False)
ax.set(xlabel='Number of DEGs', ylabel="Number of simulated experiments")
```
**Check**
As a check, we compared the number of DEGs identified here versus what was reported in the [Kim et. al. publication](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3566005/#pone.0055596.s008), which found:
* Four conditions needed to be met for the genes to be selected as differentially expressed genes (DEGs): (i) overall differential expression from the edgeR analysis with FDR < 0.001, (ii) a minimum of 3 patients with significant differential expression, as tested by edgeR for individual differential expression with FDR < 0.01, (iii) consistent up/down regulation among different patients representing more than a two-fold change, and (iv) significant expression in at least 3 patients to remove genes with large fold changes within the noise expression level (FVKM>2 in either normal or tumor tissue).
* In total, we selected 1459 genes (543 upregulated and 916 downregulated in tumors) differentially expressed in female NSCLC never-smoker patients
* Used edgeR to identify DEGs
By comparison:
* Our study found 2358 DEGs using limma and applying FDR < 0.001
* Spot checking the genes identified with their list of DEGs from S2, we found the some of the same genes and FC direction was consistent.
* Currently we are normalizing read counts [downloaded from recount2](https://bioconductor.org/packages/devel/bioc/vignettes/recount/inst/doc/recount-quickstart.html) using RPKM and piping that through limma to identify DEG (this is legacy code from when we expected microarray input instead of RNA-seq)
## Get statistics for differential expression analysis
```
# Get ranks of template experiment
# If ranking by p-value or adjusted p-value then high rank = low value
if col_to_rank in ['P.Value', 'adj.P.Val']:
template_DE_stats['ranking'] = template_DE_stats[col_to_rank].rank(ascending = False)
template_DE_stats = template_DE_stats.sort_values(by=col_to_rank, ascending=True)
# If ranking by logFC then high rank = high abs(value)
elif col_to_rank in ['logFC','t']:
template_DE_stats['ranking'] = template_DE_stats[col_to_rank].abs().rank(ascending = True)
template_DE_stats = template_DE_stats.sort_values(by=col_to_rank, ascending=False)
# If ranking by Z-score then high rank = high value
else:
template_DE_stats['ranking'] = template_DE_stats[col_to_rank].rank(ascending = True)
template_DE_stats = template_DE_stats.sort_values(by=col_to_rank, ascending=False)
template_DE_stats.head()
# Concatenate simulated experiments
simulated_DE_stats_all = pd.DataFrame()
for i in range(num_runs):
simulated_DE_stats_file = os.path.join(
local_dir,
"DE_stats",
"DE_stats_simulated_data_"+project_id+"_"+str(i)+".txt")
#Read results
simulated_DE_stats = pd.read_csv(
simulated_DE_stats_file,
header=0,
sep='\t',
index_col=0)
simulated_DE_stats.reset_index(inplace=True)
# Concatenate df
simulated_DE_stats_all = pd.concat([simulated_DE_stats_all,
simulated_DE_stats])
print(simulated_DE_stats_all.shape)
simulated_DE_stats_all.head()
# Aggregate statistics across all simulated experiments
if col_to_rank == "adj.P.Val":
simulated_DE_summary_stats = simulated_DE_stats_all.groupby(['index'])[[col_to_rank]].agg(
['median','mean', 'std','count']
)
else:
simulated_DE_summary_stats = simulated_DE_stats_all.groupby(['index'])[[col_to_rank, 'adj.P.Val']].agg({
col_to_rank:['median','mean', 'std','count'],
'adj.P.Val':['median']
})
simulated_DE_summary_stats.head()
# Rank gene by median value of col_to_rank for simulated experiments
# If ranking by p-value or adjusted p-value then high rank = low value
if col_to_rank in ['P.Value', 'adj.P.Val']:
simulated_DE_summary_stats['ranking'] = simulated_DE_summary_stats[(col_to_rank,'median')].rank(ascending = False)
simulated_DE_summary_stats = simulated_DE_summary_stats.sort_values(by=(col_to_rank,'median'), ascending=True)
# If ranking by logFC then high rank = high abs(value)
elif col_to_rank in ['logFC','t']:
simulated_DE_summary_stats['ranking'] = simulated_DE_summary_stats[(col_to_rank,'median')].abs().rank(ascending = True)
simulated_DE_summary_stats = simulated_DE_summary_stats.sort_values(by=(col_to_rank,'median'), ascending=False)
# If ranking by Z-score then high rank = high value
else:
simulated_DE_summary_stats['ranking'] = simulated_DE_summary_stats[(col_to_rank,'median')].rank(ascending = True)
simulated_DE_summary_stats = simulated_DE_summary_stats.sort_values(by=(col_to_rank,'median'), ascending=False)
simulated_DE_summary_stats.head()
simulated_DE_summary_stats.tail()
# Merge template statistics with simulated statistics
template_simulated_DE_stats = template_DE_stats.merge(simulated_DE_summary_stats,
left_index=True,
right_index=True)
print(template_simulated_DE_stats.shape)
template_simulated_DE_stats.head()
ax = sns.distplot(template_simulated_DE_stats[('ranking',"")].values, kde=False)
ax.set(xlabel='Ranking', ylabel="Number of genes")
# Parse columns
median_pval_simulated = template_simulated_DE_stats[('adj.P.Val','median')]
mean_test_simulated = template_simulated_DE_stats[(col_to_rank,'mean')]
std_test_simulated = template_simulated_DE_stats[(col_to_rank,'std')]
count_simulated = template_simulated_DE_stats[(col_to_rank,'count')]
rank_simulated = template_simulated_DE_stats[('ranking','')]
summary = pd.DataFrame(data={'Gene ID': template_simulated_DE_stats.index,
'Adj P-value (Real)': template_simulated_DE_stats['adj.P.Val'],
'Rank (Real)': template_simulated_DE_stats['ranking'],
'Test statistic (Real)': template_simulated_DE_stats[col_to_rank],
'Median adj p-value (simulated)': median_pval_simulated ,
'Rank (simulated)': rank_simulated ,
'Mean test statistic (simulated)': mean_test_simulated ,
'Std deviation (simulated)': std_test_simulated,
'Number of experiments (simulated)': count_simulated
}
)
summary['Z score'] = (summary['Test statistic (Real)'] - summary['Mean test statistic (simulated)'])/summary['Std deviation (simulated)']
summary.head()
# Save file
summary_file = os.path.join(
local_dir,
"gene_summary_table_"+col_to_rank+".tsv")
#summary.to_csv(summary_file, float_format='%.5f', sep='\t')
```
| github_jupyter |
foo.038 Nutrient Application for Major Crops
http://www.earthstat.org/data-download/
File type: geotiff
not sure which one to include - nitrogen application rate, total, and quality; phosphorus application rate, total and quality ; potassium application rate and total and quality for each crop
```
# Libraries for downloading data from remote server (may be ftp)
import requests
from urllib.request import urlopen
from contextlib import closing
import shutil
# Library for uploading/downloading data to/from S3
import boto3
# Libraries for handling data
import rasterio as rio
import numpy as np
# from netCDF4 import Dataset
# import pandas as pd
# import scipy
# Libraries for various helper functions
# from datetime import datetime
import os
import threading
import sys
from glob import glob
```
s3
```
s3_upload = boto3.client("s3")
s3_download = boto3.resource("s3")
s3_bucket = "wri-public-data"
s3_folder = "resourcewatch/raster/soc_030_gross_domestic_product/"
s3_file1 = "soc_030_mer_1990_sum.asc"
s3_file2 = "soc_030_mer_1995_sum.asc"
s3_file3 = "soc_030_mer_2000_sum.asc"
s3_file4 = "soc_030_mer_2005_sum.asc"
s3_file5 = "soc_030_ppp_1990_sum.asc"
s3_file6 = "soc_030_ppp_1995_sum.asc"
s3_file7 = "soc_030_ppp_2000_sum.asc"
s3_file8 = "soc_030_ppp_2005_sum.asc"
s3_key_orig1 = s3_folder + s3_file1
s3_key_edit1 = s3_key_orig1[0:-4] + "_edit.tif"
s3_key_orig2 = s3_folder + s3_file2
s3_key_edit2 = s3_key_orig2[0:-4] + "_edit.tif"
s3_key_orig3 = s3_folder + s3_file3
s3_key_edit3 = s3_key_orig3[0:-4] + "_edit.tif"
s3_key_orig4 = s3_folder + s3_file4
s3_key_edit4 = s3_key_orig4[0:-4] + "_edit.tif"
s3_key_orig5 = s3_folder + s3_file5
s3_key_edit5 = s3_key_orig5[0:-4] + "_edit.tif"
s3_key_orig6= s3_folder + s3_file6
s3_key_edit6 = s3_key_orig6[0:-4] + "_edit.tif"
s3_key_orig7 = s3_folder + s3_file7
s3_key_edit7 = s3_key_orig7[0:-4] + "_edit.tif"
s3_key_orig8 = s3_folder + s3_file8
s3_key_edit8 = s3_key_orig8[0:-4] + "_edit.tif"
class ProgressPercentage(object):
def __init__(self, filename):
self._filename = filename
self._size = float(os.path.getsize(filename))
self._seen_so_far = 0
self._lock = threading.Lock()
def __call__(self, bytes_amount):
# To simplify we'll assume this is hooked up
# to a single filename.
with self._lock:
self._seen_so_far += bytes_amount
percentage = (self._seen_so_far / self._size) * 100
sys.stdout.write("\r%s %s / %s (%.2f%%)"%(
self._filename, self._seen_so_far, self._size,
percentage))
sys.stdout.flush()
```
Define local file locations
```
local_folder = "/Users/Max81007/Desktop/Python/Resource_Watch/Raster/soc_030/"
file_name1 = "mer1990sum.asc"
file_name2 = "mer1995sum.asc"
file_name3 = "mer2000sum.asc"
file_name4 = "mer2005sum.asc"
file_name5 = "ppp1990sum.asc"
file_name6 = "ppp1995sum.asc"
file_name7 = "ppp2000sum.asc"
file_name8 = "ppp2005sum.asc"
local_orig1 = local_folder + file_name1
local_orig2 = local_folder + file_name2
local_orig3 = local_folder + file_name3
local_orig4 = local_folder + file_name4
local_orig5 = local_folder + file_name5
local_orig6 = local_folder + file_name6
local_orig7 = local_folder + file_name7
local_orig8 = local_folder + file_name8
orig_extension_length = 4 #4 for each char in .tif
local_edit1 = local_orig1[:-orig_extension_length] + "edit.tif"
local_edit2 = local_orig2[:-orig_extension_length] + "edit.tif"
local_edit3 = local_orig3[:-orig_extension_length] + "edit.tif"
local_edit4 = local_orig4[:-orig_extension_length] + "edit.tif"
local_edit5 = local_orig5[:-orig_extension_length] + "edit.tif"
local_edit6 = local_orig6[:-orig_extension_length] + "edit.tif"
local_edit7 = local_orig7[:-orig_extension_length] + "edit.tif"
local_edit8 = local_orig8[:-orig_extension_length] + "edit.tif"
```
Use rasterio to reproject and compress
```
files = [local_orig1, local_orig2]
for file in files:
with rio.open(file, 'r') as src:
profile = src.profile
print(profile)
# Note - this is the core of Vizz's netcdf2tif function
def convert_asc_to_tif(orig_name, edit_name):
with rio.open(orig_name, 'r') as src:
# This assumes data is readable by rasterio
# May need to open instead with netcdf4.Dataset, for example
data = src.read()[0]
rows = data.shape[0]
columns = data.shape[1]
print(rows)
print(columns)
# Latitude bounds
south_lat = -90
north_lat = 90
# Longitude bounds
west_lon = -180
east_lon = 180
transform = rio.transform.from_bounds(west_lon, south_lat, east_lon, north_lat, columns, rows)
# Profile
no_data_val = -9999.0
target_projection = 'EPSG:4326'
target_data_type = np.float64
profile = {
'driver':'GTiff',
'height':rows,
'width':columns,
'count':1,
'dtype':target_data_type,
'crs':target_projection,
'transform':transform,
'compress':'lzw',
'nodata': no_data_val
}
with rio.open(edit_name, "w", **profile) as dst:
dst.write(data.astype(profile["dtype"]), 1)
convert_asc_to_tif(local_orig1, local_edit1)
convert_asc_to_tif(local_orig2, local_edit2)
convert_asc_to_tif(local_orig3, local_edit3)
convert_asc_to_tif(local_orig4, local_edit4)
convert_asc_to_tif(local_orig5, local_edit5)
convert_asc_to_tif(local_orig6, local_edit6)
convert_asc_to_tif(local_orig7, local_edit7)
convert_asc_to_tif(local_orig8, local_edit8)
```
Upload orig and edit files to s3
```
# Original
s3_upload.upload_file(local_orig1, s3_bucket, s3_key_orig1,
Callback=ProgressPercentage(local_orig1))
s3_upload.upload_file(local_orig2, s3_bucket, s3_key_orig2,
Callback=ProgressPercentage(local_orig2))
s3_upload.upload_file(local_orig3, s3_bucket, s3_key_orig3,
Callback=ProgressPercentage(local_orig3))
s3_upload.upload_file(local_orig4, s3_bucket, s3_key_orig4,
Callback=ProgressPercentage(local_orig4))
s3_upload.upload_file(local_orig5, s3_bucket, s3_key_orig5,
Callback=ProgressPercentage(local_orig5))
s3_upload.upload_file(local_orig6, s3_bucket, s3_key_orig6,
Callback=ProgressPercentage(local_orig6))
s3_upload.upload_file(local_orig7, s3_bucket, s3_key_orig7,
Callback=ProgressPercentage(local_orig7))
s3_upload.upload_file(local_orig8, s3_bucket, s3_key_orig8,
Callback=ProgressPercentage(local_orig8))
# Edit
s3_upload.upload_file(local_edit1, s3_bucket, s3_key_edit1,
Callback=ProgressPercentage(local_edit1))
s3_upload.upload_file(local_edit2, s3_bucket, s3_key_edit2,
Callback=ProgressPercentage(local_edit2))
s3_upload.upload_file(local_edit3, s3_bucket, s3_key_edit3,
Callback=ProgressPercentage(local_edit3))
s3_upload.upload_file(local_edit4, s3_bucket, s3_key_edit4,
Callback=ProgressPercentage(local_edit4))
s3_upload.upload_file(local_edit5, s3_bucket, s3_key_edit5,
Callback=ProgressPercentage(local_edit5))
s3_upload.upload_file(local_edit6, s3_bucket, s3_key_edit6,
Callback=ProgressPercentage(local_edit6))
s3_upload.upload_file(local_edit7, s3_bucket, s3_key_edit7,
Callback=ProgressPercentage(local_edit7))
s3_upload.upload_file(local_edit8, s3_bucket, s3_key_edit8,
Callback=ProgressPercentage(local_edit8))
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
train_data = pd.read_csv("processed/train_data_filled.csv")
test_data = pd.read_csv("processed/test_data_filled.csv")
x_train = train_data.drop('traffic_volume', axis=1)
y_train = train_data['traffic_volume']
x_test = test_data
all_data = pd.concat([x_train, x_test])
```
Handle time
```
all_data['timestamp'] = pd.to_datetime(all_data['timestamp'])
all_data['weekday'] = all_data['timestamp'].dt.weekday
all_data['month'] = all_data['timestamp'].dt.month
all_data['hour'] = all_data['timestamp'].dt.hour
all_data = all_data.drop(['timestamp'], axis=1)
x_train = all_data[:len(x_train)]
x_test = all_data[len(x_train):]
print(x_train.info())
print(x_test.info())
```
Head map
```
x_y_train = x_train.copy(deep=True)
x_y_train['traffic_volume'] = train_data['traffic_volume']
corr = x_y_train.corr()
#Correlation with output variable
corr_target = abs(corr["traffic_volume"])
top_corr_features = corr_target.index
plt.figure(figsize=(10,8))
#plot heat map
heatmap=sns.heatmap(x_y_train[top_corr_features].corr(), annot=True)
plt.figure(figsize=(16, 10))
holiday = sns.stripplot(x=y_train, y="holiday", data=x_train, size=10)
plt.figure(figsize=(16, 10))
weather = sns.stripplot(x=y_train, y="weather", data=x_train, size=10)
plt.figure(figsize=(16, 10))
weather_detail = sns.stripplot(x=y_train, y="weather_detail", data=x_train, size=10)
x_train.hist(bins=50, figsize=(15,10))
plt.show()
size_resolution = 300
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax2.plot(x_train.weekday[0:size_resolution],'r')
ax1.plot(y_train[0:size_resolution])
fig.tight_layout() # otherwise the right y-label is slightly clipped
fig.set_size_inches(18.5, 10.5)
```
One-hot encoding
```
holiday_dummy = pd.get_dummies(all_data['holiday'])
weather_dummy = pd.get_dummies(all_data['weather'])
weather_detail_dummy = pd.get_dummies(all_data['weather_detail'])
all_data.drop('holiday', axis=1, inplace=True)
all_data.drop('weather', axis=1, inplace=True)
all_data.drop('weather_detail', axis=1, inplace=True)
scaler = StandardScaler()
scaler.fit(all_data)
all_data = scaler.transform(all_data)
all_data = np.hstack((all_data, holiday_dummy, weather_dummy, weather_detail_dummy))
x_train = all_data[:len(x_train)]
x_test = all_data[len(x_train):]
```
PCA analysis
```
pca = PCA().fit(x_train)
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('Number of components')
plt.ylabel('Cumulative explained variance ratio')
# from sklearn.ensemble import RandomForestRegressor
from xgboost import XGBRegressor
from sklearn.model_selection import cross_val_score
# regressor = RandomForestRegressor()
regressor = XGBRegressor(n_estimators=600, max_depth=5)
scores = cross_val_score(regressor, x_train, y_train, scoring="neg_mean_absolute_error", cv=10)
mae_scores = scores
# mae_scores = -scores
print("MAE ")
print("Scores:", mae_scores)
print("Mean:", mae_scores.mean())
print("Standard deviation:", mae_scores.std())
from sklearn.model_selection import cross_val_predict
predictions = cross_val_predict(regressor, x_train, y_train, cv=10)
size_resolution = 500
# fig, ax1 = plt.subplots()
# ax2 = ax1.twinx()
# ax2.plot( predictions[0:size_resolution],'r')
# ax1.plot(y_train[0:size_resolution],'-')
# fig.tight_layout() # otherwise the right y-label is slightly clipped
# fig.set_size_inches(18.5, 10.5)
plt.xlabel('dataset')
plt.ylabel('traffic_volume')
plt.plot(predictions[0:size_resolution], label='Prediction')
plt.plot(y_train[0:size_resolution], label='Real value')
plt.legend()
plt.show()
plt.xlabel('Prediction')
plt.ylabel('Real data')
plt.plot(predictions, y_train, 'o',markersize=1)
predictor = regressor.fit(x_train, y_train)
y_predict = predictor.predict(x_test)
np.savetxt('result.txt', y_predict, fmt='%d')
```
| github_jupyter |
```
!date
import warnings
import numpy as np
import pandas as pd
import pymc3 as pm
import arviz as az
import theano.tensor as tt
import matplotlib.pyplot as plt
%matplotlib inline
warnings.simplefilter(action="ignore", category=FutureWarning)
df_all = pd.read_csv("13-14.txt", sep="\t")
team_names = df_all["Home \ Away "].str.strip().copy(deep=True).values
team_abbrs = df_all.columns.str.strip().copy(deep=True).values[1:]
df_all = df_all.melt(id_vars=["Home \ Away "])
df_all.columns = ["home_team", "away_team", "result"]
df_all = df_all.apply(lambda x: x.str.strip(), axis="index")
df_all = df_all.replace(
{
"away_team": {abbr: team for abbr, team in zip(team_abbrs, team_names)},
"result": {"—": np.nan}
}
).dropna(subset=["result"]).reset_index(drop=True)
df_aux = df_all.result.str.split("–", expand=True)
df_aux.columns = ["home_goals", "away_goals"]
df_final = pd.concat((df_all[["home_team", "away_team"]], df_aux), axis="columns")
df_final.to_csv("premier_13-14.csv", index=False)
df = pd.read_csv("premier_13-14.csv")
num_teams = df.home_team.nunique()
with pm.Model() as m_general:
# constant data
home_team = pm.intX(pm.Data("home_team", pd.factorize(df.home_team, sort=True)[0]))
away_team = pm.intX(pm.Data("away_team", pd.factorize(df.away_team, sort=True)[0]))
# global model parameters
home = pm.Normal('home', mu=0, sigma=5)
sd_att = pm.HalfStudentT('sd_att', nu=3, sigma=2.5)
sd_def = pm.HalfStudentT('sd_def', nu=3, sigma=2.5)
intercept = pm.Normal('intercept', mu=0, sigma=5)
# team-specific model parameters
atts_star = pm.Normal("atts_star", mu=0, sigma=sd_att, shape=num_teams)
defs_star = pm.Normal("defs_star", mu=0, sigma=sd_def, shape=num_teams)
atts = pm.Deterministic('atts', atts_star - tt.mean(atts_star))
defs = pm.Deterministic('defs', defs_star - tt.mean(defs_star))
home_theta = tt.exp(intercept + home + atts[home_team] + defs[away_team])
away_theta = tt.exp(intercept + atts[away_team] + defs[home_team])
# likelihood of observed data
home_goals = pm.Poisson('home_goals', mu=home_theta, observed=df.home_goals)
away_goals = pm.Poisson('away_goals', mu=away_theta, observed=df.away_goals)
dims = {
"home_goals": ["match"],
"away_goals": ["match"],
"home_team": ["match"],
"away_team": ["match"],
"atts": ["team"],
"atts_star": ["team"],
"defs": ["team"],
"defs_star": ["team"],
}
coords = {"team": pd.factorize(df.home_team, sort=True)[1]}
with m_general:
trace = pm.sample(random_seed=1375)
idata_general = az.from_pymc3(trace, coords=coords, dims=dims)
idata_general
with pm.Model() as m_match:
# constant data
home_team = pm.intX(pm.Data("home_team", pd.factorize(df.home_team, sort=True)[0]))
away_team = pm.intX(pm.Data("away_team", pd.factorize(df.away_team, sort=True)[0]))
# global model parameters
home = pm.Normal('home', mu=0, sigma=5)
sd_att = pm.HalfStudentT('sd_att', nu=3, sigma=2.5)
sd_def = pm.HalfStudentT('sd_def', nu=3, sigma=2.5)
intercept = pm.Normal('intercept', mu=0, sigma=5)
# team-specific model parameters
atts_star = pm.Normal("atts_star", mu=0, sigma=sd_att, shape=num_teams)
defs_star = pm.Normal("defs_star", mu=0, sigma=sd_def, shape=num_teams)
atts = pm.Deterministic('atts', atts_star - tt.mean(atts_star))
defs = pm.Deterministic('defs', defs_star - tt.mean(defs_star))
home_theta = tt.exp(intercept + home + atts[home_team] + defs[away_team])
away_theta = tt.exp(intercept + atts[away_team] + defs[home_team])
# alternative likelihood of observed data
def double_poisson(h, a):
return (
pm.Poisson.dist(mu=home_theta).logp(h) +
pm.Poisson.dist(mu=away_theta).logp(a)
)
matches = pm.DensityDist('matches', double_poisson, observed={'h': df.home_goals, 'a': df.away_goals})
dims = {
"goals": ["match"],
"home_team": ["match"],
"away_team": ["match"],
"atts": ["team"],
"atts_star": ["team"],
"defs": ["team"],
"defs_star": ["team"],
}
coords = {"team": pd.factorize(df.home_team, sort=True)[1]}
with m_match:
trace = pm.sample(random_seed=1375, tune=1500)
idata_match = az.from_pymc3(trace, coords=coords, dims=dims)
idata_match
az.loo(idata_match)
idata_general.log_likelihood["match_lik"] = (
idata_general.log_likelihood.home_goals + idata_general.log_likelihood.away_goals
)
az.loo(idata_general, var_name="match_lik")
df_home = df[["home_team", "away_team", "home_goals"]].rename(columns={"home_goals": "goals"})
df_home["visitor"] = 0
df_away = df[["home_team", "away_team", "away_goals"]].rename(columns={"away_goals": "goals"})
df_away["visitor"] = 1
df_goals = pd.concat((df_home, df_away), axis="index")
df_goals
with pm.Model() as m_goals:
# constant data
home_team = pm.intX(pm.Data("home_team", pd.factorize(df_goals.home_team, sort=True)[0]))
away_team = pm.intX(pm.Data("away_team", pd.factorize(df_goals.away_team, sort=True)[0]))
is_visitor = pm.intX(pm.Data("is_visitor", df_goals.visitor))
# global model parameters
home = pm.Normal('home', mu=0, sigma=5)
sd_att = pm.HalfStudentT('sd_att', nu=3, sigma=2.5)
sd_def = pm.HalfStudentT('sd_def', nu=3, sigma=2.5)
intercept = pm.Normal('intercept', mu=0, sigma=5)
# team-specific model parameters
atts_star = pm.Normal("atts_star", mu=0, sigma=sd_att, shape=num_teams)
defs_star = pm.Normal("defs_star", mu=0, sigma=sd_def, shape=num_teams)
atts = pm.Deterministic('atts', atts_star - tt.mean(atts_star))
defs = pm.Deterministic('defs', defs_star - tt.mean(defs_star))
home_aux = tt.stack([home, 0])
pars_aux = tt.stack([defs, atts])
theta = tt.exp(
intercept + home_aux[is_visitor] + pars_aux[(is_visitor, away_team)] + pars_aux[(1-is_visitor, home_team)]
)
# alternative likelihood of observed data
goals = pm.Poisson('goals', mu=theta, observed=df_goals.goals.values)
dims = {
"goals": ["idx"],
"home_team": ["idx"],
"away_team": ["idx"],
"atts": ["team"],
"atts_star": ["team"],
"defs": ["team"],
"defs_star": ["team"],
}
coords = {"team": pd.factorize(df_goals.home_team, sort=True)[1]}
with m_goals:
trace = pm.sample(random_seed=1375, tune=1500)
idata_goals = az.from_pymc3(trace, coords=coords, dims=dims)
idata_goals
az.loo(idata_goals)
import xarray as xr
idata_general.log_likelihood["goals_lik"] = xr.concat(
(idata_general.log_likelihood.home_goals, idata_general.log_likelihood.away_goals),
"match"
).rename({"match": "observation"})
az.loo(idata_general, var_name="goals_lik")
az.plot_density(
(idata_general, idata_goals, idata_match),
var_names=["home", "intercept", "atts_star", "defs_star"],
);
```
| github_jupyter |
# Tutorial in chapter 09 - petastorm-pyspark-pytorch
### 1. load parquet data into pytorch loader
file path: `notebooks/images_data/silver/augmented`
```
# spark
from pyspark.sql.functions import lit
from pyspark.sql.types import BinaryType,StringType
from pyspark.sql import SparkSession
from pyspark.sql.functions import col
import pyspark.sql.functions
from pyspark.sql.types import *
#petastorm
from petastorm.spark import SparkDatasetConverter, make_spark_converter
from petastorm import TransformSpec
import io
import numpy as np
from PIL import Image
from functools import partial
# train images with pytorch
#from torchvision import transforms
from hyperopt import fmin, tpe, hp, SparkTrials, STATUS_OK
import torch
import torch.optim as optim
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
# import mlflow
import mlflow
import mlflow.pytorch
# start Spark session:
spark = SparkSession \
.builder \
.appName("Distributed Pytorch training") \
.config("spark.memory.offHeap.enabled",True) \
.config("spark.memory.offHeap.size","30g")\
.getOrCreate()
from petastorm.spark import SparkDatasetConverter, make_spark_converter
spark.conf.set(SparkDatasetConverter.PARENT_CACHE_DIR_URL_CONF, 'petastorm_cache')
data_path = "images_data/silver/augmented"
mlflow_model_dir_path = "/"
```
# Enable MLFlow tracking
```
import pytorch_lightning as pl
import torch
from torch.nn import functional as F
from torch.utils.data import DataLoader
from torchvision import transforms
from torchvision.datasets import MNIST
try:
from torchmetrics.functional import accuracy
except ImportError:
from pytorch_lightning.metrics.functional import accuracy
#Enable MLFlow tracking
mlflow.set_experiment(mlflow_model_dir_path)
# requires pytorch_lightning
mlflow.pytorch.autolog()
```
## params
```
IMG_SHAPE = (224, 224, 3)
BATCH_SIZE = 5
#The number of **epochs** is a hyperparameter that defines the number times that the learning algorithm will work through the entire training dataset. One epoch means that each sample in the training dataset has had an opportunity to update the internal model parameters.
SAMPLE_SIZE = 50
NUM_EPOCHS = 1
NUM_EXECUTERS = 1
```
## 2. Load preprocessed data
```
# Read the training data stored in parquet, limiting the dataset for the example
df_parquet = spark.read.parquet(data_path)
df = df_parquet.select(col("content"), col("label_index").cast(LongType())).limit(SAMPLE_SIZE)
num_classes = df.select("label_index").distinct().count()
num_classes =4
```
## 3. Split to train and test
```
df_train , df_val = df.randomSplit([0.6,0.4], seed=12345)
```
## 4. Cache the Spark DataFrame using Petastorm Spark Converter
```
tmp_path = "file:/home/jovyan/petastorm_cache/"
# Set a cache directory on DBFS FUSE for intermediate data
spark.conf.set(SparkDatasetConverter.PARENT_CACHE_DIR_URL_CONF,tmp_path)
# TIP: Use a low value for parquet_row_group_bytes. The detafault of 32 MiB can be too high for larger datasets. Using 1MB instead.
#train
converter_train = make_spark_converter(df_train, parquet_row_group_size_bytes=32000000)
#test
converter_val = make_spark_converter(df_val, parquet_row_group_size_bytes=32000000)
```
### Petastorm prepreocess
used during materlizing spark dataframe with petastorm and bridging to TensorFlow
```
import torchvision, torch
from torchvision import datasets, models, transforms
def preprocess(grayscale_image):
"""
Preprocess an image file bytes for MobileNetV2 (ImageNet).
"""
image = Image.open(io.BytesIO(grayscale_image)).resize([224, 224])
image_array = np.array(image)
#image_array = keras.preprocessing.image.img_to_array(image)
return image_array
def transform_row(pd_batch):
"""
The input and output of this function are pandas dataframes.
"""
pd_batch['features'] = pd_batch['content'].map(lambda x: preprocess(x))
pd_batch = pd_batch.drop(labels=['content'], axis=1)
return pd_batch
# The output shape of the `TransformSpec` is not automatically known by petastorm,
# so you need to specify the shape for new columns in `edit_fields` and specify the order of
# the output columns in `selected_fields`.
transform_spec_fn = TransformSpec(
func=transform_row,
edit_fields=[('features', np.uint8 , IMG_SHAPE, False)],
selected_fields=['features', 'label_index']
)
```
## 5. Get the model MobileNetV2
#### Get the model MobileNetV2 from torch hub
and only retraint it's final layer to fit our needs.
```
model = torch.hub.load('pytorch/vision:v0.10.0', 'mobilenet_v2', pretrained=True)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 32, 3, 1)
self.conv2 = nn.Conv2d(32, 64, 3, 1)
self.dropout1 = nn.Dropout2d(0.25)
self.dropout2 = nn.Dropout2d(0.5)
self.fc1 = nn.Linear(9216, 128)
self.fc2 = nn.Linear(128, 10)
def forward(self, x): # pylint: disable=arguments-differ
x = x.view((-1, 1, 28, 28))
x = self.conv1(x)
x = F.relu(x)
x = self.conv2(x)
x = F.relu(x)
x = F.max_pool2d(x, 2)
x = self.dropout1(x)
x = torch.flatten(x, 1)
x = self.fc1(x)
x = F.relu(x)
x = self.dropout2(x)
x = self.fc2(x)
output = F.log_softmax(x, dim=1)
return output
```
## 6. set the train function
```
def train(data_loader, steps=100, lr=0.0005, momentum=0.5):
model = Net()
model.train()
optimizer = optim.SGD(model.parameters(), lr=lr, momentum=momentum)
loss_hist = []
for batch_idx, batch in enumerate(data_loader):
if batch_idx > steps:
break
data, target = Variable(batch['features']), Variable(batch['label_index'])
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
if batch_idx % 10 == 0:
logging.info('[%d/%d]\tLoss: %.6f', batch_idx, steps, loss.data.item())
loss_hist.append(loss.data.item())
return model
def test(model, test_loader):
model.eval()
test_loss = 0
correct = 0
test_len = 0
with torch.no_grad():
for batch in test_loader:
data, target = batch['features'], batch['label']
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_len += data.shape[0]
test_loss /= test_len
accuracy = correct / test_len
logging.info('Test set: Average loss: %.4f, Accuracy: %d/%d (%.0f%%)',
test_loss, correct, test_len, 100. * accuracy)
return accuracy
def train_and_evaluate(_=None):
with converter_train.make_torch_dataloader(transform_spec=transform_spec_fn) as loader:
model = train(loader)
with converter_test.make_torch_dataloader(transform_spec=transform_spec_fn,num_epochs=1) as loader:
accuracy = test(model, loader)
return accuracy
accuracy = train_and_evaluate()
```
| github_jupyter |
# Community detection on the Wikipedia hyperlink graph
Team members:
* Armand Boschin
* Bojana Ranković
* Quentin Rebjock
In order to read this properly (especially the graph visualization in the end), you might want to used this [link](https://nbviewer.jupyter.org/github/armand33/wikipedia_graph/blob/master/wikipedia.ipynb?flush_cache=true) to NBViewer.
Please go through the README.md [file](https://github.com/armand33/wikipedia_graph/blob/master/README.md) beforehand.
**NB**:
* Please take the time to download the pickle file `shortest_paths.pkl` from this [link](https://drive.google.com/file/d/17bXr-OKY8xrUhCDfwR0WwP9uTOXeKG9d/view?usp=sharing). It should be put in the directory `data/`.
* Some cells are not executed and are in comments because they are really long. All the output files are directly dumped into pickle files. Those should be present in the directory `data/`.
- scraping step ($\sim 3$ hours) : `network.pkl`
- shortest-paths step ($\sim 5$ hours) : `shortest_paths.pkl`
- cross-validation step ($\sim 1$ hour) : `cross_val.pkl`
This is a reminder of the project proposal adapted to the reality of the project:
***Graph:*** Wikipedia hyperlink network
***Problem:*** Does the structure of the graph bears info on the content of the nodes ? We would like to find out if it is possible to detect communities of pages just by looking at the hyperlink connections and match these communities with real-world data such as categories of the pages. Is spectral clustering a viable possibility compared to proven method of community detection.
***Steps of the project:***
* Scraping the Wikipedia hyperlink network. Start from one node and get the pages as far as 2 or 3 hops depending on the number of nodes we get.
* Model the network by a random graph/scale-free network/something else in order to try to retrieve some of its characteristics.
* Apply Louvain algorithm for community detection to get a baseline to compare spectral clustering to. Indeed in term of community detection, Louvain is quite usual and our approach is to test the performance of spectral clustering.
* Try to apply spectral clustering in order to detect communities of pages.
* Visualize the clusters to match them with real-world categories (using some of the tools from the last guest lecture).
## Table of contents
* [**1) Data Acquisition**](#Data Acquisition)
<br><br>
* [**2) Data Exploration**](#Data Exploration)
<br><br>
* [**3) Data Exploitation**](#Data Exploitation)
- [3.1) Modelisation of the network](#Modelisation of the network)
- [*3.1.1) Exploration of the degree distribution*](#Exploration of the degree distribution)
- [*3.1.2) Modelisation with usual graphs models*](#Modelisation with usual graphs models)
- [*3.1.3) A power law network with the right exponent*](#A power law network with the right exponent)
- [*3.1.4 Comparison of the models*](#Comparison of the models)
- [*3.1.5) Comparison with common networks from the web*](#Comparison with common networks from the web) <br><br>
- [3.2) Community detection using Spectral clustering](#Community detection using Spectral clustering)
- [*3.2.1) Louvain community detection*](#Louvain community detection)
- [*3.2.2) Spectral Clustering*](#Spectral Clustering)
- [*3.2.3 Comparison of the two methods*](#Comparison of the two methods)
- [*3.2.4 Visualization*](#3.2.4Visualization)
<br><br>
* [**4) Conclusion**](#Conclusion)
```
import numpy as np
import seaborn as sns
import networkx as nx
import matplotlib.pyplot as plt
import operator
import community
import plotly
import plotly.graph_objs as go
import plotly.plotly as py
from networkx.drawing.nx_agraph import graphviz_layout
from scipy import linalg, cluster, sparse
from tqdm import tqdm_notebook
from utils import load_obj, save_obj
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
# 1) Data Acquisition <a class="anchor" id="Data Acquisition"></a>
We want to acquire a sub-network of the Wikipedia hyperlink network. In such a graph, each node is a Wikipedia page and there is a link between node $a$ and node $b$ if there is a link to page $b$ on page $a$. This is a directed network but we will make it undirected later on.
The process of the acquisition is the following :
* Start from an arbitrary root page (prefer an ambiguous page in order to get as many different communities as possible).
* Explore this page to get the intra-wiki links : the first nodes.
* For each first node, explore the intra-wiki links to get the second nodes.
* Look for inner connections (links from second nodes to first nodes tipically).
* Eventually, for each node, take the intersection of its neighbor with the collected nodes (it can be that some nodes have neighbors that have not been collected, for example a second node that has been collected from a disambiguation page).
We use the `Wikipedia` API that allows us to scrap pages and get links and categories for each one. We chose to include in our network only real pages (not the disambiguation ones). Those pages are indeed useful during the scraping because they allow us to get a larger sample of the real graph. Disambiguation pages act like bridges between pages that have nothing to do together.
For each node we need to get URL, title, categories and links to other pages.
We use as `root_node` the disambiguation page Jaguar (disambiguation) as it lists a really wide variety of themes (animals, cars, music, films, weapons...). It can lead us to a large selection of categories.
The function `explore_page` is implemented in the file `utils.py`. All implementation details are provided there.
```
from utils import explore_page
root_node = 'Jaguar (disambiguation)'
network = {} # This dict stores for each page a dictionnary containing the keys [url, links, categories]
first_nodes = []
explore_page(root_node, network, first_nodes)
second_nodes = []
for node in first_nodes:
explore_page(node, network, second_nodes)
```
Look for connections between second nodes and the rest of the nodes.
```
all_nodes = list(network.keys()) + second_nodes
for link in tqdm_notebook(second_nodes):
explore_page(link, network, [], inner=True, all_nodes=all_nodes)
```
The above cell took 2 hours and 47 minutes to run (duration of scraping).
Now we need to go through all the nodes in order to remove from their neighbors any page that has not been scraped.
```
all_nodes = list(network.keys())
for title in tqdm_notebook(network.keys()):
network[title]['links'] = list(set(network[title]['links']).intersection(set(all_nodes)))
```
The previous step can lead to pages with no neighbor (if a second node comes from a disambiguation page and all its neigbors are not in first or second nodes).
```
l = list(network.keys())
for i in l:
if len(network[i]['links']) == 0:
del network[i]
```
#### Cleaning the categories
There are some categories for each page that are irrelevant in our work (e.g. "All articles with unsourced statements"). We need to get rid of those.
```
for i, title in enumerate(network.keys()):
cats = network[title]['categories']
new_cats = []
for c in cats:
if not c.startswith('Redundant') and not c.startswith('Pages') and not c.startswith('Webarchive') and not c.startswith('Wikipedia') and not c.startswith('Articles') and not c.startswith('Coordinates on Wikidata') and not 'Wikidata' in c and not c.startswith('CS1') and not c.startswith('EngvarB') and not c.startswith('All') and not c.startswith('Good articles') and not c.startswith('Use dmy'):
new_cats.append(c)
network[title]['categories'] = new_cats
```
#### Creating pickle files
As the scraping of the network takes quite some time ($\sim$ 3 hours) (especially getting the inner connections), we store the results in pickle files.
```
# save_obj(network, 'network')
network = load_obj('network')
```
### Network creation
Let's convert the collected network into a NetworkX instance which is quite handy to manipulate.
Let's make it undirected as well.
```
neighbors = {}
for i in network.keys():
neighbors[i] = network[i]['links']
g = nx.Graph(neighbors) # undirected graph
```
# 2) Data Exploration <a class="anchor" id="Data Exploration"></a>
In this part of the notebook, we provide some indicators of the data in order to understand what we'll be working on.
* Adjacency matrix
* Degrees distribution
* Average degree
* Diameter of the collected network
* Visualization of the network
```
print('Total number of nodes : {}'.format(len(g.nodes)))
print('Total number of edges : {}'.format(len(g.edges)))
if nx.is_connected(g):
print('The graph is connected.')
else:
print('The graph is not connected.')
```
#### Adjacency Matrix
```
adj = nx.adjacency_matrix(g)
plt.spy(adj.todense())
```
Check if it's symmetric :
```
(adj != adj.T).count_nonzero() == 0
```
#### Degrees distribution
As there are some clear outliers making the visualization difficult, we can truncate the degrees or just use a box plot.
```
degrees = np.array(adj.sum(axis=1)).squeeze()
degrees_truncated = degrees[degrees < 700]
fig, ax = plt.subplots(ncols=2, sharey=True, figsize=(15,5))
ax[0].set_title('Degree distribution')
ax[0].hist(degrees, bins=50)
ax[1].set_title('Truncated degree distribution')
ax[1].hist(degrees_truncated, bins=20)
plt.tight_layout()
plt.show()
fig, ax = plt.subplots(ncols=2, sharey=True, figsize=(15,5))
ax[0].set_title('Degree box plot')
sns.boxplot(degrees, ax=ax[0])
ax[1].set_title('Truncated degree box plot')
sns.boxplot(degrees_truncated, ax=ax[1])
plt.tight_layout()
plt.show()
```
#### Average degree
```
avg_degree = np.mean(degrees)
print('The average degree of the network is {}.'.format(np.round(avg_degree, 2)))
```
#### Diameter
First we compute the shortest paths lengths. NetworkX allows us to do the computation and returns a dictionnary. This will be useful later on.
```
# shortest_paths = dict(nx.shortest_path_length(g))
# save_obj(shortest_paths, 'shortest_paths')
```
As this computation is quite long ($\sim$ 5 hours), we dumped the resulting dictionnary in a pickle file.
```
shortest_paths = load_obj('shortest_paths')
```
Now the computing the diameter of the networks comes down to finding the largest distance. Let's turn the dictionnary into a numpy array that is faster to manipulate.
```
nodes = list(network.keys())
distances = np.zeros(shape=(len(nodes), len(nodes)))
for i in range(len(nodes)):
for j in range(len(nodes)):
distances[i, j] = shortest_paths[nodes[i]][nodes[j]]
diameter = np.amax(distances)
print('The diameter of the network is {}.'.format(int(diameter)))
```
At first sight, if we had scraped first nodes and then second nodes, we should have had a diameter less than 4. Because a node should be at distance at most 2 from the root node.
Here, thanks to the use of disambiguation pages, we manage to get nodes that are further away from the root node but surprisingly our graph is connected anyway.
#### Visualization
```
nx.draw(g, node_size=5, figsize=(15, 15))
```
# 3) Data Exploitation <a class="anchor" id="Data Exploitation"></a>
## 3.1) Modelisation of the network <a class="anchor" id="Modelisation of the network"></a>
In this section, we are trying to model the collected network with a simpler one, trying to get the same main features like the number of nodes, the number of edges, the degree distribution, the shape of the giant components, and so on. Such a model is particularly useful to understand the original structure and compare it to other famous and already known networks.
In this modelisation part, we are using functions implemented in the `utils.py` in order to plot degree distributions and to get the regression coefficient of a power law.
```
from utils import get_distribution, linear_regression_coefficient
from utils import plot_degree_distribution, print_distribution, print_denoised_degree_distribution
```
### 3.1.1) Exploration of the degree distribution <a class="anchor" id="Exploration of the degree distribution"></a>
Let's first try to plot various transformations of the degree distribution in order to get a sense of a model that could fit.
```
nNodes = len(g.nodes())
nEdges = g.size()
print('The network has {0} nodes and {1} edges.'.format(nNodes, nEdges))
print('The minimum and the maximum degrees are respectively {0} and {1}.'.format(np.min(degrees), np.max(degrees)))
print_distribution(g, a=1000)
```
The previous plots show that the degree distribution of the network is complicated and doesn't fit exactly any of the basic network structures studied during the semester. However the last log-log plot suggests that a scale-free network with a power law could approximate the distribution. Let's make a regression to see what coefficient would fit. We use the `linear_model` function from sklearn.
```
linear_regression_coefficient(g, title='Linear regression of the original degree distribution')
```
The value of $R^2$ is not really close to 1 but a scale free network model does not seem too bad anyway.
We will later use that regression to build an approximation of the network. We make the assumption that the network distribution follows a power law of coefficient -1.0693.
### 3.1.2) Modelisation with usual graphs models <a class="anchor" id="Modelisation with usual graphs models"></a>
#### Erdős–Rényi graph
The Erdős–Rényi graph models a random network where each pair of nodes has a fixed probability to be linked. We want this network to have the same number of nodes as the original one, and approximate the number of edges as much as possible.
```
p = 2 * nEdges / nNodes / (nNodes - 1)
print('The probability hyper-parameter giving the best approximation of the number of edges is {}'.format(np.round(p, 4)))
er = nx.erdos_renyi_graph(nNodes, p)
plot_degree_distribution(er, title='Degree distribution of the Erdős–Rényi graph')
```
As expected, it clearly doesn't match the distribution of our network. The random networks have a Poisson degree distribution (when the number of nodes is large) and it doesn't fit to the observed distribution.
#### Barabási-Albert
The Barabási-Albert graph follows a power law distribution (in theory $p(k) = C \times k^{-3}$) so we can hope much better results than with the Erdős–Rényi model. The number of nodes that we want in the graph is fixed and we can only play with the parameter specifying the number of edges to attach from a new node to existing nodes. With the trial and error method, we found out that setting this parameter to 54 gives the closest number of edges to our original graph.
```
ba = nx.barabasi_albert_graph(nNodes, 54)
print('This Barabási-Albert network has {0} edges while our original network has {1} edges.'.format(ba.size(), nEdges))
plot_degree_distribution(ba, title='Degree distribution of the Barabási-Albert graph')
```
It indeed seems to be a power law distribution. Let's have a deeper insight and try to measure the parameter of this power law. The coefficient of such a random graph should be 3 in theory.
```
print_denoised_degree_distribution(ba, b=200, d=200)
```
Regression to measure the model law's coefficient
```
linear_regression_coefficient(ba, limit=300, title='Linear regression of the Barabási-Albert degree distribution')
```
We get a coefficient 2.7 that is close to 3 the expected value. Thus this network will be a better approximation than the random network precedently exposed but is still not ideal : we would like a power law network whose coefficient is closer to 1.0693 as computed earlier.
#### Comparison between the collected and the Barabási-Albert network distributions
```
ba_degrees = list(dict(ba.degree()).values())
f, ax = plt.subplots(figsize=(15, 6))
sns.distplot(degrees_truncated, label='Collected network', ax=ax)
sns.distplot(ba_degrees, label='Barabási-Albert network', ax=ax)
plt.legend(loc='upper right')
plt.show()
```
We clearly see here that it is a better approximation than the Erdős–Rényi graph but is still not ideal.
### 3.1.3) A power law network with the right exponent <a class="anchor" id="A power law network with the right exponent"></a>
In this section we are trying to make a power law network with a closer exponent to the one measured in the regression of the original network. We didn't find any method to make a graph with the exact exponent but we approximated it with the following code.
The `configuration_model` method from NetworkX allows us to create a graph from a given list of degrees.
In order to create a list of degrees respecting a power law distribution of coefficient $\gamma$, we use the function `powerlaw_sequence` from NetworkX. However this function can return zeros and we don't want to do that because our graph is connected. So what we do is generate each degree one at a time and check that it's not 0.
```
while True:
# Iterate the construction of a degree sequence until we find one that has a pair sum
# (this is a requirement of the function configuration_model)
s = []
while len(s) < nNodes:
# generate degrees one at a time
nextval = int(nx.utils.powerlaw_sequence(1, 1.6)[0])
if nextval != 0:
s.append(nextval)
if sum(s) % 2 == 0:
# we found a proper distribution, can break!
break
power_law = nx.configuration_model(s)
power_law = nx.Graph(power_law) # remove parallel edges
power_law.remove_edges_from(power_law.selfloop_edges())
print('This power law network has {0} nodes and {1} edges.'.format(len(power_law), power_law.size()))
```
We note right now that the number of edges in this model is really lower to the value in the collected network (367483).
It seems that the lowest coefficient we can set for the power law is 1.6. All the other attemps with smaller coefficient have crashed.
We can check that it indeed follows a power law distribution :
```
print_denoised_degree_distribution(power_law, a=100, b=200, d=200)
```
And we calculate here the coefficient of the power law :
```
linear_regression_coefficient(power_law, limit=79, title='Linear regression of the power-law degree distribution')
```
It's indeed closer to the original network but there is still a little gap (reminder, the obtjective is 1.0693). However as noted earlier, the number of edges of this power law network is extremely low compared to the original network. It seems like the Barabási-Albert network is a better approximation even if the fit of the distribution is not as good.
The following plot comparing the obtained degree distribution to the original one confirms that the Barabási-Albert network is a better approximation.
```
pl_degrees = list(dict(power_law.degree()).values())
f, ax = plt.subplots(figsize=(15, 6))
sns.distplot(degrees_truncated, label='Collected network', ax=ax)
sns.distplot(pl_degrees, label='Barabási-Albert network', ax=ax)
plt.legend(loc='upper right')
axes = plt.gca()
axes.set_xlim([-100, 1000])
plt.show()
```
### 3.1.4 Comparison of the models <a class="anchor" id="Comparison of the models"></a>
#### Giant components
In this part we are analyzing the giant components of the original network and of the models.
```
giant_g = max(nx.connected_component_subgraphs(g), key=len)
giant_er = max(nx.connected_component_subgraphs(er), key=len)
giant_ba = max(nx.connected_component_subgraphs(ba), key=len)
giant_pl = max(nx.connected_component_subgraphs(power_law), key=len)
print('Size of the giant component / Size of the network ')
print('Collected network : \t {}/{}'.format(len(giant_g.nodes()), len(g.nodes())))
print('Erdős–Rényi model : \t {}/{}'.format(len(giant_er.node()), len(er.nodes())))
print('Barabási-Albert model : {}/{}'.format(len(giant_ba.nodes()), len(ba.nodes())))
print('Power law model : \t {}/{}'.format(len(giant_pl.nodes), len(power_law.nodes)))
```
The original network, the Erdős–Rényi and the Barabási-Albert graphs are fully connected. The modelisation with the last power law network has also a very big giant component and is almost fully connected. We can conclude that the connectedness of the network is respected.
#### Clustering coefficient
The average clustering coefficient measures the overall degree of clustering in the network. Real-world networks tend to have a higher average clustering coefficient because of their ability to have compact groupements of nodes so we expect it to be greater than the models.
```
avg_clustering_g = nx.average_clustering(g)
avg_clustering_er = nx.average_clustering(er)
avg_clustering_ba = nx.average_clustering(ba)
avg_clustering_pl = nx.average_clustering(power_law)
print('Clustering coefficients')
print('Collected network : \t {}'.format(np.round(avg_clustering_g, 3)))
print('Erdős–Rényi model : \t {}'.format(np.round(avg_clustering_er, 3)))
print('Barabási-Albert model : {}'.format(np.round(avg_clustering_ba, 3)))
print('Power law model : \t {}'.format(np.round(avg_clustering_pl, 3)))
```
The last model created following a power law has the closest clustering coefficient. However, the really low number of edges is critical to make it a good model.
### 3.1.5) Comparison with common networks from the web <a class="anchor" id="Comparison with common networks from the web"></a>
Most scale-free networks follow a distribution of the form $p(k) = C\times k^{-\gamma}$ where $2 < \gamma < 3$ usually. In the approximation by a power law distribution we made, we found out that $\gamma \simeq 1.0693$ which is not really a common value as the following array shows (values seen during the lectures).
| Network | Gamma |
|----------------------|-------|
| WWW in | 2.00 |
| WWW out | 2.31 |
| Emails in | 3.43 |
| Emails out | 2.03 |
| Actor | 2.12 |
| Protein interactions | 2.12 |
| Citations in | 3.03 |
| Citations out | 4.00 |
Is a scale free network such a good model for our collected network ? We saw that the fit is not too bad but there are also more empirical reasons for such a model.
We may wonder why a scale free network seems to be a good approximation of the collected network. One of the most notable caracteristic of a scale free network is the presence of nodes with a degree much larger than the average which is the case here :
```
print('Average degree : {}'.format(np.round(np.mean(degrees), 1)))
print('{} nodes with degree 5 times bigger.'.format(np.count_nonzero(degrees > 5*np.mean(degrees))))
print('{} nodes with degree 10 times bigger.'.format(np.count_nonzero(degrees > 10*np.mean(degrees))))
```
It's in fact quite intuitive when you know that the network is composed of nodes representing wikipedia webpages and being linked if there is a link directing from one of the page to the other one. We expect a few large hubs (Wikipedia pages covering an important subject) appearing in such a network, followed by smaller ones (moderately important subjects) in a larger proportion and finally quite a lot of minor ones. The plots above show that the distribution respects that trend pretty much, except that there are fewer minor and very important topics that in a real scale free network.
This difference is likely to come directly from our **sampling method**. Indeed as we start from a central node and stop the collection somewhere, central nodes are important and at the *end* of the network we get what looks like minor pages. Those one could have been important if we had pushed the collection one hop further from the root node.
The preferential attachment process is another intuitive way to understand why the scrapped network looks like a scale free network. This process is also known as "the rich get richer and the poor get poorer" : a quantity (here the links between the nodes) is distributed according to how much they already have. It has been shown that such a process produces scale free networks and most algorithms (like the Barabási-Albert one) use this principle to create such networks. Regarding wikipedia, the more popular a page is and the more the topic is important, the more links it will have and conversely for lesser known pages. It is exactly a **preferential attachment** phenomena.
## 3.2) Community detection using Spectral clustering <a class="anchor" id="Community detection using Spectral clustering"></a>
We will try to use the collected data to answer our problem which is:
**Can we isolate communities of pages just by looking at the hyperlink graph ? **
This is the famous community detection problem for which a popular method is the [Louvain Algorithm](https://en.wikipedia.org/wiki/Louvain_Modularity).
The measure of performance we will use for the community detection is the modularity. Modularity measures the strengh of the division of a network into sub-groups. A network with high modularity has dense intra-connections (within sub-groups) and sparse inter-connections (between different groups).
Louvain as been presented in 2008 and though it has been improved [[1](https://link.springer.com/chapter/10.1007/978-3-319-11683-9_12)], we will use this as a baseline to compare the performance of spectral clustering for community detection.
The steps are the following :
* Louvain algorithm as a baseline
* Spectral clustering
* Visualization of the communities
```
from utils import get_bag_of_communities
```
### 3.2.1) Louvain community detection <a class="anchor" id="Louvain community detection"></a>
We use the Python library [`community`](https://pypi.python.org/pypi/python-louvain/0.3) that implements the Louvain algorithm.
This library also allows us to compute the modularity of a given partition of the nodes.
```
louvain_partition = community.best_partition(g)
louvain_modularity = community.modularity(louvain_partition, g)
louvain_modularity
k_louvain = len(set(louvain_partition.values()))
print('Louvain algorithm found {} communities'.format(k_louvain))
```
We can try to visualize the categories of the nodes in each of these communities. From the scraping, we got for each page a list of categories in which the page belongs. Let's compute for each community what we'll call a bag of categories that is the list of all the categories of the nodes it contains and the count of the number of nodes that belong to this category for each one.
The function can be found in `utils.py`, it's implementation is quite straight-forward.
```
louvain_bag = get_bag_of_communities(network, louvain_partition)
```
Let's get the number of pages in each community.
```
louvain_counts = [0 for _ in range(k_louvain)]
for i, title in enumerate(louvain_partition.keys()):
louvain_counts[louvain_partition[title]] += 1
```
Now we want to visualize the categories of the nodes in each community. We print for each community the 10 most represented categories of the community.
```
for i in range(k_louvain):
sorted_bag = sorted(louvain_bag[i].items(), key=operator.itemgetter(1), reverse=True)
print(' ')
print('Community {}/{} ({} pages) : '.format(i+1, k_louvain, louvain_counts[i]))
for ind in range(10):
print(sorted_bag[ind])
```
We can see that we get some nice results because it seems that a general topic can be infered for each community. The topics are:
| Alphabetical Order |
|---------------------------------|
| Aircrafts |
| American Footbal |
| Animals / mammals |
| Apple inc. |
| British ships |
| Cars |
| Comics and fictional characters |
| Electronics |
| Car racing |
| Luxury in Britain |
| Mexican soccer |
| Music instruments |
| Rugby |
| Science |
| Social science |
| Songwriters |
| Weapons |
### 3.2.2) Spectral Clustering <a class="anchor" id="Spectral Clustering"></a>
Nows let's try a spectral clustering approach for this community detection problem.
#### 3.2.2.1 Using the natural graph
The first idea is to use the natural graph, that is each node is a page and there is an edge of weight 1 between two pages if one of the pages links to the other.
We define the graph laplacian using the formula $L = D- A$ where $D$ is the diagonal matrix containing the degrees and $A$ is the adjacency matrix.
```
laplacian = np.diag(degrees) - adj.todense()
laplacian = sparse.csr_matrix(laplacian)
plt.spy(laplacian.todense())
```
In order to do spectral clustering using this Laplacian, we need to compute the $k$ first eigenvalues and corresponding eigenvectors. We get a matrix $U$ of $\mathbb{R}^{n \times k}$ where $n$ is the number of nodes in the graph. Applying a k-means algorithm in order to clusterize the $n$ vectors of $\mathbb{R}^k$ corresponding to the lines of $U$ gives us a clustering of the $n$ nodes.
Here we need to specify the number of clusters (communities) we want to look for. As a reminder, Louvain returned 17 (sometimes it gives 16) communities (it seems that it gives the maximum modularity but let's recall that Louvain is a heuristic so we are not sure of that).
Later in this notebook (at the end of the development of the model), we run some sort of cross-validation on the parameter `k_spectral`. For different values, we run the algorithm 5 times and take the mean and standard deviation of the modularity. It seems that 21 gives the best results. Please see below for details on this.
```
k_spectral = 21
eigenvalues, eigenvectors = sparse.linalg.eigsh(laplacian.asfptype(), k=k_spectral, which='SM')
plt.plot(eigenvalues, '.-', markersize=15)
eigenvalues[:2]
```
We check that the first eigenvalue is 0 but the second is not. The graph is connected.
Now we clusterize the resulting vectors in $\mathbb{R}^k$
```
centroids, labels = cluster.vq.kmeans2(eigenvectors, k_spectral)
```
This warning shows that at least one of the clusters is empty.
In order to get a first idea of how this algorithm did, let's look at the number of nodes in each cluster.
```
cc = [0 for i in range(k_spectral)]
for i in labels:
cc[i] += 1
', '.join([str(i) for i in cc])
```
We can see that with almost all the clusters containing less than 3 nodes, this first algorithm did not perform really well.
#### 3.2.2.2 Building another graph
As we have seen in class and in one of the homework. In order for spectral clustering to work, we need to assign edge weights that are stronger the closer the nodes are.
Let's build another graph with still the same vertex but some new edges between them.
We have already computed the distances in the graph let's define edges with weights using a kernel (e.g. the Gaussian kernel).
```
kernel_width = distances.mean()
weights = np.exp(-np.square(distances)/kernel_width**2)
np.fill_diagonal(weights, 0)
```
This creates a complete graph. We could sparsify it for faster computations but this is not really long and experience seems to show that results are better with the full graph.
```
degrees = np.sum(weights, axis=0)
plt.hist(degrees, bins=50);
laplacian = np.diag(1/np.sqrt(degrees)).dot((np.diag(degrees) - weights).dot(np.diag(1/np.sqrt(degrees))))
```
We can check that the obtained Laplacian matrix is symmetric.
```
tol = 1e-8
np.allclose(laplacian, laplacian.T, atol=tol)
eigenvalues, eigenvectors = linalg.eigh(laplacian, eigvals=(0, k_spectral-1))
plt.plot(eigenvalues, '.-', markersize=15)
centroids, labels = cluster.vq.kmeans2(eigenvectors, k_spectral)
cc = [0 for i in range(k_spectral)]
for i in labels:
cc[i] += 1
', '.join([str(i) for i in cc])
```
This seems better. We get pages distributed among all the clusters (with somme clusters more important than the others of course).
First let's have a look at the categories of each cluster.
```
spectral_partition = {}
for i, title in enumerate(network.keys()):
spectral_partition[title] = labels[i]
spectral_bag = get_bag_of_communities(network, spectral_partition)
spectral_counts = [0 for _ in range(k_spectral)]
for i, title in enumerate(spectral_partition.keys()):
spectral_counts[spectral_partition[title]] += 1
for i in range(k_spectral):
sorted_bag = sorted(spectral_bag[i].items(), key=operator.itemgetter(1), reverse=True)
print(' ')
print('Community {}/{} ({} pages) : '.format(i+1, k_spectral, spectral_counts[i]))
if spectral_counts[i] > 0:
for ind in range(10):
print(sorted_bag[ind])
```
It seems that we get the same results. As we asked for more communities than Louvain, some of them are split but it's either a duplicate or a finer separation.
There are some inconsistensies in the partition we get:
- two communities for Songwriters
- three communities for Ship incidents
- two communities for NFL
- two communities for mammals
but the community electronics is now split into video games and computer hardware.
So we get more communities. Sometimes its just duplicates but sometimes it is a finer separation of two groups.
```
spectral_modularity = community.modularity(spectral_partition, g)
spectral_modularity
```
The modularity coefficient is lower.
#### Testing different values of `k_spectral`
Here we test different values of k. It seems after some testing that there is a high variance in the modularity of partitions returned by the algo (for a given `k_spectral`). In order to find out if there is really a value better than the others. We compute the mean and variance of modularity for a given value by running 5 times the algorithm.
```
"""cross_val = {}
for k in tqdm_notebook(range(10, 30)):
tmp = []
for _ in range(5):
eigenvalues, eigenvectors = linalg.eigh(laplacian, eigvals=(0, k-1))
centroids, labels = cluster.vq.kmeans2(eigenvectors, k)
spectral_partition = {}
for i, title in enumerate(network.keys()):
spectral_partition[title] = labels[i]
spectral_modularity = community.modularity(spectral_partition, g)
tmp.append(spectral_modularity)
cross_val[k] = [np.mean(tmp), np.std(tmp)]
save_obj(d, 'cross_val')"""
```
As this computation takes approximately one hour to terminate, the results have been stored in a pickle file.
```
cross_val = load_obj('cross_val')
cross_val
```
We see that the best modularity seems to be achieved with the parameter k of 21. However we note that the standard deviation is quite high in all the cases.
### 3.2.3 Comparison of the two methods <a class="anchor" id="Comparison of the two methods"></a>
It seems that no matter the value of `k` we choose, we wont be able to have a higher modularity than the one achieved by the Louvain algorithm. So what could be the advantages of the spectral approach ?
* Computational cost :
- Louvain algo : greedy algorithm that appears to run in $0(n \log n)$ where $n$ is the number
of nodes in the network.
- Spectral clustering : the computation of the Laplacian is already in $O(n^3)$ and that's without counting the shortest-paths matrix whose computation is costly $O(|E|+|V|\log |V|)$ using the Dijkstra algorithm. As we have many edges this is really costly.
The spectral clustering seems really more costly than the Louvain method. That's is something we had noticed in our study.
* Better communities ?
- We have seen that the communities are not better in the sense of the modularity (measuring the internal cohesion of communities versus the separation of different ones).
- Could the partition could be better when it comes to the categories of the nodes ? In order to be able to measure that, we could apply a NLP pipeline on the categories of the pages to do Topic Selection. This could give us a more precise idea of the performance of the split when it comes to topics (and not only modularity).
### 3.2.4 Visualization <a class="anchor" id="3.2.4Visualization"></a>
#### Plotly visualization
First we want to visualize the graph in the notebook using plotly.
In order to get clean representation of how our nodes build communities, we define a color map for each community that will help us differentiate clusters in our network.
```
community2color = {
0: sns.xkcd_rgb["peach"],
1: sns.xkcd_rgb["powder blue"],
2: sns.xkcd_rgb["light pink"],
3: sns.xkcd_rgb["chocolate"],
4: sns.xkcd_rgb["orange"],
5: sns.xkcd_rgb["magenta"],
6: sns.xkcd_rgb["purple"],
7: sns.xkcd_rgb["blue"],
8: sns.xkcd_rgb["deep blue"],
9: sns.xkcd_rgb["sky blue"],
10: sns.xkcd_rgb["olive"],
11: sns.xkcd_rgb["seafoam green"],
12: sns.xkcd_rgb["tan"],
13: sns.xkcd_rgb["mauve"],
14: sns.xkcd_rgb["hot pink"],
15: sns.xkcd_rgb["pale green"],
16: sns.xkcd_rgb["indigo"],
17: sns.xkcd_rgb["lavender"],
18: sns.xkcd_rgb["eggplant"],
19: sns.xkcd_rgb["brick"],
20: sns.xkcd_rgb["light blue"],
}
```
A simple representation of our graph can be obtained using the networkx tool for drawing. Here already, we get an idea of how each category of wikipedia articles creates a clear dense connection of nodes in our graph.
```
position = nx.spring_layout(g)
for community in set(louvain_partition.values()) :
list_nodes = [nodes for nodes in louvain_partition.keys() if louvain_partition[nodes] == community]
nx.draw_networkx_nodes(g, position, list_nodes, node_size=20, node_color=community2color[int(community)])
nx.draw_networkx_edges(g, position, alpha=0.5)
plt.show()
```
Now, in order to put this into a more interactive perspective we will be using plotly scatter plots, to help us play with our network. For each of the nodes, we set up attribute as in which community it belongs to based on Louvain or spectral partition. We can also assigne positions to each node. This is important in order to find a good representation of the network. Networkx and community packages come with built in functions for positioning networks according to various algorithms. After trying out various algorithms, we chose to use spring_layout. The result is not perfect but it is easy to use and to implement.
```
nx.set_node_attributes(g, spectral_partition, 'spectral')
nx.set_node_attributes(g, louvain_partition, 'louvain')
nx.set_node_attributes(g, position, 'position')
```
We implemented two functions in utils that allow us to plot interactive graphs.
```
from utils import build_communities, set_layout
data = build_communities('louvain','position', G=g, community2color=community2color)
layout = set_layout('Louvain')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
data = build_communities('spectral', 'position', G=g, community2color=community2color)
layout = set_layout('spectral clustering')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
```
The layout of the nodes is not ideal but we could not make it as clear as on Gephi (see below).
We can go through the nodes in graph and see the titles of the pages that belong to each cluster marked with different colors. As we have already observed, we managed to connect the communities to a category of articles, with using the network based on hyperlinks.
#### Gephi visualization
Though the previous visualizations can be handy, they are quite slow and the layout is not really representative of the structure of the graph. This is why we chose to use Gephi as our main visualization tool. Here is the result of some visualizations.
Using Gephi, we were able to get some nice visualizations of our clustering.
```
for i in louvain_partition.keys():
louvain_partition[i] = str(louvain_partition[i])
for i in spectral_partition.keys():
spectral_partition[i] = str(spectral_partition[i])
```
NetworkX requires all the node attributes to be strings to export them.
```
nx.set_node_attributes(g, louvain_partition, 'louvain')
nx.set_node_attributes(g, spectral_partition, 'spectral')
for n, d in g.nodes(data=True):
del d['position']
nx.write_graphml(g, 'data/full_graph.graphml')
```
Now opened in Gephi, this is what our graph looks like.
We used a repulsive layout in order to make the graph the more readable possible. Nodes in the same community are in the same color.
<img src="images/1.png" style="width: 700px;">
We can look more closely to some communities, for example the Apple one :
<img src="images/apple.png" style="width: 500px;">
or the Mexican soccer one:
<img src="images/mexican_soccer.png" style="width: 500px;">
Some additional screenshots are provided in the folder images/. Feel free to have a look at different communities.
#### Comparison of the two communities:
Here is a view of the communities obtained by spectral clustering first and below by Louvain algorithm.
<img src="images/1.png" style="width: 200px;">
<img src="images/2.png" style="width: 200px;">
Sure the layout is the same because the graph is the same. What is interesting to note is that the two clustering look approximately alike.
# 4) Conclusion <a class="anchor" id="Conclusion"></a>
In this study, we first collected a sample network of the Wikipedia hyperlink graph. We tried different approaches using either the wikipedia API or the scraping of the raw pages. Eventually we managed to get a significant sample (6830 nodes) in a reasonable time (3 hours). The use of disambiguation pages proved to be handy to get a large variety of pages from a single root node.
Then we tried to model this collected network using usual network structures. The various plots of the degree distribution suggest that a power distribution of the degrees can be a good approximation. The Barabási-Albert model turned out to be the best approximation as it fits the collected network on many of its properties (number of nodes and edges, degrees distribution, shape of giant component).
Eventually we answered our initial question that was to know if the structure of the graph bears information on the content of the nodes. That proved to be true as we could extract communities of categories only by looking at proximity features. The Louvain algorithm seemed to provide a better community detection than the spectral clustering algorithm. It indeed gives a better modularity and is faster to run. We checked that the extracted communities share a common topic by visualizing the clusters using for example Gephi.
In order to continue our study we could try to measure properly the fit of the community detection with the categories of the nodes by implementing a natural language processing pipeline on those categories in order to extract topics.
| github_jupyter |
High Density Areas of Urban Development cit.014 http://data.jrc.ec.europa.eu/dataset/jrc-ghsl-ghs_smod_pop_globe_r2016a
```
import numpy as np
import pandas as pd
import rasterio
import boto3
import requests as req
from matplotlib import pyplot as plt
%matplotlib inline
import os
import sys
import threading
```
Establish s3 location
```
# Investigate what the data in these rasters means, and whether we can
# Display high and low density clusters separately as is
s3_bucket = "wri-public-data"
s3_folder = "resourcewatch/cit_014_areas_of_urban_development/"
s3_files = ["cit_014_areas_of_urban_development_1975.tif",
"cit_014_areas_of_urban_development_1990.tif",
"cit_014_areas_of_urban_development_2000.tif",
"cit_014_areas_of_urban_development_2015.tif",
"cit_014_areas_of_urban_development_2015_HDC.tif",
"cit_014_areas_of_urban_development_2015_LDC.tif"]
s3_file_merge = "cit_014_areas_of_urban_development_merge.tif"
s3_key_origs = []
s3_key_edits = []
for file in s3_files:
orig = s3_folder + file
s3_key_origs.append(orig)
s3_key_edits.append(orig[0:-4] + "_edit.tif")
s3_key_merge = s3_folder + s3_file_merge
s3_key_edits
```
Create local staging folder for holding data
```
!mkdir staging
os.chdir("staging")
staging_folder = os.getcwd()
os.environ["Z_STAGING_FOLDER"] = staging_folder
```
Local files
```
local_folder = "/Users/nathansuberi/Desktop/WRI_Programming/RW_Data"
rw_data_type = "/Cities/"
# Topics include: [Society, Food, Forests, Water, Energy, Climate, Cities, Biodiversity, Commerce, Disasters]
local_files = [
"GHS_SMOD_POP1975_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP1975_GLOBE_R2016A_54009_1k_v1_0.tif",
"GHS_SMOD_POP1990_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP1990_GLOBE_R2016A_54009_1k_v1_0.tif",
"GHS_SMOD_POP2000_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP2000_GLOBE_R2016A_54009_1k_v1_0.tif",
"GHS_SMOD_POP2015_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP2015_GLOBE_R2016A_54009_1k_v1_0.tif",
"GHS_SMOD_POP2015HDC_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP2015HDC_GLOBE_R2016A_54009_1k_v1_0.tif",
"GHS_SMOD_POP2015LDC_GLOBE_R2016A_54009_1k_v1_0/GHS_SMOD_POP2015LDC_GLOBE_R2016A_54009_1k_v1_0.tif"
]
local_orig_keys = []
local_edit_keys = []
for file in local_files:
local_orig_keys.append(local_folder + rw_data_type + file)
local_edit_keys.append(local_folder + rw_data_type + file[0:-4] + "_edit.tif")
local_orig_keys
```
<b>Regardless of any needed edits, upload original file</b>
<i>Upload tif to S3 folder</i>
http://boto3.readthedocs.io/en/latest/guide/s3-example-creating-buckets.html
<i>Monitor Progress of Upload</i>
http://boto3.readthedocs.io/en/latest/_modules/boto3/s3/transfer.html
https://boto3.readthedocs.io/en/latest/guide/s3.html#using-the-transfer-manager
```
s3 = boto3.client("s3")
class ProgressPercentage(object):
def __init__(self, filename):
self._filename = filename
self._size = float(os.path.getsize(filename))
self._seen_so_far = 0
self._lock = threading.Lock()
def __call__(self, bytes_amount):
# To simplify we'll assume this is hooked up
# to a single filename.
with self._lock:
self._seen_so_far += bytes_amount
percentage = (self._seen_so_far / self._size) * 100
sys.stdout.write(
"\r%s %s / %s (%.2f%%)" % (
self._filename, self._seen_so_far, self._size,
percentage))
sys.stdout.flush()
# Defined above:
# s3_bucket
# s3_key_orig
# s3_key_edit
# staging_key_orig
# staging_key_edit
for i in range(0,6):
print(i)
s3.upload_file(local_orig_keys[i], s3_bucket, s3_key_origs[i],
Callback=ProgressPercentage(local_orig_keys[i]))
```
Check for compression, projection
Create edit file if necessary
```
# Check Compression, Projection
with rasterio.open(local_orig_keys[0]) as src:
pro0 = src.profile
data0 = src.read(1)
with rasterio.open(local_orig_keys[1]) as src:
pro1 = src.profile
data1 = src.read(1)
with rasterio.open(local_orig_keys[2]) as src:
pro2 = src.profile
data2 = src.read(1)
with rasterio.open(local_orig_keys[3]) as src:
pro3 = src.profile
data3 = src.read(1)
with rasterio.open(local_orig_keys[4]) as src:
pro4 = src.profile
data4 = src.read(1)
with rasterio.open(local_orig_keys[5]) as src:
pro5 = src.profile
data5 = src.read(1)
# uniq0 = np.unique(data0, return_counts=True)
# uniq1 = np.unique(data1, return_counts=True)
# uniq2 = np.unique(data2, return_counts=True)
# uniq3 = np.unique(data3, return_counts=True)
# uniq4 = np.unique(data4, return_counts=True)
# uniq5 = np.unique(data5, return_counts=True)
uniq4
# Examine each of the profiles - are they all the same data type?
print(pro0)
print(pro1)
print(pro2)
print(pro3)
print(pro4)
print(pro5)
profiles = [pro0, pro1, pro2, pro3, pro4, pro5]
```
Upload edited files to S3
```
# Defined above:
# s3_bucket
# s3_key_orig
# s3_key_edit
# staging_key_orig
# staging_key_edit
for i in range(0,6):
orig_key = local_orig_keys[i]
edit_key = local_edit_keys[i]
# Use rasterio to reproject and store locally, then upload
with rasterio.open(orig_key) as src:
kwargs = profiles[i]
print(kwargs)
kwargs.update(
driver='GTiff',
dtype=rasterio.int32, #rasterio.int16, rasterio.int32, rasterio.uint8,rasterio.uint16, rasterio.uint32, rasterio.float32, rasterio.float64
count=1,
compress='lzw',
nodata=0,
bigtiff='NO',
crs = 'EPSG:4326',
)
windows = src.block_windows()
with rasterio.open(edit_key, 'w', **kwargs) as dst:
for idx, window in windows:
src_data = src.read(1, window=window)
formatted_data = src_data.astype("int32")
dst.write_band(1, formatted_data, window=window)
s3.upload_file(edit_key, s3_bucket, s3_key_edits[i],
Callback=ProgressPercentage(edit_key))
s3_file_merge
```
Merge files and upload to s3
```
merge_key = './'+s3_file_merge
kwargs = profiles[i]
print(kwargs)
kwargs.update(
driver='GTiff',
dtype=rasterio.int32, #rasterio.int16, rasterio.int32, rasterio.uint8,rasterio.uint16, rasterio.uint32, rasterio.float32, rasterio.float64
count=len(profiles),
compress='lzw',
nodata=0,
bigtiff='NO',
crs = 'EPSG:4326',
)
with rasterio.open(merge_key, 'w', **kwargs) as dst:
for idx, file in enumerate(local_edit_keys):
print(idx)
with rasterio.open(file) as src:
band = idx+1
windows = src.block_windows()
for win_id, window in windows:
src_data = src.read(1, window=window)
dst.write_band(band, src_data, window=window)
s3.upload_file(merge_key, s3_bucket, s3_key_merge,
Callback=ProgressPercentage(merge_key))
```
Inspect the final product
```
tmp = "./temp"
s3 = boto3.resource("s3")
s3.meta.client.download_file(s3_bucket, s3_key_merge, tmp)
with rasterio.open(tmp) as src:
print(src.profile)
data = src.read(4)
os.getcwd()
np.unique(data, return_counts=True)
plt.imshow(data)
```
| github_jupyter |
```
%%writefile mapper1.py
# Your code for mapper here.
import sys
import re
reload(sys)
sys.setdefaultencoding('utf-8') # required to convert to unicode
path = 'stop_words_en.txt'
def read_stop_words(file_path):
return set(word.strip().lower() for word in open(file_path))
stop_words = read_stop_words(path)
for line in sys.stdin:
try:
article_id, text = unicode(line.strip()).split('\t', 1)
except ValueError as e:
continue
words = re.split("\W*\s+\W*", text, flags=re.UNICODE)
for word in words:
if word.lower() not in stop_words:
print >> sys.stderr, "reporter:counter:Wiki stats,Total words,%d" % 1
print "%s\t%d" % (word.lower(), 1)
%%writefile reducer1.py
# Your code for reducer here.
import sys
current_key = None
word_sum = 0
for line in sys.stdin:
try:
key, count = line.strip().split('\t', 1)
count = int(count)
except ValueError as e:
continue
if current_key != key:
if current_key:
print "%s\t%d" % (current_key, word_sum)
word_sum = 0
current_key = key
word_sum += count
if current_key:
print "%s\t%d" % (current_key, word_sum)
%%writefile mapper2.py
import sys
reload(sys)
for line in sys.stdin:
try:
word, count = line.strip().split('\t', 1)
count = int(count)
if word.isalpha():
word_permuted = ''.join(sorted(word))
print "%s\t%s\t%d" % (word_permuted, word, count)
except ValueError as e:
continue
%%writefile reducer2.py
# Your code for reducer here.
import sys
current_key = None
permuted_sum = 0
word_group = []
for line in sys.stdin:
try:
key, word, count = line.strip().split('\t', 2)
count = int(count)
except ValueError as e:
continue
if current_key != key:
if current_key:
if len(word_group) > 1:
word_string = ','.join(sorted(word_group))
print "%d\t%d\t%s" % (permuted_sum, len(word_group), word_string)
current_key = key
permuted_sum = 0
word_group = []
permuted_sum += count
word_group.append(word)
if current_key:
if len(word_group) > 1:
word_string = ','.join(sorted(word_group))
print "%d\t%d\t%s" % (permuted_sum, len(word_group), word_string)
%%bash
OUT_DIR="wordgroup_wordcount_"$(date +"%s%6N")
NUM_REDUCERS=8
hdfs dfs -rm -r -skipTrash ${OUT_DIR} > /dev/null
# Code for your first job
# yarn jar /opt/cloudera/parcels/CDH/lib/hadoop-mapreduce/hadoop-streaming.jar ...
yarn jar /opt/cloudera/parcels/CDH/lib/hadoop-mapreduce/hadoop-streaming.jar \
-D mapred.jab.name="Streaming wordCount" \
-D mapreduce.job.reduces=${NUM_REDUCERS} \
-files mapper1.py,reducer1.py,/datasets/stop_words_en.txt \
-mapper "python mapper1.py" \
-combiner "python reducer1.py" \
-reducer "python reducer1.py" \
-input /data/wiki/en_articles_part \
-output ${OUT_DIR} > /dev/null
OUT_DIR1="wordgroup_permutation_"$(date +"%s%6N")
NUM_REDUCERS=1
hdfs dfs -rm -r -skipTrash ${OUT_DIR1} > /dev/null
# Code for your second job
# yarn jar /opt/cloudera/parcels/CDH/lib/hadoop-mapreduce/hadoop-streaming.jar ...
yarn jar /opt/cloudera/parcels/CDH/lib/hadoop-mapreduce/hadoop-streaming.jar \
-D mapred.jab.name="Streaming permutations" \
-D mapreduce.job.reduces=${NUM_REDUCERS} \
-files mapper2.py,reducer2.py \
-mapper "python mapper2.py" \
-reducer "python reducer2.py" \
-input ${OUT_DIR} \
-output ${OUT_DIR1} > /dev/null
# Code for obtaining the results
hdfs dfs -cat ${OUT_DIR1}/part-00000 | grep -P '(,|\t)english($|,)'
hdfs dfs -rm -r -skipTrash ${OUT_DIR1}* > /dev/null
```
| github_jupyter |
<font color=gray>Oracle Cloud Infrastructure Data Science Sample Notebook
Copyright (c) 2021 Oracle, Inc. All rights reserved. <br>
Licensed under the Universal Permissive License v 1.0 as shown at https://oss.oracle.com/licenses/upl.
</font>
# Uploading Larger Size Model Artifact Using OCI Pyhton SDK
This notebook demonstrates simple solution for OCI Python SDK which allows data scientists to upload larger model artifacts and eliminate the timeout error that is experienced by data scientists when the artifact is large. It shows end-to-end steps from setting up the configuration till uploading the model artifact.
## Pre-requisites to Running this Notebook
* We recommend that you run this notebook in an OCI Data Science Notebook Session. Use a conda environment that has a `oci>=2.43.2`. You can always upgrade oci by executing `!pip install oci --upgrade` in your notebook cell.
* You need access to the public internet
```
import os
import logging
import json
import oci
from oci.data_science import DataScienceClient
from oci.data_science.models import CreateModelDetails, CreateProjectDetails, \
CreateModelProvenanceDetails, Project, Model, UpdateModelDetails
data_science_models = oci.data_science.models
print(oci.__version__)
# here we assume that you are authenticating with OCI resources using user principals (config + key)
# set resource_principal == True if you are using resource principal instead of user principal
resource_principal = False
if not resource_principal:
config = oci.config.from_file()
data_science = DataScienceClient(config)
else:
config = {}
auth = oci.auth.signers.get_resource_principals_signer()
data_science = DataScienceClient(config, signer=auth)
# Creating the metadata about the model:
create_model_details_object = data_science_models.CreateModelDetails()
create_model_details_object.compartment_id = os.environ['NB_SESSION_COMPARTMENT_OCID']
create_model_details_object.display_name = f"<replace-with-your-object-display-name>"
create_model_details_object.project_id = os.environ['PROJECT_OCID']
# create the model object:
model = data_science.create_model(create_model_details_object)
# print the model OCID:
model_id = json.loads(str(model.data))['id']
print(model_id)
# next we download a sample boilerplate model artifact and setting the model artifact path.
!wget https://github.com/oracle/oci-data-science-ai-samples/blob/master/model_catalog_examples/artifact_boilerplate/artifact_boilerplate.zip
artifact_path = 'artifact_boilerplate.zip'
with open(artifact_path,'rb') as artifact_file:
artifact_bytes = artifact_file.read()
data_science.create_model_artifact(model_id,
artifact_bytes,
content_disposition='attachment; filename="{artifact_path}"')
# Go to the OCI Console and confirm that your model was successfully created.
```
| github_jupyter |
```
from __future__ import division, absolute_import
import sys
import os
import numpy as np
import random
import pickle
from plotnine import *
import math
import pandas as pd
#root
absPath = '/home/angela3/imbalance_pcm_benchmark/'
sys.path.insert(0, absPath)
from src.imbalance_functions import *
np.random.seed(8)
random.seed(8)
#LOADING DATA
activity_filename = "/home/angela3/hybrid_model/data/smiles_prots_activity.csv"
activity_df = pd.read_csv(activity_filename, sep="\t", header=0).drop("Unnamed: 0", axis=1)
activity_df.info()
activity_df.groupby("family")["DeepAffinity Protein ID"].nunique()
family_names = activity_df["family"].unique().tolist()
figures = {}
for family in family_names:
print("Protein family: ", family)
activity_fam = activity_df[activity_df.family == family]
unique_prots = activity_fam["DeepAffinity Protein ID"].drop_duplicates().tolist()
print("There are",len(unique_prots),"different proteins")
list_ratios = []
for prot in unique_prots:
ratio_actives_inactives = computing_active_inactive_ratio(activity_fam, prot)
dicti = {"DeepAffinity Protein ID" : prot, "ratio_actives_inactives": ratio_actives_inactives}
list_ratios.append(dicti)
df_ratios = pd.DataFrame(list_ratios)
p = (ggplot(df_ratios, aes("ratio_actives_inactives")) + geom_histogram()
+ xlab("Proportion of actives respect to all the interactions per protein") + ggtitle(family))
figures[family] = p
print("Family: PK")
figures["PK"]
ggsave(plot=figures["PK"],
filename="".join((absPath, "data/kinases/histogram_ratio_kinases.pdf")), device = "pdf")
print("Family: GPCR")
figures["GPCR"]
ggsave(plot=figures["GPCR"],
filename="".join((absPath, "data/GPCRs/histogram_ratio_gpcrs.pdf")), device = "pdf")
print("Family: NR")
figures["NR"]
print("Family: Non assigned")
figures["Non assigned"]
print("Family: OE")
figures["OE"]
print("Family: IC")
figures["IC"]
print("Family: TR")
figures["TR"]
print("Family: CY")
figures["CY"]
print("Family: PR")
figures["PR"]
```
## Imbalance per protein
```
protein_family = "PK" #"GPCR" #
protein_name = "kinases" #"GPCRs" #
# keeping only the protein family
activity_sub = activity_df[activity_df.family == protein_family]
activity_sub.info()
activity_sub.head()
label_count_subprot = pd.DataFrame(activity_sub[["DeepAffinity Protein ID", "label"]].groupby(["DeepAffinity Protein ID", "label"]).size()).reset_index()
label_count_subprot.head()
label_count_subprot.info()
label_count_subprot.columns = ['DeepAffinity Protein ID', 'label', "number"]
#Filling the count label dataframe
for prot in label_count_subprot["DeepAffinity Protein ID"].unique().tolist():
labels = [0, 1]
subprot = label_count_subprot[label_count_subprot["DeepAffinity Protein ID"] == prot]
#print(subprot.shape[0])
if subprot.shape[0]==2:
continue
else:
nomissing_label = subprot[["label"]].values
missing_label = [i for i in labels if i!=nomissing_label][0]
row_dict = {"DeepAffinity Protein ID": prot, "label": missing_label, "number":0}
label_count_subprot = label_count_subprot.append(row_dict, ignore_index=True)
label_count_subprot.info()
label_count_subprot.sort_values("DeepAffinity Protein ID")
p = (ggplot(label_count_subprot, aes(x="factor(label)", y="number"))
+ geom_boxplot() + ylab("Number of interactions")
+ xlab("Labels") + theme(legend_title=element_blank())
+ ggtitle(protein_name)
#+ scale_y_log10()
)
p
ggsave(plot=p,
filename="".join((absPath, "data/", protein_name, "/boxplot_ninteractions.pdf")), device = "pdf")
```
### Exporting CSV for a protein family
```
protein_family = "GPCR" #"PK"
protein_name = "GPCRs" #"kinases"
# keeping only the protein family
activity_sub = activity_df[activity_df.family == protein_family]
unique_prots = activity_sub["DeepAffinity Protein ID"].unique().tolist()
activity_sub.info()
activity_sub["label"].value_counts()
print("There are", activity_sub.shape[0], "protein-compound pairs available in ", protein_name)
#Saving final dataset to a csv
output_filename = "".join((absPath, "data/", protein_name, "_activity.csv"))
activity_sub.to_csv(output_filename, sep="\t", header=True)
unique_prots = activity_sub["DeepAffinity Protein ID"].unique().tolist()
unique_prots
len(unique_prots)
unique_comps = activity_sub["DeepAffinity Compound ID"].unique().tolist()
len(unique_comps)
#unique_prots_str = [prot.encode("utf-8") for prot in unique_prots]
#unique_prots_str[:3]
with open("".join((absPath, "data/", protein_name, "/", protein_name, "_prots.pickle")), 'wb') as handle:
pickle.dump(unique_prots, handle)
```
| github_jupyter |
```
import pandas as pd
data_files = [
"ap_2010.csv",
"class_size.csv",
"demographics.csv",
"graduation.csv",
"hs_directory.csv",
"sat_results.csv"
]
data = {}
for data_file in data_files:
#remove .csv in the file name for the variable name
df_name = data_file[:-4]
data[df_name] = pd.read_csv('../resources/' + data_file)
print(data.keys())
#make copy pointer to the sat dataframe
sat = data['sat_results']
print(sat.head())
for key,value in data.items():
print('Data set name {}'.format(key))
print(value.head())
all_survey = pd.read_csv('../resources//survey_all.txt', delimiter='\t',
encoding='windows-1252')
print(all_survey.iloc[0])
d75_survey = pd.read_csv('../resources/survey_d75.txt', delimiter='\t',
encoding='windows-1252')
print(d75_survey.iloc[0])
survey = pd.concat([all_survey,d75_survey],axis=0)
print(survey.iloc[0])
survey["DBN"] = survey["dbn"]
survey_fields = [
"DBN",
"rr_s",
"rr_t",
"rr_p",
"N_s",
"N_t",
"N_p",
"saf_p_11",
"com_p_11",
"eng_p_11",
"aca_p_11",
"saf_t_11",
"com_t_11",
"eng_t_11",
"aca_t_11",
"saf_s_11",
"com_s_11",
"eng_s_11",
"aca_s_11",
"saf_tot_11",
"com_tot_11",
"eng_tot_11",
"aca_tot_11",
]
survey = survey.loc[:,survey_fields]
data["survey"] = survey
print(survey.head())
def padded_csd(csd):
csd_str = str(csd)
if len(csd_str) == 1:
csd_str = csd_str.zfill(2)
return csd_str
def genDBN(row):
return row['padded_csd'] + row['SCHOOL CODE']
hs_dir = data['hs_directory']
hs_dir['DBN'] = hs_dir['dbn']
class_size = data['class_size']
class_size['padded_csd'] = class_size['CSD'].apply(padded_csd)
class_size['DBN'] = class_size.apply(genDBN, axis=1)
print(class_size['DBN'].iloc[0:5])
sat_results = data['sat_results']
string_to_num_cols = ['SAT Math Avg. Score', 'SAT Critical Reading Avg. Score','SAT Writing Avg. Score']
sat_results['sat_score'] = [0]*len(sat_results)
for col in string_to_num_cols:
sat_results[col] = pd.to_numeric(sat_results[col],errors='coerce')
sat_results['sat_score'] = sat_results['sat_score'] + sat_results[col]
print(sat_results[string_to_num_cols].iloc[0])
print(sat_results['sat_score'].iloc[0])
import re
def extract_lat(data):
match = re.findall('\(.+\)', data)
#splits the lat and long and remove first char '(' and last char from lat which is comma(,) and returns the string
lat_long = match[0]
return lat_long.split(' ')[0][1:-1]
hs_directory = data['hs_directory']
hs_directory['lat'] = hs_directory['Location 1'].apply(extract_lat)
print(hs_directory['lat'].iloc[0:5])
def extract_longitude(data):
match = re.findall('\(.+\)', data)
lat_long = match[0]
#split lat and long and remove last char ')' from long then return it
return lat_long.split(' ')[1][:-1]
hs_directory = data['hs_directory']
hs_directory['lon'] = hs_directory['Location 1'].apply(extract_longitude)
lat_lon = ['lat', 'lon']
for l in lat_lon:
hs_directory[l] = pd.to_numeric(hs_directory[l], errors='coerce')
print(hs_directory.iloc[0:5])
class_size = data['class_size']
#drop rows where grade is different than '09-12' and program type diff than 'GEN ED'
program_type = class_size['PROGRAM TYPE'] != 'GEN ED'
grade = class_size['GRADE '] != '09-12'
program_type_grade = (program_type | grade)
class_size.drop(class_size[program_type_grade].index, inplace=True)
print(class_size.iloc[0:5])
```
| github_jupyter |
# COVID-19
This notebook analyzes the growth of the COVID-19 pandemy. It relies on the data provided by Johns Hopkins CSSE at https://github.com/CSSEGISandData/COVID-19 . The main question is: how will the number of infected people change over time. We will use a very simple approach, that should not be used for serious predictions of spreads of deseases, but which is well supported in PySpark. For a better mathematical model, please read https://de.wikipedia.org/wiki/SIR-Modell . Unfortunately there is no support in PySpark for estimating model parameters within a more meaningful model.
So this notebook is mainly about getting some basic insights into machine learning with PySpark.
# 0. Spark Context & Imports
```
import matplotlib.pyplot as plt
from pyspark.sql import SparkSession
import pyspark.sql.functions as f
if not 'spark' in locals():
spark = SparkSession.builder \
.master("local[*]") \
.config("spark.driver.memory","64G") \
.getOrCreate()
spark
%matplotlib inline
```
# 1. Load Data
The original data is available at https://github.com/CSSEGISandData/COVID-19 provided by Johns Hopkins CSSE. There are several different representations of the data, we will peek into different versions and then select the most appropriate to work with.
```
basedir = 's3://dimajix-training/data/covid-19'
```
## 1.1 Load Time Series
The repository already contains time series data. This is nice to look at, but specifically for PySpark maybe a little bit hard to work with. Each line in the file contains a full time series of the number of positive tested persons. This means that the number of columns change with every update.
```
series = spark.read\
.option("header", True) \
.csv(basedir + "/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv")
series.toPandas()
```
## 1.2 Load Daily Reports
The repository also contains more detailed files containing the daily reports of the total number of positively tested persons. Within those files, every line represents exactly one region and one time. Therefore the schema stays stable with every update, only new records are appended. But there are some small technical challenges that we need to take.
