text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
import necessary modules
```
import vcs # For plots
import vcsaddons # module containing pcoords
import cdms2 # for data
import glob # to list files in directories
import pcmdi_metrics # for special json loader class
```
## Work around to visualize plot in Jupyter Notebook
This class allow use to use vcsaddons plots
```
import tempfile
import base64
class VCSAddonsNotebook(object):
def __init__(self, x):
self.x = x
def _repr_png_(self):
fnm = tempfile.mktemp()+".png"
x.png(fnm)
encoded = base64.b64encode(open(fnm, "rb").read())
return encoded
def __call__(self):
return self
```
# Sample Data
These files are in the test directory of pcmdi_metrics repo at:
http://github.com/PCMDI/pcmdi_metrics.git
```
# Prepare list of json files
# Location on your computer
json_pth = "/git/pcmdi_metrics/test/graphics"
# Geenrate list ofjson files
json_files = glob.glob(
os.path.join(
json_pth,
"json",
"v2.0",
"*.json"))
json_files += glob.glob(
os.path.join(
json_pth,
"json",
"v1.0",
"*.json"))
# Read them in via pmp special json class
J = pcmdi_metrics.pcmdi.io.JSONs(json_files)
# Retrieve data we need for plot
# Annual mean RMS (XYT dimensions)
# All models and all variables
rms_xyt = J(statistic=["rms_xyt"],season=["ann"],region="global")(squeeze=1)
```
Let's take a look at the array generated
Note the axis are strings of varialbes used and models
The order of the axes is the order on the plot
```
rms_xyt.info()
# Ok now let's create a VCS pcoord graphic method
# initialize a canvas
x=vcs.init(geometry=(1200,600),bg=True)
import vcsaddons
gm = vcsaddons.createparallelcoordinates(x=x)
```
# Preparing the plot
## Data
'id' is used for variable in plot the JSON class returns var as "pmp", here "RMS" is more appropriate
'title' is used to draw the plot title (location/font controlled by template)
## Template
The template section prepares where data will be rendered on plot, and the fonts used
fonts are controlled via textorientation and texttable VCS primary objects
Here we need to angle a bit the xlabels (45 degrees)
We also want to turn off the boxes around the legend and the data area.
```
# Prepare the graphics
# Set variable name
rms_xyt.id = "RMS"
# Set units of each variables on axis
# This is a trick to have units listed on plot
rms_xyt.getAxis(-2).units = ["mm/day","mm/day","hPa","W/m2","W/m2","W/m2", "K","K","K","m/s","m/s","m/s","m/s","m"]
# Sets title on the variable
rms_xyt.title = "Annual Mean Error"
# Preprare the canvas areas
t = vcs.createtemplate()
# Create a text orientation object for xlabels
to=x.createtextorientation()
to.angle=-45
to.halign="right"
# Tell template to use this orientation for x labels
t.xlabel1.textorientation = to.name
# Define area where plot will be drawn in x direction
t.reset('x',0.05,0.9,t.data.x1,t.data.x2)
ln = vcs.createline()
# Turn off box around legend
ln.color = [[0,0,0,0]]
t.legend.line = ln
# turn off box around data area
t.box1.priority=0
# Define box where legend will be drawn
t.legend.x1 = .91
t.legend.x2 = .99
# use x/y of data drawn for legend height
t.legend.y1 = t.data.y1
t.legend.y2 = t.data.y2
# Plot with default values of graphic method
# Bug vcsaddons need to return a display
# as a result it does not show up in notebook
x.clear()
show = VCSAddonsNotebook(x)
gm.plot(rms_xyt,template=t,bg=True)
show()
```
# Control various aspects of the graphic method
We want the first two model to be 'blue' and 'red' and a bit thicker
All other plots will be 'grey' and 'dashed'
```
x.clear()
gm.linecolors = ["blue","red","grey"]
gm.linestyles=["solid","solid","dot"]
gm.linewidths=[5.,5.,1.]
gm.markercolors = ["blue","red","grey"]
gm.markertypes=["triangle_up","star","dot"]
gm.markersizes=[7,5,2]
gm.plot(rms_xyt,template=t,bg=True)
show()
# change order and number of models and variables
axes = rms_xyt.getAxisList()
models = ['MIROC4h', 'HadGEM2-AO', 'GFDL-ESM2M',
'GFDL-ESM2G', 'GFDL-CM3', 'FGOALS-g2', 'CSIRO-Mk3-6-0', 'CESM1-WACCM',
'CESM1-FASTCHEM', 'CESM1-CAM5', 'CESM1-BGC', 'CCSM4', 'ACCESS1-3', 'ACCESS1-0',
'0071-0100'] # invert them
variables = ['prw', 'psl', 'rltcre', 'rlut', 'rstcre', 'ta-200', 'ta-850', 'tas',
'ua-850', 'va-850', 'zg-500']
rms_xyt = J(statistic=["rms_xyt"],season=["ann"],region="global",model=models,variable=variables)(squeeze=1)
x.clear()
gm.plot(rms_xyt,template=t,bg=True)
show()
```
| github_jupyter |
```
!pip install sentence-transformers
"""
This script trains sentence transformers with a triplet loss function.
As corpus, we use the wikipedia sections dataset that was describd by Dor et al., 2018, Learning Thematic Similarity Metric Using Triplet Networks.
"""
from sentence_transformers import SentenceTransformer, InputExample, LoggingHandler, losses, models, util
from torch.utils.data import DataLoader
from sentence_transformers.evaluation import TripletEvaluator
from datetime import datetime
from zipfile import ZipFile
from sentence_transformers.evaluation import EmbeddingSimilarityEvaluator, BinaryClassificationEvaluator
import csv
import logging
import os
logging.basicConfig(format='%(asctime)s - %(message)s',
datefmt='%Y-%m-%d %H:%M:%S',
level=logging.INFO,
handlers=[LoggingHandler()])
logger = logging.getLogger(__name__)
logging.info("Read test dataset")
test_sts_samples = []
with open("/content/sample_data/sentence_pair_unique.csv", 'rt', encoding='utf8') as fIn:
reader = csv.DictReader(fIn)
for row in reader:
score = float(row['avg']) / 4.0 # Normalize score to range 0 ... 1
inp_example = InputExample(texts=[row['sent1_text'], row['sent2_text']], label=score)
test_sts_samples.append(inp_example)
model_name = 'stsb-bert-base'
output_path = "output/training-wikipedia-sections-"+model_name+"-"+datetime.now().strftime("%Y-%m-%d_%H-%M-%S")
model = SentenceTransformer(model_name)
test_evaluator = EmbeddingSimilarityEvaluator.from_input_examples(test_sts_samples, name='sts-test')
test_evaluator(model, output_path=output_path)
import pandas as pd
# Load the dataset into a pandas dataframe.
df_test = pd.read_csv("/content/sample_data/sentence_pair_unique.csv")
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df_test.shape[0]))
# Display 5 random rows from the data.
df_test.sample(1)
# Get the lists of sentences and their labels.
sent1 = df_test.sent1_text.values
sent2 = df_test.sent2_text.values
sent1 = sent1.tolist()
sent2 = sent2.tolist()
sent1_embeddings = model.encode(sent1)
sent1_embeddings.shape
sent2_embeddings = model.encode(sent2)
sent2_embeddings.shape
from sklearn.metrics.pairwise import cosine_similarity
import scipy
import pandas as pd
import numpy as np
cos_list = []
cos2_list = []
for j in range(len(sent1_embeddings)):
sent1 = sent1_embeddings[j]
sent2 = sent2_embeddings[j]
cos = 1-scipy.spatial.distance.cdist(sent1.reshape(1, -1), sent2.reshape(1, -1), "cosine")[0][0]
cos2 = cosine_similarity(sent1.reshape(1, -1), sent2.reshape(1, -1))[0][0]
cos_list.append(cos)
cos2_list.append(cos2)
assert(len(cos_list)==len(df_test))
assert(len(cos2_list)==len(df_test))
df_test["model_similarity"] = cos_list
df_test["model_similarity2"] = cos2_list
df_test.to_csv("/content/sample_data/sentence_pair_sbert.csv",
index=None)
```
## Siamese Network Training
```
### Create a torch.DataLoader that passes training batch instances to our model
model_name = 'stsb-bert-base'
train_batch_size = 8
output_path = "output/training-yelp-pair-"+model_name+"-"+datetime.now().strftime("%Y-%m-%d_%H-%M-%S")
num_epochs = 1
# n_train = 1000
n_train = 3000
# n_train = 8000
trip_type = "firstsentence"
model = SentenceTransformer(model_name)
logger.info("Read Yelp Pair train dataset")
train_examples = []
with open("/content/sample_data/yelp_pair_train_10000_" + trip_type + ".csv", encoding="utf-8") as fIn:
reader = csv.DictReader(fIn)
for row in reader:
# print(row)
train_examples.append(InputExample(texts=[row['sent1_text'], row['sent2_text']], label=int(row['label'])))
test_examples = train_examples[0:1000]
dev_examples = train_examples[1000:2000]
train_examples = train_examples[2000:(2000+n_train)]
print(len(test_examples))
print(len(dev_examples))
print(len(train_examples))
train_dataloader = DataLoader(train_examples, shuffle=True, batch_size=train_batch_size)
train_loss = losses.SoftmaxLoss(model=model, sentence_embedding_dimension=model.get_sentence_embedding_dimension(), num_labels=2)
logger.info("Read Yelp Pair dev dataset")
evaluator = BinaryClassificationEvaluator.from_input_examples(dev_examples, name='yelp-dev_sbertft_' + str(n_train) + "_" + trip_type + "_run1")
warmup_steps = int(len(train_dataloader) * num_epochs * 0.1) #10% of train data
print(output_path)
# Train the model
model.fit(train_objectives=[(train_dataloader, train_loss)],
evaluator=evaluator,
epochs=num_epochs,
evaluation_steps=1000,
warmup_steps=warmup_steps,
output_path=output_path)
##############################################################################
#
# Load the stored model and evaluate its performance on STS benchmark dataset
#
##############################################################################
logger.info("Read test examples")
model = SentenceTransformer(output_path)
test_evaluator = BinaryClassificationEvaluator.from_input_examples(dev_examples, name='yelp-test_sbertft_' + str(n_train) + "_" + trip_type + "_run1")
test_evaluator(model, output_path=output_path)
import pandas as pd
# Load the dataset into a pandas dataframe.
df_test = pd.read_csv("/content/sample_data/sentence_pair_unique.csv")
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df_test.shape[0]))
# Display 5 random rows from the data.
df_test.sample(1)
# Get the lists of sentences and their labels.
sent1 = df_test.sent1_text.values
sent2 = df_test.sent2_text.values
sent1 = sent1.tolist()
sent2 = sent2.tolist()
sent1_embeddings = model.encode(sent1)
sent1_embeddings.shape
sent2_embeddings = model.encode(sent2)
sent2_embeddings.shape
from sklearn.metrics.pairwise import cosine_similarity
import scipy
import pandas as pd
import numpy as np
cos_list = []
cos2_list = []
for j in range(len(sent1_embeddings)):
sent1 = sent1_embeddings[j]
sent2 = sent2_embeddings[j]
cos = 1-scipy.spatial.distance.cdist(sent1.reshape(1, -1), sent2.reshape(1, -1), "cosine")[0][0]
cos2 = cosine_similarity(sent1.reshape(1, -1), sent2.reshape(1, -1))[0][0]
cos_list.append(cos)
cos2_list.append(cos2)
assert(len(cos_list)==len(df_test))
assert(len(cos2_list)==len(df_test))
df_test["model_similarity"] = cos_list
df_test["model_similarity2"] = cos2_list
df_test.to_csv("/content/sample_data/sentence_pair_sbert_yelp_"+str(n_train)+"_" + trip_type + "_pair_run1.csv",
index=None)
```
## Triplet Network Training
```
### Create a torch.DataLoader that passes training batch instances to our model
model_name = 'stsb-bert-base'
train_batch_size = 8
output_path = "output/training-yelp-triplets-"+model_name+"-"+datetime.now().strftime("%Y-%m-%d_%H-%M-%S")
num_epochs = 1
# n_train = 1000
n_train = 3000
# n_train = 8000
trip_margin = 9.0
trip_type = "firstsentence"
model = SentenceTransformer(model_name)
logger.info("Read Triplet train dataset")
train_examples = []
with open("/content/sample_data/yelp_triplets_train_10000_" + trip_type + ".csv", encoding="utf-8") as fIn:
reader = csv.DictReader(fIn)
for row in reader:
train_examples.append(InputExample(texts=[row['anchor_text'], row['same_text'], row['diff_text']], label=0))
len(train_examples)
test_examples = train_examples[0:1000]
dev_examples = train_examples[1000:2000]
train_examples = train_examples[2000:(2000+n_train)]
print(len(test_examples))
print(len(dev_examples))
print(len(train_examples))
train_dataloader = DataLoader(train_examples, shuffle=True, batch_size=train_batch_size)
train_loss = losses.TripletLoss(model=model, triplet_margin= trip_margin)
len(train_dataloader)
logger.info("Read Yelp Triplet dev dataset")
evaluator = TripletEvaluator.from_input_examples(dev_examples, name='yelp-dev_sbertft_' + str(n_train) + "_" + trip_type + "_margin" + str(int(trip_margin))+ "_run1")
warmup_steps = int(len(train_dataloader) * num_epochs * 0.1) #10% of train data
import torch
torch.cuda.empty_cache()
# Train the model
model.fit(train_objectives=[(train_dataloader, train_loss)],
evaluator=evaluator,
epochs=num_epochs,
evaluation_steps=200,
warmup_steps=warmup_steps,
output_path=output_path)
##############################################################################
#
# Load the stored model and evaluate its performance on STS benchmark dataset
#
##############################################################################
logger.info("Read test examples")
model = SentenceTransformer(output_path)
test_evaluator = TripletEvaluator.from_input_examples(test_examples, name='yelp-test_sbertft_' + str(n_train) + "_" + trip_type + "_margin" + str(int(trip_margin))+ "_run1")
test_evaluator(model, output_path=output_path)
## Baseline for SBERT##
## # logger.info("Read test examples")
# model = SentenceTransformer(model_name)
# test_evaluator = TripletEvaluator.from_input_examples(test_examples, name='yelp-test')
# test_evaluator(model, output_path=output_path)
import pandas as pd
model = SentenceTransformer(output_path)
# Load the dataset into a pandas dataframe.
df_test = pd.read_csv("/content/sample_data/sentence_pair_unique.csv")
# Report the number of sentences.
print('Number of training sentences: {:,}\n'.format(df_test.shape[0]))
# Display 5 random rows from the data.
df_test.sample(1)
# Get the lists of sentences and their labels.
sent1 = df_test.sent1_text.values
sent2 = df_test.sent2_text.values
sent1 = sent1.tolist()
sent2 = sent2.tolist()
sent1_embeddings = model.encode(sent1)
sent1_embeddings.shape
sent2_embeddings = model.encode(sent2)
sent2_embeddings.shape
from sklearn.metrics.pairwise import cosine_similarity
import scipy
import pandas as pd
import numpy as np
cos_list = []
cos2_list = []
for j in range(len(sent1_embeddings)):
sent1 = sent1_embeddings[j]
sent2 = sent2_embeddings[j]
cos = 1-scipy.spatial.distance.cdist(sent1.reshape(1, -1), sent2.reshape(1, -1), "cosine")[0][0]
cos2 = cosine_similarity(sent1.reshape(1, -1), sent2.reshape(1, -1))[0][0]
cos_list.append(cos)
cos2_list.append(cos2)
assert(len(cos_list)==len(df_test))
assert(len(cos2_list)==len(df_test))
df_test["model_similarity"] = cos_list
df_test["model_similarity2"] = cos2_list
df_test.to_csv("/content/sample_data/sentence_pair_sbert_yelp_"+str(n_train)+"_" + trip_type + "_triplet_margin" + str(int(trip_margin)) + "_run1.csv",
index=None)
import gzip
sts_dataset_path = 'datasets/stsbenchmark.tsv.gz'
if not os.path.exists(sts_dataset_path):
util.http_get('https://sbert.net/datasets/stsbenchmark.tsv.gz', sts_dataset_path)
logging.info("Read STSbenchmark train dataset")
train_sts_samples = []
dev_sts_samples = []
test_sts_samples = []
with gzip.open(sts_dataset_path, 'rt', encoding='utf8') as fIn:
reader = csv.DictReader(fIn, delimiter='\t', quoting=csv.QUOTE_NONE)
for row in reader:
score = float(row['score']) / 5.0 # Normalize score to range 0 ... 1
inp_example = InputExample(texts=[row['sentence1'], row['sentence2']], label=score)
if row['split'] == 'dev':
dev_sts_samples.append(inp_example)
elif row['split'] == 'test':
test_sts_samples.append(inp_example)
else:
train_sts_samples.append(inp_example)
##############################################################################
#
# Load the stored model and evaluate its performance on STS benchmark dataset
#
##############################################################################
model = SentenceTransformer(output_path)
test_evaluator = EmbeddingSimilarityEvaluator.from_input_examples(test_sts_samples, name='sts-test')
test_evaluator(model, output_path=output_path)
len(test_sts_samples)
```
| github_jupyter |
## Data Mining and Machine Learning
## Edgar Acuna
### February 2021
## Mean/median/mode and k-nn imputation
### Datasets: Titanic, Breastw y Segment
```
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
```
### Ejemplo1. Leyendo los datos de Titanic
```
titanic=pd.read_csv('https://raw.githubusercontent.com/eacunafer/DataAnalysiswithPython3/master/Datasets/titanic.csv',header=0,sep=',',na_values='')
titanic.info()
titanic.head(39)
```
Deleting columns: PassengerID, Name and cabin(it has a lot of missing values)
```
titanic1=titanic.iloc[:,[1,3,4,5,6,8,10]]
titanic1.info()
```
Hay missings en las variables Age, Fare que son continuas y en Embarked que es categorica
```
#computing the values to be used in the imputation
t3=titanic1['Age'].mean()
t6=titanic1['Fare'].mean()
t7=titanic1['Embarked'].mode().iloc[0]
#Performing the imputacion
values = {'Age': t3, 'Fare': t6, 'Embarked': t7}
titanic2=titanic1.fillna(value=values)
titanic2.info()
```
Imputando los missing values por la media si el atributo es continuo y por la moda si el atributo es nominal
```
#Imputando los missing values por la media si el atributo es continuo
#y por la moda si el atributo es nominal
from sklearn.impute import SimpleImputer
from sklearn.base import TransformerMixin
class DataFrameImputer(TransformerMixin):
def fit(self, X, y=None):
self.fill = pd.Series([X[c].value_counts().index[0]
if X[c].dtype == np.dtype('O') else X[c].mean() for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.fill)
X=titanic1
xt = DataFrameImputer().fit_transform(X)
xt.info()
#Imputando los missing values por la mediana si el atributo es continuo
#y por la moda si el atributo es nominal
class DataFrameImputer(TransformerMixin):
def fit(self, X, y=None):
self.fill = pd.Series([X[c].value_counts().index[0]
if X[c].dtype == np.dtype('O') else X[c].median() for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.fill)
X=titanic1
xt = DataFrameImputer().fit_transform(X)
xt.info()
```
### IMPUTACION k-nn
```
import numpy as np
from sklearn.impute import KNNImputer
X = [[4, 5, 6,np.nan], [5, 1, 1,4], [7, 9,2, 5], [8, 2,8, 5],[6,4,2,6]]
#imputer = KNNImputer(n_neighbors=3)
#imputer.fit_transform(X)
KNNImputer(n_neighbors=3).fit_transform(X)
```
### Ejemplo 2. k-nn imputation applied to the Breast-wisconsin dataset (todas las features son continuas)
```
#Reading the data from the UCI
breastdf=pd.read_csv("https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data",header=None, sep=",",na_values=['?'])
#breastdf=pd.read_csv("c://PW-PR/breast-cancer-wisconsin.data",header=None, sep=",",na_values=['?'])
breastdf.columns=['idx','v1','v2','v3','v4','v5','v6','v7','v8','v9','class']
breastdf.info()
breastdf.head()
breast1=breastdf.iloc[:,1:11]
breast1.head()
b1=np.array(breast1)
imputer = KNNImputer(n_neighbors=5)
data_imp=imputer.fit_transform(b1)
completo5=pd.DataFrame(data_imp)
completo5.info()
```
### Ejemplo 3. Imputacion k-nn a Titanic. Hay que convertir las variables categoticas a numericas porque knn-imputacion solo esta implementada para variables numericas
Convirtiendo las variables nominales en numericas, debido a que la imputacion knn de python no trabaja con variables
mezcladas Debe hacerse con cuidado porque el atributo categorico contiene missing value y el LabelEncoder
puede asignarle un valor en lugar de ignorarlo. LabelEncoder codifica los valores categoricas en numero enteros desde 0 hasta el numero de categorias-1. Similar resultado se puede conseguir con cat.codes de Pandas
```
#Guardando las posiciones donde estan los missings
titanic3=titanic1.copy()
mask = titanic3.isnull()
print(mask)
#Convirtiendo a numero los atributos categoricos
titanic1=titanic1.astype(str).apply(LabelEncoder().fit_transform)
#Sustituyendo con NaN las posiciones donde los atributos categoricos tienen missings
titanic4=titanic1.where(~mask, titanic3).copy()
titanic4.tail(7)
titanic4.info()
#Convirtiendo en Float las variables categoricas donde ahi missings
titanic4['Age'] = titanic4['Age'].astype('float64')
titanic4['Fare'] = titanic4['Fare'].astype('float64')
titanic4['Embarked'] = titanic4['Embarked'].astype('float64')
b2=np.array(titanic4)
imputer = KNNImputer(n_neighbors=5)
titanic_knnimp=imputer.fit_transform(b2)
completo=pd.DataFrame(titanic_knnimp)
completo.info()
```
### Example 4. Simulating missing values and imputing them. Dataset: segment from the UCI
This dataset is an image segmentation database similar to a database already present in the repository
(Image segmentation database) but in a slightly different form. The instances were drawn randomly from a
database of 7 outdoor images.
The images were handsegmented to create a classification for every pixel. Each instance is a 3x3 region.
```
seg=pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/statlog/segment/segment.dat',header=None, delim_whitespace=True)
#seg=seg=pd.read_table('c://PW-PR/segment.dat',header=None, delim_whitespace=True)
seg.columns=['v1','v2','v3','v4','v5','v6','v7','v8','v9','v10','v11','v12','v13','v14','v15','v16','v17','v18','v19','class']
seg.info()
```
Todos las variables de segment son cuantitativas.
```
#simulando un 5% de celdas missing
import random
seg5=seg.copy()
seg5= seg5.stack().sample(frac=0.95).unstack().reindex(index=seg5.index, columns=seg5.columns)
seg5.head()
seg5.info()
#imputando por la media
seg5.mean=seg5.apply(lambda x: x.fillna(x.mean()),axis=0)
#Imputando por la mediana
seg5.median=seg5.apply(lambda x: x.fillna(x.median()),axis=0)
b1=np.array(seg5)
imputer = KNNImputer(n_neighbors=5)
data_nimp=imputer.fit_transform(b1)
completoknn5=pd.DataFrame(data_nimp)
completoknn5.info()
from sklearn.metrics import mean_squared_error
#Calculando el cuadrado medio del error para imputacion por la media
completo5mean=pd.DataFrame(seg5.mean)
mean_squared_error(seg,completo5mean)
#Calculando el cuadrado medio del error para imputacion por la mediana
completo5median=pd.DataFrame(seg5.median)
mean_squared_error(seg,completo5median)
#Calculando el cuadrado medio del error para imputacion knn
mean_squared_error(seg,completoknn5)
```
Notice that the best imputation method was k-nn
```
#simulando un 20% de celdas missing
seg20=seg.copy()
seg20= seg20.stack().sample(frac=0.80).unstack().reindex(index=seg20.index, columns=seg20.columns)
seg20.info()
#hallando el numero de filas que tienen missings
sum(seg20.apply(lambda x: sum(x.isnull().values), axis = 1)>0)
#imputando por la media al 20%
seg20.mean=seg20.apply(lambda x: x.fillna(x.mean()),axis=0)
completo20mean=pd.DataFrame(seg20.mean)
mean_squared_error(seg,completo20mean)
#imputando por la mediana al 20%
seg20.median=seg20.apply(lambda x: x.fillna(x.median()),axis=0)
completo20median=pd.DataFrame(seg20.median)
mean_squared_error(seg,completo20median)
b2=np.array(seg20)
imputer = KNNImputer(n_neighbors=5)
data_imp=imputer.fit_transform(b2)
completoknn20=pd.DataFrame(data_imp)
mean_squared_error(seg,completoknn20)
```
Once again the best imputation method was k-nn
| github_jupyter |
<h1> <b>Homework 2</b></h1>
<i>Alejandro J. Rojas<br>
ale@ischool.berkeley.edu<br>
W261: Machine Learning at Scale<br>
Week: 02<br>
Jan 26, 2016</i></li>
<h2>HW2.0. </h2>
What is a race condition in the context of parallel computation? Give an example.
What is MapReduce?
How does it differ from Hadoop?
Which programming paradigm is Hadoop based on? Explain and give a simple example in code and show the code running.
<h2>HW2.1. Sort in Hadoop MapReduce</h2>
Given as input: Records of the form '<'integer, “NA”'>', where integer is any integer, and “NA” is just the empty string.
Output: sorted key value pairs of the form '<'integer, “NA”'>' in decreasing order; what happens if you have multiple reducers? Do you need additional steps? Explain.
Write code to generate N random records of the form '<'integer, “NA”'>'. Let N = 10,000.
Write the python Hadoop streaming map-reduce job to perform this sort. Display the top 10 biggest numbers. Display the 10 smallest numbers
# Data
```
import random
N = 10000 ### for a sample size of N
random.seed(0) ### pick a random seed to replicate results
input_file = open("numcount.txt", "w") # writing file
for i in range(N):
a = random.randint(0, 100) ### Select a random integer from 0 to 100
b = ''
input_file.write(str(a))
input_file.write(b)
input_file.write('\n')
input_file.close()
```
# Mapper
```
%%writefile mapper.py
#!/usr/bin/python
import sys
for line in sys.stdin: ### input comes from STDIN (standard input)
number = line.strip() ### remove leading and trailing whitespace
print ('%s\t%s' % (number, 1)) ### mapper out looks like 'number' \t 1
!chmod +x mapper.py
```
# Reducer
```
%%writefile reducer.py
#!/usr/bin/python
from operator import itemgetter
import sys
current_number = None
current_count = 0
number = None
numlist = []
# input comes from STDIN
for line in sys.stdin:
line = line.strip() ### remove leading and trailing whitespace
line = line.split('\t') ### parse the input we got from mapper.py
number = line[0] ### integer generated randomly we got from mapper.py
try:
count = line[1]
count = int(count) ### convert count (currently a string) to int
except ValueError: ### if count was not a number then silently
continue ### ignore/discard this line
if current_number == number: ### this IF-switch only works because Hadoop sorts map output
current_count += count ### by key (here: number) before it is passed to the reducer
else:
if current_number:
numlist.append((current_number,current_count)) ### store tuple in a list once totalize count per number
current_count = count ### set current count
current_number = number ### set current number
if current_number == number: ### do not forget to output the last word if needed!
numlist.append((current_number,current_count))
toplist = sorted(numlist,key=lambda record: record[1], reverse=True) ### sort list from largest count to smallest
bottomlist = sorted(numlist,key=lambda record: record[1]) ### sort list from smalles to largest
print '%25s' %'TOP 10', '%25s' % '', '%28s' %'BOTTOM 10'
print '%20s' %'Number', '%10s' %'Count', '%20s' % '', '%20s' %'Number','%10s' %'Count'
for i in range (10):
print '%20s%10s' % (toplist[i][0], toplist[i][1]),'%20s' % '', '%20s%10s' % (bottomlist[i][0], bottomlist[i][1])
!chmod +x reducer.py
!echo "10 \n 10\n 5\n 6\n 8\n 9 \n 10 \n 9 \n 12 \n 21 \n 22 \n 23 \n 24 \n 25" | python mapper.py | sort -k1,1 | python reducer.py
```
# Run numcount in Hadoop
<h2>start yarn and hdfs</h2>
```
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-yarn.sh ### start up yarn
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-dfs.sh ### start up dfs
```
<h2> remove files from prior runs </h2>
```
!hdfs dfs -rm -r /user/venamax ### remove prior files
```
<h2> create folder</h2>
```
!hdfs dfs -mkdir -p /user/venamax ### create hdfs folder
```
<h2> upload numcount.txt to hdfs</h2>
```
!hdfs dfs -put numcount.txt /user/venamax #### save source data file to hdfs
```
<h2> Hadoop streaming command </h2>
hadoop jar hadoopstreamingjarfile \
-D stream.num.map.output.key.fields=n \
-mapper mapperfile \
-reducer reducerfile \
-input inputfile \
-output outputfile
```
!hadoop jar hadoop-*streaming*.jar -mapper mapper.py -reducer reducer.py -input numcount.txt -output numcountOutput
```
<h2>show the results</h2>
```
!hdfs dfs -cat numcountOutput/part-00000
```
<h2>stop yarn and hdfs </h2>
```
!/usr/local/Cellar/hadoop/2.7.1/sbin/stop-yarn.sh
!/usr/local/Cellar/hadoop/2.7.1/sbin/stop-dfs.sh
```
<h2>=====================
END OF HW 2.1</h2>
<h2>HW2.2. WORDCOUNT</h2>
Using the Enron data from HW1 and Hadoop MapReduce streaming, write the mapper/reducer job that will determine the word count (number of occurrences) of each white-space delimitted token (assume spaces, fullstops, comma as delimiters). Examine the word “assistance” and report its word count results.
CROSSCHECK: >grep assistance enronemail_1h.txt|cut -d$'\t' -f4| grep assistance|wc -l
8
#NOTE "assistance" occurs on 8 lines but how many times does the token occur? 10 times! This is the number we are looking for!
# Mapper
```
%%writefile mapper.py
#!/usr/bin/python
## mapper.py
## Author: Alejandro J. Rojas
## Description: mapper code for HW2.2
import sys
import re
########## Collect user input ###############
filename = 'enronemail_1h.txt'
findwords = ['assistance', 'commercial', 'intelligence','call']
counts = {}
##filename = sys.argv[1]
##findwords = re.split(" ",sys.argv[2].lower())
for keyword in findwords: ### Initialize to zero all keywords to find
counts[keyword] = 0
## open the input file
with open (filename, "r") as myfile:
for line in myfile.readlines():
record = re.split(r'\t+', line)
if len(record) == 4: ### Take only complete records
for keyword in findwords: ### For each word to find
for i in range (2,len(record)): ### Compare it to the words from each email
bagofwords = re.split(" ",record[i]) ### Break each email records into words
for word in bagofwords:
neword = word.strip(',') ### eliminate comas
if keyword in neword:
counts[keyword] += 1
for keyword in findwords: ### output results in the form:
print ('%s\t%s'% (keyword, str(counts[keyword]))) ### word to find, count
!chmod +x mapper.py
```
# Reducer
```
%%writefile reducer.py
#!/usr/bin/python
from operator import itemgetter
import sys
current_word = None
current_count = 0
word = None
words = {}
# input comes from STDIN
# input comes from STDIN
for line in sys.stdin:
line = line.strip() ### remove leading and trailing whitespace
line = line.split('\t') ### parse the input we got from mappe
word = line[0]
try:
count = line[1]
count = int(count) ### convert count (currently a string) to int
except ValueError: ### if count was not a number then silently
continue ### ignore/discard this line
if current_word == word: ### this IF-switch only works because Hadoop sorts map output
current_count += count ### by key (here: number) before it is passed to the reducer
else:
if current_word:
words[current_word] = current_count ### store tuple in a list once totalize count per number
current_count = count ### set current count
current_word = word ### set current word
if current_word == word: ### do not forget to output the last word if needed!
words[current_word] = current_count
for word in words:
print ('We found %s' %word, ' on %s'%words[word] , 'occassions.')
!chmod +x reducer.py
```
# Run numcount in Hadoop
<h2>start yarn and hdfs</h2>
```
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-yarn.sh ### start up yarn
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-dfs.sh ### start up dfs
```
<h2> remove files from prior runs</h2>
```
!hdfs dfs -rm -r /user/venamax ### remove prior files
```
<h2> create folder</h2>
```
!hdfs dfs -mkdir -p /user/venamax ### create hdfs folder
```
<h2> upload enronemail_1h.txt to hdfs</h2>
```
!hdfs dfs -put enronemail_1h.txt /user/venamax #### save source data file to hdfs
```
<h2> Hadoop streaming command </h2>
```
!hadoop jar hadoop-*streaming*.jar -mapper mapper.py -reducer reducer.py -input enronemail_1h.txt -output wordcountOutput
```
<h2>show the results</h2>
```
!hdfs dfs -cat wordcountOutput/part-00000
```
<h2>stop yarn and hdfs </h2>
```
!/usr/local/Cellar/hadoop/2.7.1/sbin/stop-yarn.sh
!/usr/local/Cellar/hadoop/2.7.1/sbin/stop-dfs.sh
```
<h2>=====================
END OF HW 2.2</h2>
# Mapper
```
%%writefile mapper.py
#!/usr/bin/python
## mapper.py
## Author: Alejandro J. Rojas
## Description: mapper code for HW2.2
import sys
import re
########## Collect user input ###############
filename = sys.argv[1]
findwords = re.split(" ",sys.argv[2].lower())
with open (filename, "r") as myfile:
for line in myfile.readlines():
line = line.strip()
record = re.split(r'\t+', line) ### Each email is a record with 4 components
### 1) ID 2) Spam Truth 3) Subject 4) Content
if len(record)==4: ### Take only complete records
for i in range (2,len(record)): ### Starting from Subject to the Content
bagofwords = re.split(" " | "," ,record[i])### Collect all words present on each email
for word in bagofwords:
flag=0
if word in findwords:
flag=1
print '%s\t%s\t%s\t%s\t%s' % (word, 1,record[0], record[1],flag)
### output: word, 1, id, spam truth and flag
!chmod +x mapper.py
```
# Reducer
```
%%writefile reducer.py
#!/usr/bin/python
from operator import itemgetter
import sys
from itertools import groupby
current_word, word = None, None
current_wordcount, current_spam_wordcount, current_ham_wordcount = 0,0,0
current_id, record_id = None, None
current_y_true, y_true = None, None
current_flag, flag = None,None
sum_records, sum_spamrecords, sum_hamrecords = 0,0,0
sum_spamwords, sum_hamwords = 0,0
flagged_words = []
emails={} #Associative array to hold email data
words={} #Associative array for word data
# input comes from STDIN
for line in sys.stdin:
line = line.strip() ### remove leading and trailing whitespace
line = line.split('\t') ### parse the input we got from mapper.py
word = line[0] ### word we get from mapper.py
try:
count = line[1]
count = int(count) ### convert count (currently a string) to int
email = line[2] ### id that identifies each email
y_true = line[3]
y_true = int(y_true) ### spam truth as an integer
flag = line[4]
flag = int(flag) ### flags if word is in the user specified list
except ValueError: ### if count was not a number then silently
continue ### ignore/discard this line
if current_word == word: ### this IF-switch only works because Hadoop sorts map output
current_count += count ### by key (here: word) before it is passed to the reducer
if current_word not in words.keys():
words[current_word]={'ham_count':0,'spam_count':0,'flag':flag}
if email not in emails.keys():
emails[current_email]={'y_true':y_true,'word_count':0,'words':[]}
sum_records +=1
if y_true == 1:
sum_spamrecords +=1
else
sum_hamrecords +=1
if y_true == 1: ### if record where word is located is a spam
current_spamcount += count ### add to spam count of that word
sum_spamwords += 1
else:
current_hamcount += count ### if not add to ham count of thet word
sum_hamwords +=1
emails[current_email]['word_count'] += 1
emails[current_email]['words'].append(current_word)### store words in email
else:
if current_word:
if flag==1 and current_word not in flagged_words:
flagged_words.append(current_word)
words[current_word]['flag'] = flag ### denote if current word is a word specified by the user list
words[current_word]['spam_count'] += current_spamcount ### update spam count for current word
words[current_word]['ham_count'] += current_hamcount ### update ham count for current word
current_count = count ### set current count
current_spamcount, current_hamcount = 0,0 ### initialize spam and ham wordcount
current_word = word ### set current number
current_email = email ### set current id of email
current_y_true = y_true ### set current spam truth
current_flag = flag ### set current flag
if current_word == word: ### do not forget to output the last word if needed!
emails[current_email]['word_count'] += 1
emails[current_email]['words'].append(current_word)### store words in email
words[current_word]['flag'] = flag ### denote if current word is a word specified by the user list
words[current_word]['spam_count'] += current_spamcount ### update spam count for current word
words[current_word]['ham_count'] += current_hamcount ### update ham count for current word
#Calculate stats for entire corpus
prior_spam= sum_spamrecords/sum_records
prior_ham=sum_hamrecords/sum_records
vocab_count=len(words)#number of unique words in the total vocabulary
for k,word in words.iteritems():
#These versions calculate conditional probabilities WITH Laplace smoothing.
#word['p_spam']=(word['spam_count']+1)/(spam_word_count+vocab_count)
#word['p_ham']=(word['ham_count']+1)/(ham_word_count+vocab_count)
#Compute conditional probabilities WITHOUT Laplace smoothing
word['p_spam']=(word['spam_count'])/(sum_spamwords)
word['p_ham']=(word['ham_count'])/(sum_hamwords)
#At this point the model is now trained, and we can use it to make our predictions
print '%30s' %'ID', '%10s' %'TRUTH', '%10s' %'CLASS', '%20s' %'CUMULATIVE ACCURACY'
miss, sample_size = 0,0
for j,email in emails.iteritems():
#Log versions - no longer used
#p_spam=log(prior_spam)
#p_ham=log(prior_ham)
p_spam=prior_spam
p_ham=prior_ham
for word in email['words']:
if word in flagged_words:
try:
#p_spam+=log(words[word]['p_spam']) #Log version - no longer used
p_spam*=words[word]['p_spam']
except ValueError:
pass #This means that words that do not appear in a class will use the class prior
try:
#p_ham+=log(words[word]['p_ham']) #Log version - no longer used
p_ham*=words[word]['p_ham']
except ValueError:
pass
if p_spam>p_ham:
y_pred=1
else:
y_pred=0
y_true = email['y_true']
if y_pred != y_true:
miss+= 1.0
sample_size += 1.0
accuracy = ((sample_size-miss)/sample_size)*100
print '%30s' %email, '%10d' %y_true, '%10d' %y_pred, '%18.2f %%' % accuracy
!chmod +x reducer.py
```
# Run Hadoop MapReduce Streaming
<h2> start up yarn and dfs </h2>
```
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-yarn.sh ### start up yarn
!/usr/local/Cellar/hadoop/2.7.1/sbin/start-dfs.sh ### start up dfs
```
<h2> create folder </h2>
```
!hdfs dfs -mkdir -p /user/venamax ### create hdfs folder
```
<h2> upload enronmail_1h.txt file </h2>
```
!hdfs dfs -put enronemail_1h.txt /user/venamax #### save source data file to hdfs
```
<h2> Hadoop streaming </h2>
```
!hadoop jar hadoop-*streaming*.jar -mapper mapper.py -reducer reducer.py -input numcount.txt -output numcountOutput
```
<h2>HW2.2.1</h2> Using Hadoop MapReduce and your wordcount job (from HW2.2) determine the top-10 occurring tokens (most frequent tokens)
<h2>HW2.3. Multinomial NAIVE BAYES with NO Smoothing</h2>
Using the Enron data from HW1 and Hadoop MapReduce, write a mapper/reducer job(s) that
will both learn Naive Bayes classifier and classify the Enron email messages using the learnt Naive Bayes classifier. Use all white-space delimitted tokens as independent input variables (assume spaces, fullstops, commas as delimiters). Note: for multinomial Naive Bayes, the Pr(X=“assistance”|Y=SPAM) is calculated as follows:
the number of times “assistance” occurs in SPAM labeled documents / the number of words in documents labeled SPAM
E.g., “assistance” occurs 5 times in all of the documents Labeled SPAM, and the length in terms of the number of words in all documents labeled as SPAM (when concatenated) is 1,000. Then Pr(X=“assistance”|Y=SPAM) = 5/1000. Note this is a multinomial estimation of the class conditional for a Naive Bayes Classifier. No smoothing is needed in this HW. Multiplying lots of probabilities, which are between 0 and 1, can result in floating-point underflow. Since log(xy) = log(x) + log(y), it is better to perform all computations by summing logs of probabilities rather than multiplying probabilities. Please pay attention to probabilites that are zero! They will need special attention. Count up how many times you need to process a zero probabilty for each class and report.
Report the performance of your learnt classifier in terms of misclassifcation error rate of your multinomial Naive Bayes Classifier. Plot a histogram of the log posterior probabilities (i.e., Pr(Class|Doc))) for each class over the training set. Summarize what you see.
Error Rate = misclassification rate with respect to a provided set (say training set in this case). It is more formally defined here:
Let DF represent the evalution set in the following:
Err(Model, DF) = |{(X, c(X)) ∈ DF : c(X) != Model(x)}| / |DF|
Where || denotes set cardinality; c(X) denotes the class of the tuple X in DF; and Model(X) denotes the class inferred by the Model “Model”
<h2>HW2.4 Repeat HW2.3 with the following modification: use Laplace plus-one smoothing. </h2>
Compare the misclassifcation error rates for 2.3 versus 2.4 and explain the differences.
For a quick reference on the construction of the Multinomial NAIVE BAYES classifier that you will code,
please consult the "Document Classification" section of the following wikipedia page:
https://en.wikipedia.org/wiki/Naive_Bayes_classifier#Document_classification
OR the original paper by the curators of the Enron email data:
http://www.aueb.gr/users/ion/docs/ceas2006_paper.pdf
<h2>HW2.5. Repeat HW2.4. This time when modeling and classification ignore tokens with a frequency of less than three (3) in the training set. </h2>How does it affect the misclassifcation error of learnt naive multinomial Bayesian Classifier on the training dataset:
<h2>HW2.6 Benchmark your code with the Python SciKit-Learn implementation of the multinomial Naive Bayes algorithm</h2>
It always a good idea to benchmark your solutions against publicly available libraries such as SciKit-Learn, The Machine Learning toolkit available in Python. In this exercise, we benchmark ourselves against the SciKit-Learn implementation of multinomial Naive Bayes. For more information on this implementation see: http://scikit-learn.org/stable/modules/naive_bayes.html more
In this exercise, please complete the following:
— Run the Multinomial Naive Bayes algorithm (using default settings) from SciKit-Learn over the same training data used in HW2.5 and report the misclassification error (please note some data preparation might be needed to get the Multinomial Naive Bayes algorithm from SkiKit-Learn to run over this dataset)
- Prepare a table to present your results, where rows correspond to approach used (SkiKit-Learn versus your Hadoop implementation) and the column presents the training misclassification error
— Explain/justify any differences in terms of training error rates over the dataset in HW2.5 between your Multinomial Naive Bayes implementation (in Map Reduce) versus the Multinomial Naive Bayes implementation in SciKit-Learn
<h2>HHW 2.6.1 OPTIONAL (note this exercise is a stretch HW and optional)</h2>
— Run the Bernoulli Naive Bayes algorithm from SciKit-Learn (using default settings) over the same training data used in HW2.6 and report the misclassification error
- Discuss the performance differences in terms of misclassification error rates over the dataset in HW2.5 between the Multinomial Naive Bayes implementation in SciKit-Learn with the Bernoulli Naive Bayes implementation in SciKit-Learn. Why such big differences. Explain.
Which approach to Naive Bayes would you recommend for SPAM detection? Justify your selection.
<h2>HW2.7 OPTIONAL (note this exercise is a stretch HW and optional)</h2>
The Enron SPAM data in the following folder enron1-Training-Data-RAW is in raw text form (with subfolders for SPAM and HAM that contain raw email messages in the following form:
--- Line 1 contains the subject
--- The remaining lines contain the body of the email message.
In Python write a script to produce a TSV file called train-Enron-1.txt that has a similar format as the enronemail_1h.txt that you have been using so far. Please pay attend to funky characters and tabs. Check your resulting formated email data in Excel and in Python (e.g., count up the number of fields in each row; the number of SPAM mails and the number of HAM emails). Does each row correspond to an email record with four values? Note: use "NA" to denote empty field values.
<h2>HW2.8 OPTIONAL</h2>
Using Hadoop Map-Reduce write job(s) to perform the following:
-- Train a multinomial Naive Bayes Classifier with Laplace plus one smoothing using the data extracted in HW2.7 (i.e., train-Enron-1.txt). Use all white-space delimitted tokens as independent input variables (assume spaces, fullstops, commas as delimiters). Drop tokens with a frequency of less than three (3).
-- Test the learnt classifier using enronemail_1h.txt and report the misclassification error rate. Remember to use all white-space delimitted tokens as independent input variables (assume spaces, fullstops, commas as delimiters). How do we treat tokens in the test set that do not appear in the training set?
<h2>HW2.8.1 OPTIONAL</h2>
— Run both the Multinomial Naive Bayes and the Bernoulli Naive Bayes algorithms from SciKit-Learn (using default settings) over the same training data used in HW2.8 and report the misclassification error on both the training set and the testing set
- Prepare a table to present your results, where rows correspond to approach used (SciKit-Learn Multinomial NB; SciKit-Learn Bernouili NB; Your Hadoop implementation) and the columns presents the training misclassification error, and the misclassification error on the test data set
- Discuss the performance differences in terms of misclassification error rates over the test and training datasets by the different implementations. Which approch (Bernouili versus Multinomial) would you recommend for SPAM detection? Justify your selection.
<h2>=====================
END OF HOMEWORK</h2>
| github_jupyter |
# 个性化推荐
本项目使用文本卷积神经网络,并使用[`MovieLens`](https://grouplens.org/datasets/movielens/)数据集完成电影推荐的任务。
推荐系统在日常的网络应用中无处不在,比如网上购物、网上买书、新闻app、社交网络、音乐网站、电影网站等等等等,有人的地方就有推荐。根据个人的喜好,相同喜好人群的习惯等信息进行个性化的内容推荐。比如打开新闻类的app,因为有了个性化的内容,每个人看到的新闻首页都是不一样的。
这当然是很有用的,在信息爆炸的今天,获取信息的途径和方式多种多样,人们花费时间最多的不再是去哪获取信息,而是要在众多的信息中寻找自己感兴趣的,这就是信息超载问题。为了解决这个问题,推荐系统应运而生。
协同过滤是推荐系统应用较广泛的技术,该方法搜集用户的历史记录、个人喜好等信息,计算与其他用户的相似度,利用相似用户的评价来预测目标用户对特定项目的喜好程度。优点是会给用户推荐未浏览过的项目,缺点呢,对于新用户来说,没有任何与商品的交互记录和个人喜好等信息,存在冷启动问题,导致模型无法找到相似的用户或商品。
为了解决冷启动的问题,通常的做法是对于刚注册的用户,要求用户先选择自己感兴趣的话题、群组、商品、性格、喜欢的音乐类型等信息,比如豆瓣FM:
<img src="assets/IMG_6242_300.PNG"/>
## 下载数据集
运行下面代码把[`数据集`](http://files.grouplens.org/datasets/movielens/ml-1m.zip)下载下来
```
import pandas as pd
from sklearn.model_selection import train_test_split
import numpy as np
from collections import Counter
import tensorflow as tf
import os
import pickle
import re
from tensorflow.python.ops import math_ops
from urllib.request import urlretrieve
from os.path import isfile, isdir
from tqdm import tqdm
import zipfile
import hashlib
def _unzip(save_path, _, database_name, data_path):
"""
Unzip wrapper with the same interface as _ungzip
:param save_path: The path of the gzip files
:param database_name: Name of database
:param data_path: Path to extract to
:param _: HACK - Used to have to same interface as _ungzip
"""
print('Extracting {}...'.format(database_name))
with zipfile.ZipFile(save_path) as zf:
zf.extractall(data_path)
def download_extract(database_name, data_path):
"""
Download and extract database
:param database_name: Database name
"""
DATASET_ML1M = 'ml-1m'
if database_name == DATASET_ML1M:
url = 'http://files.grouplens.org/datasets/movielens/ml-1m.zip'
hash_code = 'c4d9eecfca2ab87c1945afe126590906'
extract_path = os.path.join(data_path, 'ml-1m')
save_path = os.path.join(data_path, 'ml-1m.zip')
extract_fn = _unzip
if os.path.exists(extract_path):
print('Found {} Data'.format(database_name))
return
if not os.path.exists(data_path):
os.makedirs(data_path)
if not os.path.exists(save_path):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='Downloading {}'.format(database_name)) as pbar:
urlretrieve(
url,
save_path,
pbar.hook)
assert hashlib.md5(open(save_path, 'rb').read()).hexdigest() == hash_code, \
'{} file is corrupted. Remove the file and try again.'.format(save_path)
os.makedirs(extract_path)
try:
extract_fn(save_path, extract_path, database_name, data_path)
except Exception as err:
shutil.rmtree(extract_path) # Remove extraction folder if there is an error
raise err
print('Done.')
# Remove compressed data
# os.remove(save_path)
class DLProgress(tqdm):
"""
Handle Progress Bar while Downloading
"""
last_block = 0
def hook(self, block_num=1, block_size=1, total_size=None):
"""
A hook function that will be called once on establishment of the network connection and
once after each block read thereafter.
:param block_num: A count of blocks transferred so far
:param block_size: Block size in bytes
:param total_size: The total size of the file. This may be -1 on older FTP servers which do not return
a file size in response to a retrieval request.
"""
self.total = total_size
self.update((block_num - self.last_block) * block_size)
self.last_block = block_num
data_dir = './'
download_extract('ml-1m', data_dir)
```
## 先来看看数据
本项目使用的是MovieLens 1M 数据集,包含6000个用户在近4000部电影上的1亿条评论。
数据集分为三个文件:用户数据users.dat,电影数据movies.dat和评分数据ratings.dat。
### 用户数据
分别有用户ID、性别、年龄、职业ID和邮编等字段。
数据中的格式:UserID::Gender::Age::Occupation::Zip-code
- Gender is denoted by a "M" for male and "F" for female
- Age is chosen from the following ranges:
* 1: "Under 18"
* 18: "18-24"
* 25: "25-34"
* 35: "35-44"
* 45: "45-49"
* 50: "50-55"
* 56: "56+"
- Occupation is chosen from the following choices:
* 0: "other" or not specified
* 1: "academic/educator"
* 2: "artist"
* 3: "clerical/admin"
* 4: "college/grad student"
* 5: "customer service"
* 6: "doctor/health care"
* 7: "executive/managerial"
* 8: "farmer"
* 9: "homemaker"
* 10: "K-12 student"
* 11: "lawyer"
* 12: "programmer"
* 13: "retired"
* 14: "sales/marketing"
* 15: "scientist"
* 16: "self-employed"
* 17: "technician/engineer"
* 18: "tradesman/craftsman"
* 19: "unemployed"
* 20: "writer"
```
users_title = ['UserID', 'Gender', 'Age', 'OccupationID', 'Zip-code']
users = pd.read_table('./ml-1m/users.dat', sep='::', header=None, names=users_title, engine = 'python')
users.head()
```
可以看出UserID、Gender、Age和Occupation都是类别字段,其中邮编字段是我们不使用的。
### 电影数据
分别有电影ID、电影名和电影风格等字段。
数据中的格式:MovieID::Title::Genres
- Titles are identical to titles provided by the IMDB (including
year of release)
- Genres are pipe-separated and are selected from the following genres:
* Action
* Adventure
* Animation
* Children's
* Comedy
* Crime
* Documentary
* Drama
* Fantasy
* Film-Noir
* Horror
* Musical
* Mystery
* Romance
* Sci-Fi
* Thriller
* War
* Western
```
movies_title = ['MovieID', 'Title', 'Genres']
movies = pd.read_table('./ml-1m/movies.dat', sep='::', header=None, names=movies_title, engine = 'python')
movies.head()
```
MovieID是类别字段,Title是文本,Genres也是类别字段
### 评分数据
分别有用户ID、电影ID、评分和时间戳等字段。
数据中的格式:UserID::MovieID::Rating::Timestamp
- UserIDs range between 1 and 6040
- MovieIDs range between 1 and 3952
- Ratings are made on a 5-star scale (whole-star ratings only)
- Timestamp is represented in seconds since the epoch as returned by time(2)
- Each user has at least 20 ratings
```
ratings_title = ['UserID','MovieID', 'Rating', 'timestamps']
ratings = pd.read_table('./ml-1m/ratings.dat', sep='::', header=None, names=ratings_title, engine = 'python')
ratings.head()
```
评分字段Rating就是我们要学习的targets,时间戳字段我们不使用。
## 来说说数据预处理
- UserID、Occupation和MovieID不用变。
- Gender字段:需要将‘F’和‘M’转换成0和1。
- Age字段:要转成7个连续数字0~6。
- Genres字段:是分类字段,要转成数字。首先将Genres中的类别转成字符串到数字的字典,然后再将每个电影的Genres字段转成数字列表,因为有些电影是多个Genres的组合。
- Title字段:处理方式跟Genres字段一样,首先创建文本到数字的字典,然后将Title中的描述转成数字的列表。另外Title中的年份也需要去掉。
- Genres和Title字段需要将长度统一,这样在神经网络中方便处理。空白部分用‘< PAD >’对应的数字填充。
## 实现数据预处理
```
def load_data():
"""
Load Dataset from File
"""
#读取User数据
users_title = ['UserID', 'Gender', 'Age', 'JobID', 'Zip-code']
users = pd.read_table('./ml-1m/users.dat', sep='::', header=None, names=users_title, engine = 'python')
users = users.filter(regex='UserID|Gender|Age|JobID')
users_orig = users.values
#改变User数据中性别和年龄
gender_map = {'F':0, 'M':1}
users['Gender'] = users['Gender'].map(gender_map)
age_map = {val:ii for ii,val in enumerate(set(users['Age']))}
users['Age'] = users['Age'].map(age_map)
#读取Movie数据集
movies_title = ['MovieID', 'Title', 'Genres']
movies = pd.read_table('./ml-1m/movies.dat', sep='::', header=None, names=movies_title, engine = 'python')
movies_orig = movies.values
#将Title中的年份去掉
pattern = re.compile(r'^(.*)\((\d+)\)$')
title_map = {val:pattern.match(val).group(1) for ii,val in enumerate(set(movies['Title']))}
movies['Title'] = movies['Title'].map(title_map)
#电影类型转数字字典
genres_set = set()
for val in movies['Genres'].str.split('|'):
genres_set.update(val)
genres_set.add('<PAD>')
genres2int = {val:ii for ii, val in enumerate(genres_set)}
#将电影类型转成等长数字列表,长度是18
genres_map = {val:[genres2int[row] for row in val.split('|')] for ii,val in enumerate(set(movies['Genres']))}
for key in genres_map:
for cnt in range(max(genres2int.values()) - len(genres_map[key])):
genres_map[key].insert(len(genres_map[key]) + cnt,genres2int['<PAD>'])
movies['Genres'] = movies['Genres'].map(genres_map)
#电影Title转数字字典
title_set = set()
for val in movies['Title'].str.split():
title_set.update(val)
title_set.add('<PAD>')
title2int = {val:ii for ii, val in enumerate(title_set)}
#将电影Title转成等长数字列表,长度是15
title_count = 15
title_map = {val:[title2int[row] for row in val.split()] for ii,val in enumerate(set(movies['Title']))}
for key in title_map:
for cnt in range(title_count - len(title_map[key])):
title_map[key].insert(len(title_map[key]) + cnt,title2int['<PAD>'])
movies['Title'] = movies['Title'].map(title_map)
#读取评分数据集
ratings_title = ['UserID','MovieID', 'ratings', 'timestamps']
ratings = pd.read_table('./ml-1m/ratings.dat', sep='::', header=None, names=ratings_title, engine = 'python')
ratings = ratings.filter(regex='UserID|MovieID|ratings')
#合并三个表
data = pd.merge(pd.merge(ratings, users), movies)
#将数据分成X和y两张表
target_fields = ['ratings']
features_pd, targets_pd = data.drop(target_fields, axis=1), data[target_fields]
features = features_pd.values
targets_values = targets_pd.values
return title_count, title_set, genres2int, features, targets_values, ratings, users, movies, data, movies_orig, users_orig
```
### 加载数据并保存到本地
- title_count:Title字段的长度(15)
- title_set:Title文本的集合
- genres2int:电影类型转数字的字典
- features:是输入X
- targets_values:是学习目标y
- ratings:评分数据集的Pandas对象
- users:用户数据集的Pandas对象
- movies:电影数据的Pandas对象
- data:三个数据集组合在一起的Pandas对象
- movies_orig:没有做数据处理的原始电影数据
- users_orig:没有做数据处理的原始用户数据
```
title_count, title_set, genres2int, features, targets_values, ratings, users, movies, data, movies_orig, users_orig = load_data()
pickle.dump((title_count, title_set, genres2int, features, targets_values, ratings, users, movies, data, movies_orig, users_orig), open('preprocess.p', 'wb'))
```
### 预处理后的数据
```
users.head()
movies.head()
movies.values[0]
```
### 从本地读取数据
```
title_count, title_set, genres2int, features, targets_values, ratings, users, movies, data, movies_orig, users_orig = pickle.load(open('preprocess.p', mode='rb'))
```
## 模型设计
<img src="assets/model.001.jpeg"/>
通过研究数据集中的字段类型,我们发现有一些是类别字段,通常的处理是将这些字段转成one hot编码,但是像UserID、MovieID这样的字段就会变成非常的稀疏,输入的维度急剧膨胀,这是我们不愿意见到的,毕竟我这小笔记本不像大厂动辄能处理数以亿计维度的输入:)
所以在预处理数据时将这些字段转成了数字,我们用这个数字当做嵌入矩阵的索引,在网络的第一层使用了嵌入层,维度是(N,32)和(N,16)。
电影类型的处理要多一步,有时一个电影有多个电影类型,这样从嵌入矩阵索引出来是一个(n,32)的矩阵,因为有多个类型嘛,我们要将这个矩阵求和,变成(1,32)的向量。
电影名的处理比较特殊,没有使用循环神经网络,而是用了文本卷积网络,下文会进行说明。
从嵌入层索引出特征以后,将各特征传入全连接层,将输出再次传入全连接层,最终分别得到(1,200)的用户特征和电影特征两个特征向量。
我们的目的就是要训练出用户特征和电影特征,在实现推荐功能时使用。得到这两个特征以后,就可以选择任意的方式来拟合评分了。我使用了两种方式,一个是上图中画出的将两个特征做向量乘法,将结果与真实评分做回归,采用MSE优化损失。因为本质上这是一个回归问题,另一种方式是,将两个特征作为输入,再次传入全连接层,输出一个值,将输出值回归到真实评分,采用MSE优化损失。
实际上第二个方式的MSE loss在0.8附近,第一个方式在1附近,5次迭代的结果。
## 文本卷积网络
网络看起来像下面这样
<img src="assets/text_cnn.png"/>
图片来自Kim Yoon的论文:[`Convolutional Neural Networks for Sentence Classification`](https://arxiv.org/abs/1408.5882)
将卷积神经网络用于文本的文章建议你阅读[`Understanding Convolutional Neural Networks for NLP`](http://www.wildml.com/2015/11/understanding-convolutional-neural-networks-for-nlp/)
网络的第一层是词嵌入层,由每一个单词的嵌入向量组成的嵌入矩阵。下一层使用多个不同尺寸(窗口大小)的卷积核在嵌入矩阵上做卷积,窗口大小指的是每次卷积覆盖几个单词。这里跟对图像做卷积不太一样,图像的卷积通常用2x2、3x3、5x5之类的尺寸,而文本卷积要覆盖整个单词的嵌入向量,所以尺寸是(单词数,向量维度),比如每次滑动3个,4个或者5个单词。第三层网络是max pooling得到一个长向量,最后使用dropout做正则化,最终得到了电影Title的特征。
## 辅助函数
```
import tensorflow as tf
import os
import pickle
def save_params(params):
"""
Save parameters to file
"""
pickle.dump(params, open('params.p', 'wb'))
def load_params():
"""
Load parameters from file
"""
return pickle.load(open('params.p', mode='rb'))
```
## 编码实现
```
#嵌入矩阵的维度
embed_dim = 32
#用户ID个数
uid_max = max(features.take(0,1)) + 1 # 6040
#性别个数
gender_max = max(features.take(2,1)) + 1 # 1 + 1 = 2
#年龄类别个数
age_max = max(features.take(3,1)) + 1 # 6 + 1 = 7
#职业个数
job_max = max(features.take(4,1)) + 1# 20 + 1 = 21
#电影ID个数
movie_id_max = max(features.take(1,1)) + 1 # 3952
#电影类型个数
movie_categories_max = max(genres2int.values()) + 1 # 18 + 1 = 19
#电影名单词个数
movie_title_max = len(title_set) # 5216
#对电影类型嵌入向量做加和操作的标志,考虑过使用mean做平均,但是没实现mean
combiner = "sum"
#电影名长度
sentences_size = title_count # = 15
#文本卷积滑动窗口,分别滑动2, 3, 4, 5个单词
window_sizes = {2, 3, 4, 5}
#文本卷积核数量
filter_num = 8
#电影ID转下标的字典,数据集中电影ID跟下标不一致,比如第5行的数据电影ID不一定是5
movieid2idx = {val[0]:i for i, val in enumerate(movies.values)}
```
### 超参
```
# Number of Epochs
num_epochs = 5
# Batch Size
batch_size = 256
dropout_keep = 0.5
# Learning Rate
learning_rate = 0.0001
# Show stats for every n number of batches
show_every_n_batches = 20
save_dir = './save'
```
### 输入
定义输入的占位符
```
def get_inputs():
uid = tf.placeholder(tf.int32, [None, 1], name="uid")
user_gender = tf.placeholder(tf.int32, [None, 1], name="user_gender")
user_age = tf.placeholder(tf.int32, [None, 1], name="user_age")
user_job = tf.placeholder(tf.int32, [None, 1], name="user_job")
movie_id = tf.placeholder(tf.int32, [None, 1], name="movie_id")
movie_categories = tf.placeholder(tf.int32, [None, 18], name="movie_categories")
movie_titles = tf.placeholder(tf.int32, [None, 15], name="movie_titles")
targets = tf.placeholder(tf.int32, [None, 1], name="targets")
LearningRate = tf.placeholder(tf.float32, name = "LearningRate")
dropout_keep_prob = tf.placeholder(tf.float32, name = "dropout_keep_prob")
return uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, LearningRate, dropout_keep_prob
```
## 构建神经网络
#### 定义User的嵌入矩阵
```
def get_user_embedding(uid, user_gender, user_age, user_job):
with tf.name_scope("user_embedding"):
uid_embed_matrix = tf.Variable(tf.random_uniform([uid_max, embed_dim], -1, 1), name = "uid_embed_matrix")
uid_embed_layer = tf.nn.embedding_lookup(uid_embed_matrix, uid, name = "uid_embed_layer")
gender_embed_matrix = tf.Variable(tf.random_uniform([gender_max, embed_dim // 2], -1, 1), name= "gender_embed_matrix")
gender_embed_layer = tf.nn.embedding_lookup(gender_embed_matrix, user_gender, name = "gender_embed_layer")
age_embed_matrix = tf.Variable(tf.random_uniform([age_max, embed_dim // 2], -1, 1), name="age_embed_matrix")
age_embed_layer = tf.nn.embedding_lookup(age_embed_matrix, user_age, name="age_embed_layer")
job_embed_matrix = tf.Variable(tf.random_uniform([job_max, embed_dim // 2], -1, 1), name = "job_embed_matrix")
job_embed_layer = tf.nn.embedding_lookup(job_embed_matrix, user_job, name = "job_embed_layer")
return uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer
```
#### 将User的嵌入矩阵一起全连接生成User的特征
```
def get_user_feature_layer(uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer):
with tf.name_scope("user_fc"):
#第一层全连接
uid_fc_layer = tf.layers.dense(uid_embed_layer, embed_dim, name = "uid_fc_layer", activation=tf.nn.relu)
gender_fc_layer = tf.layers.dense(gender_embed_layer, embed_dim, name = "gender_fc_layer", activation=tf.nn.relu)
age_fc_layer = tf.layers.dense(age_embed_layer, embed_dim, name ="age_fc_layer", activation=tf.nn.relu)
job_fc_layer = tf.layers.dense(job_embed_layer, embed_dim, name = "job_fc_layer", activation=tf.nn.relu)
#第二层全连接
user_combine_layer = tf.concat([uid_fc_layer, gender_fc_layer, age_fc_layer, job_fc_layer], 2) #(?, 1, 128)
user_combine_layer = tf.contrib.layers.fully_connected(user_combine_layer, 200, tf.tanh) #(?, 1, 200)
user_combine_layer_flat = tf.reshape(user_combine_layer, [-1, 200])
return user_combine_layer, user_combine_layer_flat
```
#### 定义Movie ID的嵌入矩阵
```
def get_movie_id_embed_layer(movie_id):
with tf.name_scope("movie_embedding"):
movie_id_embed_matrix = tf.Variable(tf.random_uniform([movie_id_max, embed_dim], -1, 1), name = "movie_id_embed_matrix")
movie_id_embed_layer = tf.nn.embedding_lookup(movie_id_embed_matrix, movie_id, name = "movie_id_embed_layer")
return movie_id_embed_layer
```
#### 对电影类型的多个嵌入向量做加和
```
def get_movie_categories_layers(movie_categories):
with tf.name_scope("movie_categories_layers"):
movie_categories_embed_matrix = tf.Variable(tf.random_uniform([movie_categories_max, embed_dim], -1, 1), name = "movie_categories_embed_matrix")
movie_categories_embed_layer = tf.nn.embedding_lookup(movie_categories_embed_matrix, movie_categories, name = "movie_categories_embed_layer")
if combiner == "sum":
movie_categories_embed_layer = tf.reduce_sum(movie_categories_embed_layer, axis=1, keep_dims=True)
# elif combiner == "mean":
return movie_categories_embed_layer
```
#### Movie Title的文本卷积网络实现
```
def get_movie_cnn_layer(movie_titles):
#从嵌入矩阵中得到电影名对应的各个单词的嵌入向量
with tf.name_scope("movie_embedding"):
movie_title_embed_matrix = tf.Variable(tf.random_uniform([movie_title_max, embed_dim], -1, 1), name = "movie_title_embed_matrix")
movie_title_embed_layer = tf.nn.embedding_lookup(movie_title_embed_matrix, movie_titles, name = "movie_title_embed_layer")
movie_title_embed_layer_expand = tf.expand_dims(movie_title_embed_layer, -1)
#对文本嵌入层使用不同尺寸的卷积核做卷积和最大池化
pool_layer_lst = []
for window_size in window_sizes:
with tf.name_scope("movie_txt_conv_maxpool_{}".format(window_size)):
filter_weights = tf.Variable(tf.truncated_normal([window_size, embed_dim, 1, filter_num],stddev=0.1),name = "filter_weights")
filter_bias = tf.Variable(tf.constant(0.1, shape=[filter_num]), name="filter_bias")
conv_layer = tf.nn.conv2d(movie_title_embed_layer_expand, filter_weights, [1,1,1,1], padding="VALID", name="conv_layer")
relu_layer = tf.nn.relu(tf.nn.bias_add(conv_layer,filter_bias), name ="relu_layer")
maxpool_layer = tf.nn.max_pool(relu_layer, [1,sentences_size - window_size + 1 ,1,1], [1,1,1,1], padding="VALID", name="maxpool_layer")
pool_layer_lst.append(maxpool_layer)
#Dropout层
with tf.name_scope("pool_dropout"):
pool_layer = tf.concat(pool_layer_lst, 3, name ="pool_layer")
max_num = len(window_sizes) * filter_num
pool_layer_flat = tf.reshape(pool_layer , [-1, 1, max_num], name = "pool_layer_flat")
dropout_layer = tf.nn.dropout(pool_layer_flat, dropout_keep_prob, name = "dropout_layer")
return pool_layer_flat, dropout_layer
```
#### 将Movie的各个层一起做全连接
```
def get_movie_feature_layer(movie_id_embed_layer, movie_categories_embed_layer, dropout_layer):
with tf.name_scope("movie_fc"):
#第一层全连接
movie_id_fc_layer = tf.layers.dense(movie_id_embed_layer, embed_dim, name = "movie_id_fc_layer", activation=tf.nn.relu)
movie_categories_fc_layer = tf.layers.dense(movie_categories_embed_layer, embed_dim, name = "movie_categories_fc_layer", activation=tf.nn.relu)
#第二层全连接
movie_combine_layer = tf.concat([movie_id_fc_layer, movie_categories_fc_layer, dropout_layer], 2) #(?, 1, 96)
movie_combine_layer = tf.contrib.layers.fully_connected(movie_combine_layer, 200, tf.tanh) #(?, 1, 200)
movie_combine_layer_flat = tf.reshape(movie_combine_layer, [-1, 200])
return movie_combine_layer, movie_combine_layer_flat
```
## 构建计算图
```
tf.reset_default_graph()
train_graph = tf.Graph()
with train_graph.as_default():
#获取输入占位符
uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, lr, dropout_keep_prob = get_inputs()
#获取User的4个嵌入向量
uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer = get_user_embedding(uid, user_gender, user_age, user_job)
#得到用户特征
user_combine_layer, user_combine_layer_flat = get_user_feature_layer(uid_embed_layer, gender_embed_layer, age_embed_layer, job_embed_layer)
#获取电影ID的嵌入向量
movie_id_embed_layer = get_movie_id_embed_layer(movie_id)
#获取电影类型的嵌入向量
movie_categories_embed_layer = get_movie_categories_layers(movie_categories)
#获取电影名的特征向量
pool_layer_flat, dropout_layer = get_movie_cnn_layer(movie_titles)
#得到电影特征
movie_combine_layer, movie_combine_layer_flat = get_movie_feature_layer(movie_id_embed_layer,
movie_categories_embed_layer,
dropout_layer)
#计算出评分,要注意两个不同的方案,inference的名字(name值)是不一样的,后面做推荐时要根据name取得tensor
with tf.name_scope("inference"):
#将用户特征和电影特征作为输入,经过全连接,输出一个值的方案
# inference_layer = tf.concat([user_combine_layer_flat, movie_combine_layer_flat], 1) #(?, 200)
# inference = tf.layers.dense(inference_layer, 1,
# kernel_initializer=tf.truncated_normal_initializer(stddev=0.01),
# kernel_regularizer=tf.nn.l2_loss, name="inference")
#简单的将用户特征和电影特征做矩阵乘法得到一个预测评分
# inference = tf.matmul(user_combine_layer_flat, tf.transpose(movie_combine_layer_flat))
inference = tf.reduce_sum(user_combine_layer_flat * movie_combine_layer_flat, axis=1)
inference = tf.expand_dims(inference, axis=1)
with tf.name_scope("loss"):
# MSE损失,将计算值回归到评分
cost = tf.losses.mean_squared_error(targets, inference )
loss = tf.reduce_mean(cost)
# 优化损失
# train_op = tf.train.AdamOptimizer(lr).minimize(loss) #cost
global_step = tf.Variable(0, name="global_step", trainable=False)
optimizer = tf.train.AdamOptimizer(lr)
gradients = optimizer.compute_gradients(loss) #cost
train_op = optimizer.apply_gradients(gradients, global_step=global_step)
inference
```
## 取得batch
```
def get_batches(Xs, ys, batch_size):
for start in range(0, len(Xs), batch_size):
end = min(start + batch_size, len(Xs))
yield Xs[start:end], ys[start:end]
```
## 训练网络
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import time
import datetime
losses = {'train':[], 'test':[]}
with tf.Session(graph=train_graph) as sess:
#搜集数据给tensorBoard用
# Keep track of gradient values and sparsity
grad_summaries = []
for g, v in gradients:
if g is not None:
grad_hist_summary = tf.summary.histogram("{}/grad/hist".format(v.name.replace(':', '_')), g)
sparsity_summary = tf.summary.scalar("{}/grad/sparsity".format(v.name.replace(':', '_')), tf.nn.zero_fraction(g))
grad_summaries.append(grad_hist_summary)
grad_summaries.append(sparsity_summary)
grad_summaries_merged = tf.summary.merge(grad_summaries)
# Output directory for models and summaries
timestamp = str(int(time.time()))
out_dir = os.path.abspath(os.path.join(os.path.curdir, "runs", timestamp))
print("Writing to {}\n".format(out_dir))
# Summaries for loss and accuracy
loss_summary = tf.summary.scalar("loss", loss)
# Train Summaries
train_summary_op = tf.summary.merge([loss_summary, grad_summaries_merged])
train_summary_dir = os.path.join(out_dir, "summaries", "train")
train_summary_writer = tf.summary.FileWriter(train_summary_dir, sess.graph)
# Inference summaries
inference_summary_op = tf.summary.merge([loss_summary])
inference_summary_dir = os.path.join(out_dir, "summaries", "inference")
inference_summary_writer = tf.summary.FileWriter(inference_summary_dir, sess.graph)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
for epoch_i in range(num_epochs):
#将数据集分成训练集和测试集,随机种子不固定
train_X,test_X, train_y, test_y = train_test_split(features,
targets_values,
test_size = 0.2,
random_state = 0)
train_batches = get_batches(train_X, train_y, batch_size)
test_batches = get_batches(test_X, test_y, batch_size)
#训练的迭代,保存训练损失
for batch_i in range(len(train_X) // batch_size):
x, y = next(train_batches)
categories = np.zeros([batch_size, 18])
for i in range(batch_size):
categories[i] = x.take(6,1)[i]
titles = np.zeros([batch_size, sentences_size])
for i in range(batch_size):
titles[i] = x.take(5,1)[i]
feed = {
uid: np.reshape(x.take(0,1), [batch_size, 1]),
user_gender: np.reshape(x.take(2,1), [batch_size, 1]),
user_age: np.reshape(x.take(3,1), [batch_size, 1]),
user_job: np.reshape(x.take(4,1), [batch_size, 1]),
movie_id: np.reshape(x.take(1,1), [batch_size, 1]),
movie_categories: categories, #x.take(6,1)
movie_titles: titles, #x.take(5,1)
targets: np.reshape(y, [batch_size, 1]),
dropout_keep_prob: dropout_keep, #dropout_keep
lr: learning_rate}
step, train_loss, summaries, _ = sess.run([global_step, loss, train_summary_op, train_op], feed) #cost
losses['train'].append(train_loss)
train_summary_writer.add_summary(summaries, step) #
# Show every <show_every_n_batches> batches
if (epoch_i * (len(train_X) // batch_size) + batch_i) % show_every_n_batches == 0:
time_str = datetime.datetime.now().isoformat()
print('{}: Epoch {:>3} Batch {:>4}/{} train_loss = {:.3f}'.format(
time_str,
epoch_i,
batch_i,
(len(train_X) // batch_size),
train_loss))
#使用测试数据的迭代
for batch_i in range(len(test_X) // batch_size):
x, y = next(test_batches)
categories = np.zeros([batch_size, 18])
for i in range(batch_size):
categories[i] = x.take(6,1)[i]
titles = np.zeros([batch_size, sentences_size])
for i in range(batch_size):
titles[i] = x.take(5,1)[i]
feed = {
uid: np.reshape(x.take(0,1), [batch_size, 1]),
user_gender: np.reshape(x.take(2,1), [batch_size, 1]),
user_age: np.reshape(x.take(3,1), [batch_size, 1]),
user_job: np.reshape(x.take(4,1), [batch_size, 1]),
movie_id: np.reshape(x.take(1,1), [batch_size, 1]),
movie_categories: categories, #x.take(6,1)
movie_titles: titles, #x.take(5,1)
targets: np.reshape(y, [batch_size, 1]),
dropout_keep_prob: 1,
lr: learning_rate}
step, test_loss, summaries = sess.run([global_step, loss, inference_summary_op], feed) #cost
#保存测试损失
losses['test'].append(test_loss)
inference_summary_writer.add_summary(summaries, step) #
time_str = datetime.datetime.now().isoformat()
if (epoch_i * (len(test_X) // batch_size) + batch_i) % show_every_n_batches == 0:
print('{}: Epoch {:>3} Batch {:>4}/{} test_loss = {:.3f}'.format(
time_str,
epoch_i,
batch_i,
(len(test_X) // batch_size),
test_loss))
# Save Model
saver.save(sess, save_dir) #, global_step=epoch_i
print('Model Trained and Saved')
```
## 在 TensorBoard 中查看可视化结果
tensorboard --logdir /PATH_TO_CODE/runs/1513402825/summaries/
<img src="assets/loss.png"/>
## 保存参数
保存`save_dir` 在生成预测时使用。
```
save_params((save_dir))
load_dir = load_params()
```
## 显示训练Loss
```
plt.plot(losses['train'], label='Training loss')
plt.legend()
_ = plt.ylim()
```
## 显示测试Loss
迭代次数再增加一些,下降的趋势会明显一些
```
plt.plot(losses['test'], label='Test loss')
plt.legend()
_ = plt.ylim()
```
## 获取 Tensors
使用函数 [`get_tensor_by_name()`](https://www.tensorflow.org/api_docs/python/tf/Graph#get_tensor_by_name)从 `loaded_graph` 中获取tensors,后面的推荐功能要用到。
```
def get_tensors(loaded_graph):
uid = loaded_graph.get_tensor_by_name("uid:0")
user_gender = loaded_graph.get_tensor_by_name("user_gender:0")
user_age = loaded_graph.get_tensor_by_name("user_age:0")
user_job = loaded_graph.get_tensor_by_name("user_job:0")
movie_id = loaded_graph.get_tensor_by_name("movie_id:0")
movie_categories = loaded_graph.get_tensor_by_name("movie_categories:0")
movie_titles = loaded_graph.get_tensor_by_name("movie_titles:0")
targets = loaded_graph.get_tensor_by_name("targets:0")
dropout_keep_prob = loaded_graph.get_tensor_by_name("dropout_keep_prob:0")
lr = loaded_graph.get_tensor_by_name("LearningRate:0")
#两种不同计算预测评分的方案使用不同的name获取tensor inference
# inference = loaded_graph.get_tensor_by_name("inference/inference/BiasAdd:0")
inference = loaded_graph.get_tensor_by_name("inference/ExpandDims:0") # 之前是MatMul:0 因为inference代码修改了 这里也要修改 感谢网友 @清歌 指出问题
movie_combine_layer_flat = loaded_graph.get_tensor_by_name("movie_fc/Reshape:0")
user_combine_layer_flat = loaded_graph.get_tensor_by_name("user_fc/Reshape:0")
return uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, lr, dropout_keep_prob, inference, movie_combine_layer_flat, user_combine_layer_flat
```
## 指定用户和电影进行评分
这部分就是对网络做正向传播,计算得到预测的评分
```
def rating_movie(user_id_val, movie_id_val):
loaded_graph = tf.Graph() #
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
# Get Tensors from loaded model
uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, lr, dropout_keep_prob, inference,_, __ = get_tensors(loaded_graph) #loaded_graph
categories = np.zeros([1, 18])
categories[0] = movies.values[movieid2idx[movie_id_val]][2]
titles = np.zeros([1, sentences_size])
titles[0] = movies.values[movieid2idx[movie_id_val]][1]
feed = {
uid: np.reshape(users.values[user_id_val-1][0], [1, 1]),
user_gender: np.reshape(users.values[user_id_val-1][1], [1, 1]),
user_age: np.reshape(users.values[user_id_val-1][2], [1, 1]),
user_job: np.reshape(users.values[user_id_val-1][3], [1, 1]),
movie_id: np.reshape(movies.values[movieid2idx[movie_id_val]][0], [1, 1]),
movie_categories: categories, #x.take(6,1)
movie_titles: titles, #x.take(5,1)
dropout_keep_prob: 1}
# Get Prediction
inference_val = sess.run([inference], feed)
return (inference_val)
rating_movie(234, 1401)
```
## 生成Movie特征矩阵
将训练好的电影特征组合成电影特征矩阵并保存到本地
```
loaded_graph = tf.Graph() #
movie_matrics = []
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
# Get Tensors from loaded model
uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, lr, dropout_keep_prob, _, movie_combine_layer_flat, __ = get_tensors(loaded_graph) #loaded_graph
for item in movies.values:
categories = np.zeros([1, 18])
categories[0] = item.take(2)
titles = np.zeros([1, sentences_size])
titles[0] = item.take(1)
feed = {
movie_id: np.reshape(item.take(0), [1, 1]),
movie_categories: categories, #x.take(6,1)
movie_titles: titles, #x.take(5,1)
dropout_keep_prob: 1}
movie_combine_layer_flat_val = sess.run([movie_combine_layer_flat], feed)
movie_matrics.append(movie_combine_layer_flat_val)
pickle.dump((np.array(movie_matrics).reshape(-1, 200)), open('movie_matrics.p', 'wb'))
movie_matrics = pickle.load(open('movie_matrics.p', mode='rb'))
movie_matrics = pickle.load(open('movie_matrics.p', mode='rb'))
```
## 生成User特征矩阵
将训练好的用户特征组合成用户特征矩阵并保存到本地
```
loaded_graph = tf.Graph() #
users_matrics = []
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
# Get Tensors from loaded model
uid, user_gender, user_age, user_job, movie_id, movie_categories, movie_titles, targets, lr, dropout_keep_prob, _, __,user_combine_layer_flat = get_tensors(loaded_graph) #loaded_graph
for item in users.values:
feed = {
uid: np.reshape(item.take(0), [1, 1]),
user_gender: np.reshape(item.take(1), [1, 1]),
user_age: np.reshape(item.take(2), [1, 1]),
user_job: np.reshape(item.take(3), [1, 1]),
dropout_keep_prob: 1}
user_combine_layer_flat_val = sess.run([user_combine_layer_flat], feed)
users_matrics.append(user_combine_layer_flat_val)
pickle.dump((np.array(users_matrics).reshape(-1, 200)), open('users_matrics.p', 'wb'))
users_matrics = pickle.load(open('users_matrics.p', mode='rb'))
users_matrics = pickle.load(open('users_matrics.p', mode='rb'))
```
## 开始推荐电影
使用生产的用户特征矩阵和电影特征矩阵做电影推荐
### 推荐同类型的电影
思路是计算当前看的电影特征向量与整个电影特征矩阵的余弦相似度,取相似度最大的top_k个,这里加了些随机选择在里面,保证每次的推荐稍稍有些不同。
```
def recommend_same_type_movie(movie_id_val, top_k = 20):
loaded_graph = tf.Graph() #
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
norm_movie_matrics = tf.sqrt(tf.reduce_sum(tf.square(movie_matrics), 1, keep_dims=True))
normalized_movie_matrics = movie_matrics / norm_movie_matrics
#推荐同类型的电影
probs_embeddings = (movie_matrics[movieid2idx[movie_id_val]]).reshape([1, 200])
probs_similarity = tf.matmul(probs_embeddings, tf.transpose(normalized_movie_matrics))
sim = (probs_similarity.eval())
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
print("您看的电影是:{}".format(movies_orig[movieid2idx[movie_id_val]]))
print("以下是给您的推荐:")
p = np.squeeze(sim)
p[np.argsort(p)[:-top_k]] = 0
p = p / np.sum(p)
results = set()
while len(results) != 5:
c = np.random.choice(3883, 1, p=p)[0]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
recommend_same_type_movie(1401, 20)
```
### 推荐您喜欢的电影
思路是使用用户特征向量与电影特征矩阵计算所有电影的评分,取评分最高的top_k个,同样加了些随机选择部分。
```
def recommend_your_favorite_movie(user_id_val, top_k = 10):
loaded_graph = tf.Graph() #
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
#推荐您喜欢的电影
probs_embeddings = (users_matrics[user_id_val-1]).reshape([1, 200])
probs_similarity = tf.matmul(probs_embeddings, tf.transpose(movie_matrics))
sim = (probs_similarity.eval())
# print(sim.shape)
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
# sim_norm = probs_norm_similarity.eval()
# print((-sim_norm[0]).argsort()[0:top_k])
print("以下是给您的推荐:")
p = np.squeeze(sim)
p[np.argsort(p)[:-top_k]] = 0
p = p / np.sum(p)
results = set()
while len(results) != 5:
c = np.random.choice(3883, 1, p=p)[0]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
recommend_your_favorite_movie(234, 10)
```
### 看过这个电影的人还看了(喜欢)哪些电影
- 首先选出喜欢某个电影的top_k个人,得到这几个人的用户特征向量。
- 然后计算这几个人对所有电影的评分
- 选择每个人评分最高的电影作为推荐
- 同样加入了随机选择
```
import random
def recommend_other_favorite_movie(movie_id_val, top_k = 20):
loaded_graph = tf.Graph() #
with tf.Session(graph=loaded_graph) as sess: #
# Load saved model
loader = tf.train.import_meta_graph(load_dir + '.meta')
loader.restore(sess, load_dir)
probs_movie_embeddings = (movie_matrics[movieid2idx[movie_id_val]]).reshape([1, 200])
probs_user_favorite_similarity = tf.matmul(probs_movie_embeddings, tf.transpose(users_matrics))
favorite_user_id = np.argsort(probs_user_favorite_similarity.eval())[0][-top_k:]
# print(normalized_users_matrics.eval().shape)
# print(probs_user_favorite_similarity.eval()[0][favorite_user_id])
# print(favorite_user_id.shape)
print("您看的电影是:{}".format(movies_orig[movieid2idx[movie_id_val]]))
print("喜欢看这个电影的人是:{}".format(users_orig[favorite_user_id-1]))
probs_users_embeddings = (users_matrics[favorite_user_id-1]).reshape([-1, 200])
probs_similarity = tf.matmul(probs_users_embeddings, tf.transpose(movie_matrics))
sim = (probs_similarity.eval())
# results = (-sim[0]).argsort()[0:top_k]
# print(results)
# print(sim.shape)
# print(np.argmax(sim, 1))
p = np.argmax(sim, 1)
print("喜欢看这个电影的人还喜欢看:")
results = set()
while len(results) != 5:
c = p[random.randrange(top_k)]
results.add(c)
for val in (results):
print(val)
print(movies_orig[val])
return results
recommend_other_favorite_movie(1401, 20)
```
# 结论
以上就是实现的常用的推荐功能,将网络模型作为回归问题进行训练,得到训练好的用户特征矩阵和电影特征矩阵进行推荐。
## 扩展阅读
如果你对个性化推荐感兴趣,以下资料建议你看看:
- [`Understanding Convolutional Neural Networks for NLP`](http://www.wildml.com/2015/11/understanding-convolutional-neural-networks-for-nlp/)
- [`Convolutional Neural Networks for Sentence Classification`](https://github.com/yoonkim/CNN_sentence)
- [`利用TensorFlow实现卷积神经网络做文本分类`](http://www.jianshu.com/p/ed3eac3dcb39?from=singlemessage)
- [`Convolutional Neural Network for Text Classification in Tensorflow`](https://github.com/dennybritz/cnn-text-classification-tf)
- [`SVD Implement Recommendation systems`](https://github.com/songgc/TF-recomm)
今天的分享就到这里,请多指教!
| github_jupyter |
```
import cv2
import os
import numpy as np
import tkinter as tk
from tkinter.filedialog import askopenfilename
#Global Variables
drawing = False #for drawing or not
ix,iy = -1,-1
Rectangle_all = []#for make new Rectangle file
img = np.zeros((512,512,3), np.uint8)#Initialize Images
def draw_Square(event, x,y, flags, param):
#Draw Function
#If you left mouse button down start to follow your mouse and checked drawing rectangle starting point
#and if your's button up end to follow your mouse and drawing last point
global ix,iy, drawing, Rectangle_all
if event == cv2.EVENT_LBUTTONDOWN:
drawing = True
ix, iy = x,y
# elif event == cv2.EVENT_MOUSEMOVE:
# if drawing == True:
# cv2.rectangle(img, (ix,iy),(x,y), (0,0,255), 3)
elif event == cv2.EVENT_LBUTTONUP:
drawing = False
cv2.rectangle(img, (ix, iy), (x,y), (0,0,255), 1)
label = input()
Rectangle_all.append([label, ix, iy, x - ix,y - iy])
def load_image():
global filename, img, Rectangle_all
Rectangle_all = []# Initialize list
tk.Tk().withdraw()
filename = askopenfilename()
if not filename: return None #if doesn't input file out of this function
#Set path to make Data Save folder
#If you want to change another folder, change path of you wants in parents_filename, and child_filename
filenames = filename.split('/')
parents_filename = './data/' + str(filenames[-3])
child_filename = './data/' + str(filenames[-3]) +'/'+ str(filenames[-2])
#Start to make Ground Truth
img = cv2.imread(filename)
cv2.namedWindow('image')
cv2.setMouseCallback('image',draw_Square)
#Until you push ESC, wait for 1 and retrying check
while True:
cv2.imshow('image', img) #Showing update image
k = cv2.waitKey(1) & 0xFF #wait Key
if k == 27: # If you click the ESC, End of make
break
#If directory doesn't exists in path make directory
if not os.path.exists('./data/'):os.mkdir('./data/')
if not os.path.exists(parents_filename):os.mkdir(parents_filename)
if not os.path.exists(child_filename):os.mkdir(child_filename)
#Make File or Initialize label folder
#In label follow this form [label, left up x, left up y, width, height]
print(Rectangle_all) #to check your data, If you don't want to see, take off it
f = open(child_filename +'/' + str(filenames[-1][:-4]) + '.txt', 'w')
for labels in Rectangle_all:
f.write(str(labels) + '\n')
f.close()
cv2.destroyAllWindows()
def quit():
window.destroy()
window.quit()
window=tk.Tk()
button = tk.Button(window, width=15, command = quit , text = "quit")
button2 = tk.Button(window, width=15, command = load_image , text = "file load")
button2.pack()
button.pack()
window.mainloop()
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Better ML Engineering with ML Metadata
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/mlmd/mlmd_tutorial"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/mlmd/mlmd_tutorial.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/tfx/blob/master/docs/tutorials/mlmd/mlmd_tutorial.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
<td><a target="_blank" href="https://storage.googleapis.com/tensorflow_docs/tfx/docs/tutorials/mlmd/mlmd_tutorial.ipynb">
<img width=32px src="https://www.tensorflow.org/images/download_logo_32px.png">Download notebook</a></td>
</td>
</table>
Assume a scenario where you set up a production ML pipeline to classify penguins. The pipeline ingests your training data, trains and evaluates a model, and pushes it to production.
However, when you later try using this model with a larger dataset that contains different kinds of penguins, you observe that your model does not behave as expected and starts classifying the species incorrectly.
At this point, you are interested in knowing:
* What is the most efficient way to debug the model when the only available artifact is the model in production?
* Which training dataset was used to train the model?
* Which training run led to this erroneous model?
* Where are the model evaluation results?
* Where to begin debugging?
[ML Metadata (MLMD)](https://github.com/google/ml-metadata) is a library that leverages the metadata associated with ML models to help you answer these questions and more. A helpful analogy is to think of this metadata as the equivalent of logging in software development. MLMD enables you to reliably track the artifacts and lineage associated with the various components of your ML pipeline.
In this tutorial, you set up a TFX Pipeline to create a model that classifies penguins into three species based on the body mass and the length and depth of their culmens, and the length of their flippers. You then use MLMD to track the lineage of pipeline components.
## TFX Pipelines in Colab
Colab is a lightweight development environment which differs significantly from a production environment. In production, you may have various pipeline components like data ingestion, transformation, model training, run histories, etc. across multiple, distributed systems. For this tutorial, you should be aware that siginificant differences exist in Orchestration and Metadata storage - it is all handled locally within Colab. Learn more about TFX in Colab [here](https://www.tensorflow.org/tfx/tutorials/tfx/components_keras#background).
## Setup
First, we install and import the necessary packages, set up paths, and download data.
### Upgrade Pip
To avoid upgrading Pip in a system when running locally, check to make sure that we're running in Colab. Local systems can of course be upgraded separately.
```
try:
import colab
!pip install --upgrade pip
except:
pass
```
### Install and import TFX
```
!pip install -q -U --use-feature=2020-resolver tfx
```
You must restart the Colab runtime after installing TFX. Select **Runtime > Restart runtime** from the Colab menu. This is because of the way that Colab loads packages.
Do not proceed with the rest of this tutorial without first restarting the runtime.
### Import other libraries
```
import os
import tempfile
import urllib
import pandas as pd
```
Import [TFX component](https://tensorflow.google.cn/tfx/tutorials/tfx/components_keras) classes.
```
from tfx.components.evaluator.component import Evaluator
from tfx.components.example_gen.csv_example_gen.component import CsvExampleGen
from tfx.components.pusher.component import Pusher
from tfx.components.schema_gen.component import SchemaGen
from tfx.components.statistics_gen.component import StatisticsGen
from tfx.components.trainer.component import Trainer
from tfx.components.base import executor_spec
from tfx.components.trainer.executor import GenericExecutor
from tfx.orchestration.experimental.interactive.interactive_context import InteractiveContext
from tfx.proto import evaluator_pb2
from tfx.proto import pusher_pb2
from tfx.proto import trainer_pb2
from tensorflow_metadata.proto.v0 import anomalies_pb2
from tensorflow_metadata.proto.v0 import schema_pb2
from tensorflow_metadata.proto.v0 import statistics_pb2
import tensorflow_model_analysis as tfma
from tfx.dsl.experimental import latest_blessed_model_resolver
from tfx.types import Channel
from tfx.types.standard_artifacts import Model
from tfx.types.standard_artifacts import ModelBlessing
```
Import the MLMD library.
```
import ml_metadata as mlmd
from ml_metadata.proto import metadata_store_pb2
```
## Download the dataset
In this colab, we use the [Palmer Penguins dataset](https://allisonhorst.github.io/palmerpenguins/articles/intro.html) which can be found on [Github](https://github.com/allisonhorst/palmerpenguins). We processed the dataset by leaving out any incomplete records, and drops `island` and `sex` columns, and converted labels to `int32`. The dataset contains 334 records of the body mass and the length and depth of penguins' culmens, and the length of their flippers. You use this data to classify penguins into one of three species.