### Date parser helper
First small challenge: All records contain a date, some of them a datetime. But the format has changed several times. In order to handle the different cases, we provide a small PySpark UDF (User Defined Function) that is capable of parsing all formats and which returns the extracted date.
```
import datetime
from pyspark.sql.types import *
@f.udf(DateType())
def parse_date(date):
if "/" in date:
date = date.split(" ")[0]
(m,d,y) = date.split("/")
y = int(y)
m = int(m)
d = int(d)
if (y < 2000):
y += 2000
else:
date = date[0:10]
(y,m,d) = date.split("-")
y = int(y)
m = int(m)
d = int(d)
return datetime.date(year=y,month=m,day=d)
#print(parse_date("2020-03-01"))
#print(parse_date("1/22/2020"))
#print(parse_date("2020-03-01T23:45:50"))
```
### Read in data, old schema
Next challenge is that the schema did change, namely between 2020-03-21 and 2020-03-22. The column names have changed, new columns have been added and so on. Therefore we cannot read in all files within a single `spark.read.csv`, but we need to split them up into two separate batches with different schemas.
```
# Last date to read
today = datetime.date(2020, 4, 7) #datetime.date.today()
# First date to read
start_date = datetime.date(2020, 1, 22)
# First date with new schema
schema_switch_date = datetime.date(2020, 3, 22)
```
The first bunch of files is stored as CSV and has the following columns:
* `Province_State`
* `Country_Region`
* `Last_Update` date of the last update
* `Confirmed` the number of confirmed cases
* `Deaths` the number of confirmed cases, which have died
* `Recovered` the number of recovered cases
* `Latitude` and `Longitude` geo coordinates of the province
The metrics (confirmed, deaths and recovered) are always totals, they already contain all cases from the past.
```
daily_reports_dir = basedir + "/csse_covid_19_daily_reports"
# Define old schema for first batch of files
schema_1 = StructType([
StructField("Province_State", StringType()),
StructField("Country_Region", StringType()),
StructField("Last_Update", StringType()),
StructField("Confirmed", LongType()),
StructField("Deaths", LongType()),
StructField("Recovered", LongType()),
StructField("Latitude", DoubleType()),
StructField("Longitude", DoubleType()),
])
# Generate all dates with old schema
schema_1_dates = [start_date + datetime.timedelta(days=d) for d in range(0,(schema_switch_date - start_date).days)]
# Generate file names with old schema
schema_1_files = [daily_reports_dir + "/" + d.strftime("%m-%d-%Y") + ".csv" for d in schema_1_dates]
# Read in all files with old schema
cases_1 = spark.read\
.schema(schema_1) \
.option("header", True) \
.csv(schema_1_files)
# Peek inside
cases_1.toPandas()
```
### Read in data, new schema
Now we perform exactly the same logical step, we read in all files with the new schema. The second bunch of files is stored as CSV and has the following columns:
* `FIPS` country code
* `Admin2` administrative name below province (i.e. counties)
* `Province_State`
* `Country_Region`
* `Last_Update` date of the last update
* `Latitude` and `Longitude` geo coordinates of the province
* `Confirmed` the number of confirmed cases
* `Deaths` the number of confirmed cases, which have died
* `Recovered` the number of recovered cases
* `Active` the number of currently active cases
* `Combined_Key` a combination of `Admin2`, `Province_State` and `Country_Region`
The metrics (confirmed, deaths and recovered) are always totals, they already contain all cases from the past.
```
from pyspark.sql.types import *
daily_reports_dir = basedir + "/csse_covid_19_daily_reports"
# New schema
schema_2 = StructType([
StructField("FIPS", StringType()),
StructField("Admin2", StringType()),
StructField("Province_State", StringType()),
StructField("Country_Region", StringType()),
StructField("Last_Update", StringType()),
StructField("Latitude", DoubleType()),
StructField("Longitude", DoubleType()),
StructField("Confirmed", LongType()),
StructField("Deaths", LongType()),
StructField("Recovered", LongType()),
StructField("Active", LongType()),
StructField("Combined_Key", StringType())
])
# Generate all dates with new schema
schema_2_dates = [schema_switch_date + datetime.timedelta(days=d) for d in range(0,(today- schema_switch_date).days)]
# Generate file names with new schema
schema_2_files = [daily_reports_dir + "/" + d.strftime("%m-%d-%Y") + ".csv" for d in schema_2_dates]
# Read in all CSV files with new schema
cases_2 = spark.read\
.schema(schema_2) \
.option("header", True) \
.csv(schema_2_files)
cases_2.toPandas()
```
### Unify Records
Now we union both data sets `cases_1` and `cases_2` into a bigger data set with a common schema. The target schema should contain the following columns:
* `Country_Region`
* `Province_State`
* `Admin2`
* `Last_Update`
* `Confirmed`
* `Deaths`
* `Recovered`
In case a specific column is not present in onw of the two input DataFrames, simply provide a NULL value (`None` in Python) instead.
```
all_cases = \
cases_1.select(
f.col("Country_Region"),
f.col("Province_State"),
f.lit(None).cast(StringType()).alias("Admin2"),
f.col("Last_Update"),
f.col("Confirmed"),
f.col("Deaths"),
f.col("Recovered")
).union(
cases_2.select(
f.col("Country_Region"),
f.col("Province_State"),
f.col("Admin2"),
f.col("Last_Update"),
f.col("Confirmed"),
f.col("Deaths"),
f.col("Recovered")
)
)
all_cases.toPandas()
```
## 1.3 Aggragate
The records can contain multiple updates per day. But we only want to have the latest update per administrative region on each day. Therefore we perform a simple grouped aggregation and simply pick the maximum of all metrics of interest (`Confirmed`, `Deaths`, `Recovered`). This means we require a grouped aggregation with the grouping keys `Last_Update`, `Country_Region`, `Province_State` and `Admin2`.
```
all_cases_eod = # YOUR CODE HERE
all_cases_eod.show()
```
## 1.4 Sanity Checks
Since we have now a nice data set containing all records, lets peek inside and let us perform some sanity checks if the numbers are correct.
```
all_cases_eod.where(f.col("Country_Region") == f.lit("US")) \
.orderBy(f.col("Confirmed").desc()) \
.show(truncate=False)
```
### Count cases in US
Let us count the cases in the US for a specific date, maybe compare it to some resource on the web. This can be done by summing up all confirmed case where `Country_Region` equals to `US` and where `Last_Update` equals some date of your choice (for example `2020-04-05`).
```
# YOUR CODE HERE
```
### Count cases in Germany
Let us now sum up the confirmed cases for Germany.
```
all_cases_eod.where(f.col("Country_Region") == f.lit("Germany")) \
.where(f.col("Last_Update") == f.lit("2020-04-06")) \
.select(f.sum(f.col("Confirmed"))) \
.toPandas()
```
# 2. Inspect & Visualize
Now that we have a meaningful dataset, let us create some visualizations.
## 2.1 Additional Preparations
Before doing deeper analyzis, we still need to perform some simple preparations in order to make the resuls more meaningful.
### Cases pre country and day
We are not interested in the specific numbers of different provinces or states within a single country. The problem with data per province is that they may contain too few cases for following any theoretical law or for forming any meaningful probability distribution. Therefore we sum up all cases per country per day.
```
all_country_cases = # YOUR CODE HERE
```
### Calculate age in days
Before continuing, we will add one extra column, namely the day of the epedemy for every country. The desease started on different dates in different countries (start being defined as the date of the first record in the data set). To be able to compare the development of the desease between different countries, it is advisable to add a country specific `day` column, which simply counts the days since the first infection in the particular country.
```
from pyspark.sql.window import Window
all_countries_age_cases = all_country_cases \
.withColumn("First_Update", f.min(f.col("Last_Update")).over(Window.partitionBy("Country_Region").orderBy("Last_Update"))) \
.withColumn("day", f.datediff(f.col("Last_Update"), f.col("First_Update")))
all_countries_age_cases.show()
```
## 2.2 Pick single country
For the whole analysis, we focus on a single country. I decided to pick Germany, but you can also pick a different country.
The selection can easily be done by filtering using the column `Country_Region` to contain the desired country (for example `Germany`).
```
country_cases = all_countries_age_cases.where(f.col("Country_Region") == "Germany")
# Show first 10 days of data in the correct order
country_cases.orderBy(f.col("day")).show(10)
```
## 2.3 Plot
Let us make a simple plot which shows the number of cases over time. This can be achieved by using the matplotlib function `plt.plot`, which takes two arguments: the data on the horizontal axis (x-axis) and the data on the vertical axis (y-axis). One can also specify the size of the plot by using the function `plt.figure` as below.
```
df = country_cases.toPandas()
# Set size of the figure
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
# Make an image usig plt.plot
# YOUR CODE HERE
```
### Plot on logarithmic scale
The spread of a epedemy follows an exponential pattern (specifically at the beginning), this can also be seen from the plot above. Therefore it is a good idea to change the scale from a linear scale to a logarithmic scale. With the logarithmic scale, you can spot the relativ rate of increase, which is the slope of the curve. Changing the scale can easily be done with the function `plt.yscale('log')`.
```
df = country_cases.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
plt.yscale('log')
plt.plot(df["day"], df["Confirmed"], color='blue', lw=2)
```
### Plot of daily increases
In this case, we are also interested in the number of new cases on every day. This means, that we need to subtract the current number of confirmed cases from the last number of confirmed cases. This is a good example where so called *windowed aggregation* can help us in PySpark. Normally all rows of a DataFrame are processed independently, but for this question (the difference of the number of confirmed cases between two days), we would need to access the rows from two different days. That can be done with window functions.
```
daily_increase = country_cases.withColumn(
"Confirmed_Increase",
f.col("Confirmed") - f.last(f.col("Confirmed")).over(Window.partitionBy("Country_Region").orderBy("day").rowsBetween(-100,-1))
)
daily_increase.show(10)
```
Now we have an additional column "Confirmed_Increase", which we can now plot. A continuous line plot doesn't make so much sense, since the metric is very discrete by its nature. Therefore we opt for a bar chart instead by using the method `plt.bar` instead of `plt.plot`.
```
df = daily_increase.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
# Make a bar plot using plt.bar
# YOUR CODE HERE
```
# 3. Machine Learning
Now we want to use some methods of machine learning in order to predict the further development of the desease within a single country.
```
# Import relevant packages
from pyspark.ml import *
from pyspark.ml.feature import *
from pyspark.ml.regression import *
from pyspark.ml.evaluation import *
```
## 3.1 Split Data
The very first step in every machine learning project is to split up the whole data into two data sets, the *training data* and the *validation data*. The basic idea is that in order to validate our model, we need some data that was not used during training. That is what the *validation data* will be used for. If we did not exclude the data from the training, we could only infer information about how good the model fits to our data - but we would have no information about how good the model copes with new data. And the second aspect is crucial in prediction applications, where the model will be used on new data points which have never been seen before.
There are different approaches how to do that, but not all approaches work in every scenario. In our use case we are looking at a time series, therefore we need to split the data at a specific date - we need to hide out some information for the training phase. For time series it is important not to perform a random sampling, since this would imply information creep from the future. I.e. if we exclude day 40 and include day 50 in our training data set, day 50 obviously has some information on day 40, which normally would not be available.
```
all_days = country_cases.select("day").orderBy(f.col("day")).distinct().collect()
all_days = [row[0] for row in all_days]
num_days = len(all_days)
cut_day = all_days[int(0.7*num_days)]
print("cut_day = " + str(cut_day))
# We might want to skip some days where there was no real growth
first_day = 28
# Select training records from first_day until cut_day (inclusive)
training_records = # YOUR CODE HERE
# Select validation records from cut_day (exclusive)
validation_records = # YOUR CODE HERE
```
## 3.2 Simple Regression
The most simple approach is to use a very basic linear regression. We skip this super simple approach, since we already know that our data has some exponential ingredients. Therefore we already use a so called *generalized linear model* (GLM), which transforms our values into a logarithmic space before performing a linear regression. Here we already know that this won't work out nicely, since the plots above already indicate a curved shape over time - something a trivial linear model cannot catch. We will take care of that in a later step.
### PySpark ML Pipelines
Spark ML encourages to use so called *pipelines*, which combine a list of transformation blocks. For the very first very simple example, we need two building blocks:
* `VectorAssembler` is required to collect all features into a single column of the special type `Vector`. Most machine learning algorithms require the independant variables (the predictor variables) to packaged together into a single column of type `Vector`. This can be easily done by using the `VectorAssembler`-
* `GeneralizedLinearRegression` provides the regression algorithm a a building block. It needs to be configured with the indepedant variable (the features column), the dependant variable (the label column) and the prediction column where the predictions should be stored in.
```
pipeline = Pipeline(stages=[
# YOUR CODE HERE
])
```
### Fit model
Once we have specified all building blocks in the pipeline, we can *fit* the whole pipeline to obtain a *pipeline model*. The fitting operation either applies a transformation (like the `VectorAssembler`) or recursively fits any embedded estimator (like the `GeneralizedLinearRegression`).
```
model = # YOUR CODE HERE
```
### Perform prediction
One we have obtained the model, it can be used as a transformer again in order to produce predictions. For plotting a graph, we will apply the model not only to the validation set, but to the whole data set. This can be done with the `model.transform` method applied to the `country_cases` DataFrame.
```
pred = # YOUR CODE HERE
```
### Visualize
Finally we want to visualize the real values and the predictions.
```
df = pred.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
plt.yscale('log')
plt.plot(df["day"], df["Confirmed"], color='blue', lw=2)
plt.plot(df["day"], df["Predict"], color='red', lw=2)
plt.vlines(cut_day, ymin=0, ymax=400000)
```
### Measure Perfrmance
The picture already doesn't make much hope that the model generalizes well to new data. But in order to compare different models, we should also quantify the fit. We use a built in metric called *root mean squared error* provided by the class `RegressionEvaluator`. One an instance of the class is created, you can evaluate the predictions by using the `evaluate` function.
Since we are only interested in the ability to generalize, we use the `validation_records` DataFrame for measuring the qulity of fit.
```
evaluator = # YOUR CODE HERE
pred = model.transform(validation_records)
evaluator.evaluate(pred)
```
## 3.3 Improve Model
The first shot is not very satisfactory, specifically if looking at the logarithmic plot. The data seems to describe a curve (which is good), we could try to fit a polynom of order 2. This means that we will use (day^2, day, const) as features. This *polynomial expansion* of the original feature `day` can be generated by the `PolynomialExpansion` feature transformer.
This means that we will slightly extend our pipeline as follows.
```
pipeline = Pipeline(stages=[
# YOUR CODE HERE
])
```
### Fit and predict
Now we will again fit the pipeline to retrieve a model and immediately apply the model to all cases in order to get the data for another plot.
```
model = pipeline.fit(training_records)
pred = model.transform(country_cases)
```
### Visualize
The next visualization looks better, especially the critical part of the graph is estimated much better. Note that we did not use data before day 28, since there was no real growth before that day.
Note that our predicted values are above the measured values. This can mean multiple things:
* *Pessimist*: Our model does not perform as good as desired
* *Optimist*: Actions taken by politics change the real model parameters in a favorable way, such that the real number of infections do not grow any more as predicted
```
df = pred.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
plt.yscale('log')
plt.plot(df["day"], df["Confirmed"], color='blue', lw=2)
plt.plot(df["day"], df["Predict"], color='red', lw=2)
plt.vlines(cut_day, ymin=0, ymax=400000)
```
Although the image looks quite promising on the logarithmic scale, let us have a look at the linear scale. We will notice that we overpredict the number of cases by a factor of two and our prediction will look even worse for the future.
```
df = pred.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
plt.plot(df["day"], df["Confirmed"], color='blue', lw=2)
plt.plot(df["day"], df["Predict"], color='red', lw=2)
plt.vlines(cut_day, ymin=0, ymax=300000)
```
### Measure performance
Again we will use the `RegressionEvaluator` as before to quantify the prediction error. The error should be much lower now.
```
evaluator = RegressionEvaluator(
predictionCol='Predict',
labelCol='Confirmed',
metricName='rmse'
)
pred = model.transform(validation_records)
evaluator.evaluate(pred)
```
## 3.5 Change training/validation split
If we change the split of training to validation, things look much better. Of course this might be already expected, since we predict less data, but even the non-logarithmic plot looks really good
```
# Use 80% for training (it was 70% before)
cut_day = all_days[int(0.8*num_days)]
print("cut_day = " + str(cut_day))
training_records_80 = country_cases.where((f.col("day") <= cut_day) & (f.col("day") >= first_day)).cache()
validation_records_80 = country_cases.where(f.col("day") > cut_day).cache()
model = pipeline.fit(training_records_80)
pred = model.transform(country_cases)
df = pred.toPandas()
plt.figure(figsize=(16, 6), dpi=80, facecolor='w', edgecolor='k')
plt.plot(df["day"], df["Confirmed"], color='blue', lw=2)
plt.plot(df["day"], df["Predict"], color='red', lw=2)
plt.vlines(cut_day, ymin=0, ymax=120000)
```
This result should let you look optimistic into the future, as it may indicate that the underlying process really has changed between day 50 and 60 and that the infection really slows down.
### Measure performance
Again we will use the `RegressionEvaluator` as before to quantify the prediction error. The error should be much lower now.
```
evaluator = RegressionEvaluator(
predictionCol='Predict',
labelCol='Confirmed',
metricName='rmse'
)
pred = model.transform(validation_records)
evaluator.evaluate(pred)
```
# 4. Final Note
As already mentioned in the beginning, the whole approach is somewhat questionable. We are throwing a very generic machinery at a very specific problem which has a very specific structure. Therefore other approaches involving more meaningful models like https://de.wikipedia.org/wiki/SIR-Modell could give better prediction results. But those models require a completely different numerical approach for fitting the model to the data. We used the tool at hand (in this case PySpark) to generate a model, which does only make very mild (and possibly wrong) assumptions about the development process of the desease. Nevertheless such approaches might also give good results, since on the other hand specific mathematical models also rely on very specific assumptions and simplifications, which may also not be justified.
| github_jupyter |
# Skip-gram Word2Vec
In this notebook, I'll lead you through using PyTorch to implement the [Word2Vec algorithm](https://en.wikipedia.org/wiki/Word2vec) using the skip-gram architecture. By implementing this, you'll learn about embedding words for use in natural language processing. This will come in handy when dealing with things like machine translation.
## Readings
Here are the resources I used to build this notebook. I suggest reading these either beforehand or while you're working on this material.
* A really good [conceptual overview](http://mccormickml.com/2016/04/19/word2vec-tutorial-the-skip-gram-model/) of Word2Vec from Chris McCormick
* [First Word2Vec paper](https://arxiv.org/pdf/1301.3781.pdf) from Mikolov et al.
* [Neural Information Processing Systems, paper](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) with improvements for Word2Vec also from Mikolov et al.
---
## Word embeddings
When you're dealing with words in text, you end up with tens of thousands of word classes to analyze; one for each word in a vocabulary. Trying to one-hot encode these words is massively inefficient because most values in a one-hot vector will be set to zero. So, the matrix multiplication that happens in between a one-hot input vector and a first, hidden layer will result in mostly zero-valued hidden outputs.
To solve this problem and greatly increase the efficiency of our networks, we use what are called **embeddings**. Embeddings are just a fully connected layer like you've seen before. We call this layer the embedding layer and the weights are embedding weights. We skip the multiplication into the embedding layer by instead directly grabbing the hidden layer values from the weight matrix. We can do this because the multiplication of a one-hot encoded vector with a matrix returns the row of the matrix corresponding the index of the "on" input unit.
<img src='assets/lookup_matrix.png' width=50%>
Instead of doing the matrix multiplication, we use the weight matrix as a lookup table. We encode the words as integers, for example "heart" is encoded as 958, "mind" as 18094. Then to get hidden layer values for "heart", you just take the 958th row of the embedding matrix. This process is called an **embedding lookup** and the number of hidden units is the **embedding dimension**.
There is nothing magical going on here. The embedding lookup table is just a weight matrix. The embedding layer is just a hidden layer. The lookup is just a shortcut for the matrix multiplication. The lookup table is trained just like any weight matrix.
Embeddings aren't only used for words of course. You can use them for any model where you have a massive number of classes. A particular type of model called **Word2Vec** uses the embedding layer to find vector representations of words that contain semantic meaning.
---
## Word2Vec
The Word2Vec algorithm finds much more efficient representations by finding vectors that represent the words. These vectors also contain semantic information about the words.
<img src="assets/context_drink.png" width=40%>
Words that show up in similar **contexts**, such as "coffee", "tea", and "water" will have vectors near each other. Different words will be further away from one another, and relationships can be represented by distance in vector space.
There are two architectures for implementing Word2Vec:
>* CBOW (Continuous Bag-Of-Words) and
* Skip-gram
<img src="assets/word2vec_architectures.png" width=60%>
In this implementation, we'll be using the **skip-gram architecture** with **negative sampling** because it performs better than CBOW and trains faster with negative sampling. Here, we pass in a word and try to predict the words surrounding it in the text. In this way, we can train the network to learn representations for words that show up in similar contexts.
---
## Loading Data
Next, we'll ask you to load in data and place it in the `data` directory
1. Load the [text8 dataset](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/October/5bbe6499_text8/text8.zip); a file of cleaned up *Wikipedia article text* from Matt Mahoney.
2. Place that data in the `data` folder in the home directory.
3. Then you can extract it and delete the archive, zip file to save storage space.
After following these steps, you should have one file in your data directory: `data/text8`.
```
# read in the extracted text file
with open('data/text8') as f:
text = f.read()
# print out the first 100 characters
print(text[:100])
```
## Pre-processing
Here I'm fixing up the text to make training easier. This comes from the `utils.py` file. The `preprocess` function does a few things:
>* It converts any punctuation into tokens, so a period is changed to ` <PERIOD> `. In this data set, there aren't any periods, but it will help in other NLP problems.
* It removes all words that show up five or *fewer* times in the dataset. This will greatly reduce issues due to noise in the data and improve the quality of the vector representations.
* It returns a list of words in the text.
This may take a few seconds to run, since our text file is quite large. If you want to write your own functions for this stuff, go for it!
```
import utils
# get list of words
words = utils.preprocess(text)
print(words[:30])
# print some stats about this word data
print("Total words in text: {}".format(len(words)))
print("Unique words: {}".format(len(set(words)))) # `set` removes any duplicate words
```
### Dictionaries
Next, I'm creating two dictionaries to convert words to integers and back again (integers to words). This is again done with a function in the `utils.py` file. `create_lookup_tables` takes in a list of words in a text and returns two dictionaries.
>* The integers are assigned in descending frequency order, so the most frequent word ("the") is given the integer 0 and the next most frequent is 1, and so on.
Once we have our dictionaries, the words are converted to integers and stored in the list `int_words`.
```
vocab_to_int, int_to_vocab = utils.create_lookup_tables(words)
int_words = [vocab_to_int[word] for word in words]
print(int_words[:30])
```
## Subsampling
Words that show up often such as "the", "of", and "for" don't provide much context to the nearby words. If we discard some of them, we can remove some of the noise from our data and in return get faster training and better representations. This process is called subsampling by Mikolov. For each word $w_i$ in the training set, we'll discard it with probability given by
$$ P(w_i) = 1 - \sqrt{\frac{t}{f(w_i)}} $$
where $t$ is a threshold parameter and $f(w_i)$ is the frequency of word $w_i$ in the total dataset.
> Implement subsampling for the words in `int_words`. That is, go through `int_words` and discard each word given the probablility $P(w_i)$ shown above. Note that $P(w_i)$ is the probability that a word is discarded. Assign the subsampled data to `train_words`.
```
from collections import Counter
import random
import numpy as np
threshold = 1e-5
word_counts = Counter(int_words)
#print(list(word_counts.items())[0]) # dictionary of int_words, how many times they appear
total_count = len(int_words)
freqs = {word: count/total_count for word, count in word_counts.items()}
p_drop = {word: 1 - np.sqrt(threshold/freqs[word]) for word in word_counts}
# discard some frequent words, according to the subsampling equation
# create a new list of words for training
train_words = [word for word in int_words if random.random() < (1 - p_drop[word])]
print(train_words[:30])
```
## Making batches
Now that our data is in good shape, we need to get it into the proper form to pass it into our network. With the skip-gram architecture, for each word in the text, we want to define a surrounding _context_ and grab all the words in a window around that word, with size $C$.
From [Mikolov et al.](https://arxiv.org/pdf/1301.3781.pdf):
"Since the more distant words are usually less related to the current word than those close to it, we give less weight to the distant words by sampling less from those words in our training examples... If we choose $C = 5$, for each training word we will select randomly a number $R$ in range $[ 1: C ]$, and then use $R$ words from history and $R$ words from the future of the current word as correct labels."
> **Exercise:** Implement a function `get_target` that receives a list of words, an index, and a window size, then returns a list of words in the window around the index. Make sure to use the algorithm described above, where you chose a random number of words to from the window.
Say, we have an input and we're interested in the idx=2 token, `741`:
```
[5233, 58, 741, 10571, 27349, 0, 15067, 58112, 3580, 58, 10712]
```
For `R=2`, `get_target` should return a list of four values:
```
[5233, 58, 10571, 27349]
```
```
def get_target(words, idx, window_size=5):
''' Get a list of words in a window around an index. '''
R = np.random.randint(1, window_size+1)
start = idx - R if (idx - R) > 0 else 0
stop = idx + R
target_words = words[start:idx] + words[idx+1:stop+1]
return list(target_words)
# test your code!
# run this cell multiple times to check for random window selection
int_text = [i for i in range(10)]
print('Input: ', int_text)
idx=5 # word index of interest
target = get_target(int_text, idx=idx, window_size=5)
print('Target: ', target) # you should get some indices around the idx
```
### Generating Batches
Here's a generator function that returns batches of input and target data for our model, using the `get_target` function from above. The idea is that it grabs `batch_size` words from a words list. Then for each of those batches, it gets the target words in a window.
```
def get_batches(words, batch_size, window_size=5):
''' Create a generator of word batches as a tuple (inputs, targets) '''
n_batches = len(words)//batch_size
# only full batches
words = words[:n_batches*batch_size]
for idx in range(0, len(words), batch_size):
x, y = [], []
batch = words[idx:idx+batch_size]
for ii in range(len(batch)):
batch_x = batch[ii]
batch_y = get_target(batch, ii, window_size)
y.extend(batch_y)
x.extend([batch_x]*len(batch_y))
yield x, y
int_text = [i for i in range(20)]
x,y = next(get_batches(int_text, batch_size=4, window_size=5))
print('x\n', x)
print('y\n', y)
```
---
## Validation
Here, I'm creating a function that will help us observe our model as it learns. We're going to choose a few common words and few uncommon words. Then, we'll print out the closest words to them using the cosine similarity:
<img src="assets/two_vectors.png" width=30%>
$$
\mathrm{similarity} = \cos(\theta) = \frac{\vec{a} \cdot \vec{b}}{|\vec{a}||\vec{b}|}
$$
We can encode the validation words as vectors $\vec{a}$ using the embedding table, then calculate the similarity with each word vector $\vec{b}$ in the embedding table. With the similarities, we can print out the validation words and words in our embedding table semantically similar to those words. It's a nice way to check that our embedding table is grouping together words with similar semantic meanings.
```
def cosine_similarity(embedding, valid_size=16, valid_window=100, device='cpu'):
""" Returns the cosine similarity of validation words with words in the embedding matrix.
Here, embedding should be a PyTorch embedding module.
"""
# Here we're calculating the cosine similarity between some random words and
# our embedding vectors. With the similarities, we can look at what words are
# close to our random words.
# sim = (a . b) / |a||b|
embed_vectors = embedding.weight
# magnitude of embedding vectors, |b|
magnitudes = embed_vectors.pow(2).sum(dim=1).sqrt().unsqueeze(0)
# pick N words from our ranges (0,window) and (1000,1000+window). lower id implies more frequent
valid_examples = np.array(random.sample(range(valid_window), valid_size//2))
valid_examples = np.append(valid_examples,
random.sample(range(1000,1000+valid_window), valid_size//2))
valid_examples = torch.LongTensor(valid_examples).to(device)
valid_vectors = embedding(valid_examples)
similarities = torch.mm(valid_vectors, embed_vectors.t())/magnitudes
return valid_examples, similarities
```
---
# SkipGram model
Define and train the SkipGram model.
> You'll need to define an [embedding layer](https://pytorch.org/docs/stable/nn.html#embedding) and a final, softmax output layer.
An Embedding layer takes in a number of inputs, importantly:
* **num_embeddings** – the size of the dictionary of embeddings, or how many rows you'll want in the embedding weight matrix
* **embedding_dim** – the size of each embedding vector; the embedding dimension
Below is an approximate diagram of the general structure of our network.
<img src="assets/skip_gram_arch.png" width=60%>
>* The input words are passed in as batches of input word tokens.
* This will go into a hidden layer of linear units (our embedding layer).
* Then, finally into a softmax output layer.
We'll use the softmax layer to make a prediction about the context words by sampling, as usual.
---
## Negative Sampling
For every example we give the network, we train it using the output from the softmax layer. That means for each input, we're making very small changes to millions of weights even though we only have one true example. This makes training the network very inefficient. We can approximate the loss from the softmax layer by only updating a small subset of all the weights at once. We'll update the weights for the correct example, but only a small number of incorrect, or noise, examples. This is called ["negative sampling"](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf).
There are two modifications we need to make. First, since we're not taking the softmax output over all the words, we're really only concerned with one output word at a time. Similar to how we use an embedding table to map the input word to the hidden layer, we can now use another embedding table to map the hidden layer to the output word. Now we have two embedding layers, one for input words and one for output words. Secondly, we use a modified loss function where we only care about the true example and a small subset of noise examples.
$$
- \large \log{\sigma\left(u_{w_O}\hspace{0.001em}^\top v_{w_I}\right)} -
\sum_i^N \mathbb{E}_{w_i \sim P_n(w)}\log{\sigma\left(-u_{w_i}\hspace{0.001em}^\top v_{w_I}\right)}
$$
This is a little complicated so I'll go through it bit by bit. $u_{w_O}\hspace{0.001em}^\top$ is the embedding vector for our "output" target word (transposed, that's the $^\top$ symbol) and $v_{w_I}$ is the embedding vector for the "input" word. Then the first term
$$\large \log{\sigma\left(u_{w_O}\hspace{0.001em}^\top v_{w_I}\right)}$$
says we take the log-sigmoid of the inner product of the output word vector and the input word vector. Now the second term, let's first look at
$$\large \sum_i^N \mathbb{E}_{w_i \sim P_n(w)}$$
This means we're going to take a sum over words $w_i$ drawn from a noise distribution $w_i \sim P_n(w)$. The noise distribution is basically our vocabulary of words that aren't in the context of our input word. In effect, we can randomly sample words from our vocabulary to get these words. $P_n(w)$ is an arbitrary probability distribution though, which means we get to decide how to weight the words that we're sampling. This could be a uniform distribution, where we sample all words with equal probability. Or it could be according to the frequency that each word shows up in our text corpus, the unigram distribution $U(w)$. The authors found the best distribution to be $U(w)^{3/4}$, empirically.
Finally, in
$$\large \log{\sigma\left(-u_{w_i}\hspace{0.001em}^\top v_{w_I}\right)},$$
we take the log-sigmoid of the negated inner product of a noise vector with the input vector.
<img src="assets/neg_sampling_loss.png" width=50%>
To give you an intuition for what we're doing here, remember that the sigmoid function returns a probability between 0 and 1. The first term in the loss pushes the probability that our network will predict the correct word $w_O$ towards 1. In the second term, since we are negating the sigmoid input, we're pushing the probabilities of the noise words towards 0.
```
import torch
from torch import nn
import torch.optim as optim
class SkipGramNeg(nn.Module):
def __init__(self, n_vocab, n_embed, noise_dist=None):
super().__init__()
self.n_vocab = n_vocab
self.n_embed = n_embed
self.noise_dist = noise_dist
# define embedding layers for input and output words
self.in_embed = nn.Embedding(n_vocab, n_embed)
self.out_embed = nn.Embedding(n_vocab, n_embed)
# Initialize embedding tables with uniform distribution
# I believe this helps with convergence
self.in_embed.weight.data.uniform_(-1, 1)
self.out_embed.weight.data.uniform_(-1, 1)
def forward_input(self, input_words):
input_vectors = self.in_embed(input_words)
return input_vectors
def forward_output(self, output_words):
output_vectors = self.out_embed(output_words)
return output_vectors
def forward_noise(self, batch_size, n_samples):
""" Generate noise vectors with shape (batch_size, n_samples, n_embed)"""
if self.noise_dist is None:
# Sample words uniformly
noise_dist = torch.ones(self.n_vocab)
else:
noise_dist = self.noise_dist
# Sample words from our noise distribution
noise_words = torch.multinomial(noise_dist,
batch_size * n_samples,
replacement=True)
device = "cuda" if model.out_embed.weight.is_cuda else "cpu"
noise_words = noise_words.to(device)
noise_vectors = self.out_embed(noise_words).view(batch_size, n_samples, self.n_embed)
return noise_vectors
class NegativeSamplingLoss(nn.Module):
def __init__(self):
super().__init__()
def forward(self, input_vectors, output_vectors, noise_vectors):
batch_size, embed_size = input_vectors.shape
# Input vectors should be a batch of column vectors
input_vectors = input_vectors.view(batch_size, embed_size, 1)
# Output vectors should be a batch of row vectors
output_vectors = output_vectors.view(batch_size, 1, embed_size)
# bmm = batch matrix multiplication
# correct log-sigmoid loss
out_loss = torch.bmm(output_vectors, input_vectors).sigmoid().log()
out_loss = out_loss.squeeze()
# incorrect log-sigmoid loss
noise_loss = torch.bmm(noise_vectors.neg(), input_vectors).sigmoid().log()
noise_loss = noise_loss.squeeze().sum(1) # sum the losses over the sample of noise vectors
# negate and sum correct and noisy log-sigmoid losses
# return average batch loss
return -(out_loss + noise_loss).mean()
```
### Training
Below is our training loop, and I recommend that you train on GPU, if available.
```
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# Get our noise distribution
# Using word frequencies calculated earlier in the notebook
word_freqs = np.array(sorted(freqs.values(), reverse=True))
unigram_dist = word_freqs/word_freqs.sum()
noise_dist = torch.from_numpy(unigram_dist**(0.75)/np.sum(unigram_dist**(0.75)))
# instantiating the model
embedding_dim = 300
model = SkipGramNeg(len(vocab_to_int), embedding_dim, noise_dist=noise_dist).to(device)
# using the loss that we defined
criterion = NegativeSamplingLoss()
optimizer = optim.Adam(model.parameters(), lr=0.003)
print_every = 1500
steps = 0
epochs = 5
# train for some number of epochs
for e in range(epochs):
# get our input, target batches
for input_words, target_words in get_batches(train_words, 512):
steps += 1
inputs, targets = torch.LongTensor(input_words), torch.LongTensor(target_words)
inputs, targets = inputs.to(device), targets.to(device)
# input, output, and noise vectors
input_vectors = model.forward_input(inputs)
output_vectors = model.forward_output(targets)
noise_vectors = model.forward_noise(inputs.shape[0], 5)
# negative sampling loss
loss = criterion(input_vectors, output_vectors, noise_vectors)
optimizer.zero_grad()
loss.backward()
optimizer.step()
# loss stats
if steps % print_every == 0:
print("Epoch: {}/{}".format(e+1, epochs))
print("Loss: ", loss.item()) # avg batch loss at this point in training
valid_examples, valid_similarities = cosine_similarity(model.in_embed, device=device)
_, closest_idxs = valid_similarities.topk(6)
valid_examples, closest_idxs = valid_examples.to('cpu'), closest_idxs.to('cpu')
for ii, valid_idx in enumerate(valid_examples):
closest_words = [int_to_vocab[idx.item()] for idx in closest_idxs[ii]][1:]
print(int_to_vocab[valid_idx.item()] + " | " + ', '.join(closest_words))
print("...\n")
```
## Visualizing the word vectors
Below we'll use T-SNE to visualize how our high-dimensional word vectors cluster together. T-SNE is used to project these vectors into two dimensions while preserving local stucture. Check out [this post from Christopher Olah](http://colah.github.io/posts/2014-10-Visualizing-MNIST/) to learn more about T-SNE and other ways to visualize high-dimensional data.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
# getting embeddings from the embedding layer of our model, by name
embeddings = model.in_embed.weight.to('cpu').data.numpy()
viz_words = 380
tsne = TSNE()
embed_tsne = tsne.fit_transform(embeddings[:viz_words, :])
fig, ax = plt.subplots(figsize=(16, 16))
for idx in range(viz_words):
plt.scatter(*embed_tsne[idx, :], color='steelblue')
plt.annotate(int_to_vocab[idx], (embed_tsne[idx, 0], embed_tsne[idx, 1]), alpha=0.7)
```
| github_jupyter |
# Customer Churn Prediction with XGBoost
_**Using Gradient Boosted Trees to Predict Mobile Customer Departure**_
---
---
## Contents
1. [Background](#Background)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Compile](#Compile)
1. [Host](#Host)
1. [Evaluate](#Evaluate)
1. [Relative cost of errors](#Relative-cost-of-errors)
1. [Extensions](#Extensions)
---
## Background
_This notebook has been adapted from an [AWS blog post](https://aws.amazon.com/blogs/ai/predicting-customer-churn-with-amazon-machine-learning/)_
Losing customers is costly for any business. Identifying unhappy customers early on gives you a chance to offer them incentives to stay. This notebook describes using machine learning (ML) for the automated identification of unhappy customers, also known as customer churn prediction. ML models rarely give perfect predictions though, so this notebook is also about how to incorporate the relative costs of prediction mistakes when determining the financial outcome of using ML.
We use an example of churn that is familiar to all of us–leaving a mobile phone operator. Seems like I can always find fault with my provider du jour! And if my provider knows that I’m thinking of leaving, it can offer timely incentives–I can always use a phone upgrade or perhaps have a new feature activated–and I might just stick around. Incentives are often much more cost effective than losing and reacquiring a customer.
---
## Setup
_This notebook was created and tested on an ml.m4.xlarge notebook instance._
Let's start by specifying:
- The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
- The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
```
import sagemaker
sess = sagemaker.Session()
bucket=sess.default_bucket()
prefix = 'sagemaker/DEMO-xgboost-churn'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
```
Next, we'll import the Python libraries we'll need for the remainder of the exercise.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import os
import sys
import time
import json
from IPython.display import display
from time import strftime, gmtime
import sagemaker
from sagemaker.predictor import csv_serializer
```
---
## Data
Mobile operators have historical records on which customers ultimately ended up churning and which continued using the service. We can use this historical information to construct an ML model of one mobile operator’s churn using a process called training. After training the model, we can pass the profile information of an arbitrary customer (the same profile information that we used to train the model) to the model, and have the model predict whether this customer is going to churn. Of course, we expect the model to make mistakes–after all, predicting the future is tricky business! But I’ll also show how to deal with prediction errors.
The dataset we use is publicly available and was mentioned in the book [Discovering Knowledge in Data](https://www.amazon.com/dp/0470908742/) by Daniel T. Larose. It is attributed by the author to the University of California Irvine Repository of Machine Learning Datasets. Let's download and read that dataset in now:
```
!wget http://dataminingconsultant.com/DKD2e_data_sets.zip
!unzip -o DKD2e_data_sets.zip
churn = pd.read_csv('./Data sets/churn.txt')
pd.set_option('display.max_columns', 500)
churn
```
By modern standards, it’s a relatively small dataset, with only 3,333 records, where each record uses 21 attributes to describe the profile of a customer of an unknown US mobile operator. The attributes are:
- `State`: the US state in which the customer resides, indicated by a two-letter abbreviation; for example, OH or NJ
- `Account Length`: the number of days that this account has been active
- `Area Code`: the three-digit area code of the corresponding customer’s phone number
- `Phone`: the remaining seven-digit phone number
- `Int’l Plan`: whether the customer has an international calling plan: yes/no
- `VMail Plan`: whether the customer has a voice mail feature: yes/no
- `VMail Message`: presumably the average number of voice mail messages per month
- `Day Mins`: the total number of calling minutes used during the day
- `Day Calls`: the total number of calls placed during the day
- `Day Charge`: the billed cost of daytime calls
- `Eve Mins, Eve Calls, Eve Charge`: the billed cost for calls placed during the evening
- `Night Mins`, `Night Calls`, `Night Charge`: the billed cost for calls placed during nighttime
- `Intl Mins`, `Intl Calls`, `Intl Charge`: the billed cost for international calls
- `CustServ Calls`: the number of calls placed to Customer Service
- `Churn?`: whether the customer left the service: true/false
The last attribute, `Churn?`, is known as the target attribute–the attribute that we want the ML model to predict. Because the target attribute is binary, our model will be performing binary prediction, also known as binary classification.
Let's begin exploring the data:
```
# Frequency tables for each categorical feature
for column in churn.select_dtypes(include=['object']).columns:
display(pd.crosstab(index=churn[column], columns='% observations', normalize='columns'))
# Histograms for each numeric features
display(churn.describe())
%matplotlib inline
hist = churn.hist(bins=30, sharey=True, figsize=(10, 10))
```
We can see immediately that:
- `State` appears to be quite evenly distributed
- `Phone` takes on too many unique values to be of any practical use. It's possible parsing out the prefix could have some value, but without more context on how these are allocated, we should avoid using it.
- Only 14% of customers churned, so there is some class imabalance, but nothing extreme.
- Most of the numeric features are surprisingly nicely distributed, with many showing bell-like gaussianity. `VMail Message` being a notable exception (and `Area Code` showing up as a feature we should convert to non-numeric).
```
churn = churn.drop('Phone', axis=1)
churn['Area Code'] = churn['Area Code'].astype(object)
```
Next let's look at the relationship between each of the features and our target variable.
```
for column in churn.select_dtypes(include=['object']).columns:
if column != 'Churn?':
display(pd.crosstab(index=churn[column], columns=churn['Churn?'], normalize='columns'))
for column in churn.select_dtypes(exclude=['object']).columns:
print(column)
hist = churn[[column, 'Churn?']].hist(by='Churn?', bins=30)
plt.show()
```
Interestingly we see that churners appear:
- Fairly evenly distributed geographically
- More likely to have an international plan
- Less likely to have a voicemail plan
- To exhibit some bimodality in daily minutes (either higher or lower than the average for non-churners)
- To have a larger number of customer service calls (which makes sense as we'd expect customers who experience lots of problems may be more likely to churn)
In addition, we see that churners take on very similar distributions for features like `Day Mins` and `Day Charge`. That's not surprising as we'd expect minutes spent talking to correlate with charges. Let's dig deeper into the relationships between our features.
```
display(churn.corr())
pd.plotting.scatter_matrix(churn, figsize=(12, 12))
plt.show()
```
We see several features that essentially have 100% correlation with one another. Including these feature pairs in some machine learning algorithms can create catastrophic problems, while in others it will only introduce minor redundancy and bias. Let's remove one feature from each of the highly correlated pairs: Day Charge from the pair with Day Mins, Night Charge from the pair with Night Mins, Intl Charge from the pair with Intl Mins:
```
churn = churn.drop(['Day Charge', 'Eve Charge', 'Night Charge', 'Intl Charge'], axis=1)
```
Now that we've cleaned up our dataset, let's determine which algorithm to use. As mentioned above, there appear to be some variables where both high and low (but not intermediate) values are predictive of churn. In order to accommodate this in an algorithm like linear regression, we'd need to generate polynomial (or bucketed) terms. Instead, let's attempt to model this problem using gradient boosted trees. Amazon SageMaker provides an XGBoost container that we can use to train in a managed, distributed setting, and then host as a real-time prediction endpoint. XGBoost uses gradient boosted trees which naturally account for non-linear relationships between features and the target variable, as well as accommodating complex interactions between features.
Amazon SageMaker XGBoost can train on data in either a CSV or LibSVM format. For this example, we'll stick with CSV. It should:
- Have the predictor variable in the first column
- Not have a header row
But first, let's convert our categorical features into numeric features.
```
model_data = pd.get_dummies(churn)
model_data = pd.concat([model_data['Churn?_True.'], model_data.drop(['Churn?_False.', 'Churn?_True.'], axis=1)], axis=1)
```
And now let's split the data into training, validation, and test sets. This will help prevent us from overfitting the model, and allow us to test the models accuracy on data it hasn't already seen.
```
train_data, validation_data, test_data = np.split(model_data.sample(frac=1, random_state=1729), [int(0.7 * len(model_data)), int(0.9 * len(model_data))])
train_data.to_csv('train.csv', header=False, index=False)
validation_data.to_csv('validation.csv', header=False, index=False)
```
Now we'll upload these files to S3.
```
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation/validation.csv')).upload_file('validation.csv')
```
---
## Train
Moving onto training, first we'll need to specify the locations of the XGBoost algorithm containers.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost')
```
Then, because we're training with the CSV file format, we'll create `s3_input`s that our training function can use as a pointer to the files in S3.
```
s3_input_train = sagemaker.s3_input(s3_data='s3://{}/{}/train'.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.s3_input(s3_data='s3://{}/{}/validation/'.format(bucket, prefix), content_type='csv')
```
Now, we can specify a few parameters like what type of training instances we'd like to use and how many, as well as our XGBoost hyperparameters. A few key hyperparameters are:
- `max_depth` controls how deep each tree within the algorithm can be built. Deeper trees can lead to better fit, but are more computationally expensive and can lead to overfitting. There is typically some trade-off in model performance that needs to be explored between a large number of shallow trees and a smaller number of deeper trees.
- `subsample` controls sampling of the training data. This technique can help reduce overfitting, but setting it too low can also starve the model of data.
- `num_round` controls the number of boosting rounds. This is essentially the subsequent models that are trained using the residuals of previous iterations. Again, more rounds should produce a better fit on the training data, but can be computationally expensive or lead to overfitting.
- `eta` controls how aggressive each round of boosting is. Larger values lead to more conservative boosting.
- `gamma` controls how aggressively trees are grown. Larger values lead to more conservative models.
More detail on XGBoost's hyperparmeters can be found on their GitHub [page](https://github.com/dmlc/xgboost/blob/master/doc/parameter.md).
```
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
num_round=100)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
```
---
## Compile
[Amazon SageMaker Neo](https://aws.amazon.com/sagemaker/neo/) optimizes models to run up to twice as fast, with no loss in accuracy. When calling `compile_model()` function, we specify the target instance family (m4) as well as the S3 bucket to which the compiled model would be stored.
```
compiled_model = xgb
if xgb.create_model().check_neo_region(boto3.Session().region_name) is False:
print('Neo is not currently supported in', boto3.Session().region_name)
else:
output_path = '/'.join(xgb.output_path.split('/')[:-1])
compiled_model = xgb.compile_model(target_instance_family='ml_m4',
input_shape={'data': [1, 69]},
role=role,
framework='xgboost',
framework_version='0.7',
output_path=output_path)
compiled_model.name = 'deployed-xgboost-customer-churn'
compiled_model.image = get_image_uri(sess.boto_region_name, 'xgboost-neo', repo_version='latest')
```
---
## Host
Now that we've trained the algorithm, let's create a model and deploy it to a hosted endpoint.
```
xgb_predictor = compiled_model.deploy(initial_instance_count = 1, instance_type = 'ml.m4.xlarge')
```
### Evaluate
Now that we have a hosted endpoint running, we can make real-time predictions from our model very easily, simply by making an http POST request. But first, we'll need to setup serializers and deserializers for passing our `test_data` NumPy arrays to the model behind the endpoint.
```
xgb_predictor.content_type = 'text/csv'
xgb_predictor.serializer = csv_serializer
xgb_predictor.deserializer = None
```
Now, we'll use a simple function to:
1. Loop over our test dataset
1. Split it into mini-batches of rows
1. Convert those mini-batchs to CSV string payloads
1. Retrieve mini-batch predictions by invoking the XGBoost endpoint
1. Collect predictions and convert from the CSV output our model provides into a NumPy array
```
def predict(data, rows=500):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, xgb_predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
predictions = predict(test_data.to_numpy()[:,1:])
```
There are many ways to compare the performance of a machine learning model, but let's start by simply by comparing actual to predicted values. In this case, we're simply predicting whether the customer churned (`1`) or not (`0`), which produces a simple confusion matrix.
```
pd.crosstab(index=test_data.iloc[:, 0], columns=np.round(predictions), rownames=['actual'], colnames=['predictions'])
```
_Note, due to randomized elements of the algorithm, you results may differ slightly._
Of the 48 churners, we've correctly predicted 39 of them (true positives). And, we incorrectly predicted 4 customers would churn who then ended up not doing so (false positives). There are also 9 customers who ended up churning, that we predicted would not (false negatives).
An important point here is that because of the `np.round()` function above we are using a simple threshold (or cutoff) of 0.5. Our predictions from `xgboost` come out as continuous values between 0 and 1 and we force them into the binary classes that we began with. However, because a customer that churns is expected to cost the company more than proactively trying to retain a customer who we think might churn, we should consider adjusting this cutoff. That will almost certainly increase the number of false positives, but it can also be expected to increase the number of true positives and reduce the number of false negatives.
To get a rough intuition here, let's look at the continuous values of our predictions.
```
plt.hist(predictions)
plt.show()
```
The continuous valued predictions coming from our model tend to skew toward 0 or 1, but there is sufficient mass between 0.1 and 0.9 that adjusting the cutoff should indeed shift a number of customers' predictions. For example...
```
pd.crosstab(index=test_data.iloc[:, 0], columns=np.where(predictions > 0.3, 1, 0))
```
We can see that changing the cutoff from 0.5 to 0.3 results in 1 more true positives, 3 more false positives, and 1 fewer false negatives. The numbers are small overall here, but that's 6-10% of customers overall that are shifting because of a change to the cutoff. Was this the right decision? We may end up retaining 3 extra customers, but we also unnecessarily incentivized 5 more customers who would have stayed. Determining optimal cutoffs is a key step in properly applying machine learning in a real-world setting. Let's discuss this more broadly and then apply a specific, hypothetical solution for our current problem.
### Relative cost of errors
Any practical binary classification problem is likely to produce a similarly sensitive cutoff. That by itself isn’t a problem. After all, if the scores for two classes are really easy to separate, the problem probably isn’t very hard to begin with and might even be solvable with simple rules instead of ML.
More important, if I put an ML model into production, there are costs associated with the model erroneously assigning false positives and false negatives. I also need to look at similar costs associated with correct predictions of true positives and true negatives. Because the choice of the cutoff affects all four of these statistics, I need to consider the relative costs to the business for each of these four outcomes for each prediction.
#### Assigning costs
What are the costs for our problem of mobile operator churn? The costs, of course, depend on the specific actions that the business takes. Let's make some assumptions here.
First, assign the true negatives the cost of \$0. Our model essentially correctly identified a happy customer in this case, and we don’t need to do anything.
False negatives are the most problematic, because they incorrectly predict that a churning customer will stay. We lose the customer and will have to pay all the costs of acquiring a replacement customer, including foregone revenue, advertising costs, administrative costs, point of sale costs, and likely a phone hardware subsidy. A quick search on the Internet reveals that such costs typically run in the hundreds of dollars so, for the purposes of this example, let's assume \$500. This is the cost of false negatives.
Finally, for customers that our model identifies as churning, let's assume a retention incentive in the amount of \$100. If my provider offered me such a concession, I’d certainly think twice before leaving. This is the cost of both true positive and false positive outcomes. In the case of false positives (the customer is happy, but the model mistakenly predicted churn), we will “waste” the \$100 concession. We probably could have spent that \$100 more effectively, but it's possible we increased the loyalty of an already loyal customer, so that’s not so bad.
#### Finding the optimal cutoff
It’s clear that false negatives are substantially more costly than false positives. Instead of optimizing for error based on the number of customers, we should be minimizing a cost function that looks like this:
```txt
$500 * FN(C) + $0 * TN(C) + $100 * FP(C) + $100 * TP(C)
```
FN(C) means that the false negative percentage is a function of the cutoff, C, and similar for TN, FP, and TP. We need to find the cutoff, C, where the result of the expression is smallest.
A straightforward way to do this, is to simply run a simulation over a large number of possible cutoffs. We test 100 possible values in the for loop below.
```
cutoffs = np.arange(0.01, 1, 0.01)
costs = []
for c in cutoffs:
costs.append(np.sum(np.sum(np.array([[0, 100], [500, 100]]) *
pd.crosstab(index=test_data.iloc[:, 0],
columns=np.where(predictions > c, 1, 0)))))
costs = np.array(costs)
plt.plot(cutoffs, costs)
plt.show()
print('Cost is minimized near a cutoff of:', cutoffs[np.argmin(costs)], 'for a cost of:', np.min(costs))
```
The above chart shows how picking a threshold too low results in costs skyrocketing as all customers are given a retention incentive. Meanwhile, setting the threshold too high results in too many lost customers, which ultimately grows to be nearly as costly. The overall cost can be minimized at \$8400 by setting the cutoff to 0.46, which is substantially better than the \$20k+ I would expect to lose by not taking any action.
---
## Extensions
This notebook showcased how to build a model that predicts whether a customer is likely to churn, and then how to optimally set a threshold that accounts for the cost of true positives, false positives, and false negatives. There are several means of extending it including:
- Some customers who receive retention incentives will still churn. Including a probability of churning despite receiving an incentive in our cost function would provide a better ROI on our retention programs.
- Customers who switch to a lower-priced plan or who deactivate a paid feature represent different kinds of churn that could be modeled separately.
- Modeling the evolution of customer behavior. If usage is dropping and the number of calls placed to Customer Service is increasing, you are more likely to experience churn then if the trend is the opposite. A customer profile should incorporate behavior trends.
- Actual training data and monetary cost assignments could be more complex.
- Multiple models for each type of churn could be needed.
Regardless of additional complexity, similar principles described in this notebook are likely apply.
### (Optional) Clean-up
If you're ready to be done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
```
sagemaker.Session().delete_endpoint(xgb_predictor.endpoint)
```
| github_jupyter |
```
# Import packages and libraries
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
import matplotlib
import matplotlib.pyplot as plt
from IPython.display import display, HTML
# read data
df = pd.read_csv("C:\\Users\\LylionCj\\Desktop\\Github_Projects\\ChurnPrediction\\Churn.csv")
display(df.head(6))
# Futher explore the data
print("Number of rows: ", df.shape[0])
counts = df.describe().iloc[0]
display(
pd.DataFrame(
counts.tolist(),
columns=["Count of values"],
index=counts.index.values
).transpose()
)
# Delet the columns that won't be used in the model
df = df.drop(["Phone", "Area Code", "State"], axis=1)
features = df.drop(["Churn"], axis=1).columns
# Fitting the model
df_train, df_test = train_test_split(df, test_size=0.20)
# Set up our RandomForestClassifier instance and fit to data
clf = RandomForestClassifier(n_estimators=30)
clf.fit(df_train[features], df_train["Churn"])
# Make predictions
predictions = clf.predict(df_test[features])
probs = clf.predict_proba(df_test[features])
display(predictions)
# Model evaluation
score = clf.score(df_test[features], df_test["Churn"])
print("Accuracy: ", score)
# Construct a confusion matrix and and a ROC curve to dig further into the quality of model prediction results
get_ipython().magic('matplotlib inline')
confusion_matrix = pd.DataFrame(
confusion_matrix(df_test["Churn"], predictions),
columns=["Predicted False", "Predicted True"],
index=["Actual False", "Actual True"]
)
display(confusion_matrix)
# Calculate the fpr and tpr for all thresholds of the classification
fpr, tpr, threshold = roc_curve(df_test["Churn"], probs[:,1])
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, 'b')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# Plot feature importance to the following feature selection for the imporved incoming models
fig = plt.figure(figsize=(20, 18))
ax = fig.add_subplot(111)
df_f = pd.DataFrame(clf.feature_importances_, columns=["importance"])
df_f["labels"] = features
df_f.sort_values("importance", inplace=True, ascending=False)
display(df_f.head(5))
index = np.arange(len(clf.feature_importances_))
bar_width = 0.5
rects = plt.barh(index , df_f["importance"], bar_width, alpha=0.4, color='b', label='Main')
plt.yticks(index, df_f["labels"])
plt.show()
```
| github_jupyter |
# Computing the optimal road trip across the U.S.
This notebook provides the methodology and code used in the blog post, [Computing the optimal road trip across the U.S.](http://www.randalolson.com/2015/03/08/computing-the-optimal-road-trip-across-the-u-s/)
### Notebook by [Randal S. Olson](http://www.randalolson.com)
Please see the [repository README file](https://github.com/rhiever/Data-Analysis-and-Machine-Learning-Projects#license) for the licenses and usage terms for the instructional material and code in this notebook. In general, I have licensed this material so that it is as widely useable and shareable as possible.
The code in this notebook is also available as a single Python script [here](OptimalRoadTripHtmlSaveAndDisplay.py) courtesy of [Andrew Liesinger](https://github.com/AndrewLiesinger).
### Required Python libraries
If you don't have Python on your computer, you can use the [Anaconda Python distribution](http://continuum.io/downloads) to install most of the Python packages you need. Anaconda provides a simple double-click installer for your convenience.
This code uses base Python libraries except for `googlemaps` and `pandas` packages. You can install these packages using `pip` by typing the following commands into your command line:
> pip install pandas
> pip install googlemaps
If you're on a Mac, Linux, or Unix machine, you may need to type `sudo` before the command to install the package with administrator privileges.
### Construct a list of road trip waypoints
The first step is to decide where you want to stop on your road trip.
Make sure you look all of the locations up on [Google Maps](http://maps.google.com) first so you have the correct address, city, state, etc. If the text you use to look up the location doesn't work on Google Maps, then it won't work here either.
Add all of your waypoints to the list below. Make sure they're formatted the same way as in the example below.
*Technical note: Due to daily usage limitations of the Google Maps API, you can only have a maximum of 70 waypoints. You will have to pay Google for an increased API limit if you want to add more waypoints.*
```
all_waypoints = ["USS Alabama, Battleship Parkway, Mobile, AL",
"Grand Canyon National Park, Arizona",
"Toltec Mounds, Scott, AR",
"San Andreas Fault, San Benito County, CA",
"Cable Car Museum, 94108, 1201 Mason St, San Francisco, CA 94108",
"Pikes Peak, Colorado",
"The Mark Twain House & Museum, Farmington Avenue, Hartford, CT",
"New Castle Historic District, Delaware",
"White House, Pennsylvania Avenue Northwest, Washington, DC",
"Cape Canaveral, FL",
"Okefenokee Swamp Park, Okefenokee Swamp Park Road, Waycross, GA",
"Craters of the Moon National Monument & Preserve, Arco, ID",
"Lincoln Home National Historic Site Visitor Center, 426 South 7th Street, Springfield, IL",
"West Baden Springs Hotel, West Baden Avenue, West Baden Springs, IN",
"Terrace Hill, Grand Avenue, Des Moines, IA",
"C. W. Parker Carousel Museum, South Esplanade Street, Leavenworth, KS",
"Mammoth Cave National Park, Mammoth Cave Pkwy, Mammoth Cave, KY",
"French Quarter, New Orleans, LA",
"Acadia National Park, Maine",
"Maryland State House, 100 State Cir, Annapolis, MD 21401",
"USS Constitution, Boston, MA",
"Olympia Entertainment, Woodward Avenue, Detroit, MI",
"Fort Snelling, Tower Avenue, Saint Paul, MN",
"Vicksburg National Military Park, Clay Street, Vicksburg, MS",
"Gateway Arch, Washington Avenue, St Louis, MO",
"Glacier National Park, West Glacier, MT",
"Ashfall Fossil Bed, Royal, NE",
"Hoover Dam, NV",
"Omni Mount Washington Resort, Mount Washington Hotel Road, Bretton Woods, NH",
"Congress Hall, Congress Place, Cape May, NJ 08204",
"Carlsbad Caverns National Park, Carlsbad, NM",
"Statue of Liberty, Liberty Island, NYC, NY",
"Wright Brothers National Memorial Visitor Center, Manteo, NC",
"Fort Union Trading Post National Historic Site, Williston, North Dakota 1804, ND",
"Spring Grove Cemetery, Spring Grove Avenue, Cincinnati, OH",
"Chickasaw National Recreation Area, 1008 W 2nd St, Sulphur, OK 73086",
"Columbia River Gorge National Scenic Area, Oregon",
"Liberty Bell, 6th Street, Philadelphia, PA",
"The Breakers, Ochre Point Avenue, Newport, RI",
"Fort Sumter National Monument, Sullivan's Island, SC",
"Mount Rushmore National Memorial, South Dakota 244, Keystone, SD",
"Graceland, Elvis Presley Boulevard, Memphis, TN",
"The Alamo, Alamo Plaza, San Antonio, TX",
"Bryce Canyon National Park, Hwy 63, Bryce, UT",
"Shelburne Farms, Harbor Road, Shelburne, VT",
"Mount Vernon, Fairfax County, Virginia",
"Hanford Site, Benton County, WA",
"Lost World Caverns, Lewisburg, WV",
"Taliesin, County Road C, Spring Green, Wisconsin",
"Yellowstone National Park, WY 82190"]
```
Next you'll have to register this script with the Google Maps API so they know who's hitting their servers with hundreds of Google Maps routing requests.
1) Enable the Google Maps Distance Matrix API on your Google account. Google explains how to do that [here](https://github.com/googlemaps/google-maps-services-python#api-keys).
2) Copy and paste the API key they had you create into the code below.
```
import googlemaps
gmaps = googlemaps.Client(key="PASTE YOUR API KEY HERE")
```
Now we're going to query the Google Maps API for the shortest route between all of the waypoints.
This is equivalent to doing Google Maps directions lookups on the Google Maps site, except now we're performing hundreds of lookups automatically using code.
If you get an error on this part, that most likely means one of the waypoints you entered couldn't be found on Google Maps. Another possible reason for an error here is if it's not possible to drive between the points, e.g., finding the driving directions between Hawaii and Florida will return an error until we invent flying cars.
### Gather the distance traveled on the shortest route between all waypoints
```
from itertools import combinations
waypoint_distances = {}
waypoint_durations = {}
for (waypoint1, waypoint2) in combinations(all_waypoints, 2):
try:
route = gmaps.distance_matrix(origins=[waypoint1],
destinations=[waypoint2],
mode="driving", # Change this to "walking" for walking directions,
# "bicycling" for biking directions, etc.
language="English",
units="metric")
# "distance" is in meters
distance = route["rows"][0]["elements"][0]["distance"]["value"]
# "duration" is in seconds
duration = route["rows"][0]["elements"][0]["duration"]["value"]
waypoint_distances[frozenset([waypoint1, waypoint2])] = distance
waypoint_durations[frozenset([waypoint1, waypoint2])] = duration
except Exception as e:
print("Error with finding the route between %s and %s." % (waypoint1, waypoint2))
```
Now that we have the routes between all of our waypoints, let's save them to a text file so we don't have to bother Google about them again.
```
with open("my-waypoints-dist-dur.tsv", "w") as out_file:
out_file.write("\t".join(["waypoint1",
"waypoint2",
"distance_m",
"duration_s"]))
for (waypoint1, waypoint2) in waypoint_distances.keys():
out_file.write("\n" +
"\t".join([waypoint1,
waypoint2,
str(waypoint_distances[frozenset([waypoint1, waypoint2])]),
str(waypoint_durations[frozenset([waypoint1, waypoint2])])]))
```
### Use a genetic algorithm to optimize the order to visit the waypoints in
Instead of exhaustively looking at every possible solution, genetic algorithms start with a handful of random solutions and continually tinkers with these solutions — always trying something slightly different from the current solutions and keeping the best ones — until they can’t find a better solution any more.
Below, all you need to do is make sure that the file name above matches the file name below (both currently `my-waypoints-dist-dur.tsv`) and run the code. The code will read in your route information and use a genetic algorithm to discover an optimized driving route.
```
import pandas as pd
import numpy as np
waypoint_distances = {}
waypoint_durations = {}
all_waypoints = set()
waypoint_data = pd.read_csv("my-waypoints-dist-dur.tsv", sep="\t")
for i, row in waypoint_data.iterrows():
waypoint_distances[frozenset([row.waypoint1, row.waypoint2])] = row.distance_m
waypoint_durations[frozenset([row.waypoint1, row.waypoint2])] = row.duration_s
all_waypoints.update([row.waypoint1, row.waypoint2])
import random
def compute_fitness(solution):
"""
This function returns the total distance traveled on the current road trip.
The genetic algorithm will favor road trips that have shorter
total distances traveled.
"""
solution_fitness = 0.0
for index in range(len(solution)):
waypoint1 = solution[index - 1]
waypoint2 = solution[index]
solution_fitness += waypoint_distances[frozenset([waypoint1, waypoint2])]
return solution_fitness
def generate_random_agent():
"""
Creates a random road trip from the waypoints.
"""
new_random_agent = list(all_waypoints)
random.shuffle(new_random_agent)
return tuple(new_random_agent)
def mutate_agent(agent_genome, max_mutations=3):
"""
Applies 1 - `max_mutations` point mutations to the given road trip.
A point mutation swaps the order of two waypoints in the road trip.
"""
agent_genome = list(agent_genome)
num_mutations = random.randint(1, max_mutations)
for mutation in range(num_mutations):
swap_index1 = random.randint(0, len(agent_genome) - 1)
swap_index2 = swap_index1
while swap_index1 == swap_index2:
swap_index2 = random.randint(0, len(agent_genome) - 1)
agent_genome[swap_index1], agent_genome[swap_index2] = agent_genome[swap_index2], agent_genome[swap_index1]
return tuple(agent_genome)
def shuffle_mutation(agent_genome):
"""
Applies a single shuffle mutation to the given road trip.
A shuffle mutation takes a random sub-section of the road trip
and moves it to another location in the road trip.
"""
agent_genome = list(agent_genome)
start_index = random.randint(0, len(agent_genome) - 1)
length = random.randint(2, 20)
genome_subset = agent_genome[start_index:start_index + length]
agent_genome = agent_genome[:start_index] + agent_genome[start_index + length:]
insert_index = random.randint(0, len(agent_genome) + len(genome_subset) - 1)
agent_genome = agent_genome[:insert_index] + genome_subset + agent_genome[insert_index:]
return tuple(agent_genome)
def generate_random_population(pop_size):
"""
Generates a list with `pop_size` number of random road trips.
"""
random_population = []
for agent in range(pop_size):
random_population.append(generate_random_agent())
return random_population
def run_genetic_algorithm(generations=5000, population_size=100):
"""
The core of the Genetic Algorithm.
`generations` and `population_size` must be a multiple of 10.
"""
population_subset_size = int(population_size / 10.)
generations_10pct = int(generations / 10.)
# Create a random population of `population_size` number of solutions.
population = generate_random_population(population_size)
# For `generations` number of repetitions...
for generation in range(generations):
# Compute the fitness of the entire current population
population_fitness = {}
for agent_genome in population:
if agent_genome in population_fitness:
continue
population_fitness[agent_genome] = compute_fitness(agent_genome)
# Take the top 10% shortest road trips and produce offspring each from them
new_population = []
for rank, agent_genome in enumerate(sorted(population_fitness,
key=population_fitness.get)[:population_subset_size]):
if (generation % generations_10pct == 0 or generation == generations - 1) and rank == 0:
print("Generation %d best: %d | Unique genomes: %d" % (generation,
population_fitness[agent_genome],
len(population_fitness)))
print(agent_genome)
print("")
# Create 1 exact copy of each of the top road trips
new_population.append(agent_genome)
# Create 2 offspring with 1-3 point mutations
for offspring in range(2):
new_population.append(mutate_agent(agent_genome, 3))
# Create 7 offspring with a single shuffle mutation
for offspring in range(7):
new_population.append(shuffle_mutation(agent_genome))
# Replace the old population with the new population of offspring
for i in range(len(population))[::-1]:
del population[i]
population = new_population
```
Try running the genetic algorithm a few times to see the different solutions it comes up with. It should take about a minute to finish running.
```
run_genetic_algorithm(generations=5000, population_size=100)
```
### Visualize your road trip on a Google map
Now that we have an ordered list of the waypoints, we should put them on a Google map so we can see the trip from a high level and make any extra adjustments.
There's no easy way to make this visualization in Python, but the Google Maps team provides a nice JavaScript library for visualizing routes on a Google Map.
Here's an example map with the route between 50 waypoints visualized: [link](http://rhiever.github.io/optimal-roadtrip-usa/major-landmarks.html)
The tricky part here is that the JavaScript library only plots routes with a maximum of 10 waypoints. If we want to plot a route with >10 waypoints, we need to call the route plotting function multiple times.
Thanks to some optimizations by [Nicholas Clarke](https://github.com/nicholasgodfreyclarke) to my original map, this is a simple operation:
1) Copy the final route generated by the genetic algorithm above.
2) Place brackets (`[` & `]`) around the route, e.g.,
['Graceland, Elvis Presley Boulevard, Memphis, TN', 'Vicksburg National Military Park, Clay Street, Vicksburg, MS', 'French Quarter, New Orleans, LA', 'USS Alabama, Battleship Parkway, Mobile, AL', 'Cape Canaveral, FL', 'Okefenokee Swamp Park, Okefenokee Swamp Park Road, Waycross, GA', "Fort Sumter National Monument, Sullivan's Island, SC", 'Wright Brothers National Memorial Visitor Center, Manteo, NC', 'Congress Hall, Congress Place, Cape May, NJ 08204', 'Shelburne Farms, Harbor Road, Shelburne, VT', 'Omni Mount Washington Resort, Mount Washington Hotel Road, Bretton Woods, NH', 'Acadia National Park, Maine', 'USS Constitution, Boston, MA', 'The Breakers, Ochre Point Avenue, Newport, RI', 'The Mark Twain House & Museum, Farmington Avenue, Hartford, CT', 'Statue of Liberty, Liberty Island, NYC, NY', 'Liberty Bell, 6th Street, Philadelphia, PA', 'New Castle Historic District, Delaware', 'Maryland State House, 100 State Cir, Annapolis, MD 21401', 'White House, Pennsylvania Avenue Northwest, Washington, DC', 'Mount Vernon, Fairfax County, Virginia', 'Lost World Caverns, Lewisburg, WV', 'Olympia Entertainment, Woodward Avenue, Detroit, MI', 'Spring Grove Cemetery, Spring Grove Avenue, Cincinnati, OH', 'Mammoth Cave National Park, Mammoth Cave Pkwy, Mammoth Cave, KY', 'West Baden Springs Hotel, West Baden Avenue, West Baden Springs, IN', 'Gateway Arch, Washington Avenue, St Louis, MO', 'Lincoln Home National Historic Site Visitor Center, 426 South 7th Street, Springfield, IL', 'Taliesin, County Road C, Spring Green, Wisconsin', 'Fort Snelling, Tower Avenue, Saint Paul, MN', 'Terrace Hill, Grand Avenue, Des Moines, IA', 'C. W. Parker Carousel Museum, South Esplanade Street, Leavenworth, KS', 'Ashfall Fossil Bed, Royal, NE', 'Mount Rushmore National Memorial, South Dakota 244, Keystone, SD', 'Fort Union Trading Post National Historic Site, Williston, North Dakota 1804, ND', 'Glacier National Park, West Glacier, MT', 'Yellowstone National Park, WY 82190', 'Craters of the Moon National Monument & Preserve, Arco, ID', 'Hanford Site, Benton County, WA', 'Columbia River Gorge National Scenic Area, Oregon', 'Cable Car Museum, 94108, 1201 Mason St, San Francisco, CA 94108', 'San Andreas Fault, San Benito County, CA', 'Hoover Dam, NV', 'Grand Canyon National Park, Arizona', 'Bryce Canyon National Park, Hwy 63, Bryce, UT', 'Pikes Peak, Colorado', 'Carlsbad Caverns National Park, Carlsbad, NM', 'The Alamo, Alamo Plaza, San Antonio, TX', 'Chickasaw National Recreation Area, 1008 W 2nd St, Sulphur, OK 73086', 'Toltec Mounds, Scott, AR']
3) Paste the final route with brackets into [line 93](https://github.com/rhiever/optimal-roadtrip-usa/blob/gh-pages/major-landmarks.html#L93) of my road trip map code. It should look like this:
optimal_route = [ ... ]
where `...` is your optimized road trip.
That's all it takes! Now you have your own optimized road trip ready to show off to the world.
### Some technical notes
As I mentioned in the [original article](http://www.randalolson.com/2015/03/08/computing-the-optimal-road-trip-across-the-u-s/), by the end of 5,000 generations, the genetic algorithm will very likely find a *good* but probably not the *absolute best* solution to the optimal routing problem. It is in the nature of genetic algorithms that we never know if we found the absolute best solution.
However, there exist some brilliant analytical solutions to the optimal routing problem such as the [Concorde TSP solver](http://en.wikipedia.org/wiki/Concorde_TSP_Solver). If you're interested in learning more about Concorde and how it's possible to find a perfect solution to the routing problem, I advise you check out [Bill Cook's article](http://www.math.uwaterloo.ca/tsp/usa50/index.html) on the topic.
### If you have any questions
Please feel free to:
* [Email me](http://www.randalolson.com/contact/),
* [Tweet](https://twitter.com/randal_olson) at me, or
* comment on the [blog post](http://www.randalolson.com/2015/03/08/computing-the-optimal-road-trip-across-the-u-s/)
I'm usually pretty good about getting back to people within a day or two.
| github_jupyter |
# Testing if a Distribution is Normal
## Imports
```
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
import quiz_tests
# Set plotting options
%matplotlib inline
plt.rc('figure', figsize=(16, 9))
```
## Create normal and non-normal distributions
```
# Sample A: Normal distribution
sample_a = stats.norm.rvs(loc=0.0, scale=1.0, size=(1000,))
# Sample B: Non-normal distribution
sample_b = stats.lognorm.rvs(s=0.5, loc=0.0, scale=1.0, size=(1000,))
```
## Boxplot-Whisker Plot and Histogram
We can visually check if a distribution looks normally distributed. Recall that a box whisker plot lets us check for symmetry around the mean. A histogram lets us see the overall shape. A QQ-plot lets us compare our data distribution with a normal distribution (or any other theoretical "ideal" distribution).
```
# Sample A: Normal distribution
sample_a = stats.norm.rvs(loc=0.0, scale=1.0, size=(1000,))
fig, axes = plt.subplots(2, 1, figsize=(16, 9), sharex=True)
axes[0].boxplot(sample_a, vert=False)
axes[1].hist(sample_a, bins=50)
axes[0].set_title("Boxplot of a Normal Distribution");
# Sample B: Non-normal distribution
sample_b = stats.lognorm.rvs(s=0.5, loc=0.0, scale=1.0, size=(1000,))
fig, axes = plt.subplots(2, 1, figsize=(16, 9), sharex=True)
axes[0].boxplot(sample_b, vert=False)
axes[1].hist(sample_b, bins=50)
axes[0].set_title("Boxplot of a Lognormal Distribution");
# Q-Q plot of normally-distributed sample
plt.figure(figsize=(10, 10)); plt.axis('equal')
stats.probplot(sample_a, dist='norm', plot=plt);
# Q-Q plot of non-normally-distributed sample
plt.figure(figsize=(10, 10)); plt.axis('equal')
stats.probplot(sample_b, dist='norm', plot=plt);
```
## Testing for Normality
### Shapiro-Wilk
The Shapiro-Wilk test is available in the scipy library. The null hypothesis assumes that the data distribution is normal. If the p-value is greater than the chosen p-value, we'll assume that it's normal. Otherwise we assume that it's not normal.
https://docs.scipy.org/doc/scipy-0.19.0/reference/generated/scipy.stats.shapiro.html
```
def is_normal(sample, test=stats.shapiro, p_level=0.05, **kwargs):
"""Apply a normality test to check if sample is normally distributed."""
t_stat, p_value = test(sample, **kwargs)
print("Test statistic: {}, p-value: {}".format(t_stat, p_value))
print("Is the distribution Likely Normal? {}".format(p_value > p_level))
return p_value > p_level
# Using Shapiro-Wilk test (default)
print("Sample A:-"); is_normal(sample_a);
print("Sample B:-"); is_normal(sample_b);
```
## Kolmogorov-Smirnov
The Kolmogorov-Smirnov is available in the scipy.stats library. The K-S test compares the data distribution with a theoretical distribution. We'll choose the 'norm' (normal) distribution as the theoretical distribution, and we also need to specify the mean and standard deviation of this theoretical distribution. We'll set the mean and stanadard deviation of the theoretical norm with the mean and standard deviation of the data distribution.
https://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.kstest.html
# Quiz
To use the Kolmogorov-Smirnov test, complete the function `is_normal_ks`.
To set the variable normal_args, create a tuple with two values. An example of a tuple is `("apple","banana")`
The first is the mean of the sample. The second is the standard deviation of the sample.
**hint:** Hint: Numpy has functions np.mean() and np.std()
```
def is_normal_ks(sample, test=stats.kstest, p_level=0.05, **kwargs):
"""
sample: a sample distribution
test: a function that tests for normality
p_level: if the test returns a p-value > than p_level, assume normality
return: True if distribution is normal, False otherwise
"""
normal_args = (np.mean(sample),np.std(sample))
t_stat, p_value = test(sample, 'norm', normal_args, **kwargs)
print("Test statistic: {}, p-value: {}".format(t_stat, p_value))
print("Is the distribution Likely Normal? {}".format(p_value > p_level))
return p_value > p_level
quiz_tests.test_is_normal_ks(is_normal_ks)
# Using Kolmogorov-Smirnov test
print("Sample A:-"); is_normal_ks(sample_a);
print("Sample B:-"); is_normal_ks(sample_b);
```
If you're stuck, you can also check out the solution [here](test_normality_solution.ipynb)
| github_jupyter |
## Tutorial 104: Generate 21 ADME Predictors with 10 Lines of Code
[Kexin](https://twitter.com/KexinHuang5)
In the previous set of tutorials, hopefully, you are now familiarized with TDC. In this tutorial, we show through examples how to use TDC for fast ML model prototyping using DeepPurpose. Let's start introducing what is DeepPurpose.
### DeepPurpose Overview
DeepPurpose is a scikit learn style Deep Learning Based Molecular Modeling and Prediction Toolkit on Drug-Target Interaction Prediction, Compound Property Prediction, Protein-Protein Interaction Prediction, and Protein Function prediction. Using DeepPurpose, we can rapidly build model prototypes for various drug discovery tasks covered in TDC, such as ADME, Tox, HTS, Developability prediction, DTI, DDI, PPI, Antibody Affinity predictions.
Note that DeepPurpose is developed by two of the core teams in TDC, Kexin and Tianfan, and it is now published in Bioinformatics. To start with this tutorial, please follow [DeepPurpose instructions](https://github.com/kexinhuang12345/DeepPurpose#install--usage) to set up the necessary packages. DeepPurpose also provides [tutorials](https://github.com/kexinhuang12345/DeepPurpose/blob/master/Tutorial_1_DTI_Prediction.ipynb) for you to familiarize with it.
To install DeepPurpose, in your terminal, do the following:
```bash
git clone https://github.com/kexinhuang12345/DeepPurpose.git
cd DeepPurpose
conda env create -f environment.yml
conda activate DeepPurpose
pip install PyTDC --upgrade
```
And then open this jupyter notebook in the DeepPurpose directory.
We assume now you have set up the right environment. Now, we show you how to build an ADME predictor using Message Passing Neural Network (MPNN)!
### Predicting HIA using MPNN with 10 Lines of Code
First, let's load DeepPurpose and TDC:
```
from DeepPurpose import utils, CompoundPred
from tdc.single_pred import ADME
```
Now, you can get the HIA dataset from TDC. HIA is from ADME task from Single-instance prediction and we want to predict whether or not can a compound be absorped in human intestinal, i.e. given SMILES X, predict 1/0. Note that for drug property prediction, DeepPurpose takes in an array of drug SMILES string and an array of labels. You could access that directly by setting the `get_data(format = 'dict')`:
```
data = ADME(name = 'HIA_Hou').get_data(format = 'dict')
X, y = data['Drug'], data['Y']
```
or for simplicity. We also provide a DeepPurpose format, where you can directly get the correct input data:
```
X, y = ADME(name = 'HIA_Hou').get_data(format = 'DeepPurpose')
```
DeepPurpose provides 8 encoders for compound, ranging from MLP on classic cheminformatics fingerprint such as Morgan, RDKit2D to deep learning models such as CNN, transformer, and MPNN. To specify the encoder, simply types the encoder name. Here, we use MPNN as an example:
```
drug_encoding = 'MPNN'
```
Now, we encode the data into the specified format, using `utils.data_process` function. It specifies train/validation/test split fractions, and random seed to ensure same data splits for reproducibility. **We have made DeepPurpose to accomodate the TDC benchmark split.** Simply type 'TDC' in the random seed will generate the same split as in TDC split function. The function outputs train, val, test pandas dataframes.
```
train, val, test = utils.data_process(X_drug = X,
y = y,
drug_encoding = drug_encoding,
random_seed = 'TDC')
train.head(2)
```
After we have the dataset, you can set up the model configuration such as dimension, # of layers, optimization parameters and etc. As an example, we set the following configurations (we set train epochs to be 3 here for demonstration purpose):
```
config = utils.generate_config(drug_encoding = drug_encoding,
train_epoch = 3,
LR = 0.001,
batch_size = 128,
mpnn_hidden_size = 32,
mpnn_depth = 2
)
```
Now, you need to initialize the model. You can also load from pretrained model by `CompoundPred.model_pretrained(path_dir = PATH)`. Here, we initialize a new model:
```
model = CompoundPred.model_initialize(**config)
```
That's it. Now, you can train it!
```
# Training
model.train(train, val, test)
```
In less than 1 minute with a CPU, you can already get pretty good performance! Now, you can save the model by typing:
```
model.save_model('./tutorial_model')
```
You may already notice that the whole thing can be streamlined for all the ADME tasks in TDC by simply specifying different dataset names. To generate a full set of ADME predictors, you can do:
```
from tdc.utils import retrieve_dataset_names
adme_datasets = retrieve_dataset_names('ADME')
for dataset_name in adme_datasets:
X, y = ADME(name = dataset_name).get_data(format = 'DeepPurpose')
drug_encoding = 'MPNN'
train, val, test = utils.data_process(X_drug = X,
y = y,
drug_encoding = drug_encoding,
random_seed = 'TDC')
config = utils.generate_config(drug_encoding = drug_encoding,
train_epoch = 5,
LR = 0.001,
batch_size = 128,
mpnn_hidden_size = 32,
mpnn_depth = 2
)
model = CompoundPred.model_initialize(**config)
model.train(train, val, test)
model.save_model('./' + dataset_name + '_model')
```
We have tested the above code and it would print out all the training process. We omit the output here since it is a bit long. But hopefully, you could get the gist of it. DeepPurpose also allows to do similar modeling for DTI, DDI, PPI and protein function prediction! Checkout the [repository](https://github.com/kexinhuang12345/DeepPurpose)!
That's it for this tutorial!
| github_jupyter |
# Introduction to Data Science
# Lecture 15 continued
```
import numpy as np
from sklearn import svm, metrics
from sklearn.datasets import make_moons, load_iris
from sklearn.model_selection import train_test_split, cross_val_score, cross_validate
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
%matplotlib inline
plt.rcParams['figure.figsize'] = (10, 6)
plt.style.use('ggplot')
import seaborn as sns
sns.set()
```
## Classification
Recall that in *classification* we attempt to predict a categorical variable based on several features or attributes.
We've already seen three methods for classification:
1. Logistic Regression
+ k Nearest Neighbors (k-NN)
+ Decision Trees
Another method for classification is [Support Vector Machines (SVM)](https://en.wikipedia.org/wiki/Support_vector_machine). These methods are especially good at classifying small to medium sized, complex datasets.
We'll use the [sk-learn implementation](https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html).
## Support Vector Machines
**Idea:** Use tools from optimization to find the *best* lines (or hyperplanes or curves) that divide the two datasets.
### Simplest version first: 2 classes and no "kernel transformation"
**Data:** Given $n$ samples $(\vec{x}_1,y_1), (\vec{x}_2,y_2),\ldots,(\vec{x}_n,y_n)$, where $\vec{x}_i$ are attributes or features and $y_i$ are categorical variables that you want to predict. We assume that each $\vec{x}_i$ is a real vector and the $y_i$ are either $1$ or $-1$ indicating the class.
**Goal:** Find the "maximum-margin line" or more generally, the "maximum-margin hyperplane" that divides the two classes, which is defined so that the distance between the hyperplane and the nearest point in either group is maximized.
<img src="https://upload.wikimedia.org/wikipedia/commons/2/2a/Svm_max_sep_hyperplane_with_margin.png" width="400">
$\qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad \qquad$
source: [wikipedia](https://en.wikipedia.org/wiki/Support_vector_machine)
We can write any line as the set of points $\vec{x}$ satisfying
$$\vec{w}\cdot\vec{x} = b$$
for some (normal) vector $\vec{w}$ and number $b$. The goal is to determine $\vec{w}$ and $b$. Once we have the best values of $\vec{w}$ $b$, our classifier is simply
$$
\vec{x} \mapsto \textrm{sgn}(\vec{w} \cdot \vec{x} - b).
$$
**Hard-margin.** If the data is linearly separable, then we find the separating line (hyperplane) with the *largest margin* ($b/\|w\|$).
This is given by the solution to the optimization problem
\begin{align*}
\min_{\vec{w},b} & \ \|\vec{w}\|^2 \\
\textrm{ subject to } & \ y_i(\vec{w}\cdot\vec{x}_i - b) \ge 1 , \textrm{for } i = 1,\,\ldots,\,n
\end{align*}
This can be written as a *convex optimization problem* and efficiently solved. Take Math 5770/6640: Introduction to Optimization to learn more! In this class, we'll simply use scikit-learn.
From the picture, the max-margin hyperplane is determined by the $\vec{x}_i$ which are closest to it. These $\vec{x}_i$ are called *support vectors*.
**Soft-margin.** For data that is not linearly separable (e.g., two moons dataset), we introduce a *penalty* or *loss* function for violating the constraint, $y_i(\vec{w}\cdot\vec{x}_i + b) \ge 1$. One of these is the *hinge loss* function, given by
$$
g(\vec{x}_i; \vec{w},b) = \max\left(0, 1-y_i(\vec{w}\cdot\vec{x}_i - b)\right).
$$
We can see that this function is zero if the constraint is satisfied. If the constraint is not satisfied, we pay a penatly, which is proportional to the distance to the separating hyperplane.
We then fix the parameter $C > 0$ and solve the problem
$$
\min_{\vec{w},b} \ \frac{1}{n} \sum_{i=1}^n g(\vec{x}_i; \vec{w},b) \ + \ C \|\vec{w}\|^2 .
$$
The parameter $C$ determines how much we penalize points for being on the wrong side of the line.
**Question:** How to choose the parameter $C$? Cross Validation! (more on this below)
## SVM and two moons
```
# there are two features contained in X and the labels are contained in y
X,y = make_moons(n_samples=500,random_state=1,noise=0.3)
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="DarkBlue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="DarkRed", marker="o",label="class 2")
plt.legend(scatterpoints=1)
plt.title('Two Moons Dataset')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
model = svm.SVC(kernel='linear', C=10)
# Note or a 'linear' kernel, there is a faster method:
#
# from sklearn.svm import LinearSVC
# model = LinearSVC(C=10,loss="hinge")
model.fit(X, y)
print(model)
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="DarkBlue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="DarkRed", marker="o",label="class 2")
plt.legend(scatterpoints=1)
# Plot the predictions made by SVM
x_min, x_max = X[:,0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 200),np.linspace(y_min, y_max, 200))
zz = model.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.contourf(xx, yy, zz, cmap=ListedColormap(['DarkRed', 'DarkBlue']), alpha=.2)
plt.contour(xx, yy, zz, colors="black", alpha=1, linewidths=0.2)
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Classification of Two Moons using SVM')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
print('Confusion Matrix:')
y_pred = model.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Accuracy = ', metrics.accuracy_score(y_true = y, y_pred = y_pred))
```
## Support Vector Machines
**Basic idea:** The dataset may not be linearly separable, but it may be if we apply a *nonlinear kernel transformation*.
**Example:** two "rings" of points
**Algorithmic challenge:** Not increasing the computationally complexity of the optimization problem.
**(Very) rough sketch:**
Replace the linear classifier
$$
\vec{x} \mapsto \textrm{sgn}(\vec{w} \cdot \vec{x} - b).
$$
with a nonlinear one
$$
\vec{x} \mapsto \textrm{sgn}( f(x)).
$$
Choices of the nonlinear transformation in scikit-learn are:
- linear: $\langle x, x'\rangle$.
- polynomial: $(\gamma \langle x, x'\rangle + r)^d$. $d$ is specified by keyword degree, $r$ by coef0.
- rbf: $\exp(-\gamma |x-x'|^2)$. $\gamma$ is specified by keyword gamma, must be greater than 0.
- sigmoid: $\tanh(\gamma \langle x,x'\rangle + r)$, where $r$ is specified by coef0.
The most popular choice is rbf.
```
model = svm.SVC(kernel='rbf',C=20,gamma='scale')
model.fit(X, y)
# Plot the data, color by class
plt.scatter(X[y == 1, 0], X[y == 1, 1], color="DarkBlue", marker="s",label="class 1")
plt.scatter(X[y == 0, 0], X[y == 0, 1], color="DarkRed", marker="o",label="class 2")
plt.legend(scatterpoints=1)
# Plot the predictions made by SVM
x_min, x_max = X[:,0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 200),np.linspace(y_min, y_max, 200))
zz = model.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.contourf(xx, yy, zz, cmap=ListedColormap(['DarkRed', 'DarkBlue']), alpha=.2)
plt.contour(xx, yy, zz, colors="black", alpha=1, linewidths=0.2)
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Classification of Two Moons using SVM')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
print('Confusion Matrix:')
y_pred = model.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Accuracy = ', metrics.accuracy_score(y_true = y, y_pred = y_pred))
```
## More than 2 classes?
There are a few different methods for extending binary classification to more than 2 classes.
- **one vs. one:** Consider all pairs of classes (if there are $k$ classes, then there are $\binom k 2$ different pairs of classes. For each pair, we develop a classifier. For a new point, we perform all classifications and then choose the class that was most frequently assigned. (The classifiers all vote on a class.)
- **all vs. one:** We compare one of the $k$ classes to the remaining $k-1$ classes. We assign the class to the observation which we are most confident (we have to make this precise).
### Dataset: The Iris dataset
This dataset contains 4 features (attributes) of 50 samples containing 3 different species of iris plants. The goal is to classify the species of iris plant given the attributes.
**Classes:**
1. Iris Setosa
+ Iris Versicolour
+ Iris Virginica
**Features (attributes):**
1. sepal length (cm)
+ sepal width (cm)
+ petal length (cm)
+ petal width (cm)
<img src="iris.png" title="http://mirlab.org/jang/books/dcpr/dataSetIris.asp?title=2-2%20Iris%20Dataset" width="20%">
```
df = sns.load_dataset("iris") # built-in dataset in seaborn
sns.pairplot(df, hue="species");
# import data, scikit-learn also has this dataset built-in
iris = load_iris()
# For easy plotting and interpretation, we only use first 2 features here.
# We're throwing away useful information - don't do this at home!
X = iris.data[:,:2]
y = iris.target
# Create color maps
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
# plot data
plt.scatter(X[:, 0], X[:, 1], c=y, marker="o", cmap=cmap_bold, s=30)
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.title('Iris dataset')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.show()
# SVM
svm_iris = svm.SVC(kernel='rbf',C=3,gamma='scale')
svm_iris.fit(X, y)
# plot classification
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 200),np.linspace(y_min, y_max, 200))
zz = svm_iris.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, zz, cmap=cmap_light)
# plot data
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cmap_bold, s=30)
plt.title('Classification of Iris dataset using SVM')
plt.xlabel('feature 1')
plt.ylabel('feature 2')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.show()
print('Confusion Matrix:')
y_pred = svm_iris.predict(X)
print(metrics.confusion_matrix(y_true = y, y_pred = y_pred))
print('Accuracy = ', metrics.accuracy_score(y_true = y, y_pred = y_pred))
```
## Cross Validation
**Recall:** Cross validation is a method for assessing how well a model will generalize to an independent data set.
**Idea:** We split the dataset into a *training set* and a *test set*. We train the model on the training set. We measure the accuracy of the model on the test set.
There are several different ways to do this. Two popular methods are:
- **k-fold cross validation** The data is randomly partitioned into k (approximately) equal sized subsamples (folds). For each of the k folds, the method is trained on the other k-1 folds and tested on that fold. The accuracy is computed using each of the k folds as the test set.
- **leave-one-out cross validation** Same as above with k=n
A variety of function are implemented in scikit-learn for cross validation.
- The [train_test_split](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function randomly splits the data for cross validation
- The [cross_val_score](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html) function splits the data and measures accuracy on the test set. The parameter *cv* is the k in k-fold cross validation. The parameter *scoring* specifies how you want to [evaluate the model](https://scikit-learn.org/stable/modules/model_evaluation.html) (we can just use accuracy).
- The [cross_validate](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_validate.html) function evaluate metric(s) by cross-validation and also record fit/score times. So, cross-validate is different from cross_val_score in two ways:
1. It allows specifying multiple metrics for evaluation.
+ It returns a dict containing fit-times, score-times (and optionally training scores as well as fitted estimators) in addition to the test score.
```
# note: here we are again using all 4 features of the data
scores = cross_val_score(estimator = svm_iris, X = iris.data, y = iris.target, cv=5, scoring='accuracy')
print(scores)
scoring = ['accuracy','precision_macro','recall_macro']
scores = cross_validate(estimator = svm_iris, X = iris.data, y = iris.target, cv=5, scoring=scoring)
print(sorted(scores.keys()))
print(scores['test_accuracy'])
print(scores['fit_time'])
print(scores['test_accuracy'])
print(scores['test_recall_macro'])
print(scores['test_precision_macro'])
```
## Incorporating cross-validation to choose SVM parameter, $C$
Recall above that when the data is not linearly separable, introduce a loss function $g$ which penalizes the violation of datapoints lying on the wrong side of the hyperplane and solve the problem
$$
\min_{\vec{w},b} \ \frac{1}{n} \sum_{i=1}^n g(x_i; \vec{w},b) \ + \ C \|\vec{w}\| .
$$
Here, the parameter $C>0$ determines how much we penalize points for being on the wrong side of the line.
Generally, the function *get_params()* an be used to see what parameters are available to change in a model.
We can use cross-validation to choose the parameter $C$.
```
svm_iris.get_params()
Cs = np.linspace(.01,200,100)
Accuracies = np.zeros(Cs.shape[0])
for i,C in enumerate(Cs):
svm_iris = svm.SVC(kernel='rbf', C = C,gamma='scale')
scores = cross_val_score(estimator = svm_iris, X = iris.data, y = iris.target, cv=5, scoring='accuracy')
Accuracies[i] = scores.mean()
plt.plot(Cs,Accuracies)
plt.show()
```
Based on the cross validation results, I would choose $C\approx 3$.
| github_jupyter |
# CNTK 101: Logistic Regression and ML Primer
This tutorial is targeted to individuals who are new to CNTK and to machine learning. In this tutorial, you will train a simple yet powerful machine learning model that is widely used in industry for a variety of applications. The model trained below scales to massive data sets in the most expeditious manner by harnessing computational scalability leveraging the computational resources you may have (one or more CPU cores, one or more GPUs, a cluster of CPUs or a cluster of GPUs), transparently via the CNTK library.
The following notebook users Python APIs. If you are looking for this example in BrainScript, please look [here](https://github.com/Microsoft/CNTK/tree/v2.0.beta6.0/Tutorials/HelloWorld-LogisticRegression).
## Introduction
**Problem**:
A cancer hospital has provided data and wants us to determine if a patient has a fatal [malignant][] cancer vs. a benign growth. This is known as a classification problem. To help classify each patient, we are given their age and the size of the tumor. Intuitively, one can imagine that younger patients and/or patient with small tumor size are less likely to have malignant cancer. The data set simulates this application where the each observation is a patient represented as a dot (in the plot below) where red color indicates malignant and blue indicates benign disease. Note: This is a toy example for learning, in real life there are large number of features from different tests/examination sources and doctors' experience that play into the diagnosis/treatment decision for a patient.
<img src="https://www.cntk.ai/jup/cancer_data_plot.jpg", width=400, height=400>
**Goal**:
Our goal is to learn a classifier that automatically can label any patient into either benign or malignant category given two features (age and tumor size). In this tutorial, we will create a linear classifier that is a fundamental building-block in deep networks.
<img src="https://www.cntk.ai/jup/cancer_classify_plot.jpg", width=400, height=400>
In the figure above, the green line represents the learnt model from the data and separates the blue dots from the red dots. In this tutorial, we will walk you through the steps to learn the green line. Note: this classifier does make mistakes where couple of blue dots are on the wrong side of the green line. However, there are ways to fix this and we will look into some of the techniques in later tutorials.
**Approach**:
Any learning algorithm has typically five stages. These are Data reading, Data preprocessing, Creating a model, Learning the model parameters, and Evaluating (a.k.a. testing/prediction) the model.
>1. Data reading: We generate simulated data sets with each sample having two features (plotted below) indicative of the age and tumor size.
>2. Data preprocessing: Often the individual features such as size or age needs to be scaled. Typically one would scale the data between 0 and 1. To keep things simple, we are not doing any scaling in this tutorial (for details look here: [feature scaling][]).
>3. Model creation: We introduce a basic linear model in this tutorial.
>4. Learning the model: This is also known as training. While fitting a linear model can be done in a variety of ways ([linear regression][]), in CNTK we use Stochastic Gradient Descent a.k.a. [SGD][].
>5. Evaluation: This is also known as testing where one takes data sets with known labels (a.k.a. ground-truth) that was not ever used for training. This allows us to assess how a model would perform in real world (previously unseen) observations.
## Logistic Regression
[Logistic regression][] is fundamental machine learning technique that uses a linear weighted combination of features and generates the probability of predicting different classes. In our case the classifier will generate a probability in [0,1] which can then be compared with a threshold (such as 0.5) to produce a binary label (0 or 1). However, the method shown can be extended to multiple classes easily.
<img src="https://www.cntk.ai/jup/logistic_neuron.jpg", width=300, height=200>
In the figure above, contributions from different input features are linearly weighted and aggregated. The resulting sum is mapped to a 0-1 range via a [sigmoid][] function. For classifiers with more than two output labels, one can use a [softmax][] function.
[malignant]: https://en.wikipedia.org/wiki/Malignancy
[feature scaling]: https://en.wikipedia.org/wiki/Feature_scaling
[SGD]: https://en.wikipedia.org/wiki/Stochastic_gradient_descent
[linear regression]: https://en.wikipedia.org/wiki/Linear_regression
[logistic regression]: https://en.wikipedia.org/wiki/Logistic_regression
[softmax]: https://en.wikipedia.org/wiki/Multinomial_logistic_regression
[sigmoid]: https://en.wikipedia.org/wiki/Sigmoid_function
```
# Import the relevant components
from __future__ import print_function
import numpy as np
import sys
import os
from cntk import Trainer, learning_rate_schedule, UnitType
from cntk.device import cpu, set_default_device
from cntk.learner import sgd
from cntk.ops import *
```
## Data Generation
Let us generate some synthetic data emulating the cancer example using `numpy` library. We have two features (represented in two-dimensions) each either being to one of the two classes (benign:blue dot or malignant:red dot).
In our example, each observation in the training data has a label (blue or red) corresponding to each observation (set of features - age and size). In this example, we have two classes represented by labels 0 or 1, thus a binary classification task.
```
# Define the network
input_dim = 2
num_output_classes = 2
```
### Input and Labels
In this tutorial we are generating synthetic data using `numpy` library. In real world problems, one would use a [reader][], that would read feature values (`features`: *age* and *tumor size*) corresponding to each obeservation (patient). The simulated *age* variable is scaled down to have similar range as the other variable. This is a key aspect of data pre-processing that we will learn more in later tutorials. Note, each observation can reside in a higher dimension space (when more features are available) and will be represented as a [tensor][] in CNTK. More advanced tutorials shall introduce the handling of high dimensional data.
[reader]: https://github.com/Microsoft/CNTK/search?p=1&q=reader&type=Wikis&utf8=%E2%9C%93
[tensor]: https://en.wikipedia.org/wiki/Tensor
```
# Ensure we always get the same amount of randomness
np.random.seed(0)
# Helper function to generate a random data sample
def generate_random_data_sample(sample_size, feature_dim, num_classes):
# Create synthetic data using NumPy.
Y = np.random.randint(size=(sample_size, 1), low=0, high=num_classes)
# Make sure that the data is separable
X = (np.random.randn(sample_size, feature_dim)+3) * (Y+1)
# Specify the data type to match the input variable used later in the tutorial
# (default type is double)
X = X.astype(np.float32)
# converting class 0 into the vector "1 0 0",
# class 1 into vector "0 1 0", ...
class_ind = [Y==class_number for class_number in range(num_classes)]
Y = np.asarray(np.hstack(class_ind), dtype=np.float32)
return X, Y
# Create the input variables denoting the features and the label data. Note: the input_variable
# does not need additional info on number of observations (Samples) since CNTK creates only
# the network topology first
mysamplesize = 32
features, labels = generate_random_data_sample(mysamplesize, input_dim, num_output_classes)
```
Let us visualize the input data.
**Note**: If the import of `matplotlib.pyplot` fails, please run `conda install matplotlib` which will fix the `pyplot` version dependencies. If you are on a python environment different from Anaconda, then use `pip install`.
```
# Plot the data
import matplotlib.pyplot as plt
%matplotlib inline
# given this is a 2 class ()
colors = ['r' if l == 0 else 'b' for l in labels[:,0]]
plt.scatter(features[:,0], features[:,1], c=colors)
plt.xlabel("Scaled age (in yrs)")
plt.ylabel("Tumor size (in cm)")
plt.show()
```
# Model Creation
A logistic regression (a.k.a. LR) network is the simplest building block but has been powering many ML
applications in the past decade. LR is a simple linear model that takes as input, a vector of numbers describing the properties of what we are classifying (also known as a feature vector, $\bf{x}$, the blue nodes in the figure) and emits the *evidence* ($z$) (output of the green node, a.k.a. as activation). Each feature in the input layer is connected with a output node by a corresponding weight w (indicated by the black lines of varying thickness).
<img src="https://www.cntk.ai/jup/logistic_neuron2.jpg", width=300, height=200>
The first step is to compute the evidence for an observation.
$$z = \sum_{i=1}^n w_i \times x_i + b = \textbf{w} \cdot \textbf{x} + b$$
where $\bf{w}$ is the weight vector of length $n$ and $b$ is known as the [bias][] term. Note: we use **bold** notation to denote vectors.
The computed evidence is mapped to a 0-1 scale using a [`sigmoid`][] (when the outcome can take one of two values) or a `softmax` function (when the outcome can take one of more than 2 classes value).
Network input and output:
- **input** variable (a key CNTK concept):
>An **input** variable is a user-code facing container where user-provided code fills in different observations (data point or sample, equivalent to a blue/red dot in our example) as inputs to the model function during model learning (a.k.a.training) and model evaluation (a.k.a. testing). Thus, the shape of the `input_variable` must match the shape of the data that will be provided. For example, when data are images each of height 10 pixels and width 5 pixels, the input feature dimension will be 2 (representing image height and width). Similarly, in our example the dimensions are age and tumor size, thus `input_dim` = 2. More on data and their dimensions to appear in separate tutorials.
[bias]: https://www.quora.com/What-does-the-bias-term-represent-in-logistic-regression
[`sigmoid`]: https://en.wikipedia.org/wiki/Sigmoid_function
```
input = input_variable(input_dim, np.float32)
```
## Network setup
The `linear_layer` function is a straight forward implementation of the equation above. We perform two operations:
0. multiply the weights ($\bf{w}$) with the features ($\bf{x}$) using CNTK `times` operator and add individual features' contribution,
1. add the bias term $b$.
These CNTK operations are optimized for execution on the available hardware and the implementation hides the complexity away from the user.
```
# Define a dictionary to store the model parameters
mydict = {"w":None,"b":None}
def linear_layer(input_var, output_dim):
input_dim = input_var.shape[0]
weight_param = parameter(shape=(input_dim, output_dim))
bias_param = parameter(shape=(output_dim))
mydict['w'], mydict['b'] = weight_param, bias_param
return times(input_var, weight_param) + bias_param
```
`z` will be used to represent the output of a network.
```
output_dim = num_output_classes
z = linear_layer(input, output_dim)
```
### Learning model parameters
Now that the network is setup, we would like to learn the parameters $\bf w$ and $b$ for our simple linear layer. To do so we convert, the computed evidence ($z$) into a set of predicted probabilities ($\textbf p$) using a `softmax` function.
$$ \textbf{p} = \mathrm{softmax}(z)$$
The `softmax` is an activation function that maps the accumulated evidences to a probability distribution over the classes (Details of the [softmax function][]). Other choices of activation function can be [found here][].
[softmax function]: https://www.cntk.ai/pythondocs/cntk.ops.html?highlight=softmax#cntk.ops.softmax
[found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
## Training
The output of the `softmax` is a probability of observations belonging to the respective classes. For training the classifier, we need to determine what behavior the model needs to mimic. In other words, we want the generated probabilities to be as close as possible to the observed labels. This function is called the *cost* or *loss* function and shows what is the difference between the learnt model vs. that generated by the training set.
[`Cross-entropy`][] is a popular function to measure the loss. It is defined as:
$$ H(p) = - \sum_{j=1}^C y_j \log (p_j) $$
where $p$ is our predicted probability from `softmax` function and $y$ represents the label. This label provided with the data for training is also called the ground-truth label. In the two-class example, the `label` variable has dimensions of two (equal to the `num_output_classes` or $C$). Generally speaking, if the task in hand requires classification into $C$ different classes, the label variable will have $C$ elements with 0 everywhere except for the class represented by the data point where it will be 1. Understanding the [details][] of this cross-entropy function is highly recommended.
[`cross-entropy`]: http://cntk.ai/pythondocs/cntk.ops.html#cntk.ops.cross_entropy_with_softmax
[details]: http://colah.github.io/posts/2015-09-Visual-Information/
```
label = input_variable((num_output_classes), np.float32)
loss = cross_entropy_with_softmax(z, label)
```
#### Evaluation
In order to evaluate the classification, one can compare the output of the network which for each observation emits a vector of evidences (can be converted into probabilities using `softmax` functions) with dimension equal to number of classes.
```
eval_error = classification_error(z, label)
```
### Configure training
The trainer strives to reduce the `loss` function by different optimization approaches, [Stochastic Gradient Descent][] (`sgd`) being one of the most popular one. Typically, one would start with random initialization of the model parameters. The `sgd` optimizer would calculate the `loss` or error between the predicted label against the corresponding ground-truth label and using [gradient-decent][] generate a new set model parameters in a single iteration.
The aforementioned model parameter update using a single observation at a time is attractive since it does not require the entire data set (all observation) to be loaded in memory and also requires gradient computation over fewer datapoints, thus allowing for training on large data sets. However, the updates generated using a single observation sample at a time can vary wildly between iterations. An intermediate ground is to load a small set of observations and use an average of the `loss` or error from that set to update the model parameters. This subset is called a *minibatch*.
With minibatches we often sample observation from the larger training dataset. We repeat the process of model parameters update using different combination of training samples and over a period of time minimize the `loss` (and the error). When the incremental error rates are no longer changing significantly or after a preset number of maximum minibatches to train, we claim that our model is trained.
One of the key parameter for optimization is called the `learning_rate`. For now, we can think of it as a scaling factor that modulates how much we change the parameters in any iteration. We will be covering more details in later tutorial.
With this information, we are ready to create our trainer.
[optimization]: https://en.wikipedia.org/wiki/Category:Convex_optimization
[Stochastic Gradient Descent]: https://en.wikipedia.org/wiki/Stochastic_gradient_descent
[gradient-decent]: http://www.statisticsviews.com/details/feature/5722691/Getting-to-the-Bottom-of-Regression-with-Gradient-Descent.html
```
# Instantiate the trainer object to drive the model training
learning_rate = 0.5
lr_schedule = learning_rate_schedule(learning_rate, UnitType.minibatch)
learner = sgd(z.parameters, lr_schedule)
trainer = Trainer(z, loss, eval_error, [learner])
```
First let us create some helper functions that will be needed to visualize different functions associated with training. Note these convinience functions are for understanding what goes under the hood.
```
from cntk.utils import get_train_eval_criterion, get_train_loss
# Define a utility function to compute the moving average sum.
# A more efficient implementation is possible with np.cumsum() function
def moving_average(a, w=10):
if len(a) < w:
return a[:]
return [val if idx < w else sum(a[(idx-w):idx])/w for idx, val in enumerate(a)]
# Defines a utility that prints the training progress
def print_training_progress(trainer, mb, frequency, verbose=1):
training_loss, eval_error = "NA", "NA"
if mb % frequency == 0:
training_loss = get_train_loss(trainer)
eval_error = get_train_eval_criterion(trainer)
if verbose:
print ("Minibatch: {0}, Loss: {1:.4f}, Error: {2:.2f}".format(mb, training_loss, eval_error))
return mb, training_loss, eval_error
```
### Run the trainer
We are now ready to train our Logistic Regression model. We want to decide what data we need to feed into the training engine.
In this example, each iteration of the optimizer will work on 25 samples (25 dots w.r.t. the plot above) a.k.a. `minibatch_size`. We would like to train on say 20000 observations. If the number of samples in the data is only 10000, the trainer will make 2 passes through the data. This is represented by `num_minibatches_to_train`. Note: In real world case, we would be given a certain amount of labeled data (in the context of this example, observation (age, size) and what they mean (benign / malignant)). We would use a large number of observations for training say 70% and set aside the remainder for evaluation of the trained model.
With these parameters we can proceed with training our simple feedforward network.
```
# Initialize the parameters for the trainer
minibatch_size = 25
num_samples_to_train = 20000
num_minibatches_to_train = int(num_samples_to_train / minibatch_size)
# Run the trainer and perform model training
training_progress_output_freq = 50
plotdata = {"batchsize":[], "loss":[], "error":[]}
for i in range(0, num_minibatches_to_train):
features, labels = generate_random_data_sample(minibatch_size, input_dim, num_output_classes)
# Specify input variables mapping in the model to actual minibatch data to be trained with
trainer.train_minibatch({input : features, label : labels})
batchsize, loss, error = print_training_progress(trainer, i,
training_progress_output_freq, verbose=1)
if not (loss == "NA" or error =="NA"):
plotdata["batchsize"].append(batchsize)
plotdata["loss"].append(loss)
plotdata["error"].append(error)
# Compute the moving average loss to smooth out the noise in SGD
plotdata["avgloss"] = moving_average(plotdata["loss"])
plotdata["avgerror"] = moving_average(plotdata["error"])
# Plot the training loss and the training error
import matplotlib.pyplot as plt
plt.figure(1)
plt.subplot(211)
plt.plot(plotdata["batchsize"], plotdata["avgloss"], 'b--')
plt.xlabel('Minibatch number')
plt.ylabel('Loss')
plt.title('Minibatch run vs. Training loss')
plt.show()
plt.subplot(212)
plt.plot(plotdata["batchsize"], plotdata["avgerror"], 'r--')
plt.xlabel('Minibatch number')
plt.ylabel('Label Prediction Error')
plt.title('Minibatch run vs. Label Prediction Error')
plt.show()
```
## Evaluation / Testing
Now that we have trained the network. Let us evaluate the trained network on data that hasn't been used for training. This is called **testing**. Let us create some new data and evaluate the average error and loss on this set. This is done using `trainer.test_minibatch`. Note the error on this previously unseen data is comparable to training error. This is a **key** check. Should the error be larger than the training error by a large margin, it indicates that the trained model will not perform well on data that it has not seen during training. This is known as [overfitting][]. There are several ways to address overfitting that is beyond the scope of this tutorial but the Cognitive Toolkit provides the necessary components to address overfitting.
Note: We are testing on a single minibatch for illustrative purposes. In practice one runs several minibatches of test data and reports the average.
**Question** Why is this suggested? Try plotting the test error over several set of generated data sample and plot using plotting functions used for training. Do you see a pattern?
[overfitting]: https://en.wikipedia.org/wiki/Overfitting
```
# Run the trained model on newly generated dataset
test_minibatch_size = 25
features, labels = generate_random_data_sample(test_minibatch_size, input_dim, num_output_classes)
trainer.test_minibatch({input : features, label : labels})
```
### Checking prediction / evaluation
For evaluation, we map the output of the network between 0-1 and convert them into probabilities for the two classes. This suggests the chances of each observation being malignant and benign. We use a softmax function to get the probabilities of each of the class.
```
out = softmax(z)
result = out.eval({input : features})
```
Let us compare the ground-truth label with the predictions. They should be in agreement.
**Question:**
- How many predictions were mislabeled? Can you change the code below to identify which observations were misclassified?
```
print("Label :", np.argmax(labels[:25],axis=1))
print("Predicted:", np.argmax(result[0,:25,:],axis=1))
```
### Visualization
It is desirable to visualize the results. In this example, the data is conveniently in two dimensions and can be plotted. For data with higher dimensions, visualization can be challenging. There are advanced dimensionality reduction techniques that allow for such visualizations [t-sne][].
[t-sne]: https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding
```
# Model parameters
print(mydict['b'].value)
bias_vector = mydict['b'].value
weight_matrix = mydict['w'].value
# Plot the data
import matplotlib.pyplot as plt
# given this is a 2 class
colors = ['r' if l == 0 else 'b' for l in labels[:,0]]
plt.scatter(features[:,0], features[:,1], c=colors)
plt.plot([0, bias_vector[0]/weight_matrix[0][1]],
[ bias_vector[1]/weight_matrix[0][0], 0], c = 'g', lw = 3)
plt.xlabel("Scaled age (in yrs)")
plt.ylabel("Tumor size (in cm)")
plt.show()
```
**Exploration Suggestions**
- Try exploring how the classifier behaves with different data distributions - suggest changing the `minibatch_size` parameter from 25 to say 64. Why is the error increasing?
- Try exploring different activation functions
- Try exploring different learners
- You can explore training a [multiclass logistic regression][] classifier.
[multiclass logistic regression]: https://en.wikipedia.org/wiki/Multinomial_logistic_regression
| github_jupyter |
# Evaluation
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import warnings
warnings.filterwarnings('ignore')
plt.rcParams['figure.figsize'] = [10, 5]
```
# Continual Learning Metrics
```
# Because of a mistake in my implementation
# ["no_of_test"] cannot be used but it can be calculated by ["no_of_correct_prediction"]/["accuracy"]
# but it cannot be calculated when ["accuracy"] == 0
# ((raw["no_of_correct_prediction"]/ raw["accuracy"]).apply(np.ceil))
# the mistake have been fixed now but the data have not updated
def calculateContinualMetircs(raw):
task_order = raw["task_order"].unique()
method = raw["method"].unique()
print(task_order, method)
all_MBase = {k:[] for k in method}
all_Mnew = {k:[] for k in method}
all_Mnow = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
if m=="offline":
continue
target = rows[rows["method"]==m]
# calculate m_base
_ideal = offline[offline["task_index"]==1]["accuracy"]
_m = target[target["task_index"]==1][["accuracy", "no_of_test", "no_of_correct_prediction"]]
_N = len(_m)
_m = (_m["accuracy"]/float(_ideal)).sum()
Mbase = float(_m/_N)
all_MBase[m].append(Mbase)
_sum = 0.0
train_session = target["train_session"].unique()
for s in train_session:
s = int(s)
_ideal = offline[offline["task_index"]==s]["accuracy"]
_m = target[target["train_session"]==str(s)]
_m = _m[_m["task_index"]==s]["accuracy"]
assert len(_m)==1
_sum += float(_m)/float(_ideal)
if len(train_session)==0:
all_Mnew[m].append(np.nan)
else:
Mnew = _sum/len(train_session)
all_Mnew[m].append(Mnew)
_sum = 0.0
task_index = target["task_index"].unique()
_m = target[target["train_session"]==str(len(task_index))]
for t in task_index:
t = int(t)
_ideal = offline[offline["task_index"]==t]["accuracy"]
_m1 = _m[_m["task_index"]==t]["accuracy"]
assert len(_m1)==1
_sum += float(_m1)/float(_ideal)
if len(train_session)==0:
all_Mnow[m].append(np.nan)
else:
Mnow = _sum/len(train_session)
all_Mnow[m].append(Mnow)
return all_MBase, all_Mnew, all_Mnow
from scipy import stats
def printCLMetrics(all_MBase, all_Mnew, all_Mnow):
def p(metric, name):
print("Metric: ", name)
for m in metric:
avg = np.mean(metric[m])
err = stats.sem(metric[m])
print("{0} {1:.3f} {2:.3f}".format(m, avg, err))
print("=====================")
print("")
p(all_MBase, "M base")
p(all_Mnew, "M new")
p(all_Mnow, "M now")
```
# CASAS Dataset 👌
```
# Result from newsrc/result_iter1000-1000_h500-100_all/
folder = "../Results/result_iter1000-1000_h500-100_all/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
raw.head()
b, n, nw = calculateContinualMetircs(raw)
print("")
printCLMetrics(b, n, nw)
# Result from newsrc/result_iter1000-1000_h500-100_all/
folder = "../Results/result_iter5000-1000_h500-100_all/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
raw.head()
b, n, nw = calculateContinualMetircs(raw)
print("")
printCLMetrics(b, n, nw)
```
# PAMAP2 Dataset
```
# PAMAP2 Dataset with 100 hidden units/layer
folder = "../Results/results_two_datasets/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
b, n, nw = calculateContinualMetircs(raw)
print("")
printCLMetrics(b, n, nw)
# CASAS Dataset; re-run again
folder = "../Results/results_two_datasets.s2/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
b, n, nw = calculateContinualMetircs(raw)
print("")
printCLMetrics(b, n, nw)
# PAMAP2 Dataset with 1000 hidden units/layer (bigger network)
folder = "../Results/results_two_datasets.s4/"
raw = pd.read_csv(folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
CSMbase = []
CSMnew = []
CSMnow = []
cmd = raw["cmd"].unique()
cmd = [0,1,2,3,4]
for c in cmd:
target = raw[raw["cmd"]==c]
b, n, nw = calculateContinualMetircs(target)
CSMbase.append(b)
CSMnew.append(n)
CSMnow.append(nw)
# Metric: M base
# offline nan nan
# exact 1.227 0.061
# mp-gan 0.663 0.061
# mp-wgan 0.278 0.020
# sg-cgan 0.780 0.064
# sg-cwgan 0.411 0.044
# =====================
# Metric: M new
# offline nan nan
# exact 1.135 0.014
# mp-gan 1.153 0.014
# mp-wgan 1.158 0.015
# sg-cgan 1.158 0.014
# sg-cwgan 1.159 0.015
# =====================
# Metric: M now
# offline nan nan
# exact 1.081 0.015
# mp-gan 0.537 0.025
# mp-wgan 0.268 0.018
# sg-cgan 0.681 0.031
# sg-cwgan 0.306 0.023
# =====================
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
def plotline(values, label, x=[], xticks=[], models=None):
plt.rcParams['figure.figsize'] = [10, 5]
m = []
merr = []
if models is None:
models = ["mp-gan", "mp-wgan", "sg-cgan", "sg-cwgan"]
print(models)
for model in models:
tmp = []
tmperr = []
for i, v in enumerate(values):
avg = np.nanmean(v[model])
err = stats.sem(v[model], nan_policy="omit")
tmp.append(avg)
tmperr.append(err)
m.append(tmp)
merr.append(tmperr)
ind = np.arange(len(m[0])) # the x locations for the groups
fig, ax = plt.subplots()
patterns = [ "-s" , "-o" , "-x" , "-D" , "-+" , "-*", "-2" ]
for i, model in enumerate(models):
ax.errorbar(x, m[i], yerr=merr[i], fmt=patterns[i])
ax.set_title(label)
ax.set_xticks(x)
ax.set_xticklabels(x)
ax.legend(models)
fig.tight_layout()
plt.show()
xticks = [500, 1000, 1500, 2000, 2500]
models = None
def fixbugs(data):
return data
plotline(fixbugs(CSMbase), "Stability of the model", x=xticks, models=models)
plotline(fixbugs(CSMnew), "Plasticity of the model", x=xticks, models=models)
plotline(fixbugs(CSMnow),"Overall performance of the model", x=xticks, models=models)
```
| github_jupyter |
### Using workflows in combination with deskewing
* Read a czi image and get the first timepoint
* Generate deskewing workflow, save it, reload and run it again
* Generate a deskewing workflow with filtering, segmentation and labelling and run it
```
from napari_workflows import Workflow
from napari_workflows._io_yaml_v1 import load_workflow, save_workflow
from skimage.io import imread, imshow,imsave
from skimage.filters import gaussian
import numpy as np
import pyclesperanto_prototype as cle
import matplotlib.pyplot as plt
from aicsimageio import AICSImage, writers
from aicsimageio.types import PhysicalPixelSizes
#can be any czi file from lattice
data_path = 'C:\\RBC_lattice_dataset.czi'
data = AICSImage(data_path)
voxel_size_x_in_microns = data.physical_pixel_sizes.X
voxel_size_y_in_microns = data.physical_pixel_sizes.Y
voxel_size_z_in_microns = data.physical_pixel_sizes.Z
#eventually get angle from metadata
deskewing_angle_in_degrees = 30
image_to_deskew = data.get_image_dask_data("ZYX",T=0,C=0,S=0)
image_to_deskew
deskew_workflow = Workflow()
#define deskewing workflow
#save it
deskew_workflow.set("deskewing", cle.deskew_y, "input", angle_in_degrees = deskewing_angle_in_degrees,
voxel_size_x = voxel_size_x_in_microns, voxel_size_y= voxel_size_y_in_microns,
voxel_size_z = voxel_size_z_in_microns)
deskew_workflow.set("input", image_to_deskew)
print(deskew_workflow)
filename = "deskew_workflow.yaml"
save_workflow(filename, deskew_workflow)
new_w = load_workflow(filename)
print(new_w)
new_w.set("input",image_to_deskew )
print(new_w)
new_w.get("deskewing")
```
# Deskewing and labelling workflow
* Deskew dask image
* Median filter of 2,2,2
* Background subtraction using top hat sphere (20,20,20)
* Binarisation using a fixed threshold of 5
* Connected components labeling
```
#save_path = "D:\\deskew_save\\labelled.tif"
image_to_deskew = data.get_image_dask_data("ZYX",T=0,C=0,S=0)
#define deskewing workflow
image_seg_workflow = Workflow()
image_seg_workflow.set("deskewing", cle.deskew_y, "input", angle_in_degrees = deskewing_angle_in_degrees,
voxel_size_x = voxel_size_x_in_microns, voxel_size_y= voxel_size_y_in_microns,
voxel_size_z = voxel_size_z_in_microns)
image_seg_workflow.set("input", image_to_deskew)
image_seg_workflow.set("median", cle.median_sphere,"deskewing",radius_x = 2, radius_y = 2, radius_z = 2)
image_seg_workflow.set("background_subtraction", cle.top_hat_sphere,"median",radius_x = 20, radius_y = 20, radius_z = 20)
image_seg_workflow.set("binarisation", cle.threshold,"background_subtraction",constant =5)
image_seg_workflow.set("labeling", cle.connected_components_labeling_box,"binarisation")
#image_seg_workflow.set("save_image", imsave, save_path, arr ="labeling")
print(str(image_seg_workflow))
filename = "D:\\deskew_seg_workflow.yaml"
save_workflow(filename, image_seg_workflow)
new_deskew = load_workflow(filename)
print(new_deskew)
new_deskew.set("input",image_to_deskew)
print(new_deskew)
# Run the whole workflow
deskewed_img = new_deskew.get("deskewing")
labelled = new_deskew.get("labeling")
#How do we access intermediate objects without rerunning the whole workflow???
#Can I run the labeling workflow and also get the deskewed_img?
fig,axes = plt.subplots(3,1, figsize=(30,20))
cle.imshow(cle.maximum_z_projection(image_to_deskew),max_display_intensity=500,plot=axes[0])
cle.imshow(cle.maximum_z_projection(deskewed_img),max_display_intensity=500,plot=axes[1])
cle.imshow(cle.maximum_z_projection(labelled), color_map ="nipy_spectral" ,plot=axes[2])
```
View it in napari if needed
```
import napari
viewer = napari.Viewer()
viewer.add_image(image_to_deskew)
viewer.add_image(deskewed_img)
viewer.add_image(labelled)
```
| github_jupyter |
# Create Hive schema for Parquet files
This notebook demonstrates how to create a hive table for existing Parquet files. <br>
This can be done for a single file as well as for multiple files residing under the same folder.
```
import os
```
### Creating of spark context with hive support.
```
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("Import parquet schema to hive").config("hive.metastore.uris", "thrift://hive:9083").enableHiveSupport().getOrCreate()
```
Define function below for getting sql script needed for creating table in hive using dataframe types as columns to table
```
def getCreateTableScript(databaseName, tableName, path, df, partitions=[]):
#remove partition columns from the df to avoid repetition exception
partition_names = map(lambda x: x.split(' ')[0] , partitions )
ndf = df.drop(*partition_names)
cols = ndf.dtypes
createScript = "CREATE EXTERNAL TABLE " + databaseName + "." + tableName + "("
colArray = []
for colName, colType in cols:
colArray.append(colName.replace(" ", "_") + " " + colType)
createColsScript = ", ".join(colArray ) + ") "
partitionBy = ""
if len(partitions) > 0:
partitionBy = "PARTITIONED BY (" + ", ".join(partitions) + ") "
script = createScript + createColsScript + partitionBy + " STORED AS PARQUET LOCATION '" + path + "'"
print(script)
return script
#define main function for creating table where arqument 'path' is path to parquet files
def createTable(databaseName, tableName, path, partitions=[]):
df = spark.read.parquet(path)
sqlScript = getCreateTableScript(databaseName, tableName, path, df, partitions)
spark.sql(sqlScript)
if len(partitions) > 0:
spark.sql(f'msck repair table {databaseName}.{tableName}')
```
## One file example
```
# Set the path where the parquet file is located.
my_parqute_file_path = os.path.join('v3io://users/'+os.getenv('V3IO_USERNAME')+'/examples/userdata1.parquet')
createTable("default","tab1_single_file",my_parqute_file_path)
%sql select * from hive.default.tab1_single_file limit 10
```
## One folder example for spark output job
```
# Set the path where the parquet folder is located.
folder_path = os.path.join('v3io://users/'+os.getenv('V3IO_USERNAME')+'/examples/spark-output/')
createTable("default","table_from_dir",folder_path)
%sql select * from hive.default.table_from_dir limit 10
```
# Partitioned parquet example
Table partitioning is a common optimization approach used in systems like Hive. In a partitioned table, data are usually stored in different directories, with partitioning column values encoded in the path of each partition directory.
```
# Set path where parquet folder with parquet partitions are located indside.
folder_path = os.path.join('v3io://users/'+os.getenv('V3IO_USERNAME')+'/examples/partitioned_pq')
#provide list of partitions and their type
partition_list = ["gender string"]
createTable("default", "partitioned_table", folder_path, partition_list)
%sql select * from hive.default.partitioned_table limit 10
```
# Adding new partitions
```
#Once added new partitions to the table,
# it is required to run the below command in order for the hive metastore to be aware of the new files.
spark.sql('msck repair table default.partitioned_table')
```
# Browse the Metastore
```
# test how the tables were saved
#spark.sql("drop database test CASCADE")
databaseName = "default"
spark.sql("show databases").show()
spark.sql("show tables in " + databaseName).show()
```
### Access Hive from command line
In order to run Hive from command line,open up a jupyter terminal and run "hive" <br>
To view all existing hive tables run: show tables; <br>
Here you can run queries without specifying Hive. <br>
e.g. select * from table_from_single_file2;
## Cleanup
This will only clean the metastore definitions.
<br>The underlying data won't be affected.
```
spark.sql("drop table " + databaseName + ".tab1_single_file")
spark.sql("drop table " + databaseName + ".table_from_dir")
spark.sql("drop table " + databaseName + ".partitioned_table")
```
| github_jupyter |
```
import string
import numpy as np
import matplotlib.pyplot as plt
# Spacy
import spacy
nlp = spacy.load('en_core_web_sm')
# Keras
from keras.models import Sequential
from keras.layers import Dense
# Read the data
text = ''
with open('data/royal-family.txt', 'r') as f:
text = f.read()
print('Number of words: ', len(text.split()))
# Initial configurations and properties
window_size = 5 # Must be odd
embedding_dim = 2
stop_words = nlp.Defaults.stop_words
# Function to remove punctuation
def remove_punctutation(text):
return text.translate(str.maketrans('', '', string.punctuation))
# Function to remove stop words
def remove_stopwords(text):
return ' '.join([word for word in text.split() if word not in stop_words])
# Do a little preprocessing
text = remove_punctutation(text)
text = remove_stopwords(text)
# Create vocabulary
words = set(text.split())
vocab = {word: i for i, word in enumerate(words)}
# Create windows
windows = []
words = text.split()
bin = window_size // 2
for i in range(len(words)):
# left
for j in range(i - bin, i):
if j >= 0:
windows.append([words[j], words[bin]])
# right
for j in range(i + 1, i + bin + 1):
if j < len(words):
windows.append([words[bin], words[j]])
# One hot encoding
def one_hot_encoding(token):
token_vector = np.zeros(len(vocab))
token_vector[vocab[token]] = 1
return token_vector
X, Y = [], []
for window in windows:
# Encode
X_one_hot = one_hot_encoding(window[0])
Y_one_hot = one_hot_encoding(window[1])
# Append to corresponding lists
X.append(X_one_hot)
Y.append(Y_one_hot)
# Conver to numpy arrays
X = np.array(X)
Y = np.array(Y)
# Build the model
model = Sequential()
model.add(Dense(units = embedding_dim, input_dim = X.shape[1], activation = 'linear'))
model.add(Dense(units = Y.shape[1], activation = 'softmax'))
# Compile the model
model.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
# Train the model
model.fit(
X,
Y,
epochs = 100,
batch_size = 256,
verbose = 0
)
# model.save_weights('embedding_dict.h5')
# Get word embedding vectors
weights = model.get_weights()[0]
# Create Embedding dictionary
embedding_dict = {}
for word in words:
embedding_dict.update({word: weights[vocab.get(word)]})
# Plot the model pari-wise
plt.figure(figsize = (10, 10))
for word in list(vocab.keys()):
coord = embedding_dict.get(word)
plt.scatter(coord[0], coord[1])
plt.annotate(word, (coord[0], coord[1]))
```
| github_jupyter |
# 'Targeted Killing': The Drone War in Pakistan
**By Anna Mongillo**
The first CIA drone strike hit the tribal belt (otherwise known as the "lawless frontier") in the Northwest of Pakistan in 2004. Since then, drones have become America's vehicle of choice for eliminating terrorist targets in Pakistan. However, the use of drone strikes has come under fire for being ineffective and immoral as a counterterrorism tactic. It has been said that, although drone strikes certainly have a lower casualty rate and civilian casualty rate than any form of traditional warfare, ["the inherently secret nature of the weapon creates a persistent feeling of fear in the areas where drones hover in the sky, and the hopelessness of communities that are on the receiving end of strikes causes severe backlash -- both in terms of anti-U.S. opinion and violence"](http://www.theatlantic.com/international/archive/2013/08/how-drones-create-more-terrorists/278743/). In other words, drone strikes may create the opportunity for terrorists to sympathize with civilians and recruit from pools of frightened Pakistani citizens, creating terrorists rather than combating them. Now, in 2016, can we show that over a decade of drone strikes have had any effect on the number of terrorist attacks? Can we actually find a correlation between the two? And how do drone strike casualties actually look in terms of morality?
For this exploration, I have used the Bureau of Investigative Journalism's Pakistan data sheets on drone strikes as well as the Global Terrorism Database information on terrorist attacks, read into Python below.
### Import Packages
```
import sys
import matplotlib.pyplot as plt
import datetime as dt
import numpy as np
from mpl_toolkits.basemap import Basemap
import pandas as pd
import seaborn as sns
from scipy.stats.stats import pearsonr
print('Python version: ', sys.version)
print('Pandas version: ', pd.__version__)
print('Today: ', dt.date.today())
```
### Reading in the Data/Data Cleaning
```
# url1: Pakistan latitude and longitude data for map visualization of drone strikes
# https://docs.google.com/spreadsheets/d/1P1TSWNwr1j-0pX022Q2iLEZd2-IGoswD08unYSvCYaU/edit#gid=9
# save as csv onto desktop
path = '/Users/anna/Desktop/pakistan.csv' # read in Pakistan drone strike data
df = pd.read_csv(path) # read into Python from desktop
df=df.replace(to_replace=['Unreported'], value=[np.nan]) # data cleaning for NaN values
df["Date"] = pd.to_datetime(df["Date"]) # change this column to datetime
df["Date"]=df["Date"].dt.year # change datetimes to years
# url2: More in-depth Pakistan drone strike data (casualty rates/summary tables, etc.) for plots
# https://docs.google.com/spreadsheets/d/1NAfjFonM-Tn7fziqiv33HlGt09wgLZDSCP-BQaux51w/edit#gid=694046452
# save as excel file onto desktop
drones=pd.read_excel('/Users/anna/Desktop/CIAdrones.xlsx',
sheetname="Summary tables and casualty rates", skip_footer=25)
drones["Year"]=drones["Year"].dt.year # change this column from datetimes to years
drones=drones.set_index('Year') # read in drone strike summary table and set the index to year
casualtyrates=pd.read_excel('/Users/anna/Desktop/CIAdrones.xlsx',
sheetname="Summary tables and casualty rates", skiprows=21)
casualtyrates=casualtyrates.drop([12,13,14,15,16])
casualtyrates["Year"]=casualtyrates["Year"].dt.year # change this column from datetimes to years
casualtyrates=casualtyrates.set_index('Year') # read in drone strike casualty rate table and set the index to year
casualtyrates=casualtyrates.dropna(axis=1)
# url3: Global Terrorism data
# http://www.start.umd.edu/gtd/contact/
# save as excel files onto desktop
gtdto11= pd.read_excel('/Users/anna/Desktop/gtd_92to11_0616dist.xlsx') # read in Global Terrorism Data 1992-2011
pakistan=gtdto11['country_txt'] == 'Pakistan' # create a Boolean variable for country name Pakistan
year=gtdto11['iyear']>2003 # create a Boolean variable for years after 2003
gtdto11count=gtdto11[pakistan & year] # create a new dataframe containing only Pakistan data and years after 2003
gtdto15=pd.read_excel('/Users/anna/Desktop/gtd_12to15_0616dist.xlsx') # read in Global Terrorism Data 2011-2015
pakistan2=gtdto15['country_txt']=='Pakistan' # create a dataframe with only Pakistan data
gtdto15count=gtdto15[pakistan2] # create a series
gtd=gtdto11count.append(gtdto15count, ignore_index=True) # append dataframes from 2004-2011 and 2011-2015
```
### Plot 1: Number of Drone Strikes and Number of Terrorist Attacks
*Do drone strikes actually have an effect on terrorism?*
Although the below chart seems to visually show that an increase in drone strikes led to a later increase in terrorist attacks, analysis of number of drone strikes compared to number of terrorist attacks reveals a low Pearson's correlation coefficient (0.19), while the p value exceeds threshold (0.55), making a convincing argument that drone strikes and terrorist attacks are not related.
```
numgtd=gtd.groupby('iyear').count() # group global terrorism data by year and set to variable numgtd
numter=numgtd['eventid'] # set new variable numter as number of terrorist attacks by year
numdrones=drones['CIA Drone Strikes'] # set new variable numdrones as number of drone strikes by year
numdrones=numdrones.drop(numdrones.index[[12]]) # drop year 2016 using the index
chartdf=pd.concat([numter, numdrones], axis=1) # create new chart with only numter and numgtd
chartdf=chartdf.dropna() # clean the chart of NaN
eventid=chartdf['eventid']
chartdf['eventid']=eventid.astype(int) # convert number of terrorist attacks from float to integer
sns.set_style('darkgrid') # set darkgrid style using Seaborn
fig, (ax1, ax2)=plt.subplots(2, 1, figsize=(8,7)) # create figure with two axes
chartdf['eventid'].plot(ax=ax2, color='r', title='Terrorist Attacks Per Year',
y='Number of Attacks', kind='bar', alpha=0.6)
chartdf['CIA Drone Strikes'].plot(ax=ax1, title='Drone Strikes Per Year',
y="Number of Strikes", kind='bar', alpha=0.5)
ax2.set_ylabel('Number of Attacks')
ax1.set_ylabel('Number of Strikes')
ax2.set_xlabel('Year')
ax1.set_xlabel('Year')
plt.tight_layout()
plt.show()
print('Correlation:',pearsonr(chartdf['eventid'],chartdf['CIA Drone Strikes'])) #Pearson's correlation and p value
```
### Plot 2: Monthwise Strikes/Attacks
We have to wonder: perhaps the amount of strikes in a given month will lead to more strikes in that month? However, this plot shows that not even 0.5% of the data is correlated on a monthly level, along with another high p value. Drone strikes are further proven to be unrelated to terrorist attacks.
```
# below is a long process of converting date columns in the Global Terrorism database to datetime:
year=list(gtd['iyear'])
month=list(gtd['imonth'])
day=list(gtd['iday'])
date=(zip(year, month, day))
date=list(date)
date=[str(x)for x in date]
date=[x.replace(', ','-') for x in date]
date=[x.replace('(', '') for x in date]
date=[x.replace(')', '') for x in date]
gtd['idate']=date
gtd["idate"] = pd.to_datetime(gtd["idate"], format='%Y-%m-%d', errors='coerce') # change this column to datetime
per2=gtd.idate.dt.to_period("M") # convert to monthly data
ter2=gtd.groupby(per2) # group by month
ter2=ter2.count() # count number of attacks per month
termonths=ter2['eventid'] # save in new variable
cia='/Users/anna/Desktop/pakistan.csv' # read in Pakistan drone strike data one more time in order to adjust "Date"
cia = pd.read_csv(cia)
cia["Date"] = pd.to_datetime(cia["Date"]) # change this column to datetime
per=cia.Date.dt.to_period("M") # convert to monthly data
cia=cia.groupby(per) #group by month
cia=cia.count() #count number of strikes in each month
dronemonths=cia['Strike'] # save in new variable
totalmonths=pd.concat([termonths, dronemonths], axis=1) # create new dataframe for plotting
totalmonths=totalmonths.dropna() # clean the data frame
totalmonths=totalmonths.rename(columns={'eventid': 'Attacks', 'Strike':'Strikes'})
totalmonths.plot()
plt.title('Strikes/Attacks by Month')
plt.show()
print('Correlation:',pearsonr(totalmonths['Attacks'],totalmonths['Strikes'])) # Pearson's correlation and p value
```
### Plot 3: Drone Strikes vs. Terrorist Attacks (Normalized)
A better (normalized) visualization of the data:
```
x=np.array(numgtd.index) # create x array (years, taken from Global Terrorism Data)
y=np.array(numdrones) # create y array (number of drone attacks)
y=y/y.max().astype(np.float64) # normalize drone strike data on a scale of 0-1 and convert to float
z=np.array(numter) # create x array (number of terrorist attacks)
z=z/z.max().astype(np.float64) # normalize terrorist attack data on a scale of 0-1 and convert to float
plt.figure(figsize=(15,6)) # create a figure of size 15, 6
plt.scatter(x,y, zorder=2, label="Drone Strikes") # plot x and y arrays as a scatter plot (Drone Strikes)
plt.plot(x, y, zorder=1, lw=3) # connect dots with line
plt.bar(x,z, color='red', alpha=0.6, label='Terrorist Attacks', align='center') # plot x and z arrays as a bar chart
plt.title('Drone Strikes vs. Terrorist Attacks: Normalized', fontsize=15)
plt.xlim(2003,2016) # set x upper and lower limits
plt.xlabel("Year", fontsize=12)
plt.ylabel("Strikes/Attacks", fontsize=12)
plt.tick_params(axis='y', labelleft='off', labelright='off') # turn off y axis labels because data is normalized
plt.ylim(0,1) # set y upper and lower limits
plt.legend(loc='best', fontsize='large')
plt.show()
print('Correlation:',pearsonr(y,z)) # Pearson's correlation and p value
```
### Plot 4: Percent Change of Drone Strikes and Terrorist Attacks
*Is terrorism related to retaliation after a higher/lower increase in drone strikes?*
This plot shows that while terrorism levels certainly aren't decreased by drone strikes, they aren't increased by them directly either...however, a slightly prolonged period of lower percent change in drone strikes (which finally drops below zero at the end of the dataset, meaning that drone strike numbers are decreasing) seems to result in a lower percent change of terrorist attacks, which also drops below zero at the end.
```
attacks=list(chartdf['eventid'])
diff=[100*(y - x)/x for x, y in zip(attacks[::1], attacks[1::1])] # percent change of terrorist attacks
diff=pd.Series(diff) # turn into series
diff=pd.Series.to_frame(diff) # turn into dataframe
years=['2004-2005', '2005-2006', '2006-2007', '2007-2008', '2008-2009', '2009-2010',
'2010-2011', '2011-2012', '2012-2013', '2013-2014', '2014-2015']
years=pd.Series(years) # convert list object to series
years=pd.Series.to_frame(years) # convert series to dataframe
years=years.rename(columns={0: "Year"})
diff=pd.concat([diff,years], axis=1) # create a dataframe with terrorism % change and years
diff=diff.set_index('Year')
diff=diff.rename(columns={0:'Percent Change of Terrorist Attacks'})
strikes=list(chartdf['CIA Drone Strikes'])
dronediff=[100*(y - x)/x for x, y in zip(strikes[::1], strikes[1::1])] # percent change of terrorist attacks
dronediff=pd.Series(dronediff) # turn into series
dronediff=pd.Series.to_frame(dronediff) # turn into dataframe
years=['2004-2005', '2005-2006', '2006-2007', '2007-2008', '2008-2009', '2009-2010',
'2010-2011', '2011-2012', '2012-2013', '2013-2014', '2014-2015']
years=pd.Series(years) # convert list object to series
years=pd.Series.to_frame(years) # convert series to dataframe
years=years.rename(columns={0: "Year"})
dronediff=pd.concat([dronediff,years], axis=1) # create a dataframe with years and drone strike % change
dronediff=dronediff.set_index('Year')
dronediff=dronediff.rename(columns={0:'Percent Change of Drone Strikes'})
combined=pd.concat([dronediff, diff], axis=1) # create a dataframe with drone % change and terrorism % change
combined.plot()
plt.show()
```
### Plot 5: Drone Strike Casualty Details
*Who is dying?*
Below are two plots that may give insight into the morality of drone strikes: one showing the demographics of those killed in drone strikes and another detailing casualty rates. The demographics show that the most people died between 2007-2012, which is right before terrorist attacks peaked in number. Casualty rates in 2006 were markedly higher than the rest of the years.
```
dcasualties=drones["Minimum people killed"]
dcivilians=drones["Minimum civilians killed"]
dchildren=drones['Minimum children killed']
civcas=pd.concat([dcasualties, dcivilians], axis=1)
dronedeaths=pd.concat([civcas, dchildren], axis=1)
dronedeaths=dronedeaths.drop([2016]) # new Dataframe for total, civilian and child casualty rates during drone strikes
dronedeaths.plot.area(stacked=False)
plt.title('Drone Strike Casualties')
plt.ylabel("Number Killed", fontsize=12)
plt.xlabel("Year")
plt.ylabel("Number Killed", fontsize=12)
plt.xlabel("Year")
plt.title('Drone Strike Casualties')
plt.show() # plot the new Dataframe
casualtyrates.plot.bar()
plt.title('Drone Strike Casualty Rates')
plt.show()
```
### Plot 6: Casualties and Terrorist Attacks
*Are higher drone strike casualties/number of people killed seemingly related to an increase in terrorist attacks?*
Once again, Pearson's r reveals a low correlation and a high p value between casualties and terrorist attacks, making the calculations untrustworthy.
```
dcasualties=dcasualties.drop([2016]) # drop 2016 from casualty because it is not included in numter
x=np.array(numgtd.index) # create x array (years, taken from Global Terrorism Data)
y=np.array(dcasualties) # create y array (casualty rates)
y=y/y.max().astype(np.float64) # normalize casualty rate data on a scale of 0-1 and convert to float
z=np.array(numter) # create x array (number of terrorist attacks)
z=z/z.max().astype(np.float64) # normalize terrorist attack data on a scale of 0-1 and convert to float
plt.figure(figsize=(15,6)) # create a figure of size 15, 6
plt.scatter(x,y, zorder=2,
label="Drone Casualties") # plot x and y arrays as a scatter plot (casualty rates)
plt.plot(x, y, zorder=1, lw=3) # connect dots with line
plt.bar(x,z, color='red', alpha=0.6, label='Terrorist Attacks', align='center') # plot x and z arrays as a bar chart
plt.title('Drone Casualties vs. Terrorist Attacks: Normalized', fontsize=15)
plt.xlim(2003.5,2015.5) # set x upper and lower limits
plt.xlabel("Year", fontsize=12)
plt.ylabel("Casualties/Attacks", fontsize=12)
plt.tick_params(axis='y', labelleft='off', labelright='off') # turn off y axis labels because data is normalized
plt.ylim(0,1) # set y upper and lower limits
plt.legend(loc='best', fontsize='large')
plt.show()
print('Correlation:',pearsonr(y,z)) # Pearson's correlation and p value
```
# Appendix
*Map Visualizations: where are the drone strikes targeted? Where are the terrorist attacks centered?*
The below vizualization seems to show that drone strikes, unlike terrorist attacks, are concentrated in the tribal belt of Pakistan in the Northwest.
```
lat=df["Latitude"]
long=df["Longitude"]
coordinates=pd.concat([lat, long], axis=1) # new DataFrame for latitude and longitude of drone strikes
lat2=gtd["latitude"]
long2=gtd["longitude"]
coordinates2=pd.concat([lat2, long2], axis=1) # new DataFrame for latitude and longitude of terrorist attacks
fig = plt.figure(figsize=(10,5)) # create a figure with size 10,5
map = Basemap(projection='gall', # make the figure a Basemap map (Gall projection)
resolution = 'h', # high image resolution, with latitude and longitude upper and lower bounds
area_thresh = 0.1,
llcrnrlon=60, llcrnrlat=24,
urcrnrlon=80, urcrnrlat=40,
lat_0=30, lon_0=70)
map.drawcoastlines()
map.drawcountries()
map.fillcontinents(color = 'tan')
map.drawmapboundary(fill_color='#f4f4f4')
x,y = map(list(coordinates['Longitude']), list(coordinates['Latitude'])) # set my coordinates df as x and y values
map.plot(x, y, 'ro', markersize=4) # plot my coordinates df points onto the map with point size 4 and color red
plt.title('Drone Strikes 2004-2013')
plt.show()
fig = plt.figure(figsize=(10,5)) # create a figure with size 10,5
map = Basemap(projection='gall', # make the figure a Basemap map
resolution = 'h', # high resolution, with latitude and longitude upper and lower bounds
area_thresh = 0.1,
llcrnrlon=60, llcrnrlat=24,
urcrnrlon=80, urcrnrlat=40,
lat_0=30, lon_0=70)
map.drawcoastlines()
map.drawcountries()
map.fillcontinents(color = 'tan')
map.drawmapboundary(fill_color='#f4f4f4')
x,y = map(list(coordinates2['longitude']), list(coordinates2['latitude'])) # set my coordinates gtd as x and y values
map.plot(x, y, 'ro', markersize=4) # plot my coordinates gtd points onto the map
plt.title('Terrorist Attacks 2004-2015')
plt.show()
```
## Change of Location?
*Map visualization: Are the terrorists just moving locations?*
The below map visualization seems to show a spreading-out of terrorist attacks from 2004 to 2015, growing more frequent in more areas. Perhaps the terrorists have been simply moving locations or spreading out throughout the years we have analyzed, and that is why attacks have been increasing.
```
terattacks=gtd.set_index(['iyear'])
ter2004=terattacks.loc[2004] # terrorist attacks in 2004
ter2015=terattacks.loc[2015] # terrorist attacks in 2015
lat2004=ter2004['latitude']
long2004=ter2004['longitude']
terfinal2004=pd.concat([lat2004, long2004], axis=1) # new Dataframe for coordinates of 2004 attacks
lat2015=ter2015['latitude']
long2015=ter2015['longitude']
terfinal2015=pd.concat([lat2015, long2015], axis=1) # new Dataframe for coordinates of 2015 attackss
fig = plt.figure(figsize=(10,5)) # create a figure with size 10,5
map = Basemap(projection='gall', # make the figure a Basemap map (Gall projection)
resolution = 'h', # high image resolution, with latitude and longitude upper and lower bounds
area_thresh = 0.1,
llcrnrlon=60, llcrnrlat=24,
urcrnrlon=80, urcrnrlat=40,
lat_0=30, lon_0=70)
map.drawcoastlines()
map.drawcountries()
map.fillcontinents(color = 'tan')
map.drawmapboundary(fill_color='#f4f4f4')
x,y = map(list(terfinal2004['longitude']), list(terfinal2004['latitude'])) # set my coordinates as x and y values
map.plot(x, y, 'ro', markersize=4) # plot my coordinates 2004 points onto the map with point size 4 and color red
plt.title('Terrorist Attack Locations 2004')
plt.show()
fig = plt.figure(figsize=(10,5)) # create a figure with size 10,5
map = Basemap(projection='gall', # make the figure a Basemap map
resolution = 'h', # high resolution, with latitude and longitude upper and lower bounds
area_thresh = 0.1,
llcrnrlon=60, llcrnrlat=24,
urcrnrlon=80, urcrnrlat=40,
lat_0=30, lon_0=70)
map.drawcoastlines()
map.drawcountries()
map.fillcontinents(color = 'tan')
map.drawmapboundary(fill_color='#f4f4f4')
x,y = map(list(terfinal2015['longitude']), list(terfinal2015['latitude'])) # set my coordinates as x and y values
map.plot(x, y, 'ro', markersize=4) # plot my coordinates 2015 points onto the map
plt.title('Terrorist Attack Locations 2015')
plt.show()
```
# Summary
In summary, while drone strikes cannot necessarily be proven to correlate with increases in terrorist attacks due to factors like retaliation, the data visualizations point more to that than to the idea that drone strikes prevent or combat terrorism. This could be due to the fact that, as seen in the Appendix, terrorist attacks are more spread out than drone strikes, so the drone strikes are hardly affecting terrorists. Nevertheless, after many years of consecutively 'targeting' terrorism in Pakistan using drone strikes, their effectiveness as a counterterrorism tactic cannot be proven--a worrisome concept.
| github_jupyter |
# Data Mining Challange: *Reddit Gender Text-Classification*
### Modules
```
# Numpy & matplotlib for notebooks
%pylab inline
# Pandas
import pandas as pd # Data analysis and manipulation
# Sklearn
from sklearn import utils
from sklearn.preprocessing import StandardScaler # to standardize features by removing the mean and scaling to unit variance (z=(x-u)/s)
from sklearn.neural_network import MLPClassifier # Multi-layer Perceptron classifier which optimizes the log-loss function using LBFGS or sdg.
from sklearn.model_selection import train_test_split # to split arrays or matrices into random train and test subsets
from sklearn.model_selection import KFold # K-Folds cross-validator providing train/test indices to split data in train/test sets.
from sklearn.decomposition import PCA, TruncatedSVD # Principal component analysis (PCA); dimensionality reduction using truncated SVD.
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import MultinomialNB # Naive Bayes classifier for multinomial models
from sklearn.feature_extraction.text import CountVectorizer # Convert a collection of text documents to a matrix of token counts
from sklearn.metrics import roc_auc_score as roc # Compute Area Under the Receiver Operating Characteristic Curve from prediction scores
from sklearn.metrics import roc_curve, auc # Compute ROC; Compute Area Under the Curve (AUC) using the trapezoidal rule
# Matplotlib
import matplotlib # Data visualization
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# Seaborn
import seaborn as sns # Statistical data visualization (based on matplotlib)
# Tqdm
from tqdm import tqdm
tqdm.pandas(desc="progress-bar")
# Gensim
import gensim
from gensim.models import Doc2Vec
from gensim.models.doc2vec import TaggedDocument
# Regular Expressions
import re # String manipulation
# Nltk
import nltk # lemmatization
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk import pos_tag
from nltk.corpus import wordnet as wn
from nltk.stem.snowball import SnowballStemmer # stemmer
from bs4 import BeautifulSoup
from collections import defaultdict
```
## Data Loading and Manipulation
```
# Load datasets
train_data = pd.read_csv("../input/dataset/train_data.csv")
target = pd.read_csv("../input/dataset/train_target.csv")
test_data = pd.read_csv("../input/dataset/test_data.csv")
# Create author's gender dictionary
author_gender = {}
for i in range(len(target)):
author_gender[target.author[i]] = target.gender[i]
# X is the list of aggregated comments
X = []
# y is the list of genders
y = []
# Populate the dictionary with keys ("authors") and values ("gender")
for author, group in train_data.groupby("author"):
X.append(group.body.str.cat(sep = " "))
y.append(author_gender[author])
```
## Preprocessing, Optimize Input for `doc2vec` Training
```
# Create pre-processing functions
def remove_number(text):
num = re.compile(r'[-+]?[.\d]*[\d]+[:,.\d]*')
return num.sub(r'NUMBER', text)
def remove_URL(text):
url = re.compile(r'https?://\S+|www\.\S+')
return url.sub(r'URL',text)
def remove_repeat_punct(text):
rep = re.compile(r'([!?.]){2,}')
return rep.sub(r'\1 REPEAT', text)
def remove_elongated_words(text):
rep = re.compile(r'\b(\S*?)([a-z])\2{2,}\b')
return rep.sub(r'\1\2 ELONG', text)
def remove_allcaps(text):
caps = re.compile(r'([^a-z0-9()<>\'`\-]){2,}')
return caps.sub(r'ALLCAPS', text)
def transcription_smile(text):
eyes = "[8:=;]"
nose = "['`\-]"
smiley = re.compile(r'[8:=;][\'\-]?[)dDp]')
#smiley = re.compile(r'#{eyes}#{nose}[)d]+|[)d]+#{nose}#{eyes}/i')
return smiley.sub(r'SMILE', text)
def transcription_sad(text):
eyes = "[8:=;]"
nose = "['`\-]"
smiley = re.compile(r'[8:=;][\'\-]?[(\\/]')
return smiley.sub(r'SADFACE', text)
def transcription_heart(text):
heart = re.compile(r'<3')
return heart.sub(r'HEART', text)
# Tags Part of Speech (POS), because the lemmatizer requires it
tag_map = defaultdict(lambda : wn.NOUN)
tag_map['J'] = wn.ADJ
tag_map['V'] = wn.VERB
tag_map['R'] = wn.ADV
# Create lemmatizer
word_Lemmatized = WordNetLemmatizer()
def review_to_words1(raw_body):
# remove html tags
body_text = BeautifulSoup(raw_body).get_text()
#letters_only = re.sub("[^a-zA-Z]", " ", body_text)
# lowercase all text
words = body_text.lower()
# remove urls
text = remove_URL(words)
# remove numbers
text = remove_number(text)
# remove smiles
text = transcription_sad(text)
text = transcription_smile(text)
text = transcription_heart(text)
text = remove_elongated_words(text)
words = remove_repeat_punct(text)
# tokenizes and pass to lemmatizer, which lemmatizes taking tags into account (see before)
words = word_tokenize(words)
# we don't remove stop words, because doing it on combination with removing the 40 (trial & error estimated parameter) most utilized words (see below) decreases performance
#stops = set(stopwords.words("english"))
#meaningful_words = [w for w in words if not w in stops]
Final_words = []
for word, tag in pos_tag(words):
word_Final = word_Lemmatized.lemmatize(word,tag_map[tag[0]])
Final_words.append(word_Final)
#if len(Final_words)<11: return -1
# returns lemmatized texts as strings
return( " ".join(Final_words))
lemmatizer = WordNetLemmatizer()
stemmer = SnowballStemmer("english")
# Another lemmatizer function. We verified it gives worse performance
def review_to_words(raw_body):
body_text = BeautifulSoup(raw_body).get_text()
letters_only = re.sub("[^a-zA-Z]", " ", body_text)
words = letters_only.lower().split()
stops = set(stopwords.words("english"))
meaningful_words = [w for w in words if not w in stops]
final = [lemmatizer.lemmatize(w,pos="v") for w in meaningful_words]
#if len(final)<10: return -1
return( " ".join(final))
clean_train_comments = [review_to_words1(x) for x in X]
# This function formats the input dor doc2vec
def label_sentences(corpus, label_type):
labeled = []
for i, v in enumerate(corpus):
label = label_type + '_' + str(i)
labeled.append(TaggedDocument(v.split(), [label]))
return labeled
y = np.array(y)
# Splitting
X_train, X_valid, y_train, y_valid = train_test_split(clean_train_comments, y,
train_size=0.8, test_size=0.2,
random_state=0)
# createdoc2vec input
X_train = label_sentences(X_train, 'Train')
X_valid = label_sentences(X_valid, 'Valid')
all_data = X_train + X_valid
```
## `doc2vec`: Model Definition and Training
```
# define the model
# window: qhow many neighboring words should the moel look at
# negative :som words are negatively weighted
# min_count: once-appearing words are discarded
model_dbow = Doc2Vec(dm=1, vector_size=400, window=7, negative=5, min_count=1, alpha=0.065)
# creates the vocabulary. tdqm is the progress bar
model_dbow.build_vocab([x for x in tqdm(all_data)])
# trianing. The sub doc2vec is trianed an training and test set
for epoch in range(30):
model_dbow.train(utils.shuffle([x for x in tqdm(all_data)]), total_examples=len(all_data), epochs=1)
model_dbow.alpha -= 0.002
model_dbow.min_alpha = model_dbow.alpha
# Returns vectorized aggragated texts
def get_vectors(model, corpus_size, vectors_size, vectors_type):
"""
Get vectors from trained doc2vec model
:param doc2vec_model: Trained Doc2Vec model
:param corpus_size: Size of the data
:param vectors_size: Size of the embedding vectors
:param vectors_type: Training or Testing vectors
:return: list of vectors
"""
vectors = np.zeros((corpus_size, vectors_size))
for i in range(0, corpus_size):
prefix = vectors_type + '_' + str(i)
# builds teh vector extracting it from model.docvecs indexed by label
vectors[i] = model.docvecs[prefix]
return vectors
train_vectors_dbow = get_vectors(model_dbow, len(X_train), 400, 'Train')
valid_vectors_dbow = get_vectors(model_dbow, len(X_valid), 400, 'Valid')
```
## `TruncatedSVD`: Train Data Visualization
```
# Plot along the two dimensions with most variance
def plot_LSA(test_data, test_labels, savepath="PCA_demo.csv", plot=True):
lsa = TruncatedSVD(n_components=2)
lsa.fit(test_data)
lsa_scores = lsa.transform(test_data)
color_mapper = {label:idx for idx,label in enumerate(set(test_labels))}
color_column = [color_mapper[label] for label in test_labels]
colors = ['orange','blue']
if plot:
plt.scatter(lsa_scores[:,0], lsa_scores[:,1], s=8, alpha=.8, c=test_labels, cmap=matplotlib.colors.ListedColormap(colors))
orange_patch = mpatches.Patch(color='orange', label='M')
blue_patch = mpatches.Patch(color='blue', label='F')
plt.legend(handles=[orange_patch, blue_patch], prop={'size': 20})
plt.title('Doc2Vec comments only')
plt.savefig('foo.pdf')
fig = plt.figure(figsize=(8, 8))
plot_LSA(train_vectors_dbow,y_train)
plt.show()
```
## Classifier: Model Definition and Validation
```
# Define MLP Classifier:
## Activation function for the hidden layer: "rectified linear unit function"
## Solver for weight optimization: "stochastic gradient-based optimizer"
## Alpha: regularization parameter
## Learning rate schedule for weight updates: "gradually decreases the learning rate at each time step t using an inverse scaling exponent of power_t"
## Verbose: "True" in order to print progress messages to stdout.
## Early stopping: "True" in order to use early stopping to terminate training when validation score is not improving. It automatically sets aside 10% of training data as validation and terminate training when validation score is not improving by at least tol for n_iter_no_change consecutive epochs.
mlpClf = MLPClassifier(solver = 'adam', activation= 'relu' ,alpha = 0.0005, verbose = True, early_stopping = True,
learning_rate = 'invscaling', max_iter = 400,random_state=0)
# K fold (cross validation)
kf = KFold(n_splits = 10)
# Training and validation on all subsets
results = cross_val_score(mlpClf, np.concatenate((train_vectors_dbow, valid_vectors_dbow)), np.concatenate((y_train , y_valid)), cv=kf, scoring='roc_auc')
print("roc = ", np.mean(results))
# Final fit
mlpClf.fit(train_vectors_dbow, y_train)
# XGBoost Regressor
from xgboost import XGBRegressor
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
# let's also try XGBoost (but MLP performs better)
my_model1 = XGBRegressor(objective = "reg:logistic",n_estimators=10000, learning_rate=0.01, n_jobs=4,subsample = 0.9,
min_child_weight = 1,max_depth=4,gamma=1.5,colsample_bytree=0.6,random_state=0)
# K fold (cross validation)
kf = KFold(n_splits = 10)
# Training and validation on all subsets
results = cross_val_score(my_model1, np.concatenate((train_vectors_dbow, valid_vectors_dbow)), np.concatenate((y_train , y_valid)), cv=kf, scoring='roc_auc')
print("roc = ", np.mean(results))
# Model fit
my_model1.fit(train_vectors_dbow, y_train,
early_stopping_rounds=80,
#sample_weight = w,
eval_set=[(valid_vectors_dbow, y_valid)],
verbose=False)
# In the fit function there is the early stop, that one may set iff there is a validation set.
# The early stop interrupts the training when themodel starts overfitting.
# But, the model that will predict the test will have no validation during training,
# so we get here a value and heuristicallly use it also when predicting test
print(my_model1.best_iteration)
# roc plot
y_score = my_model1.predict(valid_vectors_dbow)
# Roc Curve for validation data
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y_valid, y_score)
roc_auc = auc(fpr, tpr)
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.4f)'% roc_auc )
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('XGB')
plt.legend(loc="lower right")
plt.show()
# Save XGB predictions
np.save("../working/y_scoremXGB",y_score)
y_score = mlpClf.predict_proba(valid_vectors_dbow)[:,1]
# Roc Curve for validation data
from sklearn.metrics import roc_curve, auc
fpr, tpr, thresholds = roc_curve(y_valid, y_score)
roc_auc = auc(fpr, tpr)
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.4f)'% roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('mlpClf')
plt.legend(loc="lower right")
plt.show()
# Save MLP predictions
np.save("../working/y_scoremlpClf",y_score)
```
| github_jupyter |
# Train Task2Baseline model
This is PyTorch version trainer code, will basically do the same with the original baseline written with Keras.
- Train with all machine type data, then save each models.
- Tensorboard visualization will show you training trajectories, thanks to [PyTorch Lightning](https://github.com/PyTorchLightning/pytorch-lightning).
- And some extra visualizations.
```
# public modules
from dlcliche.notebook import *
from dlcliche.utils import (
sys, random, Path, np, plt, EasyDict,
ensure_folder, deterministic_everything,
)
from argparse import Namespace
# private modules
sys.path.append('..')
import common as com
from pytorch_common import *
from model import Task2Baseline
# loading parameters -> hparams (argparse compatible)
params = EasyDict(com.yaml_load('config.yaml'))
# create working directory
ensure_folder(params.model_directory)
# test directories
dirs = com.select_dirs(param=params, mode='development')
# fix random seeds
deterministic_everything(2020, pytorch=True)
# PyTorch device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
%load_ext tensorboard
%tensorboard --logdir lightning_logs/
# train models
for target_dir in dirs:
target = str(target_dir).split('/')[-1]
print(f'==== Start training [{target}] with {torch.cuda.device_count()} GPU(s). ====')
files = com.file_list_generator(target_dir)
model = Task2Baseline().to(device)
if target == 'ToyCar': summary(device, model)
task2 = Task2Lightning(device, model, params, files)
trainer = pl.Trainer(max_epochs=params.fit.epochs, gpus=torch.cuda.device_count())
trainer.fit(task2)
model_file = f'{params.model_directory}/model_{target}.pth'
torch.save(task2.model.state_dict(), model_file)
print(f'saved {model_file}.\n')
```
## Visualize
```
#load_weights(task2.model, 'model/model_ToyCar.pth')
show_some_predictions(task2.train_dataloader(), task2.model, 0, 3)
# Validation set samples
show_some_predictions(task2.val_dataloader(), task2.model, 0, 3)
```
## Data structure check
Curious about data structure? here's how `file_to_vector_array()` does.
```
# simplify problem. if we use n_mels and frames as follows,
n_mels = 2
frames = 5
# and let's set wave length is short and mel spectrogram array length is,
vector_array_size = 50
# log_mel_spectrogram will be like this.
dims = frames * n_mels
tmp = np.array(list(range(vector_array_size + frames - 1)))
log_mel_spectrogram = np.c_[tmp, tmp+0.1].T
print('simply after converting wave into log mel spectrogram, log_mel_spectrogram will have shape like this:', log_mel_spectrogram.shape)
print(log_mel_spectrogram)
vector_array = np.zeros((vector_array_size, dims))
print(f"we'd like to arrange data structure and copy into a vector_array with shape {vector_array.shape} as follows:\n")
for t in range(frames):
vector_array[:, n_mels * t: n_mels * (t + 1)] = log_mel_spectrogram[:, t: t + vector_array_size].T
print(vector_array)
```
| github_jupyter |
```
import open3d as o3d
import numpy as np
import os
import sys
# monkey patches visualization and provides helpers to load geometries
sys.path.append('..')
import open3d_tutorial as o3dtut
# change to True if you want to interact with the visualization windows
o3dtut.interactive = not "CI" in os.environ
```
# RGBD integration
Open3D implements a scalable RGBD image integration algorithm. The algorithm is based on the technique presented in [\[Curless1996\]](../reference.html#curless1996) and [\[Newcombe2011\]](../reference.html#newcombe2011). In order to support large scenes, we use a hierarchical hashing structure introduced in [Integrater in ElasticReconstruction](https://github.com/qianyizh/ElasticReconstruction/tree/master/Integrate).
## Read trajectory from .log file
This tutorial uses the function `read_trajectory` to read a camera trajectory from a [.log file](http://redwood-data.org/indoor/fileformat.html). A sample `.log` file is as follows.
```
# examples/test_data/RGBD/odometry.log
0 0 1
1 0 0 2
0 1 0 2
0 0 1 -0.3
0 0 0 1
1 1 2
0.999988 3.08668e-005 0.0049181 1.99962
-8.84184e-005 0.999932 0.0117022 1.97704
-0.0049174 -0.0117024 0.999919 -0.300486
0 0 0 1
```
```
class CameraPose:
def __init__(self, meta, mat):
self.metadata = meta
self.pose = mat
def __str__(self):
return 'Metadata : ' + ' '.join(map(str, self.metadata)) + '\n' + \
"Pose : " + "\n" + np.array_str(self.pose)
def read_trajectory(filename):
traj = []
with open(filename, 'r') as f:
metastr = f.readline()
while metastr:
metadata = list(map(int, metastr.split()))
mat = np.zeros(shape=(4, 4))
for i in range(4):
matstr = f.readline()
mat[i, :] = np.fromstring(matstr, dtype=float, sep=' \t')
traj.append(CameraPose(metadata, mat))
metastr = f.readline()
return traj
redwood_rgbd = o3d.data.SampleRGBDDatasetRedwood()
camera_poses = read_trajectory(redwood_rgbd.odometry_log_path)
```
## TSDF volume integration
Open3D provides two types of TSDF volumes: `UniformTSDFVolume` and `ScalableTSDFVolume`. The latter is recommended since it uses a hierarchical structure and thus supports larger scenes.
`ScalableTSDFVolume` has several parameters. `voxel_length = 4.0 / 512.0` means a single voxel size for TSDF volume is $\frac{4.0\mathrm{m}}{512.0} = 7.8125\mathrm{mm}$. Lowering this value makes a high-resolution TSDF volume, but the integration result can be susceptible to depth noise. `sdf_trunc = 0.04` specifies the truncation value for the signed distance function (SDF). When `color_type = TSDFVolumeColorType.RGB8`, 8 bit RGB color is also integrated as part of the TSDF volume. Float type intensity can be integrated with `color_type = TSDFVolumeColorType.Gray32` and `convert_rgb_to_intensity = True`. The color integration is inspired by [PCL](http://pointclouds.org/).
```
volume = o3d.pipelines.integration.ScalableTSDFVolume(
voxel_length=4.0 / 512.0,
sdf_trunc=0.04,
color_type=o3d.pipelines.integration.TSDFVolumeColorType.RGB8)
for i in range(len(camera_poses)):
print("Integrate {:d}-th image into the volume.".format(i))
color = o3d.io.read_image(redwood_rgbd.color_paths[i])
depth = o3d.io.read_image(redwood_rgbd.depth_paths[i])
rgbd = o3d.geometry.RGBDImage.create_from_color_and_depth(
color, depth, depth_trunc=4.0, convert_rgb_to_intensity=False)
volume.integrate(
rgbd,
o3d.camera.PinholeCameraIntrinsic(
o3d.camera.PinholeCameraIntrinsicParameters.PrimeSenseDefault),
np.linalg.inv(camera_poses[i].pose))
```
## Extract a mesh
Mesh extraction uses the marching cubes algorithm [\[LorensenAndCline1987\]](../reference.html#lorensenandcline1987).
```
print("Extract a triangle mesh from the volume and visualize it.")
mesh = volume.extract_triangle_mesh()
mesh.compute_vertex_normals()
o3d.visualization.draw_geometries([mesh],
front=[0.5297, -0.1873, -0.8272],
lookat=[2.0712, 2.0312, 1.7251],
up=[-0.0558, -0.9809, 0.1864],
zoom=0.47)
```
<div class="alert alert-info">
**Note:**
TSDF volume works like a weighted average filter in 3D space. If more frames are integrated, the volume produces a smoother and nicer mesh. Please check [Make fragments](../reconstruction_system/make_fragments.rst) for more examples.
</div>
| github_jupyter |
```
s = 25
d=2
x=s+d
print(x)
print(" * ")
print(" *** ")
print(" ***** ")
print(" ******* ")
print(" ***** ")
print(" *** ")
print(" * ")
"fred"
str("judy")
int('4')
int(5.4)
float("33")
int('55')
s=22.2
str(s)
s,d,f =2.22,3,4
print(s)
print(d)
print(f)
a = 2
b = 5
print(a)
a = 3
a = b
print(a)
b = 7
print(a)
a=22
print(a)
del a
print(a)
a,s,d = 1,2,3
print(a,s,d)
del a,s,d
print(a,s,d)
a=5
aa=int(2+5)
a+aa
type(print)
print('Our good friend print')
from math import pi
print("Pi = {:.5}".format(pi) )
print("or", 3.14, "for short")
print("hello there my number is ",3.46563," and i am here")
round(33.33)
round(23.7)
x = 28793.54836
a = round(x,-1)
type(a)
print("A\nB\nC")
print("A\tB\tC")
# The \b and \a do not produce the desired results in the interactive shell, but they work properly in a command shell
print('WX\bYZ')
print('1\a2\a3\a4\a5\a6')
filename = 'C:\\Users\\rick'
print(filename)
print(r"F:\music\audio\songs")
print('Please enter some text:')
x = input()
print('Text entered:', x)
print('Type:', type(x))
a = input("enter a number : ")
print(int(a))
x = eval(input('Please enter an integer value: ') )
y = eval(input('Please enter another integer value: ') )
num1 = int(x)
num2 = int(y)
print(num1, '+', num2, '=', num1 + num2)
print('Please enter an integer value : ', end='')
a=input()
print(a)
print(end='Please enter an integer value :')
a=input()
print(a)
print('Please enter an integer value:', end='\n')
a=input()
print(a)
print('{0} {1}'.format(0, 10**0))
print('{0} {1}'.format(1, 10**1))
print('{0} {1}'.format(2, 10**2))
print('{0} {1}'.format(3, 10**3))
print('{0} {1}'.format(4, 10**4))
print('{0} {1}'.format(5, 10**5))
print('{0} {1}'.format(6, 10**6))
print('{0} {1}'.format(7, 10**7))
print('{0} {1}'.format(8, 10**8))
print('{0} {1}'.format(9, 10**9))
print('{0} {1}'.format(10, 10**10))
print('{0} {1}'.format(11, 10**11))
print('{0} {1}'.format(12, 10**12))
print('{0} {1}'.format(13, 10**13))
print('{0:>3} {1:>16}'.format(0, 10**0))
print('{0:>3} {1:>16}'.format(1, 10**1))
print('{0:>3} {1:>16}'.format(2, 10**2))
print('{0:>3} {1:>16}'.format(3, 10**3))
print('{0:>3} {1:>16}'.format(4, 10**4))
print('{0:>3} {1:>16}'.format(5, 10**5))
print('{0:>3} {1:>16}'.format(6, 10**6))
print('{0:>3} {1:>16}'.format(7, 10**7))
print('{0:>3} {1:>16}'.format(8, 10**8))
print('{0:>3} {1:>16}'.format(9, 10**9))
print('{0:>3} {1:>16}'.format(10, 10**10))
print('{0:>3} {1:>16}'.format(11, 10**11))
print('{0:>3} {1:>16}'.format(12, 10**12))
print('{0:>3} {1:>16}'.format(13, 10**13))
print('{0:>3} {1:>16}'.format(14, 10**14))
print('{0:>3} {1:>16}'.format(15, 10**15))
x = '''
A cube has 8 corners:
7------8
/| / |
3------4 |
| | | |
| 5----|-6
|/ |/
1------2
'''
print(x)
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
### Exercise 1
```
### Chapter 3 : Expressions and Arithmetic
| github_jupyter |
# Google Colab Init
```
from google.colab import drive
drive.mount('/content/drive')
# !pip install transformers==2.8.0
# !pip install deeppavlov
# !pip uninstall -y tensorflow tensorflow-gpu
# !pip install tensorflow-gpu==1.15.2
# !python -m deeppavlov install squad_bert
# !pip uninstall -y scikit-learn
# !pip install scikit-learn
# !pip install pandas
```
# RuBert DeepPavlov
```
import os
import pickle
import warnings
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from deeppavlov.core.common.file import read_json
from deeppavlov import build_model, configs, train_model
from deeppavlov.models.torch_bert.torch_transformers_classifier import TorchTransformersClassifierModel
from deeppavlov.models.preprocessors.torch_transformers_preprocessor import TorchTransformersPreprocessor
from sklearn.metrics import (f1_score, precision_score, average_precision_score, roc_auc_score,
classification_report, plot_roc_curve, accuracy_score, make_scorer,
plot_precision_recall_curve, precision_recall_curve, recall_score)
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
from tqdm.auto import tqdm
warnings.filterwarnings('ignore')
```
## Get data
```
data_path = '/content/drive/MyDrive/SarcasmDetection/data/Quotes'
dataname = 'rus-train-balanced-sarcasm-ling_feat.pkl'
with open(os.path.join(data_path, dataname), 'rb') as f:
df = shuffle(pickle.load(f), random_state=8)
train_df, test_df = train_test_split(df, test_size=0.3, random_state=8)
train_df, valid_df = train_test_split(train_df, test_size=0.1, random_state=8)
train_df = train_df.groupby(
'label', group_keys=False
).apply(lambda x: x.sample(n=(train_df.label == 0).sum())).sample(frac=1).reset_index(drop=True)
train_df[['rus_comment', 'label']].to_csv('train.csv', index=False)
valid_df[['rus_comment', 'label']].to_csv('valid.csv', index=False)
test_df[['rus_comment', 'label']].to_csv('test.csv', index=False)
```
## RuBert
```
def chunks(list_like, n):
for i in range(0, len(list_like), n):
yield list_like[i:i + n]
def show_test_classification_metrics(y_test, y_pred, y_pred_prob, X_test=None, classifier=None):
print(f"F1: {f1_score(y_test, y_pred):.5}")
print(f"PREC: {precision_score(y_test, y_pred):.5}")
print(f"PR-AUC: {average_precision_score(y_test, y_pred_prob):.5}")
print(f"ROC-AUC: {roc_auc_score(y_test, y_pred_prob):.5}")
print('-------------------------------------------------------')
print(classification_report(y_test, y_pred, labels=[0, 1]))
print('-------------------------------------------------------')
if classifier:
fig, ax = plt.subplots(1, 2, figsize=(15, 5))
ax[0].set_title('Precision-Recall curve')
plot_precision_recall_curve(classifier, X_test, y_test, ax=ax[0])
ax[1].set_title('ROC-AUC curve')
plot_roc_curve(classifier, X_test, y_test, ax=ax[1])
plt.show()
!python -m deeppavlov download paraphraser_rubert
bert_config = read_json(configs.classifiers.rusentiment_bert)
bert_config['dataset_reader']['x'] = 'rus_comment'
bert_config['dataset_reader']['y'] = 'label'
bert_config['dataset_reader']['data_path'] = './'
bert_config['dataset_reader']['train'] = 'train.csv'
bert_config['dataset_reader']['valid'] = 'valid.csv'
bert_config['dataset_reader']['test'] = 'test.csv'
del bert_config['dataset_iterator']['split_seed']
del bert_config['dataset_iterator']['field_to_split']
del bert_config['dataset_iterator']['split_fields']
del bert_config['dataset_iterator']['split_proportions']
bert_config['metadata']['variables']['MODEL_PATH'] = '/content/drive/MyDrive/SarcasmDetection/data/Models/classifiers/bert_classifier_rus_comment_ling_feat/'
del bert_config['chainer']['pipe'][-2:]
del bert_config['chainer']['pipe'][1]
bert_config['chainer']['pipe'][1]['in'] = 'y'
bert_config['chainer']['pipe'][1]['depth'] = 2
bert_config['chainer']['pipe'][2]['n_classes'] = 2
bert_config['train']['metrics'] = [bert_config['train']['metrics'][-1]]
bert_config['chainer']['out'] = ['y_pred_probas']
bert_config['train']['epochs'] = 2
bert_config['train']['batch_size'] = 32
bert_config['train']['show_examples'] = True
vocab_file = '{DOWNLOADS_PATH}/bert_models/rubert_cased_L-12_H-768_A-12_v1/vocab.txt'
bert_config_file = "{DOWNLOADS_PATH}/bert_models/rubert_cased_L-12_H-768_A-12_v1/bert_config.json"
pretrained_bert = "{DOWNLOADS_PATH}/bert_models/rubert_cased_L-12_H-768_A-12_v1/bert_model.ckpt"
bert_config['chainer']['pipe'][0]['vocab_file'] = vocab_file
bert_config['chainer']['pipe'][1]['bert_config_file'] = bert_config_file
bert_config['chainer']['pipe'][1]['pretrained_bert'] = pretrained_bert
bert_config['chainer']['pipe'][2]['bert_config_file'] = bert_config_file
bert_config['chainer']['pipe'][2]['pretrained_bert'] = pretrained_bert
m = train_model(bert_config)
preds_proba = []
for batch in tqdm(chunks(test_df['rus_comment'].values, 64), total=int(test_df.index.size / 64)):
preds_proba.append(m(batch))
preds = np.concatenate(preds_proba)
show_test_classification_metrics(
test_df.label.values,
(preds[:, 1] > 0.5).astype(int),
preds[:, 1]
)
```
| github_jupyter |
```
import numpy as np
from skimage import io
import glob
from measures import compute_ave_MAE_of_methods
print("------0. set the data path------")
# >>>>>>> Follows have to be manually configured <<<<<<< #
data_name = 'TEST-DATA' # this will be drawn on the bottom center of the figures
data_dir = '/content/drive/My Drive/COVID-SemiSeg/COVID-SemiSeg/Dataset/TestingSet/LungInfection-Test' # set the data directory,
# ground truth and results to-be-evaluated should be in this directory
# the figures of PR and F-measure curves will be saved in this directory as well
gt_dir = 'GT' # set the ground truth folder name
rs_dirs = ['FCN_pred','Res2Net_pred'] # set the folder names of different methods
# 'rs1' contains the result of method1
# 'rs2' contains the result of method 2
# we suggest to name the folder as the method names because they will be shown in the figures' legend
lineSylClr = ['r-','b-'] # curve style, same size with rs_dirs
linewidth = [2,1] # line width, same size with rs_dirs
gt_name_list = glob.glob(data_dir+'/'+gt_dir+'/'+'*.jpg')
# gt = io.imread(gt_name_list[0]) # read ground truth
# gt_name = gt_name_list[0].split('/')[-1]
# gt.shape
# from PIL import Image
# import os, sys
# path = '/content/drive/My Drive/COVID-SemiSeg/COVID-SemiSeg/Dataset/TestingSet/LungInfection-Test/GT/'
# dirs = os.listdir( path )
# def resize():
# for item in dirs:
# if os.path.isfile(path+item):
# im = Image.open(path+item)
# f, e = os.path.splitext(path+item)
# # print(f)
# imResize = im.resize((256,256), Image.ANTIALIAS)
# imResize.save(f + '.jpg', 'JPEG', quality=90)
# resize()
## get directory list of predicted maps
rs_dir_lists = []
for i in range(len(rs_dirs)):
rs_dir_lists.append(data_dir+'/'+rs_dirs[i]+'/')
print('\n')
rs_dir_lists
```
#For resnet and densenet
```
## 1. =======compute the average MAE of methods=========
print("------1. Compute the average MAE of Methods------")
aveMAE, gt2rs_mae = compute_ave_MAE_of_methods(gt_name_list,rs_dir_lists)
print('\n')
for i in range(0,len(rs_dirs)):
print('>>%s: num_rs/num_gt-> %d/%d, aveMAE-> %.3f'%(rs_dirs[i], gt2rs_mae[i], len(gt_name_list), aveMAE[i]))
## 2. =======compute the Precision, Recall and F-measure of methods=========
from measures import compute_PRE_REC_FM_of_methods,plot_save_pr_curves,plot_save_fm_curves
print('\n')
print("------2. Compute the Precision, Recall and F-measure of Methods------")
PRE, REC, FM, gt2rs_fm = compute_PRE_REC_FM_of_methods(gt_name_list,rs_dir_lists,beta=0.3)
for i in range(0,FM.shape[0]):
print(">>", rs_dirs[i],":", "num_rs/num_gt-> %d/%d,"%(int(gt2rs_fm[i][0]),len(gt_name_list)), "maxF->%.3f, "%(np.max(FM,1)[i]), "meanF->%.3f, "%(np.mean(FM,1)[i]))
print("Precision:{}".format(np.mean(PRE,1)[i]))
print('\n')
print("Recall:{}".format(np.mean(REC,1)[i]))
print('\n')
## 3. =======Plot and save precision-recall curves=========
print("------ 3. Plot and save precision-recall curves------")
plot_save_pr_curves(PRE, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
REC, # numpy array (num_rs_dir,255)
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.5,1.0), # the showing range of x-axis
yrange = (0.5,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
## 4. =======Plot and save F-measure curves=========
print("------ 4. Plot and save F-measure curves------")
plot_save_fm_curves(FM, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
mybins = np.arange(0,256),
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.0,1.0), # the showing range of x-axis
yrange = (0.0,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
print('Done!!!')
```
FOR Predictions and Resnet_Unet_Predictions
```
## 1. =======compute the average MAE of methods=========
print("------1. Compute the average MAE of Methods------")
aveMAE, gt2rs_mae = compute_ave_MAE_of_methods(gt_name_list,rs_dir_lists)
print('\n')
for i in range(0,len(rs_dirs)):
print('>>%s: num_rs/num_gt-> %d/%d, aveMAE-> %.3f'%(rs_dirs[i], gt2rs_mae[i], len(gt_name_list), aveMAE[i]))
## 2. =======compute the Precision, Recall and F-measure of methods=========
from measures import compute_PRE_REC_FM_of_methods,plot_save_pr_curves,plot_save_fm_curves
print('\n')
print("------2. Compute the Precision, Recall and F-measure of Methods------")
PRE, REC, FM, gt2rs_fm = compute_PRE_REC_FM_of_methods(gt_name_list,rs_dir_lists,beta=0.3)
for i in range(0,FM.shape[0]):
print(">>", rs_dirs[i],":", "num_rs/num_gt-> %d/%d,"%(int(gt2rs_fm[i][0]),len(gt_name_list)), "maxF->%.3f, "%(np.max(FM,1)[i]), "meanF->%.3f, "%(np.mean(FM,1)[i]))
print('\n')
## 3. =======Plot and save precision-recall curves=========
print("------ 3. Plot and save precision-recall curves------")
plot_save_pr_curves(PRE, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
REC, # numpy array (num_rs_dir,255)
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.5,1.0), # the showing range of x-axis
yrange = (0.5,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
## 4. =======Plot and save F-measure curves=========
print("------ 4. Plot and save F-measure curves------")
plot_save_fm_curves(FM, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
mybins = np.arange(0,256),
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.0,1.0), # the showing range of x-axis
yrange = (0.0,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
print('Done!!!')
```
FOR res_pred and Unet_predictions2 (Dont use pred and Predictions2)
```
rs_dir_lists = []
for i in range(len(rs_dirs)):
rs_dir_lists.append(data_dir+'/'+rs_dirs[i]+'/')
print('\n')
rs_dir_lists
## 1. =======compute the average MAE of methods=========
print("------1. Compute the average MAE of Methods------")
aveMAE, gt2rs_mae = compute_ave_MAE_of_methods(gt_name_list,rs_dir_lists)
print('\n')
for i in range(0,len(rs_dirs)):
print('>>%s: num_rs/num_gt-> %d/%d, aveMAE-> %.3f'%(rs_dirs[i], gt2rs_mae[i], len(gt_name_list), aveMAE[i]))
## 2. =======compute the Precision, Recall and F-measure of methods=========
from measures import compute_PRE_REC_FM_of_methods,plot_save_pr_curves,plot_save_fm_curves
print('\n')
print("------2. Compute the Precision, Recall and F-measure of Methods------")
PRE, REC, FM, gt2rs_fm = compute_PRE_REC_FM_of_methods(gt_name_list,rs_dir_lists,beta=0.3)
for i in range(0,FM.shape[0]):
print(">>", rs_dirs[i],":", "num_rs/num_gt-> %d/%d,"%(int(gt2rs_fm[i][0]),len(gt_name_list)), "maxF->%.3f, "%(np.max(FM,1)[i]), "meanF->%.3f, "%(np.mean(FM,1)[i]))
print('\n')
## 3. =======Plot and save precision-recall curves=========
print("------ 3. Plot and save precision-recall curves------")
plot_save_pr_curves(PRE, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
REC, # numpy array (num_rs_dir,255)
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.5,1.0), # the showing range of x-axis
yrange = (0.5,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
## 4. =======Plot and save F-measure curves=========
print("------ 4. Plot and save F-measure curves------")
plot_save_fm_curves(FM, # numpy array (num_rs_dir,255), num_rs_dir curves will be drawn
mybins = np.arange(0,256),
method_names = rs_dirs, # method names, shape (num_rs_dir), will be included in the figure legend
lineSylClr = lineSylClr, # curve styles, shape (num_rs_dir)
linewidth = linewidth, # curve width, shape (num_rs_dir)
xrange = (0.0,1.0), # the showing range of x-axis
yrange = (0.0,1.0), # the showing range of y-axis
dataset_name = data_name, # dataset name will be drawn on the bottom center position
save_dir = data_dir, # figure save directory
save_fmt = 'png') # format of the to-be-saved figure
print('\n')
print('Done!!!')