```
DATA_PATH = 'https://raw.githubusercontent.com/tensorflow/tfx/master/tfx/examples/penguin/data/penguins_processed.csv'
_data_root = tempfile.mkdtemp(prefix='tfx-data')
_data_filepath = os.path.join(_data_root, "penguins_processed.csv")
urllib.request.urlretrieve(DATA_PATH, _data_filepath)
```
## Create an InteractiveContext
To run TFX components interactively in this notebook, create an `InteractiveContext`. The `InteractiveContext` uses a temporary directory with an ephemeral MLMD database instance. Note that calls to `InteractiveContext` are no-ops outside the Colab environment.
In general, it is a good practice to group similar pipeline runs under a `Context`.
```
interactive_context = InteractiveContext()
```
## Construct the TFX Pipeline
A TFX pipeline consists of several components that perform different aspects of the ML workflow. In this notebook, you create and run the `ExampleGen`, `StatisticsGen`, `SchemaGen`, and `Trainer` components and use the `Evaluator` and `Pusher` component to evaluate and push the trained model.
Refer to the [components tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/components_keras) for more information on TFX pipeline components.
Note: Constructing a TFX Pipeline by setting up the individual components involves a lot of boilerplate code. For the purpose of this tutorial, it is alright if you do not fully understand every line of code in the pipeline setup.
### Instantiate and run the ExampleGen Component
```
example_gen = CsvExampleGen(input_base=_data_root)
interactive_context.run(example_gen)
```
### Instantiate and run the StatisticsGen Component
```
statistics_gen = StatisticsGen(examples=example_gen.outputs['examples'])
interactive_context.run(statistics_gen)
```
### Instantiate and run the SchemaGen Component
```
infer_schema = SchemaGen(
statistics=statistics_gen.outputs['statistics'], infer_feature_shape=True)
interactive_context.run(infer_schema)
```
### Instantiate and run the Trainer Component
```
# Define the module file for the Trainer component
trainer_module_file = 'penguin_trainer.py'
%%writefile {trainer_module_file}
# Define the training algorithm for the Trainer module file
import os
from typing import List, Text
import tensorflow as tf
from tensorflow import keras
from tfx.components.trainer.fn_args_utils import DataAccessor
from tfx.components.trainer.fn_args_utils import FnArgs
from tfx.utils import io_utils
from tfx_bsl.tfxio import dataset_options
from tensorflow_metadata.proto.v0 import schema_pb2
# Features used for classification - culmen length and depth, flipper length,
# body mass, and species.
_LABEL_KEY = 'species'
_FEATURE_KEYS = [
'culmen_length_mm', 'culmen_depth_mm', 'flipper_length_mm', 'body_mass_g'
]
def _input_fn(file_pattern: List[Text], data_accessor: DataAccessor,
schema: schema_pb2.Schema, batch_size: int) -> tf.data.Dataset:
return data_accessor.tf_dataset_factory(
file_pattern,
dataset_options.TensorFlowDatasetOptions(
batch_size=batch_size, label_key=_LABEL_KEY), schema).repeat()
def _build_keras_model():
inputs = [keras.layers.Input(shape=(1,), name=f) for f in _FEATURE_KEYS]
d = keras.layers.concatenate(inputs)
d = keras.layers.Dense(8, activation='relu')(d)
d = keras.layers.Dense(8, activation='relu')(d)
outputs = keras.layers.Dense(3, activation='softmax')(d)
model = keras.Model(inputs=inputs, outputs=outputs)
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=[keras.metrics.SparseCategoricalAccuracy()])
return model
def run_fn(fn_args: FnArgs):
schema = schema_pb2.Schema()
io_utils.parse_pbtxt_file(fn_args.schema_path, schema)
train_dataset = _input_fn(
fn_args.train_files, fn_args.data_accessor, schema, batch_size=10)
eval_dataset = _input_fn(
fn_args.eval_files, fn_args.data_accessor, schema, batch_size=10)
model = _build_keras_model()
model.fit(
train_dataset,
epochs=int(fn_args.train_steps / 20),
steps_per_epoch=20,
validation_data=eval_dataset,
validation_steps=fn_args.eval_steps)
model.save(fn_args.serving_model_dir, save_format='tf')
```
Run the `Trainer` component.
```
trainer = Trainer(
module_file=os.path.abspath(trainer_module_file),
custom_executor_spec=executor_spec.ExecutorClassSpec(GenericExecutor),
examples=example_gen.outputs['examples'],
schema=infer_schema.outputs['schema'],
train_args=trainer_pb2.TrainArgs(num_steps=100),
eval_args=trainer_pb2.EvalArgs(num_steps=50))
interactive_context.run(trainer)
```
### Evaluate and push the model
Use the `Evaluator` component to evaluate and 'bless' the model before using the `Pusher` component to push the model to a serving directory.
```
_serving_model_dir = os.path.join(tempfile.mkdtemp(),
'serving_model/penguins_classification')
eval_config = tfma.EvalConfig(
model_specs=[
tfma.ModelSpec(label_key='species', signature_name='serving_default')
],
metrics_specs=[
tfma.MetricsSpec(metrics=[
tfma.MetricConfig(
class_name='SparseCategoricalAccuracy',
threshold=tfma.MetricThreshold(
value_threshold=tfma.GenericValueThreshold(
lower_bound={'value': 0.6})))
])
],
slicing_specs=[tfma.SlicingSpec()])
evaluator = Evaluator(
examples=example_gen.outputs['examples'],
model=trainer.outputs['model'],
schema=infer_schema.outputs['schema'],
eval_config=eval_config)
interactive_context.run(evaluator)
pusher = Pusher(
model=trainer.outputs['model'],
model_blessing=evaluator.outputs['blessing'],
push_destination=pusher_pb2.PushDestination(
filesystem=pusher_pb2.PushDestination.Filesystem(
base_directory=_serving_model_dir)))
interactive_context.run(pusher)
```
Running the TFX pipeline populates the MLMD Database. In the next section, you use the MLMD API to query this database for metadata information.
## Query the MLMD Database
The MLMD database stores three types of metadata:
* Metadata about the pipeline and lineage information associated with the pipeline components
* Metadata about artifacts that were generated during the pipeline run
* Metadata about the executions of the pipeline
A typical production environment pipeline serves multiple models as new data arrives. When you encounter erroneous results in served models, you can query the MLMD database to isolate the erroneous models. You can then trace the lineage of the pipeline components that correspond to these models to debug your models
Set up the metadata (MD) store with the `InteractiveContext` defined previously to query the MLMD database.
```
connection_config = interactive_context.metadata_connection_config
store = mlmd.MetadataStore(connection_config)
# All TFX artifacts are stored in the base directory
base_dir = connection_config.sqlite.filename_uri.split('metadata.sqlite')[0]
```
Create some helper functions to view the data from the MD store.
```
def display_types(types):
# Helper function to render dataframes for the artifact and execution types
table = {'id': [], 'name': []}
for a_type in types:
table['id'].append(a_type.id)
table['name'].append(a_type.name)
return pd.DataFrame(data=table)
def display_artifacts(store, artifacts):
# Helper function to render dataframes for the input artifacts
table = {'artifact id': [], 'type': [], 'uri': []}
for a in artifacts:
table['artifact id'].append(a.id)
artifact_type = store.get_artifact_types_by_id([a.type_id])[0]
table['type'].append(artifact_type.name)
table['uri'].append(a.uri.replace(base_dir, './'))
return pd.DataFrame(data=table)
def display_properties(store, node):
# Helper function to render dataframes for artifact and execution properties
table = {'property': [], 'value': []}
for k, v in node.properties.items():
table['property'].append(k)
table['value'].append(
v.string_value if v.HasField('string_value') else v.int_value)
for k, v in node.custom_properties.items():
table['property'].append(k)
table['value'].append(
v.string_value if v.HasField('string_value') else v.int_value)
return pd.DataFrame(data=table)
```
First, query the MD store for a list of all its stored `ArtifactTypes`.
```
display_types(store.get_artifact_types())
```
Next, query all `PushedModel` artifacts.
```
pushed_models = store.get_artifacts_by_type("PushedModel")
display_artifacts(store, pushed_models)
```
Query the MD store for the latest pushed model. This tutorial has only one pushed model.
```
pushed_model = pushed_models[-1]
display_properties(store, pushed_model)
```
One of the first steps in debugging a pushed model is to look at which trained model is pushed and to see which training data is used to train that model.
MLMD provides traversal APIs to walk through the provenance graph, which you can use to analyze the model provenance.
```
def get_one_hop_parent_artifacts(store, artifacts):
# Get a list of artifacts within a 1-hop of the artifacts of interest
artifact_ids = [artifact.id for artifact in artifacts]
executions_ids = set(
event.execution_id
for event in store.get_events_by_artifact_ids(artifact_ids)
if event.type == metadata_store_pb2.Event.OUTPUT)
artifacts_ids = set(
event.artifact_id
for event in store.get_events_by_execution_ids(executions_ids)
if event.type == metadata_store_pb2.Event.INPUT)
return [artifact for artifact in store.get_artifacts_by_id(artifacts_ids)]
```
Query the parent artifacts for the pushed model.
```
parent_artifacts = get_one_hop_parent_artifacts(store, [pushed_model])
display_artifacts(store, parent_artifacts)
```
Query the properties for the model.
```
exported_model = parent_artifacts[0]
display_properties(store, exported_model)
```
Query the upstream artifacts for the model.
```
model_parents = get_one_hop_parent_artifacts(store, [exported_model])
display_artifacts(store, model_parents)
```
Get the training data the model trained with.
```
used_data = model_parents[0]
display_properties(store, used_data)
```
Now that you have the training data that the model trained with, query the database again to find the training step (execution). Query the MD store for a list of the registered execution types.
```
display_types(store.get_execution_types())
```
The training step is the `ExecutionType` named `tfx.components.trainer.component.Trainer`. Traverse the MD store to get the trainer run that corresponds to the pushed model.
```
def find_producer_execution(store, artifact):
executions_ids = set(
event.execution_id
for event in store.get_events_by_artifact_ids([artifact.id])
if event.type == metadata_store_pb2.Event.OUTPUT)
return store.get_executions_by_id(executions_ids)[0]
trainer = find_producer_execution(store, exported_model)
display_properties(store, trainer)
```
## Summary
In this tutorial, you learned about how you can leverage MLMD to trace the lineage of your TFX pipeline components and resolve issues.
To learn more about how to use MLMD, check out these additional resources:
* [MLMD API documentation](https://www.tensorflow.org/tfx/ml_metadata/api_docs/python/mlmd)
* [MLMD guide](https://www.tensorflow.org/tfx/guide/mlmd)
| github_jupyter |
###### 22 November 2018, by Jeroen van Lidth de Jeude - [NETWORKS](http://networks.imtlucca.it/) - [IMT School for Advanced Studies Lucca](https://www.imtlucca.it/jeroen.vanlidth)
# Maximum Entropy null models: the Directed Configuration Model and the Reciprocated Configuration Model
In this notebook we provide the code for the analytical solution of the Directed Configurgion Model and the Reciprocated Configuration Model. For a full introduction and explanation of these models, please see: [Analytical maximum-likelihood method to detect patterns in real networks - T. Squartini and D. Garlaschelli, *New Journal of Physics* (03/08/2011)](http://iopscience.iop.org/article/10.1088/1367-2630/13/8/083001/meta).
The Maximum Entropy Network framework is a very successful class of null models because of its versatilaty to incorporate different model constraints, and the fact that it provides the least biased distribution of graph given those constraints.
Roughly the procedure follows:

# Code
While the actual specifications of the system of equations and the numerical solver can be very short, this code adds tricks to help the numerical solving, mainly by seperating zero-degree nodes.
```
import numpy as np
from numba import jit
from scipy.optimize import least_squares
@jit
def equations_to_solve_dcm(p, k_out, k_in):
"""DCM equations for numerical solver.
Args:
p: list of independent variables [x y]
adjacency_matrix: numpy.array adjacency matrix to be solved
Returns:
numpy array of observed degree - expected degree
"""
n_nodes = len(k_out)
p = np.array(p)
num_x_nonzero_nodes = np.count_nonzero(k_out)
x_nonzero = p[0:num_x_nonzero_nodes]
y_nonzero = p[num_x_nonzero_nodes:len(p)]
x = np.zeros(n_nodes)
x[k_out != 0] = x_nonzero
y = np.zeros(n_nodes)
y[k_in != 0] = y_nonzero
# Expected degrees
k_out_exp = np.zeros(x.shape[0])
k_in_exp = np.zeros(x.shape[0])
for i in np.arange(x.shape[0]):
for j in np.arange(x.shape[0]):
if i != j:
k_out_exp[i] += (x[i] * y[j]) / (1 + x[i] * y[j])
k_in_exp[i] += (x[j] * y[i]) / (1 + x[j] * y[i])
k_out_nonzero = k_out[k_out != 0]
k_in_nonzero = k_in[k_in != 0]
k_out_exp_nonzero = k_out_exp[k_out != 0]
k_in_exp_nonzero = k_in_exp[k_in != 0]
f1 = k_out_nonzero - k_out_exp_nonzero
f2 = k_in_nonzero - k_in_exp_nonzero
return np.concatenate((f1, f2))
def numerically_solve_dcm(adjacency_matrix):
"""Solves the DCM numerically with least squares.
Directed Binary Configuration Model is solved using the
system of equations. The optimization is done using
scipy.optimize.least_squares on the system of equations.
Args:
adjacency_matrix : numpy.array adjacency matrix (binary, square)
Returns:
numpy.array probability matrix with dcm probabilities
"""
n_nodes = len(adjacency_matrix)
# Rough estimate of initial values
k_in = np.sum(adjacency_matrix, 0)
k_out = np.sum(adjacency_matrix, 1)
x_initial_values = k_out / np.sqrt(np.sum(k_out) + 1) # plus one to prevent dividing by zero
y_initial_values = k_in / np.sqrt(np.sum(k_in) + 1)
x_initial_values = x_initial_values[k_out != 0]
y_initial_values = y_initial_values[k_in != 0]
initial_values = np.concatenate((x_initial_values, y_initial_values))
# print(len(adjacency_matrix), len(x_initial_values), len(y_initial_values))
boundslu = tuple([0] * len(initial_values)), tuple([np.inf] * len(initial_values))
x_solved = least_squares(fun=equations_to_solve_dcm,
x0=initial_values,
args=(k_out, k_in,),
bounds=boundslu,
max_nfev=1e2,
ftol=1e-5, xtol=1e-5, gtol=1e-5)
print(x_solved.cost, x_solved.message)
# Numerical solution checks
assert x_solved.cost < 0.1, 'Numerical convergence problem: final cost function evaluation > 1'
# Set extremely small values to zero
# x_solved.x[x_solved.x < 1e-8] = 0
p = x_solved.x
p = np.array(p)
num_x_nonzero_nodes = np.count_nonzero(k_out)
x_nonzero = p[0:num_x_nonzero_nodes]
y_nonzero = p[num_x_nonzero_nodes:len(p)]
x = np.zeros(n_nodes)
x[k_out != 0] = x_nonzero
y = np.zeros(n_nodes)
y[k_in != 0] = y_nonzero
x_array = x
y_array = y
p_adjacency = np.zeros([n_nodes, n_nodes])
for i in np.arange(n_nodes):
for j in np.arange(n_nodes):
if i == j:
continue
p_adjacency[i, j] = x_array[i] * y_array[j] / (1 + x_array[i] * y_array[j])
return p_adjacency
@jit
def equations_to_solve_rcm(p, k_out, k_in, k_rec):
"""RCM equations for numerical solver.
Args:
p: list of independent variables [x y z]
adjacency_matrix: adjacency matrix to be solved
Returns:
numpy array of observed degree - expected degree
"""
n_nodes = len(k_out)
p = np.array(p)
num_x_nonzero_nodes = np.count_nonzero(k_out)
num_y_nonzero_nodes = np.count_nonzero(k_in)
x_nonzero = p[0:num_x_nonzero_nodes]
y_nonzero = p[num_x_nonzero_nodes:num_x_nonzero_nodes + num_y_nonzero_nodes]
z_nonzero = p[num_x_nonzero_nodes + num_y_nonzero_nodes:len(p)]
# print(len(k_out),len(k_in),len(k_rec),len(x_nonzero),len(y_nonzero),len(z_nonzero))
x = np.zeros(n_nodes)
x[k_out != 0] = x_nonzero
y = np.zeros(n_nodes)
y[k_in != 0] = y_nonzero
z = np.zeros(n_nodes)
z[k_rec != 0] = z_nonzero
# Expected degrees
k_out_exp = np.zeros(x.shape[0])
k_in_exp = np.zeros(x.shape[0])
k_rec_exp = np.zeros(x.shape[0])
for i in np.arange(x.shape[0]):
for j in np.arange(x.shape[0]):
if i != j:
k_out_exp[i] += (x[i] * y[j]) / (1 + x[i] * y[j] + x[j] * y[i] + z[i] * z[j])
k_in_exp[i] += (x[j] * y[i]) / (1 + x[i] * y[j] + x[j] * y[i] + z[i] * z[j])
k_rec_exp[i] += (z[i] * z[j]) / (1 + x[i] * y[j] + x[j] * y[i] + z[i] * z[j])
k_out_nonzero = k_out[k_out != 0]
k_in_nonzero = k_in[k_in != 0]
k_rec_nonzero = k_rec[k_rec != 0]
k_out_exp_nonzero = k_out_exp[k_out != 0]
k_in_exp_nonzero = k_in_exp[k_in != 0]
k_rec_exp_nonzero = k_rec_exp[k_rec != 0]
f1 = k_out_nonzero - k_out_exp_nonzero
f2 = k_in_nonzero - k_in_exp_nonzero
f3 = k_rec_nonzero - k_rec_exp_nonzero
return np.concatenate((f1, f2, f3))
def numerically_solve_rcm(adjacency_matrix):
"""Solves the RCM numerically with least squares.
Reciprocated Binary Configuration Model is solved using the
system of equations. The optimization is done using
scipy.optimize.least_squares on the system of equations.
Args:
adjacency_matrix : numpy.array adjacency matrix (binary, square)
Returns:
numpy.array probability matrices with dcm probabilities:
p_out_edges, p_in_edges, p_reciprocated_edges
"""
n_nodes = len(adjacency_matrix)
# Observed degrees
k_rec = np.zeros(adjacency_matrix.shape[0], dtype=np.int)
k_out = np.zeros(adjacency_matrix.shape[0], dtype=np.int)
k_in = np.zeros(adjacency_matrix.shape[0], dtype=np.int)
for i in np.arange(adjacency_matrix.shape[0]):
for j in np.arange(adjacency_matrix.shape[0]):
if i != j:
k_rec_i = np.min((adjacency_matrix[i, j], adjacency_matrix[j, i]))
k_out[i] += adjacency_matrix[i, j] - k_rec_i
k_in[i] += adjacency_matrix[j, i] - k_rec_i
k_rec[i] += k_rec_i
# Rough estimate of initial values
x_initial_values = k_out / np.sqrt(np.sum(k_out) + 1) # plus one to prevent dividing by zero
y_initial_values = k_in / np.sqrt(np.sum(k_in) + 1)
z_initial_values = k_rec / np.sqrt(np.sum(k_rec) + 1)
x_initial_values = x_initial_values[k_out != 0]
y_initial_values = y_initial_values[k_in != 0]
z_initial_values = z_initial_values[k_rec != 0]
initial_values = np.concatenate((x_initial_values, y_initial_values, z_initial_values))
boundslu = tuple([0] * len(initial_values)), tuple([np.inf] * len(initial_values))
x_solved = least_squares(fun=equations_to_solve_rcm,
x0=initial_values,
args=(k_out, k_in, k_rec,),
bounds=boundslu,
max_nfev=1e4,
ftol=1e-15, xtol=1e-15, gtol=1e-15)
print(x_solved.cost, x_solved.message)
# Numerical solution checks
assert x_solved.cost < 1, 'Numerical convergence problem: final cost function evaluation > 1'
p = np.array(x_solved.x)
num_x_nonzero_nodes = np.count_nonzero(k_out)
num_y_nonzero_nodes = np.count_nonzero(k_in)
x_nonzero = p[0:num_x_nonzero_nodes]
y_nonzero = p[num_x_nonzero_nodes:num_x_nonzero_nodes + num_y_nonzero_nodes]
z_nonzero = p[num_x_nonzero_nodes + num_y_nonzero_nodes:len(p)]
x = np.zeros(n_nodes)
x[k_out != 0] = x_nonzero
y = np.zeros(n_nodes)
y[k_in != 0] = y_nonzero
z = np.zeros(n_nodes)
z[k_rec != 0] = z_nonzero
x_array = x
y_array = y
z_array = z
p_out = np.zeros([n_nodes, n_nodes])
for i in np.arange(n_nodes):
for j in np.arange(n_nodes):
if i == j:
continue
p_out[i, j] = x_array[i] * y_array[j] / (
1 + x_array[i] * y_array[j] + x_array[j] * y_array[i] + z_array[i] * z_array[j])
p_in = np.zeros([n_nodes, n_nodes])
for i in np.arange(n_nodes):
for j in np.arange(n_nodes):
if i == j:
continue
p_in[i, j] = x_array[j] * y_array[i] / (
1 + x_array[i] * y_array[j] + x_array[j] * y_array[i] + z_array[i] * z_array[j])
p_rec = np.zeros([n_nodes, n_nodes])
for i in np.arange(n_nodes):
for j in np.arange(n_nodes):
if i == j:
continue
p_rec[i, j] = z_array[i] * z_array[j] / (
1 + x_array[i] * y_array[j] + x_array[j] * y_array[i] + z_array[i] * z_array[j])
return p_out, p_in, p_rec
```
# Applying the code
With a simple random matrix we now show how this code works.
The input to the algorithm is always an adjacency matrix; in this case a binary, unweighted one without self-edges. This should be supplied as a `numpy.array`.
The code is as simple as running the function `numerically_solve_dcm` with the adjacency matrix as argument. This will return an adjacency matrix where the matrix entries are the edge-probabilities under the given null model.
```
# First we generate a random matrix
n = 15
l = 0.6
adjacency_matrix = (np.random.rand(n,n) < l) # Generate random matrix with given edge density
np.fill_diagonal(adjacency_matrix,0) # Prevent self-loops
adjacency_matrix = adjacency_matrix.astype(int) # Force binary links
adjacency_matrix = np.asarray(adjacency_matrix) # Format as np.ndarray instead of np.matrix
```
Now we solve both the DCM and the RCM null models
```
probabilities_dcm = numerically_solve_dcm(adjacency_matrix=adjacency_matrix)
probabilities_rcm_out, probabilities_rcm_in, probabilities_rcm_rec = numerically_solve_rcm(adjacency_matrix=adjacency_matrix)
probabilities_rcm = probabilities_rcm_out + probabilities_rcm_in + probabilities_rcm_rec
```
The solving prodedures will display the final cost and the termination condition of the numerical solver of the system of equations. The final cost is the difference between the degree sequence of the original graph and that of the expected graph under the null model. In general a small error (<1e-3) is considered enough to claim a good solution of system of equations.
Now we visualise the result:
```
# Loading matplotlib library for visualisation
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
adjacencies = [adjacency_matrix, probabilities_dcm, probabilities_rcm]
titles = ['Original matrix', 'DCM probabilities', 'RCM probabilities']
colormap = 'Blues'
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(15,5))
for i, ax in enumerate(axes.flat):
im = ax.imshow(adjacencies[i], vmin=0, vmax=1, cmap = colormap)
ax.set_title(titles[i])
fig.subplots_adjust(right=0.9)
cbar_ax = fig.add_axes([0.95, 0.15, 0.05, 0.7])
fig.colorbar(im, cax=cbar_ax)
cbar_ax.set_title('edge-probability')
plt.show()
```
### Useful links
This work was done within the [NETWORKS](http://networks.imtlucca.it/) research group at [IMT School for Advanced Studies Lucca](http://www.imtlucca.it/).
The Maximum Entropy null models are based on the many works of the collaborators mentioned above, see also [Maximum-Entropy Networks](https://www.springer.com/it/book/9783319694368), [The Statistical Physics of Real-World Networks](https://arxiv.org/abs/1810.05095) and [Analytical maximum-likelihood method to detect patterns in real networks](http://iopscience.iop.org/article/10.1088/1367-2630/13/8/083001/meta).
| github_jupyter |
```
import ipywidgets as widgets
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import random
import itertools
from sklearn.metrics import accuracy_score, auc, roc_auc_score
%run c:/code/python/pyIDS/main.py
from pyids.data_structures.ids_rule import IDSRule
from pyids.data_structures.ids_ruleset import IDSRuleSet
from pyids.data_structures.ids_objective_function import ObjectiveFunctionParameters, IDSObjectiveFunction
from pyids.data_structures.ids_optimizer import RSOptimizer, SLSOptimizer
from pyids.data_structures.ids_cacher import IDSCacher
from pyids.data_structures.ids_classifier import IDS, mine_CARs
from matplotlib.collections import PatchCollection
from matplotlib.patches import Rectangle
from matplotlib.colors import ListedColormap
from pyarc.qcba import *
from pyarc.qcba.data_structures import (
IntervalReader,
Interval,
QuantitativeDataFrame,
QuantitativeCAR
)
interval_reader = IntervalReader()
interval_reader.closed_bracket = "<", ">"
interval_reader.open_bracket = "(", ")"
interval_reader.infinity_symbol = "-inf", "+inf"
interval_reader.members_separator = ";"
interval_reader.compile_reader()
QuantitativeCAR.interval_reader = interval_reader
%matplotlib inline
import sys
class IDSVisualization:
def __init__(self, rules, quant_dataframe, figsize=(10, 10)):
if type(quant_dataframe) != QuantitativeDataFrame:
raise Exception("type of quant_dataframe must be QuantitativeDataFrame")
self.rules = rules
self.pd_dataframe = quant_dataframe.dataframe
self.colnames = list(self.pd_dataframe.columns)
self.colnames_len = len(self.colnames)
self.colnames_x = self.colnames[:self.colnames_len - 1]
self.colnames_y = self.colnames[self.colnames_len - 1]
self.colors = ["red", "green", "blue", "black", "cyan", "yellow", "magenta", "lightblue", "lightgreen"]
self.classes = list(self.pd_dataframe[self.colnames_y].values)
self.classes_numbers_encoding = list(range(len(self.classes)))
self.classes_unique = list(set(self.classes))
self.classes_numbers = [ self.classes_numbers_encoding[self.classes_unique.index(clazz)] for clazz in self.classes ]
self.classes_numbers_dict = { clazz:self.classes_numbers_encoding[self.classes_unique.index(clazz)] for clazz in self.classes_unique }
self.colors_map = self._get_color_map()
self.colors_unique = [ self.colors_map[clazz] for clazz in self.classes_unique ]
self.colnames_x_combinations = list(itertools.combinations(self.colnames_x, 2))
self.column_extremas = self._get_column_extremas()
self.figsize = figsize
self.figure = None
self.axes = None
def _get_color_map(self):
classes_unique = set(self.classes)
colors_map = dict()
for idx, class_label in enumerate(classes_unique):
colors_map.update({class_label: self.colors[idx]})
return colors_map
def _get_column_extremas(self):
column_extremas = dict()
for column in self.pd_dataframe.columns:
minval = self.pd_dataframe[column].min()
maxval = self.pd_dataframe[column].max()
column_extremas.update({column: (minval, maxval)})
for rule in self.rules:
for column, interval in rule.antecedent:
minval, maxval = interval.minval, interval.maxval
current_minval, current_maxval = column_extremas.get(
column,
(self.pd_dataframe[column].min(), self.pd_dataframe[column].max())
)
if minval > current_minval:
minval = current_minval
if maxval < current_maxval:
maxval = current_maxval
column_extremas.update({column: (minval, maxval)})
return column_extremas
def _prepare_figure(self):
fig_columns = 2
fig_rows = int(np.ceil(self.colnames_len / 2))
if len(self.colnames) - 1 == 2:
fig_columns = 1
fig_rows = 1
fig, axes = plt.subplots(fig_rows, fig_columns, figsize=self.figsize)
if len(self.colnames) - 1 == 2:
axes = np.array([axes])
axes = axes.reshape(fig_columns * fig_rows)
#fig.set_size_inches(*figsize)
self.fig = fig
self.axes = axes
def visualize_IDS(self):
self._prepare_figure()
self._visualize_rules()
self._visualize_dataframe()
def visualize_dataframe(self):
self._prepare_figure()
self._visualize_dataframe()
def _visualize_dataframe(self):
for idx, ax in enumerate(self.axes):
col_x, col_y = self.colnames_x_combinations[idx]
x = self.pd_dataframe[col_x]
y = self.pd_dataframe[col_y]
colours = ListedColormap(self.colors_unique)
scatter = ax.scatter(x, y, c=self.classes_numbers, cmap=colours)
ax.set_xlabel(col_x)
ax.set_ylabel(col_y)
ax.set_title("Scatter of '{}' vs '{}'".format(col_x, col_y))
ax.grid()
ax.legend(handles=scatter.legend_elements()[0], labels=self.classes_numbers_dict.keys())
if len(self.colnames_x_combinations) == 1 and idx == 0:
break
def _rule_covers_whole_subspace(self):
return False
def _substitute_infinities(self, colname, minval, maxval):
if minval == -np.inf:
minval = self.column_extremas[colname][0]
if maxval == np.inf:
maxval = self.column_extremas[colname][1]
return minval, maxval
def _create_rule_rectangle(self, ax_idx, antecedent_dict):
col_x, col_y = self.colnames_x_combinations[ax_idx]
column_x_extrema = self.column_extremas[col_x]
column_y_extrema = self.column_extremas[col_y]
interval_x = antecedent_dict.get(col_x, Interval(*column_x_extrema, True, True))
interval_y = antecedent_dict.get(col_y, Interval(*column_y_extrema, True, True))
rule_x_min, rule_x_max = interval_x.minval, interval_x.maxval
rule_y_min, rule_y_max = interval_y.minval, interval_y.maxval
rule_x_min, rule_x_max = self._substitute_infinities(col_x, rule_x_min, rule_x_max)
rule_y_min, rule_y_max = self._substitute_infinities(col_y, rule_y_min, rule_y_max)
if (rule_x_min, rule_x_max) == column_x_extrema and (rule_y_min, rule_y_max) == column_y_extrema:
return None
rect_width = rule_x_max - rule_x_min
rect_height = rule_y_max - rule_y_min
rect = Rectangle((rule_x_min, rule_y_min), rect_width, rect_height)
return rect
def _visualize_rules(self):
for ax_idx, ax in enumerate(self.axes):
ax_rule_patches = []
for rule in self.rules:
antecedent_dict = dict(rule.antecedent)
rule_rect = self._create_rule_rectangle(ax_idx, antecedent_dict)
if rule_rect:
rule_class = rule.consequent.value
class_color = self.colors_map[rule_class]
rule_alpha = rule.confidence * 0.25
rule_rect.set_fill(True)
rule_rect.set_color(class_color)
rule_rect.set_alpha(rule_alpha)
ax_rule_patches.append(rule_rect)
for rect in ax_rule_patches:
ax.add_patch(rect)
if len(self.colnames_x_combinations) == 1 and ax_idx == 0:
break
df_undiscr = pd.read_csv("../data/movies.csv", sep=";")
df_discr = pd.read_csv("../data/movies_discr.csv", sep=";")
df_discr = df_discr.iloc[:, 1:]
cars0 = mine_CARs(df_discr, 3, sample=True)
cars1 = mine_CARs(df_discr, 7, sample=False)
cars = cars0 + cars1
quant_cars = list(map(QuantitativeCAR, cars))
quant_df = QuantitativeDataFrame(df_undiscr)
ids = IDS()
ids.fit(quant_df, cars, debug=False)
viz = IDSVisualization(quant_cars, quant_df, figsize=(6, 6))
viz.visualize_IDS()
viz.visualize_dataframe()
def print_cars(ids_cars):
cars = [ r.car for r in ids_cars ]
latex = []
for car in cars:
latex.append(car_to_latex(car))
return "\n\n".join(latex)
def car_to_latex(car):
latex = []
latex.append("If")
for predicate in car.antecedent:
name, interval = predicate
name = name.translate(str.maketrans({"-": r"\-",
"_": r"\_",
"]": r"\]",
"\\": r"\\",
"^": r"\^",
"$": r"\$",
"*": r"\*",
".": r"\."}))
interval_string = interval
latex.append(
f"\\textcolor{{blue}}{{{name}}}=\\textcolor{{gray}}{{{interval_string}}}"
)
class_name, class_value = car.consequent
latex.append("then")
latex.append(
f"\\textcolor{{red}}{{{class_name}}}=\\textcolor{{gray}}{{{class_value}}}"
)
return " ".join(latex)
print(print_cars(ids.clf.rules))
viz.visualize_dataframe()
```
## f0
```
viz_f0_0 = IDSVisualization(quant_cars, quant_df, figsize=(6, 6))
viz_f0_1 = IDSVisualization(quant_cars[:4], quant_df, figsize=(6, 6))
viz_f0_0.visualize_IDS()
viz_f0_1.visualize_IDS()
quant_cars
```
## f1
```
viz_f1_0 = IDSVisualization(quant_cars, quant_df, figsize=(6, 6))
viz_f1_1 = IDSVisualization([ car for car in quant_cars if len(car.antecedent) == 1 ], quant_df, figsize=(6, 6))
viz_f1_0.visualize_IDS()
viz_f1_1.visualize_IDS()
```
## f2
```
rules_f2_0 = [car for car in quant_cars if car.consequent[1] == "box-office-bomb" ]
rules_f2_1 = [ car for car in quant_cars if car.consequent[1] == "main-stream-hit" ]
viz_f2_0 = IDSVisualization(rules_f2_0, quant_df, figsize=(6, 6))
viz_f2_1 = IDSVisualization(rules_f2_1, quant_df, figsize=(6, 6))
viz_f2_0.visualize_IDS()
viz_f2_1.visualize_IDS()
```
## f3
```
def two_classes_predicate(car):
return car.consequent[1] in ["critical-success", "box-office-bomb"]
def minimize_overlap_heuristically_predicate(car):
return len(car.antecedent) == 2
rules_f3_0 = [car for car in quant_cars if two_classes_predicate(car) and minimize_overlap_heuristically_predicate(car) ]
rules_f3_1 = [ car for car in quant_cars[:5] ]
viz_f3_0 = IDSVisualization(rules_f3_0, quant_df, figsize=(6, 6))
viz_f3_1 = IDSVisualization(rules_f3_1, quant_df, figsize=(6, 6))
viz_f3_0.visualize_IDS()
viz_f3_1.visualize_IDS()
```
## f4
```
viz_f4_0 = IDSVisualization(quant_cars, quant_df, figsize=(6, 6))
viz_f4_1 = IDSVisualization([ car for car in quant_cars if car.consequent[1] != "critical-success" ], quant_df, figsize=(6, 6))
viz_f4_0.visualize_IDS()
viz_f4_1.visualize_IDS()
```
## f5
```
viz_f5_0 = IDSVisualization([ car for car in quant_cars if car.confidence >= 0.9 ], quant_df, figsize=(6, 6))
viz_f5_1 = IDSVisualization([ car for car in quant_cars if car.confidence < 0.9 ], quant_df, figsize=(6, 6))
viz_f5_0.visualize_IDS()
viz_f5_1.visualize_IDS()
```
## f6
```
viz_f6_0 = IDSVisualization(quant_cars, quant_df, figsize=(6, 6))
viz_f6_1 = IDSVisualization([ car for car in quant_cars if car.support < 0.05 ], quant_df, figsize=(6, 6))
viz_f6_0.visualize_IDS()
viz_f6_1.visualize_IDS()
```
| github_jupyter |
```
import json
import pandas as pd
import requests
import synapseclient
from synapseclient import File
syn = synapseclient.Synapse()
syn.login()
```
Load the categories from the Yelp documentation at https://www.yelp.com/developers/documentation/v3/all_category_list
```
r = requests.get('https://www.yelp.com/developers/documentation/v3/all_category_list/categories.json')
if r.ok:
j = r.json()
```
Manual mappings based primarily on the Yelp defined hieararchy but several have been manually tuned.
```
YELP_TYPE_MAPPINGS = dict(
government_offices=[
'publicservicesgovt', 'animalshelters', 'government_offices'
],
place_of_mourning=[
'funeralservices', 'place_of_mourning'
],
education=[
'education', 'tastingclasses', 'specialtyschools', 'adultedu'
],
place_of_worship=[
'religiousorgs', 'place_of_worship'
],
lodging=[
'hotels', 'hotelstravel', 'agriturismi', 'apartments', 'condominiums', 'university_housing',
'homelessshelters', 'lodging'
],
entertainment=[
'active', 'adultentertainment', 'artclasses', 'arts', 'artsandcrafts', 'entertainment',
'bars', 'breweries', 'diving', 'festivals', 'martialarts', 'movietheaters',
'museums', 'nightlife', 'tours', 'wineries', 'zoos', 'social_clubs', 'localflavor'
],
health=[
'dentalhygienists', 'dentists', 'fitness', 'gyms', 'health', 'medcenters', 'medicalspa',
'opthamalogists', 'opticians', 'physicians', 'tcm', 'c_and_mh', 'acnetreatment', 'acupuncture',
'addictionmedicine', 'allergist', 'alternativemedicine', 'anesthesiologists',
'animalassistedtherapy'
],
finance=[
'estateplanning', 'financialservices', 'insurance', 'accountants', 'finance'
],
repair=[
'autoglass', 'autorepair', 'diagnosticservices', 'itservices', 'homeservices', 'repair',
'junkremovalandhauling', 'laundryservices', 'localservices', 'musicinstrumentservices',
'plumbing', 'professional'
],
transit=[
'airports', 'transport', 'travelservices', 'carrental', 'motorcycle_rental', 'trainstations', 'transit',
],
dining_out=[
'cafes', 'food', 'restaurants', 'jpsweets', 'african', 'arabian', 'dining_out',
'belgian', 'brazilian', 'caribbean', 'chinese', 'donburi', 'french', 'german', 'turkish',
'italian', 'japanse', 'latin', 'malaysian', 'mediterranean', 'mexican', 'mideastern', 'polish',
'portugese', 'spanish', 'portuguese', 'japanese'
],
home_store=[
'gardening', 'homeandgarden', 'professional', 'kitchenandbath', 'landscaping', 'realestate', 'home_store'
'realestateagents', 'realestatesvcs', 'apartmentagents', 'apartments', 'appliances', 'appraisalservices'
],
supermarket=[
'food, shopping', 'farms', 'wholesale_stores', 'wholesalers', 'gourmet', 'grocery', 'ethicgrocery', 'beverage_stores',
'butcher', 'csa', 'convenience', 'farmersmarket', 'organic_stores', 'supermarket'
],
automotive=[
'auto', 'automotive'
],
consumer_goods=[
'flowers', 'bicycles', 'cannabis_clinics', 'fashion', 'shopping', 'partyequipmentrentals',
'sportgoods', 'sportswear', 'wholesalers', 'pets', 'petstore', 'petservices',
'accessories', 'petbreeders', 'antiques', 'shoppingcenters'
],
personal_services=[
'eventservices', 'beautysvc', 'hair', 'hairremoval', 'othersalons', 'psychic_astrology', 'skincare',
'tanning', 'photographers', 'utilities', 'pet_sitting', 'aestheticians', 'animalholistic',
'animalphysicaltherapy', '3dprinting', 'personal_services', 'lawyers', 'legalservices'
],
park=[
'parks', 'park'
],
other_organization=[
'massmedia', 'media', 'nonprofit', 'adoptionservices', 'advertising', 'other', 'other_organization'
]
)
```
Now, we iterate first through each alias looking for a mapping but if the alias isn't already in one of our predefined categories then check the parent. Mostly, the parent will be our source but in some instances we use the alias as it is a 'finer-grained' mapping.
```
yelp_mappings = pd.DataFrame(columns=['cat', 'mapping'])
for i, r in enumerate(j):
alias = r.get('alias')
parent = r.get('parents')
mapping = 'none'
for k, v in YELP_TYPE_MAPPINGS.items():
if alias in v:
mapping = k
break
if mapping == 'none':
for k, v in YELP_TYPE_MAPPINGS.items():
if any([p in v for p in parent]):
mapping = k
yelp_mappings.loc[i] = (alias, mapping)
yelp_mappings.to_csv('yelp_mappings.csv', index=None)
t = syn.store(File(name='yelp_mappings', path='yelp_mappings.csv', parent='syn16816579')); del t
GMAP_TYPE_MAPPINGS = dict(
government_offices=[
'post_office', 'city_hall', 'courthouse', 'embassy',
'local_government_office', 'police', 'fire_station',
'government_office'
],
place_of_mourning=[
'cemetery', 'funeral_home', 'place_of_mourning'
],
education=[
'school', 'university', 'education'
],
place_of_worship=[
'church', 'hindu_temple', 'mosque', 'synagogue', 'place_of_worship'
],
lodging=[
'campground', 'lodging', 'rv_park'
],
entertainment=[
'bar', 'amusement_park', 'aquarium', 'art_gallery', 'bowling_alley',
'casino', 'movie_rental', 'movie_theater', 'museum', 'night_club',
'stadium', 'zoo', 'library', 'entertainment'
],
health=[
'dentist', 'doctor', 'gym', 'hospital', 'pharmacy', 'physiotherapist', 'health'
],
finance=[
'atm', 'bank', 'insurance_agency', 'finance', 'accounting'
],
repair=[
'car_repair', 'car_wash', 'electrician', 'plumber', 'general_contractor',
'roofing_contractor', 'painter', 'locksmith', 'travel_agency', 'repair'
],
transit=[
'airport', 'bus_station', 'taxi_stand', 'train_station',
'transit_station', 'subway_station', 'travel_agency', 'transit'
],
dining_out=[
'bakery', 'cafe', 'meal_delivery', 'meal_takeaway', 'restaurant', 'dining_out', 'food'
],
home_store=[
'furniture_store', 'electronics_store', 'hardware_store',
'home_goods_store', 'moving_company', 'real_estate_agency',
'storage', 'laundry', 'home_store'
],
supermarket=[
'convenience_store', 'liquor_store', 'supermarket',
'grocery_or_supermarket'
],
automotive=[
'car_dealer', 'car_rental', 'gas_station', 'parking', 'automotive'
],
consumer_goods=[
'book_store', 'bicycle_store', 'clothing_store', 'department_store',
'florist', 'jewelry_store', 'pet_store', 'shoe_store', 'shopping_mall',
'consumer_goods', 'store'
],
personal_services=[
'beauty_salon', 'hair_care', 'spa', 'personal_services', 'lawyer', 'veterinary_care'
],
other_organization=[
'other'
],
park=[
'parks', 'park'
]
)
gmap_mappings = pd.DataFrame(columns=['cat', 'mapping'])
i = 0
for k, v in GMAP_TYPE_MAPPINGS.items():
for vi in v:
gmap_mappings.loc[i] = (vi, k)
i += 1
gmap_mappings.to_csv('gmap_mappings.csv', index=None)
t = syn.store(File(name='gmap_mappings', path='gmap_mappings.csv', parent='syn16816579')); del t
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Pruning in Keras example
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/model_optimization/guide/pruning/pruning_with_keras"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/model-optimization/blob/master/tensorflow_model_optimization/g3doc/guide/pruning/pruning_with_keras.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/model-optimization/blob/master/tensorflow_model_optimization/g3doc/guide/pruning/pruning_with_keras.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/model-optimization/tensorflow_model_optimization/g3doc/guide/pruning/pruning_with_keras.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
## Overview
Welcome to an end-to-end example for magnitude-based *weight pruning*.
### Other pages
For an introduction to what pruning is and to determine if you should use it (including what's supported), see the [overview page](https://www.tensorflow.org/model_optimization/guide/pruning).
To quickly find the APIs you need for your use case (beyond fully pruning a model with 80% sparsity), see the
[comprehensive guide](https://www.tensorflow.org/model_optimization/guide/pruning/comprehensive_guide.md).
### Summary
In this tutorial, you will:
1. Train a `tf.keras` model for MNIST from scratch.
2. Fine tune the model by applying the pruning API and see the accuracy.
3. Create 3x smaller TF and TFLite models from pruning.
4. Create a 10x smaller TFLite model from combining pruning and post-training quantization.
5. See the persistence of accuracy from TF to TFLite.
## Setup
```
! pip install -q tensorflow-model-optimization
import tempfile
import os
import tensorflow as tf
import numpy as np
from tensorflow import keras
%load_ext tensorboard
```
## Train a model for MNIST without pruning
```
# Load MNIST dataset
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# Normalize the input image so that each pixel value is between 0 to 1.
train_images = train_images / 255.0
test_images = test_images / 255.0
# Define the model architecture.
model = keras.Sequential([
keras.layers.InputLayer(input_shape=(28, 28)),
keras.layers.Reshape(target_shape=(28, 28, 1)),
keras.layers.Conv2D(filters=12, kernel_size=(3, 3), activation='relu'),
keras.layers.MaxPooling2D(pool_size=(2, 2)),
keras.layers.Flatten(),
keras.layers.Dense(10)
])
# Train the digit classification model
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(
train_images,
train_labels,
epochs=4,
validation_split=0.1,
)
```
Evaluate baseline test accuracy and save the model for later usage.
```
_, baseline_model_accuracy = model.evaluate(
test_images, test_labels, verbose=0)
print('Baseline test accuracy:', baseline_model_accuracy)
_, keras_file = tempfile.mkstemp('.h5')
tf.keras.models.save_model(model, keras_file, include_optimizer=False)
print('Saved baseline model to:', keras_file)
```
## Fine-tune pre-trained model with pruning
### Define the model
You will apply pruning to the whole model and see this in the model summary.
In this example, you start the model with 50% sparsity (50% zeros in weights)
and end with 80% sparsity.
In the [comprehensive guide](https://www.tensorflow.org/model_optimization/guide/pruning/comprehensive_guide.md), you can see how to prune some layers for model accuracy improvements.
```
import tensorflow_model_optimization as tfmot
prune_low_magnitude = tfmot.sparsity.keras.prune_low_magnitude
# Compute end step to finish pruning after 2 epochs.
batch_size = 128
epochs = 2
validation_split = 0.1 # 10% of training set will be used for validation set.
num_images = train_images.shape[0] * (1 - validation_split)
end_step = np.ceil(num_images / batch_size).astype(np.int32) * epochs
# Define model for pruning.
pruning_params = {
'pruning_schedule': tfmot.sparsity.keras.PolynomialDecay(initial_sparsity=0.50,
final_sparsity=0.80,
begin_step=0,
end_step=end_step)
}
model_for_pruning = prune_low_magnitude(model, **pruning_params)
# `prune_low_magnitude` requires a recompile.
model_for_pruning.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model_for_pruning.summary()
```
### Train and evaluate the model against baseline
Fine tune with pruning for two epochs.
`tfmot.sparsity.keras.UpdatePruningStep` is required during training, and `tfmot.sparsity.keras.PruningSummaries` provides logs for tracking progress and debugging.
```
logdir = tempfile.mkdtemp()
callbacks = [
tfmot.sparsity.keras.UpdatePruningStep(),
tfmot.sparsity.keras.PruningSummaries(log_dir=logdir),
]
model_for_pruning.fit(train_images, train_labels,
batch_size=batch_size, epochs=epochs, validation_split=validation_split,
callbacks=callbacks)
```
For this example, there is minimal loss in test accuracy after pruning, compared to the baseline.
```
_, model_for_pruning_accuracy = model_for_pruning.evaluate(
test_images, test_labels, verbose=0)
print('Baseline test accuracy:', baseline_model_accuracy)
print('Pruned test accuracy:', model_for_pruning_accuracy)
```
The logs show the progression of sparsity on a per-layer basis.
```
#docs_infra: no_execute
%tensorboard --logdir={logdir}
```
For non-Colab users, you can see [the results of a previous run](https://tensorboard.dev/experiment/sRQnrycaTMWQOaswXzClYA/#scalars&_smoothingWeight=0) of this code block on [TensorBoard.dev](https://tensorboard.dev/).
## Create 3x smaller models from pruning
Both `tfmot.sparsity.keras.strip_pruning` and applying a standard compression algorithm (e.g. via gzip) are necessary to see the compression
benefits of pruning.
* `strip_pruning` is necessary since it removes every tf.Variable that pruning only needs during training, which would otherwise add to model size during inference
* Applying a standard compression algorithm is necessary since the serialized weight matrices are the same size as they were before pruning. However, pruning makes most of the weights zeros, which is
added redundancy that algorithms can utilize to further compress the model.
First, create a compressible model for TensorFlow.
```
model_for_export = tfmot.sparsity.keras.strip_pruning(model_for_pruning)
_, pruned_keras_file = tempfile.mkstemp('.h5')
tf.keras.models.save_model(model_for_export, pruned_keras_file, include_optimizer=False)
print('Saved pruned Keras model to:', pruned_keras_file)
```
Then, create a compressible model for TFLite.
```
converter = tf.lite.TFLiteConverter.from_keras_model(model_for_export)
pruned_tflite_model = converter.convert()
_, pruned_tflite_file = tempfile.mkstemp('.tflite')
with open(pruned_tflite_file, 'wb') as f:
f.write(pruned_tflite_model)
print('Saved pruned TFLite model to:', pruned_tflite_file)
```
Define a helper function to actually compress the models via gzip and measure the zipped size.
```
def get_gzipped_model_size(file):
# Returns size of gzipped model, in bytes.
import os
import zipfile
_, zipped_file = tempfile.mkstemp('.zip')
with zipfile.ZipFile(zipped_file, 'w', compression=zipfile.ZIP_DEFLATED) as f:
f.write(file)
return os.path.getsize(zipped_file)
```
Compare and see that the models are 3x smaller from pruning.
```
print("Size of gzipped baseline Keras model: %.2f bytes" % (get_gzipped_model_size(keras_file)))
print("Size of gzipped pruned Keras model: %.2f bytes" % (get_gzipped_model_size(pruned_keras_file)))
print("Size of gzipped pruned TFlite model: %.2f bytes" % (get_gzipped_model_size(pruned_tflite_file)))
```
## Create a 10x smaller model from combining pruning and quantization
You can apply post-training quantization to the pruned model for additional benefits.
```
converter = tf.lite.TFLiteConverter.from_keras_model(model_for_export)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
quantized_and_pruned_tflite_model = converter.convert()
_, quantized_and_pruned_tflite_file = tempfile.mkstemp('.tflite')
with open(quantized_and_pruned_tflite_file, 'wb') as f:
f.write(quantized_and_pruned_tflite_model)
print('Saved quantized and pruned TFLite model to:', quantized_and_pruned_tflite_file)
print("Size of gzipped baseline Keras model: %.2f bytes" % (get_gzipped_model_size(keras_file)))
print("Size of gzipped pruned and quantized TFlite model: %.2f bytes" % (get_gzipped_model_size(quantized_and_pruned_tflite_file)))
```
## See persistence of accuracy from TF to TFLite
Define a helper function to evaluate the TF Lite model on the test dataset.
```
import numpy as np
def evaluate_model(interpreter):
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
# Run predictions on ever y image in the "test" dataset.
prediction_digits = []
for i, test_image in enumerate(test_images):
if i % 1000 == 0:
print('Evaluated on {n} results so far.'.format(n=i))
# Pre-processing: add batch dimension and convert to float32 to match with
# the model's input data format.
test_image = np.expand_dims(test_image, axis=0).astype(np.float32)
interpreter.set_tensor(input_index, test_image)
# Run inference.
interpreter.invoke()
# Post-processing: remove batch dimension and find the digit with highest
# probability.
output = interpreter.tensor(output_index)
digit = np.argmax(output()[0])
prediction_digits.append(digit)
print('\n')
# Compare prediction results with ground truth labels to calculate accuracy.
prediction_digits = np.array(prediction_digits)
accuracy = (prediction_digits == test_labels).mean()
return accuracy
```
You evaluate the pruned and quantized model and see that the accuracy from TensorFlow persists to the TFLite backend.
```
interpreter = tf.lite.Interpreter(model_content=quantized_and_pruned_tflite_model)
interpreter.allocate_tensors()
test_accuracy = evaluate_model(interpreter)
print('Pruned and quantized TFLite test_accuracy:', test_accuracy)
print('Pruned TF test accuracy:', model_for_pruning_accuracy)
```
## Conclusion
In this tutorial, you saw how to create sparse models with the TensorFlow Model Optimization Toolkit API for both TensorFlow and TFLite. You
then combined pruning with post-training quantization for additional benefits.
You created a 10x smaller model for MNIST, with minimal accuracy difference.
We encourage you to try this new capability, which can be particularly important for deployment in resource-constrained environments.
| github_jupyter |
```
import boto3
import sagemaker
print(sagemaker.__version__)
session = sagemaker.Session()
bucket = session.default_bucket()
prefix = 'dogscats'
s3_train_path = 's3://{}/{}/input/train/'.format(bucket, prefix)
s3_val_path = 's3://{}/{}/input/validation/'.format(bucket, prefix)
s3_output = 's3://{}/{}/output/'.format(bucket, prefix)
print(s3_train_path)
print(s3_val_path)
print(s3_output)
```
### Get the name of the image classification algorithm in our region
```
from sagemaker import image_uris
region = boto3.Session().region_name
container = image_uris.retrieve('image-classification', region)
print(container)
```
### Configure the training job
```
role = sagemaker.get_execution_role()
ic = sagemaker.estimator.Estimator(container,
role,
instance_count=1,
instance_type='ml.p3.2xlarge',
output_path=s3_output)
```
### Set algorithm parameters
```
#precision_dtype = 'float16'
precision_dtype = 'float32'
ic.set_hyperparameters(num_layers=18, # Train a Resnet-18 model
use_pretrained_model=0, # Train from scratch
num_classes=2, # Dogs and cats
num_training_samples=22500, # Number of training samples
mini_batch_size=128,
precision_dtype=precision_dtype,
epochs=10) # Learn the training samples 10 times
```
### Set dataset parameters
```
train_data = sagemaker.TrainingInput(s3_train_path,
distribution='FullyReplicated',
content_type='application/x-recordio',
s3_data_type='S3Prefix')
validation_data = sagemaker.TrainingInput(s3_val_path,
distribution='FullyReplicated',
content_type='application/x-recordio',
s3_data_type='S3Prefix')
s3_channels = {'train': train_data, 'validation': validation_data}
```
### Train the model
```
ic.fit(inputs=s3_channels)
```
### Deploy the model
```
import time
endpoint_name = 'c5-'+time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
c5_predictor = ic.deploy(initial_instance_count=1,
instance_type='ml.c5.large',
endpoint_name=endpoint_name,
wait=False)
endpoint_name = 'g4-'+time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
g4_predictor = ic.deploy(initial_instance_count=1,
instance_type='ml.g4dn.xlarge',
endpoint_name=endpoint_name,
wait=False)
endpoint_name = 'c5-medium-'+time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
c5_medium_predictor = ic.deploy(initial_instance_count=1,
instance_type='ml.c5.large',
accelerator_type='ml.eia2.medium',
endpoint_name=endpoint_name,
wait=False)
endpoint_name = 'c5-large-'+time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
c5_large_predictor = ic.deploy(initial_instance_count=1,
instance_type='ml.c5.large',
accelerator_type='ml.eia2.large',
endpoint_name=endpoint_name,
wait=False)
endpoint_name = 'c5-xlarge-'+time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
c5_xlarge_predictor = ic.deploy(initial_instance_count=1,
instance_type='ml.c5.large',
accelerator_type='ml.eia2.xlarge',
endpoint_name=endpoint_name,
wait=False)
```
### Download a test image
```
# Dog
!wget -O /tmp/test.jpg https://upload.wikimedia.org/wikipedia/commons/b/b7/LabradorWeaving.jpg
file_name = '/tmp/test.jpg'
from IPython.display import Image
Image(file_name)
```
### Predict test image
```
# Load test image from file
with open(file_name, 'rb') as f:
payload = f.read()
payload = bytearray(payload)
def predict_images(predictor, iterations=1000):
total = 0
predictor.content_type = 'application/x-image'
for i in range(0, iterations):
tick = time.time()
response = predictor.predict(payload)
tock = time.time()
total += tock-tick
return total/iterations
%%time
predict_images(c5_predictor)
%%time
predict_images(g4_predictor)
%%time
predict_images(c5_medium_predictor)
%%time
predict_images(c5_large_predictor)
%%time
predict_images(c5_xlarge_predictor)
```
### Delete endpoints
```
c5_predictor.delete_endpoint()
g4_predictor.delete_endpoint()
c5_medium_predictor.delete_endpoint()
c5_large_predictor.delete_endpoint()
c5_xlarge_predictor.delete_endpoint()
```
| github_jupyter |
## Trial Recurrent Neural Network
With the change to the data set (i.e. dropping betting data), it seems that we have to go back to basics, because earlier experiments are no longer valid, resulting in a different set of algorithms having stronger performance. The experiments with deep learning in 2019 yielded performance comparable to simpler models, and I decided not to pursue them at the time, because comparable performance isn't worth the extra time for training/param-tuning required. With the current benchmark being a bit lower, however, it's worth seeing if a deep neural net can surpass simpler models.
To avoid spending too much time on this, I'm going to focus on RNNs, because it was the best-performing architecture from earlier experiments and lends itself to the time-based nature of AFL matches.
## Code setup
```
%load_ext autoreload
%autoreload 2
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from sklearn.experimental import enable_halving_search_cv
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import make_pipeline
from sklearn.metrics import get_scorer
from sklearn.model_selection import HalvingRandomSearchCV, RandomizedSearchCV, cross_validate
from sklearn.preprocessing import StandardScaler, OrdinalEncoder, FunctionTransformer
from mlxtend.feature_selection import ColumnSelector
from augury.ml_data import MLData
from augury.model_tracking import graph_tf_model_history
from augury.settings import SEED, CV_YEAR_RANGE, CATEGORY_COLS, TEAM_NAMES, ROUND_TYPES, VENUES
from augury.sklearn.metrics import match_accuracy_scorer
from augury.sklearn.models import RNNRegressor
from augury.sklearn.model_selection import year_cv_split
from augury.sklearn.preprocessing import ColumnDropper
np.random.seed(SEED)
data = MLData(train_year_range=(max(CV_YEAR_RANGE),))
data.data
```
### Create pipelines
## Runn RNN
### Basic RNN for baseline performance
RNNRegressor mostly uses the defaults for a Keras RNN model with LSTM cells. It has one hidden layer and measures accuracy, but optimises toward MAE, because accuracy doesn't lend itself easily to a loss function for a regressor. I chose 6 steps for the time-series, because that's a quarter of an average season, which seems like a decent starting point.
```
ELO_MODEL_COLS = [
"prev_match_oppo_team",
"oppo_prev_match_oppo_team",
"prev_match_at_home",
"oppo_prev_match_at_home",
"date",
]
N_STEPS = 6
X_train, _ = data.train_data
n_features = X_train.shape[1] - len(ELO_MODEL_COLS)
rnn_pipeline = make_pipeline(
ColumnDropper(cols_to_drop=ELO_MODEL_COLS),
ColumnTransformer(
[
(
"ordinalencoder",
OrdinalEncoder(categories=[TEAM_NAMES, TEAM_NAMES, ROUND_TYPES, VENUES]),
CATEGORY_COLS,
)
],
remainder=StandardScaler(),
),
RNNRegressor(
n_features=n_features,
n_steps=N_STEPS,
verbose=1,
epochs=50,
),
)
rnn_pipeline.fit(*data.train_data)
None
graph_tf_model_history(rnn_pipeline.steps[-1][1].history, metrics=['regressor_team_match_accuracy'])
```
Performance is in the ballpark with other top models, though about 2% lower accuracy (this is from being trained on a full training data set and validated on 2019 data, however, rather than using a multi-fold CV from recent years, so it's not apples-to-apples).
### SearchCV for a better model structure
Given all the potential parameters for an RNN, a little trial-and-error isn't going to give us a realistic indication of the model's performance, so we'll do a small parameter search to hopefully give us a better structure (i.e. # layers, # cells per layer) to start fine tuning.
```
[param for param in rnn_pipeline.get_params() if 'rnnregressor__' in param]
PARAM_GRID = {
'rnnregressor__dropout': stats.uniform(0.0, 0.5),
'rnnregressor__n_cells': np.arange(20, 201),
'rnnregressor__n_hidden_layers': np.arange(1, 11),
'rnnregressor__n_steps': np.arange(2, 13), # ~10% to ~50% of a season
'rnnregressor__recurrent_dropout': stats.uniform(0.0, 0.5),
}
rnn_pipeline.set_params(rnnregressor__verbose=0)
X_train, y_train = data.train_data
random_search = RandomizedSearchCV(
rnn_pipeline,
PARAM_GRID,
scoring=match_accuracy_scorer,
n_jobs=-1,
cv=year_cv_split(X_train, CV_YEAR_RANGE),
refit=False,
verbose=2,
error_score='raise',
n_iter=5,
)
random_search.fit(X_train, y_train)
random_search.cv_results_
results = (
pd
.DataFrame(random_search.cv_results_)
.sort_values('rank_test_score')
.filter(regex='mean_test_score|param_')
.sort_index(axis=1)
)
results.head(20)
results.to_json('2.1-recurrent-neural-net-param-scores.json', indent=2, orient='records')
```
Well, that was underwhelming. I got better performance in 2019 with some configurations that I arrived at via trial-and-error. I'll give the best of those a try to see if I just got a bad batch of params.
### Try RNN from 2019
This was more-or-less the configuration for the final RNN tested in `notebooks/2019_season/6.1-all-data-rnn.ipynb`, which got accuracies in the low 70s. The mix of validation seasons that we're using here are harder to predict, and thus produce lower accuracy across models, but we'll see if we can still get comparable performance.
```
existing_rnn_pipeline = make_pipeline(
ColumnDropper(cols_to_drop=ELO_MODEL_COLS),
ColumnTransformer(
[
(
"ordinalencoder",
OrdinalEncoder(categories=[TEAM_NAMES, TEAM_NAMES, ROUND_TYPES, VENUES]),
CATEGORY_COLS,
)
],
remainder=StandardScaler(),
),
RNNRegressor(
n_features=n_features,
verbose=1,
epochs=50,
n_hidden_layers=2,
n_cells=100,
n_steps=5,
dropout=0.3,
recurrent_dropout=0.6,
patience=5,
),
)
existing_rnn = cross_validate(
rnn_pipeline,
X_train,
y_train,
scoring=match_accuracy_scorer,
n_jobs=None,
cv=year_cv_split(X_train, CV_YEAR_RANGE),
verbose=2,
error_score='raise',
)
existing_rnn
existing_rnn['test_score'].mean()
```
We have slightly better accuracy than from the best params from the CV, but it's still quite a bit worse than simple `Ridge`. I'm running low on time and patience, so I'm just going to go linear this season and look into deep learning model best practices to maybe come up with something useful for next season.
| github_jupyter |
# **Assignment - 2: Basic Data Understanding**
---
This assignment will get you familiarized with Python libraries and functions required for data visualization.
---
## Part 1 - Loading data
---
###Import the following libraries:
* ```numpy``` with an alias name ```np```,
* ```pandas``` with an alias name ```pd```,
* ```matplotlib.pyplot``` with an alias name ```plt```, and
* ```seaborn``` with an alias name ```sns```.
```
# Load the four libraries with their aliases
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
```
### Using the files ```train.csv``` and ```moviesData.csv```, peform the following:
* Load these file as ```pandas``` dataframes and store it in variables named ```df``` and ```movies``` respectively.
* Print the first ten rows of ```df```.
```
# Load the file as a dataframe
df = pd.read_csv('train.csv')
movies = pd.read_csv("moviesData.csv")
movies.head(5)
# Print the first ten rows of df
df.head(10)
```
### Using the dataframe ```df```, perform the following:
* Print the first five rows of the column ```MonthlyRate```.
* Find out the details of the column ```MonthlyRate``` like mean, maximum value, minimum value, etc.
```
# Print the first five rows of MonthlyRate
df['MonthlyRate'].head(10)
# Find the details of MonthlyRate
df['MonthlyRate'].describe()
df['MonthlyRate'].shape
```
---
## Part 2 - Cleaning and manipulating data
---
### Using the dataframe ```df```, peform the following:
* Check whether there are any missing values in ```df```.
* If yes, drop those values and print the size of ```df``` after dropping these.
```
# Check for missing values
df.isna()
# Drop the missing values
df.dropna()
# Print the size of df after dropping
print("shape : {} and size : {}".format(df.shape,df.size))
```
### Using the dataframe ```df```, peform the following:
* Add another column named ```MonthRateNew``` in ```df``` by subtracting the mean from ```MonthlyRate``` and dividing it by standard deviation.
```
# Add a column named MonthRateNew
df['MonthRateNew'] = ((df['MonthlyRate'].mean()-df['MonthlyRate'])/(df['MonthlyRate'].std()))
df
```
### Using the dataframe ```movies```, perform the following:
* Check whether there are any missing values in ```movies```.
* Find out the number of observations/rows having any of their features/columns missing.
* Drop the missing values and print the size of ```movies``` after dropping these.
* Instead of dropping the missing values, replace the missing values by their mean (or some suitable value).
```
# Check for missing values
movies.isna()
# Drop the missing values
movies.dropna()
print("shape : {} and size : {}".format(movies.shape,movies.size))
# Replace the missing values
# You can use SimpleImputer of sklearn for this
from sklearn.impute import SimpleImputer
imputer= SimpleImputer(missing_values=np.nan, strategy='mean')
Dependent = movies.iloc[:, 10].values
imputer=imputer.fit(Dependent)
Dependent=imputer.transform(Dependent)
movies
```
---
## Part 3 - Visualizing data
---
### Visualize the ```df``` by drawing the following plots:
* Plot a histogram of ```Age``` and find the range in which most people are there.
* Modify the histogram of ```Age``` by adding 30 bins.
* Draw a scatter plot between ```Age``` and ```Attrition``` and suitable labels to the axes. Find out whether people more than 50 years are more likely to leave the company. (```Attrition``` = 1 means people have left the company).
```
plt.hist(df.Age)
df.Age.describe()
# Plot and modify the histogram of Age
plt.hist(df.Age,color='red',orientation='vertical')
plt.xlabel('Age')
plt.ylabel('Frequancy of age')
plt.title("Histogram of Age")
plt.show()
#with bins
plt.hist(df.Age,color='blue',bins = 30,orientation='vertical')
plt.xlabel('Age')
plt.ylabel('Frequancy of age')
plt.title("Histogram of Age with bins = 30")
plt.show()
# Draw a scatter plot between Age and Attrition
plt.scatter(df.Age,df.Attrition,color="green")
plt.xlabel('Age')
plt.ylabel('Attrition')
plt.title("scatter plot between Age and Attrition")
plt.show()
```
### Visualize the ```df``` by following the steps given below:
* Get a series containing counts of unique values of ```Attrition```.
* Draw a countplot for ```Attrition``` using ```sns.countplot()```.
```
# Get a series of counts of values of Attrition
df["Attrition"].value_counts()
# Draw a countplot for Attrition
# You may use countplot of seaborn for this
sns.set(style="darkgrid")
sns.countplot(x="Attrition",data=df)
```
### Visualize the ```df``` by following the steps given below:
* Draw a cross tabulation of ```Attrition``` and ```BusinessTravel``` as bar charts. Find which value of ```BusinessTravel``` has highest number of people.
```
# Draw a cross tab of Attritiona and BusinessTravel
# You may use crosstab of pandas for this
pd.crosstab(df.Attrition, df.BusinessTravel)
```
### Visualize the ```df``` by drawing the following plot:
* Draw a stacked bar chart between ```Attrition``` and ```Gender``` columns.
```
# Draw a stacked bar chart between Attrition and Gender
plt.bar(df.Gender,df.Attrition,width=0.9)
plt.show()
```
### Visualize the ```df``` by drawing the following histogram:
* Draw a histogram of ```TotalWorkingYears``` with 30 bins.
* Draw a histogram of ```YearsAtCompany``` with 30 bins and find whether the values in ```YearsAtCompany``` are skewed.
```
# Draw a histogram of TotalWorkingYears with 30 bins
plt.hist(df.TotalWorkingYears,bins=30,color="blue")
plt.show()
# Draw a histogram of YearsAtCompany
plt.hist(df.YearsAtCompany,bins=30,color="yellow")
plt.show()
```
### Visualize the ```df``` by drawing the following boxplot:
* Draw a boxplot of ```MonthlyIncome``` for each ```Department``` and report whether there is/are outlier(s).
```
# Draw a boxplot of MonthlyIncome for each Department and report outliers
x1 = df[df['Department']=='Sales']
print(x1['Department'].count())
x2 = df[df['Department']=='Research & Development']
x2['Department'].count()
plt.boxplot(x1.MonthlyIncome)
plt.show()
plt.boxplot(x2.MonthlyIncome)
plt.show()
```
### Visualize the ```df``` by drawing the following piechart:
* Create a pie chart of the values in ```JobRole``` with suitable label and report which role has highest number of persons.
```
# Create a piechart of JobRole
# You will need to find the counts of unique values in JobRole.
job_count = df.JobRole.value_counts()
plt.pie(job_count
,autopct='%1.1f%%',
shadow=True)
plt.show()
```
| github_jupyter |
# Amino Acid Decode Prep
This notebook documents steps taken to create a set of keys for converting amino acid one letter codes to a larger set of information to be used by the CNN.
Data references:
Nelson, David L.; Cox, Michael M. (2000). Lehninger Principles of Biochemistry (3rd ed.). Worth Publishers. ISBN 978-1-57259-153-0.
Kyte J, Doolittle RF (May 1982). "A simple method for displaying the hydropathic character of a protein". Journal of Molecular Biology. 157 (1): 105–32. CiteSeerX 10.1.1.458.454. doi:10.1016/0022-2836(82)90515-0. PMID 7108955.
Meierhenrich, Uwe J. (2008). Amino acids and the asymmetry of life (1st ed.). Springer. ISBN 978-3-540-76885-2.
Biochemistry, Harpers (2015). Harpers Illustrated Biochemistry (30st ed.). Lange. ISBN 978-0-07-182534-4.
```
import pandas as pd
import numpy as np
data_r = pd.read_csv('AA_info.csv')
data_r.head()
data_r.columns
data_r.index = data_r['One Letter']
data_r.drop(['O', 'U'], inplace=True)
```
Numeric columns taken as is and scaled with standard scaler. Note that for pka.1, some amino acids do not have reported vlues. They are given a value of 30, which is consistent with their side chains expected to have significantly higher pka's than the ones with reported values.
```
data = data_r[['mass', 'PI', 'pka', 'pkb','pka.1','van der waal volume', 'Hydrophobicity']]
data.index = data_r['One Letter']
data
from sklearn.preprocessing import StandardScaler, MinMaxScaler
ss = MinMaxScaler()
data = pd.DataFrame(ss.fit_transform(data), columns=data.columns, index=data.index)
data
```
Boolean type columns are converted to 1,0 columns
```
# Hydrophobic is simple yes or no
for col in ['hydrophobic', 'polar']:
data[col] = pd.get_dummies(data_r[col])['Yes']
all(data['hydrophobic'] != data['polar'])
# dropping polar as it is an inverse of hydrophobic
data.drop('polar', axis=1, inplace=True)
# Aromatic/Aliphatic are each yes or no, '-' is a 0 for both
data = pd.concat((data, pd.get_dummies(data_r['aromatic or aliphatic'])[['Aliphatic', 'Aromatic']]), axis=1)
data
```
The ph column can be converted to an ordinal as 'basic' and 'acidic' represent two ends of a scale.
```
ordinal = []
for i in data_r['ph']:
if i == 'acidic':
ordinal.append(-2)
elif i == 'weak acidic':
ordinal.append(-1)
elif i == 'weak basic':
ordinal.append(1)
elif i == 'basic':
ordinal.append(2)
elif i == 'strongly basic':
ordinal.append(3)
else:
ordinal.append(0)
data['ph'] = ordinal
pd.concat((data['ph'], data_r['ph']), axis=1)
```
Finally, the amino acids contain much more information not captured in the above columns. Their one letter codes are encoded in dummy variables for the CNN to have the opportunity to 'learn' more information from these.
```
letters = pd.DataFrame(data=pd.get_dummies(data_r.index))
letters.index = data_r.index
letters
for c in letters.columns:
if letters[c][letters[c] == 1].name != c:
print(c)
data = pd.concat((data, letters), axis=1)
data
data.shape
data.to_csv('AA_keys.csv')
```
# Visualizations for write-up
```
import os
os.chdir('..')
from seq2mat import DataGenerator, seq_class, label_dict, aa_keys, aa_to_map
import numpy as np
params = {
'batch_size': 128,
'dim': (1502, 31),
'n_classes': 25,
'shuffle': True,
'struct': '1d',
'random_insert': False
}
data_gen_1d = DataGenerator([seq_class.index[10]], seq_class['label'], **params)
test = data_gen_1d[0][0][0]
test.shape
def get_mat_1d(sequence, dim):
mat = np.array([aa_keys.loc[aa] for aa in sequence])
# Allows for Conv2D or Conv1D layers
if len(dim) == 2:
mat = mat.reshape((mat.shape[0], mat.shape[1]))
elif len(dim) == 3:
mat = mat.reshape((mat.shape[0], mat.shape[1], 1))
else:
print("Incompatible Dimensions")
mat = 0
return mat
def get_mat_2d(sequence):
mat = np.zeros((2*len(sequence), 8, 6))
for i, aa in enumerate(sequence):
mat[2*i] = aa_to_map(aa)
return mat
seq_class['len'] = seq_class['sequence'].str.len()
seq = seq_class.sort_values('len')['sequence'][1000]
mat_1d = get_mat_1d(seq, dim=(1,31))
mat_1d.shape
import matplotlib.pyplot as plt
import pandas as pd
print(pd.DataFrame(mat_1d)[10])
for i in range(mat_1d.shape[1]):
if (mat_1d[:,i].max() - mat_1d[:,i].min()) == 0:
mat_1d[:,i] = (mat_1d[:,i] - mat_1d[:,i].min())
else:
mat_1d[:,i] = (mat_1d[:,i] - mat_1d[:,i].min())/(mat_1d[:,i].max() - mat_1d[:,i].min())
plt.figure()
plt.imshow(mat_1d, cmap='afmhot')
plt.savefig('1d_sequnce.png', dpi=300)
mat_2d = get_mat_2d(seq)
for i, a in enumerate(seq):
if a == 'W':
j = i
mat_2d[j*2].T
s = [255, 255, 0]
n = [0, 0, 255]
o = [255, 0, 0]
h = [0, 0, 0]
c = [50, 50, 50]
r = [1, 1, 1]
cmat = np.array([c, h, n, o, s, r])
cmat
mat_2d[:,:6,:].reshape(98,36)[14]
mat_2d[0,:6,:].shape
test = np.arange(6*3).reshape((6,3))
np.matmul(mat_2d, test).shape
image = np.matmul(mat_2d, cmat)
image = image/image.max()
image_sum = image.sum(axis=2)
for i in range(3):
image[:,:,i] = np.where(image_sum == 0, 1, image[:,:,i])
plt.figure(figsize=(5,10))
plt.imshow(image)
plt.savefig('2d_sequnce.png', dpi=300)
seq[:20]
mat_2d[0].T
np.where(image.sum(axis=2) == 0, 1, 0)
all_2d = get_mat_2d(aa_keys.index)
image = np.matmul(all_2d, cmat)
image = image/image.max()
image_sum = image.sum(axis=2)
for i in range(3):
image[:,:,i] = np.where(image_sum == 0, 1, image[:,:,i])
plt.figure(figsize=(5,10))
plt.imshow(image)
plt.yticks(np.arange(20)*2, aa_keys.index)
plt.savefig('2d_all.png', dpi=300)
aa_keys[aa_keys.columns[:11]]
all_1d = get_mat_1d(aa_keys.index, (1,1))
for i in range(all_1d.shape[1]):
if (all_1d[:,i].max() - all_1d[:,i].min()) == 0:
all_1d[:,i] = (all_1d[:,i] - all_1d[:,i].min())
else:
all_1d[:,i] = (all_1d[:,i] - all_1d[:,i].min())/(all_1d[:,i].max() - all_1d[:,i].min())
plt.rcParams['xtick.bottom'] = plt.rcParams['xtick.labelbottom'] = False
plt.rcParams['xtick.top'] = plt.rcParams['xtick.labeltop'] = True
plt.figure(figsize=(8,8))
plt.imshow(all_1d, cmap='afmhot')
plt.yticks(np.arange(20), aa_keys.index)
plt.xticks(np.arange(31), aa_keys.columns, rotation=90)
plt.savefig('1d_all.png', dpi=300)
```
| github_jupyter |
```
#hide
#skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
#export
from fastai.data.all import *
from fastai.optimizer import *
from fastai.learner import *
#hide
from nbdev.showdoc import *
#default_exp metrics
# default_cls_lvl 3
```
# Metrics
> Definition of the metrics that can be used in training models
## Core metric
This is where the function that converts scikit-learn metrics to fastai metrics is defined. You should skip this section unless you want to know all about the internals of fastai.
```
#export
import sklearn.metrics as skm
import scipy.stats as scs
#export torch_core
def flatten_check(inp, targ):
"Check that `out` and `targ` have the same number of elements and flatten them."
inp,targ = TensorBase(inp.contiguous()).view(-1),TensorBase(targ.contiguous()).view(-1)
test_eq(len(inp), len(targ))
return inp,targ
x1,x2 = torch.randn(5,4),torch.randn(20)
x1,x2 = flatten_check(x1,x2)
test_eq(x1.shape, [20])
test_eq(x2.shape, [20])
x1,x2 = torch.randn(5,4),torch.randn(21)
test_fail(lambda: flatten_check(x1,x2))
#export
mk_class('ActivationType', **{o:o.lower() for o in ['No', 'Sigmoid', 'Softmax', 'BinarySoftmax']},
doc="All possible activation classes for `AccumMetric")
#export
class AccumMetric(Metric):
"Stores predictions and targets on CPU in accumulate to perform final calculations with `func`."
def __init__(self, func, dim_argmax=None, activation=ActivationType.No, thresh=None, to_np=False,
invert_arg=False, flatten=True, **kwargs):
store_attr('func,dim_argmax,activation,thresh,flatten')
self.to_np,self.invert_args,self.kwargs = to_np,invert_arg,kwargs
def reset(self):
"Clear all targs and preds"
self.targs,self.preds = [],[]
def accumulate(self, learn):
"Store targs and preds from `learn`, using activation function and argmax as appropriate"
pred = learn.pred
if self.activation in [ActivationType.Softmax, ActivationType.BinarySoftmax]:
pred = F.softmax(pred, dim=self.dim_argmax)
if self.activation == ActivationType.BinarySoftmax: pred = pred[:, -1]
elif self.activation == ActivationType.Sigmoid: pred = torch.sigmoid(pred)
elif self.dim_argmax: pred = pred.argmax(dim=self.dim_argmax)
if self.thresh: pred = (pred >= self.thresh)
self.accum_values(pred,learn.y,learn)
def accum_values(self, preds, targs,learn=None):
"Store targs and preds"
to_d = learn.to_detach if learn is not None else to_detach
preds,targs = to_d(preds),to_d(targs)
if self.flatten: preds,targs = flatten_check(preds,targs)
self.preds.append(preds)
self.targs.append(targs)
def __call__(self, preds, targs):
"Calculate metric on one batch of data"
self.reset()
self.accum_values(preds,targs)
return self.value
@property
def value(self):
"Value of the metric using accumulated preds and targs"
if len(self.preds) == 0: return
preds,targs = torch.cat(self.preds),torch.cat(self.targs)
if self.to_np: preds,targs = preds.numpy(),targs.numpy()
return self.func(targs, preds, **self.kwargs) if self.invert_args else self.func(preds, targs, **self.kwargs)
@property
def name(self): return self.func.func.__name__ if hasattr(self.func, 'func') else self.func.__name__
```
`func` is only applied to the accumulated predictions/targets when the `value` attribute is asked for (so at the end of a validation/training phase, in use with `Learner` and its `Recorder`).The signature of `func` should be `inp,targ` (where `inp` are the predictions of the model and `targ` the corresponding labels).
For classification problems with single label, predictions need to be transformed with a softmax then an argmax before being compared to the targets. Since a softmax doesn't change the order of the numbers, we can just apply the argmax. Pass along `dim_argmax` to have this done by `AccumMetric` (usually -1 will work pretty well). If you need to pass to your metrics the probabilities and not the predictions, use `softmax=True`.
For classification problems with multiple labels, or if your targets are one-hot encoded, predictions may need to pass through a sigmoid (if it wasn't included in your model) then be compared to a given threshold (to decide between 0 and 1), this is done by `AccumMetric` if you pass `sigmoid=True` and/or a value for `thresh`.
If you want to use a metric function sklearn.metrics, you will need to convert predictions and labels to numpy arrays with `to_np=True`. Also, scikit-learn metrics adopt the convention `y_true`, `y_preds` which is the opposite from us, so you will need to pass `invert_arg=True` to make `AccumMetric` do the inversion for you.
```
#For testing: a fake learner and a metric that isn't an average
@delegates()
class TstLearner(Learner):
def __init__(self,dls=None,model=None,**kwargs): self.pred,self.xb,self.yb = None,None,None
def _l2_mean(x,y): return torch.sqrt((x.float()-y.float()).pow(2).mean())
#Go through a fake cycle with various batch sizes and computes the value of met
def compute_val(met, x1, x2):
met.reset()
vals = [0,6,15,20]
learn = TstLearner()
for i in range(3):
learn.pred,learn.yb = x1[vals[i]:vals[i+1]],(x2[vals[i]:vals[i+1]],)
met.accumulate(learn)
return met.value
x1,x2 = torch.randn(20,5),torch.randn(20,5)
tst = AccumMetric(_l2_mean)
test_close(compute_val(tst, x1, x2), _l2_mean(x1, x2))
test_eq(torch.cat(tst.preds), x1.view(-1))
test_eq(torch.cat(tst.targs), x2.view(-1))
#test argmax
x1,x2 = torch.randn(20,5),torch.randint(0, 5, (20,))
tst = AccumMetric(_l2_mean, dim_argmax=-1)
test_close(compute_val(tst, x1, x2), _l2_mean(x1.argmax(dim=-1), x2))
#test thresh
x1,x2 = torch.randn(20,5),torch.randint(0, 2, (20,5)).bool()
tst = AccumMetric(_l2_mean, thresh=0.5)
test_close(compute_val(tst, x1, x2), _l2_mean((x1 >= 0.5), x2))
#test sigmoid
x1,x2 = torch.randn(20,5),torch.randn(20,5)
tst = AccumMetric(_l2_mean, activation=ActivationType.Sigmoid)
test_close(compute_val(tst, x1, x2), _l2_mean(torch.sigmoid(x1), x2))
#test to_np
x1,x2 = torch.randn(20,5),torch.randn(20,5)
tst = AccumMetric(lambda x,y: isinstance(x, np.ndarray) and isinstance(y, np.ndarray), to_np=True)
assert compute_val(tst, x1, x2)
#test invert_arg
x1,x2 = torch.randn(20,5),torch.randn(20,5)
tst = AccumMetric(lambda x,y: torch.sqrt(x.pow(2).mean()))
test_close(compute_val(tst, x1, x2), torch.sqrt(x1.pow(2).mean()))
tst = AccumMetric(lambda x,y: torch.sqrt(x.pow(2).mean()), invert_arg=True)
test_close(compute_val(tst, x1, x2), torch.sqrt(x2.pow(2).mean()))
#hide
def _l2_mean(x,y): return torch.sqrt((x.argmax(dim=-1).float()-y.float()).pow(2).mean())
x1,x2 = torch.randn(20,5),torch.randint(0, 5, (20,))
tst = AccumMetric(_l2_mean, dim_argmax=-1, flatten=False, activation=ActivationType.Softmax)
test_close(compute_val(tst, x1, x2), _l2_mean(F.softmax(x1, dim=-1), x2))
#export
def skm_to_fastai(func, is_class=True, thresh=None, axis=-1, activation=None, **kwargs):
"Convert `func` from sklearn.metrics to a fastai metric"
dim_argmax = axis if is_class and thresh is None else None
if activation is None:
activation = ActivationType.Sigmoid if (is_class and thresh is not None) else ActivationType.No
return AccumMetric(func, dim_argmax=dim_argmax, activation=activation, thresh=thresh,
to_np=True, invert_arg=True, **kwargs)
```
This is the quickest way to use a scikit-learn metric in a fastai training loop. `is_class` indicates if you are in a classification problem or not. In this case:
- leaving `thresh` to `None` indicates it's a single-label classification problem and predictions will pass through an argmax over `axis` before being compared to the targets
- setting a value for `thresh` indicates it's a multi-label classification problem and predictions will pass through a sigmoid (can be deactivated with `sigmoid=False`) and be compared to `thresh` before being compared to the targets
If `is_class=False`, it indicates you are in a regression problem, and predictions are compared to the targets without being modified. In all cases, `kwargs` are extra keyword arguments passed to `func`.
```
tst_single = skm_to_fastai(skm.precision_score)
x1,x2 = torch.randn(20,2),torch.randint(0, 2, (20,))
test_close(compute_val(tst_single, x1, x2), skm.precision_score(x2, x1.argmax(dim=-1)))
tst_multi = skm_to_fastai(skm.precision_score, thresh=0.2)
x1,x2 = torch.randn(20),torch.randint(0, 2, (20,))
test_close(compute_val(tst_multi, x1, x2), skm.precision_score(x2, torch.sigmoid(x1) >= 0.2))
tst_multi = skm_to_fastai(skm.precision_score, thresh=0.2, activation=ActivationType.No)
x1,x2 = torch.randn(20),torch.randint(0, 2, (20,))
test_close(compute_val(tst_multi, x1, x2), skm.precision_score(x2, x1 >= 0.2))
tst_reg = skm_to_fastai(skm.r2_score, is_class=False)
x1,x2 = torch.randn(20,5),torch.randn(20,5)
test_close(compute_val(tst_reg, x1, x2), skm.r2_score(x2.view(-1), x1.view(-1)))
test_close(tst_reg(x1, x2), skm.r2_score(x2.view(-1), x1.view(-1)))
#export
def optim_metric(f, argname, bounds, tol=0.01, do_neg=True, get_x=False):
"Replace metric `f` with a version that optimizes argument `argname`"
def _f(preds, targs):
def minfunc(x):
kwargs = {argname:x}
res = f(preds, targs, **kwargs)
return -res if do_neg else res
optres = scipy.optimize.minimize_scalar(minfunc, bounds=bounds, method='bounded',
options={'xatol':0.01})
fun = -optres.fun if do_neg else optres.fun
return (fun,optres.x) if get_x else fun
_f.__name__ = f'opt_{f.__name__}'
return _f
```
## Single-label classification
> Warning: All functions defined in this section are intended for single-label classification and targets that are not one-hot encoded. For multi-label problems or one-hot encoded targets, use the version suffixed with multi.
> Warning: Many metrics in fastai are thin wrappers around sklearn functionality. However, sklearn metrics can handle python list strings, amongst other things, whereas fastai metrics work with PyTorch, and thus require tensors. The arguments that are passed to metrics are after all transformations, such as categories being converted to indices, have occurred. This means that when you pass a label of a metric, for instance, that you must pass indices, not strings. This can be converted with `vocab.map_obj`.
```
#export
def accuracy(inp, targ, axis=-1):
"Compute accuracy with `targ` when `pred` is bs * n_classes"
pred,targ = flatten_check(inp.argmax(dim=axis), targ)
return (pred == targ).float().mean()
#For testing
def change_targ(targ, n, c):
idx = torch.randperm(len(targ))[:n]
res = targ.clone()
for i in idx: res[i] = (res[i]+random.randint(1,c-1))%c
return res
x = torch.randn(4,5)
y = x.argmax(dim=1)
test_eq(accuracy(x,y), 1)
y1 = change_targ(y, 2, 5)
test_eq(accuracy(x,y1), 0.5)
test_eq(accuracy(x.unsqueeze(1).expand(4,2,5), torch.stack([y,y1], dim=1)), 0.75)
#export
def error_rate(inp, targ, axis=-1):
"1 - `accuracy`"
return 1 - accuracy(inp, targ, axis=axis)
x = torch.randn(4,5)
y = x.argmax(dim=1)
test_eq(error_rate(x,y), 0)
y1 = change_targ(y, 2, 5)
test_eq(error_rate(x,y1), 0.5)
test_eq(error_rate(x.unsqueeze(1).expand(4,2,5), torch.stack([y,y1], dim=1)), 0.25)
#export
def top_k_accuracy(inp, targ, k=5, axis=-1):
"Computes the Top-k accuracy (`targ` is in the top `k` predictions of `inp`)"
inp = inp.topk(k=k, dim=axis)[1]
targ = targ.unsqueeze(dim=axis).expand_as(inp)
return (inp == targ).sum(dim=-1).float().mean()
x = torch.randn(6,5)
y = torch.arange(0,6)
test_eq(top_k_accuracy(x[:5],y[:5]), 1)
test_eq(top_k_accuracy(x, y), 5/6)
#export
def APScoreBinary(axis=-1, average='macro', pos_label=1, sample_weight=None):
"Average Precision for single-label binary classification problems"
return skm_to_fastai(skm.average_precision_score, axis=axis, activation=ActivationType.BinarySoftmax,
average=average, pos_label=pos_label, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.average_precision_score.html#sklearn.metrics.average_precision_score) for more details.
```
#export
def BalancedAccuracy(axis=-1, sample_weight=None, adjusted=False):
"Balanced Accuracy for single-label binary classification problems"
return skm_to_fastai(skm.balanced_accuracy_score, axis=axis,
sample_weight=sample_weight, adjusted=adjusted)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.balanced_accuracy_score.html#sklearn.metrics.balanced_accuracy_score) for more details.
```
#export
def BrierScore(axis=-1, sample_weight=None, pos_label=None):
"Brier score for single-label classification problems"
return skm_to_fastai(skm.brier_score_loss, axis=axis,
sample_weight=sample_weight, pos_label=pos_label)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.brier_score_loss.html#sklearn.metrics.brier_score_loss) for more details.
```
#export
def CohenKappa(axis=-1, labels=None, weights=None, sample_weight=None):
"Cohen kappa for single-label classification problems"
return skm_to_fastai(skm.cohen_kappa_score, axis=axis, labels=labels, weights=weights,
sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.cohen_kappa_score.html#sklearn.metrics.cohen_kappa_score) for more details.
```
#export
def F1Score(axis=-1, labels=None, pos_label=1, average='binary', sample_weight=None):
"F1 score for single-label classification problems"
return skm_to_fastai(skm.f1_score, axis=axis,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html#sklearn.metrics.f1_score) for more details.
```
#export
def FBeta(beta, axis=-1, labels=None, pos_label=1, average='binary', sample_weight=None):
"FBeta score with `beta` for single-label classification problems"
return skm_to_fastai(skm.fbeta_score, axis=axis,
beta=beta, labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.fbeta_score.html#sklearn.metrics.fbeta_score) for more details.
```
#export
def HammingLoss(axis=-1, sample_weight=None):
"Hamming loss for single-label classification problems"
return skm_to_fastai(skm.hamming_loss, axis=axis,
sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.hamming_loss.html#sklearn.metrics.hamming_loss) for more details.
```
#export
def Jaccard(axis=-1, labels=None, pos_label=1, average='binary', sample_weight=None):
"Jaccard score for single-label classification problems"
return skm_to_fastai(skm.jaccard_score, axis=axis,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.jaccard_score.html#sklearn.metrics.jaccard_score) for more details.
```
#export
def Precision(axis=-1, labels=None, pos_label=1, average='binary', sample_weight=None):
"Precision for single-label classification problems"
return skm_to_fastai(skm.precision_score, axis=axis,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_score.html#sklearn.metrics.precision_score) for more details.
```
#export
def Recall(axis=-1, labels=None, pos_label=1, average='binary', sample_weight=None):
"Recall for single-label classification problems"
return skm_to_fastai(skm.recall_score, axis=axis,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html#sklearn.metrics.recall_score) for more details.
```
#export
def RocAuc(axis=-1, average='macro', sample_weight=None, max_fpr=None, multi_class='ovr'):
"Area Under the Receiver Operating Characteristic Curve for single-label multiclass classification problems"
assert multi_class in ['ovr', 'ovo']
return skm_to_fastai(skm.roc_auc_score, axis=axis, activation=ActivationType.Softmax, flatten=False,
average=average, sample_weight=sample_weight, max_fpr=max_fpr, multi_class=multi_class)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html#sklearn.metrics.roc_auc_score) for more details.
```
#export
def RocAucBinary(axis=-1, average='macro', sample_weight=None, max_fpr=None, multi_class='raise'):
"Area Under the Receiver Operating Characteristic Curve for single-label binary classification problems"
return skm_to_fastai(skm.roc_auc_score, axis=axis, activation=ActivationType.BinarySoftmax,
average=average, sample_weight=sample_weight, max_fpr=max_fpr, multi_class=multi_class)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html#sklearn.metrics.roc_auc_score) for more details.
```
#export
def MatthewsCorrCoef(sample_weight=None, **kwargs):
"Matthews correlation coefficient for single-label classification problems"
return skm_to_fastai(skm.matthews_corrcoef, sample_weight=sample_weight, **kwargs)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.matthews_corrcoef.html#sklearn.metrics.matthews_corrcoef) for more details.