```
| github_jupyter |
# Commodity price forecasting using CNN-GRU
```
import pandas as pd
import numpy as np
import os
import time
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
# preprocessing methods
from sklearn.preprocessing import StandardScaler
# accuracy measures and data spliting
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import train_test_split
# deep learning libraries
from keras.models import Input, Model
from keras.models import Sequential
from keras.layers import SimpleRNN, LSTM, Dense, GRU
from keras.layers import Conv1D, MaxPooling1D
from keras.layers import Flatten
from keras import layers
from keras import losses
from keras import optimizers
from keras import metrics
from keras import callbacks
from keras import initializers
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = 15, 7
```
## 1. Data import and basic analysis
```
DATADIR = 'data/'
MODELDIR = '../checkpoints/commodity/nn/'
path = os.path.join(DATADIR, 'gold-silver.csv')
data = pd.read_csv(path, header=0, index_col=[0], infer_datetime_format=True, sep=';')
data.head()
data[['gold', 'silver']].plot();
plt.plot(np.log(data.gold), label='log(gold)')
plt.plot(np.log(data.silver), label='log(silver)')
plt.title('Commodity data', fontsize='14')
plt.show()
```
## 2. Data preparation
```
# function to prepare x and y variable
# for the univariate series
def prepare_data(df, steps=1):
temp = df.shift(-steps).copy()
y = temp[:-steps].copy()
X = df[:-steps].copy()
return X, y
gold_X, gold_y = prepare_data(np.log(data[['gold']]), steps=1)
silver_X, silver_y = prepare_data(np.log(data[['silver']]), steps=1)
len(gold_X), len(gold_y), len(silver_X), len(silver_y)
X = pd.concat([gold_X, silver_X], axis=1)
y = pd.concat([gold_y, silver_y], axis=1)
X.head()
y.head()
seed = 42
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.05,
random_state=seed, shuffle=False)
print('Training and test data shape:')
X_train.shape, y_train.shape, X_test.shape, y_test.shape
timesteps = 1
features = X_train.shape[1]
xavier = initializers.glorot_normal()
X_train = np.reshape(X_train.values, (X_train.shape[0], timesteps, features))
X_test = np.reshape(X_test.values, (X_test.shape[0], timesteps, features))
X_train.shape, X_test.shape, y_train.shape, y_test.shape
```
## 3. Model building
```
def model_evaluation(y_train, y_test, y_train_pred, y_test_pred):
y_train_inv, y_test_inv = np.exp(y_train), np.exp(y_test)
y_train_pred_inv, y_test_pred_inv = np.exp(y_train_pred), np.exp(y_test_pred)
# MAE and NRMSE calculation for gold
y_train_gold = y_train_inv.values[:, 0]
y_train_pred_gold = y_train_pred_inv[:, 0]
y_test_gold = y_test_inv.values[:, 0]
y_test_pred_gold = y_test_pred_inv[:, 0]
train_rmse_g = np.sqrt(mean_squared_error(y_train_gold, y_train_pred_gold))
train_mae_g = np.round(mean_absolute_error(y_train_gold, y_train_pred_gold), 3)
train_nrmse_g = np.round(train_rmse_g/np.std(y_train_gold), 3)
test_rmse_g = np.sqrt(mean_squared_error(y_test_gold, y_test_pred_gold))
test_mae_g = np.round(mean_absolute_error(y_test_gold, y_test_pred_gold), 3)
test_nrmse_g = np.round(test_rmse_g/np.std(y_test_gold), 3)
print('Training and test result for gold:')
print(f'Training MAE: {train_mae_g}')
print(f'Trainig NRMSE: {train_nrmse_g}')
print(f'Test MAE: {test_mae_g}')
print(f'Test NRMSE: {test_nrmse_g}')
print()
# MAE and NRMSE calculation for silver
y_train_silver = y_train_inv.values[:, 1]
y_train_pred_silver = y_train_pred_inv[:, 1]
y_test_silver = y_test_inv.values[:, 1]
y_test_pred_silver = y_test_pred_inv[:, 1]
train_rmse_s = np.sqrt(mean_squared_error(y_train_silver, y_train_pred_silver))
train_mae_s = np.round(mean_absolute_error(y_train_silver, y_train_pred_silver), 3)
train_nrmse_s = np.round(train_rmse_s/np.std(y_train_silver), 3)
test_rmse_s = np.sqrt(mean_squared_error(y_test_silver, y_test_pred_silver))
test_mae_s = np.round(mean_absolute_error(y_test_silver, y_test_pred_silver), 3)
test_nrmse_s = np.round(test_rmse_s/np.std(y_test_silver), 3)
print('Training and test result for silver:')
print(f'Training MAE: {train_mae_s}')
print(f'Trainig NRMSE: {train_nrmse_s}')
print(f'Test MAE: {test_mae_s}')
print(f'Test NRMSE: {test_nrmse_s}')
return y_train_pred_inv, y_test_pred_inv
def model_training(X_train, X_test, y_train, model, batch=4, name='m'):
start = time.time()
loss = losses.mean_squared_error
opt = optimizers.Adam()
metric = [metrics.mean_absolute_error]
model.compile(loss=loss, optimizer=opt, metrics=metric)
callbacks_list = [callbacks.ReduceLROnPlateau(monitor='loss', factor=0.2,
patience=5, min_lr=0.001)]
history = model.fit(X_train, y_train,
epochs=100,
batch_size=batch,
verbose=0,
shuffle=False,
callbacks=callbacks_list
)
# save model weights and
if os.path.exists(MODELDIR):
pass
else:
os.makedirs(MODELDIR)
m_name = name + str('.h5')
w_name = name + str('_w.h5')
model.save(os.path.join(MODELDIR, m_name))
model.save_weights(os.path.join(MODELDIR, w_name))
# prediction
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
end = time.time()
time_taken = np.round((end-start), 3)
print(f'Time taken to complete the process: {time_taken} seconds')
return y_train_pred, y_test_pred, history
```
### CNN-GRU - v1
```
model = Sequential()
model.add(Conv1D(3, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(GRU(3, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=4, name='cnngru-v1')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU- v2
```
model = Sequential()
model.add(Conv1D(8, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(GRU(8, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=4, name='cnngru-v2')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU - v3 (Final Model)
```
model = Sequential()
model.add(Conv1D(12, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(GRU(12, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=4, name='cnngru-v3')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU - v4
```
model = Sequential()
model.add(Conv1D(8, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(Conv1D(8, kernel_size=3, activation='relu', padding='same', strides=1,
kernel_initializer=xavier))
model.add(GRU(8, kernel_initializer=xavier, activation='relu', return_sequences=True))
model.add(GRU(8, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=4, name='cnngru-v4')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU - v5
```
model = Sequential()
model.add(Conv1D(3, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(Conv1D(3, kernel_size=3, activation='relu', padding='same', strides=1,
kernel_initializer=xavier))
model.add(GRU(3, kernel_initializer=xavier, activation='relu', return_sequences=True))
model.add(GRU(3, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=4, name='cnngru-v5')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU - v6
```
model = Sequential()
model.add(Conv1D(8, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(GRU(8, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=1, name='cnngru-v6')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
### CNN-GRU - v7
```
model = Sequential()
model.add(Conv1D(8, kernel_size=3, activation='relu', padding='same', strides=1,
input_shape=(timesteps, features), kernel_initializer=xavier))
model.add(GRU(8, kernel_initializer=xavier, activation='relu'))
model.add(Dense(2, kernel_initializer=xavier))
model.summary()
# training
y_train_pred, y_test_pred, history = model_training(X_train, X_test, y_train, model, batch=2, name='cnngru-v7')
# evaluation
y_train_pred, y_test_pred = model_evaluation(y_train, y_test, y_train_pred, y_test_pred)
# plotting
plt.subplot(211)
plt.plot(np.exp(y_test.values[:, 0]), label='actual')
plt.plot(y_test_pred[:, 0], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for gold using CNN-GRU', fontsize=14)
plt.legend()
plt.subplot(212)
plt.plot(np.exp(y_test.values[:, 1]), label='actual')
plt.plot(y_test_pred[:, 1], label='predicted')
plt.ylabel('$')
plt.xlabel('sample')
plt.title('Test prediction for silver using CNN-GRU', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
```
| github_jupyter |
```
import numpy as np
import random
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data.dataset import Dataset
from torch.utils.data import DataLoader
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, roc_auc_score
from secure_ml.attack import ShadowModel, AttackerModel, Membership_Inference
from secure_ml.utils import DataSet
```
# Check GPU
```
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
print(device)
```
# Load data
```
# トレーニングデータをダウンロード
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True)
# テストデータをダウンロード
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True)
X_train = trainset.data
y_train = np.array(trainset.targets)
X_test = testset.data
y_test = np.array(testset.targets)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
victim_idx = random.sample(range(X_train.shape[0]), k=2000)
attack_idx = random.sample(range(X_test.shape[0]), k=6000)
shadow_idx = attack_idx[:4000]
eval_idx = attack_idx[4000:]
X_victim = X_train[victim_idx]
y_victim = y_train[victim_idx]
X_shadow = X_test[shadow_idx]
y_shadow = y_test[shadow_idx]
X_eval = X_test[eval_idx]
y_eval = y_test[eval_idx]
print(X_victim.shape, y_victim.shape)
print(X_shadow.shape, y_shadow.shape)
print(X_eval.shape, y_eval.shape)
# ToTensor:画像のグレースケール化(RGBの0~255を0~1の範囲に正規化)、Normalize:Z値化(RGBの平均と標準偏差を0.5で決め打ちして正規化)
transform = transforms.Compose([transforms.ToTensor(), transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
victimset = DataSet(X_victim, y_victim, transform=transform)
victimloader = torch.utils.data.DataLoader(victimset, batch_size=4, shuffle=True, num_workers=2)
valset = DataSet(X_eval, y_eval, transform=transform)
valloader = torch.utils.data.DataLoader(valset, batch_size=4, shuffle=True, num_workers=2)
X_victim.shape
```
# Define and Train a victim Model
```
# CNNを実装する
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=64, kernel_size=3, padding=1, stride=1)
self.bn1 = nn.BatchNorm2d(64)
self.conv2 = nn.Conv2d(64, 128, 3, padding=1)
self.bn2 = nn.BatchNorm2d(128)
self.conv3 = nn.Conv2d(128, 256, 3, padding=1)
self.bn3 = nn.BatchNorm2d(256)
self.conv4 = nn.Conv2d(256, 512, 3, padding=1)
self.bn4 = nn.BatchNorm2d(512)
self.L1 = nn.Linear(2048, 10) # 10クラス分類
def forward(self, x):
# 3ch > 64ch, shape 32 x 32 > 16 x 16
x = self.conv1(x) # [64,32,32]
x = self.bn1(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [64,16,16]
# 64ch > 128ch, shape 16 x 16 > 8 x 8
x = self.conv2(x) # [128,16,16]
x = self.bn2(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [128,8,8]
# 128ch > 256ch, shape 8 x 8 > 4 x 4
x = self.conv3(x) # [256,8,8]
x = self.bn3(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [256,4,4]
# 256ch > 512ch, shape 4 x 4 > 2 x 2
x = self.conv4(x) # [512,4,4]
x = self.bn4(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [512,2,2]
# 全結合層
x = x.view( -1, 2048) # [256,2048]
x = self.L1(x)
#x = F.softmax(x, dim=0)
return x
victim_net = Net()
victim_net = victim_net.to(device)
# 交差エントロピー
criterion = nn.CrossEntropyLoss()
# 確率的勾配降下法
optimizer = optim.SGD(victim_net.parameters(), lr=0.005, momentum=0.9)
for epoch in range(20): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(victimloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
labels = labels.to(torch.int64)
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = victim_net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
test_preds = []
test_label = []
with torch.no_grad():
for data in valloader:
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
outputs = victim_net(inputs)
test_preds.append(outputs)
test_label.append(labels)
test_preds = torch.cat(test_preds)
test_label = torch.cat(test_label)
print(accuracy_score(torch.argmax(test_preds, axis=1).cpu().detach().numpy(),
test_label.cpu().detach().numpy()))
print('Finished Training')
in_preds = []
in_label = []
with torch.no_grad():
for data in victimloader:
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
outputs = victim_net(inputs)
in_preds.append(outputs)
in_label.append(labels)
in_preds = torch.cat(in_preds)
in_label = torch.cat(in_label)
print("train_accuracy: ",
accuracy_score(torch.argmax(in_preds, axis=1).cpu().detach().numpy(),
in_label.cpu().detach().numpy()))
out_preds = []
out_label = []
with torch.no_grad():
for data in valloader:
inputs, labels = data
inputs = inputs.to(device)
labels = labels.to(device)
outputs = victim_net(inputs)
out_preds.append(outputs)
out_label.append(labels)
out_preds = torch.cat(out_preds)
out_label = torch.cat(out_label)
print("test_accuracy: ",
accuracy_score(torch.argmax(out_preds, axis=1).cpu().detach().numpy(),
out_label.cpu().detach().numpy()))
```
you can see how overfitting the victim model is for each label
```
in_pred_numpy = torch.argmax(in_preds, axis=1).cpu().detach().numpy()
in_label_numpy = in_label.cpu().detach().numpy()
out_pred_numpy = torch.argmax(out_preds, axis=1).cpu().detach().numpy()
out_label_numpy = out_label.cpu().detach().numpy()
target_model_accuracy_per_label = {}
print("train_accuracy - test_accuracy")
for label in np.unique(in_label_numpy):
in_label_idx = np.where(in_label_numpy == label)
out_label_idx = np.where(out_label_numpy == label)
train_score = accuracy_score(in_pred_numpy[in_label_idx],
in_label_numpy[in_label_idx])
test_score = accuracy_score(out_pred_numpy[out_label_idx],
out_label_numpy[out_label_idx])
print(f"label {label}: ", train_score - test_score)
target_model_accuracy_per_label[label] = train_score - test_score
```
# Memership inference (shadow models + attack models)
```
# CNNを実装する
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=64, kernel_size=3, padding=1, stride=1)
self.bn1 = nn.BatchNorm2d(64)
self.conv2 = nn.Conv2d(64, 128, 3, padding=1)
self.bn2 = nn.BatchNorm2d(128)
self.conv3 = nn.Conv2d(128, 256, 3, padding=1)
self.bn3 = nn.BatchNorm2d(256)
self.conv4 = nn.Conv2d(256, 512, 3, padding=1)
self.bn4 = nn.BatchNorm2d(512)
self.L1 = nn.Linear(2048, 10) # 10クラス分類
def forward(self, x):
# 3ch > 64ch, shape 32 x 32 > 16 x 16
x = self.conv1(x) # [64,32,32]
x = self.bn1(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [64,16,16]
# 64ch > 128ch, shape 16 x 16 > 8 x 8
x = self.conv2(x) # [128,16,16]
x = self.bn2(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [128,8,8]
# 128ch > 256ch, shape 8 x 8 > 4 x 4
x = self.conv3(x) # [256,8,8]
x = self.bn3(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [256,4,4]
# 256ch > 512ch, shape 4 x 4 > 2 x 2
x = self.conv4(x) # [512,4,4]
x = self.bn4(x)
x = F.relu(x)
x = F.max_pool2d(x, 2, 2) # [512,2,2]
# 全結合層
x = x.view( -1, 2048) # [256,2048]
x = self.L1(x)
#x = F.softmax(x, dim=0)
return x
shadow_models = [Net().to(device),
Net().to(device)]
shadow_data_size = 2000
shadow_transform = transform
num_label = 10
attack_models = [SVC(probability=True) for i in range(num_label)]
y_test = np.array(y_test).astype(np.int64)
mi = Membership_Inference(victim_net, shadow_models, attack_models,
shadow_data_size, shadow_transform)
mi.train_shadow(X_test, y_test, num_itr=1)
mi.train_attacker()
attacked_pred_in_prob = mi.predict_proba(in_preds, in_label)
attacked_pred_out_prob = mi.predict_proba(out_preds, out_label)
score = roc_auc_score(np.concatenate([np.ones_like(attacked_pred_in_prob),
np.zeros_like(attacked_pred_out_prob)]),
np.concatenate([attacked_pred_in_prob, attacked_pred_out_prob])
)
print("overall auc is ", score)
in_label_numpy = in_label.cpu().numpy()
out_label_numpy = out_label.cpu().numpy()
attack_model_auc_per_label = {}
for label in np.unique(in_label_numpy):
in_label_idx = np.where(in_label_numpy == label)
out_label_idx = np.where(out_label_numpy == label)
score = roc_auc_score(np.concatenate([np.ones_like(attacked_pred_in_prob[in_label_idx]),
np.zeros_like(attacked_pred_out_prob[out_label_idx])]),
np.concatenate([attacked_pred_in_prob[in_label_idx],
attacked_pred_out_prob[out_label_idx]])
)
print(f"label {label}: ", score)
attack_model_auc_per_label[label] = score
for i in range(10):
plt.scatter(list(target_model_accuracy_per_label.values())[i],
list(attack_model_auc_per_label.values())[i],
marker=f"${i}$")
plt.title("overfitting - membership inference performance")
plt.xlabel("victim model: trian_accuracy - test_accuracy per class")
plt.ylabel("attack model: auc per clas# s")
plt.savefig("membership_inference_overfitting.png")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import cv2
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
from keras.models import Sequential
from keras.layers import Dense, Flatten, Conv2D, MaxPool2D, Dropout
from tensorflow.keras.optimizers import SGD
from keras import optimizers
from keras.callbacks import ReduceLROnPlateau, EarlyStopping
from keras.utils.np_utils import to_categorical
data = pd.read_csv(r"L:\Handwriting Recognition\archive\A_Z Handwritten Data.csv").astype('float32')
data.head()
df = pd.DataFrame(data)
df.head()
df.shape
df.info()
df.describe()
df.isnull().sum()
#splitting
x = df.drop('0', axis = 1)
y = df['0']
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.2)
x_train = np.reshape(x_train.values, (x_train.shape[0], 28, 28))
x_test = np.reshape(x_test.values, (x_test.shape[0], 28, 28))
print('Train Data Shape:', x_train.shape)
print('Test Data Shape:', x_test.shape)
word_dict = {
0:'A',1:'B',2:'C',3:'D',4:'E',5:'F',6:'G',7:'H',8:'I',9:'J',10:'K',11:'L',12:'M',13:'N',14:'O',15:'P',16:'Q',17:'R',18:'S',19:'T',20:'U',21:'V',22:'W',23:'X', 24:'Y',25:'Z'
}
#plt.style.use('fivethirtyeight')
#plt.xkcd()
y_integer = np.int0(y)
count = np.zeros(26, dtype = 'int')
for i in y_integer:
count[i] += 1
alphabets = []
for i in word_dict.values():
alphabets.append(i)
fig, ax = plt.subplots(1, 1, figsize = (15, 15))
ax.barh(alphabets, count)
plt.xlabel('Number Of Elements..!!', fontsize = 20, fontweight = 'bold', color = 'green')
plt.ylabel('Alphabets..!!', fontsize = 30, fontweight = 'bold', color = 'green')
plt.grid()
plt.show()
#plotting and shuffling
#plt.style.use('fivethirtyeight')
#plt.xkcd()
shuff = shuffle(x_train[:100])
fig, ax = plt.subplots(3, 3, figsize = (15, 15))
axes = ax.flatten()
for i in range(9):
shu = cv2.threshold(shuff[i], 30, 200, cv2.THRESH_BINARY)
axes[i].imshow(np.reshape(shuff[i], (28, 28)), cmap = 'Greys')
plt.show()
x_train = x_train.reshape(x_train.shape[0], x_train.shape[1], x_train.shape[2], 1)
print("New shape of train data:", x_train.shape)
x_test = x_test.reshape(x_test.shape[0], x_test.shape[1], x_test.shape[2], 1)
print("New shape of test data:", x_test.shape)
categorical_train = to_categorical(y_train, num_classes = 26, dtype = 'int')
print("New shape of train labels:", categorical_train.shape)
categorical_test = to_categorical(y_test, num_classes = 26, dtype = 'int')
print("New shape of test labels:", categorical_test.shape)
my_model = Sequential()
my_model.add(Conv2D(filters = 32, kernel_size = (3, 3), activation = 'relu', input_shape = (28, 28, 1)))
my_model.add(MaxPool2D(pool_size = (2, 2), strides = 2))
my_model.add(Conv2D(filters = 64, kernel_size = (3, 3), activation = 'relu', padding = 'same'))
my_model.add(MaxPool2D(pool_size = (2, 2), strides = 2))
my_model.add(Conv2D(filters = 128, kernel_size = (3, 3), activation = 'relu', padding = 'valid'))
my_model.add(MaxPool2D(pool_size = (2, 2), strides = 2))
my_model.add(Flatten())
my_model.add(Dense(64, activation = "relu"))
my_model.add(Dense(128, activation = "relu"))
my_model.add(Dense(26, activation = "softmax"))
my_model.compile(optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['accuracy'])
history = my_model.fit(x_train, categorical_train, epochs = 1, validation_data = (x_test, categorical_test))
my_model.summary()
my_model.save(r'Handwriting_rec.h5')
print("The validation accuracy is :", history.history['val_accuracy'])
print("The training accuracy is :", history.history['accuracy'])
print("The validation loss is :", history.history['val_loss'])
print("The training loss is :", history.history['loss'])
fig, axes = plt.subplots(3, 3, figsize = (12, 15))
axes = axes.flatten()
for i, ax in enumerate(axes):
img = np.reshape(x_test[i], (28, 28))
ax.imshow(img, cmap = 'Greys')
pred = word_dict[np.argmax(categorical_test[i])]
ax.set_title("Prediction: " + pred, fontsize = 20, fontweight = 'bold', color = 'red')
ax.grid()
```
| github_jupyter |
```
# Load dependencies
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
pd.options.display.float_format = '{:,.1e}'.format
from scipy.stats import gmean
import sys
sys.path.insert(0, '../../../statistics_helper')
from CI_helper import *
from excel_utils import *
```
# Estimating the biomass of fish
To estimate the biomass of fish, we first estimate the total biomass of mesopelagic fish, and then add to this estimate the estmimate for the non-mesopelagic fish made by [Wilson et al.](http://dx.doi.org/10.1126/science.1157972).
In order to estimate the biomass of mesopelagic fish, we rely on two independent methods - and estimate based on trawling by [Lam & Pauly](http://www.seaaroundus.org/doc/Researcher+Publications/dpauly/PDF/2005/OtherItems/MappingGlobalBiomassMesopelagicFishes.pdf), and an estimate based on sonar.
## Sonar-based estimate
We generate the sonar-based estimate relying on data from [Irigoien et al.](http://dx.doi.org/10.1038/ncomms4271) and [Proud et al.](http://dx.doi.org/10.1016/j.cub.2016.11.003).
Estimating the biomass of mesopelagic fish using sonar is a two step process. First we use estimates of the global backscatter of mesopelagic fish. This backscatter is converted to an estimate of the global biomass of mesopelagic fish by using estimates for the target strength of a single mesopelagic fish.
### Total backscatter
To estimate the total backscatter of mesopelagic fish, we rely on [Irigoien et al.](http://dx.doi.org/10.1038/ncomms4271) and [Proud et al.](http://dx.doi.org/10.1016/j.cub.2016.11.003). Irigoien et al. generates several different estimates for the global nautical area scatter of mesopelagic fish. We use the geometric mean of the estimates of Irigoien et al. as one source for estimating the total backscatter of mesopelagic fish. We note that the units used by Irigoien et al. are incorrect, as nautical area scatteing coefficient (NASC) is measured in $\frac{m^2}{nm^2}$, but the order of magnitude of the values estimated by Irigoien et al. implies that they multiplied the NASC by the surface area of the ocean in units of $m^2$. This means that the values reported by Irigoien et al. are in fact in units of $\frac{m^4}{nm^2}$. We convert the values reported by Irigoein et al. from the total scatter to the total backscatter by using the equation: $$global \: backscatter \: [m^2] = \frac{global \: scatter \: [\frac{m^4}{nmi^2}]}{4\pi×\frac{1852^2 m^2}{nmi^2}}$$
```
# Load scatter data from Irigoien et al.
scatter = pd.read_excel('fish_biomass_data.xlsx', 'Total scatter',skiprows=1)
# convert scater to backscatter
scatter['Total backscatter [m^2]'] = scatter['Total sA [m^4 nmi^-2]']/(4*np.pi*1852**2)
scatter['Total sA [m^4 nmi^-2]'] = scatter['Total sA [m^4 nmi^-2]'].astype(float)
scatter
# Calculate the geometric mean of values from Irigoien et al.
irigoien_backscatter = gmean(scatter['Total backscatter [m^2]'])
print('The geometric mean of global backscatter from Irigoien et al. is ≈%.1e m^2' %irigoien_backscatter)
```
As our best estimate for the global backscatter of mesopelagic fish, we use the geometric mean of the average value from Irigoien et al. and the value reported in Proud et al.
```
# The global backscatter reported by Proud et al.
proud_backscatter = 6.02e9
# Our best estimate
best_backscatter = gmean([irigoien_backscatter,proud_backscatter])
print('Our best estimate for the global backscatter of mesapelagic fish is %.0e m^2' %best_backscatter)
```
### Target strength
In order to convert the global backscatter into biomass, we use reported values for the target strength per unit biomass of mesopelagic fish. The target strength is a measure of the the backscattering cross-section in dB, which is defined as $TS = 10 \times log_{10}(\sigma_{bs})$ with units of dB 1 re $m^2$. By measuring the relation between the target strength and biomass of mesopelagic fish, one can calculate the target strength per unit biomass in units of db 1 re $\frac{m^2}{kg}$. We can use the global backscatter to calculate the total biomass of mesopelagic fish based on the equation provided in [MacLennan et al.](https://doi.org/10.1006/jmsc.2001.1158): $$biomass_{fish} \:[kg]= \frac{global \: backscatter \: [m^2]}{10^{\frac{TS_{kg}}{10}} [m^2 kg^{-1}]}$$
Where $TS_{kg}$ is the terget strength per kilogram biomass.
The main source affecting the target strength of mesopelagic fish is their swimbaldder, as the swimbladder serves as a strong acoustic reflector at the frequencies used to measure the backscattering of mesopelagic fish. Irigoien et al. provide a list of values from the literature of target strength per unit biomass for mesopelagic fish with or without swimbladder. It is clear from the data that the presence or absence of swimbladder segregates the data into two distinct groups:
```
# Load terget strength data
ts = pd.read_excel('fish_biomass_data.xlsx', 'Target strength',skiprows=1)
# Plot the distribution of TS for fish with or without swimbladder
ts[ts['Swimbladder']=='No']['dB kg^-1'].hist(label='No swimbladder', bins=3)
ts[ts['Swimbladder']=='Yes']['dB kg^-1'].hist(label='With swimbladder', bins=3)
plt.legend()
plt.xlabel(r'Target strength per unit biomass dB kg$^{-1}$')
plt.ylabel('Counts')
```
To estimate the characteristic target strength per unit biomass of mesopelagic fish, we first estiamte the characteristic target strength per unit biomass of fish with or without swimbladder. We assume that fish with and without swimbladder represent an equal portion of the population of mesopelagic fish. We test the uncertainty associated with this assumption in the uncertainty analysis section.
```
# Calculate the average TS per kg for fish with and without swimbladder
TS_bin = ts.groupby('Swimbladder').mean()
TS_bin['dB kg^-1']
```
We use our best estimate for the target strength per unit biomass to estimate the total biomass of mesopelagic fish. We transform the TS to backscattering cross-section, and then calculate the effective population backscattering cross-section based on the assumption that fish with or without swimbladder represent equal portions of the population.
```
# The conversion equation from global backscatter and terget strength per unit biomass
biomass_estimator = lambda TS1,TS2,bs,frac: bs/(frac*10**(TS1/10.) + (1.-frac)*10**(TS2/10.))
# Estimate biomass and convert to g C, assuming fish with or without swimbladder are both 50% of the population
sonar_biomass = biomass_estimator(*TS_bin['dB kg^-1'],best_backscatter,frac=0.5)*1000*0.15
print('Our best sonar-based estimate for the biomass of mesopelagic fish is ≈%.1f Gt C' %(sonar_biomass/1e15))
```
As noted in the Supplementary Information, there are several caveats which might bias the results. We use the geometric mean of estimates based on sonar and earlier estimates based on trawling to generate a robust estimate for the biomass of mesopelagic fish.
```
# The estimate of the global biomass of mesopelagic fish based on trawling reported in Lan & Pauly
trawling_biomass = 1.5e14
# Estimate the biomass of mesopelagic fish based on the geometric mean of sonar-based and trawling-based estimates
best_mesopelagic_biomass = gmean([sonar_biomass,trawling_biomass])
print('Our best estimate for the biomass of mesopelagic fish is ≈%.1f Gt C' %(best_mesopelagic_biomass/1e15))
```
Finally, we add to our estimate of the biomass of mesopelagic fish the estimate of biomass of non-mesopelagic fish made by [Wilson et al.](http://dx.doi.org/10.1126/science.1157972) to generate our estimate for the total biomass of fish.
```
# The estimate of non-mesopelagic fish based on Wilson et al.
non_mesopelagic_fish_biomass = 1.5e14
best_estimate = best_mesopelagic_biomass+non_mesopelagic_fish_biomass
print('Our best estimate for the biomass of fish is ≈%.1f Gt C' %(best_estimate/1e15))
```
# Uncertainty analysis
In order to assess the uncertainty associated with our estimate for the biomass of fish, we assess the uncertainty associated with the sonar-based estimate of the biomass of mesopelagic fish, as well as for the non-mesopelagic fish biomass.
## Mesopelagic fish uncertainty
To quantify the uncertainty associated with our estimate of the biomass of mesopelagic fish, we assess the uncertainty associated with the sonar-based estimate, and the inter-method uncertainty between the sonar-based and trawling-based estimates. We do not assess the uncertainty of the trawling-based estimate as no data regarding the uncertainty of the estimate is available.
### Sonar-based estimate uncertainty
The main parameters influencing the uncertainty of the sonar-based estimates are the global backscatter and the characteristic target-strength per unit biomass. We calculate the uncertainty associated with each one of those parameters, and them combine these uncertainties to quantify the uncertainty of the sonar-based estimate.
#### Global Backscatter
For calculating the global backscatter, we rely in two sources of data - Data from Irigoien et al. and data from Proud et al. We survery both the intra-study uncertainty and interstudy uncertainty associated with the global backscatter.
##### Intra-study uncertainty
Irigoien et al. provides several estimates for the global scatter based on several different types of equations characterizing the relationship between primary productivity and the NASC. We calculate the 95% confidence interval of the geometric mean of these different estimates.
Proud et al. estimate a global backscatter of 6.02×$10^9$ $m^2$ ± 1.4×$10^9$ $m^2$. We thus use this range as a measure of the intra-study uncertainty in the estimate of Proud et al.
```
# Calculate the intra-study uncertainty of Irigoien et al.
irigoien_CI = geo_CI_calc(scatter['Total backscatter [m^2]'])
# Calculate the intra-study uncertainty of Proud et al.
proud_CI = (1.4e9+6.02e9)/6.02e9
print('The intra-study uncertainty of the total backscatter estimate of Irigoien et al. is ≈%.1f-fold' %irigoien_CI)
print('The intra-study uncertainty of the total backscatter estimate of Proud et al. is ≈%.1f-fold' %proud_CI)
```
##### Interstudy uncertainty
As a measure of the interstudy uncertainty of the global backscatter, we calculate the 95% confidence interval of the geometric mean of the estimate from Irigoien et al. and Proud et al.:
```
# Calculate the interstudy uncertainty of the global backscatter
bs_inter_CI = geo_CI_calc([irigoien_backscatter,proud_backscatter])
print('The interstudy uncertainty of the total backscatter is ≈%.1f-fold' %bs_inter_CI)
# Take the highest uncertainty as our best projection of the uncertainty associates with the global backscatter
bs_CI = np.max([irigoien_CI,proud_CI,bs_inter_CI])
```
We use the highest uncertainty among these different kinds of uncertainty measures as our best projection of the uncertainty of the global backscatter, which is ≈1.7-fold.
#### Target strength per unit biomass
To assess the uncertainty associated with the target strength per unit biomass, we calculate the uncertainty in estimating the characteristic target strength per unit biomass of fish with or without swimbladders, adn the uncertainty associated with the fraction of the population that either has or lacks swimbladder
##### Uncertainty of characteristic target strength per unit biomass of fish with or without swimbladder
We calculate the 95% confidence interval of the target strength of fish with or withour swimbladder, and propagate this confidence interval to the total estimate of biomass to assess the uncertainty associated with the estimate of the target strength. We calculated an uncertainty of ≈1.3-fold. associated with te estimate of the target strength per unit biomass of fish.
```
# Define the function that will estimate the 95% confidence interval
def CI_groupby(input):
return input['dB kg^-1'].std(ddof=1)/np.sqrt(input['dB kg^-1'].shape[0])
# Group target strength values by the presence of absence of swimbladder
ts_bin = ts.groupby('Swimbladder')
# Calculate sandard error of those values
ts_bin_CI = ts_bin.apply(CI_groupby)
ts_CI = []
# For the target strength of fish with or without swimbladder, sample 1000 times from the distribution
# of target strengths, and calculate the estimate of the total biomass of fish. Then calcualte the 95%
# confidence interval of the resulting distribution as a measure of the uncertainty in the biomass
# estimate resulting from the uncertainty in the target strength
for x, instance in enumerate(ts_bin_CI):
ts_dist = np.random.normal(TS_bin['dB kg^-1'][x],instance,1000)
biomass_dist = biomass_estimator(ts_dist,TS_bin['dB kg^-1'][1-x],best_backscatter,frac=0.5)*1000*0.15
upper_CI = np.percentile(biomass_dist,97.5)/np.mean(biomass_dist)
lower_CI = np.mean(biomass_dist)/np.percentile(biomass_dist,2.5)
ts_CI.append(np.mean([upper_CI,lower_CI]))
# Take the maximum uncertainty of the with or with out swimbladder as our best projection
ts_CI = np.max(ts_CI)
print('Our best projection for the uncertainty associated with the estimate of the target strength per unit biomass is ≈%.1f-fold' %ts_CI)
```
##### Uncertainty of the fraction of the population possessing swimbladder
As a measure of the uncertainty associated with the assumption that fish with or without swimbladder contributed similar portions to the total population of mesopelagic fish, we sample different ratios of fish with and without swimbladder, and calculate the biomass estimate for those fractions.
```
# Sample different fractions of fish with swimbladder
ratio_range = np.linspace(0,1,1000)
# Estiamte the biomass of mesopelagic fish using the sampled fraction
biomass_ratio_dist = biomass_estimator(*TS_bin['dB kg^-1'],best_backscatter,ratio_range)*1000*0.15/1e15
# Plot the results for all fractions
plt.plot(ratio_range,biomass_ratio_dist)
plt.xlabel('Fraction of the population possessing swimbladder')
plt.ylabel('Biomass estimate [Gt C]')
```
We take the 95% range of distribution of fraction of fish with swimbladder account and calculate the uncertainty this fraction introduces into the sonar-based estimate of mesopelagic fish biomass. In this range the confidence interval of the biomass estimate is ≈8.7-fold.
```
# Calculate the upper and lower bounds of the influence of the fraction of fish with swimbladder on biomass estimate
ratio_upper_CI = (biomass_estimator(*TS_bin['dB kg^-1'],best_backscatter,0.975)*1000*0.15)/sonar_biomass
ratio_lower_CI = sonar_biomass/(biomass_estimator(*TS_bin['dB kg^-1'],best_backscatter,0)*1000*0.15)
ratio_CI = np.max([ratio_upper_CI,ratio_lower_CI])
print('Our best projection for the uncertainty associated with the fraction of fish possessing swimbladder is ≈%.1f-fold' %ratio_CI)
```
To calculate the total uncertainty associated with the sonar-based estimate, we propagate the uncertainties associated with the total backscatter, the target strength per unit biomass and the fraction of fish with swimbladder.
```
sonar_CI = CI_prod_prop(np.array([ratio_CI,ts_CI,bs_CI]))
print('Our best projection for the uncertainty associated with the sonar-based estimate for the biomass of mesopelagic fish is ≈%.1f-fold' %sonar_CI)
```
### Inter-method uncertainty
As a measure of the inter-method uncertainty of our estimate of the biomass of mesopelagic fish, we calculate the 95% confidence interval of the geometric mean of the sonar-based estimate and the trawling-based estimate.
```
meso_inter_CI = geo_CI_calc(np.array([sonar_biomass,trawling_biomass]))
print('Our best projection for the inter method uncertainty associated with estimate of the biomass of mesopelagic fish is ≈%.1f-fold' %meso_inter_CI)
# Take the highest uncertainty as our best projection for the uncertainty associated with the estimate
# of the biomass of mesopelagic fish
meso_CI = np.max([meso_inter_CI,sonar_CI])
```
Comparing our projections for the uncertainty of the sonar-based estimate of the biomass of mesopelagic fish and the inter-method uncertainty, our best projection for the biomass of mesopelagic fish is about one order of magnitude.
## Non-mesopelagic fish biomass uncertainty
For estimating the biomass of non-mesopelagic fish, we rely on estimates by Wilson et al., which does not report an uncertainty range for the biomass of non-meso pelagic fish. A later study ([Jennings et al.](https://doi.org/10.1371/journal.pone.0133794), gave an estimate for the total biomass of fish with body weight of 1 g to 1000 kg, based on ecological models. Jenning et al. reports a 90% confidence interval of 0.34-26.12 Gt wet weight, with a median estimate of ≈5 Gt wet weight. We take this range as a crude measure of the uncertainty associated with the estimate of the biomass of non-mesopelagic fish.
```
# Calculate the uncertainty of the non-mesopelagic fish biomass
non_meso_CI = np.max([26.12/5,5/0.34])
# Propagate the uncertainties of mesopelagic fish biomass and non-mesopelagic fish biomass to the total estimate
# of fish biomass
mul_CI = CI_sum_prop(estimates=np.array([best_mesopelagic_biomass,non_mesopelagic_fish_biomass]), mul_CIs=np.array([meso_CI,non_meso_CI]))
print('Our best projection for the uncertainty associated with the estimate of the biomass of fish is ≈%.1f-fold' %mul_CI)
```
# Prehuman fish biomass
To estimate the prehuman fish biomass, we rely on a study ([Costello et al.](http://dx.doi.org/10.1073/pnas.1520420113)) which states that fish stocks in global fisheries are 1.17 of the Maximal Sustainable Yield biomass, when looking at all fisheries and calculating a catch-weighted average global fishery (Figure S12 in the SI Appendix of Costello et al.). Costello et al. also reports the total biomass of present day fisheries at 0.84 Gt wet weight (Table S15 in the SI Appendix of Costello et al.). Assuming 70% water content and 50% carbon content out of wet weight, this translates to:
```
costello_ww = 0.84
wet_to_c = 0.3*0.5
costello_cc = costello_ww*wet_to_c
print('Costello et al. estimate ≈%.2f Gt C of current fisheries' %costello_cc)
```
This number is close to the number reported by Wilson et al. Using a database of published landings data and stock assessment biomass estimates, [Thorson et al.](http://dx.doi.org/10.1139/f2012-077) estimate that the biomass of fish at the maximum sustainable yield represent ≈40% of the biomass the population would have reached in case of no fishing. From these two numbers, we can estimate the prehuamn biomass of fish in fisheries. We use the total biomass of fisheries reported in Costello et al., divide it the bte ratio reported in Costello et al. to estimate the Maximal Sustainable Yield biomass, and then divide this number by 0.4 to arrive at the prehuman biomass of fish in fisheries. We add to this estimate the estimate of the total biomass of mesopelagic fish, assuming their biomass wasn't affected by humans.
```
costello_ratio = 1.17
thorson_ratio = 0.4
prehuman_biomass_fisheries = costello_cc*1e15/costello_ratio/thorson_ratio
prehuman_fish_biomass = prehuman_biomass_fisheries+best_mesopelagic_biomass
print('Our estimate for the total prehuman biomass of fish is ≈%.1f Gt C' %(prehuman_fish_biomass/1e15))
```
Comparing the prehuman fish biomass to the present day fish biomass, we can estimate the human associated reduction in fish biomass:
```
fish_biomass_decrease = prehuman_fish_biomass-best_estimate
print('Our estimate for the decrease in the total biomass of fish is ≈%.2f Gt C' %(fish_biomass_decrease/1e15))
```
Which means that, based on the assumptions in our calculation, the decrease in the total biomass of fish is about the same as the remaining total mass of fish in all fisheries (disregarding mesopalegic fish).
# Estimating the total number of fish
To estimate the total number of fish, we divide our estimate of the total biomass of mesopelagic fish by an estimate for the characteristic carbon content of a single mesopelagic fish.
To estimate the mean weight of mesopelagic fish, we rely on data reported in [Fock & Ehrich](https://doi.org/10.1111/j.1439-0426.2010.01450.x) for the family Myctophidae (Lanternfish), which dominate the mesopelagic fish species. Fock & Ehrich report the length range of each fish species, as well as allometric relations between fish length and weight for each species. Here is a sample of the data:
```
# Load the data from Fock & Ehrich
fe_data = pd.read_excel('fish_biomass_data.xlsx','Fock & Ehrich', skiprows=1)
# Use only data for the Myctophidae family
fe_mycto = fe_data[fe_data['Family'] == 'Myctophidae']
fe_mycto.head()
```
The allometric parameters a and b are plugged into the following equation to produce the weight of each fish species based on the length of each fish: $$ W = a*L^b$$
Where W is the fish weight and L is the fish length. For each fish species, we calculate the characteristic fish length by using the mean of the minimum and maximum reported fish lengths:
```
fe_mean_length = np.mean([fe_mycto['Maximum length (mm)'].astype(float),fe_mycto['Minimum length (mm)'].astype(float)])
```
We plug the mean length of each fish species into the allometric equation along with its specific parameters a and b to generate the mean wet weight of each fish. We use the geometric mean of the weights of all species as our best estimate of the weight of a single mesopelagic fish. We convert wet weight to carbon mass assuming 70% water content and 50% carbon our of the dry weight.
```
# The allometric equation to convert fish length into fish weight. The equation takes values
# in cm and the data is given in mm so we divide the length by a factor of 10
calculate_weight = lambda x,a,b: a*(x/10)**b
# Transform the mean lengths of each fish species into a characteristic weight of each fish species
fe_mean_weight = calculate_weight(fe_mean_length,fe_mycto['a(SL)'],fe_mycto['b(SL)'])
# Conversion factor from wet weight to carbon mass
wet_to_c = 0.15
# Calculate the mean carbon content of a single mesopelagic fish
fish_cc = gmean(fe_mean_weight.astype(float))*wet_to_c
print('Our best estimate for the carbon content of a single mesopelagic fish is ≈%.2f g C' %fish_cc)
```
We estimate the total number of mesopelagic fish by dividing our best estimate for the total biomass of mesopelagic fish by our estimate for the carbon content of a single mesopelagic fish:
```
# Estimate the total number of fish
tot_fish_num = best_mesopelagic_biomass/fish_cc
print('Our best estimate for the total number of individual fish is ≈%.0e.' %tot_fish_num)
# Feed results to the chordate biomass data
old_results = pd.read_excel('../../animal_biomass_estimate.xlsx',index_col=0)
result = old_results.copy()
result.loc['Fish',(['Biomass [Gt C]','Uncertainty'])] = (best_estimate/1e15,mul_CI)
result.to_excel('../../animal_biomass_estimate.xlsx')
# Feed results to Table 1 & Fig. 1
update_results(sheet='Table1 & Fig1',
row=('Animals','Fish'),
col=['Biomass [Gt C]', 'Uncertainty'],
values=[best_estimate/1e15,mul_CI],
path='../../../results.xlsx')
# Feed results to Table S1
update_results(sheet='Table S1',
row=('Animals','Fish'),
col=['Number of individuals'],
values=tot_fish_num,
path='../../../results.xlsx')
# Update the data mentioned in the MS
update_MS_data(row ='Decrease in biomass of fish',
values=fish_biomass_decrease/1e15,
path='../../../results.xlsx')
```
| github_jupyter |
# Implementing the Gradient Descent Algorithm
##### _Written by Owais Gondal_
In this lab, we'll implement the basic functions of the Gradient Descent algorithm to find the boundary in a small dataset. First, we'll start with some functions that will help us plot and visualize the data.
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
#Some helper functions for plotting and drawing lines
def plot_points(X, y):
admitted = X[np.argwhere(y==1)]
rejected = X[np.argwhere(y==0)]
plt.scatter([s[0][0] for s in rejected], [s[0][1] for s in rejected], s = 25, color = 'blue', edgecolor = 'k')
plt.scatter([s[0][0] for s in admitted], [s[0][1] for s in admitted], s = 25, color = 'red', edgecolor = 'k')
def display(m, b, color='g--'):
plt.xlim(-0.05,1.05)
plt.ylim(-0.05,1.05)
x = np.arange(-10, 10, 0.1)
plt.plot(x, m*x+b, color)
```
## Reading and plotting the data
```
# data = pd.read_csv('data.csv', header=None)
data=[]
for i in range(100):
new
print(data.shape())
# X = np.array(data[[0, 1]])
# y = np.array(data[2])
# plot_points(X, y)
# plt.show()
# print(data)
```
## TODO: Implementing the basic functions
Here is your turn to shine. Implement the following formulas, as explained in the text.
- Sigmoid activation function
$$\sigma(x) = \frac{1}{1+e^{-x}}$$
- Output (prediction) formula
$$\hat{y} = \sigma(w_1 x_1 + w_2 x_2 + b)$$
- Error function
$$Error(y, \hat{y}) = - y \log(\hat{y}) - (1-y) \log(1-\hat{y})$$
- The function that updates the weights
$$ w_i \longrightarrow w_i + \alpha (y - \hat{y}) x_i$$
$$ b \longrightarrow b + \alpha (y - \hat{y})$$
```
# Implement the following functions
# Activation (sigmoid) function
def sigmoid(x):
# print(x)
return 1/(1+np.exp(-x))
# Output (prediction) formula
def output_formula(features, weights, bias):
return sigmoid(np.dot(features, weights)+bias)
# Error (log-loss) formula
def error_formula(y, output):
return -y*np.log(output)-(1-y)*np.log(1-output)
# Gradient descent step
def update_weights(x, y, weights, bias, learnrate):
# learnrate=learnrate*1/len(x)
for i in range(0, len(x)):
weights[i] = weights[i] + learnrate * \
(y-output_formula(x, weights, bias))*x[i]
bias = bias+learnrate*(y-output_formula(x, weights, bias))
return weights, bias
```
## Training function
This function will help us iterate the gradient descent algorithm through all the data, for a number of epochs. It will also plot the data, and some of the boundary lines obtained as we run the algorithm.
```
np.random.seed(44)
epochs = 100
learnrate = 0.01
def train(features, targets, epochs, learnrate, graph_lines=False):
errors = []
n_records, n_features = features.shape
last_loss = None
weights = np.random.normal(scale=1 / n_features**.5, size=n_features)
bias = 0
for e in range(epochs):
del_w = np.zeros(weights.shape)
for x, y in zip(features, targets):
output = output_formula(x, weights, bias)
error = error_formula(y, output)
weights, bias = update_weights(x, y, weights, bias, learnrate)
# Printing out the log-loss error on the training set
out = output_formula(features, weights, bias)
loss = np.mean(error_formula(targets, out))
errors.append(loss)
if e % (epochs / 10) == 0:
print("\n========== Epoch", e, "==========")
if last_loss and last_loss < loss:
print("Train loss: ", loss, " WARNING - Loss Increasing")
else:
print("Train loss: ", loss)
last_loss = loss
predictions = out > 0.5
accuracy = np.mean(predictions == targets)
print("Accuracy: ", accuracy)
if graph_lines and e % (epochs / 100) == 0:
display(-weights[0]/weights[1], -bias/weights[1])
# Plotting the solution boundary
plt.title("Solution boundary")
display(-weights[0]/weights[1], -bias/weights[1], 'black')
# Plotting the data
plot_points(features, targets)
plt.show()
# Plotting the error
plt.title("Error Plot")
plt.xlabel('Number of epochs')
plt.ylabel('Error')
plt.plot(errors)
plt.show()
```
## Time to train the algorithm!
When we run the function, we'll obtain the following:
- 10 updates with the current training loss and accuracy
- A plot of the data and some of the boundary lines obtained. The final one is in black. Notice how the lines get closer and closer to the best fit, as we go through more epochs.
- A plot of the error function. Notice how it decreases as we go through more epochs.
```
train(X, y, epochs, learnrate, True)
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/official/feature_store/gapic-feature-store.ipynb"">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/blob/master/notebooks/official/feature_store/gapic-feature-store.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
</table>
## Overview
This Colab introduces Feature Store, a managed cloud service for machine learning engineers and data scientists to store, serve, manage and share machine learning features at a large scale.
This Colab assumes that you understand basic Google Cloud concepts such as [Project](https://cloud.google.com/storage/docs/projects), [Storage](https://cloud.google.com/storage) and [Vertex AI](https://cloud.google.com/vertex-ai/docs). Some machine learning knowledge is also helpful but not required.
### Dataset
This Colab uses a movie recommendation dataset as an example throughout all the sessions. The task is to train a model to predict if a user is going to watch a movie and serve this model online.
### Objective
In this notebook, you will learn how to:
* How to import your features into Feature Store.
* How to serve online prediction requests using the imported features.
* How to access imported features in offline jobs, such as training jobs.
### Costs
This tutorial uses billable components of Google Cloud:
* Vertex AI
* Cloud Storage
* Cloud Bigtable
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
### Set up your local development environment
**If you are using Colab or Google Cloud Notebooks**, your environment already meets
all the requirements to run this notebook. You can skip this step.
**Otherwise**, make sure your environment meets this notebook's requirements.
You need the following:
* The Google Cloud SDK
* Git
* Python 3
* virtualenv
* Jupyter notebook running in a virtual environment with Python 3
The Google Cloud guide to [Setting up a Python development
environment](https://cloud.google.com/python/setup) and the [Jupyter
installation guide](https://jupyter.org/install) provide detailed instructions
for meeting these requirements. The following steps provide a condensed set of
instructions:
1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
1. [Install
virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
and create a virtual environment that uses Python 3. Activate the virtual environment.
1. To install Jupyter, run `pip3 install jupyter` on the
command-line in a terminal shell.
1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
1. Open this notebook in the Jupyter Notebook Dashboard.
### Install additional packages
For this Colab, you need the Vertex SDK for Python.
```
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
! pip3 install {USER_FLAG} --upgrade git+https://github.com/googleapis/python-aiplatform.git@main-test
```
### Restart the kernel
After you install the SDK, you need to restart the notebook kernel so it can find the packages. You can restart kernel from *Kernel -> Restart Kernel*, or running the following:
```
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
## Before you begin
### Select a GPU runtime
**Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select "Runtime --> Change runtime type > GPU"**
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
1. [Enable the Vertex AI API and Compute Engine API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com,compute_component).
1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
1. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
#### Set your project ID
**If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
```
import os
PROJECT_ID = ""
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
shell_output=!gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
```
Otherwise, set your project ID here.
```
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "python-docs-samples-tests" # @param {type:"string"}
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebooks**, your environment is already
authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions
when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
1. In the Cloud Console, go to the [**Create service account key**
page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
2. Click **Create service account**.
3. In the **Service account name** field, enter a name, and
click **Create**.
4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
into the filter box, and select
**Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
5. Click *Create*. A JSON file that contains your key downloads to your
local environment.
6. Enter the path to your service account key as the
`GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
```
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# If on Google Cloud Notebooks, then don't execute this code
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
## Prepare for output
### Step 1. Create dataset for output
You need a BigQuery dataset to host the output data in `us-central1`. Input the name of the dataset you want to created and specify the name of the table you want to store the output later. These will be used later in the notebook.
**Make sure that the table name does NOT already exist**.
```
from datetime import datetime
from google.cloud import bigquery
# Output dataset
DESTINATION_DATA_SET = "movie_predictions" # @param {type:"string"}
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
DESTINATION_DATA_SET = "{prefix}_{timestamp}".format(
prefix=DESTINATION_DATA_SET, timestamp=TIMESTAMP
)
# Output table. Make sure that the table does NOT already exist; the BatchReadFeatureValues API cannot overwrite an existing table
DESTINATION_TABLE_NAME = "training_data" # @param {type:"string"}
DESTINATION_PATTERN = "bq://{project}.{dataset}.{table}"
DESTINATION_TABLE_URI = DESTINATION_PATTERN.format(
project=PROJECT_ID, dataset=DESTINATION_DATA_SET, table=DESTINATION_TABLE_NAME
)
# Create dataset
REGION = "us-central1" # @param {type:"string"}
client = bigquery.Client()
dataset_id = "{}.{}".format(client.project, DESTINATION_DATA_SET)
dataset = bigquery.Dataset(dataset_id)
dataset.location = REGION
dataset = client.create_dataset(dataset, timeout=30)
print("Created dataset {}.{}".format(client.project, dataset.dataset_id))
```
### Import libraries and define constants
```
# Other than project ID and featurestore ID and endpoints needs to be set
API_ENDPOINT = "us-central1-aiplatform.googleapis.com" # @param {type:"string"}
INPUT_CSV_FILE = "gs://cloud-samples-data-us-central1/ai-platform-unified/datasets/featurestore/movie_prediction.csv"
from google.cloud.aiplatform_v1beta1 import (
FeaturestoreOnlineServingServiceClient, FeaturestoreServiceClient)
from google.cloud.aiplatform_v1beta1.types import FeatureSelector, IdMatcher
from google.cloud.aiplatform_v1beta1.types import \
entity_type as entity_type_pb2
from google.cloud.aiplatform_v1beta1.types import feature as feature_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore as featurestore_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore_monitoring as featurestore_monitoring_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore_online_service as featurestore_online_service_pb2
from google.cloud.aiplatform_v1beta1.types import \
featurestore_service as featurestore_service_pb2
from google.cloud.aiplatform_v1beta1.types import io as io_pb2
from google.protobuf.duration_pb2 import Duration
# Create admin_client for CRUD and data_client for reading feature values.
admin_client = FeaturestoreServiceClient(client_options={"api_endpoint": API_ENDPOINT})
data_client = FeaturestoreOnlineServingServiceClient(
client_options={"api_endpoint": API_ENDPOINT}
)
# Represents featurestore resource path.
BASE_RESOURCE_PATH = admin_client.common_location_path(PROJECT_ID, REGION)
```
## Terminology and Concept
### Featurestore Data model
Feature Store organizes data with the following 3 important hierarchical concepts:
```
Featurestore -> EntityType -> Feature
```
* **Featurestore**: the place to store your features
* **EntityType**: under a Featurestore, an *EntityType* describes an object to be modeled, real one or virtual one.
* **Feature**: under an EntityType, a *feature* describes an attribute of the EntityType
In the movie prediction example, you will create a featurestore called *movie_prediction*. This store has 2 entity types: *Users* and *Movies*. The Users entity type has the age, gender, and like genres features. The Movies entity type has the genres and average rating features.
## Create Featurestore and Define Schemas
### Create Featurestore
The method to create a featurestore returns a
[long-running operation](https://google.aip.dev/151) (LRO). An LRO starts an asynchronous job. LROs are returned for other API
methods too, such as updating or deleting a featurestore. Calling
`create_fs_lro.result()` waits for the LRO to complete.
```
FEATURESTORE_ID = "movie_prediction_{timestamp}".format(timestamp=TIMESTAMP)
create_lro = admin_client.create_featurestore(
featurestore_service_pb2.CreateFeaturestoreRequest(
parent=BASE_RESOURCE_PATH,
featurestore_id=FEATURESTORE_ID,
featurestore=featurestore_pb2.Featurestore(
display_name="Featurestore for movie prediction",
online_serving_config=featurestore_pb2.Featurestore.OnlineServingConfig(
fixed_node_count=3
),
),
)
)
# Wait for LRO to finish and get the LRO result.
print(create_lro.result())
```
You can use [GetFeaturestore](https://cloud.google.com/vertex-ai/docs/reference/rpc/google.cloud.aiplatform.v1beta1#google.cloud.aiplatform.v1beta1.FeaturestoreService.GetFeaturestore) or [ListFeaturestores](https://cloud.google.com/vertex-ai/docs/reference/rpc/google.cloud.aiplatform.v1beta1#google.cloud.aiplatform.v1beta1.FeaturestoreService.ListFeaturestores) to check if the Featurestore was successfully created. The following example gets the details of the Featurestore.
```
admin_client.get_featurestore(
name=admin_client.featurestore_path(PROJECT_ID, REGION, FEATURESTORE_ID)
)
```
### Create Entity Type
You can specify a monitoring config which will by default be inherited by all Features under this EntityType.
```
# Create users entity type with monitoring enabled.
# All Features belonging to this EntityType will by default inherit the monitoring config.
users_entity_type_lro = admin_client.create_entity_type(
featurestore_service_pb2.CreateEntityTypeRequest(
parent=admin_client.featurestore_path(PROJECT_ID, REGION, FEATURESTORE_ID),
entity_type_id="users",
entity_type=entity_type_pb2.EntityType(
description="Users entity",
monitoring_config=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig(
snapshot_analysis=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig.SnapshotAnalysis(
monitoring_interval=Duration(seconds=86400), # 1 day
),
),
),
)
)
# Similarly, wait for EntityType creation operation.
print(users_entity_type_lro.result())
# Create movies entity type without a monitoring configuration.
movies_entity_type_lro = admin_client.create_entity_type(
featurestore_service_pb2.CreateEntityTypeRequest(
parent=admin_client.featurestore_path(PROJECT_ID, REGION, FEATURESTORE_ID),
entity_type_id="movies",
entity_type=entity_type_pb2.EntityType(description="Movies entity"),
)
)
# Similarly, wait for EntityType creation operation.
print(movies_entity_type_lro.result())
```
### Create Feature
You can also set a custom monitoring configuration at the Feature level, and view the properties and metrics in the console: sample [properties](https://storage.googleapis.com/cloud-samples-data/ai-platform-unified/datasets/featurestore/Feature%20Properties.png), sample [metrics](https://storage.googleapis.com/cloud-samples-data/ai-platform-unified/datasets/featurestore/Feature%20Snapshot%20Distribution.png).
```
# Create features for the 'users' entity.
# 'age' Feature leaves the monitoring config unset, which means it'll inherit the config from EntityType.
# 'gender' Feature explicitly disables monitoring.
# 'liked_genres' Feature is a STRING_ARRAY type, so it is automatically excluded from monitoring.
# For Features with monitoring enabled, distribution statistics are updated periodically in the console.
admin_client.batch_create_features(
parent=admin_client.entity_type_path(PROJECT_ID, REGION, FEATURESTORE_ID, "users"),
requests=[
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.INT64,
description="User age",
),
feature_id="age",
),
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.STRING,
description="User gender",
monitoring_config=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig(
snapshot_analysis=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig.SnapshotAnalysis(
disabled=True,
),
),
),
feature_id="gender",
),
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.STRING_ARRAY,
description="An array of genres that this user liked",
),
feature_id="liked_genres",
),
],
).result()
# Create features for movies type.
# 'title' Feature enables monitoring.
admin_client.batch_create_features(
parent=admin_client.entity_type_path(PROJECT_ID, REGION, FEATURESTORE_ID, "movies"),
requests=[
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.STRING,
description="The title of the movie",
monitoring_config=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig(
snapshot_analysis=featurestore_monitoring_pb2.FeaturestoreMonitoringConfig.SnapshotAnalysis(
monitoring_interval=Duration(seconds=172800), # 2 days
),
),
),
feature_id="title",
),
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.STRING,
description="The genres of the movie",
),
feature_id="genres",
),
featurestore_service_pb2.CreateFeatureRequest(
feature=feature_pb2.Feature(
value_type=feature_pb2.Feature.ValueType.DOUBLE,
description="The average rating for the movie, range is [1.0-5.0]",
),
feature_id="average_rating",
),
],
).result()
```
## Search created features
While the [ListFeatures](https://cloud.google.com/vertex-ai/docs/reference/rpc/google.cloud.aiplatform.v1beta1#google.cloud.aiplatform.v1beta1.FeaturestoreService.ListFeatures) method allows you to easily view all features of a single
entity type, the [SearchFeatures](https://cloud.google.com/vertex-ai/docs/reference/rpc/google.cloud.aiplatform.v1beta1#google.cloud.aiplatform.v1beta1.FeaturestoreService.SearchFeatures) method searches across all featurestores
and entity types in a given location (such as `us-central1`). This can help you discover features that were created by someone else.
You can query based on feature properties including feature ID, entity type ID,
and feature description. You can also limit results by filtering on a specific
featurestore, feature value type, and/or labels.
```
# Search for all features across all featurestores.
list(admin_client.search_features(location=BASE_RESOURCE_PATH))
```
Now, narrow down the search to features that are of type `DOUBLE`
```
# Search for all features with value type `DOUBLE`
list(
admin_client.search_features(
featurestore_service_pb2.SearchFeaturesRequest(
location=BASE_RESOURCE_PATH, query="value_type=DOUBLE"
)
)
)
```
Or, limit the search results to features with specific keywords in their ID and type.
```
# Filter on feature value type and keywords.
list(
admin_client.search_features(
featurestore_service_pb2.SearchFeaturesRequest(
location=BASE_RESOURCE_PATH, query="feature_id:title AND value_type=STRING"
)
)
)
```
## Import Feature Values
You need to import feature values before you can use them for online/offline serving. In this step, you will learn how to import feature values by calling the ImportFeatureValues API using the Python SDK.
### Source Data Format and Layout
As mentioned above, BigQuery table/Avro/CSV are supported. No matter what format you are using, each imported entity *must* have an ID; also, each entity can *optionally* have a timestamp, sepecifying when the feature values are generated. This Colab uses Avro as an input, located at this public [bucket](https://pantheon.corp.google.com/storage/browser/cloud-samples-data/ai-platform-unified/datasets/featurestore;tab=objects?project=storage-samples&prefix=&forceOnObjectsSortingFiltering=false). The Avro schemas are as follows:
**For the Users entity**:
```
schema = {
"type": "record",
"name": "User",
"fields": [
{
"name":"user_id",
"type":["null","string"]
},
{
"name":"age",
"type":["null","long"]
},
{
"name":"gender",
"type":["null","string"]
},
{
"name":"liked_genres",
"type":{"type":"array","items":"string"}
},
{
"name":"update_time",
"type":["null",{"type":"long","logicalType":"timestamp-micros"}]
},
]
}
```
**For the Movies entity**
```
schema = {
"type": "record",
"name": "Movie",
"fields": [
{
"name":"movie_id",
"type":["null","string"]
},
{
"name":"average_rating",
"type":["null","double"]
},
{
"name":"title",
"type":["null","string"]
},
{
"name":"genres",
"type":["null","string"]
},
{
"name":"update_time",
"type":["null",{"type":"long","logicalType":"timestamp-micros"}]
},
]
}
```
### Import feature values for Users
When importing, specify the following in your request:
* Data source format: BigQuery Table/Avro/CSV
* Data source URL
* Destination: featurestore/entity types/features to be imported
```
import_users_request = featurestore_service_pb2.ImportFeatureValuesRequest(
entity_type=admin_client.entity_type_path(
PROJECT_ID, REGION, FEATURESTORE_ID, "users"
),
avro_source=io_pb2.AvroSource(
# Source
gcs_source=io_pb2.GcsSource(
uris=[
"gs://cloud-samples-data-us-central1/ai-platform-unified/datasets/featurestore/users.avro"
]
)
),
entity_id_field="user_id",
feature_specs=[
# Features
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(id="age"),
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(id="gender"),
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(
id="liked_genres"
),
],
feature_time_field="update_time",
worker_count=10,
)
# Start to import, will take a couple of minutes
ingestion_lro = admin_client.import_feature_values(import_users_request)
# Polls for the LRO status and prints when the LRO has completed
ingestion_lro.result()
```
### Import feature values for Movies
Similarly, import feature values for 'movies' into the featurestore.
```
import_movie_request = featurestore_service_pb2.ImportFeatureValuesRequest(
entity_type=admin_client.entity_type_path(
PROJECT_ID, REGION, FEATURESTORE_ID, "movies"
),
avro_source=io_pb2.AvroSource(
gcs_source=io_pb2.GcsSource(
uris=[
"gs://cloud-samples-data-us-central1/ai-platform-unified/datasets/featurestore/movies.avro"
]
)
),
entity_id_field="movie_id",
feature_specs=[
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(id="title"),
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(id="genres"),
featurestore_service_pb2.ImportFeatureValuesRequest.FeatureSpec(
id="average_rating"
),
],
feature_time_field="update_time",
worker_count=10,
)
# Start to import, will take a couple of minutes
ingestion_lro = admin_client.import_feature_values(import_movie_request)
# Polls for the LRO status and prints when the LRO has completed
ingestion_lro.result()
```
## Online serving
The
[Online Serving APIs](https://cloud.google.com/vertex-ai/docs/reference/rpc/google.cloud.aiplatform.v1beta1#featurestoreonlineservingservice)
lets you serve feature values for small batches of entities. It's designed for latency-sensitive service, such as online model prediction. For example, for a movie service, you might want to quickly shows movies that the current user would most likely watch by using online predictions.
### Read one entity per request
The ReadFeatureValues API is used to read feature values of one entity; hence
its custom HTTP verb is `readFeatureValues`. By default, the API will return the latest value of each feature, meaning the feature values with the most recent timestamp.
To read feature values, specify the entity ID and features to read. The response
contains a `header` and an `entity_view`. Each row of data in the `entity_view`
contains one feature value, in the same order of features as listed in the response header.
```
# Fetch the following 3 features.
feature_selector = FeatureSelector(
id_matcher=IdMatcher(ids=["age", "gender", "liked_genres"])
)
data_client.read_feature_values(
featurestore_online_service_pb2.ReadFeatureValuesRequest(
# Fetch from the following feature store/entity type
entity_type=admin_client.entity_type_path(
PROJECT_ID, REGION, FEATURESTORE_ID, "users"
),
# Fetch the user features whose ID is "alice"
entity_id="alice",
feature_selector=feature_selector,
)
)
```
### Read multiple entities per request
To read feature values from multiple entities, use the
StreamingReadFeatureValues API, which is almost identical to the previous
ReadFeatureValues API. Note that fetching only a small number of entities is recomended when using this API due to its latency-sensitive nature.
```
# Read the same set of features as above, but for multiple entities.
response_stream = data_client.streaming_read_feature_values(
featurestore_online_service_pb2.StreamingReadFeatureValuesRequest(
entity_type=admin_client.entity_type_path(
PROJECT_ID, REGION, FEATURESTORE_ID, "users"
),
entity_ids=["alice", "bob"],
feature_selector=feature_selector,
)
)
# Iterate and process response. Note the first one is always the header only.
for response in response_stream:
print(response)
```
Now that you have learned how to featch imported feature values for online serving, the next step is learning how to use imported feature values for offline use cases.
## Batch Serving
Batch Serving is used to fetch a large batch of feature values for high-throughput, typically for training a model or batch prediction. In this section, you will learn how to prepare for training examples by calling the BatchReadFeatureValues API.
### Use case
**The task** is to prepare a training dataset to train a model, which predicts if a given user will watch a given movie. To achieve this, you need 2 sets of input:
* Features: you already imported into the featurestore.
* Labels: the groud-truth data recorded that user X has watched movie Y.
To be more specific, the ground-truth observation is described in Table 1 and the desired training dataset is described in Table 2. Each row in Table 2 is a result of joining the imported feature values from Feature Store according to the entity IDs and timestamps in Table 1. In this example, the `age`, `gender` and `liked_genres` features from `users` and
the `genres` and `average_rating` features from `movies` are chosen to train the model. Note that only positive examples are shown in these 2 tables, i.e., you can imagine there is a label column whose values are all `True`.
BatchReadFeatureValues API takes Table 1 as
input, joins all required feature values from the featurestore, and returns Table 2 for training.
<h4 align="center">Table 1. Ground-truth Data</h4>
users | movies | timestamp
----- | -------- | --------------------
alice | Cinema Paradiso | 2019-11-01T00:00:00Z
bob | The Shining | 2019-11-15T18:09:43Z
... | ... | ...
<h4 align="center">Table 2. Expected Training Data Generated by Batch Read API (Positive Samples)</h4>
timestamp | entity_type_users | age | gender | liked_genres | entity_type_movies | genres | average_rating
-------------------- | ----------------- | --------------- | ---------------- | -------------------- | -------- | --------- | -----
2019-11-01T00:00:00Z | bob | 35 | M | [Action, Crime] | The Shining | Horror | 4.8
2019-11-01T00:00:00Z | alice | 55 | F | [Drama, Comedy] | Cinema Paradiso | Romance | 4.5
... | ... | ... | ... | ... | ... | ... | ...
#### Why timestamp?
Note that there is a `timestamp` column in Table 2. This indicates the time when the ground-truth was observed. This is to avoid data inconsistency.
For example, the 1st row of Table 2 indicates that user `alice` watched movie `Cinema Paradiso` on `2019-11-01T00:00:00Z`. The featurestore keeps feature values for all timestamps but fetches feature values *only* at the given timestamp during batch serving. On 2019-11-01 alice might be 54 years old, but now alice might be 56; featurestore returns `age=54` as alice's age, instead of `age=56`, because that is the value of the feature at the observation time. Similarly, other features might be time-variant as well, such as liked_genres.
### Batch Read Feature Values
Assemble the request which specify the following info:
* Where is the label data, i.e., Table 1.
* Which features are read, i.e., the column names in Table 2.
The output is stored in a BigQuery table.
```
batch_serving_request = featurestore_service_pb2.BatchReadFeatureValuesRequest(
# featurestore info
featurestore=admin_client.featurestore_path(PROJECT_ID, REGION, FEATURESTORE_ID),
# URL for the label data, i.e., Table 1.
csv_read_instances=io_pb2.CsvSource(
gcs_source=io_pb2.GcsSource(uris=[INPUT_CSV_FILE])
),
destination=featurestore_service_pb2.FeatureValueDestination(
bigquery_destination=io_pb2.BigQueryDestination(
# Output to BigQuery table created earlier
output_uri=DESTINATION_TABLE_URI
)
),
entity_type_specs=[
featurestore_service_pb2.BatchReadFeatureValuesRequest.EntityTypeSpec(
# Read the 'age', 'gender' and 'liked_genres' features from the 'users' entity
entity_type_id="users",
feature_selector=FeatureSelector(
id_matcher=IdMatcher(
ids=[
# features, use "*" if you want to select all features within this entity type
"age",
"gender",
"liked_genres",
]
)
),
),
featurestore_service_pb2.BatchReadFeatureValuesRequest.EntityTypeSpec(
# Read the 'average_rating' and 'genres' feature values of the 'movies' entity
entity_type_id="movies",
feature_selector=FeatureSelector(
id_matcher=IdMatcher(ids=["average_rating", "genres"])
),
),
],
)
# Execute the batch read
batch_serving_lro = admin_client.batch_read_feature_values(batch_serving_request)
# This long runing operation will poll until the batch read finishes.
batch_serving_lro.result()
```
After the LRO finishes, you should be able to see the result from the [BigQuery console](https://console.cloud.google.com/bigquery), in the dataset created earlier.
## Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
You can also keep the project but delete the featurestore:
```
admin_client.delete_featurestore(
request=featurestore_service_pb2.DeleteFeaturestoreRequest(
name=admin_client.featurestore_path(PROJECT_ID, REGION, FEATURESTORE_ID),
force=True,
)
).result()
client.delete_dataset(
DESTINATION_DATA_SET, delete_contents=True, not_found_ok=True
) # Make an API request.
print("Deleted dataset '{}'.".format(DESTINATION_DATA_SET))
```
| github_jupyter |
##### Copyright 2020 The Cirq Developers
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Simulation
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.example.org/cirq/simulation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on QuantumLib</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/quantumlib/Cirq/blob/master/docs/simulation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/quantumlib/Cirq/blob/master/docs/simulation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/Cirq/docs/simulation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
```
try:
import cirq
except ImportError:
print("installing cirq...")
!pip install --quiet cirq
print("installed cirq.")
```
Cirq comes with built-in Python simulators for testing
small circuits. The two main types of simulations that Cirq
supports are pure state and mixed state. The pure state simulations
are supported by ``cirq.Simulator`` and the mixed state
simulators are supported by ``cirq.DensityMatrixSimulator``.
The names *pure state simulator* and *mixed state
simulators* refer to the fact that these simulations are
for quantum circuits; including unitary, measurements, and noise
that keeps the evolution in a pure state or a mixed state.
In other words, there are some noisy evolutions
that are supported by the pure state simulator as long as they
preserve the purity of the state.
Some external high-performance simulators also provide an interface
to Cirq. These can sometimes provide results faster than Cirq's
built-in simulators, especially when working with larger circuits.
For details on these tools, see the
[external simulators section](#external-simulators).
## Introduction to pure state simulation
Here is a simple circuit:
```
!pip install cirq --quiet
import cirq
q0 = cirq.GridQubit(0, 0)
q1 = cirq.GridQubit(1, 0)
def basic_circuit(meas=True):
sqrt_x = cirq.X**0.5
yield sqrt_x(q0), sqrt_x(q1)
yield cirq.CZ(q0, q1)
yield sqrt_x(q0), sqrt_x(q1)
if meas:
yield cirq.measure(q0, key='q0'), cirq.measure(q1, key='q1')
circuit = cirq.Circuit()
circuit.append(basic_circuit())
print(circuit)
```
We can simulate this by creating a ``cirq.Simulator`` and
passing the circuit into its ``run`` method:
```
from cirq import Simulator
simulator = Simulator()
result = simulator.run(circuit)
print(result)
```
The method `run()` returns an ``Result``. As you can see, the object `result` contains the result of any measurements for the simulation run.
The actual measurement results depend on the seeding from `random` seed generator (`numpy`) . You can set this using ``numpy.random_seed``.
Another run, can result in a different set of measurement results:
```
result = simulator.run(circuit)
print(result)
```
The simulator is designed to mimic what running a program
on a quantum computer is actually like.
In particular, the ``run()`` methods (``run()`` and ``run_sweep()``) on the simulator do not give access to the wave function of the quantum computer (since one doesn't have access to this on the actual quantum
hardware). Instead, the ``simulate()`` methods (``simulate()``,
``simulate_sweep()``, ``simulate_moment_steps()``) should be used
if one wants to debug the circuit and get access to the full
wave function:
```
import numpy as np
circuit = cirq.Circuit()
circuit.append(basic_circuit(False))
result = simulator.simulate(circuit, qubit_order=[q0, q1])
print(np.around(result.final_state_vector, 3))
```
Note that the simulator uses numpy's ``float32`` precision
(which is ``complex64`` for complex numbers) by default,
but that the simulator can take in a `dtype` of `np.complex128`
if higher precision is needed.
## Qubit and Amplitude Ordering
The `qubit_order` argument to the simulator's `run()` method
determines the ordering of some results, such as the
amplitudes in the final wave function. The `qubit_order` argument is optional: when it is omitted, qubits are ordered
ascending by their name (i.e., what their `__str__` method returns).
The simplest `qubit_order` value you can provide is a list of
the qubits in the desired ordered. Any qubits from the circuit
that are not in the list will be ordered using the
default `__str__` ordering, but come after qubits that are in
the list. Be aware that all qubits in the list are included in
the simulation, even if they are not operated on by the circuit.
The mapping from the order of the qubits to the order of the
amplitudes in the wave function can be tricky to understand.
Basically, it is the same as the ordering used by `numpy.kron`:
```
outside = [1, 10]
inside = [1, 2]
print(np.kron(outside, inside))
```
More concretely, `k`'th amplitude in the wave function
will correspond to the `k`'th case that would be encountered
when nesting loops over the possible values of each qubit.
The first qubit's computational basis values are looped over
in the outermost loop, the last qubit's computational basis
values are looped over in the inner-most loop, etc.:
```
i = 0
for first in [0, 1]:
for second in [0, 1]:
print('amps[{}] is for first={}, second={}'.format(i, first, second))
i += 1
```
We can check that this is in fact the ordering with a
circuit that flips one qubit out of two:
```
q_stay = cirq.NamedQubit('q_stay')
q_flip = cirq.NamedQubit('q_flip')
c = cirq.Circuit(cirq.X(q_flip))
# first qubit in order flipped
result = simulator.simulate(c, qubit_order=[q_flip, q_stay])
print(abs(result.final_state_vector).round(3))
# second qubit in order flipped
result = simulator.simulate(c, qubit_order=[q_stay, q_flip])
print(abs(result.final_state_vector).round(3))
```
## Stepping through a circuit
When debugging, it is useful to not just see the end
result of a circuit, but to inspect, or even modify, the
state of the system at different steps in the circuit.
To support this, Cirq provides a method to return an iterator
over a ``Moment`` by ``Moment`` simulation. This method is named ``simulate_moment_steps``:
```
circuit = cirq.Circuit()
circuit.append(basic_circuit())
for i, step in enumerate(simulator.simulate_moment_steps(circuit)):
print('state at step %d: %s' % (i, np.around(step.state_vector(), 3)))
```
The object returned by the ``moment_steps`` iterator is a
``StepResult``. This object has the state along with any
measurements that occurred **during** that step (so does
not include measurement results from previous ``Moments``).
In addition, the ``StepResult`` contains ``set_state()``,
which can be used to set the ``state``. One can pass a valid
full state to this method by passing a numpy array. Or,
alternatively, one can pass an integer, and then the state
will be set lie entirely in the computation basis state
for the binary expansion of the passed integer.
## Parameterized values and studies
In addition to circuit gates with fixed values, Cirq also
supports gates which can have ``Symbol`` value (see
[Gates](gates.ipynb)). These are values that can be resolved
at *run-time*.
For simulators, these values are resolved by
providing a ``ParamResolver``. A ``ParamResolver`` provides
a map from the ``Symbol``'s name to its assigned value.
```
import sympy
rot_w_gate = cirq.X**sympy.Symbol('x')
circuit = cirq.Circuit()
circuit.append([rot_w_gate(q0), rot_w_gate(q1)])
for y in range(5):
resolver = cirq.ParamResolver({'x': y / 4.0})
result = simulator.simulate(circuit, resolver)
print(np.round(result.final_state_vector, 2))
```
Here we see that the ``Symbol`` is used in two gates, and then the resolver provides this value at run time.
Parameterized values are most useful in defining what we call a
``sweep``. A ``sweep`` is a sequence of trials, where each
trial is a run with a particular set of parameter values.
Running a ``sweep`` returns a ``Result`` for each set of fixed parameter values and repetitions.
For instance:
```
resolvers = [cirq.ParamResolver({'x': y / 2.0}) for y in range(3)]
circuit = cirq.Circuit()
circuit.append([rot_w_gate(q0), rot_w_gate(q1)])
circuit.append([cirq.measure(q0, key='q0'), cirq.measure(q1, key='q1')])
results = simulator.run_sweep(program=circuit,
params=resolvers,
repetitions=2)
for result in results:
print(result)
```
Above we see that assigning different values to gate parameters yields
different results for each trial in the sweep, and that each trial is repeated
``repetitions`` times.
## Mixed state simulations
In addition to pure state simulation, Cirq also supports
simulation of mixed states.
Even though this simulator is not as efficient as the pure state simulators, they allow for a larger class of noisy circuits to be run as well as keeping track of the simulation's density matrix. This fact can allow for more exact simulations (for example,
the pure state simulator's Monte Carlo simulation only
allows sampling from the density matrix, not explicitly giving
the entries of the density matrix like the mixed state simulator
can do).
Mixed state simulation is supported by the
``cirq.DensityMatrixSimulator`` class.
Here is a simple example of simulating a channel using the
mixed state simulator:
```
q = cirq.NamedQubit('a')
circuit = cirq.Circuit(cirq.H(q), cirq.amplitude_damp(0.2)(q), cirq.measure(q))
simulator = cirq.DensityMatrixSimulator()
result = simulator.run(circuit, repetitions=100)
print(result.histogram(key='a'))
```
We create a state in an equal superposition of 0 and 1,
then apply amplitude damping which takes 1 to 0 with
something like a probability of 0.2.
We see that instead of about 50 percent of the timing being in 0, about 20 percent of the 1 has been converted into 0, so we end up with total
around 60 percent in the 0 state.
Like the pure state simulators, the mixed state simulator
supports ``run()`` and ``run_sweeps()`` methods.
The ``cirq.DensityMatrixSimulator`` also supports getting access
to the density matrix of the circuit at the end of simulating
the circuit, or when stepping through the circuit. These are
done by the ``simulate()`` and ``simulate_sweep()`` methods, or,
for stepping through the circuit, via the ``simulate_moment_steps``
method. For example, we can simulate creating an equal
superposition followed by an amplitude damping channel with a
gamma of 0.2 by:
```
q = cirq.NamedQubit('a')
circuit = cirq.Circuit(cirq.H(q), cirq.amplitude_damp(0.2)(q))
simulator = cirq.DensityMatrixSimulator()
result = simulator.simulate(circuit)
print(np.around(result.final_density_matrix, 3))
```
We see that we have access to the density matrix at the
end of the simulation via ``final_density_matrix``.
## External simulators
There are a few high-performance circuit simulators which
provide an interface for simulating Cirq ``Circuit``s.
These projects are listed below, along with their PyPI package
name and a description of simulator methods that they support.
**Note:** In general, these simulators are optimized for
specific use cases. Before choosing a simulator, make sure it
supports the behavior that you need!
| Project name | PyPI package | Description |
| --- | --- | --- |
| [qsim](https://github.com/quantumlib/qsim) | qsimcirq | Implements ``SimulatesAmplitudes`` and ``SimulatesFinalState``. Recommended for deep circuits with up to 30 qubits (consumes 8GB RAM). Larger circuits are possible, but RAM usage doubles with each additional qubit. |
| [qsimh](https://github.com/quantumlib/qsim/blob/master/qsimcirq/qsimh_simulator.py) | qsimcirq | Implements ``SimulatesAmplitudes``. Intended for heavy parallelization across several computers; Cirq users should generally prefer qsim. |
| [qFlex](https://github.com/ngnrsaa/qflex) | qflexcirq | Implements ``SimulatesAmplitudes``. Recommended for shallow / low-entanglement circuits with more than 30 qubits. RAM usage is highly dependent on the number of two-qubit gates in the circuit. |
| [quimb](https://quimb.readthedocs.io/en/latest/) | quimb | Cirq-to-quimb translation layer provided in `contrib/quimb`. In addition to circuit simulation, this allows the use of quimb circuit-analysis tools on Cirq circuits. |
| github_jupyter |
## Random Forest Classifier
The model predicts the severity of the landslide (or if there will even be one) within the next 2 days, based on weather data from the past 5 days.
A Random Forest model with 113 trees yielded an accuracy of 81.21% when trained on slope data and precipitation and wind data over a 5 day period.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import sklearn
from sklearn.utils import shuffle
import pickle
df = pd.read_csv("dataset.csv")
len(df)
df['severity'].value_counts()
df['severity'].value_counts()
df = shuffle(df)
df.reset_index(inplace=True, drop=True)
print(len(df))
X = df.copy()
y = X.landslide
columns=[]
for i in range(9, 4, -1):
columns.append('humidity' + str(i))
columns.append('ARI' + str(i))
columns.append('slope')
columns.append('forest2')
# columns.append('osm')
X = X[columns]
X
```
## Scaling
```
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
X_train
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
```
## Prediction
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
model = RandomForestClassifier()
model.fit(X_train, y_train)
pred = model.predict(X_test)
print("ACCURACY:", accuracy_score(pred, y_test))
best = 1
highest = 0
for i in range(85, 150, 2):
rf = RandomForestClassifier(n_estimators = i)
rf.fit(X_train, y_train)
pred = rf.predict(X_test)
score = round(accuracy_score(pred, y_test)*10000)/100
print("n_estimators =", i, " ACCURACY:", score)
if score > highest:
highest = score
best = i
print("# of trees:", best, highest)
rf = RandomForestClassifier(n_estimators = 139)
rf.fit(X_train, y_train)
pred = rf.predict(X_test)
print(accuracy_score(pred, y_test))
from sklearn.metrics import confusion_matrix
array = confusion_matrix(y_test, pred)
array
array = [[1254,245],[161,1902]]
import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
binary=True
if binary:
df_cm = pd.DataFrame(array, index = [i for i in ["No", "Yes"]],
columns = [i for i in ["No", "Yes"]])
else:
df_cm = pd.DataFrame(array, index = [i for i in ["None", "Small", "Medium", "Large", "Very Large"]],
columns = [i for i in ["None", "Small", "Medium", "Large", "Very Large"]])
plt.figure(figsize = (10,7))
ax = sn.heatmap(df_cm, cmap="Greens", annot=True, annot_kws={"size":50}, fmt='g')
ax.tick_params(axis='both', which='major', labelsize=27)
plt.xlabel('Predicted', fontsize = 40)
# plt.title("KNN Confusion Matrix", fontsize = 50)
plt.ylabel('Actual', fontsize = 40)
plt.savefig("RF Matrix", bbox_inches="tight")
plt.show()
```
| github_jupyter |
```
import os
import pickle
import numpy as np
import astropy.table as atable
from astropy.io import fits
import matplotlib.pyplot as plt
# read in exposures outputed from desisurvey run
fsurveysim = os.path.join(os.environ['CSCRATCH'], 'desisurvey_output', 'exposures_280s_bgs14000_skybranch_v8.brightsky.fits')
exposures = fits.getdata(fsurveysim, 'exposures')
# get observing conditions of BGS exposures calculated from TILE position and MJD output
isbgs = pickle.load(open(fsurveysim.replace('.fits', '.is_bgs.p'), 'rb'))
fsky = 'sv1.bright_exps.20210217.Isky.npy'
fwave = 'sv1.bright_exps.20210217.wave.npy'
fobscond = 'sv1.bright_exps.20210217.obs_cond.hdf5'
bright_exps = atable.Table.read(fobscond, format='hdf5')
Iskies = np.load(fsky)
wave = np.load(fwave)
# observing conditions from median value of the GFAs
sv1_airmass = bright_exps['AIRMASS']
sv1_moon_frac = bright_exps['MOON_ILLUMINATION']
sv1_moon_sep = bright_exps['MOON_SEP_DEG']
sv1_moon_alt = 90. - bright_exps['MOON_ZD_DEG']
sv1_transp = bright_exps['TRANSPARENCY']
from scipy.ndimage import gaussian_filter1d
def smooth_sky(sky, smoothing=100.):
return gaussian_filter1d(sky, smoothing)
Isky5000_sv1 = np.array([np.interp(5000, wave, smooth_sky(Isky)) for Isky in Iskies])
fig = plt.figure(figsize=(15,5))
sub = fig.add_subplot(131)
sub.scatter(exposures['AIRMASS'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2, label='BGS surveysim')
sub.scatter(sv1_airmass, sv1_moon_frac, c='C1', label='SV1 BRIGHT')
sub.legend(loc='lower right', fontsize=20, handletextpad=0, markerscale=2)
sub.set_xlabel('airmass', fontsize=20)
sub.set_xlim(1., 2.)
sub.set_ylabel('moon illumination', fontsize=20)
sub = fig.add_subplot(132)
sub.scatter(exposures['MOON_ALT'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2)
sub.scatter(sv1_moon_alt, sv1_moon_frac, c='C1')
sub.set_xlabel('moon atlitude', fontsize=20)
sub.set_xlim(-90., 90.)
sub.set_yticklabels([])
sub = fig.add_subplot(133)
sub.scatter(exposures['MOON_SEP'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2)
sub.scatter(sv1_moon_sep, sv1_moon_frac, c='C1')
sub.set_xlabel('moon separation', fontsize=20)
sub.set_xlim(0., 180.)
sub.set_yticklabels([])
from itertools import chain, combinations_with_replacement
def bright_Isky5000_regression(airmass, moon_frac, moon_sep, moon_alt):
''' polynomial regression model for bright sky surface brightness at 5000A
*without twilight*. The regression model was fit using observed sky surface
brightnesses from DESI SV1, DESI CMX, and BOSS.
see
https://github.com/desi-bgs/bgs-cmxsv/blob/4c5f124164b649c595cd2dca87d14ba9f3b2c64d/doc/nb/sv1_sky_model_fit.ipynb
for detials.
'''
# polynomial regression cofficients for estimating exposure time factor during
# non-twilight from airmass, moon_frac, moon_sep, moon_alt
coeffs = np.array([
1.11964670e+00, 1.89072762e-01, 3.20306279e+00, 4.10688340e-02,
-2.66073069e-02, -5.44857514e-01, 4.15680599e+00, 1.75625108e-02,
-5.01143360e-03, 1.30579080e+00, 6.17225096e-02, -1.07765709e-01,
-7.23089844e-04, -5.42455907e-04, 6.32035728e-04])
theta = np.atleast_2d(np.array([airmass, moon_frac, moon_alt, moon_sep]).T)
combs = chain.from_iterable(combinations_with_replacement(range(4), i) for i in range(0, 3))
theta_transform = np.empty((theta.shape[0], len(coeffs)))
for i, comb in enumerate(combs):
theta_transform[:, i] = theta[:, comb].prod(1)
return np.dot(theta_transform, coeffs.T)
Isky5000_surveysim = bright_Isky5000_regression(exposures['AIRMASS'][isbgs], exposures['MOON_ILL'][isbgs], exposures['MOON_SEP'][isbgs], exposures['MOON_ALT'][isbgs])
import desisim.simexp
import specsim.instrument
from desimodel.io import load_throughput
# get nominal dark sky surface brightness
wavemin = load_throughput('b').wavemin - 10.0
wavemax = load_throughput('z').wavemax + 10.0
_wave = np.arange(round(wavemin, 1), wavemax, 0.8)
config = desisim.simexp._specsim_config_for_wave(_wave, dwave_out=0.8, specsim_config_file='desi')
sb_dict = config.load_table(config.atmosphere.sky, 'surface_brightness', as_dict=True)
Isky_dark = sb_dict['dark'] # nominal dark sky
Isky5000_dark = np.interp(5000, _wave, smooth_sky(Isky_dark))
not_actually_bright = (Isky5000_surveysim / Isky5000_dark < 2.5)
print('%i of %i exposures are <2.5x dark sky' % (np.sum(not_actually_bright), len(not_actually_bright)))
print('%.3f percent' % (np.sum(not_actually_bright)/len(not_actually_bright)*100.))
fig = plt.figure(figsize=(15,5))
sub = fig.add_subplot(131)
sub.scatter(exposures['AIRMASS'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2, label='> 2.5 x dark')
sub.scatter(exposures['AIRMASS'][isbgs][not_actually_bright], exposures['MOON_ILL'][isbgs][not_actually_bright], c='k', s=2, label='< 2.5 x dark')
sub.legend(loc='lower right', fontsize=15, handletextpad=0, markerscale=5)
sub.set_xlabel('airmass', fontsize=20)
sub.set_xlim(1., 2.)
sub.set_ylabel('moon illumination', fontsize=20)
sub = fig.add_subplot(132)
sub.scatter(exposures['MOON_ALT'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2)
sub.scatter(exposures['MOON_ALT'][isbgs][not_actually_bright], exposures['MOON_ILL'][isbgs][not_actually_bright], c='k', s=2)
sub.set_xlabel('moon atlitude', fontsize=20)
sub.set_xlim(-90., 90.)
sub.set_yticklabels([])
sub = fig.add_subplot(133)
sub.scatter(exposures['MOON_SEP'][isbgs], exposures['MOON_ILL'][isbgs], c='C0', s=2)
sub.scatter(exposures['MOON_SEP'][isbgs][not_actually_bright], exposures['MOON_ILL'][isbgs][not_actually_bright], c='k', s=2)
sub.set_xlabel('moon separation', fontsize=20)
sub.set_xlim(0., 180.)
sub.set_yticklabels([])
fig = plt.figure(figsize=(6,6))
sub = fig.add_subplot(111)
sub.hist(Isky5000_surveysim, range=(0., 20), bins=40, density=True, alpha=0.5)
sub.hist(Isky5000_sv1, range=(0., 20.), bins=40, density=True, alpha=0.5)
sub.set_xlabel('sky surface brightness', fontsize=25)
```
| github_jupyter |
```
import os
hostname = os.popen("hostname").read().split("\n")[0]
if(hostname != "reckoner1429-Predator-PH315-52" and hostname != "janhvijo"):
from google.colab import drive
from google.colab import drive
drive.mount('/content/gdrive')
! chmod 755 "/content/gdrive/My Drive/collab-var.sh"
! "/content/gdrive/My Drive/collab-var.sh"
%cd "/content/gdrive/My Drive/github/video-emotion-recognition"
import librosa
import librosa.display
import numpy as np
import utils.data_util as data_util
import utils.preprocess_util as preprocess_util
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Dense, Flatten, Concatenate
from tensorflow.keras import Model
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow.keras.models import load_model
import tensorflow.keras as keras
import time
import utils.config as config
from utils.hyparam_util import load_fusion_hyparam
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
print(gpu)
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
else:
print("no gpu available")
```
# Define Functions for Training
## Define function to pretrain the model
```
def fine_tune(model, dataset, train_data_gen, val_data_gen, epochs, batch_size, seed, network):
output_layer = Dense(len(dataset.EMOTION_CLASSES),
activation = 'softmax')(model.layers[-2].output)
model = Model(model.input,
output_layer,
name = 'ftm-' + model.name + '-' + network + '-' + str(batch_size))
model.compile(optimizer = 'rmsprop',
loss = 'categorical_crossentropy',
metrics = ['categorical_accuracy'])
model_save_dir_path = os.path.join(dataset.MODEL_SAVE_DIR,
'ftm-' + str(seed),
'saved_models')
model_save_path = os.path.join(model_save_dir_path,
model.name + '.h5')
if(not os.path.exists(model_save_path)):
history = model.fit(train_data_gen,
validation_data = val_data_gen,
epochs = epochs,
verbose = 2)
plt.plot(history.history['categorical_accuracy'])
if(not os.path.exists(model_save_dir_path)):
os.makedirs(model_save_dir_path)
model.save(model_save_path)
model_history_path = os.path.join(dataset.MODEL_SAVE_DIR,
'ftm-' + str(seed),
'history')
if(not os.path.exists(model_history_path)):
os.makedirs(model_history_path)
np.save(os.path.join(model_history_path,
model.name + '-history.npy'),
history.history)
model.evaluate(val_data_gen)
```
## Define the function for tranining fusion network
```
def train_fusion_network(ptrn_model_name, ptrn_face_batch_size, ptrn_audio_batch_size, X_train_gen, X_val_gen, iteration, dataset, hyparams, seed):
#=============================================== make layers non trainable =======================================================
model_face = keras.models.load_model(os.path.join(dataset.MODEL_SAVE_DIR, 'ftm-' + str(seed), 'saved_models/ftm-' \
+ ptrn_model_name + '-face-' + str(ptrn_face_batch_size) + '.h5'))
for layer in model_face.layers:
layer._name = layer.name + '-face'
layer.trainable = False
output_layer_face = (model_face.layers[-2].output)
model_audio = keras.models.load_model(os.path.join(dataset.MODEL_SAVE_DIR, 'ftm-' + str(seed), 'saved_models/ftm-' \
+ ptrn_model_name + '-audio-' + str(ptrn_face_batch_size) + '.h5'))
for layer in model_audio.layers:
layer._name = layer.name + '-audio'
layer.trainable = False
output_layer_audio = (model_audio.layers[-2].output)
# ==============================================Construct the fusion network ===================================================
layer = Concatenate()([output_layer_face, output_layer_audio])
l1_hyparams = load_fusion_hyparam(iteration, "layer1")
layer = Dense(2048, activation=l1_hyparams['activation'], kernel_initializer = l1_hyparams['kernel_initializer'], \
kernel_regularizer = l1_hyparams['kernel_regularizer'], activity_regularizer = l1_hyparams['activity_regularizer'])(layer)
layer = l1_hyparams['dropout_layer'](layer)
layer = Dense(len(dataset.EMOTION_CLASSES), activation = 'softmax')(layer)
model = Model([model_face.input, model_audio.input], layer, name = ptrn_model_name + '-' + str(ptrn_face_batch_size) + '-' \
+ str(ptrn_audio_batch_size) + '-' + str(hyparams['batch_size']) + '-' + str(iteration))
# ========================================= Compile the fusion network model ================================================
model.compile(optimizer = l1_hyparams['optimizer'], loss = 'categorical_crossentropy', metrics = ['categorical_accuracy'])
history = model.fit(X_train_gen, validation_data = X_val_gen, epochs=hyparams['epochs'], verbose = 2)
model_save_dir_path = os.path.join(dataset.MODEL_SAVE_DIR, 'iteration-' + str(iteration), 'saved_models')
if(not os.path.exists(model_save_dir_path)):
os.makedirs(model_save_dir_path)
model.save(os.path.join(model_save_dir_path, model.name + '.h5'))
model_history_path = os.path.join(dataset.MODEL_SAVE_DIR, 'iteration-' + str(iteration), 'history')
if(not os.path.exists(model_history_path)):
os.makedirs(model_history_path)
np.save(os.path.join(model_history_path, model.name + '-history.npy'),history.history)
plt.plot(history.history['loss'])
plt.plot(history.history['categorical_accuracy'])
model.evaluate(X_val_gen)
```
# Train the Networks
### Train the fusion Network
```
ptrn_face_batch_size = 8
ptrn_audio_batch_size = 8
ptrn_epochs = 200
def train(dataset, base_model, iteration):
#=============================================== load the dataset ========================================================================
SEED = 0
X_train_audio, X_test_audio, Y_train, Y_test = dataset.load_audio_filenames(SEED, 0.2)
X_train_face, X_test_face, Y_train, Y_test = dataset.load_visual_filenames(SEED, 0.2)
#=============================================== load the hyperparameters ==============================================================
hyparams = load_fusion_hyparam(iteration)
#======================================= fine tuning of facial model ===================================================================
print('training using ' + base_model.name + ', iteration = ' + str(iteration))
X_train_face_gen = data_util.FaceDataGenerator(X_train_face,
Y_train,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
X_val_face_gen = data_util.FaceDataGenerator(X_test_face,
Y_test,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
fine_tune(base_model, dataset, X_train_face_gen, X_val_face_gen, ptrn_epochs, ptrn_face_batch_size, SEED, 'face')
print("face fine tuned")
#=========================== fine tuning of audio model ====================================================================
X_train_audio_gen = data_util.AudioDataGenerator(X_train_audio,
Y_train,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
X_val_audio_gen = data_util.AudioDataGenerator(X_test_audio,
Y_test,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
fine_tune(base_model, dataset, X_train_audio_gen, X_val_audio_gen, ptrn_epochs, ptrn_audio_batch_size, SEED, 'audio')
print("audio fine tuned")
#=================================== training of fusion network ==============================================================
X_train_gen = data_util.MultimodalDataGenerator(X_train_face,
X_train_audio,
Y_train,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
X_val_gen = data_util.MultimodalDataGenerator(X_test_face,
X_test_audio,
Y_test,
hyparams['batch_size'],
hyparams['input_width'],
hyparams['input_height'])
train_fusion_network(base_model.name, ptrn_face_batch_size, ptrn_audio_batch_size, X_train_gen, X_val_gen, iteration, dataset, hyparams, SEED)
train(preprocess_util.RML(), tf.keras.applications.Xception(),'test')
```
| github_jupyter |
# PROJECT 2 : TEAM 11
Members: Talia Tandler, SeungU Lyu
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
import math
```
http://www.worldometers.info/world-population/us-population/
US pop in 2017 = 324,459,463
https://wwwnc.cdc.gov/travel/yellowbook/2018/infectious-diseases-related-to-travel/measles-rubeola
Measles incubation period 11 days average, infectious period 2-4 days before rash to after rash.
https://www.cdc.gov/vaccines/imz-managers/coverage/childvaxview/data-reports/mmr/trend/index.html
MMR immunization rate in 2017 = 90.7%
```
pop = 999 #population
init_im = 0.907 #initial immunity of the US population
beta = 0.9 #assumed contact rate
gamma = 1/7 #US recovery rate from measles
sigma = 0.091; #US rate from exposure period of 11 days to infected
```
## Question
### What is the result of lowering the measles immunity rate in a small community during a outbreak?
Measles is a highly infectious disease that can infect about 90% of people that come into contact with the patient. However, the disease is not common these days because of the MMR vaccination, which can effectively prevent people getting the disease. Due to the high vaccination rate, the United States was declared free of circulating measles in 2000. However there were 911 cases of measles between 2001 and 2011. These occurences arose due to individuals from other countries entering the U.S. with measles. Because of the disease's high infectious rate upon contact, herd immunity is considered very important for measles.
In 2015, a measles outbreak occured at Disney World causing more than 159 people to be infected during a single outbreak. Only 50~86% people exposed to this outbreak were vaccinated, causing an even bigger outbreak. This vaccination was lower than it should have been due to Anti-Vaccination Movements in the U.S. These lower rates lowered the population immunity rate and caused the herd immunity to not function as expected. The starter of this movement, Andrew Wakefield, stated that the MMR vaccination can cause autism in newborn children because of the mercury content inside the specific vaccine. Due to this false research, many parents became concerned with the side effects of the vaccination and opted to not vaccinate their children with MMR. As a result, there was a decently sized generation of children susceptible to measles because they did not receive the vaccination at birth.
This simulation utilizes an SEIR model to understand how varying the measles immunity rate in a community effects herd immunity.
## Methodology
In order to create this model, we:
1. Did background research on the MMR vaccination and the measles diseases and found a set of constants we would implement in our model.
2. Put the variables into a state function.
3. Set the total population to 1000, with initial infection number as one person infected with measles.
4. Ran the simulation based on the number measles infections every day.
5. Set a condition where the measles outbreak ends when the number infected people is less than one person.
6. Created graphs to visually represent our results.
```
def make_system (pop, init_im, beta, gamma, sigma):
"""Make a system object for the SCIR model
pop: Total US population
init_im: Initial Population Immunity
beta: effective contact number for patient
gamma: recovery rate for infected people
sigma: rate of incubation group moving to infectious group
return: System object"""
init = State(S = int(pop*(1 - init_im)), E = 0, I = 1, R = int(pop*init_im))
init /= np.sum(init)
#S: susceptible, E: exposed period, I: infected, R: recovered(immune to disease)
t0 = 0
t_end = 365 #number of days in 1 year
return System(init = init,
beta = beta,
gamma = gamma,
sigma = sigma,
t0 = t0,
t_end = t_end,
init_im = init_im)
```
make_system function sets the initial values for the state and returns it with other necessary variables. Since the model is a SEIR model, initial state init contains four values, S, E, I, R where S and R is determined by the initial size and immunization rate of the community, and I is set to 1 to show that one person is infected at the start. Time span for the simulation was set to a year, since every outbreak in this simulation ends within the period.
```
def update_func(state, time, system):
"""Update the SEIR model
state: starting variables of SEIR
t: time step
system: includes alpha,beta,gamma,omega rates
contact: current contact number for the state
"""
unpack(system)
s,e,i,r = state
#current population
total_pop = s+e+i+r
#change rate for each status
ds = (-beta*s*i)/total_pop #change in number of people susceptible
de = ((beta*s*i)/total_pop) - sigma*e #change in number of people moving to exposed period
di = sigma*e - gamma*i #change in people moving to infectious period
dr = gamma*i #change in people recovered
s += ds #number of people susceptible
e += de #number of people exposed
i += di #number of people infected
r += dr #number of people recovered
return State(S=s, E=e, I=i, R=r)
```
update_func function updates the state with four different differential equations. System object was unpacked at the beginning of the code to make it easy to read. Change in susceptible group is affected only by the number of people in infected group, which will raise the number of people in exposed group. There is no direct transition from susceptible group to the infected group, because measles have average of 11 days incubation period, where the person does not spread the disease during that period. Therefore, about 1/11 (sigma value) of people in the exposed group move to the infected group every day, showing that their incubatoin period has ended. It takes about 7 days in average for people to get recoverd, so 1/7 (gamma) of people infected is recovered every day.
```
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: TimeFrame
"""
unpack(system)
#creates timeframe to save daily states
frame = TimeFrame(columns=init.index)
frame.row[t0] = init
for time in linrange(t0, t_end):
frame.row[time+1] = update_func(frame.row[time], time, system)
return frame
```
run_simulation function takes a system object with a update_func function, and simulates the state for the duration of the time span set at the make_system function. It returns a TimeFrame object with all the state values for each time step.
```
def plot_results (S,E,I,R):
plot(S, '--', label = 'Susceptible')
plot(E, '-', label = 'Exposed')
plot(I, '.', label = 'Infected')
plot(R, ':', label = 'Recovered')
decorate(xlabel='Time (days)',
ylabel = 'Fraction of population')
```
A plotting function was made for convenience.
```
init_im = 0.907
system = make_system(pop, init_im, beta, gamma, sigma)
results = run_simulation(system, update_func);
```
The code was tested with 2017 average immunization rate for the U.S (90.7%), testing out what will happen if a measles infected person is introduced to a community of 1000 people in a real world situation.
```
plot_results(results.S, results.E, results.I, results.R)
decorate(title ='Figure 1')
```
The result shows that even though measles is a highly contagious disease, the measles outbreak ends without infecting number of people due high immunity rate. We call this herd immunity, because immunized people acts as a barrier that prevents disease to spread among the susceptible people. For each disease, there is specific percentage of people needed to create a herd immunity. Lowering the immunity rate will show abrupt change in infected people, once the herd immunity stops working.
```
init_im2 = 0.3
system = make_system(pop, init_im2, beta, gamma, sigma)
results2 = run_simulation(system, update_func)
results2;
```
Next, the code was tested with lowered initial immunity rate of 30%.
```
plot_results(results2.S, results2.E, results2.I, results2.R)
decorate (title = 'Figure 2')
```
The result is a lot different from the one above, showing that most of susceptible people become infected before the outbreak ends. This shows that the community with only 30% immunity rate has lost their herd immunity, because the number of immuned (recovered) people is too small to act as a barrier that protects the susceptible people. Seeing the result, we can assume that there must be a point between immunity rate of 30% to 90% where the herd immunity fails to function.
```
def calc_highest_infected(results):
"""Fraction of population infected during the simulation.
results: DataFrame with columns S, E, I, R
returns: fraction of population
"""
return max(results.I)
def sweep_init_im(imun_rate_array):
"""Sweep a range of values for beta.
beta_array: array of beta values
gamma: recovery rate
returns: SweepSeries that maps from beta to total infected
"""
sweep = SweepSeries()
for init_im in imun_rate_array:
system = make_system(pop, init_im, beta, gamma, sigma)
results = run_simulation(system, update_func)
sweep[system.init_im] = calc_highest_infected(results)*pop
return sweep
```
To carefully check out the impact due to the change of initial immunity for the community, a sweep_init_im function was created. The function checks out the highest number of people infected to the disease during the simulation. Since the number of people being infected at a day is proportional to the number of currently infected people, higher numbers means that the disease is spreading faster.
```
imun_rate_array = linspace(0, 1, 21)
sweep = sweep_init_im(imun_rate_array)
sweep
plot(sweep)
decorate(xlabel='Immunity Rate',
ylabel = 'Highest number of people infected during 1 outbreak',
title = 'Figure 3')
```
Looking at the table and the plot, we can examine that the speed of infection decreases almost linearly until the immunity rate reachs 80%. Actually, the table states that the maximum number of people infected after the initial immunization rate of 85% is 1, meaning that no one except for the initially infected person was infected during the outbreak. We guessed that the herd immunity for measles in this simulation must be around 80~85% range.
```
def calc_fraction_infected(results):
"""Fraction of susceptible population infected during the simulation.
results: DataFrame with columns S, E, I, R
returns: fraction of susceptible group population
"""
return (get_first_value(results.S) - get_last_value(results.S))/get_first_value(results.S)
def sweep_init_im2(imun_rate_array):
"""Sweep a range of values for beta.
beta_array: array of beta values
gamma: recovery rate
returns: SweepSeries that maps from beta to total infected
"""
sweep = SweepSeries()
for init_im in imun_rate_array:
system = make_system(pop, init_im, beta, gamma, sigma)
results = run_simulation(system, update_func)
sweep[system.init_im] = calc_fraction_infected(results) * 100
return sweep
```
To do a deeper analysis, another sweep_init_im function was created to check out the percentage of people in the susceptible group infected during the outbreak. It will give us more clear view toward the herd immunity for measles and hopefully reveal the danger of lowering immunity rate for a community.
```
imun_rate_array = linspace(0, 0.99, 34)
sweep2 = sweep_init_im2(imun_rate_array)
sweep2
plot(sweep2)
decorate(xlabel='Immunity Rate',
ylabel = '% of susceptible people getting measles during an outbreak',
title = 'Figure 4')
```
Until the immunity rate reaches 60%, more than 90% of people in the susceptible group is infected by the measles. However, the percentage drops abruptly after that, hitting less than 10% on immunity rate of 84%. This graph clearly shows the importance of herd immunity, and the threat people can face due to the lowering of the immunity rate.
## Results
This model uses SEIR methodology to examine how measels would spread throughout a community of 1000 individuals with varying immunity rates. Figure 1 depicts an SEIR representation based on a 90.7% measles immunity rate, equivalent to that of the immunity rate in the United States. Due to the high immunity rate, susceptible people are protected by the herd immunity, and the number of individuals in each of the categories, susceptible, recovered, and infected remains constant throughout the simulated outbreak.
Figure 2 represents an example of the SEIR model with an immunity rate of 30%. In this model, we can see that as the number of susceptible individuals decreases, the number of recovered individuals increases at an equal and opposite rate. The entire population get infected and later recovered from this measles outbreak within 150 days of the start.
Figure 3 depicts the predicted outcome of this model that as the immunity rate in a community increases, rate of infection decreases, thus the number of people infected during an outbreak will decrease. We see the number of infected individuals plateau around 80%~85% immunity.
Figure 4 depicts the percent of susceptible individuals that do contact measles during an outbreak. At low immunity rates (without herd immunity) a large percent of susceptible individuals do contact measles. As the immunity rate increases, this percentage decreases.
## Interpretation
As expected, as the immunity rate in the community increased, the highest number of people infected with measles during an outbreak decreased. The number of people infected with measles begins to plateau between an 80 - 85% immunity rate. From the data that Figure 4 is based on we can see that the ideal immunity rate for a community should be more than 80 - 85%, because the herd immunity is lost at the lowered immunity rate. Between these 2 numbers, the percent of susceptible individuals that contract measles drops sharply from 36% to 6%.
Our model does have several limitations:
1. We were unable to find an effective contact number or contact rate for measles within the United States. Having this number would have enabled us to calculate beta instead of just assuming it to be 0.9.
2. The model gets to a point where less than 1 person is infected with measles. This is physically impossible as you cannot have less than one person. In our results, we interpreted less than 1 to mean the individual did not have measles.
3. The outbreak usually happens country wide, not restricted into a single community. Due to the fact that the simulation was done in a close community, the results may vary in real world situation.
4. People who get measles are usually quarantined before they start infecting other people. One special feature about measles is the rash, which usuaully appears 14 days after exposure. In real world, people get quarantined right away when they get the rash. In this simulation, the factor was ignored. People can also get a MMR vaccination while they are exposed, meaning that not every exposed people move to the infected stage.
5. Measles spread differently among different age groups. Usually, it spread easily among the younger children. The age factor was ignored in this simulation due to its complexity.
## Abstract
In this model, we were seeking to find out the result of lowering the measles immunity rate in a small community during a outbreak. As predicted, we found that as the immunity rate in a community is lowered, the number of infections in a community increases. We also found that when immunity is between 80-85%, the number of individuals infected in a population begins to plateau. This finding indicated that the ideal immunity rate for a community of 1000 individuals is between 80-85%.
```
plot(sweep)
decorate(xlabel='Immunity Rate',
ylabel = 'Highest number of people infected during 1 outbreak',
title = 'Figure 3')
plot(sweep2)
decorate(xlabel='Immunity Rate',
ylabel = '% of susceptible people getting measles during an outbreak',
title = 'Figure 4')
```
| github_jupyter |
<img src="http://dask.readthedocs.io/en/latest/_images/dask_horizontal.svg"
align="right"
width="30%"
alt="Dask logo\">
# Dask DataFrames
We finished Chapter 1 by building a parallel dataframe computation over a directory of CSV files using `dask.delayed`. In this section we use `dask.dataframe` to automatically build similiar computations, for the common case of tabular computations. Dask dataframes look and feel like Pandas dataframes but they run on the same infrastructure that powers `dask.delayed`.
In this notebook we use the same airline data as before, but now rather than write for-loops we let `dask.dataframe` construct our computations for us. The `dask.dataframe.read_csv` function can take a globstring like `"data/nycflights/*.csv"` and build parallel computations on all of our data at once.
## When to use `dask.dataframe`
Pandas is great for tabular datasets that fit in memory. Dask becomes useful when the dataset you want to analyze is larger than your machine's RAM. The demo dataset we're working with is only about 200MB, so that you can download it in a reasonable time, but `dask.dataframe` will scale to datasets much larger than memory.
<img src="images/pandas_logo.png" align="right" width="28%">
The `dask.dataframe` module implements a blocked parallel `DataFrame` object that mimics a large subset of the Pandas `DataFrame` API. One Dask `DataFrame` is comprised of many in-memory pandas `DataFrames` separated along the index. One operation on a Dask `DataFrame` triggers many pandas operations on the constituent pandas `DataFrame`s in a way that is mindful of potential parallelism and memory constraints.
**Related Documentation**
* [DataFrame documentation](https://docs.dask.org/en/latest/dataframe.html)
* [DataFrame screencast](https://youtu.be/AT2XtFehFSQ)
* [DataFrame API](https://docs.dask.org/en/latest/dataframe-api.html)
* [DataFrame examples](https://examples.dask.org/dataframe.html)
* [Pandas documentation](https://pandas.pydata.org/pandas-docs/stable/)
**Main Take-aways**
1. Dask DataFrame should be familiar to Pandas users
2. The partitioning of dataframes is important for efficient execution
## Create data
```
%run prep.py -d flights
```
## Setup
```
from dask.distributed import Client
client = Client(n_workers=4)
```
We create artifical data.
```
from prep import accounts_csvs
accounts_csvs()
import os
import dask
filename = os.path.join('data', 'accounts.*.csv')
filename
```
Filename includes a glob pattern `*`, so all files in the path matching that pattern will be read into the same Dask DataFrame.
```
import dask.dataframe as dd
df = dd.read_csv(filename)
df.head()
# load and count number of rows
len(df)
```
What happened here?
- Dask investigated the input path and found that there are three matching files
- a set of jobs was intelligently created for each chunk - one per original CSV file in this case
- each file was loaded into a pandas dataframe, had `len()` applied to it
- the subtotals were combined to give you the final grand total.
### Real Data
Lets try this with an extract of flights in the USA across several years. This data is specific to flights out of the three airports in the New York City area.
```
df = dd.read_csv(os.path.join('data', 'nycflights', '*.csv'),
parse_dates={'Date': [0, 1, 2]})
```
Notice that the respresentation of the dataframe object contains no data - Dask has just done enough to read the start of the first file, and infer the column names and dtypes.
```
df
```
We can view the start and end of the data
```
df.head()
df.tail() # this fails
```
### What just happened?
Unlike `pandas.read_csv` which reads in the entire file before inferring datatypes, `dask.dataframe.read_csv` only reads in a sample from the beginning of the file (or first file if using a glob). These inferred datatypes are then enforced when reading all partitions.
In this case, the datatypes inferred in the sample are incorrect. The first `n` rows have no value for `CRSElapsedTime` (which pandas infers as a `float`), and later on turn out to be strings (`object` dtype). Note that Dask gives an informative error message about the mismatch. When this happens you have a few options:
- Specify dtypes directly using the `dtype` keyword. This is the recommended solution, as it's the least error prone (better to be explicit than implicit) and also the most performant.
- Increase the size of the `sample` keyword (in bytes)
- Use `assume_missing` to make `dask` assume that columns inferred to be `int` (which don't allow missing values) are actually floats (which do allow missing values). In our particular case this doesn't apply.
In our case we'll use the first option and directly specify the `dtypes` of the offending columns.
```
df = dd.read_csv(os.path.join('data', 'nycflights', '*.csv'),
parse_dates={'Date': [0, 1, 2]},
dtype={'TailNum': str,
'CRSElapsedTime': float,
'Cancelled': bool})
df.tail() # now works
```
## Computations with `dask.dataframe`
We compute the maximum of the `DepDelay` column. With just pandas, we would loop over each file to find the individual maximums, then find the final maximum over all the individual maximums
```python
maxes = []
for fn in filenames:
df = pd.read_csv(fn)
maxes.append(df.DepDelay.max())
final_max = max(maxes)
```
We could wrap that `pd.read_csv` with `dask.delayed` so that it runs in parallel. Regardless, we're still having to think about loops, intermediate results (one per file) and the final reduction (`max` of the intermediate maxes). This is just noise around the real task, which pandas solves with
```python
df = pd.read_csv(filename, dtype=dtype)
df.DepDelay.max()
```
`dask.dataframe` lets us write pandas-like code, that operates on larger than memory datasets in parallel.
```
%time df.DepDelay.max().compute()
```
This writes the delayed computation for us and then runs it.
Some things to note:
1. As with `dask.delayed`, we need to call `.compute()` when we're done. Up until this point everything is lazy.
2. Dask will delete intermediate results (like the full pandas dataframe for each file) as soon as possible.
- This lets us handle datasets that are larger than memory
- This means that repeated computations will have to load all of the data in each time (run the code above again, is it faster or slower than you would expect?)
As with `Delayed` objects, you can view the underlying task graph using the `.visualize` method:
```
# notice the parallelism
df.DepDelay.max().visualize()
```
## Exercises
In this section we do a few `dask.dataframe` computations. If you are comfortable with Pandas then these should be familiar. You will have to think about when to call `compute`.
### 1.) How many rows are in our dataset?
If you aren't familiar with pandas, how would you check how many records are in a list of tuples?
```
# Your code here
len(df)
```
### 2.) In total, how many non-canceled flights were taken?
With pandas, you would use [boolean indexing](https://pandas.pydata.org/pandas-docs/stable/indexing.html#boolean-indexing).
```
# Your code here
len(df[~df.Cancelled])
```
### 3.) In total, how many non-cancelled flights were taken from each airport?
*Hint*: use [`df.groupby`](https://pandas.pydata.org/pandas-docs/stable/groupby.html).
```
# Your code here
df[~df.Cancelled].groupby('Origin').Origin.count().compute()
```
### 4.) What was the average departure delay from each airport?
Note, this is the same computation you did in the previous notebook (is this approach faster or slower?)
```
# Your code here
df.groupby("Origin").DepDelay.mean().compute()
```
### 5.) What day of the week has the worst average departure delay?
```
# Your code here
df.groupby("DayOfWeek").DepDelay.mean().compute()
```
## Sharing Intermediate Results
When computing all of the above, we sometimes did the same operation more than once. For most operations, `dask.dataframe` hashes the arguments, allowing duplicate computations to be shared, and only computed once.
For example, lets compute the mean and standard deviation for departure delay of all non-canceled flights. Since dask operations are lazy, those values aren't the final results yet. They're just the recipe required to get the result.
If we compute them with two calls to compute, there is no sharing of intermediate computations.
```
non_cancelled = df[~df.Cancelled]
mean_delay = non_cancelled.DepDelay.mean()
std_delay = non_cancelled.DepDelay.std()
%%time
mean_delay_res = mean_delay.compute()
std_delay_res = std_delay.compute()
```
But let's try by passing both to a single `compute` call.
```
%%time
mean_delay_res, std_delay_res = dask.compute(mean_delay, std_delay)
```
Using `dask.compute` takes roughly 1/2 the time. This is because the task graphs for both results are merged when calling `dask.compute`, allowing shared operations to only be done once instead of twice. In particular, using `dask.compute` only does the following once:
- the calls to `read_csv`
- the filter (`df[~df.Cancelled]`)
- some of the necessary reductions (`sum`, `count`)
To see what the merged task graphs between multiple results look like (and what's shared), you can use the `dask.visualize` function (we might want to use `filename='graph.pdf'` to save the graph to disk so that we can zoom in more easily):
```
dask.visualize(mean_delay, std_delay)
```
## How does this compare to Pandas?
Pandas is more mature and fully featured than `dask.dataframe`. If your data fits in memory then you should use Pandas. The `dask.dataframe` module gives you a limited `pandas` experience when you operate on datasets that don't fit comfortably in memory.
During this tutorial we provide a small dataset consisting of a few CSV files. This dataset is 45MB on disk that expands to about 400MB in memory. This dataset is small enough that you would normally use Pandas.
We've chosen this size so that exercises finish quickly. Dask.dataframe only really becomes meaningful for problems significantly larger than this, when Pandas breaks with the dreaded
MemoryError: ...
Furthermore, the distributed scheduler allows the same dataframe expressions to be executed across a cluster. To enable massive "big data" processing, one could execute data ingestion functions such as `read_csv`, where the data is held on storage accessible to every worker node (e.g., amazon's S3), and because most operations begin by selecting only some columns, transforming and filtering the data, only relatively small amounts of data need to be communicated between the machines.
Dask.dataframe operations use `pandas` operations internally. Generally they run at about the same speed except in the following two cases:
1. Dask introduces a bit of overhead, around 1ms per task. This is usually negligible.
2. When Pandas releases the GIL `dask.dataframe` can call several pandas operations in parallel within a process, increasing speed somewhat proportional to the number of cores. For operations which don't release the GIL, multiple processes would be needed to get the same speedup.
## Dask DataFrame Data Model
For the most part, a Dask DataFrame feels like a pandas DataFrame.
So far, the biggest difference we've seen is that Dask operations are lazy; they build up a task graph instead of executing immediately (more details coming in [Schedulers](05_distributed.ipynb)).
This lets Dask do operations in parallel and out of core.
In [Dask Arrays](03_array.ipynb), we saw that a `dask.array` was composed of many NumPy arrays, chunked along one or more dimensions.
It's similar for `dask.dataframe`: a Dask DataFrame is composed of many pandas DataFrames. For `dask.dataframe` the chunking happens only along the index.
<img src="http://dask.pydata.org/en/latest/_images/dask-dataframe.svg" width="30%">
We call each chunk a *partition*, and the upper / lower bounds are *divisions*.
Dask *can* store information about the divisions. For now, partitions come up when you write custom functions to apply to Dask DataFrames
## Converting `CRSDepTime` to a timestamp
This dataset stores timestamps as `HHMM`, which are read in as integers in `read_csv`:
```
crs_dep_time = df.CRSDepTime.head(10)
crs_dep_time
```
To convert these to timestamps of scheduled departure time, we need to convert these integers into `pd.Timedelta` objects, and then combine them with the `Date` column.
In pandas we'd do this using the `pd.to_timedelta` function, and a bit of arithmetic:
```
import pandas as pd
# Get the first 10 dates to complement our `crs_dep_time`
date = df.Date.head(10)
# Get hours as an integer, convert to a timedelta
hours = crs_dep_time // 100
hours_timedelta = pd.to_timedelta(hours, unit='h')
# Get minutes as an integer, convert to a timedelta
minutes = crs_dep_time % 100
minutes_timedelta = pd.to_timedelta(minutes, unit='m')
# Apply the timedeltas to offset the dates by the departure time
departure_timestamp = date + hours_timedelta + minutes_timedelta
departure_timestamp
```
### Custom code and Dask Dataframe
We could swap out `pd.to_timedelta` for `dd.to_timedelta` and do the same operations on the entire dask DataFrame. But let's say that Dask hadn't implemented a `dd.to_timedelta` that works on Dask DataFrames. What would you do then?
`dask.dataframe` provides a few methods to make applying custom functions to Dask DataFrames easier:
- [`map_partitions`](http://dask.pydata.org/en/latest/dataframe-api.html#dask.dataframe.DataFrame.map_partitions)
- [`map_overlap`](http://dask.pydata.org/en/latest/dataframe-api.html#dask.dataframe.DataFrame.map_overlap)
- [`reduction`](http://dask.pydata.org/en/latest/dataframe-api.html#dask.dataframe.DataFrame.reduction)
Here we'll just be discussing `map_partitions`, which we can use to implement `to_timedelta` on our own:
```
# Look at the docs for `map_partitions`
help(df.CRSDepTime.map_partitions)
```
The basic idea is to apply a function that operates on a DataFrame to each partition.
In this case, we'll apply `pd.to_timedelta`.
```
hours = df.CRSDepTime // 100
# hours_timedelta = pd.to_timedelta(hours, unit='h')
hours_timedelta = hours.map_partitions(pd.to_timedelta, unit='h')
minutes = df.CRSDepTime % 100
# minutes_timedelta = pd.to_timedelta(minutes, unit='m')
minutes_timedelta = minutes.map_partitions(pd.to_timedelta, unit='m')
departure_timestamp = df.Date + hours_timedelta + minutes_timedelta
departure_timestamp
departure_timestamp.head()
```
### Exercise: Rewrite above to use a single call to `map_partitions`
This will be slightly more efficient than two separate calls, as it reduces the number of tasks in the graph.
```
def compute_departure_timestamp(df):
pass # TODO: implement this
departure_timestamp = df.map_partitions(compute_departure_timestamp)
departure_timestamp.head()
def compute_departure_timestamp(df):
hours = df.CRSDepTime // 100
hours_timedelta = pd.to_timedelta(hours, unit='h')
minutes = df.CRSDepTime % 100
minutes_timedelta = pd.to_timedelta(minutes, unit='m')
return df.Date + hours_timedelta + minutes_timedelta
departure_timestamp = df.map_partitions(compute_departure_timestamp)
departure_timestamp.head()
```
## Limitations
### What doesn't work?
Dask.dataframe only covers a small but well-used portion of the Pandas API.
This limitation is for two reasons:
1. The Pandas API is *huge*
2. Some operations are genuinely hard to do in parallel (e.g. sort)
Additionally, some important operations like ``set_index`` work, but are slower
than in Pandas because they include substantial shuffling of data, and may write out to disk.
## Learn More
* [DataFrame documentation](https://docs.dask.org/en/latest/dataframe.html)
* [DataFrame screencast](https://youtu.be/AT2XtFehFSQ)
* [DataFrame API](https://docs.dask.org/en/latest/dataframe-api.html)
* [DataFrame examples](https://examples.dask.org/dataframe.html)
* [Pandas documentation](https://pandas.pydata.org/pandas-docs/stable/)
```
client.shutdown()
```
| github_jupyter |
# Simulate data
This notebook generates simulated experiments by sampling from the VAE using the user selected template experiment as a guide for sampling
```
%load_ext autoreload
%autoreload 2
import os
import sys
import glob
import pandas as pd
import numpy as np
import random
import seaborn as sns
import umap
from keras.models import load_model
from sklearn.decomposition import PCA
import pickle
from plotnine import (ggplot,
labs,
geom_line,
geom_point,
geom_errorbar,
aes,
ggsave,
theme_bw,
theme,
xlim,
ylim,
facet_wrap,
scale_color_manual,
guides,
guide_legend,
element_blank,
element_text,
element_rect,
element_line,
coords)
from ponyo import utils, generate_template_data
from numpy.random import seed
randomState = 123
seed(randomState)
# Read in config variables
base_dir = os.path.abspath(os.path.join(os.getcwd(),"../"))
config_file = os.path.abspath(os.path.join(base_dir,
"config_human.tsv"))
params = utils.read_config(config_file)
# Load params
local_dir = params["local_dir"]
dataset_name = params['dataset_name']
NN_architecture = params['NN_architecture']
num_runs = params['num_simulated']
project_id = params['project_id']
template_data_file = params['template_data_file']
original_compendium_file = params['compendium_data_file']
normalized_compendium_file = params['normalized_compendium_data_file']
scaler_file = params['scaler_transform_file']
NN_dir = os.path.join(
base_dir,
dataset_name,
"models",
NN_architecture)
shared_genes_file = os.path.join(
local_dir,
"shared_gene_ids.pickle")
# Load pickled file
scaler = pickle.load(open(scaler_file, "rb"))
shared_genes = pickle.load(open(shared_genes_file, "rb"))
# Read data
compendium = pd.read_csv(
original_compendium_file,
header=0,
sep='\t',
index_col=0)
normalized_compendium = pd.read_csv(
normalized_compendium_file,
header=0,
sep='\t',
index_col=0)
template_data = pd.read_csv(
template_data_file,
header=0,
sep='\t',
index_col=0)
```
### Simulate experiments using selected template experiment
```
# Simulate experiments
# Make sure range is correct
# Generate multiple simulated datasets
for i in range(num_runs):
generate_template_data.shift_template_experiment(
normalized_compendium_file,
project_id,
NN_architecture,
dataset_name,
scaler,
local_dir,
base_dir,
i)
# Truncate simulated experiments
smRNA_samples = ["SRR493961",
"SRR493962",
"SRR493963",
"SRR493964",
"SRR493965",
"SRR493966",
"SRR493967",
"SRR493968",
"SRR493969",
"SRR493970",
"SRR493971",
"SRR493972"]
for i in range(num_runs):
simulated_data_file = os.path.join(
local_dir,
"pseudo_experiment",
"selected_simulated_data_"+project_id+"_"+str(i)+".txt")
# Read simulated data
simulated_data = pd.read_csv(
simulated_data_file,
header=0,
sep='\t',
index_col=0)
# Drop samples
simulated_data = simulated_data.drop(smRNA_samples)
# Drop genes
#simulated_data = simulated_data[shared_genes]
# Save
simulated_data.to_csv(simulated_data_file, float_format='%.5f', sep='\t')
```
### Quick validation of simulated experiments
**Spot check expression values**
1. Values are different between different simulated data files (meaning it was a different simulated dataset), and different from the template experiment
2. Range of values is scaled the same as the compendium
```
# Compendium
print(compendium.shape)
compendium.head()
sns.distplot(compendium['GPR176'])
# Template experiment
print(template_data.shape)
template_data.head()
sns.distplot(template_data['GPR176'])
# Manual select one simulated experiment
simulated_file_1 = os.path.join(
local_dir,
"pseudo_experiment",
"selected_simulated_data_"+project_id+"_0.txt")
# Read data
simulated_test_1 = pd.read_csv(
simulated_file_1,
header=0,
sep='\t',
index_col=0)
print(simulated_test_1.shape)
simulated_test_1.head()
sns.distplot(simulated_test_1['GPR176'])
# Manual select another simulated experiment
simulated_file_2 = os.path.join(
local_dir,
"pseudo_experiment",
"selected_simulated_data_"+project_id+"_10.txt")
# Read data
simulated_test_2 = pd.read_csv(
simulated_file_2,
header=0,
sep='\t',
index_col=0)
print(simulated_test_2.shape)
simulated_test_2.head()
sns.distplot(simulated_test_2['GPR176'])
```
**Check clustering of simulated samples**
Check PCA embedding of original experiment and simulated experiments. We expect to see a similar structure in the template and simulated experiments. Also expect to see that the simulated experiment follows the distribution of the compendium.
```
# Load VAE models
model_encoder_file = glob.glob(os.path.join(
NN_dir,
"*_encoder_model.h5"))[0]
weights_encoder_file = glob.glob(os.path.join(
NN_dir,
"*_encoder_weights.h5"))[0]
model_decoder_file = glob.glob(os.path.join(
NN_dir,
"*_decoder_model.h5"))[0]
weights_decoder_file = glob.glob(os.path.join(
NN_dir,
"*_decoder_weights.h5"))[0]
# Load saved models
loaded_model = load_model(model_encoder_file)
loaded_decode_model = load_model(model_decoder_file)
loaded_model.load_weights(weights_encoder_file)
loaded_decode_model.load_weights(weights_decoder_file)
pca = PCA(n_components=2)
# Embedding of real compendium (encoded)
# Scale compendium
#compendium_scaled = scaler.transform(compendium)
#compendium_scaled_df = pd.DataFrame(compendium_scaled,
# columns=compendium.columns,
# index=compendium.index)
# Encode normalized compendium into latent space
compendium_encoded = loaded_model.predict_on_batch(normalized_compendium)
compendium_encoded_df = pd.DataFrame(data=compendium_encoded,
index=compendium.index)
# Get and save PCA model
model = pca.fit(compendium_encoded_df)
compendium_PCAencoded = model.transform(compendium_encoded_df)
compendium_PCAencoded_df = pd.DataFrame(data=compendium_PCAencoded,
index=compendium_encoded_df.index,
columns=['1','2'])
# Add label
compendium_PCAencoded_df['experiment_id'] = 'background'
# Embedding of real template experiment (encoded)
# Scale template data
template_data_scaled = scaler.transform(template_data)
template_data_scaled_df = pd.DataFrame(template_data_scaled,
columns=template_data.columns,
index=template_data.index)
# Encode template experiment into latent space
template_encoded = loaded_model.predict_on_batch(template_data_scaled_df)
template_encoded_df = pd.DataFrame(data=template_encoded,
index=template_data.index)
template_PCAencoded = model.transform(template_encoded_df)
template_PCAencoded_df = pd.DataFrame(data=template_PCAencoded,
index=template_encoded_df.index,
columns=['1','2'])
# Add back label column
template_PCAencoded_df['experiment_id'] = 'template_experiment'
```
**Visualization in latent space**
```
encoded_simulated_file = os.path.join(local_dir,
"pseudo_experiment",
"selected_simulated_encoded_data_"+project_id+"_10.txt")
# Embedding of simulated experiment (encoded)
simulated_encoded_df = pd.read_csv(
encoded_simulated_file,
header=0,
sep='\t',
index_col=0)
# Drop samples
simulated_encoded_df = simulated_encoded_df.drop(smRNA_samples)
simulated_PCAencoded = model.transform(simulated_encoded_df)
simulated_PCAencoded_df = pd.DataFrame(data=simulated_PCAencoded,
index=simulated_encoded_df.index,
columns=['1','2'])
# Add back label column
simulated_PCAencoded_df['experiment_id'] = 'simulated_experiment'
# Concatenate dataframes
combined_PCAencoded_df = pd.concat([compendium_PCAencoded_df,
template_PCAencoded_df,
simulated_PCAencoded_df])
print(combined_PCAencoded_df.shape)
combined_PCAencoded_df.head()
# Plot
fig = ggplot(combined_PCAencoded_df, aes(x='1', y='2'))
fig += geom_point(aes(color='experiment_id'), alpha=0.2)
fig += labs(x ='PCA 1',
y = 'PCA 2',
title = 'PCA original data with experiments (latent space)')
fig += theme_bw()
fig += theme(
legend_title_align = "center",
plot_background=element_rect(fill='white'),
legend_key=element_rect(fill='white', colour='white'),
legend_title=element_text(family='sans-serif', size=15),
legend_text=element_text(family='sans-serif', size=12),
plot_title=element_text(family='sans-serif', size=15),
axis_text=element_text(family='sans-serif', size=12),
axis_title=element_text(family='sans-serif', size=15)
)
fig += guides(colour=guide_legend(override_aes={'alpha': 1}))
fig += scale_color_manual(['#bdbdbd', 'red', 'blue'])
fig += geom_point(data=combined_PCAencoded_df[combined_PCAencoded_df['experiment_id'] == 'template_experiment'],
alpha=0.2,
color='blue')
fig += geom_point(data=combined_PCAencoded_df[combined_PCAencoded_df['experiment_id'] == 'simulated_experiment'],
alpha=0.1,
color='red')
print(fig)
```
**Visualization in latent space (re-encoded)**
```
# Embedding of simulated experiment (encoded)
# Scale simulated data
simulated_test_2_scaled = scaler.transform(simulated_test_2)
simulated_test_2_scaled_df = pd.DataFrame(simulated_test_2_scaled,
columns=simulated_test_2.columns,
index=simulated_test_2.index)
# Encode simulated experiment into latent space
simulated_encoded = loaded_model.predict_on_batch(simulated_test_2_scaled_df)
simulated_encoded_df = pd.DataFrame(simulated_encoded,
index=simulated_test_2.index)
simulated_PCAencoded = model.transform(simulated_encoded_df)
simulated_PCAencoded_df = pd.DataFrame(data=simulated_PCAencoded,
index=simulated_encoded_df.index,
columns=['1','2'])
# Add back label column
simulated_PCAencoded_df['experiment_id'] = 'simulated_experiment'
# Concatenate dataframes
combined_PCAencoded_df = pd.concat([compendium_PCAencoded_df,
template_PCAencoded_df,
simulated_PCAencoded_df])
print(combined_PCAencoded_df.shape)
combined_PCAencoded_df.head()
# Plot
fig = ggplot(combined_PCAencoded_df, aes(x='1', y='2'))
fig += geom_point(aes(color='experiment_id'), alpha=0.2)
fig += labs(x ='PCA 1',
y = 'PCA 2',
title = 'PCA original data with experiments (latent space)')
fig += theme_bw()
fig += theme(
legend_title_align = "center",
plot_background=element_rect(fill='white'),
legend_key=element_rect(fill='white', colour='white'),
legend_title=element_text(family='sans-serif', size=15),
legend_text=element_text(family='sans-serif', size=12),
plot_title=element_text(family='sans-serif', size=15),
axis_text=element_text(family='sans-serif', size=12),
axis_title=element_text(family='sans-serif', size=15)
)
fig += guides(colour=guide_legend(override_aes={'alpha': 1}))
fig += scale_color_manual(['#bdbdbd', 'red', 'blue'])
fig += geom_point(data=combined_PCAencoded_df[combined_PCAencoded_df['experiment_id'] == 'template_experiment'],
alpha=0.2,
color='blue')
fig += geom_point(data=combined_PCAencoded_df[combined_PCAencoded_df['experiment_id'] == 'simulated_experiment'],
alpha=0.1,
color='red')
print(fig)
```
**Visualization in gene space**
```
# Embedding of real compendium
# Get and save model
model = umap.UMAP(random_state=randomState).fit(normalized_compendium)
compendium_UMAPencoded = model.transform(normalized_compendium)
compendium_UMAPencoded_df = pd.DataFrame(data=compendium_UMAPencoded,
index=normalized_compendium.index,
columns=['1','2'])
# Add label
compendium_UMAPencoded_df['experiment_id'] = 'background'
# Embedding of real template experiment
template_UMAPencoded = model.transform(template_data_scaled_df)
template_UMAPencoded_df = pd.DataFrame(data=template_UMAPencoded,
index=template_data_scaled_df.index,
columns=['1','2'])
# Add back label column
template_UMAPencoded_df['experiment_id'] = 'template_experiment'
# Embedding of simulated template experiment
simulated_UMAPencoded = model.transform(simulated_test_2_scaled_df)
simulated_UMAPencoded_df = pd.DataFrame(data=simulated_UMAPencoded,
index=simulated_test_2_scaled_df.index,
columns=['1','2'])
# Add back label column
simulated_UMAPencoded_df['experiment_id'] = 'simulated_experiment'
# Concatenate dataframes
combined_UMAPencoded_df = pd.concat([compendium_UMAPencoded_df,
template_UMAPencoded_df,
simulated_UMAPencoded_df])
combined_UMAPencoded_df.shape
# Plot
fig = ggplot(combined_UMAPencoded_df, aes(x='1', y='2'))
fig += geom_point(aes(color='experiment_id'), alpha=0.2)
fig += labs(x ='UMAP 1',
y = 'UMAP 2',
title = 'UMAP original data with experiments (gene space)')
fig += theme_bw()
fig += theme(
legend_title_align = "center",
plot_background=element_rect(fill='white'),
legend_key=element_rect(fill='white', colour='white'),
legend_title=element_text(family='sans-serif', size=15),
legend_text=element_text(family='sans-serif', size=12),
plot_title=element_text(family='sans-serif', size=15),
axis_text=element_text(family='sans-serif', size=12),
axis_title=element_text(family='sans-serif', size=15)
)
fig += guides(colour=guide_legend(override_aes={'alpha': 1}))
fig += scale_color_manual(['#bdbdbd', 'red', 'blue'])
fig += geom_point(data=combined_UMAPencoded_df[combined_UMAPencoded_df['experiment_id'] == 'template_experiment'],
alpha=0.2,
color='blue')
fig += geom_point(data=combined_UMAPencoded_df[combined_UMAPencoded_df['experiment_id'] == 'simulated_experiment'],
alpha=0.2,
color='red')
print(fig)
```
**Observation:**
The latent space encoded simulated data embedded into the first 2 PCs shows the linear shift of the simulated experiment compared to the template experiment, as expected. However, we noticed that the shift is fairly large and moves the experiment outside of the background distribution. This is something to consider for future iterations of this simulation.
The latent space visualization (re-encoded) is another visualization using PCA. In this case, the simulated data, which was shifted in the latent space and decoded into gene space, is encoded into latent space. We can see the shift fo the simulated data, however the relationship between samples are not an exact match to the template experiment because we have decoded the simulated data, which is a nonlinear transformation. This nonlinear transformation of the data is even more apparent when we embed the simulated data in gene space into UMAP space.
| github_jupyter |
```
#manually mount the drive!
HOME="/content/drive/My Drive"
#@title installs
%cd /content
!rm -rf ddsp_gm2
!git clone --single-branch --branch using_vocoder https://github.com/gianmarcohutter/ddsp_gm2
%cd ddsp_gm2
%tensorflow_version 2.x
!pip install -e /content/ddsp_gm2[data_preparation]
!pip install -e /content/ddsp_gm2[ddsp]
!pip install mir_eval
#for phonemes
!apt-get update
!apt-get install -y swig libpulse-dev
!swig -version
!pip3 install pocketsphinx
!pip3 list | grep pocketsphinx
#@title imports
#dont know if this is needed?
import ddsp
'''
import time
from ddsp.training import (data, decoders, encoders, models, preprocessing,
train_util, trainers)
from ddsp.colab.colab_utils import play, specplot, DEFAULT_SAMPLE_RATE
import gin
import matplotlib.pyplot as plt
import numpy as np
import tensorflow.compat.v2 as tf
import tensorflow_datasets as tfds
'''
#for phonemes
import itertools
import pocketsphinx
import os
#@title create test folder
#the last one showing up in the list below is where its actually saved
%cd {HOME}
#make a new testx folder:
i=0
while(True):
test_folder="test"
i=i+1
test_folder=test_folder+str(i)
print(test_folder)
var=![ -d {test_folder} ] && echo 'does exist'
if(len(var)==0):
!mkdir {test_folder}
#%cd test_folder
break
# copy audio to the test folder
DRIVE_DIR=os.path.join(HOME,test_folder)
#copy the all songs from that folder into the drive_dir
AUDIO_STORAGE=SAVE_DIR = os.path.join(HOME,"vocoder_alternative_jack_spleeter/.")
!cp -r "$AUDIO_STORAGE" "$DRIVE_DIR"
!pwd
print(DRIVE_DIR)
#going back to /content is crucial! otherwise it gets saved in drive and reused the next time the script is run
%cd /content/
import glob
import os
AUDIO_DIR = 'data/audio'
AUDIO_FILEPATTERN = AUDIO_DIR + '/*'
!mkdir -p $AUDIO_DIR
if DRIVE_DIR:
SAVE_DIR = os.path.join(DRIVE_DIR, 'GM-Voice')
else:
SAVE_DIR = '/content/models/GM-Voice'
!mkdir -p "$SAVE_DIR"
if DRIVE_DIR:
mp3_files = glob.glob(os.path.join(DRIVE_DIR, '*.mp3'))
wav_files = glob.glob(os.path.join(DRIVE_DIR, '*.wav'))
audio_files = mp3_files + wav_files
for fname in audio_files:
target_name = os.path.join(AUDIO_DIR,
os.path.basename(fname).replace(' ', '_'))
print('Copying {} to {}'.format(fname, target_name))
!cp "$fname" $target_name
import glob
import os
TRAIN_TFRECORD = 'data/train.tfrecord'
TRAIN_TFRECORD_FILEPATTERN = TRAIN_TFRECORD + '*'
# Copy dataset from drive if dataset has already been created.
drive_data_dir = os.path.join(DRIVE_DIR, 'data')
drive_dataset_files = glob.glob(drive_data_dir + '/*')
if DRIVE_DIR and len(drive_dataset_files) > 0:
!cp "$drive_data_dir"/* data/
print("did not prepare the tfrecord new")
else:
# Make a new dataset.
print("prepare the tfrecord new")
if not glob.glob(AUDIO_FILEPATTERN):
raise ValueError('No audio files found. Please use the previous cell to '
'upload.')
!ddsp_prepare_tfrecord_phonemes \
--input_audio_filepatterns=$AUDIO_FILEPATTERN \
--output_tfrecord_path=$TRAIN_TFRECORD \
--num_shards=10 \
--alsologtostderr
# Copy dataset to drive for safe-keeping.
if DRIVE_DIR:
!mkdir "$drive_data_dir"/
print('Saving to {}'.format(drive_data_dir))
!cp $TRAIN_TFRECORD_FILEPATTERN "$drive_data_dir"/
# for Quantile normalization
from ddsp.colab import colab_utils
import ddsp.training
#not sure what provider i need to choose here
data_provider = ddsp.training.data.TFRecordProvider(TRAIN_TFRECORD_FILEPATTERN)
#data_provider = ddsp.training.data_phoneme.TFRecordProviderPhoneme(TRAIN_TFRECORD_FILEPATTERN)
dataset = data_provider.get_dataset(shuffle=False)
PICKLE_FILE_PATH = os.path.join(SAVE_DIR, 'dataset_statistics.pkl')
colab_utils.save_dataset_statistics(data_provider, PICKLE_FILE_PATH,batch_size=16)
from ddsp.colab import colab_utils
import ddsp.training
from matplotlib import pyplot as plt
import numpy as np
data_provider = ddsp.training.data_phoneme.TFRecordProviderPhoneme(TRAIN_TFRECORD_FILEPATTERN)
dataset = data_provider.get_dataset(shuffle=False)
try:
ex = next(iter(dataset))
except StopIteration:
raise ValueError(
'TFRecord contains no examples. Please try re-running the pipeline with '
'different audio file(s).')
colab_utils.specplot(ex['audio'])
colab_utils.play(ex['audio'])
for key, value in ex.items():
print (key)
colab_utils.specplot(ex['alternative_audio'])
colab_utils.play(ex['alternative_audio'])
f, ax = plt.subplots(4, 1, figsize=(14, 4))
x = np.linspace(0, 4.0, 1000)
ax[0].set_ylabel('loudness_db')
ax[0].plot(x, ex['loudness_db'])
ax[1].set_ylabel('F0_Hz')
ax[1].set_xlabel('seconds')
ax[1].plot(x, ex['f0_hz'])
ax[2].set_ylabel('F0_confidence')
ax[2].set_xlabel('seconds')
ax[2].plot(x, ex['f0_confidence'])
ax[3].set_ylabel('phonemes')
ax[3].plot(x,ex['phoneme'])
```
#training
```
%reload_ext tensorboard
import tensorboard as tb
tb.notebook.start('--logdir "{}"'.format(SAVE_DIR))
#run the actual training
!ddsp_run \
--mode=train \
--alsologtostderr \
--save_dir="$SAVE_DIR" \
--gin_file=models/ae_phoneme.gin \
--gin_file=datasets/tfrecord_phoneme.gin \
--gin_param="TFRecordProviderPhoneme.file_pattern='$TRAIN_TFRECORD_FILEPATTERN'" \
--gin_param="batch_size=16" \
--gin_param="train_util.train.num_steps=24000" \
--gin_param="train_util.train.steps_per_save=2400" \
--gin_param="trainers.Trainer.checkpoints_to_keep=10"
from google.colab import output
def beep():
output.eval_js('new Audio("https://upload.wikimedia.org/wikipedia/commons/0/05/Beep-09.ogg").play()')
beep()
from ddsp.colab.colab_utils import play, specplot
import ddsp.training
import gin
from matplotlib import pyplot as plt
import numpy as np
data_provider = ddsp.training.data_phoneme.TFRecordProviderPhoneme(TRAIN_TFRECORD_FILEPATTERN)
dataset = data_provider.get_batch(batch_size=1, shuffle=True)
try:
batch = next(iter(dataset))
except OutOfRangeError:
raise ValueError(
'TFRecord contains no examples. Please try re-running the pipeline with '
'different audio file(s).')
# Parse the gin config.
gin_file = os.path.join(SAVE_DIR, 'operative_config-0.gin')
gin.parse_config_file(gin_file)
# Load model
model = ddsp.training.models.Autoencoder()
model.restore(SAVE_DIR)
# Resynthesize audio.
audio_gen = model(batch, training=False)
audio = batch['audio']
print('Original Audio')
specplot(audio)
play(audio)
print('Resynthesis')
specplot(audio_gen)
play(audio_gen)
from ddsp.colab import colab_utils
import tensorflow as tf
import os
CHECKPOINT_ZIP = 'GM-Voice.zip'
latest_checkpoint_fname = os.path.basename(tf.train.latest_checkpoint(SAVE_DIR))
!cd "$SAVE_DIR" && zip $CHECKPOINT_ZIP $latest_checkpoint_fname* operative_config-0.gin dataset_statistics.pkl
!cp "$SAVE_DIR/$CHECKPOINT_ZIP" ./
#colab_utils.download(CHECKPOINT_ZIP)
```
| github_jupyter |
```
import gc
from datetime import datetime, timedelta,date
import warnings
import itertools
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
from sklearn.random_projection import GaussianRandomProjection
from sklearn.random_projection import SparseRandomProjection,johnson_lindenstrauss_min_dim
from sklearn.decomposition import PCA, FastICA,NMF,LatentDirichletAllocation,IncrementalPCA,MiniBatchSparsePCA
from sklearn.decomposition import TruncatedSVD,FactorAnalysis,KernelPCA
import seaborn as sns
import lightgbm as lgb
import matplotlib.pyplot as plt
from sklearn.metrics import mean_squared_error, log_loss, mean_absolute_error
from sklearn.model_selection import StratifiedKFold, KFold
from scipy.stats import ks_2samp
import tqdm
#settings
warnings.filterwarnings('ignore')
np.random.seed(2018)
pd.set_option("display.max_columns", None)
pd.set_option('display.max_rows',130)
def get_prefix(group_col, target_col, prefix=None):
if isinstance(group_col, list) is True:
g = '_'.join(group_col)
else:
g = group_col
if isinstance(target_col, list) is True:
t = '_'.join(target_col)
else:
t = target_col
if prefix is not None:
return prefix + '_' + g + '_' + t
return g + '_' + t
def groupby_helper(df, group_col, target_col, agg_method, prefix_param=None):
try:
prefix = get_prefix(group_col, target_col, prefix_param)
print(group_col, target_col, agg_method)
group_df = df.groupby(group_col)[target_col].agg(agg_method)
group_df.columns = ['{}_{}'.format(prefix, m) for m in agg_method]
except BaseException as e:
print(e)
return group_df.reset_index()
def create_new_columns(name,aggs):
return [name + '_' + k + '_' + agg for k in aggs.keys() for agg in aggs[k]]
historical_trans_df = pd.read_csv('input/historical_transactions.csv')
new_merchant_trans_df = pd.read_csv('input/new_merchant_transactions.csv')
merchant_df = pd.read_csv('input/merchants.csv')
train_df = pd.read_csv('input/train.csv')
test_df = pd.read_csv('input/test.csv')
def get_hist_default_prorcessing(df):
df['purchase_date'] = pd.to_datetime(df['purchase_date'])
df['year'] = df['purchase_date'].dt.year
df['weekofyear'] = df['purchase_date'].dt.weekofyear
df['month'] = df['purchase_date'].dt.month
df['dayofweek'] = df['purchase_date'].dt.dayofweek
df['weekend'] = (df.purchase_date.dt.weekday >=5).astype(int)
df['hour'] = df['purchase_date'].dt.hour
df['authorized_flag'] = df['authorized_flag'].map({'Y':1, 'N':0})
df['category_1'] = df['category_1'].map({'Y':1, 'N':0})
df['category_3'] = df['category_3'].map({'A':0, 'B':1, 'C':2})
df['month_diff'] = ((datetime(2012,4,1) - df['purchase_date']).dt.days)//30
df['month_diff'] += df['month_lag']
df['reference_date'] = (df['year']+(df['month'] - df['month_lag'])//12)*100 + (((df['month'] - df['month_lag'])%12) + 1)*1
return df
historical_trans_df = get_hist_default_prorcessing(historical_trans_df)
new_merchant_trans_df = get_hist_default_prorcessing(new_merchant_trans_df)
historical_trans_df = historical_trans_df.sort_values('purchase_date')
new_merchant_trans_df = new_merchant_trans_df.sort_values('purchase_date')
historical_trans_df.loc[historical_trans_df['installments']==999,'installments_999'] = 1
new_merchant_trans_df.loc[new_merchant_trans_df['installments']==999,'installments_999'] = 1
historical_trans_df.loc[historical_trans_df['installments']==999,'installments'] = -1
new_merchant_trans_df.loc[new_merchant_trans_df['installments']==999,'installments_999'] = -1
```
### Feature Engineering
```
all_df = pd.concat([train_df,test_df])
group_df = groupby_helper(historical_trans_df,['card_id','month_lag'], 'purchase_amount',['count','mean'])
group_df['card_id_month_lag_purchase_amount_count'] = group_df['card_id_month_lag_purchase_amount_count']/(1-group_df['month_lag'])
group_df['card_id_month_lag_purchase_amount_mean'] = group_df['card_id_month_lag_purchase_amount_mean']/(1-group_df['month_lag'])
del group_df['month_lag']
count_df = groupby_helper(group_df,['card_id'], 'card_id_month_lag_purchase_amount_count',['sum','mean','std'])
mean_df = groupby_helper(group_df,['card_id'], 'card_id_month_lag_purchase_amount_mean',['sum','mean','std'])
all_df = all_df.merge(count_df, on=['card_id'], how='left')
all_df = all_df.merge(mean_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'month',['nunique','max','min','mean','std'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'merchant_id',['nunique'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'merchant_category_id',['nunique'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'subsector_id',['nunique'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'state_id',['nunique'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id'], 'city_id',['nunique'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
aggs = {}
for col in ['hour', 'weekofyear', 'dayofweek', 'year']:
aggs[col] = ['nunique', 'mean', 'min', 'max']
aggs['purchase_amount'] = ['sum','max','min','mean','var']
aggs['installments'] = ['sum','max','min','mean','var']
aggs['purchase_date'] = ['max','min']
aggs['month_lag'] = ['max','min','mean','var']
aggs['month_diff'] = ['mean', 'max', 'min', 'var']
aggs['weekend'] = ['sum', 'mean', 'min', 'max']
aggs['category_1'] = ['sum', 'mean', 'min', 'max']
aggs['authorized_flag'] = ['sum', 'mean', 'min', 'max']
#aggs['category_2'] = ['sum', 'mean', 'min', 'max']
#aggs['category_3'] = ['sum', 'mean', 'min', 'max']
aggs['card_id'] = ['size']
aggs['reference_date'] = ['median']
new_columns = create_new_columns('hist',aggs)
historical_trans_group_df = historical_trans_df.groupby('card_id').agg(aggs)
historical_trans_group_df.columns = new_columns
historical_trans_group_df.reset_index(drop=False,inplace=True)
historical_trans_group_df['hist_purchase_date_diff'] = (historical_trans_group_df['hist_purchase_date_max'] - historical_trans_group_df['hist_purchase_date_min']).dt.days
historical_trans_group_df['hist_purchase_date_average'] = historical_trans_group_df['hist_purchase_date_diff']/historical_trans_group_df['hist_card_id_size']
historical_trans_group_df['hist_purchase_date_uptonow'] = (datetime(2012,4,1) - historical_trans_group_df['hist_purchase_date_max']).dt.days
historical_trans_group_df['hist_purchase_date_uptomin'] = (datetime(2012,4,1) - historical_trans_group_df['hist_purchase_date_min']).dt.days
all_df = all_df.merge(historical_trans_group_df, on=['card_id'], how='left')
def get_train_default_prorcessing(df):
df['first_active_month'] = pd.to_datetime(df['first_active_month'])
df['dayofweek'] = df['first_active_month'].dt.dayofweek
df['weekofyear'] = df['first_active_month'].dt.weekofyear
df['dayofyear'] = df['first_active_month'].dt.dayofyear
df['quarter'] = df['first_active_month'].dt.quarter
#df['is_month_start'] = df['first_active_month'].dt.is_month_start
df['month'] = df['first_active_month'].dt.month
df['year'] = df['first_active_month'].dt.year
#df['elapsed_time'] = (datetime(2018, 2, 1).date() - df['first_active_month'].dt.date).dt.days
df['elapsed_time'] = (datetime(2019,1, 20) - df['first_active_month']).dt.days
#df['after_big_event'] = (datetime(2012,4, 1) - df['first_active_month']).dt.days
df['hist_first_buy'] = (df['hist_purchase_date_min'] - df['first_active_month']).dt.days
df['hist_last_buy'] = (df['hist_purchase_date_max'] - df['first_active_month']).dt.days
df['year_month'] = df['year']*100 + df['month']
df['hist_diff_reference_date_first'] = 12*(df['hist_reference_date_median']//100 - df['year_month']//100) + (df['hist_reference_date_median']%100 - df['year_month']%100)
df['hist_diff_reference_date_last'] = 12*(df['hist_purchase_date_max'].dt.year - df['year_month']//100) + (df['hist_purchase_date_max'].dt.month - df['year_month']%100)
df['hist_diff_first_last'] = df['hist_diff_reference_date_first'] - df['hist_diff_reference_date_last']
df['hist_flag_ratio'] = df['hist_authorized_flag_sum'] / df['hist_card_id_size']
#df['new_flag_ratio'] = df['new_hist_authorized_flag_sum'] / df['new_hist_card_id_size']
#df['new_hist_flag_ratio'] = 1/(1+df['hist_flag_ratio'])
for f in ['hist_purchase_date_max','hist_purchase_date_min']:
df[f] = df[f].astype(np.int64) * 1e-9
df['hist_CLV'] = df['hist_card_id_size'] * df['hist_purchase_amount_sum'] / df['hist_month_diff_mean']
del df['year']
del df['year_month']
return df
all_df = get_train_default_prorcessing(all_df)
all_df['feature123'] = all_df['feature_1'].astype(str) +'_'+all_df['feature_2'].astype(str)+'_'+all_df['feature_3'].astype(str)
all_df['feature123'] = pd.factorize(all_df['feature123'])[0]
group_df = groupby_helper(historical_trans_df,['card_id','month_lag'], 'subsector_id',['nunique'])
group_df['card_id_month_lag_subsector_id_nunique'] = group_df['card_id_month_lag_subsector_id_nunique']/(1-group_df['month_lag'])
del group_df['month_lag']
count_df = groupby_helper(group_df,['card_id'], 'card_id_month_lag_subsector_id_nunique',['sum','mean','std'])
all_df = all_df.merge(count_df, on=['card_id'], how='left')
historical_trans_df =historical_trans_df.merge(all_df[['card_id','first_active_month','target','feature123']], on='card_id',how='left')
historical_trans_df['ym'] = historical_trans_df['purchase_date'].dt.year*100 + historical_trans_df['purchase_date'].dt.month
historical_trans_df['first_active_month_ym'] = historical_trans_df['first_active_month'].dt.year*100 + historical_trans_df['first_active_month'].dt.month
historical_trans_df['is_smaller_firstactive'] = 0
historical_trans_df.loc[historical_trans_df['ym']<=historical_trans_df['first_active_month_ym'],'is_smaller_firstactive'] = 1
group_df = groupby_helper(historical_trans_df,'card_id', 'is_smaller_firstactive',['sum'])
temp = groupby_helper(historical_trans_df,'card_id', 'card_id',['size'])
group_df = group_df.merge(temp, on='card_id', how='left')
group_df['before_firstactive_purchase_ratio']=group_df['card_id_is_smaller_firstactive_sum'] /group_df['card_id_card_id_size']
all_df = all_df.merge(group_df[['card_id','before_firstactive_purchase_ratio']], on=['card_id'], how='left')
all_df['feature123_frequency_encoding']= all_df['feature123'].map(all_df['feature123'].value_counts()/all_df.shape[0])
```
마지막 1달, 마지막 3달, 마지막 6달 masking
```
group_df = groupby_helper(historical_trans_df,['card_id'], 'month_lag',['min'])
historical_trans_df = historical_trans_df.merge(group_df, on='card_id', how='left')
group_df = historical_trans_df.groupby(['card_id'])['month_lag'].unique().reset_index()
group_df['month_lag_last'] = group_df['month_lag'].apply(lambda x: x[-1])
group_df['month_lag_last3month'] =group_df['month_lag'].apply(lambda x: x[-3:])
group_df['month_lag_last3month'] = group_df['month_lag_last3month'].apply(lambda x: x[0])
group_df['month_lag_last6month'] =group_df['month_lag'].apply(lambda x: x[-6:])
group_df['month_lag_last6month'] = group_df['month_lag_last6month'].apply(lambda x: x[0])
historical_trans_df = historical_trans_df.merge(group_df, on='card_id', how='left')
del historical_trans_df['month_lag_y']
historical_trans_df.rename(columns={'month_lag_x':'month_lag'},inplace=True)
last_month = historical_trans_df.loc[historical_trans_df['month_lag']==historical_trans_df['month_lag_last']]
last_3month = historical_trans_df.loc[historical_trans_df['month_lag']>=historical_trans_df['month_lag_last3month']]
last_6month = historical_trans_df.loc[historical_trans_df['month_lag']>=historical_trans_df['month_lag_last6month']]
group_df = groupby_helper(last_month,['card_id'], 'merchant_id',['nunique'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month,['card_id'], 'subsector_id',['nunique'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month,['card_id'], 'city_id',['nunique'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month,['card_id'], 'card_id',['size'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month,['card_id'], 'purchase_amount',['sum','min','mean','max','std'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_3month,['card_id'], 'card_id',['size'],'last_3month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_6month,['card_id'], 'card_id',['size'],'last_6month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
historical_trans_df_merchant = historical_trans_df.merge(merchant_df[['merchant_id', 'merchant_group_id','numerical_1', 'numerical_2',
'most_recent_sales_range', 'most_recent_purchases_range',
'avg_sales_lag3', 'avg_purchases_lag3', 'active_months_lag3',
'avg_sales_lag6', 'avg_purchases_lag6', 'active_months_lag6',
'avg_sales_lag12', 'avg_purchases_lag12', 'active_months_lag12',
'category_4']], on='merchant_id', how='left')
historical_trans_df_merchant['most_sales_purchases_range'] = historical_trans_df_merchant['most_recent_sales_range'] +historical_trans_df_merchant['most_recent_purchases_range']
historical_trans_df_merchant['avg_lag3_aov'] = historical_trans_df_merchant['avg_sales_lag3']/historical_trans_df_merchant['avg_purchases_lag3']
historical_trans_df_merchant['avg_lag3_conv'] = historical_trans_df_merchant['avg_purchases_lag3']/historical_trans_df_merchant['active_months_lag3']
historical_trans_df_merchant['avg_lag3_rps'] = historical_trans_df_merchant['avg_sales_lag3']/historical_trans_df_merchant['active_months_lag3']
historical_trans_df_merchant['avg_lag6_aov'] = historical_trans_df_merchant['avg_sales_lag6']/historical_trans_df_merchant['avg_purchases_lag6']
historical_trans_df_merchant['avg_lag6_conv'] = historical_trans_df_merchant['avg_purchases_lag6']/historical_trans_df_merchant['active_months_lag6']
historical_trans_df_merchant['avg_lag6_rps'] = historical_trans_df_merchant['avg_sales_lag6']/historical_trans_df_merchant['active_months_lag6']
historical_trans_df_merchant['avg_lag12_aov'] = historical_trans_df_merchant['avg_sales_lag12']/historical_trans_df_merchant['avg_purchases_lag12']
historical_trans_df_merchant['avg_lag12_conv'] = historical_trans_df_merchant['avg_purchases_lag12']/historical_trans_df_merchant['active_months_lag12']
historical_trans_df_merchant['avg_lag12_rps'] = historical_trans_df_merchant['avg_sales_lag12']/historical_trans_df_merchant['active_months_lag12']
group_df = groupby_helper(historical_trans_df_merchant,['card_id'], 'avg_lag3_rps',['min','max','sum','mean','std'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
historical_trans_df_merchant.columns
historical_trans_df_merchant['category_4'] = historical_trans_df_merchant['category_4'].map({'Y':1,'N':0})
group_df = groupby_helper(historical_trans_df_merchant,['card_id'], 'category_4',['sum','mean'])
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df_merchant.loc[historical_trans_df_merchant['month_lag']==historical_trans_df_merchant['month_lag_last']],['card_id'], 'category_4',['sum','mean'],'last_month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
```
### 실패 피쳐
```
last_month_auth1 = last_month.loc[last_month['authorized_flag']==1]
group_df = groupby_helper(last_month_auth1,['card_id'], 'merchant_id',['nunique'],'last_month_auth1')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month_auth1,['card_id'], 'subsector_id',['nunique'],'last_month_auth1')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_month_auth1,['card_id'], 'city_id',['nunique'],'last_month_auth1')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_3month,['card_id'], 'city_id',['nunique'],'last_3month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(last_6month,['card_id'], 'merchant_id',['nunique'],'last_6month')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id','ym'], 'merchant_id',['nunique'])
group_df_pivot = group_df.pivot('card_id','ym','card_id_ym_merchant_id_nunique').reset_index()
group_df_pivot.columns = ['card_id'] + ['ym_{}_card_id_ym_merchant_id_nunique'.format(m) for m in [201701, 201702, 201703, 201704, 201705,
201706, 201707, 201708, 201709, 201710, 201711,
201712, 201801, 201802]]
all_df = all_df.merge(group_df_pivot, on=['card_id'], how='left')
group_df = groupby_helper(historical_trans_df,['card_id','month_lag'], 'merchant_id',['nunique'])
group_df['card_id_month_lag_merchant_id_nunique'] = group_df['card_id_month_lag_merchant_id_nunique']/(1-group_df['month_lag'])
del group_df['month_lag']
count_df = groupby_helper(group_df,['card_id'], 'card_id_month_lag_merchant_id_nunique',['sum','mean','std'])
all_df = all_df.merge(count_df, on=['card_id'], how='left')
group_df_mode = pd.read_csv('input/subsector_id_mode.csv')
group_df = groupby_helper(group_df_mode.reset_index(),'card_id', 'subsector_id',['nunique'],'mode')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df_mode_mci = pd.read_csv('input/merchant_category_id_mode.csv')
group_df = groupby_helper(group_df_mode_mci.reset_index(),'card_id', 'merchant_category_id',['nunique'],'mode')
all_df = all_df.merge(group_df, on=['card_id'], how='left')
```
### Modeling
target value 생성
```
group_df = groupby_helper(new_merchant_trans_df,['card_id'], 'purchase_date',['max'],'new_hist')
group_df['new_hist_card_id_purchase_date_max'] = np.sqrt(group_df['new_hist_card_id_purchase_date_max'].astype(np.int64) * 1e-9)
all_df = all_df.merge(group_df, on=['card_id'], how='left')
group_df = groupby_helper(new_merchant_trans_df,['card_id'], 'purchase_date',['min'],'new_hist')
group_df['new_hist_card_id_purchase_date_min'] = np.sqrt(group_df['new_hist_card_id_purchase_date_min'].astype(np.int64) * 1e-9)
all_df = all_df.merge(group_df, on=['card_id'], how='left')
for col in all_df.columns:
if col.find('card_id_avg_lag6_rps') !=-1:
print(col)
del all_df[col]
for col in all_df.columns:
if all_df[col].nunique() == 1:
print(col)
del all_df[col]
```
(325540, 104)
(290001, 104)
(35539, 104)
```
print(all_df.shape)
train_df = all_df.loc[all_df['new_hist_card_id_purchase_date_min'].notnull()]
test_df = all_df.loc[all_df['new_hist_card_id_purchase_date_min'].isnull()]
print(train_df.shape)
print(test_df.shape)
train_columns = [c for c in train_df.columns if c not in ['card_id','first_active_month','new_hist_card_id_size','target','outliers'
,'hist_weekend_max','hist_year_max','before_firstactive_purchase_ratio',
'hist_weekend_min','hist_year_nunique','new_hist_card_id_purchase_date_max',
'new_hist_card_id_purchase_date_min']]
new_col = []
for col in train_columns:
if col.find('new_')==-1:
print(col)
new_col.append(col)
train_columns = new_col.copy()
train_columns
train = train_df.copy()
target = train['new_hist_card_id_purchase_date_min']
del train['new_hist_card_id_purchase_date_min']
param = {'num_leaves': 24,
'min_data_in_leaf': 30,
'objective':'regression',
'max_depth': 5,
'learning_rate': 0.02,
"min_child_samples": 50,
"boosting": "gbdt",
"feature_fraction": 0.6,
"bagging_fraction": 0.7 ,
"metric": 'mae',
"lambda_l1": 0.1,
"verbosity": -1,
"nthread": 24,
"seed": 6}
#prepare fit model with cross-validation
np.random.seed(2019)
feature_importance_df = pd.DataFrame()
folds = KFold(n_splits=9, shuffle=True, random_state=4950)
oof = np.zeros(len(train))
predictions = np.zeros(len(test_df))
cv_score_list = []
for fold_, (trn_idx, val_idx) in enumerate(folds.split(train)):
strLog = "fold {}".format(fold_+1)
print(strLog)
trn_data = lgb.Dataset(train.iloc[trn_idx][train_columns], label=target.iloc[trn_idx])#, categorical_feature=categorical_feats)
val_data = lgb.Dataset(train.iloc[val_idx][train_columns], label=target.iloc[val_idx])#, categorical_feature=categorical_feats)
num_round = 10000
clf = lgb.train(param, trn_data, num_round, valid_sets = [trn_data, val_data], verbose_eval=100, early_stopping_rounds = 100)
oof[val_idx] = clf.predict(train.iloc[val_idx][train_columns], num_iteration=clf.best_iteration)
cv_score = mean_absolute_error(oof[val_idx], target.iloc[val_idx])
cv_score_list.append(cv_score)
#feature importance
fold_importance_df = pd.DataFrame()
fold_importance_df["Feature"] = train_columns
fold_importance_df["importance"] = clf.feature_importance()
fold_importance_df["fold"] = fold_ + 1
feature_importance_df = pd.concat([feature_importance_df, fold_importance_df], axis=0)
#predictions
predictions += clf.predict(test_df[train_columns], num_iteration=clf.best_iteration) / folds.n_splits
cv_score = mean_absolute_error(oof, target)
print(cv_score)
print(cv_score_list)
print(np.std(cv_score_list))
withoutoutlier_predictions = predictions.copy()
```
3.3979265177382523
[3.410920807322843, 3.3781160121042837, 3.390035302847772, 3.3980637829332077, 3.41249645944368]
0.012929663651872132
```
filename = '{}_cv{:.6f}'.format(datetime.now().strftime('%Y%m%d_%H%M%S'), cv_score)
cols = (feature_importance_df[["Feature", "importance"]]
.groupby("Feature")
.mean()
.sort_values(by="importance", ascending=False)[:1000].index)
best_features = feature_importance_df.loc[feature_importance_df.Feature.isin(cols)]
plt.figure(figsize=(14,26))
sns.barplot(x="importance", y="Feature", data=best_features.sort_values(by="importance",ascending=False))
plt.title('LightGBM Features (averaged over folds)')
plt.tight_layout()
plt.savefig('fi/{}_lgbm_importances.png'.format(filename))
feature_importance_df.groupby('Feature')['importance'].sum().reset_index().sort_values('importance')
mean_absolute_error(np.floor(oof), target)
result = pd.concat([pd.DataFrame(np.power(oof,2),columns=['oof']),pd.DataFrame(np.power(target,2)).reset_index(drop=True)],axis=1)
sns.distplot(result['oof'])
sns.distplot(result['new_hist_card_id_purchase_date_min'])
result['oof'] = result['oof'].apply(lambda x: datetime.fromtimestamp(x))
result['new_hist_card_id_purchase_date_min'] = result['new_hist_card_id_purchase_date_min'].apply(lambda x: datetime.fromtimestamp(x))
convert_prediction = [datetime.fromtimestamp(x) for x in np.power(predictions,2)]
sub_df = pd.DataFrame({"card_id":test_df["card_id"].values})
sub_df['new_hist_card_id_purchase_date_max'] = convert_prediction
sub_df = pd.DataFrame({"card_id":test_df["card_id"].values})
sub_df['new_hist_purchase_date_min'] = convert_prediction
#sub_df.loc[sub_df['target']<-9,'target'] = -33.21928095
sub_df.to_csv("input/fill_new_hist_card_id_purchase_date_min_{}.csv".format(filename), index=False)
```
| github_jupyter |
# 📃 Solution for Exercise M1.03
The goal of this exercise is to compare the performance of our classifier in
the previous notebook (roughly 81% accuracy with `LogisticRegression`) to
some simple baseline classifiers. The simplest baseline classifier is one
that always predicts the same class, irrespective of the input data.
- What would be the score of a model that always predicts `' >50K'`?
- What would be the score of a model that always predicts `' <=50K'`?
- Is 81% or 82% accuracy a good score for this problem?
Use a `DummyClassifier` and do a train-test split to evaluate
its accuracy on the test set. This
[link](https://scikit-learn.org/stable/modules/model_evaluation.html#dummy-estimators)
shows a few examples of how to evaluate the generalization performance of these
baseline models.
```
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
```
We will first split our dataset to have the target separated from the data
used to train our predictive model.
```
target_name = "class"
target = adult_census[target_name]
data = adult_census.drop(columns=target_name)
```
We start by selecting only the numerical columns as seen in the previous
notebook.
```
numerical_columns = [
"age", "capital-gain", "capital-loss", "hours-per-week"]
data_numeric = data[numerical_columns]
```
Split the data and target into a train and test set.
```
from sklearn.model_selection import train_test_split
# solution
data_numeric_train, data_numeric_test, target_train, target_test = \
train_test_split(data_numeric, target, random_state=42)
```
Use a `DummyClassifier` such that the resulting classifier will always
predict the class `' >50K'`. What is the accuracy score on the test set?
Repeat the experiment by always predicting the class `' <=50K'`.
Hint: you can set the `strategy` parameter of the `DummyClassifier` to
achieve the desired behavior.
```
from sklearn.dummy import DummyClassifier
# solution
class_to_predict = " >50K"
high_revenue_clf = DummyClassifier(strategy="constant",
constant=class_to_predict)
high_revenue_clf.fit(data_numeric_train, target_train)
score = high_revenue_clf.score(data_numeric_test, target_test)
print(f"Accuracy of a model predicting only high revenue: {score:.3f}")
```
We clearly see that the score is below 0.5 which might be surprising at
first. We will now check the generalization performance of a model which always
predict the low revenue class, i.e. `" <=50K"`.
```
class_to_predict = " <=50K"
low_revenue_clf = DummyClassifier(strategy="constant",
constant=class_to_predict)
low_revenue_clf.fit(data_numeric_train, target_train)
score = low_revenue_clf.score(data_numeric_test, target_test)
print(f"Accuracy of a model predicting only low revenue: {score:.3f}")
```
We observe that this model has an accuracy higher than 0.5. This is due to
the fact that we have 3/4 of the target belonging to low-revenue class.
Therefore, any predictive model giving results below this dummy classifier
will not be helpful.
```
adult_census["class"].value_counts()
(target == " <=50K").mean()
```
In practice, we could have the strategy `"most_frequent"` to predict the
class that appears the most in the training target.
```
most_freq_revenue_clf = DummyClassifier(strategy="most_frequent")
most_freq_revenue_clf.fit(data_numeric_train, target_train)
score = most_freq_revenue_clf.score(data_numeric_test, target_test)
print(f"Accuracy of a model predicting the most frequent class: {score:.3f}")
```
So the `LogisticRegression` accuracy (roughly 81%) seems better than the
`DummyClassifier` accuracy (roughly 76%). In a way it is a bit reassuring,
using a machine learning model gives you a better performance than always
predicting the majority class, i.e. the low income class `" <=50K"`.
| github_jupyter |
# Day 1
#Numpy
```
#Exercise 1 answer
def p(x,coeff):
a = np.array(coeff)
for i in range(len(a)):
p[i] = a[i]*(x**i)
f = np.sum(p)
return f
#Exercise 2
"""
we want to use the inverse cdf method to draw values from a discrete distribution
"""
import numpy as np
import scipy as sc
"""
we want to write our method such that for any given probability vector q we can obtain
k draws from the discrete distribution corresponding to q
"""
def invcdf(q,k):
prob = np.array(q)
p = np.cumsum(prob)
x = []
for i in range(k):
u = np.random.uniform()
a = p.searchsorted(u)
x.append(a)
return x
```
#Scipy
```
#Exercise 1 Answer
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
"""
plotting the profit function
"""
p=5
q = np.linspace(0,2,1000)
pi = p*q - np.exp(q) - 0.5*(q**2)
plt.figure
plt.plot(q,pi)
plt.xlabel('q', fontsize=20)
plt.ylabel('$\pi$(q)', fontsize=20)
plt.title(f'Profit Function, price = {p}', fontsize=20)
plt.show()
import scipy as sc
from scipy import optimize
def profitmax(p,x):
answer = optimize.fminbound(lambda q: -p*q + np.exp(q) + 0.5*(q**2), 0, x)
return answer
def profit(p,q):
pi = p*q - np.exp(q) - 0.5*(q**2)
return pi
q = profitmax(5,5)
print(f'The maximised level of profit is {profit(5,q)}')
print(f'Quantity of output at profit maximum is {q}')
```
#Numba
```
#Exercise 1
import numpy as np
from numba import jit
@jit
def estimate_pi(n):
U = np.random.uniform(-1,1,n)
V = np.random.uniform(-1,1,n)
count = 0
for i in range(n):
d = np.sqrt(U[i]**2 + V[i]**2)
if d <= 1:
count += 1
pi = 4*count/n
return pi
estimate_pi(1000000)
#Exercise 2
import numpy as np
"""
denote the low state at 0 and the high state as 1
"""
state = 1
x = []
n=1000000
for i in range(n):
u = np.random.uniform()
x.append(state)
if state == 1:
if u<0.2: #probability of moving from high to low is 0.2
state = 0
else:
state = 1
else:
if u<0.1: #probability of moving from low to high is 0.1
state = 1
else:
state = 0
timeH = sum(x)/n
print(timeH)
```
# Day 2
#Writing Good Code
```
#Exercise 1
#Improving the code
import numpy as np
from scipy import optimize
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
#parameters
α = 0.1
β = 1
γ = 1
δ = 1
lb, ub = 2,4
#defining the excess demand
def excess_demand(p):
h = γ*(p**(-δ)) - np.exp(α*p) + β
return h
#solving numerically for the equilibrium price and quantity
price_eq = sc.optimize.brentq(excess_demand, lb,ub)
quantity_eq = γ*(price_eq**(-δ))
grid = np.linspace(2, 4, 100)
print(f'Equilibrium price is {price_eq: .2f}')
print(f'Equilibrium quantity is {quantity_eq: .2f}')
#plotting the demand and supply curves
price = np.linspace(lb,ub,100)
qd = γ*(price**(-δ))
qs = np.exp(α*price) - β
fig, ax = plt.subplots()
ax.plot(qd, grid, 'b-', lw=2, label='Demand')
ax.plot(qs, grid, 'g-', lw=2, label='Supply')
fig.set_size_inches(10.5, 7.5)
ax.set_xlabel('Quantity',fontsize=20)
ax.set_ylabel('Price',fontsize=20)
ax.legend(loc='upper right')
ax.set_title('Market Diagram',fontsize=20)
plt.axhline(y=price_eq, color='r', linestyle='--')
plt.axvline(x=quantity_eq, color='r', linestyle='--')
plt.show()
```
#Parallelisation
```
#Exercise 1
"""
To estimate π I draw a circle of radius one inside a square with side length 2. The ratio of areas of the circle
to the square is π/4. Therefore, by 'throwing' dots at the 2x2 grid centred at 0, we can estimate π by the
proportion of dots that lie inside the circle.
"""
import numpy as np
from numba import prange
n = 1_000_000
count = 0
for i in prange(n):
u,v = np.random.uniform(-1,1),np.random.uniform(-1,1)
d = np.sqrt(u**2 + v**2)
if d<=1:
count += 1
pi = 4*count/n
print(f'The estimated value of π is {pi}')
```
#AR1 Processes
```
#Exercise 1
import numpy as np
mu = 3
sigma = 0.6
m = 10_000_000
moment = 10
def doublefactorial(n):
if n <= 0:
return 1
else:
return n * doublefactorial(n-2)
X = mu + sigma*np.random.randn(m)
Xbar = sum(X)/len(X)
k = np.arange(1,moment+1,1)
M = np.empty(len(k))
error = np.empty(len(k))
for i in range(len(k)):
if k[i]%2 != 0:
k[i] = 0
M[i] = (sigma**k[i])*doublefactorial(k[i]-1)
error[i] = sum((X-Xbar)**k[i])/len(X) - M[i]
print(f'The error in approximation for the {i+1}th moment, {M[i]}, is {error[i]}')
#Exercise 2
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
def kerneldensity(a,b,n=500):
X = sc.stats.beta.rvs(a, b, size=n)
grid_size = 1000
x = np.linspace(min(X)-np.std(X),max(X)+np.std(X),grid_size)
#1.06 rule of thumb
h = ((4/3)**1.5)*(n**-0.2)*np.std(X)
non_parametric = np.empty(grid_size)
for i, numbers in enumerate(x):
n_p = (1/(n*h))*sum(sc.stats.norm.pdf((X-numbers)/h))
non_parametric[i] = n_p
fig, ax = plt.subplots()
ax.plot(x,non_parametric)
ax.plot(x,sc.stats.beta.pdf(x,a,b))
ax.set_xlabel('x')
ax.set_ylabel('KDE(x),f(x)')
ax.set_title(f'Kerndel Density Estimate of Beta Distribution with Parameters {a,b}')
plt.show()
α = [2,5,0.5]
β = [2,5,0.5]
for i in range(len(α)):
kerneldensity(α[i],β[i])
#Exercise 3
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
n=2000
a = 0.9
b= 0
c = 0.1
mu = -3
s = 0.2
W = np.random.randn(n)
X = mu + s*np.random.randn(n)
Y = a*mu+b + np.sqrt((a**2)*(s**2)+(c**2))*np.random.randn(n)
X_t = a*X + b + c*W
#kernel density estimation
grid_size = n
x = np.linspace(min(X_t)-np.std(X_t),max(X_t)+np.std(X_t),grid_size)
#1.06 rule of thumb
h = ((4/3)**1.5)*(n**-0.2)*np.std(X_t)
non_parametric = np.empty(grid_size)
for i, numbers in enumerate(x):
n_p = (1/(n*h))*sum(sc.stats.norm.pdf((X_t-numbers)/h))
non_parametric[i] = n_p
fig, ax = plt.subplots()
ax.plot(x,non_parametric)
ax.plot(x,sc.stats.norm.pdf(x,a*mu+b,(a**2)*(s**2)+(c**2)))
ax.set_xlabel('x')
ax.set_ylabel('KDE(x),f(x)')
ax.set_title(f'Kerndel Density Estimate of One Step Ahead of AR(1) with N({a*mu+b,(a**2)*(s**2)+(c**2)})')
ax.legend(['Kernel Density Estimate','True Distribution'])
plt.show()
```
#Heavy Tails
```
#Exercise 1
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
np.random.seed(11)
n = 120
norm_draws_s2 = 2*np.random.randn(n)
norm_draws_s12 = 12*np.random.randn(n)
cauchy_draws = np.random.standard_cauchy(size=n)
fig, ax = plt.subplots()
plt.subplot(3,1,1)
plt.scatter(range(n),norm_draws_s2)
plt.subplot(3,1,2)
plt.scatter(range(n),norm_draws_s12)
plt.subplot(3,1,3)
plt.scatter(range(n),cauchy_draws)
#Exercise 3
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
np.random.seed(11)
α = [1.15,1.5,1.75]
n=120
def plotter(a):
X = np.random.pareto(a,n)
fig,ax = plt.subplots()
ax.plot(X)
for a in α:
plotter(a)
```
#Scalar Dynamics
```
#Exercise 1
"""
For the difference equation x_{t+1} = ax_t + b study the dynamics over time series length
T for a variety of a \in (-1,1), b=1
"""
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
b=1
a_pos = np.arange(0.1,1,0.1)
a_neg = np.arange(-0.9,0,0.1)
T = 20
x = np.random.randn()
y = np.random.randn()
X = np.empty((int(len(a_pos)),T))
Y = np.empty((int(len(a_neg)),T))
for a in range(len(a_pos)):
for t in range(T):
x = a_pos[a]*x+b
y = a_neg[a]*y+b
X[a,t] = x
Y[a,t] = y
fig
fig = plt.figure()
plt.subplot(2, 1, 1)
fig.set_size_inches(18.5, 10.5)
for a in range(len(a_pos)):
plt.plot(range(T),X[a,:])
plt.subplot(2, 1, 2)
fig.set_size_inches(18.5, 10.5)
for a in range(len(a_neg)):
plt.plot(range(T),Y[a,:])
plt.show()
```
# Day 3
#Kesten Processes
```
#Exercise 1
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
%matplotlib inline
def garch(time, σ_0 = 0, α_0 = 0.00001, α_1 = 0.1, β = 0.9):
stock_returns = []
sigma = np.empty(time)
sigma[0] = σ_0
for t in range(time-1):
ε = np.random.randn()
sigma[t+1] = α_0 + sigma[t]*(α_1*ε + β)
ϵ = np.random.randn()
r = np.sqrt(sigma[t+1])*ϵ
stock_returns.append(r)
return stock_returns
time = 15*250
returns = garch(time)
###---------------------Comparison to Nasdaq Composite Index
import yfinance as yf
import pandas as pd
s = yf.download('^IXIC', '2006-1-1', '2019-11-1')['Adj Close']
r = s.pct_change()
fig, axes = plt.subplots(2, 1, figsize=(11, 6))
ax = axes[0]
ax.plot(returns,alpha=0.7)
ax.set_ylabel('returns', fontsize=12)
ax.set_xlabel('date', fontsize=12)
axes[1].plot(r,alpha=0.7)
axes[1].set_ylabel('returns', fontsize=12)
axes[1].set_xlabel('date', fontsize=12)
plt.show()
#Exercise 4
μ_a = -0.5 # location parameter for a
σ_a = 0.1 # scale parameter for a
μ_b = 0.0 # location parameter for b
σ_b = 0.5 # scale parameter for b
μ_e = 0.0 # location parameter for e
σ_e = 0.5 # scale parameter for e
s_bar = 1.0 # threshold
s_init = 1.0 # initial condition for each firm
import numpy as np
from numba import prange, jit
import matplotlib.pyplot as plt
%matplotlib inline
@jit
def firm_dynamics(T,M):
firm = np.empty((M,T))
firm[:,0] = s_init
for m in prange(M):
for t in range(T-1):
if firm[m,t] < s_bar:
e = np.exp(μ_e + σ_e*np.random.randn())
firm[m,t+1] = e
else:
a = np.exp(μ_a + σ_a*np.random.randn())
b = np.exp(μ_b + σ_b*np.random.randn())
firm[m,t+1] = a*firm[m,t] + b
return firm
T = 500 # sampling date
M = 10_000_000 # number of firms
firm = firm_dynamics(T,M)
size = 1000 #size of the size rank plot
log_size = np.log(sorted(firm[0:size,-1],reverse = True))
log_rank = np.log(range(1,size+1))
fig, ax = plt.subplots()
ax.plot(log_size,log_rank)
ax.set(xlabel='log size', ylabel='log rank')
plt.show()
```
# Day 4
#Short Path
```
#Exercise 1
num_nodes = 100
destination_node = 99
def map_graph_to_distance_matrix(in_file):
# First let's set of the distance matrix Q with inf everywhere
Q = np.ones((num_nodes, num_nodes))
Q = Q * np.inf
# Now we read in the data and modify Q
infile = open(in_file)
for line in infile:
elements = line.split(',')
node = elements.pop(0)
node = int(node[4:]) # convert node description to integer
if node != destination_node:
for element in elements:
destination, cost = element.split()
destination = int(destination[4:])
Q[node, destination] = float(cost)
Q[destination_node, destination_node] = 0
infile.close()
return Q
Q = map_graph_to_distance_matrix('graph.txt')
print(Q)
num_nodes = len(Q)
J = np.zeros(num_nodes, dtype=np.float64) # Initial guess
next_J = np.empty(num_nodes, dtype=np.float64) # Stores updated guess
max_iter = 500
i = 0
while i < max_iter:
for v in range(num_nodes):
next_J[v] = np.min(Q[v, :] + J)
if np.equal(next_J, J).all():
break
else:
J[:] = next_J # Copy contents of next_J to J
i += 1
print("The cost-to-go function is", J)
path = [0]
number = 0
while number != 99:
k = Q[number,:] + J
number = k.argmin()
path.append(number)
print(path)
```
#McCall Model
```
#Exercise 1
import numpy as np
from numba import jit, jitclass, float64
import matplotlib.pyplot as plt
%matplotlib inline
import quantecon as qe
from quantecon.distributions import BetaBinomial
mcm = McCallModel()
@jit
def invcdf(q):
prob = np.array(q)
p = np.cumsum(prob)
u = np.random.uniform()
a = p.searchsorted(u)
return a
from numba import jit, njit, prange
c_vals = np.linspace(10,40,25)
wages_c = []
for C_value in c_vals:
mcm.c = C_value
wage = compute_reservation_wage_two(mcm)
wages_c.append(wage)
print(wages_c)
number_agents = 20000
time = np.empty((20000,len(wages_c)))
n,a,b = 50, 200, 100
x = []
for i in range(50):
e = BetaBinomial(n,a,b).pdf()[i]
x.append(e)
@jit
def unemployment(reserve_wage,number_agents=20000):
agents = np.zeros(number_agents)
for i in prange(len(agents)):
w = invcdf(x) + 10
while w < reserve_wage:
w = invcdf(x) + 10
agents[i] += 1
return agents
average_length = []
for reserve_wage in wages_c:
unemployment_length = np.mean(unemployment(reserve_wage))
average_length.append(unemployment_length)
print(f'The average unemployment length for reservation wage {reserve_wage} is {unemployment_length} days')
plt.plot(c_vals,average_length)
plt.xlabel('c values')
plt.ylabel('average length of unemployment')
plt.show()
#Exercise 2
#Exercise 2
import numpy as np
from numba import jit, jitclass, float64
import matplotlib.pyplot as plt
%matplotlib inline
import quantecon as qe
from quantecon.distributions import BetaBinomial
mccall_data = [
('c', float64), # unemployment compensation
('β', float64), # discount factor
('w', float64[:]), # array of wage values, w[i] = wage at state i
('q', float64[:]) # array of probabilities
]
@jitclass(mccall_data)
class McCallModel:
def __init__(self, c=25, β=0.99, w=w_default, q=q_default):
self.c, self.β = c, β
self.w, self.q = w_default, q_default
def bellman(self, i, v):
"""
The r.h.s. of the Bellman equation at state i.