## Multi-label classification
```
#export
def accuracy_multi(inp, targ, thresh=0.5, sigmoid=True):
"Compute accuracy when `inp` and `targ` are the same size."
inp,targ = flatten_check(inp,targ)
if sigmoid: inp = inp.sigmoid()
return ((inp>thresh)==targ.bool()).float().mean()
#For testing
def change_1h_targ(targ, n):
idx = torch.randperm(targ.numel())[:n]
res = targ.clone().view(-1)
for i in idx: res[i] = 1-res[i]
return res.view(targ.shape)
x = torch.randn(4,5)
y = (torch.sigmoid(x) >= 0.5).byte()
test_eq(accuracy_multi(x,y), 1)
test_eq(accuracy_multi(x,1-y), 0)
y1 = change_1h_targ(y, 5)
test_eq(accuracy_multi(x,y1), 0.75)
#Different thresh
y = (torch.sigmoid(x) >= 0.2).byte()
test_eq(accuracy_multi(x,y, thresh=0.2), 1)
test_eq(accuracy_multi(x,1-y, thresh=0.2), 0)
y1 = change_1h_targ(y, 5)
test_eq(accuracy_multi(x,y1, thresh=0.2), 0.75)
#No sigmoid
y = (x >= 0.5).byte()
test_eq(accuracy_multi(x,y, sigmoid=False), 1)
test_eq(accuracy_multi(x,1-y, sigmoid=False), 0)
y1 = change_1h_targ(y, 5)
test_eq(accuracy_multi(x,y1, sigmoid=False), 0.75)
#export
def APScoreMulti(sigmoid=True, average='macro', pos_label=1, sample_weight=None):
"Average Precision for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.average_precision_score, activation=activation, flatten=False,
average=average, pos_label=pos_label, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.average_precision_score.html#sklearn.metrics.average_precision_score) for more details.
```
#export
def BrierScoreMulti(thresh=0.5, sigmoid=True, sample_weight=None, pos_label=None):
"Brier score for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.brier_score_loss, thresh=thresh, activation=activation, flatten=False,
sample_weight=sample_weight, pos_label=pos_label)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.brier_score_loss.html#sklearn.metrics.brier_score_loss) for more details.
```
#export
def F1ScoreMulti(thresh=0.5, sigmoid=True, labels=None, pos_label=1, average='macro', sample_weight=None):
"F1 score for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.f1_score, thresh=thresh, activation=activation, flatten=False,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html#sklearn.metrics.f1_score) for more details.
```
#export
def FBetaMulti(beta, thresh=0.5, sigmoid=True, labels=None, pos_label=1, average='macro', sample_weight=None):
"FBeta score with `beta` for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.fbeta_score, thresh=thresh, activation=activation, flatten=False,
beta=beta, labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.fbeta_score.html#sklearn.metrics.fbeta_score) for more details.
```
#export
def HammingLossMulti(thresh=0.5, sigmoid=True, labels=None, sample_weight=None):
"Hamming loss for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.hamming_loss, thresh=thresh, activation=activation, flatten=False,
sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.hamming_loss.html#sklearn.metrics.hamming_loss) for more details.
```
#export
def JaccardMulti(thresh=0.5, sigmoid=True, labels=None, pos_label=1, average='macro', sample_weight=None):
"Jaccard score for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.jaccard_score, thresh=thresh, activation=activation, flatten=False,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.jaccard_score.html#sklearn.metrics.jaccard_score) for more details.
```
#export
def MatthewsCorrCoefMulti(thresh=0.5, sigmoid=True, sample_weight=None):
"Matthews correlation coefficient for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.matthews_corrcoef, thresh=thresh, activation=activation, flatten=False, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.matthews_corrcoef.html#sklearn.metrics.matthews_corrcoef) for more details.
```
#export
def PrecisionMulti(thresh=0.5, sigmoid=True, labels=None, pos_label=1, average='macro', sample_weight=None):
"Precision for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.precision_score, thresh=thresh, activation=activation, flatten=False,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_score.html#sklearn.metrics.precision_score) for more details.
```
#export
def RecallMulti(thresh=0.5, sigmoid=True, labels=None, pos_label=1, average='macro', sample_weight=None):
"Recall for multi-label classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.recall_score, thresh=thresh, activation=activation, flatten=False,
labels=labels, pos_label=pos_label, average=average, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html#sklearn.metrics.recall_score) for more details.
```
#export
def RocAucMulti(sigmoid=True, average='macro', sample_weight=None, max_fpr=None):
"Area Under the Receiver Operating Characteristic Curve for multi-label binary classification problems"
activation = ActivationType.Sigmoid if sigmoid else ActivationType.No
return skm_to_fastai(skm.roc_auc_score, activation=activation, flatten=False,
average=average, sample_weight=sample_weight, max_fpr=max_fpr)
roc_auc_metric = RocAucMulti(sigmoid=False)
x,y = torch.tensor([np.arange(start=0, stop=0.2, step=0.04)]*20), torch.tensor([0, 0, 1, 1]).repeat(5)
assert compute_val(roc_auc_metric, x, y) == 0.5
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html#sklearn.metrics.roc_auc_score) for more details.
## Regression
```
#export
def mse(inp,targ):
"Mean squared error between `inp` and `targ`."
return F.mse_loss(*flatten_check(inp,targ))
x1,x2 = torch.randn(4,5),torch.randn(4,5)
test_close(mse(x1,x2), (x1-x2).pow(2).mean())
#export
def _rmse(inp, targ): return torch.sqrt(F.mse_loss(inp, targ))
rmse = AccumMetric(_rmse)
rmse.__doc__ = "Root mean squared error"
show_doc(rmse, name="rmse")
x1,x2 = torch.randn(20,5),torch.randn(20,5)
test_eq(compute_val(rmse, x1, x2), torch.sqrt(F.mse_loss(x1,x2)))
#export
def mae(inp,targ):
"Mean absolute error between `inp` and `targ`."
inp,targ = flatten_check(inp,targ)
return torch.abs(inp - targ).mean()
x1,x2 = torch.randn(4,5),torch.randn(4,5)
test_eq(mae(x1,x2), torch.abs(x1-x2).mean())
#export
def msle(inp, targ):
"Mean squared logarithmic error between `inp` and `targ`."
inp,targ = flatten_check(inp,targ)
return F.mse_loss(torch.log(1 + inp), torch.log(1 + targ))
x1,x2 = torch.randn(4,5),torch.randn(4,5)
x1,x2 = torch.relu(x1),torch.relu(x2)
test_close(msle(x1,x2), (torch.log(x1+1)-torch.log(x2+1)).pow(2).mean())
#export
def _exp_rmspe(inp,targ):
inp,targ = torch.exp(inp),torch.exp(targ)
return torch.sqrt(((targ - inp)/targ).pow(2).mean())
exp_rmspe = AccumMetric(_exp_rmspe)
exp_rmspe.__doc__ = "Root mean square percentage error of the exponential of predictions and targets"
show_doc(exp_rmspe, name="exp_rmspe")
x1,x2 = torch.randn(20,5),torch.randn(20,5)
test_eq(compute_val(exp_rmspe, x1, x2), torch.sqrt((((torch.exp(x2) - torch.exp(x1))/torch.exp(x2))**2).mean()))
#export
def ExplainedVariance(sample_weight=None):
"Explained variance between predictions and targets"
return skm_to_fastai(skm.explained_variance_score, is_class=False, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.explained_variance_score.html#sklearn.metrics.explained_variance_score) for more details.
```
#export
def R2Score(sample_weight=None):
"R2 score between predictions and targets"
return skm_to_fastai(skm.r2_score, is_class=False, sample_weight=sample_weight)
```
See the [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.r2_score.html#sklearn.metrics.r2_score) for more details.
```
#export
@delegates(AccumMetric)
def PearsonCorrCoef(dim_argmax=None, **kwargs):
"Pearson correlation coefficient for regression problem"
def pearsonr(x,y): return scs.pearsonr(x,y)[0]
return AccumMetric(pearsonr, invert_arg=False, dim_argmax=dim_argmax, **kwargs)
```
See the [scipy documentation](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.pearsonr.html?highlight=pearson#scipy.stats.pearsonr) for more details.
```
x = torch.randint(-999, 999,(20,))
y = torch.randint(-999, 999,(20,))
test_eq(compute_val(PearsonCorrCoef(), x, y), scs.pearsonr(x.view(-1), y.view(-1))[0])
#export
@delegates(AccumMetric)
def SpearmanCorrCoef(dim_argmax=None, axis=0, nan_policy='propagate', **kwargs):
"Spearman correlation coefficient for regression problem"
def spearmanr(a,b=None,**kwargs): return scs.spearmanr(a,b,**kwargs)[0]
return AccumMetric(partial(spearmanr, axis=axis, nan_policy=nan_policy),
invert_arg=False, dim_argmax=dim_argmax, **kwargs)
```
See the [scipy documentation](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.spearmanr.html?highlight=spearman#scipy.stats.spearmanr) for more details.
```
x = torch.randint(-999, 999,(20,))
y = torch.randint(-999, 999,(20,))
test_eq(compute_val(SpearmanCorrCoef(), x, y), scs.spearmanr(x.view(-1), y.view(-1))[0])
```
## Segmentation
```
from fastai.vision.all import *
model = resnet34()
x = cast(torch.rand(1,3,128,128), TensorImage)
type(model(x))
#export
def foreground_acc(inp, targ, bkg_idx=0, axis=1):
"Computes non-background accuracy for multiclass segmentation"
targ = cast(targ.squeeze(1), TensorBase)
mask = targ != bkg_idx
return (inp.argmax(dim=axis)[mask]==targ[mask]).float().mean()
x = cast(torch.randn(4,5,3,3), TensorImage)
y = cast(x, TensorMask).argmax(dim=1)[:,None]
test_eq(foreground_acc(x,y), 1)
y[0] = 0 #the 0s are ignored so we get the same value
test_eq(foreground_acc(x,y), 1)
#export
class Dice(Metric):
"Dice coefficient metric for binary target in segmentation"
def __init__(self, axis=1): self.axis = axis
def reset(self): self.inter,self.union = 0,0
def accumulate(self, learn):
pred,targ = flatten_check(learn.pred.argmax(dim=self.axis), learn.y)
self.inter += (pred*targ).float().sum().item()
self.union += (pred+targ).float().sum().item()
@property
def value(self): return 2. * self.inter/self.union if self.union > 0 else None
x1 = cast(torch.randn(20,2,3,3), TensorImage)
x2 = cast(torch.randint(0, 2, (20, 3, 3)), TensorMask)
pred = x1.argmax(1)
inter = (pred*x2).float().sum().item()
union = (pred+x2).float().sum().item()
test_eq(compute_val(Dice(), x1, x2), 2*inter/union)
#export
class DiceMulti(Metric):
"Averaged Dice metric (Macro F1) for multiclass target in segmentation"
def __init__(self, axis=1): self.axis = axis
def reset(self): self.inter,self.union = {},{}
def accumulate(self, learn):
pred,targ = flatten_check(learn.pred.argmax(dim=self.axis), learn.y)
for c in range(learn.pred.shape[self.axis]):
p = torch.where(pred == c, 1, 0)
t = torch.where(targ == c, 1, 0)
c_inter = (p*t).float().sum().item()
c_union = (p+t).float().sum().item()
if c in self.inter:
self.inter[c] += c_inter
self.union[c] += c_union
else:
self.inter[c] = c_inter
self.union[c] = c_union
@property
def value(self):
binary_dice_scores = np.array([])
for c in self.inter:
binary_dice_scores = np.append(binary_dice_scores, 2.*self.inter[c]/self.union[c] if self.union[c] > 0 else np.nan)
return np.nanmean(binary_dice_scores)
```
The DiceMulti method implements the "Averaged F1: arithmetic mean over harmonic means" described in this publication: https://arxiv.org/pdf/1911.03347.pdf
```
x1a = torch.ones(20,1,1,1)
x1b = torch.clone(x1a)*0.5
x1c = torch.clone(x1a)*0.3
x1 = torch.cat((x1a,x1b,x1c),dim=1) # Prediction: 20xClass0
x2 = torch.zeros(20,1,1) # Target: 20xClass0
test_eq(compute_val(DiceMulti(), x1, x2), 1.)
x2 = torch.ones(20,1,1) # Target: 20xClass1
test_eq(compute_val(DiceMulti(), x1, x2), 0.)
x2a = torch.zeros(10,1,1)
x2b = torch.ones(5,1,1)
x2c = torch.ones(5,1,1) * 2
x2 = torch.cat((x2a,x2b,x2c),dim=0) # Target: 10xClass0, 5xClass1, 5xClass2
dice1 = (2*10)/(2*10+10) # Dice: 2*TP/(2*TP+FP+FN)
dice2 = 0
dice3 = 0
test_eq(compute_val(DiceMulti(), x1, x2), (dice1+dice2+dice3)/3)
#export
class JaccardCoeff(Dice):
"Implementation of the Jaccard coefficient that is lighter in RAM"
@property
def value(self): return self.inter/(self.union-self.inter) if self.union > 0 else None
x1 = cast(torch.randn(20,2,3,3), TensorImage)
x2 = cast(torch.randint(0, 2, (20, 3, 3)), TensorMask)
pred = x1.argmax(1)
inter = (pred*x2).float().sum().item()
union = (pred+x2).float().sum().item()
test_eq(compute_val(JaccardCoeff(), x1, x2), inter/(union-inter))
```
## NLP
```
#export
class CorpusBLEUMetric(Metric):
def __init__(self, vocab_sz=5000, axis=-1):
"BLEU Metric calculated over the validation corpus"
self.metric_name = 'CorpusBLEU'
self.axis, self.vocab_sz = axis, vocab_sz
self.pred_len,self.targ_len,self.samp_idx,self.corrects,self.counts, = 0,0,0,[0]*4,[0]*4
def reset(self):
self.pred_len,self.targ_len,self.corrects,self.counts = 0,0,[0]*4,[0]*4
class NGram():
def __init__(self, ngram, max_n=5000): self.ngram,self.max_n = ngram,max_n
def __eq__(self, other):
if len(self.ngram) != len(other.ngram): return False
return np.all(np.array(self.ngram) == np.array(other.ngram))
def __hash__(self): return int(sum([o * self.max_n**i for i,o in enumerate(self.ngram)]))
def get_grams(self, x, n, max_n=5000):
return x if n==1 else [self.NGram(x[i:i+n], max_n=max_n) for i in range(len(x)-n+1)]
def get_correct_ngrams(self, pred, targ, n, max_n=5000):
pred_grams,targ_grams = self.get_grams(pred, n, max_n=max_n),self.get_grams(targ, n, max_n=max_n)
pred_cnt,targ_cnt = Counter(pred_grams),Counter(targ_grams)
return sum([min(c, targ_cnt[g]) for g,c in pred_cnt.items()]),len(pred_grams)
def accumulate(self, learn):
if learn.training: return None
else:
last_output = learn.pred.argmax(dim=self.axis)
last_target = learn.y
for pred,targ in zip(last_output.cpu().numpy(),last_target.cpu().numpy()):
self.pred_len += len(pred)
self.targ_len += len(targ)
smooth_mteval = 1
for i in range(4):
c,t = self.get_correct_ngrams(pred, targ, i+1, max_n=self.vocab_sz)
if c == 0:
smooth_mteval *= 2
c = 1 / smooth_mteval # exp smoothing, method 3 from http://acl2014.org/acl2014/W14-33/pdf/W14-3346.pdf
self.corrects[i] += c
self.counts[i] += t
@property
def value(self):
if self.counts == 0: return None
elif max(self.corrects) == 0: return 0.0
else:
precs = [c/t for c,t in zip(self.corrects,self.counts)]
len_penalty = math.exp(1 - self.targ_len/self.pred_len) if self.pred_len < self.targ_len else 1
return len_penalty * ((precs[0]*precs[1]*precs[2]*precs[3]) ** 0.25)
def create_vcb_emb(pred, targ):
# create vocab "embedding" for predictions
vcb_sz = max(torch.unique(torch.cat([pred, targ])))+1
pred_emb=torch.zeros(pred.size()[0], pred.size()[1] ,vcb_sz)
for i,v in enumerate(pred):
pred_emb[i].scatter_(1, v.view(len(v),1),1)
return pred_emb
def compute_bleu_val(met, x1, x2):
met.reset()
learn = TstLearner()
learn.training=False
for i in range(len(x1)):
learn.pred,learn.yb = x1, (x2,)
met.accumulate(learn)
return met.value
targ = torch.tensor([[1,2,3,4,5,6,1,7,8]])
pred = torch.tensor([[1,9,3,4,5,6,1,10,8]])
pred_emb = create_vcb_emb(pred, targ)
test_close(compute_bleu_val(CorpusBLEUMetric(), pred_emb, targ), 0.48549)
targ = torch.tensor([[1,2,3,4,5,6,1,7,8],[1,2,3,4,5,6,1,7,8]])
pred = torch.tensor([[1,9,3,4,5,6,1,10,8],[1,9,3,4,5,6,1,10,8]])
pred_emb = create_vcb_emb(pred, targ)
test_close(compute_bleu_val(CorpusBLEUMetric(), pred_emb, targ), 0.48549)
```
The BLEU metric was introduced in [this article](https://www.aclweb.org/anthology/P02-1040) to come up with a way to evaluate the performance of translation models. It's based on the precision of n-grams in your prediction compared to your target. See the [fastai NLP course BLEU notebook](https://github.com/fastai/course-nlp/blob/master/bleu_metric.ipynb) for a more detailed description of BLEU.
The smoothing used in the precision calculation is the same as in [SacreBLEU](https://github.com/mjpost/sacrebleu/blob/32c54cdd0dfd6a9fadd5805f2ea189ac0df63907/sacrebleu/sacrebleu.py#L540-L542), which in turn is "method 3" from the [Chen & Cherry, 2014](http://acl2014.org/acl2014/W14-33/pdf/W14-3346.pdf) paper.
```
#export
class Perplexity(AvgLoss):
"Perplexity (exponential of cross-entropy loss) for Language Models"
@property
def value(self): return torch.exp(self.total/self.count) if self.count != 0 else None
@property
def name(self): return "perplexity"
perplexity = Perplexity()
x1,x2 = torch.randn(20,5),torch.randint(0, 5, (20,))
tst = perplexity
tst.reset()
vals = [0,6,15,20]
learn = TstLearner()
for i in range(3):
learn.yb = (x2[vals[i]:vals[i+1]],)
learn.loss = F.cross_entropy(x1[vals[i]:vals[i+1]],x2[vals[i]:vals[i+1]])
tst.accumulate(learn)
test_close(tst.value, torch.exp(F.cross_entropy(x1,x2)))
```
## LossMetrics -
```
#export
class LossMetric(AvgMetric):
"Create a metric from `loss_func.attr` named `nm`"
def __init__(self, attr, nm=None): store_attr('attr,nm')
def accumulate(self, learn):
bs = find_bs(learn.yb)
self.total += learn.to_detach(getattr(learn.loss_func, self.attr, 0))*bs
self.count += bs
@property
def name(self): return self.attr if self.nm is None else self.nm
#export
def LossMetrics(attrs, nms=None):
"List of `LossMetric` for each of `attrs` and `nms`"
if isinstance(attrs, str): attrs = attrs.split(',')
nms = attrs if nms is None else nms.split(',') if isinstance(nms, str) else nms
return [LossMetric(a, n) for a,n in zip(attrs,nms)]
#hide
from fastai.test_utils import *
class CombineL1L2(Module):
def forward(self, out, targ):
self.l1 = F.l1_loss(out, targ)
self.l2 = F.mse_loss(out, targ)
return self.l1+self.l2
learn = synth_learner(metrics=LossMetrics('l1,l2'))
learn.loss_func = CombineL1L2()
learn.fit(2)
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# Transformers by HuggingFace 🤗
In this module, we will get familiar with HuggingFace’s `transformers` library. We will first set up `transformers` package, and then check out some existing pretrained models. We will explore the magical world of the latest chatbot models, sentiment analyzers, and summarizers!
__What you will learn:__
As an ML practitioner, you will often want to explore existing models before developing your own models. By the end of this notebook, you will learn how to use existing models for your tasks using the `pipeline` API and `AutoClass` features.
The goal of this notebook is not to get familiar with how the library is implemented. Therefore, it is okay if you don't fully understand some details.
This notebook covers following topics:
- `transformers` library
- ProphetNet
- GPT-2
#### Before we start...
__transformer architecture:__
It would be good to review transformers architecture basics. The notebook assumes you have some basic understanding of what transformers are, and the theory is beyond the scope of this tutorial.
__transformer library:__
To get an idea about the how the `transformers` library is laid out, do check the [transformers](https://github.com/huggingface/transformers/) repo out . That's where the latest code `transformers` code exists!
__HuggingFace models:__
Do checkout all models supported by the `transformers` [library](https://huggingface.co/models). We will be using models from this throughout the notebook.
__setup:__
Alright, let's go ahead and install the `transformers` library
```
!pip --quiet install transformers
#recommended to have PyTorch, TensorFlow >= 2.0, or Flax installed
import transformers
```
# Pipeline
Sometimes, we might just want to access off-the-shelf NLP tools like Question Answering modules, sentiment analyzers etc. Transformers package makes it easy to access the existing models through `pipeline`.
```
from transformers import pipeline
classifier = pipeline("sentiment-analysis")
classifier("I am extremely happy today!")
classifier("alright, I want to go for a walk")
```
Notice how in the above example `alright, I want to go for a walk` is considered a `POSITIVE` sentiment. In reality, this sentence should be detected as a `NEUTRAL` sentiment.
Why is that? Let's dig into the implementation of `pipeline`. All the implementation related details are in the [`transformers/src/transformers/`](https://github.com/huggingface/transformers/tree/master/src/transformers). The `pipeline` implementation is in the `pipelines.py` file. In [this](https://github.com/huggingface/transformers/blob/443f67e887a030d8254eba126e5f2cdb8b70eb63/src/transformers/pipelines.py#L2729) line we see that the default sentimental analysis model is trained on [SST-2](https://paperswithcode.com/sota/sentiment-analysis-on-sst-2-binary) a binary sentiment classification dataset, and therefore the labels are either `POSITIVE` or `NEGATIVE`.
As a Machine Learning practitioner, you might want to tweak this model to fit to your own data distribution. In the subsequent modules of this course, we will learn more about pretraining and fine-tuning existing models; in particular, we will use `BERT` based models as it is the most popular NLP model in the last few years.
# HuggingFace Models
There's one really cool place to find all the latest models that are supported by `transformers`. It's the [huggingface models](https://huggingface.co/models) repo!. Let's get the `GPT-2` model, the predecessor of `GPT-3` and do some poetry generation! 🎶
Since the task is text generation, we will pass `text-generation` as the pipeline in our model and use `gpt2` model for generation. You can safely ignore the warning messages.
```
from transformers import pipeline, set_seed
generator = pipeline("text-generation", model="gpt2")
set_seed(1234)
my_poem = """
something in you wakes up
in the light of the day
and in the quite of the night
and I wonder, where did the years go?
I try to find them in the
chasms of your breath
and in the crevices of your broken voice
"""
generated_poem = generator(
my_poem,
max_length=200
)[0]["generated_text"]
print(generated_poem)
```
Also, try replacing `gpt2` with the following models and compare the results with different models:
- `lvwerra/gpt2-imdb`
- `sshleifer/tiny-gpt2`
`pipeline` supports a whole slew of tasks some of the following tasks:
- Named Entity Recognition
- Question-answering
- Summarization
- Translation
- Text Generation
- Text2Text Generation
- Zero-shot classification, and
- Conversational
## Summarization
Alright, let's move on to another interesting task. I often find it so cumbersome to go through giganitic emails or a huge news article. Thankfully for us, there are models that summarize at human-level competency.
To this end, we will use another `transformer` based model called `ProphetNet` to generate summaries. At the moment of writing this notebook, `ProphetNet` happens to be the state-of-the-art model for generating abstractive summaries. You can read more about the model in [this](https://arxiv.org/pdf/2001.04063.pdf) paper.
From here onwards, we will avoid using `pipeline` and get more familiar with some of underlying implementation.
Okay, first things first, find that long email or a text you have been meaning to read for a long time and paste it below:
```
TEXT_TO_SUMMARIZE = """
copy-paste-your-text-here
"""
# if your text is relatively shorter,
# choose a smaller summary length
summary_len = 20
from transformers import (
ProphetNetTokenizer,
ProphetNetForConditionalGeneration,
ProphetNetConfig
)
# download and load the pre-trained model
model = ProphetNetForConditionalGeneration.from_pretrained(
'microsoft/prophetnet-large-uncased-cnndm'
)
# download and load the tokenizer
tokenizer = ProphetNetTokenizer.from_pretrained(
'microsoft/prophetnet-large-uncased-cnndm'
)
# tokenize the input text
inputs = tokenizer([TEXT_TO_SUMMARIZE], max_length=1000, return_tensors='pt', truncation=True)
# Generate Summary
summary_ids = model.generate(
inputs['input_ids'],
num_beams=4,
max_length=summary_len,
early_stopping=True
)
tokenizer.batch_decode(summary_ids, skip_special_tokens=True)
```
A few things to note here:
- you are not expected to understand the workings of these models. For now, you should consider these models as a black-box and get comfortable with just using existing models in HuggingFace's model repository. Go to [HuggingFace models](https://huggingface.co/models) and replace `microsoft/prophetnet-large-uncased-cnndm` and other summarization models.
- Observe that our summarizer uses a `model` and a `tokenizer`. This is a common pattern for any projects built with transformers as we will see in our next lesson.
In the subsequent notebooks, we will get familiar with `transformers` library. We will get to know the library better by addressing sentence classification and token classification taks.Text generation and summarization models that we have seen in this notebook will not be explored further as they are beyond the scope of this learning package.
# Homework Time! ☕️
Facebook recently released a great open-domain chatbot called Blenderbot. You can read the paper [here](https://arxiv.org/pdf/2004.13637.pdf). The exciting news is that a HuggingFace model exists for Blenderbot!
The homework for this module is to use Blenderbot to create a small-talk chatbot!
Here is your [hint](https://huggingface.co/transformers/master/model_doc/blenderbot.html)!
__Note regarding the exercise:__
Since this exercise was first written, the tokenizers corresponding to Blenderbot have been updated, you might run into the following error: `TypeError: forward() got an unexpected keyword argument 'token_type_ids'`. To resolve it simply, remove the `token_type_ids` from the input:
```python
inputs.pop("token_type_ids")
```
## Additional Resources
- Here is a great [video](https://www.youtube.com/watch?v=KMY2Knr4iAs_) covering the implementational aspects of transformers with a code walk through
- [Attention is All You Need](https://papers.nips.cc/paper/2017/file/3f5ee243547dee91fbd053c1c4a845aa-Paper.pdf)---the paper that introduced the transformer architecture
| github_jupyter |
# Autofaiss getting started
## Information
**This Demo notebook automatically creates a Faiss knn indices with the most optimal similarity search parameters.**
It selects the best indexing parameters to achieve the highest recalls given memory and query speed constraints.
Github: https://github.com/criteo/autofaiss
## Parameters
```
#@title Index parameters
max_index_query_time_ms = 10 #@param {type: "number"}
max_index_memory_usage = "10MB" #@param
metric_type = "l2" #@param ['ip', 'l2']
```
## Embeddings creation (add your own embeddings here)
```
import numpy as np
# Create embeddings
embeddings = np.float32(np.random.rand(4000, 100))
```
## Save your embeddings on the disk
```
# Create a new folder
import os
import shutil
embeddings_dir = "embeddings_folder"
if os.path.exists(embeddings_dir):
shutil.rmtree(embeddings_dir)
os.makedirs(embeddings_dir)
# Save your embeddings
# You can split you embeddings in several parts if it is too big
# The data will be read in the lexicographical order of the filenames
np.save(f"{embeddings_dir}/part1.npy", embeddings[:2000])
np.save(f"{embeddings_dir}/part2.npy", embeddings[2000:])
```
## Build the KNN index with Autofaiss
```
os.makedirs("my_index_folder", exist_ok=True)
# Install autofaiss
!pip install autofaiss &> /dev/null
# Build a KNN index
!autofaiss build_index --embeddings={embeddings_dir} \
--index_path="knn.index" \
--index_infos_path="infos.json" \
--metric_type={metric_type} \
--max_index_query_time_ms=5 \
--max_index_memory_usage={max_index_memory_usage}
```
## Load the index and play with it
```
import faiss
import glob
import numpy as np
my_index = faiss.read_index("knn.index")
query_vector = np.float32(np.random.rand(1, 100))
k = 5
distances, indices = my_index.search(query_vector, k)
print(f"Top {k} elements in the dataset for max inner product search:")
for i, (dist, indice) in enumerate(zip(distances[0], indices[0])):
print(f"{i+1}: Vector number {indice:4} with distance {dist}")
```
## (Bonus) Python version of the CLI
```
from autofaiss import build_index
build_index(embeddings="embeddings_folder",
index_path="knn.index",
index_infos_path="infos.json",
max_index_query_time_ms = max_index_query_time_ms,
max_index_memory_usage = max_index_memory_usage,
metric_type=metric_type)
```
| github_jupyter |
# Quickstart to `torchdyn`
`torchdyn` is a PyTorch library dedicated to neural differential equations and equilibrium models.
Central to the `torchdyn` approach are continuous and implicit neural networks, where *depth* is taken to its infinite limit.
This notebook serves as a gentle introduction to NeuralODE, concluding with a small overview of `torchdyn` features.
```
from torchdyn.core import NeuralODE
from torchdyn.datasets import *
from torchdyn import *
%load_ext autoreload
%autoreload 2
# quick run for automated notebook validation
dry_run = False
```
## Generate data from a static toy dataset
We’ll be generating data from toy datasets. In torchdyn, we provide a wide range of datasets often use to benchmark and understand Neural ODEs. Here we will use the classic moons dataset and train a Neural ODE for binary classification
```
d = ToyDataset()
X, yn = d.generate(n_samples=512, noise=1e-1, dataset_type='moons')
import matplotlib.pyplot as plt
colors = ['orange', 'blue']
fig = plt.figure(figsize=(3,3))
ax = fig.add_subplot(111)
for i in range(len(X)):
ax.scatter(X[i,0], X[i,1], s=1, color=colors[yn[i].int()])
```
Generated data can be easily loaded in the dataloader with standard `PyTorch` calls
```
import torch
import torch.utils.data as data
device = torch.device("cpu") # all of this works in GPU as well :)
X_train = torch.Tensor(X).to(device)
y_train = torch.LongTensor(yn.long()).to(device)
train = data.TensorDataset(X_train, y_train)
trainloader = data.DataLoader(train, batch_size=len(X), shuffle=True)
```
We utilize [Pytorch Lightning](https://github.com/PyTorchLightning/pytorch-lightning) to handle training loops, logging and general bookkeeping. This allows `torchdyn` and Neural Differential Equations to have access to modern best practices for training and experiment reproducibility.
In particular, we combine modular `torchdyn` models with `LightningModules` via a `Learner` class:
```
import torch.nn as nn
import pytorch_lightning as pl
class Learner(pl.LightningModule):
def __init__(self, t_span:torch.Tensor, model:nn.Module):
super().__init__()
self.model, self.t_span = model, t_span
def forward(self, x):
return self.model(x)
def training_step(self, batch, batch_idx):
x, y = batch
t_eval, y_hat = self.model(x, t_span)
y_hat = y_hat[-1] # select last point of solution trajectory
loss = nn.CrossEntropyLoss()(y_hat, y)
return {'loss': loss}
def configure_optimizers(self):
return torch.optim.Adam(self.model.parameters(), lr=0.01)
def train_dataloader(self):
return trainloader
```
## Define a Neural ODE
Analogously to most forward neural models we want to realize a map
$$
x \mapsto \hat y
$$
where $\hat y$ becomes the best approximation of a true output $y$ given an input $x$.
In torchdyn you can define very simple Neural ODE models of the form
$$ \left\{
\begin{aligned}
\dot{z}(t) &= f(z(t), \theta)\\
z(0) &= x\\
\hat y & = z(1)
\end{aligned}
\right. \quad t\in[0,1]
$$
by just specifying a neural network $f$ and giving some simple settings.
**Note:** This Neural ODE model is of *depth-invariant* type as neither $f$ explicitly depend on $s$ nor the parameters $\theta$ are depth-varying. Together with their *depth-variant* counterpart with $s$ concatenated in the vector field was first proposed and implemented by [[Chen T. Q. et al, 2018]](https://arxiv.org/abs/1806.07366)
### Define the vector field (DEFunc)
The first step is to define any PyTorch `torch.nn.Module`. This takes the role of the Neural ODE vector field $f(h,\theta)$
```
f = nn.Sequential(
nn.Linear(2, 16),
nn.Tanh(),
nn.Linear(16, 2)
)
t_span = torch.linspace(0, 1, 5)
```
In this case we chose $f$ to be a simple MLP with one hidden layer and $\tanh$ activation
### Define the NeuralDE
The final step to define a Neural ODE is to instantiate the torchdyn's class `NeuralDE` passing some customization arguments and `f` itself.
In this case we specify:
* we compute backward gradients with the `'adjoint'` method.
* we will use the `'dopri5'` (Dormand-Prince) ODE solver from `torchdyn`, with no additional options;
```
model = NeuralODE(f, sensitivity='adjoint', solver='dopri5').to(device)
```
## Train the Model
With the same forward method of `NeuralDE` objects you can quickly evaluate the entire trajectory of each data point in `X_train` on an interval `t_span`
```
t_span = torch.linspace(0,1,100)
t_eval, trajectory = model(X_train, t_span)
trajectory = trajectory.detach().cpu()
```
The numerical method used to solve a `NeuralODE` have great effect on its speed. Try retraining with the following
```
f = nn.Sequential(
nn.Linear(2, 16),
nn.Tanh(),
nn.Linear(16, 2)
)
model = NeuralODE(f, sensitivity='adjoint', solver='rk4', solver_adjoint='dopri5', atol_adjoint=1e-4, rtol_adjoint=1e-4).to(device)
learn = Learner(t_span, model)
if dry_run: trainer = pl.Trainer(min_epochs=1, max_epochs=1)
else: trainer = pl.Trainer(min_epochs=200, max_epochs=300)
trainer.fit(learn)
```
### Plot the Training Results
We can first plot the trajectories of the data points in the depth domain $s$
```
t_eval, trajectory = model(X_train, t_span)
trajectory = trajectory.detach().cpu()
color=['orange', 'blue']
fig = plt.figure(figsize=(10,2))
ax0 = fig.add_subplot(121)
ax1 = fig.add_subplot(122)
for i in range(500):
ax0.plot(t_span, trajectory[:,i,0], color=color[int(yn[i])], alpha=.1);
ax1.plot(t_span, trajectory[:,i,1], color=color[int(yn[i])], alpha=.1);
ax0.set_xlabel(r"$t$ [Depth]") ; ax0.set_ylabel(r"$h_0(t)$")
ax1.set_xlabel(r"$t$ [Depth]") ; ax1.set_ylabel(r"$z_1(t)$")
ax0.set_title("Dimension 0") ; ax1.set_title("Dimension 1")
```
Then the trajectory in the *state-space*
As you can see, the Neural ODE steers the data-points into regions of null loss with a continuous flow in the depth domain. Finally, we can also plot the learned vector field $f$
```
# evaluate vector field
n_pts = 50
x = torch.linspace(trajectory[:,:,0].min(), trajectory[:,:,0].max(), n_pts)
y = torch.linspace(trajectory[:,:,1].min(), trajectory[:,:,1].max(), n_pts)
X, Y = torch.meshgrid(x, y) ; z = torch.cat([X.reshape(-1,1), Y.reshape(-1,1)], 1)
f = model.vf(0,z.to(device)).cpu().detach()
fx, fy = f[:,0], f[:,1] ; fx, fy = fx.reshape(n_pts , n_pts), fy.reshape(n_pts, n_pts)
# plot vector field and its intensity
fig = plt.figure(figsize=(4, 4)) ; ax = fig.add_subplot(111)
ax.streamplot(X.numpy().T, Y.numpy().T, fx.numpy().T, fy.numpy().T, color='black')
ax.contourf(X.T, Y.T, torch.sqrt(fx.T**2+fy.T**2), cmap='RdYlBu')
```
**Sweet! You trained your first Neural ODEs! Now you can proceed and learn about more advanced models with the next tutorials**
## More about `torchdyn`
```
import time
from torchdyn.numerics import Euler, RungeKutta4, Tsitouras45, DormandPrince45, MSZero, Euler, HyperEuler
from torchdyn.numerics import odeint, odeint_mshooting, Lorenz
from torchdyn.core import ODEProblem, MultipleShootingProblem
```
But wait! `torchdyn` has way more than `NeuralODEs`. If you wish to solve generic differential equations parallelizable both in space (initial conditions) as well in time, with parallel, but do not need neural networks inside the vector field, you can use our functional API like so:
```
x0 = torch.randn(8, 3) + 15
t_span = torch.linspace(0, 3, 2000)
sys = Lorenz()
t0 = time.time()
t_eval, accurate_sol = odeint(sys, x0, t_span, solver='dopri5', atol=1e-6, rtol=1e-6)
accurate_sol_time = time.time() - t0
t0 = time.time()
t_eval, base_sol = odeint(sys, x0, t_span, solver='euler')
base_sol_time = time.time() - t0
t0 = time.time()
t_eval, rk4_sol = odeint(sys, x0, t_span, solver='rk4')
rk4_sol_time = time.time() - t0
t0 = time.time()
t_eval, dp5_low_sol = odeint(sys, x0, t_span, solver='dopri5', atol=1e-3, rtol=1e-3)
dp5_low_time = time.time() - t0
t0 = time.time()
t_eval, ms_sol = odeint_mshooting(sys, x0, t_span, solver='mszero', fine_steps=2, maxiter=4)
ms_sol_time = time.time() - t0
```
Alternatively, you can wrap your vector field in a specific `*Problem` to perform sensitivity analysis and optimize for terminal as well as integral objectives:
```
prob = ODEProblem(sys, sensitivity='interpolated_adjoint', solver='dopri5', atol=1e-3, rtol=1e-3,
solver_adjoint='tsit5', atol_adjoint=1e-3, rtol_adjoint=1e-3)
t0 = time.time()
t_eval, sol_torchdyn = prob.odeint(x0, t_span)
t_end1 = time.time() - t0
```
Our numerics suite includes other tools, such as a `odeint_hybrid` for hybrid systems (potentially stochastic and multi-mode). We have built our numerics suite from the ground up to be compatible with hybridized methods such as hypersolvers, where a base solver works in tandem with neural approximators to increase accuracy while retaining improved extrapolation capabilities. In fact, these methods can be called from the same API:
```
class VanillaHyperNet(nn.Module):
def __init__(self, net):
super().__init__()
self.net = net
for p in self.net.parameters():
torch.nn.init.uniform_(p, 0, 1e-5)
def forward(self, t, x):
return self.net(x)
net = nn.Sequential(nn.Linear(3, 64), nn.Softplus(), nn.Linear(64, 64), nn.Softplus(), nn.Linear(64, 3))
hypersolver = HyperEuler(VanillaHyperNet(net))
t_eval, sol = odeint(sys, x0, t_span, solver=hypersolver) # note: this has to be trained!
```
We also provide an extensive set of tutorial subdivided into modules. Each tutorials deals with a specific aspect of continuous or implicit models, or showcases applications (control, generative modeling, forecasting, optimal control of ODEs and PDEs, graph node classification). Check `torchdyn/tutorials` for more information.
| github_jupyter |
**<div style='font-size:200%'>Batch Transform using the gluonts entrypoint</div>**
In this notebook, we first register a model artifact into a SageMaker model, then perform a batch evaluation. Optionally, we deregister the model.
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
import logging
import sagemaker as sm
from sagemaker.mxnet.model import MXNetModel
from smallmatter.sm import get_sm_execution_role, get_model_and_output_tgz
# smallmatter.sm.get_sm_execution_role() will:
# - on SageMaker classic notebook instance, simply call sagemaker.get_execution_role()
# - outside of SageMaker classic notebook instance, return the first role whose name
# startswith "AmazonSageMaker-ExecutionRole-"
role: str = get_sm_execution_role()
sess = sm.Session()
region: str = sess.boto_session.region_name
```
# Global config
```
bucket = 'BUCKETNAME'
# I/O S3 paths MUST have trailing '/'
bt_input = f's3://{bucket}/gluonts-examples-dataset/synthetic-dataset/test/' # Reuse test-split from notebook 01.
bt_output = f's3://{bucket}/bt_output/'
# Use artifacts from this training job.
train_job = "mxnet-training-2021-09-29-08-04-10-326"
```
# Observe training results
As in any SageMaker training job, entrypoint script will generate two artifacts in the S3: `model.tar.gz` and `output.tar.gz`.
The `model.tar.gz` contains the persisted model that can be used later on for inference.
The `output.tar.gz` contains the following:
- individual plot of each test timeseries
- montage of plots of all test timeseries
- backtest evaluation metrics.
```
model_tgz, output_tgz = (str(path) for path in get_model_and_output_tgz(train_job))
%set_env MODEL_S3=$model_tgz
%set_env OUTPUT_S3=$output_tgz
%%bash
echo -e "\nModel artifacts $MODEL_S3:"
aws s3 cp $MODEL_S3 - | tar -tzvf -
echo -e "\nOutput $OUTPUT_S3:"
aws s3 cp $OUTPUT_S3 - | tar -tzvf - | head # NOTE: "[Errno 32] Broken pipe" can be safely ignored.
```
# Create model
Let SDK auto-generates the new model name, so we can safely make this notebook reentrant.
```
mxnet_model = MXNetModel(
model_data=model_tgz,
role=role,
entry_point='inference.py',
source_dir='../src/entrypoint',
py_version="py3",
framework_version="1.7.0",
sagemaker_session=sess,
container_log_level=logging.DEBUG, # Comment this line to reduce the amount of logs in CloudWatch.
)
```
A bit of reverse engineering, to confirm env. vars that the model will end-up using. Will be useful when the time comes where I need to do all these in boto3 or botocore.
```
# Before create model
mxnet_model._framework_env_vars()
# Create model
mxnet_model._create_sagemaker_model(instance_type='ml.m5.xlarge')
# Model name
mxnet_model.name
mxnet_model._framework_env_vars()
# Peek into model's model.tar.gz (which is different from training artifact model.tar.gz).
model_s3 = mxnet_model._framework_env_vars()['SAGEMAKER_SUBMIT_DIRECTORY']
%set_env MODEL_S3=$model_s3
!aws s3 cp $MODEL_S3 - | tar -tzvf -
```
# Batch Transform
```
instance_type = 'ml.m5.4xlarge'
# By default, GluonTS runs inference with multiple cores.
# On ml.m5.4xlarge with 8 cpu cores (= vcpu_count / 2), a single request
# already reported 75% CPU utilization (viewed in CloudWatch metrics; measured
# with gluonts-0.5).
#
# Note that this number was specific to the gluonts-0.5's DeepAR example.
# Other algorithms and gluonts versions may need different configurations.
max_concurrent_transforms = 1
bt = mxnet_model.transformer(
instance_count=1,
instance_type='ml.m5.4xlarge',
strategy='MultiRecord',
assemble_with='Line',
output_path=bt_output,
accept='application/json',
env={
'SAGEMAKER_MODEL_SERVER_TIMEOUT': '3600',
'SAGEMAKER_MODEL_SERVER_WORKERS': str(max_concurrent_transforms),
},
max_payload=1,
max_concurrent_transforms=max_concurrent_transforms,
)
bt.base_transform_job_name
# Setting wait=False (which is the default) frees this notebook
# from getting blocked by the transform job.
bt.transform(
data=bt_input,
data_type='S3Prefix',
content_type='application/json',
split_type='Line',
join_source='Input',
output_filter='$',
wait=False,
logs=False,
)
```
By setting `wait=False` (which is the default for transform jobs), while the transform job is running, you can may shutdown this notebook's kernel, close this notebook, and go to the SageMaker console to monitor the batch-transform progress. The batch-transform job's console also contains links to CloudWatch log.
Once the job finishes, from the batch-transform job's console, you can follow through the S3 output location, where you can preview or download the output.
# Delete model
Uncomment and execute cell to "deregister" the model from SageMaker. The inference model artifacts remain untouched in S3.
```
#mxnet_model.delete_model()
```
| github_jupyter |
# Isa Create Mode
```
from isatools.model import *
from isatools.create.models import *
from isatools.create.models import StudyCell
```
# Treatments
Here we will try to create a few treatments:
```
NAME = 'name'
treatment_0_conf = dict(TYPE=INTERVENTIONS['CHEMICAL'], FACTORS_0_VALUE='nitroglycerin',FACTORS_1_VALUE=5, FACTORS_1_UNIT='kg/m^3',
FACTORS_2_VALUE=100.0, FACTORS_2_UNIT = 's')
treatment_1_conf = dict(TYPE=INTERVENTIONS['CHEMICAL'], FACTORS_0_VALUE='aether',FACTORS_1_VALUE=1.25, FACTORS_1_UNIT='g',
FACTORS_2_VALUE=60000.0, FACTORS_2_UNIT = 's')
treatment_2_conf = dict(TYPE=INTERVENTIONS['RADIOLOGICAL'], FACTORS_0_VALUE='gamma ray',FACTORS_1_VALUE=10, FACTORS_1_UNIT='gy',
FACTORS_2_VALUE=60000.0, FACTORS_2_UNIT = 's')
treatment_3_conf = dict(TYPE=INTERVENTIONS['DIET'], FACTORS_0_VALUE='glucose',FACTORS_1_VALUE=0.25, FACTORS_1_UNIT='kg',
FACTORS_2_VALUE=30, FACTORS_2_UNIT = 'day')
treatment_3_conf.keys()
Treatment(treatment_3_conf.get('TYPE', None))
Treatment(treatment_3_conf['TYPE']).treatment_type
treatment_3_conf.items()
treatments = [Treatment(treatment_type=conf.get('TYPE',None), factor_values=(
FactorValue(factor_name=StudyFactor(name=BASE_FACTORS_[0]['name']), value=conf['FACTORS_0_VALUE']),
FactorValue(factor_name=StudyFactor(BASE_FACTORS_[1]['name']), value=conf['FACTORS_1_VALUE'], unit=conf['FACTORS_1_UNIT']),
FactorValue(factor_name=StudyFactor(BASE_FACTORS_[2]['name']), value=conf['FACTORS_2_VALUE'], unit=conf['FACTORS_2_UNIT'])
)) for conf in [treatment_0_conf, treatment_1_conf, treatment_2_conf, treatment_3_conf]]
treatments
```
## Study Epochs:
Here we will create three epochs. The central epoch will have two concomitant treatments.
```
sample_plan = SampleAssayPlan()
sample_plan
epoch_0 = StudyCell(name='first', elements=[treatments[0]])
epoch_1 = StudyCell(name='second', elements=(treatments[1], treatments[2]))
epoch_2 = StudyCell(name='third', elements=(treatments[2], treatments[2]))
epoch_0
epoch_1
epoch_2
```
# Study Design
Here we compose a study design with the three study epochs
```
study_design = StudyDesign()
```
# Study Design Factory
Here we use a `StudyDesignFactory` class to generate study designs:
```
factory = StudyDesignFactory(treatments, sample_plan)
factory
crossover_design = factory.compute_crossover_design(screen=True, follow_up=True)
parallel_design = factory.compute_parallel_design(screen=True, follow_up=True)
crossover_design
study_design = StudyDesign(crossover_design)
study_design
```
| github_jupyter |
# Generational changes among the religious and non-religious
Allen Downey
Copyright 2020
[MIT License](https://en.wikipedia.org/wiki/MIT_License)
## Introduction
In this notebook I use data from the GSS to explore differences in beliefs and attitudes between Christians and Nones (people with no religious affiliation) and look at generational changes in those differences.
### Setup
If you are running this notebook in Colab, the following cell downloads the `empiricaldist` library.
If you are running in another environment, you will need to install it yourself.
```
# If we're running in Colab, install empiricaldist
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
!pip install empiricaldist
```
The following cell downloads `utils.py`, which contains function I use in many data science projects.
```
# Load some utility code
import os
file = 'utils.py'
if not os.path.exists(file):
!wget https://github.com/AllenDowney/PoliticalAlignmentCaseStudy/raw/master/utils.py
```
If everything we need is installed, the following cell should run without error.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from empiricaldist import Cdf
from utils import decorate
from utils import underride
from utils import values
```
### Loading the data
The following cell downloads an HDF5 file that contains the data we need.
The HDF file is created by a notebook called `01_clean.ipynb`, which you should find in the same directory as this notebook, if you want to see the details.
```
# Load the data file
import os
datafile = 'gss_eda.hdf5'
if not os.path.exists(datafile):
!wget https://github.com/AllenDowney/PoliticalAlignmentCaseStudy/raw/master/gss_eda.hdf5
```
Now we can read the data.
```
gss = pd.read_hdf(datafile, 'gss')
gss.shape
values(gss['year'])
```
For modeling purposes, I'll use data from 1998 to 2018 and respondents born during or after 1940.
These choices are a compromise between using more data, so the results are more likely to be statistically valid, and using recent data, so the models are not too influenced by irrelevant history.
I classify as "Christian" anyone who reports that their religious affiliation is Catholic, Protestant, or Christian.
```
def prepare_df(df):
# compute quadratic and cubic terms for the models
df['y2'] = (df['year']-2004)**2
df['c2'] = (df['cohort']-1970)**2
df['c3'] = (df['cohort']-1970)**3
# classify religious affiliation
df['christian'] = df['relig'].isin([1,2,11])
df['none'] = (df['relig'] == 4)
# select recent years and generations
recent = (df['year'] >= 1998)
notold = (df['cohort'] >= 1940)
return df[recent & notold]
```
For exploration, I'll use this subset of the data without resampling, which means it oversamples some groups.
```
df = prepare_df(gss)
df.shape
```
For inference, I'll use 101 resamplings of the data, weighted to be representative of the adult population in the U.S.
```
from utils import resample_rows_weighted
dfs = None
dfs = [resample_rows_weighted(df, df['wtssall'])
for i in range(101)]
```
## Exploring the groups
To see what the groups look like, I select Christians and Nones.
```
christian = df[df['christian']]
christian.shape
none = df[df['none']]
none.shape
```
Here's what the distribution of birth years looks like for the two groups:
```
Cdf.from_seq(christian['cohort']).plot(label='Christian')
Cdf.from_seq(none['cohort']).plot(label='None')
decorate(xlabel='Birth year',
ylabel='CDF',
title='Distribution of birth year')
plt.legend();
```
The Christians in this dataset come from earlier cohorts, so we have to control for that.
```
Cdf.from_seq(christian['year']).plot(label='Christian')
Cdf.from_seq(none['year']).plot(label='None')
decorate(xlabel='Interview year',
ylabel='CDF',
title='Distribution of interview year')
plt.legend();
```
Also, the fraction of Christians was declining over the observation period, so the Christians in the dataset were more likely to be observed earlier. We'll have to control for that, too.
## Selecting variables
The following is a list of the variables we'll explore.
For each variable, I identify a response or list of responses to consider, and provide text that describes what that response means.
I do my best to paraphrase the wording of the question accurately, but if you have any questions, you can [consult the documentation](https://gssdataexplorer.norc.org/projects/52787/variables/data_cart).
```
variables = [
# allow to speak
('spkmil', 1, 'Anti-democratic\nmilitarist'),
('spkmslm', 1, 'Anti-U.S.\nMuslim clergyman'),
('spkath', 1, 'Opponent of churches\nand religion'),
('spkcom', 1, 'Communist'),
('spkrac', 1, 'Racist'),
('spkhomo', 1, 'Homosexual'),
# allow to teach at a college or university
('colmil', 4, 'Anti-democratic\nmilitarist'),
('colmslm', 4, 'Anti-U.S.\nMuslim clergyman'),
('colath', 4, 'Opponent of churches\nand religion'),
('colcom', 5, 'Communist'), # not fired
('colrac', 4, 'Racist'),
('colhomo', 4, 'Homosexual'),
# do not remove from library
('libmil', 2, 'Anti-democratic\nmilitarist'),
('libmslm', 2, 'Anti-U.S.\nMuslim clergyman'),
('libath', 2, 'Opponent of churches\nand religion'),
('libcom', 2, 'Communist'),
('librac', 2, 'Racist'),
('libhomo', 2, 'Homosexual'),
# items related to sex
('homosex', 1, 'Same-sex relations\nalways wrong'),
('premarsx', 1, 'Premarital sex\nalways wrong'),
('xmarsex', 1, 'Extramarital sex\nalways wrong'),
('teensex', 1, 'Teen sex\nalways wrong'),
('sexeduc', 1, 'Favor sex education\nin schools'),
# items related to abortion
('abany', 1, 'A pregnant woman wants it\nfor any reason'),
('abdefect', 1, 'There is a strong chance\nof a serious defect'),
('abnomore', 1, 'She is married and\ndoes not want more children'),
('abhlth', 1, 'Her health is seriously endangered'),
('abpoor', 1, 'She has very low income and\ncannot afford more children'),
('abrape', 1, 'She became pregnant\nas a result of rape'),
('absingle', 1, 'She is not married and\ndoes not want to marry the man'),
# other items related to public policy
('cappun', 2, 'Oppose death penalty\nfor murder'),
('gunlaw', 1, 'Favor permit to buy gun'),
('grass', 1, 'Marijuana should be\nmade legal'),
('divlaw', 1, 'Divorce should be\neasier to obtain'),
('prayer', 1, 'Approve SCOTUS ruling\nprohibiting school prayer'),
('letdie1', 1, 'Allow doctor to end life\nof terminal patient'),
('racopen', 2, 'Favor law barring\nhousing discrimination'),
('pornlaw', [2,3], 'Pornography should be legal'),
('affrmact', [1,2], 'Favor affirmative action\nfor blacks'),
# items related to spending
('natroad', 1, 'Highways and bridges'),
('natsoc', 1, 'Social Security'),
('natmass', 1, 'Mass transportation'),
('natpark', 1, 'Parks and recreation'),
('natchld', 1, 'Assistance for child care'),
('natsci', 1, 'Supporting scientific research'),
('natenrgy', 1, 'Developing alternative\nenergy sources'),
('natspac', 1, 'Space exploration'),
('natenvir', 1, 'Improving and protecting\nthe environment'),
('natheal', 1, "Improving and protecting\nthe nation's health"),
('natcity', 1, 'Solving the problems of\nthe big cities'),
('natcrime', 1, 'Halting the rising\ncrime rate'),
('natdrug', 1, 'Dealing with drug addiction'),
('nateduc', 1, "Improving the nation's\neducation system"),
('natrace', 1, 'Improving the conditions\nof Blacks'),
('natarms', 1, 'The military, armaments\nand defense'),
('nataid', 1, 'Foreign aid'),
('natfare', 1, 'Welfare'),
# confidence in institutions
('confinan', 1, 'Banks and financial instituions'),
('conbus', 1, 'Major companies'),
('conclerg', 1, 'Organized religion'),
('coneduc', 1, 'Education'),
('confed', 1, 'Executive branch of\nthe federal government'),
('conlabor', 1, 'Organized labor'),
('conpress', 1, 'Press'),
('conmedic', 1, 'Medicine'),
('contv', 1, 'Television'),
('conjudge', 1, 'U.S. Supreme Court'),
('consci', 1, 'Scientific community'),
('conlegis', 1, 'Congress'),
('conarmy', 1, 'Military'),
# religious beliefs
('god', 6, 'Know God exists with no doubts'),
('reborn', 1, 'Had a born again experience'),
('savesoul', 1, 'Tried to convince others\nto accept Jesus'),
('bible', 1, 'Bible is the actual word of God\nto be taken literally'),
('postlife', 1, 'Believe there is a life after death'),
('relpersn', [1,2], 'Considers self very or\nmoderately religious'),
('sprtprsn', [1,2], 'Considers self very or\nmoderately spiritual'),
('relexp', 1, 'Had a religious or spiritual\nexperience that changed life'),
('relactiv', [8,9,10,11], 'Church activities weekly or more\nnot including services'),
('pray', [1,2,3,4], 'Prayer weekly or more often'),
('attend', [7,8], 'Attend religious services\nweekly or more often'),
# outlook on people
('helpful', 1, 'People try to be helpful'),
('fair', 2, 'People try to be fair'),
('trust', 1, 'People can be trusted'),
('fear', 2, 'Not afraid to walk alone at night'),
# miscellaneous
('spanking', [1,2], 'Spanking sometimes\nnecessary'),
# gender roles and work
('fepol', 1, 'Agree men are emotionally\nmore suited for politics'),
('fejobaff', [1,2], 'Favor preferential hiring\nand promotion of women'),
('fehire', [1,2], 'Favor special effort to hire\nand promote qualified women'),
('fechld', [1,2], 'Working mother can have warm\nsecure relationship with child'),
('fepresch', [1,2], 'Preschool child is likely\nto suffer if mother works'),
('fefam', [1,2], 'Much better if man achieves and\nwoman takes care of family'),
('discaffm', [1,2], 'Likely that a less qualified\nwoman gets job or promotion'),
('discaffw', [1,2], 'Likely that a less qualified\nman gets job or promotion'),
('meovrwrk', [1,2], 'Family life suffers because men\nconcentrate too much on work'),
]
```
The following cell makes maps from variable names to values and labels.
```
label_map = {}
value_map = {}
for varname, value, label in variables:
value_map[varname] = value
label_map[varname] = label
```
## Period and cohort effects
First I select the subset of one group where the dependent variable is valid.
```
varname = 'natmass'
value = value_map[varname]
print(varname, value)
group = christian.copy()
valid = group.dropna(subset=[varname]).copy()
valid.shape
```
Here's the fraction of this variable that has the selected value.
```
value = np.atleast_1d(value)
(valid[varname].isin(value)).mean()
```
For logistic regression, we need the dependent variable to be 0 or 1.
```
valid['y'] = (valid[varname].isin(value)).astype(int)
valid['y'].value_counts()
```
Here's what the changes look like over time.
```
from utils import plot_series_lowess
by_year = valid.groupby('year')['y'].mean()
plot_series_lowess(by_year, 'C2')
```
And here's what they look like by cohort.
```
by_cohort = valid.groupby('cohort')['y'].mean()
plot_series_lowess(by_cohort, 'C3')
```
## Testing the model
Now we can run logistic regression with year and cohort as explanatory variables.
I consider two versions of this model, with and without quadratic terms. It seems like the model with quadratic terms does a better job of capturing the period and cohort effects.
```
import statsmodels.formula.api as smf
formula = ('y ~ year + cohort')
model = smf.logit(formula, data=valid).fit()
model.summary()
formula = ('y ~ year + y2 + cohort + c2')
model = smf.logit(formula, data=valid).fit()
model.summary()
formula = ('y ~ year + y2 + cohort + c2 + c3')
model = smf.logit(formula, data=valid).fit()
model.summary()
```
The following plot shows the data grouped by cohort and the model for someone interviewed in 2008.
We can use it to confirm that the model is capturing the cohort effect.
```
xs = np.arange(1940, 2000)
dfp = pd.DataFrame()
dfp['cohort'] = xs
dfp['year'] = 2008
dfp['y2'] = (dfp['year']-2004)**2
dfp['c2'] = (dfp['cohort']-1970)**2
dfp['c3'] = (dfp['cohort']-1970)**3
plot_series_lowess(by_cohort, 'C3')
ys = model.predict(dfp)
plt.plot(xs, ys, color='C7', label='Model at year 2008')
plt.xlim(1938, 2002)
decorate(xlabel='Birth year',
ylabel='Fraction',
title='Mean response by year of birth',
loc='lower left')
plt.tight_layout()
plt.savefig('generation_by_cohort.png', dpi=150)
```
The following plot shows the data grouped by year along with predictions for people born in 1968 and 1993.
We can use it to confirm that the model captures the period effect, and we can see the generational difference as measured by the model.
```
plot_series_lowess(by_year, 'C2')
xs = np.arange(1998, 2020)
dfp = pd.DataFrame()
dfp['year'] = xs
dfp['cohort'] = 1968
dfp['y2'] = (dfp['year']-2004)**2
dfp['c2'] = (dfp['cohort']-1970)**2
dfp['c3'] = (dfp['cohort']-1970)**3
ys = model.predict(dfp)
plt.plot(xs, ys, color='C4', label='Born 1968')
dfp['cohort'] = 1993
ys = model.predict(dfp)
plt.plot(xs, ys, color='C6', label='Born 1993')
decorate(xlabel='Interview year',
ylabel='Fraction',
title='Mean response by year of interview',
loc='lower left')
plt.xlim(1996, 2020)
plt.savefig('generation_by_year', dpi=150)
```
## Comparing generations
Now let's see how things change from one generation to the next, controlling to period effects.
I'll use the model to generate predictions for two hypothetical members of this group, born in 1968 and 1993, both interviewed in 2018.
Here's a DataFrame that describes these hypothetical people.
```
df_pred = pd.DataFrame()
df_pred['cohort'] = [1968, 1993]
df_pred['year'] = 2018
df_pred['y2'] = (df_pred['year']-2004)**2
df_pred['c2'] = (df_pred['cohort']-1970)**2
df_pred['c3'] = (df_pred['cohort']-1970)**3
df_pred
```
And here are the predictions.
```
model.predict(df_pred)
```
## Running the model
The following function encapsulates the steps in the previous section.
```
def run_model(df, varname, value, formula):
"""Runs the model and generates predictions.
df: DataFrame
varname: string variable name
value: value or list of values considered "yes"
formula: string patsy model
returns: array of predicted values based on df_pred
"""
value = np.atleast_1d(value)
valid = df.dropna(subset=[varname]).copy()
valid['y'] = (valid[varname].isin(value)).astype(int)
model = smf.logit(formula, data=valid).fit(disp=0)
res = model.predict(df_pred)
return res.values
```
Depending on `formula`, we can run either the linear or quadratic version of the model.
```
#formula = 'y ~ year + cohort'
#formula = 'y ~ year + y2 + cohort + c2'
formula = 'y ~ year + y2 + cohort + c2 + c3'
```
Here are the results for Christians and Nones.
```
run_model(christian, varname, value, formula)
run_model(none, varname, value, formula)
```
## Comparing results
The following function runs the analysis for the two groups and return an array with predictions for 4 hypothetical people:
* Christian born in 1968
* Christian born in 1993
* None born in 1968
* None born in 1993
```
def compare(df, varname, value):
christian = df[df['christian']]
none = df[df['none']]
c = run_model(christian, varname, value, formula)
n = run_model(none, varname, value, formula)
return np.hstack([c, n]) * 100
compare(df, varname, value)
```
The following function runs the same analysis 101 times, using each of the resampled datasets.
It computes the 5th, 50th, and 95th percentiles in each column and returns an array with one row for each percentile and one column for each of the 4 hypothetical people.
```
def compare_iter(dfs, varname, value):
t = [compare(df, varname, value) for df in dfs]
a = np.array(t)
percentiles = np.percentile(a, [5, 50, 95], axis=0)
return percentiles
percentiles = compare_iter(dfs, varname, value)
percentiles
```
## Plotting the results
The following functions visualize the results.
```
def plot_interval(y, interval, dy=0.16, **options):
"""Show a confidence interval.
y: vertical coordinate
interval: triple of low, med, and high
dy: height of the error bars
options: passed to plt
"""
low, mid, high = interval
plt.hlines(y+dy, low, high, alpha=0.6, **options)
plt.vlines([low, high], y+dy, y+dy/2, alpha=0.6, **options)
color_map = {'Christian':'C0',
'None':'C1'}
def plot_arrow(y, row, group):
"""Plot an arrow showing generational changes.
y: vertical coordinate
row: x1, x2 pair
group: string group name
"""
color = color_map[group]
label1 = f'{group} born 1968'
label2 = f'{group} born 1993'
x1, x2 = row
dx = x2 - x1
plt.hlines(y, x1, x2, color=color, alpha=0.3)
plt.plot(x1, y, 'o', color=color, label=label1)
style = '>' if x2 > x1 else '<'
plt.plot(x2, y, style, color=color, label=label2)
def plot_percentiles(y, percentiles, dy=0.12):
"""Plot the results from the resampled analysis.
y: vertical coordinate
percentiles: array with a row for each percentile and
column for each hypothetical person
dy: vertical offset for the first group
"""
plot_interval(y+dy, percentiles[:, 2], color='C1')
plot_arrow(y+dy, percentiles[1, 2:4], group='None')
plot_interval(y-dy, percentiles[:, 0], color='C0')
plot_arrow(y-dy, percentiles[1, 0:2], group='Christian')
def miniplot(percentiles):
"""Make a plot with just one variable"""
plot_percentiles(0, percentiles)
plt.ylim(-1.5, 1.5)
plt.gca().invert_yaxis()
miniplot(percentiles)
plt.legend()
plt.yticks([0], [varname])
decorate(xlabel='Percentage')
```
## Beliefs and attitudes
Now let's see what the results look like for a variety of beliefs and attitudes.
The following list contains variable names, the response I've selected, and a label that summarizes the selected response.
As an example, here's a complete analysis of a single variable.
```
varname = 'pray'
value = value_map[varname]
print(varname, value)
res = compare(df, varname, value)
print(res)
percentiles = compare_iter(dfs, varname, value)
print(percentiles)
miniplot(percentiles)
```
For testing, we can loop through the variables and run one analysis with the unresampled data.
```
for varname, value, _ in variables:
print(varname)
res = compare(df, varname, value)
print(res)
```
And here's how we run the complete analysis for all variables.
It takes a few minutes.
```
def generate_results(variables, results=None):
if results is None:
results = {}
for varname, value, _ in variables:
if varname in results:
continue
print(varname)
percentiles = compare_iter(dfs, varname, value)
results[varname] = percentiles
return results
# uncomment to clear saved results
# results = None
results = generate_results(variables, results)
```
The following function generates a plot for a collection of variable names.
```
def multiplot(results, varnames, **options):
"""Make a plot showing results for several variables.
results: map from varname to array of percentiles
varnames: list of string varnames
loc: string location for the legend.
"""
plt.figure(figsize=(8, 4.5))
for i, varname in enumerate(varnames):
percentiles = results[varname]
plot_percentiles(i, percentiles)
# label the y axis
labels = [label_map[varname] for varname in varnames]
plt.yticks(range(len(varnames)), labels)
# make a legend with just the first four entries
ax = plt.gca()
handles, labels = ax.get_legend_handles_labels()
loc = options.pop('loc', 'best')
ax.legend(handles[:4], labels[:4], loc=loc)
# flip the axes so the results go from top to bottom
plt.gca().invert_yaxis()
underride(options, xlabel='Percentage', legend=False)
decorate(**options)
plt.tight_layout()
```
## Free speech
There are always some people whose ideas are considered bad or dangerous by other people.
* For instance, somebody who is against all churches and religion . . .
If such a person wanted to make a speech in your (city/town/community) against churches and religion, should he be allowed to speak, or not?
* Or consider a person who believes that Blacks are genetically inferior...
If such a person wanted to make a speech in your community claiming that Blacks are inferior, should he be allowed to speak, or not?
* Now, I should like to ask you some questions about a man who admits he is a Communist.
Suppose this admitted Communist wanted to make a speech in your community. Should he be allowed to speak, or not?
* Consider a person who advocates doing away with elections and letting the military run the country.
If such a person wanted to make a speech in your community, should he be allowed to speak, or not?
* And what about a man who admits that he is a homosexual?
Suppose this admitted homosexual wanted to make a speech in your community. Should he be allowed to speak, or not?
* Now consider a Muslim clergyman who preaches hatred of the United States.
If such a person wanted to make a speech in your community preaching hatred of the United States, should he be allowed to speak, or not?
```
varnames = ['spkmslm', 'spkrac', 'spkcom', 'spkmil', 'spkath', 'spkhomo']
multiplot(results, varnames, title='Allow to speak')
```
There are always some people whose ideas are considered bad or dangerous by other people.
* For instance, somebody who is against all churches and religion . . .
Should such a person be allowed to teach in a college or university, or not?
* Now consider a Muslim clergyman who preaches hatred of the United States.
Should such a person be allowed to teach in a college or university, or not?
* Questions associated with this variable:
Or consider a person who believes that Blacks are genetically inferior....
Should such a person be allowed to teach in a college or university, or not?
* Now, I should like to ask you some questions about a man who admits he is a Communist.
Suppose he is teaching in a college. Should he be fired, or not?
* Consider a person who advocates doing away with elections and letting the military run the country.
Should such a person be allowed to teach in a college or university, or not?
* And what about a man who admits that he is a homosexual?
Should such a person be allowed to teach in a college or university, or not?
```
varnames = ['colmslm', 'colrac', 'colcom', 'colmil', 'colath', 'colhomo']
multiplot(results, varnames, title='Allow to teach at college or university');
```
There are always some people whose ideas are considered bad or dangerous by other people.
* libath: For instance, somebody who is against all churches and religion . . .
If some people in your community suggested that a book he wrote against churches and religion should be taken out of your public library, would you favor removing this book, or not?
* librac: Or consider a person who believes that Blacks are genetically inferior.
If some people in your community suggested that a book he wrote which said Blacks are inferior should be taken out of your public library, would you favor removing this book, or not?
* libcom: Now, I should like to ask you some questions about a man who admits he is a Communist.
Suppose he wrote a book which is in your public library. Somebody in your community suggests that the book should be removed from the library. Would you favor removing it, or not?
* libmil: Consider a person who advocates doing away with elections and letting the military run the country.
Suppose he wrote a book advocating doing away with elections and letting the military run the country. Somebody in your community suggests that the book be removed from the public library. Would you favor removing it, or not?
* libhomo: And what about a man who admits that he is a homosexual?
If some people in your community suggested that a book he wrote in favor of homosexuality should be taken out of your public library, would you favor removing this book, or not?
* libmslm: Now consider a Muslim clergyman who preaches hatred of the United States.
If some people in your community suggested that a book he wrote which preaches hatred of the United States should be taken out of your public library, would you favor removing this book, or not?
```
varnames = ['libmslm', 'librac', 'libcom', 'libmil', 'libath', 'libhomo']
multiplot(results, varnames, title='Keep book in library written by');
```
## Confidence in institutions
"I am going to name some institutions in this country. As far as the people running these institutions are concerned, would you say you have a great deal of confidence, only some confidence, or hardly any confidence at all in them?"
```
varnames = ['conbus', 'coneduc', 'conjudge', 'conmedic', 'consci', 'conarmy']
multiplot(results, varnames, loc='upper right', title='Great deal of confidence in');
varnames = ['contv', 'conpress', 'conlegis', 'confed', 'conclerg', 'confinan', 'conlabor']
multiplot(results, varnames, loc='upper right', title='Great deal of confidence in');
```
## Allocation of resources
"We are faced with many problems in this country, none of which can be solved easily or inexpensively. I'm going to name some of these problems, and for each one I'd like you to tell me whether you think we're spending too much money on it, too little money, or about the right amount."
```
varnames = ['nataid', 'natfare', 'natpark', 'natrace', 'natchld', 'natenvir']
multiplot(results, varnames, title='We spend too little money on')
plt.savefig('generation5.png', dpi=150)
varnames = ['natarms', 'natmass', 'natsci', 'natroad', 'natsoc', 'natcrime']
multiplot(results, varnames, loc='upper right', title='We spend too little money on')
plt.savefig('generation6.png', dpi=150)
varnames = ['natspac', 'natcity', 'natenrgy', 'natdrug', 'natheal', 'nateduc']
multiplot(results, varnames, loc='upper right', title='We spend too little money on')
plt.savefig('generation7.png', dpi=150)
```
## Issues related to sex
```
varnames = ['premarsx', 'homosex', 'teensex', 'xmarsex', 'sexeduc']
multiplot(results, varnames, loc='upper right')
plt.savefig('generation8.png', dpi=150)
```
## Outlook
```
varnames = ['trust', 'helpful', 'fair', 'fear']
multiplot(results, varnames)
```
## Public policy
```
varnames = ['affrmact', 'cappun', 'divlaw', 'prayer']
multiplot(results, varnames, loc='upper right', title='Law and public policy')
plt.savefig('generation3.png', dpi=150)
varnames = ['grass', 'pornlaw', 'gunlaw', 'letdie1', 'racopen']
multiplot(results, varnames, title='Law and public policy')
plt.savefig('generation2.png', dpi=150)
```
## Abortion
"Please tell me whether or not you think it should be possible for a pregnant woman to obtain a legal abortion if ...
```
varnames = ['abany', 'abnomore', 'absingle', 'abpoor', 'abdefect', 'abrape', 'abhlth']
multiplot(results, varnames, title='Abortion should be legal if', loc='lower left')
plt.savefig('generation4.png', dpi=150)
```
## Religion
```
varnames = ['bible', 'reborn', 'savesoul', 'god', 'postlife']
multiplot(results, varnames,
loc='upper right',
title='Religious beliefs',
xlim=[0, 110])
plt.savefig('generation1.png', dpi=150)
varnames = ['relactiv', 'attend', 'relpersn', 'sprtprsn', 'relexp', 'pray']
multiplot(results, varnames,
loc='upper right',
title='Religious beliefs',
#xlim=[0, 110]
)
plt.savefig('generation1b.png', dpi=150)
```
## Gender roles and work
"Tell me if you agree or disagree with this statement: Most men are better suited emotionally for politics than are most women." Select "agree"
df.fepol.replace([0, 8, 9], np.nan, inplace=True)
df.fejobaff.replace([0, 8, 9], np.nan, inplace=True)
df.fehire.replace([0, 8, 9], np.nan, inplace=True)
df.fechld.replace([0, 8, 9], np.nan, inplace=True)
df.fepresch.replace([0, 8, 9], np.nan, inplace=True)
df.fefam.replace([0, 8, 9], np.nan, inplace=True)
df.discaffm.replace([0, 8, 9], np.nan, inplace=True)
df.discaffw.replace([0, 8, 9], np.nan, inplace=True)
df.meovrwrk.replace([0, 8, 9], np.nan, inplace=True)
```
varnames = ['fepol', 'fefam', 'fepresch', 'fejobaff', 'discaffm', 'meovrwrk']
multiplot(results, varnames, title='Agree or strongly agree');
varnames = ['discaffw', 'fehire', 'fechld']
multiplot(results, varnames, title='Agree or strongly agree', loc='lower left')
plt.xlim(60,85);
```
## Misc
"Do you strongly agree, agree, disagree, or strongly disagree that it is sometimes necessary to discipline a child with a good, hard spanking?"
"Tell me if you agree or disagree with this statement: Most men are better suited emotionally for politics than are most women." Select "agree"
```
varnames = ['fepol', 'spanking']
multiplot(results, varnames, title='Agree or strongly agree');
t = []
for varname, _, _ in variables:
if varname.startswith('con'):
t.append(varname)
t
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from itertools import combinations
from collections import Counter
df = pd.read_csv('movie_bd_v5.csv')
df.sample(5)
df.describe()
```
# Предобработка
```
answers = {}
# some questions required season grouping
def get_season_by_month(month):
if month in [12, 1, 2]:
return 'winter'
if month in [3, 4, 5]:
return 'spring'
if month in [6, 7, 8]:
return 'summer'
if month in [9, 10, 11]:
return 'fall'
raise RuntimeError('invalid month provided: ' + month)
# to group by actor combinations we need to split cast list and sort it
def split_sorted(string):
result = string.split('|')
result.sort()
return result
df['full_title'] = df['original_title'] + ' (' + df['imdb_id'] + ')'
df['original_title_len'] = df['original_title'].apply(lambda s: len(s))
# profit = revenue - budget
df['profit'] = df['revenue'] - df['budget']
df['genres_split'] = df['genres'].apply(lambda x: x.split('|'))
df['director_split'] = df['director'].apply(lambda x: x.split('|'))
# use split_sorted in order to make actors combinations similar
df['cast_split'] = df['cast'].apply(split_sorted)
df['cast_split_combinations'] = df['cast_split'].apply(lambda x: list(combinations(x, 2)))
df['prod_companies_split'] = df['production_companies'].apply(lambda x: x.split('|'))
df['release_month'] = df['release_date'].apply(lambda x: int(x.split('/')[0]))
df['release_season'] = df['release_month'].apply(get_season_by_month)
df['overview_words_count'] = df['overview'].apply(lambda s: len(s.split()))
df.sample(5)
genres_split_df = df.explode('genres_split')
prod_companies_split_df = df.explode('prod_companies_split')
director_split_df = df.explode('director_split')
director_genres_split_df = director_split_df.explode('genres_split')
cast_split_df = df.explode('cast_split')
cast_genres_split_df = cast_split_df.explode('genres_split')
cast_split_combinations_df = df.explode('cast_split_combinations')
```
# 1. У какого фильма из списка самый большой бюджет?
Использовать варианты ответов в коде решения запрещено.
Вы думаете и в жизни у вас будут варианты ответов?)
```
answers['1'] = 'Pirates of the Caribbean: On Stranger Tides (tt1298650)'
df[df['budget'] == max(df['budget'])]['full_title'].iloc[0]
```
# 2. Какой из фильмов самый длительный (в минутах)?
```
answers['2'] = 'Gods and Generals (tt0279111)'
df[df['runtime'] == max(df['runtime'])]['full_title'].iloc[0]
```
# 3. Какой из фильмов самый короткий (в минутах)?
```
answers['3'] = 'Winnie the Pooh (tt1449283)'
df[df['runtime'] == min(df['runtime'])]['full_title'].iloc[0]
```
# 4. Какова средняя длительность фильмов?
```
answers['4'] = 110
np.mean(df['runtime'])
```
# 5. Каково медианное значение длительности фильмов?
```
answers['5'] = 107.0
np.median(df['runtime'])
```
# 6. Какой самый прибыльный фильм?
#### Внимание! Здесь и далее под «прибылью» или «убытками» понимается разность между сборами и бюджетом фильма. (прибыль = сборы - бюджет) в нашем датасете это будет (profit = revenue - budget)
```
answers['6'] = 'Avatar (tt0499549)'
df[df['profit'] == max(df['profit'])]['full_title'].iloc[0]
```
# 7. Какой фильм самый убыточный?
```
answers['7'] = 'The Lone Ranger (tt1210819)'
df[df['profit'] == min(df['profit'])]['full_title'].iloc[0]
```
# 8. У скольких фильмов из датасета объем сборов оказался выше бюджета?
```
answers['8'] = 1478
len(df[df['revenue'] > df['budget']])
```
# 9. Какой фильм оказался самым кассовым в 2008 году?
```
answers['9'] = 'The Dark Knight (tt0468569)'
release_2008_df = df[(df['release_year'] == 2008)]
release_2008_max_profit_df = release_2008_df[release_2008_df['profit'] == max(release_2008_df['profit'])]
release_2008_max_profit_df['full_title'].iloc[0]
```
# 10. Самый убыточный фильм за период с 2012 по 2014 г. (включительно)?
```
answers['10'] = 'The Lone Ranger (tt1210819)'
release_2012_2014_df = df[(df['release_year'] >= 2012) & (df['release_year'] <= 2014)]
min_profit_df = release_2012_2014_df[release_2012_2014_df['profit'] == min(release_2012_2014_df['profit'])]
min_profit_df['full_title'].iloc[0]
```
# 11. Какого жанра фильмов больше всего?
```
answers['11'] = 'Drama'
```
ВАРИАНТ 1
```
Counter(df['genres'].str.cat(sep='|').split('|')).most_common()[0][0]
```
ВАРИАНТ 2
```
Counter(genres_split_df['genres_split']).most_common()[0][0]
```
# 12. Фильмы какого жанра чаще всего становятся прибыльными?
```
answers['12'] = 'Drama'
```
ВАРИАНТ 1
```
profitable_df = df[df['profit'] > 0]
Counter(profitable_df['genres'].str.cat(sep='|').split('|')).most_common()[0][0]
```
ВАРИАНТ 2
```
profitable_df = genres_split_df[genres_split_df['profit'] > 0]
Counter(profitable_df['genres_split']).most_common()[0][0]
```
# 13. У какого режиссера самые большие суммарные кассовые сборы?
```
answers['13'] = 'Peter Jackson'
revenue_df = director_split_df.groupby('director_split')['revenue']
revenue_df.apply(sum).sort_values(ascending=False).keys()[0]
```
# 14. Какой режиссер снял больше всего фильмов в стиле Action?
```
answers['14'] = 'Robert Rodriguez'
action_df = director_genres_split_df[director_genres_split_df['genres_split'] == 'Action']
action_df['director_split'].value_counts().sort_values(ascending=False).keys()[0]
```
# 15. Фильмы с каким актером принесли самые высокие кассовые сборы в 2012 году?
```
answers['15'] = 'Chris Hemsworth'
release_2012_df = cast_split_df[cast_split_df['release_year'] == 2012]
revenue_df = release_2012_df.groupby('cast_split')['revenue']
revenue_df.apply(sum).sort_values(ascending=False).keys()[0]
```
# 16. Какой актер снялся в большем количестве высокобюджетных фильмов?
```
answers['16'] = 'Matt Damon'
high_budget_df = cast_split_df[cast_split_df['budget'] >= np.mean(cast_split_df['budget'])]
high_budget_df['cast_split'].value_counts().sort_values(ascending=False).keys()[0]
```
# 17. В фильмах какого жанра больше всего снимался Nicolas Cage?
```
answers['17'] = 'Action'
ncage_df = cast_genres_split_df[cast_genres_split_df['cast_split'] == 'Nicolas Cage']
ncage_df['genres_split'].value_counts().sort_values(ascending=False).keys()[0]
```
# 18. Самый убыточный фильм от Paramount Pictures
```
answers['18'] = 'K-19: The Widowmaker (tt0267626)'
para_df = prod_companies_split_df[prod_companies_split_df['prod_companies_split'] == 'Paramount Pictures']
para_df[para_df['profit'] == min(para_df['profit'])]['full_title'].iloc[0]
```
# 19. Какой год стал самым успешным по суммарным кассовым сборам?
```
answers['19'] = 2015
df.groupby('release_year')['revenue'].apply(sum).sort_values(ascending=False).keys()[0]
```
# 20. Какой самый прибыльный год для студии Warner Bros?
```
answers['20'] = 2014
warner_df = prod_companies_split_df[prod_companies_split_df['prod_companies_split'].str.startswith('Warner Bros', na=False)]
warner_df.groupby('release_year')['profit'].apply(sum).sort_values(ascending=False).keys()[0]
```
# 21. В каком месяце за все годы суммарно вышло больше всего фильмов?
```
answers['21'] = 'Сентябрь'
df['release_month'].value_counts().sort_values(ascending=False).keys()[0]
```
# 22. Сколько суммарно вышло фильмов летом? (за июнь, июль, август)
```
answers['22'] = 450
df['release_season'].value_counts()['summer']
```
# 23. Для какого режиссера зима – самое продуктивное время года?
```
answers['23'] = 'Peter Jackson'
winter_df = director_split_df[director_split_df['release_season'] == 'winter']['director_split']
winter_df.value_counts().sort_values(ascending=False).keys()[0]
```
# 24. Какая студия дает самые длинные названия своим фильмам по количеству символов?
```
answers['24'] = 'Four By Two Productions'
title_len_df = prod_companies_split_df.groupby('prod_companies_split')['original_title_len']
title_len_df.apply(np.mean).sort_values(ascending=False).keys()[0]
```
# 25. Описание фильмов какой студии в среднем самые длинные по количеству слов?
```
answers['25'] = 'Midnight Picture Show'
overview_words_count_df = prod_companies_split_df.groupby('prod_companies_split')['overview_words_count']
overview_words_count_df.apply(np.mean).sort_values(ascending=False).keys()[0]
```
# 26. Какие фильмы входят в 1 процент лучших по рейтингу?
по vote_average
```
answers['26'] = 'Inside Out, The Dark Knight, 12 Years a Slave'
df[df['vote_average'] >= df['vote_average'].quantile(0.99)]['original_title']
```
# 27. Какие актеры чаще всего снимаются в одном фильме вместе?
```
answers['27'] = 'Daniel Radcliffe & Rupert Grint'
cast_combinations_df = cast_split_combinations_df.groupby('cast_split_combinations')['cast_split_combinations']
cast_combinations_df.agg(['count']).sort_values(by='count',ascending=False)
```
# Submission
```
answers
len(answers)
```
| github_jupyter |
# 多因子模型MFM
理论上多因子模型应涵盖包含股票、债券、大宗商品、地产等所有资产,但我们的讨论仅局限在股票的二级市场。
作为量化选股多因子𝐴𝑙𝑝ℎ𝑎模型构建环节中最重要的一部分,如何寻找具有逻辑支撑且能有效区分和预测股票收益的因子是主要内容。
大部分机构投资者主要研究Alpha模型,风险模型和组合优化采用外购商业软件(BARRA,Axioma等)
风险模型:任意股票同一时刻都暴露于多种不同的风险因素下,它们之间的共同作用形成了股票价格的波动,为了定量研究各种风险因素的作用,量化风险模型应运而生。
风险模型的意义在于找到股票价格波动的成因,并将股票收益来源进行剥离,并实行对未来股票波动价格的预测。
Alpha因子:是对股票收益率具有明显显著且稳定影响的某一变量,同时该影响是剔除其余所有因子对收益的作用而独立存在的。
“群众型”:更符合分散风险的量化投资基本原理,大型的Alpha因子库在提供多样的Alpha源的而同时,也带来了大量的无效和重复信息,会使得不同的Alpha源被人为的放大或缩小权重,最终影响组合表现。
对于一个新的因子,需要思考:是否提供了新的Alpha源?或只是把现有的因子库信息进行了切分重组?