"""
# Simplify names
c, β, w, q = self.c, self.β, self.w, self.q
# Evaluate right hand side of Bellman equation
X = np.exp(μ + σ*np.random.randn(S))
max_value = max(w[i] / (1 - β), c + β * np.sum(v * q))
return(max_value)
mcm = McCallModel()
def compute_reservation_wage_two(mcm, max_iter=500, tol=1e-5, σ = 0.5, μ = 2.5): #tol = tolerance
# Simplify names
c, β, w, q = mcm.c, mcm.β, mcm.w, mcm.q
# == First compute h == #
h = np.sum(w * q) / (1 - β)
i = 0
error = tol + 1
S = 10_000
w = np.exp(μ + σ*np.random.randn(S))
simul = []
while i < max_iter and error > tol:
for wage in w:
s = np.maximum(wage/(1-β),h)
simul.append(s)
h_next = c + β * np.mean(simul)
error = np.abs(h_next - h)
i += 1
h = h_next
# == Now compute the reservation wage == #
return (1 - β) * h
compute_reservation_wage_two(mcm)
```
| github_jupyter |
# NEXUS tool: case study for the NWSAS basin - irrigation water demand
In this notebook a case study for the NWSAS basin is covered, using the `nexus_tool` package. The water requirements for agricultural irrigation are calculated, then the energy requirements for pumping and desalination of brackish water are estimated and then least-cost options to supply such energy are identified between selected technologies.
First import the package by running the following block:
```
%load_ext autoreload
%autoreload
import nexus_tool
```
After importing all required packages, the input GIS data is loaded into the variable `df`. Change the `file_path` variable to reflect the name and relative location of your data file.
```
file_path = r'../nwsas_1km_pre_water_model.gz'
df = nexus_tool.read_csv(file_path)
```
## 1. Calculating irrigation water demand
To be able to calculate the water demand for agricultural irrigation, it is required to define crop irrigation calendars for each crop type to be assessed. Then an excel file containing the information of the crop calendars is needed. Such file should look something like this:
|crop|init_start|init_end|dev_start|dev_end|mid_start|mid_end|late_start|late_end|
|:---|:---:|:---:|:---:|:---:|:---:|:---:|:---:|:---:|
|dates|01/11|30/03|31/03|04/05|05/05|30/09|01/10|31/10|
|vegetables|01/11|25/11|26/11|31/12|01/01|07/02|08/02|28/02|
|olives|01/03|30/03|31/06|30/06|01/07|31/08|01/09|30/11|
Change the `file_path` variable to reflect the name and relative location of your data file.
```
file_path = r'NWSAS_crop_calendar.xlsx'
crop_calendar = nexus_tool.read_excel(file_path)
```
### 1.1. Creating the model
Once all input data is loaded. To create a model simply create an instance of the `nexus_tool.Model()` class and store it in a variable name. The `nexus_tool.Model()` class requires a dataframe as input data and another dataframe as crop calendar data. Several other properties and parameter values can be defined by explicitly passing values to them. To see a full list of parameters and their explaination refer to the documentation of the package.
```
nwsas = nexus_tool.Model(df, crop_calendar = crop_calendar,
pumping_hours_per_day=10, deff= 1, aeff= 1)
```
After creating the model you can see the default values of the properties by running `nwsas.print_properties()`. Moreover, to define values or property names after creating the model, each property can be called individually and its value can be overwrited as:
```python
nwsas.eto = "ETo_"
nwsas.pumping_hours_per_day = 10
```
```
nwsas.print_properties()
nwsas.prec = 'prec_'
nwsas.wind = 'wind_'
nwsas.srad = 'srad_'
nwsas.tmin = 'tmin_'
nwsas.tmax = 'tmax_'
nwsas.tavg = 'tavg_'
nwsas.gw_depth = 'GroundwaterDepth'
nwsas.crop_area = 'IrrigatedArea'
```
### 1.2. Setting required model parameters
To compute the irrigation water requierements, the share of cropland needs to be defined for each data point. That is, to specify the share each croptype has within each data point. To achieve this, first create a dictionary containing all the croplands of the region and assign a share for each. This share should be the default value that most of the data points should have. Specific values for different regions can also be defined, as explined later:
```python
crop_dic = {'crop1':0.5,'crop2':0.5,'crop3':0,...}
```
Then, use the `.set_cropland_share()` method to pass this dictionary to the model like:
```python
nwsas.set_cropland_share(crop_dic, inplace = True)
```
The option `inplace = True` is used to tell the model to store the dictionary in it.
Moreover, to define different cropland share values from the default one, a new dictionary can be passed to specific provinces, cities or regions, by passing a `geo_boundary` and a `boundary_name` for the region in question:
```python
nwsas.set_cropland_share({'crop1':0.7,'crop2':0.3,'crop3':0,...},
geo_boundary = 'province',
boundary_name = ['province name'], inplace = True)
```
The `geo_boundary` value needs to match an existent variable in the input dataframe and the `boundary_name` value should exist within the `geo_boundary` column.
```
crop_dic = {'dates': 0.5, 'vegetables': 0.50}
nwsas.set_cropland_share(crop_dic, inplace=True)
# nwsas.set_cropland_share({'dates': 0.65, 'vegetables': 0.35, 'herbaceous': 0.0},
# geo_boundary='Province',
# boundary_name=['El Oued'],
# inplace = True)
# nwsas.set_cropland_share({'dates': 0.54, 'vegetables': 0.02, 'herbaceous': 0.44},
# geo_boundary='Province',
# boundary_name=['Ouargla'],
# inplace = True)
```
### 1.3. Setting the ky and kc values
The yield responese factor (*ky*), is a coefficient that relates the water uses by a crop throughout the different growing seassons. A definition by the [FAO Irrigation and Drainage Paper](http://www.fao.org/3/i2800e/i2800e.pdf) is a follows:
>The yield response factor (Ky) captures the essence of the complex linkages
between production and water use by a crop, where many biological,
physical and chemical processes are involved.
The Ky values are crop specific as:
>**Ky > 1**: crop response is very sensitive to water deficit with proportional larger yield reductions
when water use is reduced because of stress.
**Ky < 1**: crop is more tolerant to water deficit, and recovers partially from stress, exhibiting less than proportional reductions in yield with reduced water use.
**Ky = 1**: yield reduction is directly proportional to reduced water use.
The crop coefficient (*kc*) is a factor that relates the water requirements of a cropland during a specific growing seasson. A definition by the [FAO Irrigation and drainage paper 56](http://www.fao.org/3/x0490e/x0490e0a.htm) goes as follows:
>The coefficient integrates differences in the soil evaporation and crop transpiration rate between the crop and the grass reference surface. As soil evaporation may fluctuate daily as a result of rainfall or irrigation, the single crop coefficient expresses only the time-averaged (multi-day) effects of crop evapotranspiration.
To define the *ky* values, a dictionary containing the values for each crop type evaluated in the region needs to be passed to the `.ky_dict` parameter of the model. Similarly the *kc* values are passed to the `.kc_dict` parameter as a dictionary containing a list of values for each croptype (i.e. one for each season in order, i.e initial, development, mid and late season).
```
nwsas.ky_dict = {'dates':0.5,
'vegetables':1.1,
# 'herbaceous':1.1
}
nwsas.kc_dict = {'dates': [0.8,0.9,1,0.8],
'vegetables':[0.5,1,1,0.8],
# 'herbaceous':[0.3,1.15,1.15,0.4]
}
```
### 1.4. Calculating the reference evapotranspiration
To calculate the reference evapotranspiration, make sure you have the correct definitions por all the properties in the model (check them with `nwsas.print_properties()`) and the correct input values check them with (check them with `nwsas.print_inputs()`). Then, run the `nwsas.get_eto(inplace = True)` method.
```
nwsas.get_eto(inplace = True)
```
### 1.5. Calculating the effective rainfall
The effective rainfall stands for the actuall usable water that is stored in the root zone of the plant. Then, it substract all runoff, evapotranspiration and water that is percolated deeper in the soil and can not be reached by the plant. There are several methods available to compute the effective rainfall, depending on the soil type, climatic region, among other parameters. The one used by the `nexus_tool` package is the **(reference here)**.
Get the effective rainfall for al the region by running the method `nwsas.get_effective_rainfall(inplace = True)`.
```
nwsas.get_effective_rainfall(inplace=True)
```
### 1.6. Calculating the kc values and standard evapotranspiration
To calculate the kc values and get the standar evapotranspiration, run the methods `nwsas.get_calendar_days(inplace = True)` and `nwsas.get_kc_values(inplace = True)` in that order. The former, will map the crop calendars of the crops to every region, and compute the duration of each seasson in days. Then, the *kc* values are calculated according to the days transcurred in each seassons and the values passed in the `kc_dict` input.
```
nwsas.get_calendar_days(inplace=True)
nwsas.get_kc_values(inplace=True)
```
### 1.7. Geting the irrigation water demand
Then everything should be setup to compute the irrigation water demand. For that run the `nwsas.get_water_demand(inplace = True)` method.
```
nwsas.get_water_demand(inplace=True)
```
## 2. Displaying and saving the results
After the calculations are completed, display a summary of results by running the `nwsas.print_summary()` method. If you run the method without any argument, then the summary values will be dispayed for the entire region, under the label of "Glogal". However, if you like to summarize by regions, then pass the argument `geo_boundary` to the function, specifing the variable that you want the results to be grouped by.
```
nwsas.print_summary(geo_boundary = ['Region'])
nwsas.df['AgWaterReq'] = nwsas.df.filter(like='SSWD').sum(axis=1) / nwsas.df['IrrigatedArea']
```
Finally, save the results in .csv format, by specifing an output file name and location (`output_file`) and running the `nwsas.df.to_csv(output_file, index = False)` method.
```
output_file = r'../nwsas_1km_input_data.gz'
nwsas.df.to_csv(output_file, index = False)
nwsas.df.groupby('Province')[['AgWaterReq']].mean().sort_values('AgWaterReq')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/HansHenseler/masdav/blob/main/Part_4_Elasticsearch_and_log2timeline.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Elasticsearch and log2timeline
Exercise 4:
Master of Advanced Studies in Digital Forensics & Cyber Investigation
Data Analytics and Visualization for Digital Forensics
(c) Hans Henseler, 2021
## 1 Installing plaso tools in the colab notebook
First install Plaso-tools as we did in exercise 3
```
# various install steps to install plaso tools and dependencies to get plaso working in colab
# -y option is to skip user interaction
# some packages need to be deinstalled and reinstalled to resolve dependencies
# these steps take app. 3 minutes to complete on a fresh colab instance
!add-apt-repository -y ppa:gift/stable
!apt update
!apt-get update
!apt install plaso-tools
!pip uninstall -y pytsk3
!pip install pytsk3
!pip uninstall -y yara-python
!pip install yara-python
!pip uninstall -y lz4
!pip install lz4
# check is plasoo tools were installed by running psort.py
!psort.py -h
```
## 2 Download and setup the Elasticsearch instance
For demo purposes, the open-source version of the elasticsearch package is used.
```
%%bash
wget -q https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-oss-7.9.2-linux-x86_64.tar.gz
wget -q https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-oss-7.9.2-linux-x86_64.tar.gz.sha512
tar -xzf elasticsearch-oss-7.9.2-linux-x86_64.tar.gz
sudo chown -R daemon:daemon elasticsearch-7.9.2/
shasum -a 512 -c elasticsearch-oss-7.9.2-linux-x86_64.tar.gz.sha512
```
Run Elasticsearch as a daemon process
```
import time
%%bash --bg
sudo -H -u daemon elasticsearch-7.9.2/bin/elasticsearch
# Sleep for few seconds to let the instance start.
time.sleep(20)
```
Once the instance has been started, grep for elasticsearch in the processes list to confirm the availability.
```
%%bash
ps -ef | grep elasticsearch
```
query the base endpoint to retrieve information about the cluster.
```
%%bash
curl -sX GET "localhost:9200/"
# This command created an index in Elasticsearch
#
# this is a useful page to look up Elasticsearch REST api calls :
# https://www.elastic.co/guide/en/elasticsearch/reference/6.8/cat-indices.html
#
!curl -X GET "localhost:9200/_cat/indices?format=json&pretty"
```
## 3 Use Log2timeline.py and Psort.py to load data in Elasticsearch
```
from google.colab import drive
drive.mount('/content/gdrive')
# In part 3 (step 3) we stored the mus2019ctf.plaso file in your drive.
#
plaso_file = 'gdrive/MyDrive/mus2019ctf.plaso'
#
# and check if it's there
#
!ls -l $plaso_file
# If it's not there you can create it by repeating the following steps
#
# The complete mus2019ctf.plaso file is 450MB and takes a while. After you have created it
# it makes sense to store it in your gdrive so you can reuse it:
plaso_file = 'gdrive/MyDrive/Colab\ Notebooks/Data\ Analytics\ and\ Visualisation\ Course/mus2019ctf.plaso'
#
# if not you need to create it with log2timeline.py using the complete windows_filter.txt filter
#
# add a shortcut in your Google drive to this shared drive https://drive.google.com/drive/folders/1KUlZUl4Sy2JzgbuRW-oHjIGFClY2bl75?usp=sharing
# then mount you google drive in this colab (you need to authorize this colab to access your google drive)
#
# disk_image = "/content/gdrive/MyDrive/Images/MUS-CTF-19-DESKTOP-001.E01"
# plaso_gdrive_folder = 'gdrive'
# !wget "https://raw.githubusercontent.com/mark-hallman/plaso_filters/master/filter_windows.txt"
# !log2timeline.py -f filter_windows.txt mus2019ctf.plaso $disk_image
# !cp mus2019ctf.plaso $plaso_gdrive_folder
# plaso_file = 'gdrive/MyDrive/mus2019ctf.plaso'
# !ls -l $plaso_file
```
Use psort to write events to Elasticsearch that we setup earlier. We can use the elastic output format
```
# Before we do that, let's take a look at the elasticsearch.mappings file that comes with plaso
# actually there is more in that folder that you may be interested in
#
!ls /usr/share/plaso
# let's take a look at the elasticsearch.mappings
#
!cat /usr/share/plaso/elasticsearch.mappings
# run psort.py. It takes about 10 minutes to export all rows from the 430MB plaso file
#
!psort.py -o elastic --server localhost --port 9200 --elastic_mappings /usr/share/plaso/elasticsearch.mappings --index_name mus2019ctf $plaso_file --status_view none
# Let's take a look again at the indices in our Elasticsearch instance
#
!curl -X GET "localhost:9200/_cat/indices?format=json&pretty"
# we can also see what fields were mapped in this index by Psort.py
#
!curl -X GET "localhost:9200/mus2019ctf/_mapping?format=json&pretty"
```
## 4 Accessing Elasticsearch via the REST API
```
!curl -sX GET "localhost:9200/_search?format=json&pretty"
!curl -sX GET "localhost:9200/mus2019ctf/_settings?format=json&pretty"
!curl -sX GET "localhost:9200/mus2019ctf/_mapping?format=json&pretty"
```
## 5 Accessing the Elasticsearch API in Python
```
# So far we have been accessing information directly with curl from the Elasticsearch REST API
# The is also an Elasticsearch Python API that we can use
#
from elasticsearch import Elasticsearch
es = Elasticsearch([{'host': 'localhost', 'port': 9200}])
```
Which indexes are available?
```
es.indices.get_alias("*")
```
Search the index with a full-text query
```
# https://www.elastic.co/guide/en/elasticsearch/reference/current/query-dsl-match-query.html
response = es.search(index="mus2019ctf", body={"query": {"match": {"message": { "query": "selmabouvier" }}}}, size=5)
elastic_docs = response['hits']['hits']
elastic_docs
for num, doc in enumerate(elastic_docs):
print(num, '-->', doc['_source'], "\n")
# We define a Python function to list results
#
def print_results(response):
for num, doc in enumerate(response['hits']['hits']):
print(num, '-->', doc['_source'], "\n")
# we can try this function on the response we got earlier
#
print_results(response)
# Elasticsearch query syntax is quite elaborate. We will provide some examples in this colab
# For a complete overview see the Elasticsearch reference documents
#
# https://www.elastic.co/guide/en/elasticsearch/reference/current/query-dsl-query-string-query.html#query-string-syntax
#
query = '{"query": { "query_string": {"query": "source_short: WEBHIST" }}}'
query = '{"query": { "query_string": {"query": "data_type: windows*link" }}}'
query = '{"query": { "query_string": {"query": "drive_type: 3" }}}'
query = '{"query": { "query_string": {"query": "drive_type: 3 AND data_type: windows*link" }}}'
query = '{"query": { "range": {"drive_type": { "gte":0 , "lte":2 } }}}'
query = '{"query": { "range": {"drive_type": { "gte":1 , "lte":3 } }}}'
query = '{"query": { "query_string": {"query": "drive_type:>=0 and drive_type:<2" }}}'
query = '{"query": { "query_string": {"query": "file_size:>=10000 and file_size:<100000" }}}'
query = '{"query": { "query_string": {"query": "data_type: msie\\\\:*" }}}'
response = es.search(index="mus2019ctf", body=query, size=15)
print_results(response)
```
## 6 Elasticsearch field aggregation
```
# First we define some helper functions:
def print_facets(agg_dict):
sum=0
for field, val in agg_dict:
print("facets of field ", field,':')
for bucket in val['buckets']:
for key in bucket:
if key=='key':
print('\t',bucket[key],end='=')
else:
print(bucket[key],end='')
sum = sum + bucket[key]
print()
print("total number of hits for ",field," is ",sum)
def print_hit_stats(response):
print('hit stats:')
for key, val in response['hits'].items():
print(key, val)
print('\n')
querystring = '{ "query_string": {"query": "source_short: WEBHIST" }}'
query = '{"query": %s}' % querystring
print(query)
# Aggregating results is one of the most powerful options in Elasticsearch
#
# https://www.elastic.co/guide/en/elasticsearch/reference/current/search-aggregations.html
#
querystring = '{ "query_string": {"query": "SelmaBouvier" }}'
facets = '"aggs": { "data_type": { "terms": { "field": "data_type.keyword"}}}'
query = '{"query": %s,%s}' % (querystring,facets)
print(query)
response = es.search(index="mus2019ctf", body=query, size=0)
print_hit_stats(response)
print_facets(response['aggregations'].items())
# print_results(response)
# Aggregate accross multiple facets
#
querystring = '{ "query_string": {"query": "SelmaBouvier" }}'
facets = '"aggs": { "parser": { "terms": { "field": "parser.keyword"}}, "data_type": { "terms": { "field": "data_type.keyword"}}}'
query = '{"query": %s,%s}' % (querystring,facets)
response = es.search(index="mus2019ctf", body=query, size=0)
print_facets(response['aggregations'].items())
# date range search
query = '{"query": { "query_string": {"query": "datetime:[2019-03-12 TO 2019-03-22]" }}}'
print(query)
response = es.search(index="mus2019ctf", body=query, size=10)
print_results(response)
```
## 7 Putting Elasticsearch json out into a Pandas dataframe
```
# the out is json format which we can store in a pandas dataframe
import pandas as pd
import json
from io import StringIO
output = !curl -sX GET "localhost:9200/_search?q=logon"
df = pd.read_json(StringIO(output[0]))
df.head
df['hits']['hits'][:1]
```
# Exercises
## 1 Use elasticsearch to filter events in between 2019-03-12 and 2019-03-22
```
# Your answer
```
## 2 Write a query that performs an aggregation on source_long and source_short (can you find the right field names?)
```
# Your answer
```
## 3 Combine your date range filter from exercise 1 with facet aggregation in exercise 2
```
# Your answer
```
## 4 ***Advanced*** Create an aggregation accross 3 fields and visualise them in a treemap or sunburst plot
```
# Your answer
```
| github_jupyter |
<a href="https://www.skills.network/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01"><img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DL0120ENedX/labs/Template%20for%20Instructional%20Hands-on%20Labs/images/IDSNlogo.png" width="400px" align="center"></a>
<h1 align="center"><font size="5">RECURRENT NETWORKS and LSTM IN DEEP LEARNING</font></h1>
<h2>Applying Recurrent Neural Networks/LSTM for Language Modeling</h2>
Hello and welcome to this part. In this notebook, we will go over the topic of Language Modelling, and create a Recurrent Neural Network model based on the Long Short-Term Memory unit to train and benchmark on the Penn Treebank dataset. By the end of this notebook, you should be able to understand how TensorFlow builds and executes a RNN model for Language Modelling.
<h2>The Objective</h2>
By now, you should have an understanding of how Recurrent Networks work -- a specialized model to process sequential data by keeping track of the "state" or context. In this notebook, we go over a TensorFlow code snippet for creating a model focused on <b>Language Modelling</b> -- a very relevant task that is the cornerstone of many different linguistic problems such as <b>Speech Recognition, Machine Translation and Image Captioning</b>. For this, we will be using the Penn Treebank dataset, which is an often-used dataset for benchmarking Language Modelling models.
<h2>Table of Contents</h2>
<ol>
<li><a href="https://#language_modelling">What exactly is Language Modelling?</a></li>
<li><a href="https://#treebank_dataset">The Penn Treebank dataset</a></li>
<li><a href="https://#word_embedding">Word Embedding</a></li>
<li><a href="https://#building_lstm_model">Building the LSTM model for Language Modeling</a></li>
<li><a href="https://#ltsm">LTSM</a></li>
</ol>
<p></p>
</div>
<br>
***
<a id="language_modelling"></a>
<h2>What exactly is Language Modelling?</h2>
Language Modelling, to put it simply, <b>is the task of assigning probabilities to sequences of words</b>. This means that, given a context of one or a sequence of words in the language the model was trained on, the model should provide the next most probable words or sequence of words that follows from the given sequence of words the sentence. Language Modelling is one of the most important tasks in Natural Language Processing.
<img src="https://ibm.box.com/shared/static/1d1i5gub6wljby2vani2vzxp0xsph702.png" width="1080">
<center><i>Example of a sentence being predicted</i></center>
<br><br>
In this example, one can see the predictions for the next word of a sentence, given the context "This is an". As you can see, this boils down to a sequential data analysis task -- you are given a word or a sequence of words (the input data), and, given the context (the state), you need to find out what is the next word (the prediction). This kind of analysis is very important for language-related tasks such as <b>Speech Recognition, Machine Translation, Image Captioning, Text Correction</b> and many other very relevant problems.
<img src="https://ibm.box.com/shared/static/az39idf9ipfdpc5ugifpgxnydelhyf3i.png" width="1080">
<center><i>The above example is a schema of an RNN in execution</i></center>
<br><br>
As the above image shows, Recurrent Network models fit this problem like a glove. Alongside LSTM and its capacity to maintain the model's state for over one thousand time steps, we have all the tools we need to undertake this problem. The goal for this notebook is to create a model that can reach <b>low levels of perplexity</b> on our desired dataset.
For Language Modelling problems, <b>perplexity</b> is the way to gauge efficiency. Perplexity is simply a measure of how well a probabilistic model is able to predict its sample. A higher-level way to explain this would be saying that <b>low perplexity means a higher degree of trust in the predictions the model makes</b>. Therefore, the lower perplexity is, the better.
<a id="treebank_dataset"></a>
<h2>The Penn Treebank dataset</h2>
Historically, datasets big enough for Natural Language Processing are hard to come by. This is in part due to the necessity of the sentences to be broken down and tagged with a certain degree of correctness -- or else the models trained on it won't be able to be correct at all. This means that we need a <b>large amount of data, annotated by or at least corrected by humans</b>. This is, of course, not an easy task at all.
The Penn Treebank, or PTB for short, is a dataset maintained by the University of Pennsylvania. It is <i>huge</i> -- there are over <b>four million and eight hundred thousand</b> annotated words in it, all corrected by humans. It is composed of many different sources, from abstracts of Department of Energy papers to texts from the Library of America. Since it is verifiably correct and of such a huge size, the Penn Treebank is commonly used as a benchmark dataset for Language Modelling.
The dataset is divided in different kinds of annotations, such as Piece-of-Speech, Syntactic and Semantic skeletons. For this example, we will simply use a sample of clean, non-annotated words (with the exception of one tag --<code>\<unk></code>
, which is used for rare words such as uncommon proper nouns) for our model. This means that we just want to predict what the next words would be, not what they mean in context or their classes on a given sentence.
<center>Example of text from the dataset we are going to use, <b>ptb.train</b></center>
<br><br>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<center>the percentage of lung cancer deaths among the workers at the west <code><unk></code> mass. paper factory appears to be the highest for any asbestos workers studied in western industrialized countries he said
the plant which is owned by <code><unk></code> & <code><unk></code> co. was under contract with <code><unk></code> to make the cigarette filters
the finding probably will support those who argue that the U.S. should regulate the class of asbestos including <code><unk></code> more <code><unk></code> than the common kind of asbestos <code><unk></code> found in most schools and other buildings dr. <code><unk></code> said</center>
</div>
<a id="word_embedding"></a>
<h2>Word Embeddings</h2><br/>
For better processing, in this example, we will make use of <a href="https://www.tensorflow.org/tutorials/word2vec/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01"><b>word embeddings</b></a>, which is <b>a way of representing sentence structures or words as n-dimensional vectors (where n is a reasonably high number, such as 200 or 500) of real numbers</b>. Basically, we will assign each word a randomly-initialized vector, and input those into the network to be processed. After a number of iterations, these vectors are expected to assume values that help the network to correctly predict what it needs to -- in our case, the probable next word in the sentence. This is shown to be a very effective task in Natural Language Processing, and is a commonplace practice. <br><br> <font size="4"><strong>
$$Vec("Example") = \[0.02, 0.00, 0.00, 0.92, 0.30, \ldots]$$ </strong></font> <br>
Word Embedding tends to group up similarly used words <i>reasonably</i> close together in the vectorial space. For example, if we use T-SNE (a dimensional reduction visualization algorithm) to flatten the dimensions of our vectors into a 2-dimensional space and plot these words in a 2-dimensional space, we might see something like this:
<img src="https://ibm.box.com/shared/static/bqhc5dg879gcoabzhxra1w8rkg3od1cu.png" width="800">
<center><i>T-SNE Mockup with clusters marked for easier visualization</i></center>
<br><br>
As you can see, words that are frequently used together, in place of each other, or in the same places as them tend to be grouped together -- being closer together the higher they are correlated. For example, "None" is pretty semantically close to "Zero", while a phrase that uses "Italy", you could probably also fit "Germany" in it, with little damage to the sentence structure. The vectorial "closeness" for similar words like this is a great indicator of a well-built model.
<hr>
We need to import the necessary modules for our code. We need <b><code>numpy</code></b> and <b><code>tensorflow</code></b>, obviously. Additionally, we can import directly the <b><code>tensorflow\.models.rnn</code></b> model, which includes the function for building RNNs, and <b><code>tensorflow\.models.rnn.ptb.reader</code></b> which is the helper module for getting the input data from the dataset we just downloaded.
If you want to learn more take a look at <https://github.com/tensorflow/models/blob/master/tutorials/rnn/ptb/reader.py>
```
!pip install tensorflow==2.2.0rc0
!pip install numpy
import time
import numpy as np
import tensorflow as tf
if not tf.__version__ == '2.2.0-rc0':
print(tf.__version__)
raise ValueError('please upgrade to TensorFlow 2.2.0-rc0, or restart your Kernel (Kernel->Restart & Clear Output)')
```
IMPORTANT! => Please restart the kernel by clicking on "Kernel"->"Restart and Clear Outout" and wait until all output disapears. Then your changes are beeing picked up
```
!mkdir data
!mkdir data/ptb
!wget -q -O data/ptb/reader.py https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DL0120EN-SkillsNetwork/labs/Week3/data/ptb/reader.py
!cp data/ptb/reader.py .
import reader
```
<a id="building_lstm_model"></a>
<h2>Building the LSTM model for Language Modeling</h2>
Now that we know exactly what we are doing, we can start building our model using TensorFlow. The very first thing we need to do is download and extract the <code>simple-examples</code> dataset, which can be done by executing the code cell below.
```
!wget http://www.fit.vutbr.cz/~imikolov/rnnlm/simple-examples.tgz
!tar xzf simple-examples.tgz -C data/
```
Additionally, for the sake of making it easy to play around with the model's hyperparameters, we can declare them beforehand. Feel free to change these -- you will see a difference in performance each time you change those!
```
#Initial weight scale
init_scale = 0.1
#Initial learning rate
learning_rate = 1.0
#Maximum permissible norm for the gradient (For gradient clipping -- another measure against Exploding Gradients)
max_grad_norm = 5
#The number of layers in our model
num_layers = 2
#The total number of recurrence steps, also known as the number of layers when our RNN is "unfolded"
num_steps = 20
#The number of processing units (neurons) in the hidden layers
hidden_size_l1 = 256
hidden_size_l2 = 128
#The maximum number of epochs trained with the initial learning rate
max_epoch_decay_lr = 4
#The total number of epochs in training
max_epoch = 15
#The probability for keeping data in the Dropout Layer (This is an optimization, but is outside our scope for this notebook!)
#At 1, we ignore the Dropout Layer wrapping.
keep_prob = 1
#The decay for the learning rate
decay = 0.5
#The size for each batch of data
batch_size = 30
#The size of our vocabulary
vocab_size = 10000
embeding_vector_size= 200
#Training flag to separate training from testing
is_training = 1
#Data directory for our dataset
data_dir = "data/simple-examples/data/"
```
Some clarifications for LSTM architecture based on the arguments:
Network structure:
<ul>
<li>In this network, the number of LSTM cells are 2. To give the model more expressive power, we can add multiple layers of LSTMs to process the data. The output of the first layer will become the input of the second and so on.
</li>
<li>The recurrence steps is 20, that is, when our RNN is "Unfolded", the recurrence step is 20.</li>
<li>the structure is like:
<ul>
<li>200 input units -> [200x200] Weight -> 200 Hidden units (first layer) -> [200x200] Weight matrix -> 200 Hidden units (second layer) -> [200] weight Matrix -> 200 unit output</li>
</ul>
</li>
</ul>
<br>
Input layer:
<ul>
<li>The network has 200 input units.</li>
<li>Suppose each word is represented by an embedding vector of dimensionality e=200. The input layer of each cell will have 200 linear units. These e=200 linear units are connected to each of the h=200 LSTM units in the hidden layer (assuming there is only one hidden layer, though our case has 2 layers).
</li>
<li>The input shape is [batch_size, num_steps], that is [30x20]. It will turn into [30x20x200] after embedding, and then 20x[30x200]
</li>
</ul>
<br>
Hidden layer:
<ul>
<li>Each LSTM has 200 hidden units which is equivalent to the dimensionality of the embedding words and output.</li>
</ul>
<br>
There is a lot to be done and a ton of information to process at the same time, so go over this code slowly. It may seem complex at first, but if you try to apply what you just learned about language modelling to the code you see, you should be able to understand it.
This code is adapted from the <a href="https://github.com/tensorflow/models?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01">PTBModel</a> example bundled with the TensorFlow source code.
<h3>Training data</h3>
The story starts from data:
<ul>
<li>Train data is a list of words, of size 929589, represented by numbers, e.g. [9971, 9972, 9974, 9975,...]</li>
<li>We read data as mini-batch of size b=30. Assume the size of each sentence is 20 words (num_steps = 20). Then it will take $$floor(\frac{N}{b \times h})+1=1548$$ iterations for the learner to go through all sentences once. Where N is the size of the list of words, b is batch size, and h is size of each sentence. So, the number of iterators is 1548
</li>
<li>Each batch data is read from train dataset of size 600, and shape of [30x20]</li>
</ul>
```
# Reads the data and separates it into training data, validation data and testing data
raw_data = reader.ptb_raw_data(data_dir)
train_data, valid_data, test_data, vocab, word_to_id = raw_data
len(train_data)
def id_to_word(id_list):
line = []
for w in id_list:
for word, wid in word_to_id.items():
if wid == w:
line.append(word)
return line
print(id_to_word(train_data[0:100]))
```
Lets just read one mini-batch now and feed our network:
```
itera = reader.ptb_iterator(train_data, batch_size, num_steps)
first_touple = itera.__next__()
_input_data = first_touple[0]
_targets = first_touple[1]
_input_data.shape
_targets.shape
```
Lets look at 3 sentences of our input x:
```
_input_data[0:3]
print(id_to_word(_input_data[0,:]))
```
<h3>Embeddings</h3>
We have to convert the words in our dataset to vectors of numbers. The traditional approach is to use one-hot encoding method that is usually used for converting categorical values to numerical values. However, One-hot encoded vectors are high-dimensional, sparse and in a big dataset, computationally inefficient. So, we use word2vec approach. It is, in fact, a layer in our LSTM network, where the word IDs will be represented as a dense representation before feeding to the LSTM.
The embedded vectors also get updated during the training process of the deep neural network.
We create the embeddings for our input data. <b>embedding_vocab</b> is matrix of \[10000x200] for all 10000 unique words.
<b>embedding_lookup()</b> finds the embedded values for our batch of 30x20 words. It goes to each row of <code>input_data</code>, and for each word in the row/sentence, finds the correspond vector in <code>embedding_dic<code>. <br>
It creates a \[30x20x200] tensor, so, the first element of <b>inputs</b> (the first sentence), is a matrix of 20x200, which each row of it, is vector representing a word in the sentence.
```
embedding_layer = tf.keras.layers.Embedding(vocab_size, embeding_vector_size,batch_input_shape=(batch_size, num_steps),trainable=True,name="embedding_vocab")
# Define where to get the data for our embeddings from
inputs = embedding_layer(_input_data)
inputs
```
<h3>Constructing Recurrent Neural Networks</h3>
In this step, we create the stacked LSTM using <b>tf.keras.layers.StackedRNNCells</b>, which is a 2 layer LSTM network:
```
lstm_cell_l1 = tf.keras.layers.LSTMCell(hidden_size_l1)
lstm_cell_l2 = tf.keras.layers.LSTMCell(hidden_size_l2)
stacked_lstm = tf.keras.layers.StackedRNNCells([lstm_cell_l1, lstm_cell_l2])
```
<b>tf.keras.layers.RNN</b> creates a recurrent neural network using <b>stacked_lstm</b>.
The input should be a Tensor of shape: \[batch_size, max_time, embedding_vector_size], in our case it would be (30, 20, 200)
```
layer = tf.keras.layers.RNN(stacked_lstm,[batch_size, num_steps],return_state=False,stateful=True,trainable=True)
```
Also, we initialize the states of the nework:
<h4>_initial_state</h4>
For each LSTM, there are 2 state matrices, c_state and m_state. c_state and m_state represent "Memory State" and "Cell State". Each hidden layer, has a vector of size 30, which keeps the states. so, for 200 hidden units in each LSTM, we have a matrix of size \[30x200]
```
init_state = tf.Variable(tf.zeros([batch_size,embeding_vector_size]),trainable=False)
layer.inital_state = init_state
layer.inital_state
```
so, lets look at the outputs. The output of the stackedLSTM comes from 128 hidden_layer, and in each time step(=20), one of them get activated. we use the linear activation to map the 128 hidden layer to a \[30X20 matrix]
```
outputs = layer(inputs)
outputs
```
<h2>Dense layer</h2>
We now create densely-connected neural network layer that would reshape the outputs tensor from [30 x 20 x 128] to [30 x 20 x 10000].
```
dense = tf.keras.layers.Dense(vocab_size)
logits_outputs = dense(outputs)
print("shape of the output from dense layer: ", logits_outputs.shape) #(batch_size, sequence_length, vocab_size)
```
<h2>Activation layer</h2>
A softmax activation layers is also then applied to derive the probability of the output being in any of the multiclass(10000 in this case) possibilities.
```
activation = tf.keras.layers.Activation('softmax')
output_words_prob = activation(logits_outputs)
print("shape of the output from the activation layer: ", output_words_prob.shape) #(batch_size, sequence_length, vocab_size)
```
Lets look at the probability of observing words for t=0 to t=20:
```
print("The probability of observing words in t=0 to t=20", output_words_prob[0,0:num_steps])
```
<h3>Prediction</h3>
What is the word correspond to the probability output? Lets use the maximum probability:
```
np.argmax(output_words_prob[0,0:num_steps], axis=1)
```
So, what is the ground truth for the first word of first sentence? You can get it from target tensor, if you want to find the embedding vector:
```
_targets[0]
```
<h4>Objective function</h4>
How similar the predicted words are to the target words?
Now we have to define our objective function, to calculate the similarity of predicted values to ground truth, and then, penalize the model with the error. Our objective is to minimize loss function, that is, to minimize the average negative log probability of the target words:
$$\text{loss} = -\frac{1}{N}\sum\_{i=1}^{N} \ln p\_{\text{target}\_i}$$
This function is already implemented and available in TensorFlow through *tf.keras.losses.sparse_categorical_crossentropy*. It calculates the categorical cross-entropy loss for <b>logits</b> and the <b>target</b> sequence.
The arguments of this function are:
<ul>
<li>logits: List of 2D Tensors of shape [batch_size x num_decoder_symbols].</li>
<li>targets: List of 1D batch-sized int32 Tensors of the same length as logits.</li>
</ul>
```
def crossentropy(y_true, y_pred):
return tf.keras.losses.sparse_categorical_crossentropy(y_true, y_pred)
loss = crossentropy(_targets, output_words_prob)
```
Lets look at the first 10 values of loss:
```
loss[0,:10]
```
Now, we define cost as average of the losses:
```
cost = tf.reduce_sum(loss / batch_size)
cost
```
<h3>Training</h3>
To do training for our network, we have to take the following steps:
<ol>
<li>Define the optimizer.</li>
<li>Assemble layers to build model.</li>
<li>Calculate the gradients based on the loss function.</li>
<li>Apply the optimizer to the variables/gradients tuple.</li>
</ol>
<h4>1. Define Optimizer</h4>
```
# Create a variable for the learning rate
lr = tf.Variable(0.0, trainable=False)
optimizer = tf.keras.optimizers.SGD(lr=lr, clipnorm=max_grad_norm)
```
<h4>2. Assemble layers to build model.</h4>
```
model = tf.keras.Sequential()
model.add(embedding_layer)
model.add(layer)
model.add(dense)
model.add(activation)
model.compile(loss=crossentropy, optimizer=optimizer)
model.summary()
```
<h4>2. Trainable Variables</h4>
Defining a variable, if you passed <i>trainable=True</i>, the variable constructor automatically adds new variables to the graph collection <b>GraphKeys.TRAINABLE_VARIABLES</b>. Now, using <i>tf.trainable_variables()</i> you can get all variables created with <b>trainable=True</b>.
```
# Get all TensorFlow variables marked as "trainable" (i.e. all of them except _lr, which we just created)
tvars = model.trainable_variables
```
Note: we can find the name and scope of all variables:
```
[v.name for v in tvars]
```
<h4>3. Calculate the gradients based on the loss function</h4>
**Gradient**: The gradient of a function is the slope of its derivative (line), or in other words, the rate of change of a function. It's a vector (a direction to move) that points in the direction of greatest increase of the function, and calculated by the <b>derivative</b> operation.
First lets recall the gradient function using an toy example:
$$ z = \left(2x^2 + 3xy\right)$$
```
x = tf.constant(1.0)
y = tf.constant(2.0)
with tf.GradientTape(persistent=True) as g:
g.watch(x)
g.watch(y)
func_test = 2 * x * x + 3 * x * y
```
The <b>tf.gradients()</b> function allows you to compute the symbolic gradient of one tensor with respect to one or more other tensors—including variables. <b>tf.gradients(func, xs)</b> constructs symbolic partial derivatives of sum of <b>func</b> w\.r.t. <i>x</i> in <b>xs</b>.
Now, lets look at the derivitive w\.r.t. <b>var_x</b>:
$$ \frac{\partial :}{\partial \:x}\left(2x^2 + 3xy\right) = 4x + 3y $$
```
var_grad = g.gradient(func_test, x) # Will compute to 10.0
print(var_grad)
```
the derivative w\.r.t. <b>var_y</b>:
$$ \frac{\partial :}{\partial \:y}\left(2x^2 + 3xy\right) = 3x $$
```
var_grad = g.gradient(func_test, y) # Will compute to 3.0
print(var_grad)
```
Now, we can look at gradients w\.r.t all variables:
```
with tf.GradientTape() as tape:
# Forward pass.
output_words_prob = model(_input_data)
# Loss value for this batch.
loss = crossentropy(_targets, output_words_prob)
cost = tf.reduce_sum(loss,axis=0) / batch_size
# Get gradients of loss wrt the trainable variables.
grad_t_list = tape.gradient(cost, tvars)
print(grad_t_list)
```
now, we have a list of tensors, t-list. We can use it to find clipped tensors. <b>clip_by_global_norm</b> clips values of multiple tensors by the ratio of the sum of their norms.
<b>clip_by_global_norm</b> get <i>t-list</i> as input and returns 2 things:
<ul>
<li>a list of clipped tensors, so called <i>list_clipped</i></li>
<li>the global norm (global_norm) of all tensors in t_list</li>
</ul>
```
# Define the gradient clipping threshold
grads, _ = tf.clip_by_global_norm(grad_t_list, max_grad_norm)
grads
```
<h4> 4.Apply the optimizer to the variables/gradients tuple. </h4>
```
# Create the training TensorFlow Operation through our optimizer
train_op = optimizer.apply_gradients(zip(grads, tvars))
```
<a id="ltsm"></a>
<h2>LSTM</h2>
We learned how the model is build step by step. Noe, let's then create a Class that represents our model. This class needs a few things:
<ul>
<li>We have to create the model in accordance with our defined hyperparameters</li>
<li>We have to create the LSTM cell structure and connect them with our RNN structure</li>
<li>We have to create the word embeddings and point them to the input data</li>
<li>We have to create the input structure for our RNN</li>
<li>We need to create a logistic structure to return the probability of our words</li>
<li>We need to create the loss and cost functions for our optimizer to work, and then create the optimizer</li>
<li>And finally, we need to create a training operation that can be run to actually train our model</li>
</ul>
```
class PTBModel(object):
def __init__(self):
######################################
# Setting parameters for ease of use #
######################################
self.batch_size = batch_size
self.num_steps = num_steps
self.hidden_size_l1 = hidden_size_l1
self.hidden_size_l2 = hidden_size_l2
self.vocab_size = vocab_size
self.embeding_vector_size = embeding_vector_size
# Create a variable for the learning rate
self._lr = 1.0
###############################################################################
# Initializing the model using keras Sequential API #
###############################################################################
self._model = tf.keras.models.Sequential()
####################################################################
# Creating the word embeddings layer and adding it to the sequence #
####################################################################
with tf.device("/cpu:0"):
# Create the embeddings for our input data. Size is hidden size.
self._embedding_layer = tf.keras.layers.Embedding(self.vocab_size, self.embeding_vector_size,batch_input_shape=(self.batch_size, self.num_steps),trainable=True,name="embedding_vocab") #[10000x200]
self._model.add(self._embedding_layer)
##########################################################################
# Creating the LSTM cell structure and connect it with the RNN structure #
##########################################################################
# Create the LSTM Cells.
# This creates only the structure for the LSTM and has to be associated with a RNN unit still.
# The argument of LSTMCell is size of hidden layer, that is, the number of hidden units of the LSTM (inside A).
# LSTM cell processes one word at a time and computes probabilities of the possible continuations of the sentence.
lstm_cell_l1 = tf.keras.layers.LSTMCell(hidden_size_l1)
lstm_cell_l2 = tf.keras.layers.LSTMCell(hidden_size_l2)
# By taking in the LSTM cells as parameters, the StackedRNNCells function junctions the LSTM units to the RNN units.
# RNN cell composed sequentially of stacked simple cells.
stacked_lstm = tf.keras.layers.StackedRNNCells([lstm_cell_l1, lstm_cell_l2])
############################################
# Creating the input structure for our RNN #
############################################
# Input structure is 20x[30x200]
# Considering each word is represended by a 200 dimentional vector, and we have 30 batchs, we create 30 word-vectors of size [30xx2000]
# The input structure is fed from the embeddings, which are filled in by the input data
# Feeding a batch of b sentences to a RNN:
# In step 1, first word of each of the b sentences (in a batch) is input in parallel.
# In step 2, second word of each of the b sentences is input in parallel.
# The parallelism is only for efficiency.
# Each sentence in a batch is handled in parallel, but the network sees one word of a sentence at a time and does the computations accordingly.
# All the computations involving the words of all sentences in a batch at a given time step are done in parallel.
########################################################################################################
# Instantiating our RNN model and setting stateful to True to feed forward the state to the next layer #
########################################################################################################
self._RNNlayer = tf.keras.layers.RNN(stacked_lstm,[batch_size, num_steps],return_state=False,stateful=True,trainable=True)
# Define the initial state, i.e., the model state for the very first data point
# It initialize the state of the LSTM memory. The memory state of the network is initialized with a vector of zeros and gets updated after reading each word.
self._initial_state = tf.Variable(tf.zeros([batch_size,embeding_vector_size]),trainable=False)
self._RNNlayer.inital_state = self._initial_state
############################################
# Adding RNN layer to keras sequential API #
############################################
self._model.add(self._RNNlayer)
#self._model.add(tf.keras.layers.LSTM(hidden_size_l1,return_sequences=True,stateful=True))
#self._model.add(tf.keras.layers.LSTM(hidden_size_l2,return_sequences=True))
####################################################################################################
# Instantiating a Dense layer that connects the output to the vocab_size and adding layer to model#
####################################################################################################
self._dense = tf.keras.layers.Dense(self.vocab_size)
self._model.add(self._dense)
####################################################################################################
# Adding softmax activation layer and deriving probability to each class and adding layer to model #
####################################################################################################
self._activation = tf.keras.layers.Activation('softmax')
self._model.add(self._activation)
##########################################################
# Instantiating the stochastic gradient decent optimizer #
##########################################################
self._optimizer = tf.keras.optimizers.SGD(lr=self._lr, clipnorm=max_grad_norm)
##############################################################################
# Compiling and summarizing the model stacked using the keras sequential API #
##############################################################################
self._model.compile(loss=self.crossentropy, optimizer=self._optimizer)
self._model.summary()
def crossentropy(self,y_true, y_pred):
return tf.keras.losses.sparse_categorical_crossentropy(y_true, y_pred)
def train_batch(self,_input_data,_targets):
#################################################
# Creating the Training Operation for our Model #
#################################################
# Create a variable for the learning rate
self._lr = tf.Variable(0.0, trainable=False)
# Get all TensorFlow variables marked as "trainable" (i.e. all of them except _lr, which we just created)
tvars = self._model.trainable_variables
# Define the gradient clipping threshold
with tf.GradientTape() as tape:
# Forward pass.
output_words_prob = self._model(_input_data)
# Loss value for this batch.
loss = self.crossentropy(_targets, output_words_prob)
# average across batch and reduce sum
cost = tf.reduce_sum(loss/ self.batch_size)
# Get gradients of loss wrt the trainable variables.
grad_t_list = tape.gradient(cost, tvars)
# Define the gradient clipping threshold
grads, _ = tf.clip_by_global_norm(grad_t_list, max_grad_norm)
# Create the training TensorFlow Operation through our optimizer
train_op = self._optimizer.apply_gradients(zip(grads, tvars))
return cost
def test_batch(self,_input_data,_targets):
#################################################
# Creating the Testing Operation for our Model #
#################################################
output_words_prob = self._model(_input_data)
loss = self.crossentropy(_targets, output_words_prob)
# average across batch and reduce sum
cost = tf.reduce_sum(loss/ self.batch_size)
return cost
@classmethod
def instance(cls) :
return PTBModel()
```
With that, the actual structure of our Recurrent Neural Network with Long Short-Term Memory is finished. What remains for us to do is to actually create the methods to run through time -- that is, the <code>run_epoch</code> method to be run at each epoch and a <code>main</code> script which ties all of this together.
What our <code>run_epoch</code> method should do is take our input data and feed it to the relevant operations. This will return at the very least the current result for the cost function.
```
########################################################################################################################
# run_one_epoch takes as parameters the model instance, the data to be fed, training or testing mode and verbose info #
########################################################################################################################
def run_one_epoch(m, data,is_training=True,verbose=False):
#Define the epoch size based on the length of the data, batch size and the number of steps
epoch_size = ((len(data) // m.batch_size) - 1) // m.num_steps
start_time = time.time()
costs = 0.
iters = 0
m._model.reset_states()
#For each step and data point
for step, (x, y) in enumerate(reader.ptb_iterator(data, m.batch_size, m.num_steps)):
#Evaluate and return cost, state by running cost, final_state and the function passed as parameter
#y = tf.keras.utils.to_categorical(y, num_classes=vocab_size)
if is_training :
loss= m.train_batch(x, y)
else :
loss = m.test_batch(x, y)
#Add returned cost to costs (which keeps track of the total costs for this epoch)
costs += loss
#Add number of steps to iteration counter
iters += m.num_steps
if verbose and step % (epoch_size // 10) == 10:
print("Itr %d of %d, perplexity: %.3f speed: %.0f wps" % (step , epoch_size, np.exp(costs / iters), iters * m.batch_size / (time.time() - start_time)))
# Returns the Perplexity rating for us to keep track of how the model is evolving
return np.exp(costs / iters)
```
Now, we create the <code>main</code> method to tie everything together. The code here reads the data from the directory, using the <code>reader</code> helper module, and then trains and evaluates the model on both a testing and a validating subset of data.
```
# Reads the data and separates it into training data, validation data and testing data
raw_data = reader.ptb_raw_data(data_dir)
train_data, valid_data, test_data, _, _ = raw_data
# Instantiates the PTBModel class
m=PTBModel.instance()
K = tf.keras.backend
for i in range(max_epoch):
# Define the decay for this epoch
lr_decay = decay ** max(i - max_epoch_decay_lr, 0.0)
dcr = learning_rate * lr_decay
m._lr = dcr
K.set_value(m._model.optimizer.learning_rate,m._lr)
print("Epoch %d : Learning rate: %.3f" % (i + 1, m._model.optimizer.learning_rate))
# Run the loop for this epoch in the training mode
train_perplexity = run_one_epoch(m, train_data,is_training=True,verbose=True)
print("Epoch %d : Train Perplexity: %.3f" % (i + 1, train_perplexity))
# Run the loop for this epoch in the validation mode
valid_perplexity = run_one_epoch(m, valid_data,is_training=False,verbose=False)
print("Epoch %d : Valid Perplexity: %.3f" % (i + 1, valid_perplexity))
# Run the loop in the testing mode to see how effective was our training
test_perplexity = run_one_epoch(m, test_data,is_training=False,verbose=False)
print("Test Perplexity: %.3f" % test_perplexity)
```
As you can see, the model's perplexity rating drops very quickly after a few iterations. As was elaborated before, <b>lower Perplexity means that the model is more certain about its prediction</b>. As such, we can be sure that this model is performing well!
***
This is the end of the <b>Applying Recurrent Neural Networks to Text Processing</b> notebook. Hopefully you now have a better understanding of Recurrent Neural Networks and how to implement one utilizing TensorFlow. Thank you for reading this notebook, and good luck on your studies.
## Want to learn more?
Also, you can use **Watson Studio** to run these notebooks faster with bigger datasets.**Watson Studio** is IBM’s leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, **Watson Studio** enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of **Watson Studio** users today with a free account at [Watson Studio](https://cocl.us/ML0120EN_DSX).This is the end of this lesson. Thank you for reading this notebook, and good luck on your studies.
### Thanks for completing this lesson!
Notebook created by <a href="https://br.linkedin.com/in/walter-gomes-de-amorim-junior-624726121?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01">Walter Gomes de Amorim Junior</a>, <a href = "https://linkedin.com/in/saeedaghabozorgi?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01"> Saeed Aghabozorgi </a></h4>
Updated to TF 2.X by <a href="https://www.linkedin.com/in/samaya-madhavan?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01"> Samaya Madhavan </a>
<hr>
Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0120ENSkillsNetwork20629446-2021-01-01).
| github_jupyter |
## Run GraphScope like NetworkX
Graphscope provides a set of graph analysis interfaces compatible with Networkx.
In this article, we will show how to use graphscope to perform graph analysis like Networkx.
### How does Networkx perform graph analysis ?
Usually, the graph analysis process of NetworkX starts with the construction of a graph.
In the following example, we create an empty graph first, and then expand the data through the interface of NetworkX.
```
# Install graphscope package if you are NOT in the Playground
!pip3 install graphscope
import networkx
# Initialize an empty graph
G = networkx.Graph()
# Add edges (1, 2)and(1 3) by `add_edges_from` interface
G.add_edges_from([(1, 2), (1, 3)])
# Add vertex "4" by `add_node` interface
G.add_node(4)
```
Then we can query the graph information.
```
# Query the number of vertices by `number_of_nodes` interface.
G.number_of_nodes()
# Similarly, query the number of edges by `number_of_edges` interface.
G.number_of_edges()
# Query the degree of each vertex by `degree` interface.
sorted(d for n, d in G.degree())
```
Finally, calling the builtin algorithm of NetworkX to analysis the graph `G`.
```
# Run 'connected components' algorithm
list(networkx.connected_components(G))
# Run 'clustering' algorithm
networkx.clustering(G)
```
### How to use NetworkX interface from GraphScope
**Graph Building**
To use NetworkX interface from graphscope, we just need to replace `import networkx as nx` with `import graphscope.nx as nx `.
Here we use `nx.Graph()` interace to create an empty undirected graph.
```
import graphscope
graphscope.set_option(show_log=True)
import graphscope.nx as nx
# Initialize an empty graph
G = nx.Graph()
```
**Add edges and vertices**
Just like operating NetworkX, you can add vertices by `add_node` `add_nodes_from` and add edges by `add_edge` `add_edges_from`.
```
# Add one vertex by `add_node` interface
G.add_node(1)
# Or add a batch of vertices from iterable list
G.add_nodes_from([2, 3])
# Also you can add attributes while adding vertices
G.add_nodes_from([(4, {"color": "red"}), (5, {"color": "green"})])
# Similarly, add one edge by `add_edge` interface
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e)
# Or add a batch of edges from iterable list
G.add_edges_from([(1, 2), (1, 3)])
# Add attributes while adding edges
G.add_edges_from([(1, 2), (2, 3, {'weight': 3.1415})])
```
**Query Graph**
Just like operating NetworkX, you can search the number of vertices/edge by `number_of_nodes`/`number_of_edges` interface, or query the neighbor of vertex by `adj` interface.
```
# Query the number of vertices by `number_of_nodes` interface.
G.number_of_nodes()
# Similarly, query the number of edges by `number_of_edges` interface.
G.number_of_edges()
# list the vertices in graph `G`
list(G.nodes)
# list the edges in graph `G`
list(G.edges)
# query the nerghbors of vertex '1'
list(G.adj[1])
# search the degree of vertex '1'
G.degree(1)
```
**Delete**
Just like operating NetworkX, you can remove vertices by `remove_node`or `remove_nodes_from` interface, and remove edges by `remove_edge` or `remove_edges_from` interface.
```
# remove one vertex by `remove_node` interface
G.remove_node(5)
list(G.nodes)
# remove a batch of vertices by `remove_nodes_from` interface
G.remove_nodes_from([4, 5])
list(G.nodes)
# remove one edge by `remove_edge` interface
G.remove_edge(1, 2)
list(G.edges)
# remove a batch of edges by `remove_edges_from` interface
G.remove_edges_from([(1, 3), (2, 3)])
list(G.edges)
# query the number of vertices after removal
G.number_of_nodes()
# query the number of edges after removal
G.number_of_edges()
```
**Graph Analysis**
The interface of graph analysis module in graphscope is also compatible with NetworkX.
In following examples, we use `connected_components` to analyze the connected components of the graph, use `clustering` to get the clustering coefficient of each vertex, and `all_pairs_shortest_path` to compute the shortest path between any two vertices.
```
# Building graph
G = nx.Graph()
G.add_edges_from([(1, 2), (1, 3)])
G.add_node(4)
# Run connected_components
list(nx.connected_components(G))
# Run clustering
nx.clustering(G)
# Run all_pairs_shortest_path
sp = dict(nx.all_pairs_shortest_path(G))
sp[3]
```
**Graph Display**
Like NetworkX, you can draw a graph by `draw` interface, which relies on the drawing function of 'Matplotlib'.
You should install `matplotlib` first if you are not in playground environment.
```
!pip3 install matplotlib
```
使用 GraphScope 来进行简单地绘制图
```
# Create a star graph with 5 vertices
G = nx.star_graph(5)
# Sraw
nx.draw(G, with_labels=True, font_weight='bold')
```
### The performance speed-up of GraphScope over NetworkX can reach up to several orders of magnitudes.
Let's see how much GraphScope improves the algorithm performance compared with NetworkX by a simple experiment.
We run [clustering](https://networkx.org/documentation/stable/reference/algorithms/generated/networkx.algorithms.cluster.clustering.html#networkx.algorithms.cluster.clustering) algorithm on [twitter datasets](https://snap.stanford.edu/data/ego-Twitter.html).
Download dataset if you are not in playground environment
```
!wget https://raw.githubusercontent.com/GraphScope/gstest/master/twitter.e -P /tmp
```
Loading dataset both in GraphScope and NetwrokX.
```
import os
import graphscope.nx as gs_nx
import networkx as nx
# loading graph in NetworkX
g1 = nx.read_edgelist(
os.path.expandvars('/tmp/twitter.e'), nodetype=int, data=False, create_using=nx.Graph
)
type(g1)
# Loading graph in GraphScope
g2 = gs_nx.read_edgelist(
os.path.expandvars('/tmp/twitter.e'), nodetype=int, data=False, create_using=gs_nx.Graph
)
type(g2)
```
Run algorithm and display time both in GraphScope and NetworkX.
```
%%time
# GraphScope
ret_gs = gs_nx.clustering(g2)
%%time
# NetworkX
ret_nx = nx.clustering(g1)
# Result comparison
ret_gs == ret_nx
```
| github_jupyter |
# AnalysePhoneticsBetweenSkills
```
#laod CMU dict
CMUpath = 'cmudict_SPHINX_40.txt'
cmu_dict = {}
with open(CMUpath, "r", encoding='utf8') as f:
for line in f:
#A line looks like this
#ABORT AH B AO R T
if line and len(line.strip())>0:
spl = line.split('\t')
word = spl[0].strip()
phonemes = spl[1].strip()
cmu_dict[word] = phonemes.split(' ')
'''
Call getPhoneticTranslation to get the phonetic trasnlation of a sentence.
It cleans the word and obtains a CMU translation of the sentence.
Replaces the non existent words in the dictionary by a word of the same length formed by '_'
'''
import string
invalidChars = set(string.punctuation)
numbers = {'0':'ZERO','1':'ONE','2':'TWO','3':'THREE','4':'FOUR','5':'FIVE',
'6':'SIX','7':'SEVEN','8':'EIGHT','9':'NINE'}
def getPhoneticTranslation(sentence):
#Transcriptions to be compared were converted to uppercase and all non-ASCII characters and punctuation were
#removed. Digits were replaced with their corresponding texts (e.g., “2” became “two”)
def PreProcessWord(word):
for c in invalidChars:
if c in word:
word = word.replace(c,'')
for n in numbers.keys():
if n in word:
word = word.replace(n, ' '+numbers[n]+' ')
word = word.upper()
word = word.replace(' ', ' ')
return word.strip()
a=PreProcessWord(sentence)
cmutranslated = []
for w in a.split(' '):
if w in cmu_dict:
b = cmu_dict[w]
else:
b = ['_']*len(w)
cmutranslated+= b+[' ']
return cmutranslated[:len(cmutranslated)-1]
#example of the process
word = '123feeli_ng down!'
print('*', word)
print(getPhoneticTranslation(word))
word = 'Apple watch --!'
print('*', word)
print(getPhoneticTranslation(word))
word = 'Unexxxxisting word but the rest is fine'
print('*', word)
print(getPhoneticTranslation(word))
```
### Load new Alexa Dataset
```
#Load dataset
import os
import json
def reloadData():
def getFiles(dirName):
listOfFile = os.listdir(dirName)
completeFileList = list()
for file in listOfFile:
completePath = os.path.join(dirName, file)
if os.path.isdir(completePath):
completeFileList = completeFileList + getFiles(completePath)
elif '.json' in completePath:
completeFileList.append(completePath.replace('\\', '/'))
return completeFileList
allf = getFiles( "../NewFullDataset/")
print(allf)
#create dictionary of markets and skills
skillsmarketd = {}
for f in allf:
marketname = f.split('/')[2].split('.json')[0]
with open(f, 'r') as json_file:
skillsmarketd[marketname] = json.load(json_file)
print()
print('Total unique skills per market as per last week')
for key,value in skillsmarketd.items():
print(key, ": ", len(value))
print()
print()
return skillsmarketd
allskills = reloadData()
```
### Levenshtein Distance
```
import editdistance
#luckily I found this iplementation of levehenstein distance that goes almost 100 times faster than the defined above...
def levdistance(w1,w2):
#if there is a difference of more than 3 words in the skil names,
#we are not interested in the real distance as they are not interesting
if(w1 == w2):
return 0
if(len(w1) > len(w2)+2):
return 1
if(len(w2) > len(w1)+2):
return 1
return editdistance.eval(w1, w2)/ max(len(w1), len(w2))
levdistance(['ck', 'a', 't'], ['k', 'a', 't'])
```
### Perform experiment for TABLE III.SKILL NAMES WITHLEVENSHTEIN DISTANCE≤0.2, considering english markets and ignoring words that do not have a CMU translation
#### NOTE: Do not run this, it takes a while!
```
import datetime
#explore the different markets per separate, otherwis eI get memory errors
data = reloadData()
englishmarkets = ['UK','AU', 'CA', 'IN', 'US']
for market in englishmarkets:
uniqskcount = {} #dictionary that contains the quantity of skills for each element
skilltocmu = {} #set of unique skill names considered and their translation to phonetics
'''
Create skilltocmu dict
'''
print('Exploring market', market)
skills = data[market]
for skillid, skillobj in skills.items():
name1 = skillobj['name'].lower()
cmuname = getPhoneticTranslation(name1)
#we ignore skills we couldnt completely trasnalte (include '_') [also to speed up the process]
if '_' not in cmuname:
skilltocmu[name1] = cmuname
if(name1 in uniqskcount):
uniqskcount[name1] += 1
else:
uniqskcount[name1] = 1
print('finished creating crate skilltocmu dict for market', market)
'''
Calculate levehenstein distances:
This process is super slow. We do many tricks to speed up the process:
1- since we are only interested in super similar skill names, we ignore words with different lengths (see lev function above)
2- we only iterate trhough unique skill names transformed to lower, hence the results may be different as
in other figures since 'catFacts' and 'CatFacts' are considered the same skill
3- We only identify which skill names have levehenstein distances < 0.1 or 0.2 (to reduce the final json files).
Then, we count how many of skills with this name.lower() exist in the market to get hte data for table III
'''
alldistances = set([])
uniqsk = list(skilltocmu.keys())
totalanalysed = 0
for i in range(len(uniqsk)):
name1 = uniqsk[i].lower()
if(name1 not in skilltocmu):
continue
cmuname1 = skilltocmu[name1]
for j in range(i, len(uniqsk)):
name2 = uniqsk[j].lower()
if(name2 not in skilltocmu):
continue
cmuname2 = skilltocmu[name2]
d= levdistance(cmuname1, cmuname2)
if(d<0.31):
#only save these skills with smaller lev distances (otherwise memory error)
alldistances.add( (d,name1,name2) )
#count all distances processed
totalanalysed +=1
if(i%1000 == 0):
now = datetime.datetime.now()
print('\t[',now.hour,':',now.minute,'.',now.second,'] Distances computed ', i)
#save analysis in PhonDict json
path = 'PhonDict/'+market+'.json'
with open(path, 'w') as json_file:
print('Saving json in ',path)
json.dump( {'phonetic_distances_minororequal03':list(alldistances), 'all_distances_processed':totalanalysed, 'skillnames_count_d':uniqskcount} , json_file)
#min01 = [d for d in alldistances if d <= 0.1 and d!=0]
#min02 = [d for d in alldistances if d <= 0.2 and d!=0]
##we should count how many times this skills appear in the market!
#print('There are ', len(min01), 'skills within a ld of 0.1')
#print('There are ', len(min02), 'skills within a ld of 0.2')
#print('Total unique skill names ', len(uniqsk))
#
now = datetime.datetime.now()
print('[',now.hour,':',now.minute,'.',now.second,'] Finished estiamting skill names levehenstein distances for market', market)
```
### Load the dictionaries of words and get the data for table III
```
def getFiles(dirName):
listOfFile = os.listdir(dirName)
completeFileList = list()
for file in listOfFile:
completePath = os.path.join(dirName, file)
if os.path.isdir(completePath):
completeFileList = completeFileList + getFiles(completePath)
elif '.json' in completePath:
completeFileList.append(completePath.replace('\\', '/'))
return completeFileList
#load the jsons
alljsons = getFiles('PhonDict/')
phondict = {}
for f in alljsons:
market = f.split('/')[1].split('.json')[0]
print('Loaded json for ', market, ' :', f)
with open(f, 'r') as json_file:
phondict[market] = json.load(json_file)
#get the data
for market in phondict:
print('Studying market', market)
phondist = phondict[market]['phonetic_distances_minororequal03']
totalcomparisons = phondict[market]['all_distances_processed']
skillnamecount_d = phondict[market]['skillnames_count_d']
totalskills_hits = sum(skillnamecount_d.values())
print('Total hits for unique skill names.lower() ', totalskills_hits)
'''
find, for every unique name skill.lower(), the minimum distance to any other skill.lower()
'''
min_d_skillname = {}
for skillname in skillnamecount_d.keys():
#print('exploring skill ', skillname)
alldforskill = [(d, sk1, sk2) for (d, sk1, sk2) in phondist if (sk1 == skillname or sk2 == skillname) and d!=0]
if(len(alldforskill)>0):
min_d_skillname[skillname] = min([d for (d, sk1, sk2) in alldforskill ])
else:
min_d_skillname[skillname] = 1
'''
now, iterate over all unique skills.lower(), and bin them < 0.1 and < 0.2
'''
min_d_skillname_l = [ (k,v) for (k,v) in min_d_skillname.items()]
#get these skills with a minimum lev distance to any other skill name of 0.1
min01 = [(skname, skmin) for (skname, skmin) in min_d_skillname_l if skmin <= 0.1]
#get how many skills are aggregated with that specific name
total01 = sum([skillnamecount_d[skname] for (skname, skmin) in min01])
print('There are ', len(min01), 'skills within a ld of 0.1 to any other skillname that does not contain '_''', addding a total of hits ', total01, '(',total01/totalskills_hits,'%)')
#get these skills with a minimum lev distance to any other skill name of 0.1
min02 = [(skname, skmin) for (skname, skmin) in min_d_skillname_l if skmin <= 0.2]
#get how many skills are aggregated with that specific name
total02 = sum([skillnamecount_d[skname] for (skname, skmin) in min02])
print('There are ', len(min02), 'skills within a ld of 0.1 to any other skillname that does not contain '_', addding a total of hits ', total02, '(',total02/totalskills_hits,'%)')
min01
min01 = [(d, sk1, sk2) for (d, sk1, sk2) in phondist if d <= 0.1 and d!=0]
total = 0
for m in min01:
name1,name2 = m[1],m[2]
total += skillnamecount_d[name1]
total += skillnamecount_d[name2]
print('There are ', len(min01), 'skills within a ld of 0.1, addding a total of hits ', total, '(',total/totalanalysis,'%)')
min02 = [(d, sk1, sk2) for (d, sk1, sk2) in phondist if d <= 0.2 and d!=0]
total = 0
for m in min02:
name1,name2 = m[1],m[2]
total += skillnamecount_d[name1]
total += skillnamecount_d[name2]
print('There are ', len(min02), 'skills within a ld of 0.1, addding a total of hits ', total, '(',total/totalanalysis,'%)')
phondist
# #first create a dictionary with all skillname to CMU tranlations to speed up the process
#skilltocmu = {}
#uniqueskillnames = {}
#for market in ['UK','US', 'CA', 'IN', 'AU']:
# uniqsk = set([])
# skills = data[market]
# for skillid, skillobj in skills.items():
# name1 = skillobj['name'].lower()
# uniqsk.add(name1)
# cmuname = getPhoneticTranslation(name1)
# if '_' not in cmuname:
# skilltocmu[name1] = cmuname
#
# uniqueskillnames[market] = uniqsk
# print('finished translating skill names to cmu for market', market)
#second find levehenstein distances between all uinque skill names of every market
#ldd = {}
for market in ['UK']:#,'US', 'CA', 'IN', 'AU']:
alldistances = []
print('Processing market ', market)
uniqsk = list(uniqueskillnames[market])
#ldp = {}
for i in range(len(uniqsk)):
name1 = uniqsk[i].lower()
if(name1 not in skilltocmu):
continue
cmuname1 = skilltocmu[name1]
for j in range(i, len(uniqsk)):
name2 = uniqsk[j].lower()
if(name2 not in skilltocmu):
continue
cmuname2 = skilltocmu[name2]
d= levdistance(cmuname1, cmuname2)
#alldistances.append( (d, name1, name2) )
alldistances.append( d )
#ldp[name1+'_'+name2] = d
if(i%1000 == 0):
print('\tDistances computed ', i)
#comment this part
#if(i > 2000):
# ldd[market] = ldp
# break
#ldd[market] = ldp
min01 = [d for d in alldistances if d <= 0.1 and d!=0]
min02 = [d for d in alldistances if d <= 0.2 and d!=0]
#we should count how many times this skills appear in the market!
print('There are ', len(min01), 'skills within a ld of 0.1')
print('There are ', len(min02), 'skills within a ld of 0.2')
print('Total unique skill names ', len(uniqsk))
print('Finished estiamting skill names levehenstein distances for market', market)
start 00.19
```
```
data = reloadData()
for market in ['UK']:#, 'AU', 'CA', 'IN', 'US']:
print('Exploring market', market)
sknames = list(uniqueskillnames[market])
for i in range(len(sknames)):
name1 = sknames[i].lower()
for j in range(len(sknames)):
name2 = sknames[j].lower()
combo = name1+'_'+name2
combo2 = name2+'_'+name1
if(combo in ldd[market]):
ld = ldd[market][combo]
alldistances.append( (ld, name1, name2) )
elif(combo2 in ldd[market]):
ld = ldd[market][combo2]
alldistances.append( (ld, name1, name2) )
if(i%100 == 0):
print('Analisys computed for ', i, 'skills')
#print('explord skill', i)
i+=1
min01 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.1 and d!=0]
min02 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.2 and d!=0]
#skills = data[market]
#alldistances = []
#i = 0
#for skillid, skillobj in skills.items():
# name1 = skillobj['name'].lower()
# #cmuname1 = skilltocmu[name1]
# for skillid2, skillobj2 in skills.items():
# name2 = skillobj2['name'].lower()
# #cmuname2 = skilltocmu[name2]
# combo = name1+'_'+name2
# combo2 = name2+'_'+name1
# if(combo in ldd[market]):
# ld = ldd[market][combo]
# alldistances.append( (ld, name1, name2) )
# elif(combo2 in ldd[market]):
# ld = ldd[market][combo2]
# alldistances.append( (ld, name1, name2) )
#collected all releveant data for the market
min01 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.1 and d!=0]
min02 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.2 and d!=0]
print( len(min01) )
print( len(min02) )
min01 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.1 and d!=0]
min02 = [(d,n1,n2) for (d,n1,n2) in alldistances if d <= 0.2 and d!=0]
print( len(min01) )
print( len(min02) )
```
```
min01
```
| github_jupyter |
# Face Generation
In this project, you'll define and train a DCGAN on a dataset of faces. Your goal is to get a generator network to generate *new* images of faces that look as realistic as possible!
The project will be broken down into a series of tasks from **loading in data to defining and training adversarial networks**. At the end of the notebook, you'll be able to visualize the results of your trained Generator to see how it performs; your generated samples should look like fairly realistic faces with small amounts of noise.
### Get the Data
You'll be using the [CelebFaces Attributes Dataset (CelebA)](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) to train your adversarial networks.
This dataset is more complex than the number datasets (like MNIST or SVHN) you've been working with, and so, you should prepare to define deeper networks and train them for a longer time to get good results. It is suggested that you utilize a GPU for training.
### Pre-processed Data
Since the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. Some sample data is show below.
<img src='assets/processed_face_data.png' width=60% />
> If you are working locally, you can download this data [by clicking here](https://s3.amazonaws.com/video.udacity-data.com/topher/2018/November/5be7eb6f_processed-celeba-small/processed-celeba-small.zip)
This is a zip file that you'll need to extract in the home directory of this notebook for further loading and processing. After extracting the data, you should be left with a directory of data `processed_celeba_small/`
```
# can comment out after executing
#!unzip processed_celeba_small.zip
data_dir = 'processed_celeba_small/'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import pickle as pkl
import matplotlib.pyplot as plt
import numpy as np
import problem_unittests as tests
#import helper
%matplotlib inline
```
## Visualize the CelebA Data
The [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html) dataset contains over 200,000 celebrity images with annotations. Since you're going to be generating faces, you won't need the annotations, you'll only need the images. Note that these are color images with [3 color channels (RGB)](https://en.wikipedia.org/wiki/Channel_(digital_image)#RGB_Images) each.
### Pre-process and Load the Data
Since the project's main focus is on building the GANs, we've done *some* of the pre-processing for you. Each of the CelebA images has been cropped to remove parts of the image that don't include a face, then resized down to 64x64x3 NumPy images. This *pre-processed* dataset is a smaller subset of the very large CelebA data.
> There are a few other steps that you'll need to **transform** this data and create a **DataLoader**.
#### Exercise: Complete the following `get_dataloader` function, such that it satisfies these requirements:
* Your images should be square, Tensor images of size `image_size x image_size` in the x and y dimension.
* Your function should return a DataLoader that shuffles and batches these Tensor images.
#### ImageFolder
To create a dataset given a directory of images, it's recommended that you use PyTorch's [ImageFolder](https://pytorch.org/docs/stable/torchvision/datasets.html#imagefolder) wrapper, with a root directory `processed_celeba_small/` and data transformation passed in.
```
# necessary imports
import torch
from torchvision import datasets
from torchvision import transforms
def get_dataloader(batch_size, image_size, data_dir='processed_celeba_small/'):
"""
Batch the neural network data using DataLoader
:param batch_size: The size of each batch; the number of images in a batch
:param img_size: The square size of the image data (x, y)
:param data_dir: Directory where image data is located
:return: DataLoader with batched data
"""
# TODO: Implement function and return a dataloader
transform = transforms.Compose([transforms.Resize(image_size), transforms.ToTensor()])
dataset = datasets.ImageFolder(data_dir, transform)
data_loader = torch.utils.data.DataLoader(dataset, batch_size, shuffle=True)
return data_loader
```
## Create a DataLoader
#### Exercise: Create a DataLoader `celeba_train_loader` with appropriate hyperparameters.
Call the above function and create a dataloader to view images.
* You can decide on any reasonable `batch_size` parameter
* Your `image_size` **must be** `32`. Resizing the data to a smaller size will make for faster training, while still creating convincing images of faces!
```
# Define function hyperparameters
batch_size = 64
img_size = 32
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# Call your function and get a dataloader
celeba_train_loader = get_dataloader(batch_size, img_size)
```
Next, you can view some images! You should seen square images of somewhat-centered faces.
Note: You'll need to convert the Tensor images into a NumPy type and transpose the dimensions to correctly display an image, suggested `imshow` code is below, but it may not be perfect.
```
# helper display function
def imshow(img):
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# obtain one batch of training images
dataiter = iter(celeba_train_loader)
images, _ = dataiter.next() # _ for no labels
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(20, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
```
#### Exercise: Pre-process your image data and scale it to a pixel range of -1 to 1
You need to do a bit of pre-processing; you know that the output of a `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
```
# TODO: Complete the scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# assume x is scaled to (0, 1)
# scale to feature_range and return scaled x
x = (x * (feature_range[1] - feature_range[0])) + feature_range[0]
return x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
# check scaled range
# should be close to -1 to 1
img = images[0]
scaled_img = scale(img)
print('Min: ', scaled_img.min())
print('Max: ', scaled_img.max())
```
---
# Define the Model
A GAN is comprised of two adversarial networks, a discriminator and a generator.
## Discriminator
Your first task will be to define the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers. To deal with this complex data, it's suggested you use a deep network with **normalization**. You are also allowed to create any helper functions that may be useful.
#### Exercise: Complete the Discriminator class
* The inputs to the discriminator are 32x32x3 tensor images
* The output should be a single value that will indicate whether a given image is real or fake
```
import torch.nn as nn
import torch.nn.functional as F
# From CycleGAN notebook
def conv(in_channels, out_channels, kernel_size=4, stride=2, padding=1, batch_norm=True):
layers = []
conv_layer = nn.Conv2d(in_channels=in_channels, out_channels=out_channels,
kernel_size=kernel_size, stride=stride, padding=padding, bias=False)
layers.append(conv_layer)
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
class Discriminator(nn.Module):
def __init__(self, conv_dim):
"""
Initialize the Discriminator Module
:param conv_dim: The depth of the first convolutional layer
"""
super(Discriminator, self).__init__()
# complete init function
self.conv_dim = conv_dim
self.conv1 = conv(3, conv_dim, 4, batch_norm=False) # first layer, no batch_norm
self.conv2 = conv(conv_dim, conv_dim*2, 4)
self.conv3 = conv(conv_dim*2, conv_dim*4, 4)
self.fc = nn.Linear(conv_dim*4*4*4, 1)
def forward(self, x):
"""
Forward propagation of the neural network
:param x: The input to the neural network
:return: Discriminator logits; the output of the neural network
"""
# define feedforward behavior
x = F.leaky_relu(self.conv1(x), 0.2)
x = F.leaky_relu(self.conv2(x), 0.2)
x = F.leaky_relu(self.conv3(x), 0.2)
x = x.view(-1, self.conv_dim*4*4*4)
x = self.fc(x)
return x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_discriminator(Discriminator)
```
## Generator
The generator should upsample an input and generate a *new* image of the same size as our training data `32x32x3`. This should be mostly transpose convolutional layers with normalization applied to the outputs.
#### Exercise: Complete the Generator class
* The inputs to the generator are vectors of some length `z_size`
* The output should be a image of shape `32x32x3`
```
def deconv(in_channels, out_channels, kernel_size=4, stride=2, padding=1, batch_norm=True, activate=True):
layers = []
layers.append(nn.ConvTranspose2d(in_channels, out_channels, kernel_size,
stride, padding, bias=False))
if batch_norm:
layers.append(nn.BatchNorm2d(out_channels))
if activate:
layers.append(nn.ReLU())
return nn.Sequential(*layers)
from collections import OrderedDict
class Generator(nn.Module):
def __init__(self, z_size, conv_dim):
"""
Initialize the Generator Module
:param z_size: The length of the input latent vector, z
:param conv_dim: The depth of the inputs to the *last* transpose convolutional layer
"""
super(Generator, self).__init__()
# complete init function
out_size = 32
n_deep_conv_layers = 2
self.conv_t_w_in = 4
self.conv_t_dim_in = conv_dim * (n_deep_conv_layers + 1) * 2
fc_out = self.conv_t_dim_in * self.conv_t_w_in**2
self.fc1 = nn.Linear(z_size, fc_out)
self.conv_t_in = deconv(self.conv_t_dim_in,
conv_dim * ((n_deep_conv_layers) * 2))
deep_conv_t_layers = []
for layer in reversed(range(1, n_deep_conv_layers + 1)):
i = abs(layer - n_deep_conv_layers)
conv_in = conv_dim * layer * 2
conv_out = conv_dim * (((layer - 1) * 2) or 1)
deep_conv_t_layers.append((f'conv_t{i}', deconv(conv_in, conv_out,
kernel_size=4 if bool(i%2) else 3,
stride=2 if bool(i%2) else 1)))
self.deep_conv_t = nn.Sequential(OrderedDict(deep_conv_t_layers))
self.conv_t_out = deconv(conv_dim, 3, batch_norm=False, activate=False)
def forward(self, x):
"""
Forward propagation of the neural network
:param x: The input to the neural network
:return: A 32x32x3 Tensor image as output
"""
# define feedforward behavior
x = self.fc1(x)
x = x.view(x.size(0), self.conv_t_dim_in, self.conv_t_w_in, self.conv_t_w_in)
x = self.conv_t_in(x)
x = self.deep_conv_t(x)
x = torch.tanh(self.conv_t_out(x))
return x
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_generator(Generator)
```
## Initialize the weights of your networks
To help your models converge, you should initialize the weights of the convolutional and linear layers in your model. From reading the [original DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf), they say:
> All weights were initialized from a zero-centered Normal distribution with standard deviation 0.02.
So, your next task will be to define a weight initialization function that does just this!
You can refer back to the lesson on weight initialization or even consult existing model code, such as that from [the `networks.py` file in CycleGAN Github repository](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/blob/master/models/networks.py) to help you complete this function.
#### Exercise: Complete the weight initialization function
* This should initialize only **convolutional** and **linear** layers
* Initialize the weights to a normal distribution, centered around 0, with a standard deviation of 0.02.
* The bias terms, if they exist, may be left alone or set to 0.
```
def weights_init_normal(m):
"""
Applies initial weights to certain layers in a model .
The weights are taken from a normal distribution
with mean = 0, std dev = 0.02.
:param m: A module or layer in a network
"""
# classname will be something like:
# `Conv`, `BatchNorm2d`, `Linear`, etc.
classname = m.__class__.__name__
# TODO: Apply initial weights to convolutional and linear layers
if classname.find('Linear') != -1 or classname.find('Conv') != -1:
m.weight.data.normal_(0, 0.02)
```
## Build complete network
Define your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
def build_network(d_conv_dim, g_conv_dim, z_size):
# define discriminator and generator
D = Discriminator(d_conv_dim)
G = Generator(z_size=z_size, conv_dim=g_conv_dim)
# initialize model weights
D.apply(weights_init_normal)
G.apply(weights_init_normal)
print(D)
print()
print(G)
return D, G
```
#### Exercise: Define model hyperparameters
```
# Define model hyperparams
d_conv_dim = 128
g_conv_dim = 128
z_size = 96
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
D, G = build_network(d_conv_dim, g_conv_dim, z_size)
```
### Training on GPU
Check if you can train on GPU. Here, we'll set this as a boolean variable `train_on_gpu`. Later, you'll be responsible for making sure that
>* Models,
* Model inputs, and
* Loss function arguments
Are moved to GPU, where appropriate.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import torch
# Check for a GPU
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('No GPU found. Please use a GPU to train your neural network.')
else:
print('Training on GPU!')
```
---
## Discriminator and Generator Losses
Now we need to calculate the losses for both types of adversarial networks.
### Discriminator Losses
> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
### Generator Loss
The generator loss will look similar only with flipped labels. The generator's goal is to get the discriminator to *think* its generated images are *real*.
#### Exercise: Complete real and fake loss functions
**You may choose to use either cross entropy or a least squares error loss to complete the following `real_loss` and `fake_loss` functions.**
```
def real_loss(D_out, smooth=False):
'''Calculates how close discriminator outputs are to being real.
param, D_out: discriminator logits
return: real loss'''
D_out = D_out.squeeze()
target = torch.ones(D_out.size())
if smooth:
target = target * 0.9
if train_on_gpu:
target = target.cuda()
loss = F.binary_cross_entropy_with_logits(D_out, target)
return loss
def fake_loss(D_out):
'''Calculates how close discriminator outputs are to being fake.
param, D_out: discriminator logits
return: fake loss'''
D_out = D_out.squeeze()
target = torch.zeros(D_out.size())
if train_on_gpu:
target = target.cuda()
loss = F.binary_cross_entropy_with_logits(D_out, target)
return loss
```
## Optimizers
#### Exercise: Define optimizers for your Discriminator (D) and Generator (G)
Define optimizers for your models with appropriate hyperparameters.
```
import torch.optim as optim
# Create optimizers for the discriminator D and generator G
d_optimizer = optim.Adam(D.parameters(), 0.0002, [0.5, 0.999])
g_optimizer = optim.Adam(G.parameters(), 0.0002, [0.5, 0.999])
```
---
## Training
Training will involve alternating between training the discriminator and the generator. You'll use your functions `real_loss` and `fake_loss` to help you calculate the discriminator losses.
* You should train the discriminator by alternating on real and fake images
* Then the generator, which tries to trick the discriminator and should have an opposing loss function
#### Saving Samples
You've been given some code to print out some loss statistics and save some generated "fake" samples.
#### Exercise: Complete the training function
Keep in mind that, if you've moved your models to GPU, you'll also have to move any model inputs to GPU.
```
def train(D, G, n_epochs, print_every=50):
'''Trains adversarial networks for some number of epochs
param, D: the discriminator network
param, G: the generator network
param, n_epochs: number of epochs to train for
param, print_every: when to print and record the models' losses
return: D and G losses'''
# move models to GPU
if train_on_gpu:
D.cuda()
G.cuda()
# keep track of loss and generated, "fake" samples
samples = []
losses = []
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# move z to GPU if available
if train_on_gpu:
fixed_z = fixed_z.cuda()
# epoch training loop
for epoch in range(n_epochs):
# batch training loop
for batch_i, (real_images, _) in enumerate(celeba_train_loader):
batch_size = real_images.size(0)
real_images = scale(real_images)
# ===============================================
# YOUR CODE HERE: TRAIN THE NETWORKS
# ===============================================
if train_on_gpu:
real_images = real_images.cuda()
# 1. Train the discriminator on real and fake images
d_optimizer.zero_grad()
d_real_out = D(real_images)
d_real_loss = real_loss(d_real_out, True)
fake_images = G(fixed_z)
d_fake_out = D(fake_images)
d_fake_loss = fake_loss(d_fake_out)
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# 2. Train the generator with an adversarial loss
g_optimizer.zero_grad()
fake_images = G(fixed_z)
g_fake_out = D(fake_images)
g_loss = real_loss(g_fake_out)
g_loss.backward()
g_optimizer.step()
# ===============================================
# END OF YOUR CODE
# ===============================================
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, n_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# this code assumes your generator is named G, feel free to change the name
# generate and save sample, fake images
G.eval() # for generating samples
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
# finally return losses
return losses
```
Set your number of training epochs and train your GAN!
```
# set number of epochs
#n_epochs = 100
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# call training function
#losses = train(D, G, n_epochs=n_epochs)
#!curl -O https://raw.githubusercontent.com/udacity/workspaces-student-support/master/jupyter/workspace_utils.py
# set number of epochs
n_epochs = 40
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# call training function
losses = train(D, G, n_epochs=n_epochs)
```
## Training loss
Plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
```
## Generator samples from training
View samples of images from the generator, and answer a question about the strengths and weaknesses of your trained models.
```
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img + 1)*255 / (2)).astype(np.uint8)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
# Load samples from generator, taken while training
with open('train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
_ = view_samples(-1, samples)
# Load samples from generator, taken while training
with open('simple_40_epoch_train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
_ = view_samples(-1, samples)
# Load samples from generator, taken while training
with open('80_epoch_train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
_ = view_samples(-1, samples)
```
### Question: What do you notice about your generated samples and how might you improve this model?
When you answer this question, consider the following factors:
* The dataset is biased; it is made of "celebrity" faces that are mostly white
* Model size; larger models have the opportunity to learn more features in a data feature space
* Optimization strategy; optimizers and number of epochs affect your final result
**Answer:** Considering the bias of the train images, it is clear that the resulting faces are not very diverse. I think having a broader range of faces could make for a more interesting result. The resulting model might even produce more realistic images.
Considering that my model is not especially complex, it is also not surprising that the generated images are not especially convincing. Additionally the images are low in quality. I think a more complex model, with a higher output size could be interesting, though computationally challenging.
On epochs, after testing a simple Generator on a single epoch, the generated images were not very nice to look at. Though, if you squint, they do resemble human faces. After 100 epochs on the same model, the faces appear corrupted not at all like faces, but all sharing many "features". I suspect the Generator will not reach an optimal solution.
A more complex model produced better single epoch results, but after 20 epochs, the outcome was similarly not optimal.
A similarly complex model, with fewer conv dimensions (down to 64 from 256) using binary cross entropy loss, with smoothing (previously, mean squared error, no smoothing) produces the decent results after 5 epochs, snd seems to have resolved the corruption issue. After 50 epochs corruption issue returns. Note: D loss is much lower than G loss.
A simplified D produced identical results at 5 (compared to prev model, 5 epochs) epochs, but the G and D loss appear more natural. After 50 epochs, uncorrupted, but unconvincing faces are generated.
Asimplified D, with tweaked hyperparams, batch 96=>64, shows good signs at 40 epochs. At 80 epochs, the outcome is passable, though not impressive. Some faces look realistic.
Further simplified model produces good results at 40 epochs (5 conv layers => 4 conv layers.)
### Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_face_generation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "problem_unittests.py" files in your submission.
| github_jupyter |
Topic: NMT Re-Ranking and Paraphrase Detection Using TreeLSTM-based Pointer Network and Attention Mechanism on Dependency Trees
## NMT Re-Ranking
Instead of solely relying on an integrated search for the best translation, <br>
we may introduce a second decoding pass <br>
in which the best translation is chosen from <br>
the set of the most likely candidates <br>
generated by a traditional decoder. <br>
This allows more features or alternate decision rules.
- [Reranking (Statmt)](http://www.statmt.org/survey/Topic/Reranking)
- [Energy-Based Reranking: Improving Neural Machine Translation Using Energy-Based Models (Sumanta Bhattacharyya et al.)](https://arxiv.org/abs/2009.13267)
- [Neural Machine Translation: A Review and Survey (Felix Stahlberg)](https://arxiv.org/pdf/1912.02047.pdf)
- [A Comparable Study on Model Averaging, Ensembling and Reranking in NMT (Y. Liu et al.)](https://www.semanticscholar.org/paper/A-Comparable-Study-on-Model-Averaging%2C-Ensembling-Liu-Zhou/1df3667e81e37fd6fcb9485514622b9e78fa8161?sort=relevance&page=2)
## Study Resources
- [Unsupervised Sub-tree Alignment for Tree-to-Tree Translation (paper)](https://www.jair.org/index.php/jair/article/view/10850/25893)
- [PyTorch for Deep Learning - Full Course (FreeCodeCamp Video)](https://www.youtube.com/watch?v=GIsg-ZUy0MY&t=205s)
- [Jovian.ai Notebooks](https://jovian.ai/aakashns/03-logistic-regression)
- [Understanding LSTMs (Colah)](http://colah.github.io/posts/2015-08-Understanding-LSTMs/)
- [Attention Mechanism (Wikidocs 딥 러닝을 이용한 자연어 처리 입문)](https://wikidocs.net/22893)
- [BLEU (Wikidocs)](https://wikidocs.net/31695)
- [TreeLSTM (Slideshare)](https://www.slideshare.net/tuvistavie/tree-lstm)
- [TreeLSTM pytorch implementation](https://github.com/dasguptar/treelstm.pytorch/blob/master/treelstm/dataset.py)
- [Dependency vs Constituency Parsing](https://www.baeldung.com/cs/constituency-vs-dependency-parsing)
- [Pointer Networks 논문 리뷰 (티스토리)](https://ropiens.tistory.com/57)
## Dependency Tree-LSTM Structure
<p style="display:inline-block"><img src="attachment:image.png" width="426" ></p> <p style="display:inline-block"><img src="attachment:image-2.png" width="426"><p>
## Activation/Loss Functions
- [Why Sigmoid?](https://stats.stackexchange.com/questions/162988/why-sigmoid-function-instead-of-anything-else/318209#318209)
- [Why Softmax?](https://stackoverflow.com/questions/17187507/why-use-softmax-as-opposed-to-standard-normalization)
- [Cross Entropy](https://machinelearningmastery.com/cross-entropy-for-machine-learning)
Uncategorized
- [NYU Drive (ms9144)](https://drive.google.com/drive/folders/1zhXsGz1C9AikH6t1ADj0IosTQGaVn9kV?usp=sharing)
- [Neural Networks, Manifolds, and Topology (colah)](http://colah.github.io/posts/2014-03-NN-Manifolds-Topology/)
- [How to make a C/C++ extension to Python](https://medium.com/@matthiasbitzer94/how-to-extend-python-with-c-c-code-aa205417b2aa)
- [Git 서브모듈](https://ohgyun.com/711)
- [deeplearningtheory.com (physics-based interpretations)](https://deeplearningtheory.com/PDLT.pdf)
- [Graph neural networks: A review of methods and applications (Elsevier paper)](https://arxiv.org/ftp/arxiv/papers/1812/1812.08434.pdf)
- [Boyd CVXOPT (book, slides)](https://web.stanford.edu/~boyd/cvxbook/)
## Some Combinatorial Problems
<p style="display:inline-block"><img src="attachment:image.png" width="600" ></p>
$$
\begin{align*}
\mathbf{u}_i &= A \mathbf{c}_i & \; i = 1, \ldots, n \\n
\mathbf{v} &= B \mathbf{q} \\n
\mathbf{p} &= \text{softmax}_i(\mathbf{v}^T \mathbf{u}_i) \\n
\mathbf{z} &= \sum_i p_i \mathbf{c}_i
\end{align*}
$$
| github_jupyter |
Extinction Profile
=====================
Extinction profiles is a recently proposed technique used for the classification of remote sensing data. The main publication describing the method is the following:
**P Ghamisi, R Souza, JA Benediktsson, XX Zhu, L Rittner, RA Lotufo. " Extinction profiles for the classification of remote sensing data," IEEE Transactions on Geoscience and Remote Sensing 54 (10), 5631-5645.**
In this demo, we illustrate the computation of an area extinction profile of a satellite image.
Loading the image and defining profile connectivity
----------------------------------------------------------
Lines 1 htrough 13 import the necessary libraries to run this demo. Line 17 load the satellite image. Lines 26 through 29 define the structuring element with connectivity-4. Line 32 defines the parameters to be used to compute the extinction profile. Line 39 declares the array to store the extinction profile.
```
# This makes plots appear in the notebook
%matplotlib inline
import numpy as np # numpy is the major library in which siamxt was built upon
# we like the array programming style =)
# We are using PIL to read images
from PIL import Image
# and matplotlib to display images
import matplotlib.pyplot as plt
import siamxt
# Loading the image.
# Make sure the image you read is either uint8 or uint16
data = np.asarray(Image.open("./Sample-images/sattelite.jpg").convert("L"))
print("Image dimensions: %dx%d pixels" %data.shape)
#Displaying the image
fig = plt.figure()
plt.imshow(data, cmap='Greys_r')
plt.axis('off')
plt.title("Original sattelite image")
#Structuring element. connectivity-4
Bc = np.zeros((3,3),dtype = bool)
Bc[1,:] = True
Bc[:,1] = True
# Parameters used to compute the extinction profile
nextrema = [int(2**jj) for jj in range(7)][::-1]
print("Nb. of extrema used to compute the profile:")
print(nextrema)
# Array to store the profile
H,W = data.shape
Z = 2*len(nextrema)+1
ep = np.zeros((H,W,Z))
```
Min-tree Profile
------------------
Initially, we compute the profile for the min-tree (max-tre of the negated image). Lines 1 through 3 negate the image. Line 6 builds the min-tree. Line 9 extracts the area attribute from the node array structure and line 10 computes the area extinction values.
The loop in lines 14 through 18 compute the extinction filter for the different parameters of the profile and assign the results to the variable *ep*.
```
#Negating the image
max_value = data.max()
data_neg = (max_value - data)
# Building the max-tree of the negated image, i.e. min-tree
mxt = siamxt.MaxTreeAlpha(data_neg,Bc)
# Area attribute extraction and computation of area extinction values
area = mxt.node_array[3,:]
Aext = mxt.computeExtinctionValues(area,"area")
# Min-tree profile
i = len(nextrema) - 1
for n in nextrema:
mxt2 = mxt.clone()
mxt2.extinctionFilter(Aext,n)
ep[:,:,i] = max_value - mxt2.getImage()
i-=1
# Putting the original image in the profile
i = len(nextrema)
ep[:,:,i] = data
i +=1
```
Max-tree Profile
------------------
Then, we compute the profile for the max-tree. Lines 1 through 3 negate the image. Line 2 builds the min-tree. Line 5 extracts the area attribute from the node array structure and line 6 computes the area extinction values.
The loop in lines 9 through 13 computes the extinction filter for the different parameters of the profile and assign the results to the variable *ep*.
```
#Building the max-tree
mxt = siamxt.MaxTreeAlpha(data,Bc)
# Area attribute extraction and computation of area extinction values
area = mxt.node_array[3,:]
Aext = mxt.computeExtinctionValues(area,"area")
# Max-tree profile
for n in nextrema:
mxt2 = mxt.clone()
mxt2.extinctionFilter(Aext,n)
ep[:,:,i] = mxt2.getImage()
i+=1
```
Displaying the profile mosaic
--------------------------------
Finally, the code below puts the images in the profile side by side and dispaly it. SInce, this code is not related to max-tree, we will ommit its explanation.
```
# Number of cloumns in the mosaic
N = 3
# Computing the dimensions of the mosiac image
H2 = Z//N
if (Z%N != 0):
H2+=1
H2 = H2*H
W2 = N*W
# Mosaic image initialization
mosaic = np.zeros((H2,W2), dtype = ep.dtype)
i,j = 0,0
for ii in range(Z):
mosaic[i*H:(i+1)*H,j*W:(j+1)*W] = ep[:,:,ii]
j+=1
if (j%N == 0):
j = 0
i+=1
plt.rcParams['figure.figsize'] = 16, 12
#Displaying the profile mosaic
fig = plt.figure()
plt.imshow(mosaic, cmap='Greys_r')
plt.axis('off')
plt.title("Extinction Profile Mosaic")
```
| github_jupyter |
# image classification
We will use the CIFAR10 dataset: https://www.cs.toronto.edu/~kriz/cifar.html
```
from tensorflow import keras
keras.__version__
(train_images, train_labels), (test_images, test_labels) = keras.datasets.cifar10.load_data()
train_images.shape
train_images.dtype
n_images = 5000
train_images = train_images[:n_images]
train_labels = train_labels[:n_images]
train_images.shape
train_images.min(), train_images.max()
train_labels.shape
train_labels.min(), train_labels.max()
train_labels.dtype
train_images = train_images / 255.
test_images = test_images / 255.
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
import matplotlib.pyplot as plt
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.axis('off')
plt.title(class_names[train_labels[i,0]])
plt.show()
image_dim = train_images.shape[1]*train_images.shape[2]*train_images.shape[3]
print(image_dim)
```
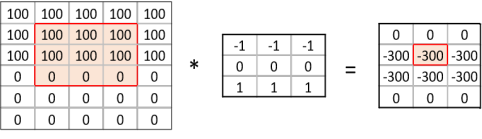

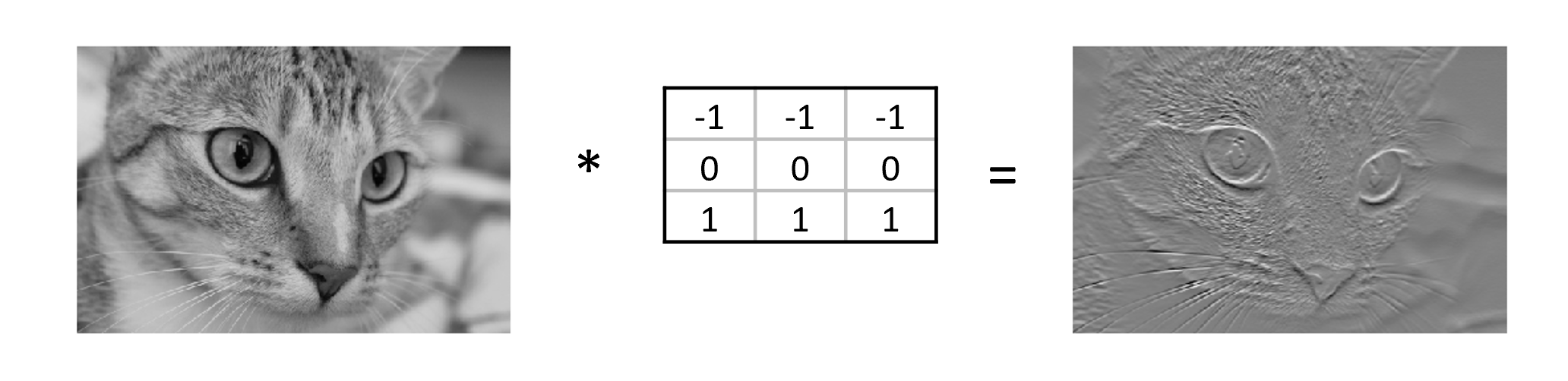
```
def create_cnn1():
inputs = keras.Input(shape=train_images.shape[1:])
conv1 = keras.layers.Conv2D(32, (3,3), activation='relu')(inputs)
conv2 = keras.layers.Conv2D(32, (3,3), activation='relu')(conv1)
flat = keras.layers.Flatten()(conv2)
outputs = keras.layers.Dense(10)(flat)
return keras.Model(inputs=inputs, outputs=outputs, name="cifar_model_small")
model = create_cnn1()
model.summary()
def create_cnn2():
inputs = keras.Input(shape=train_images.shape[1:])
conv1 = keras.layers.Conv2D(32, (3,3), activation='relu')(inputs)
pool1 = keras.layers.MaxPool2D((2,2))(conv1)
conv2 = keras.layers.Conv2D(32, (3,3), activation='relu')(pool1)
pool2 = keras.layers.MaxPool2D((2,2))(conv2)
flat = keras.layers.Flatten()(pool2)
outputs = keras.layers.Dense(10)(flat)
return keras.Model(inputs=inputs, outputs=outputs, name="cifar_model_small")
model = create_cnn2()
model.summary()
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(train_images, train_labels, epochs=10,
validation_data=(test_images, test_labels))
import seaborn as sns
import pandas as pd
history_df = pd.DataFrame.from_dict(history.history)
print(history_df.columns)
sns.lineplot(data=history_df[['accuracy','val_accuracy']])
sns.lineplot(data=history_df[['loss','val_loss']])
```
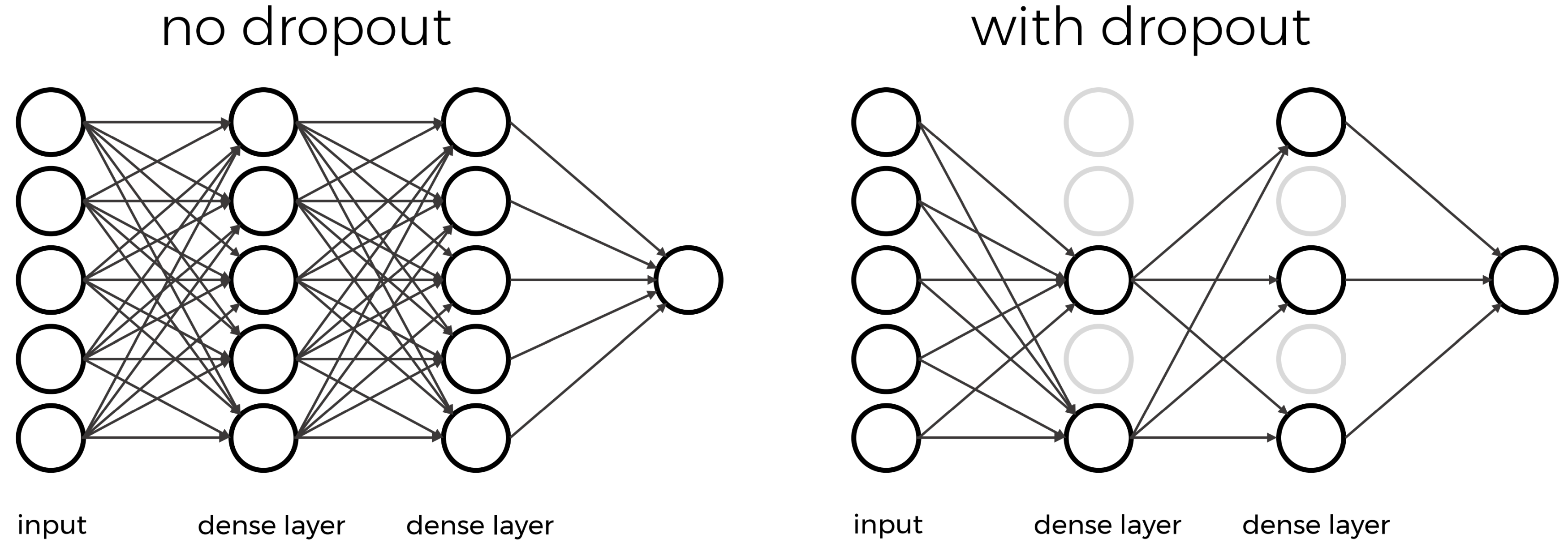
```
def create_cnn3():
inputs = keras.Input(shape=train_images.shape[1:])
conv1 = keras.layers.Conv2D(32, (3,3), activation='relu')(inputs)
pool1 = keras.layers.MaxPool2D((2,2))(conv1)
conv2 = keras.layers.Conv2D(32, (3,3), activation='relu')(pool1)
pool2 = keras.layers.MaxPool2D((2,2))(conv2)
dropped = keras.layers.Dropout(0.2)(pool2)
flat = keras.layers.Flatten()(dropped)
outputs = keras.layers.Dense(10)(flat)
return keras.Model(inputs=inputs, outputs=outputs, name="cifar_model_small_withdropout")
model = create_cnn3()
model.summary()
model.compile(optimizer='adam',
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(train_images, train_labels, epochs=10,
validation_data=(test_images, test_labels))
history_df = pd.DataFrame.from_dict(history.history)
history_df['epoch'] = range(1,len(history_df)+1)
history_df = history_df.set_index('epoch')
sns.lineplot(data=history_df[['accuracy', 'val_accuracy']])
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
sns.lineplot(data=history_df[['loss', 'val_loss']])
```
| github_jupyter |
# ***Introduction to Radar Using Python and MATLAB***
## Andy Harrison - Copyright (C) 2019 Artech House
<br/>
# **Apparent Elevation**
***
Referring to Section 2.7.1, a signal transmitted through the atmosphere bends toward the earth. In many radar applications, this would require transmission of the signal at a higher elevation angle in order to have the energy intercept the target, as shown in Figure 2.9. This angle is referred to as the apparent elevation angle, as the target appears to be at a different angle than its true position.
The amount of correction to the true target elevation is
$$
\Delta \theta = -\int\limits_{h}^{\infty} \frac{n'(z)}{n(z) \tan \phi}\,dz \hspace{0.5in} \mathrm{(deg)}
$$
where
$$
\cos \phi = \frac{c}{(r_e+z) \, n(z)}, \hspace{0.2in} c = (r_e + h) \, n(h) \cos \theta.
$$
Since the refraction in the atmosphere is largely determined by the lower layers, a model based on the exponential atmosphere for terrestrial propagation allows the index of refraction at some altitude, $z$, to be expressed as
$$
n(z) = 1 + \alpha \, e^{-\beta z},
$$
where $\alpha = 0.000315$, and $\beta = 0.1361$.
***
Begin by getting the library path
```
import lib_path
```
Set the true elevation (degrees) and the height (km) and create an array of 100 values using the `linspace` routine from `scipy`
```
from numpy import linspace
true_elevation = 20.0
max_height = 5
height = linspace(0, max_height, 100)
```
Calculate the apparent elevation for each height value
```
from Libs.wave_propagation import refraction
apparent_elevation = [refraction.apparent_elevation(true_elevation, h) for h in height]
```
Also calculate the apparent elevation with the approximate method
```
kwargs = {'theta_true': true_elevation, 'height': height}
apparent_elevation_approximate = refraction.apparent_elevation_approximate(**kwargs)
```
Display the apparent elevation due to refraction using the `matplotlib` routines
```
from matplotlib import pyplot as plt
# Set the figure size
plt.rcParams["figure.figsize"] = (15, 10)
# Display the results
plt.plot(height, apparent_elevation, label='Integration')
plt.plot(height, apparent_elevation_approximate, '--', label = 'Approximate')
# Set the plot title and labels
plt.title('Apparent Elevation due to Refraction', size=14)
plt.xlabel('Height (km)', size=12)
plt.ylabel('Apparent Elevation Angle (degrees)', size=12)
# Set the tick label size
plt.tick_params(labelsize=12)
# Turn on the legend
plt.legend(loc='upper right', prop={'size': 10})
# Turn on the grid
plt.grid(linestyle=':', linewidth=0.5)
```
| github_jupyter |
# ENDF Files
## Library class
Below is an example of how to grab and graph cross section data from ENDF files using the `Library` class.
```
%matplotlib inline
import os
import requests
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import HTML
from tabulate import tabulate
from pyne.endf import Library, Evaluation
if not os.path.isfile("U235-VII.txt"):
url = "http://t2.lanl.gov/nis/data/data/ENDFB-VII.1-neutron/U/235"
r = requests.get(url, allow_redirects=True)
with open("U235-VII.txt", "wb") as outfile:
outfile.write(r.content)
u235 = Library("U235-VII.txt")
xs_data = u235.get_xs(922350000, 16)[0]
fig = plt.figure()
Eints, sigmas = xs_data['e_int'], xs_data['xs']
plt.step(Eints, sigmas, where = "pre")
plt.suptitle(r'(n, 2n) Reaction in $^{235}$U')
plt.ylabel(r'$\sigma(E)$ (barns)')
plt.xlabel(r'$E_{int} (eV)$')
plt.xscale('log')
plt.yscale('log')
plt.savefig('u235_2n.eps')
if not os.path.isfile("U238-VII.txt"):
url = "http://t2.lanl.gov/nis/data/data/ENDFB-VII.1-neutron/U/238"
r = requests.get(url, allow_redirects=True)
with open("U238-VII.txt", "wb") as outfile:
outfile.write(r.content)
u238 = Library("U238-VII.txt")
xs_data = u238.get_xs(922380000, 1)[0]
fig = plt.figure()
Eints, sigmas = xs_data['e_int'], xs_data['xs']
plt.step(Eints, sigmas, where = "pre")
plt.suptitle(r'Total Cross Section for $^{238}$U')
plt.ylabel(r'$\sigma(E)$ (barns)')
plt.xlabel(r'$E_{int} (eV)$')
plt.xlim(xmin = 10000)
plt.xscale('log')
plt.yscale('log')
```
## Evaluation class
The `pyne.endf.Evaluation` class provides a facility for parsing data in an ENDF file. Parsing of all data other than covariances (MF=30+) is supported has been tested against the ENDF/B-VII.1 neutron, photoatomic, electroatomic, atomic relaxation, and photonuclear sublibraries. In this example, we will use the `Evaluation` class to look at typical data in the ENDF/B-VII.1 evaluation of U-235.
```
u235 = Evaluation("U235-VII.txt")
```
By default, when an `Evaluation` is instantiated, only the descriptive data in MF=1, MT=451 is parsed. This allows us to get basic information about an evaluation without necessarily reading the whole thing. This useful data can be found in the `info` and `target` attributes.
```
u235.info
u235.target
```
To look at cross sections, secondary energy and angle distributions, and resonance data, we need to parse the rest of the data in the file, which can be done through the `Evaluation.read(...)` method.
```
u235.read()
```
Most of the data that is parsed resides in the `reactions` attribute, which is a dictionary that is keyed by the MT value.
```
elastic = u235.reactions[2]
print('Elastic scattering has the following attributes:')
for attr in elastic.__dict__:
if elastic.__dict__[attr]:
print(' ' + attr)
```
Now with our reaction we can look at the cross section and any other associated data. The cross section `elastic.xs` is a `Tab1` object whose (x,y) pairs can be accessed from the `x` and `y` attributes. The first ten values of the cross section are:
```
zip(elastic.xs.x[:10], elastic.xs.y[:10])
```
Since resonances haven't been reconstructed, everything below the unresolved resonance range at 2250 keV is zero. Above that energy, we can use `elastic.xs` like a function to get a value at a particular energy. For example, to get the elastic cross section at 1 MeV:
```
elastic.xs(1.0e6)
```
We can also take a look at the angular distribution for elastic scattering.
```
esad = elastic.angular_distribution
print(esad)
# Elastic scattering angular distribution at 100 keV
E = esad.energy[5]
pdf = esad.probability[5]
theta = np.linspace(0., 2*np.pi, 1000)
mu = np.cos(theta)
plt.subplot(111, polar=True)
plt.plot(theta, pdf(mu))
```
Ah, but elastic scattering is a simple reaction you say. What if I want information about something more complicated like fission! In the special case of fission, there is the normal reaction data as well as a special attribute on the Evaluation class called `fission`:
```
print(u235.reactions[18])
print(u235.fission.keys())
```
We can look at the neutrons released per fission:
```
E = np.logspace(-5, 6)
plt.semilogx(E, u235.fission['nu']['total'](E))
plt.xlabel('Energy (eV)')
plt.ylabel('Neutrons per fission')
```
The components of energy release from fission are also available to us:
```
for component, coefficients in u235.fission['energy_release'].items():
if component != 'order':
print('{}: {} +/- {} MeV'.format(component, coefficients[0,0], coefficients[1,0]))
```
To look at the fission energy distribution, we must use the normal reaction data:
```
# Get prompt fission neutron spectra
fission = u235.reactions[18]
pfns = fission.energy_distribution[0]
# Plot the distribution for the lowest incoming energy
plt.semilogx(pfns.pdf[0].x, pfns.pdf[0].y)
plt.xlabel('Energy (eV)')
plt.ylabel('Probability / eV')
plt.title('Neutron spectrum at E={} eV'.format(pfns.energy[0]))
```
Finally, let's take a look at resolved resonance data, which can be found in the `resonances` dictionary.
```
rrr = u235.resonances['resolved']
print(rrr)
# Show all (l,J) combinations
print(rrr.resonances.keys())
# Set up headers for table
headers = ['Energy', 'Neutron width', 'Capture width', 'FissionA width', 'FissionB width']
# Get resonance data for l=0, J=3
l = 0
J = 3.0
# Create table data
data = [[r.energy, r.width_neutron, r.width_gamma, r.width_fissionA, r.width_fissionB]
for r in rrr.resonances[l,J]]
# Render table
HTML(tabulate(data, headers=headers, tablefmt='html'))
```
| github_jupyter |
# Inverted Indexing and Index Compression
Build the inverted index for the following documents:
* ID1 : Selenium is a portable framework for testing web applications
* ID2 : Beautiful Soup is useful for web scraping
* ID3: It is a python package for parsing the pages
Perform Index Compression for the integer values in the inverted index (duplicates to be eliminated) using Elias delta coding and variable byte scheme.
```
# Import statements
import math
import re
import string
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
# Documents list
documents = [
"Selenium is a portable framework for testing web applications",
"Beautiful Soup is useful for web scraping",
"It is a python package for parsing the pages"
]
```
## 1. Inverted Index Construction
### 1.1 Pre-processing
Refer to this link for more details : [Text Preprocessing Reference](https://medium.com/@datamonsters/text-preprocessing-in-python-steps-tools-and-examples-bf025f872908)
```
def preprocess(text) :
"""
Given a text, we pre-process and return an array of the words from the text.
"""
s = text
# Convert to lower case
s = s.lower()
# Removing numbers and other numerical data
# We substitute all the occurances of numbers by an empty string, thereby effectively removing them.
s = re.sub(r'\d+', '', s)
# Remove punctuation signs
#s = s.translate(string.maketrans("",""), string.punctuation)
s = s.replace('/[^A-Za-z0-9]/g', '')
# Trim the leading and trailing spaces
s = s.strip()
# Tokenize the text
words = word_tokenize(s)
# Stop Word Removal
stop_words = set(stopwords.words('english'))
words = [word for word in words if word not in stop_words]
# Return the word list
return words
```
### 1.2 Find Occurance Function
```
def findOccurance(text, word) :
"""
Given a text and the word to be found, we partially pre-process the text and then return the count of occurances of the word,
and the positions in the text where they occur.
"""
# Split the text into tokens and remove the punctuation signs and convert to lower case
# This is to find the position of the words to have in the inverted index
text = text.replace('/[^A-Za-z0-9]/g', '')
text = text.replace(' ', ' ')
text = text.lower()
text_words = text.strip().split()
word_count = 0
word_positions = []
for i in range(len(text_words)) :
if word == text_words[i] :
word_count += 1
word_positions.append(i)
return (word_count, word_positions)
```
### 1.3 Inverted Indexing
```
inverted_index = {}
# Process each of the documnet
for (i, doc) in enumerate(documents) :
# Pre-Processing of each individual document
words = preprocess(doc)
# Add the words into the inverted index
for word in words :
# Create an entry for the word if one does not exist
if word not in inverted_index :
inverted_index[word] = []
# Find all the occurances of the word in the doc.
occurance_count, occurance_pos_list = findOccurance(doc, word)
# Add these details into the inverted index
inverted_index[word].append(((i+1), occurance_count, occurance_pos_list))
```
Format for the inverted index is :
* inverted index :
```python
{
word : [
(document_id, number_of_occurances_in_document, [offset_of_occurances]),
...
],
...
}
```
```
print('Inverted Index : ')
for item in inverted_index.items() :
print(item)
```
## 2. Index Compression
For every unique number in the index, we create a map of the number to a encoded version of the number which occupies a lower size, thereby ensuring compression.
### 2.1 Binary Conversion
```
def binary(n) :
"""
Given an integer number returns the equivalent binary string.
"""
# Convert to binary string
num = bin(n)
# Remove the `0b` which is present in the front of the string
num = num[2:]
return num
```
### 2.2 Elias Gamma Encoding
```
def eliasGammaEncoding(n) :
"""
Given an integer number `n`, we encode the number using the `Elias Gamma Encoding` scheme, and return the compressed value as a string.
"""
# Zero is already encoded
if n == 0 :
return "0"
# Find the binary value of number
num = binary(n)
# Prepend the value with (length-1) zeros
num = ('0' * (len(num) - 1)) + num
return num
```
### 2.3 Elias Delta Encoding
```
def eliasDeltaEncoding(n) :
"""
Given an integer number `n`, we encode the number using the `Elias Delta Encoding` scheme, and return the compressed value as a string.
"""
# Zero is already encoded
if n == 0 :
return "0"
# Find the gamma code for (1 + log2(n))
num1 = 1 + int(math.log2(n))
num1 = eliasGammaEncoding(num1)
# Number in binary form after removing the MSB
num2 = binary(n)
num2 = str(num2)[1:]
# Combine the gamma code and the other code value
num = num1 + num2
return num
```
### 2.4 Variable Byte Encoding Scheme
```
def variableByteEncoding(n) :
"""
Given an integer number `n`, we encode the number using the `Variable Byte Encoding` scheme, and return the compressed value as a string.
"""
# Convert the number into binary form
s = binary(n)
result = ""
while len(s) > 0 :
# Get the term and update the binary string
if len(s) > 7 :
term = s[-7:]
s = s[:-7]
else :
term = s
s = ""
term = ("0" * (7 - len(term))) + term
if len(result) == 0 :
result = term + "0"
else :
result = term + "1" + result
return result
```
### 2.5 Index Compression Function
```
def indexCompression(inverted_index, encoding_scheme) :
"""
Given an inverted index, we perform compression for all the integers in the inverted index and return the encoding map.
"""
compression_map = {}
for word_indices in inverted_index.values() :
for word_index in word_indices :
# Prepare an array to have all the numbers involved in this
i, count, positions = word_index
arr = [i, count] + positions
# For each number compute and store the elias delta encoded value if not already present
for n in arr :
if n not in compression_map :
if encoding_scheme == 'ELIAS_DELTA' :
compression_map[n] = eliasDeltaEncoding(n)
elif encoding_scheme == 'VARIABLE_BYTE' :
compression_map[n] = variableByteEncoding(n)
return compression_map
```
### 2.6 Index Compression By Elias Delta
We perform compression for all the numbers using `Elias Delta` encoding scheme in the inverted index created in `Section 1`
```
elias_delta_compression_map = indexCompression(inverted_index, 'ELIAS_DELTA')
print("Elias Delta Encoding Map :")
for item in elias_delta_compression_map.items() :
print(item)
```
### 2.7 Index Compression By Variable Byte Encoding
We perform compression for all the numbers using `Variable Byte` encoding scheme in the inverted index created in `Section 1`
```
variable_byte_compression_map = indexCompression(inverted_index, 'VARIABLE_BYTE')
print("Variable Byte Encoding Map :")
for item in variable_byte_compression_map.items() :
print(item)
```
| github_jupyter |
```
from IPython.core.display import display, HTML, Markdown
import copy
p_original_instance = [
[0, 0, 1, 0, 1, 1, 0, 1],
[0, 0, 1, 0, 0, 1, 1, 0],
[1, 1, 0, 1, 1, 0, 0, 0],
[1, 1, 0, 1, 0, 1, 1, 0],
[0, 0, 1, 0, 0, 1, 0, 1],
[1, 0, 1, 0, 1, 1, 0, 1]
]
def visualizza(p):
num_accese = 0;
for r in range(len(p)):
for c in range(len(p[0])):
print(p[r][c],end=" ")
num_accese += p[r][c]
print()
display(Markdown(f"Attualmente ci sono <b>{num_accese} luci accese</b>."))
def visualizza_lo_stato_che_si_ottiene_con_gli_interruttori_impostati(agisci_riga, agisci_col):
new_p = copy.deepcopy(p_original_instance)
num_accese = 0;
for r in range(len(agisci_riga)):
for c in range(len(agisci_col)):
if (agisci_riga[r] + agisci_col[c]) % 2 != 0:
new_p[r][c] = 1-new_p[r][c]
num_accese += new_p[r][c]
display(Markdown(f"Agendo sugli interruttori settati come segue:<br><b> Interruttori di riga:</b> {agisci_riga}<br><b> Interruttori di colonna:</b> {agisci_col}"))
display(Markdown(f"Ti porti dalla configurazione iniziale riportata in $p$ nella seguente configurazione finale:<br>"))
visualizza(new_p)
```
## Esercizio \[60 pts\]
(pirellone) Partendo dalla seguente matrice di $m\times n$ valori booleani (acceso/spento), utilizza quelle che credi delle $m+n$ mosse che invertono tutta una riga oppure tutta una colonna. Il tuo scopo è minimizzare il numero di luci che trovi accese alla fine.
```
# Ecco il pirellone in input:
visualizza(p)
```
__Richieste__:
[15pt] Portati in una configurazione col minor numero possibile di luci accese. Puoi effettuare una mossa alla volta utilizzando le funzioni $flippa\_riga(indice\_riga)$ e $flippa\_colonna(indice\_colonna)$ per spostarti un pò alla volta fino ad una configurazione che reputi ottima.
```
# Setta su quali degli m+n interruttori di riga e di colonna intendi agire.
# Puoi fare questo sostituendo gli 0 (=non agire) con degli 1 (=agire) nelle posizioni corrispondenti
# (stiamo parlando dei soli m+n numeri di colore diverso).
# Quando hai finito premi Shift-Invio per visualizzare lo stato che otterresti ed ottenere una valutazione.
agisci_col = [0, 0, 1, 0, 0, 1, 0, 1]
agisci_riga = [
0, # [0, 0, 1, 0, 1, 1, 0, 1],
0, # [0, 0, 1, 0, 0, 1, 1, 0],
1, # [1, 1, 0, 1, 1, 0, 0, 0],
1, # [1, 1, 0, 1, 0, 1, 1, 0],
0, # [0, 0, 1, 0, 0, 1, 0, 1],
0 # [1, 0, 1, 0, 1, 1, 0, 1]
]
visualizza_lo_stato_che_si_ottiene_con_gli_interruttori_impostati(agisci_riga, agisci_col)
```
[30pt]
Nel quadro sotto, fornisci argomenti conclusivi per certificare che non è possibile spegnere un numero maggiore di celle. Faccio presente che questo problema di ottimizzazione non è noto essere in P e pertanto, ove fosse NP-hard, resterebbe allora inesauribile spazio alla tua creatività nel produrre questi argomenti.
Si considerino le seguenti $8$ matrici cattive $2\times 2$ (e si noti che non hanno alcuna cella in comune):
righe: 5,4 colonne: 0,1
righe: 5,4 colonne: 4,2
righe: 3,2 colonne: 4,3
righe: 3,2 colonne: 5,2
righe: 2,3 colonne: 6,7
righe: 1,2 colonne: 6,1
righe: 1,2 colonne: 7,2
righe: 0,1 colonne: 4,3
Osserviamo infine, cosa che rende il nostro certificato ancora più piacevole e magico, che ogni luce che abbiamo lasciato accesa nella soluzione sopra data trova posto in precisamente una delle $8$ sottomatrici $2\times 2$ quì listate (abbiamo avuto cura di porre per prima la riga e la colonna dell'unico $1$ in ciascuna di queste matrici). Valgono cioè quì, almeno su questa istanza, le condizioni degli scarti complementari.
Si potrebbe sperare di mettere il problema in P (ma ricordo che un tempo, che mi era stato chiesto, ero riuscito a dare una qualche dimostrazione che mi pareva convincermi dell'NP-hardness di questo problema di ottimizzazione).
| github_jupyter |
```
import pandas as pd
import numpy as np
import random as rnd
from sklearn.cross_validation import KFold, cross_val_score
# machine learning
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.tree import DecisionTreeClassifier
train_=pd.read_csv('../train_allcols.csv')
validate_=pd.read_csv('../validate_allcols.csv')
#test=pd.read_csv('../testwDSM.csv')
train_.shape, validate_.shape, #test.shape
train = train_.query('DSMCRIT in [4,9]')
validate = validate_.query('DSMCRIT in [4,9]')
print train['DSMCRIT'].value_counts()
print validate.shape
#alcohol
#print train['DSMCRIT'].value_counts() / train['DSMCRIT'].count()
#print train['SUB1'].value_counts() / train['SUB1'].count()
#train.query('SUB1 == 4')['DSMCRIT'].value_counts() / train.query('SUB1 == 4')['DSMCRIT'].count()
#train.describe()
train = train.sample(10000)
validate = validate.sample(3000)
train.shape, #validate.shape, #validate.head(2)
#train = train.query('SUB1 <= 10').query('SUB2 <= 10')
#validate = validate.query('SUB1 <= 10').query('SUB2 <= 10')
drop_list = ['DSMCRIT', #'NUMSUBS'
]
drop_list_select = ['RACE', 'PREG', 'ARRESTS', 'PSYPROB', 'DETNLF', 'ETHNIC', 'MARSTAT', 'GENDER', 'EDUC'
,'LIVARAG', 'EMPLOY', 'SUB3']
retain_list = ['RACE','PCPFLG','PRIMINC','LIVARAG','BENZFLG','HLTHINS','GENDER','ROUTE3','PRIMPAY',
'MARSTAT','PSYPROB','ROUTE2','EMPLOY','SUB2','FRSTUSE3','FREQ3','FRSTUSE2','OTHERFLG',
'EDUC','FREQ2','FREQ1','YEAR',
'PSOURCE','DETCRIM','DIVISION','REGION','NOPRIOR','NUMSUBS','ALCDRUG',
'METHUSE','FRSTUSE1','AGE','COKEFLG','OPSYNFLG','IDU','SERVSETA','ROUTE1','MARFLG',
'MTHAMFLG','HERFLG',
'ALCFLG','SUB1']
X_train = train[retain_list]
#X_train = train.drop(drop_list + drop_list_select, axis=1)
Y_train = train["DSMCRIT"]
X_validate = validate[retain_list]
Y_validate = validate["DSMCRIT"]
#X_test = test.drop(drop_list, axis=1)
X_train.shape, #X_validate.shape, #X_test.shape
print X_train.columns.tolist()
from sklearn.feature_selection import SelectKBest, SelectPercentile
from sklearn.feature_selection import f_classif,chi2
#Selector_f = SelectPercentile(f_classif, percentile=25)
Selector_f = SelectKBest(f_classif, k=10)
Selector_f.fit(X_train,Y_train)
zipped = zip(X_train.columns.tolist(),Selector_f.scores_)
ans = sorted(zipped, key=lambda x: x[1])
for n,s in ans:
print 'F-score: %3.2ft for feature %s' % (s,n)
#X_train= SelectKBest(f_classif, k=10).fit_transform(X_train, Y_train)
#one hot
from sklearn import preprocessing
# 1. INSTANTIATE
enc = preprocessing.OneHotEncoder()
# 2. FIT
enc.fit(X_train)
# 3. Transform
onehotlabels = enc.transform(X_train).toarray()
X_train = onehotlabels
onehotlabels = enc.transform(X_validate).toarray()
X_validate = onehotlabels
X_train.shape, #X_validate.shape
#kfold
kf = 3
# Logistic Regression
logreg = LogisticRegression(n_jobs=-1)
logreg.fit(X_train, Y_train)
#Y_pred = logreg.predict(X_test)
l_acc_log = cross_val_score(logreg, X_train, Y_train, cv=kf)
acc_log = round(np.mean(l_acc_log), 3)
l_acc_log = ['%.3f' % elem for elem in l_acc_log]
print l_acc_log
print acc_log
# Random Forest (slow)
random_forest = RandomForestClassifier(n_estimators=200, max_depth=20, n_jobs=-1)
random_forest.fit(X_train, Y_train)
#Y_pred = random_forest.predict(X_test)
l_acc_random_forest = cross_val_score(random_forest, X_train, Y_train, cv=kf)
acc_random_forest = round(np.mean(l_acc_random_forest), 3)
l_acc_random_forest = ['%.3f' % elem for elem in l_acc_random_forest]
print l_acc_random_forest
print acc_random_forest
# Linear SVC
linear_svc = LinearSVC(C=1.0)
linear_svc.fit(X_train, Y_train)
#Y_pred = linear_svc.predict(X_test)
l_acc_linear_svc = cross_val_score(linear_svc, X_train, Y_train, cv=kf)
acc_linear_svc = round(np.mean(l_acc_linear_svc), 3)
l_acc_linear_svc = ['%.3f' % elem for elem in l_acc_linear_svc]
print l_acc_linear_svc
print acc_linear_svc
print 'predict-sub2-woflags-newsplit-sample20000'
models = pd.DataFrame({
'Model': ['Logistic Regression',
'Random Forest','Linear SVC'],
'Cross Validation': [l_acc_log,
l_acc_random_forest, l_acc_linear_svc],
'Cross Validation Mean': [acc_log,
acc_random_forest, acc_linear_svc]
})
print models.sort_values(by='Cross Validation Mean', ascending=False)
import matplotlib.pyplot as plt
import seaborn as sns
Y_pred = random_forest.predict(X_validate)
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(Y_validate, Y_pred, labels=[3,4,5,6,7,8,9,10])
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Compute confusion matrix
cnf_matrix = cm #confusion_matrix(y_test, Y_pred)
#class_names = ["ANXIETY","DEPRESS","SCHIZOPHRENIA","BIPOLAR","ATTENTION DEFICIT"]
class_names = [3,4,5,6,7,8,9,10]
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, class_names,
title='Confusion Matrix, without normalization')
#plt.savefig('cnf matrix', dpi=150)
plt.show()
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=class_names, normalize=True,
title='Normalized Confusion Matrix')
#plt.figure(figsize=(16,8))
#plt.savefig('cnf matrix norm', dpi=150)
plt.show()
print X_validate.shape,Y_pred.shape, Y_validate.shape
print round(random_forest.score(X_validate, Y_validate) * 100, 2)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
**BikeShare Demand Forecasting**
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Compute](#Compute)
1. [Data](#Data)
1. [Train](#Train)
1. [Featurization](#Featurization)
1. [Evaluate](#Evaluate)
## Introduction
This notebook demonstrates demand forecasting for a bike-sharing service using AutoML.
AutoML highlights here include built-in holiday featurization, accessing engineered feature names, and working with the `forecast` function. Please also look at the additional forecasting notebooks, which document lagging, rolling windows, forecast quantiles, other ways to use the forecast function, and forecaster deployment.
Make sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.
Notebook synopsis:
1. Creating an Experiment in an existing Workspace
2. Configuration and local run of AutoML for a time-series model with lag and holiday features
3. Viewing the engineered names for featurized data and featurization summary for all raw features
4. Evaluating the fitted model using a rolling test
## Setup
```
import azureml.core
import pandas as pd
import numpy as np
import logging
from azureml.core import Workspace, Experiment, Dataset
from azureml.train.automl import AutoMLConfig
from datetime import datetime
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.23.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
```
As part of the setup you have already created a <b>Workspace</b>. To run AutoML, you also need to create an <b>Experiment</b>. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.
```
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'automl-bikeshareforecasting'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['SKU'] = ws.sku
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
## Compute
You will need to create a [compute target](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
#### Creation of AmlCompute takes approximately 5 minutes.
If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your cluster.
amlcompute_cluster_name = "bike-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
## Data
The [Machine Learning service workspace](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-workspace) is paired with the storage account, which contains the default data store. We will use it to upload the bike share data and create [tabular dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) for training. A tabular dataset defines a series of lazily-evaluated, immutable operations to load data from the data source into tabular representation.
```
datastore = ws.get_default_datastore()
datastore.upload_files(files = ['./bike-no.csv'], target_path = 'dataset/', overwrite = True,show_progress = True)
```
Let's set up what we know about the dataset.
**Target column** is what we want to forecast.
**Time column** is the time axis along which to predict.
```
target_column_name = 'cnt'
time_column_name = 'date'
dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'dataset/bike-no.csv')]).with_timestamp_columns(fine_grain_timestamp=time_column_name)
# Drop the columns 'casual' and 'registered' as these columns are a breakdown of the total and therefore a leak.
dataset = dataset.drop_columns(columns=['casual', 'registered'])
dataset.take(5).to_pandas_dataframe().reset_index(drop=True)
```
### Split the data
The first split we make is into train and test sets. Note we are splitting on time. Data before 9/1 will be used for training, and data after and including 9/1 will be used for testing.
```
# select data that occurs before a specified date
train = dataset.time_before(datetime(2012, 8, 31), include_boundary=True)
train.to_pandas_dataframe().tail(5).reset_index(drop=True)
test = dataset.time_after(datetime(2012, 9, 1), include_boundary=True)
test.to_pandas_dataframe().head(5).reset_index(drop=True)
```
## Forecasting Parameters
To define forecasting parameters for your experiment training, you can leverage the ForecastingParameters class. The table below details the forecasting parameter we will be passing into our experiment.
|Property|Description|
|-|-|
|**time_column_name**|The name of your time column.|
|**forecast_horizon**|The forecast horizon is how many periods forward you would like to forecast. This integer horizon is in units of the timeseries frequency (e.g. daily, weekly).|
|**country_or_region_for_holidays**|The country/region used to generate holiday features. These should be ISO 3166 two-letter country/region codes (i.e. 'US', 'GB').|
|**target_lags**|The target_lags specifies how far back we will construct the lags of the target variable.|
|**freq**|Forecast frequency. This optional parameter represents the period with which the forecast is desired, for example, daily, weekly, yearly, etc. Use this parameter for the correction of time series containing irregular data points or for padding of short time series. The frequency needs to be a pandas offset alias. Please refer to [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) for more information.
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|forecasting|
|**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>
|**blocked_models**|Models in blocked_models won't be used by AutoML. All supported models can be found at [here](https://docs.microsoft.com/en-us/python/api/azureml-train-automl-client/azureml.train.automl.constants.supportedmodels.forecasting?view=azure-ml-py).|
|**experiment_timeout_hours**|Experimentation timeout in hours.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
|**compute_target**|The remote compute for training.|
|**n_cross_validations**|Number of cross validation splits.|
|**enable_early_stopping**|If early stopping is on, training will stop when the primary metric is no longer improving.|
|**forecasting_parameters**|A class that holds all the forecasting related parameters.|
This notebook uses the blocked_models parameter to exclude some models that take a longer time to train on this dataset. You can choose to remove models from the blocked_models list but you may need to increase the experiment_timeout_hours parameter value to get results.
### Setting forecaster maximum horizon
The forecast horizon is the number of periods into the future that the model should predict. Here, we set the horizon to 14 periods (i.e. 14 days). Notice that this is much shorter than the number of days in the test set; we will need to use a rolling test to evaluate the performance on the whole test set. For more discussion of forecast horizons and guiding principles for setting them, please see the [energy demand notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand).
```
forecast_horizon = 14
```
### Config AutoML
```
from azureml.automl.core.forecasting_parameters import ForecastingParameters
forecasting_parameters = ForecastingParameters(
time_column_name=time_column_name,
forecast_horizon=forecast_horizon,
country_or_region_for_holidays='US', # set country_or_region will trigger holiday featurizer
target_lags='auto' # use heuristic based lag setting
)
automl_config = AutoMLConfig(task='forecasting',
primary_metric='normalized_root_mean_squared_error',
blocked_models = ['ExtremeRandomTrees'],
experiment_timeout_hours=0.3,
training_data=train,
label_column_name=target_column_name,
compute_target=compute_target,
enable_early_stopping=True,
n_cross_validations=3,
max_concurrent_iterations=4,
max_cores_per_iteration=-1,
verbosity=logging.INFO,
forecasting_parameters=forecasting_parameters)
```
We will now run the experiment, you can go to Azure ML portal to view the run details.
```
remote_run = experiment.submit(automl_config, show_output=False)
remote_run
remote_run.wait_for_completion()
```
### Retrieve the Best Model
Below we select the best model from all the training iterations using get_output method.
```
best_run, fitted_model = remote_run.get_output()
fitted_model.steps
```
## Featurization
You can access the engineered feature names generated in time-series featurization. Note that a number of named holiday periods are represented. We recommend that you have at least one year of data when using this feature to ensure that all yearly holidays are captured in the training featurization.
```
fitted_model.named_steps['timeseriestransformer'].get_engineered_feature_names()
```
### View the featurization summary
You can also see what featurization steps were performed on different raw features in the user data. For each raw feature in the user data, the following information is displayed:
- Raw feature name
- Number of engineered features formed out of this raw feature
- Type detected
- If feature was dropped
- List of feature transformations for the raw feature
```
# Get the featurization summary as a list of JSON
featurization_summary = fitted_model.named_steps['timeseriestransformer'].get_featurization_summary()
# View the featurization summary as a pandas dataframe
pd.DataFrame.from_records(featurization_summary)
```
## Evaluate
We now use the best fitted model from the AutoML Run to make forecasts for the test set. We will do batch scoring on the test dataset which should have the same schema as training dataset.
The scoring will run on a remote compute. In this example, it will reuse the training compute.
```
test_experiment = Experiment(ws, experiment_name + "_test")
```
### Retrieving forecasts from the model
To run the forecast on the remote compute we will use a helper script: forecasting_script. This script contains the utility methods which will be used by the remote estimator. We copy the script to the project folder to upload it to remote compute.
```
import os
import shutil
script_folder = os.path.join(os.getcwd(), 'forecast')
os.makedirs(script_folder, exist_ok=True)
shutil.copy('forecasting_script.py', script_folder)
```
For brevity, we have created a function called run_forecast that submits the test data to the best model determined during the training run and retrieves forecasts. The test set is longer than the forecast horizon specified at train time, so the forecasting script uses a so-called rolling evaluation to generate predictions over the whole test set. A rolling evaluation iterates the forecaster over the test set, using the actuals in the test set to make lag features as needed.
```
from run_forecast import run_rolling_forecast
remote_run = run_rolling_forecast(test_experiment, compute_target, best_run, test, target_column_name)
remote_run
remote_run.wait_for_completion(show_output=False)
```
### Download the prediction result for metrics calcuation
The test data with predictions are saved in artifact outputs/predictions.csv. You can download it and calculation some error metrics for the forecasts and vizualize the predictions vs. the actuals.
```
remote_run.download_file('outputs/predictions.csv', 'predictions.csv')
df_all = pd.read_csv('predictions.csv')
from azureml.automl.core.shared import constants
from azureml.automl.runtime.shared.score import scoring
from sklearn.metrics import mean_absolute_error, mean_squared_error
from matplotlib import pyplot as plt
# use automl metrics module
scores = scoring.score_regression(
y_test=df_all[target_column_name],
y_pred=df_all['predicted'],
metrics=list(constants.Metric.SCALAR_REGRESSION_SET))
print("[Test data scores]\n")
for key, value in scores.items():
print('{}: {:.3f}'.format(key, value))
# Plot outputs
%matplotlib inline
test_pred = plt.scatter(df_all[target_column_name], df_all['predicted'], color='b')
test_test = plt.scatter(df_all[target_column_name], df_all[target_column_name], color='g')
plt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)
plt.show()
```
For more details on what metrics are included and how they are calculated, please refer to [supported metrics](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-understand-automated-ml#regressionforecasting-metrics). You could also calculate residuals, like described [here](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-understand-automated-ml#residuals).
Since we did a rolling evaluation on the test set, we can analyze the predictions by their forecast horizon relative to the rolling origin. The model was initially trained at a forecast horizon of 14, so each prediction from the model is associated with a horizon value from 1 to 14. The horizon values are in a column named, "horizon_origin," in the prediction set. For example, we can calculate some of the error metrics grouped by the horizon:
```
from metrics_helper import MAPE, APE
df_all.groupby('horizon_origin').apply(
lambda df: pd.Series({'MAPE': MAPE(df[target_column_name], df['predicted']),
'RMSE': np.sqrt(mean_squared_error(df[target_column_name], df['predicted'])),
'MAE': mean_absolute_error(df[target_column_name], df['predicted'])}))
```
To drill down more, we can look at the distributions of APE (absolute percentage error) by horizon. From the chart, it is clear that the overall MAPE is being skewed by one particular point where the actual value is of small absolute value.
```
df_all_APE = df_all.assign(APE=APE(df_all[target_column_name], df_all['predicted']))
APEs = [df_all_APE[df_all['horizon_origin'] == h].APE.values for h in range(1, forecast_horizon + 1)]
%matplotlib inline
plt.boxplot(APEs)
plt.yscale('log')
plt.xlabel('horizon')
plt.ylabel('APE (%)')
plt.title('Absolute Percentage Errors by Forecast Horizon')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/pachterlab/GRNP_2020/blob/master/notebooks/figure_generation/GenFig4AC_S23.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**Generates figure 4 A-C and supplementary figure 23**
This notebook generates figures for showing the improvement of BUTTERFLY correction on scRNA-Seq data, by comparing a downsampled and full dataset. Furthermore, we show the effect of "borrowing" CU histogram information from similar datasets, and determine the sampling noise, which sets a theoretical maximum performance for the prediction in downsampling scenarios such as this.
Steps:
1. Download the code and processed data
2. Setup the R environment
3. Generate the figures
The data for these figures is produced by the following notebooks:
Processing of FASTQ files with kallisto and bustools:
https://github.com/pachterlab/GRNP_2020/blob/master/notebooks/FASTQ_processing/ProcessPBMC_V3_3.ipynb
Preprocessing of BUG files:
https://github.com/pachterlab/GRNP_2020/blob/master/notebooks/R_processing/ProcessR_PBMC_V3_3.ipynb
Precalculation of figure data:
https://github.com/pachterlab/GRNP_2020/blob/master/notebooks/figure_generation/GenFig4AC_S23Data.ipynb
**1. Download the code and processed data**
```
#download the R code
![ -d "GRNP_2020" ] && rm -r GRNP_2020
!git clone https://github.com/pachterlab/GRNP_2020.git
#download processed data from Zenodo for all datasets
![ -d "figureData" ] && rm -r figureData
!mkdir figureData
!cd figureData && wget https://zenodo.org/record/4661263/files/FigureData.zip?download=1 && unzip 'FigureData.zip?download=1' && rm 'FigureData.zip?download=1'
#Check that download worked
!cd figureData && ls -l && cd PBMC_V3_3 && ls -l
```
**2. Prepare the R environment**
```
#switch to R mode
%reload_ext rpy2.ipython
#install the R packages and setup paths
%%R
install.packages("dplyr")
install.packages("ggplot2")
install.packages("DescTools")
install.packages("ggpubr")
install.packages("hexbin")
install.packages("reshape2")
install.packages("farver")
```
**3. Generate the figures**
```
#First set some path variables
%%R
source("GRNP_2020/RCode/pathsGoogleColab.R")
#Import the code for prediction (available in other notebooks)
%%R
source(paste0(sourcePath,"ButterflyHelpers.R"))
#source(paste0(sourcePath,"preseqHelpers.R"))
source(paste0(sourcePath,"CCCHelpers.R"))
source(paste0(sourcePath,"ggplotHelpers.R"))
#create figure directory
![ -d "figures" ] && rm -r figures
!mkdir figures
#Create and save the figures
%%R
library(ggplot2)
library(ggpubr)
library(hexbin)
library(dplyr)
ldata = readRDS(paste0(figure_data_path, "Fig4AC_ldata.RDS"))
ldata2 = readRDS(paste0(figure_data_path, "Fig4AC_ldata2.RDS"))
#generate plot data
plotdata = tibble(gene=ldata$gene,
x=ldata$x,
nopred=ldata$nopred - ldata$trueval,
pred=ldata$pred - ldata$trueval,
poolpred=ldata$poolpred - ldata$trueval)
#melt
plotdata.m = reshape2::melt(plotdata, id.vars=c("gene","x"), measure.vars = c("nopred", "pred", "poolpred"))
labl = labeller(variable =
c("nopred" = "No Correction",
"pred" = "Correction",
"poolpred" = "Correction using Pooling"))
dfline = data.frame(x=c(0,16), y=c(0,0))
dummyData = data.frame(x=c(0,0), y=c(1.1, -1.5)) #used in a trick to set y axis range below
fig4AC = ggplot(plotdata.m) +
stat_binhex(bins=60,na.rm = TRUE, mapping=aes(x = x, y=value, fill = log(..count..))) + # opts(aspect.ratio = 1) +
facet_wrap(facets = ~ variable, scales = "free_x", labeller = labl, ncol=3) +
geom_line(data=dfline, mapping = aes(x=x, y=y), color="black", size=1) +
geom_blank(data = dummyData, mapping = aes(x=x, y=y)) + #trick to set y axis range
labs(y=expression(Log[2]*" fold change (CPM)"), x=expression(Log[2]*"(CPM + 1)")) +
theme(panel.background = element_rect("white", "white", 0, 0, "white"),
legend.position= "bottom", legend.direction = "horizontal",#, legend.title = element_blank())
strip.text.x = element_text(size = 12, face = "bold"),
#legend.position= "none",
strip.background = element_blank())
print(fig4AC)
ggsave(
paste0(figure_path, "Fig4AC.png"),
plot = fig4AC, device = "png",
width = 7, height = 4, dpi = 300)
%%R
#########################
# Fig S23 (Sampling noise)
#########################
#cpm and log transform
plotdata2 = tibble(gene=ldata2$gene,
x=ldata2$x,
y=ldata2$sampling - ldata$nopred)
dfline = data.frame(x=c(0,16), y=c(0,0))
dummyData = data.frame(x=c(0,0), y=c(1.1, -1.5))
figS23 = ggplot(plotdata2) +
stat_binhex(bins=60,na.rm = TRUE, mapping=aes(x = x, y=y, fill = log(..count..))) + # opts(aspect.ratio = 1) +
#facet_wrap(facets = ~ variable, scales = "free_x", labeller = labl, ncol=3) +
geom_line(data=dfline, mapping = aes(x=x, y=y), color="black", size=1) +
geom_blank(data = dummyData, mapping = aes(x=x, y=y)) + #trick to set y axis range
labs(y=expression(Log[2]*" fold change (CPM)"), x=expression(Log[2]*"(CPM + 1)")) +
theme(panel.background = element_rect("white", "white", 0, 0, "white"),
legend.position= "bottom", legend.direction = "horizontal",#, legend.title = element_blank())
strip.text.x = element_text(size = 12, face = "bold"),
#legend.position= "none",
strip.background = element_blank())
print(figS23)
ggsave(
paste0(figure_path, "FigS23.png"),
plot = figS23, device = "png",
width = 3, height = 4, dpi = 300)
#The data to present over the plots
%%R
print(paste0("CCC, no pred: ", getCCC(ldata$nopred, ldata$trueval))) #0.981275291888894
print(paste0("CCC, pred no pooling: ", getCCC(ldata$pred, ldata$trueval))) #0.993829998877551
print(paste0("CCC, pred with pooling: ", getCCC(ldata$poolpred, ldata$trueval))) #0.997030743452436
print(paste0("CCC, no pred, bin ds vs ds: ", getCCC(ldata2$nopred, ldata2$sampling))) #0.99895348369159
#also get mean squared error
print(paste0("MSE, no pred: ", getMSE(ldata$nopred, ldata$trueval))) #0.192957136231134
print(paste0("MSE, pred no pooling: ", getMSE(ldata$pred, ldata$trueval))) #0.0624471167682468
print(paste0("MSE, pred with pooling: ", getMSE(ldata$poolpred, ldata$trueval))) #0.0299541970779269
print(paste0("MSE, no pred, ds 100 times vs ds: ", getMSE(ldata2$nopred, ldata2$sampling))) #0.0108884743502623
```
| github_jupyter |
```
import math
import torch
from torch import nn
from torchvision import models
import matplotlib.pyplot as plt
import torch.nn.functional as F
from torchvision import datasets, transforms
from torch.utils.data import DataLoader, random_split
```
# Transforms, DataLoader, DataSet
```
transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
])
target_transform = transforms.Compose([
transforms.Lambda(lambda y: torch.zeros(4, dtype=torch.float) \
.scatter_(0, torch.tensor(y), value=1))
])
data_dir = '../data/images'
raw_data = datasets.ImageFolder(data_dir,
transform=transform,
target_transform=target_transform)
classes = raw_data.classes
sz = len(raw_data)
train_sz = math.floor(.8 * sz)
val_sz = sz - train_sz
train_dataset, val_dataset = random_split(raw_data, [train_sz, val_sz])
```
# Model Arch
```
# Get cpu or gpu device for training.
device = "cuda" if torch.cuda.is_available() else "cpu"
print("Using {} device".format(device))
classes = 4
# Define model
class VisionModel(nn.Module):
def __init__(self):
super(VisionModel, self).__init__()
self.xfer = models.mobilenet_v3_small(pretrained=True)
self.fc1 = nn.Linear(1000, classes)
def forward(self, x):
x = F.relu(self.xfer(x))
return F.softmax(self.fc1(x), dim=1)
model = VisionModel().to(device)
print(model)
```
# Training
```
def train(dataloader, model, loss_fn, optimizer):
size = len(dataloader.dataset)
model.train()
current = 0
for batch, (X, y) in enumerate(dataloader):
X, y = X.to(device), y.to(device)
# Compute prediction error
pred = model(X)
loss = loss_fn(pred, y)
# Backpropagation
optimizer.zero_grad()
loss.backward()
optimizer.step()
current += len(X)
print(f"loss: {loss.item():>7f} [{current:>5d}/{size:>5d}], {batch}")
def test(dataloader, model, loss_fn):
size = len(dataloader.dataset)
num_batches = len(dataloader)
model.eval()
test_loss, correct = 0, 0
with torch.no_grad():
for X, y in dataloader:
X, y = X.to(device), y.to(device)
pred = model(X)
test_loss += loss_fn(pred, y).item()
correct += (pred.argmax(1) == y.argmax(1)).type(torch.float).sum().item()
test_loss /= num_batches
correct /= size
print(f"Test Error: \n Accuracy: {(100*correct):>0.1f}%, Avg loss: {test_loss:>8f} \n")
epochs = 5
batch_size = 32
loss_fn = nn.BCELoss()
optimizer = torch.optim.SGD(model.parameters(), lr=.01)
train_dataloader = DataLoader(train_dataset, batch_size=batch_size)
test_dataloader = DataLoader(val_dataset, batch_size=batch_size)
for t in range(epochs):
print(f"Epoch {t+1}\n-------------------------------")
train(train_dataloader, model, loss_fn, optimizer)
test(test_dataloader, model, loss_fn)
print("Done!")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/gmagannaDevelop/Taller-DCI-NET/blob/master/Ejercicios_funcionales.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Programación funcional en Python
Veamos a nuestros nuevos amigos. He aquí una lista con una descripción increíblemente útil. Posteriormente los veremos en acción.
1. ```lambda``` : Declarar una función anónima.
2. ```map``` : Mapear, se especifica primero la función y después el objeto.
3. ```filter``` : Filtrar para mantener elementos que cumplan con un criterio.
4. ```reduce``` : Aplicar una función cumulativa.
```
from functools import reduce
```
La función reduce no es parte del espacio de nombres por defecto, por decisión del creador de Python : [Guido](https://github.com/gvanrossum). Por eso debemos importarla del módulo de herramientas para programación funcional functools.
```
import numpy as np
import seaborn as sns
import pandas as pd
```
## Funciones anónimas
**Utilidad** : Digamos que queremos calcular algo rápidamente, pero no queremos guardar una función que lo haga. Tal vez es una operación que se hará sólo una vez y no queremos "ocupar ese nombre", ahí usamos una función anónima o expresión lambda.
1. **Sintaxis** :
$$ f(x) \; = \; x $$
```
lambda x: x
```
Ahora con varios argumentos :
$$ f(x,y,x) \; = \; x\cdot y\cdot z $$
```
lambda x, y, z: x*y*z
```
2. **Evaluación**
$$ f(x) = x^{x}\vert_{3} = 27 $$
```
(lambda x: x**x)(3)
```
Está muy bien eso de que sean anónimas pero, ¿y si yo quisiera guardar mi función?
3. **Asignación**
```
cuadrado = lambda x: x**2
cuadrado(3)
```
4. Funciones de orden superior
```
aplica_función = lambda x, y: x(y)
aplica_función(cuadrado, 3)
```
5. **Condicionales**
Digamos que quisiésemos saber si un valor es positivo.
```
es_positivo = lambda x: True if x > 0 else False
es_positivo(3)
es_positivo(-np.pi)
```
## Mapear
Hay diversas formas de llevar a cabo la misma operación. A continuación las abordaremos, pasando por clásicos hasta la forma funcional.
**Nuestra tarea :** Elevar una lista de números al cuadrado.
```python
x = [1, 2, 3, 4, 5, 6, 7, 8]
```
1. La forma tradicional, no pitónica :
```
x = [1, 2, 3, 4, 5, 6, 7, 8]
y = x.copy()
for i in range(len(x)):
y[i] = x[i] ** 2
print(x)
print(y)
```
2. Una forma más pitónica :
```
x = [1, 2, 3, 4, 5, 6, 7, 8]
y = [ valor**2 for valor in x ]
print(x)
print(y)
```
3. La forma funcional :
```
x = [1, 2, 3, 4, 5, 6, 7, 8]
y = list(map(lambda x: x**2, x))
print(x)
print(y)
```
¿Por qué tuvimos que hacer ``` list(map(...)) ``` en vez de sólo ``` map(...) ```? Porque map crea un iterador que va generando los elementos mientras se los vamos pidiendo. En este caso la función (constructor) ```list()``` crea una lista a partir del iterador.
```
x = [1, 2, 3, 4, 5, 6, 7, 8]
y = [*map(lambda x: x**2, x)]
print(x)
print(y)
```
También podemos utilizar funciones que toman más de un argumento, aplicándolas elemento a elemento.
```
list(map(lambda x, y: x - y, [1, 2, 3], [4, 5, 6j]))
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
#sample 1
elements = ['Sodium', 'Potassium', 'Cobalt', 'Bromine', 'Rubidium', 'Antimony',
'Cesium', 'Mercury']
ppm = [980.2, 2588.97, 0.043, 1.4, 2.22, 2.57, 0.21, 0.035]
sys_unc = [428.53, 1325.27, 0.022, 0.45, 1.056, 0.62, 0.061, 0.0043]
stat_unc = [5.66, 98.93, 0.0056, 0.07, 0.058, 0.17, 0.016, 0.0026]
x = np.linspace(1,len(ppm),len(ppm))
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(10,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [1120, 4912, 0.045, 13, 14.7, 1.5, 0.01, 0.314]
#regulatory limits for the elements
limits = [0, 0, 8.48, 0, 23, 0.1, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 1: Swai Basa Fillet from Vietnam')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 2
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Arsenic', 'Bromine',
'Rubidium', 'Antimony','Barium', 'Cesium', 'Gold', 'Mercury']
ppm = [64.70, 3780.58, 0.052, 1.63, 1.45, 4.15, 0.14, 0.31, 10.48, 0.212, 0.00075, 0.234]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [28.48, 1935.24, 0.027, 0.18, 0.43, 1.32, 0.068, 0.08, 1.26, 0.057, 0.00014, 0.029]
stat_unc = [2.06, 790.84, 0.0059, 0.19, 0.14, 0.12, 0.014, 0.07, 1.81, 0.038, 0.00009, 0.031]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [1120, 4912, 0.045, 0.142, 0.03, 13, 14.7, 1.5, 14.3, 0.01, 0.0005, 0.5]
#regulatory limits for the elements
limits = [0, 0, 8.48, 11.3, 3.5, 0, 20, 0.1, 14.3, 0, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 2: Hamachi Fillet from JAPAN')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 3
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Arsenic', 'Bromine',
'Rubidium', 'Antimony', 'Cesium', 'Mercury']
ppm = [69.16, 2650.95, 0.274, 0.923, 0.62, 2.16, 0.15, 0.0277, 0.032, 0.024]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [30.68, 1357, 0.14, 0.087, 0.19, 0.7, 0.07, 0.01, 0.0094, 0.003]
stat_unc = [2.54, 95.49, 0.031, 0.072, 0.092, 0.07, 0.014, 0.01, 0.0048, 0.0022]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 12345, 0.3, 0.58, 3.5, 13, 15.65, 1.5, 0.01, 0.773]
#regulatory limits for the elements
limits = [0, 0, 8.48, 8, 3.5, 0, 20, 0.1, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Isotope ppm of Sample 3: Cardinal snapper from MEXICO')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 4
elements = ['Sodium', 'Potassium', 'Cobalt','Selenium', 'Arsenic', 'Bromine',
'Rubidium', 'Barium', 'Cesium', 'Mercury']
ppm = [164.3, 3322.37, 1.52, 2.10, 0.7, 5.5, 0.144, 25.22, 0.25, 0.22]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [70.72, 1700.69, 0.77, 0.2, 0.21, 1.8, 0.069, 2.03, 0.1, 0.027]
stat_unc = [7.94, 88.56, 0.11, 0.25, 0.081, 0.13, 0.014, 5.3, 0.091, 0.0065]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 1000, 0.3, 0.58, 3.5, 13, 20, 14.3, 0.01, 0.514]
#regulatory limits for the elements
limits = [0, 0, 8.48, 11.3, 3.5, 0, 20, 14.3, 0.0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 4: Shortraker steak from CANADA')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 5
elements = ['Sodium', 'Potassium', 'Iron', 'Cobalt','Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Cesium']
ppm = [143.56, 3556.56, 3.58, 0.885, 1.77, 0.83, 4.61, 0.25, 0.06]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [62.91, 1820.56, 1.83, 0.45, 0.17, 0.23, 1.45, 0.12, 0.019]
stat_unc = [1.24, 144.19, 0.55, 0.087, 0.13, 0.08, 0.24, 0.020, 0.006]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 12345, 13.5, 0.3, 0.58, 3.5, 13, 15.65, 0.01]
#regulatory limits for the elements
limits = [0, 0, 1, 8.48, 8, 3.5, 0, 23, 0]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 5: Yellow tail snapper from Brazil')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 6
elements = ['Sodium', 'Potassium', 'Iron', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Cesium']
ppm = [510.92, 5369.57, 6.48, 0.05, 0.79, 0.64, 2.19, 1.82, 0.25]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [204.54, 2748.11, 3.12, 0.026, 0.07, 0.19, 0.24, 0.87, 0.073]
stat_unc = [154.81, 174.78, 0.73, 0.0058, 0.088, 0.09, 0.21, 0.092, 0.038]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [860, 4912, 13.5, 0.3, 0.58, 3.5, 13, 20, 0.01]
#regulatory limits for the elements
limits = [0, 0, 1, 8.48, 8, 3.5, 0, 23, 0]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 6: Tilapia from TAIWAN')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 7
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Bromine',
'Rubidium', 'Gold', 'Mercury']
ppm = [153.35, 2436.11, 0.1, 0.78, 1.46, 0.78, 0.00086, 0.09]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [66.67, 1247, 0.05, 0.077, 0.48, 0.37, 0.00016, 0.011]
stat_unc = [1.03, 96.54, 0.0087, 0.06, 0.06, 0.037, 0.000067, 0.0043]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(10,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 1000, 0.03, 0.237, 13, 20, 0.0005, 0.217]
#regulatory limits for the elements
limits = [0, 0, 8.48, 11.3, 0, 23, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 7: Grass carp steak from TAIWAN')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 8
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Cesium', 'Mercury']
ppm = [9384.64, 2360.73, 0.22, 0.39, 2.73, 3.33, 0.114, 0.033, 0.064]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [3168.57, 1208.43, 0.11, 0.04, 0.74, 1.034, 0.054, 0.011, 0.008]
stat_unc = [3237, 163.37, 0.032, 0.024, 0.56, 0.02, 0.011, 0.0045, 0.0048]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 1000, 0.03, 0.237, 0.01, 13, 20, 0.01, 0.271]
#regulatory limits for the elements
limits = [0, 0, 8.48, 11.3, 3.5, 13, 23, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 8: Grass carp steak from TAIWAN')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 9
elements = ['Sodium', 'Potassium', 'Scandium', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Cesium', 'Gold', 'Mercury']
ppm = [207.54, 2006.06, 0.00055, 0.03, 1.93, 7.04, 7.77, 0.125, 0.085, 0.0025, 0.09]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [91.25, 1026.88, 0.00023, 0.013, 0.18, 2.07, 2.39, 0.059, 0.027, 0.00046, 0.011]
stat_unc = [2.25, 194.5, 0.000073, 0.0029, 0.14, 0.36, 0.47, 0.011, 0.0064, 0.00029, 0.003]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 1000, 0, 0.03, 0.58, 3.5, 13, 20, 0.01, 0.0005, 2.18]
#regulatory limits for the elements
limits = [0, 0, 0, 8.48, 8, 3.5, 0, 23, 0, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 9: Chilean sea bass from KOREA')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 10
elements = ['Sodium', 'Potassium', 'Scandium', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Antimony', 'Barium', 'Cesium', 'Mercury']
ppm = [50.48, 2164.19, 0.00027, 0.17, 0.34, 0.56, 2.92, 0.11, 0.064, 51.01, 0.091, 0.0085]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [22.26, 1107.83, 0.00011, 0.085, 0.033, 0.17, 0.96, 0.11, 0.017, 7.22, 0.027, 0.001]
stat_unc = [1.0, 181.28, 0.000061, 0.018, 0.021, 0.095, 0.09, 0.019, 0.0084, 7.23, 0.0093, 0.0011]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [1120, 1000, 0, 0.3, 0.58, 3.5, 13, 20, 1.5, 14.3, 0.01, 0.19]
#regulatory limits for the elements
limits = [0, 0, 0, 8.48, 11.3, 3.5, 0, 23, 0.1, 14.3, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 10: Wild isle salmon from SCOTLAND')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 11
elements = ['Sodium', 'Potassium', 'Iron', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Mercury']
ppm = [110.81, 1736.26, 7.88, 0.19, 1.78, 1.16, 13.22, 0.203, 0.066]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [48.87, 888.78, 3.27, 0.098, 0.16, 0.34, 6.54, 0.11, 0.0097]
stat_unc = [1.07, 112.71, 0.92, 0.012, 0.08, 0.1, 2.22, 0.025, 0.0038]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [860, 1000, 13.5, 0.3, 0.58, 3.5, 13, 15.65, 0.17]
#regulatory limits for the elements
limits = [0, 0, 1, 8.48, 8, 3.5, 0, 23, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 11: Mackerel from INDIA')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 12
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Arsenic', 'Bromine',
'Rubidium', 'Antimony', 'Cesium', 'Mercury']
ppm = [85.33, 1652.5, 0.127, 0.56, 2.14, 3.38, 0.084, 0.5736, 0.02, 0.048]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [37.58, 845.9, 0.064, 0.053, 0.64, 1.06, 0.04, 0.25, 0.0057, 0.006]
stat_unc = [1.83, 185.65, 0.017, 0.029, 0.22, 0.13, 0.013, 0.25, 0.0029, 0.0029]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [1120, 4912, 0.3, 0.58, 3.5, 13, 14.7, 1.5, 0.01, 0.314]
#regulatory limits for the elements
limits = [0, 0, 8.48, 8, 3.5, 0, 23, 0.1, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 12: Norwegian basa from NORWAY')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 13
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Arsenic', 'Bromine',
'Rubidium', 'Cesium', 'Mercury']
ppm = [95.74, 2103.34, 0.244, 0.42, 1.65, 3.68, 0.25, 0.11, 0.02]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [40.13, 1076.68, 0.12, 0.041, 0.49, 1.16, 0.12, 0.045, 0.0023]
stat_unc = [2.6, 322.2, 0.043, 0.045, 0.21, 0.14, 0.034, 0.024, 0.002]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [860, 11693, 0.3, 0.58, 0.1, 13, 125.2, 0.01, 0.21]
#regulatory limits for the elements
limits = [0, 0, 8.48, 8, 3.5, 13, 23, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 13: Golden pompano from CHINA')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 14
elements = ['Sodium', 'Potassium', 'Scandium', 'Cobalt', 'Selenium', 'Arsenic',
'Bromine', 'Rubidium', 'Cesium', 'Mercury']
ppm = [91.89, 1930.47, 0.001, 0.08, 0.96, 0.64, 23.19, 0.22, 0.22, 0.15]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [40.3, 834.62, 0.0007, 0.04, 0.092, 0.19, 6.97, 0.1, 0.067, 0.018]
stat_unc = [1.93, 123.31, 0.00034, 0.012, 0.057, 0.099, 3.75, 0.027, 0.025, 0.0079]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [1120, 1000, 0, 0.3, 0.58, 3.5, 13, 30, 0.01, 2.18]
#regulatory limits for the elements
limits = [0, 0, 0, 8.48, 11.3, 3.5, 0, 23, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 14: Chilean sea bass from the ARTIC')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 15
elements = ['Sodium', 'Potassium', 'Scandium', 'Iron', 'Cobalt', 'Selenium',
'Arsenic', 'Bromine', 'Strontium', 'Rubidium', 'Mercury']
ppm = [303.97, 1856.51, 0.0084, 15.57, 1.45, 0.72, 1.19, 61.80, 34.34, 0.31, 0.047]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [133.68, 950.33, 0.00037, 7.78, 0.74, 0.066, 0.37, 19.83, 12.23, 0.15, 0.0058]
stat_unc = [3.37, 136.6, 0.00047, 0.83, 0.038, 0.045, 0.14, 3.45, 1.79, 0.019, 0.0022]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [3000, 1000, 0, 13.5, 0.3, 0.58, 3.5, 13, 8, 20, 0.033]
#regulatory limits for the elements
limits = [0, 0, 0, 1, 8.48, 8, 3.5, 0, 4, 23, 13]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 15: HD shrimp from VENEZUELA')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#sample 16
elements = ['Sodium', 'Potassium', 'Cobalt', 'Selenium', 'Bromine',
'Cesium', 'Mercury']
ppm = [176.69, 1473.66, 0.166, 1.51, 5.83, 0.09, 0.75]
x = np.linspace(1,len(ppm),len(ppm))
sys_unc = [80.61, 754.35, 0.084, 0.15, 1.92, 0.024, 0.093]
stat_unc = [4.79, 98.09, 0.022, 0.10, 0.21, 0.013, 0.023]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.25
p1 = ax.bar(x, ppm, width, color='powderblue', label='Element')
#reference values (based on averages)
ppm2 = [860, 1000, 0.3, 0.58, 13, 0.01, 1.205]
#regulatory limits for the elements
limits = [0, 0, 8.48, 11.3, 0, 0, 1]
#reference graph
p2 = ax.bar(x+0.25, ppm2, width, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
#error bars
p3 = plt.errorbar(x, ppm, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
#limits graph
p4 = ax.bar(x-0.25, limits, width, color='lightslategray', edgecolor='k', label='Limit')
plt.xlabel('Element')
plt.ylabel('Concentration in ppm')
plt.title('Element ppm of Sample 16: Pink grouper steak from MEXICO')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#Pottery comparison: comparing the calculated grams of isotopes and
#comparing it to the pottery's original grams
#sample 16
elements = ['Sb-121', 'As-75','Ba-130','Br-81','Ce-140','Cs-133','Co-59',
'Fe-58','Cu-63','Ga-71','La-139','K-41','Rb-85','Sc-45',
'Na-23','Sr-84','Hf-180']
grams = [9.97e-7,4.74e-6,3.08e-5,1.33e-6,4.97e-6,9.24e-5,3.00e-4,
2.04e-6, 5.82e-5, 6.06e-6, 5.97e-6, 6.85e-5, 1.52e-6,
5.36e-6, 8.16e-5, 9.55e-7, 2.4e-7]
x = np.linspace(1,len(grams),len(grams))
sys_unc = [2.58e-7,1.2e-6,2.45e-6,1.13e-6,4.1e-7,2.34e-5,1.49e-4,1.5e-8,
9.8e-7, 3.01e-5,2.38e-6,8.7e-7,3.49e-5,1.21e-6,2.22e-6,4.23e-5,
2.25e-7,5.3e-8]
stat_unc = [1.1e-8,4.37e-8,6.07e-8,4.05e-8,5.48e-8,1.06e-6,8.89e-7,8.78e-10,
3.57e-8,9.04e-6, 1.51e-7,3.65e-8,1.66e-6,4.98e-8,1.93e-8,4.27e-7,
2.21e-8,6.13e-9]
tot_unc = []
for i in x:
val = (sys_unc[int(i)-1]**2+stat_unc[int(i)-1]**2)**0.5
tot_unc.append(val)
mpl_fig = plt.figure(figsize=(12,5))
ax = mpl_fig.add_subplot(111)
width = 0.5
p1 = ax.bar(x, grams, width, color='powderblue', label='Element')
#reference values (based on averages)
pot_grams = [1.66e-6,3.08e-5,7.12e-4,2.3e-6,8.03e-5,8.31e-5,1.406e-5,
1.017e-2,6.0e-5,4.44e-5,4.49e-5,1.45e-2,7.0e-5,2.055e-5,
2.61e-3,1.45e-4,6.23e-6]
#reference graph
p2 = ax.bar(x, pot_grams, width=0.35, color='lightsalmon', edgecolor='red', alpha=0.3, label='Reference')
p3 = plt.errorbar(x, grams, yerr=tot_unc, fmt='.', ecolor='steelblue', capthick=1, capsize=2)
plt.xlabel('Isotope')
plt.ylabel('Grams')
plt.title('Detected grams in Pottery sample')
plt.legend()
plt.xticks(x, elements)
ax.set_yscale('log')
plt.show()
#The reason why some isotopes seem to have a higher amount than the
#starting existing amount is because the pottery was previously irradiated
#therefore the model is predicting more of the pre-existing isotope due to
#the radioactive isotopes created in the first irradiation.
#co-59 can be from the brick contamination
```
| github_jupyter |
## מגישות
טוהר רחמין וליאור דדון
#### קישור
https://www.census.gov/foreign-trade/statistics/historical/index.html
```
from IPython.display import Image
Image("DHL.jpg")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
data = pd.read_csv('exhibit_history_project (2).csv')
data= data.iloc[:,0:10]
data['month']= data.iloc[:,0]
data['years']= data.iloc[:,0]
data['years']=pd.to_numeric(data['years'], errors='coerce')
data['years']=data['years'].fillna(method='ffill')
data_n=data[:].dropna()
data_n['years']=data_n['years'].astype(int)
data_n
data_n['Balance_Services']=data_n['Balance_Services'].astype(int)
data_n['Balance_Goods']=data_n['Balance_Goods'].astype(int)
data_n['Balance_Total']=data_n['Balance_Total'].astype(int)
data_n['Exports_Total']=data_n['Exports_Total'].astype(int)
data_n['Exports_Goods']=data_n['Exports_Goods'].astype(int)
data_n['Exports_Services']=data_n['Exports_Services'].astype(int)
data_n['Imports_Total']=data_n['Imports_Total'].astype(int)
data_n['Imports_Goods']=data_n['Imports_Goods'].astype(int)
data_n['Imports_Services']=data_n['Imports_Services'].astype(int)
Data= data_n.loc[data_n.month!='Jan. - Dec.']
```
# יצוא ויבוא לפי עונות
```
data_n['month']=data_n['month'].convert_dtypes()
Spring=data_n.loc[(data_n.month=='April')|(data_n.month== 'March') |(data_n.month== 'May')].copy()
Spring['s']='Spring'
Summer=data_n.loc[(data_n.month=='June')|(data_n.month== 'July') |(data_n.month== 'August')].copy()
Summer['s']='summer'
Autumn=data_n.loc[(data_n.month=='Septembe')|(data_n.month== 'October') |(data_n.month== 'November')].copy()
Autumn['s']='Autumn'
Winter=data_n.loc[(data_n.month=='December')|(data_n.month== 'January') |(data_n.month== 'February')].copy()
Winter['s']='winter'
data_n['winter']=Winter[['s']]
data_n['spring']=Spring[['s']]
data_n['summer']=Summer[['s']]
data_n['autumn']=Autumn[['s']]
data_n[['winter','autumn','summer','spring']]
data_n["seasons"]=data_n[['winter']]
data_n["seasons"].fillna(data_n["autumn"], inplace=True)
data_n["seasons"].fillna(data_n["summer"], inplace=True)
data_n["seasons"].fillna(data_n["spring"], inplace=True)
sns.catplot(x="seasons", y='Exports_Total', data=data_n, jitter=0.1)
plt.title('Exports_Total',fontsize=20)
plt.ylim([190000,220000])
plt.ylabel("")
sns.catplot(x="seasons", y='Imports_Total', data=data_n, jitter=0.1)
plt.title('Imports_Total',fontsize=20)
plt.ylim([100000,220000])
plt.ylabel("")
```
### מסקנה
כמו שניתן לראות העונה בא יש את היצוא הכי גדול היא אביב ואילו הכי נמוכה היא החורף
בנוסף העונות אביב וקיץ היו בעלות היבוא הגדול ביותר אך העונות חורף וסתיו היו הקטנות יות
# רגעי השיא והשפל
```
years2020=Data.loc[(data_n.years==2020.0)]
years2001=Data.loc[(data_n.years==2001.0)]
```
# קורונה
```
fig, axes = plt.subplots(figsize=(30,10), ncols=2)
sns.barplot(x= 'month', y='Exports_Total', data=years2020, ax = axes[0],palette=("YlOrRd"))
plt.xticks(rotation='vertical')
plt.title('Exports Total 2020',fontsize=35)
sns.barplot(x= 'month', y='Imports_Total', data=years2020, ax = axes[1],palette=("YlOrRd"))
plt.xticks(rotation='vertical')
plt.title('Imports Total 2020',fontsize=45)
```
## הערה
ניתן לראות שבחודשים מרץ-אפריל-מאי חלה ירידה גם ביבוא וגם ביצוא
# בספטמבר 11
```
fig, axes = plt.subplots(figsize=(30,10), ncols=2)
sns.barplot(x= 'month', y='Exports_Total', data=years2001, ax = axes[0],palette=("bone"))
plt.xticks(rotation='vertical')
plt.title('Exports Total 2001',fontsize=30)
sns.set_context("poster")
sns.barplot(x= 'month', y='Imports_Total', data=years2001, ax = axes[1],palette=("bone"))
plt.xticks(rotation='vertical')
plt.title('Imports Total 2001',fontsize=30)
```
## הערה
אפשר לראות כי מחודש ספטמבר חלה ירידה
# Hanjin Shipping פשיטת הרגל של חברת הענק
```
years_total= data_n.loc[data.month== 'Jan. - Dec.']
years_total=years_total.loc[(years_total.years>2014.0)& (years_total.years<2020.0)]
fig, axes = plt.subplots(figsize=(24, 5), ncols=2)
sns.barplot(x= 'years', y='Imports_Total', data=years_total, ax = axes[0],palette=("icefire_r"))
plt.ylim([0,3500000])
plt.title('Imports Total 2015-2019',fontsize=24)
sns.barplot(x= 'years', y='Exports_Total', data=years_total, ax = axes[1],palette=("icefire_r"))
plt.ylim([0,3000000])
plt.title('Exports Total 2015-2019',fontsize=24)
```
## הערה
ניתן לראות כי בשנים 2017-2018 חלה עליה ביבוא ויבצוא
## מסקנה
בעזרת שלושת האירועים כי הכלכלה העולמית משפיעה על היבוא והיצוא באופן ישיר
# חיזוי המאזן בעזרת ריגרסיה לינארית
```
balance_f=data_n.loc[:,["Balance_Total","years"]]
predict_balance=balance_f.groupby("years")[["Balance_Total"]].mean()
predict_balance["years"]=range(1992,2022)
sum_xi_yi=0
sum_xi=0
sum_yi=0
sum_xi_2=0
for i in range(1992,2022):
sum_xi_yi+=(predict_balance.loc[i])*i
sum_xi+=i
sum_yi+=predict_balance.loc[i]
sum_xi_2+=(i**2)
sum_2xi=sum_xi**2
n=len(predict_balance)
b=((n*sum_xi_yi)-(sum_xi*sum_yi))/((n*sum_xi_2)-sum_2xi)
mean_x=sum_xi/n
mean_y=sum_yi/n
a=mean_y-(b*mean_x)
balance_2022=a+(b*2022)
rigras={1992:0,1993:0,1994:0,1995:0,1996:0,1997:0,1998:0,1999:0,2000:0,2001:0,2002:0,2003:0,2004:0,2005:0,2006:0,2007:0,2008:0,2009:0,2010:0,2011:0,2012:0,2013:0,2014:0,2015:0,2016:0,2017:0,2018:0,2019:0,2020:0,2021:0,2022:0}
for j in range(1992,2023):
rigras[j]=a+(b*j)
r_data=pd.DataFrame(rigras)
rigras_data=r_data.T
rigras_data["years"]=range(1992,2023)
XValues=rigras_data["years"]
YValues=rigras_data["Balance_Total"]
xValues=predict_balance["years"]
yValues=predict_balance["Balance_Total"]
fig, ax1 = plt.subplots()
ax1.set_xlabel('years')
ax1.set_ylabel('Balance Total', color="purple")
ax1.plot(XValues,YValues,color="purple")
ax1.tick_params(axis='y', labelcolor="purple")
ax2 = ax1.twinx()
ax2.set_ylabel('Balance Total regression', color="blue")
ax2.plot(xValues,yValues,color="blue")
ax2.tick_params(axis='y', labelcolor="blue")
fig.tight_layout()
plt.title("Reality vs. Regression",fontsize=24)
plt.show()
```
## הערה
בגרף ניתן לראות את ההפרשים ונקודות החיתוך בין הרגרסיה לנתוני המציאות
## חישוב סטיות
```
ms_sum=rigras_data[["Balance_Total"]]
ms_sum["Balance_Total_rill"]=predict_balance[["Balance_Total"]]
ms_sum["MSE"]=(ms_sum["Balance_Total"]-ms_sum["Balance_Total_rill"])**2
MSE=(ms_sum["MSE"].sum())/n
ms_sum["MAD"]=abs(ms_sum["Balance_Total"]-ms_sum["Balance_Total_rill"])
MAD=(ms_sum["MAD"].sum())/n
ms_sum=ms_sum.dropna()
devi=ms_sum.copy()
devi["MAD"]=sorted(devi["MAD"])
devi["MSE"]=sorted(devi["MSE"])
devi=devi.loc[:2019,["MAD","MSE"]]
fig, axes = plt.subplots(figsize=(10,7), ncols=2)
sns.boxplot(y="MSE", data=devi , ax = axes[0])
plt.title("MSE",fontsize=24)
sns.boxplot(y="MAD", data=devi,ax = axes[1])
plt.title("MAD",fontsize=24)
```
## מסקנה
לפי חישובי הרגרסיה ובעזרת הסטיות ניתן לחזות כי בשנת 2022 סכום המאזן יהיה 114501.292813
| github_jupyter |
```
import torch
import torch.nn as nn
import numpy as np
import os
import matplotlib.pyplot as plt
from torch.autograd import Variable
import torch.utils.data as data_utils
train_path = '/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/new/'
#input_ = np.load('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_input.npy')
#output_ = np.load('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_output.npy')
input_dev = np.load('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_input_dev.npy')
output_dev = np.load('/home/vishesh/TUM/Thesis/Coreference-Resolution/data/processed/ffnn_output_dev.npy')
list_of_train_files = []
list_of_label_files = []
for path, subdirs, files in os.walk(train_path):
for name in files:
if name.startswith("ffnn_train"):
list_of_train_files.append(os.path.join(path, name))
if name.startswith("ffnn_labels"):
list_of_label_files.append(os.path.join(path, name))
list_of_train_files = sorted(list_of_train_files)
list_of_label_files = sorted(list_of_label_files)
len(list_of_train_files)
len(list_of_label_files)
inp = np.load(list_of_train_files[0])
inp[0].shape
input_dev[0].shape
INPUT_DIM = 1237
HIDDEN_DIM1 = 1000
HIDDEN_DIM2 = 500
HIDDEN_DIM3 = 500
OUTPUT_DIM = 2
BATCH_SIZE = 32
NUM_EPOCHS = 10
def dataLoader(start_index, batch_size, inputs, labels, input_dim):
'''
This function provides the data to the network in batches from the starting position start_index.
Args:
start_index (int): Position to start the batch from the dataset.
batch_size (int): Batch size.
inputs: The inputs to the model.
labels: The expected outputs from the model.
Returns:
train_tensor(torch tensor): Tensor of batch size.
label_tensor(torch tensor): Tensor of batch size.
'''
inputs_tensor_list = []
labels_tensor_list = []
if len(inputs[i:]) < batch_size:
batch_size = len(inputs[i:])
for pos in range(start_index, start_index + batch_size):
inputs_tensor_list.append((torch.from_numpy(inputs[pos].reshape(input_dim))).float())
labels_tensor_list.append((torch.from_numpy(labels[pos])))
inputs_tensor = torch.Tensor(batch_size, input_dim)
labels_tensor = torch.LongTensor(batch_size, 1)
torch.cat(inputs_tensor_list, out = inputs_tensor)
torch.cat(labels_tensor_list, out = labels_tensor)
inputs_tensor = inputs_tensor.view(batch_size, 1237)
return inputs_tensor, labels_tensor
def computeF1(model, inputs, labels):
'''
This function computes the F1 score of the model.
Args:
model: The model which is being used to compute the F1 score.
inputs: The inputs to the model.
labels: The expected outputs from the model.
Returns:
tp(int): True Positives.
fp(int): False Positives.
fn(int): False Negaitves.
precision(int): The Precision of the F1 score.
recall(int): The recall of the F1 score.
f1(int): The computed F1 score.