对于因子相关性检验:IC协方差矩阵的估计(改进:压缩估计量、Boostrap方法)

# **多因子模型的构建**
多因子模型是个较为复杂的体系,模型的构建流程往往包括以下几个方面:

## **样本筛选**
全体 A 股 ,为了使测试结果更符合投资逻辑,设定了三条样本筛选规则:
(1) 剔除选股日的 ST/PT 股票;
(2) 剔除上市不满一年的股票;
(3) 剔除选股日由于停牌等原因而无法买入的股票。
## **数据清洗**
避免可能的数据错误和极端数据对测试结果产生影响,使用标准化后的数据保证最终得到的模型的稳健性。数据清洗的内容主要包括两部分,异常值和缺失值的处理。
常见的因子标准化方法包括:Z 值标准化(Z-Score),Rank 标准化,风格 标准化等等。 由于 Rank 标准化后的数据会丢失原始样本的一些重要信息,这里我们仍然 选择 Z 值标准化来处理因子数据。
## **因子测试**
有效的单因子首先应该具有一定的逻辑支撑,其次则是与股票收益率的相关性较为显著
单因子测试
截面回归(Cross-Section Regression)是目前业界较常用于因子测试的方法,相比全样本面板回归(Panel Data Regression)的方法,截面回归更有利于对因子变化趋势的捕捉。
我们选择每期针对全体样本做一次回归,回归时因子暴露为已知变量,回归 得到每期的一个因子收益值𝑓𝑗,在通过多期回归后我们就可以得到因子值𝑓𝑗 的序列,也就是因子收益率序列,同时可以得到 t 值序列,也就是因子值与 股票收益率相关性的t检验得到的t值。
针对这两个序列我们将通过以下几个指标来判断 该因子的有效性以及稳定性:
(1) 因子收益序列𝑓𝑖的假设检验 t 值
(2) 因子收益序列𝑓𝑖大于 0 的概率
(3) t 值绝对值的均值
(4) t 值绝对值大于等于 2 的概率
IC 值(信息系数)是指个股第 t 期在因子 i 上的因子暴露(剔除行业与市值 后)与 t + 1 期的收益率的相关系数。通过计算 IC 值可以有效的观察到某个因子收益率预测的稳定性和动量特征,以便在优化组合时用作筛选的指标。 常见的计算 IC 值方法有两种:相关系数(Pearson Correlation) 和秩相关系数( Spearman Rank Correlation)。
由于 Pearson 相关系数计算时假设变量具有相等间隔以及服从正态分布,而 这一假设往往与因子值和股票收益率的分布情况相左。因此我们将采用 Spearman 的方法计算因子暴露与下期收益率的秩相关性 IC 值。类似回归法 的因子测试流程,我们在计算 IC 时同样考虑剔除了行业因素与市值因素。
同样我们会得到一个 IC 值序列,类似的,我们将关注以下几个与 IC 值相关 的指标来判断因子的有效性和预测能力:
(1) IC 值的均值
(2) IC 值的标准差
(3) IC 大于 0 的比例
(4) IC 绝对值大于 0.02 的比例
(5) IR (IR = IC 均值/IC 标准差)
### **多因子测试**
在完成单个因子的测试之后,就为多因子模型的构建打下了坚实的基础。我们可以通过下面几个步骤来剔除同类因子之间的多重共线性影响,筛选出同时具有良好的单调性和预测性的有效因子,构造我们的多因子模型:
根据上文的截面回归因子测试方法,我们可以轻松的得到每个因子的因子暴 露值序列和因子 IC 值序列。在研究因子间共线性时,就可以通过计算因子 间 IC 值和因子暴露值得相关性来求证因子间的共线性。
需要注意的是,经济含义相似度较高的同类型因子往往存在明显的正相关性, 在处理此类因子时,我们可以通过一些方法将因子进行合并;而如果是经济 含义差异较大因子之间存在明显相关性,就需要有所取舍。
消除共线性的方法包括以下几种:
(1) 在同类因子的共线性较大的几个因子中,保留有效性最高的因子, 剔除余下的因子
(2) 因子组合:方法包括等权加权,以因子收益 f 为权重加权,以及 PCA 主成分分析法等等
(3) 暴力迭代法,即将因子两两组合暴力迭代得到表现最好的组合方法。
在对因子集做残差的异方差分析处理之后,就可以进行多元线性回归,估计每期的因子收益序列。

## **因子的初步筛选**
对每个因子做详尽的分析,具体测试了包括因子收益,因子收益显著性, 因子 IC、IR,分层回溯收益、多空收益,历史 IC 序列相关性等等指标,一般根据前期的测试结果,筛选出了收益率较显著,高 IC、IR 并且单调性得分较高的因子。
### **因子权重的优化—基于因子 IC**
静态因子加权 v.s.动态调整权重
静态优化IC
复合因子是M 个因子的一个线形组合:因子的 IR 值为因子 IC 的均值与因子 IC 的标准差的比值。因子 IR 值越高, 代表因子综合考虑区分度和稳定性后效果越好,优化目标是使复合因子的信息比 IR 取到最大值。
动态最优化IR
虑到市场环境变化和风格转变等原因会使得因子的有效性和 IC 值出现波 动,仅仅使用静态的 IC 优化方法给定复合因子中的因子权重会容易导致组 合受风格变化影响而出现较大的波动和回撤,构造了一个动态调整的基于 IC 值的复合因子 IR 最优化模型。
动态的最优化权重模型是建立在上面我们提到的静态最优化 IR 模型的基础上的,假设每期优化因子权重时所参考的历史 IC 序列长度为 N 个月。(首先假设组合股票数量为 M=100,手续费单边 0.3%,每月初调仓,则在不 同的参数 N 下)
一般动态的最优化 IR 赋权方法要优于静态的因子赋权法
## **因子赋权组合**
组合内等权 v.s.复合因子得分加权 在构建每月调仓组合时,入选标的的赋权方式包括等权和按复合因子的得分加权两种方法。
(EW 为等权,SW 为复合因子得分加权)
# **因子中性化**
原因:A股的行业轮动明显,行业热点之间切换迅速,并且A股小市值个股占比明显,高波动率高收益率
目的:
* 降低投资组合的波动性和回测
* 为了在用某一因子时能够剔除其他因素的影响,使得选出的股票更加分散,需要进行中性化处理
* 为了消除因子中的偏差和不必要的影响
对于因子来说,中性化主要考虑市场风险和行业风险。
行业中性:多头组合的行业配置与对冲基准的行业配置一致,目的在于剔除行业因子对策略收益的影响,仅考察行业内部个股的超额收益。(行业中性策略的净值曲线往往比较平稳,回测较小)
风格因子中性:多头组合的风格因子较之对冲基准的风险暴露为0,目的在于使多头组合的风格特征完全与对冲基准相匹配,使得组合的超额收益不是来自于某类风格
1. 行业中性化
行业中性化一般采用两种方式处理
* 简单的标准化法
利用申万行业指数,将各个行业的因子进行标准化处理,即减去均值除以标准差
* 回归取残差法
将因子值作为y,行业哑变量作为x,进行线性回归,回归模型的残差即为行业中性化后的因子值
2. 市值中性化
因为市值因子是连续的,采用回归残差法
| github_jupyter |
```
#importing libraries
import PIL
from PIL import Image,ImageTk,ImageEnhance
import cv2
import sys
if "Tkinter" not in sys.modules:
import tkinter as tk
from tkinter import *
from tkinter import messagebox
import os
import csv
import pandas as pd
import numpy as np
import statistics
from statistics import mode
import shutil
#global variable
global Name
global sample_count
global ID
global DetectMode
global W
global lside
global lbright
global s1
global ListDetect
global Info
global label2
global label3
global label4
global label5
global label6
global Ck2
global Ck3
global Ck4
global Ck7
global face_count
face_count=0
sample_count=100
DetectMode=False
W=False
###############################################################
#this function extracts the person's unique iD from the image name and return it with the image
#Use to make the dataset for the recognizer to train upon
def getImagesAndLabels(path):
# get the path of all the files in the folder
imagePaths =[os.path.join(path, f) for f in os.listdir(path)]
faces =[]
# creating empty ID list
Ids =[]
# now looping through all the image paths and loading the
# Ids and the images saved in the folder
for imagePath in imagePaths:
if not(imagePath[15:]=='.DS_Store'):
# loading the image and converting it to gray scale
pilImage = PIL.Image.open(imagePath).convert('L')
# Now we are converting the PIL image into numpy array
imageNp = np.array(pilImage, 'uint8')
# getting the Id from the image
Id = int(os.path.split(imagePath)[-1].split(".")[1])
# extract the face from the training image sample
faces.append(imageNp)
Ids.append(Id)
return faces, Ids
################################################################
#these are used to handle the text entry boxes
def focus1(event):
# set focus on the name_field box
name_field.focus_set()
# Function to set focus
def focus2(event):
# set focus on the age_field box
age_field.focus_set()
# Function to set focus
def focus3(event):
# set focus on the address_field box
address_field.focus_set()
def clear():
# clear the content of text entry box
username_field.delete(0, END)
name_field.delete(0, END)
age_field.delete(0, END)
address_field.delete(0, END)
print("Done")
#################################################################
#inserting user information into the csv file
def insert():
global Name
global sample_count
global ID
global En
data=pd.read_csv("Data.csv")
# if user not fill any entry
# then show error message
if (username_field.get() == "" or
name_field.get() == "" or
age_field.get() == "" or
address_field.get() == "" ):
messagebox.showerror(title="Message", message="Please fill out all the information",icon='error')
elif(username_field.get() in list(data['UserID']) ): #if the username is used again by someone else error message
messagebox.showerror(title="Message", message="This UserID is already taken",icon='error')
else:
ID+=1
row = [ID,username_field.get(),name_field.get(),age_field.get(),address_field.get()]
with open('Data.csv', 'a+') as csvFile:
writer = csv.writer(csvFile)
# Entry of the row in csv file
writer.writerow(row)
csvFile.close()
Name=name_field.get()
username_field.focus_set()
#By setting sample_count=0, you can snap some pictures of particular people.
sample_count=0
clear()
#####################################################################
#enabling decting mode. Also train the recognizer
def Detect():
global DetectMode
global ListDetect
global Info
ListDetect=[]
Info="Detecting......."
data=pd.read_csv("Data.csv")
Count=len(data["ID"])
if Count==0:
messagebox.showerror(title="Message", message="Database is Empty",icon='error')
elif Count==1:
messagebox.showerror(title="Message", message="Add at least two data records",icon='error')
else:
faces, Id = getImagesAndLabels("TrainingImages")
if 'Trainner.yml' not in os.listdir('TrainingImageLabel'):
recognizer.train(faces, np.array(Id))
recognizer.save("TrainingImageLabel/Trainner.yml")
messagebox.showinfo(title="Message", message="Finished Training",icon='info')
file1 = open("TrainingImageLabel/Counter.txt","w")
file1.write(str(Count))
file1.close()
else:
file1 = open("TrainingImageLabel/Counter.txt")
R=file1.readlines(0)
file1.close()
CountV=int(R[0])
if Count==CountV:
recognizer.read("TrainingImageLabel/Trainner.yml")
else:
file1 = open("TrainingImageLabel/Counter.txt","w")
file1.write(str(Count))
file1.close()
recognizer.train(faces, np.array(Id))
recognizer.save("TrainingImageLabel/Trainner.yml")
messagebox.showinfo(title="Message", message="Finished Training",icon='info')
DetectMode=True
########################################################################
#accessing Monitor Mode
def Monitor():
global DetectMode
DetectMode=False
########################################################################
#loading images from profile folders (to show when a person is recognized)
def ProfileLoad(N):
try:
pilImage = PIL.Image.open('ProfileImages/ '+str(N)+".jpg")
pilImage = pilImage.resize((250, 250), PIL.Image.ANTIALIAS)
imgTK = ImageTk.PhotoImage(image=pilImage)
return imgTK
except:
pass
#loading images from profile folders (to show in the database management system)
def ProfileLoad1(N):
try:
pilImage = PIL.Image.open('ProfileImages/ '+str(N)+".jpg")
pilImage = pilImage.resize((150, 150), PIL.Image.ANTIALIAS)
imgTK = ImageTk.PhotoImage(image=pilImage)
return imgTK
except:
pass
########################################################################
#showing the information and the photos of the system users
def DataBaseManagement():
global DetectMode
DetectMode=False
data4=pd.read_csv("Data.csv")
Plist=[]
#loading images to be shown from the ProfileImages folder
for i in range(len(data4["ID"])):
Plist.append(ProfileLoad1(i+1))
def data():
for i in range(len(data4["ID"])):
label=Label(frame)
label.grid(row=i,column=0)
imgx = Plist[i]
label.imgx=imgx
label.configure(image=imgx)
Txt='Username : '+str(data4.at[i,"UserID"])+"\n"+'Name : '+str(data4.at[i,"Name"])+"\n"+'Age : '+str(data4.at[i,"Age"])+"\n"+'Address : '+str(data4.at[i,"Address"])
Label(frame,text=Txt,height=4,bg='light blue',justify=tk.LEFT,font=("Helvetica", 20)).grid(row=i,column=1)
def myfunction(event):
canvas.configure(scrollregion=canvas.bbox("all"),width=500,height=500)
root=Toplevel(gui)
sizex = 590
sizey = 625
posx = 100
posy = 100
root.wm_geometry("%dx%d+%d+%d" % (sizex, sizey, posx, posy))
#frame
myframe=Frame(root,relief=GROOVE,width=100,height=100,bd=1)
myframe.place(x=35,y=50)
#label
label0=Label(root,bg='black',fg='white',text="Database Management System", font=("Helvetica", 25))
label0.place(x=300, y=25,anchor='c')
#button
DeleteRecord = Button(root, text="Delete last record", fg="black",
bg="red", command=Delete,height=2, width=20)
DeleteRecord.place(x=175, y=590,anchor='c')
EmptyData = Button(root,bg="red", text="Empty Database",command=Empty, fg="black",
height=2, width=20)
EmptyData.place(x=425, y=590,anchor='c')
#canvas and scroll bar
canvas=Canvas(myframe)
frame=Frame(canvas)
myscrollbar=Scrollbar(myframe,orient="vertical",command=canvas.yview)
canvas.configure(yscrollcommand=myscrollbar.set)
myscrollbar.pack(side="right",fill="y")
canvas.pack(side="left")
canvas.create_window((0,0),window=frame,anchor='nw')
frame.bind("<Configure>",myfunction)
data()
root.mainloop()
########################################################################
#Delete the last record and the corresponding images
def Delete():
global ID
Pdata=pd.read_csv("Data.csv")
ln=len(Pdata['ID'])
if ln==0:
messagebox.showerror(title="Message", message="No records left",icon='error')
else:
Nme=Pdata.at[ln-1,"Name"]
Pdata=Pdata.drop(Pdata.index[ln-1])
Pdata.to_csv("Data.csv",index=False,header=True)
for j in range(1,32):
os.remove("TrainingImages/ "+str(Nme)+"."+str(ln)+"."+str(j)+".jpg")
os.remove("ProfileImages/ "+str(ln)+".jpg")
ID=ID-1
########################################################################
#reset the database deleting all the records
def Empty():
shutil.rmtree('TrainingImageLabel')
os.makedirs('TrainingImageLabel')
shutil.rmtree('ProfileImages')
os.makedirs('ProfileImages')
shutil.rmtree('TrainingImages')
os.makedirs('TrainingImages')
with open('Data.csv', 'w', newline='') as file:
writer = csv.writer(file)
writer.writerow(["ID","UserID", "Name", "Age","Address"])
########################################################################
#function to close the application
def Quit():
gui.destroy()
capmain.release()
########################################################################
#this is a recursive function. at the end of the function it will call the next function
def show_Mainframe():
global DetectMode
global sample_count
global W
global lside
global lbright
global s1
global ListDetect
global label2
global label3
global label4
global label5
global label6
global label7
global Info
global Ck2
global Ck3
global Ck4
global Ck7
global face_count
isValid=True
try :
ret, framemain = capmain.read()
except:
print("error the take a image")
isValid = False
if isValid == True:
framemain = cv2.flip(framemain, 1)
framemain = cv2.resize(framemain, (400, 300))
frameside= cv2.resize(framemain, (200, 150))
framebright= cv2.resize(framemain, (200, 150))
#coverting image to grayscale
gray = cv2.cvtColor(framemain, cv2.COLOR_BGR2GRAY)
faces = faceCascade.detectMultiScale(
gray,
scaleFactor=1.3,
minNeighbors=3,
minSize=(30, 30),
flags=cv2.CASCADE_SCALE_IMAGE
)
#counting number of faces
if len(faces)>0 and face_count>0:
face_count-=1
elif len(faces)==0 and face_count<=16:
face_count+=1
# Draw a rectangle around the faces
for (x, y, w, h) in faces:
cv2.rectangle(framemain, (x, y), (x+w, y+h), (0, 255, 0), 2)
if DetectMode:
data2=pd.read_csv("Data.csv")
#predicting the face
iDs, confidence = recognizer.predict(gray[y:y+h,x:x+w])
# If confidence is less them 100 ==> "0" : perfect match
if (confidence < 100):
iD = data2.at[iDs-1,"Name"]
confidence = " {0}%".format(round(100 - confidence))
else:
iD = "unknown"
confidence = " {0}%".format(round(100 - confidence))
#collecting 10 IDs of the detected faces
if len(ListDetect)<10:
if iD=='unknown':
ListDetect.append(str(iD))
else:
ListDetect.append(str(iDs))
else:
ListDetect=[]
#putting a label of the detected persons name
cv2.putText(
framemain,
str(iD),
(x+5,y-5),
cv2.FONT_HERSHEY_SIMPLEX,
1,
(255,255,255),
2
)
#putting a label of the probability
cv2.putText(
framemain,
str(confidence),
(x+5,y+h-5),
cv2.FONT_HERSHEY_SIMPLEX,
1,
(255,255,0),
1
)
#to collect data for training
if sample_count<=30:
sample_count+=1
cv2.imwrite(
"TrainingImages/ "+Name +'.'+str(ID)+"."+ str(
sample_count) + ".jpg", gray[y:y + h, x:x + w])
# this image is used to represent the person detected
#this saves in the ProfileImages folder
if sample_count==10:
propic=cv2.resize(frameside, (400, 300))
cv2.imwrite(
"ProfileImages/ "+str(ID)+".jpg", propic[y-int(h/10):y + h+int(h/10), x-int(w/10):x + w+int(w/10)])
if sample_count==31:
messagebox.showinfo(title="Message", message="Finished taking photos",icon='info')
cv2image = cv2.cvtColor(framemain, cv2.COLOR_BGR2RGBA)
cv2image1 = cv2.cvtColor(frameside, cv2.COLOR_BGR2RGBA)
cv2image2 = cv2.cvtColor(framebright, cv2.COLOR_BGR2RGBA)
img = PIL.Image.fromarray(cv2image)
img1 = PIL.Image.fromarray(cv2image1)
img2 = PIL.Image.fromarray(cv2image2)
#this is for changing the brightness of the frames
if v1.get()==0.0:
img2=img2
else:
img2=ImageEnhance.Brightness(img2).enhance(1+2/10*v1.get() if v1.get()>0.0 else 1-1/10*(-1*v1.get()))
imgtk = ImageTk.PhotoImage(image=img)
imgtk1 = ImageTk.PhotoImage(image=img1)
imgtk2 = ImageTk.PhotoImage(image=img2)
lmain.imgtk = imgtk
lmain.configure(image=imgtk)
if not DetectMode:
if label2.winfo_exists():
label2.destroy()
Ck2=True
if label3.winfo_exists():
label3.destroy()
Ck3=True
if label4.winfo_exists():
print("Test")
label4.destroy()
label5.destroy()
label6.destroy()
Ck4=True
if label7.winfo_exists():
label7.destroy()
Ck7=True
if W:
lside=Label(gui)
lside.place(x=650, y=200,anchor='c')
lbright=Label(gui)
lbright.place(x=900, y=200,anchor='c')
s1 = Scale( gui, variable = v1,
from_ = -10, to = 10,
orient = HORIZONTAL,length=300,width=20,tickinterval=1)
s1.place(x=775, y=325,anchor='c')
W=False
lside.imgtk1 = imgtk1
lbright.imgtk2 = imgtk2
lside.configure(image=imgtk1)
lbright.configure(image=imgtk2)
else:
data3=pd.read_csv("Data.csv")
if lside.winfo_exists() and lbright.winfo_exists() and s1.winfo_exists():
lside.destroy()
lbright.destroy()
s1.destroy()
W=True
if face_count>=8:
if label3.winfo_exists():
label3.destroy()
Ck3=True
if label4.winfo_exists():
label4.destroy()
label5.destroy()
label6.destroy()
Ck4=True
if label2.winfo_exists():
label2.destroy()
Ck2=True
if Ck7:
label7=Label(gui, height=1,bg="light green",justify=tk.CENTER,font=("Helvetica", 30))
label7.place(x=775, y=400,anchor='c')
Ck7=False
label7.configure(text="No Face Detected")
else:
if len(ListDetect)==10:
Info=max(set(ListDetect), key=ListDetect.count)
if Info=='unknown':
if label3.winfo_exists():
label3.destroy()
Ck3=True
if label4.winfo_exists():
label4.destroy()
label5.destroy()
label6.destroy()
Ck4=True
if label7.winfo_exists():
label7.destroy()
Ck7=True
if Ck2:
label2=Label(gui, bg="light green",font=("Helvetica", 45))
label2.place(x=775, y=400,anchor='c')
Ck2=False
label2.configure(text="Unknown")
elif Info=="Detecting.......":
if Ck3:
label3=Label(gui, bg="light green",font=("Helvetica", 45))
label3.place(x=775, y=400,anchor='c')
Ck3=False
label3.configure(text='Detecting.......')
else:
if label2.winfo_exists():
label2.destroy()
Ck2=True
if label3.winfo_exists():
label3.destroy()
Ck3=True
if label7.winfo_exists():
label7.destroy()
Ck7=True
if Ck4:
label4=Label(gui, height=4,bg="light green",justify=tk.LEFT,font=("Helvetica", 25))
label4.place(x=775, y=550,anchor='c')
label5=Label(gui)
label5.place(x=775, y=290,anchor='c')
label6=Label(gui, height=1,bg="light green",justify=tk.CENTER,font=("Helvetica", 30))
label6.place(x=775, y=100,anchor='c')
Ck4=False
Txt='Username : '+str(data3.at[int(Info)-1,"UserID"])+"\n"+'Name : '+str(data3.at[int(Info)-1,"Name"])+"\n"+'Age : '+str(data3.at[int(Info)-1,"Age"])+"\n"+'Address : '+str(data3.at[int(Info)-1,"Address"])
label4.configure(text=Txt)
label6.configure(text="Face Detected")
try:
imgtk5=ProfileLoad(int(Info))
label5.imgtk5 = imgtk5
label5.configure(image=imgtk5)
except:
pass
#this calls the function again. So this time we will process the next frame coming from the web cam stream
lmain.after(30, show_Mainframe)
########################################################################
#path to the pre-trained face detector
cascPath=os.path.dirname(cv2.__file__)+"/data/haarcascade_frontalface_default.xml"
#loading the face detector
faceCascade = cv2.CascadeClassifier(cascPath)
#creating a recognizer(LBPH)
recognizer = cv2.face.LBPHFaceRecognizer_create()
#setting up the web cam
width, height = 400, 300
capmain = cv2.VideoCapture(0)
capmain.set(cv2.CAP_PROP_FRAME_WIDTH, width)
capmain.set(cv2.CAP_PROP_FRAME_HEIGHT, height)
data1=pd.read_csv("Data.csv")
ID=len(data1['ID'])
#staring the main window
gui = Tk()
v1 = DoubleVar()
gui.configure(background="light green")
gui.title("Face Recognition System")
gui.geometry("1050x775")
#setting up the title
label1=Label(gui,bg='black',fg='white',text="FACE RECOGNITION SYSTEM", font=("Helvetica", 30))
label1.place(x=525, y=25,anchor='c')
#setting up the main display unit
lmain = Label(gui)
lmain.place(x=250, y=275,anchor='c')
if not DetectMode:
lside=Label(gui)
lside.place(x=650, y=200,anchor='c')
lbright=Label(gui)
lbright.place(x=900, y=200,anchor='c')
s1 = Scale( gui, variable = v1,
from_ = -10, to = 10,
orient = HORIZONTAL,length=300,width=20,tickinterval=1)
s1.place(x=775, y=325,anchor='c')
print(lside.winfo_exists())
label2=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 25))
label2.place(x=775, y=500,anchor='c')
label3=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 25))
label3.place(x=775, y=550,anchor='c')
label4=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 16))
label4.place(x=775, y=600,anchor='c')
label5=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 16))
label5.place(x=775, y=600,anchor='c')
label6=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 16))
label6.place(x=775, y=600,anchor='c')
label7=Label(gui, text="Initial Testing", bg="light green",font=("Helvetica", 16))
label7.place(x=775, y=600,anchor='c')
#setting up labels for the text entry boxes
heading = Label(gui, text="Form", bg="light green",font=("Helvetica", 20))
heading.place(x=250, y=505,anchor='c')
username = Label(gui, text="Username", bg="light green",font=("Helvetica", 16))
username.place(x=125, y=540,anchor='c')
name = Label(gui, text="Name", bg="light green",font=("Helvetica", 16))
name.place(x=125, y=590,anchor='c')
age = Label(gui, text="Age", bg="light green",font=("Helvetica", 16))
age.place(x=125, y=640,anchor='c')
address = Label(gui, text="Address", bg="light green",font=("Helvetica", 16))
address.place(x=125, y=690,anchor='c')
#setting up text entry boxes
username_field = Entry(gui)
username_field.place(x=300, y=540,anchor='c')
name_field = Entry(gui)
name_field.place(x=300, y=590,anchor='c')
age_field = Entry(gui)
age_field.place(x=300, y=640,anchor='c')
address_field = Entry(gui)
address_field.place(x=300, y=690,anchor='c')
username_field.bind("<Return>", focus1)
name_field.bind("<Return>", focus2)
age_field.bind("<Return>", focus3)
#setting up buttons of the main window
addperson = Button(gui, text="Add Person", fg="Black",
bg="Red", command=insert,height=2, width=20)
addperson.place(x=250, y=745,anchor='c')
DetectFace = Button(gui, text="Detect Face", fg="Black",
bg="Red", command=Detect,height=2, width=20)
DetectFace.place(x=250, y=460,anchor='c')
MonitorB = Button(gui, text="Monitor", fg="Black",
bg="Red", command=Monitor,height=2, width=20)
MonitorB.place(x=360, y=85,anchor='c')
UR = Button(gui, text="User Registration",command=DataBaseManagement, fg="Black",
bg="Red",height=2, width=20)
UR.place(x=140, y=85,anchor='c')
QuitAll=Button(gui, text="Quit",command=Quit, fg="Black",
bg="Red",height=2, width=20)
QuitAll.place(x=925, y=745,anchor='c')
show_Mainframe()
gui.mainloop()
########################################################################
capmain.release()
data=pd.read_csv("Data.csv")
list(data["UserID"])
len(data["ID"])
sample_count
os.listdir('TrainingImageLabel')
file1 = open("TrainingImageLabel/Counter.txt","w")
file1.write(str(1))
file1.close()
file1 = open("TrainingImageLabel/Counter.txt","r")
CountV=file1.readlines(0)
file1.close()
print(CountV[0])
file1 = open("TrainingImageLabel/Counter.txt","w")
file1.write(str(2))
file1.close()
data3=pd.read_csv("Data.csv")
print(data3)
ID=1
data3.at[ID-1,"UserID"]
DetectMode
lst=['a','a','b','c','c']
print(max(set(lst), key=lst.count))
imagePaths =[os.path.join('ProfileImages', f) for f in os.listdir('ProfileImages')]
for imagePath in imagePaths:
print(imagePath)
from tkinter import *
cv2.__version__
b=pd.read_csv("Data.csv")
b[b['ID']==3]["Name"]
b.at[2,"Name"]
Pdata=pd.read_csv("Data.csv")
Pdata=Pdata.drop(Pdata.index[4])
Pdata.to_csv("Data.csv",index=False,header=True)
```
| github_jupyter |
# Load Specter Embeddings
```
%load_ext autoreload
%autoreload 2
import pandas as pd
from pathlib import PurePath
VECTOR_DOWNSAMPLED = 192
```
## 1. Load Embeddings
```
VECTOR_COLS = [str(i) for i in range(768)]
COLUMNS = ['cord_uid'] + VECTOR_COLS
spector_path = PurePath('../data/CORD-19-research-challenge') / "cord_19_embeddings_4_24/cord_19_embeddings_4_24.csv"
%time embeddings = pd.read_csv(spector_path, names=COLUMNS).set_index('cord_uid')
embeddings
embeddings.shape
```
## 2. Downsample Document Vectors
```
import numpy as np
from sklearn.decomposition import PCA
RANDOM_STATE = 42
N_CLUSTERS = 6
def kmean_labels(docvectors, n_clusters=6, random_state=RANDOM_STATE):
print('Setting cluster labels')
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=n_clusters,
random_state=random_state).fit(docvectors)
return kmeans.labels_, kmeans
def downsample(docvectors, dimensions=2):
print(f'Downsampling to {dimensions}D embeddings')
pca = PCA(n_components=dimensions, svd_solver='full')
docvectors_downsampled = pca.fit_transform(docvectors)
return np.squeeze(docvectors_downsampled), pca
vectors_downsampled, pca_downsampled = downsample(embeddings,VECTOR_DOWNSAMPLED)
```
## 2. Create Full Spector Annoy Index
```
from annoy import AnnoyIndex
from pathlib import Path
def cord_support_dir():
return Path('../cord') / 'cordsupport'
def create_annoy_index(vectors, num_trees=30):
print('Building Annoy index from vectors', vectors.shape)
num_papers, vector_length = vectors.shape
annoy_index = AnnoyIndex(vector_length, 'angular')
for i in range(num_papers):
annoy_index.add_item(i, vectors[i])
annoy_index.build(num_trees)
index_path = str((Path(cord_support_dir()) / f'DocumentIndex{vector_length}.ann').resolve())
print('Saving annoy index to', index_path)
annoy_index.save(index_path)
del annoy_index
create_annoy_index(vectors_downsampled)
```
## 3. Downsample to 2d and 1d, and get cluster ids
```
vectors_2d, pca_2d = downsample(embeddings,2)
vectors_1d, pca_1d = downsample(embeddings,1)
cluster_ids, kmeans = kmean_labels(embeddings)
```
## 4. Save to cord support dir
```
from pathlib import PurePath
document_vectors = pd.DataFrame({'cluster': cluster_ids,
'x': vectors_2d[:,0],
'y': vectors_2d[:,1],
'1d': vectors_1d.tolist(),
'2d': vectors_2d.tolist()
}, index=embeddings.index)
document_vectors.to_parquet(PurePath(cord_support_dir()) / 'DocumentVectors.pq', compression='gzip')
```
## 5. Load Document Vectors
```
document_vectors = pd.read_parquet(PurePath(cord_support_dir()) / 'DocumentVectors.pq')
assert len(document_vectors) == len(embeddings), \
f'The document vectors {len(document_vectors)} and the embeddings {len(embeddings)} are not the same length'
document_vectors
```
## Chart
```
import altair as alt
alt.Chart(document_vectors.sample(5000)).mark_circle(opacity=0.5, size=25).encode(
x=alt.X('x', axis=None),
y=alt.Y('y', axis=None),
color=alt.Color('cluster:N', legend=None),
).configure_axis(
grid=False
).configure_view(
strokeWidth=0
).properties(
title='COVID-19 Research Papers'
)
from cord import ResearchPapers
metadata = ResearchPapers.load_metadata()
COLS = ['cord_uid', 'title','covid_related','virus','coronavirus','sars', 'cluster']
docs = document_vectors.merge(metadata, on='cord_uid', how='left')[COLS]
cluster_pct = docs[['cluster', 'covid_related']].groupby('cluster').sum() * 100 / \
docs[['cluster', 'covid_related']].groupby('cluster').count()
import matplotlib.style as style
style.use('fivethirtyeight')
cluster_pct.sort_values('covid_related').plot.barh(grid=False);
```
### Top Cluster
```
top_cluster = cluster_pct[cluster_pct.covid_related ==cluster_pct.covid_related.max()].index[0]
pd.options.display.max_rows = 400
a = 'time ; well; spent'
a.partition(';')
```
| github_jupyter |
## RIHAD VARIAWA, Data Scientist - Who has fun LEARNING, EXPLORING & GROWING
## Currency Conversion with Matrix Multiplication
In this notebook you will solve a currency problem using matrix multiplication and the python package [NumPy](http://www.numpy.org/). This demonstration is provided to prepare you for using matrix multiplication to solve more complex problems.
## Currency Conversion Problem
Over the years you have traveled to eight different countries and just happen to have leftover local currency from each of your trips.
You are planning to return to one of the eight countries, but you aren't sure which one just yet.
You are waiting to find out which will have the cheapest airfare.
In preparation, for the trip you *will* want convert *all* your local currency into the currency local of the place you will be traveling to.
Therefore, to double check the bank's conversion of your currency, you want to compute the total amount of currency you would expect for each of the eight countries.
To compute the conversion you first need to import a matrix that contains the currency conversion rates for each of the eight countries. The data we will be use comes from the [Overview Matrix of Exchange Rates from Bloomberg Cross-Rates _Overall Chart_](https://www.bloomberg.com/markets/currencies/cross-rates) on January, 10 2018.
<img src="currencyProbImage.png" height=300 width=750>
You can think about this problem as taking a _vector of **inputs**_ (the currencies from the 8 countries) and applying a _matrix of **weights**_ (the conversion rates matrix) to these inputs to produce a _vector of **outputs**_ (total amount of currency for each country) using matrix multiplication with the NumPy package.
### Coding the Currency Conversion Problem
First you will need to create the _**inputs** vector_ that holds the currency you have from the eight countries into a numpy vector. To begin, first import the NumPy package and then use the package to create a vector from a list. Next we convert the vector into a pandas dataframe so that it will print out nicely below with column labels to indicate the country the currency amount is associated to.
```
import numpy as np
import pandas as pd
# Creates numpy vector from a list to represent money (inputs) vector.
money = np.asarray([70, 100, 20, 80, 40, 70, 60, 100])
# Creates pandas dataframe with column labels(currency_label) from the numpy vector for printing.
currency_label = ["USD", "EUR", "JPY", "GBP", "CHF", "CAD", "AUD", "HKD"]
money_df = pd.DataFrame(data=money, index=currency_label, columns=["Amounts"])
print("Inputs Vector:")
money_df.T
```
Next we need to create the _**weights** matrix_ by importing the currency conversion rates matrix. We will use python package [Pandas](https://pandas.pydata.org/) to quickly read in the matrix and approriately assign row and colunm labels. Additionally, we define a variable **_path_** to define the location of the currency conversion matrix. The code below imports this weights matrix, converts the dataframe into a numpy matrix, and displays its content to help you determine how to solve the problem using matrix multiplication.
```
# Sets path variable to the 'path' of the CSV file that contains the conversion rates(weights) matrix.
path = %pwd
# Imports conversion rates(weights) matrix as a pandas dataframe.
conversion_rates_df = pd.read_csv(path+"/currencyConversionMatrix.csv",header=0,index_col=0)
# Creates numpy matrix from a pandas dataframe to create the conversion rates(weights) matrix.
conversion_rates = conversion_rates_df.values
# Prints conversion rates matrix.
print("Weights Matrix:")
conversion_rates_df
```
The _**weights** matrix_ above provides the conversion rates between each of the eight countries. For example, in row 1, column 1 the value **1.0000** represents the conversion rate from US dollars to US dollars. In row 2, column 1 the value **1.1956** represents that 1 Euro is worth **1.1956** US dollars. In row 1, column 2 the value **0.8364** represents that 1 US dollar is only worth **0.8364** Euro.
The _**outputs** vector_ is computed below using matrix multiplication. The numpy package provides the [function _**matmul**_](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.matmul.html) for multiplying two matrices (or a vector and a matrix). Below you will find the equation for matrix multiplication as it applies to AI, where the _**inputs** vector_($x_{1}...x_{n}$) multiplied by the _**weights** matrix_($w_{11}...w_{nm}$) to compute the _**outputs** vector_($y_{1}...y_{m}$).
$\hspace{4cm} \begin{bmatrix} x_{1}&x_{2}&...&x_{n}\end{bmatrix} \begin{bmatrix} w_{11}&w_{12}&...&w_{1m}\\ w_{21}&w_{22}&...&w_{2m}\\ ...&...&...&... \\ w_{n1}&w_{n2}&...&w_{nm}\end{bmatrix} = \begin{bmatrix} y_{1}&y_{2}&...&y_{m}\end{bmatrix}$
The example matrix multiplication below, has $n$ as 4 in **inputs** and **weights** and $m$ as 3 in **weights** and **outputs**.
$\hspace{4cm} \begin{bmatrix} 10 & 2 & 1 & 5\end{bmatrix} \begin{bmatrix} 1 & 20 & 7\\ 3 & 15 & 6 \\ 2 & 5 & 12 \\ 4 & 25 & 9 \end{bmatrix} = \begin{bmatrix} 38 & 360 & 139 \end{bmatrix}$
As seen with the example above, matrix multiplication resulting matrix(_**outputs** vector_) will have same row dimension as the first matrix(_**inputs** vector_) and the same column dimension as the second matrix(_**weights** matrix_). With the currency example the number of columns in the inputs and weights matrices are the same, but this won't always be the case in AI.
## TODO: Matrix Multiplication
Replace the **None** below with code that uses the [function _**matmul**_](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.matmul.html) for multiplying **money** and **conversion_rates** to compute the vector **money_totals**. Recall that we used the alias _**np**_ when we imported the Numpy package above, so be certain to use the _**np**_ alias when calling the _**matmul**_ function below. Additionally, be certain to select _'Cell'_ and _'Run All'_ to check the code you insert below.
```
# TODO 1.: Calculates the money totals(outputs) vector using matrix multiplication in numpy.
money_totals = None
# Converts the resulting money totals vector into a dataframe for printing.
money_totals_df = pd.DataFrame(data = money_totals, index = currency_label, columns = ["Money Totals"])
print("Outputs Vector:")
money_totals_df.T
```
### Solution for Currrency Conversion with Matrix Multiplication
Your output from above should match the **Money Totals** displayed below. If you need any help or want to check your answer, feel free to check out the solution notebook by clicking [here](matrixMultCurrencySolution.ipynb). The results can be interperted as converting all the currency to US dollars(**USD**) would provide **454.28** US dollars, converting all the currency to Euros(**EUR**) would provide **379.96** Euros, and etc.
<img src="money_totals.png" height=225 width=563>
### Solution Video for Currrency Conversion with Matrix Multiplication
The solution video can be found in the **Linear Mapping Lab Solution** section. You may want to open another browser window to allow you to easily toggle between the Vector's Lab Jupyter Notebook and the solution videos for this lab.
| github_jupyter |
```
import numpy as np
import pandas as pd
import linearsolve as ls
import matplotlib.pyplot as plt
plt.style.use('classic')
%matplotlib inline
```
# Class 17: Introduction to New-Keynesian Business Cycle Modeling
In this notebook, we will use `linearsolve` to compute impulse responses of output, inflation, and the nominal interest rate to an aggregate demand shock in the New-Keynesian model.
## The New-Keynesian Model
The most basic version of the New-Keynesian Model can be expressed as:
\begin{align}
y_t & = E_t y_{t+1} - \left( r_{t} - \bar{r}\right) + g_t\\
i_{t} & = r_{t} + E_t \pi_{t+1}\\
i_{t} & = \bar{r} + \pi^T + \phi_{\pi}\big(\pi_t - \pi^T\big) + \phi_{y}\big(y_t - \bar{y}\big) + v_t\\
\pi_t -\pi^T & = \beta \left( E_t\pi_{t+1} - \pi^T\right) + \kappa (y_t -\bar{y})+ u_t,
\end{align}
where: $y_t$ is (log) output, $r_t$ is the real interest rate, $i_t$ is the nominal interest rate, $\pi_t$ is the rate of inflation between periods $t-1$ and $t$, $\bar{r}$ is the long-run average real interest rate or the *natural rate of interest*, $\beta$ is the household's subjective discount factor, and $\pi^T$ is the central bank's inflation target. The coeffieints $\phi_{\pi}$ and $\phi_{y}$ reflect the degree of intensity to which the central bank *endogenously* adjusts the nominal interest rate in response to movements in inflation and output.
The variables $g_t$, $u_t$, and $v_t$ represent exogenous shocks to aggregate demand, inflation, and monetary policy. They follow AR(1) processes:
\begin{align}
g_{t+1} & = \rho_g g_{t} + \epsilon^g_{t+1}\\
u_{t+1} & = \rho_u u_{t} + \epsilon^u_{t+1}\\
v_{t+1} & = \rho_v v_{t} + \epsilon^v_{t+1}.
\end{align}
The goal is to compute impulse responses in the model to a one percent exogenous increase in the nominal interest rate. We will use the following parameterization:
| $$\bar{y}$$ | $$\beta$$ | $$\bar{r}$$ | $$\kappa$$ | $$\pi^T$$ | $$\phi_{\pi}$$ | $$\phi_y$$ | $$\rho_g$$ | $$\rho_u$$ | $$\rho_v$$ |
|-------------|-----------|----------------|------------|-----------|----------------|------------|------------|------------|------------|
| 0 | 0.995 | $$-\log\beta$$ | 0.25 | 0.02/4 | 1.5 | 0.5/4 | 0.5 | 0.5 | 0.5 |
```
# Create a variable called 'parameters' that stores the model parameter values in a Pandas Series
parameters = pd.Series(dtype=float)
parameters['y_bar'] = 0
parameters['beta'] = 0.995
parameters['r_bar'] = -np.log(parameters.beta)
parameters['kappa'] = 0.25
parameters['pi_T'] = 0.02/4
parameters['phi_pi'] = 1.5
parameters['phi_y'] = 0.5/4
parameters['rho_g'] = 0.5
parameters['rho_u'] = 0.5
parameters['rho_v'] = 0.5
# Print the model's parameters
print(parameters)
# Create variable called 'var_names' that stores the variable names in a list with state variables ordered first
var_names = ['g','u','v','y','pi','i','r']
# Create variable called 'shock_names' that stores an exogenous shock name for each state variable.
shock_names = ['e_g','e_u','e_v']
# Define a function that evaluates the equilibrium conditions of the model solved for zero. PROVIDED
def equilibrium_equations(variables_forward,variables_current,parameters):
# Parameters. PROVIDED
p = parameters
# Current variables. PROVIDED
cur = variables_current
# Forward variables. PROVIDED
fwd = variables_forward
# IS equation
is_equation = fwd.y - (cur.r -p.r_bar) + cur.g - cur.y
# Fisher_equation
fisher_equation = cur.r + fwd.pi - cur.i
# Monetary policy
monetary_policy = p.r_bar + p.pi_T + p.phi_pi*(cur.pi - p.pi_T) + p.phi_y*cur.y + cur.v - cur.i
# Phillips curve
phillips_curve = p.beta*(fwd.pi- p.pi_T) + p.kappa*cur.y + cur.u - (cur.pi-p.pi_T)
# Demand process
demand_process = p.rho_g*cur.g - fwd.g
# Monetary policy process
monetary_policy_process = p.rho_v*cur.v - fwd.v
# Inflation process
inflation_process = p.rho_u*cur.u - fwd.u
# Stack equilibrium conditions into a numpy array
return np.array([
is_equation,
fisher_equation,
monetary_policy,
phillips_curve,
demand_process,
monetary_policy_process,
inflation_process
])
# Initialize the model into a variable named 'nk_model'
nk_model = ls.model(equations = equilibrium_equations,
n_states=3,
var_names=var_names,
shock_names=shock_names,
parameters = parameters)
# Compute the steady state numerically using .compute_ss() method of nk_model
guess = [0,0,0,0,0.01,0.01,0.01]
nk_model.compute_ss(guess)
# Print the computed steady state
print(nk_model.ss)
# Find the log-linear approximation around the non-stochastic steady state and solve using .approximate_and_solve() method of nk_model
# set argumement 'log_linear' to False because the model is already log-linear.
nk_model.approximate_and_solve(log_linear=False)
print(nk_model.approximated())
```
### Impulse Responses
Compute a 21 period impulse response of the model's variables to a 0.01 unit shock to the aggregate demand for goods ($g_t$) in period 5.
```
# Compute impulse responses
nk_model.impulse(T=21,t0=5,shocks=[0.01,0,0])
# Print the first 10 rows of the computed impulse responses to the demand shock
print(nk_model.irs['e_g'].head(10))
```
Plot the computed impulses responses of the nominal interest rate, the real interest rate, output, and inflation. Express inflation and interest rates in *annualized* (e.g., multiplied by 4) terms.
```
# Create figure. PROVIDED
fig = plt.figure(figsize=(12,8))
# Create upper-left axis. PROVIDED
ax1 = fig.add_subplot(2,2,1)
# Create upper-right axis. PROVIDED
ax2 = fig.add_subplot(2,2,2)
# Create lower-left axis. PROVIDED
ax3 = fig.add_subplot(2,2,3)
# Create lower-right axis. PROVIDED
ax4 = fig.add_subplot(2,2,4)
# Set axis 1 ylabel
ax1.set_ylabel('% dev from steady state')
# Set axis 2 ylabel
ax2.set_ylabel('% dev from steady state')
# Set axis 3 ylabel
ax3.set_ylabel('% dev from steady state')
# Set axis 4 ylabel
ax4.set_ylabel('% dev from steady state')
# Set axis 1 limits
ax1.set_ylim([-0.5,3.5])
# Set axis 2 limits
ax2.set_ylim([-0.5,3.5])
# Set axis 3 limits
ax3.set_ylim([-0.5,2])
# Set axis 4 limits
ax4.set_ylim([-0.5,2])
# Plot the nominal interest rate, real interest rate, output, and inflation
(nk_model.irs['e_g']['i']*400).plot(ax=ax1,lw=4,alpha=0.75,title='Nominal Interest (annualized)',grid=True)
(nk_model.irs['e_g']['r']*400).plot(ax=ax2,lw=4,alpha=0.75,title='Real Interest (annualized)',grid=True)
(nk_model.irs['e_g']['y']*100).plot(ax=ax3,lw=4,alpha=0.75,title='Output',grid=True)
(nk_model.irs['e_g']['pi']*400).plot(ax=ax4,lw=4,alpha=0.75,title='Inflation (annualized)',grid=True)
```
| github_jupyter |
#### Importieren der Libraries
```
!pip install deeplearning2020
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
from deeplearning2020 import helpers
```
#### Laden des MNIST Dataset
Als Erstes wollen wir den Datensatz wie im Video "Laden und Bearbeiten des MNIST Datensatz" laden.
```
data = keras.datasets.mnist
(train_images, train_labels), (test_images, test_labels) = data.load_data()
```
Wir errinnern uns, dass die Pixelwerte noch nicht in normaliserter Form vorliegen. Wir normalisieren diese also zunächst indem wir durch den maximalen Pixelwert 255 teilen:
```
train_images = train_images / 255.0
test_images = test_images / 255.0
```
Zuletzt hatten wir noch das Problem, dass die Labels des Datensatzes einfach nur Zahlen waren.
- Das Bild der handgeschriebenen 5 hat das Label `5`.
- Wir hätten in diesem Fall jedoch gerne den Vektor `[0. 0. 0. 0. 0. 1. 0. 0. 0. 0.]` als Label, welcher der von unserem Netz gewünschten Ausgabe entspricht. Dieser Vektor hat nur an Stelle 5 (beginnend bei 0) eine 1.
Für detailliertere Erklärungen sei an dieser Stelle wieder auf das Video "Laden und Bearbeiten des MNIST Datensatzes" verwiesen.
```
total_classes = 10
train_vec_labels = keras.utils.to_categorical(train_labels, total_classes)
test_vec_labels = keras.utils.to_categorical(test_labels, total_classes)
```
#### Entwurf eines Netzes
Nun haben wir die Eingabedaten normalisiert und die Labels als Vektoren vorliegen. Wir können also endlich anfangen ein Netz für die Erkennung der handgeschriebenen Zahlen zu bauen! :)
Wir wollen dafür ein sehr einfaches Netz mit 3 Layern definieren (Eingabelayer, Hidden Layer und Ausgabelayer):
- Als **Input-Layer** verwenden wir einen `keras.layers.Flatten` Layer, der die 28x28 Matrizen, die wir als Eingaben erhalten auf $28\cdot 28 = 784$ Neuronen verteilt
- Als nächstes verwenden wir für den **Hidden-Layer** einen `keras.layers.Dense` Layer mit 128 Neuronen, wobei wir 128 als eine gute Anzahl empfinden
- Als **Output-Layer** verwenden wir einen `keras.layers.Dense` Layer mit 10 Neuronen, da wir 10 Klassen (Ziffern von 0-9) erkennen wollen
```
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation='sigmoid'),
keras.layers.Dense(10, activation='sigmoid')
])
```
#### Kompilieren des Netzes
Nach dem wir unser Netz definiert haben, müssen wir es *kompilieren*, bevor wir mit dem Training beginnen können.
In diesem Schritt legen wir wichtige Parameter für die Trainingsphase fest:
- Der **Optimizer** ist der im Training verwendete Lernalgorithmus zur Verbesserung des Netzes. In der letzen Woche haben wir ja bereits *Gradient Descent* und dessen Optimierung *Stochastic Gradient Descent* (SGD, siehe *Deep Dive: Backpropagation*) kennengelernt.
- Der **Loss** ist die verwendete Kostenfunktion. Ziel während des Trainings ist es, diese zu minimieren. Wir haben in Woche 1 bereits die Quadratische Fehlerfunktion (*Squared Error*) kennengelernt.
- Die **Metrics** sind die während des Trainings ausgewerteten Metrics. Bei allen Klassifikationsproblemen interessiert uns hier die `"accuracy"`.
In diesem Beispiel verwenden wir
- Den *Stochastic Gradient Descent* (`"sgd"`) Lernalgorithmus als unseren Optimizer.
- Die `"mean_squared_error"` Kostenfunktion, welche im Vergleich zur normalen *Squared Error* Kostenfunktion nicht die Summe, sondern den Mittelwert der Fehler der Ausgabeneuronen berechnet.
```
# sgd = keras.optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(
optimizer='sgd',
loss='mean_squared_error',
metrics=['accuracy'])
```
#### Trainieren des Netzes
Jetzt können wir endlich unser Netz tranieren. Dazu verwenden wir die `fit` Methode und übergeben unsere Trainingsbilder als Eingaben mit den dazugehörigen Labels als gewünschte Ausgaben. Die Anzahl der `epochs` gibt an, wie oft das Netz das gesamte Trainingsset zu sehen bekommt. Erhöhen wir die Anzahl der Epochen lassen wir unser Netz länger lernen.
```
model.fit(train_images, train_vec_labels, epochs=10, verbose=True)
```
#### Evaluieren des Netzes
Bisher hat das Netz nur Trainingsbilder gesehen und damit gelernt. Ziel ist es aber ja, mit unserem Netz neue Bilder von handgeschriebenen Ziffern zu erkennen. Dafür gibt es ja die Testdaten, mit denen wir unser Netz nun auf die Genauigkeit bei ungesehenen Daten überprüfen wollen.
```
eval_loss, eval_accuracy = model.evaluate(test_images, test_vec_labels, verbose=False)
print("Model accuracy: %.2f" % eval_accuracy)
```
#### Ausgaben des Netzes
```
helpers.plot_predictions(model, test_images[:20], labels=test_vec_labels[:20])
```
| github_jupyter |
```
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
"""
# If you're using Google Colab and not running locally, run this cell
# install NeMo
BRANCH = 'main'
!python -m pip install git+https://github.com/NVIDIA/NeMo.git@$BRANCH#egg=nemo_toolkit[nlp]
# If you're not using Colab, you might need to upgrade jupyter notebook to avoid the following error:
# 'ImportError: IProgress not found. Please update jupyter and ipywidgets.'
! pip install ipywidgets
! jupyter nbextension enable --py widgetsnbextension
# Please restart the kernel after running this cell
from nemo.collections import nlp as nemo_nlp
from nemo.utils.exp_manager import exp_manager
import os
import wget
import torch
import pytorch_lightning as pl
from omegaconf import OmegaConf
```
# Task Description
**Sentiment Analysis** is the task of detecting the sentiment in text. We model this problem as a simple form of a text classification problem. For example `Gollum's performance is incredible!` has a positive sentiment while `It's neither as romantic nor as thrilling as it should be.` has a negative sentiment.
.
# Dataset
In this tutorial we going to use [The Stanford Sentiment Treebank (SST-2)](https://nlp.stanford.edu/sentiment/index.html) corpus for sentiment analysis. This version of the dataset contains a collection of sentences with binary labels of positive and negative. It is a standard benchmark for sentence classification and is part of the GLUE Benchmark: https://gluebenchmark.com/tasks. Please download and unzip the SST-2 dataset from GLUE. It should contain three files of train.tsv, dev.tsv, and test.tsv which can be used for training, validation, and test respectively.
# NeMo Text Classification Data Format
[TextClassificationModel](https://github.com/NVIDIA/NeMo/blob/stable/nemo/collections/nlp/models/text_classification/text_classification_model.py) in NeMo supports text classification problems such as sentiment analysis or domain/intent detection for dialogue systems, as long as the data follows the format specified below.
TextClassificationModel requires the data to be stored in TAB separated files (.tsv) with two columns of sentence and label. Each line of the data file contains text sequences, where words are separated with spaces and label separated with [TAB], i.e.:
```
[WORD][SPACE][WORD][SPACE][WORD][TAB][LABEL]
```
For example:
```
hide new secretions from the parental units[TAB]0
that loves its characters and communicates something rather beautiful about human nature[TAB]1
...
```
If your dataset is stored in another format, you need to convert it to this format to use the TextClassificationModel.
## Download and Preprocess the Data
First, you need to download the zipped file of the SST-2 dataset from the GLUE Benchmark website: https://gluebenchmark.com/tasks, and put it in the current folder. Then the following script would extract it into the data path specified by `DATA_DIR`:
```
DATA_DIR = "DATA_DIR"
WORK_DIR = "WORK_DIR"
os.environ['DATA_DIR'] = DATA_DIR
os.makedirs(WORK_DIR, exist_ok=True)
os.makedirs(DATA_DIR, exist_ok=True)
! unzip -o SST-2.zip -d {DATA_DIR}
```
Now, the data folder should contain the following files:
* train.tsv
* dev.tsv
* test.tsv
The format of `train.tsv` and `dev.tsv` is close to NeMo's format except to have an extra header line at the beginning of the files. We would remove these extra lines. But `test.tsv` has different format and labels are missing for this part of the data.
```
! sed 1d {DATA_DIR}/SST-2/train.tsv > {DATA_DIR}/SST-2/train_nemo_format.tsv
! sed 1d {DATA_DIR}/SST-2/dev.tsv > {DATA_DIR}/SST-2/dev_nemo_format.tsv
! ls -l {DATA_DIR}/SST-2
# let's take a look at the data
print('Contents (first 5 lines) of train.tsv:')
! head -n 5 {DATA_DIR}/SST-2/train_nemo_format.tsv
print('\nContents (first 5 lines) of test.tsv:')
! head -n 5 {DATA_DIR}/SST-2/test.tsv
```
# Model Configuration
Now, let's take a closer look at the model's configuration and learn to train the model from scratch and finetune the pretrained model.
Our text classification model uses a pretrained [BERT](https://arxiv.org/pdf/1810.04805.pdf) model (or other BERT-like models) followed by a classification layer on the output of the first token ([CLS]).
The model is defined in a config file which declares multiple important sections. The most important ones are:
- **model**: All arguments that are related to the Model - language model, tokenizer, head classifier, optimizer, schedulers, and datasets/data loaders.
- **trainer**: Any argument to be passed to PyTorch Lightning including number of epochs, number of GPUs, precision level, etc.
```
# download the model's configuration file
MODEL_CONFIG = "text_classification_config.yaml"
CONFIG_DIR = WORK_DIR + '/configs/'
os.makedirs(CONFIG_DIR, exist_ok=True)
if not os.path.exists(CONFIG_DIR + MODEL_CONFIG):
print('Downloading config file...')
wget.download(f'https://raw.githubusercontent.com/NVIDIA/NeMo/{BRANCH}/examples/nlp/text_classification/conf/' + MODEL_CONFIG, CONFIG_DIR)
print('Config file downloaded!')
else:
print ('config file already exists')
config_path = f'{WORK_DIR}/configs/{MODEL_CONFIG}'
print(config_path)
config = OmegaConf.load(config_path)
```
## this line will print the entire config of the model
print(OmegaConf.to_yaml(config))
# Model Training From Scratch
## Setting up data within the config
We first need to set the num_classes in the config file which specifies the number of classes in the dataset. For SST-2, we have just two classes (0-positive and 1-negative). So we set the num_classes to 2. The model supports more than 2 classes too.
```
config.model.dataset.num_classes=2
```
Among other things, the config file contains dictionaries called dataset, train_ds and validation_ds. These are configurations used to setup the Dataset and DataLoaders of the corresponding config.
Notice that some config lines, including `model.dataset.classes_num`, have `???` as their value, this means that values for these fields are required to be to be specified by the user. We need to specify and set the `model.train_ds.file_name`, `model.validation_ds.file_name`, and `model.test_ds.file_name` in the config file to the paths of the train, validation, and test files if they exist. We may do it by updating the config file or by setting them from the command line.
Let's now set the train and validation paths in the config.
```
config.model.train_ds.file_path = os.path.join(DATA_DIR, 'SST-2/train_nemo_format.tsv')
config.model.validation_ds.file_path = os.path.join(DATA_DIR, 'SST-2/dev_nemo_format.tsv')
# Name of the .nemo file where trained model will be saved.
config.save_to = 'trained-model.nemo'
config.export_to = 'trained-model.onnx'
print("Train dataloader's config: \n")
# OmegaConf.to_yaml() is used to create a proper format for printing the train dataloader's config
# You may change other params like batch size or the number of samples to be considered (-1 means all the samples)
print(OmegaConf.to_yaml(config.model.train_ds))
```
## Building the PyTorch Lightning Trainer
NeMo models are primarily PyTorch Lightning (PT) modules - and therefore are entirely compatible with the PyTorch Lightning ecosystem.
Let's first instantiate a PT Trainer object by using the trainer section of the config.
```
print("Trainer config - \n")
# OmegaConf.to_yaml() is used to create a proper format for printing the trainer config
print(OmegaConf.to_yaml(config.trainer))
```
First you need to create a PT trainer with the params stored in the trainer's config. You may set the number of steps for training with max_steps or number of epochs with max_epochs in the trainer's config.
```
# lets modify some trainer configs
# checks if we have GPU available and uses it
config.trainer.gpus = 1 if torch.cuda.is_available() else 0
# for mixed precision training, uncomment the lines below (precision should be set to 16 and amp_level to O1):
# config.trainer.precision = 16
# config.trainer.amp_level = O1
# disable distributed training when using Colab to prevent the errors
config.trainer.accelerator = None
# setup max number of steps to reduce training time for demonstration purposes of this tutorial
# Training stops when max_step or max_epochs is reached (earliest)
config.trainer.max_epochs = 5
# instantiates a PT Trainer object by using the trainer section of the config
trainer = pl.Trainer(**config.trainer)
```
## Setting up the NeMo Experiment¶
NeMo has an experiment manager that handles the logging and saving checkpoints for us, so let's setup it. We need the PT trainer and the exp_manager config:
```
# The experiment manager of a trainer object can not be set twice. We repeat the trainer creation code again here to prevent getting error when this cell is executed more than once.
trainer = pl.Trainer(**config.trainer)
# exp_dir specifies the path to store the the checkpoints and also the logs, it's default is "./nemo_experiments"
# You may set it by uncommentig the following line
# config.exp_manager.exp_dir = 'LOG_CHECKPOINT_DIR'
# OmegaConf.to_yaml() is used to create a proper format for printing the trainer config
print(OmegaConf.to_yaml(config.exp_manager))
exp_dir = exp_manager(trainer, config.exp_manager)
# the exp_dir provides a path to the current experiment for easy access
print(exp_dir)
```
Before initializing the model, we might want to modify some of the model configs. For example, we might want to modify the pretrained BERT model to another model. The default model is `bert-base-uncased`. We support a variety of models including all the models available in HuggingFace, and Megatron.
```
# complete list of supported BERT-like models
print(nemo_nlp.modules.get_pretrained_lm_models_list())
# specify the BERT-like model, you want to use
# set the `model.language_modelpretrained_model_name' parameter in the config to the model you want to use
config.model.language_model.pretrained_model_name = "bert-base-uncased"
```
Now, we are ready to initialize our model. During the model initialization call, the dataset and data loaders will also be prepared for the training and validation.
Also, the pretrained BERT model will be automatically downloaded. Note it can take up to a few minutes depending on the size of the chosen BERT model for the first time you create the model. If your dataset is large, it also may take some time to read and process all the datasets.
Now we can create the model with the model config and the trainer object like this:
```
model = nemo_nlp.models.TextClassificationModel(cfg=config.model, trainer=trainer)
```
## Monitoring Training Progress
Optionally, you can create a Tensorboard visualization to monitor training progress.
```
try:
from google import colab
COLAB_ENV = True
except (ImportError, ModuleNotFoundError):
COLAB_ENV = False
# Load the TensorBoard notebook extension
if COLAB_ENV:
%load_ext tensorboard
%tensorboard --logdir {exp_dir}
else:
print("To use tensorboard, please use this notebook in a Google Colab environment.")
```
## Training
You may start the training by using the trainer.fit() method. The number of steps/epochs of the training are specified already in the config of the trainer and you may update them before creating the trainer.
```
# start model training
trainer.fit(model)
model.save_to(config.save_to)
```
# Evaluation
To see how the model performs, we can run evaluate and test the performance of the trained model on a data file. Here we would load the best checkpoint (the one with the lowest validation loss) and create a model (eval_model) from the checkpoint. We would also create a new trainer (eval_trainer) to show how it is done when training is done and you have just the checkpoints. If you want to perform the evaluation in the same script as the training's script, you may still use the same model and trainer you used for training.
```
# extract the path of the best checkpoint from the training, you may update it to any checkpoint
checkpoint_path = trainer.checkpoint_callback.best_model_path
# Create an evaluation model and load the checkpoint
eval_model = nemo_nlp.models.TextClassificationModel.load_from_checkpoint(checkpoint_path=checkpoint_path)
# create a dataloader config for evaluation, the same data file provided in validation_ds is used here
# file_path can get updated with any file
eval_config = OmegaConf.create({'file_path': config.model.validation_ds.file_path, 'batch_size': 64, 'shuffle': False, 'num_samples': -1})
eval_model.setup_test_data(test_data_config=eval_config)
#eval_dataloader = eval_model._create_dataloader_from_config(cfg=eval_config, mode='test')
# a new trainer is created to show how to evaluate a checkpoint from an already trained model
# create a copy of the trainer config and update it to be used for final evaluation
eval_trainer_cfg = config.trainer.copy()
eval_trainer_cfg.gpus = 1 if torch.cuda.is_available() else 0 # it is safer to perform evaluation on single GPU as PT is buggy with the last batch on multi-GPUs
eval_trainer_cfg.accelerator = None # 'ddp' is buggy with test process in the current PT, it looks like it has been fixed in the latest master
eval_trainer = pl.Trainer(**eval_trainer_cfg)
eval_trainer.test(model=eval_model, verbose=False) # test_dataloaders=eval_dataloader,
```
# Inference
You may create a model from a saved checkpoint and use the model.infer() method to perform inference on a list of queries. There is no need of any trainer for inference.
```
# extract the path of the best checkpoint from the training, you may update it to any other checkpoint file
checkpoint_path = trainer.checkpoint_callback.best_model_path
# Create an evaluation model and load the checkpoint
infer_model = nemo_nlp.models.TextClassificationModel.load_from_checkpoint(checkpoint_path=checkpoint_path)
```
To see how the model performs, let’s get model's predictions for a few examples:
```
# move the model to the desired device for inference
# we move the model to "cuda" if available otherwise "cpu" would be used
if torch.cuda.is_available():
infer_model.to("cuda")
else:
infer_model.to("cpu")
# define the list of queries for inference
queries = ['by the end of no such thing the audience , like beatrice , has a watchful affection for the monster .',
'director rob marshall went out gunning to make a great one .',
'uneasy mishmash of styles and genres .']
# max_seq_length=512 is the maximum length BERT supports.
results = infer_model.classifytext(queries=queries, batch_size=3, max_seq_length=512)
print('The prediction results of some sample queries with the trained model:')
for query, result in zip(queries, results):
print(f'Query : {query}')
print(f'Predicted label: {result}')
```
## Training Script
If you have NeMo installed locally (eg. cloned from the Github), you can also train the model with `examples/nlp/text_classification/text_classifciation_with_bert.py`. This script contains an example on how to train, evaluate and perform inference with the TextClassificationModel.
For example the following would train a model for 50 epochs in 2 GPUs on a classification task with 2 classes:
```
# python text_classification_with_bert.py
model.dataset.num_classes=2
model.train_ds=PATH_TO_TRAIN_FILE
model.validation_ds=PATH_TO_VAL_FILE
trainer.max_epochs=50
trainer.gpus=2
```
This script would also reload the best checkpoint after the training is done and does evaluation on the dev set. Then perform inference on some sample queries.
By default, this script uses `examples/nlp/text_classification/conf/text_classifciation_config.py` config file, and you may update all the params in the config file from the command line. You may also use another config file like this:
```
# python text_classification_with_bert.py --config-name==PATH_TO_CONFIG_FILE
model.dataset.num_classes=2
model.train_ds=PATH_TO_TRAIN_FILE
model.validation_ds=PATH_TO_VAL_FILE
trainer.max_epochs=50
trainer.gpus=2
```
## Deployment
You can also deploy a model to an inference engine (like TensorRT or ONNXRuntime) using ONNX exporter.
If you don't have one, let's install it:
```
!pip install --upgrade onnxruntime # for gpu, use onnxruntime-gpu
# !mkdir -p ort
# %cd ort
# !git clean -xfd
# !git clone --depth 1 --branch v1.8.0 https://github.com/microsoft/onnxruntime.git .
# !./build.sh --skip_tests --config Release --build_shared_lib --parallel --use_cuda --cuda_home /usr/local/cuda --cudnn_home /usr/lib/x86_64-linux-gnu --build_wheel
# !pip uninstall -y onnxruntime
# !pip uninstall -y onnxruntime-gpu
# !pip install --upgrade --force-reinstall ./build/Linux/Release/dist/onnxruntime*.whl
# %cd ..
```
Then export
```
model.export(config.export_to)
```
And run some queries
```
import numpy as np
import torch
from nemo.utils import logging
from nemo.collections.nlp.parts.utils_funcs import tensor2list
from nemo.collections.nlp.models.text_classification import TextClassificationModel
from nemo.collections.nlp.data.text_classification import TextClassificationDataset
import onnxruntime
def to_numpy(tensor):
return tensor.detach().cpu().numpy() if tensor.requires_grad else tensor.cpu().numpy()
def postprocessing(results, labels):
return [labels[str(r)] for r in results]
def create_infer_dataloader(model, queries):
batch_size = len(queries)
dataset = TextClassificationDataset(tokenizer=model.tokenizer, queries=queries, max_seq_length=512)
return torch.utils.data.DataLoader(
dataset=dataset,
batch_size=batch_size,
shuffle=False,
num_workers=2,
pin_memory=True,
drop_last=False,
collate_fn=dataset.collate_fn,
)
queries = ["by the end of no such thing the audience , like beatrice , has a watchful affection for the monster .",
"director rob marshall went out gunning to make a great one .",
"uneasy mishmash of styles and genres .",
"I love exotic science fiction / fantasy movies but this one was very unpleasant to watch . Suggestions and images of child abuse , mutilated bodies (live or dead) , other gruesome scenes , plot holes , boring acting made this a regretable experience , The basic idea of entering another person's mind is not even new to the movies or TV (An Outer Limits episode was better at exploring this idea) . i gave it 4 / 10 since some special effects were nice ."]
model.eval()
infer_datalayer = create_infer_dataloader(model, queries)
ort_session = onnxruntime.InferenceSession(config.export_to)
for batch in infer_datalayer:
input_ids, input_type_ids, input_mask, subtokens_mask = batch
ort_inputs = {ort_session.get_inputs()[0].name: to_numpy(input_ids),
ort_session.get_inputs()[1].name: to_numpy(input_mask),
ort_session.get_inputs()[2].name: to_numpy(input_type_ids),}
ologits = ort_session.run(None, ort_inputs)
alogits = np.asarray(ologits)
logits = torch.from_numpy(alogits[0])
preds = tensor2list(torch.argmax(logits, dim = -1))
processed_results = postprocessing(preds, {"0": "negative", "1": "positive"})
logging.info('The prediction results of some sample queries with the trained model:')
for query, result in zip(queries, processed_results):
logging.info(f'Query : {query}')
logging.info(f'Predicted label: {result}')
break
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Reducer/mean_std_image.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Reducer/mean_std_image.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Reducer/mean_std_image.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Load a Landsat 8 image.
image = ee.Image('LANDSAT/LC08/C01/T1/LC08_044034_20140318')
# Combine the mean and standard deviation reducers.
reducers = ee.Reducer.mean().combine(**{
'reducer2': ee.Reducer.stdDev(),
'sharedInputs': True
})
# Use the combined reducer to get the mean and SD of the image.
stats = image.reduceRegion(**{
'reducer': reducers,
'bestEffort': True,
})
# Display the dictionary of band means and SDs.
print(stats.getInfo())
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Chapter 8 - Movie Review Example
```
%pylab inline
import pandas
d = pandas.read_csv("data/movie_reviews.tsv", delimiter="\t")
# Holdout split
split = 0.7
d_train = d[:int(split*len(d))]
d_test = d[int((1-split)*len(d)):]
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer()
features = vectorizer.fit_transform(d_train.review)
i = 45000
j = 10
words = vectorizer.get_feature_names()[i:i+10]
pandas.DataFrame(features[j:j+7,i:i+10].todense(), columns=words)
float(features.getnnz())*100 / (features.shape[0]*features.shape[1])
from sklearn.naive_bayes import MultinomialNB
model1 = MultinomialNB()
model1.fit(features, d_train.sentiment)
pred1 = model1.predict_proba(vectorizer.transform(d_test.review))
from sklearn.metrics import accuracy_score, roc_auc_score, classification_report, roc_curve
def performance(y_true, pred, color="g", ann=True):
acc = accuracy_score(y_true, pred[:,1] > 0.5)
auc = roc_auc_score(y_true, pred[:,1])
fpr, tpr, thr = roc_curve(y_true, pred[:,1])
plot(fpr, tpr, color, linewidth="3")
xlabel("False positive rate")
ylabel("True positive rate")
if ann:
annotate("Acc: %0.2f" % acc, (0.1,0.8), size=14)
annotate("AUC: %0.2f" % auc, (0.1,0.7), size=14)
performance(d_test.sentiment, pred1)
```
## tf-idf features
```
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer()
features = vectorizer.fit_transform(d_train.review)
pred2 = model1.predict_proba(vectorizer.transform(d_test.review))
performance(d_test.sentiment, pred1, ann=False)
performance(d_test.sentiment, pred2, color="b")
xlim(0,0.5)
ylim(0.5,1)
```
## Parameter optimization
```
param_ranges = {
"max_features": [10000, 30000, 50000, None],
"min_df": [1,2,3],
"nb_alpha": [0.01, 0.1, 1.0]
}
def build_model(max_features=None, min_df=1, nb_alpha=1.0, return_preds=False):
vectorizer = TfidfVectorizer(max_features=max_features, min_df=min_df)
features = vectorizer.fit_transform(d_train.review)
model = MultinomialNB(alpha=nb_alpha)
model.fit(features, d_train.sentiment)
pred = model.predict_proba(vectorizer.transform(d_test.review))
res = {
"max_features": max_features,
"min_df": min_df,
"nb_alpha": nb_alpha,
"auc": roc_auc_score(d_test.sentiment, pred[:,1])
}
if return_preds:
res['preds'] = pred
return res
from itertools import product
results = []
for p in product(*param_ranges.values()):
res = build_model(**dict(zip(param_ranges.keys(), p)))
results.append( res )
print res
opt = pandas.DataFrame(results)
mf_idx = [0,9,18,27]
plot(opt.max_features[mf_idx], opt.auc[mf_idx], linewidth=2)
title("AUC vs max_features")
mdf_idx = [27,28,29]
plot(opt.min_df[mdf_idx], opt.auc[mdf_idx], linewidth=2)
title("AUC vs min_df")
nba_idx = [27,30,33]
plot(opt.nb_alpha[nba_idx], opt.auc[nba_idx], linewidth=2)
title("AUC vs alpha")
pred3 = build_model(nb_alpha=0.01, return_preds=True)['preds']
performance(d_test.sentiment, pred1, ann=False)
performance(d_test.sentiment, pred2, color="b", ann=False)
performance(d_test.sentiment, pred3, color="r")
xlim(0,0.5)
ylim(0.5,1)
```
## Random Forest
```
vectorizer = TfidfVectorizer(strip_accents='unicode', stop_words='english', min_df=3, max_features=30000, norm="l2")
features = vectorizer.fit_transform(d_train.review)
model3 = MultinomialNB()
model3.fit(features, d_train.sentiment)
pred3 = model3.predict_proba(vectorizer.transform(d_test.review))
performance(d_test.sentiment, pred3)
from sklearn.ensemble import RandomForestClassifier
model2 = RandomForestClassifier(n_estimators=100)
model2.fit(features, d_train.sentiment)
pred2 = model2.predict_proba(vectorizer.transform(d_test.review))
performance(d_test.sentiment, pred2)
```
## Word2Vec
```
import re, string
stop_words = set(['all', "she'll", "don't", 'being', 'over', 'through', 'yourselves', 'its', 'before', "he's", "when's", "we've", 'had', 'should', "he'd", 'to', 'only', "there's", 'those', 'under', 'ours', 'has', "haven't", 'do', 'them', 'his', "they'll", 'very', "who's", "they'd", 'cannot', "you've", 'they', 'not', 'during', 'yourself', 'him', 'nor', "we'll", 'did', "they've", 'this', 'she', 'each', "won't", 'where', "mustn't", "isn't", "i'll", "why's", 'because', "you'd", 'doing', 'some', 'up', 'are', 'further', 'ourselves', 'out', 'what', 'for', 'while', "wasn't", 'does', "shouldn't", 'above', 'between', 'be', 'we', 'who', "you're", 'were', 'here', 'hers', "aren't", 'by', 'both', 'about', 'would', 'of', 'could', 'against', "i'd", "weren't", "i'm", 'or', "can't", 'own', 'into', 'whom', 'down', "hadn't", "couldn't", 'your', "doesn't", 'from', "how's", 'her', 'their', "it's", 'there', 'been', 'why', 'few', 'too', 'themselves', 'was', 'until', 'more', 'himself', "where's", "i've", 'with', "didn't", "what's", 'but', 'herself', 'than', "here's", 'he', 'me', "they're", 'myself', 'these', "hasn't", 'below', 'ought', 'theirs', 'my', "wouldn't", "we'd", 'and', 'then', 'is', 'am', 'it', 'an', 'as', 'itself', 'at', 'have', 'in', 'any', 'if', 'again', 'no', 'that', 'when', 'same', 'how', 'other', 'which', 'you', "shan't", 'our', 'after', "let's", 'most', 'such', 'on', "he'll", 'a', 'off', 'i', "she'd", 'yours', "you'll", 'so', "we're", "she's", 'the', "that's", 'having', 'once'])
def tokenize(docs):
pattern = re.compile('[\W_]+', re.UNICODE)
sentences = []
for d in docs:
sentence = d.lower().split(" ")
sentence = [pattern.sub('', w) for w in sentence]
sentences.append( [w for w in sentence if w not in stop_words] )
return sentences
print list(stop_words)
sentences = tokenize(d_train.review)
from gensim.models.word2vec import Word2Vec
model = Word2Vec(sentences, size=300, window=10, min_count=1, sample=1e-3, workers=2)
model.init_sims(replace=True)
model['movie']
def featurize_w2v(model, sentences):
f = zeros((len(sentences), model.vector_size))
for i,s in enumerate(sentences):
for w in s:
try:
vec = model[w]
except KeyError:
continue
f[i,:] = f[i,:] + vec
f[i,:] = f[i,:] / len(s)
return f
features_w2v = featurize_w2v(model, sentences)
model4 = RandomForestClassifier(n_estimators=100, n_jobs=-1)
model4.fit(features_w2v, d_train.sentiment)
test_sentences = tokenize(d_test.review)
test_features_w2v = featurize_w2v(model, test_sentences)
pred4 = model4.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred1, ann=False)
performance(d_test.sentiment, pred2, color="b", ann=False)
performance(d_test.sentiment, pred3, color="r", ann=False)
performance(d_test.sentiment, pred4, color="c")
xlim(0,0.3)
ylim(0.6,1)
examples = [
"This movie is bad",
"This movie is great",
"I was going to say something awesome, but I simply can't because the movie is so bad.",
"I was going to say something awesome or great or good, but I simply can't because the movie is so bad.",
"It might have bad actors, but everything else is good."
]
example_feat4 = featurize_w2v(model, tokenize(examples))
model4.predict(example_feat4)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/hansong0219/Advanced-Deep-learning-Notebooks/blob/master/Cross-Domain/CycleGAN_ResNet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# ResNet 생성자를 이용한 Cycle GAN 모델
ResNet 구조는 이전 층의 전보를 네트워 앞쪽에 한개 이상의 층으로 스킵한다는 부분에서 U-Net 과 비슷하나, U-Net 은 다운샘플링 층을 이에 상응하는 업샘플링 층으로 연결하여 U 모양을 구성하는 대신 ResNet은 Residual block 을 차례대로 쌓아 구성하게된다.
각 블록은 다음의 층으로 출력을 전달하기 전에 입력과 출력을 합하는 스킵 연결층을 가지고 있다.
ResNet 구조는 수백 또는 수천개의 층도 훈련할 수 있는데 앞쪽에 층에 도달하는 그레디언트가 매우 작아져 매우 느리게 훈련되는 vanishing gradient 문제가 없고,Error gradient 가 Residual Block 의 스킵 연결을 통해 네트워크에 그대로 역전파 되기 때문이다. 또, 층을 추가해도 모델의 정확도를 떨어뜨리지 않는데 추가적인 특성이 추출되지 않는다면, 스킵연결로 인해 언제든지 이전 층의 특성이 identify mapping 을 통과하기 때문이다.
본 Notebook 에서는 Residual block 을 사용한 생성자를 구성하여 Image Style Transfer 를 수행할 예정이다.
```
import numpy as np
import os
import sys
from tensorflow.keras.layers import Input, Dropout, concatenate, add, Layer
from tensorflow.keras.layers import Conv2DTranspose, Conv2D
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import LeakyReLU, Activation
from tensorflow.keras.layers import BatchNormalization
from tensorflow_addons.layers import InstanceNormalization
from tensorflow.keras.optimizers import Adam, RMSprop
from tensorflow.keras.utils import plot_model
from tensorflow.keras.losses import BinaryCrossentropy
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
import cv2
import math
import datetime
import imageio
from glob import glob
```
# GPU 할당
```
import tensorflow as tf
physical_devices =tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0],True)
```
# 유틸 함수 정의
### 이미지 및 데이터 재구성 유틸
```
def display_images(imgs, filename, title='', imgs_dir=None, show=False):
#이미지를 nxn 으로 나타냄
rows = imgs.shape[1]
cols = imgs.shape[2]
channels = imgs.shape[3]
side = int(math.sqrt(imgs.shape[0]))
assert int(side * side) == imgs.shape[0]
# 이미지 저장을 위한 폴더를 만듦
if imgs_dir is None:
imgs_dir = 'saved_images'
save_dir = os.path.join(os.getcwd(), imgs_dir)
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
filename = os.path.join(imgs_dir, filename)
# 이미지의 shape 을 지정
if channels==1:
imgs = imgs.reshape((side, side, rows, cols))
else:
imgs = imgs.reshape((side, side, rows, cols, channels))
imgs = np.vstack([np.hstack(i) for i in imgs])
if np.min(imgs)< 0:
imgs = imgs * 0.5 + 0.5
plt.figure(figsize = (8,8))
plt.axis('off')
plt.title(title)
if channels==1:
plt.imshow(imgs, interpolation='none', cmap='gray')
else:
plt.imshow(imgs, interpolation='none')
plt.savefig(filename)
if show:
plt.show()
plt.close('all')
def test_generator(generators, test_data, step, titles, dirs, todisplay=4, show=False):
# generator 모델을 테스트함
# 입력 인수
"""
generators (tuple): 소스와 타겟 생성기
test_data (tuple): 소스와 타겟 데이터
step (int): 진행 단계
titles (tuple): 표시 이미지의 타이틀
dirs (tuple): 이미지 저장 폴더
todisplay (int): 저장이미지의 수 (정사각형 형태로 생성되어야 한다.)
show (bool): 이미지 표시 여부
"""
# test data 로 부터 output 예측
g_source, g_target = generators
test_source_data, test_target_data = test_data
t1, t2, t3, t4 = titles
title_pred_source = t1
title_pred_target = t2
title_reco_source = t3
title_reco_target = t4
dir_pred_source, dir_pred_target = dirs
pred_target_data = g_target.predict(test_source_data)
pred_source_data = g_source.predict(test_target_data)
reco_source_data = g_source.predict(pred_target_data)
reco_target_data = g_target.predict(pred_source_data)
# 정사각형 형태의 하나의 이미지로 나타냄
imgs = pred_target_data[:todisplay]
filename = '%06d.png' % step
step = " Step: {:,}".format(step)
title = title_pred_target + step
display_images(imgs,
filename=filename,
imgs_dir=dir_pred_target,
title=title,
show=show)
imgs = pred_source_data[:todisplay]
title = title_pred_source
display_images(imgs,
filename=filename,
imgs_dir=dir_pred_source,
title=title,
show=show)
imgs = reco_source_data[:todisplay]
title = title_reco_source
filename = "reconstructed_source.png"
display_images(imgs,
filename=filename,
imgs_dir=dir_pred_source,
title=title,
show=show)
imgs = reco_target_data[:todisplay]
title = title_reco_target
filename = "reconstructed_target.png"
display_images(imgs,
filename=filename,
imgs_dir=dir_pred_target,
title=title,
show=show)
def process_data(data, titles, filenames, todisplay=4):
source_data, target_data, test_source_data, test_target_data = data
test_source_filename, test_target_filename = filenames
test_source_title, test_target_title = titles
# 테스트 타겟 이미지 표시
imgs = test_target_data[:todisplay]
display_images(imgs,
filename=test_target_filename,
title=test_target_title)
# 테스트 소스이미지 표시
imgs = test_source_data[:todisplay]
display_images(imgs,
filename=test_source_filename,
title=test_source_title)
# 이미지 표시 정리
target_data = target_data.astype('float32') / 127.5 - 1
test_target_data = test_target_data.astype('float32') / 127.5 - 1
source_data = source_data.astype('float32') / 127.5 - 1
test_source_data = test_source_data.astype('float32') / 127.5 - 1
# 소스, 타겟, 테스트 데이터
data = (source_data, target_data, test_source_data, test_target_data)
rows = source_data.shape[1]
cols = source_data.shape[2]
channels = source_data.shape[3]
source_shape = (rows, cols, channels)
rows = target_data.shape[1]
cols = target_data.shape[2]
channels = target_data.shape[3]
target_shape = (rows, cols, channels)
shapes = (source_shape, target_shape)
return data, shapes
```
### 데이터 불러오기 유틸
```
def imread(path):
return imageio.imread(path,as_gray=False,pilmode='RGB').astype(np.float)
def load_data(dataset_path, is_test=False):
data_type = "train" if not is_test else "test"
path_source = glob(dataset_path + '/%sA/*' %(data_type))
path_target = glob(dataset_path + '/%sB/*' %(data_type))
source_data, target_data = [], []
for source, target in zip(path_source, path_target):
img_source = imread(source)
img_target = imread(target)
img_source = np.array(img_source)
img_target = np.array(img_target)
if is_test and np.random.random()>0.5:
img_source = np.fliplr(img_source)
img_target = np.fliplr(img_target)
source_data.append(img_source)
target_data.append(img_target)
return np.array(source_data), np.array(target_data)
# 데이터 불러오기 및 확인
dataset_path = "D:/data/vangogh2photo"
source_data, target_data = load_data(dataset_path)
plt.figure()
plt.imshow(source_data[0]/255.0)
plt.figure()
plt.imshow(target_data[0]/255.0)
```
# 모델 구성
### ReflectionPadding2D
CycleGAN 논문 및 Source 를 통해 ResNet 생성기의 Layer 구조에서 Reflection Padding을 사용하는 것이 ZerosPadding을 사용하는 Conv2D의 padding=same 조건에 비해 권장 되었다. 따라서 ResNet 생성기에는 ReflectionPadding2D의 Class를 지정하여 사용한다.
```
class ReflectionPadding2D(Layer):
# ReflectionPadding Layer 를 실행할 수 있도록 구성된 class
# 입력인수
"""
padding(tuple): padding을 위한 특정 차원의 크기로 지정된다.
"""
# 출력 : padding 이 실행된 텐서를 출력한다.
def __init__(self, padding=(1, 1), **kwargs):
self.padding = tuple(padding)
super(ReflectionPadding2D, self).__init__(**kwargs)
def get_config(self):
config = super().get_config().copy()
config.update({'padding' : self.padding,})
return config
def call(self, input_tensor, mask=None):
padding_width, padding_height = self.padding
padding_tensor = [
[0, 0],
[padding_height, padding_height],
[padding_width, padding_width],
[0, 0],
]
return tf.pad(input_tensor, padding_tensor, mode="REFLECT")
```
# 모델 Build 함수 정의
### 모델 구성층 구성 함수
ResNet 을 생성자로 하는 Cycle GAN 의 구성요소는 크게 3가지로 분류할 수 있다.
1. 생성자와 판별자의 encoder(downsampling) layer
2. 생성자의 Residual Block Unit
3. 생성자의 decoder(upsampling) layer
```
def encoder_layer(inputs, filters=16, kernel_size=3, strides = 2, activation='relu', instance_norm=True):
#Conv2D -IN - ReLU(LeakyReLU) 의 인코더 층을 구성한다.
kernel_init = keras.initializers.RandomNormal(mean=0.0, stddev=0.02)
x = inputs
x = Conv2D(filters, kernel_size=kernel_size, strides=strides,kernel_initializer=kernel_init, padding = 'same')(x)
if instance_norm:
x = InstanceNormalization(axis = -1, center = False, scale = False)(x)
if activation=='relu':
x = Activation(activation)(x)
else:
x = LeakyReLU(alpha=0.2)(x)
return x
def decoder_layer(inputs, filters=16, kernel_size=3, strides=2, activation='relu', instance_norm = True):
#Conv2DTranspose-IN-LeakyReLU로 구성된 디코더 계층구성, 활성화 함수는 ReLU 로 교체될 수 있음
kernel_init = keras.initializers.RandomNormal(mean=0.0, stddev=0.02)
x = inputs
x = Conv2DTranspose(filters, kernel_size=kernel_size, strides=strides, kernel_initializer = kernel_init, padding='same')(x)
if instance_norm:
x = InstanceNormalization(axis = -1, center = False, scale = False)(x)
if activation=='relu':
x = Activation(activation)(x)
else:
x = LeakyReLU(alpha=0.2)(x)
return x
def residual_block(inputs, filters=64, kernel_size=3, resblock=2):
# shorcut 연결 구현한 Residual Block
kernel_init = keras.initializers.RandomNormal(mean=0.0, stddev=0.02)
shortcut = inputs
for num_unit in range(resblock):
if num_unit == 0:
x = ReflectionPadding2D(padding=(1,1))(inputs)
else:
x = ReflectionPadding2D(padding=(1,1))(x)
x = Conv2D(filters, kernel_size=kernel_size, strides=1, padding='valid', kernel_initializer=kernel_init)(x)
x = InstanceNormalization(axis = -1, center = False, scale = False)(x)
if num_unit != resblock-1:
x = Activation('relu')(x)
return add([shortcut, x])
```
### 생성기 Build 함수 구현
```
# 생성기 Build 함수
def build_generator(input_shape, output_shape=None, filters = 64, num_encoders = 2, num_residual_blocks= 9, num_decoders = 2, name=None):
inputs = Input(shape=input_shape)
channels = int(output_shape[-1])
kernel_init = keras.initializers.RandomNormal(mean=0.0, stddev=0.02)
x = ReflectionPadding2D(padding=(3, 3))(inputs)
x = Conv2D(filters, kernel_size=7, strides=1, padding='valid', kernel_initializer = kernel_init, use_bias=False)(x)
x = InstanceNormalization()(x)
x = Activation('relu')(x)
#DownSampling
for _ in range(num_encoders):
filters *=2
x = encoder_layer(x, filters=filters)
#Residual Block 층 구성
for _ in range(num_residual_blocks):
x = residual_block(x, filters=filters)
#UpSampling
for _ in range(num_decoders):
filters//=2
x = decoder_layer(x, filters=filters)
#Final Block
x = ReflectionPadding2D(padding=(3, 3))(x)
x = Conv2D(channels, (7, 7), strides=1, padding="valid")(x)
x = Activation("tanh")(x)
return Model(inputs, x, name=name)
```
### 판별자 Build 함수 (Patch GAN)
```
def build_discriminator(input_shape, filters=64, kernel_size=4, num_encoder=4, name=None):
kernel_init = keras.initializers.RandomNormal(mean=0.0, stddev=0.02)
inputs = Input(shape=input_shape)
x = encoder_layer(inputs, filters = filters, kernel_size = kernel_size, strides=2, activation='leaky_relu', instance_norm=False)
num_filters=filters
for num_block in range(num_encoder):
num_filters *= 2
if num_block < num_encoder-1:
x = encoder_layer(x, filters=num_filters, kernel_size = kernel_size, strides = 2, activation='leaky_relu')
else:
x = encoder_layer(x, filters=num_filters, kernel_size = kernel_size, strides = 1, activation='leaky_relu')
outputs = Conv2D(1, (4,4), strides=1, padding='same', kernel_initializer = kernel_init)(x)
return Model(inputs, outputs, name=name)
```
### Cycle GAN Build 함수 지정
```
def build_cyclegan(shape, source_name='source', target_name='target', kernel_size=3, patchgan=False, identify=False):
#Cycle GAN 의 구성
"""
1) 타깃과 소스의 판별기
2) 타깃과 소스의 생성기
3) 적대적 네트워크 구성
"""
#입력 인수
"""
shape(tuple): 소스와 타깃 형상
source_name (string): 판별기/생성기 모델 이름 뒤에 붙는 소스이름 문자열
target_name (string): 판별기/생성기 모델 이름 뒤에 붙는 타깃이름 문자열
target_size (int): 인코더/디코더 혹은 판별기/생성기 모델에 사용될 커널 크기
patchgan(bool): 판별기에 patchgan 사용 여부
identify(bool): 동질성 사용 여부
"""
#출력 결과:
#list : 2개의 생성기, 2개의 판별기, 1개의 적대적 모델
source_shape, target_shape = shapes
lr = 2e-4
decay = 6e-8
gt_name = "gen_" + target_name
gs_name = "gen_" + source_name
dt_name = "dis_" + target_name
ds_name = "dis_" + source_name
#타깃과 소스 생성기 구성
g_target = build_generator(source_shape, target_shape, name=gt_name)
g_source = build_generator(target_shape, source_shape, name=gs_name)
print('-----Target Generator-----')
g_target.summary()
print('-----Source Generator-----')
g_source.summary()
#타깃과 소스 판별기 구성
d_target = build_discriminator(target_shape, name=dt_name)
d_source = build_discriminator(source_shape, name=ds_name)
print('-----Targent Discriminator-----')
d_target.summary()
print('-----Source Discriminator-----')
d_source.summary()
optimizer = RMSprop(lr=lr, decay=decay)
d_target.compile(loss='mse',optimizer=optimizer,metrics=['accuracy'])
d_source.compile(loss='mse',optimizer=optimizer,metrics=['accuracy'])
#적대적 모델에서 판별기 가중치 고정
d_target.trainable=False
d_source.trainable=False
#적대적 모델의 계산 그래프 구성
#전방순환 네트워크와 타깃 판별기
source_input = Input(shape=source_shape)
fake_target = g_target(source_input)
preal_target = d_target(fake_target)
reco_source = g_source(fake_target)
#후방순환 네트워크와 타깃 판별기
target_input = Input(shape=target_shape)
fake_source = g_source(target_input)
preal_source = d_source(fake_source)
reco_target = g_target(fake_source)
#동질성 손실 사용 시, 두개의 손실항과 출력을 추가한다.
if identify:
iden_source = g_source(source_input)
iden_target = g_target(target_input)
loss = ['mse', 'mse', 'mae', 'mae', 'mae', 'mae']
loss_weights = [1., 1., 10., 10., 0.5, 0.5]
inputs = [source_input, target_input]
outputs = [preal_source, preal_target, reco_source, reco_target, iden_source, iden_target]
else:
loss = ['mse', 'mse', 'mae', 'mae']
loss_weights = [1., 1., 10., 10.]
inputs = [source_input, target_input]
outputs = [preal_source, preal_target, reco_source, reco_target]
#적대적 모델 구성
adv = Model(inputs, outputs, name='adversarial')
optimizer = RMSprop(lr=lr*0.5, decay=decay*0.5)
adv.compile(loss=loss, loss_weights=loss_weights, optimizer=optimizer)
print('-----Adversarial Network-----')
adv.summary()
return g_source, g_target, d_source, d_target, adv
def train_cyclegan(models, data, params, test_params, test_generator, identify=False):
#Cycle GAN 훈련
"""
1) 타깃 판별기 훈련
2) 소스 판별기 훈련
3) 적대적 네트워크의 전방/ 후방 순환 훈련
"""
# 입력 인수:
"""
models (list): 소스/타깃에 대한 판별기/생성기, 적대적 모델
data (tuple): 소스와 타깃 훈련데이터
params (tuple): 네트워크 매개변수
test_params (tuple): 테스트 매개변수
test_generator (function): 예측 타깃/소스 이미지생성에 사용됨.
"""
#모델
g_source, g_target, d_source, d_target, adv = models
#네트워크 매개변수 지정
batch_size, train_steps, patch, model_name = params
#훈련 데이터 세트
source_data, target_data, test_source_data, test_target_data = data
titles, dirs = test_params
#생성기 이미지는 2000단계마다 저장됨
save_interval = 2000
target_size = target_data.shape[0]
source_size = source_data.shape[0]
#patchgan 사용 여부
if patch > 1:
d_patch = (patch, patch, 1)
valid = np.ones((batch_size,) + d_patch)
fake = np.zeros((batch_size,) + d_patch)
else:
valid = np.ones([batch_size, 1])
fake = np.zeros([batch_size, 1])
valid_fake = np.concatenate((valid,fake))
start_time = datetime.datetime.now()
for step in range(train_steps):
#실제 타깃 데이터 배치 샘플링
rand_indices = np.random.randint(0, target_size, size=batch_size)
real_target = target_data[rand_indices]
#실제 소스 데이터 배치 샘플링
rand_indices = np.random.randint(0, source_size, size=batch_size)
real_source = source_data[rand_indices]
#실제 소스 데이터에서 가짜 타깃 데이터 배치를 생성
fake_target = g_target.predict(real_source)
#실제 타겟 데이터와 가짜 타겟데이터를 하나의 배치로 결합
x = np.concatenate((real_target, fake_target))
#타겟 판별자를 훈련시킴
metrics = d_target.train_on_batch(x, valid_fake)
log = "%d: [d_target loss: %f]" %(step, metrics[0])
#실제 타깃 데이터에서 가짜 소스 데이터 배치 생성
fake_source = g_source.predict(real_target)
x = np.concatenate((real_source, fake_source))
#가짜/실제 데이터를 사용해 소스 판별기 훈련
metrics = d_source.train_on_batch(x, valid_fake)
log = "%s [d_source loss: %f]" %(log, metrics[0])
#전방/후방 순환을 사용해 적대적 네트워크 훈련
#생성된 가짜 소스/타깃 데이터는 판별기를 속이려고 시도함
if identify:
x = [real_source, real_target]
y = [valid, valid, real_source, real_target, real_source, real_target]
else:
x = [real_source, real_target]
y = [valid, valid, real_source, real_target]
metrics = adv.train_on_batch(x, y)
elapsed_time = datetime.datetime.now()-start_time
fmt = "%s [adv loss: %f] [time: %s]"
log = fmt %(log, metrics[0], elapsed_time)
print(log)
if (step+1) % save_interval == 0:
if (step+1) == train_steps:
show = True
else:
show = False
test_generator((g_source, g_target), (test_source_data, test_target_data), step = step+1, titles=titles, dirs=dirs, show=show)
g_source.save(model_name + "-g_source.h5")
g_target.save(model_name + "-g_target.h5")
```
# 데이터 전처리
```
test_source_data, test_target_data = load_data(dataset_path, is_test=True)
filenames = ('vangogh_test_source.png', 'picture_test_target.png')
titles = ('Van Gogh test source images', 'picture test target images')
data = (source_data, target_data, test_source_data, test_target_data)
# 이미지 저장 및 shape 재지정
data, shapes = process_data(data, titles, filenames)
#model_fine tuning 변수 지정
model_name = 'cyclegan_vangogh'
batch_size = 1
train_steps = 100000
patchgan = True
kernel_size = 3
postfix = ('%dp' % kernel_size) if patchgan else ('%d' % kernel_size)
titles = ('vangogh2pic predicted source images.',
'vangogh2pic predicted target images.',
'vangogh2pic reconstructed source images.',
'vangogh2pic reconstructed target images.')
dirs = ('vangogh2pic_source-%s' % postfix, 'vangogh2pic_target-%s' % postfix)
models = build_cyclegan(shapes, "vangogh-%s" % postfix, "picture-%s" % postfix, kernel_size=kernel_size, patchgan=patchgan)
#판별기의 입력을 2^n 만큼 척도를 줄임 -> patch 크기를 2^n 으로 나눔(즉 strides=2를 n회 사용함)
patch = int(source_data.shape[1] / 2**4) if patchgan else 1
params = (batch_size, train_steps, patch, model_name)
test_params = (titles, dirs)
#cyclegan 훈련
train_cyclegan(models, data, params, test_params, test_generator)
```
| github_jupyter |
[View in Colaboratory](https://colab.research.google.com/github/douglaswchung/california-housing-value/blob/master/neural_nets_regression.ipynb)
#### Copyright 2017 Google LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Intro to Neural Networks
**Learning Objectives:**
* Define a neural network (NN) and its hidden layers using the TensorFlow `DNNRegressor` class
* Train a neural network to learn nonlinearities in a dataset and achieve better performance than a linear regression model
In the previous exercises, we used synthetic features to help our model incorporate nonlinearities.
One important set of nonlinearities was around latitude and longitude, but there may be others.
We'll also switch back, for now, to a standard regression task, rather than the logistic regression task from the previous exercise. That is, we'll be predicting `median_house_value` directly.
## Setup
First, let's load and prepare the data.
```
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
from scipy.stats import zscore
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy()
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
"""processed_features = processed_features.apply(zscore)"""
"""processed_features["latitude"] = california_housing_dataframe["latitude"]
processed_features["longitude"] = california_housing_dataframe["longitude"]"""
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Scale the target to be in units of thousands of dollars.
output_targets["median_house_value"] = (
california_housing_dataframe["median_house_value"] / 1000.0)
return output_targets
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(california_housing_dataframe.head(13600))
training_targets = preprocess_targets(california_housing_dataframe.head(13600))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(california_housing_dataframe.tail(3400))
validation_targets = preprocess_targets(california_housing_dataframe.tail(3400))
# Double-check that we've done the right thing.
print "Training examples summary:"
display.display(training_examples.describe())
print "Validation examples summary:"
display.display(validation_examples.describe())
print "Training targets summary:"
display.display(training_targets.describe())
print "Validation targets summary:"
display.display(validation_targets.describe())
```
## Building a Neural Network
The NN is defined by the [DNNRegressor](https://www.tensorflow.org/api_docs/python/tf/contrib/learn/DNNRegressor) class.
Use **`hidden_units`** to define the structure of the NN. The `hidden_units` argument provides a list of ints, where each int corresponds to a hidden layer and indicates the number of nodes in it. For example, consider the following assignment:
`hidden_units=[3,10]`
The preceding assignment specifies a neural net with two hidden layers:
* The first hidden layer contains 3 nodes.
* The second hidden layer contains 10 nodes.
If we wanted to add more layers, we'd add more ints to the list. For example, `hidden_units=[10,20,30,40]` would create four layers with ten, twenty, thirty, and forty units, respectively.
By default, all hidden layers will use ReLu activation and will be fully connected.
```
def construct_feature_columns(input_features):
"""Construct the TensorFlow Feature Columns.
Args:
input_features: The names of the numerical input features to use.
Returns:
A set of feature columns
"""
return set([tf.feature_column.numeric_column(my_feature)
for my_feature in input_features])
def get_quantile_based_boundaries(feature_values, num_buckets):
boundaries = np.arange(1.0, num_buckets) / num_buckets
quantiles = feature_values.quantile(boundaries)
return [quantiles[q] for q in quantiles.keys()]
def construct_feature_columns():
"""Construct the TensorFlow Feature Columns.
Returns:
A set of feature columns
"""
households = tf.feature_column.numeric_column("households")
longitude = tf.feature_column.numeric_column("longitude")
latitude = tf.feature_column.numeric_column("latitude")
housing_median_age = tf.feature_column.numeric_column("housing_median_age")
median_income = tf.feature_column.numeric_column("median_income")
rooms_per_person = tf.feature_column.numeric_column("rooms_per_person")
# Divide households into 7 buckets.
bucketized_households = tf.feature_column.bucketized_column(
households, boundaries=get_quantile_based_boundaries(
training_examples["households"], 6))
bucketized_households=tf.feature_column.embedding_column(
categorical_column = bucketized_households,
dimension = math.ceil(6**0.25))
# Divide longitude into 10 buckets.
bucketized_longitude = tf.feature_column.bucketized_column(
longitude, boundaries=get_quantile_based_boundaries(
training_examples["longitude"], 10))
# Divide latitude into 10 buckets.
bucketized_latitude = tf.feature_column.bucketized_column(
latitude, boundaries=get_quantile_based_boundaries(
training_examples["latitude"], 10))
# Divide housing_median_age into 7 buckets.
bucketized_housing_median_age = tf.feature_column.bucketized_column(
housing_median_age, boundaries=get_quantile_based_boundaries(
training_examples["housing_median_age"], 6))
bucketized_housing_median_age=tf.feature_column.embedding_column(
categorical_column = bucketized_housing_median_age,
dimension = math.ceil(6**0.25))
# Divide median_income into 7 buckets.
bucketized_median_income = tf.feature_column.bucketized_column(
median_income, boundaries=get_quantile_based_boundaries(
training_examples["median_income"], 8))
bucketized_median_income=tf.feature_column.embedding_column(
categorical_column = bucketized_median_income,
dimension = math.ceil(8**0.25))
# Divide rooms_per_person into 7 buckets.
bucketized_rooms_per_person = tf.feature_column.bucketized_column(
rooms_per_person, boundaries=get_quantile_based_boundaries(
training_examples["rooms_per_person"], 6))
bucketized_rooms_per_person=tf.feature_column.embedding_column(
categorical_column = bucketized_rooms_per_person,
dimension = math.ceil(6**0.25))
# YOUR CODE HERE: Make a feature column for the long_x_lat feature cross
long_x_lat = tf.feature_column.crossed_column([bucketized_longitude, bucketized_latitude], hash_bucket_size = 1000)
long_x_lat=tf.feature_column.embedding_column(
categorical_column = long_x_lat,
dimension = math.ceil(100**0.25))
feature_columns = set([
bucketized_median_income,
bucketized_rooms_per_person,
long_x_lat])
return feature_columns
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a neural net regression model.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
def train_nn_regression_model(
learning_rate,
steps,
batch_size,
hidden_units,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a neural network regression model.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
hidden_units: A `list` of int values, specifying the number of neurons in each layer.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `DNNRegressor` object trained on the training data.