'''
tp = 0
fp = 0
fn = 0
for j in range(0, len(inputs), BATCH_SIZE):
inputs, labels = dataLoader(j, BATCH_SIZE, input_, output_, INPUT_DIM)
inputs = Variable(inputs)
outputs = model(inputs)
_, predicted = torch.max(outputs.data, 1)
for l, p in zip(labels, predicted):
if l == 1 and p == 1:
tp += 1
if l == 0 and p == 1:
fp += 1
if l == 1 and p == 0:
fn += 1
if tp == 0 and fp == 0:
precision = 0
else:
precision = tp / (tp + fp)
if tp == 0 and fn == 0:
recall = 0
else:
recall = tp / (tp + fn)
if precision == 0 and recall == 0:
f1 = 0
else:
f1 = (2 * precision * recall) / (precision + recall)
return tp, fp, fn, precision, recall, f1
class FFNN(nn.Module):
def __init__(self, input_dim, hidden_dim1, hidden_dim2, hidden_dim3, output_dim):
super(FFNN, self).__init__()
self.fc1 = nn.Linear(input_dim, hidden_dim1)
self.relu1 = nn.ReLU()
self.fc2 = nn.Linear(hidden_dim1, hidden_dim2)
self.relu2 = nn.ReLU()
self.fc3 = nn.Linear(hidden_dim2, hidden_dim3)
self.relu3 = nn.ReLU()
self.fc4 = nn.Linear(hidden_dim2, output_dim)
def forward(self, x):
#x = x.view(-1, 1337)
out = self.fc1(x)
out = self.relu1(out)
out = self.fc2(out)
out = self.relu2(out)
out = self.fc3(out)
out = self.relu3(out)
out = self.fc4(out)
return out
model = FFNN(INPUT_DIM, HIDDEN_DIM1, HIDDEN_DIM2, HIDDEN_DIM3, OUTPUT_DIM)
LEARNING_RATE = 1e-1
optimizer = torch.optim.SGD(model.parameters(), lr = LEARNING_RATE)
# Loss Function
criterion = nn.CrossEntropyLoss()
iterations = 0
for epoch in range (1):
for t, l in zip(list_of_train_files, list_of_label_files):
input_ = np.load(t)
output_ = np.load(l)
for i in range(0, len(input_), BATCH_SIZE):
inputs, labels = dataLoader(i, BATCH_SIZE, input_, output_, INPUT_DIM)
inputs , labels = Variable(inputs), Variable(labels)
optimizer.zero_grad()
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
iterations += 1
if iterations % 500 == 0:
tp, fp, fn, precision, recall, f1 = computeF1(model, input_dev, output_dev)
print ('Epoch: ' + str(epoch))
print ('Iteration: ' + str(iterations))
print ('True Positives: ' + str(tp))
print ('False Positives: ' + str(fp))
print ('False Negatives: ' + str(fn))
print ('F1: ' + str(f1))
for i in range(0, 5, 2):
print (i)
```
| github_jupyter |
# Swiggy Data Science Assessment - LinkedIn, MTV Get a Job
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
from sklearn.cluster import KMeans
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
```
## Import Dataset and Quick Inspection
```
dataset_path = './data/SampleAssessment.csv'
df = pd.read_csv(dataset_path)
df.head()
df.tail()
df.columns = ['customer_id', 'first_time', 'recent_time', 'no_of_orders', 'orders_7_days', 'orders_last_4_weeks', 'amount', 'amt_last_7_days', 'amt_in_last_4_weeks', 'avg_dist_rest', 'avg_del_time']
df.info()
df.describe().round(2)
df['first_time'] = pd.to_datetime(df['first_time'])
df['recent_time'] = pd.to_datetime(df['recent_time'])
df.head()
```
## Number of missing values (orders in 7 days and last 4 weeks)
```
df.isna().sum()
```
## Verifying whether the amount is zero for no orders
```
(df['amt_last_7_days'] == 0).sum(axis=0)
(df['amt_in_last_4_weeks'] == 0).sum(axis=0)
```
## Description of the subset of dataset with negative restaurant distances
```
df[df['avg_dist_rest'] < 0].describe().round(2)
```
## Description of the original dataset with negative restaurant distances
```
df.describe().round(2)
```
## Description of the dataset without negative restaurant distances
```
df[df['avg_dist_rest'] > 0].describe().round(2)
```
* The mean and standard deviation are not much affected by the removal of negative disatances
* The above dataset will be use for further evaluations
* The rows with negative values are thereby discarded
```
df = df[df['avg_dist_rest'] > 0]
df.describe().round(2)
```
## Correlation
```
df.corr()
```
* It appears that there are no other important correlations other than order vs amount
```
plt.figure(figsize=(20,10))
plt.title('Number of orders vs Amount spent', fontsize=26)
plt.xlabel('Number of Orders', fontsize=24)
plt.ylabel('Amount Spent', fontsize=24)
plt.scatter(df['no_of_orders'], df['amount'])
plt.savefig('./plots/orders_vs_amount.png', format='png', dpi=1000)
plt.show()
plt.figure(figsize=(20,10))
plt.title('Distribution of amount', fontsize=26)
plt.ylabel('Amount Spent', fontsize=24)
plt.hist(df['amount'], bins=[0, 10000, 20000, 30000, 40000, 50000, 60000, 70000])
plt.savefig('./plots/distribution_amount.png', format='png', dpi=300)
plt.show()
plt.figure(figsize=(20,10))
plt.title('Distribution of distances', fontsize=26)
plt.ylabel('Average Distances', fontsize=24)
plt.hist(df['avg_dist_rest'])
plt.savefig('./plots/distribution_distances.png', format='png', dpi=300)
plt.show()
plt.figure(figsize=(20,10))
plt.title('Distribution of delivery time', fontsize=26)
plt.ylabel('Average Delivery Time', fontsize=24)
plt.hist(df['avg_del_time'])
plt.savefig('./plots/distribution_delivery.png', format='png', dpi=300)
plt.show()
df.head()
```
# Calculating customer Recency
```
latest_recent_time = df['recent_time'].max()
df['recency'] = df['recent_time'].apply(lambda x: (latest_recent_time - x).days)
```
## Delivery score
* A custom score given to every customer
```
df['delivery_score'] = df['avg_del_time'] / df['avg_dist_rest']
df.head()
quantiles = df.quantile(q=[0.25,0.5,0.75])
quantiles.to_dict()
# Arguments (x = value, p = recency, monetary_value, frequency, d = quartiles dict)
def RScore(x, p, d):
if x <= d[p][0.25]:
return 4
elif x <= d[p][0.50]:
return 3
elif x <= d[p][0.75]:
return 2
else:
return 1
# Arguments (x = value, p = recency, monetary_value, frequency, d = quartiles dict)
def FMScore(x, p, d):
if x <= d[p][0.25]:
return 1
elif x <= d[p][0.50]:
return 2
elif x <= d[p][0.75]:
return 3
else:
return 4
```
## RFM Segemtation for last 7 days of customer orders
```
rfm_segmentation_7_days = df
rfm_segmentation_7_days['r_quartile_7_days'] = rfm_segmentation_7_days['recency'].apply(RScore, args=('recency',quantiles,))
rfm_segmentation_7_days['f_quartile_7_days'] = rfm_segmentation_7_days['orders_7_days'].apply(FMScore, args=('orders_7_days',quantiles,))
rfm_segmentation_7_days['m_quartile_7_days'] = rfm_segmentation_7_days['amt_last_7_days'].apply(FMScore, args=('amt_last_7_days',quantiles,))
rfm_segmentation_7_days.head()
```
## RFM Segemtation for last 4 weeks of customer orders
```
rfm_segmentation_4_weeks = df
rfm_segmentation_4_weeks['r_quartile_4_weeks'] = rfm_segmentation_4_weeks['recency'].apply(RScore, args=('recency',quantiles,))
rfm_segmentation_4_weeks['f_quartile_4_weeks'] = rfm_segmentation_4_weeks['orders_last_4_weeks'].apply(FMScore, args=('orders_last_4_weeks',quantiles,))
rfm_segmentation_4_weeks['m_quartile_4_weeks'] = rfm_segmentation_4_weeks['amt_in_last_4_weeks'].apply(FMScore, args=('amt_in_last_4_weeks',quantiles,))
rfm_segmentation_4_weeks.head()
```
## RFMScore = (r * f * m) * delivery_score
```
rfm_segmentation_7_days['RFMScore_7_days'] = rfm_segmentation_7_days.r_quartile_7_days \
* rfm_segmentation_7_days.f_quartile_7_days \
* rfm_segmentation_7_days.m_quartile_7_days \
* rfm_segmentation_7_days.delivery_score
rfm_segmentation_7_days.head()
rfm_segmentation_7_days.describe()
```
## RFMScore = (r * f * m) * delivery_score
```
rfm_segmentation_4_weeks['RFMScore_4_weeks'] = rfm_segmentation_4_weeks.r_quartile_4_weeks \
* rfm_segmentation_4_weeks.f_quartile_4_weeks \
* rfm_segmentation_4_weeks.m_quartile_4_weeks \
* rfm_segmentation_4_weeks.delivery_score
rfm_segmentation_4_weeks.head()
rfm_segmentation_7_days.describe()['RFMScore_7_days']
rfm_segmentation_4_weeks.describe()['RFMScore_4_weeks']
```
## Moving on to K Means Clustering
```
rfm_segmentation_7_days_values = rfm_segmentation_7_days[['amt_last_7_days', 'RFMScore_7_days']].values
rfm_segmentation_4_weeks_values = rfm_segmentation_4_weeks[['amt_in_last_4_weeks', 'RFMScore_4_weeks']].values
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters=i, init='k-means++', max_iter=300, n_init=10, random_state=0)
kmeans.fit(rfm_segmentation_7_days_values)
wcss.append(kmeans.inertia_)
plt.figure(figsize=(20,10))
plt.plot(range(1, 11), wcss)
plt.title('Finding the number of clusters - Elbow curve - last 7 days', fontsize=26)
plt.xlabel('Number of clusters', fontsize=24)
plt.ylabel('WCSS')
plt.savefig('./plots/7_days_elbow.png', format='png', dpi=300)
plt.show()
```
### The curve says that 2 would be the ideal number of clusters
```
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters=i, init='k-means++', max_iter=300, n_init=10, random_state=0)
kmeans.fit(rfm_segmentation_4_weeks_values)
wcss.append(kmeans.inertia_)
plt.figure(figsize=(20,10))
plt.plot(range(1, 11), wcss)
plt.title('Finding the number of clusters - Elbow curve - last 4 weeks', fontsize=26)
plt.xlabel('Number of clusters', fontsize=24)
plt.ylabel('WCSS')
plt.savefig('./plots/4_weeks_elbow.png', format='png', dpi=300)
plt.show()
```
### The curve says that 3 would be the ideal number of clusters
## Applying K Means to customer data of past 7 days
```
kmeans = KMeans(n_clusters=2, init='k-means++', max_iter=300, n_init=10, random_state=0)
y_kmeans = kmeans.fit_predict(rfm_segmentation_7_days_values)
# Visualizing the clusters
plt.figure(figsize=(20,10))
plt.scatter(rfm_segmentation_7_days_values[y_kmeans==0, 0], rfm_segmentation_7_days_values[y_kmeans==0, 1], s=100, c='red', label='Cluster 1', alpha=0.5)
plt.scatter(rfm_segmentation_7_days_values[y_kmeans==1, 0], rfm_segmentation_7_days_values[y_kmeans==1, 1], s=100, c='blue', label='Cluster 2', alpha=0.5)
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:, 1], s=300, c='yellow', label='Centroids')
plt.title('Clusters of customers for last 7 days', fontsize=26)
plt.xlabel('Amount', fontsize=24)
plt.ylabel('RFMScore', fontsize=24)
plt.legend(prop={'size': 30})
plt.savefig('./plots/k_means_7_days.png', format='png', dpi=300)
plt.show()
```
## Applying K Means to customer data of past 4 weeks
```
kmeans = KMeans(n_clusters=3, init='k-means++', max_iter=300, n_init=10, random_state=0)
y_kmeans = kmeans.fit_predict(rfm_segmentation_4_weeks_values)
# Visualizing the clusters
plt.figure(figsize=(20,10))
plt.scatter(rfm_segmentation_4_weeks_values[y_kmeans==0, 0], rfm_segmentation_4_weeks_values[y_kmeans==0, 1], s=100, c='red', label='Cluster 1', alpha=0.5)
plt.scatter(rfm_segmentation_4_weeks_values[y_kmeans==1, 0], rfm_segmentation_4_weeks_values[y_kmeans==1, 1], s=100, c='blue', label='Cluster 2', alpha=0.5)
plt.scatter(rfm_segmentation_4_weeks_values[y_kmeans==2, 0], rfm_segmentation_4_weeks_values[y_kmeans==2, 1], s=100, c='green', label='Cluster 3', alpha=0.5)
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:, 1], s=300, c='yellow', label='Centroids')
plt.title('Clusters of customers for last 4 weeks', fontsize=26)
plt.xlabel('Amount', fontsize=24)
plt.ylabel('RFMScore', fontsize=24)
plt.legend(prop={'size': 30})
plt.savefig('./plots/k_means_4_weeks.png', format='png', dpi=300)
plt.show()
```
# Conclusion
* In a dataset of 10000 rows
* Recent orders (Last 7 days) can be categorized into two parts. One being least spending customers and the others are returning customers
* Past orders (Last 4 weeks) can be categorized into three parts. One being the least spending, another being loyal customers and the other being returning customers
* The customers in least spending customers (Red Cluster) have more orders but less amount spent
* The customers who are loyal (Green Cluster) are more like to spend money along with more orders
* The customers who are returning (Blue Cluster) are more likely to spend money eventhough their order quantity is less
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn import model_selection
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
#from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import GradientBoostingClassifier
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
#time
from datetime import datetime
from datetime import timedelta
import jieba
import jieba.analyse
jieba.set_dictionary('dict.idkrsi.txt') # 改預設字典
jieba.analyse.set_stop_words("stopword.goatwang.kang.txt") #指定stopwords字典
# get data
# ! conda install pandas-datareader s
#import pandas_datareader as pdr
# visual
# ! pip install mpl-finance
#import matplotlib.pyplot as plt
#import mpl_finance as mpf
#import seaborn as sns
# https://github.com/mrjbq7/ta-lib
# ! pip install ta-lib
#import talib
df_bbs = pd.read_csv("bda2019_dataset/bbs2.csv",encoding="utf-8")
df_forum = pd.read_csv("bda2019_dataset/forum2.csv",encoding="utf-8")
df_news = pd.read_csv("bda2019_dataset/news2.csv",encoding="utf-8")
df_news['comment_count']=0
df_article = pd.concat([df_forum, df_bbs, df_news]) #三個合併
del df_bbs, df_forum, df_news
df_article['post_time'] = pd.to_datetime(df_article['post_time'])
df_article['post_time2'] = df_article['post_time'].dt.date # .dt.date用在dataframe .date()用在一個 #只留日期
#df_article['label'] = 'even'
df_article['content'] = df_article['content'].astype(str).str.replace(',' , ' ').str.replace('\n' , ' ').str.replace('"' , ' ').str.replace("'" , ' ')
df_article['title'] = df_article['title'].astype(str).str.replace(',' , ' ').str.replace('\n' , ' ').str.replace('"' , ' ').str.replace("'" , ' ')
df_article = df_article.sort_values(by=['post_time']).reset_index(drop=True) # 用post_time排序 # 在重設index
df_article = df_article.rename(index=str, columns={"author": "author_", "content": "content_", "id": "id_", "title": "title_"}) # 換column名 以免跟切詞重複
df_article.head(2)
#df_article2 = df_article[['post_time2','title','content']]
df_TWSE2018 = pd.read_csv("bda2019_dataset/TWSE2018.csv",encoding="utf-8")
df_TWSE2017 = pd.read_csv("bda2019_dataset/TWSE2017.csv",encoding="utf-8")
df_TWSE2016 = pd.read_csv("bda2019_dataset/TWSE2016.csv",encoding="utf-8")
df_TWSE = pd.concat([df_TWSE2016, df_TWSE2017, df_TWSE2018]) #三年合併
del df_TWSE2016, df_TWSE2017, df_TWSE2018
# ['開盤價(元)', '最高價(元)', '最低價(元)', '收盤價(元)', '成交量(千股)', '成交值(千元)', '成交筆數(筆)', '流通在外股數(千股)', '本益比-TSE', '股價淨值比-TSE']
df_TWSE['證券代碼'] = df_TWSE['證券代碼'].astype(str)
df_TWSE['年月日'] = pd.to_datetime(df_TWSE['年月日'])
df_TWSE['開盤價(元)'] = df_TWSE['開盤價(元)'].str.replace(',' , '').astype('float64') # 1,000 to 1000 to float
df_TWSE['最高價(元)'] = df_TWSE['最高價(元)'].str.replace(',' , '').astype('float64')
df_TWSE['最低價(元)'] = df_TWSE['最低價(元)'].str.replace(',' , '').astype('float64')
df_TWSE['收盤價(元)'] = df_TWSE['收盤價(元)'].str.replace(',' , '').astype('float64')
df_TWSE['成交量(千股)'] = df_TWSE['成交量(千股)'].str.replace(',' , '').astype('float64')
df_TWSE['成交值(千元)'] = df_TWSE['成交值(千元)'].str.replace(',' , '').astype('float64')
df_TWSE['成交筆數(筆)'] = df_TWSE['成交筆數(筆)'].str.replace(',' , '').astype('int64')
df_TWSE['流通在外股數(千股)'] = df_TWSE['流通在外股數(千股)'].str.replace(',' , '').astype('float64')
df_TWSE['本益比-TSE'] = df_TWSE['本益比-TSE'].str.replace(',' , '').astype('float64')
df_TWSE['股價淨值比-TSE'] = df_TWSE['股價淨值比-TSE'].astype('float64')
df_TWSE.head(2)
# 選那家股票
#company_name = '國巨'
company_name = '奇力新'
# 文章包含那家字
#company_words = '被動元件|積層陶瓷電容|MLCC|電感|晶片電阻|車用電子|凱美|同欣電|大毅|君耀|普斯|國巨'
company_words = '被動元件|積層陶瓷電容|MLCC|電感|晶片電阻|車用電子|飛磁|旺詮|美磊|美桀|向華科技|奇力新'
# 漲跌幾%
PA = 0.05
# even幾%
PAE = 0.003
# 用日期排序 再把index重排
#2327
#df_trend = df_TWSE[df_TWSE['證券代碼'].str.contains('國巨')].sort_values(by=['年月日']).reset_index(drop=True)
#2456
#df_trend = df_TWSE[df_TWSE['證券代碼'].str.contains('奇力新')].sort_values(by=['年月日']).reset_index(drop=True)
#2478
#df_trend = df_TWSE[df_TWSE['證券代碼'].str.contains('大毅')].sort_values(by=['年月日']).reset_index(drop=True)
#6271
#df_trend = df_TWSE[df_TWSE['證券代碼'].str.contains('同欣電')].sort_values(by=['年月日']).reset_index(drop=True)
df_trend = df_TWSE[df_TWSE['證券代碼'].str.contains(company_name)].sort_values(by=['年月日']).reset_index(drop=True)
del df_TWSE
df_trend.head(2)
##增欄位:fluctuation幅度 tag漲跌平
df_trend['fluctuation'] = 0.0
df_trend['tag']='--'
df_trend['closeshift'] = 0.0
df_trend.head(2)
# ##增欄位:fluctuation幅度 tag漲跌平
# df_trend['fluctuation'] = 0.0
# df_trend['tag']='--'
# ###計算漲跌
# for index, row in df_trend.iterrows():
# try:
# margin =(float(df_trend.loc[index,'收盤價(元)']) - float(df_trend.loc[index-1,'收盤價(元)']) )/ float(df_trend.loc[index-1,'收盤價(元)'])
# df_trend.loc[index,'fluctuation']=margin
# if margin >=0.03:
# df_trend.loc[index,'tag']='up'
# elif margin <= -0.03:
# df_trend.loc[index,'tag']='down'
# else:
# df_trend.loc[index,'tag']='even'
# except:
# continue
df_trend['closeshift'] = df_trend['收盤價(元)'].shift(periods=1)#.fillna(value=0.0, inplace=True)
#df_trend['closeshift'].fillna(value= 0.0, inplace=True)
df_trend.head(2)
df_trend['fluctuation'] = (df_trend['收盤價(元)'] - df_trend['closeshift']) / df_trend['closeshift']
df_trend.head(2)
print('fluctuation std = ',df_trend['fluctuation'].std(axis=0))
print('fluctuation mean = ',df_trend['fluctuation'].mean(axis=0))
df_trend.loc[df_trend['fluctuation'] >= PA, 'tag'] = 'up'
df_trend.loc[df_trend['fluctuation'] <= -PA, 'tag'] = 'down'
df_trend.loc[(df_trend['fluctuation'] >= -PAE) & (df_trend['fluctuation'] <= PAE), 'tag'] = 'even'
df_trend.head(2)
len(df_trend[df_trend['tag']=='up'])
len(df_trend[df_trend['tag']=='down'])
len(df_trend[df_trend['tag']=='even'])
#df_company = df_article[ df_article['content'].str.contains('國巨')] # df 某欄位 string contains "國巨"
#df_company = df_article[ df_article['content'].str.contains('奇力新')]
#df_company = df_article[ df_article['content'].str.contains('大毅')]
#df_company = df_article[ df_article['content'].str.contains('同欣電 ')]
df_company = df_article[ df_article['content_'].str.contains(company_words)]
print(len(df_company))
del df_article
df_company.head(2)
stopwords=list()
with open('stopword.goatwang.kang.txt', 'r',encoding='utf-8') as data:
for stopword in data:
stopwords.append(stopword.strip('\n'))
# 'content'全部切詞
corpus = [] # array
for index, row in df_company.iterrows():
not_cut = df_company.loc[index,'content_']
# not_cut = row['description'] # 跟上一行一樣意思
seg_generator = jieba.cut(not_cut, cut_all=False) # genarator
seglist = list(seg_generator) # 整篇文章string切出來的list
# seglist = list(filter(lambda a: a not in stopwords and a != '\n', seglist )) #去除停用詞 #未必需要這步驟
corpus.append(' '.join(seglist)) # ' '.join(seg_generator)也可
df_company["content2"]=corpus
df_company.head(2)
df_trend.loc[2,'年月日'].date() + timedelta(days=-1) == df_trend.loc[1,'年月日'].date()
df_trend.loc[5,'年月日'].date() + timedelta(days=-1) == df_trend.loc[4,'年月日'].date()
d = df_trend.loc[1,'年月日'].date() - df_trend.loc[ 1-1 ,'年月日'].date() #相減差幾天
d
d.days #只取天數
int(d.days) #幾天 轉整數
df_trend.loc[3,'年月日'].date()
df_company[ df_company['post_time2'] == df_trend.loc[3,'年月日'].date() ].head() # 某欄位 == n 的 全部撈出來
# # 演算法
# for index, row in df_2327.iterrows():
# try:
# if df_2327.loc[index,'年月日'].date() + timedelta(days=-1) == df_2327.loc[index-1,'年月日'].date():
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-1), 'label'] = df_2327.loc[index,'tag']
# # 如果股票前一筆差1天 # 那前1天的文章標上當天的漲跌
# elif df_2327.loc[index,'年月日'].date() + timedelta(days=-2) == df_2327.loc[index-1,'年月日'].date():
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-1), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-2), 'label'] = df_2327.loc[index,'tag']
# # 如果股票前一筆差2天 #那前2天的文章標上當天的漲跌
# elif df_2327.loc[index,'年月日'].date() + timedelta(days=-3) == df_2327.loc[index-1,'年月日'].date():
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-1), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-2), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-3), 'label'] = df_2327.loc[index,'tag']
# elif df_2327.loc[index,'年月日'].date() + timedelta(days=-4) == df_2327.loc[index-1,'年月日'].date():
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-1), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-2), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-3), 'label'] = df_2327.loc[index,'tag']
# df_forum.loc[df_forum['post_time2'] == df_2327.loc[index,'年月日'].date() + timedelta(days=-4), 'label'] = df_2327.loc[index,'tag']
# except:
# continue
# 看所有相差的天數
# for index, row in df_2327.iterrows():
# try:
# n = df_2327.loc[index,'年月日'].date() - df_2327.loc[index-1,'年月日'].date()
# print(n)
# except:
# continue
# 最多12天
# 如果股票前一筆差n天 # 那前n天的文章標上當天的漲跌
df_company['label5566']='--'
for index, row in df_trend.iterrows():
try:
n = int((df_trend.loc[index,'年月日'].date() - df_trend.loc[index-1,'年月日'].date()).days ) # 差幾個datetime # 轉天數 # 再轉整數
# print(n)
for i in range(1, n+1):
# print(i)
df_company.loc[df_company['post_time2'] == df_trend.loc[index,'年月日'].date() + timedelta(days=-i), 'label5566'] = df_trend.loc[index,'tag']
except:
continue
print(len(df_company[df_company['label5566']=='down']))
df_company[df_company['label5566']=='down'].head(2)
print(len(df_company[df_company['label5566']=='up']))
df_company[df_company['label5566']=='up'].head(2)
print(len(df_company[df_company['label5566']=='even']))
df_company[df_company['label5566']=='even'].head(2)
#df_company2 = df_company[df_company['label5566'].str.contains('up|down|even')]
#df_company2.to_csv('5pa.csv')
df_keyword = pd.read_csv("chi_5pa_word.csv",encoding="utf-8")
df_keyword.head()
features = df_keyword['word'].astype(str).to_numpy()
features = list(features)
# df_keyword1 = pd.read_csv("final_higher_tf_idf_part.csv",encoding="utf-8") #上漲形容詞
# df_keyword2 = pd.read_csv("final_lower_tf_idf_part.csv",encoding="utf-8") #下跌形容詞
# df_keyword = pd.concat([df_keyword1,df_keyword2])
# del df_keyword1,df_keyword2
# df_keyword.head()
# features = df_keyword['key'].astype(str).to_numpy()
# features = list(features)
# import re
# features = [] # features=list()
# with open('finance.words.txt', 'r',encoding='utf-8') as data:
# for line in data:
# # line = re.sub('[a-zA-Z0-9\W]', '', line) # 把數字英文去掉
# line = re.sub('[0-9]', '', line) # 把數字去掉
# features.append(line.replace('\n', '').replace(' ', '')) # 空格 \n去掉
# print(len(features))
# print(type(features))
# features[:10]
from sklearn.feature_extraction.text import TfidfVectorizer
#features = [ '上漲','下跌','看好','走高','走低','漲停','跌停']
features = features[:20000]
#cv = TfidfVectorizer() #預設有空格就一個feature
cv = TfidfVectorizer(vocabulary = features) # 設定自己要的詞
r = pd.SparseDataFrame(cv.fit_transform(df_company['content2']),
df_company.index,
cv.get_feature_names(),
default_fill_value=0.0)
r.fillna(value=0.0, inplace=True)
r.head(2)
# from sklearn.feature_extraction.text import CountVectorizer
# #features = [ '上漲','下跌','看好','走高','走低','漲停','跌停']
# #features = features[:1000]
# #cv = CountVectorizer() #預設有空格就一個feature
# cv = CountVectorizer(vocabulary = features) # 設定自己要的詞
# r = pd.SparseDataFrame(cv.fit_transform(df_company['content2']),
# df_company.index,
# cv.get_feature_names(),
# default_fill_value=0)
# r.head(2)
df_company2 = pd.concat([df_company,r], axis=1)
df_company2.head(2)
df_company2 = df_company2[df_company2['label5566'].str.contains('up|down')] #只取漲跌
df_train = df_company2[(df_company2['post_time'] >= '2016-1-1 00:00:00') & (df_company2['post_time'] < '2018-10-1 00:00:00')]
df_validation = df_company2[(df_company2['post_time'] >= '2018-10-1 00:00:00') & (df_company2['post_time'] < '2019-1-1 00:00:00')]
print(len(df_train))
print(len(df_validation))
print(df_train['label5566'].value_counts()) # 類別數 count
print(df_validation['label5566'].value_counts())
seed = 7
df_train_up = df_train[df_train['label5566']=='up'] # oversampling 讓類別比例平均
df_train_up = df_train_up.sample(998, replace=True, random_state=seed)
df_train_down = df_train[df_train['label5566']=='down']
df_train_down = df_train_down.sample(998, replace=True, random_state=seed)
#df_train_even = df_train[df_train['label5566']=='even']
#df_train_even = df_train_even.sample(998, replace=True, random_state=seed)
#df_train = pd.concat( [df_train_up, df_train_down, df_train_even] )
df_train = pd.concat( [df_train_up, df_train_down] )
df_train['label5566'].value_counts()
# seed = 7
# df_validation_up = df_validation[df_validation['label5566']=='up'] # oversampling 讓類別比例平均
# df_validation_up = df_validation_up.sample(286, replace=True, random_state=seed)
# df_validation_down = df_validation[df_validation['label5566']=='down']
# df_validation_down = df_validation_down.sample(286, replace=True, random_state=seed)
# #df_validation_even = df_validation[df_validation['label5566']=='even']
# #df_validation_even = df_validation_even.sample(286, replace=True, random_state=seed)
# #df_validation = pd.concat( [df_validation_up, df_validation_down, df_validation_even] )
# df_validation = pd.concat( [df_validation_up, df_validation_down,] )
# df_validation['label5566'].value_counts()
X_train = df_train[features] # features 要 轉成list 用numpy出了問題
X_train.fillna(value=0.0, inplace=True)
X_train = X_train.to_numpy()
X_validation = df_validation[features]
X_validation.fillna(value=0.0, inplace=True)
X_validation = X_validation.to_numpy()
Y_train = df_train['label5566']
Y_validation = df_validation['label5566']
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
model_RandomForest = RandomForestClassifier()
name = 'RandomForest'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_RandomForest, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
model_RandomForest = RandomForestClassifier()
model_RandomForest.fit(X_train, Y_train)
print(model_RandomForest.score(X_train, Y_train))
predictions = model_RandomForest.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
from lightgbm.sklearn import LGBMClassifier
model_LGBMClassifier = LGBMClassifier()
name = 'LGBMClassifier'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_LGBMClassifier, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
from lightgbm.sklearn import LGBMClassifier
model_LGBMClassifier = LGBMClassifier()
model_LGBMClassifier.fit(X_train, Y_train)
print(model_LGBMClassifier.score(X_train, Y_train))
predictions = model_LGBMClassifier.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
model_XGBClassifier = XGBClassifier()
model_XGBClassifier.fit(X_train, Y_train)
print(model_XGBClassifier.score(X_train, Y_train))
predictions = model_XGBClassifier.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
model_GradientBoost = GradientBoostingClassifier()
model_GradientBoost.fit(X_train, Y_train)
print(model_GradientBoost.score(X_train, Y_train))
predictions = model_GradientBoost.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
model_DecisionTree = DecisionTreeClassifier()
model_DecisionTree.fit(X_train, Y_train)
print(model_DecisionTree.score(X_train, Y_train))
predictions = model_DecisionTree.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
# # 用10-Fold CV並且列出平均的效率
# from sklearn.model_selection import cross_validate
# from sklearn.model_selection import RepeatedStratifiedKFold
# # 呼叫單個model MLP
# model_KNeighbors = KNeighborsClassifier()
# name = 'KNeighbors'
# seed = 7
# kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
# cv_results = model_selection.cross_validate(model_KNeighbors, X_train, Y_train, cv=kfold, scoring='accuracy')
# #model用MLP() cross valitation
# print(cv_results['test_score'])
# print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
# print(cv_results['train_score'])
# print(cv_results['fit_time'])
# print(cv_results['score_time'])
model_KNeighbors = KNeighborsClassifier()
model_KNeighbors.fit(X_train, Y_train)
print(model_KNeighbors.score(X_train, Y_train))
predictions = model_KNeighbors.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
model_GaussianNB = GaussianNB()
model_GaussianNB.fit(X_train, Y_train)
print(model_GaussianNB.score(X_train, Y_train))
predictions = model_GaussianNB.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
model_MultinomialNB = MultinomialNB()
name = 'MultinomialNB'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_MultinomialNB, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
model_MultinomialNB = MultinomialNB()
model_MultinomialNB.fit(X_train, Y_train)
print(model_MultinomialNB.score(X_train, Y_train) )
predictions = model_MultinomialNB.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
print(model_MultinomialNB.coef_)
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
model_LogisticRegression = LogisticRegression()
name = 'LogisticRegression'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_LogisticRegression, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
model_LogisticRegression = LogisticRegression()
model_LogisticRegression.fit(X_train, Y_train)
print(model_LogisticRegression.score(X_train, Y_train))
predictions = model_LogisticRegression.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
print(model_LogisticRegression.coef_)
model_svclinear = SVC(kernel='linear')
model_svclinear.fit(X_train, Y_train)
print(model_svclinear.score(X_train, Y_train))
predictions = model_svclinear.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
print(model_svclinear.coef_)
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
from sklearn.svm import LinearSVC
model_LinearSVC = LinearSVC()
name = 'LinearSVC'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_LinearSVC, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
from sklearn.svm import LinearSVC
model_LinearSVC = LinearSVC()
model_LinearSVC.fit(X_train, Y_train)
print(model_LinearSVC.score(X_train, Y_train))
predictions = model_LinearSVC.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
print(model_LinearSVC.coef_)
# 用10-Fold CV並且列出平均的效率
from sklearn.model_selection import cross_validate
from sklearn.model_selection import RepeatedStratifiedKFold
# 呼叫單個model MLP
from sklearn.linear_model import SGDClassifier
model_SGDClassifier = SGDClassifier(loss='hinge')
name = 'SGDClassifier'
seed = 7
kfold = model_selection.RepeatedStratifiedKFold(n_splits=5, n_repeats=1, random_state=seed) #分割 10% cross validation
cv_results = model_selection.cross_validate(model_SGDClassifier, X_train, Y_train, cv=kfold, scoring='accuracy')
#model用MLP() cross valitation
print(cv_results['test_score'])
print("%s: %f (%f)" % (name, cv_results['test_score'].mean(), cv_results['test_score'].std()))
print(cv_results['train_score'])
print(cv_results['fit_time'])
print(cv_results['score_time'])
from sklearn.linear_model import SGDClassifier
model_SGDClassifier = SGDClassifier(loss='hinge')
model_SGDClassifier.fit(X_train, Y_train)
print(model_SGDClassifier.score(X_train, Y_train))
predictions = model_SGDClassifier.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
print(model_SGDClassifier.coef_)
model_MLP = MLPClassifier(hidden_layer_sizes=(256, 256,), max_iter=256)
model_MLP.fit(X_train, Y_train)
print(model_MLP.score(X_train, Y_train))
predictions = model_MLP.predict(X_validation)
print(accuracy_score(Y_validation, predictions))
print(confusion_matrix(Y_validation, predictions))
print(classification_report(Y_validation, predictions))
#print(model_MLP.coefs_)
#df_company2.to_csv('5pa.csv')
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# GR HD Equations
## Author: Zach Etienne
### Formatting improvements courtesy Brandon Clark
$\newcommand{\be}{\begin{equation}}$
$\newcommand{\ee}{\end{equation}}$
$\newcommand{\grad}{{\boldsymbol{\nabla}}}$
$\newcommand{\vel}{{\boldsymbol{v}}}$
$\newcommand{\mom}{{\boldsymbol{p}}}$
$\newcommand{\ddt}[1]{{\frac{\partial #1}{\partial t}}}$
$\newcommand{\ddx}[1]{{\frac{\partial #1}{\partial x}}}$
$\newcommand{\state}{{\boldsymbol{\mathcal{U}}}}$
$\newcommand{\charge}{{\boldsymbol{U}}}$
$\newcommand{\psicharge}{{\boldsymbol{\psi}}}$
$\newcommand{\lapse}{\alpha}$
$\newcommand{\shift}{\boldsymbol{\beta}}$
$\newcommand{\rhostar}{{\rho_*}}$
$\newcommand{\tautilde}{{\tilde{\tau}}}$
$\newcommand{\Svectilde}{{\tilde{\boldsymbol{S}}}}$
$\newcommand{\rtgamma}{{\sqrt{\gamma}}}$
$\newcommand{\T}[2]{{T^{#1 #2}}}$
$\newcommand{\uvec}{{\boldsymbol{u}}}$
$\newcommand{\Vvec}{{\boldsymbol{\mathcal{V}}}}$
$\newcommand{\vfluid}{{\boldsymbol{v}_{\rm f}}}$
$\newcommand{\vVal}{{\tilde{\boldsymbol{v}}}}$
$\newcommand{\flux}{{\boldsymbol{\mathcal{F}}}}$
$\newcommand{\fluxV}{{\boldsymbol{F}}}$
$\newcommand{\source}{{\boldsymbol{\mathcal{S}}}}$
$\newcommand{\sourceV}{{\boldsymbol{S}}}$
$\newcommand{\area}{{\boldsymbol{A}}}$
$\newcommand{\normal}{{\hat{\boldsymbol{n}}}}$
$\newcommand{\pt}{{\boldsymbol{p}}}$
$\newcommand{\nb}{{\boldsymbol{n}}}$
$\newcommand{\meshv}{{\boldsymbol{w}}}$
$\newcommand{\facev}{{\boldsymbol{\tilde{w}}_{ij}}}$
$\newcommand{\facer}{{\boldsymbol{\tilde{r}}_{ij}}}$
$\newcommand{\meshr}{{\boldsymbol{r}}}$
$\newcommand{\cmr}{{\boldsymbol{c}}}$
## Introduction:
We start out with the ** GRHD ** equations in conservative form with the state vector $\state=(\rhostar, \Svectilde, \tautilde)$:
\begin{equation}
\ddt{\state} + \grad\cdot\flux = \source,
\end{equation}
where $\rhostar = \lapse\rho\rtgamma u^0$, $\Svectilde = \rhostar h \uvec$, $\tautilde = \lapse^2\rtgamma \T00 - \rhostar$. The associated set of primitive variables are $(\rho, \vel, \epsilon)$, which are the rest mass density, fluid 3-velocity, and internal energy (measured in the rest frame).
The flux, $\flux$ is given by
\begin{equation}
\flux=(\rhostar \vel, \lapse\rtgamma\T{j}{\beta}g_{\beta i}, \lapse^2\rtgamma\T0j - \rhostar\vel
\end{equation}
where $\vel$ is the 3-velocity, and $\source = (0, \frac 1 2 \lapse\rtgamma \T{\lapse}{\beta}g_{\lapse\beta,i}, s)$ is the source function, and
\begin{equation}
s = \lapse\rtgamma\left[\left(\T00\beta^i\beta^j + 2\T0i\beta^j\right)K_{ij} - \left(\T00\beta^i + \T0i\right)\partial_i\lapse\right]
\end{equation}
The stress energy tensor for a perfect fluid is written as
\begin{equation}
\T{\mu}{\nu} = \rho h u^{\mu} u^{\nu} + P g^{\mu\nu},
\end{equation}
where $h = 1 + \epsilon + P/\rho$ is the specific enthalpy and $u^{\mu}$ are the respective components of the four velocity.
Noting that the mass $\flux$ is defined in terms of $\rhostar$ and $\vel$, we need to first find a mapping between $\vel$ and $u$.
### Alternative formulation
The Athena++ folks have an alternative formulations that might be superior.
Begin with the continuity equation
\begin{equation}
\grad_{\mu}\rho u^{\mu} = 0,
\end{equation}
where $\grad$ is the covariant derivative. This can be mapped directly to
\begin{equation}
\partial_{0} \sqrt{-g}\rho u^0 + \partial_i\sqrt{-g} \rho u^0 v^i = 0
\end{equation}
which we can identify with $\rhostar = \alpha\rtgamma \rho u^0$ because $\sqrt{-g} = \alpha\rtgamma$.
Now the second equation is conservation of energy-momentum which we write as
\begin{equation}
\grad_{\nu}T^{\nu}_{\mu} = 0
\end{equation}
writing this out we have
\begin{equation}
\partial_0 g_{\mu\alpha}T^{\alpha 0} + \partial_i g_{\mu\alpha}T^{\alpha i} - \Gamma_{\mu\alpha}^{\gamma} g_{\gamma\beta}T^{\alpha\beta} = 0
\end{equation}
Noting that
\begin{equation}
\Gamma^{\alpha}_{\beta\gamma} = \frac 1 2 g^{\alpha\delta}\left(\partial_{\gamma}g_{\beta\delta} + \partial_{\beta}g_{\gamma\delta} - \partial_{\delta}g_{\beta\gamma}\right)
\end{equation}
Writing this all out, we note the last term is
\begin{equation}
\Gamma_{\mu\alpha}^{\gamma} g_{\gamma\beta}T^{\alpha\beta} =
\frac 1 2 g^{\gamma\delta}\left(\partial_{\alpha}g_{\mu\delta} + \partial_{\mu}g_{\alpha \delta} - \partial_{\delta}g_{\mu \alpha}\right) T_{\gamma}^{\alpha} =
\frac 1 2 \left(\partial_{\alpha}g_{\mu\delta} + \partial_{\mu}g_{\alpha \delta} - \partial_{\delta}g_{\mu \alpha}\right)
T^{\alpha\delta}
\end{equation}
We sum over $\alpha$ and $\delta$, but noting that we are antisymmetric in first and last terms in $\alpha$ and $\delta$ in the () but symmetric in $T_{\alpha\delta}$ so we have
\begin{equation}
\Gamma_{\mu\alpha}^{\gamma} g_{\gamma\beta}T^{\alpha\beta} = \frac 1 2 \partial_{\mu}g_{\alpha \delta} T^{\alpha\delta}
\end{equation}
Thus we have
\begin{equation}
\partial_0 T^{0}_{\mu} + \partial_i T^{i}_{\mu} = \frac 1 2 \partial_{\mu}g_{\alpha \delta} T^{\alpha\delta}
\end{equation}
The $\mu = (1,2,3)$, we almost get back the equations in the standard formulation
\begin{equation}
\partial_0 \rho h u^0 u_i + \partial_j T^j_i = \frac 1 2 \partial_{i}g_{\alpha \delta} T^{\alpha\delta},
\end{equation}
which modulo a factors of $\lapse\rtgamma$ in front is the same as the "standard" equations.
The $T^0_0$ term is more interesting. Here we have
\begin{equation}
\partial_0 (\rho h u^0 u_0 + + \partial_j T^j_i = \frac 1 2 \partial_{0}g_{\alpha \delta} T^{\alpha\delta},
\end{equation}
However the disadvantage is that we need the time derivative of the metric.
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#mapping): Primitive to Conservative Mapping
1. [Step 2](#zach): Compute $u^0$ from the Valencia 3-velocity (Zach step)
1. [Step 3](#flux): Compute the flux
1. [Step 4](#source): Source Terms
1. [Step 5](#rotation): Rotation
1. [Step 6](#solver): Conservative to Primitive Solver
1. [Step 7](#lorentz): Lorentz Boosts
1. [Step 8](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='mapping'></a>
# Step 1: Primitive to Conservative Mapping
$$\label{mapping}$$
We want to make a mapping from the primitives to conserved variables:
\begin{equation}
(\rho, \vel, \epsilon) \rightarrow (\rhostar = \lapse\rho\rtgamma u^0, \Svectilde = \rhostar h \uvec, \tautilde = \lapse^2\rtgamma \T00 - \rhostar).
\end{equation}
To do so, we first need to determine $u^0$ and $\uvec$. Noting that $g_{\mu\nu} u^{\mu} u^{\nu} = -1$, we find
\begin{equation}
u^0 = \left(-g_{00} - 2g_{i0} v^i - g_{ij}v^iv^j\right)^{-1/2},
\end{equation}
where we have used $\vel = \uvec/u^0$. This gives me $\rhostar$ and $\uvec$. We note that the metric is (B&S 2.122)
\begin{equation}
g_{\mu\nu} = \begin{pmatrix}
-\lapse^2 + \shift\cdot\shift & \beta_i \\
\beta_j & \gamma_{ij}
\end{pmatrix},
\end{equation}
Lets write some code to define metric contraction on four vectors in this context:
```
import NRPy_param_funcs as par
import indexedexp as ixp
import sympy as sp
from outputC import *
import NRPy_param_funcs as par
DIM = 3
# Declare rank-2 covariant gmunu
gmunuDD = ixp.declarerank2("gmunuDD","sym01",DIM=4)
gammaDD = ixp.declarerank2("gammaDD","sym01")
components = ["xx", "xy", "xz", "yy", "yz", "zz"]
names = ""
for comp in components :
names = names + "mi.gamDD{0} ".format(comp)
gxx, gxy, gxz, gyy, gyz, gzz = sp.symbols( names)
gammaDD[0][0] = gxx
gammaDD[0][1] = gxy
gammaDD[0][2] = gxz
gammaDD[1][0] = gxy
gammaDD[1][1] = gyy
gammaDD[1][2] = gyz
gammaDD[2][0] = gxz
gammaDD[2][1] = gyz
gammaDD[2][2] = gzz
lapse, rtgamma, beta_x, beta_y, beta_z = sp.symbols( "mi.alpha mi.rtDetGamma mi.betaX mi.betaY mi.betaZ")
u10, u1x, u1y, u1z = sp.symbols("u1[0] u1[1] u1[2] u1[3]")
u20, u2x, u2y, u2z = sp.symbols("u2[0] u2[1] u2[2] u2[3]")
u1U = ixp.declarerank1("u1Vector", DIM=4)
u2U = ixp.declarerank1("u2Vector", DIM=4)
u1U[0] = u10
u1U[1] = u1x
u1U[2] = u1y
u1U[3] = u1z
u2U[0] = u20
u2U[1] = u2x
u2U[2] = u2y
u2U[3] = u2z
shiftU = ixp.declarerank1("shiftU")
shiftU[0] = beta_x
shiftU[1] = beta_y
shiftU[2] = beta_z
beta2 = 0
for i in range(DIM) :
for j in range(DIM) :
beta2 += gammaDD[i][j] * shiftU[i]*shiftU[j]
gmunuDD[0][0] = -lapse*lapse + beta2
for i in range(DIM) :
gmunuDD[i+1][0] = shiftU[i]
gmunuDD[0][i+1] = shiftU[i]
for j in range(DIM) :
gmunuDD[i+1][j+1] = gammaDD[i][j]
dot4Product = 0
for i in range(4):
for j in range(4):
dot4Product += gmunuDD[i][j]*u1U[i]*u2U[j]
str = outputC( dot4Product, "dotProduct", filename="returnstring")
print(str)
```
which then gives
\begin{equation}
u^0 = \left(\lapse^2 - \shift\cdot\shift - 2\shift\cdot\vel - \gamma_{ij}v^iv^j\right)^{-1/2},
\end{equation}
The other thing is $\uvec = u^0\vel$. So then we can proceed and spit out the conservative variables: $\rhostar, \Svectilde$.
To get $\tau$, we note that we have defined the metric as the covariant form, e.g., lower indices. The upper form of $g^{\mu\nu}$ is found in [B&S]() 2.119 and is given by
\begin{equation}
g^{\mu\nu} =
\begin{pmatrix}
-\lapse^{-2} & \lapse^{-2}\beta^i \\
\lapse^{-2}\beta^j & \gamma^{ij} - \lapse^{-2} \beta^i\beta^j
\end{pmatrix}
\end{equation}
Lets get the form of this in code:
The main challenge is the calculation of the inverse of the 3x3 matrix $\gamma_{ij}$. To do so we note:
```
import indexedexp as ixp
gammaUU, gammabarDet = ixp.symm_matrix_inverter3x3(gammaDD)
gUUxx = gammaUU[0][0]
gUUxy = gammaUU[0][1]
gUUxz = gammaUU[0][2]
gUUyy = gammaUU[1][1]
gUUyz = gammaUU[1][2]
gUUzz = gammaUU[2][2]
rtDetGamma = sp.sqrt(gammabarDet)
outputC( [gUUxx,gUUxy,gUUxz,gUUyy,gUUyz,gUUzz, rtDetGamma], ["mi.gamUUxx", "mi.gamUUxy","mi.gamUUxz","mi.gamUUyy","mi.gamUUyz","mi.gamUUzz","mi.rtDetGamma"], filename="NRPY+gmunuUU_and_det.h")
#print str
```
<a id='zach'></a>
# Step 2: Compute $u^0$ from the Valencia 3-velocity (Zach step)
$$\label{zach}$$
According to Eqs. 9-11 of [the IllinoisGRMHD paper](https://arxiv.org/pdf/1501.07276.pdf), the Valencia 3-velocity $v^i_{(n)}$ is related to the 4-velocity $u^\mu$ via
\begin{align}
\alpha v^i_{(n)} &= \frac{u^i}{u^0} + \beta^i \\
\implies u^i &= u^0 \left(\alpha v^i_{(n)} - \beta^i\right)
\end{align}
Defining $v^i = \frac{u^i}{u^0}$, we get
$$v^i = \alpha v^i_{(n)} - \beta^i,$$
Or in other words in terms of the 3 velocity
$$v^i_{(n)} = \alpha^{-1}\left(v^i + \beta^i\right)$$
and in terms of this variable we get
\begin{align}
g_{00} \left(u^0\right)^2 + 2 g_{0i} u^0 u^i + g_{ij} u^i u^j &= \left(u^0\right)^2 \left(g_{00} + 2 g_{0i} v^i + g_{ij} v^i v^j\right)\\
\implies u^0 &= \pm \sqrt{\frac{-1}{g_{00} + 2 g_{0i} v^i + g_{ij} v^i v^j}} \\
&= \pm \sqrt{\frac{-1}{(-\alpha^2 + \beta^2) + 2 \beta_i v^i + \gamma_{ij} v^i v^j}} \\
&= \pm \sqrt{\frac{1}{\alpha^2 - \gamma_{ij}\left(\beta^i + v^i\right)\left(\beta^j + v^j\right)}}\\
&= \pm \sqrt{\frac{1}{\alpha^2 - \alpha^2 \gamma_{ij}v^i_{(n)}v^j_{(n)}}}\\
&= \pm \frac{1}{\alpha}\sqrt{\frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}}}
\end{align}
Generally speaking, numerical errors will occasionally drive expressions under the radical to either negative values or potentially enormous values (corresponding to enormous Lorentz factors). Thus a reliable approach for computing $u^0$ requires that we first rewrite the above expression in terms of the Lorentz factor squared: $\Gamma^2=\left(\alpha u^0\right)^2$:
\begin{align}
u^0 &= \pm \frac{1}{\alpha}\sqrt{\frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}}}\\
\implies \left(\alpha u^0\right)^2 &= \frac{1}{1 - \gamma_{ij}v^i_{(n)}v^j_{(n)}} \\
\implies \gamma_{ij}v^i_{(n)}v^j_{(n)} &= 1 - \frac{1}{\left(\alpha u^0\right)^2}
\end{align}
In order for the bottom expression to hold true, the left-hand side must be between 0 and 1. Again, this is not guaranteed due to the appearance of numerical errors. In fact, a robust algorithm will not allow $\Gamma^2$ to become too large (which might contribute greatly to the stress-energy of a given gridpoint), so let's define $\Gamma_{\rm max}$, the largest allowed Lorentz factor.
Then our algorithm for computing $u^0$ is as follows:
If
$$R=\gamma_{ij}v^i_{(n)}v^j_{(n)}>1 - \frac{1}{\Gamma_{\rm max}},$$
then adjust the 3-velocity $v^i$ as follows:
$$v^i_{(n)} = \sqrt{\frac{1 - \frac{1}{\Gamma_{\rm max}}}{R}}v^i_{(n)}.$$
After this rescaling, we are then guaranteed that if $R$ is recomputed, it will be set to its ceiling value $R=1 - \frac{1}{\Gamma_{\rm max}}$.
Then $u^0$ can be safely computed via
$$
u^0 = \frac{1}{\alpha \sqrt{1-R}}.
$$
```
import sympy as sp
import NRPy_param_funcs as par
import grid as gri
import indexedexp as ixp
import reference_metric as rfm
from outputC import *
vx, vy, vz = sp.symbols( "vx vy vz")
vU = ixp.declarerank1("vU")
vU[0] = vx
vU[1] = vy
vU[2] = vz
ValenciavU = ixp.register_gridfunctions_for_single_rank1("AUX","ValenciavU",DIM=3)
ValenciavU[0] = (vx + beta_x)/lapse
ValenciavU[1] = (vy + beta_y)/lapse
ValenciavU[2] = (vz + beta_z)/lapse
# Step 1: Compute R = 1 - 1/max(Gamma)
R = sp.sympify(0)
for i in range(DIM):
for j in range(DIM):
R += gammaDD[i][j]*ValenciavU[i]*ValenciavU[j]
GAMMA_SPEED_LIMIT = par.Cparameters("REAL","GRMHD_equations","GAMMA_SPEED_LIMIT", 10.0) # 10.0 is default for IllinoisGRMHD
Rmax = 1 - 1/(GAMMA_SPEED_LIMIT*GAMMA_SPEED_LIMIT)
rescaledValenciavU = ixp.zerorank1()
for i in range(DIM):
rescaledValenciavU[i] = ValenciavU[i]*sp.sqrt(Rmax/R)
rescaledu0 = 1/(lapse*sp.sqrt(1-Rmax))
regularu0 = 1/(lapse*sp.sqrt(1-R))
computeu0_Cfunction = "/* Function for computing u^0 from Valencia 3-velocity. */\n"
computeu0_Cfunction += "/* Inputs: vx, vy, vz, lapse, MetricInformation, GAMMA_SPEED_LIMIT (C parameter) */\n"
computeu0_Cfunction += "/* Output: u0=u^0 */\n\n"
computeu0_Cfunction += outputC([R,Rmax],["const double R","const double Rmax"],"returnstring",
params="includebraces=False,CSE_varprefix=tmpR,outCverbose=False")
computeu0_Cfunction += "if(R <= Rmax) "
computeu0_Cfunction += outputC(regularu0,"u0","returnstring",
params="includebraces=True,CSE_varprefix=tmpnorescale,outCverbose=False")
computeu0_Cfunction += " else "
computeu0_Cfunction += outputC([rescaledu0],
["u0"],"returnstring",
params="includebraces=True,CSE_varprefix=tmprescale,outCverbose=False")
print(computeu0_Cfunction)
```
We now note that $\tau = \lapse^2\rtgamma T^{00} - \rhostar$, which gives
\begin{equation}
\tau = \lapse\rhostar h u^0 - P\rtgamma - \rhostar
\end{equation}
The code for this is
```
rho, epsilon, gamma1, p = sp.symbols("rho ie gamma p")
betaDotV = 0
for i in range(DIM) :
for j in range(DIM) :
betaDotV += gammaDD[i][j] * shiftU[i]*vU[j]
v2 = 0
for i in range(DIM) :
for j in range(DIM) :
v2 += gammaDD[i][j] * vU[i]*vU[j]
u0 = sp.symbols("u0")
uvec4U = ixp.zerorank1(DIM=4)
uvec4D = ixp.zerorank1(DIM=4)
#StildeU = ixp.declarerank1("StildeU")
StildeD = ixp.zerorank1()
rhostar = lapse*rtgamma*rho*u0
h = 1. + epsilon + p/rho
for i in range(1,4):
uvec4U[i] = vU[i-1]*u0
uvec4U[0] = u0
for mu in range(4) :
for nu in range(4) :
uvec4D[mu] += gmunuDD[mu][nu]*uvec4U[nu]
for i in range(DIM):
StildeD[i] = uvec4D[i+1]*rhostar*h
tau = lapse*rhostar*h*u0 - rtgamma*p - rhostar
cFunction = "double u0 = 0.;\n" + computeu0_Cfunction
str = outputC([rhostar, StildeD[0], StildeD[1], StildeD[2], tau], ["con[iRhoStar]", "con[iSx]", "con[iSy]", "con[iSz]", "con[iTau]"], filename="returnstring")
print(str)
f = open("NRPY+prim2Con.h", "w")
f.write( cFunction + str)
f.close()
```
<a id='flux'></a>
# Step 3: Compute the flux
$$\label{flux}$$
The fluxes are as follows
\begin{equation}
\frac{\partial}{\partial t}
\begin{pmatrix}
\rhostar\\
\Svectilde\\
\tautilde
\end{pmatrix} + \frac{\partial}{\partial x^j}\begin{pmatrix} \rhostar v^j\\
\lapse\rtgamma T^j_i\\ \lapse^2\rtgamma T^{0j} - \rhostar v^j
\end{pmatrix} = \begin{pmatrix} 0 \\ \frac 1 2 \lapse\rtgamma T^{\alpha\beta}g_{\alpha\beta,i} \\ s \end{pmatrix}
\end{equation}
so the flux is
\begin{equation}
\mathcal{F} = \begin{pmatrix} \rhostar v^i \\ \lapse\rtgamma T^i_k \\ \lapse^2\rtgamma T^{0i} - \rhostar v^i
\end{pmatrix}
\end{equation}
In the moving-mesh formalism, the flux is just taken along the x directions so we have
\begin{equation}
\mathcal{F} = \begin{pmatrix} \rhostar v^1 \\ \lapse\rtgamma T^1_k \\ \lapse^2\rtgamma T^{01} - \rhostar v^1
\end{pmatrix}
\end{equation}
Note that we will need to rotate $T^{\mu\nu}$ and $g_{\mu\nu}$ to get the right orientation.
In order to do this, we must first compute the stress energy tensor:
\begin{equation}
T^{\mu\nu} = \rho h u^{\mu}u^{\nu} + Pg^{\mu\nu} = \rho h (u^0)^2v^iv^j + P g^{\mu\nu}
\end{equation}
```
TmunuUU = ixp.declarerank2("TmunuUU","sym01",DIM=4)
uvecU = ixp.zerorank1()
for i in range(3) :
uvecU[i] = uvec4U[i+1]
TmunuUU[0][0] = rho*h*u0*u0 - p/(lapse*lapse) #is this \pm?
for i in range(3):
TmunuUU[0][i+1] = rho*h*u0*uvecU[i] + p/(lapse*lapse)*shiftU[i]
TmunuUU[i+1][0] = rho*h*u0*uvecU[i] + p/(lapse*lapse)*shiftU[i]
for i in range(3):
for j in range(3):
TmunuUU[i+1][j+1] = rho*h*uvecU[i]*uvecU[j] + p*(gammaUU[i][j] - 1./(lapse*lapse)*shiftU[i]*shiftU[j])
#str = outputC([TmunuUU[1][0], TmunuUU[1][1], TmunuUU[1][2], TmunuUU[1][3]], ["Tmunu10", "Tmunu11", "Tmunu12", "Tmunu13"], filename="returnstring")
#print(str)
#str = outputC([gmunuDD[1][0], gmunuDD[1][1], gmunuDD[1][2], gmunuDD[1][3]], ["gmunu10", "gmunu11", "gmunu12", "gmunu13"], filename="returnstring")
#print(str)
#calculate Tmunu^1_i
Tmunu1D = ixp.zerorank1()
for i in range(3):
for j in range(0,4) :
Tmunu1D[i] += gmunuDD[i+1][j] * TmunuUU[1][j]
#str = outputC([Tmunu1D[0], Tmunu1D[1], Tmunu1D[2]], ["Tmunu1Dx", "Tmunu1Dy", "Tmunu1Dz"], filename="returnstring")
#print str
# now get the flux
fluxRho, fluxMomX, fluxMomY, fluxMomZ, fluxEnergy = sp.symbols("flux[iRhoStar] flux[iSx] flux[iSy] flux[iSz] flux[iTau]")
fluxRho = rhostar * vU[0]
fluxMomX = lapse*rtgamma*Tmunu1D[0]
fluxMomY = lapse*rtgamma*Tmunu1D[1]
fluxMomZ = lapse*rtgamma*Tmunu1D[2]
fluxEnergy = lapse*lapse*rtgamma*TmunuUU[0][1] - rhostar*vU[0]
cFunction = "double u0 = 0.;\n" + computeu0_Cfunction
str = outputC([fluxRho, fluxMomX, fluxMomY, fluxMomZ, fluxEnergy], ["flux[iRhoStar]", "flux[iSx]", "flux[iSy]", "flux[iSz]", "flux[iTau]"], filename="returnstring")
print(str)
f = open("NRPY+calFlux.h", "w")
f.write( cFunction + str)
f.close()
```
<a id='source'></a>
# Step 4: Source Terms
$$\label{source}$$
The sources terms are for mass, momentum and energy are:
\begin{equation}
\source = (0, \frac 1 2 \lapse\rtgamma \T{\alpha}{\beta}g_{\alpha\beta,i}, s),
\end{equation}
For a time stationary metric $s\neq 0$, so we will ignore this until the next section. As for the rest, we need to define derivatives of the metric. Suppose I have done this already. Then the code for the source terms is:
```
gmunuDDind = [0,0]
gammaDDind = [0,0]
lapseind = [0,0]
alpha = [0.,0.]
h = sp.symbols( "h")
for ind in range(2) :
gmunuDDind[ind] = ixp.zerorank2(DIM=4) #derivative of gmunu in some direction
gammaDDind[ind] = ixp.zerorank2()
components = ["xx", "xy", "xz", "yy", "yz", "zz"]
names = ""
for comp in components :
names = names + "mi{1}.gamDD{0} ".format(comp, ind+1)
gxx, gxy, gxz, gyy, gyz, gzz = sp.symbols( names)
gammaDDind[ind][0][0] = gxx
gammaDDind[ind][0][1] = gxy
gammaDDind[ind][0][2] = gxz
gammaDDind[ind][1][0] = gxy
gammaDDind[ind][1][1] = gyy
gammaDDind[ind][1][2] = gyz
gammaDDind[ind][2][0] = gxz
gammaDDind[ind][2][1] = gyz
gammaDDind[ind][2][2] = gzz
lapse, rtgamma, beta_x, beta_y, beta_z = sp.symbols( "mi{0}.alpha mi{0}.rtDetGamma mi{0}.betaX mi{0}.betaY mi{0}.betaZ".format(ind+1))
u10, u1x, u1y, u1z = sp.symbols("u1[0] u1[1] u1[2] u1[3]")
u20, u2x, u2y, u2z = sp.symbols("u2[0] u2[1] u2[2] u2[3]")
lapseind[ind] = lapse
shiftU = ixp.zerorank1()
shiftU[0] = beta_x
shiftU[1] = beta_y
shiftU[2] = beta_z
beta2 = 0
for i in range(DIM) :
for j in range(DIM) :
beta2 += gammaDDind[ind][i][j] * shiftU[i]*shiftU[j]
gmunuDDind[ind][0][0] = -lapse*lapse + beta2
for i in range(DIM) :
gmunuDDind[ind][i+1][0] = shiftU[i]
gmunuDDind[ind][0][i+1] = shiftU[i]
for j in range(DIM) :
gmunuDDind[ind][i+1][j+1] = gammaDDind[ind][i][j]
dgmunuDD = ixp.zerorank2(DIM=4)
source = 0
for mu in range(4) :
for nu in range(4) :
dgmunuDD[mu][nu] = (gmunuDDind[1][mu][nu] - gmunuDDind[0][mu][nu])/(2*h) # /(2*h) = 1.0/(2.0*h); for the finite differene
source = source + TmunuUU[mu][nu]*dgmunuDD[mu][nu]
# precalculate dalpha/di for next calculation
dalpha = (lapseind[1] - lapseind[0])/(2*h)
#print TmunuUU[2][1]
cFunction = "double u0 = 0.;\n" + computeu0_Cfunction
str = outputC( [source, dalpha, gmunuDDind[1][0][0]], ["source", "dlapse", "dgmunu"], filename="returnstring")
print(str)
f = open("NRPY+calMomSources.h", "w")
f.write( cFunction + str)
f.close()
```
So now we need to include source term for the energy. It is
\begin{equation}
s = \lapse\rtgamma\left[\left(\T00\beta^i\beta^j + 2\T0i\beta^j\right)K_{ij} - \left(\T00\beta^i + \T0i\right)\partial_i\lapse\right]
\end{equation}
For a stationary metric. $\beta^i = 0$, so all the terms go away except for the final term. So we have
\begin{equation}
s = -\lapse\rtgamma\T0i\partial_i\lapse
\end{equation}
```
dalpha = ixp.zerorank1()
for i in range(3) :
dalpha[i] = sp.symbols("dalpha[{0}]".format(i))
esource = 0
for j in range(3) :
esource += TmunuUU[0][j+1]*dalpha[j]
cFunction = "double u0 = 0.;\n" + computeu0_Cfunction
str = outputC( [esource], ["eSource"], filename="returnstring")
f = open("NRPY+calTauSource.h", "w")
f.write( cFunction + str)
f.close()
```
<a id='rotation'></a>
# Step 5: Rotation
$$\label{rotation}$$
One of the key ideas behind the moving-mesh idea is that we must rotate the vector to the appropriate direction such that the Riemann solve is along the x-direction. This is done by computing the normal vector also the "x-direction" and two normal vectors orthogonal to it. In MANGA they are labeled $n_0$, $n_1$, and $n_2$. And the rotation matrix looks like
\begin{equation}
R = \begin{pmatrix}
n_{0,x} & n_{0,y} & n_{0,z} \\
n_{1,x} & n_{1,y} & n_{1,z} \\
n_{2,x} & n_{2,y} & n_{2,z}
\end{pmatrix}
\end{equation}
Likewise, I also define an inverse Rotation as
\begin{equation}
R^{-1} = \begin{pmatrix}
n'_{0,x} & n'_{0,y} & n'_{0,z} \\
n'_{1,x} & n'_{1,y} & n'_{1,z} \\
n'_{2,x} & n'_{2,y} & n'_{2,z}
\end{pmatrix}
\end{equation}
Base on this how does $g_{\mu\nu}$ and $T_{\mu\nu}$ transform under rotation. We begin by defining an extended rotation matrix:
\begin{equation}
\mathcal{R} = \begin{pmatrix}
1 &0 &0 &0 \\
0 &n_{0,x} & n_{0,y} & n_{0,z} \\
0 &n_{1,x} & n_{1,y} & n_{1,z} \\
0 & n_{2,x} & n_{2,y} & n_{2,z}
\end{pmatrix}
\end{equation}
Now we note that the term $g_{\mu\nu} x^{\mu} x^{\nu}$ is invariant under rotation. Defining $x' = R x$, we then note
\begin{equation}
g_{\mu\nu} x^{\mu} x^{\nu} = g_{\mu\nu} \mathcal{R}^{-1\mu}_{\alpha}x'^{\alpha} \mathcal{R}^{-1\nu}_{\beta}x'^{\beta} = g_{\alpha\beta} \mathcal{R}^{-1\alpha}_{\mu}\mathcal{R}^{-1\beta}_{\nu}x'^{\mu} x'^{\nu} \rightarrow g'_{\mu\nu} = g_{\alpha\beta} \mathcal{R}^{-1\alpha}_{\mu}\mathcal{R}^{-1\beta}_{\nu},
\end{equation}
which gives us the appropriate rotated metric.
To get the similar transformation for $T^{\mu\nu}$, we note the $T = T^{\mu}_{\mu} = g_{\mu\nu}T^{\mu\nu}$ tranforms as a scalar. So we have
\begin{equation}
T = g_{\mu\nu}T^{\mu\nu} = g_{\alpha\beta}\mathcal{R}^{-1,\alpha}_{\gamma}\mathcal{R}^{-1,\beta}_{\delta}\mathcal{R}^{\gamma}_{\mu}\mathcal{R}^{\delta}_{\nu} T^{\mu\nu} = \rightarrow g'_{\mu\nu} T'^{\mu\nu}
\end{equation}
which provide the identity
\begin{equation}
T'^{\mu\nu} = \mathcal{R}^{\mu}_{\alpha}\mathcal{R}^{\nu}_{\beta} T^{\alpha\beta}
\end{equation}
```
RotUD = ixp.declarerank2("RotUD", "nosym", DIM=4)
RotInvUD = ixp.declarerank2("RotInvUD", "nosym", DIM=4)
#declare normal primed vectors for rotation
n00, n01, n02 = sp.symbols("n0[0] n0[1] n0[2]")
n10, n11, n12 = sp.symbols("n1[0] n1[1] n1[2]")
n20, n21, n22 = sp.symbols("n2[0] n2[1] n2[2]")
#declare normal primed vectors for inverse rotation
n0p0, n0p1, n0p2 = sp.symbols("n0p[0] n0p[1] n0p[2]")
n1p0, n1p1, n1p2 = sp.symbols("n1p[0] n1p[1] n1p[2]")
n2p0, n2p1, n2p2 = sp.symbols("n2p[0] n2p[1] n2p[2]")
for i in range(4):
RotUD[0][i] = 0.
RotUD[i][0] = 0.
RotInvUD[0][i] = 0.
RotInvUD[i][0] = 0.
RotUD[0][0] = 1.
RotInvUD[0][0] = 1.
RotUD[1][1] = n00
RotUD[1][2] = n01
RotUD[1][3] = n02
RotUD[2][1] = n10
RotUD[2][2] = n11
RotUD[2][3] = n12
RotUD[3][1] = n20
RotUD[3][2] = n21
RotUD[3][3] = n22
RotInvUD[1][1] = n0p0
RotInvUD[1][2] = n0p1
RotInvUD[1][3] = n0p2
RotInvUD[2][1] = n1p0
RotInvUD[2][2] = n1p1
RotInvUD[2][3] = n1p2
RotInvUD[3][1] = n2p0
RotInvUD[3][2] = n2p1
RotInvUD[3][3] = n2p2
gmunuRotDD = ixp.declarerank2("gmunuRotDD", "sym01", DIM=4)
for i in range(4) :
for j in range(4) :
gmunuRotDD[i][j] = 0.
for k in range(4) :
for l in range(4) :
gmunuRotDD[i][j] += gmunuDD[l][k]*RotInvUD[l][i]*RotInvUD[k][j]
outputC([gmunuRotDD[1][1], gmunuRotDD[1][2], gmunuRotDD[1][3],gmunuRotDD[2][2], gmunuRotDD[2][3], gmunuRotDD[3][3]], ["metricInfo.gamDDxx", "metricInfo.gamDDxy","metricInfo.gamDDxz","metricInfo.gamDDyy","metricInfo.gamDDyz","metricInfo.gamDDzz"], filename="NRPY+rotateMetric.h")
# provie
TmunuRotUU = ixp.declarerank2("TmunuRotUU", "sym01", DIM=4)
for i in range(4) :
for j in range(4) :
TmunuRotUU[i][j] = 0.
for k in range(4) :
for l in range(4) :
TmunuRotUU[i][j] += TmunuUU[l][k]*RotUD[i][l]*RotUD[j][k]
str = outputC([TmunuRotUU[0][0], TmunuRotUU[0][1], TmunuRotUU[1][0]], ["Tmunu00", "Tmunu12", "Tmunu21"], filename="returnstring")
print(str)
```
<a id='solver'></a>
# Step 6: Conservative to Primitive Solver
$$\label{solver}$$
We now discuss the reverse mapping from conservative to primitive variables.
Given the lapse, shift vector and $\rtgamma$, the mapping between primitive and conserved variable is straightforward. However, the reverse is not as simple. In GRMHD, the conservative to primitive solver is amplified by the inclusion of the magnetic field, leading to rather sophisticated root finding strategies. The failure rates of these algorithms are low (??), but since this algorithm may be executed several times per timestep for every gridpoint, even a low failure can give unacceptable collective failure rates. However, for purely polytropic equations of state, e.g., $P\propto\rho^{\Gamma_1}$, the convervative to primitive variable solver is greatly simplified.
To construct the conservative-to-primitive variable solver, we restrict ourselves to polytropic equations of states
\begin{equation}
P = P_0\left(\frac{\rho}{\rho_0}\right)^{\Gamma_1} \quad\textrm{and}\quad \epsilon = \epsilon_0\left(\frac{\rho}{\rho_0}\right)^{\Gamma_1-1},
\end{equation}
where $P_0$, $\rho_0$, and $\epsilon_0$ are the fiducial pressure, density, and internal energy, and we have used the relation $P = (\Gamma_1 - 1)\rho\epsilon$.
For such a polytropic equation of state, the energy equation is redundant and effectively we are only concerned with the continuity and momentum equations. The conservative variables of concern are $\rhostar$ and $\Svectilde$. Noting that the shift, $\alpha$, and $\rtgamma$ are provided by the Einsteins field equation solver, we can write
\begin{equation}
u^0 = \frac{\rhostar}{\alpha\rtgamma\rho} = u^0(\rho) \quad\textrm{and}\quad \uvec = \frac{\Svectilde}{\alpha\rtgamma\rho h} = \uvec(\rho).
\end{equation}
Noting that the four velocity $u^2 = g_{\mu\nu}u^{\mu}u^{\nu} = g^{00}u^0u^0 + 2g^{0i}u^0\uvec^i + g_{ij}\uvec^i\uvec^j = -1$, we have
\begin{equation}
0 = f(\rho)\equiv \alpha^2\gamma\rho^2h^2 + \left(-\lapse^2 + \shift\cdot\shift\right)\rhostar^2h^2 + 2h\rhostar\shift\cdot\Svectilde + \Svectilde\cdot\Svectilde,
\end{equation}
which is an implicit equation of either $\rho$ or $u^0$, where $h(\rho = \rhostar/(\alpha\rtgamma u^0)) = 1 + \gamma_1 \epsilon$ which can be inverted by standard nonlinear root finding algorithms, e.g., Newton-raphson.
We put this all together to define a function, $f(\rho)$, whose root is zero that we will find via Newton-raphson.
Several checks must be performed:
1. $\rhostar > 0$ : This check is performed at the very beginning
2. $\rho > \rho_{\rm min}$ : This check is performed after the fact
3. $u_0 < \alpha^{-1}\Gamma_{\rm max}$ : This check is performed after the fact as well
```
DIM = 3
# Declare rank-1 contravariant ("v") vector
vU = ixp.declarerank1("vU")
shiftU = ixp.zerorank1()
rho, gamma1 = sp.symbols("rho gamma")
Sx, Sy, Sz = sp.symbols("con[iSx] con[iSy] con[iSz]")
p0, rho0, rhostar = sp.symbols("p_0 rho_0 rhostar")
# Declare rank-2 covariant gmunu
#gammaDD = ixp.declarerank2("gammaDD","sym01")
StildeD = ixp.declarerank1("StildeD")
lapse, rtgamma, beta_x, beta_y, beta_z = sp.symbols( "mi.alpha mi.rtDetGamma mi.betaX mi.betaY mi.betaZ")
shiftU[0] = beta_x
shiftU[1] = beta_y
shiftU[2] = beta_z
StildeD[0] = Sx
StildeD[1] = Sy
StildeD[2] = Sz
gamma = rtgamma*rtgamma
lapse2 = lapse*lapse
uU0 = rhostar/(lapse*rtgamma*rho)
epsilon = p0/rho0*(rho/rho0)**(gamma1 - 1)/(gamma1 - 1)
h = 1. + gamma1*epsilon
beta2 = 0.
for i in range(DIM) :
for j in range(DIM) :
beta2 += gammaDD[i][j] * shiftU[i]*shiftU[j]
betaDotStilde = 0
for i in range(DIM) :
betaDotStilde += shiftU[i]*StildeD[i]
Stilde2 = 0
for i in range(DIM) :
for j in range(DIM) :
Stilde2 += gammaUU[i][j] * StildeD[i]*StildeD[j]
f = rhostar**2*h**2 + (-lapse2 + beta2)*rhostar**2.*h**2.*uU0**2 + 2.*h*rhostar*betaDotStilde*uU0 + Stilde2
outputC(f,"rootRho",filename="NRPY+rhoRoot.h")
outputC(Stilde2, "Stilde2", filename="NRPY+Stilde2.h")
```
The root solve above finds $\rho$, which then allows us to get
\begin{equation}
u^0 = \frac{\rhostar}{\alpha\rtgamma\rho}\quad\textrm{and}\quad \vel = \frac{\uvec}{u^0} = \frac{\Svectilde}{\rhostar h(\rho)}.
\end{equation}
and thus we can find the rest of the primitives.
```
#rhostar = sp.symbols("rhostar")
#StildeU = ixp.declarerank1("StildeU")
velU = ixp.zerorank1()
#lapse, rtgamma, rho, gamma1, c = sp.symbols("lapse rtgamma rho gamma1 c")
rho, rhostar = sp.symbols("testPrim[iRho] con[iRhoStar]")
u0 = rhostar/(lapse*rtgamma*rho)
epsilon = p0/rho0*(rho/rho0)**(gamma1 - 1)/(gamma1 - 1)
h = 1. + gamma1*epsilon
for i in range(DIM) :
for j in range(DIM) :
velU[i] += gammaUU[i][j]*StildeD[j]/(rhostar * h)/u0
outputC([h,u0,velU[0],velU[1],velU[2]], ["h", "u0","testPrim[ivx]", "testPrim[ivy]", "testPrim[ivz]"],filename="NRPY+getv.h")
```
<a id='lorentz'></a>
# Step 7: Lorentz Boosts
$$\label{lorentz}$$
We need to boost to the frame of the moving face. The boost is
\begin{equation}
B(\beta) =\begin{pmatrix}
\gamma & -\beta\gamma n_x & -\beta\gamma n_y & -\beta\gamma n_z \\
-\beta\gamma n_x & 1 + (\gamma-1)n_x^2 & (\gamma-1)n_x n_y & (\gamma-1)n_x n_z\\
-\beta\gamma n_x & (\gamma-1)n_y n_x & 1 + (\gamma-1)n_y^2 & (\gamma-1)n_y n_z\\
-\beta\gamma n_x & (\gamma-1) n_z n_x & (\gamma-1)n_z n_x & 1 + (\gamma-1)n_z^2
\end{pmatrix}
\end{equation}
And the boost is $X' = B(\beta) X$, where $X'$ and $X$ are four vectors.
So the rest of this is straightforward.
<a id='latex_pdf_output'></a>
# Step 8: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-GRMHD_Equations-Cartesian.pdf](Tutorial-GRMHD_Equations-Cartesian.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
!jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-GRMHD_Equations-Cartesian.ipynb
!pdflatex -interaction=batchmode Tutorial-GRMHD_Equations-Cartesian.tex
!pdflatex -interaction=batchmode Tutorial-GRMHD_Equations-Cartesian.tex
!pdflatex -interaction=batchmode Tutorial-GRMHD_Equations-Cartesian.tex
!rm -f Tut*.out Tut*.aux Tut*.log
```
| github_jupyter |
```
n=int(input("请输入一个数\n"))
sum=1
i=1
for i in range(n):
sum*=(i+1)
print(sum)
```
### *如何无限循环*
```
sum=0
m=int(input("请输入一个整数,输入0表示结束\n"))
for i in range(1000):
sum=sum+m
m=int(input("请输入一个整数,输入0表示结束\n"))
if(m==0):
break
print(sum)
m=int(input("请输入一个整数,以回车结束"))
sum=m
mul=m
for i in range(1000):
if(m<=sum and mul>=m**2):
sum=sum+m
mul=mul*m
break
m=int(input("请输入一个整数,以回车结束\n"))
print("积为",mul)
m=int(input("Please enter the rows of blank places"))
i=0
for i in range(m):
print()
print("end")
m=int(input('请输入要输入的整数个数,回车结束。'))
max1=int(input('请输入一个整数,回车结束'))
max2=int(input('请输入一个整数,回车结束'))
if max2>max1:
temp=max1
max1=max2
max2=temp
for i in range(m-2):
n=int(input('请输入一个整数,回车结束'))
if n>max2 and n<max1:
max2=n
elif n>max1:
temp=max2
max2=max1
max1=n
print("第二大的数为",max2)
def fact (end):
j=1
for num in range(end):
j=(num+1)*j
return j
n = int(input('请输入第1个整数,以回车结束。'))
m = int(input('请输入第2个整数,以回车结束。'))
k = int(input('请输入第3个整数,以回车结束。'))
print('最终的和是:', fact(m) + fact(n) + fact(k))
def fun (end):
i=1
j=1
m=1
sum=0
for p in range(end):
m=j/i
i=i+2
j=-1*j
sum=sum+m
return sum
n=int(input("请输入一个整数"))
print("最终的和为:",4*fun(n))
n=1000
print("最终的和为:",4*fun(n))
n=100000
print("最终的和为:",4*fun(n))
def total (m,n,k):
sum=0
for i in range(m,n,k):
sum=sum+i
return sum
a=int(input("plz enter an positive interger\n"))
b=int(input("plz enter an positive interger and bigger than a\n"))
c=int(input("plz enter an positive interger\n"))
print("The sum is",total (a,b,c))
import math,random
m=int(input("请输入一个整数作为上界\n"))
k=int(input("请输入一个整数作为下界\n"))
n=int(input("请输入你要随机生成的整数的个数\n"))
def fun ():
i=0
total=0
for i in range (n):
num=random.randint(k,m)
print("第",i+1,"次随机生成的数为:",num)
total=total+num
aver=total/n
root=math.sqrt(aver)
print(n,"个平均数的平方根为",root)
fun()
import math,random
n=int(input("请输入你要随机生成的整数的个数\n"))
max=int(input("请输入一个整数作为上界\n"))
min=int(input("请输入一个整数作为下界\n"))
i=0
sum1=0
sum2=0
for i in range(n):
num=random.randint(min,max)
print("第",i+1,"次随机生成的数为",num)
a=math.log10(num)
sum1=sum1+a
sum2=sum2+1/a
i+=1
print("∑log(random,2)=",sum1,"∑1/log(random,2)=",sum2)
import math,random
n=int(input("请输入你要相加的数字的个数\n"))
a=random.randint(1,9)
print("本次随机生成的数为:",a)
b=math.pow(10,0)*a
i=0
sum=0
for i in range(n):
sum=sum+b
print(b)
b=math.pow(10,i+1)*a+b
print(n,"个数相加的和为",sum)
def total(m):
sum=0
numbers=[]
for i in range(m):
n=int(input("请输入一个整数:"))
numbers.append(n)
for i in range(m):
sum=sum+numbers[i]
return(sum)
m = int(input('请输入一个整数,表示将要输入的数字个数,回车结束。\n'))
print ("和为",total(m))
def Min(a):
n=[]
for i in range(a):
num=int(input("请输入一个整数"))
n.append(num)
min=n[i]
for i in range(a):
if n[i]<min:
min=n[i]
print(n)
return(min)
a=int(input("请输入将要输入的数字的个数\n"))
print("最小数为",Min(a))
def find(m):
n=[1,2,5,7,8,9,10,15,18]
for i in range(len(n)):
if m!=n[i]:
continue
else: return(i+1)
if(i==len(n)-1):
if(m!=n[i]):
return(-1)
m=int(input("请输入你要寻找的整数"))
a=find(m)
if(a!=-1):
print(m,"的位置是","第",a,"个")
else: print("查无此数")
import math
def cos(d):
product=0
len1=len2=0
Vector=[]
for i in range(2*d):
m=int(input("Plz enter the number:"))
Vector.append(m)
for i in range(d):
product=Vector[i]*Vector[i+d]+product
for i in range(d):
len1=Vector[i]**2+len1
len2=Vector[i+d]**2+len2
cos=product/(math.sqrt(len1)*math.sqrt(len2))
return(cos)
d=int(input("Plz enter the deminsion of the vector:"))
print(cos(d))
num=0
for i in range (5,90):
for j in range (5,90):
for p in range (5,90):
if(i+j+p==100):
##print("分别分给三个班级",i,j,p,"个")
num+=1
print("共",num,"种分法",sep=' ')
```
| github_jupyter |
## Outline
1. Loading datasets - Transforming images
2. VGG-16 with modification to network head
3. Using pre-trained models
4. Storing intermediate models
5. Resnet
6. Inception v3
7. Exercises
```
import matplotlib.pyplot as plt
import numpy as np
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.optim as optim
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
```
## Dataset, transforms, and visualisation
```
transform_train = transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
])
transform_test = transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True,
transform=transform_train)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True,
transform=transform_test)
num_classes = 10
batch_size = 4
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True)
dataiter = iter(trainloader)
images, labels = dataiter.next()
print(images.shape)
print(images[1].shape)
print(labels[1].item())
def imshow(img, title):
npimg = img.numpy() / 2 + 0.5
plt.figure(figsize=(batch_size, 1))
plt.axis('off')
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.title(title)
plt.show()
def show_batch_images(dataloader):
images, labels = next(iter(dataloader))
img = torchvision.utils.make_grid(images)
imshow(img, title=[str(x.item()) for x in labels])
for i in range(4):
show_batch_images(trainloader)
```
## Creating VGG-16
https://pytorch.org/docs/master/_modules/torchvision/models/vgg.html
```
from torchvision import models
vgg = models.vgg16_bn()
print(vgg)
print(vgg.features[0])
print(vgg.classifier[6])
final_in_features = vgg.classifier[6].in_features
mod_classifier = list(vgg.classifier.children())[:-1]
mod_classifier.extend([nn.Linear(final_in_features, num_classes)])
print(mod_classifier)
vgg.classifier = nn.Sequential(*mod_classifier)
print(vgg)
```
### Train CIFAR10
```
batch_size = 16
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size, shuffle=False)
def evaluation(dataloader, model):
total, correct = 0, 0
for data in dataloader:
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
_, pred = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (pred == labels).sum().item()
return 100 * correct / total
vgg = vgg.to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.SGD(vgg.parameters(), lr=0.05)
loss_epoch_arr = []
max_epochs = 1
n_iters = np.ceil(50000/batch_size)
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs = vgg(inputs)
loss = loss_fn(outputs, labels)
loss.backward()
opt.step()
del inputs, labels, outputs
torch.cuda.empty_cache()
if i % 100 == 0:
print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
loss_epoch_arr.append(loss.item())
print('Epoch: %d/%d, Test acc: %0.2f, Train acc: %0.2f' % (
epoch, max_epochs,
evaluation(testloader, vgg), evaluation(trainloader, vgg)))
plt.plot(loss_epoch_arr)
plt.show()
```
### Freeze layers of Convolutional Operations
```
batch_size = 16
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size, shuffle=False)
vgg = models.vgg16_bn(pretrained=True)
for param in vgg.parameters():
param.requires_grad = False
final_in_features = vgg.classifier[6].in_features
vgg.classifier[6] = nn.Linear(final_in_features, num_classes)
for param in vgg.parameters():
if param.requires_grad:
print(param.shape)
vgg = vgg.to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.SGD(vgg.parameters(), lr=0.05)
loss_epoch_arr = []
max_epochs = 1
n_iters = np.ceil(50000/batch_size)
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs = vgg(inputs)
loss = loss_fn(outputs, labels)
loss.backward()
opt.step()
if i % 100 == 0:
print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
del inputs, labels, outputs
torch.cuda.empty_cache()
loss_epoch_arr.append(loss.item())
print('Epoch: %d/%d, Test acc: %0.2f, Train acc: %0.2f' % (
epoch, max_epochs,
evaluation(testloader, vgg), evaluation(trainloader, vgg)))
plt.plot(loss_epoch_arr)
plt.show()
```
### With model copies
```
import copy
loss_epoch_arr = []
max_epochs = 1
min_loss = 1000
n_iters = np.ceil(50000/batch_size)
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs = vgg(inputs)
loss = loss_fn(outputs, labels)
loss.backward()
opt.step()
if min_loss > loss.item():
min_loss = loss.item()
best_model = copy.deepcopy(vgg.state_dict())
print('Min loss %0.2f' % min_loss)
if i % 100 == 0:
print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
del inputs, labels, outputs
torch.cuda.empty_cache()
loss_epoch_arr.append(loss.item())
vgg.load_state_dict(best_model)
print(evaluation(trainloader, vgg), evaluation(testloader, vgg))
```
## ResNet Model
https://pytorch.org/docs/master/_modules/torchvision/models/resnet.html
```
resnet = models.resnet18(pretrained=True)
print(resnet)
for param in resnet.parameters():
param.requires_grad = False
in_features = resnet.fc.in_features
resnet.fc = nn.Linear(in_features, num_classes)
for param in resnet.parameters():
if param.requires_grad:
print(param.shape)
resnet = resnet.to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.SGD(resnet.parameters(), lr=0.01)
loss_epoch_arr = []
max_epochs = 4
min_loss = 1000
n_iters = np.ceil(50000/batch_size)
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs = resnet(inputs)
loss = loss_fn(outputs, labels)
loss.backward()
opt.step()
if min_loss > loss.item():
min_loss = loss.item()
best_model = copy.deepcopy(resnet.state_dict())
print('Min loss %0.2f' % min_loss)
if i % 100 == 0:
print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
del inputs, labels, outputs
torch.cuda.empty_cache()
loss_epoch_arr.append(loss.item())
print('Epoch: %d/%d, Test acc: %0.2f, Train acc: %0.2f' % (
epoch, max_epochs,
evaluation(testloader, resnet), evaluation(trainloader, resnet)))
plt.plot(loss_epoch_arr)
plt.show()
resnet.load_state_dict(best_model)
print(evaluation(trainloader, resnet), evaluation(testloader, resnet))
```
## Inception Model
https://pytorch.org/docs/master/_modules/torchvision/models/inception.html
```
inception = models.inception_v3(pretrained=True)
print(inception)
for param in inception.parameters():
param.requires_grad = False
aux_in_features = inception.AuxLogits.fc.in_features
inception.AuxLogits.fc = nn.Linear(aux_in_features, num_classes)
for param in inception.parameters():
if param.requires_grad:
print(param.shape)
in_features = inception.fc.in_features
inception.fc = nn.Linear(in_features, num_classes)
for param in inception.parameters():
if param.requires_grad:
print(param.shape)
transform_train = transforms.Compose([
transforms.RandomResizedCrop(299),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
])
transform_test = transforms.Compose([
transforms.RandomResizedCrop(299),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True,
transform=transform_train)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True,
transform=transform_test)
batch_size=16
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size, shuffle=False)
inception = inception.to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.SGD(inception.parameters(), lr=0.01)
def evaluation_inception(dataloader, model):
total, correct = 0, 0
for data in dataloader:
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs, aux_outputs = model(inputs)
_, pred = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (pred == labels).sum().item()
return 100 * correct / total
loss_epoch_arr = []
max_epochs = 1
min_loss = 1000
n_iters = np.ceil(50000/batch_size)
for epoch in range(max_epochs):
for i, data in enumerate(trainloader, 0):
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
opt.zero_grad()
outputs, aux_outputs = inception(inputs)
loss = loss_fn(outputs, labels) + 0.3 * loss_fn(aux_outputs, labels)
loss.backward()
opt.step()
if min_loss > loss.item():
min_loss = loss.item()
best_model = copy.deepcopy(inception.state_dict())
print('Min loss %0.2f' % min_loss)
if i % 100 == 0:
print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
del inputs, labels, outputs
torch.cuda.empty_cache()
loss_epoch_arr.append(loss.item())
print('Epoch: %d/%d, Test acc: %0.2f, Train acc: %0.2f' % (
epoch, max_epochs,
evaluation_inception(testloader, inception),
evaluation_inception(trainloader, inception)))
plt.plot(loss_epoch_arr)
plt.show()
inception.load_state_dict(best_model)
print(evaluation_inception(trainloader, inception), evaluation_inception(testloader, inception))
```
## Exercises
1. Structure the above code into a series of functions and then call each model
2. Try out different hyperparameter combinations and try to achieve published results on different networks
3. Try out the CIFAR100 and STL10 datasets
4. Try out another model - SqueezeNet
5. Try training multiple layers and not just the last one
```
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score
import pdb
```
## 【問題1】ラグランジュの未定乗数法による最急降下
SVMの学習は、ラグランジュの未定乗数法を用います。サンプル数分のラグランジュ乗数 $ \lambda $ を用意して、以下の式により更新していきます。
この計算を行うメソッドをScratchSVMClassifierクラスに実装してください。
$$
\lambda_i^{new} = \lambda_i + \alpha(1 - \sum_{j=1}^{n}{\lambda_j y_i y_j k(x_i, x_j)})
$$
ここで $ k(x_i, x_j) $ はカーネル関数です。線形カーネルの場合は次のようになります。
他のカーネル関数にも対応できるように、この部分は独立したメソッドとしておきましょう。
条件として、更新毎に $ \lambda_i >= 0 $ を満たす必要があります。満たさない場合は $ \lambda_i = 0 $とします。
## 【問題2】サポートベクターの決定
計算したラグランジュ乗数 $ \lambda $ が設定した閾値より大きいサンプルをサポートベクターとして扱います。
推定時にサポートベクターが必要になります。サポートベクターを決定し、インスタンス変数として保持しておくコードを書いてください。
閾値はハイパーパラメータですが、1e-5程度からはじめると良いでしょう。サポートベクターの数を出力させられるようにしておくと学習がうまく行えているかを確認できます。
## 【問題3】推定
推定時には、推定したいデータの特徴量とサポートベクターの特徴量をカーネル関数によって計算します。求めた $ f(x) $ の符号が分類結果です。
$$
f(x) = \sum_{n=1}^{N}\lambda_n y_{sv\_n} k(x, s_n)
$$
```
from matplotlib.colors import ListedColormap
import matplotlib.patches as mpatches
def decision_region(X_train, y_train, model, step=0.01, target_names=['0', '1']):
scatter_color = ['red', 'blue']
contourf_color = ['pink', 'skyblue']
n_class = 2
# pred
mesh_f0, mesh_f1 = np.meshgrid(np.arange(np.min(X_train[:,0])-0.5, np.max(X_train[:,0])+0.5, step),
np.arange(np.min(X_train[:,1])-0.5, np.max(X_train[:,1])+0.5, step))
mesh = np.c_[np.ravel(mesh_f0),np.ravel(mesh_f1)]
pred = model.predict(mesh).reshape(mesh_f0.shape)
# plot
plt.title('dicision region')
plt.xlabel('f0')
plt.ylabel('f1')
plt.contourf(mesh_f0, mesh_f1, pred, n_class-1, cmap=ListedColormap(contourf_color))
plt.contour(mesh_f0, mesh_f1, pred, n_class-1, colors='y', linewidths=3, alpha=0.5)
for i, target in enumerate(set(y_train)):
plt.scatter(X_train[y_train==target][:, 0], X_train[y_train==target][:, 1], s=80, color=scatter_color[i], label=target_names[i], marker='o')
patches = [mpatches.Patch(color=scatter_color[i], label=target_names[i]) for i in range(n_class)]
plt.scatter(X_train[svm.support_vec_idx,0], X_train[svm.support_vec_idx,1], s=80, c='yellow')
#plt.xlim([-10, 10])
#plt.ylim([-10, 10])
plt.legend(handles=patches)
plt.legend()
plt.show()
class ScratchSVMClassifier():
def __init__(self, num_iter=5000, lr=0.001, kernel='linear', threshold=1e-5):
# ハイパーパラメータを属性として記録
self.iter = num_iter
self.lr = lr
self.kernel = kernel
self.threshold = threshold
self.X = None
self.y = None
self.m = None # number of samples
self.lmd = None # lambda
self.support_vec_idx = None
def kernel_func(self, X1, X2):
if self.kernel == 'linear':
return np.dot(X1, X2.T)
elif self.kernel == 'poly':
gamma = 5
theta = 2
d = 2
return gamma * (np.dot(X1, X2.T) + theta)**d
def fit(self, X, y):
self.X = X
self.y = y.reshape(-1,)
self.m = len(self.X)
self.lmd = np.random.rand(self.m) * 0.1
for _ in range(self.iter):
self.lmd += self.lr * (1 - self.y*np.sum(self.y*self.lmd*self.kernel_func(self.X, self.X), axis=1))
self.lmd[self.lmd<0] = 0
self.support_vec_idx = np.where(self.lmd>self.threshold)[0]
#print(len(self.support_vec_idx))
#print(self.lmd)
def predict(self, X_test):
f = np.sum(self.y[self.support_vec_idx]*self.lmd[self.support_vec_idx]*self.kernel_func(X_test, self.X[self.support_vec_idx,:]), axis=1)
f = np.where(f<0, -1, 1)
return f
```
## 【問題4】学習と推定
機械学習スクラッチ入門のSprintで用意したシンプルデータセット1の2値分類に対してスクラッチ実装の学習と推定を行なってください。
scikit-learnによる実装と比べ、正しく動いているかを確認してください。
AccuracyやPrecision、Recallなどの指標値はscikit-learnを使用してください。
## 【問題5】決定領域の可視化
決定領域を可視化してください。
サポートベクトルは異なる色で示してください。
## 【問題6】多項式カーネル関数の作成
最初に作成した実装では線形カーネルを使用していました。多項式カーネルにも切り替えられるようにしましょう。
$$
k(x_i, x_j) = \gamma(x_{i}^{T} x_j + \theta_0)^{d}
$$
```
np.random.seed(seed=0)
n_samples = 500
f0 = [-1, 2]
f1 = [2, -1]
cov = [[1.0,0.8], [0.8, 1.0]]
f0 = np.random.multivariate_normal(f0, cov, int(n_samples/2))
f1 = np.random.multivariate_normal(f1, cov, int(n_samples/2))
X = np.concatenate((f0, f1))
y = np.concatenate((np.ones((int(n_samples/2))), np.ones((int(n_samples/2))) *(-1))).astype(np.int)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
svm = ScratchSVMClassifier(kernel='linear')
svm.fit(X_train, y_train)
y_pred = svm.predict(X_test)
accuracy_score(y_test, y_pred)
decision_region(X_train, y_train, svm)
svm = ScratchSVMClassifier(num_iter=5000, lr=0.000001, kernel='poly')
svm.fit(X_train, y_train)
y_pred = svm.predict(X_test)
accuracy_score(y_test, y_pred)
decision_region(X_train, y_train, svm)
```
### sklearnによる実装との比較
```
svc = SVC(kernel='linear')
svc.fit(X_train, y_train)
y_pred = svc.predict(X_test)
accuracy_score(y_test, y_pred)
decision_region(X_train, y_train, svc)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.