"""
periods = 100
steps_per_period = steps / periods
# Create a DNNRegressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
dnn_regressor = tf.estimator.DNNRegressor(
feature_columns=construct_feature_columns(),
hidden_units=hidden_units,
optimizer=my_optimizer,
)
# Create input functions.
training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
num_epochs=1,
shuffle=False)
predict_validation_input_fn = lambda: my_input_fn(validation_examples,
validation_targets["median_house_value"],
num_epochs=1,
shuffle=False)
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print "Training model..."
print "RMSE (on training data):"
training_rmse = []
validation_rmse = []
for period in range (0, periods):
# Train the model, starting from the prior state.
dnn_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period
)
# Take a break and compute predictions.
training_predictions = dnn_regressor.predict(input_fn=predict_training_input_fn)
training_predictions = np.array([item['predictions'][0] for item in training_predictions])
validation_predictions = dnn_regressor.predict(input_fn=predict_validation_input_fn)
validation_predictions = np.array([item['predictions'][0] for item in validation_predictions])
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, training_targets))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, validation_targets))
# Occasionally print the current loss.
print " period %02d : %0.2f" % (period, training_root_mean_squared_error)
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print "Model training finished."
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
print "Final RMSE (on training data): %0.2f" % training_root_mean_squared_error
print "Final RMSE (on validation data): %0.2f" % validation_root_mean_squared_error
return dnn_regressor
```
## Task 1: Train a NN Model
**Adjust hyperparameters, aiming to drop RMSE below 110.**
Run the following block to train a NN model.
Recall that in the linear regression exercise with many features, an RMSE of 110 or so was pretty good. We'll aim to beat that.
Your task here is to modify various learning settings to improve accuracy on validation data.
Overfitting is a real potential hazard for NNs. You can look at the gap between loss on training data and loss on validation data to help judge if your model is starting to overfit. If the gap starts to grow, that is usually a sure sign of overfitting.
Because of the number of different possible settings, it's strongly recommended that you take notes on each trial to help guide your development process.
Also, when you get a good setting, try running it multiple times and see how repeatable your result is. NN weights are typically initialized to small random values, so you should see differences from run to run.
```
dnn_regressor = train_nn_regression_model(
learning_rate=0.005,
steps=2000,
batch_size=136,
hidden_units=[13,7],
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
```
### Solution
Click below to see a possible solution
**NOTE:** This selection of parameters is somewhat arbitrary. Here we've tried combinations that are increasingly complex, combined with training for longer, until the error falls below our objective. This may not be the best combination; others may attain an even lower RMSE. If your aim is to find the model that can attain the best error, then you'll want to use a more rigorous process, like a parameter search.
```
dnn_regressor = train_nn_regression_model(
learning_rate=0.001,
steps=2000,
batch_size=100,
hidden_units=[10, 10],
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
```
## Task 2: Evaluate on Test Data
**Confirm that your validation performance results hold up on test data.**
Once you have a model you're happy with, evaluate it on test data to compare that to validation performance.
Reminder, the test data set is located [here](https://storage.googleapis.com/mledu-datasets/california_housing_test.csv).
```
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
# YOUR CODE HERE
test_examples = preprocess_features(california_housing_test_data)
test_targets = preprocess_targets(california_housing_test_data)
predict_test_input_fn = lambda: my_input_fn(test_examples,
test_targets["median_house_value"],
num_epochs=1,
shuffle=False)
test_predictions = dnn_regressor.predict(input_fn=predict_test_input_fn)
test_predictions = np.array([item['predictions'][0] for item in test_predictions])
test_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(test_predictions, test_targets))
print "Final RMSE (on test data): %0.2f" % test_root_mean_squared_error
```
### Solution
Click below to see a possible solution.
Similar to what the code at the top does, we just need to load the appropriate data file, preprocess it and call predict and mean_squared_error.
Note that we don't have to randomize the test data, since we will use all records.
```
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
test_examples = preprocess_features(california_housing_test_data)
test_targets = preprocess_targets(california_housing_test_data)
predict_testing_input_fn = lambda: my_input_fn(test_examples,
test_targets["median_house_value"],
num_epochs=1,
shuffle=False)
test_predictions = dnn_regressor.predict(input_fn=predict_testing_input_fn)
test_predictions = np.array([item['predictions'][0] for item in test_predictions])
root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(test_predictions, test_targets))
print "Final RMSE (on test data): %0.2f" % root_mean_squared_error
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 텐서 소개
<table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/guide/tensor"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/guide/tensor.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행하기</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/guide/tensor.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서소스 보기</a></td>
<td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/guide/tensor.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드하기</a></td>
</table>
```
import tensorflow as tf
import numpy as np
```
텐서는 일관된 유형(`dtype`이라고 불림)을 가진 다차원 배열입니다. 지원되는 모든 `dtypes`은 `tf.dtypes.DType`에서 볼 수 있습니다.
[NumPy](https://numpy.org/devdocs/user/quickstart.html)에 익숙하다면, 텐서는 일종의 `np.arrays`와 같습니다.
모든 텐서는 Python 숫자 및 문자열과 같이 변경할 수 없습니다. 텐서의 내용을 업데이트할 수 없으며 새로운 텐서를 만들 수만 있습니다.
## 기초
기본 텐서를 만들어 봅시다.
다음은 "스칼라" 또는 "순위-0" 텐서입니다. 스칼라는 단일 값을 포함하며 "축"은 없습니다.
```
# This will be an int32 tensor by default; see "dtypes" below.
rank_0_tensor = tf.constant(4)
print(rank_0_tensor)
```
"벡터" 또는 "순위-1" 텐서는 값의 목록과 같습니다. 벡터는 1축입니다.
```
# Let's make this a float tensor.
rank_1_tensor = tf.constant([2.0, 3.0, 4.0])
print(rank_1_tensor)
```
"행렬" 또는 "rank-2" 텐서에는 2축이 있습니다.
```
# If we want to be specific, we can set the dtype (see below) at creation time
rank_2_tensor = tf.constant([[1, 2],
[3, 4],
[5, 6]], dtype=tf.float16)
print(rank_2_tensor)
```
<table>
<tr>
<th>스칼라, 형상: <code>[]</code>
</th>
<th>벡터, 형상: <code>[3]</code>
</th>
<th>행렬, 형상: <code>[3, 2]</code>
</th>
</tr>
<tr>
<td><img alt="A scalar, the number 4" src="images/tensor/scalar.png"></td>
<td><img alt="The line with 3 sections, each one containing a number." src="images/tensor/vector.png"></td>
<td><img alt="A 3x2 grid, with each cell containing a number." src="images/tensor/matrix.png"></td>
</tr>
</table>
텐서에는 더 많은 축이 있을 수 있습니다. 여기에는 3축 텐서가 사용됩니다.
```
# There can be an arbitrary number of
# axes (sometimes called "dimensions")
rank_3_tensor = tf.constant([
[[0, 1, 2, 3, 4],
[5, 6, 7, 8, 9]],
[[10, 11, 12, 13, 14],
[15, 16, 17, 18, 19]],
[[20, 21, 22, 23, 24],
[25, 26, 27, 28, 29]],])
print(rank_3_tensor)
```
2축 이상의 텐서를 시각화하는 방법에는 여러 가지가 있습니다.
<table>
<tr>
<th colspan="3">3축 텐서, 형상: <code>[3, 2, 5]</code>
</th>
</tr>
<tr>
</tr>
<tr>
<td><img src="images/tensor/3-axis_numpy.png"></td>
<td><img src="images/tensor/3-axis_front.png"></td>
<td><img src="images/tensor/3-axis_block.png"></td>
</tr>
</table>
`np.array` 또는 `tensor.numpy` 메서드를 사용하여 텐서를 NumPy 배열로 변환할 수 있습니다.
```
np.array(rank_2_tensor)
rank_2_tensor.numpy()
```
텐서에는 종종 float와 int가 포함되지만, 다음과 같은 다른 유형도 있습니다.
- 복소수
- 문자열
기본 `tf.Tensor` 클래스에서는 텐서가 "직사각형"이어야 합니다. 즉, 각 축을 따라 모든 요소의 크기가 같습니다. 그러나 다양한 형상을 처리할 수 있는 특수 유형의 텐서가 있습니다.
- 비정형(아래의 [RaggedTensor](#ragged_tensors) 참조)
- 희소(아래의 [SparseTensor](#sparse_tensors) 참조)
덧셈, 요소별 곱셈 및 행렬 곱셈을 포함하여 텐서에 대한 기본 수학을 수행 할 수 있습니다.
```
a = tf.constant([[1, 2],
[3, 4]])
b = tf.constant([[1, 1],
[1, 1]]) # Could have also said `tf.ones([2,2])`
print(tf.add(a, b), "\n")
print(tf.multiply(a, b), "\n")
print(tf.matmul(a, b), "\n")
print(a + b, "\n") # element-wise addition
print(a * b, "\n") # element-wise multiplication
print(a @ b, "\n") # matrix multiplication
```
텐서는 모든 종류의 연산(ops)에 사용됩니다.
```
c = tf.constant([[4.0, 5.0], [10.0, 1.0]])
# Find the largest value
print(tf.reduce_max(c))
# Find the index of the largest value
print(tf.argmax(c))
# Compute the softmax
print(tf.nn.softmax(c))
```
## 형상 정보
텐서는 형상이 있습니다. 사용되는 일부 용어는 다음과 같습니다.
- **형상**: 텐서의 각 차원의 길이(요소의 수)
- **순위**: 텐서 차원의 수입니다. 스칼라는 순위가 0이고 벡터의 순위는 1이며 행렬의 순위는 2입니다.
- **축** 또는 **차원**: 텐서의 특정 차원
- **크기**: 텐서의 총 항목 수, 곱 형상 벡터
참고: "2차원 텐서"에 대한 참조가 있을 수 있지만, 순위-2 텐서는 일반적으로 2D 공간을 설명하지 않습니다.
텐서 및 `tf.TensorShape` 객체에는 다음에 액세스하기 위한 편리한 속성이 있습니다.
```
rank_4_tensor = tf.zeros([3, 2, 4, 5])
```
<table>
<tr>
<th colspan="2">순위-4 텐서, 형상: <code>[3, 2, 4, 5]</code>
</th>
</tr>
<tr>
<td> <img alt="A tensor shape is like a vector." src="images/tensor/shape.png">
</td>
<td> <img alt="A 4-axis tensor" src="images/tensor/4-axis_block.png">
</td>
</tr>
</table>
```
print("Type of every element:", rank_4_tensor.dtype)
print("Number of dimensions:", rank_4_tensor.ndim)
print("Shape of tensor:", rank_4_tensor.shape)
print("Elements along axis 0 of tensor:", rank_4_tensor.shape[0])
print("Elements along the last axis of tensor:", rank_4_tensor.shape[-1])
print("Total number of elements (3*2*4*5): ", tf.size(rank_4_tensor).numpy())
```
축은 종종 인덱스로 참조하지만, 항상 각 축의 의미를 추적해야 합니다. 축이 전역에서 로컬로 정렬되는 경우가 종종 있습니다. 배치 축이 먼저 오고 그 다음에 공간 차원과 각 위치의 특성이 마지막에 옵니다. 이러한 방식으로 특성 벡터는 연속적인 메모리 영역입니다.
<table>
<tr>
<th>일반적인 축 순서</th>
</tr>
<tr>
<td> <img alt="Keep track of what each axis is. A 4-axis tensor might be: Batch, Width, Height, Freatures" src="images/tensor/shape2.png">
</td>
</tr>
</table>
## 인덱싱
### 단일 축 인덱싱
TensorFlow는 [파이썬의 목록 또는 문자열 인덱싱](https://docs.python.org/3/tutorial/introduction.html#strings)과 마찬가지로 표준 파이썬 인덱싱 규칙과 numpy 인덱싱의 기본 규칙을 따릅니다.
- 인덱스는 `0`에서 시작합니다.
- 음수 인덱스는 끝에서부터 거꾸로 계산합니다.
- 콜론, `:`은 슬라이스 `start:stop:step`에 사용됩니다.
```
rank_1_tensor = tf.constant([0, 1, 1, 2, 3, 5, 8, 13, 21, 34])
print(rank_1_tensor.numpy())
```
스칼라를 사용하여 인덱싱하면 차원이 제거됩니다.
```
print("First:", rank_1_tensor[0].numpy())
print("Second:", rank_1_tensor[1].numpy())
print("Last:", rank_1_tensor[-1].numpy())
```
`:` 조각으로 인덱싱하면 차원이 유지됩니다.
```
print("Everything:", rank_1_tensor[:].numpy())
print("Before 4:", rank_1_tensor[:4].numpy())
print("From 4 to the end:", rank_1_tensor[4:].numpy())
print("From 2, before 7:", rank_1_tensor[2:7].numpy())
print("Every other item:", rank_1_tensor[::2].numpy())
print("Reversed:", rank_1_tensor[::-1].numpy())
```
### 다축 인덱싱
더 높은 순위의 텐서는 여러 인덱스를 전달하여 인덱싱됩니다.
단일 축의 경우에서와 정확히 같은 단일 축 규칙이 각 축에 독립적으로 적용됩니다.
```
print(rank_2_tensor.numpy())
```
각 인덱스에 정수를 전달하면 결과는 스칼라입니다.
```
# Pull out a single value from a 2-rank tensor
print(rank_2_tensor[1, 1].numpy())
```
정수와 슬라이스를 조합하여 인덱싱할 수 있습니다.
```
# Get row and column tensors
print("Second row:", rank_2_tensor[1, :].numpy())
print("Second column:", rank_2_tensor[:, 1].numpy())
print("Last row:", rank_2_tensor[-1, :].numpy())
print("First item in last column:", rank_2_tensor[0, -1].numpy())
print("Skip the first row:")
print(rank_2_tensor[1:, :].numpy(), "\n")
```
다음은 3축 텐서의 예입니다.
```
print(rank_3_tensor[:, :, 4])
```
<table>
<tr>
<th colspan="2">배치에서 각 예의 모든 위치에서 마지막 특성 선택하기</th>
</tr>
<tr>
<td><img alt="A 3x2x5 tensor with all the values at the index-4 of the last axis selected." src="images/tensor/index1.png"></td>
<td><img alt="The selected values packed into a 2-axis tensor." src="images/tensor/index2.png"></td>
</tr>
</table>
## 형상 조작하기
텐서의 형상을 바꾸는 것은 매우 유용합니다.
```
# Shape returns a `TensorShape` object that shows the size on each dimension
var_x = tf.Variable(tf.constant([[1], [2], [3]]))
print(var_x.shape)
# You can convert this object into a Python list, too
print(var_x.shape.as_list())
```
텐서를 새로운 형상으로 바꿀 수 있습니다. 기본 데이터를 복제할 필요가 없으므로 재구성이 빠르고 저렴합니다.
```
# We can reshape a tensor to a new shape.
# Note that we're passing in a list
reshaped = tf.reshape(var_x, [1, 3])
print(var_x.shape)
print(reshaped.shape)
```
데이터의 레이아웃은 메모리에서 유지되고 요청된 형상이 같은 데이터를 가리키는 새 텐서가 작성됩니다. TensorFlow는 C 스타일 "행 중심" 메모리 순서를 사용합니다. 여기에서 가장 오른쪽에 있는 인덱스를 증가시키면 메모리의 단일 단계에 해당합니다.
```
print(rank_3_tensor)
```
텐서를 평평하게 하면 어떤 순서로 메모리에 배치되어 있는지 확인할 수 있습니다.
```
# A `-1` passed in the `shape` argument says "Whatever fits".
print(tf.reshape(rank_3_tensor, [-1]))
```
일반적으로, `tf.reshape`의 합리적인 용도는 인접한 축을 결합하거나 분할하는 것(또는 `1`을 추가/제거)입니다.
이 3x2x5 텐서의 경우, 슬라이스가 혼합되지 않으므로 (3x2)x5 또는 3x (2x5)로 재구성하는 것이 합리적입니다.
```
print(tf.reshape(rank_3_tensor, [3*2, 5]), "\n")
print(tf.reshape(rank_3_tensor, [3, -1]))
```
<table>
<th colspan="3">몇 가지 좋은 재구성</th>
<tr>
<td><img alt="A 3x2x5 tensor" src="images/tensor/reshape-before.png"></td>
<td><img alt="The same data reshaped to (3x2)x5" src="images/tensor/reshape-good1.png"></td>
<td><img alt="The same data reshaped to 3x(2x5)" src="images/tensor/reshape-good2.png"></td>
</tr>
</table>
형상을 변경하면 같은 총 요소 수를 가진 새로운 형상에 대해 "작동"하지만, 축의 순서를 고려하지 않으면 별로 쓸모가 없습니다.
`tf.reshape`에서 축 교환이 작동하지 않으면, `tf.transpose`를 수행해야 합니다.
```
# Bad examples: don't do this
# You can't reorder axes with reshape.
print(tf.reshape(rank_3_tensor, [2, 3, 5]), "\n")
# This is a mess
print(tf.reshape(rank_3_tensor, [5, 6]), "\n")
# This doesn't work at all
try:
tf.reshape(rank_3_tensor, [7, -1])
except Exception as e:
print(f"{type(e).__name__}: {e}")
```
<table>
<th colspan="3">몇 가지 잘못된 재구성</th>
<tr>
<td><img alt="You can't reorder axes, use tf.transpose for that" src="images/tensor/reshape-bad.png"></td>
<td><img alt="Anything that mixes the slices of data together is probably wrong." src="images/tensor/reshape-bad4.png"></td>
<td><img alt="The new shape must fit exactly." src="images/tensor/reshape-bad2.png"></td>
</tr>
</table>
완전히 지정되지 않은 형상에서 실행할 수 있습니다. 형상에 `None`(차원의 길이를 알 수 없음)이 포함되거나 형상이`None`(텐서의 순위를 알 수 없음)입니다.
[tf.RaggedTensor](#ragged_tensors)를 제외하고, TensorFlow의 상징적인 그래프 빌딩 API의 컨텍스트에서만 발생합니다.
- [tf.function](function.ipynb)
- [keras 함수형 API](keras/functional.ipynb)
## `DTypes`에 대한 추가 정보
`tf.Tensor`의 데이터 유형을 검사하려면, `Tensor.dtype` 속성을 사용합니다.
Python 객체에서 `tf.Tensor`를 만들 때 선택적으로 데이터 유형을 지정할 수 있습니다.
그렇지 않으면, TensorFlow는 데이터를 나타낼 수 있는 데이터 유형을 선택합니다. TensorFlow는 Python 정수를 `tf.int32`로, Python 부동 소수점 숫자를 `tf.float32`로 변환합니다. 그렇지 않으면, TensorFlow는 NumPy가 배열로 변환할 때 사용하는 것과 같은 규칙을 사용합니다.
유형별로 캐스팅할 수 있습니다.
```
the_f64_tensor = tf.constant([2.2, 3.3, 4.4], dtype=tf.float64)
the_f16_tensor = tf.cast(the_f64_tensor, dtype=tf.float16)
# Now, let's cast to an uint8 and lose the decimal precision
the_u8_tensor = tf.cast(the_f16_tensor, dtype=tf.uint8)
print(the_u8_tensor)
```
## 브로드캐스팅
브로드캐스팅은 [NumPy의 해당 특성](https://numpy.org/doc/stable/user/basics.html)에서 빌린 개념입니다. 요컨대, 특정 조건에서 작은 텐서는 결합된 연산을 실행할 때 더 큰 텐서에 맞게 자동으로 "확장(streched)"됩니다.
가장 간단하고 가장 일반적인 경우는 스칼라에 텐서를 곱하거나 추가하려고 할 때입니다. 이 경우, 스칼라는 다른 인수와 같은 형상으로 브로드캐스트됩니다.
```
x = tf.constant([1, 2, 3])
y = tf.constant(2)
z = tf.constant([2, 2, 2])
# All of these are the same computation
print(tf.multiply(x, 2))
print(x * y)
print(x * z)
```
마찬가지로, 크기가 1인 차원은 다른 인수와 일치하도록 확장될 수 있습니다. 두 인수 모두 같은 계산으로 확장될 수 있습니다.
이 경우, 3x1 행렬에 요소별로 1x4 행렬을 곱하여 3x4 행렬을 만듭니다. 선행 1이 선택 사항인 점에 유의하세요. y의 형상은 `[4]`입니다.
```
# These are the same computations
x = tf.reshape(x,[3,1])
y = tf.range(1, 5)
print(x, "\n")
print(y, "\n")
print(tf.multiply(x, y))
```
<table>
<tr>
<th>추가 시 브로드캐스팅: <code>[1, 4]</code>와 <code>[3, 1]</code>의 곱하기는 <code>[3,4]</code>입니다.</th>
</tr>
<tr>
<td><img alt="Adding a 3x1 matrix to a 4x1 matrix results in a 3x4 matrix" src="images/tensor/broadcasting.png"></td>
</tr>
</table>
브로드캐스팅이 없는 같은 연산이 여기 있습니다.
```
x_stretch = tf.constant([[1, 1, 1, 1],
[2, 2, 2, 2],
[3, 3, 3, 3]])
y_stretch = tf.constant([[1, 2, 3, 4],
[1, 2, 3, 4],
[1, 2, 3, 4]])
print(x_stretch * y_stretch) # Again, operator overloading
```
대부분의 경우 브로드캐스팅은 브로드캐스트 연산으로 메모리에서 확장된 텐서를 구체화하지 않으므로 시간과 공간 효율적입니다.
`tf.broadcast_to`를 사용하여 브로드캐스팅이 어떤 모습인지 알 수 있습니다.
```
print(tf.broadcast_to(tf.constant([1, 2, 3]), [3, 3]))
```
예를 들어, `broadcast_to`는 수학적인 op와 달리 메모리를 절약하기 위해 특별한 연산을 수행하지 않습니다. 여기에서 텐서를 구체화합니다.
훨씬 더 복잡해질 수 있습니다. Jake VanderPlas의 저서 *Python Data Science Handbook*의 [해당 섹션](https://jakevdp.github.io/PythonDataScienceHandbook/02.05-computation-on-arrays-broadcasting.html)에서는 더 많은 브로드캐스팅 트릭을 보여줍니다(NumPy에서).
## tf.convert_to_tensor
`tf.matmul` 및 `tf.reshape`와 같은 대부분의 ops는 클래스 `tf.Tensor`의 인수를 사용합니다. 그러나 위의 경우, 텐서 형상의 Python 객체를 자주 전달합니다.
전부는 아니지만 대부분의 ops는 텐서가 아닌 인수에 대해 `convert_to_tensor`를 호출합니다. 변환 레지스트리가 있어 NumPy의 `ndarray`, `TensorShape` , Python 목록 및 `tf.Variable`과 같은 대부분의 객체 클래스는 모두 자동으로 변환됩니다.
자세한 내용은 `tf.register_tensor_conversion_function`을 참조하세요. 자신만의 유형이 있으면 자동으로 텐서로 변환할 수 있습니다.
## 비정형 텐서
어떤 축을 따라 다양한 수의 요소를 가진 텐서를 "비정형(ragged)"이라고 합니다. 비정형 데이터에는 `tf.ragged.RaggedTensor`를 사용합니다.
예를 들어, 비정형 텐서는 정규 텐서로 표현할 수 없습니다.
<table>
<tr>
<th>`tf.RaggedTensor`, 형상: <code>[4, None]</code>
</th>
</tr>
<tr>
<td><img alt="A 2-axis ragged tensor, each row can have a different length." src="images/tensor/ragged.png"></td>
</tr>
</table>
```
ragged_list = [
[0, 1, 2, 3],
[4, 5],
[6, 7, 8],
[9]]
try:
tensor = tf.constant(ragged_list)
except Exception as e:
print(f"{type(e).__name__}: {e}")
```
대신 `tf.ragged.constant`를 사용하여 `tf.RaggedTensor`를 작성합니다.
```
ragged_tensor = tf.ragged.constant(ragged_list)
print(ragged_tensor)
```
`tf.RaggedTensor`의 형상에는 알 수 없는 차원이 있습니다.
```
print(ragged_tensor.shape)
```
## 문자열 텐서
`tf.string`은 `dtype`이며, 텐서에서 문자열(가변 길이의 바이트 배열)과 같은 데이터를 나타낼 수 있습니다.
문자열은 원자성이므로 Python 문자열과 같은 방식으로 인덱싱할 수 없습니다. 문자열의 길이는 텐서의 차원 중의 하나가 아닙니다. 문자열을 조작하는 함수에 대해서는 `tf.strings`를 참조하세요.
다음은 스칼라 문자열 텐서입니다.
```
# Tensors can be strings, too here is a scalar string.
scalar_string_tensor = tf.constant("Gray wolf")
print(scalar_string_tensor)
```
문자열의 벡터는 다음과 같습니다.
<table>
<tr>
<th>문자열의 벡터, 형상: <code>[3,]</code>
</th>
</tr>
<tr>
<td><img alt="The string length is not one of the tensor's axes." src="images/tensor/strings.png"></td>
</tr>
</table>
```
# If we have two string tensors of different lengths, this is OK.
tensor_of_strings = tf.constant(["Gray wolf",
"Quick brown fox",
"Lazy dog"])
# Note that the shape is (2,), indicating that it is 2 x unknown.
print(tensor_of_strings)
```
위의 출력에서 `b` 접두사는 `tf.string` dtype이 유니코드 문자열이 아니라 바이트 문자열임을 나타냅니다. TensorFlow에서 유니코드 텍스트를 처리하는 자세한 내용은 [유니코드 튜토리얼](https://www.tensorflow.org/tutorials/load_data/unicode)을 참조하세요.
유니코드 문자를 전달하면 UTF-8로 인코딩됩니다.
```
tf.constant("🥳👍")
```
문자열이 있는 일부 기본 함수는 `tf.strings`을 포함하여 `tf.strings.split`에서 찾을 수 있습니다.
```
# We can use split to split a string into a set of tensors
print(tf.strings.split(scalar_string_tensor, sep=" "))
# ...but it turns into a `RaggedTensor` if we split up a tensor of strings,
# as each string might be split into a different number of parts.
print(tf.strings.split(tensor_of_strings))
```
<table>
<tr>
<th>세 개의 분할된 문자열, 형상: <code>[3, None]</code>
</th>
</tr>
<tr>
<td><img alt="Splitting multiple strings returns a tf.RaggedTensor" src="images/tensor/string-split.png"></td>
</tr>
</table>
`tf.string.to_number`:
```
text = tf.constant("1 10 100")
print(tf.strings.to_number(tf.strings.split(text, " ")))
```
`tf.cast`를 사용하여 문자열 텐서를 숫자로 변환할 수는 없지만, 바이트로 변환한 다음 숫자로 변환할 수 있습니다.
```
byte_strings = tf.strings.bytes_split(tf.constant("Duck"))
byte_ints = tf.io.decode_raw(tf.constant("Duck"), tf.uint8)
print("Byte strings:", byte_strings)
print("Bytes:", byte_ints)
# Or split it up as unicode and then decode it
unicode_bytes = tf.constant("アヒル 🦆")
unicode_char_bytes = tf.strings.unicode_split(unicode_bytes, "UTF-8")
unicode_values = tf.strings.unicode_decode(unicode_bytes, "UTF-8")
print("\nUnicode bytes:", unicode_bytes)
print("\nUnicode chars:", unicode_char_bytes)
print("\nUnicode values:", unicode_values)
```
`tf.string` dtype은 TensorFlow의 모든 원시 바이트 데이터에 사용됩니다. `tf.io` 모듈에는 이미지 디코딩 및 csv 구문 분석을 포함하여 데이터를 바이트로 변환하거나 바이트에서 변환하는 함수가 포함되어 있습니다.
## 희소 텐서
때로는 매우 넓은 임베드 공간과 같이 데이터가 희소합니다. TensorFlow는 `tf.sparse.SparseTensor` 및 관련 연산을 지원하여 희소 데이터를 효율적으로 저장합니다.
<table>
<tr>
<th>`tf.SparseTensor`, 형상: <code>[3, 4]</code>
</th>
</tr>
<tr>
<td><img alt="An 3x4 grid, with values in only two of the cells." src="images/tensor/sparse.png"></td>
</tr>
</table>
```
# Sparse tensors store values by index in a memory-efficient manner
sparse_tensor = tf.sparse.SparseTensor(indices=[[0, 0], [1, 2]],
values=[1, 2],
dense_shape=[3, 4])
print(sparse_tensor, "\n")
# We can convert sparse tensors to dense
print(tf.sparse.to_dense(sparse_tensor))
```
| github_jupyter |
```
import os
import numpy as np
import torch
from utils import load_model
model, _ = load_model('pretrained/tsp_20/')
model.eval() # Put in evaluation mode to not track gradients
xy = np.random.rand(100, 2)
def make_oracle(model, xy, temperature=1.0):
num_nodes = len(xy)
xyt = torch.tensor(xy).float()[None] # Add batch dimension
with torch.no_grad(): # Inference only
embeddings, _ = model.embedder(model._init_embed(xyt))
# Compute keys, values for the glimpse and keys for the logits once as they can be reused in every step
fixed = model._precompute(embeddings)
def oracle(tour):
with torch.no_grad(): # Inference only
# Input tour with 0 based indices
# Output vector with probabilities for locations not in tour
tour = torch.tensor(tour).long()
if len(tour) == 0:
step_context = model.W_placeholder
else:
step_context = torch.cat((embeddings[0, tour[0]], embeddings[0, tour[-1]]), -1)
# Compute query = context node embedding, add batch and step dimensions (both 1)
query = fixed.context_node_projected + model.project_step_context(step_context[None, None, :])
# Create the mask and convert to bool depending on PyTorch version
mask = torch.zeros(num_nodes, dtype=torch.uint8) > 0
mask[tour] = 1
mask = mask[None, None, :] # Add batch and step dimension
log_p, _ = model._one_to_many_logits(query, fixed.glimpse_key, fixed.glimpse_val, fixed.logit_key, mask)
p = torch.softmax(log_p / temperature, -1)[0, 0]
assert (p[tour] == 0).all()
assert (p.sum() - 1).abs() < 1e-5
#assert np.allclose(p.sum().item(), 1)
return p.numpy()
return oracle
oracle = make_oracle(model, xy)
sample = False
tour = []
tour_p = []
while(len(tour) < len(xy)):
p = oracle(tour)
if sample:
# Advertising the Gumbel-Max trick
g = -np.log(-np.log(np.random.rand(*p.shape)))
i = np.argmax(np.log(p) + g)
# i = np.random.multinomial(1, p)
else:
# Greedy
i = np.argmax(p)
tour.append(i)
tour_p.append(p)
print(tour)
%matplotlib inline
from matplotlib import pyplot as plt
from matplotlib.collections import PatchCollection
from matplotlib.patches import Rectangle
from matplotlib.lines import Line2D
# Code inspired by Google OR Tools plot:
# https://github.com/google/or-tools/blob/fb12c5ded7423d524fc6c95656a9bdc290a81d4d/examples/python/cvrptw_plot.py
def plot_tsp(xy, tour, ax1):
"""
Plot the TSP tour on matplotlib axis ax1.
"""
ax1.set_xlim(0, 1)
ax1.set_ylim(0, 1)
xs, ys = xy[tour].transpose()
xs, ys = xy[tour].transpose()
dx = np.roll(xs, -1) - xs
dy = np.roll(ys, -1) - ys
d = np.sqrt(dx * dx + dy * dy)
lengths = d.cumsum()
# Scatter nodes
ax1.scatter(xs, ys, s=40, color='blue')
# Starting node
ax1.scatter([xs[0]], [ys[0]], s=100, color='red')
# Arcs
qv = ax1.quiver(
xs, ys, dx, dy,
scale_units='xy',
angles='xy',
scale=1,
)
ax1.set_title('{} nodes, total length {:.2f}'.format(len(tour), lengths[-1]))
fig, ax = plt.subplots(figsize=(10, 10))
plot_tsp(xy, tour, ax)
from matplotlib.collections import PatchCollection
from matplotlib.patches import Rectangle
from matplotlib.lines import Line2D
from IPython.display import HTML
from celluloid import Camera # pip install celluloid
def format_prob(prob):
return ('{:.6f}' if prob > 1e-5 else '{:.2E}').format(prob)
def plot_tsp_ani(xy, tour, tour_p=None, max_steps=1000):
n = len(tour)
fig, ax1 = plt.subplots(figsize=(10, 10))
xs, ys = xy[tour].transpose()
dx = np.roll(xs, -1) - xs
dy = np.roll(ys, -1) - ys
d = np.sqrt(dx * dx + dy * dy)
lengths = d.cumsum()
ax1.set_xlim(0, 1)
ax1.set_ylim(0, 1)
camera = Camera(fig)
total_length = 0
cum_log_prob = 0
for i in range(n + 1):
for plot_probs in [False] if tour_p is None or i >= n else [False, True]:
# Title
title = 'Nodes: {:3d}, length: {:.4f}, prob: {}'.format(
i, lengths[i - 2] if i > 1 else 0., format_prob(np.exp(cum_log_prob))
)
ax1.text(0.6, 0.97, title, transform=ax.transAxes)
# First print current node and next candidates
ax1.scatter(xs, ys, s=40, color='blue')
if i > 0:
ax1.scatter([xs[i - 1]], [ys[i - 1]], s=100, color='red')
if i > 1:
qv = ax1.quiver(
xs[:i-1],
ys[:i-1],
dx[:i-1],
dy[:i-1],
scale_units='xy',
angles='xy',
scale=1,
)
if plot_probs:
prob_rects = [Rectangle((x, y), 0.01, 0.1 * p) for (x, y), p in zip(xy, tour_p[i]) if p > 0.01]
pc = PatchCollection(prob_rects, facecolor='lightgray', alpha=1.0, edgecolor='lightgray')
ax1.add_collection(pc)
camera.snap()
if i < n and tour_p is not None:
# Add cumulative_probability
cum_log_prob += np.log(tour_p[i][tour[i]])
if i > max_steps:
break
# Plot final tour
# Scatter nodes
ax1.scatter(xs, ys, s=40, color='blue')
# Starting node
ax1.scatter([xs[0]], [ys[0]], s=100, color='red')
# Arcs
qv = ax1.quiver(
xs, ys, dx, dy,
scale_units='xy',
angles='xy',
scale=1,
)
if tour_p is not None:
# Note this does not use stable logsumexp trick
cum_log_prob = format_prob(np.exp(sum([np.log(p[node]) for node, p in zip(tour, tour_p)])))
else:
cum_log_prob = '?'
ax1.set_title('{} nodes, total length {:.4f}, prob: {}'.format(len(tour), lengths[-1], cum_log_prob))
camera.snap()
return camera
animation = plot_tsp_ani(xy, tour, tour_p).animate(interval=500)
# animation.save('images/tsp.gif', writer='imagemagick', fps=2) # requires imagemagick
# compress by running 'convert tsp.gif -strip -coalesce -layers Optimize tsp.gif'
HTML(animation.to_html5_video()) # requires ffmpeg
```
| github_jupyter |
# Sentence Generation from Language Model
This tutorial demonstrates how to generate text using a pre-trained language model in the following two ways:
- with sequence sampler
- with beam search sampler
Variables to configure when generating sequences:
- V = vocabulary size
- T = sequence length
- the number of possible outcomes to consider a sequence = V^T.
Given a language model, we can generate sequences according to the probability that they would occur according to our model. At each time step, a language model predicts the likelihood of each word occuring, given the context from prior time steps. The outputs at any time step can be any word from the vocabulary whose size is V and thus the number of all possible outcomes for a sequence of length T is thus V^T.
While sometimes we might want to generate sentences according to their probability of occuring, at other times we want to find the sentences that *are most likely to occur*. This is especially true in the case of language translation where we don't just want to see *a* translation. We want the *best* translation. While finding the optimal outcome quickly becomes intractable as time step increases, there are still many ways to sample reasonably good sequences. GluonNLP provides two samplers for generating from a language model: SequenceSampler and BeamSearchSampler.
First import the libraries:
```
import numpy as np
import mxnet as mx
import gluonnlp as nlp
import text_generation.model
```
## Load Pretrained Language Model
```
# change to mx.cpu() if GPU is not present
ctx = mx.cpu()
model, vocab = text_generation.model.get_model(name='gpt2_117m',
dataset_name='openai_webtext',
pretrained=True,
ctx=ctx)
tokenizer = nlp.data.GPT2BPETokenizer()
detokenizer = nlp.data.GPT2BPEDetokenizer()
eos_id = vocab[vocab.eos_token]
print(vocab.eos_token)
```
## Sampling a Sequence
### Sequence Sampler
A SequenceSampler samples from the contextual multinomial distribution produced by the language model at each time step. Since we may want to control how "sharp" the distribution is to tradeoff diversity with correctness, we can use the temperature option in SequenceSampler, which controls the temperature of the softmax function.
For each input same, sequence sampler can sample multiple independent sequences at once. The number of independent sequences to sample can be specified through the argument `beam_size`.
```
bos_str = 'Deep learning and natural language processing'
if not bos_str.startswith(' '):
bos_str = ' ' + bos_str
bos_tokens = tokenizer(bos_str)
bos_ids = vocab[bos_tokens]
print(bos_tokens)
```
#### Define the Decoder
```
class GPT2Decoder(text_generation.model.LMDecoder):
def __call__(self, inputs, states):
inputs = inputs.expand_dims(axis=1)
out, new_states = self.net(inputs, states)
out = mx.nd.slice_axis(out, axis=1, begin=0, end=1).reshape((inputs.shape[0], -1))
return out, new_states
decoder = GPT2Decoder(model)
```
#### Define the initial state
```
def get_initial_input_state(decoder, bos_ids, temperature):
inputs, begin_states = decoder.net(
mx.nd.array([bos_ids], dtype=np.int32, ctx=ctx), None)
inputs = inputs[:, -1, :]
smoothed_probs = (inputs / temperature).softmax(axis=1)
inputs = mx.nd.sample_multinomial(smoothed_probs, dtype=np.int32)
return inputs, begin_states
```
### Define the Sampler
```
# number of independent sequences to search
beam_size = 2
temperature = 0.97
num_results = 2
# must be less than 1024
max_len = 256 - len(bos_tokens)
sampler = nlp.model.SequenceSampler(beam_size=beam_size,
decoder=decoder,
eos_id=eos_id,
max_length=max_len,
temperature=temperature)
```
#### Generate result
```
def generate(decoder, bos_ids, temperature, sampler, num_results, vocab):
inputs, begin_states = get_initial_input_state(decoder, bos_ids, temperature)
# samples have shape (1, beam_size, length), scores have shape (1, beam_size)
samples, scores, valid_lengths = sampler(inputs, begin_states)
samples = samples[0].asnumpy()
scores = scores[0].asnumpy()
valid_lengths = valid_lengths[0].asnumpy()
print('Generation Result:')
for i in range(num_results):
generated_tokens = [vocab.idx_to_token[ele] for ele in samples[i][:valid_lengths[i]]]
tokens = bos_tokens + generated_tokens[1:]
print([detokenizer(tokens).strip(), scores[i]])
generate(decoder, bos_ids, temperature, sampler, num_results, vocab)
```
### Beam Search Sampler
To overcome the exponential complexity in sequence decoding, beam search decodes greedily, keeping those sequences that are most likely based on the probability up to the current time step. The size of this subset is called the *beam size*. Suppose the beam size is K and the output vocabulary size is V. When selecting the beams to keep, the beam search algorithm first predict all possible successor words from the previous K beams, each of which has V possible outputs. This becomes a total of K\*V paths. Out of these K\*V paths, beam search ranks them by their score keeping only the top K paths.
#### Generate Sequences w/ Beam Search
Next, we are going to generate sentences starting with "I love it" using beam search first. We feed ['I', 'Love'] to the language model to get the initial states and set the initial input to be the word 'it'. We will then print the top-3 generations.
#### Scorer Function
The BeamSearchScorer is a simple HybridBlock that implements the scoring function with length penalty in Google NMT paper.
```
scores = (log_probs + scores) / length_penalty
length_penalty = (K + length)^alpha / (K + 1)^alpha
```
```
scorer = nlp.model.BeamSearchScorer(alpha=0.5, K=5, from_logits=False)
```
#### Beam Search Sampler
Given a scorer and decoder, we are ready to create a sampler. We use symbol '.' to indicate the end of sentence (EOS). We can use vocab to get the index of the EOS, and then feed the index to the sampler. The following codes shows how to construct a beam search sampler. We will create a sampler with 4 beams and a maximum sample length of 20.
```
beam_sampler = nlp.model.BeamSearchSampler(beam_size=3,
decoder=decoder,
eos_id=eos_id,
scorer=scorer,
max_length=max_len)
```
#### Generate Sequences w/ Sequence Sampler
Now, use the sequence sampler created to sample sequences based on the same inputs used previously.
```
generate(decoder, bos_ids, temperature, beam_sampler, num_results, vocab)
```
### Practice
- Tweak alpha and K in BeamSearchScorer, how are the results changed?
- Try different samples to decode.
| github_jupyter |
<a href="https://colab.research.google.com/github/chemaar/python-programming-course/blob/master/Lab_6_Functions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lab 6: Functions
In this notebook, we propose and solve some exercises about functions in Python.
* **In these exercises, we can always proceed solving the problems in a generic way or taking advantage of Python capabilities. As a recommendation, first, try the generic way (applicable to any programming language) and, then, using Python**
* **As a good programming practice, our test cases should ensure that all branches of the code are executed at least once.**
* **In the specific case of functions, we have always to keep in mind the next keypoints:**"
* Design the functions defining a proper domain and range.
* Think in the pre-conditions to execute the function.
* Design (pure) functions without side-effects.
## List of exercises
1. Write a function that defines a set of input parameters and displays their identifiers, types and values. Invoke this function from the `main` function.
* Input: my_function(1,"Hello",[1,2,3])
* Expected output:
```
1 of type <class 'int'> with id: 10914496
Hello of type <class 'str'> with id: 139654081206232
[1, 2, 3] of type <class 'list'> with id: 139654081356488
```
* Make use of the Python functions:
```
type(object)
id(object)
```
```
def my_fun(value, message, alist):
print(value, "of type ", type(value), " with id: ", id(value))
print(message, "of type ", type(message), " with id: ", id(message))
print(alist, "of type ", type(alist), " with id: ", id(alist))
if __name__=="__main__":
my_fun(1,"Hello", [1,2,3])
```
2. Write a function that takes two arguments, at lest one parameter with a default boolean value True, and prints out the values of all parameters.
* Input:
* `default_parameters(1)`
* `default_parameters(1, False)`
* Expected output:
```
1
True
1
False
```
```
def default_parameters(a, b = True):
print(a)
print(b)
if __name__=="__main__":
default_parameters(1)
default_parameters(1, False)
```
3. Write a function that takes three arguments (integer, string and list), modifies the value of such argument (displaying the id) and checks the value in the invocation point (displaying the id again).
* Input:
```
a = 2
msg = "Hello"
alist = [1,2,3]
my_fun(a, msg, alist)
```
* Expected output (the ids may change):
```
10914528
139654071637640
139654071236424
Call function...
10914528
4
139654071637640
Other
139654071236424
[1, 3]
After calling function...
2
Hello
[1, 2, 3]
```
```
def my_fun(a,msg,alist):
print(id(a))
a = 4
print(a)
print(id(msg))
msg = "Other"
print(msg)
print(id(alist))
alist = [1,3]
print(alist)
if __name__=="__main__":
a = 2
msg = "Hello"
alist = [1,2,3]
print(id(a))
print(id(msg))
print(id(alist))
print("Call function...")
my_fun(a, msg, alist)
print("After calling function...")
print(a)
print(msg)
print(alist)
```
4. Write a function that takes a parameter (a list), appends a new element and prints out the content of the list in the invocation point.
* Input: [1,2,3], a new element 4 is added.
* Expected output:
```
Before calling...
Hello
After calling...
Hello Mary
```
```
def add_element(alist):
alist.append(4)
if __name__=="__main__":
alist = [1,2,3]
print("Before calling...")
print(alist)
add_element(alist)
print("After calling...")
print(alist)
```
5. Write a function that takes a parameter (a string), appends a new string and prints out the content of the string in the invocation point.
* Input: "Hello", a new string " Mary".
* Expected output:
```
Before calling...
Hello
After calling...
Hello
```
```
def add_message(msg):
msg = msg + " Mary"
if __name__=="__main__":
msg = "Hello"
print("Before calling...")
print(msg)
add_message(msg)
print("After calling...")
print(msg)
```
6. Write a function that takes two integer numbers and returns tha addition of both numbers (an integer number).
* Input: my_add(1,2).
* Expected output:
```
3
```
```
def my_add(a,b):
return a+b
if __name__=="__main__":
print(my_add(1,2))
```
7. Write a function to compare two integer numbers. The function shall return:
* 0 if both values are equal.
* 1 if the first parameter is greater than the second.
* -1 if the second parameter is greater than the first.
* Input:
* are_equal(1,2)
* are_equal(2,1)
* are_equal(1,1)
* Expected output:
```
1
-1
0
```
```
def are_equal(a,b):
if a == b:
return 0
elif a > b:
return 1
else:
return -1
if __name__=="__main__":
print(are_equal(1,2))
print(are_equal(2,1))
print(are_equal(1,1))
```
8. Write a function to implement the absolute value of an integer number.
* Input:
* my_abs(5)
* my_abs(-5)
* Expected output:
```
5
5
```
```
def my_abs(a):
return ( -a if a<0 else a)
if __name__=="__main__":
print(my_abs(5))
print(my_abs(-5))
```
9. Write a function that takes as an argument one tuple packing argument (*args) and diplays the values.
* Input:
* my_f(2,"Hello",[2,3])
* Expected output:
```
2
Hello
[2,3]
```
```
def my_f(*args):
if args:
for v in args:
print(v)
if __name__=="__main__":
my_f(2,"Hello",[2,3])
```
10. Write a function that takes as an argument one dictionary argument (**kwargs) and diplays the values.
* Input:
* my_f(name="Mary", age=25)
* Expected output:
```
Key: name , value: Mary
Key: age , value: 25
```
```
def my_f(**kwargs):
if kwargs:
for k,v in kwargs.items():
print("Key: ", k, ", value: ", v)
if __name__=="__main__":
my_f(name="Mary", age=25)
```
11. Write a function, `is_leap`, that given a year number, returns whether is a leap year (reuse the code in the previous notebook: *Lab_3_Control_Flow_Conditional_Statements*).
* Input: -1
* Expected output:
```
False
```
* Input: 2019
* Expected output:
```
False
```
* Input: 2020
* Expected output:
```
True
```
```
def is_leap(year):
is_leap_year = False
if year >= 0:
is_leap_year = (year % 4 == 0)
is_leap_year = is_leap_year and (year % 100 != 0)
is_leap_year = is_leap_year or (year % 400 == 0)
return is_leap_year
if __name__=="__main__":
print(is_leap(-1))
print(is_leap(2019))
print(is_leap(2020))
```
12. Check the next function in which a documentation string is added.
* The documentation is enclosed between `"""`.
* The documentation string is multiline.
* The documentation is string is situated after the function signature.
* The documentation can be checked in two manners:
* help(function_name): displays the function signature and the doc string.
* `function_name.__doc__`: returns the doc string.
* The reference for documentation strings is defined in the next PEP: https://www.python.org/dev/peps/pep-0257/.
```
def my_sum(alist):
"""Returns the sum of a list of numbers."""
aggregated = 0
if alist:
for v in alist:
aggregated += aggregated + v
return aggregated
def my_sum2(alist):
"""Returns the sum of a list of numbers.
Keyword arguments:
alist: is a non null list of integer numbers.
Last update: January 2020
"""
aggregated = 0
if alist:
for v in alist:
aggregated += aggregated + v
return aggregated
help(my_sum)
print(my_sum.__doc__)
help(my_sum2)
```
13. Check the next function in which **metadata** to describe the function is added.
* Internally, the annotations are added as a dictionary to the function object.
* The reference for annotations is defined in the next PEP: https://www.python.org/dev/peps/pep-3107/
```
#Annotations can be just string values.
def my_add(a: '<a>', b: '<b>') -> '<return_value>':
return a+b
print(my_add.__annotations__)
#Annotations can also include types. However, this is only documentation. it does not impose any restriction on the parameters.
def my_add2(a: int, b: int) -> float:
return a+b
print(my_add2.__annotations__)
```
14. Write a program to calculate the factorial of an integer number.
* factorial (n) = n * factorial (n-1)
* Input: an integer number, 5
* Expected output:
```
120
```
```
def factorial(number):
if number < 0:
fact = -1
elif number == 0 or number == 1:
fact = 1
else:
fact = 1
for i in range(1,number+1):
fact *= i
return fact
if __name__=="__main__":
print(factorial(5))
```
15. Write a program to calculate the factorial of an integer number using a recursive function.
* factorial (n) = n * factorial (n-1)
A recursive function has two main parts:
* The basic case. In the factorial function, when $n=0$ or $n=1$.
* The recursive case. In the factorial function, when $n>1$.
* Input: an integer number, 5
* Expected output:
```
120
```
```
def factorial(number):
if number < 0:
return -1
elif number == 0 or number == 1:
return 1
else:
return number * factorial(number-1)
if __name__=="__main__":
print(factorial(5))
```
16. Write a function to calculate the exponentiation of a number with base b, and exponent n.
* $base^{exponent} = base * base* base...*base$
* Input: base 2, exponent 3
* Expected output:
```
8
```
```
def my_pow (base, exponent):
mypow = 1
if exponent > 0 :
for i in range(0,exponent):
mypow *= base
return mypow
if __name__=="__main__":
print(my_pow(2,3))
```
17. Write a function to detect if a number is a prime number.
* Input: 5, 8
* Expected output:
```
The number 5 is prime.
The number 8 is not prime.
```
```
def is_prime(n):
n_divisors = 1
divisor = 2
while divisor < n and n_divisors <= 2:
if n % divisor == 0:
n_divisors = n_divisors + 1
divisor = divisor + 1
return n_divisors > 2
if __name__=="__main__":
n = 8
if is_prime(n):
print("The number {} is not prime.".format(n))
else:
print("The number {} is prime.".format(n))
```
18. Write a function to sum up the Fibonacci sequence of the first n numbers.
fibonacci(n) =
fibonacci (0) = 0
fibonacci (1) = 1
fibonacci (n) = fibonacci(n-1) + fibonacci (n-2)
* Input: a positive number $n = 4$
* Expected output:
```
Fibonacci of: 0 , sequence: [0] , sum = 0
Fibonacci of: 1 , sequence: [0, 1] , sum = 1
Fibonacci of: 2 , sequence: [0, 1, 1] , sum = 2
Fibonacci of: 3 , sequence: [0, 1, 1, 2] , sum = 4
Fibonacci of: 4 , sequence: [0, 1, 1, 2, 3] , sum = 7
```
```
def fibonacci_seq(n):
fib_seq = []
if n == 0:
return [0]
elif n == 1:
return [0, 1]
else:
fn2 = 0
fn1 = 1
fib_seq.append(fn2)
fib_seq.append(fn1)
for i in range(2, n+1):
fcurrent = fn1 + fn2
fib_seq.append(fcurrent)
temp = fn1
fn1 = fcurrent
fn2 = temp
return fib_seq
if __name__=="__main__":
for n in range(5):
seq = fibonacci_seq(n)
print("Fibonacci of:",n,", sequence: ", seq,", sum = ",sum(seq))
```
19. Write a recursive function to calculate the next Fibonacci number of the first n numbers.
fibonacci(n) =
fibonacci (0) = 0
fibonacci (1) = 1
fibonacci (n) = fibonacci(n-1) + fibonacci (n-2)
* Input: a positive number $n = 4$
* Expected output:
```
Fib at position 0 is 0
Fib at position 1 is 1
Fib at position 2 is 1
Fib at position 3 is 2
Fib at position 4 is 3
```
```
def fibonacci_r(n):
if n == 0:
result = 0
elif n == 1:
result = 1
elif n > 1:
result = fibonacci_r(n-1) + fibonacci_r(n-2)
return result
if __name__=="__main__":
for n in range(5):
print("Fib at position ",n," is ",fibonacci_r(n))
```
20. Write a function to calculate the combinatorial number (reuse your previous factorial function):
$\binom{m}{n} = \frac{m!}{n! (m-n)!}$
* Input: m = 10, n = 5
* Expected output:
```
252
```
```
def factorial(number):
if number < 0:
return -1
elif number == 0 or number == 1:
return 1
else:
return number * factorial(number-1)
def combinatorial_number(m, n):
factorialm = factorial(m)
factorialn = factorial(n)
factorialmn = factorial(m-n)
return (factorialm / (factorialn * factorialmn))
if __name__=="__main__":
print(combinatorial_number(10,5))
```
21. Write a program to display a menu with 3 options (to say hello in English, Spanish and French) and to finish, the user shall introduce the keyword "quit". If any other option is introduced, the program shall display that the input value is not a valid option.
* **Refactor your previous code to provide a function that displays the menu and returns the selected option.**
* Input: test the options
* Expected output:
```
----------MENU OPTIONS----------
1-Say Hello!
2-Say ¡Hola!
3-Say Salut!
> introduce an option or quit to exit...
```
```
def menu():
option = "-1"
while option not in {"1","2","3","quit"}:
print("----------MENU OPTIONS----------")
print("1-Say Hello!")
print("2-Say ¡Hola!")
print("3-Say Salut!")
option = input("> introduce an option or quit to exit...")
return option
if __name__=="__main__":
option = "-1"
while option != "quit":
option = menu()
if option == "1":
print("Hello!")
elif option == "2":
print("¡Hola!")
elif option == "3":
print("Salut!")
elif option == "quit":
print("...finishing...")
else:
print("Not a valid option: ", option)
```
22. Write a function to detect whether a number is a perfect number.
* A number is perfect if the addition of all positive divisors is equal to the number.
* Input: n = 6
* Expected output:
```
The number 6 is perfect.
```
```
def is_perfect(n):
perfect = 0
for i in range (1,n):
if n%i == 0:
perfect = perfect + i
return perfect == n
if __name__=="__main__":
n = 6
if is_perfect(n):
print("The number {} is perfect.".format(n))
else:
print("The number {} is NOT perfect.".format(n))
```
23. Write a function to calculate the length of a sequence. If the list is None, the function shall return -1.
* Input: [3,4,5,6]
* Expected output:
```
The length of the list is: 4
```
```
def my_len(alist):
if alist:
size = 0
for v in alist:
size += 1
else:
size = -1
return size
if __name__=="__main__":
length = my_len([1,2,3,4])
print("The length of the list is: ", length)
```
24. Write a function that given a list of $n$ numbers, calculates and returns the max value within the list and its position.
* Input: [8, 1, 9, 2]
* Expected output:
```
The max value is: 9 in position: 2.
```
```
def max_position(values):
max_value = -1
position = -1
if values and len(values) > 0:
max_value = values[0]
position = 0
for i in range(1, len(values)):
if values [i] > max_value:
max_value = values [i]
position = i
return (max_value, position)
if __name__=="__main__":
values = [8,1,9,2]
max_value, position = max_position(values)
print("The max value is {} in position: {}".format(max_value, position))
```
25. Write a function that given a list of $n$ numbers and a target number $k$, counts the number of occurrences of $k$ in the list.
* Input: [8, 1, 9, 1], $k=1$
* Expected output:
```
The number 1 has 2 occurrences.
```
```
def my_count(values, k):
occurrences = 0
for v in values:
if k == v:
occurrences += 1
return occurrences
if __name__=="__main__":
values = [8,1,9,1]
k = 1
count = my_count(values,k)
print("The number {} has {} occurrences.".format(k, count))
```
26. Write a module named `my_list_functions.py` that contains functions for:
* Counting the number of occurrences of a value $k$ in a list.
* Finding the position of the first/last/all apparition (this shall be a parameter) of the value $k$.
* Creating a new list in reverse order.
* Returning the first/last (this shall be a parameter) $k$ numbers of the list.
* Making the union of two lists.
* Making the intersection of two lists.
* Creating chunks of $k$ elements.
These functions will be invoked from a program including the module with the next directive:
`from my_list_functions import *`
```
#Content of the file my_list_functions.py
def count(alist, k):
occurrences = 0
if alist:
for v in alist:
if k == v:
occurrences += 1
return occurrences
def find_last(values, k):
last_occur = -1
if values:
i = len(values)-1
while i>=0 and last_occur == -1:
if values[i] == k:
last_occur = i
i -= 1
return last_occur
def find_first(values, k):
first_occur = -1
i = 0
if values:
size = len(values)
while i<size and first_occur == -1:
if values[i] == k:
first_occur = i
i += 1
return first_occur
def find_all(values, k):
positions = []
if values:
size = len(values)
for i in range(size):
if values[i] == k:
positions.append(i)
return positions
def find(values, k, strategy=0):
if strategy == 0:
return [find_first(values, k)]
elif strategy == 1:
return [find_last(values, k)]
else:
return find_all(values, k)
def reverse(values):
if values:
return values[::-1]
return []
def take_first(values, k):
if values and k>0:
return values[:k:]
return []
def take_last(values, k):
if values and k>0:
return values[len(values)-1:len(values)-k-1:-1]
return []
def union(l1, l2):
union_list = []
if l1 and l2:
union_list.extend(l1)
for v in l2:
if union_list.count(v) == 0:
union_list.append(v)
return union_list
def intersection(l1, l2):
intersection_list = []
if l1 and l2:
for v in l1:
if l2.count(v) != 0:
intersection_list.append(v)
return intersection_list
def chunks(values, k):
chunks = []
if values and k > 0:
size = len(values)
chunks = [values[i:k+i] for i in range(0, size, k)]
return chunks
#Content of the file app.py: both files must be in the same directory
from my_list_functions import *
if __name__=="__main__":
values = [8,1,9,1]
k = 1
print("All the assertions must be true") #Change to assert
print(count(values, k) == 2)
print(len(find(values, k)) == 1)
print(len(find(values, k, 1)) == 1)
print(len(find(values, k, 2)) == 2)
print(values[::-1] == reverse(values))
print(len(take_first(values, 2)) == 2)
print(len(take_last(values, 2)) == 2)
print(union([4,5,6], [5,7,8,9]) == [4, 5, 6, 7, 8, 9])
print(intersection([4,5,6], [5,7,8,9]) == [5])
print(len(chunks(values,2)) == 2)
```
27. Refactor the TIC, TAC, TOE game implementing the following functions.
* The program shall have a function to get the position to situate a new value.
* The program shall have function to detect whether a player is winner by rows.
* The program shall have function to detect whether a player is winner by columns.
* The program shall have function to detect whether a player is winner in the main diagonal.
* The program shall have function to detect whether a player is winner in the secondary diagonal.
* The program shall have function to print the board.
```
#Version using only one vector to store the positions
def print_board(board):
for i in range(n):
print(board[n*i:n*(i+1)])
def get_free_position(board):
set_position = False
x = -1
y = -1
size = len(board)
n = len(board) // 3
while not set_position:
x =int(input("Select position x:"))
y =int(input("Select position y:"))
if x>=0 and y >= 0 and (x*n+y)<size and board[x*n+y] == "":
set_position = True
else:
print("The position is already set.")
return (x,y,)
def is_winner_by_rows(board, current_player):
i = 0
winner = False
n = len(board) // 3
while not winner and i<n:
winner = board[n*i:n*(i+1)].count(current_player) == n
i = i + 1
return winner
def is_winner_by_cols(board, current_player):
winner = False
i = 0
size = len(board)
n = len(board) // 3
while not winner and i<n:
winner = board[i:size:n].count(current_player) == n
i = i + 1
return winner
def is_winner_main_diagonal(board, current_player):
size = len(board)
n = len(board) // 3
return board[:size:n+1].count(current_player) == n
def is_winner_secondary_diagonal(board, current_player):
size = len(board)
n = len(board) // 3
return board[n-1:size-1:n-1].count(current_player) == n
if __name__=="__main__":
n = 3
#Management in just one vector
board = ["" for x in range(n*n)]
current_player = "X"
other_player = "O"
end_game = False
situated = 0
while not end_game:
print("Turn of player: "+current_player)
print_board(board)
(x,y) = get_free_position(board)
board[x*n+y] = current_player
situated += 1
winner = is_winner_by_rows(board, current_player) or is_winner_by_cols(board, current_player) or is_winner_main_diagonal(board, current_player) or is_winner_secondary_diagonal(board, current_player)
end_game = winner or situated == 9
current_player, other_player = other_player, current_player
if winner:
print("The winner is: ", other_player)
else:
print("Draw")
print_board(board)
```
28. As a refactoring exercise, take exercises of previous notebooks and select the parts of code that can be a function, e.g. string functions, vector operations, matrix operations, etc.
## Advanced function concepts (not for studying, only information)
1. **High-order functions**. A high-order function is a function that:
1. takes a function as a parameter or
2. returns a function.
For instance, we are going to define a function that receives as parameters:
* The function, $f$, to be applied to.
* The list of elements, alist
and returns, a list of the values after applying $f$ to each of the elements in alist.
In this case, the example will return the square of the elements of the list.
* Input: $f$ my own function to calculate the square of a number, and the list [1,2,3].
* Output:
```
[1, 4, 9]
```
* Modify the program to make a function that adds 2 to each of the elements of the list.
* Output: `[3, 4, 5]`
```
#We define a function that takes as a parameter a function f and a list, a list,
#then applies the function f to any element in the list.
def apply_f_to_list(f, alist):
results = []
for v in alist:
value = f(v)
results.append(value)
return results
def my_square(n):
return n**2
def add_2(n):
return n+2
if __name__=="__main__":
values = [1,2,3]
results = apply_f_to_list(my_square,values)
print(results)
results = apply_f_to_list(add_2,values)
print(results)
```
2. **Lambda functions**. A **lambda function** is an anonymous function declared online.
* A lambda function can take any number of arguments.
* A lambda function can only return one expression.
* A lambda function is a Python function, so anything regarding parameters, annotations, etc. are applicable to lambda functions.
The theory behind lambda functions comes from the "[Lambda Calculus](https://en.wikipedia.org/wiki/Lambda_calculus)".
According to the [official documentation](https://docs.python.org/3/reference/expressions.html#grammar-token-lambda-expr), the Lambda functions follow the next grammar:
>lambda_expr ::= "lambda" [parameter_list] ":" expression
>lambda_expr_nocond ::= "lambda" [parameter_list] ":" expression_nocond
Lambda expressions (sometimes called lambda forms) are used to create anonymous functions. The expression lambda parameters: expression yields a function object. The unnamed object behaves like a function object defined with:
>def <lambda>(parameters):
>
> return expression
Lambda functions are mainly used in the following scenarios:
* Simple functions that we want to apply inline and we do not plan to reuse.
Lambda functions also come with some drawbacks:
* Syntax can be complex, it is not so intuitive as a regular function.
* Readability and understandability of the source code become complex.
* Need of thinking in a functional way (not intuitive when coming from imperative program.
In the Python PEP8 document recommends the following:
>Always use a def statement instead of an assignment statement that binds a lambda expression directly to an identifier.
>
>Yes:
>
>`def f(x): return 2*x`
>
>No:
>
>`f = lambda x: 2*x`
>
>The first form means that the name of the resulting function object is specifically 'f' instead of the generic '<lambda>'. This is more useful for tracebacks and string representations in general. The use of the assignment statement eliminates the sole benefit a lambda expression can offer over an explicit def statement (i.e. that it can be embedded inside a larger expression).
**However, Lambda functions are elegant to solve specific problems and parametrize some expressions in functional programming.**
Learn more: https://www.python.org/dev/peps/pep-0008/
```
(lambda x, y: x + y)(2, 3)
#We can assign the lambda function to a variable that will become a variable of type function.
f_add_two_numbers = (lambda x, y: x + y)
print(type(f_add_two_numbers))
result = f_add_two_numbers(2,3)
print(result)
values = [1,2,3]
#Let's make the function squares inline with a lambda expression
results = apply_f_to_list(lambda x: x**2 ,values)
print(results)
```
3. **Functional programming in Python**.
* `map`. Given a function, $f$, and a sequence, $S$, it returns a new iterator, $f(S)$, where each element in $f(S)$ is the result of applying the function $f$ to each element in $S$.
* `reduce`. Given an operator, $p$, and a sequence, $S$, it returns a value, $v$, after aggregating the items in $S$ using the operator $p$.
* `filter`. Given a filter function, $filter$, and a sequence, $S$, it returns a new iterator, $filtered(S)$, where each element in $filtered(S)$ meets the conditions established in the filter $filter$.
* `zip`. *The zip() function returns a zip object, which is an iterator of tuples where the first item in each passed iterator is paired together, and then the second item in each passed iterator are paired together etc.* (source: [W3C Schools](https://www.w3schools.com/python/ref_func_zip.asp))
```
#Applying a function through the map
values = [1,2,3]
squared_values = [x for x in map(lambda x: x**2, values)]
print(squared_values)
import functools
import itertools
import operator
#Aggregating values with reduce: sum of values
values = [1,2,3]
print (functools.reduce(lambda a,b : a+b, values))
#Get the max of a list
print (functools.reduce(lambda a,b : a if a > b else b, values))
#Count words frequency
words = [('grape', 2), ('grape', 3), ('apple', 5), ('apple', 1), ('banana', 2)]
word_frequency = {key: sum(list([v[1] for v in group])) for key, group in itertools.groupby(words, operator.itemgetter(0))}
print(word_frequency)
```
##References
* Interesting discussion about Lambda expressions: https://treyhunner.com/2018/09/stop-writing-lambda-expressions/
* Functional programming in Python: https://docs.python.org/3/howto/functional.html
* The Zip function: https://realpython.com/python-zip-function/
| github_jupyter |
```
try:
%load_ext autoreload
%autoreload 2
except:
pass
%matplotlib ipympl
import functions.add_path
import functions.plotnine_theme
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from plotnine import *
from plotnine.data import *
import functions.et_condition_df as condition_df
import functions.et_make_df as make_df
import be_load
import os
print(os.getcwd())
if os.getcwd()[-4:] != 'code':
try:
os.chdir('./code')
except:
os.chdir('/net/store/nbp/users/behinger/projects/etcomp/code')
from functions import et_import
from functions import et_helper
import av
from functions.et_import import raw_pl_data
from lib.pupil.pupil_src.shared_modules import file_methods as pl_file_methods
from functions import nbp_recalib
tmp_dat = raw_pl_data(subject="VP24")
# recalib to get flgas whether stimulus was fused or not
print(et_helper.size_px2deg(30)) # markersize
print(et_helper.size_px2deg(30*0.24)) # white frame
print(et_helper.size_px2deg(1920)) # complete monitor
print(et_helper.size_px2deg((30*0.24 * 2 + 30))) # marker border (twice white frame + marker)
print(et_helper.px2deg(0,orientation="horizontal"))
tmp_dat['gaze_positions_recalib'] = nbp_recalib.nbp_recalib(tmp_dat)
gaze_topic = [p['topic'] for p in tmp_dat['gaze_positions_recalib']]
plt.plot([p['gx'] for p in tmp_dat['gaze_positions_recalib'][22115819:22115819+50]])
plt.figure()
plt.plot([p['norm_pos'][0] for p in tmp_dat['gaze_positions_recalib'][94100:(94100+100)]],'o-')
ix = range()
dat = pd.DataFrame(tmp_dat['gaze_positions_recalib'][94105:(94105+10)])
dat[['gx','gy']] = pd.DataFrame(dat.norm_pos.values.tolist(), index= dat.index)
dat[['base0','base1']] = pd.DataFrame(dat.base_data.values.tolist(), index= dat.index)
dat[['p0_x','p0_y']] = pd.DataFrame([p['norm_pos'] for p in dat.base0.values.tolist()], index= dat.index)
dat[['p1_x','p1_y']] = pd.DataFrame([p['norm_pos'] for p in dat.base1.values.tolist()], index= dat.index)
dat[['p0_timestamp']] = pd.DataFrame([p['timestamp'] for p in dat.base0.values.tolist()], index= dat.index)
dat[['p1_timestamp']] = pd.DataFrame([p['timestamp'] for p in dat.base1.values.tolist()], index= dat.index)
plt.figure()
# gaze-mapped land
plt.plot(dat.timestamp,dat.gx-dat.gx[0],'o-')
plt.plot(dat.timestamp,dat.p0_x-dat.p0_x[0],'o-')
plt.plot(dat.timestamp,dat.p1_x-dat.p1_x[0],'o-')
#pupil land
plt.plot(dat.p0_timestamp,dat.p0_x-dat.p0_x[0],'o-')
plt.plot(dat.p1_timestamp,dat.p1_x-dat.p1_x[0],'o-')
plt.legend(['both','both_p0','both_p1','true_p0','true_p1'])
etsamples,etevents,etmsgs = et_import.import_pl('VP3',recalib=False,surfaceMap=False)
A = np.array([[0,0],[0,0],[10,10]])
B = np.array([[0,0],[0,10],[0,0]])
locations = np.array([(*e, *f) for e,f in zip(A,B)])
from camera_models import load_intrinsics
intrinsics = load_intrinsics('','Pupil Cam1 ID2',(1280, 720))
width,height = intrinsics.resolution
locations[:, ::2] *=width
locations[:, 1::2] = (1. - locations[:, 1::2]) * height
undistorted = intrinsics.unprojectPoints(locations)
undistorted.shape = -1, 2
undistorted_3d = np.ones((undistorted.shape[0], 3)) # shape: 2n x 3
undistorted_3d[:, :-1] = undistorted
undistorted_3d /= np.linalg.norm(undistorted_3d, axis=1)[:, np.newaxis]
# Cosine distance of A and B: (A @ B) / (||A|| * ||B||)
# No need to calculate norms, since A and B are normalized in our case.
# np.einsum('ij,ij->i', A, B) equivalent to np.diagonal(A @ B.T) but faster.
angular_err = np.einsum('ij,ij->i', undistorted_3d[::2, :], undistorted_3d[1::2, :])
np.rad2deg(np.arccos(angular_err.clip(-1.0, 1.0)))
from functions.et_make_df import calc_3d_angle_points
location = np.array([(*e, *f) for e,f in zip(A,B)])
location
[calc_3d_angle_points(l[0], l[1], l[2], l[3]) for l in location]
x_0,y_0 = 0,0
x_1,y_1 = 90,90
calc_3d_angle_points(x_0,y_0,x_1,y_1)
pi = np.pi
vec1 = sph2cart(x_0/360*2*pi+pi/2, y_0/360*2*pi+pi/2)
vec2 = sph2cart(x_1/360*2*pi+pi/2, y_1/360*2*pi+pi/2)
# pupillabs : precision = np.sqrt(np.mean(np.rad2deg(np.arccos(succesive_distances.clip(-1., 1.))) ** 2))
cosdistance = np.dot(vec1,vec2)/(np.linalg.norm(vec1)*np.linalg.norm(vec2))
angle = np.arccos(np.clip(cosdistance,-1., 1.))
angle = angle * 360/(2*pi) # radian to degree
angle
np.sqrt(2)
from functions.et_preprocess import preprocess_et
%config InlineBackend.figure_format = 'svg'
import pandas as pd
import matplotlib
from plotnine import *
# Say, "the default sans-serif font is COMIC SANS"
matplotlib.rcParams['svg.fonttype'] = 'none'
matplotlib.rcParams['font.sans-serif'] = "Helvetica Neue LT Pro"
# Then, "ALWAYS use sans-serif fonts"
matplotlib.rcParams['font.family'] = "sans-serif"
from functions import plotnine_theme
ggplot(pd.DataFrame({'x':[1,2,3],'y':[2,3,4]},columns=['x','y']),aes(x='x',y='y'))+geom_point()+ggtitle('hello')
import matplotlib.font_manager
from IPython.core.display import HTML
def make_html(fontname):
return "<p>{font}: <span style='font-family:{font}; font-size: 24px;'>{font}</p>".format(font=fontname)
code = "\n".join([make_html(font) for font in sorted(set([f.name for f in matplotlib.font_manager.fontManager.ttflist]))])
HTML("<div style='column-count: 2;'>{}</div>".format(code))
matplotlib.rcParams['svg.fonttype'] = 'path'
from matplotlib.font_manager import FontProperties
import matplotlib.pyplot as plt
plt.subplot(111, facecolor='w')
font0 = FontProperties()
alignment = {'horizontalalignment': 'center', 'verticalalignment': 'baseline'}
# Show family options
families = ['serif', 'sans-serif', 'cursive', 'fantasy', 'monospace']
font1 = font0.copy()
font1.set_size('large')
t = plt.text(-0.8, 0.9, 'family', fontproperties=font1,
**alignment)
yp = [0.8, 0.7, 0.6, 0.5, 0.4, 0.3, 0.2]
for k, family in enumerate(families):
font = font0.copy()
font.set_family(family)
t = plt.text(-0.8, yp[k], family, fontproperties=font,
**alignment)
# Show style options
styles = ['normal', 'italic', 'oblique']
t = plt.text(-0.4, 0.9, 'style', fontproperties=font1,
**alignment)
for k, style in enumerate(styles):
font = font0.copy()
font.set_family('sans-serif')
font.set_style(style)
t = plt.text(-0.4, yp[k], style, fontproperties=font,
**alignment)
# Show variant options
variants = ['normal', 'small-caps']
t = plt.text(0.0, 0.9, 'variant', fontproperties=font1,
**alignment)
for k, variant in enumerate(variants):
font = font0.copy()
font.set_family('serif')
font.set_variant(variant)
t = plt.text(0.0, yp[k], variant, fontproperties=font,
**alignment)
# Show weight options
weights = ['light', 'medium', 'semibold', 'bold', 'heavy', 'black']
t = plt.text(0.4, 0.9, 'weight', fontproperties=font1,
**alignment)
for k, weight in enumerate(weights):
font = font0.copy()
font.set_weight(weight)
t = plt.text(0.4, yp[k], weight, fontproperties=font,
**alignment)
# Show size options
sizes = ['xx-small', 'x-small', 'small', 'medium', 'large',
'x-large', 'xx-large']
t = plt.text(0.8, 0.9, 'size', fontproperties=font1,
**alignment)
for k, size in enumerate(sizes):
font = font0.copy()
font.set_size(size)
t = plt.text(0.8, yp[k], size, fontproperties=font,
**alignment)
# Show bold italic
font = font0.copy()
font.set_style('italic')
font.set_weight('bold')
font.set_size('x-small')
t = plt.text(-0.4, 0.1, 'bold italic', fontproperties=font,
**alignment)
font = font0.copy()
font.set_style('italic')
font.set_weight('bold')
font.set_size('medium')
t = plt.text(-0.4, 0.2, 'bold italic', fontproperties=font,
**alignment)
font = font0.copy()
font.set_style('italic')
font.set_weight('bold')
font.set_size('x-large')
t = plt.text(-0.4, 0.3, 'bold italic', fontproperties=font,
**alignment)
plt.axis([-1, 1, 0, 1])
plt.show()
matplotlib.font_manager.fontManager.ttflist
etsamples,etevents,etmsgs = preprocess_et(et='pl',subject='VP3')
etmsgs.rms.describe()
etmsgs.sd.describe()
# add labels blink and saccade information from the event df to sample df
etsamples = et_helper.add_events_to_samples(etsamples, etevents)
# get all nan index (not a blink neither a saccade) and pupil has to be detected and no negative time
ix_fix = pd.isnull(etsamples.type) & (etsamples.zero_pa==False) & (etsamples.neg_time==False)
# mark them as fixations
etsamples.loc[ix_fix, 'type'] = 'fixation'
# use magic to get start and end times of fixations in a temporary column
etsamples['tmp_fix'] = ((1*(etsamples['type'] == 'fixation')).diff())
etsamples['tmp_fix'].iloc[0] = 0
etsamples['tmp_fix'] = etsamples['tmp_fix'].astype(int)
# first sample should be fix start?
if etsamples['tmp_fix'][np.argmax(etsamples['tmp_fix'] != 0)] == -1: #argmax stops at first true
# if we only find an fixation end, add a start at the beginning
etsamples.iloc[0, etsamples.columns.get_loc('tmp_fix')] = 1
# make a list of the start and end times
start_times_list = list(etsamples.loc[etsamples['tmp_fix'] == 1, 'smpl_time'].astype(float))
end_times_list = list(etsamples.loc[etsamples['tmp_fix'] == -1, 'smpl_time'].astype(float))
if len(start_times_list) == len(end_times_list)+1:
# drop the last one if not finished
start_times_list = start_times_list[0:-1]
# drop the temporary column
etsamples.drop('tmp_fix', axis=1, inplace=True)
# add them as columns to a fixationevent df
fixationevents = pd.DataFrame([start_times_list, end_times_list], ['start_time', 'end_time']).T
# delete event if start or end is NaN
fixationevents.dropna(subset=['start_time', 'end_time'], inplace=True)
# add the type
fixationevents['type'] = 'fixation'
fixationevents['duration'] = fixationevents['end_time'] - fixationevents['start_time']
# delete fixationevents shorter than 50 ms
logger.warning("Deleted %s fixationsevents of %s fixationsevents in total cause they were shorter than 50ms", np.sum(fixationevents.duration <= 0.05), len(fixationevents))
fixationevents = fixationevents[fixationevents.duration > 0.05]
etsamples, etmsgs,etevents = et_helper.load_file('el','VP3',cleaned=False)
from lib.pupil.pupil_src.shared_modules import file_methods as pl_file_methods
data = pl_file_methods.load_object('/net/store/nbp/projects/IntoTheWild/Daten/Eyetracking/Wild/VP21/2018_05_24/001/offline_data/offline_calibration_gaze')
data
data['circle_marker_positions']
etevents.loc[:,'duration'] = etevents.end_time - etevents.start_time #24 is a 0.0 duratio blink
et_helper.plot_around_event(etsamples,etmsgs,etevents,etevents.query("type=='blink'").iloc[24],plusminus=(-2,5))
plsamples,plevents,plmsgs = et_import.import_pl(subject='VP3',recalib=False,surfaceMap=False)
1
original_pldata = et_import.raw_pl_data(subject)
from functions import pl_surface
folder = '/net/store/nbp/projects/etcomp/VP3/raw'
fake_gpool = pl_surface.fake_gpool_surface(folder)
import functions.add_path
import numpy as np
import time
import os
import av # important to load this library before pupil-library! (even though we dont use it...)
from lib.pupil.pupil_src.shared_modules import offline_surface_tracker
tracker = offline_surface_tracker.Offline_Surface_Tracker(fake_gpool,min_marker_perimeter=30,robust_detection=False)
from offline_reference_surface import Offline_Reference_Surface
surface = Offline_Reference_Surface(tracker.g_pool)
# First define the markers that should be used for the surface
# find a frame where there are 16 markers and all of them have high confidence
ix = 0
while True:
if len(tracker.cache[ix]) == 16:
usable_markers = [m for m in tracker.cache[ix] if m['id_confidence'] >= 0.8]
if len(usable_markers) == 16:
break
ix +=1
# Step 3
# This dissables pupil-labs functionality. They ask for 90 frames with the markers. but because we know there will be 16 markers, we dont need it (toitoitoi)
print('Defining & Finding Surface')
surface.required_build_up = 1
surface.build_correspondence(tracker.cache[ix],0.3,0.7)
if not surface.defined:
raise('Oh oh trouble ahead. The surface was not defined')
type(tracker.cache)
surface._get_location(tracker.cache[0],0.3,0.7)
surface.init_cache(tracker.cache,0.3,0.7)
# Step 4
tracker.surfaces = [surface];
tracker
from functions import pl_surface
datapath = '/net/store/nbp/projects/etcomp/'
subject = 'VP3'
folder= os.path.join(datapath,subject,'raw')
tracker = pl_surface.map_surface(folder,loadSurface=False,loadCache=True)
tracker
tracker = pl_surface.map_surface(folder,loadSurface=True)
# use pupilhelper func to make samples df (confidence, gx, gy, smpl_time, diameter)
pldata = et_helper.gaze_to_pandas(original_pldata['gaze_positions'])
p_unproj = intrinsics.unprojectPoints(p,use_distortion=True)
p_proj = intrinsics.projectPoints(p_unproj,use_distortion=True)
p_proj_nodist = intrinsics.projectPoints(p_unproj,use_distortion=False)
plt.figure()
#plt.plot(p[:,0],p[:,1],'go')
plt.plot(p_unproj[:,0],p_unproj[:,1],'ro')
#plt.plot(p_proj[:,0],p_proj[:,1],'bo')
#plt.plot(p_proj_nodist[:,0],p_proj_nodist[:,1],'ko')
# undistort gaze postitions
pldata.loc[:,'gx_px'] = pldata.loc[:,'gx']*(2/3.)*1920 + (1/6.*1920)
pldata.loc[:,'gy_px'] = pldata.loc[:,'gy']*(2/3.)*1080 + (1/6.*1080)
undistorted_gazepoint_array = intrinsics.unprojectPoints(pldata.loc[:, ['gx_px','gy_px']].values, use_distortion=True,normalize=False)
undistorted_gazepoint_array = intrinsics.projectPoints(undistorted_gazepoint_array, use_distortion=False)
pldata.loc[:,'gx_new'] = (undistorted_gazepoint_array[:,0]-(1/6.*1920)) /((2/3.)*1080)
pldata.loc[:,'gy_new'] = (undistorted_gazepoint_array[:,1]-(1/6.*1080)) /((2/3.)*1080)
plt.figure()
plt.plot(undistorted_gazepoint_array[1:10000,1])
ggplot(pldata.iloc[8000:8400])+geom_point(aes(x='gx',y='gy'),color='black')+geom_point(aes(x='gx_new',y='gy_new'),color='red')
import lib.pupil.pupil_src.shared_modules.file_methods as file_methods
file_methods.Incremental_Legacy_Pupil_Data_Loader('test')
```
## Testing many saccades in HMM
```
from functions.detect_events import make_blinks,make_saccades,make_fixations
from functions.detect_events_hmm import detect_events_hmm,detect_events_hmm_nosmooth
import functions.et_preprocess as preprocess
etsamples, etmsgs, etevents = preprocess.preprocess_et('pl','VP11',load=False,save=False,eventfunctions=(make_blinks,detect_events_hmm_nosmooth),outputprefix='hmmnosmooth_')
etsamples, etmsgs, etevents = et_helper.load_file('el','VP3',cleaned=True)
import numpy as np
##################################
########### Parameters ###########
sampfreq = 500 #Hz
weights_name = os.path.abspath('../local/build/src_uneye/training/weights_Andersson')
min_sacc_dur = 6 # in ms
min_sacc_dist = 10 #in ms
# load data
Xtest = etsamples.gx.iloc[30000:32000]
Ytest = etsamples.gy.iloc[30000:32000]
import uneye
# Prediction
et_helper.tic()
model = uneye.DNN(weights_name=weights_name,
sampfreq=sampfreq,
min_sacc_dur=min_sacc_dur,
min_sacc_dist=min_sacc_dist,classes=5)
et_helper.toc()
Prediction,Probability = model.predict(Xtest,Ytest)
et_helper.toc()
import matplotlib.pyplot as plt
# plot example
fig = plt.figure(figsize=(8,4))
plt.suptitle('Example Trial')
ax = fig.add_subplot(611)
x_trace = Xtest
y_trace = Ytest
plt.plot(x_trace,label='X position',c=[0,0.5,0.8])
plt.plot(y_trace,label='Y position',c=[0,0.8,0.3])
plt.xticks(())
plt.ylabel('Relative eye position (°)')
plt.legend()
labellist = ['fixation','saccade','pso','blink','other']
# prediction
for k in range(5):
ax = fig.add_subplot(6,1,k+2)
#plt.plot(Prediction,label='Binary prediction',c=[0,0.6,0.3])
plt.plot(Probability[k,:],c='k',alpha=.3)
plt.yticks((0,1),['',labellist[k]])
plt.show()
```
| github_jupyter |
# Custom derivative rules for JAX-transformable Python functions
*mattjj@ Mar 19 2020, last updated Oct 14 2020*
There are two ways to define differentiation rules in JAX:
1. using `jax.custom_jvp` and `jax.custom_vjp` to define custom differentiation rules for Python functions that are already JAX-transformable; and
2. defining new `core.Primitive` instances along with all their transformation rules, for example to call into functions from other systems like solvers, simulators, or general numerical computing systems.
This notebook is about #1. To read instead about #2, see the [notebook on adding primitives](https://jax.readthedocs.io/en/latest/notebooks/How_JAX_primitives_work.html).
For an introduction to JAX's automatic differentiation API, see [The Autodiff Cookbook](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html). This notebook assumes some familiarity with [jax.jvp](https://jax.readthedocs.io/en/latest/jax.html#jax.jvp) and [jax.grad](https://jax.readthedocs.io/en/latest/jax.html#jax.grad), and the mathematical meaning of JVPs and VJPs.
## TL;DR
### Custom JVPs with `jax.custom_jvp`
```
import jax.numpy as jnp
from jax import custom_jvp
@custom_jvp
def f(x, y):
return jnp.sin(x) * y
@f.defjvp
def f_jvp(primals, tangents):
x, y = primals
x_dot, y_dot = tangents
primal_out = f(x, y)
tangent_out = jnp.cos(x) * x_dot * y + jnp.sin(x) * y_dot
return primal_out, tangent_out
from jax import jvp, grad
print(f(2., 3.))
y, y_dot = jvp(f, (2., 3.), (1., 0.))
print(y)
print(y_dot)
print(grad(f)(2., 3.))
# Equivalent alternative using the defjvps convenience wrapper
@custom_jvp
def f(x, y):
return jnp.sin(x) * y
f.defjvps(lambda x_dot, primal_out, x, y: jnp.cos(x) * x_dot * y,
lambda y_dot, primal_out, x, y: jnp.sin(x) * y_dot)
print(f(2., 3.))
y, y_dot = jvp(f, (2., 3.), (1., 0.))
print(y)
print(y_dot)
print(grad(f)(2., 3.))
```
### Custom VJPs with `jax.custom_vjp`
```
from jax import custom_vjp
@custom_vjp
def f(x, y):
return jnp.sin(x) * y
def f_fwd(x, y):
# Returns primal output and residuals to be used in backward pass by f_bwd.
return f(x, y), (jnp.cos(x), jnp.sin(x), y)
def f_bwd(res, g):
cos_x, sin_x, y = res # Gets residuals computed in f_fwd
return (cos_x * g * y, sin_x * g)
f.defvjp(f_fwd, f_bwd)
print(grad(f)(2., 3.))
```
## Example problems
To get an idea of what problems `jax.custom_jvp` and `jax.custom_vjp` are meant to solve, let's go over a few examples. A more thorough introduction to the `jax.custom_jvp` and `jax.custom_vjp` APIs is in [the next section](#scrollTo=Dr0aNkBslfQf).
### Numerical stability
One application of `jax.custom_jvp` is to improve the numerical stability of differentiation.
Say we want to write a function called `log1pexp`, which computes $x \mapsto \log ( 1 + e^x )$. We can write that using `jax.numpy`:
```
import jax.numpy as jnp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
log1pexp(3.)
```
Since it's written in terms of `jax.numpy`, it's JAX-transformable:
```
from jax import jit, grad, vmap
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
But there's a numerical stability problem lurking here:
```
print(grad(log1pexp)(100.))
```
That doesn't seem right! After all, the derivative of $x \mapsto \log (1 + e^x)$ is $x \mapsto \frac{e^x}{1 + e^x}$, and so for large values of $x$ we'd expect the value to be about 1.
We can get a bit more insight into what's going on by looking at the jaxpr for the gradient computation:
```
from jax import make_jaxpr
make_jaxpr(grad(log1pexp))(100.)
```
Stepping through how the jaxpr would be evaluated, we can see that the last line would involve multiplying values that floating point math will round to 0 and $\infty$, respectively, which is never a good idea. That is, we're effectively evaluating `lambda x: (1 / (1 + jnp.exp(x))) * jnp.exp(x)` for large `x`, which effectively turns into `0. * jnp.inf`.
Instead of generating such large and small values, hoping for a cancellation that floats can't always provide, we'd rather just express the derivative function as a more numerically stable program. In particular, we can write a program that more closely evaluates the equal mathematical expression $1 - \frac{1}{1 + e^x}$, with no cancellation in sight.
This problem is interesting because even though our definition of `log1pexp` could already be JAX-differentiated (and transformed with `jit`, `vmap`, ...), we're not happy with the result of applying standard autodiff rules to the primitives comprising `log1pexp` and composing the result. Instead, we'd like to specify how the whole function `log1pexp` should be differentiated, as a unit, and thus arrange those exponentials better.
This is one application of custom derivative rules for Python functions that are already JAX transformable: specifying how a composite function should be differentiated, while still using its original Python definition for other transformations (like `jit`, `vmap`, ...).
Here's a solution using `jax.custom_jvp`:
```
from jax import custom_jvp
@custom_jvp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
@log1pexp.defjvp
def log1pexp_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = log1pexp(x)
ans_dot = (1 - 1/(1 + jnp.exp(x))) * x_dot
return ans, ans_dot
print(grad(log1pexp)(100.))
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
Here's a `defjvps` convenience wrapper to express the same thing:
```
@custom_jvp
def log1pexp(x):
return jnp.log(1. + jnp.exp(x))
log1pexp.defjvps(lambda t, ans, x: (1 - 1/(1 + jnp.exp(x))) * t)
print(grad(log1pexp)(100.))
print(jit(log1pexp)(3.))
print(jit(grad(log1pexp))(3.))
print(vmap(jit(grad(log1pexp)))(jnp.arange(3.)))
```
### Enforcing a differentiation convention
A related application is to enforce a differentiation convention, perhaps at a boundary.
Consider the function $f : \mathbb{R}_+ \mapsto \mathbb{R}_+$ with $f(x) = \frac{x}{1 + \sqrt{x}}$, where we take $\mathbb{R}_+ = [0, \infty)$. We might implement $f$ as a program like this:
```
def f(x):
return x / (1 + jnp.sqrt(x))
```
As a mathematical function on $\mathbb{R}$ (the full real line), $f$ is not differentiable at zero (because the limit defining the derivative doesn't exist from the left). Correspondingly, autodiff produces a `nan` value:
```
print(grad(f)(0.))
```
But mathematically if we think of $f$ as a function on $\mathbb{R}_+$ then it is differentiable at 0 [Rudin's Principles of Mathematical Analysis Definition 5.1, or Tao's Analysis I 3rd ed. Definition 10.1.1 and Example 10.1.6]. Alternatively, we might say as a convention we want to consider the directional derivative from the right. So there is a sensible value for the Python function `grad(f)` to return at `0.0`, namely `1.0`. By default, JAX's machinery for differentiation assumes all functions are defined over $\mathbb{R}$ and thus doesn't produce `1.0` here.
We can use a custom JVP rule! In particular, we can define the JVP rule in terms of the derivative function $x \mapsto \frac{\sqrt{x} + 2}{2(\sqrt{x} + 1)^2}$ on $\mathbb{R}_+$,
```
@custom_jvp
def f(x):
return x / (1 + jnp.sqrt(x))
@f.defjvp
def f_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = f(x)
ans_dot = ((jnp.sqrt(x) + 2) / (2 * (jnp.sqrt(x) + 1)**2)) * x_dot
return ans, ans_dot
print(grad(f)(0.))
```
Here's the convenience wrapper version:
```
@custom_jvp
def f(x):
return x / (1 + jnp.sqrt(x))
f.defjvps(lambda t, ans, x: ((jnp.sqrt(x) + 2) / (2 * (jnp.sqrt(x) + 1)**2)) * t)
print(grad(f)(0.))
```
### Gradient clipping
While in some cases we want to express a mathematical differentiation computation, in other cases we may even want to take a step away from mathematics to adjust the computation autodiff performs. One canonical example is reverse-mode gradient clipping.
For gradient clipping, we can use `jnp.clip` together with a `jax.custom_vjp` reverse-mode-only rule:
```
from functools import partial
from jax import custom_vjp
@custom_vjp
def clip_gradient(lo, hi, x):
return x # identity function
def clip_gradient_fwd(lo, hi, x):
return x, (lo, hi) # save bounds as residuals
def clip_gradient_bwd(res, g):
lo, hi = res
return (None, None, jnp.clip(g, lo, hi)) # use None to indicate zero cotangents for lo and hi
clip_gradient.defvjp(clip_gradient_fwd, clip_gradient_bwd)
import matplotlib.pyplot as plt
from jax import vmap
t = jnp.linspace(0, 10, 1000)
plt.plot(jnp.sin(t))
plt.plot(vmap(grad(jnp.sin))(t))
def clip_sin(x):
x = clip_gradient(-0.75, 0.75, x)
return jnp.sin(x)
plt.plot(clip_sin(t))
plt.plot(vmap(grad(clip_sin))(t))
```
### Python debugging
Another application that is motivated by development workflow rather than numerics is to set a `pdb` debugger trace in the backward pass of reverse-mode autodiff.
When trying to track down the source of a `nan` runtime error, or just examine carefully the cotangent (gradient) values being propagated, it can be useful to insert a debugger at a point in the backward pass that corresponds to a specific point in the primal computation. You can do that with `jax.custom_vjp`.
We'll defer an example until the next section.
### Implicit function differentiation of iterative implementations
This example gets pretty deep in the mathematical weeds!
Another application for `jax.custom_vjp` is reverse-mode differentiation of functions that are JAX-transformable (by `jit`, `vmap`, ...) but not efficiently JAX-differentiable for some reason, perhaps because they involve `lax.while_loop`. (It's not possible to produce an XLA HLO program that efficiently computes the reverse-mode derivative of an XLA HLO While loop because that would require a program with unbounded memory use, which isn't possible to express in XLA HLO, at least without side-effecting interactions through infeed/outfeed.)
For example, consider this `fixed_point` routine which computes a fixed point by iteratively applying a function in a `while_loop`:
```
from jax.lax import while_loop
def fixed_point(f, a, x_guess):
def cond_fun(carry):
x_prev, x = carry
return jnp.abs(x_prev - x) > 1e-6
def body_fun(carry):
_, x = carry
return x, f(a, x)
_, x_star = while_loop(cond_fun, body_fun, (x_guess, f(a, x_guess)))
return x_star
```
This is an iterative procedure for numerically solving the equation $x = f(a, x)$ for $x$, by iterating $x_{t+1} = f(a, x_t)$ until $x_{t+1}$ is sufficiently close to $x_t$. The result $x^*$ depends on the parameters $a$, and so we can think of there being a function $a \mapsto x^*(a)$ that is implicity defined by equation $x = f(a, x)$.
We can use `fixed_point` to run iterative procedures to convergence, for example running Newton's method to calculate square roots while only executing adds, multiplies, and divides:
```
def newton_sqrt(a):
update = lambda a, x: 0.5 * (x + a / x)
return fixed_point(update, a, a)
print(newton_sqrt(2.))
```
We can `vmap` or `jit` the function as well:
```
print(jit(vmap(newton_sqrt))(jnp.array([1., 2., 3., 4.])))
```
We can't apply reverse-mode automatic differentiation because of the `while_loop`, but it turns out we wouldn't want to anyway: instead of differentiating through the implementation of `fixed_point` and all its iterations, we can exploit the mathematical structure to do something that is much more memory-efficient (and FLOP-efficient in this case, too!). We can instead use the implicit function theorem [Prop A.25 of Bertsekas's Nonlinear Programming, 2nd ed.], which guarantees (under some conditions) the existence of the mathematical objects we're about to use. In essence, we linearize at the solution and solve those linear equations iteratively to compute the derivatives we want.
Consider again the equation $x = f(a, x)$ and the function $x^*$. We want to evaluate vector-Jacobian products like $v^\mathsf{T} \mapsto v^\mathsf{T} \partial x^*(a_0)$.
At least in an open neighborhood around the point $a_0$ at which we want to differentiate, let's assume that the equation $x^*(a) = f(a, x^*(a))$ holds for all $a$. Since the two sides are equal as functions of $a$, their derivatives must be equal as well, so let's differentiate both sides:
$\qquad \partial x^*(a) = \partial_0 f(a, x^*(a)) + \partial_1 f(a, x^*(a)) \partial x^*(a)$.
Setting $A = \partial_1 f(a_0, x^*(a_0))$ and $B = \partial_0 f(a_0, x^*(a_0))$, we can write the quantity we're after more simply as
$\qquad \partial x^*(a_0) = B + A \partial x^*(a_0)$,
or, by rearranging,
$\qquad \partial x^*(a_0) = (I - A)^{-1} B$.
That means we can evaluate vector-Jacobian products like
$\qquad v^\mathsf{T} \partial x^*(a_0) = v^\mathsf{T} (I - A)^{-1} B = w^\mathsf{T} B$,
where $w^\mathsf{T} = v^\mathsf{T} (I - A)^{-1}$, or equivalently $w^\mathsf{T} = v^\mathsf{T} + w^\mathsf{T} A$, or equivalently $w^\mathsf{T}$ is the fixed point of the map $u^\mathsf{T} \mapsto v^\mathsf{T} + u^\mathsf{T} A$. That last characterization gives us a way to write the VJP for `fixed_point` in terms of a call to `fixed_point`! Moreover, after expanding $A$ and $B$ back out, we can see we need only to evaluate VJPs of $f$ at $(a_0, x^*(a_0))$.
Here's the upshot:
```
from jax import vjp
@partial(custom_vjp, nondiff_argnums=(0,))
def fixed_point(f, a, x_guess):
def cond_fun(carry):
x_prev, x = carry
return jnp.abs(x_prev - x) > 1e-6
def body_fun(carry):
_, x = carry
return x, f(a, x)
_, x_star = while_loop(cond_fun, body_fun, (x_guess, f(a, x_guess)))
return x_star
def fixed_point_fwd(f, a, x_init):
x_star = fixed_point(f, a, x_init)
return x_star, (a, x_star)
def fixed_point_rev(f, res, x_star_bar):
a, x_star = res
_, vjp_a = vjp(lambda a: f(a, x_star), a)
a_bar, = vjp_a(fixed_point(partial(rev_iter, f),
(a, x_star, x_star_bar),
x_star_bar))
return a_bar, jnp.zeros_like(x_star)
def rev_iter(f, packed, u):
a, x_star, x_star_bar = packed
_, vjp_x = vjp(lambda x: f(a, x), x_star)
return x_star_bar + vjp_x(u)[0]
fixed_point.defvjp(fixed_point_fwd, fixed_point_rev)
print(newton_sqrt(2.))
print(grad(newton_sqrt)(2.))
print(grad(grad(newton_sqrt))(2.))
```
We can check our answers by differentiating `jnp.sqrt`, which uses a totally different implementation:
```
print(grad(jnp.sqrt)(2.))
print(grad(grad(jnp.sqrt))(2.))
```
A limitation to this approach is that the argument `f` can't close over any values involved in differentiation. That is, you might notice that we kept the parameter `a` explicit in the argument list of `fixed_point`. While other JAX mechanisms can handle closed-over transformation-traced values in the arguments to higher-order functions (as is done for the control flow primitives like `lax.cond`, `lax.scan`, and `lax.while_loop` itself), `jax.custom_vjp` used as above cannot. A `fixed_point` routine that used a bit more of JAX's internals could have a more convenient and robust API.
## Basic usage of `jax.custom_jvp` and `jax.custom_vjp` APIs
### Use `jax.custom_jvp` to define forward-mode (and, indirectly, reverse-mode) rules
Here's a canonical basic example of using `jax.custom_jvp`:
```
from jax import custom_jvp
import jax.numpy as jnp
# f :: a -> b
@custom_jvp
def f(x):
return jnp.sin(x)
# f_jvp :: (a, T a) -> (b, T b)
def f_jvp(primals, tangents):
x, = primals
t, = tangents
return f(x), jnp.cos(x) * t
f.defjvp(f_jvp)
from jax import jvp
print(f(3.))
y, y_dot = jvp(f, (3.,), (1.,))
print(y)
print(y_dot)
```
In words, we start with a a primal function `f` that takes inputs of type `a` and produces outputs of type `b`. We associate with it a JVP rule function `f_jvp` that takes a pair of inputs representing the primal inputs of type `a` and the corresponding tangent inputs of type `T a`, and produces a pair of outputs representing the primal outputs of type `b` and tangent outputs of type `T b`. The tangent outputs should be a linear function of the tangent inputs.
You can also use `f.defjvp` as a decorator, as in
```python
@custom_jvp
def f(x):
...
@f.defjvp
def f_jvp(primals, tangents):
...
```
Even though we defined only a JVP rule and no VJP rule, we can use both forward- and reverse-mode differentiation on `f`. JAX will automatically transpose the linear computation on tangent values from our custom JVP rule, computing the VJP as efficiently as if we had written the rule by hand:
```
from jax import grad
print(grad(f)(3.))
print(grad(grad(f))(3.))
```
For automatic transposition to work, the JVP rule's output tangents must be linear as a function of the input tangents. Otherwise a transposition error is raised.
Multiple arguments work like this:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
@f.defjvp
def f_jvp(primals, tangents):
x, y = primals
x_dot, y_dot = tangents
primal_out = f(x, y)
tangent_out = 2 * x * y * x_dot + x ** 2 * y_dot
return primal_out, tangent_out
print(grad(f)(2., 3.))
```
The `defjvps` convenience wrapper lets us define a JVP for each argument separately, and the results are computed separately then summed:
```
@custom_jvp
def f(x):
return jnp.sin(x)
f.defjvps(lambda t, ans, x: jnp.cos(x) * t)
print(grad(f)(3.))
```
Here's a `defjvps` example with multiple arguments:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
f.defjvps(lambda x_dot, primal_out, x, y: 2 * x * y * x_dot,
lambda y_dot, primal_out, x, y: x ** 2 * y_dot)
print(grad(f)(2., 3.))
print(grad(f, 0)(2., 3.)) # same as above
print(grad(f, 1)(2., 3.))
```
As a shorthand, with `defjvps` you can pass a `None` value to indicate that the JVP for a particular argument is zero:
```
@custom_jvp
def f(x, y):
return x ** 2 * y
f.defjvps(lambda x_dot, primal_out, x, y: 2 * x * y * x_dot,
None)
print(grad(f)(2., 3.))
print(grad(f, 0)(2., 3.)) # same as above
print(grad(f, 1)(2., 3.))
```
Calling a `jax.custom_jvp` function with keyword arguments, or writing a `jax.custom_jvp` function definition with default arguments, are both allowed so long as they can be unambiguosly mapped to positional arguments based on the function signature retrieved by the standard library `inspect.signature` mechanism.
When you're not performing differentiation, the function `f` is called just as if it weren't decorated by `jax.custom_jvp`:
```
@custom_jvp
def f(x):
print('called f!') # a harmless side-effect
return jnp.sin(x)
@f.defjvp
def f_jvp(primals, tangents):
print('called f_jvp!') # a harmless side-effect
x, = primals
t, = tangents
return f(x), jnp.cos(x) * t
from jax import vmap, jit
print(f(3.))
print(vmap(f)(jnp.arange(3.)))
print(jit(f)(3.))
```
The custom JVP rule is invoked during differentiation, whether forward or reverse:
```
y, y_dot = jvp(f, (3.,), (1.,))
print(y_dot)
print(grad(f)(3.))
```
Notice that `f_jvp` calls `f` to compute the primal outputs. In the context of higher-order differentiation, each application of a differentiation transform will use the custom JVP rule if and only if the rule calls the original `f` to compute the primal outputs. (This represents a kind of fundamental tradeoff, where we can't make use of intermediate values from the evaluation of `f` in our rule _and also_ have the rule apply in all orders of higher-order differentiation.)
```
grad(grad(f))(3.)
```
You can use Python control flow with `jax.custom_jvp`:
```
@custom_jvp
def f(x):
if x > 0:
return jnp.sin(x)
else:
return jnp.cos(x)
@f.defjvp
def f_jvp(primals, tangents):
x, = primals
x_dot, = tangents
ans = f(x)
if x > 0:
return ans, 2 * x_dot
else:
return ans, 3 * x_dot
print(grad(f)(1.))
print(grad(f)(-1.))
```
### Use `jax.custom_vjp` to define custom reverse-mode-only rules
While `jax.custom_jvp` suffices for controlling both forward- and, via JAX's automatic transposition, reverse-mode differentiation behavior, in some cases we may want to directly control a VJP rule, for example in the latter two example problems presented above. We can do that with `jax.custom_vjp`:
```
from jax import custom_vjp
import jax.numpy as jnp
# f :: a -> b
@custom_vjp
def f(x):
return jnp.sin(x)
# f_fwd :: a -> (b, c)
def f_fwd(x):
return f(x), jnp.cos(x)
# f_bwd :: (c, CT b) -> CT a
def f_bwd(cos_x, y_bar):
return (cos_x * y_bar,)
f.defvjp(f_fwd, f_bwd)
from jax import grad
print(f(3.))
print(grad(f)(3.))
```
In words, we again start with a a primal function `f` that takes inputs of type `a` and produces outputs of type `b`. We associate with it two functions, `f_fwd` and `f_bwd`, which describe how to perform the forward- and backward-passes of reverse-mode autodiff, respectively.
The function `f_fwd` describes the forward pass, not only the primal computation but also what values to save for use on the backward pass. Its input signature is just like that of the primal function `f`, in that it takes a primal input of type `a`. But as output it produces a pair, where the first element is the primal output `b` and the second element is any "residual" data of type `c` to be stored for use by the backward pass. (This second output is analogous to [PyTorch's save_for_backward mechanism](https://pytorch.org/tutorials/beginner/examples_autograd/two_layer_net_custom_function.html).)
The function `f_bwd` describes the backward pass. It takes two inputs, where the first is the residual data of type `c` produced by `f_fwd` and the second is the output cotangents of type `CT b` corresponding to the output of the primal function. It produces an output of type `CT a` representing the cotangents corresponding to the input of the primal function. In particular, the output of `f_bwd` must be a sequence (e.g. a tuple) of length equal to the number of arguments to the primal function.
So multiple arguments work like this:
```
from jax import custom_vjp
@custom_vjp
def f(x, y):
return jnp.sin(x) * y
def f_fwd(x, y):
return f(x, y), (jnp.cos(x), jnp.sin(x), y)
def f_bwd(res, g):
cos_x, sin_x, y = res
return (cos_x * g * y, -sin_x * g)
f.defvjp(f_fwd, f_bwd)
print(grad(f)(2., 3.))
```
Calling a `jax.custom_vjp` function with keyword arguments, or writing a `jax.custom_vjp` function definition with default arguments, are both allowed so long as they can be unambiguosly mapped to positional arguments based on the function signature retrieved by the standard library `inspect.signature` mechanism.
As with `jax.custom_jvp`, the custom VJP rule comprised by `f_fwd` and `f_bwd` is not invoked if differentiation is not applied. If function is evaluated, or transformed with `jit`, `vmap`, or other non-differentiation transformations, then only `f` is called.
```
@custom_vjp
def f(x):
print("called f!")
return jnp.sin(x)
def f_fwd(x):
print("called f_fwd!")
return f(x), jnp.cos(x)
def f_bwd(cos_x, y_bar):
print("called f_bwd!")
return (cos_x * y_bar,)
f.defvjp(f_fwd, f_bwd)
print(f(3.))
print(grad(f)(3.))
from jax import vjp
y, f_vjp = vjp(f, 3.)
print(y)
print(f_vjp(1.))
```
**Forward-mode autodiff cannot be used on the** `jax.custom_vjp` **function** and will raise an error:
```
from jax import jvp
try:
jvp(f, (3.,), (1.,))
except TypeError as e:
print('ERROR! {}'.format(e))
```
If you want to use both forward- and reverse-mode, use `jax.custom_jvp` instead.
We can use `jax.custom_vjp` together with `pdb` to insert a debugger trace in the backward pass:
```
import pdb
@custom_vjp
def debug(x):
return x # acts like identity
def debug_fwd(x):
return x, x
def debug_bwd(x, g):
import pdb; pdb.set_trace()
return g
debug.defvjp(debug_fwd, debug_bwd)
def foo(x):
y = x ** 2
y = debug(y) # insert pdb in corresponding backward pass step
return jnp.sin(y)
```
```python
jax.grad(foo)(3.)
> <ipython-input-113-b19a2dc1abf7>(12)debug_bwd()
-> return g
(Pdb) p x
DeviceArray(9., dtype=float32)
(Pdb) p g
DeviceArray(-0.91113025, dtype=float32)
(Pdb) q
```
## More features and details
### Working with `list` / `tuple` / `dict` containers (and other pytrees)
You should expect standard Python containers like lists, tuples, namedtuples, and dicts to just work, along with nested versions of those. In general, any [pytrees](https://github.com/google/jax/blob/master/docs/notebooks/JAX_pytrees.ipynb) are permissible, so long as their structures are consistent according to the type constraints.
Here's a contrived example with `jax.custom_jvp`:
```
from collections import namedtuple
Point = namedtuple("Point", ["x", "y"])
@custom_jvp
def f(pt):
x, y = pt.x, pt.y
return {'a': x ** 2,
'b': (jnp.sin(x), jnp.cos(y))}
@f.defjvp
def f_jvp(primals, tangents):
pt, = primals
pt_dot, = tangents
ans = f(pt)
ans_dot = {'a': 2 * pt.x * pt_dot.x,
'b': (jnp.cos(pt.x) * pt_dot.x, -jnp.sin(pt.y) * pt_dot.y)}
return ans, ans_dot
def fun(pt):
dct = f(pt)
return dct['a'] + dct['b'][0]
pt = Point(1., 2.)
print(f(pt))
print(grad(fun)(pt))
```
And an analogous contrived example with `jax.custom_vjp`:
```
@custom_vjp
def f(pt):
x, y = pt.x, pt.y
return {'a': x ** 2,
'b': (jnp.sin(x), jnp.cos(y))}
def f_fwd(pt):
return f(pt), pt
def f_bwd(pt, g):
a_bar, (b0_bar, b1_bar) = g['a'], g['b']
x_bar = 2 * pt.x * a_bar + jnp.cos(pt.x) * b0_bar
y_bar = -jnp.sin(pt.y) * b1_bar
return (Point(x_bar, y_bar),)
f.defvjp(f_fwd, f_bwd)
def fun(pt):
dct = f(pt)
return dct['a'] + dct['b'][0]
pt = Point(1., 2.)
print(f(pt))
print(grad(fun)(pt))
```
### Handling non-differentiable arguments
Some use cases, like the final example problem, call for non-differentiable arguments like function-valued arguments to be passed to functions with custom differentiation rules, and for those arguments to also be passed to the rules themselves. In the case of `fixed_point`, the function argument `f` was such a non-differentiable argument. A similar situation arises with `jax.experimental.odeint`.
#### `jax.custom_jvp` with `nondiff_argnums`
Use the optional `nondiff_argnums` parameter to `jax.custom_jvp` to indicate arguments like these. Here's an example with `jax.custom_jvp`:
```
from functools import partial
@partial(custom_jvp, nondiff_argnums=(0,))
def app(f, x):
return f(x)
@app.defjvp
def app_jvp(f, primals, tangents):
x, = primals
x_dot, = tangents
return f(x), 2. * x_dot
print(app(lambda x: x ** 3, 3.))
print(grad(app, 1)(lambda x: x ** 3, 3.))
```
Notice the gotcha here: no matter where in the argument list these parameters appear, they're placed at the *start* of the signature of the corresponding JVP rule. Here's another example:
```
@partial(custom_jvp, nondiff_argnums=(0, 2))
def app2(f, x, g):
return f(g((x)))
@app2.defjvp
def app2_jvp(f, g, primals, tangents):
x, = primals
x_dot, = tangents
return f(g(x)), 3. * x_dot
print(app2(lambda x: x ** 3, 3., lambda y: 5 * y))
print(grad(app2, 1)(lambda x: x ** 3, 3., lambda y: 5 * y))
```
#### `jax.custom_vjp` with `nondiff_argnums`
A similar option exists for `jax.custom_vjp`, and similarly the convention is that the non-differentiable arguments are passed as the first arguments to the rules, no matter where they appear in the original function's signature. Here's an example:
```
@partial(custom_vjp, nondiff_argnums=(0,))
def app(f, x):
return f(x)
def app_fwd(f, x):
return f(x), x
def app_bwd(f, x, g):
return (5 * g,)
app.defvjp(app_fwd, app_bwd)
print(app(lambda x: x ** 2, 4.))
print(grad(app, 1)(lambda x: x ** 2, 4.))
```
See `fixed_point` above for another usage example.
**You don't need to use** `nondiff_argnums` **with array-valued arguments**, for example ones with integer dtype. Instead, `nondiff_argnums` should only be used for argument values that don't correspond to JAX types (essentially don't correspond to array types), like Python callables or strings. If JAX detects that an argument indicated by `nondiff_argnums` contains a JAX Tracer, then an error is raised. The `clip_gradient` function above is a good example of not using `nondiff_argnums` for integer-dtype array arguments.
| github_jupyter |
# Introduction to statistical mechanics of fluids
## Classical parition function of fluids
- For a homogenous fluid particles
$$Z(\beta, N, V) = \frac{1}{N! h^N}\int e^{-\beta H(p,x)}dp^N dx^N$$
- For a system composed of three types A B and C for instance the partion function would be:
$$Z(\beta, N, V) = \frac{1}{N_A! N_B! N_C! h^N}\int e^{-\beta H(p,x)}dp^N dx^N$$
With $N=N_A+N_B+N_C$
## General exprssion of probability distribution of fluids in phase space
- The probability of a state in a classical fluid system is $f(x^N, p^N)$
$$\boxed{f(x^N, p^N) = \frac{e^{-\beta H(x^N p^N)}}{Z}}$$
### Classical hamitlonian breaks into functions of momenta and positions respectively
- $$H(x^N, p^N) = K(p^N) + U(x^N)$$
<br><br>
- $$f(x^N, p^N) = \Phi(p^N) P(r^N)$$
<br><br>
- $$\boxed{\Phi(p^N) = \frac{e^{-\beta K(p^N)}}{\int dp^N e^{-\beta K(p^N)}}}$$
<br><br>
- $$\boxed{P(x^N) = \frac{e^{-\beta U(x^N)}}{\int dp^N e^{-\beta U(x^N)}}}$$
<br><br>
#### Maxwell-Boltzman distirbution for single particles
Momentum distribution further factorizes into sum of single particle kinetic energies:
- $$K(p^N) = \sum^N_{i=1} \frac{p^2_i}{2m_i}$$
<br><br>
- $$\Phi(p^N) = \prod^N_{i=1} \phi(p_i)$$
<br><br>
- $$\boxed{\phi(p_i) = \frac{e^{-\beta \frac{p^2_i}{2m}}}{\int dp e^{-\beta \frac{p^2}{2m}}}}$$
Note that $p^2_i = p^2_x+p^2_y+p^2_z$ and $\phi(p_i)$ is what is known as **Maxwell-Boltzmann distribution**. The classical system obyes the same distribution reglardles of whivh phase it is (solid, liquid, gas).
### Configurational partion function
$$Z(\beta, V, N) = Z_{p} \cdot Z_x = \frac{1}{\lambda^3N} Q(\beta, V, N) $$
$$Q = \int dx^N e^{-\beta U(x^N)}\,\, , \,\,\, F_{conf} = -{\beta}^{-1} log Q$$
Pressue is related to Free energy as
- $$p= - \frac{\partial F}{\partial V} = \frac{\partial }{\partial V} log \int dx^N e^{-\beta U(x^N)}$$
**Volume dependence of partition function is in the integration limits!** As volume grows so does partition function. Therefore p is always positive. We can thus conclude that **in equilibirum pressure is always a positive quantity**
### Thermal wavelength and broad validity of classical picture of fluids
$$Z = \frac{1}{N!} \frac{1}{\lambda_T^{3N}} \int dx^N e^{-\beta U(x^N)}$$
**Classical mechanics is valid as long as De Broglie wavelength is small compared to intermolecular length scales**
$$\lambda_{DB} = \frac{h}{p} \sim \frac{h}{\langle p \rangle} = \frac{h}{(8k_B T m/\pi)^{1/2}} \approx \lambda_T$$
For a dilute gas, relevant lengths are $\rho^{-1/3}$ (typical distance) and $\sigma$ (diamater of particles)
- $\lambda_T < \sigma $ (classical regime is good)
#### $H_2O$ vs $D_2O$ melting and freezing points
- Classical descirption predicts configurational parition function to be independent of paritcle masses. Hence equation of state $p = p(\beta, \rho)$ is independent of masses.
<br><br>
- Thus if translationa, rotational and virbational degrees of freedo are well described by classical mechanics, $H_2O$ and $D_2O$ will have the same equation of state. Experiments however predict desnity maximum $\rho(T)$ at $p=1 atm$ at 4 C for $H_2O$ and 10C in $D_2O$. The freezing point is higher for $D_2 O$ as well
<br><br>
- Water is a weird fluid. Size is about $3 A$ and thermal wavelength is around $0.3 A$. The role of quantum fluctuations is to blurr the positions of water molecules. As the atomic mass increases, bluring goes down and fluids gets ordered. This is why $D_2 O$ ice melts at a higher temperature, e.g need to supply more energy to disrupt the order.
### Reduced configruational distribution functions
- **Hard fact:** $P(r^N)$ and Q do not factorize because of inteparticle interactions with stronger interactions implying stronger correlations in positions.
<br><br>
- **Probability of many-body systems:** to find particle $1$ at $r_1$, $2$ and $r_2$ ...:
$$\boxed{P(r^N) = P(r_1, r_2,...r_N)}$$
- **Marginal(ized) probability**
$$\boxed{\rho^{2/N} (r_1, r_2) = \int dr_3... dr_N P(r_1, r_2,...r_N)}$$
- **Marginal(ized) probability for any particle 1 and 2**
$$\boxed{\rho^{2/N} (r_1, r_2) = N(N-1)\int dr_3... dr_N P(r_1, r_2,...r_N)}$$
- **Marginal(ized) probability for any particle 1 and 2...n**
$$\boxed{\rho^{n/N} (r_1, r_2,... r_n) = \frac{N!}{(N-n)!}\int dr^{N-n} P(r_1, r_2,...r_N)}$$
### Radial distribution function (RDF)
For an isotropic liquid:
$$\rho = \rho = \frac{N}{V}$$
For an ideal gas:
$$\rho^{2/N} = \frac{N(N-1)}{V^2} = \rho^2 (1-N^{-1}) \approx \rho^2$$
- To measure degree of spatial correlations we introduce Radial Distribution function (RDF):
$$\boxed{g(r_1, r_2) = \frac{\rho^{2/N}(r_1, r_2)}{\rho^2}}$$
- RDF for isotropic fluids is $\boxed{g(r) = g(|r_2-r_1|)}$
<br><br>
- Conditional probabilioty density of **finding a particle at r distance away from a tagged particle** placed at origin:
$$\rho^{2/N}(0,r) = \rho g(r)$$
### Coordination shells and structure in fluids




### Reversible work theorem and potential of mean force
$$\boxed{g(r) = e^{-\beta w(r)}}$$
$$\boxed{w(r) = - \beta^{-1} log [g(r)]}$$
- $w(r)$ Reversible work to bring two particles from infinity to distance r
- $g(r)$ Radial distribution function
$$\Big \langle -\frac{d U(r^N)}{dr_1} \Big \rangle_{r_1, r_2} = -\frac{\int dr_3...dr_N \frac{dU(r^N)}{dr_1} e^{-\beta U}}{\int dr_3...r_N e^{-\beta U}} = \frac{\beta^{-1} \frac{d}{d r_1} \int dr_3...dr_N e^{-\beta U}}{\int dr_3...dr_N e^{-\beta U}} = \beta^{-1} \frac{d}{d r_1} log \int dr_3...dr_N e^{-\beta U} $$
$$\Big \langle -\frac{d U(r^N)}{dr_1} \Big \rangle_{r_1, r_2} = \beta^{-1} \frac{d}{d r_1} log N (N-1)\int dr_3...dr_N e^{-\beta U} = \beta^{-1} g(r_1, r_2)$$
### Thermodynamic properites of $g(r)$
$$\langle E \rangle = N \Big\langle \frac{p^2}{2m} \Big\rangle + \Big\langle \sum_{j>i} u(|r_i - r_j|)\Big \rangle$$
$$\boxed{E/N = \frac{3}{2}k_B T +\frac{1}{2}\rho \int dr g(r) u(r) }$$
### Low density approximation for $g(r)$
$$w(r) =u(r) +\Delta w(r) $$
$$ g(r) = e^{-\beta u(r)} \Big (1 +O(\rho) \Big)$$
### Density expanesion and virial coefficients
$$\beta p = \rho + B_2(T) \rho^2+ O(\rho^3)$$
$$B_2(T) = -\frac{1}{2} \int dr (e^{-\beta u(r)}-1) $$
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
from google.colab import drive
drive.mount('/content/drive')
path="/content/drive/MyDrive/Research/alternate_minimisation/"
name="50_50_10runs"
# mu1 = np.array([3,3,3,3,0])
# sigma1 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu2 = np.array([4,4,4,4,0])
# sigma2 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu3 = np.array([10,5,5,10,0])
# sigma3 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu4 = np.array([-10,-10,-10,-10,0])
# sigma4 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu5 = np.array([-21,4,4,-21,0])
# sigma5 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu6 = np.array([-10,18,18,-10,0])
# sigma6 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu7 = np.array([4,20,4,20,0])
# sigma7 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu8 = np.array([4,-20,-20,4,0])
# sigma8 = np.array([[16,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu9 = np.array([20,20,20,20,0])
# sigma9 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# mu10 = np.array([20,-10,-10,20,0])
# sigma10 = np.array([[1,1,1,1,1],[1,16,1,1,1],[1,1,1,1,1],[1,1,1,1,1],[1,1,1,1,1]])
# sample1 = np.random.multivariate_normal(mean=mu1,cov= sigma1,size=500)
# sample2 = np.random.multivariate_normal(mean=mu2,cov= sigma2,size=500)
# sample3 = np.random.multivariate_normal(mean=mu3,cov= sigma3,size=500)
# sample4 = np.random.multivariate_normal(mean=mu4,cov= sigma4,size=500)
# sample5 = np.random.multivariate_normal(mean=mu5,cov= sigma5,size=500)
# sample6 = np.random.multivariate_normal(mean=mu6,cov= sigma6,size=500)
# sample7 = np.random.multivariate_normal(mean=mu7,cov= sigma7,size=500)
# sample8 = np.random.multivariate_normal(mean=mu8,cov= sigma8,size=500)
# sample9 = np.random.multivariate_normal(mean=mu9,cov= sigma9,size=500)
# sample10 = np.random.multivariate_normal(mean=mu10,cov= sigma10,size=500)
# X = np.concatenate((sample1,sample2,sample3,sample4,sample5,sample6,sample7,sample8,sample9,sample10),axis=0)
# Y = np.concatenate((np.zeros((500,1)),np.ones((500,1)),2*np.ones((500,1)),3*np.ones((500,1)),4*np.ones((500,1)),
# 5*np.ones((500,1)),6*np.ones((500,1)),7*np.ones((500,1)),8*np.ones((500,1)),9*np.ones((500,1))),axis=0).astype(int)
# print(X.shape,Y.shape)
# # plt.scatter(sample1[:,0],sample1[:,1],label="class_0")
# # plt.scatter(sample2[:,0],sample2[:,1],label="class_1")
# # plt.scatter(sample3[:,0],sample3[:,1],label="class_2")
# # plt.scatter(sample4[:,0],sample4[:,1],label="class_3")
# # plt.scatter(sample5[:,0],sample5[:,1],label="class_4")
# # plt.scatter(sample6[:,0],sample6[:,1],label="class_5")
# # plt.scatter(sample7[:,0],sample7[:,1],label="class_6")
# # plt.scatter(sample8[:,0],sample8[:,1],label="class_7")
# # plt.scatter(sample9[:,0],sample9[:,1],label="class_8")
# # plt.scatter(sample10[:,0],sample10[:,1],label="class_9")
# # plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left')
# class SyntheticDataset(Dataset):
# """MosaicDataset dataset."""
# def __init__(self, x, y):
# """
# Args:
# csv_file (string): Path to the csv file with annotations.
# root_dir (string): Directory with all the images.
# transform (callable, optional): Optional transform to be applied
# on a sample.
# """
# self.x = x
# self.y = y
# #self.fore_idx = fore_idx
# def __len__(self):
# return len(self.y)
# def __getitem__(self, idx):
# return self.x[idx] , self.y[idx] #, self.fore_idx[idx]
# trainset = SyntheticDataset(X,Y)
# # testset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
# classes = ('zero','one','two','three','four','five','six','seven','eight','nine')
# foreground_classes = {'zero','one','two'}
# fg_used = '012'
# fg1, fg2, fg3 = 0,1,2
# all_classes = {'zero','one','two','three','four','five','six','seven','eight','nine'}
# background_classes = all_classes - foreground_classes
# background_classes
# trainloader = torch.utils.data.DataLoader(trainset, batch_size=100, shuffle=True)
# dataiter = iter(trainloader)
# background_data=[]
# background_label=[]
# foreground_data=[]
# foreground_label=[]
# batch_size=100
# for i in range(50):
# images, labels = dataiter.next()
# for j in range(batch_size):
# if(classes[labels[j]] in background_classes):
# img = images[j].tolist()
# background_data.append(img)
# background_label.append(labels[j])
# else:
# img = images[j].tolist()
# foreground_data.append(img)
# foreground_label.append(labels[j])
# foreground_data = torch.tensor(foreground_data)
# foreground_label = torch.tensor(foreground_label)
# background_data = torch.tensor(background_data)
# background_label = torch.tensor(background_label)
# def create_mosaic_img(bg_idx,fg_idx,fg):
# """
# bg_idx : list of indexes of background_data[] to be used as background images in mosaic
# fg_idx : index of image to be used as foreground image from foreground data
# fg : at what position/index foreground image has to be stored out of 0-8
# """
# image_list=[]
# j=0
# for i in range(9):
# if i != fg:
# image_list.append(background_data[bg_idx[j]])
# j+=1
# else:
# image_list.append(foreground_data[fg_idx])
# label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
# #image_list = np.concatenate(image_list ,axis=0)
# image_list = torch.stack(image_list)
# return image_list,label
# desired_num = 3000
# mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
# fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
# mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
# list_set_labels = []
# for i in range(desired_num):
# set_idx = set()
# np.random.seed(i)
# bg_idx = np.random.randint(0,3500,8)
# set_idx = set(background_label[bg_idx].tolist())
# fg_idx = np.random.randint(0,1500)
# set_idx.add(foreground_label[fg_idx].item())
# fg = np.random.randint(0,9)
# fore_idx.append(fg)
# image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
# mosaic_list_of_images.append(image_list)
# mosaic_label.append(label)
# list_set_labels.append(set_idx)
# def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number):
# """
# mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
# labels : mosaic_dataset labels
# foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
# dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
# """
# avg_image_dataset = []
# for i in range(len(mosaic_dataset)):
# img = torch.zeros([5], dtype=torch.float64)
# for j in range(9):
# if j == foreground_index[i]:
# img = img + mosaic_dataset[i][j]*dataset_number/9
# else :
# img = img + mosaic_dataset[i][j]*(9-dataset_number)/(8*9)
# avg_image_dataset.append(img)
# return torch.stack(avg_image_dataset) , torch.stack(labels) , foreground_index
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/i
class MosaicDataset1(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list, mosaic_label,fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] , self.fore_idx[idx]
# data = [{"mosaic_list":mosaic_list_of_images, "mosaic_label": mosaic_label, "fore_idx":fore_idx}]
# np.save("mosaic_data.npy",data)
data = np.load(path+"mosaic_data.npy",allow_pickle=True)
mosaic_list_of_images = data[0]["mosaic_list"]
mosaic_label = data[0]["mosaic_label"]
fore_idx = data[0]["fore_idx"]
batch = 250
msd = MosaicDataset1(mosaic_list_of_images, mosaic_label, fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
```
**Focus Net**
```
class Focus_deep(nn.Module):
'''
deep focus network averaged at zeroth layer
input : elemental data
'''
def __init__(self,inputs,output,K,d):
super(Focus_deep,self).__init__()
self.inputs = inputs
self.output = output
self.K = K
self.d = d
self.linear1 = nn.Linear(self.inputs,50) #,self.output)
self.linear2 = nn.Linear(50,self.output)
def forward(self,z):
batch = z.shape[0]
x = torch.zeros([batch,self.K],dtype=torch.float64)
y = torch.zeros([batch,self.d], dtype=torch.float64)
x,y = x.to("cuda"),y.to("cuda")
for i in range(self.K):
x[:,i] = self.helper(z[:,i] )[:,0] # self.d*i:self.d*i+self.d
x = F.softmax(x,dim=1) # alphas
x1 = x[:,0]
for i in range(self.K):
x1 = x[:,i]
y = y+torch.mul(x1[:,None],z[:,i]) # self.d*i:self.d*i+self.d
return y , x
def helper(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
```
**Classification Net**
```
class Classification_deep(nn.Module):
'''
input : elemental data
deep classification module data averaged at zeroth layer
'''
def __init__(self,inputs,output):
super(Classification_deep,self).__init__()
self.inputs = inputs
self.output = output
self.linear1 = nn.Linear(self.inputs,50)
self.linear2 = nn.Linear(50,self.output)
def forward(self,x):
x = F.relu(self.linear1(x))
x = self.linear2(x)
return x
def calculate_attn_loss(dataloader,what,where,criter):
what.eval()
where.eval()
r_loss = 0
alphas = []
lbls = []
pred = []
fidices = []
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx = data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
avg,alpha = where(inputs)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
loss = criter(outputs, labels)
r_loss += loss.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/i,analysis
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
number_runs = 10
full_analysis =[]
FTPT_analysis = pd.DataFrame(columns = ["FTPT","FFPT", "FTPF","FFPF"])
every_what_epoch = 10
for n in range(number_runs):
print("--"*40)
# instantiate focus and classification Model
torch.manual_seed(n)
where = Focus_deep(5,1,9,5).double()
torch.manual_seed(n)
what = Classification_deep(5,3).double()
where = where.to("cuda")
what = what.to("cuda")
# instantiate optimizer
optimizer_where = optim.Adam(where.parameters(),lr =0.01)
optimizer_what = optim.Adam(what.parameters(), lr=0.01)
criterion = nn.CrossEntropyLoss()
acti = []
analysis_data = []
loss_curi = []
epochs = 1000
# calculate zeroth epoch loss and FTPT values
running_loss,anlys_data = calculate_attn_loss(train_loader,what,where,criterion)
loss_curi.append(running_loss)
analysis_data.append(anlys_data)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what.train()
where.train()
if ((epoch) % (every_what_epoch*2) ) <= every_what_epoch-1 :
print(epoch+1,"updating what_net, where_net is freezed")
print("--"*40)
elif ((epoch) % (every_what_epoch*2)) > every_what_epoch-1 :
print(epoch+1,"updating where_net, what_net is freezed")
print("--"*40)
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_where.zero_grad()
optimizer_what.zero_grad()
# forward + backward + optimize
avg, alpha = where(inputs)
outputs = what(avg)
loss = criterion(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
if ((epoch) % (every_what_epoch*2) ) <= every_what_epoch-1 :
optimizer_what.step()
elif ( (epoch) % (every_what_epoch*2)) > every_what_epoch-1 :
optimizer_where.step()
# optimizer_where.step()
# optimizer_what.step()
running_loss,anls_data = calculate_attn_loss(train_loader,what,where,criterion)
analysis_data.append(anls_data)
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.01:
break
print('Finished Training run ' +str(n))
analysis_data = np.array(analysis_data)
FTPT_analysis.loc[n] = analysis_data[-1,:4]/30
full_analysis.append((epoch, analysis_data))
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
images, labels,_ = data
images = images.double()
images, labels = images.to("cuda"), labels.to("cuda")
avg, alpha = where(images)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 3000 train images: %d %%' % ( 100 * correct / total))
a,b= full_analysis[0]
print(a)
cnt=1
for epoch, analysis_data in full_analysis:
analysis_data = np.array(analysis_data)
# print("="*20+"run ",cnt,"="*20)
plt.figure(figsize=(6,6))
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,0],label="ftpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,1],label="ffpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,2],label="ftpf")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,3],label="ffpf")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title("Training trends for run "+str(cnt))
plt.savefig(path+"50_50_10runs/every10/run"+str(cnt)+".png",bbox_inches="tight")
plt.savefig(path+"50_50_10runs/every10/run"+str(cnt)+".pdf",bbox_inches="tight")
cnt+=1
np.mean(np.array(FTPT_analysis),axis=0)
FTPT_analysis.to_csv(path+"50_50_10runs/FTPT_analysis_every10"+name+".csv",index=False)
FTPT_analysis
```
| github_jupyter |
# Keras Backend
In this notebook we will be using the [Keras backend module](http://keras.io/backend/), which provides an abstraction over both Theano and Tensorflow.
Let's try to re-implement the Logistic Regression Model using the `keras.backend` APIs.
The following code will look like very similar to what we would write in Theano or Tensorflow (with the *only difference* that it may run on both the two backends).
```
import keras.backend as K
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from kaggle_data import load_data, preprocess_data, preprocess_labels
X_train, labels = load_data('../data/kaggle_ottogroup/train.csv', train=True)
X_train, scaler = preprocess_data(X_train)
Y_train, encoder = preprocess_labels(labels)
X_test, ids = load_data('../data/kaggle_ottogroup/test.csv', train=False)
X_test, _ = preprocess_data(X_test, scaler)
nb_classes = Y_train.shape[1]
print(nb_classes, 'classes')
dims = X_train.shape[1]
print(dims, 'dims')
feats = dims
training_steps = 25
x = K.placeholder(dtype="float", shape=X_train.shape)
target = K.placeholder(dtype="float", shape=Y_train.shape)
# Set model weights
W = K.variable(np.random.rand(dims, nb_classes))
b = K.variable(np.random.rand(nb_classes))
# Define model and loss
y = K.dot(x, W) + b
loss = K.categorical_crossentropy(y, target)
activation = K.softmax(y) # Softmax
lr = K.constant(0.01)
grads = K.gradients(loss, [W,b])
updates = [(W, W-lr*grads[0]), (b, b-lr*grads[1])]
train = K.function(inputs=[x, target], outputs=[loss], updates=updates)
# Training
loss_history = []
for epoch in range(training_steps):
current_loss = train([X_train, Y_train])[0]
loss_history.append(current_loss)
if epoch % 20 == 0:
print("Loss: {}".format(current_loss))
loss_history = [np.mean(lh) for lh in loss_history]
# plotting
plt.plot(range(len(loss_history)), loss_history, 'o', label='Logistic Regression Training phase')
plt.ylabel('cost')
plt.xlabel('epoch')
plt.legend()
plt.show()
```
## Your Turn
Please switch to the **Theano** backend and **restart** the notebook.
You _should_ see no difference in the execution!
**Reminder**: please keep in mind that you *can* execute shell commands from a notebook (pre-pending a `!` sign).
Thus:
```shell
!cat ~/.keras/keras.json
```
should show you the content of your keras configuration file.
### Moreover
Try to play a bit with the **learning reate** parameter to see how the loss history floats...
---
## Exercise: Linear Regression
To get familiar with automatic differentiation, we start by learning a simple linear regression model using Stochastic Gradient Descent (SGD).
Recall that given a dataset $\{(x_i, y_i)\}_{i=0}^N$, with $x_i, y_i \in \mathbb{R}$, the objective of linear regression is to find two scalars $w$ and $b$ such that $y = w\cdot x + b$ fits the dataset. In this tutorial we will learn $w$ and $b$ using SGD and a Mean Square Error (MSE) loss:
$$\mathcal{l} = \frac{1}{N} \sum_{i=0}^N (w\cdot x_i + b - y_i)^2$$
Starting from random values, parameters $w$ and $b$ will be updated at each iteration via the following rule:
$$w_t = w_{t-1} - \eta \frac{\partial \mathcal{l}}{\partial w}$$
<br>
$$b_t = b_{t-1} - \eta \frac{\partial \mathcal{l}}{\partial b}$$
where $\eta$ is the learning rate.
**NOTE:** Recall that **linear regression** is indeed a **simple neuron** with a linear activation function!!
### Definition: Placeholders and Variables
First of all, we define the necessary variables and placeholders for our computational graph. Variables maintain state across executions of the computational graph, while placeholders are ways to feed the graph with external data.
For the linear regression example, we need three variables: `w`, `b`, and the learning rate for SGD, `lr`.
Two placeholders `x` and `target` are created to store $x_i$ and $y_i$ values.
```
# Placeholders and variables
x = K.placeholder()
target = K.placeholder()
w = K.variable(np.random.rand())
b = K.variable(np.random.rand())
```
#### Notes:
In case you're wondering what's the difference between a **placeholder** and a **variable**, in short:
* Use `K.variable()` for trainable variables such as weights (`W`) and biases (`b`) for your model.
* Use `K.placeholder()` to feed actual data (e.g. training examples)
## Model definition
Now we can define the $y = w\cdot x + b$ relation as well as the MSE loss in the computational graph.
```
# Define model and loss
# %load ../solutions/sol_2311.py
```
Then, given the gradient of MSE wrt to `w` and `b`, we can define how we update the parameters via SGD:
```
# %load ../solutions/sol_2312.py
```
The whole model can be encapsulated in a `function`, which takes as input `x` and `target`, returns the current loss value and updates its parameter according to `updates`.
```
train = K.function(inputs=[x, target], outputs=[loss], updates=updates)
```
## Training
Training is now just a matter of calling the `function` we have just defined. Each time `train` is called, indeed, `w` and `b` will be updated using the SGD rule.
Having generated some random training data, we will feed the `train` function for several epochs and observe the values of `w`, `b`, and loss.
```
# Generate data
np_x = np.random.rand(1000)
np_target = 0.96*np_x + 0.24
# Training
loss_history = []
for epoch in range(200):
current_loss = train([np_x, np_target])[0]
loss_history.append(current_loss)
if epoch % 20 == 0:
print("Loss: %.03f, w, b: [%.02f, %.02f]" % (current_loss, K.eval(w), K.eval(b)))
```
We can also plot the loss history:
```
# Plot loss history
# %load ../solutions/sol_2313.py
```
### Final Note:
Please switch back your backend to `tensorflow` before moving on. It may be useful for next notebooks !-)
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Build a Convolutional Neural Network using Estimators
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r1/tutorials/estimators/cnn.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/estimators/cnn.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
The `tf.layers` module provides a high-level API that makes
it easy to construct a neural network. It provides methods that facilitate the
creation of dense (fully connected) layers and convolutional layers, adding
activation functions, and applying dropout regularization. In this tutorial,
you'll learn how to use `layers` to build a convolutional neural network model
to recognize the handwritten digits in the MNIST data set.

The [MNIST dataset](http://yann.lecun.com/exdb/mnist/) comprises 60,000
training examples and 10,000 test examples of the handwritten digits 0–9,
formatted as 28x28-pixel monochrome images.
## Get Started
Let's set up the imports for our TensorFlow program:
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow.compat.v1 as tf
import numpy as np
tf.logging.set_verbosity(tf.logging.INFO)
```
## Intro to Convolutional Neural Networks
Convolutional neural networks (CNNs) are the current state-of-the-art model
architecture for image classification tasks. CNNs apply a series of filters to
the raw pixel data of an image to extract and learn higher-level features, which
the model can then use for classification. CNNs contains three components:
* **Convolutional layers**, which apply a specified number of convolution
filters to the image. For each subregion, the layer performs a set of
mathematical operations to produce a single value in the output feature map.
Convolutional layers then typically apply a
[ReLU activation function](https://en.wikipedia.org/wiki/Rectifier_\(neural_networks\)) to
the output to introduce nonlinearities into the model.
* **Pooling layers**, which
[downsample the image data](https://en.wikipedia.org/wiki/Convolutional_neural_network#Pooling_layer)
extracted by the convolutional layers to reduce the dimensionality of the
feature map in order to decrease processing time. A commonly used pooling
algorithm is max pooling, which extracts subregions of the feature map
(e.g., 2x2-pixel tiles), keeps their maximum value, and discards all other
values.
* **Dense (fully connected) layers**, which perform classification on the
features extracted by the convolutional layers and downsampled by the
pooling layers. In a dense layer, every node in the layer is connected to
every node in the preceding layer.
Typically, a CNN is composed of a stack of convolutional modules that perform
feature extraction. Each module consists of a convolutional layer followed by a
pooling layer. The last convolutional module is followed by one or more dense
layers that perform classification. The final dense layer in a CNN contains a
single node for each target class in the model (all the possible classes the
model may predict), with a
[softmax](https://en.wikipedia.org/wiki/Softmax_function) activation function to
generate a value between 0–1 for each node (the sum of all these softmax values
is equal to 1). We can interpret the softmax values for a given image as
relative measurements of how likely it is that the image falls into each target
class.
Note: For a more comprehensive walkthrough of CNN architecture, see Stanford University's [Convolutional Neural Networks for Visual Recognition course material](https://cs231n.github.io/convolutional-networks/).
## Building the CNN MNIST Classifier
Let's build a model to classify the images in the MNIST dataset using the
following CNN architecture:
1. **Convolutional Layer #1**: Applies 32 5x5 filters (extracting 5x5-pixel
subregions), with ReLU activation function
2. **Pooling Layer #1**: Performs max pooling with a 2x2 filter and stride of 2
(which specifies that pooled regions do not overlap)
3. **Convolutional Layer #2**: Applies 64 5x5 filters, with ReLU activation
function
4. **Pooling Layer #2**: Again, performs max pooling with a 2x2 filter and
stride of 2
5. **Dense Layer #1**: 1,024 neurons, with dropout regularization rate of 0.4
(probability of 0.4 that any given element will be dropped during training)
6. **Dense Layer #2 (Logits Layer)**: 10 neurons, one for each digit target
class (0–9).
The `tf.layers` module contains methods to create each of the three layer types
above:
* `conv2d()`. Constructs a two-dimensional convolutional layer. Takes number
of filters, filter kernel size, padding, and activation function as
arguments.
* `max_pooling2d()`. Constructs a two-dimensional pooling layer using the
max-pooling algorithm. Takes pooling filter size and stride as arguments.
* `dense()`. Constructs a dense layer. Takes number of neurons and activation
function as arguments.
Each of these methods accepts a tensor as input and returns a transformed tensor
as output. This makes it easy to connect one layer to another: just take the
output from one layer-creation method and supply it as input to another.
Add the following `cnn_model_fn` function, which
conforms to the interface expected by TensorFlow's Estimator API (more on this
later in [Create the Estimator](#create-the-estimator)). This function takes
MNIST feature data, labels, and mode (from
`tf.estimator.ModeKeys`: `TRAIN`, `EVAL`, `PREDICT`) as arguments;
configures the CNN; and returns predictions, loss, and a training operation:
```
def cnn_model_fn(features, labels, mode):
"""Model function for CNN."""
# Input Layer
input_layer = tf.reshape(features["x"], [-1, 28, 28, 1])
# Convolutional Layer #1
conv1 = tf.layers.conv2d(
inputs=input_layer,
filters=32,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Pooling Layer #1
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2], strides=2)
# Convolutional Layer #2 and Pooling Layer #2
conv2 = tf.layers.conv2d(
inputs=pool1,
filters=64,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2], strides=2)
# Dense Layer
pool2_flat = tf.reshape(pool2, [-1, 7 * 7 * 64])
dense = tf.layers.dense(inputs=pool2_flat, units=1024, activation=tf.nn.relu)
dropout = tf.layers.dropout(
inputs=dense, rate=0.4, training=mode == tf.estimator.ModeKeys.TRAIN)
# Logits Layer
logits = tf.layers.dense(inputs=dropout, units=10)
predictions = {
# Generate predictions (for PREDICT and EVAL mode)
"classes": tf.argmax(input=logits, axis=1),
# Add `softmax_tensor` to the graph. It is used for PREDICT and by the
# `logging_hook`.
"probabilities": tf.nn.softmax(logits, name="softmax_tensor")
}
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions)
# Calculate Loss (for both TRAIN and EVAL modes)
loss = tf.losses.sparse_softmax_cross_entropy(labels=labels, logits=logits)
# Configure the Training Op (for TRAIN mode)
if mode == tf.estimator.ModeKeys.TRAIN:
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(
loss=loss,
global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
# Add evaluation metrics (for EVAL mode)
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(
labels=labels, predictions=predictions["classes"])
}
return tf.estimator.EstimatorSpec(
mode=mode, loss=loss, eval_metric_ops=eval_metric_ops)
```
The following sections (with headings corresponding to each code block above)
dive deeper into the `tf.layers` code used to create each layer, as well as how
to calculate loss, configure the training op, and generate predictions. If
you're already experienced with CNNs and [TensorFlow `Estimator`s](../../guide/custom_estimators.md),
and find the above code intuitive, you may want to skim these sections or just
skip ahead to ["Training and Evaluating the CNN MNIST Classifier"](#train_eval_mnist).
### Input Layer
The methods in the `layers` module for creating convolutional and pooling layers
for two-dimensional image data expect input tensors to have a shape of
<code>[<em>batch_size</em>, <em>image_height</em>, <em>image_width</em>,
<em>channels</em>]</code> by default. This behavior can be changed using the
<code><em>data_format</em></code> parameter; defined as follows:
* `batch_size` —Size of the subset of examples to use when performing
gradient descent during training.
* `image_height` —Height of the example images.
* `image_width` —Width of the example images.
* `channels` —Number of color channels in the example images. For color
images, the number of channels is 3 (red, green, blue). For monochrome
images, there is just 1 channel (black).
* `data_format` —A string, one of `channels_last` (default) or `channels_first`.
`channels_last` corresponds to inputs with shape
`(batch, ..., channels)` while `channels_first` corresponds to
inputs with shape `(batch, channels, ...)`.
Here, our MNIST dataset is composed of monochrome 28x28 pixel images, so the
desired shape for our input layer is <code>[<em>batch_size</em>, 28, 28,
1]</code>.
To convert our input feature map (`features`) to this shape, we can perform the
following `reshape` operation:
```
input_layer = tf.reshape(features["x"], [-1, 28, 28, 1])
```
Note that we've indicated `-1` for batch size, which specifies that this
dimension should be dynamically computed based on the number of input values in
`features["x"]`, holding the size of all other dimensions constant. This allows
us to treat `batch_size` as a hyperparameter that we can tune. For example, if
we feed examples into our model in batches of 5, `features["x"]` will contain
3,920 values (one value for each pixel in each image), and `input_layer` will
have a shape of `[5, 28, 28, 1]`. Similarly, if we feed examples in batches of
100, `features["x"]` will contain 78,400 values, and `input_layer` will have a
shape of `[100, 28, 28, 1]`.
### Convolutional Layer #1
In our first convolutional layer, we want to apply 32 5x5 filters to the input
layer, with a ReLU activation function. We can use the `conv2d()` method in the
`layers` module to create this layer as follows:
```
conv1 = tf.layers.conv2d(
inputs=input_layer,
filters=32,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
```
The `inputs` argument specifies our input tensor, which must have the shape
<code>[<em>batch_size</em>, <em>image_height</em>, <em>image_width</em>,
<em>channels</em>]</code>. Here, we're connecting our first convolutional layer
to `input_layer`, which has the shape <code>[<em>batch_size</em>, 28, 28,
1]</code>.
Note: `conv2d()` will instead accept a shape of `[<em>batch_size</em>, <em>channels</em>, <em>image_height</em>, <em>image_width</em>]` when passed the argument `data_format=channels_first`.
The `filters` argument specifies the number of filters to apply (here, 32), and
`kernel_size` specifies the dimensions of the filters as `[<em>height</em>,
<em>width</em>]</code> (here, <code>[5, 5]`).
<p class="tip"><b>TIP:</b> If filter height and width have the same value, you can instead specify a
single integer for <code>kernel_size</code>—e.g., <code>kernel_size=5</code>.</p>
The `padding` argument specifies one of two enumerated values
(case-insensitive): `valid` (default value) or `same`. To specify that the
output tensor should have the same height and width values as the input tensor,
we set `padding=same` here, which instructs TensorFlow to add 0 values to the
edges of the input tensor to preserve height and width of 28. (Without padding,
a 5x5 convolution over a 28x28 tensor will produce a 24x24 tensor, as there are
24x24 locations to extract a 5x5 tile from a 28x28 grid.)
The `activation` argument specifies the activation function to apply to the
output of the convolution. Here, we specify ReLU activation with
`tf.nn.relu`.
Our output tensor produced by `conv2d()` has a shape of
<code>[<em>batch_size</em>, 28, 28, 32]</code>: the same height and width
dimensions as the input, but now with 32 channels holding the output from each
of the filters.
### Pooling Layer #1
Next, we connect our first pooling layer to the convolutional layer we just
created. We can use the `max_pooling2d()` method in `layers` to construct a
layer that performs max pooling with a 2x2 filter and stride of 2:
```
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2], strides=2)
```
Again, `inputs` specifies the input tensor, with a shape of
<code>[<em>batch_size</em>, <em>image_height</em>, <em>image_width</em>,
<em>channels</em>]</code>. Here, our input tensor is `conv1`, the output from
the first convolutional layer, which has a shape of <code>[<em>batch_size</em>,
28, 28, 32]</code>.
Note: As with <code>conv2d()</code>, <code>max_pooling2d()</code> will instead
accept a shape of <code>[<em>batch_size</em>, <em>channels</em>,
<em>image_height</em>, <em>image_width</em>]</code> when passed the argument
<code>data_format=channels_first</code>.
The `pool_size` argument specifies the size of the max pooling filter as
<code>[<em>height</em>, <em>width</em>]</code> (here, `[2, 2]`). If both
dimensions have the same value, you can instead specify a single integer (e.g.,
`pool_size=2`).
The `strides` argument specifies the size of the stride. Here, we set a stride
of 2, which indicates that the subregions extracted by the filter should be
separated by 2 pixels in both the height and width dimensions (for a 2x2 filter,
this means that none of the regions extracted will overlap). If you want to set
different stride values for height and width, you can instead specify a tuple or
list (e.g., `stride=[3, 6]`).
Our output tensor produced by `max_pooling2d()` (`pool1`) has a shape of
<code>[<em>batch_size</em>, 14, 14, 32]</code>: the 2x2 filter reduces height and width by 50% each.
### Convolutional Layer #2 and Pooling Layer #2
We can connect a second convolutional and pooling layer to our CNN using
`conv2d()` and `max_pooling2d()` as before. For convolutional layer #2, we
configure 64 5x5 filters with ReLU activation, and for pooling layer #2, we use
the same specs as pooling layer #1 (a 2x2 max pooling filter with stride of 2):
```
conv2 = tf.layers.conv2d(
inputs=pool1,
filters=64,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2], strides=2)
```
Note that convolutional layer #2 takes the output tensor of our first pooling
layer (`pool1`) as input, and produces the tensor `conv2` as output. `conv2`
has a shape of <code>[<em>batch_size</em>, 14, 14, 64]</code>, the same height and width as `pool1` (due to `padding="same"`), and 64 channels for the 64
filters applied.
Pooling layer #2 takes `conv2` as input, producing `pool2` as output. `pool2`
has shape <code>[<em>batch_size</em>, 7, 7, 64]</code> (50% reduction of height and width from `conv2`).
### Dense Layer
Next, we want to add a dense layer (with 1,024 neurons and ReLU activation) to
our CNN to perform classification on the features extracted by the
convolution/pooling layers. Before we connect the layer, however, we'll flatten
our feature map (`pool2`) to shape <code>[<em>batch_size</em>,
<em>features</em>]</code>, so that our tensor has only two dimensions:
```
pool2_flat = tf.reshape(pool2, [-1, 7 * 7 * 64])
```
In the `reshape()` operation above, the `-1` signifies that the *`batch_size`*
dimension will be dynamically calculated based on the number of examples in our
input data. Each example has 7 (`pool2` height) * 7 (`pool2` width) * 64
(`pool2` channels) features, so we want the `features` dimension to have a value
of 7 * 7 * 64 (3136 in total). The output tensor, `pool2_flat`, has shape
<code>[<em>batch_size</em>, 3136]</code>.
Now, we can use the `dense()` method in `layers` to connect our dense layer as
follows:
```
dense = tf.layers.dense(inputs=pool2_flat, units=1024, activation=tf.nn.relu)
```
The `inputs` argument specifies the input tensor: our flattened feature map,
`pool2_flat`. The `units` argument specifies the number of neurons in the dense
layer (1,024). The `activation` argument takes the activation function; again,
we'll use `tf.nn.relu` to add ReLU activation.
To help improve the results of our model, we also apply dropout regularization
to our dense layer, using the `dropout` method in `layers`:
```
dropout = tf.layers.dropout(
inputs=dense, rate=0.4, training=mode == tf.estimator.ModeKeys.TRAIN)
```
Again, `inputs` specifies the input tensor, which is the output tensor from our
dense layer (`dense`).
The `rate` argument specifies the dropout rate; here, we use `0.4`, which means
40% of the elements will be randomly dropped out during training.
The `training` argument takes a boolean specifying whether or not the model is
currently being run in training mode; dropout will only be performed if
`training` is `True`. Here, we check if the `mode` passed to our model function
`cnn_model_fn` is `TRAIN` mode.
Our output tensor `dropout` has shape <code>[<em>batch_size</em>, 1024]</code>.
### Logits Layer
The final layer in our neural network is the logits layer, which will return the
raw values for our predictions. We create a dense layer with 10 neurons (one for
each target class 0–9), with linear activation (the default):
```
logits = tf.layers.dense(inputs=dropout, units=10)
```
Our final output tensor of the CNN, `logits`, has shape `[batch_size, 10]`.
### Generate Predictions {#generate_predictions}
The logits layer of our model returns our predictions as raw values in a
<code>[<em>batch_size</em>, 10]</code>-dimensional tensor. Let's convert these
raw values into two different formats that our model function can return:
* The **predicted class** for each example: a digit from 0–9.
* The **probabilities** for each possible target class for each example: the
probability that the example is a 0, is a 1, is a 2, etc.
For a given example, our predicted class is the element in the corresponding row
of the logits tensor with the highest raw value. We can find the index of this
element using the `tf.argmax`
function:
```
tf.argmax(input=logits, axis=1)
```
The `input` argument specifies the tensor from which to extract maximum
values—here `logits`. The `axis` argument specifies the axis of the `input`
tensor along which to find the greatest value. Here, we want to find the largest
value along the dimension with index of 1, which corresponds to our predictions
(recall that our logits tensor has shape <code>[<em>batch_size</em>,
10]</code>).
We can derive probabilities from our logits layer by applying softmax activation
using `tf.nn.softmax`:
```
tf.nn.softmax(logits, name="softmax_tensor")
```
Note: We use the `name` argument to explicitly name this operation `softmax_tensor`, so we can reference it later. (We'll set up logging for the softmax values in ["Set Up a Logging Hook"](#set-up-a-logging-hook)).
We compile our predictions in a dict, and return an `EstimatorSpec` object:
```
predictions = {
"classes": tf.argmax(input=logits, axis=1),
"probabilities": tf.nn.softmax(logits, name="softmax_tensor")
}
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions)
```
### Calculate Loss {#calculating-loss}
For both training and evaluation, we need to define a
[loss function](https://en.wikipedia.org/wiki/Loss_function)
that measures how closely the model's predictions match the target classes. For
multiclass classification problems like MNIST,
[cross entropy](https://en.wikipedia.org/wiki/Cross_entropy) is typically used
as the loss metric. The following code calculates cross entropy when the model
runs in either `TRAIN` or `EVAL` mode:
```
loss = tf.losses.sparse_softmax_cross_entropy(labels=labels, logits=logits)
```
Let's take a closer look at what's happening above.
Our `labels` tensor contains a list of prediction indices for our examples, e.g. `[1,
9, ...]`. `logits` contains the linear outputs of our last layer.
`tf.losses.sparse_softmax_cross_entropy`, calculates the softmax crossentropy
(aka: categorical crossentropy, negative log-likelihood) from these two inputs
in an efficient, numerically stable way.
### Configure the Training Op
In the previous section, we defined loss for our CNN as the softmax
cross-entropy of the logits layer and our labels. Let's configure our model to
optimize this loss value during training. We'll use a learning rate of 0.001 and
[stochastic gradient descent](https://en.wikipedia.org/wiki/Stochastic_gradient_descent)
as the optimization algorithm:
```
if mode == tf.estimator.ModeKeys.TRAIN:
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(
loss=loss,
global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
```
Note: For a more in-depth look at configuring training ops for Estimator model functions, see ["Defining the training op for the model"](../../guide/custom_estimators.md#defining-the-training-op-for-the-model) in the ["Creating Estimations in tf.estimator"](../../guide/custom_estimators.md) tutorial.
### Add evaluation metrics
To add accuracy metric in our model, we define `eval_metric_ops` dict in EVAL
mode as follows:
```
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(
labels=labels, predictions=predictions["classes"])
}
return tf.estimator.EstimatorSpec(
mode=mode, loss=loss, eval_metric_ops=eval_metric_ops)
```
<a id="train_eval_mnist"></a>
## Training and Evaluating the CNN MNIST Classifier
We've coded our MNIST CNN model function; now we're ready to train and evaluate
it.
### Load Training and Test Data
First, let's load our training and test data with the following code:
```
# Load training and eval data
((train_data, train_labels),
(eval_data, eval_labels)) = tf.keras.datasets.mnist.load_data()
train_data = train_data/np.float32(255)
train_labels = train_labels.astype(np.int32) # not required
eval_data = eval_data/np.float32(255)
eval_labels = eval_labels.astype(np.int32) # not required
```
We store the training feature data (the raw pixel values for 55,000 images of
hand-drawn digits) and training labels (the corresponding value from 0–9 for
each image) as [numpy
arrays](https://docs.scipy.org/doc/numpy/reference/generated/numpy.array.html)
in `train_data` and `train_labels`, respectively. Similarly, we store the
evaluation feature data (10,000 images) and evaluation labels in `eval_data`
and `eval_labels`, respectively.
### Create the Estimator {#create-the-estimator}
Next, let's create an `Estimator` (a TensorFlow class for performing high-level
model training, evaluation, and inference) for our model. Add the following code
to `main()`:
```
# Create the Estimator
mnist_classifier = tf.estimator.Estimator(
model_fn=cnn_model_fn, model_dir="/tmp/mnist_convnet_model")
```
The `model_fn` argument specifies the model function to use for training,
evaluation, and prediction; we pass it the `cnn_model_fn` we created in
["Building the CNN MNIST Classifier."](#building-the-cnn-mnist-classifier) The
`model_dir` argument specifies the directory where model data (checkpoints) will
be saved (here, we specify the temp directory `/tmp/mnist_convnet_model`, but
feel free to change to another directory of your choice).
Note: For an in-depth walkthrough of the TensorFlow `Estimator` API, see the tutorial [Creating Estimators in tf.estimator](../../guide/custom_estimators.md).
### Set Up a Logging Hook {#set_up_a_logging_hook}
Since CNNs can take a while to train, let's set up some logging so we can track
progress during training. We can use TensorFlow's `tf.train.SessionRunHook` to create a
`tf.train.LoggingTensorHook`
that will log the probability values from the softmax layer of our CNN. Add the
following to `main()`:
```
# Set up logging for predictions
tensors_to_log = {"probabilities": "softmax_tensor"}
logging_hook = tf.train.LoggingTensorHook(
tensors=tensors_to_log, every_n_iter=50)
```
We store a dict of the tensors we want to log in `tensors_to_log`. Each key is a
label of our choice that will be printed in the log output, and the
corresponding label is the name of a `Tensor` in the TensorFlow graph. Here, our
`probabilities` can be found in `softmax_tensor`, the name we gave our softmax
operation earlier when we generated the probabilities in `cnn_model_fn`.
Note: If you don't explicitly assign a name to an operation via the `name` argument, TensorFlow will assign a default name. A couple easy ways to discover the names applied to operations are to visualize your graph on [TensorBoard](../../guide/graph_viz.md)) or to enable the [TensorFlow Debugger (tfdbg)](../../guide/debugger.md).
Next, we create the `LoggingTensorHook`, passing `tensors_to_log` to the
`tensors` argument. We set `every_n_iter=50`, which specifies that probabilities
should be logged after every 50 steps of training.
### Train the Model
Now we're ready to train our model, which we can do by creating `train_input_fn`
and calling `train()` on `mnist_classifier`. In the `numpy_input_fn` call, we pass the training feature data and labels to
`x` (as a dict) and `y`, respectively. We set a `batch_size` of `100` (which
means that the model will train on minibatches of 100 examples at each step).
`num_epochs=None` means that the model will train until the specified number of
steps is reached. We also set `shuffle=True` to shuffle the training data. Then train the model a single step and log the output:
```
# Train the model
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": train_data},
y=train_labels,
batch_size=100,
num_epochs=None,
shuffle=True)
# train one step and display the probabilties
mnist_classifier.train(
input_fn=train_input_fn,
steps=1,
hooks=[logging_hook])
```
Now—without logging each step—set `steps=1000` to train the model longer, but in a reasonable time to run this example. Training CNNs is computationally intensive. To increase the accuracy of your model, increase the number of `steps` passed to `train()`, like 20,000 steps.
```
mnist_classifier.train(input_fn=train_input_fn, steps=1000)
```
### Evaluate the Model
Once training is complete, we want to evaluate our model to determine its
accuracy on the MNIST test set. We call the `evaluate` method, which evaluates
the metrics we specified in `eval_metric_ops` argument in the `model_fn`.
Add the following to `main()`:
```
eval_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": eval_data},
y=eval_labels,
num_epochs=1,
shuffle=False)
eval_results = mnist_classifier.evaluate(input_fn=eval_input_fn)
print(eval_results)
```
To create `eval_input_fn`, we set `num_epochs=1`, so that the model evaluates
the metrics over one epoch of data and returns the result. We also set
`shuffle=False` to iterate through the data sequentially.
## Additional Resources
To learn more about TensorFlow Estimators and CNNs in TensorFlow, see the
following resources:
* [Creating Estimators in tf.estimator](../../guide/custom_estimators.md)
provides an introduction to the TensorFlow Estimator API. It walks through
configuring an Estimator, writing a model function, calculating loss, and
defining a training op.
* [Advanced Convolutional Neural Networks](../../tutorials/images/deep_cnn.md) walks through how to build a MNIST CNN classification model
*without estimators* using lower-level TensorFlow operations.
| github_jupyter |
# Policy Iteration Algorithm
Table of contents
* [Abstract](#abstract)
* [The environment](#the-environment)
* [Imports](#imports)
* [Measuring execution time](#measuring-execution-time)
* [The Policy Evaluation Function](#the-policy-evaluation-function)
* [The Policy Improvement Function](#the-policy-improvement-function)
* [Interpreting the Output](#interpreting-the-output)
### Abstract <a name="abstract"></a>
1. Generate random policy
2. Generate random value function
3. ???
4. Profit
Policy iteration is a technique for finding an optimal* policy in a given, fully known environment. It is neither fast nor efficient, and the practical applications are therefore limited. However, it is guaranteed to find an optimal policy in finite time, and serves as an important starting point for many reinforcement learning algorithms.
The algorithm generates a random policy and evaluates it by calculating the value function (with the help of a policy evaluation algorithm).
It then improves the random policy by changing its guidelines according to the newly obtainted value function (choosing actions which lead to the states with the maximum values). This does not yield an optimal policy yet, since the value function is based on a random, (probably**) suboptimal policy.
It then again submits the new, improved policy for review to the evaluation algorithm, receiving the new value function. Again, the policy is updated based on the value function. This process is repeated (iteration!) until there's nothing left to improve - optimality is achieved!
In this example, we will try to find the optimal policy for navigating Jack's car rental problem, an excercise provided in the standard reference of Reinforcement Learning, ["Reinforcement Learning: An Introduction"](http://incompleteideas.net/book/bookdraft2017nov5.pdf) by Sutton and Barto.
The policy iteration code was copied (and slightly modified) from [Denny Britz' GitHub](https://github.com/dennybritz/reinforcement-learning/blob/master/DP/Policy%20Iteration%20Solution.ipynb), but is applied to another problem in this tutorial.
_________
<sup id="fn1">
*why not THE optimal policy? Because technically, there could be several different policies that yield the same reward, and if there's no *better* policy, then multiple optimal policies exist.
</sup>
<br>
<sup id="fn2">
**you could, by chance, have guessed an optimal policy. Gladstone Gander's policy iteration algorithm ends here.
</sup>
### The environment <a name="the-environment"></a>
You don't need to know the details of the environment in order to understand policy iteration. You can find the full description [here](http://incompleteideas.net/book/bookdraft2017nov5.pdf).
In brief, Jack runs two car rentals. He earns money (reward) when someone requests a car, and the location is able to provide a car. If he perceives an imbalance between the two locations, he has the possibility to move cars from one spot to another at a minor cost while the business is closed at night.
But because the requests are randomly and unequally distributed over both locations, it is not obvious how this balancing should be done.
This is where reinforcement learning comes in; if we see earned money as reward, and expenditures as punishment (or negative reward), we are able to calculate guidelines for balancing the count of cars that maximize expected reward (which in this case equals maximizing expected income).
### Imports <a name="imports"></a>
First, we need to import some libraries.
`time` is imported to measure run times.
`numpy` makes some numerical operations easier.
`CarRentalEnvironment` provides the environment to which we want to apply our policy iteration algorithm.
`matplotlib.pyplot` will be used to visualize our purely numerical results.
This page is a Jupyter Notebook that allows you to run and also modify the code that's written in code cells. Go ahead and execute the imports below by either clicking on the "Run this cell"-button, or by clicking in the cell and pressing shift + enter. As I said, you are able to modify the code, but I recommend not messing with the imports. You will get to play later!
```
import time
import sys
!{sys.executable} -m pip install numpy
!{sys.executable} -m pip install matplotlib
import numpy as np
import CarRentalEnvironment as jack
import matplotlib.pyplot as plt
```
### Measuring execution time <a name="measuring-execution-time"></a>
Let us begin by storing the time at the start to keep track of the time it takes to execute the code.
Next, we initialize the environment. This might take a while, since a transition list of all possible transitions is generated. Because in our example - Jack's Car Rental - there is at least a tiny chance to land in almost any state, regardless of the action we take, the transition list is huge (relative to the number of states). It contains every possible combination of starting state, target state, reward and action.
When this is done, we measure the time again to find out how long this step took. This is relevant if we want to optimize our algorithms, because we need to know how long it took to generate the transition list.
jack.env() sets up the environment and return an object of the environment class.
Let's have a look at the environment class as defined in the imported CarRentalEnvironment. Pay special attention to the attributes; they will be of key importance in the next steps!
class environment:
"""
Complete information environment class with attributes that fully define the
environment. The environment is a Markov decision process.
env() should be used to initialize an environment object according to the
Jack's Car Rental excercise.
Attributes:
P[s][a] is a list of transition tuples (prob, next_state, reward, done).
nS is a number of states in the environment.
nA is a number of actions in the environment.
shape[] is optional for visualization purposes. It is a list of
integers representing the edge lengths of a matrix that contains the
states. E.g. if the states are best mapped on a 21x21 matrix, the list
should state [21, 21]
"""
def __init__(self, P, nS, nA):
self.P = P
self.nS = nS
self.nA = nA
self.shape = [nrOfStates, nrOfStates]
```
start_time = time.time()
print("start!")
#env = GridworldEnv()
env = jack.env()
env_time = time.time()
iterationCount = 1
print("Run time of transition list generation = {}s".format(env_time-start_time))
```
### The Policy Evaluation Function <a name="the-policy-evaluation-function"></a>
Now we can start writing our algorithm. At first we need to implement a policy evaluation function, which is a necessary tool for a policy iteration algorithm.
The policy evaluation function is supposed to evaluate a policy that it is given (as an argument), by producing a value function (or utility function). The value function has the form of a vector, with values corresponding to every state. Since the values represent the expected reward (utility) of a state, not of a transition, the vector is relatively small, containing "only" as many values as there are states. This is in contrast to the number of state-action-reward-state combinations, which is much higher.
#### Initializing the Value Function
At first we initialize a value function filled with zeros, whose values are iteratively improved until they become the true values. It doesn't matter with which values you start, they will always converge to the correct solution. In theory, you can speed up the process if you know how the final values roughly look like - by starting with values which are close to that.
For example, if you know the final values will mostly be around 20, you could initialize the value function with 20s at each point to achieve fewer iterations. It will make almost no difference in execution time though, because the final value is approximated quadratically.
But go ahead and try it out yourself by changing the `0` in `V = np.full(env.nS, 0, float)`!
You might need to pick extreme values to get a noticable difference in speed. Note that the same code will have slight variations in execution time when run several times.
```
# Taken from Policy Evaluation Exercise!
def policy_eval(policy, env, discount_factor, theta=0.00001): #original theta value theta=0.00001
"""
Evaluate a policy given an environment and a full description of the environment's dynamics.
Args:
policy: [S, A] shaped matrix representing the policy.
env: OpenAI env. env.P represents the transition probabilities of the environment.
env.P[s][a] is a list of transition tuples (prob, next_state, reward, done).
env.nS is a number of states in the environment.
env.nA is a number of actions in the environment.
theta: We stop evaluation once our value function change is less than theta for all states.
discount_factor: Gamma discount factor.
Returns:
Vector of length env.nS representing the value function.
"""
# Some visualization of the policy iteration steps
global iterationCount
print("evaluating policy iteration " + str(iterationCount) + ", discount factor = " + str(discount_factor) + ", theta = " + str(theta))
print("Policy:")
visualizePretty(np.reshape(np.argmax(policy, axis=1)-5, env.shape), "policy " + str(iterationCount))
print("")
iterationCount += 1
##
# Start with a random (all 0) value function
V = np.full(env.nS, 0, float) # was originally: V = np.zeros(env.nS)
while True:
delta = 0
for s in range(env.nS):
v = 0
# Look at the possible next actions
for a, action_prob in enumerate(policy[s]):
# For each action, look at the possible next states...
#for tupel in env.P[s][a]:
for prob, next_state, reward, done in env.P[s][a]:
#for prob, next_state, reward, done in tupel:
# Calculate the expected value
v += action_prob * prob * (reward + discount_factor * V[next_state])
# How much our value function changed (across any states)
delta = max(delta, np.abs(v - V[s]))
V[s] = v
# Stop evaluating once our value function change is below a threshold
if delta < theta:
break
return np.array(V)
```
#### The For-Each Loop Over `env.P`
`env.P[s][a]` is a list of all possible transitions from state $s$, under the condition that action $a$ is selected. The list contains target states (possible next states) with their respective rewards and probabilities.
This for-each loop calculates the value (expected future reward) of a state under the current policy by summing up the possible rewards which are discounted by the probability of
* the transition itself
* choosing the action under the given policy
and adding the value of the next state, discounted by the discount factor.
#for prob, next_state, reward, done in tupel:
# Calculate the expected value
v += action_prob * prob * (reward + discount_factor * V[next_state])
#### Exit Condition For The Outer Loop
So why is there a `while True:` loop? Shouldn't the loop end once the for-each loop has iterated through every element and has therefore considered all transitions?
The answer to why the latter doesn't work is that when we calculate the value of a state, we need to take into account the value of the next states. We do that by simply adding the value that is given by our value function. But the values of our value function are arbitrary and very likely not correct - we initialized them to be 0 or another guessed value. This means after our first loop through all transitions, the values are all based on a (probably) wrong value function.
We did, however, add some truth to our value function: we considered the (correct) rewards and transition probabilities. This means our value function is not completely arbitrary anymore, it gained some "knowledge" about the true values. By circling through this process often enough, we get closer and closer to the correct values in our value function. This is why we have the "while" loop in the beginning.
We know that we have the correct values in our value function when an update cycle completes without any changes.<sup><a href="#fn1" id="ref1">1</a></sup>
______
<sup id="fn1">1. For proof that convergence is guaranteed with any starting values, see Theorem 3.6 in ["From ants to safe Artificial Intelligence: Reinforcement Learning" (Lang, 2018)](https://github.com/Favodar/Reinforcement-Learning/blob/master/Reinforcement%20Learning%2C%20Part%201.pdf)<a href="#ref1" title="Jump back to footnote 1 in the text.">↩</a></sup>
### The Policy Improvement Function <a name="the-policy-improvement-function"></a>
We will now get to the core of the policy iteration algorithm: the policy_improvement function improves a policy until it becomes an optimal policy for maximizing utility in a given environment.
#### Initializing the policy
As a first step, we generate a random policy, similar to what we did earlier when we set up a random value function `policy = np.ones([env.nS, env.nA]) / env.nA`
#### Evaluating the policy
Next, we let our evaluation function do the job of evaluating this policy. How *value*able are the states if we act according to this policy (in terms of long term expected reward)? This doesn't tell us yet how good our policy is. If we have a utility of, say, 914 in one state, is that good? We don't know, because it's all relative. But we do have a reference now. Future versions of our policy can be measured against this, and that's exactly what we're gonna do!
#### One step lookahead
What we'll do next to improve our policy is to calculate the values of the available actions. So instead of just looking at the state value (which we already did), we look at a state-action pair and calculate the long-term expected reward of this pair. We can do this by looking at where the action could potentially bring us, and summing up the values of those states, multiplied (meaning discounted) by the probability of getting there.
For example, let's say we are in state $s_2$ and if we take action $a_3$, there's a 20% chance to get to state $s_8$, and $s_8$ has a value of $v(s_8)=10$. Then we start by multiplying $v(s_8)$ with the 20% chance:
$v(s_8)*p(s_8|s_2, a_3) = 10*0.2 = 2$
for a in range(env.nA):
for prob, next_state, reward, done in env.P[state][a]:
A[a] += prob * (reward + discount_factor * V[next_state])
return A
So $2$ is the first value we have to keep in mind. But let's say action 3 also yields a 10% chance to arrive in state 20, and state 20 has an astounding value of 360. Then we need to add $360*0.1 = 36$ to what we already got, which is 2. If we keep doing this with every state that action 3 could bring us to, and add up all these values, then we have the value of the state action pair "state 2 - action 3" (under our current policy). Now if we do this for **all** the actions that are available in state 2 (you probably realize by now that this is a lot of calculations), we can chose the one with the highest value and put *this* has our action-recommendation in our policy. [I think some magic happens here before the next step]
Since our policy has now changed (improved!), we need to re-evaluate it to get our value-function up to date as well, and then we can rinse and repeat.
If we do this often enough, we will eventually get an optimal policy. How do we know we have an optimal policy? Same as with the policy evaluation function - if the update cycle completes without changes, we know we're done!
```
def policy_improvement(env, discount_factor, theta=0.00001, policy_eval_fn=policy_eval):
"""
Policy Improvement Algorithm. Iteratively evaluates and improves a policy
until an optimal policy is found.
Args:
env: The OpenAI environment.
policy_eval_fn: Policy Evaluation function that takes 3 arguments:
policy, env, discount_factor.
discount_factor: gamma discount factor.
Returns:
A tuple (policy, V).
policy is the optimal policy, a matrix of shape [S, A] where each state s
contains a valid probability distribution over actions.
V is the value function for the optimal policy.
"""
def one_step_lookahead(state, V):
"""
Helper function to calculate the value for all action in a given state.
Args:
state: The state to consider (int)
V: The value to use as an estimator, Vector of length env.nS
Returns:
A vector of length env.nA containing the expected value of each action.
"""
A = np.zeros(env.nA)
for a in range(env.nA):
for prob, next_state, reward, done in env.P[state][a]:
A[a] += prob * (reward + discount_factor * V[next_state])
return A
# Start with a random policy
policy = np.ones([env.nS, env.nA]) / env.nA
while True:
# Evaluate the current policy
V = policy_eval_fn(policy, env, discount_factor, theta)
# Will be set to false if we make any changes to the policy
policy_stable = True
# For each state...
for s in range(env.nS):
# The best action we would take under the currect policy
chosen_a = np.argmax(policy[s])
# Find the best action by one-step lookahead
# Ties are resolved arbitarily
action_values = one_step_lookahead(s, V)
best_a = np.argmax(action_values)
# Greedily update the policy
if chosen_a != best_a:
policy_stable = False
policy[s] = np.eye(env.nA)[best_a]
# If the policy is stable we've found an optimal policy. Return it
if policy_stable:
return policy, V
```
### Interpreting the Output <a name="interpreting-the-output"></a>
#### Policy Probability Distribution
Each row of the policy probability distribution represents a state, while the columns represent the actions.
The numbers signify the probability of taking the action in that state under the policy, with a value between 0 and 1.
For example, a `1` in position $(2,6)$ means a 100% percent chance to take the 6th action in the 2nd state (which is the state where c1 = 0 and c2 = 1). The 6th action is the action "0" or "move no cars".
#### Reshaped Policy
The reshaped policy shapes the policy into a matrix where the x-axis represents the number of cars at c1 and the _inverted_ y-axis signifies c2.
The numbers in the matrix are the action with the highest probability at that state, under the final policy.
#### Value Function
The value function matrix has the same structure as the reshaped policy, but the numbers signify the value (expected future reward, or utility) of that state, if the final policy is followed. The value function is not a policy as it doesn't contain instructions. It tells you how valuable a state is *if* you choose optimal actions from there on, but it doesn't tell you what those actions are (assuming it is the value function of the optimal policy, as it is the case here).
```
def visualizePretty(array, title="title goes here"):
##
dim_1, dim_2 = array.shape
fig, ax = plt.subplots(figsize=(7,7))
im = ax.imshow(array)
# Loop over data dimensions and create text annotations.
for i in range(dim_1):
for j in range(dim_2):
text = ax.text(j, i, array[i, j],
ha="center", va="center", color="w")
ax.set_title(title)
fig.tight_layout()
plt.show()
##
plt.rcParams.update({'font.size': 22}) #this just sets the font size for the visualizing plots
policy, v = policy_improvement(env, 0.9, 0.00001)
iteration_time = time.time()
print("Run time of policy iteration = {} sec".format(iteration_time-env_time))
print("Policy Probability Distribution:")
print(policy)
print("")
print("Reshaped Policy (-5 = move 5 cars from B to A, 0 = move no cars, 5 = move 5 cars from A to B):")
#print("(not implemented)")
print(np.reshape(np.argmax(policy, axis=1)-5, env.shape))
print("")
print("visualizePretty:")
visualizePretty(np.reshape(np.argmax(policy, axis=1)-5, env.shape), "final policy")
print("Value Function:")
print(v)
print("")
print("Reshaped Grid Value Function:")
#print("(not implemented)")
list3 = []
for number in v:
list3.append(round(number, 3))
#print("(not implemented)")
print(str(np.reshape(list3, env.shape)).replace("'", ""))
print("")
plt.rcParams.update({'font.size': 5})
print("visualizePretty:")
visualizePretty(np.reshape(v, env.shape), "value function")
end_time = time.time()
print("Run time total = {} sec".format(end_time - start_time))
print("Run time of transition list generation = {} sec".format(env_time-start_time))
print("Run time of policy iteration = {} sec".format(iteration_time-env_time))
```
As you can see, convergence is *fast*. While the initial, arbitrary policy always recommended the action "-5", the first improvement (policy 2) is already very similar to the final policy, with only minor deviations. Of course, lots of operations take place between every iteration of the policy, but I still find it fascinating how a value function based on an awfully bad policy ("move 5 cars from A to B, regardless of the current distribution of cars") can guide us to an almost optimal policy in just one iteration.
If you haven't done so already, you can play around with the discount rate and the theta value in `policy, v = policy_improvement(env, 0.9, 0.00001)`. You may notice that changing the discount rate doesn't change much of the result. Can you figure out why?
*Hint: think about what the discount rate actually does, and how optimizing for short-term reward and long-term reward would differ in our environment*.
| github_jupyter |
```
# https://github.com/LankyCyril/pyvenn/blob/master/pyvenn-demo.ipynb
# steps of making venns:
# read in h5ad files and name them
# make dict with the h5ad file for specific label and corresponding name of that column
# remove genes from all ages from each individual age
# take set of those
# venn(dict)
```
## setup
```
# from anndata import read_h5ad
# from scRFE.scRFE import scRFE
# cellscRFE = scRFE(adata =adata, classOfInterest = 'cell_ontology_class', nEstimators = 10, Cv=3)
# cellscRFE[0]
adata = read_h5ad('/Users/madelinepark/downloads/Liver_droplet.h5ad')
# adata.obs
# Venn Diagrams
from matplotlib import pyplot as plt
from venn import venn
%matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams['pdf.fonttype'] = 42
plt.rcParams['ps.fonttype'] = 42
# saves text as editable text for illustrator
import pandas as pd
from venn import pseudovenn
# general format:
# make dictionary where the keys are labels and values are set of column of a csv
# venn(dict, cmap = '')
```
## facs/droplet comparisons
```
cd /users/madelinepark/Downloads/differentTreesFacsTfsResults
facs1000 = pd.read_csv('1000_facs_tfs_age.csv')
cd /users/madelinepark/Downloads/differentTreesDropletTfsResults
droplet1000 = pd.read_csv('1000_droplet_tfs_age.csv')
facsDroplet24m = {
'facs24m' : set(facs1000['24m']),
'droplet24m' : set(droplet1000['24m'])
}
cd /Users/madelinepark/Desktop
venn(facsDroplet24m)
plt.savefig('facsDroplet24m.pdf')
droplet1000['3m']
facsDroplet3m = {
'Facs 3m Genes' : set(facs1000['3m']),
'Droplet 3m Genes' : set(droplet1000['3m'])
}
venn(facsDroplet3m)
facsDroplet18m = {
'Facs 18m Genes' : set(facs1000['18m']),
'Droplet 18m Genes' : set(droplet1000['18m'])
}
venn(facsDroplet18m)
mammaryGlandDroplet = pd.read_csv('/Users/madelinepark/Downloads/Mammary_Gland_droplet_tf_age_5000.csv')
mammaryGlandFacs = pd.read_csv('/Users/madelinepark/Downloads/Mammary_Gland_facs_tf_age_5000.csv')
cd /Users/madelinepark/Desktop
mammaryGlandfacsDroplet3m = {
'Mammary_Gland_droplet_3m' : set(mammaryGlandDroplet['3m']),
'Mammary_Gland_facs_3m' : set(mammaryGlandFacs['3m'])
}
venn(mammaryGlandfacsDroplet3m)
plt.savefig('mammaryGlandfacsDroplet3m.pdf')
LungDroplet = pd.read_csv('/Users/madelinepark/Downloads/Lung_droplet_tf_age_5000.csv')
LungFacs = pd.read_csv('/Users/madelinepark/Downloads/Lung_facs_tf_age_5000.csv')
pwd
LungfacsDroplet18m = {
'Lung_droplet_18m' : set(LungDroplet['18m']),
'Lung_facs_18m' : set(LungFacs['18m'])
}
venn(LungfacsDroplet18m)
plt.savefig('LungfacsDroplet18m.pdf')
cd /users/madelinepark/downloads
liverCellFacs = pd.read_csv('Liverfacs1000tfcelltype2.csv')
liverCellDrop = pd.read_csv('Liverdroplet1000tfcelltype2.csv')
LiverFacsDroplet1 = {
'Liver Facs NK Cell Genes' : set(liverCellFacs['NK cell']),
'Liver Droplet NK Cell Genes' : set(liverCellDrop['NK cell'])
}
venn(LiverFacsDroplet1)
LiverFacsDroplet2 = {
'Liver Facs Hepatocyte Genes' : set(liverCellFacs['hepatocyte']),
'Liver Droplet Hepatocyte Genes' : set(liverCellDrop['hepatocyte'])
}
venn(LiverFacsDroplet2)
```
## Tissue at different ages venn
### Liver
```
cd /Users/madelinepark/downloads
Liver1m = pd.read_csv('1mLiverdroplet1000tfcelltype.csv')
Liver3m = pd.read_csv('3mLiverdroplet1000tfcelltype.csv')
Liver18m = pd.read_csv('18mLiverdroplet1000tfcelltype.csv')
Liver21m = pd.read_csv('21mLiverdroplet1000tfcelltype.csv')
Liver24m = pd.read_csv('24mLiverdroplet1000tfcelltype.csv')
Liver30m = pd.read_csv('30mLiverdroplet1000tfcelltype.csv')
LiverAllAge = pd.read_csv('Liverdroplet1000tfcelltype.csv')
# list1 = [11, 5, 17, 18, 23, 50]
LiverAllAgeBCell = list(set(LiverAllAge['B cell']))
Liver1mBcell = list(set(Liver1m['B cell']))
Liver18mBcell = list(set(Liver18m['B cell']))
Liver21mBcell = list(set(Liver21m['B cell']))
Liver24mBcell = list(set(Liver24m['B cell']))
Liver30mBcell = list(set(Liver30m['B cell']))
Liver1mBcell = [ele for ele in Liver1mBcell if ele not in LiverAllAgeBCell]
Liver18mBcell = [ele for ele in Liver18mBcell if ele not in LiverAllAgeBCell]
Liver21mBcell = [ele for ele in Liver21mBcell if ele not in LiverAllAgeBCell]
Liver24mBcell = [ele for ele in Liver24mBcell if ele not in LiverAllAgeBCell]
Liver30mBcell = [ele for ele in Liver30mBcell if ele not in LiverAllAgeBCell]
LiverDroplet = {
'1m_Liver_Bcell Genes' : set(Liver1mBcell),
# '3mLiverBcell Genes' : set(Liver3m['B cell']), (must not have Bcell)
'18m_Liver_Bcell Genes' : set(Liver18mBcell),
'21m_Liver_Bcell Genes' : set(Liver21mBcell),
'24m_Liver_Bcell Genes' : set(Liver24mBcell),
'30m_Liver_Bcell Genes' : set(Liver30mBcell)#,
# 'AllAge_Liver_Bcell' : set(LiverAllAge['B cell'])
}
cd /Users/madelinepark/desktop
venn (LiverDroplet, cmap = 'plasma')
plt.savefig('LiverDropletBcellAge.pdf')
# FOR THIS, ADD ALL AGE ONCE IT FINISHES RUNNING
# save as pdf
# remove 1m ones from all, 18m from all, etc, get rid of allage
```
### Marrow
```
cd /Users/madelinepark/downloads
Marrow1m = pd.read_csv('1mMarrowdroplet1000tfcelltype.csv')
Marrow3m = pd.read_csv('3mMarrowdroplet1000tfcelltype.csv')
Marrow18m = pd.read_csv('18mMarrowdroplet1000tfcelltype.csv')
Marrow21m = pd.read_csv('21mMarrowdroplet1000tfcelltype.csv')
Marrow24m = pd.read_csv('24mMarrowdroplet1000tfcelltype.csv')
Marrow30m = pd.read_csv('30mMarrowdroplet1000tfcelltype.csv')
MarrowAllAge = pd.read_csv('Marrowdroplet1000tfcelltype.csv')
# list1 = [11, 5, 17, 18, 23, 50]
MarrowAllAgeBasophil = list(set(MarrowAllAge['basophil']))
Marrow1mBasophil = list(set(Marrow1m['basophil']))
Marrow1mBasophil = list(set(Marrow1m['basophil']))
Marrow3mBasophil = list(set(Marrow3m['basophil']))
Marrow18mBasophil = list(set(Marrow18m['basophil']))
Marrow21mBasophil = list(set(Marrow21m['basophil']))
Marrow24mBasophil = list(set(Marrow24m['basophil']))
Marrow30mBasophil = list(set(Marrow30m['basophil']))
Marrow1mBasophil = [ele for ele in Marrow1mBasophil if ele not in MarrowAllAgeBasophil]
Marrow3mBasophil = [ele for ele in Marrow3mBasophil if ele not in MarrowAllAgeBasophil]
Marrow18mBasophil = [ele for ele in Marrow18mBasophil if ele not in MarrowAllAgeBasophil]
Marrow21mBasophil = [ele for ele in Marrow21mBasophil if ele not in MarrowAllAgeBasophil]
Marrow24mBasophil = [ele for ele in Marrow24mBasophil if ele not in MarrowAllAgeBasophil]
Marrow30mBasophil = [ele for ele in Marrow30mBasophil if ele not in MarrowAllAgeBasophil]
MarrowDropletBasophil = {
'1m_Marrow_Basophil Genes' : set(Marrow1mBasophil),
'3m_Marrow_Basophil Genes' : set(Marrow3mBasophil),
'18m_Marrow_Basophil Genes' : set(Marrow18mBasophil),
# '21m_Marrow_Basophil Genes' : set(Marrow21mBasophil),
'24m_Marrow_Basophil Genes' : set(Marrow24mBasophil),
'30m_Marrow_Basophil Genes' : set(Marrow30mBasophil)
}
cd /Users/madelinepark/desktop
venn (MarrowDropletBasophil, cmap = 'plasma')
plt.savefig('MarrowDropletBasophilAge.pdf')
```
### Kidney
```
cd /Users/madelinepark/downloads
Kidney1m = pd.read_csv('1mKidneydroplet1000tfcelltype.csv')
Kidney3m = pd.read_csv('3mKidneydroplet1000tfcelltype.csv')
Kidney18m = pd.read_csv('18mKidneydroplet1000tfcelltype.csv')
Kidney21m = pd.read_csv('21mKidneydroplet1000tfcelltype.csv')
Kidney24m = pd.read_csv('24mKidneydroplet1000tfcelltype.csv')
Kidney30m = pd.read_csv('30mKidneydroplet1000tfcelltype.csv')
KidneyAllAge = pd.read_csv('Kidneydroplet1000tfcelltype.csv')
# list1 = [11, 5, 17, 18, 23, 50]
KidneyAllAgeNK = list(set(KidneyAllAge['NK cell']))
Kidney1mNK = list(set(Kidney1m['NK cell']))
Kidney3mNK = list(set(Kidney3m['NK cell']))
Kidney18mNK = list(set(Kidney18m['NK cell']))
Kidney21mNK = list(set(Kidney21m['NK cell']))
Kidney24mNK = list(set(Kidney24m['NK cell']))
Kidney30mNK = list(set(Kidney30m['NK cell']))
Kidney1mNK = [ele for ele in Kidney1mNK if ele not in KidneyAllAgeNK]
Kidney3mNK = [ele for ele in Kidney3mNK if ele not in KidneyAllAgeNK]
Kidney18mNK = [ele for ele in Kidney18mNK if ele not in KidneyAllAgeNK]
Kidney21mNK = [ele for ele in Kidney21mNK if ele not in KidneyAllAgeNK]
Kidney24mNK = [ele for ele in Kidney24mNK if ele not in KidneyAllAgeNK]
Kidney30mNK = [ele for ele in Kidney30mNK if ele not in KidneyAllAgeNK]
KidneyDropletNK = {
'1m_Kidney_NKcell Genes' : set(Kidney1mNK),
'3m_Kidney_NKcell Genes' : set(Kidney3mNK),
'18m_Kidney_NKcell Genes' : set(Kidney18mNK),
# '21m_Kidney_NKcell Genes' : set(Kidney21mNK),
'24m_Kidney_NKcell Genes' : set(Kidney24mNK),
'30m_Kidney_NKcell Genes' : set(Kidney30mNK)
}
cd /Users/madelinepark/desktop
venn (KidneyDropletNK, cmap = 'plasma')
plt.savefig('KidneyDropletNkcellAge.pdf')
```
### Heart
```
cd /users/madelinepark/downloads
Heart1m = pd.read_csv('1mHeart_and_Aortadroplet1000tfcelltype.csv')
Heart3m = pd.read_csv('3mHeart_and_Aortadroplet1000tfcelltype.csv')
Heart18m = pd.read_csv('18mHeart_and_Aortadroplet1000tfcelltype.csv')
Heart21m = pd.read_csv('21mHeart_and_Aortadroplet1000tfcelltype.csv')
Heart24m = pd.read_csv('24mHeart_and_Aortadroplet1000tfcelltype.csv')
Heart30m = pd.read_csv('30mHeart_and_Aortadroplet1000tfcelltype.csv')
HeartAllAge = pd.read_csv('Heart_and_Aortadroplet1000tfcelltype.csv')
HeartAllAgeCardiacNeuron = list(set(HeartAllAge['cardiac neuron']))
Heart1mCardiacNeuron = list(set(Heart1m['cardiac neuron']))
Heart3mCardiacNeuron = list(set(Heart3m['cardiac neuron']))
Heart18mCardiacNeuron = list(set(Heart18m['cardiac neuron']))
Heart21mCardiacNeuron = list(set(Heart21m['cardiac neuron']))
# Heart24mCardiacNeuron = list(set(Heart24m['cardiac neuron']))
Heart30mCardiacNeuron = list(set(Heart30m['cardiac neuron']))
Heart1mCardiacNeuron = [ele for ele in Heart1mCardiacNeuron if ele not in Heart1mCardiacNeuron]
Heart3mCardiacNeuron = [ele for ele in Heart1mCardiacNeuron if ele not in Heart3mCardiacNeuron]
Heart18mCardiacNeuron = [ele for ele in Heart1mCardiacNeuron if ele not in Heart18mCardiacNeuron]
Heart21mCardiacNeuron = [ele for ele in Heart1mCardiacNeuron if ele not in Heart21mCardiacNeuron]
Heart30mCardiacNeuron = [ele for ele in Heart1mCardiacNeuron if ele not in Heart30mCardiacNeuron]
Heart_and_AortaDroplet = {
'1m_Heart_CardiacNeuron_Genes' : set(Heart1mCardiacNeuron),
'3m_Heart_CardiacNeuron_Genes' : set(Heart3mCardiacNeuron),
'18m_Heart_CardiacNeuron_Genes' : set(Heart18mCardiacNeuron),
'21m_Heart_CardiacNeuron_Genes' : set(Heart21mCardiacNeuron),
'30m_Heart_CardiacNeuron_Genes' : set(Heart30mCardiacNeuron)
}
venn(Heart_and_AortaDroplet, cmap = 'plasma')
# plt.savefig('LiverBcellAge.pdf')
```
| github_jupyter |
```
import sys
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
sys.path.append('../../Model')
from Inclusion import Inclusion
a=pd.read_pickle('../../Data/Plate_Alpha.pkl')
b=pd.read_pickle('../../Data/Plate_Beta.pkl')
#Calculate inclusion rate for each plate
labels=['Alpha','Beta']
plate_data=[]
for i,df in enumerate([a,b]):
data=[]
efficiency=[]
for x in df.Plate.unique(): #For each plate
plate=df[df.Plate==x]
singles=np.unique(plate['Productive'],return_counts=1)[1][1] #wells with 1 productive TCR
try:
doubles=np.unique(plate['Productive'],return_counts=1)[1][2] #wells with 2 productive TCRs
except:
doubles=0 #Case that a plate has no wells with 2 TCRs
data.append(doubles/(singles+doubles)) #allelic inclusion frequency
efficiency.append((singles+doubles)/96) #percentage of wells with >=1 productive TCR
plate_data.append(data)
print(labels[i])
print(f" Inclusion rate: {np.mean(data)} +/- {np.std(data)}")
print(f" Capture efficiency: {np.mean(efficiency)} +/- {np.std(efficiency)}")
print(f"Efficiency per plate: {efficiency}")
print('\n')
#import allelic inclusion data
a=np.load('../../Data/Alpha_distribution.npy')
b=np.load('../../Data/Beta_distribution.npy')
alphas=pd.DataFrame(a,columns=['S'+str(x) for x in range(1,a.shape[1]+1)]).iloc[:12]
betas=pd.DataFrame(b,columns=['S'+str(x) for x in range(1,a.shape[1]+1)]).iloc[:12]
#Run allelic inclusion model for S8
counts_alpha=np.array(alphas['S8'])
counts_beta=np.array(betas['S8'])
df=Inclusion(alpha=counts_alpha,beta=counts_beta)
alpha_map=df.return_map[2]
beta_map=df.return_map[4]
alpha_lower=df.return_map_interval_lower[2]
beta_lower=df.return_map_interval_lower[4]
alpha_upper=df.return_map_interval_upper[2]
beta_upper=df.return_map_interval_upper[4]
#Transform data for plotting
all_data=[]
all_data.append([alpha_map,alpha_lower,alpha_upper,'Alpha','model'])
all_data.append([np.mean(plate_data[0]),np.mean(plate_data[0])-np.std(plate_data[0]),np.mean(plate_data[0])+np.std(plate_data[0]),'Alpha','plate'])
all_data.append([beta_map,beta_lower,beta_upper,'Beta','model'])
all_data.append([np.mean(plate_data[1]),np.mean(plate_data[1])-np.std(plate_data[1]),np.mean(plate_data[1])+np.std(plate_data[1]),'Beta','plate'])
all_data=pd.DataFrame(all_data,columns=['Mean','Lower','Upper','Chain','Method'])
all_data[['Mean','Lower','Upper']]=all_data[['Mean','Lower','Upper']].astype(float)*100
plt.bar([0,.5,1.1,1.6],all_data.Mean,color=['DarkGray','Gray'],alpha=0.75,width=0.5)
plt.errorbar([0,.5,1.1,1.6],all_data.Mean,yerr=[all_data.Mean-all_data.Lower,all_data.Upper-all_data.Mean],fmt='o', ecolor='black',markersize=0,markeredgecolor='gray', markerfacecolor='gray',capthick=2)
plt.ylim([0,18])
plt.yticks(fontsize=20)
plt.xticks([.25,1.35],[r'$\alpha$',r'$\beta$'],fontsize=25)
plt.show()
plt.close()
```
| github_jupyter |
## Sequence Classification
## Task 1.1: Document-level Sentiment Classification
Build a Bidirectional Recurrent Neural Network (RNN) model for multi-class sentiment classification. Compare the performance with a Unidirectional RNN model. Your model (each) shall
include:
- RNN network that learns sentence representation from input sequences.
- Fully connected network that predicts sentiment label, given the learnt state representation.
Train the model by using data iterator and batch generator. Evaluate the trained model on
the provided test set.
## Unidirectional RNN Model for document level sentiment classification
```
import os
import sys
import codecs
import operator
import numpy as np
import re
from time import time
import _pickle as cPickle
from keras.preprocessing import sequence
from keras.utils.np_utils import to_categorical
import operator
from keras.layers import Dense, Dropout, Activation, Embedding, LSTM, Input, Bidirectional
from keras.models import Model
import keras.optimizers as opt
from keras.callbacks import EarlyStopping, ModelCheckpoint
from google.colab import drive
drive.mount('/content/drive/')
cd drive/My Drive/Colab Notebooks
data_path = 'data/doc_level'
num_regex = re.compile('^[+-]?[0-9]+\.?[0-9]*$')
def create_vocab(domain, data_path, maxlen=0, vocab_size=0):
print('Creating vocab ...')
f = os.path.join(data_path,'%s_text.txt'%(domain))
total_words, unique_words = 0, 0
word_freqs = {}
fin = codecs.open(f, 'r', 'utf-8')
for line in fin:
words = line.split()
if maxlen > 0 and len(words) > maxlen:
continue
for w in words:
if not bool(num_regex.match(w)):
try:
word_freqs[w] += 1
except KeyError:
unique_words += 1
word_freqs[w] = 1
total_words += 1
print (' %i total words, %i unique words' % (total_words, unique_words))
sorted_word_freqs = sorted(word_freqs.items(), key=operator.itemgetter(1), reverse=True)
vocab = {'<pad>':0, '<unk>':1, '<num>':2}
index = len(vocab)
for word, _ in sorted_word_freqs:
vocab[word] = index
index += 1
if vocab_size > 0 and index > vocab_size + 2:
break
if vocab_size > 0:
print (' keep the top %i words' % vocab_size)
return vocab
def create_data(vocab, text_path, label_path, domain, skip_top, skip_len, replace_non_vocab):
data = []
label = [] # {pos: 0, neg: 1, neu: 2}
f = codecs.open(text_path, 'r', 'utf-8')
f_l = codecs.open(label_path, 'r', 'utf-8')
num_hit, unk_hit, skip_top_hit, total = 0., 0., 0., 0.
pos_count, neg_count, neu_count = 0, 0, 0
max_len = 0
for line, score in zip(f, f_l):
word_indices = []
words = line.split()
if skip_len > 0 and len(words) > skip_len:
continue
score = float(score.strip())
if score < 3:
neg_count += 1
label.append(1)
elif score > 3:
pos_count += 1
label.append(0)
else:
neu_count += 1
label.append(2)
for word in words:
if bool(num_regex.match(word)):
word_indices.append(vocab['<num>'])
num_hit += 1
elif word in vocab:
word_ind = vocab[word]
if skip_top > 0 and word_ind < skip_top + 3:
skip_top_hit += 1
else:
word_indices.append(word_ind)
else:
if replace_non_vocab:
word_indices.append(vocab['<unk>'])
unk_hit += 1
total += 1
if len(word_indices) > max_len:
max_len = len(word_indices)
data.append(word_indices)
f.close()
f_l.close()
print(' <num> hit rate: %.2f%%, <unk> hit rate: %.2f%%' % (100*num_hit/total, 100*unk_hit/total))
print (domain)
print( 'pos count: ', pos_count )
print( 'neg count: ', neg_count )
print( 'neu count: ', neu_count )
return np.array(data), np.array(label), max_len
def prepare_data(domain, data_path, vocab_size, skip_top=0, skip_len=0, replace_non_vocab=1):
print(domain)
assert domain in ['amazon_electronics', 'yelp14']
vocab = create_vocab(domain, data_path, skip_len, vocab_size)
#print(vocab)
text_path = os.path.join(data_path,'%s_text.txt'%(domain))
score_path = os.path.join(data_path,'%s_label.txt'%(domain))
data, label, max_len = create_data(vocab, text_path, score_path, domain, skip_top, \
skip_len, replace_non_vocab)
return vocab, data, label, max_len
# choose domain data to train
domain_name = 'amazon_electronics'
vocab, data_list, label_list, overall_maxlen = prepare_data(domain_name, data_path, 10000)
idx_words = dict((v,k) for (k,v) in vocab.items())
data_path_save = 'Assign3DataStorage/'
def read_pickle(path_data, file_name):
f = open(os.path.join(path_data, file_name), 'rb')
read_file = cPickle.load(f)
f.close()
return read_file
def save_pickle(path_data, file_name, data):
f = open(os.path.join(path_data, file_name), 'wb')
cPickle.dump(data, f)
print(" file saved to: %s"%(os.path.join(path_data, file_name)))
f.close()
save_pickle(data_path_save, 'words_idx.pkl', vocab)
save_pickle(data_path_save, 'idx_words.pkl', idx_words)
save_pickle(data_path_save, 'data.pkl', data_list)
save_pickle(data_path_save, 'label.pkl', label_list)
```
### End of Preprocessing
### Model training, testing and conclusion summary of these are as follows
```
words_idx = read_pickle(data_path, 'words_idx.pkl')
idx_words = read_pickle(data_path, 'idx_words.pkl')
data = read_pickle(data_path, 'data.pkl')
label = read_pickle(data_path, 'label.pkl')
rand_idx = np.arange(len(data))
np.random.shuffle(rand_idx)
data = data[rand_idx]
label = to_categorical(label)[rand_idx]
data_size = len(data)
test_x = data[0:6000]
test_y = label[0:6000]
dev_x = data[6000:10800]
dev_y = label[6000:10800]
train_x = data[10800:int(data_size)]
train_y = label[10800:int(data_size)]
maxlen = 300
words_idx = [x for (x, _) in sorted(words_idx.items(), key=operator.itemgetter(1))]
train_x_ = sequence.pad_sequences(train_x, maxlen)
dev_x_ = sequence.pad_sequences(dev_x, maxlen)
test_x_ = sequence.pad_sequences(test_x, maxlen)
train_x_ = np.array(train_x_)
train_y = np.array(train_y)
dev_x_ = np.array(dev_x_)
dev_y = np.array(dev_y)
test_x_ = np.array(test_x_)
test_y = np.array(test_y)
class Dataiterator():
'''
1) Iteration over minibatches using next(); call reset() between epochs to randomly shuffle the data
2) Access to the entire dataset using all()
'''
def __init__(self, X, y, seq_length=32, decoder_dim=300, batch_size=32):
self.X = X
self.y = y
self.num_data = len(X) # total number of examples
self.batch_size = batch_size # batch size
self.reset() # initial: shuffling examples and set index to 0
def __iter__(self): # iterates data
return self
def reset(self): # initials
self.idx = 0
self.order = np.random.permutation(self.num_data) # shuffling examples by providing randomized ids
def __next__(self): # return model inputs - outputs per batch
X_ids = [] # hold ids per batch
while len(X_ids) < self.batch_size:
X_id = self.order[self.idx] # copy random id from initial shuffling
X_ids.append(X_id)
self.idx += 1 #
if self.idx >= self.num_data: # exception if all examples of data have been seen (iterated)
self.reset()
raise StopIteration()
batch_X = self.X[np.array(X_ids)] # X values (encoder input) per batch
batch_y = self.y[np.array(X_ids)] # y_in values (decoder input) per batch
return batch_X, batch_y
def all(self): # return all data examples
return self.X, self.y
```
## Model
```
sentence_input = Input(shape=(300,), dtype='int32', name='sentence_input')
vocab_size = len(words_idx)
word_emb = Embedding(vocab_size, 300, mask_zero=True, name='word_emb')
emb_output = word_emb(sentence_input)
drop = Dropout(0.25)(emb_output)
# 1 no embd drop, lstm dropout = 0.5, reccr drop = 0.1
dropout6 = 0.5
recurrent_dropout6 = 0.1
lstm_layer6 = LSTM(300, return_sequences=False, dropout=dropout6, \
recurrent_dropout=recurrent_dropout6, name='lstm6')(emb_output)
# 2 embd drop = 0.25, lstm dropout = 0.5, reccr drop = 0.1
dropout4 = 0.5
recurrent_dropout4 = 0.1
lstm_layer4 = LSTM(300, return_sequences=False, dropout=dropout4, \
recurrent_dropout=recurrent_dropout4, name='lstm4')(drop)
# 3 embd drop = 0.25, lstm dropout = 0.5, reccr drop = 0.2
dropout5 = 0.5
recurrent_dropout5 = 0.2
lstm_layer5 = LSTM(300, return_sequences=False, dropout=dropout5, \
recurrent_dropout=recurrent_dropout5, name='lstm5')(drop)
densed6 = Dense(3, name='dense')(lstm_layer6)
probs6 = Activation('softmax')(densed6)
densed4 = Dense(3, name='dense')(lstm_layer4)
probs4 = Activation('softmax')(densed4)
densed5 = Dense(3, name='dense')(lstm_layer5)
probs5 = Activation('softmax')(densed5)
model6 = Model(inputs=[sentence_input], outputs=probs6)
model4 = Model(inputs=[sentence_input], outputs=probs4)
model5 = Model(inputs=[sentence_input], outputs=probs5)
optimizer = opt.RMSprop(lr=0.001, rho=0.9, epsilon=1e-06, clipnorm=10, clipvalue=0)
model6.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("summary of Model6")
model6.summary()
model4.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("summary of Model4")
model4.summary()
model5.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("summary of Model5")
model5.summary()
batch_size = 32
train_x_[1].shape
train_steps_epoch = len(train_x_)/batch_size
batch_train_iter = Dataiterator(train_x_, train_y, batch_size)
val_steps_epoch = len(dev_x_)/batch_size
batch_val_iter = Dataiterator(dev_x_, dev_y, batch_size)
test_steps_epoch = len(test_x_)/batch_size
batch_test_iter = Dataiterator(test_x_, test_y, batch_size)
def train_generator(model, batch_train_iter, batch_val_iter):
earlystop_callbacks = [EarlyStopping(monitor='val_loss', patience=12),
ModelCheckpoint(filepath=os.path.join('./','{epoch:02d}-{loss:.2f}.check'), \
monitor='val_loss', save_best_only=False, \
save_weights_only=True)
]
def train_gen():
while True:
train_batches = [[X, y] for X, y in batch_train_iter]
for train_batch in train_batches:
yield train_batch
def val_gen():
while True:
val_batches = [[X, y] for X, y in batch_val_iter]
for val_batch in val_batches:
yield val_batch
history = model.fit_generator(train_gen(), validation_data=val_gen(), \
validation_steps=val_steps_epoch, steps_per_epoch=train_steps_epoch, \
epochs = 20, callbacks = earlystop_callbacks)
```
## Training
```
#Input shape as (300, )
print("Training for model6")
train_generator(model3, batch_train_iter, batch_val_iter)
print("Training for model4")
train_generator(model4, batch_train_iter, batch_val_iter)
print("Training for model5")
train_generator(model5, batch_train_iter, batch_val_iter)
```
## Evaluation
```
#Input shape as (300, )
print("Testing for model6")
loss, accuracy = model3.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy) #was 63.33, re-execute
print("Testing for model4")
loss, accuracy = model4.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy)
print("Testing for model5")
loss, accuracy = model5.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy)
```
## Bidirectional RNN Model for document level sentiment classification
## Model
```
sentence_input = Input(shape = (maxlen, ), dtype = 'int32', name = 'sentence_input')
vocab_size = len(words_idx)
word_emb = Embedding(vocab_size, maxlen, mask_zero=True, name='word_emb')
emb_output = word_emb(sentence_input)
drop = Dropout(0.25)(emb_output)
# 1 no embd drop, lstmdrop = 0.5, recdrop = 0.1
dropout1 = 0.5
recurrent_dropout1 = 0.1
lstm_layer1 = Bidirectional(LSTM(maxlen, return_sequences=False, dropout=dropout1, \
recurrent_dropout=recurrent_dropout1, name='lstm1'))(emb_output)
# 2 embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.1
dropout2 = 0.5
recurrent_dropout2 = 0.1
lstm_layer2 = Bidirectional(LSTM(maxlen, return_sequences=False, dropout=dropout2, \
recurrent_dropout=recurrent_dropout2, name='lstm2'))(drop)
# 3 embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.2
dropout3 = 0.5
recurrent_dropout3 = 0.2
lstm_layer3 = Bidirectional(LSTM(maxlen, return_sequences=False, dropout=dropout3, \
recurrent_dropout=recurrent_dropout3, name='lstm3'))(drop)
# 7 embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.2, merge_mode = 'ave'
dropout7 = 0.5
recurrent_dropout7 = 0.2
lstm_layer7 = Bidirectional(LSTM(maxlen, return_sequences=False, dropout=dropout7, \
recurrent_dropout=recurrent_dropout7, name='lstm3'), merge_mode = 'ave')(drop)
densed1 = Dense(3, name='dense1')(lstm_layer1)
probs1 = Activation('softmax')(densed1)
densed2 = Dense(3, name='dense2')(lstm_layer2)
probs2 = Activation('softmax')(densed2)
densed3 = Dense(3, name='dense3')(lstm_layer3)
probs3 = Activation('softmax')(densed3)
densed7 = Dense(3, name='dense3')(lstm_layer7)
probs7 = Activation('softmax')(densed7)
model1 = Model(inputs=[sentence_input], outputs=probs1)
model2 = Model(inputs=[sentence_input], outputs=probs2)
model3 = Model(inputs=[sentence_input], outputs=probs3)
model7 = Model(inputs=[sentence_input], outputs=probs7)
optimizer = opt.RMSprop(lr=0.001, rho=0.9, epsilon=1e-06, clipnorm=10, clipvalue=0)
model1.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("The summary for Model1")
model1.summary()
model2.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("The summary for Model2")
model2.summary()
model3.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("The summary for Model3")
model3.summary()
model7.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['categorical_accuracy'])
print("The summary for Model7")
model7.summary()
batch_size = 32
train_steps_epoch = len(train_x_)/batch_size
batch_train_iter = Dataiterator(train_x_, train_y, batch_size)
val_steps_epoch = len(dev_x_)/batch_size
batch_val_iter = Dataiterator(dev_x_, dev_y, batch_size)
test_steps_epoch = len(test_x_)/batch_size
batch_test_iter = Dataiterator(test_x_, test_y, batch_size)
def train_generator(model, batch_train_iter, batch_val_iter):
earlystop_callbacks = [EarlyStopping(monitor='val_loss', patience=10),
ModelCheckpoint(filepath=os.path.join('./','{epoch:02d}-{loss:.2f}.check'), \
monitor='val_loss', save_best_only=False, \
save_weights_only=True)
]
def train_gen():
while True:
train_batches = [[X, y] for X, y in batch_train_iter]
for train_batch in train_batches:
yield train_batch
def val_gen():
while True:
val_batches = [[X, y] for X, y in batch_val_iter]
for val_batch in val_batches:
yield val_batch
history = model.fit_generator(train_gen(), validation_data=val_gen(), \
validation_steps=val_steps_epoch, steps_per_epoch=train_steps_epoch, \
epochs = 20, callbacks = earlystop_callbacks)
```
## Training
```
#Without drop 0.25
print("TRAINING FOR MODEL1")
train_generator(model1, batch_train_iter, batch_val_iter)
print("TRAINING FOR MODEL2")
train_generator(model2, batch_train_iter, batch_val_iter)
print("TRAINING FOR MODEL3")
train_generator(model3, batch_train_iter, batch_val_iter)
print("TRAINING FOR MODEL7")
train_generator(model7, batch_train_iter, batch_val_iter)
```
### Evaluate
```
#Without drop 0.25
print("TESTING FOR MODEL1")
loss, accuracy = model1.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy) # test acc is 64.9%
print("TESTING FOR MODEL2")
loss, accuracy = model2.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy)
print("TESTING FOR MODEL3")
loss, accuracy = model3.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy)
print("TESTING FOR MODEL7")
loss, accuracy = model7.evaluate(x = test_x_, y = test_y, verbose = 1)
print(loss)
print(accuracy)
```
### Summary of the model
#### UniDirectional RNN
model6: acc:61.45 (no embd drop, lstm dropout = 0.5, reccr drop = 0.1), approx. training time: 96mins
model4: acc:64.03 (embd drop = 0.25, lstm dropout = 0.5, reccr drop = 0.1), approx. training time: 75mins
model5: acc:61.75 (embd drop = 0.25, lstm dropout = 0.5, reccr drop = 0.2), approx. training time: 85mins
#### BiDirectional RNN
model1: acc:61.38 (no embd drop, lstmdrop = 0.5, recdrop = 0.1), approx. training time: 145mins
model2: acc:63.93 (embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.1), approx. training time: 145mins
model3: acc:63.88 (embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.2), approx. training time: 140mins
model7: acc:65.81 (embd drop = 0.25, lstmdrop = 0.5, recdrop = 0.2, merge_mode = 'ave'), approx. training time: 150mins
We can see that in Uni Directional when having dropout for each embedding layer, lastm layer and reccurent dropout we get the highest accuracy during evaluation as overfitting in the training set has been reduced and thus the model is able to predict the unseen with higher accuracy. Also has less training time than the other models as the number of neurons that are less significant are dropped out.
In Bi Directional we can see that model7 achieves high accuracy with merge mode as average but takes more time for training. Considering the tradeoff between accuracy and training time we can conclude that the configuration in model2 is best performing.
The above reported are the models that give some meaningful change/insight into the performance of the model for small variation in hyper parameters among all the other models that were tried.
| github_jupyter |
# Module 1 Example
The dataset is known to contain information about the total nitrogen content, in mg N per $m^3$, in the effluent stream of a wastewater treatment plant. The required limit value is 18 mg N per $m^3$. The dataset contains daily samples for approximately a three week period.
We are interested in seeing if the effluent complies with regulations, and if there may be problems complying with regulations in future.
We will begin by importing the relevant libraries and modules for our problem. We could also import these as we discover our need for them, but if we know what we're going to be doing we can go ahead and import the ones we know we will need.
```
import pandas
import numpy
from scipy import stats
from matplotlib import pyplot
```
We load in our data, as in the tutorial:
```
N_data = pandas.read_csv(
r'https://raw.githubusercontent.com/imheidimarais/Engineering-Statistics/master/data/Ntot_Data.csv'
)
```
And we can begin by making sure that our data imported properly and try understand what we're working with. We will do this by looking at the size of the dataframe, and the column names. These we can compare to our raw data file and make sure that the import went smoothly.
```
print(f"Size of data: {N_data.shape[0]} rows, {N_data.shape[1]} column(s).")
print("The column title(s) in the dataset are:")
for c in N_data.columns:
print(c)
```
Having confirmed that the data imported as expected, perhaps a good place to start is with visualising our data.
We are interested in:
+ the spread of data
+ the statistics (mean, variance, etc.)
+ if there are any outliers/noise/missing data.
We will use an index plot and a boxplot to accomplish this, the code is the same as in the tutorial, but here we specify the column name instead of keeping it completely general.
```
fig = pyplot.Figure(figsize=(12, 4))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
N_data.reset_index().plot(x='index', y='Ntot', kind='scatter', ax=ax1)
N_data.boxplot(column='Ntot', ax=ax2)
ax1.set(
xlabel='Index',
ylabel='$x$'
)
ax2.set(
xlabel='Sample',
ylabel='$x$'
)
fig
```
We see that there are three suspected outliers, all below our regulation limit. And we can see the median is around 16.6. The data appears well behaved aside from the outliers.
We can calculate the sample statistics:
```
descriptive_statistics = N_data.describe()
print(descriptive_statistics)
```
And we can immediately see the calculated mean is much lower than the median, and does not appear to represent the majority of the data. The outliers are dragging it down significantly.
However, for the sake of completeness we will look at the distribution of the data before we remove any outliers. We will do this with a histogram, and an empirical cumulative distribution function. We will overlay the pdf and cdf of a true normal distribution with the same mean and variance as our data.
```
number_bins = 10
fig = pyplot.Figure(figsize=(12, 4))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
N_data.hist(column='Ntot', bins=number_bins, density=True, ax=ax1)
xrange1 = (N_data['Ntot'].min(), N_data['Ntot'].max())
normal_pdf_xs = numpy.linspace(xrange1[0], xrange1[1], 100)
normal_pdf_ys = stats.norm.pdf(
normal_pdf_xs, loc=descriptive_statistics['Ntot']['mean'], scale=descriptive_statistics['Ntot']['std']
)
ax1.plot(normal_pdf_xs, normal_pdf_ys, c="orange")
N_data_sorted = N_data.sort_values(by='Ntot', ascending=True)
ecdf = [n/N_data_sorted.shape[0] for n in range(1, N_data_sorted.shape[0]+1)]
N_data_sorted['ecdf'] = numpy.array(ecdf)
normal_cdf_ys = stats.norm.cdf(
normal_pdf_xs, loc=descriptive_statistics['Ntot']['mean'], scale=descriptive_statistics['Ntot']['std']
)
N_data_sorted.plot(x='Ntot', y='ecdf', kind='scatter', ax=ax2)
ax2.plot(normal_pdf_xs, normal_cdf_ys, c="orange")
ax1.set(
xlabel='$x$',
ylabel='Frequency'
)
ax2.set(
xlabel='$x$',
ylabel='ecdf'
)
fig
```
The outliers make it impossible to compare our data to the theoretical normal distribution, and they severely skew the histogram and ecdf plots.
For now, we can temporarily remove the outliers. Exercise caution when removing outliers in general as they may contain important information. In this case they were below the regulation limit (not concerning) and an analysis of additional data showed that they came from days with extremely high rainfall. So, for our purposes we can consider it safe to remove the outliers.
```
N_data_no_outliers = N_data[N_data['Ntot']>12]
```
We will now look again at the descriptive statistics, and then at the distributions.
```
descriptive_statistics_new = N_data_no_outliers.describe()
print(descriptive_statistics_new)
number_bins = 10
fig = pyplot.Figure(figsize=(12, 4))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
N_data_no_outliers.hist(column='Ntot', bins=number_bins, density=True, ax=ax1)
xrange1 = (N_data_no_outliers['Ntot'].min(), N_data_no_outliers['Ntot'].max())
normal_pdf_xs = numpy.linspace(xrange1[0], xrange1[1], 100)
normal_pdf_ys = stats.norm.pdf(
normal_pdf_xs, loc=descriptive_statistics_new['Ntot']['mean'], scale=descriptive_statistics_new['Ntot']['std']
)
ax1.plot(normal_pdf_xs, normal_pdf_ys, c="orange")
N_data_sorted = N_data_no_outliers.sort_values(by='Ntot', ascending=True)
ecdf = [n/N_data_sorted.shape[0] for n in range(1, N_data_sorted.shape[0]+1)]
N_data_sorted['ecdf'] = numpy.array(ecdf)
normal_cdf_ys = stats.norm.cdf(
normal_pdf_xs, loc=descriptive_statistics_new['Ntot']['mean'], scale=descriptive_statistics_new['Ntot']['std']
)
N_data_sorted.plot(x='Ntot', y='ecdf', kind='scatter', ax=ax2)
ax2.plot(normal_pdf_xs, normal_cdf_ys, c="orange")
ax1.set(
xlabel='$x$',
ylabel='Frequency'
)
ax2.set(
xlabel='$x$',
ylabel='ecdf'
)
fig
```
We can see that the mean is much closer to the median value we observed in the boxplot, which is a good sign!
We also see that, especially the ecdf, appears fairly normal. While this does not pass as any true test of normallity, it is still a useful thing to do to get an idea of the distribution of your data so that you can better interpret the results of statistical tests that you may carry out.
| github_jupyter |
<a href="https://pymt.readthedocs.io"><img style="float: right" src="images/pymt-logo-header-text.png"></a>
# Dynamically changing a running PyMT model
In this tutorial we will learn how to:
* Use the `update_until` method
* The model grid
* Change the input values of a model while it's running
```
import matplotlib.pyplot as plt
import numpy as np
from tqdm import tqdm
```
# Create and initialize a model
For this simulation, we'll be using the *Child* model with some non-default parameters.
```
from pymt import MODELS
child = MODELS.Child()
```
Have a look under the *Parameters* help section (you may have to scroll down - it's the section after the citations). The *Parameters* section describes optional keywords that you can pass the the `setup` method. In the previous example we just used defaults. Below we'll see how to set input file parameters programmatically through keywords.
```
config_file, config_dir = child.setup(
grid_node_spacing=750.,
grid_x_size=20000.,
grid_y_size=40000.,
run_duration=1e6,
)
child.initialize(config_file, dir=config_dir)
```
To begin with, we'll advance the model through 10 time steps.
```
for t in tqdm(range(10)):
child.update()
```
Update until some time in the future. Notice that, in this case, we update to a partial time step. Child is fine with this however some other models may not be. For models that can not update to times that are not full time steps, PyMT will advance to the next time step and interpolate values to the requested time.
```
child.update_until(201.5)
child.time
```
Child offers different output variables but we get them in the same way as before.
```
child.output_var_names
```
As before, we can get values of a variable with *get_value* (in whatever units we like).
```
child.get_value("land_surface__elevation", units="cm")
```
We can query each input and output variable. PyMT attaches a dictionary to each component called `var` that provides information about each variable. For instance we can see that `"land_surface__elevation"` has units of meters, is an input and output variable, and is defined on the nodes of grid with id 0.
```
child.var["land_surface__elevation"]
```
Notice that this variable is defined on grid with ID 0. We can get information about this grid through the `grid` attribute.
```
child.grid[0]
```
We can also get grid information through method functions of the model. For example, the number of **nodes** that define each **face** of the grid.
```
child.grid_nodes_per_face(0)
```
If we plot this variable, we can see the unsructured triangular grid that Child has decomposed its grid into.
```
child.quick_plot('land_surface__elevation', edgecolors='k', vmin=-200, vmax=200, cmap='BrBG_r')
```
Child initializes it's elevations with random noise centered around 0. We would like instead to give it elevations that have some land and some sea. First we'll get the x and y coordinates for each node along with their elevations.
```
x, y = child.grid_x(0), child.grid_y(0)
z = child.get_value('land_surface__elevation')
```
All nodes above `y=y_shore` will be land, and all nodes below `y=y_shore` will be sea.
```
y_shore = 15000.
z[y < y_shore] -= 100
z[y >= y_shore] += 100
```
We now use the model's **set_value** method to change its current elevation values.
```
child.set_value('land_surface__elevation', z)
```
Just to verify we set things up correctly, we'll create a plot.
```
child.quick_plot('land_surface__elevation', edgecolors='k', vmin=-200, vmax=200, cmap='BrBG_r')
```
To get things going, we'll run the model for 5000 years and see what things look like.
```
child.update_until(5000.)
child.quick_plot("land_surface__elevation", edgecolors="k", vmin=-200, vmax=200, cmap="BrBG_r")
```
## Exercise
We'll have some fun now by adding a simple uplift component. We'll run the component for another 5000 years but this time uplifting a corner of the grid by `dz_dt`. First, use the **get_value** method to create a new array of uplift values.
For this example, make the uplift zero everywhere except for *y>15km* and *x>10km* where it will be *0.02*.
```
# Your code here
x, y = child.grid_x(0), child.grid_y(0)
dz_dt = np.zeros_like(child.get_value("land_surface__elevation"))
dz_dt[(y > 15000.) & (x > 10000.)] = 0.02
```
## Exercise
Now with the uplift, we'll run the component for another 5000 years but this time uplifting a corner of the grid by `dz_dt` every time step.
```
# Your code here
# dz_dt = .02
now = child.time
times, dt = np.linspace(now, now + 5000., 50, retstep=True)
for time in tqdm(times):
child.update_until(time)
z = child.get_value('land_surface__elevation')
z += dz_dt * dt
# z[(y > 15000.) & (x > 10000.)] += dz_dt * dt
child.set_value('land_surface__elevation', z)
child.quick_plot("land_surface__elevation", edgecolors="k", vmin=-200, vmax=200, cmap="BrBG_r")
```
We now stop the uplift and run it for an additional 5000 years.
```
for time in tqdm(np.linspace(child.time, child.time + 5000.)):
child.update_until(time)
child.quick_plot("land_surface__elevation", edgecolors="k", vmin=-200, vmax=200, cmap="BrBG_r")
```
| github_jupyter |
# Bite Size Bayes
Copyright 2020 Allen B. Downey
License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
```
# Load utils.py
import os
if not os.path.exists('utils.py'):
!wget https://github.com/AllenDowney/BiteSizeBayes/raw/master/utils.py
# Load the data file
if not os.path.exists('gss_bayes.csv'):
!wget https://github.com/AllenDowney/BiteSizeBayes/raw/master/gss_bayes.csv
import pandas as pd
import numpy as np
from utils import values
```
## Introduction
This notebook takes a computational approach to understanding probability. We'll use data from the General Social Survey to compute the probability of propositions like:
* If I choose a random survey respondent, what is the probability they are female?
* If I choose a random survey respondent, what is the probability they work in banking?
From there, we will explore two related concepts:
* Conjunction, which is the probability that two propositions are both true; for example, what is the probability of choosing a female banker?
* Conditional probability, which is the probability that one proposition is true, given that another is true; for example, given than a respondent is female, what is the probability that she is a banker?
I chose these examples because they are related to a famous experiment by Tversky and Kahneman, who posed the following question:
> Linda is 31 years old, single, outspoken, and very bright. She majored in philosophy. As a student, she was deeply concerned with issues of discrimination and social justice, and also participated in anti-nuclear demonstrations. Which is more probable?
1. Linda is a bank teller.
2. Linda is a bank teller and is active in the feminist movement.
Many people choose the second answer, presumably because it seems more consistent with the description. It seems unlikely that Linda would be *just* a bank teller; if she is a bank teller, it seems likely that she would also be a feminist.
But the second answer cannot be "more probable", as the question asks. Suppose we find 1000 people who fit Linda's description and 10 of them work as bank tellers. How many of them are also feminists? At most, all 10 of them are; in that case, the two options are *equally* likely. More likely, only some of them are; in that case the second option is *less* likely. But there can't be more than 10 out of 10, so the second option cannot be more likely.
The error people make if they choose the second option is called the [conjunction fallacy](https://en.wikipedia.org/wiki/Conjunction_fallacy). It's called a [fallacy](https://en.wikipedia.org/wiki/Fallacy) because it's a logical error and "conjunction" because "bank teller AND feminist" is a [logical conjunction](https://en.wikipedia.org/wiki/Logical_conjunction).
If this example makes you uncomfortable, you are in good company. The biologist [Stephen J. Gould wrote](https://sci-hub.tw/https://doi.org/10.1080/09332480.1989.10554932) :
> I am particularly fond of this example because I know that the [second] statement is least probable, yet a little [homunculus](https://en.wikipedia.org/wiki/Homunculus_argument) in my head continues to jump up and down, shouting at me, "but she can't just be a bank teller; read the description."
If the little person in your head is still unhappy, maybe this notebook will help.
## Probability
At this point I should define probability, but that [turns out to be surprisingly difficult](https://en.wikipedia.org/wiki/Probability_interpretations). To avoid getting bogged down before we get started, I'll start with a simple definition: a **probability** is a **fraction** of a dataset.
For example, if we survey 1000 people, and 20 of them are bank tellers, the fraction that work as bank tellers is 0.02 or 2\%. If we choose a person from this population at random, the probability that they are a bank teller is 2\%.
(By "at random" I mean that every person in the dataset has the same chance of being chosen, and by "they" I mean the [singular, gender-neutral pronoun](https://en.wikipedia.org/wiki/Singular_they), which is a correct and useful feature of English.)
With this definition and an appropriate dataset, we can compute probabilities by counting.
To demonstrate, I'll use a data set from the [General Social Survey](http://gss.norc.org/) or GSS. The following cell reads the data.
```
gss = pd.read_csv('gss_bayes.csv', index_col=0)
```
The results is a Pandas DataFrame with one row for each person surveyed and one column for each variable I selected.
Here are the number of rows and columns:
```
gss.shape
```
And here are the first few rows:
```
gss.head()
```
The columns are
* `caseid`: Respondent id (which is the index of the table).
* `year`: Year when the respondent was surveyed.
* `age`: Respondent's age when surveyed.
* `sex`: Male or female.
* `polviews`: Political views on a range from liberal to conservative.
* `partyid`: Political party affiliation, Democrat, Independent, or Republican.
* `indus10`: [Code](https://www.census.gov/cgi-bin/sssd/naics/naicsrch?chart=2007) for the industry the respondent works in.
Let's look at these variables in more detail, starting with `indus10`.
## Banking
The code for "Banking and related activities" is 6870, so we can select bankers like this:
```
banker = (gss['indus10'] == 6870)
```
The result is a Boolean series, which is a Pandas Series that contains the values `True` and `False`. Here are the first few entries:
```
banker.head()
```
We can use `values` to see how many times each value appears.
```
values(banker)
```
In this dataset, there are 728 bankers.
If we use the `sum` function on this Series, it treats `True` as 1 and `False` as 0, so the total is the number of bankers.
```
banker.sum()
```
To compute the *fraction* of bankers, we can divide by the number of people in the dataset:
```
banker.sum() / banker.size
```
But we can also use the `mean` function, which computes the fraction of `True` values in the Series:
```
banker.mean()
```
About 1.5% of the respondents work in banking.
That means if we choose a random person from the dataset, the probability they are a banker is about 1.5%.
**Exercise**: The values of the column `sex` are encoded like this:
```
1 Male
2 Female
```
The following cell creates a Boolean series that is `True` for female respondents and `False` otherwise.
```
female = (gss['sex'] == 2)
```
* Use `values` to display the number of `True` and `False` values in `female`.
* Use `sum` to count the number of female respondents.
* Use `mean` to compute the fraction of female respondents.
```
# Solution
values(gss['sex'])
# Solution
female.sum()
# Solution
female.mean()
```
The fraction of women in this dataset is higher than in the adult U.S. population because [the GSS does not include people living in institutions](https://gss.norc.org/faq), including prisons and military housing, and those populations are more likely to be male.
**Exercise:** The designers of the General Social Survey chose to represent sex as a binary variable. What alternatives might they have considered? What are the advantages and disadvantages of their choice?
For more on this topic, you might be interested in this article: Westbrook and Saperstein, [New categories are not enough: rethinking the measurement of sex and gender in social surveys](https://sci-hub.tw/10.1177/0891243215584758)
## Political views
The values of `polviews` are on a seven-point scale:
```
1 Extremely liberal
2 Liberal
3 Slightly liberal
4 Moderate
5 Slightly conservative
6 Conservative
7 Extremely conservative
```
Here are the number of people who gave each response:
```
values(gss['polviews'])
```
I'll define `liberal` to be `True` for anyone whose response is "Extremely liberal", "Liberal", or "Slightly liberal".
```
liberal = (gss['polviews'] < 4)
```
Here are the number of `True` and `False` values:
```
values(liberal)
```
And the fraction of respondents who are "liberal".
```
liberal.mean()
```
If we choose a random person in this dataset, the probability they are liberal is about 27%.
## The probability function
To summarize what we have done so far:
* To represent a logical proposition like "this respondent is liberal", we are using a Boolean series, which contains the values `True` and `False`.
* To compute the probability that a proposition is true, we are using the `mean` function, which computes the fraction of `True` values in a series.
To make this computation more explicit, I'll define a function that takes a Boolean series and returns a probability:
```
def prob(A):
"""Computes the probability of a proposition, A.
A: Boolean series
returns: probability
"""
assert isinstance(A, pd.Series)
assert A.dtype == 'bool'
return A.mean()
```
The `assert` statements check whether `A` is a Boolean series. If not, they display an error message.
Using this function to compute probabilities makes the code more readable. Here are the probabilities for the propositions we have computed so far.
```
prob(banker)
prob(female)
prob(liberal)
```
**Exercise**: The values of `partyid` are encoded like this:
```
0 Strong democrat
1 Not str democrat
2 Ind,near dem
3 Independent
4 Ind,near rep
5 Not str republican
6 Strong republican
7 Other party
```
I'll define `democrat` to include respondents who chose "Strong democrat" or "Not str democrat":
```
democrat = (gss['partyid'] <= 1)
```
* Use `mean` to compute the fraction of Democrats in this dataset.
* Use `prob` to compute the same fraction, which we will think of as a probability.
```
# Solution
democrat.mean()
# Solution
prob(democrat)
```
## Conjunction
Now that we have a definition of probability and a function that computes it, let's move on to conjunction.
"Conjunction" is another name for the logical `and` operation. If you have two propositions, `A` and `B`, the conjunction `A and B` is `True` if both `A` and `B` are `True`, and `False` otherwise.
I'll demonstrate using two Boolean series constructed to enumerate every combination of `True` and `False`:
```
A = pd.Series((True, True, False, False))
A
B = pd.Series((True, False, True, False))
B
```
To compute the conjunction of `A` and `B`, we can use the `&` operator, like this:
```
A & B
```
The result is `True` only when `A` and `B` are `True`.
To show this operation more clearly, I'll put the operands and the result in a DataFrame:
```
table = pd.DataFrame()
table['A'] = A
table['B'] = B
table['A & B'] = A & B
table
```
This way of representing a logical operation is called a [truth table](https://en.wikipedia.org/wiki/Truth_table).
In a previous section, we computed the probability that a random respondent is a banker:
```
prob(banker)
```
And the probability that a respondent is a Democrat:
```
prob(democrat)
```
Now we can compute the probability that a random respondent is a banker *and* a Democrat:
```
prob(banker & democrat)
```
As we should expect, `prob(banker & democrat)` is less than `prob(banker)`, because not all bankers are Democrats.
**Exercise:** Use `prob` and the `&` operator to compute the following probabilities.
* What is the probability that a random respondent is a banker and liberal?
* What is the probability that a random respondent is female, a banker, and liberal?
* What is the probability that a random respondent is female, a banker, and a liberal Democrat?
Notice that as we add more conjunctions, the probabilities get smaller.
```
# Solution
prob(banker & liberal)
# Solution
prob(female & banker & liberal)
# Solution
prob(female & banker & liberal & democrat)
```
**Exercise:** We expect conjunction to be commutative; that is, `A & B` should be the same as `B & A`.
To check, compute these two probabilies:
* What is the probability that a random respondent is a banker and liberal?
* What is the probability that a random respondent is liberal and a banker?
```
prob(banker & liberal)
prob(liberal & banker)
```
If they are not the same, something has gone very wrong!
## Conditional probability
Conditional probability is a probability that depends on a condition, but that might not be the most helpful definition. Here are some examples:
* What is the probability that a respondent is a Democrat, given that they are liberal?
* What is the probability that a respondent is female, given that they are a banker?
* What is the probability that a respondent is liberal, given that they are female?
Let's start with the first one, which we can interpret like this: "Of all the respondents who are liberal, what fraction are Democrats?"
We can compute this probability in two steps:
1. Select all respondents who are liberal.
2. Compute the fraction of the selected respondents who are Democrats.
To select liberal respondents, we can use the bracket operator, `[]`, like this:
```
selected = democrat[liberal]
```
The result is a Boolean series that contains a subset of the values in `democrat`. Specifically, it contains only the values where `liberal` is `True`.
To confirm that, let's check the length of the result:
```
len(selected)
```
If things have gone according to plan, that should be the same as the number of `True` values in `liberal`:
```
liberal.sum()
```
Good.
`selected` contains the value of `democrat` for liberal respondents, so the mean of `selected` is the fraction of liberals who are Democrats:
```
selected.mean()
```
A little more than half of liberals are Democrats. If the result is lower than you expected, keep in mind:
1. We used a somewhat strict definition of "Democrat", excluding Independents who "lean" democratic.
2. The dataset includes respondents as far back as 1974; in the early part of this interval, there was less alignment between political views and party affiliation, compared to the present.
Let's try the second example, "What is the probability that a respondent is female, given that they are a banker?"
We can interpret that to mean, "Of all respondents who are bankers, what fraction are female?"
Again, we'll use the bracket operator to select only the bankers:
```
selected = female[banker]
len(selected)
```
As we've seen, there are 728 bankers in the dataset.
Now we can use `mean` to compute the conditional probability that a respondent is female, given that they are a banker:
```
selected.mean()
```
About 77% of the bankers in this dataset are female.
We can get the same result using `prob`:
```
prob(selected)
```
Remember that we defined `prob` to make the code easier to read. We can do the same thing with conditional probability.
I'll define `conditional` to take two Boolean series, `A` and `B`, and compute the conditional probability of `A` given `B`:
```
def conditional(A, B):
"""Conditional probability of A given B.
A: Boolean series
B: Boolean series
returns: probability
"""
return prob(A[B])
```
Now we can use it to compute the probability that a liberal is a Democrat:
```
conditional(democrat, liberal)
```
And the probability that a banker is female:
```
conditional(female, banker)
```
The results are the same as what we computed above.
**Exercise:** Use `conditional` to compute the probability that a respondent is liberal given that they are female.
Hint: The answer should be less than 30%. If your answer is about 54%, you have made a mistake (see the next exercise).
```
# Solution
conditional(liberal, female)
```
**Exercise:** In a previous exercise, we saw that conjunction is commutative; that is, `prob(A & B)` is always equal to `prob(B & A)`.
But conditional probability is NOT commutative; that is, `conditional(A, B)` is not the same as `conditional(B, A)`.
That should be clear if we look at an example. Previously, we computed the probability a respondent is female, given that they are banker.
```
conditional(female, banker)
```
The result shows that the majority of bankers are female. That is not the same as the probability that a respondent is a banker, given that they are female:
```
conditional(banker, female)
```
Only about 2% of female respondents are bankers.
**Exercise:** Use `conditional` to compute the following probabilities:
* What is the probability that a respondent is liberal, given that they are a Democrat?
* What is the probability that a respondent is a Democrat, given that they are liberal?
Think carefully about the order of the series you pass to `conditional`.
```
conditional(liberal, democrat)
conditional(democrat, liberal)
```
## Conditions and conjunctions
We can combine conditional probability and conjunction. For example, here's the probability a respondent is female, given that they are a liberal Democrat.
```
conditional(female, liberal & democrat)
```
Almost 57% of liberal Democrats are female.
And here's the probability they are a liberal female, given that they are a banker:
```
conditional(liberal & female, banker)
```
About 17% of bankers are liberal women.
**Exercise:** What fraction of female bankers are liberal Democrats?
Hint: If your answer is less than 1%, you have it backwards. Remember that conditional probability is not commutative.
```
# Solution
conditional(liberal & democrat, female & banker)
```
## Summary
At this point, you should understand the definition of probability, at least in the simple case where we have a finite dataset. Later we will consider cases where the definition of probability is more controversial.
And you should understand conjunction and conditional probability. [In the next notebook](https://colab.research.google.com/github/AllenDowney/BiteSizeBayes/blob/master/02_bayes.ipynb), we will explore the relationship between conjunction and conditional probability, and use it to derive Bayes's Theorem, which is the foundation of Bayesian statistics.
| github_jupyter |
```
import sys, os, zipfile
import urllib.request
pwd
#if you want to specify a particular location for file, give it here:
#where_to_save = '/Users/gw18g940/Desktop/ImageProcessingLectureData/'
#regular case
where_to_save = '../../Data/'
#create data directory
if not os.path.exists(where_to_save):
os.makedirs(where_to_save)
#import Klee painting
url = 'https://img.myswitzerland.com/671846/407'
urllib.request.urlretrieve(url, where_to_save+'Klee.jpg')
#import zebrafish embry
os.makedirs(where_to_save+'30567')
url = 'https://cildata.crbs.ucsd.edu/media/images/30567/30567.tif'
urllib.request.urlretrieve(url, where_to_save+'30567/30567.tif')
#import scifio wtembryo
os.makedirs(where_to_save+'2chZT')
url = 'https://samples.scif.io/2chZT.zip'
urllib.request.urlretrieve(url, where_to_save+'2chZT.zip')
#unzip
with zipfile.ZipFile(where_to_save+'2chZT.zip', 'r') as zip_ref:
zip_ref.extractall(where_to_save+'2chZT')
os.remove(where_to_save+'2chZT.zip')
#import landsat images
os.makedirs(where_to_save+'geography')
url = 'https://ndownloader.figshare.com/files/7677208'
urllib.request.urlretrieve(url, where_to_save+'geography.zip')
#unzip
with zipfile.ZipFile(where_to_save+'geography.zip', 'r') as zip_ref:
zip_ref.extractall(where_to_save+'geography')
os.remove(where_to_save+'geography.zip')
#import BBBC007
url = 'https://data.broadinstitute.org/bbbc/BBBC007/BBBC007_v1_images.zip'
urllib.request.urlretrieve(url, where_to_save+'BBBC007_v1_images.zip')
#unzip
with zipfile.ZipFile(where_to_save+'BBBC007_v1_images.zip', 'r') as zip_ref:
zip_ref.extractall(where_to_save)
os.remove(where_to_save+'BBBC007_v1_images.zip')
#import BBBC032
os.makedirs(where_to_save+'BBBC032_v1_dataset')
url = 'https://data.broadinstitute.org/bbbc/BBBC032/BBBC032_v1_dataset.zip'
urllib.request.urlretrieve(url, where_to_save+'BBBC032_v1_dataset.zip')
#unzip
with zipfile.ZipFile(where_to_save+'BBBC032_v1_dataset.zip', 'r') as zip_ref:
zip_ref.extractall(where_to_save+'BBBC032_v1_dataset')
os.remove(where_to_save+'BBBC032_v1_dataset.zip')
#import BBBC034
os.makedirs(where_to_save+'BBBC034_v1_dataset')
url = 'https://data.broadinstitute.org/bbbc/BBBC034/BBBC034_v1_dataset.zip'
urllib.request.urlretrieve(url, where_to_save+'BBBC034_v1_dataset.zip')
#unzip
with zipfile.ZipFile(where_to_save+'BBBC034_v1_dataset.zip', 'r') as zip_ref:
zip_ref.extractall(where_to_save+'BBBC034_v1_dataset')
os.remove(where_to_save+'BBBC034_v1_dataset.zip')
#import channels
os.makedirs(where_to_save+'channels')
url = 'https://drive.google.com/uc?id=1kNzXN_FkRflU4uNOpNfmpK8hUcJ1Dz6R'
urllib.request.urlretrieve(url, where_to_save+'channels/channels1.tif')
url = 'https://drive.google.com/uc?id=1OMBGdO3t_RvCIcmTLPX6zBfRiWt5KP3Z'
urllib.request.urlretrieve(url, where_to_save+'channels/channels2.tif')
#create a share folder and move data there
!sudo mkdir -p /srv/data/
!sudo mv '../../Data' '/srv/data/'
!sudo ln -s /srv/data/Data /etc/skel/Data
```
| github_jupyter |
```
# ------------------------------------------------------------------------------
# An example of training and evaluating a model for the prostate dataset in the
# Medical Segmentation Decathlon. To execute, download the data for task 5 from
# http://medicaldecathlon.com/ and specify the path in mp.paths.py.
# ------------------------------------------------------------------------------
# 1. Imports
# Reload changes modules
from IPython import get_ipython
get_ipython().magic('load_ext autoreload')
get_ipython().magic('autoreload 2')
# Import modules
import torch
from torch.utils.data import DataLoader
import torch.optim as optim
from mp.experiments.experiment import Experiment
from mp.data.data import Data
from mp.data.datasets.ds_mr_prostate_decathlon import DecathlonProstateT2
import mp.visualization.visualize_imgs as vis
from mp.data.pytorch.pytorch_seg_dataset import PytorchSeg2DDataset
from mp.models.segmentation.unet_fepegar import UNet2D
from mp.eval.losses.losses_segmentation import LossClassWeighted, LossDiceBCE
from mp.agents.segmentation_agent import SegmentationAgent
from mp.eval.result import Result
from mp.utils.load_restore import nifty_dump
# 2. Define configuration
# The configuration dictionary boundles training parameters and anything
# that can be manually defined. The config file is stored within the directory
# created for a given experiment. By specifying arguments in config files,
# higher-level modules don't need to be adapted for hyperparameter searches
# and experiments are more easily reproducible.
config = {'experiment_name':'test_exp', 'device':'cuda:0',
'nr_runs': 1, 'cross_validation': False, 'val_ratio': 0.0, 'test_ratio': 0.3,
'input_shape': (1, 256, 256), 'resize': False, 'augmentation': 'none',
'class_weights': (0.,1.), 'lr': 0.0001, 'batch_size': 8
}
device = config['device']
device_name = torch.cuda.get_device_name(device)
print('Device name: {}'.format(device_name))
input_shape = config['input_shape']
# 3. Create experiment directories
# Initializing an experiment creates a directory exp/<exp name> in the
# storage directory defined in mp.paths.py. All files created during that
# experiment are stored there.
exp = Experiment(config=config, name=config['experiment_name'], notes='', reload_exp=True)
# 4. Define data
# A data object can be initialized with multiple datasets. The idea behind this
# is to more easily test with o.o.d data, or simulate continual learning.
# In this example, we only add one dataset.
data = Data()
data.add_dataset(DecathlonProstateT2(merge_labels=True))
nr_labels = data.nr_labels
label_names = data.label_names
train_ds = ('DecathlonProstateT2', 'train')
test_ds = ('DecathlonProstateT2', 'test')
# 5. Create data splits for each repetition
# For each dataset, the instance indexes are divided into train, validation
# and test sets. The values specified in the config are used to determine
# the number of runs and how indexes are divided. For more details, look at the
# method definition for 'set_data_splits'. The index splitting is also stored
# within the experiment directory. For each repetition, a subdirectory is
# created named after the repetition index.
exp.set_data_splits(data)
# Get the experiment run respective to the first data split. This would usually
# be repeated for all 'nr_runs' runs
exp_run = exp.get_run(run_ix=0)
# 6. Bring data to Pytorch format
# Transform data to PyTorch format
datasets = dict()
for ds_name, ds in data.datasets.items():
for split, data_ixs in exp.splits[ds_name][exp_run.run_ix].items():
if len(data_ixs) > 0: # Sometimes val indexes may be an empty list
aug = config['augmentation'] if not('test' in split) else 'none'
datasets[(ds_name, split)] = PytorchSeg2DDataset(ds,
ix_lst=data_ixs, size=input_shape, aug_key=aug,
resize=config['resize'])
# Visualize the first instance from the training data
subject_ix = 0
subject = datasets[train_ds].instances[subject_ix].get_subject()
vis.plot_3d_subject_gt(subject)
# 7. Build train dataloader, and visualize
dl = DataLoader(datasets[(train_ds)],
batch_size=config['batch_size'], shuffle=True)
vis.visualize_dataloader_with_masks(dl, img_size=(128, 128))
# 8. Initialize model
model = UNet2D(input_shape, nr_labels)
model.to(device)
# 9. Define loss and optimizer
# Define loss and optimizer. In this case, the loss is a combination of Dice
# and binary cross-entropy, weighted so that only the dice on the 'prostate'
# class is considered and not that for 'background'
loss_g = LossDiceBCE(bce_weight=1., smooth=1., device=device)
loss_f = LossClassWeighted(loss=loss_g, weights=config['class_weights'],
device=device)
optimizer = optim.Adam(model.parameters(), lr=config['lr'])
# 10. Train model
# We train here only for 10 epochs for testing purposes, altough in practice
# many more are necessary to reach convergence
results = Result(name='training_trajectory')
agent = SegmentationAgent(model=model, label_names=label_names, device=device)
agent.train(results, optimizer, loss_f, train_dataloader=dl,
init_epoch=0, nr_epochs=10, run_loss_print_interval=5,
eval_datasets=datasets, eval_interval=5,
save_path=exp_run.paths['states'], save_interval=5)
# 11. Save and print results for this experiment run
exp_run.finish(results=results, plot_metrics=['Mean_ScoreDice', 'Mean_ScoreDice[prostate]'])
test_ds_key = '_'.join(test_ds)
metric = 'Mean_ScoreDice[prostate]'
last_dice = results.get_epoch_metric(
results.get_max_epoch(metric, data=test_ds_key), metric, data=test_ds_key)
print('Last Dice score for prostate class: {}'.format(last_dice))
# 12. Visualize results qualitatively
# Visualize result for the first subject in the test dataset
subject_ix = 0
subject = datasets[test_ds].instances[subject_ix].get_subject()
pred = datasets[test_ds].predictor.get_subject_prediction(agent, subject_ix)
vis.plot_3d_subject_pred(subject, pred)
```
| github_jupyter |
# Multivariate Linear Regression
$ h_\theta(x) = \theta_0 * x_0 + \theta_1 * x_1 + \theta_2 * x_2 + ... + \theta_n * x_n $
$ x_0 = 1$
## Cost function
$ J(\theta) = \frac{1}{2m} \sum_{i=1}^{m} (h_\theta(x^{(i)}) - y^{(i)})^2 $
## Gradient descent
repeat until convergence {
>$ \theta_j = \theta_j - \alpha \frac{\partial}{\partial \theta_j} J(\theta) $
}
$ \frac{\partial}{\partial \theta_j} J(\theta) = \frac{1}{m} \sum_{i=1}^{m} (h_\theta(x^{(i)}) - y^{(i)}) x_j^{(i)} $
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
def std_normalize(axis):
"""Rescale numpy 1-D array between <0, 1>"""
a1 = axis.A1
a_mean = np.mean(a1)
a_std = np.std(a1)
a_prim = list(map(lambda x: 1.0 * (x-a_mean) / a_std, a1))
return a_prim, a_mean, a_std
# We will use simpler version when applying to matrix
def std_n(axis):
a_prim, mean_a, std_a = std_normalize(axis)
return a_prim
```
## Hypothesis
$ h_\theta(x) = \theta_0 * x_0 + \theta_1 * x_1 + \theta_2 * x_2 $
,where:
$ x_0 = 1 $ (used to make math easier)
$ x_1 $ - Weight (kilograms)
$ x_2 $ - Age (years)
| Weight (kilograms) | Age (years) | Blood fat content |
|--------------------|-------------|-------------------|
| 84 | 46 | 354 |
| 73 | 20 | 190 |
| 65 | 52 | 405 |
| 70 | 30 | 263 |
| 76 | 57 | 451 |
| 69 | 25 | 302 |
| 63 | 28 | 288 |
| 72 | 36 | 385 |
| 79 | 57 | 402 |
| 75 | 44 | 365 |
| 27 | 24 | 209 |
| 89 | 31 | 290 |
| 65 | 52 | 346 |
| 57 | 23 | 254 |
| 59 | 60 | 395 |
| 69 | 48 | 434 |
| 60 | 34 | 220 |
| 79 | 51 | 374 |
| 75 | 50 | 308 |
| 82 | 34 | 220 |
| 59 | 46 | 311 |
| 67 | 23 | 181 |
| 85 | 37 | 274 |
| 55 | 40 | 303 |
| 63 | 30 | 244 |
```
X = np.matrix([[84, 46],
[73, 20],
[65, 52],
[70, 30],
[76, 57],
[69, 25],
[63, 28],
[72, 36],
[79, 57],
[75, 44],
[27, 24],
[89, 31],
[65, 52],
[57, 23],
[59, 60],
[69, 48],
[60, 34],
[79, 51],
[75, 50],
[82, 34],
[59, 46],
[67, 23],
[85, 37],
[55, 40],
[63, 30]])
# We need to store mean and standard deviation value
_, WEIGHT_MEAN, WEIGHT_STD = std_normalize(X.T[0])
_, AGE_MEAN, AGE_STD = std_normalize(X.T[1])
# Rescale X values
X = np.apply_along_axis(std_n, axis=0, arr=X)
X.shape
Y = np.matrix('354; 190; 405; 263; 451; 302; 288; 385; 402; 365; 209; 290; 346; 254; 395; 434; 220; 374; 308; 220; 311; 181; 274; 303; 244')
Y.shape
theta = np.matrix([[0.1], [0.1], [0.1]])
theta.shape
# Before we start, we need to add extra column for x_0
# It will allow us to use matrix multiplication while
# calculating our hypothesis
ones = np.ones((X.shape[0], 1))
X = np.append(ones, X, axis=1)
X.shape
# Initial hypothesis
h_theta = X * theta
h_theta.shape
# Number of training examples
m = X.shape[0]
# Initial cost
J = np.sum(np.power(h_theta - Y, 2)) * 1/(2*m)
J
alpha = 0.01
J_arr = []
n = 0
while J > 0.001 and n < 10000:
J_arr.append(J)
t = alpha * ((h_theta - Y).T.dot(X)) * 1/m
theta = theta - t.T
h_theta = X * theta
J = np.sum(np.power(h_theta - Y, 2)) * 1/(2*m)
n += 1
theta
plt.plot(xrange(len(J_arr)), J_arr, 'g')
plt.grid(True)
plt.xlabel('Iterations')
plt.ylabel('Cost')
plt.title('Cost function')
plt.show()
print('Final cost = {0}'.format(J_arr[-1]))
```
Now, we can predict value using learned $\theta$ params.
New features must be scaled with saved mean value and standard deviation.
```
def predict(weight, age, theta):
w = 1.0 * (weight-WEIGHT_MEAN) / WEIGHT_STD
a = 1.0 * (age-AGE_MEAN) / AGE_STD
a = np.array([1, w, a])
return round(np.dot(a, theta.A1), 2)
data = [(55, 21), (65, 31), (75, 41), (85, 51), (95, 61)]
for w, a in data:
predicted_fat = predict(weight=w, age=a, theta=theta)
print('Weight={0}; Age={1}; Predicted blood fat content={2}'.format(w, a, predicted_fat))
```
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Unredacted Mueller Report with TensorFlow
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.manceps.com"><img src="https://github.com/manceps/brand/raw/master/logo/Manceps_Logo_3c_32p.jpeg" /> Assembled by the Manceps team</a>
</td>
<td>
<a target="_blank" href="https://www.tensorflow.org/tutorials/sequences/text_generation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" /> Based on Text Generation with a RNN</a>
</td>
<td> <p>Preprocessed dataset and concept by </p>
<a target="_blank" href="http://www.viralml.com">Manuel Amunategui</a>
and
<a target="_blank" href="https://www.kaggle.com/paultimothymooney"> Paul Mooney</a>
</td>
</table>
With the largely redacted release of the [Robert Mueller's Report](https://www.justice.gov/storage/report.pdf) on the investigation into Russian interference in the 2016 Presidential Election, we will use Tensorflow to predict the missing text using a character-based RNN. Given a sequence of characters from this dataset, we'll train a model to predict the next character in the sequence ("e"). Longer sequences of text can be generated by calling the model repeatedly.
Note: Enable GPU acceleration to execute this notebook faster. In Colab: *Runtime > Change runtime type > Hardware acclerator > GPU*. If running locally make sure TensorFlow version >= 1.11.
This tutorial includes runnable code implemented using [tf.keras](https://www.tensorflow.org/programmers_guide/keras) and [eager execution](https://www.tensorflow.org/programmers_guide/eager).
The following is sample output when the model in this tutorial trained for 30 epochs, and started with the string "Potential Section 1030 Violation By ":
<pre>
Potential Section 1030 Violation By 11, 2016, Saunders began receiving Page to and this relationshics with Ambassador Arransky (aguiliety Trump, Vorga, Dvorkovich at an underlying conduct
</pre>
While some of the sentences are grammatical, most do not make sense. The model has not learned the meaning of words, but consider:
* The model is character-based. When training started, the model did not know how to spell an English word, or that words were even a unit of text.
* The structure of the output resembles a play—blocks of text generally begin with a speaker name, in all capital letters similar to the dataset.
* As demonstrated below, the model is trained on small batches of text (100 characters each), and is still able to generate a longer sequence of text with coherent structure.
## Setup
### Mount your Google Drive to save your model
To authenticate, the link generated will open in another page. Login with your Google credentials and authorize this notebook, then copy the code generated back in the space below..
```
from google.colab import drive
drive.mount('/content/gdrive')
```
Let's keep tidy in a separate directory in Google Drive
```
import os
os.chdir('/content/gdrive/My Drive')
os.makedirs('mueller', exist_ok=True)
os.chdir('mueller')
# Directory where the checkpoints will be saved
checkpoint_dir = './training_checkpoints'
```
### Import TensorFlow and other libraries
```
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
tf.enable_eager_execution()
import numpy as np
import time
```
### Download the preprocessed Mueller Report dataset
```
path_to_file = tf.keras.utils.get_file('mueller.txt', 'http://www.viralml.com/static/code/mueller.txt')
```
### Read the data
First, look in the text.
```
# Read, then decode for py2 compat.
text = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# length of text is the number of characters in it
print ('Length of text: {} characters'.format(len(text)))
# Take a look at some of the text
print(text[123:208])
# The unique characters in the file
vocab = sorted(set(text))
print ('{} unique characters'.format(len(vocab)))
```
## Process the text
### Vectorize the text
Before training, we need to map strings to a numerical representation. Create two lookup tables: one mapping characters to numbers, and another for numbers to characters.
```
# Creating a mapping from unique characters to indices
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
text_as_int = np.array([char2idx[c] for c in text])
```
Now we have an integer representation for each character. Notice that we mapped the character as indexes from 0 to `len(unique)`.
```
print('{')
for char,_ in zip(char2idx, range(20)):
print(' {:4s}: {:3d},'.format(repr(char), char2idx[char]))
print(' ...\n}')
# Show how the first few characters from the text are mapped to integers
print ('{} ---- characters mapped to int ---- > {}'.format(repr(text[123:151]), text_as_int[123:151]))
```
### The prediction task
Given a character, or a sequence of characters, what is the most probable next character? This is the task we're training the model to perform. The input to the model will be a sequence of characters, and we train the model to predict the output—the following character at each time step.
Since RNNs maintain an internal state that depends on the previously seen elements, given all the characters computed until this moment, what is the next character?
### Create training examples and targets
Next divide the text into example sequences. Each input sequence will contain `seq_length` characters from the text.
For each input sequence, the corresponding targets contain the same length of text, except shifted one character to the right.
So break the text into chunks of `seq_length+1`. For example, say `seq_length` is 4 and our text is "Hello". The input sequence would be "Hell", and the target sequence "ello".
To do this first use the `tf.data.Dataset.from_tensor_slices` function to convert the text vector into a stream of character indices.
```
# The maximum length sentence we want for a single input in characters
seq_length = 100
examples_per_epoch = int(len(text_as_int) / seq_length)
def show_text_as_ints(text_as_int=text_as_int):
# Create training examples / targets
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
for i in char_dataset.take(5):
print(idx2char[i.numpy()])
show_text_as_ints()
```
The `batch` method lets us easily convert these individual characters to sequences of the desired size.
```
sequences = char_dataset.batch(seq_length+1, drop_remainder=True)
for item in sequences.take(5):
print(repr(''.join(idx2char[item.numpy()])))
```
For each sequence, duplicate and shift it to form the input and target text by using the `map` method to apply a simple function to each batch:
```
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
dataset = sequences.map(split_input_target)
```
Print the first examples input and target values:
```
for input_example, target_example in dataset.take(1):
print ('Input data: ', repr(''.join(idx2char[input_example.numpy()])))
print ('Target data:', repr(''.join(idx2char[target_example.numpy()])))
```
Each index of these vectors are processed as one time step. For the input at time step 0, the model receives the index for "F" and tries to predict the index for "i" as the next character. At the next timestep, it does the same thing but the `RNN` considers the previous step context in addition to the current input character.
```
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {:4d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
```
### Create training batches
We used `tf.data` to split the text into manageable sequences. But before feeding this data into the model, we need to shuffle the data and pack it into batches.
```
# Batch size
BATCH_SIZE = 64
steps_per_epoch = examples_per_epoch//BATCH_SIZE
# Buffer size to shuffle the dataset
# (TF data is designed to work with possibly infinite sequences,
# so it doesn't attempt to shuffle the entire sequence in memory. Instead,
# it maintains a buffer in which it shuffles elements).
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
dataset
```
## Build The Model
Use `tf.keras.Sequential` to define the model. For this simple example three layers are used to define our model:
* `tf.keras.layers.Embedding`: The input layer. A trainable lookup table that will map the numbers of each character to a vector with `embedding_dim` dimensions;
* `tf.keras.layers.GRU`: A type of RNN with size `units=rnn_units` (You can also use a LSTM layer here.)
* `tf.keras.layers.Dense`: The output layer, with `vocab_size` outputs.
```
# Length of the vocabulary in chars
vocab_size = len(vocab)
# The embedding dimension
embedding_dim = 256
# Number of RNN units
rnn_units = 1024
```
Next define a function to build the model.
Use `CuDNNGRU` if running on GPU.
```
if tf.test.is_gpu_available():
rnn = tf.keras.layers.CuDNNGRU
else:
import functools
rnn = functools.partial(
tf.keras.layers.GRU, recurrent_activation='sigmoid')
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
rnn(rnn_units,
return_sequences=True,
recurrent_initializer='glorot_uniform',
stateful=True),
tf.keras.layers.Dense(vocab_size)
])
return model
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
```
For each character the model looks up the embedding, runs the GRU one timestep with the embedding as input, and applies the dense layer to generate logits predicting the log-liklihood of the next character:

## Try the model
Now run the model to see that it behaves as expected.
First check the shape of the output:
```
for input_example_batch, target_example_batch in dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(example_batch_predictions.shape, "# (batch_size, sequence_length, vocab_size)")
```
In the above example the sequence length of the input is `100` but the model can be run on inputs of any length:
```
model.summary()
```
To get actual predictions from the model we need to sample from the output distribution, to get actual character indices. This distribution is defined by the logits over the character vocabulary.
Note: It is important to _sample_ from this distribution as taking the _argmax_ of the distribution can easily get the model stuck in a loop.
Try it for the first example in the batch:
```
sampled_indices = tf.random.categorical(example_batch_predictions[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices,axis=-1).numpy()
```
This gives us, at each timestep, a prediction of the next character index:
```
sampled_indices
```
Decode these to see the text predicted by this untrained model:
```
print("Input: \n", repr("".join(idx2char[input_example_batch[0]])))
print()
print("Next Char Predictions: \n", repr("".join(idx2char[sampled_indices ])))
```
## Train the model
At this point the problem can be treated as a standard classification problem. Given the previous RNN state, and the input this time step, predict the class of the next character.
### Attach an optimizer, and a loss function
The standard `tf.keras.losses.sparse_softmax_crossentropy` loss function works in this case because it is applied across the last dimension of the predictions.
Because our model returns logits, we need to set the `from_logits` flag.
```
def loss(labels, logits):
return tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True)
example_batch_loss = loss(target_example_batch, example_batch_predictions)
print("Prediction shape: ", example_batch_predictions.shape, " # (batch_size, sequence_length, vocab_size)")
print("scalar_loss: ", example_batch_loss.numpy().mean())
```
Configure the training procedure using the `tf.keras.Model.compile` method. We'll use `tf.train.AdamOptimizer` with default arguments and the loss function.
```
model.compile(
optimizer = tf.train.AdamOptimizer(),
loss = loss)
```
### Configure checkpoints
Use a `tf.keras.callbacks.ModelCheckpoint` to ensure that checkpoints are saved during training:
```
# Name of the checkpoint files
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
```
### Execute the training
To keep training time reasonable, use 30 epochs to train the model. In Colab, set the runtime to GPU for faster training.
```
EPOCHS=30
history = model.fit(dataset.repeat(), epochs=EPOCHS, steps_per_epoch=steps_per_epoch, callbacks=[checkpoint_callback])
```
## Generate text
Time to have some fun. We need a few words right before a redacted section to predict what's concealed.
### Restore the latest checkpoint
To keep this prediction step simple, use a batch size of 1.
Because of the way the RNN state is passed from timestep to timestep, the model only accepts a fixed batch size once built.
To run the model with a different `batch_size`, we need to rebuild the model and restore the weights from the checkpoint.
```
tf.train.latest_checkpoint(checkpoint_dir)
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model.build(tf.TensorShape([1, None]))
model.summary()
```
### The prediction loop
The following code block generates the text:
* It Starts by choosing a start string, initializing the RNN state and setting the number of characters to generate.
* Get the prediction distribution of the next character using the start string and the RNN state.
* Then, use a multinomial distribution to calculate the index of the predicted character. Use this predicted character as our next input to the model.
* The RNN state returned by the model is fed back into the model so that it now has more context, instead than only one word. After predicting the next word, the modified RNN states are again fed back into the model, which is how it learns as it gets more context from the previously predicted words.

Looking at the generated text, you'll see the model knows when to capitalize, make paragraphs and imitates original text writing vocabulary. With the small number of training epochs, it has not yet learned to form coherent sentences.
```
def generate_text(model, start_string):
# Evaluation step (generating text using the learned model)
# Number of characters to generate
num_generate = 200
# Converting our start string to numbers (vectorizing)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Low temperatures results in more predictable text.
# Higher temperatures results in more surprising text.
# Experiment to find the best setting.
temperature = 1.0
# Here batch size == 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# remove the batch dimension
predictions = tf.squeeze(predictions, 0)
# using a multinomial distribution to predict the word returned by the model
predictions = predictions / temperature
predicted_id = tf.multinomial(predictions, num_samples=1)[-1,0].numpy()
# We pass the predicted word as the next input to the model
# along with the previous hidden state
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
start_string = u"Potential Section 1030 Violation By "
print(generate_text(model, start_string))
```
The easiest thing you can do to improve the results it to train it for longer (try `EPOCHS=30`).
You can also experiment with a different start string, or try adding another RNN layer to improve the model's accuracy, or adjusting the temperature parameter to generate more or less random predictions.
## Advanced: Customized Training
The above training procedure is simple, but does not give you much control.
So now that you've seen how to run the model manually let's unpack the training loop, and implement it ourselves. This gives a starting point if, for example, to implement _curriculum learning_ to help stabilize the model's open-loop output.
We will use `tf.GradientTape` to track the gradiends. You can learn more about this approach by reading the [eager execution guide](https://www.tensorflow.org/guide/eager).
The procedure works as follows:
* First, initialize the RNN state. We do this by calling the `tf.keras.Model.reset_states` method.
* Next, iterate over the dataset (batch by batch) and calculate the *predictions* associated with each.
* Open a `tf.GradientTape`, and calculate the predictions and loss in that context.
* Calculate the gradients of the loss with respect to the model variables using the `tf.GradientTape.grads` method.
* Finally, take a step downwards by using the optimizer's `tf.train.Optimizer.apply_gradients` method.
```
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
optimizer = tf.train.AdamOptimizer()
# Training step
EPOCHS = 30
for epoch in range(EPOCHS):
start = time.time()
# initializing the hidden state at the start of every epoch
# initally hidden is None
hidden = model.reset_states()
for (batch_n, (inp, target)) in enumerate(dataset):
with tf.GradientTape() as tape:
# feeding the hidden state back into the model
# This is the interesting step
predictions = model(inp)
loss = tf.losses.sparse_softmax_cross_entropy(target, predictions)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
if batch_n % 100 == 0:
template = 'Epoch {} Batch {} Loss {:.4f}'
print(template.format(epoch+1, batch_n, loss))
# saving (checkpoint) the model every 5 epochs
if (epoch + 1) % 5 == 0:
model.save_weights(checkpoint_prefix.format(epoch=epoch))
print ('Epoch {} Loss {:.4f}'.format(epoch+1, loss))
print ('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
model.save_weights(checkpoint_prefix.format(epoch=epoch))
```
| github_jupyter |
# Tuning
This notebook will demonstrate how to do tuning for models and controllers.
## Set-Up
As always, we begin by obtaining our system, model, and task from the benchmarks package.
```
import autompc as ampc
import numpy as np
from autompc.benchmarks import CartpoleSwingupBenchmark
benchmark = CartpoleSwingupBenchmark()
# Get system and task specification
system = benchmark.system
task = benchmark.task
# Generate benchmark dataset
trajs = benchmark.gen_trajs(seed=100, n_trajs=500, traj_len=200)
```
## Model Tuning
First, we will demonstrate how to automatically tune the hyperparameters of a system ID model.
First, we have to define the model evaluator, which handles the training and evaluation of individual model configurations. Here, use the `HoldoutEvaluator` which randomly splits the dataset into a training set and holdout set for evaluation. We will tune using the `RMSE` metric over a 20 step prediction horizon. We also have to provide the evaluator with the trajectory dataset.
```
from autompc.evaluation import HoldoutModelEvaluator
model_evaluator = HoldoutModelEvaluator(holdout_prop=0.25, metric="rmse", horizon=20, trajs=trajs,
system=system, rng=np.random.default_rng(100))
```
We also need to construct a model factory to tune. Model tuning also supports automatic model selection, so here will create two model factories to select between
```
from autompc.sysid import MLPFactory, SINDyFactory
model_factory_1 = MLPFactory(system)
model_factory_2 = SINDyFactory(system)
```
Now that we have created our evaluator and our factories, we instantiate the `ModelTuner` and add both factories.
```
from autompc.tuning import ModelTuner
model_tuner = ModelTuner(system, model_evaluator)
model_tuner.add_model_factory(model_factory_1)
model_tuner.add_model_factory(model_factory_2)
```
We now run tuning for 100 iterations
```
model, model_tune_result = model_tuner.run(rng=np.random.default_rng(100), n_iters=100)
```
We can then plot the accuracy of the best model found so far over the course of the tuning process.
```
from autompc.graphs import TuningCurveGraph
import matplotlib.pyplot as plt
graph = TuningCurveGraph()
fig = plt.figure()
ax = fig.gca()
graph(ax, model_tune_result)
ax.set_title("Model Tuning Curve")
plt.show()
```
## Pipeline Tuning
Next, we will demonstrate how to do hyperparameter tuning for the entire MPC pipeline. First, we create our factories and pipeline object. Here, we use an MLP system ID model, Iterative LQR control optimizer, and a quadratic cost. AutoMPC does not currently support automatic selection of pipeline components, but we hope to add this feature soon.
```
from autompc.control import IterativeLQRFactory
from autompc.costs import QuadCostFactory
ctrlr_factory = IterativeLQRFactory(system)
cost_factory = QuadCostFactory(system)
model_factory = MLPFactory(system)
pipeline = ampc.Pipeline(system, model_factory, cost_factory, ctrlr_factory)
```
Next, we instantiate the `PipelineTuner`. AutoMPC performs tuning without access to the system dynamics, so the tuner has to train a surrogate dynamics model to use a simulator for controller evaluation. In this example, we use an MLP surrogate model, so we pass in `MLPFactory`. The `surrogate_split` tells what proportion of the data to use for surrogate training vs system ID training. Here we use a 50/50 split.
Finally, AutoMPC supports several methods of selecting the surrogate model, controller by the `surrogate_mode` argument. Here we use `defaultcfg` which means the surrogate is trained using the default MLP configuration. Other modes include `fixedcfg`, where the user specifies the surrogate configuration, `autotune`, where the tuner first tunes the surrogate factory before running the pipeline tuning, and `autoselect`, where the tuner both automatically selects the model type from the availble system ID algorithms and tunes the model hyperparameters.
For more details on using these modes, see the [documentation](https://autompc.readthedocs.io/en/latest/source/tuning.html#pipelinetuner) for the `PipelineTuner` class.
```
from autompc.tuning import PipelineTuner
tuner = PipelineTuner(surrogate_factory=MLPFactory(system), surrogate_mode="defaultcfg", surrogate_split=0.5)
```
Now that we have created the tuner, we run it for 100 iterations. We pass in the ground truth dynamics to keep track of the performance, but in a real application, we don't expect to have access to this, so this information is not used for tuning.
```
controller, tune_result = tuner.run(pipeline, task, trajs, n_iters=100, rng=np.random.default_rng(100),
truedyn=benchmark.dynamics)
```
After running the tuning process, we can graph the performance of the best controller found so far, both with respect to the surrogate dynamics and the true dynamics
```
from autompc.graphs import TuningCurveGraph
import matplotlib.pyplot as plt
graph = TuningCurveGraph()
fig = plt.figure()
ax = fig.gca()
graph(ax, tune_result)
ax.set_title("Cart-Pole Tuning Curve")
plt.show()
```
## Decoupled Tuning
The above example is full pipeline tuning, which searches the configuration space of all components simultaneously. Alternatively, we can take a decoupled tuning approach, where the model is first tuned based on prediction accuracy, then the objective function and optimizer are tuned. Since full pipeline tuning requires us to train a system ID model and simulate the controller at every iteration, decoupled tuning may produce faster tuning.
First, we'll need to handle the data split between system ID training set and surrogate training set manually.
```
rng = np.random.default_rng(100)
rng.shuffle(trajs)
surr_trajs = trajs[:250]
sysid_trajs = trajs[250:]
```
Now, we tune our system ID model for 75 iterations using the same model tuning method as above
```
model_evaluator = HoldoutModelEvaluator(holdout_prop=0.25, metric="rmse", horizon=20, trajs=sysid_trajs,
system=system, rng=np.random.default_rng(100))
model_tuner = ModelTuner(system, model_evaluator)
model_tuner.add_model_factory(model_factory)
model, model_tune_result = model_tuner.run(rng=np.random.default_rng(100), n_iters=75)
```
Now, we construct a new pipeline which fixes the model to be the result of the tuning process we just ran
```
pipeline_fixed_model = ampc.Pipeline(system, model, ctrlr_factory, cost_factory)
```
And finally, we run our pipeline tuning in a similar manner to above. We now use tell the tuner to use 100% of data for tuning the surrogate, since we already handled the data split and did the system ID training
```
tuner2 = PipelineTuner(surrogate_factory=MLPFactory(system), surrogate_split=1.0)
controller2, tune_result2 = tuner2.run(pipeline_fixed_model, task, surr_trajs, n_iters=75,
rng=np.random.default_rng(100), truedyn=benchmark.dynamics)
```
Finally, we can plot the result of our decoupled pipeline tune
```
from autompc.graphs import TuningCurveGraph
import matplotlib.pyplot as plt
graph = TuningCurveGraph()
fig = plt.figure()
ax = fig.gca()
graph(ax, tune_result2)
ax.set_title("Cart-Pole Decoupled Tuning Curve")
plt.show()
```
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
b_times = np.load('transit_times_b.npy')[0, :]
b_uncertainties = np.load('transit_times_b.npy')[1, :]
b_epoch = np.arange(len(b_times))
c_times = np.load('transit_times_c.npy')[0, :]
c_uncertainties = np.load('transit_times_c.npy')[1, :]
c_epoch = np.arange(len(c_times))
plt.errorbar(b_times, 24*60*(b_times - np.polyval(np.polyfit(b_epoch, b_times, 1), b_epoch)), 24*60*b_uncertainties)
plt.errorbar(c_times, 24*60*(c_times - np.polyval(np.polyfit(c_epoch, c_times, 1), c_epoch)), 24*60*c_uncertainties)
plt.xlabel('TJD')
plt.ylabel('O-C [min]')
from ttvfast import ttvfast
from ttvfast.models import Planet
from astropy.constants import M_sun, M_earth
import numpy as np
import astropy.units as u
period_b_init = 4.653811
period_c_init = 9.150607
mass_b_init = 1 * float(M_earth/M_sun)
mass_c_init = 1 * float(M_earth/M_sun)
# b
a = 12.10
b = 0.27
inclination_b = np.degrees(np.arccos(b/a))
# c
a = 25.12
b = 0.22
inclination_c = np.degrees(np.arccos(b/a))
# mass_b_init_err = 0.154 * float(M_earth/M_sun)
# mass_c_init_err = 0.142 * float(M_earth/M_sun)
phase = 360 * (((b_times.min() - c_times.min())/period_c_init) % 1 )
b = Planet(mass=mass_b_init, period=period_b_init, eccentricity=0, inclination=inclination_b,
longnode=0, argument=90, mean_anomaly=0)
c = Planet(mass=mass_c_init, period=period_c_init, eccentricity=0, inclination=inclination_c,
longnode=0, argument=90, mean_anomaly=phase)
result = ttvfast(planets=[b, c], stellar_mass=0.90, time=b_times.min(), dt=period_b_init/100, total=b_times.min()+1000)
planet_ind, epoch, times, rsky, vsky = [np.array(i) for i in result['positions']]
condition = (times != -2)
planet_ind, epoch, times, rsky, vsky = [i[condition] for i in [planet_ind, epoch, times, rsky, vsky]]
indices_b = (planet_ind == 0) & np.in1d(epoch, b_epoch)
indices_c = (planet_ind == 1) & np.in1d(epoch, c_epoch)
epoch_c = epoch[indices_c]
model_times_b = times[indices_b]
model_times_c = times[indices_c]
# logchi2_e = -0.5 * np.sum( (model_times_e - times_e)**2 / errs_e**2 )
# logchi2_f = -0.5 * np.sum( (model_times_f - times_f)**2 / errs_f**2 )
model_times_b - b_times, model_times_c - c_times
import emcee
from corner import corner
from astropy.constants import G, M_sun
M_starM_sun = 1.0
M_star = M_starM_sun * M_sun
a_b = ( ( (G * M_star)/(4 * np.pi**2) * (period_b_init*u.day)**2 )**(1/3) ).decompose()
a_c = ( ( (G * M_star)/(4 * np.pi**2) * (period_c_init*u.day)**2 )**(1/3) ).decompose()
def lnprob(p):
mass_b, mass_c, period_b, period_c, lam_b, lam_c, w_b, w_c = p
if (mass_b < 0 or mass_c < 0 or mass_b > 1e-5 or mass_c > 1e-5):# or
#ecc_b < 0 or ecc_b > 0.01 or ecc_c < 0 or ecc_c > 0.01):
return -np.inf
ecc_b = 0
ecc_c = 0
# Hill stability criterion from Gladman 1993 Eqn. 21
gamma1 = np.sqrt(1 - ecc_b**2)
gamma2 = np.sqrt(1 - ecc_c**2)
mu1 = mass_b * M_starM_sun
mu2 = mass_c * M_starM_sun
alpha = mu1 + mu2
Delta = (a_c - a_b)/a_b
delta = np.sqrt(1 + Delta)
if (alpha**-3 * (mu1 + mu2/delta**2)*(mu1*gamma1 + mu2*gamma2*delta)**2 <=
1 + 3**(4/3) * (mu1 * mu2) / alpha**(4/3)):
return -np.inf
b = Planet(mass=mass_b_init, period=period_b_init, eccentricity=0, inclination=inclination_b,
longnode=0, argument=90, mean_anomaly=0)
c = Planet(mass=mass_c_init, period=period_c_init, eccentricity=0, inclination=inclination_c,
longnode=0, argument=90, mean_anomaly=phase)
result = ttvfast(planets=[b, c], stellar_mass=1, time=b_times.min(), dt=period_b_init/100, total=b_times.min()+1000)
planet_ind, epoch, times, rsky, vsky = [np.array(i) for i in result['positions']]
condition = (times != -2)
planet_ind, epoch, times, rsky, vsky = [i[condition] for i in [planet_ind, epoch, times, rsky, vsky]]
# Parse results for planets e and f
indices_b = (planet_ind == 0) & np.in1d(epoch, b_epoch)
indices_c = (planet_ind == 1) & np.in1d(epoch, c_epoch)
epoch_b = epoch[indices_b]
epoch_c = epoch[indices_c]
model_times_b = times[indices_b]
model_times_c = times[indices_c]
# Compute chi^2 for transit times of e and f
logchi2_b = -0.5 * np.sum( (model_times_b - b_times)**2 / b_uncertainties**2 )
logchi2_c = -0.5 * np.sum( (model_times_c - c_times)**2 / c_uncertainties**2 )
# print(model_times_b.shape, times_b.shape)
# Add log prior for planet mass from Gaia + forecaster
# lnprior = (-0.5 * (mass_b - mass_b_init)**2 / mass_b_init_err**2
# -0.5 * (mass_c - mass_c_init)**2 / mass_c_init_err**2)
return logchi2_b + logchi2_c #+ lnprior
ndim, nwalkers = 8, 16
init_masses = [mass_b_init, mass_c_init, period_b_init, period_c_init, 0, 0, 0, phase]
p0 = [init_masses + 1e-8 * np.random.randn(ndim) for i in range(nwalkers)]
sampler = emcee.EnsembleSampler(nwalkers, ndim, lnprob, threads=8)
p1 = sampler.run_mcmc(p0, 10000)[0];
sampler.reset()
p2 = sampler.run_mcmc(p1, 10000);
chains = np.copy(sampler.flatchain)
chains[:, :2] /= float(M_earth/M_sun)
corner(chains,
truths=[mass_b_init/float(M_earth/M_sun), mass_c_init/float(M_earth/M_sun),
period_b_init, period_c_init, None, None, None, None, None, None],
labels=['$M_b$', '$M_c$', "$P_b$", "$P_c$", '$\lambda_b$', "$\lambda_c$",
"$\omega_b$", "$\omega_c$"]);
plt.savefig('ttv_recovery.png', bbox_inches='tight')
```
| github_jupyter |
```
# import the wine dataset
import pandas as pd
df_wine = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/wine/wine.data', header=None)
# separate the Wine data into training and test sets
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X, y = df_wine.iloc[:, 1:].values, df_wine.iloc[:, 0].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
sc = StandardScaler()
X_train_std = sc.fit_transform(X_train)
X_test_std = sc.transform(X_test)
# obtaining eigenpairs of the Wine dataset
import numpy as np
cov_mat = np.cov(X_train_std.T)
eigen_vals, eigen_vecs = np.linalg.eig(cov_mat)
print('\nEigenvalues \n%s' % eigen_vals)
# plotting the variance explained ratios of the eigenvalues
tot = sum(eigen_vals)
var_exp = [(i / tot) for i in sorted(eigen_vals, reverse=True)]
cum_var_exp = np.cumsum(var_exp)
import matplotlib.pyplot as plt
plt.bar(range(1,14), var_exp, alpha=0.5, align='center', label='cumulative explained variance')
plt.step(range(1,14), cum_var_exp, where='mid', label='individual explained variance')
plt.ylabel('Explained variance ratio')
plt.xlabel('Principal components')
plt.legend(loc='best')
plt.show()
eigen_pairs = [(np.abs(eigen_vals[i]), eigen_vecs[:,i]) for i in range(len(eigen_vals))]
eigen_pairs.sort(key=lambda k: k[0], reverse=True)
w = np.hstack((eigen_pairs[0][1][:, np.newaxis], eigen_pairs[1][1][:, np.newaxis]))
print('Matrix W:\n',w)
# trnsforming the intire training dataset onto the to principal components
X_train_pca = X_train_std.dot(w)
colors = ['r', 'b', 'g']
markers = ['s', 'x', 'o']
for l, c, m in zip(np.unique(y_train), colors, markers):
plt.scatter(X_train_pca[y_train==l, 0], X_train_pca[y_train==l, 1], c=c, label=l, marker=m)
plt.xlabel('PC 1')
plt.ylabel('PC 2')
plt.legend(loc='lower left')
plt.show()
# Principal Component Analysis in scikit-learn
from matplotlib.colors import ListedColormap
def plot_decision_regions(X, y, classifier, resolution=0.02):
# marker generator and color map
markers = ('s', 'x', 'o', '^', 'v')
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
cmap = ListedColormap(colors[:len(np.unique(y))])
# plot the desicion surface
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution), np.arange(x2_min, x2_max, resolution))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.4, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
# plot class samples
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0], y=X[y == cl, 1], alpha=0.8, c=cmap(idx), marker=markers[idx], label=cl)
from sklearn.linear_model import LogisticRegression
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
lr = LogisticRegression()
X_train_pca = pca.fit_transform(X_train_std)
X_test_pca = pca.transform(X_test_std)
lr.fit(X_train_pca, y_train)
plot_decision_regions(X_train_pca, y_train, classifier=lr)
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.legend(loc='lower left')
plt.show()
# separating test dataset
plot_decision_regions(X_test_pca, y_test, classifier=lr)
plt.xlabel('PC1')
plt.ylabel('PC2')
plt.legend(loc='lower left')
plt.show()
# show the explained variance ratio
pca = PCA(n_components=None)
X_train_pca = pca.fit_transform(X_train_std)
pca.explained_variance_ratio_
```
| github_jupyter |
# XGBoost Built-in Algorithm - Bike Rental Regression Example
```
import numpy as np
import pandas as pd
import boto3
import re
import sagemaker
from sagemaker import get_execution_role
# SageMaker SDK Documentation: http://sagemaker.readthedocs.io/en/latest/estimators.html
```
## Upload Data to S3
```
# Specify your bucket name
bucket_name = 'chandra-ml-sagemaker'
training_folder = r'bikerental/training/'
validation_folder = r'bikerental/validation/'
test_folder = r'bikerental/test/'
s3_model_output_location = r's3://{0}/bikerental/model'.format(bucket_name)
s3_training_file_location = r's3://{0}/{1}'.format(bucket_name,training_folder)
s3_validation_file_location = r's3://{0}/{1}'.format(bucket_name,validation_folder)
s3_test_file_location = r's3://{0}/{1}'.format(bucket_name,test_folder)
print(s3_model_output_location)
print(s3_training_file_location)
print(s3_validation_file_location)
print(s3_test_file_location)
# Write and Reading from S3 is just as easy
# files are referred as objects in S3.
# file name is referred as key name in S3
# File stored in S3 is automatically replicated across 3 different availability zones
# in the region where the bucket was created.
# http://boto3.readthedocs.io/en/latest/guide/s3.html
def write_to_s3(filename, bucket, key):
with open(filename,'rb') as f: # Read in binary mode
return boto3.Session().resource('s3').Bucket(bucket).Object(key).upload_fileobj(f)
write_to_s3('bike_train.csv',
bucket_name,
training_folder + 'bike_train.csv')
write_to_s3('bike_validation.csv',
bucket_name,
validation_folder + 'bike_validation.csv')
write_to_s3('bike_test.csv',
bucket_name,
test_folder + 'bike_test.csv')
```
## Training Algorithm Docker Image
### SageMaker maintains a separate image for algorithm and region
https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html
```
# Use Spot Instance - Save up to 90% of training cost by using spot instances when compared to on-demand instances
# Reference: https://github.com/aws-samples/amazon-sagemaker-managed-spot-training/blob/main/xgboost_built_in_managed_spot_training_checkpointing/xgboost_built_in_managed_spot_training_checkpointing.ipynb
# if you are still on two-month free-tier you can use the on-demand instance by setting:
# use_spot_instances = False
# We will use spot for training
use_spot_instances = True
max_run = 3600 # in seconds
max_wait = 7200 if use_spot_instances else None # in seconds
job_name = 'xgboost-bikerental-v1'
checkpoint_s3_uri = None
if use_spot_instances:
checkpoint_s3_uri = f's3://{bucket_name}/bikerental/checkpoints/{job_name}'
print (f'Checkpoint uri: {checkpoint_s3_uri}')
# Establish a session with AWS
sess = sagemaker.Session()
role = get_execution_role()
# This role contains the permissions needed to train, deploy models
# SageMaker Service is trusted to assume this role
print(role)
# https://sagemaker.readthedocs.io/en/stable/api/utility/image_uris.html#sagemaker.image_uris.retrieve
# SDK 2 uses image_uris.retrieve the container image location
# Use XGBoost 1.2 version
container = sagemaker.image_uris.retrieve("xgboost",sess.boto_region_name,version="1.2-2")
print (f'Using XGBoost Container {container}')
```
## Build Model
```
# Configure the training job
# Specify type and number of instances to use
# S3 location where final artifacts needs to be stored
# Reference: http://sagemaker.readthedocs.io/en/latest/estimators.html
# for managed spot training, specify the use_spot_instances flag, max_run, max_wait and checkpoint_s3_uri
# SDK 2.x version does not require train prefix for instance count and type
estimator = sagemaker.estimator.Estimator(
container,
role,
instance_count=1,
instance_type='ml.m5.xlarge',
output_path=s3_model_output_location,
sagemaker_session=sess,
base_job_name = job_name,
use_spot_instances=use_spot_instances,
max_run=max_run,
max_wait=max_wait,
checkpoint_s3_uri=checkpoint_s3_uri)
# Specify hyper parameters that appropriate for the training algorithm
# XGBoost Training Parameter Reference
# https://github.com/dmlc/xgboost/blob/master/doc/parameter.rst#learning-task-parameters
estimator.set_hyperparameters(max_depth=5,
objective="reg:squarederror",
eta=0.1,
num_round=150)
estimator.hyperparameters()
```
### Specify Training Data Location and Optionally, Validation Data Location
```
# content type can be libsvm or csv for XGBoost
training_input_config = sagemaker.session.TrainingInput(
s3_data=s3_training_file_location,
content_type='csv',
s3_data_type='S3Prefix')
validation_input_config = sagemaker.session.TrainingInput(
s3_data=s3_validation_file_location,
content_type='csv',
s3_data_type='S3Prefix'
)
data_channels = {'train': training_input_config, 'validation': validation_input_config}
print(training_input_config.config)
print(validation_input_config.config)
```
### Train the model
```
# XGBoost supports "train", "validation" channels
# Reference: Supported channels by algorithm
# https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html
estimator.fit(data_channels)
```
## Deploy Model
```
# Ref: http://sagemaker.readthedocs.io/en/latest/estimators.html
predictor = estimator.deploy(initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name = job_name)
```
## Run Predictions
```
# SDK 2.0 serializers
from sagemaker.serializers import CSVSerializer
predictor.serializer = CSVSerializer()
predictor.predict([[3,0,1,2,28.7,33.335,79,12.998,2011,7,7,3]])
```
## Summary
1. Ensure Training, Test and Validation data are in S3 Bucket
2. Select Algorithm Container Registry Path - Path varies by region
3. Configure Estimator for training - Specify Algorithm container, instance count, instance type, model output location
4. Specify algorithm specific hyper parameters
5. Train model
6. Deploy model - Specify instance count, instance type and endpoint name
7. Run Predictions
| github_jupyter |
# Approximating the Hessian for large neural networks.
This notebook describes how to use the spectral-density package with Tensorflow2. The main entry point of this package is the `lanczos_algorithm.approximate_hessian` function, compatible with Keras models. This function takes the following arguments:
- `model`: The Keras model for which we want to compute the Hessian.
- `dataset`: Dataset on which the model is trained. Can be a Tensorflow dataset, or more generally any iterator yielding tuples of data (X, y). If a Tensorflow dataset is used, it should be batched beforehand.
- `order`: Rank of the approximation of the Hessian. The higher the better the approximation. See paper for more details.
- `reduce_op`: Whether the loss function averages or sums the per sample loss. The default value is `MEAN` and should be compatible with most Keras losses, provided you didn't specify another reduction when instantiating it.
- `random_seed`: Seed to use to sample the first vector in the Lanczos algorithm.
## Example 1: Full rank estimation for linear model.
We start with a simplistic usecase: we wish to train the following model:
$$ \mbox{arg}\max_\beta \sum_i (y_i - \beta^Tx_i)^2$$
As this optimization problem is quadratic, the Hessian of the loss is independent of $\beta$ and is equal to $2X^TX$. Let's verify this using `lanczos_algorithm.approximate_hessian`, and setting the order of the approximation to the number of features, thus recovering the exact Hessian.
We first generate some random inputs and outputs:
```
import tensorflow.compat.v2 as tf
import tensorflow_datasets as tfds
from matplotlib import pyplot as plt
import seaborn as sns
tf.enable_v2_behavior()
import lanczos_algorithm
num_samples = 50
num_features = 16
X = tf.random.normal([num_samples, num_features])
y = tf.random.normal([num_samples])
```
We then define a linear model using the Keras API:
```
linear_model = tf.keras.Sequential(
[tf.keras.Input(shape=[num_features]),
tf.keras.layers.Dense(1, use_bias=False)])
```
Finally, we define a loss function that takes as input the model and a batch of examples, and return a scalar loss. Here, we simply compute the mean squared error between the predictions of the model and the desired output.
```
def loss_fn(model, inputs):
x, y = inputs
preds = linear_model(x)
return tf.keras.losses.mse(y, preds)
```
Fnally, we call `approximate_hessian`, setting order to the number of parameters to compute the exact Hessian. This function returns two tensors $(V, T)$ of shapes (num_parameters, order) and (order, order), such that :
$$ H \approx V T V^T $$
with an equality if order = num_parameters.
```
V, T = lanczos_algorithm.approximate_hessian(
linear_model,
loss_fn,
[(X,y)],
order=num_features)
```
We can check that the reconstructed Hessian is indeed equal to $2X^TX$:
```
plt.figure(figsize=(14, 5))
plt.subplot(1,2,1)
H = tf.matmul(V, tf.matmul(T, V, transpose_b=True))
plt.title("Hessian as estimated by Lanczos")
sns.heatmap(H)
plt.subplot(1,2,2)
plt.title("$2X^TX$")
sns.heatmap(2 * tf.matmul(X, X, transpose_a=True))
plt.show()
```
## Example 2: Convnet on Cifar10
We first define a VGG16-like model (15.2M parameters) that we train a bit on Cifar10:
```
def preprocess_images(tfrecord):
image, label = tfrecord['image'], tfrecord['label']
image = tf.cast(image, tf.float32) / 255.0
return image, label
cifar_dataset_train = tfds.load("cifar10", split="train").map(preprocess_images).cache()
cifar_dataset_test = tfds.load("cifar10", split="test").map(preprocess_images).cache()
model = tf.keras.Sequential([
tf.keras.Input([32, 32, 3]),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, activation='relu'),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, activation='relu'),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(filters=128, kernel_size=3, activation='relu'),
tf.keras.layers.Conv2D(filters=128, kernel_size=3, activation='relu'),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(1024, activation='relu'),
tf.keras.layers.Dense(10)])
model.compile(optimizer=tf.keras.optimizers.Adam(lr=0.001),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
print(model.summary())
_ = model.fit(cifar_dataset_train.batch(32),
validation_data=cifar_dataset_test.batch(128),
epochs=5)
```
Our loss function is a bit different from the previous one, as we now use cross-entropy to train our model. Don't forget to set `training=False` to deactivate dropouts and similar mechanisms.
Computing an estimation of the Hessian will take a bit of time. A good rule of thumb is that the algorithm will take $T = order \times 2 T_{epoch}$ units of time, where $T_{epoch}$ stands for the time needed to perform one training epoch.
```
SCCE = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
def loss_fn(model, inputs):
x, y = inputs
preds = model(x, training=False)
return SCCE(y, preds)
V, T = lanczos_algorithm.approximate_hessian(
model,
loss_fn,
mnist_dataset.batch(128),
order=90,
random_seed=1)
```
Finally, you can use the visualization functions provided in `jax.density` to plot the spectum (no actual JAX code is involved in this operation).
```
import ..jax.density as density_lib
def plot(grids, density, label=None):
plt.semilogy(grids, density, label=label)
plt.ylim(1e-10, 1e2)
plt.ylabel("Density")
plt.xlabel("Eigenvalue")
plt.legend()
density, grids = density_lib.tridiag_to_density(
[T.numpy()], grid_len=10000, sigma_squared=1e-3)
plot(grids, density)
```
| github_jupyter |
```
import pymysql
# MySQL Connection 연결
conn = pymysql.connect(host='localhost', user='root', password='123456789',
db='lafamila', charset='utf8')
# Connection 으로부터 일반 Cursor 생성
curs = conn.cursor()
# Connection 으로부터 Dictoionary Cursor 생성
#curs = conn.cursor(pymysql.cursors.DictCursor)
# SQL문 실행
sql = "select * from test"
curs.execute(sql)
#insert 나 update와 같은 query는 commit 과정이 필요하다
#sql = "insert into test values('asdf', 'kk', '@mai.com', 4321)"
#curs.execute(sql)
#conn.commit()
# sql에 변수 넣기
# sql query를 구성할 때 만약 String Formatting 을 사용하게되면 injection 등의 문제를 일으킬 수 있다.
# 따라서 데이터가 들어갈 자리를 %s로 대체해두고 cursor를 실행할 때 tuple 형식으로 첨부한다.
# (이때 %s는 일반적인 formatting 이 아님)
#sql = "select * from test where PW=%s"
#curs.execute(sql, ('adf'))
# 데이타 Fetch
rows = curs.fetchall()
# 하나 가져올때는 fetchone()
# 여러개 가져올때는 fetchmany(n)
# 가져온 data는 Cursor의 종류에 따라 tuple 또는 dict 가 된다.
for row in rows:
print(row, type(row)) # 전체 rows 출력
# Connection 닫기
conn.close()
import pymysql
conn = pymysql.connect(host='localhost', user='root', password='123456789',
db='lafamila', charset='utf8')
curs = conn.cursor(pymysql.cursors.DictCursor)
sql = "select * from test"
curs.execute(sql)
rows = curs.fetchall()
print(type(rows))
for row in rows:
print(row["ID"])
#print(row, type(row))
conn.close()
import pymysql
conn = pymysql.connect(host='localhost', user='root', password='123456789',
db='shoppingmall', charset='utf8')
curs = conn.cursor(pymysql.cursors.DictCursor)
sql = "insert into category_update values(NOW(), %s, %s, %s)"
curs.execute(sql, ("name", 0, 0))
conn.commit()
#rows = curs.fetchall()
#print(type(rows))
#for row in rows:
# print(row["ID"])
#print(row, type(row))
conn.close()
```
# Shopping Mall
```
import requests
from bs4 import BeautifulSoup
import urllib
import pymysql
IMAGE_PATH = "C:\\Users\\lafamila\\Downloads\\"
conn = pymysql.connect(host='localhost', user='root', password='123456789',
db='shoppingmall', charset='utf8')
curs = conn.cursor()
sql = "select p_number from product_update"
# sql query를 구성할 때 만약 String Formatting 을 사용하게되면 injection 등의 문제를 일으킬 수 있다.
# 따라서 데이터가 들어갈 자리를 %s로 대체해두고 cursor를 실행할 때 tuple 형식으로 첨부한다.
curs.execute(sql)
rows = curs.fetchall()
#top에서 에러가 나서 top만 실행해보았음
url_d = {
'top':'http://www.secretlabel.co.kr/shop/shopbrand.html?xcode=047',
}
for url in url_d.values():
p_nums = set()
category_update = 0
response = requests.get(url)
soup = BeautifulSoup(response.content,'html.parser')
#print(soup.title)
p_count = 1
page_url = url + soup.select_one('.now > a').get('href')[0:-1]
#print(page_url)
product_count = 0
while True:
page_url_v = page_url + str(p_count)
p_count += 1
res = requests.get(page_url_v)
soup = BeautifulSoup(res.content,'html.parser')
soup_p = soup.select_one('#SMS_prd_container')
product_lists = soup_p.select('.item_list')
product_count += len(product_lists)
if len(soup_p.select('.item_list')) == 0:
break
for y in product_lists:
li = y.find_all("li")
#상품이미지 다운로드 및 경로
image_src = y.select_one('.MS_prod_img_s').get('src').split('?')[0]
start = image_src.rindex("/")+1
#urllib.request.urlretrieve( image_src , IMAGE_PATH+image_src[start:] )
#이름
name = y.select_one('.pname').text.strip('\n').split()[0]
#금액
price = y.select_one('.price').text.strip()
try:
price = int(price[:-1].replace(",",""))
except:
print(price)
break
#url
product_url = url + y.select_one('a').get('href')
#상품후기수
#상품번호를 받아와서 하면 된다.
p_num = y.select_one('a').get('href')[31:37]
#cre = 'http://widgets3.cre.ma/secretlabel.co.kr/products/reviews?product_code=' + p_num
#cre_r = requests.get(cre)
#cre_soup = BeautifulSoup(cre_r.content,'html.parser')
#총리뷰를 불러와라
#review_count = cre_soup.select_one('.reviews-count').text
#상품키
p_num = url.split(".")[1] + p_num
p_nums.add((p_num,))
if (p_num,) in rows:
print("update")
else:
"""
###########################################################
## DB ##
###########################################################
# p_num
# name
# price
# product_url
# IMAGE_PATH+image_src[start:]
# 'top'
# review_count
# '시크릿라벨'
curs = conn.cursor(pymysql.cursors.DictCursor)
#INSERT
sql = "insert into product_update values(NOW(), %s, %s, %s, %s, %s, %s, %s, %s)"
curs.execute(sql, (p_num, name, price, product_url, IMAGE_PATH+image_src[start:], 'top', review_count, "시크릿라벨"))
conn.commit()
"""
category_update += 1
#print('카테고리상품수 :' + str(product_count) + '개')
#기존에 있던 상품key 들의 집합에서 새로 크롤링한 상품key 들의 차집합을 구함
#--> 기존에는 있었으나 새로 크롤링한 데이터에는 없는 개수 (품절개수)
print(len(set(list(rows)).difference(p_nums)))
###########################################################
## DB ##
###########################################################
# 'top'
# category_update
# len(set(list(rows)).difference(p_nums))
curs = conn.cursor(pymysql.cursors.DictCursor)
#INSERT
sql = "insert into category_update values(NOW(), %s, %s, %s)"
curs.execute(sql, ('top', category_update, len(set(list(rows)).difference(p_nums))))
conn.commit()
conn.close()
print(set(list(rows)))
print(p_nums)
```
| github_jupyter |
```
import random
def get_random_number():
return random.randrange(100, 1000)
def is_digit(user_input_number):
user_input_number = str(user_input_number)
result = user_input_number.isdigit()
return result
def is_between_100_and_999(user_input_number):
if int(user_input_number)>=100 and int(user_input_number)<999:
result = True
else:
result = False
return result
def is_duplicated_number(three_digit):
three_digit = str(three_digit)
s = set([])
for digit in three_digit:
s.add(digit)
if len(s) < 3:
result = True
else:
result = False
return result
def is_validated_number(user_input_number):
if (is_digit(user_input_number) == True and is_between_100_and_999(user_input_number) == True and
is_duplicated_number(user_input_number) == False):
result = True
else:
result = False
return result
def get_not_duplicated_three_digit_number():
random_number = get_random_number()
while(is_duplicated_number(random_number) == True):
random_number = get_random_number()
result = random_number
return result
def get_strikes_or_ball(user_input_number, random_number):
user_input_number = str(user_input_number)
random_number = str(random_number)
strikes = 0
ball = 0
for i in range(3):
for j in range(3):
if user_input_number[i] == random_number[j] and i == j:
strikes += 1
elif user_input_number[i] == random_number[j]:
ball += 1
else:
continue
result = [strikes, ball]
return result
def is_yes(one_more_input):
one_more_input = one_more_input.upper()
if one_more_input == "Y" or one_more_input == "YES":
result = True
else:
result = False
return result
def is_no(one_more_input):
one_more_input = str(one_more_input)
one_more_input = one_more_input.upper()
if one_more_input == "N" or one_more_input == "NO":
result = True
else:
result = False
return result
def main():
print("Play Baseball")
user_input = 999
while(user_input is not 0 and is_no(user_input) == False): #둘중 하나라도 만족 못할때 loop 나옴
random_number = str(get_not_duplicated_three_digit_number())
print("Random Number is : ", random_number)
user_input = input("Input guess number : ")
while(is_validated_number(user_input) == False and user_input != "0"): #0넣으면 loop 나옴
print("Wrong Input, Input again")
user_input = input("Input guess number : ")
if user_input == "0":
user_input = 0
else:
#user_input 에 validated_number 들어간 상태
# user_input = str(user_inpput)?
strikes = 0
balls = 0
while(strikes < 3 and user_input != "0"):
strikes = get_strikes_or_ball(user_input, random_number)[0]
balls = get_strikes_or_ball(user_input, random_number)[1]
print("Strikes : ", strikes, "Balls : ", balls)
user_input = input("Input guess number : ")
if user_input == "0":
user_input = 0
else:
print("Strikes : ", strikes, "Balls : ", balls)
#3 Strike 상태, user_input 에는 정답이 있음
user_input = input("You win, one more(Y/N)?") #n입력하면 꺼짐
while(is_yes(user_input) == False and is_no(user_input) == False):
print("Wrong Input, Input again")
user_input = input("You win, one more(Y/N)?")
print("Thank you for using this program")
print("End of the Game")
if __name__ == "__main__":
main()
strikes = get_strikes_or_ball(user_input, random_number)[0]
balls = get_strikes_or_ball(user_input, random_number)[1]
print("Strikes : ", strikes, "Balls : ", balls)
if strikes <3:
user_input = input("Input guess number : ")
while(strikes < 3 and user_input!="0"):
strikes = get_strikes_or_ball(user_input, random_number)[0]
balls = get_strikes_or_ball(user_input, random_number)[1]
print("Strikes : ", strikes, "Balls : ", balls)
user_input = input("Input guess number : ")
if user_input == "0":
user_input = 0
else:
#한번에 맞췄을때
else:
```
| github_jupyter |
## Introduction
This tutorial is a brief introduction to music generation using **Generative Adversarial Networks** (**GAN**s).
The goal of this tutorial is to train a machine learning model using a dataset of Bach compositions so that the model learns to add accompaniments to a single track input melody. In other words, if the user provides a single piano track of a song such as "twinkle twinkle little star", the GAN model would add three other piano tracks to make the music sound more Bach-inspired.
The proposed algorithm consists of two competing networks: a generator and a critic (discriminator). A generator is a deep neural network that learns to create new synthetic data that resembles the distribution of the dataset on which it was trained. A critic is another deep neural network that is trained to differentiate between real and synthetic data. The generator and the critic are trained in alternating cycles such that the generator learns to produce more and more realistic data (Bach-like music in this use case) while the critic iteratively gets better at learning to differentiate real data (Bach music) from the synthetic ones.
As a result, the quality of music produced by the generator gets more and more realistic with time.

## Dependencies
First, let's import all of the python packages we will use throughout the tutorial.
```
# Copyright 2019 Amazon.com, Inc. or its affiliates. All Rights Reserved.
# Permission is hereby granted, free of charge, to any person obtaining a copy of
# this software and associated documentation files (the "Software"), to deal in
# the Software without restriction, including without limitation the rights to
# use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of
# the Software, and to permit persons to whom the Software is furnished to do so.
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS
# FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR
# COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER
# IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
# CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# Create the environment
import subprocess
print("Please wait, while the required packages are being installed...")
subprocess.call(['./requirements.sh'], shell=True)
print("All the required packages are installed successfully...")
# IMPORTS
import os
import numpy as np
from PIL import Image
import logging
import pypianoroll
import scipy.stats
import pickle
import music21
from IPython import display
import matplotlib.pyplot as plt
# Configure Tensorflow
import tensorflow as tf
print(tf.__version__)
tf.logging.set_verbosity(tf.logging.ERROR)
tf.enable_eager_execution()
# Use this command to make a subset of GPUS visible to the jupyter notebook.
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
# Utils library for plotting, loading and saving midi among other functions
from utils import display_utils, metrics_utils, path_utils, inference_utils, midi_utils
LOGGER = logging.getLogger("gan.train")
%matplotlib inline
```
## Configuration
Here we configure paths to retrieve our dataset and save our experiments.
```
root_dir = './Experiments'
# Directory to save checkpoints
model_dir = os.path.join(root_dir,'2Bar') # JSP: 229, Bach: 19199
# Directory to save pianorolls during training
train_dir = os.path.join(model_dir, 'train')
# Directory to save checkpoint generated during training
check_dir = os.path.join(model_dir, 'preload')
# Directory to save midi during training
sample_dir = os.path.join(model_dir, 'sample')
# Directory to save samples generated during inference
eval_dir = os.path.join(model_dir, 'eval')
os.makedirs(train_dir, exist_ok=True)
os.makedirs(eval_dir, exist_ok=True)
os.makedirs(sample_dir, exist_ok=True)
```
## Data Preparation
### Dataset summary
In this tutorial, we use the [`JSB-Chorales-dataset`](http://www-etud.iro.umontreal.ca/~boulanni/icml2012), comprising 229 chorale snippets. A chorale is a hymn that is usually sung with a single voice playing a simple melody and three lower voices providing harmony. In this dataset, these voices are represented by four piano tracks.
Let's listen to a song from this dataset.
```
display_utils.playmidi('./original_midi/MIDI-0.mid')
```
### Data format - piano roll
For the purpose of this tutorial, we represent music from the JSB-Chorales dataset in the piano roll format.
**Piano roll** is a discrete representation of music which is intelligible by many machine learning algorithms. Piano rolls can be viewed as a two-dimensional grid with "Time" on the horizontal axis and "Pitch" on the vertical axis. A one or zero in any particular cell in this grid indicates if a note was played or not at that time for that pitch.
Let us look at a few piano rolls in our dataset. In this example, a single piano roll track has 32 discrete time steps and 128 pitches. We see four piano rolls here, each one representing a separate piano track in the song.
<img src="images/pianoroll2.png" alt="Dataset summary" width="800">
You might notice this representation looks similar to an image. While the sequence of notes is often the natural way that people view music, many modern machine learning models instead treat music as images and leverage existing techniques within the computer vision domain. You will see such techniques used in our architecture later in this tutorial.
**Why 32 time steps?**
For the purpose of this tutorial, we sample two non-empty bars (https://en.wikipedia.org/wiki/Bar_(music)) from each song in the JSB-Chorales dataset. A **bar** (or **measure**) is a unit of composition and contains four beats for songs in our particular dataset (our songs are all in 4/4 time) :
We’ve found that using a resolution of four time steps per beat captures enough of the musical detail in this dataset.
This yields...
$$ \frac{4\;timesteps}{1\;beat} * \frac{4\;beats}{1\;bar} * \frac{2\;bars}{1} = 32\;timesteps $$
Let us now load our dataset as a numpy array. Our dataset comprises 229 samples of 4 tracks (all tracks are piano). Each sample is a 32 time-step snippet of a song, so our dataset has a shape of...
(num_samples, time_steps, pitch_range, tracks) = (229, 32, 128, 4).
```
training_data = np.load('./dataset/train.npy')
print(training_data.shape)
```
Let's see a sample of the data we'll feed into our model. The four graphs represent the four tracks.
```
display_utils.show_pianoroll(training_data)
```
### Load data
We now create a Tensorflow dataset object from our numpy array to feed into our model. The dataset object helps us feed batches of data into our model. A batch is a subset of the data that is passed through the deep learning network before the weights are updated. Batching data is necessary in most training scenarios as our training environment might not be able to load the entire dataset into memory at once.
```
#Number of input data samples in a batch
BATCH_SIZE = 64
#Shuffle buffer size for shuffling data
SHUFFLE_BUFFER_SIZE = 1000
#Preloads PREFETCH_SIZE batches so that there is no idle time between batches
PREFETCH_SIZE = 4
def prepare_dataset(filename):
"""Load the samples used for training."""
data = np.load(filename)
data = np.asarray(data, dtype=np.float32) # {-1, 1}
print('data shape = {}'.format(data.shape))
dataset = tf.data.Dataset.from_tensor_slices(data)
dataset = dataset.shuffle(SHUFFLE_BUFFER_SIZE).repeat()
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
dataset = dataset.prefetch(PREFETCH_SIZE)
return dataset
dataset = prepare_dataset('./dataset/train.npy')
```
## Model architecture
In this section, we will walk through the architecture of the proposed GAN.
The model consists of two networks, a generator and a critic. These two networks work in a tight loop as following:
* Generator:
1. The generator takes in a batch of single-track piano rolls (melody) as the input and generates a batch of multi-track piano rolls as the output by adding accompaniments to each of the input music tracks.
2. The critic then takes these generated music tracks and predicts how far it deviates from the real data present in your training dataset.
3. This feedback from the critic is used by the generator to update its weights.
* Critic: As the generator gets better at creating better music accompaniments using the feedback from the critic, the critic needs to be retrained as well.
1. Train the critic with the music tracks just generated by the generator as fake inputs and an equivalent number of songs from the original dataset as the real input.
* Alternate between training these two networks until the model converges and produces realistic music, beginning with the critic on the first iteration.
We use a special type of GAN called the **Wasserstein GAN with Gradient Penalty** (or **WGAN-GP**) to generate music. While the underlying architecture of a WGAN-GP is very similar to vanilla variants of GAN, WGAN-GPs help overcome some of the commonly seen defects in GANs such as the vanishing gradient problem and mode collapse (see appendix for more details).
Note our "critic" network is more generally called a "discriminator" network in the more general context of vanilla GANs.
### Generator
The generator is adapted from the U-Net architecture (a popular CNN that is used extensively in the computer vision domain), consisting of an “encoder” that maps the single track music data (represented as piano roll images) to a relatively lower dimensional “latent space“ and a ”decoder“ that maps the latent space back to multi-track music data.
Here are the inputs provided to the generator:
**Single-track piano roll input**: A single melody track of size (32, 128, 1) => (TimeStep, NumPitches, NumTracks) is provided as the input to the generator.
**Latent noise vector**: A latent noise vector z of dimension (2, 8, 512) is also passed in as input and this is responsible for ensuring that there is a distinctive flavor to each output generated by the generator, even when the same input is provided.
Notice from the figure below that the encoding layers of the generator on the left side and decoder layer on on the right side are connected to create a U-shape, thereby giving the name U-Net to this architecture.
<img src="images/dgen.png" alt="Generator architecture" width="800">
In this implementation, we build the generator following a simple four-level Unet architecture by combining `_conv2d`s and `_deconv2d`, where `_conv2d` compose the contracting path and `_deconv2d` forms the expansive path.
```
def _conv2d(layer_input, filters, f_size=4, bn=True):
"""Generator Basic Downsampling Block"""
d = tf.keras.layers.Conv2D(filters, kernel_size=f_size, strides=2,
padding='same')(layer_input)
d = tf.keras.layers.LeakyReLU(alpha=0.2)(d)
if bn:
d = tf.keras.layers.BatchNormalization(momentum=0.8)(d)
return d
def _deconv2d(layer_input, pre_input, filters, f_size=4, dropout_rate=0):
"""Generator Basic Upsampling Block"""
u = tf.keras.layers.UpSampling2D(size=2)(layer_input)
u = tf.keras.layers.Conv2D(filters, kernel_size=f_size, strides=1,
padding='same')(u)
u = tf.keras.layers.BatchNormalization(momentum=0.8)(u)
u = tf.keras.layers.ReLU()(u)
if dropout_rate:
u = tf.keras.layers.Dropout(dropout_rate)(u)
u = tf.keras.layers.Concatenate()([u, pre_input])
return u
def build_generator(condition_input_shape=(32, 128, 1), filters=64,
instruments=4, latent_shape=(2, 8, 512)):
"""Buld Generator"""
c_input = tf.keras.layers.Input(shape=condition_input_shape)
z_input = tf.keras.layers.Input(shape=latent_shape)
d1 = _conv2d(c_input, filters, bn=False)
d2 = _conv2d(d1, filters * 2)
d3 = _conv2d(d2, filters * 4)
d4 = _conv2d(d3, filters * 8)
d4 = tf.keras.layers.Concatenate(axis=-1)([d4, z_input])
u4 = _deconv2d(d4, d3, filters * 4)
u5 = _deconv2d(u4, d2, filters * 2)
u6 = _deconv2d(u5, d1, filters)
u7 = tf.keras.layers.UpSampling2D(size=2)(u6)
output = tf.keras.layers.Conv2D(instruments, kernel_size=4, strides=1,
padding='same', activation='tanh')(u7) # 32, 128, 4
generator = tf.keras.models.Model([c_input, z_input], output, name='Generator')
return generator
```
Let us now dive into each layer of the generator to see the inputs/outputs at each layer.
```
# Models
generator = build_generator()
generator.summary()
```
### Critic (Discriminator)
The goal of the critic is to provide feedback to the generator about how realistic the generated piano rolls are, so that the generator can learn to produce more realistic data. The critic provides this feedback by outputting a scalar that represents how “real” or “fake” a piano roll is.
Since the critic tries to classify data as “real” or “fake”, it is not very different from commonly used binary classifiers. We use a simple architecture for the critic, composed of four convolutional layers and a dense layer at the end.
<img src="images/ddis.png" alt="Discriminator architecture" width="800">
```
def _build_critic_layer(layer_input, filters, f_size=4):
"""
This layer decreases the spatial resolution by 2:
input: [batch_size, in_channels, H, W]
output: [batch_size, out_channels, H/2, W/2]
"""
d = tf.keras.layers.Conv2D(filters, kernel_size=f_size, strides=2,
padding='same')(layer_input)
# Critic does not use batch-norm
d = tf.keras.layers.LeakyReLU(alpha=0.2)(d)
return d
def build_critic(pianoroll_shape=(32, 128, 4), filters=64):
"""WGAN critic."""
condition_input_shape = (32,128,1)
groundtruth_pianoroll = tf.keras.layers.Input(shape=pianoroll_shape)
condition_input = tf.keras.layers.Input(shape=condition_input_shape)
combined_imgs = tf.keras.layers.Concatenate(axis=-1)([groundtruth_pianoroll, condition_input])
d1 = _build_critic_layer(combined_imgs, filters)
d2 = _build_critic_layer(d1, filters * 2)
d3 = _build_critic_layer(d2, filters * 4)
d4 = _build_critic_layer(d3, filters * 8)
x = tf.keras.layers.Flatten()(d4)
logit = tf.keras.layers.Dense(1)(x)
critic = tf.keras.models.Model([groundtruth_pianoroll,condition_input], logit,
name='Critic')
return critic
# Create the Discriminator
critic = build_critic()
critic.summary() # View discriminator architecture.
```
## Training
We train our models by searching for model parameters which optimize an objective function. For our WGAN-GP, we have special loss functions that we minimize as we alternate between training our generator and critic networks:
*Generator Loss:*
* We use the Wasserstein (Generator) loss function which is negative of the Critic Loss function. The generator is trained to bring the generated pianoroll as close to the real pianoroll as possible.
* $\frac{1}{m} \sum_{i=1}^{m} -D_w(G(z^{i}|c^{i})|c^{i})$
*Critic Loss:*
* We begin with the Wasserstein (Critic) loss function designed to maximize the distance between the real piano roll distribution and generated (fake) piano roll distribution.
* $\frac{1}{m} \sum_{i=1}^{m} [D_w(G(z^{i}|c^{i})|c^{i}) - D_w(x^{i}|c^{i})]$
* We add a gradient penalty loss function term designed to control how the gradient of the critic with respect to its input behaves. This makes optimization of the generator easier.
* $\frac{1}{m} \sum_{i=1}^{m}(\lVert \nabla_{\hat{x}^i}D_w(\hat{x}^i|c^{i}) \rVert_2 - 1)^2 $
```
# Define the different loss functions
def generator_loss(critic_fake_output):
""" Wasserstein GAN loss
(Generator) -D(G(z|c))
"""
return -tf.reduce_mean(critic_fake_output)
def wasserstein_loss(critic_real_output, critic_fake_output):
""" Wasserstein GAN loss
(Critic) D(G(z|c)) - D(x|c)
"""
return tf.reduce_mean(critic_fake_output) - tf.reduce_mean(
critic_real_output)
def compute_gradient_penalty(critic, x, fake_x):
c = tf.expand_dims(x[..., 0], -1)
batch_size = x.get_shape().as_list()[0]
eps_x = tf.random.uniform(
[batch_size] + [1] * (len(x.get_shape()) - 1)) # B, 1, 1, 1, 1
inter = eps_x * x + (1.0 - eps_x) * fake_x
with tf.GradientTape() as g:
g.watch(inter)
disc_inter_output = critic((inter,c), training=True)
grads = g.gradient(disc_inter_output, inter)
slopes = tf.sqrt(1e-8 + tf.reduce_sum(
tf.square(grads),
reduction_indices=tf.range(1, grads.get_shape().ndims)))
gradient_penalty = tf.reduce_mean(tf.square(slopes - 1.0))
return gradient_penalty
```
With our loss functions defined, we associate them with Tensorflow optimizers to define how our model will search for a good set of model parameters. We use the *Adam* algorithm, a commonly used general-purpose optimizer. We also set up checkpoints to save our progress as we train.
```
# Setup Adam optimizers for both G and D
generator_optimizer = tf.keras.optimizers.Adam(1e-3, beta_1=0.5, beta_2=0.9)
critic_optimizer = tf.keras.optimizers.Adam(1e-3, beta_1=0.5, beta_2=0.9)
# We define our checkpoint directory and where to save trained checkpoints
ckpt = tf.train.Checkpoint(generator=generator,
generator_optimizer=generator_optimizer,
critic=critic,
critic_optimizer=critic_optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, check_dir, max_to_keep=5)
```
Now we define the `generator_train_step` and `critic_train_step` functions, each of which performs a single forward pass on a batch and returns the corresponding loss.
```
@tf.function
def generator_train_step(x, condition_track_idx=0):
############################################
#(1) Update G network: maximize D(G(z|c))
############################################
# Extract condition track to make real batches pianoroll
c = tf.expand_dims(x[..., condition_track_idx], -1)
# Generate batch of latent vectors
z = tf.random.truncated_normal([BATCH_SIZE, 2, 8, 512])
with tf.GradientTape() as tape:
fake_x = generator((c, z), training=True)
fake_output = critic((fake_x,c), training=False)
# Calculate Generator's loss based on this generated output
gen_loss = generator_loss(fake_output)
# Calculate gradients for Generator
gradients_of_generator = tape.gradient(gen_loss,
generator.trainable_variables)
# Update Generator
generator_optimizer.apply_gradients(
zip(gradients_of_generator, generator.trainable_variables))
return gen_loss
@tf.function
def critic_train_step(x, condition_track_idx=0):
############################################################################
#(2) Update D network: maximize (D(x|c)) + (1 - D(G(z|c))|c) + GradientPenality()
############################################################################
# Extract condition track to make real batches pianoroll
c = tf.expand_dims(x[..., condition_track_idx], -1)
# Generate batch of latent vectors
z = tf.random.truncated_normal([BATCH_SIZE, 2, 8, 512])
# Generated fake pianoroll
fake_x = generator((c, z), training=False)
# Update critic parameters
with tf.GradientTape() as tape:
real_output = critic((x,c), training=True)
fake_output = critic((fake_x,c), training=True)
critic_loss = wasserstein_loss(real_output, fake_output)
# Caculate the gradients from the real and fake batches
grads_of_critic = tape.gradient(critic_loss,
critic.trainable_variables)
with tf.GradientTape() as tape:
gp_loss = compute_gradient_penalty(critic, x, fake_x)
gp_loss *= 10.0
# Calculate the gradients penalty from the real and fake batches
grads_gp = tape.gradient(gp_loss, critic.trainable_variables)
gradients_of_critic = [g + ggp for g, ggp in
zip(grads_of_critic, grads_gp)
if ggp is not None]
# Update Critic
critic_optimizer.apply_gradients(
zip(gradients_of_critic, critic.trainable_variables))
return critic_loss + gp_loss
```
Before we begin training, let's define some training configuration parameters and prepare to monitor important quantities. Here we log the losses and metrics which we can use to determine when to stop training. Consider coming back here to tweak these parameters and explore how your model responds.
```
# We use load_melody_samples() to load 10 input data samples from our dataset into sample_x
# and 10 random noise latent vectors into sample_z
sample_x, sample_z = inference_utils.load_melody_samples(n_sample=10)
# Number of iterations to train for
iterations = 1000
# Update critic n times per generator update
n_dis_updates_per_gen_update = 5
# Determine input track in sample_x that we condition on
condition_track_idx = 0
sample_c = tf.expand_dims(sample_x[..., condition_track_idx], -1)
```
Let us now train our model!
```
# Clear out any old metrics we've collected
metrics_utils.metrics_manager.initialize()
# Keep a running list of various quantities:
c_losses = []
g_losses = []
# Data iterator to iterate over our dataset
it = iter(dataset)
for iteration in range(iterations):
# Train critic
for _ in range(n_dis_updates_per_gen_update):
c_loss = critic_train_step(next(it))
# Train generator
g_loss = generator_train_step(next(it))
# Save Losses for plotting later
c_losses.append(c_loss)
g_losses.append(g_loss)
display.clear_output(wait=True)
fig = plt.figure(figsize=(15, 5))
line1, = plt.plot(range(iteration+1), c_losses, 'r')
line2, = plt.plot(range(iteration+1), g_losses, 'k')
plt.xlabel('Iterations')
plt.ylabel('Losses')
plt.legend((line1, line2), ('C-loss', 'G-loss'))
display.display(fig)
plt.close(fig)
# Output training stats
print('Iteration {}, c_loss={:.2f}, g_loss={:.2f}'.format(iteration, c_loss, g_loss))
# Save checkpoints, music metrics, generated output
if iteration < 100 or iteration % 50 == 0 :
# Check how the generator is doing by saving G's samples on fixed_noise
fake_sample_x = generator((sample_c, sample_z), training=False)
metrics_utils.metrics_manager.append_metrics_for_iteration(fake_sample_x.numpy(), iteration)
if iteration % 50 == 0:
# Save the checkpoint to disk.
ckpt_manager.save(checkpoint_number=iteration)
fake_sample_x = fake_sample_x.numpy()
# plot the pianoroll
display_utils.plot_pianoroll(iteration, sample_x[:4], fake_sample_x[:4], save_dir=train_dir)
# generate the midi
destination_path = path_utils.generated_midi_path_for_iteration(iteration, saveto_dir=sample_dir)
midi_utils.save_pianoroll_as_midi(fake_sample_x[:4], destination_path=destination_path)
```
### We have started training!
When using the Wasserstein loss function, we should train the critic to converge to ensure that the gradients for the generator update are accurate. This is in contrast to a standard GAN, where it is important not to let the critic get too strong, to avoid vanishing gradients.
Therefore, using the Wasserstein loss removes one of the key difficulties of training GANs—how to balance the training of the discriminator and generator. With WGANs, we can simply train the critic several times between generator updates, to ensure it is close to convergence. A typical ratio used is five critic updates to one generator update.
### "Babysitting" the learning process
Given that training these models can be an investment in time and resources, we must to continuously monitor training in order to catch and address anomalies if/when they occur. Here are some things to look out for:
**What should the losses look like?**
The adversarial learning process is highly dynamic and high-frequency oscillations are quite common. However if either loss (critic or generator) skyrockets to huge values, plunges to 0, or get stuck on a single value, there is likely an issue somewhere.
**Is my model learning?**
- Monitor the critic loss and other music quality metrics (if applicable). Are they following the expected trajectories?
- Monitor the generated samples (piano rolls). Are they improving over time? Do you see evidence of mode collapse? Have you tried listening to your samples?
**How do I know when to stop?**
- If the samples meet your expectations
- Critic loss no longer improving
- The expected value of the musical quality metrics converge to the corresponding expected value of the same metric on the training data
### How to measure sample quality during training
Typically, when training any sort of neural networks, it is standard practice to monitor the value of the loss function throughout the duration of the training. The critic loss in WGANs has been found to correlate well with sample quality.
While standard mechanisms exist for evaluating the accuracy of more traditional models like classifiers or regressors, evaluating generative models is an active area of research. Within the domain of music generation, this hard problem is even less well-understood.
To address this, we take high-level measurements of our data and show how well our model produces music that aligns with those measurements. If our model produces music which is close to the mean value of these measurements for our training dataset, our music should match on general “shape”.
We’ll look at three such measurements:
- **Empty bar rate:** The ratio of empty bars to total number of bars.
- **Pitch histogram distance:** A metric that captures the distribution and position of pitches.
- **In Scale Ratio:** Ratio of the number of notes that are in C major key, which is a common key found in music, to the total number of notes.
## Evaluate results
Now that we have finished training, let's find out how we did. We will analyze our model in several ways:
1. Examine how the generator and critic losses changed while training
2. Understand how certain musical metrics changed while training
3. Visualize generated piano roll output for a fixed input at every iteration and create a video
Let us first restore our last saved checkpoint. If you did not complete training but still want to continue with a pre-trained version, set `TRAIN = False`.
```
ckpt = tf.train.Checkpoint(generator=generator)
ckpt_manager = tf.train.CheckpointManager(ckpt, check_dir, max_to_keep=5)
ckpt.restore(ckpt_manager.latest_checkpoint).expect_partial()
print('Latest checkpoint {} restored.'.format(ckpt_manager.latest_checkpoint))
```
### Plot losses
```
display_utils.plot_loss_logs(g_losses, c_losses, figsize=(15, 5), smoothing=0.01)
```
Observe how the critic loss (C_loss in the graph) decays to zero as we train. In WGAN-GPs, the critic loss decreases (almost) monotonically as you train.
### Plot metrics
```
metrics_utils.metrics_manager.set_reference_metrics(training_data)
metrics_utils.metrics_manager.plot_metrics()
```
Each row here corresponds to a different music quality metric and each column denotes an instrument track.
Observe how the expected value of the different metrics (blue scatter) approach the corresponding training set expected values (red) as the number of iterations increase. You might expect to see diminishing returns as the model converges.
### Generated samples during training
The function below helps you probe intermediate samples generated in the training process. Remember that the conditioned input here is sampled from our training data. Let's start by listening to and observing a sample at iteration 0 and then iteration 100. Notice the difference!
```
# Enter an iteration number (can be divided by 50) and listen to the midi at that iteration
iteration = 50
midi_file = os.path.join(sample_dir, 'iteration-{}.mid'.format(iteration))
display_utils.playmidi(midi_file)
# Enter an iteration number (can be divided by 50) and look at the generated pianorolls at that iteration
iteration = 50
pianoroll_png = os.path.join(train_dir, 'sample_iteration_%05d.png' % iteration)
display.Image(filename=pianoroll_png)
```
Let's see how the generated piano rolls change with the number of iterations.
```
from IPython.display import Video
display_utils.make_training_video(train_dir)
video_path = "movie.mp4"
Video(video_path)
```
## Inference
### Generating accompaniment for custom input
Congratulations! You have trained your very own WGAN-GP to generate music. Let us see how our generator performs on a custom input.
The function below generates a new song based on "Twinkle Twinkle Little Star".
```
latest_midi = inference_utils.generate_midi(generator, eval_dir, input_midi_file='./input_twinkle_twinkle.mid')
display_utils.playmidi(latest_midi)
```
We can also take a look at the generated piano rolls for a certain sample, to see how diverse they are!
```
inference_utils.show_generated_pianorolls(generator, eval_dir, input_midi_file='./input_twinkle_twinkle.mid')
```
# What's next?
### Using your own data (Optional)
To create your own dataset you can extract the piano roll from MIDI data. An example of creating a piano roll from a MIDI file is given below
```
import numpy as np
from pypianoroll import Multitrack
midi_data = Multitrack('./input_twinkle_twinkle.mid')
tracks = [track.pianoroll for track in midi_data.tracks]
sample = np.stack(tracks, axis=-1)
print(sample.shape)
```
# Appendix
### Open source implementations
For more open-source implementations of generative models for music, check out:
- [MuseGAN](https://github.com/salu133445/musegan): Official TensorFlow Implementation that uses GANs to generate multi track polyphonic music
- [GANSynth](https://github.com/tensorflow/magenta/tree/master/magenta/models/gansynth): GANSynth uses a Progressive GAN architecture to incrementally upsample with convolution from a single vector to the full audio spectrogram
- [Music Transformer](https://github.com/tensorflow/magenta/tree/master/magenta/models/score2perf): Uses transformers to generate music!
GANs have also achieved state of the generative modeling in several other domains including cross domain image tranfer, celebrity face generation, super resolution text to image and image inpainting.
- [Keras-GAN](https://github.com/eriklindernoren/Keras-GAN): Library of reference implementations in Keras for image generation(good for educational purposes).
There's an ocean of literatures out there that use GANs for modeling distributions across fields! If you are interested, [Gan Zoo](https://github.com/hindupuravinash/the-gan-zoo) is a good place to start.
### References
<a id='references'></a>
1. [Dong, H.W., Hsiao, W.Y., Yang, L.C. and Yang, Y.H., 2018, April. MuseGAN: Multi-track sequential generative adversarial networks for symbolic music generation and accompaniment. In Thirty-Second AAAI Conference on Artificial Intelligence.](https://arxiv.org/abs/1709.06298)
2. [Ishaan, G., Faruk, A., Martin, A., Vincent, D. and Aaron, C., 2017. Improved training of wasserstein gans. In Advances in Neural Information Processing Systems.](https://arxiv.org/abs/1704.00028)
3. [Arjovsky, M., Chintala, S. and Bottou, L., 2017. Wasserstein gan. arXiv preprint arXiv:1701.07875.](https://arxiv.org/abs/1701.07875)
4. [Foster, D., 2019. Generative Deep Learning: Teaching Machines to Paint, Write, Compose, and Play. O'Reilly Media.](https://www.amazon.com/Generative-Deep-Learning-Teaching-Machines/dp/1492041947)
### More on Wassertein GAN with Gradient Penalty (optional)
While GANs are a major breakthrough for generative modeling, plain GANs are also notoriously difficult to train. Some common problems encountered are:
* **Oscillating loss:** The loss of the discriminator and generator can start to oscillate without exhibiting any long term stability.
* **Mode collapse:** The generator may get stuck on a small set of samples that always fool the discriminator. This reduces the capability of the network to produce novel samples.
* **Uninformative loss:** The lack of correlation between the generator loss and quality of generated output makes plain GAN training difficult to interpret.
The [Wasserstein GAN](#references) was a major advancement in GANs and helped mitigate to some of these issues. Some of its features are:
1. It significantly improves the interpretability of loss functions and provides clearer stopping criteria
2. WGANs generally produce results of higher quality (demonstrated within the image generation domain)
**Mathematics of Wasserstein GAN with Gradient Penalty**
The [Wasserstein distance](https://en.wikipedia.org/wiki/Wasserstein_metric) between the true distribution $P_r$ and generated piano roll distribution $P_g$ is defined as follows:
$$\mathbb{W}(P_{r},P_{g})=\sup_{\lVert{f} \rVert_{L} \le 1} \mathbb{E}_{x \sim \mathbb{P}_r}(f(x)) - \mathbb{E}_{x \sim \mathbb{P}_g}(f(x)) $$
In this equation we are trying to minimize the distance between the expectation of the real distribution and the expectation of the generation distribution. $f$ is subject to a technical constraint in that it must be [1-Lipschitz](https://en.wikipedia.org/wiki/Lipschitz_continuity).
To enforce the 1-Lipschitz condition that basically constraints the gradients from varying too rapidly we use the gradient penalty.
**Gradient penalty**: We want to penalize the gradients of the critic. We implicitly define $P_{\hat{x}}$ by sampling uniformly along straight lines between pairs of points sampled from the data distribution $P_r$ and the generator distribution $P_g$. This was originally motivated by the fact that the optimal critic contains straight lines with gradient norm 1 connecting coupled points from $P_r$ and $P_g$. We use a penalty coefficient $\lambda$= 10 as was recommended in the original paper.
The loss with gradient penalty is:
$$\mathbb{L}(P_{r},P_{g},P_{\hat{x}} )= \mathbb{W}(P_{r},P_{g}) + \lambda \mathbb{E}_{\hat{x} \sim \mathbb{P}_\hat{x}}[(\lVert \nabla_{\hat{x}}D(\hat{x}) \rVert_2 - 1)^2]$$
|
This loss can be parametrized in terms of $w$ and $\theta$. We then use neural networks to learn the functions $f_w$ (discriminator) and $g_\theta$ (generator).
$$\mathbb{W}(P_{r},P_{\theta})=\max_{w \in \mathbb{W}} \mathbb{E}_{x \sim \mathbb{P}_r}(D_w(x)) - \mathbb{E}_{z \sim p(z)}(D_w(G_{\theta}(z)) $$
$$\mathbb{L}(P_{r},P_{\theta},P_{\hat{x}})=\max_{w \in \mathbb{W}} \mathbb{E}_{x \sim \mathbb{P}_r}(D_w(x)) - \mathbb{E}_{z \sim p(z)}(D_w(G_{\theta}(z)) + \lambda \mathbb{E}_{\hat{x} \sim \mathbb{P}_\hat{x}}[(\lVert \nabla_{\hat{x}}D_w(\hat{x}) \rVert_2 - 1)^2]$$
where $$ \hat{x} = \epsilon x + (1- \epsilon) G(z) $$ and $$\epsilon \sim Unif(0,1)$$
The basic procedure to train is as following:
1. We draw real_x from the real distribution $P_r$ and fake_x from the generated distribution $G_{\theta}(z)$ where $z \sim p(z)$
2. The latent vectors are sampled from z and then tranformed using the generator $G_{\theta}$ to get the fake samples fake_x. They are evaluated using the critic function $D_w$
3. We are trying to minimize the Wasserstein distance between the two distributions
Both the generator and critic are conditioned on the input pianoroll melody.
| github_jupyter |
# Email Content: Isolation Forest based Anomaly Detection
**Attention: **
Because of the limitation of Colab, we cannot call the customized Spark backend. Therefore, this notebook can not run successfully yet.
```
from google.colab import drive
drive.mount('/content/drive')
!apt update
!apt install openjdk-8-jdk-headless
!wget -q https://www-us.apache.org/dist/spark/spark-2.4.7/spark-2.4.7-bin-hadoop2.7.tgz
!tar xf spark-2.4.7-bin-hadoop2.7.tgz
!pip install -q findspark
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.4.7-bin-hadoop2.7"
!update-alternatives --config java
#select openjdk1.8 enter
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
cd /content/drive/MyDrive/Insider-Risk-in-PySpark/spark-iforest/
!apt-get install maven
!mvn clean package -DskipTests
pwd
import os
os.environ["SPARK_HOME"] = "/content/spark-2.4.7-bin-hadoop2.7"
!cp target/spark-iforest-2.4.0.jar $SPARK_HOME/jars/
cd /content/drive/MyDrive/Insider-Risk-in-PySpark/spark-iforest/python/
!python setup.py sdist
!pip install dist/pyspark-iforest-2.4.0.tar.gz
cd /content/drive/MyDrive/Insider-Risk-in-PySpark/
ls /content/spark-2.4.7-bin-hadoop2.7/jars/spark-iforest-2.4.0.jar
import findspark
findspark.init('/content/spark-2.4.7-bin-hadoop2.7')
ls
from pyspark import SparkContext, SparkConf
conf = SparkConf()
conf.set('spark.jars', '/content/drive/MyDrive/Insider-Risk-in-PySpark/spark-iforest/target/spark-iforest-2.4.0.jar')
spark_session = SparkSession.builder.config(conf=conf).appName('IForest').getOrCreate()
cd /content/drive/MyDrive/Insider-Risk-in-PySpark/spark-iforest/python/
!pip install -e .
!spark-submit --jars /content/spark-2.4.7-bin-hadoop2.7/jars/spark-iforest-2.4.0.jar ...
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('proj_if').config("spark.jars", "/content/spark-2.4.7-bin-hadoop2.7/jars/spark-iforest-2.4.0.jar").getOrCreate()
from pyspark_iforest.ml.iforest import *
# Init an IForest Object
iforest = IForest(contamination=0.3, maxDepth=2)
# Fit on a given data frame
model = iforest.fit(df)
# Check if the model has summary or not, the newly trained model has the summary info
model.hasSummary
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pyspark.ml.feature import Tokenizer, StopWordsRemover, NGram, CountVectorizer, StandardScaler
email = spark.read.csv( './data/email.csv',inferSchema=True,header=True)
email.printSchema()
email.show(5)
tokenizer = Tokenizer(inputCol="content", outputCol="words")
wordsData = tokenizer.transform(email)
remover = StopWordsRemover(inputCol="words", outputCol="clean_words")
wordsData = remover.transform(wordsData)
wordsData.show()
cv = CountVectorizer(inputCol="clean_words", outputCol="features", vocabSize=1000, minDF=2.0)
model = cv.fit(wordsData)
wordsCV = model.transform(wordsData)
wordsCV.show()
from pyspark.ml.feature import MinHashLSH
mh = MinHashLSH(inputCol="features", outputCol="hashes", numHashTables=50)
model = mh.fit(wordsCV)
wordsHash = model.transform(wordsCV)
wordsHash.show()
from pyspark.sql.functions import udf
id_hash = wordsHash.select('id','hashes')
id_hash.count()
import pyspark.sql.functions as F
import pyspark.sql.types as T
to_array = F.udf(lambda v: v.toArray().tolist(), T.ArrayType(T.FloatType()))
sc = spark.sparkContext
numAttrs = 50
attrs = sc.parallelize(["hash_" + str(i) for i in range(numAttrs)]).zipWithIndex().collect()
for name, index in attrs:
id_hash = id_hash.withColumn(name, id_hash['hashes'].getItem(index))
id_hash.show()
id_hash.head(1)
```
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/streamlit_notebooks/NER_DE.ipynb)
# **Detect entities in German language**
## 0. Colab Setup
```
!sudo apt-get install openjdk-8-jdk
!java -version
!pip install --ignore-installed -q pyspark==2.4.4
!pip install spark-nlp
import pandas as pd
import numpy as np
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
import json
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
from sparknlp.annotator import *
from sparknlp.base import *
import sparknlp
from sparknlp.pretrained import PretrainedPipeline
```
## 1. Start Spark Session
```
spark = sparknlp.start()
```
## 2. Select the DL model
```
### Select the model and re-run all the cells below ####
MODEL_NAME = 'wikiner_840B_300'
```
## 3. Some sample examples
```
## Generating Example Files ##
text_list = ["""William Henry Gates III (* 28. Oktober 1955 in London) ist ein US-amerikanischer Geschäftsmann, Softwareentwickler, Investor und Philanthrop. Er ist bekannt als Mitbegründer der Microsoft Corporation. Während seiner Karriere bei Microsoft war Gates Vorsitzender, Chief Executive Officer (CEO), Präsident und Chief Software Architect und bis Mai 2014 der größte Einzelaktionär. Er ist einer der bekanntesten Unternehmer und Pioniere der Mikrocomputer-Revolution der 1970er und 1980er Jahre. Gates wurde in Seattle, Washington, geboren und wuchs dort auf. 1975 gründete er Microsoft zusammen mit seinem Freund aus Kindertagen, Paul Allen, in Albuquerque, New Mexico. Es entwickelte sich zum weltweit größten Unternehmen für Personal-Computer-Software. Gates leitete das Unternehmen als Chairman und CEO, bis er im Januar 2000 als CEO zurücktrat. Er blieb jedoch Chairman und wurde Chief Software Architect. In den späten neunziger Jahren wurde Gates für seine Geschäftstaktiken kritisiert, die als wettbewerbswidrig angesehen wurden. Diese Meinung wurde durch zahlreiche Gerichtsurteile bestätigt. Im Juni 2006 gab Gates bekannt, dass er eine Teilzeitstelle bei Microsoft und eine Vollzeitstelle bei der Bill & Melinda Gates Foundation, der privaten gemeinnützigen Stiftung, die er und seine Frau Melinda Gates im Jahr 2000 gegründet haben, übernehmen wird. [ 9] Er übertrug seine Aufgaben nach und nach auf Ray Ozzie und Craig Mundie. Im Februar 2014 trat er als Vorsitzender von Microsoft zurück und übernahm eine neue Position als Technologieberater, um den neu ernannten CEO Satya Nadella zu unterstützen.""",
"""Die Mona Lisa ist ein Ölgemälde aus dem 16. Jahrhundert, das von Leonardo geschaffen wurde. Es findet im Louvre in Paris statt.""",
]
```
## 4. Define Spark NLP pipeline
```
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
tokenizer = Tokenizer() \
.setInputCols(["document"]) \
.setOutputCol("token")
embeddings = WordEmbeddingsModel.pretrained('glove_840B_300', lang='xx').\
setInputCols(["document", 'token']).\
setOutputCol("embeddings")
public_ner = NerDLModel.pretrained(MODEL_NAME, 'de') \
.setInputCols(["document", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter() \
.setInputCols(["document", "token", "ner"]) \
.setOutputCol("ner_chunk")
nlpPipeline = Pipeline(stages=[ documentAssembler,
tokenizer,
embeddings,
public_ner,
ner_converter
])
```
## 5. Run the pipeline
```
empty_df = spark.createDataFrame([['']]).toDF("text")
pipelineModel = nlpPipeline.fit(empty_df)
df = spark.createDataFrame(pd.DataFrame({"text":text_list}))
result = pipelineModel.transform(df)
```
## 6. Visualize results
```
result.select(F.explode(F.arrays_zip('ner_chunk.result', 'ner_chunk.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
```
| github_jupyter |
# Energy auto-encoder: classification
* Technically akin to "transductive learning" because the sparse auto-encoder dictionary is learned over the whole dataset, not only the training data. The LeCun paper we compare with does the same. This could be solved by including the dictionary learning step in the classifier. Technical solutions:
1. Compute the dictionary in our custom classifier.
2. Create a scikit-learn Pipeline which includes the whole preprocessing and feature extraction steps.
In either case the ability to import functions from other notebooks would help. This would be very slow due to the tremendous amount of time needed to train the auto-encoder.
* We should use "grid search" to find the optimal hyper-parameters (auto-encoders, frames, feature vectors, SVM).
* We may use a validation set to mitigate the leak of the testing set in the model as we tune the hyper-parameters.
* Even if not stated in LeCun's paper we should rescale the data before SVM classification.
For the auto-encoder learning to converge we should rescale beforehand. As the transformation preserves energy, there is no need to rescale again.
## Hyper-parameters
* `scale`: scaling (None, minmax, std)
* `Nvectors`: number of feature vectors per song.
* `svm_type`: C-SVM (C) or nu-SVM (nu).
* `kernel`: C-SVM kernel (linear, rbf).
* `C`: penalty parameter C of the error term.
* `nu`: an upper bound on the fraction of training errors and a lower bound of the fraction of support vectors.
* `majority_voting`: When `True`, each of the 2`Nvectors` votes for one label and the accuracy is computed on the classification of the whole clips. When `False`, the accuracy is computed on the classification of the feature vectors.
* `test_size`: proportion of testing data for cross-validation.
* `Ncv`: number of cross-validation runs, in multiple of 10.
* `dataset_classification`: the dataset to use for classification (X, Z). It allows to compare with the baseline, i.e. spectrograms.
* `Ngenres, Nclips, Nframes`: a way to reduce the size of the dataset.
* `folder`: relative path to HDF5 files.
* `filename_*`: name of the HDF5 file.
```
if 'p' in globals().keys():
# Hyper-parameters passed by the experiment runner.
for key, value in p.items():
globals()[key] = value
else:
scale = 'minmax'
Nvectors = 6
svm_type = 'C'
kernel = 'linear'
C = 1
nu = 0.5
majority_voting = True
test_size = 0.1
Ncv = 20
dataset_classification = 'Z'
Ngenres, Nclips, Nframes = 10, 100, 644
folder = 'data'
filename_features = 'features.hdf5'
```
## Setup
```
import os, time
import numpy as np
import sklearn
from sklearn import svm
from sklearn import cross_validation
from sklearn import metrics
from sklearn import preprocessing
import h5py
import matplotlib.pyplot as plt
%matplotlib inline
print('Software versions:')
for pkg in [np, sklearn]:
print(' {}: {}'.format(pkg.__name__, pkg.__version__))
toverall = time.time()
```
## Input data
1. Retrieve data from the HDF5 data store.
2. Choose the data we want to work with:
* raw audio $X_a$,
* CQT spectrograms $X_s$,
* normalized spectrograms $X_n$,
* sparse codes $Z$.
3. Eventually reduce the number $N_{genres} \cdot N_{clips}$ of samples for quicker analysis.
```
def datinfo(X, name='Dataset'):
r"""Print dataset size and dimensionality"""
print('{}:\n'
' size: N={:,} x n={} -> {:,} floats\n'
' dim: {:,} features per clip\n'
' shape: {}'
.format(name, np.prod(X.shape[:-1]), X.shape[-1],
np.prod(X.shape), np.prod(X.shape[2:]), X.shape))
filename = os.path.join(folder, filename_features)
with h5py.File(filename, 'r') as audio:
# Display HDF5 attributes.
print('Attributes:')
for attr in audio.attrs:
print(' {} = {}'.format(attr, audio.attrs[attr]))
labels = audio.attrs['labels']
# Show datasets, their dimensionality and data type.
print('Datasets:')
for dname, dset in audio.items():
print(' {:2}: {:24}, {}'.format(dname, dset.shape, dset.dtype))
# Choose dataset: Xa, Xs, Z.
X = audio.get(dataset_classification)
# Full dataset.
n = X.shape[-1]
datinfo(X, 'Full dataset')
print(type(X))
# Load data into memory as a standard NumPy array.
X = X[:Ngenres,:Nclips,:Nframes,...]
datinfo(X, 'Reduced dataset')
print(type(X))
# Resize in place without memory loading via hyperslab.
# Require chunked datasets.
#X.resize((Ngenres, Nclips, Nframes, 2, n))
```
## Feature vectors through aggregation
Yet another (hopefully intelligent) dimensionality reduction:
* Aggregation of features from various frames to make up $2N_{vectors} = 12$ feature vectors per clip. Each vector represents approximatly 5 seconds of audio which is way longer than single frames while shorter than the whole clip.
* There is again a 50% overlap between those feature vectors.
* Absolute value rectification to prevent components of different sign from canceling each other out. May be worth disabling when working with raw audio ($X_a$). It then makes sense to rescale with 'std' instead of 'minmax'.
* Can be thought of as an histogram of used dictionary atoms (if using $Z$) or frequency bins (if using $X_s$) along the chosen time window.
```
# Flatten consecutive frames in time.
X.resize((Ngenres, Nclips, 2*Nframes, n))
#assert np.all(X1[1,4,3,:] == X[1,4,1,1,:])
datinfo(X, 'Flattened frames')
# Parameters.
Nframes_per_vector = int(np.floor(2 * Nframes / (Nvectors+0.5)))
def aggregate(X, absrect=True):
# Truncate.
X = X[:,:,:Nvectors*Nframes_per_vector,:]
# Group.
X = X.reshape((Ngenres, Nclips, Nvectors, Nframes_per_vector, n))
datinfo(X, 'Truncated and grouped')
# Aggregate.
if absrect:
return np.sum(np.abs(X), axis=3)
else:
return np.sum(X, axis=3)
# Feature vectors.
Y = np.empty((Ngenres, Nclips, Nvectors, 2, n))
Y[:,:,:,0,:] = aggregate(X) # Aligned.
Y[:,:,:,1,:] = aggregate(X[:,:,Nframes_per_vector/2:,:]) # Ovelapped.
datinfo(Y, 'Feature vectors')
# Free memory.
del(X)
```
## Feature vectors visualization
Visualize all feature vectors of a given clip.
Observations:
* Classical music seems to have a much denser spectrum than blues, which may explain why these two classes are easily identifiable using $X_s$.
* Country seems to have strong low frequencies.
```
genre, clip = 0, 7
fig = plt.figure(figsize=(8,5))
fig.suptitle('12 feature vectors each covering 5 seconds with 50% overlap')
for vector in range(Nvectors):
for k in range(2):
i = vector*2+k
ax = fig.add_subplot(4, 3, i)
ax.plot(Y[genre,clip,vector,k,:])
ax.set_xlim((0, n))
ax.set_xticks([])
ax.set_yticks([])
```
## Data preparation for classification
1. Rearrange dataset as a 2D array: number of samples x dimensionality.
2. Optionally scale the data.
3. Generate labels.
4. Optionally split in training and testing sets.
5. Optionally randomize labels for testing.
Observations:
* Scaling is necessary for classification performance (both accuracy and speed). 'std' scaling is not well suited to our histogram-like feature vectors which are not at all Gaussian distributions. Prefer 'minmax', i.e. scale features in [0,1]. Moreover this scaling will preserve the sparsity when dealing with sparse codes $Z$.
```
def prepdata(a, b, c, test_size=None, scale=None, rand=False):
"""Prepare data for classification."""
# Squeeze dataset to a 2D array.
data = Y.reshape((a*b), c)
if c == n:
assert np.all(data[31,:] == Y[0,2,3,1,:])
elif c == Nvectors*2*n:
assert np.all(data[Nclips+2,:] == Y[1,2,:,:,:].reshape(-1))
# Independently rescale each feature.
# To be put in an sklearn Pipeline to avoid transductive learning.
if scale is 'std':
# Features have zero norm and unit standard deviation.
data = preprocessing.scale(data, axis=0)
elif scale is 'minmax':
# Features in [0,1].
data -= np.min(data, axis=0)
data /= np.max(data, axis=0)
#print(np.min(data, axis=0))
#print(np.max(data, axis=0))
# Labels.
target = np.empty((a, b), dtype=np.uint8)
for genre in range(Ngenres):
target[genre,:] = genre
target.resize(data.shape[0])
print('{} genres: {}'.format(Ngenres, ', '.join(labels[:Ngenres])))
# Be sure that classification with random labels is no better than random.
if rand:
target = np.floor(np.random.uniform(0, Ngenres, target.shape))
print('Balance: {} {}'.format(np.sum(target == 0), np.sum(target == 1)))
# Training and testing sets.
if test_size is not None:
X_train, X_test, y_train, y_test = cross_validation.train_test_split(
data, target, test_size=test_size) # random_state=1
print('Training data: {}, {}'.format(X_train.shape, X_train.dtype))
print('Testing data: {}, {}'.format(X_test.shape, X_test.dtype))
print('Training labels: {}, {}'.format(y_train.shape, y_train.dtype))
print('Testing labels: {}, {}'.format(y_test.shape, y_test.dtype))
return X_train, X_test, y_train, y_test
else:
print('Data: {}, {}'.format(data.shape, data.dtype))
print('Labels: {}, {}'.format(target.shape, target.dtype))
return data, target
```
## Linear SVM
* Each feature vector gets a genre label.
* Classification with linear Vector Support Machine (SVM).
* Fast to train.
* Scale well to large dataset.
* Two implementations: liblinear (sklearn LinearSVC) and libsvm (sklearn SVC and NuSVC)
* Multi-class: "one-vs-one" approach (Knerr et al., 1990) (sklearn SVC and NuSVC) and "one-vs-the-rest" (sklearn LinearSVC)
Observations:
* We can predict genre labels of individual frames with good accuracy using CQT spectrograms only.
* SVC vs NuSVC vs LinearSVC:
* 10-fold cross-validation with 10 classes (default $C=1$ and $\nu=0.5$):
* SVC (0.56) yields better accuracy than LinearSVC (0.53) than NuSVC (0.51)
* SVC (303s) and LinearSVC (296s) faster than NuSVC (501s)
* SVC does often not converge if data is not scaled
* LinearSVC may be more scalable (in the number of samples)
* Hyper-parameters:
* $C$ seems to have little impact.
* $\nu$ has a great impact on speed: lower is slower
Open questions:
* Which multi-class strategy to adopt: one-vs-all or one-vs-one ?
* sklearn states that one-vs-all is the most common strategy
* Determine $C$ or $\nu$.
```
# Instantiate a classifier.
if svm_type is 'C':
clf_svm = svm.SVC(kernel=kernel, C=C)
elif svm_type is 'nu':
clf_svm = svm.NuSVC(kernel=kernel, nu=nu)
#clf_svm = svm.LinearSVC(C=1)
# Try the single feature vector classifier (linear SVM).
if True:
# Split data.
X_train, X_test, y_train, y_test = prepdata(
Ngenres, Nclips*Nvectors*2, n, test_size=0.4,
scale=scale, rand=False)
# Train.
clf_svm.fit(X_train, y_train)
# Test.
y_predict = clf_svm.predict(X_test)
acc = metrics.accuracy_score(y_test, y_predict)
print('Accuracy: {:.1f} %'.format(acc*100))
```
## Majority voting
Final dimensionality reduction step:
* Each of the 12 feature vectors of a clip gives a vote. We choose the genre with the highest number of votes.
* Implemented as a custom classifier which embeds an SVM for individual feature vectors classification.
* Alternative implementation: insert in a sklearn pipeline after SVC.
Observations:
* Accuracy on whole clips is indeed better than accuracy on individual feature vectors.
* *clf_svm_vote.confidence* is useful to observe if a class is harder to differentiate.
```
# Define and instantiate our custom classifier.
class svm_vote(sklearn.base.BaseEstimator):
def __init__(self, svm):
self.svm = svm
def _vectors(self, X, y=None):
"""Rearrange data in feature vectors for SVM."""
X = X.reshape(X.shape[0]*Nvectors*2, n)
if y is not None:
y = np.repeat(y, Nvectors*2, axis=0)
assert y.shape[0] == X.shape[0]
return (X, y)
else:
return (X,)
def fit(self, X, y):
"""Fit the embedded SVC."""
self.svm.fit(*self._vectors(X, y))
def svm_score(self, X, y):
"""Return SVC accuracy on feature vectors."""
return self.svm.score(*self._vectors(X, y))
def svm_predict(self, X):
"""Return SVC predictions on feature vectors."""
y = self.svm.predict(*self._vectors(X))
y.resize(X.shape[0], Nvectors*2)
return y
def confidence(self, X):
"""Return the number of votes for each class."""
def bincount(x):
return np.bincount(x, minlength=Ngenres)
y = np.apply_along_axis(bincount, 1, self.svm_predict(X))
assert np.all(np.sum(y, axis=1) == Nvectors*2)
return y
def predict(self, X):
"""Return predictions on whole clips."""
y = self.svm_predict(X)
return np.apply_along_axis(lambda x: np.bincount(x).argmax(), 1, y)
#return np.zeros(X.shape[0]) # Pretty bad prediction.
def score(self, X, y):
"""Return the accuracy score. Used by sklearn cross-validation."""
return metrics.accuracy_score(y, self.predict(X))
clf_svm_vote = svm_vote(clf_svm)
# Try the whole clip classifier (linear SVM and majority voting).
if True:
# Split data.
X_train, X_test, y_train, y_test = prepdata(
Ngenres, Nclips, Nvectors*2*n, test_size=0.4,
scale=scale, rand=False)
# Train.
clf_svm_vote.fit(X_train, y_train)
# Test on single vectors.
acc = clf_svm_vote.svm_score(X_test, y_test)
print('Feature vectors accuracy: {:.1f} %'.format(acc*100))
# Observe individual votes.
#print(clf_svm_vote.svm_predict(X_test))
#print(clf_svm_vote.confidence(X_test))
# Test on whole clips.
y_predict = clf_svm_vote.predict(X_test)
acc = metrics.accuracy_score(y_test, y_predict)
assert acc == clf_svm_vote.score(X_test, y_test)
print('Clips accuracy: {:.1f} %'.format(acc*100))
```
## Cross-validation
* 10-fold cross-validation.
* 100 randomly chosen clips per fold.
* 9 folds (900 clips) for training, 1 fold (100 clips) for testing.
* Determine a classification accuracy using testing set.
* Repeat 10 times: mean and standard deviation.
Observations:
* Data should be shuffled as samples with the same label are contiguous, i.e. data ordering is not arbitrary.
* *ShuffleSplit*, *StratifiedShuffleSplit*, *KFold* and *StratifiedKFold* yields similar results as long as data is shuffeld.
* (Lots of variance between runs.)
* Data should be rescaled for good performance (both accuracy and speed).
Results:
* With $X_a$ (best observed)
* Accuracy of 89 (+/- 5.0) for 2 genres (SVC, abs, minmax) (50s)
* Accuracy of 60 (+/- 7.9) for 2 genres (SVC, noabs, minmax) (100s)
* Accuracy of 64 (+/- 11.1) for 2 genres (SVC, noabs, std) (1000s)
* With $X_s$ (best observed)
* Accuracy of 96 (+/- 4.7) for 2 genres (SVC, minmax) (2s, CDK 1s)
* Accuracy of 81 (+/- 4) for 4 genres (SVC, minmax) (14s)
* Accuracy of 76 (+/- 3.1) for 5 genres (SVC, minmax) (CDK 12s)
* Accuracy of 56 (+/- 5) for 10 genres (SVC, minmax) (300s)
* Accuracy of 53 (+/- 3) for 10 genres (LinearSVC, minmax) (300s)
* Accuracy of 51 (+/- 5) for 10 genres (NuSVC, minmax) (500s)
* With $Z$ (best observed) (all with SVC, no normalization if not mentioned)
* Accuracy of 96 (+/- 3.2) for 2 genres (ld=10, m=128, minmax, 10 outer) (CDK 1s)
* Accuracy of 98 (+/- 2.4) for 2 genres (ld=10, m=128, 10 outer) (CDK 1s)
* Accuracy of 98 (+/- 2.5) for 2 genres (ld=10, m=128, encoder, 10 outer) (CDK 1s)
* Accuracy of 98 (+/- 2.5) for 2 genres (ld=10, m=128, 20 outer) (CDK 1s)
* Accuracy of 98 (+/- 2.5) for 2 genres (ld=100, m=128, 15 outer) (CDK 1s)
* Accuracy of 58 (+/- 10.5) for 2 genres (ld=1, m=128, 15 outer) (CDK 4s)
* Accuracy of 99 (+/- 3.2) for 2 genres (ld=10, m=512, 15 outer) (CDK 2s)
* Accuracy of 79 (+/- 2.7) for 5 genres (ld=10, m=512, 20 outer) (CDK 28s)
* Accuracy of 65 (+/- 3.6) for 10 genres (ld=10, m=512, 15 outer) (CDK 167s)
Ideas:
* Use the area under the receiver operating characteristing (ROC) curve (AUC). Not sure if applicable to multi-class.
```
if majority_voting:
clf = clf_svm_vote
b = Nclips
c = Nvectors*2*n
else:
clf = clf_svm
b = Nclips*Nvectors*2
c = n
data, target = prepdata(Ngenres, b, c, scale=scale)
print('Ratio: {} training, {} testing'.format(
(1-test_size)*target.size, test_size*target.size))
tstart = time.time()
scores = np.empty(shape=(Ncv, 10))
# Cross-validation iterators.
cv = cross_validation.ShuffleSplit(target.size, n_iter=10, test_size=test_size)
#cv = cross_validation.StratifiedShuffleSplit(target.size, n_iter=10, test_size=test_size)
#cv = cross_validation.KFold(target.size, shuffle=True, n_folds=10)
#cv = cross_validation.StratifiedKFold(target, shuffle=True, n_folds=10)
for i in range(Ncv):
scores[i,:] = cross_validation.cross_val_score(
clf, data, target, cv=cv, n_jobs=1)
# Performance: accuracy.
mean, std = scores[i,:].mean()*100, scores[i,:].std()*100
print(' {:3.0f} (+/-{:4.1f}) <- {}'.format(mean, std, (scores[i,:]*100).astype(np.int)))
accuracy, accuracy_std = scores.mean()*100, scores.std()*100
print('Accuracy: {:.1f} (+/- {:.2f})'.format(accuracy, accuracy_std))
meantime = (time.time() - tstart) / Ncv
print('Mean time ({} cv): {:.2f} seconds'.format(Ncv, meantime))
print('Overall time: {:.2f} seconds'.format(time.time() - toverall))
```
| github_jupyter |
# Quick-Start Guide
This documents presents an overview of how to run a simple example using the `Minas` classifier. It was implemented by extending the base estimator from `scikit-multiflow`, so it works similarly to the popular `scikit-learn` API.
The `Minas` classifier defines the following methods:
- `fit` – Trains a model in the offline phase, in a batch fashion.
- `partial_fit` – Incrementally trains the stream model.
- `predict` – Predicts the target’s value.
## Train and test a stream classification model using `Minas`
1. Before we start, we have to import the classifier.
```
from minas import Minas
```
2. Create a stream
We create a stream using `RandomRBFGenerator`, the Random Radial Basis Function stream generator from `scikit-multiflow`. In this example, we will create a data stream with 3 classes, 4 features, and 6 centroids.
Also, before using the stream, we need to prepare it by calling `prepare_for_use()`.
```
from skmultiflow.data.random_rbf_generator import RandomRBFGenerator
stream = RandomRBFGenerator(model_random_state=123,
sample_random_state=12,
n_classes=3,
n_features=4,
n_centroids=6)
stream.prepare_for_use()
```
3. Instantiate the `Minas` classifier
The classifier takes the following parameters:
- `kini`: Number of clusters for each class to be found during clustering (offline phase and novelty detection process).
- `cluster_algorithm`: A string containing the clustering algorithm to use. Currently only supports `'kmeans'`.
- `random_state`: Seed to use for random number generation.
- `min_short_mem_trigger`: Minimum number of samples in the short term memory required to trigger a novelty detection process.
- `min_examples_cluster`: Minimum number of examples required to form a cluster.
- `threshold_strategy`: Strategy used to compute the threshold for differentiating between novelty classes and concept extensions. Accepts `1`, `2`, or `3`. The strategies are defined in the [MINAS paper](http://www.liaad.up.pt/area/jgama/MINAS.pdf).
- `threshold_factor`: Factor to use for calculating thresholds.
- `window_size`: Window size (an integer representing the number of samples) used by the forgetting mechanism.
- `update_summary`: Defaults to `False`. If `True`, the summary statistics for a cluster are updated when a new point is added to it.
- `animation`: Defaults to `False`. If `True`, a plot is created showing the current state of the model (points and clusters). It only works if the examples have two dimensions.
For this example, we will set 10 clusters per class, with at least 30 examples required in the short term memory before triggering a novelty detection procedure, and a minimum of 10 examples per cluster.
```
clf = Minas(kini=10,
min_short_mem_trigger=30,
min_examples_cluster=10)
```
4. Get data from the stream
Next, we will get the data from the stream. For this example, we use 500 samples to train our model in the offline phase. Then, we will use the next 500 samples for the online phase.
```
n_samples = 1000
offline_size = 500
X_all, y_all = stream.next_sample(n_samples)
X_train = X_all[:offline_size]
y_train = y_all[:offline_size]
X_test = X_all[offline_size:n_samples]
y_test = y_all[offline_size:n_samples]
```
5. Offline phase
The next step corresponds to the offline phase. We run it by calling `fit()` with the training data from the last step.
```
# OFFLINE phase
clf.fit(X_train, y_train)
```
6. Online phase
Now we get to the online phase. We feed each example at a time to the model, and collect the results from the `predict()` calls. In each iteration, we call `partial_fit()` to update the model with the data from the sample it has just seen.
```
# ONLINE phase
y_preds = []
for X, y in zip(X_test, y_test):
y_preds.append(clf.predict([X])[0])
clf.partial_fit([X], [y])
```
7. Evaluate performance
Finally, we will see how our model performed. We create a confusion matrix with the following definition:
- Each row represents the true label of the examples from the stream seen during the online phase.
- Each column represents the label predicted by the model. The numbers greater than the maximum labels shown in the rows represent the novelty patterns detected. Unknown samples are represented by `-1`.
In the confusion matrix below, for our example, the labels from our data set are 0, 1, and 2. The columns 3, 4, and 5 represent 3 novelty patterns discovered by the model. We can see that the model identified classes 0 and 2 quite well, and had more trouble distinguishing examples from class 1.
```
clf.confusion_matrix(X_test, y_test)
```
| github_jupyter |
# Gradient Checking
Welcome to the final assignment for this week! In this assignment you will learn to implement and use gradient checking.
You are part of a team working to make mobile payments available globally, and are asked to build a deep learning model to detect fraud--whenever someone makes a payment, you want to see if the payment might be fraudulent, such as if the user's account has been taken over by a hacker.
But backpropagation is quite challenging to implement, and sometimes has bugs. Because this is a mission-critical application, your company's CEO wants to be really certain that your implementation of backpropagation is correct. Your CEO says, "Give me a proof that your backpropagation is actually working!" To give this reassurance, you are going to use "gradient checking".
Let's do it!
```
# Packages
import numpy as np
from testCases import *
from gc_utils import sigmoid, relu, dictionary_to_vector, vector_to_dictionary, gradients_to_vector
```
## 1) How does gradient checking work?
Backpropagation computes the gradients $\frac{\partial J}{\partial \theta}$, where $\theta$ denotes the parameters of the model. $J$ is computed using forward propagation and your loss function.
Because forward propagation is relatively easy to implement, you're confident you got that right, and so you're almost 100% sure that you're computing the cost $J$ correctly. Thus, you can use your code for computing $J$ to verify the code for computing $\frac{\partial J}{\partial \theta}$.
Let's look back at the definition of a derivative (or gradient):
$$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
If you're not familiar with the "$\displaystyle \lim_{\varepsilon \to 0}$" notation, it's just a way of saying "when $\varepsilon$ is really really small."
We know the following:
- $\frac{\partial J}{\partial \theta}$ is what you want to make sure you're computing correctly.
- You can compute $J(\theta + \varepsilon)$ and $J(\theta - \varepsilon)$ (in the case that $\theta$ is a real number), since you're confident your implementation for $J$ is correct.
Lets use equation (1) and a small value for $\varepsilon$ to convince your CEO that your code for computing $\frac{\partial J}{\partial \theta}$ is correct!
## 2) 1-dimensional gradient checking
Consider a 1D linear function $J(\theta) = \theta x$. The model contains only a single real-valued parameter $\theta$, and takes $x$ as input.
You will implement code to compute $J(.)$ and its derivative $\frac{\partial J}{\partial \theta}$. You will then use gradient checking to make sure your derivative computation for $J$ is correct.
<img src="images/1Dgrad_kiank.png" style="width:600px;height:250px;">
<caption><center> <u> **Figure 1** </u>: **1D linear model**<br> </center></caption>
The diagram above shows the key computation steps: First start with $x$, then evaluate the function $J(x)$ ("forward propagation"). Then compute the derivative $\frac{\partial J}{\partial \theta}$ ("backward propagation").
**Exercise**: implement "forward propagation" and "backward propagation" for this simple function. I.e., compute both $J(.)$ ("forward propagation") and its derivative with respect to $\theta$ ("backward propagation"), in two separate functions.
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(x, theta):
"""
Implement the linear forward propagation (compute J) presented in Figure 1 (J(theta) = theta * x)
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
J -- the value of function J, computed using the formula J(theta) = theta * x
"""
### START CODE HERE ### (approx. 1 line)
J = theta * x
### END CODE HERE ###
return J
x, theta = 2, 4
J = forward_propagation(x, theta)
print ("J = " + str(J))
```
**Expected Output**:
<table style=>
<tr>
<td> ** J ** </td>
<td> 8</td>
</tr>
</table>
**Exercise**: Now, implement the backward propagation step (derivative computation) of Figure 1. That is, compute the derivative of $J(\theta) = \theta x$ with respect to $\theta$. To save you from doing the calculus, you should get $dtheta = \frac { \partial J }{ \partial \theta} = x$.
```
# GRADED FUNCTION: backward_propagation
def backward_propagation(x, theta):
"""
Computes the derivative of J with respect to theta (see Figure 1).
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
Returns:
dtheta -- the gradient of the cost with respect to theta
"""
### START CODE HERE ### (approx. 1 line)
dtheta = x
### END CODE HERE ###
return dtheta
x, theta = 2, 4
dtheta = backward_propagation(x, theta)
print ("dtheta = " + str(dtheta))
```
**Expected Output**:
<table>
<tr>
<td> ** dtheta ** </td>
<td> 2 </td>
</tr>
</table>
**Exercise**: To show that the `backward_propagation()` function is correctly computing the gradient $\frac{\partial J}{\partial \theta}$, let's implement gradient checking.
**Instructions**:
- First compute "gradapprox" using the formula above (1) and a small value of $\varepsilon$. Here are the Steps to follow:
1. $\theta^{+} = \theta + \varepsilon$
2. $\theta^{-} = \theta - \varepsilon$
3. $J^{+} = J(\theta^{+})$
4. $J^{-} = J(\theta^{-})$
5. $gradapprox = \frac{J^{+} - J^{-}}{2 \varepsilon}$
- Then compute the gradient using backward propagation, and store the result in a variable "grad"
- Finally, compute the relative difference between "gradapprox" and the "grad" using the following formula:
$$ difference = \frac {\mid\mid grad - gradapprox \mid\mid_2}{\mid\mid grad \mid\mid_2 + \mid\mid gradapprox \mid\mid_2} \tag{2}$$
You will need 3 Steps to compute this formula:
- 1'. compute the numerator using np.linalg.norm(...)
- 2'. compute the denominator. You will need to call np.linalg.norm(...) twice.
- 3'. divide them.
- If this difference is small (say less than $10^{-7}$), you can be quite confident that you have computed your gradient correctly. Otherwise, there may be a mistake in the gradient computation.
```
# GRADED FUNCTION: gradient_check
def gradient_check(x, theta, epsilon = 1e-7):
"""
Implement the backward propagation presented in Figure 1.
Arguments:
x -- a real-valued input
theta -- our parameter, a real number as well
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Compute gradapprox using left side of formula (1). epsilon is small enough, you don't need to worry about the limit.
### START CODE HERE ### (approx. 5 lines)
thetaplus = theta + epsilon # Step 1
thetaminus = theta - epsilon # Step 2
J_plus = thetaplus * x # Step 3
J_minus = thetaminus * x # Step 4
gradapprox = (J_plus - J_minus) / (2*epsilon) # Step 5
### END CODE HERE ###
# Check if gradapprox is close enough to the output of backward_propagation()
### START CODE HERE ### (approx. 1 line)
grad = backward_propagation(x, theta)
### END CODE HERE ###
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator / denominator # Step 3'
### END CODE HERE ###
if difference < 1e-7:
print ("The gradient is correct!")
else:
print ("The gradient is wrong!")
return difference
x, theta = 2, 4
difference = gradient_check(x, theta)
print("difference = " + str(difference))
```
**Expected Output**:
The gradient is correct!
<table>
<tr>
<td> ** difference ** </td>
<td> 2.9193358103083e-10 </td>
</tr>
</table>
Congrats, the difference is smaller than the $10^{-7}$ threshold. So you can have high confidence that you've correctly computed the gradient in `backward_propagation()`.
Now, in the more general case, your cost function $J$ has more than a single 1D input. When you are training a neural network, $\theta$ actually consists of multiple matrices $W^{[l]}$ and biases $b^{[l]}$! It is important to know how to do a gradient check with higher-dimensional inputs. Let's do it!
## 3) N-dimensional gradient checking
The following figure describes the forward and backward propagation of your fraud detection model.
<img src="images/NDgrad_kiank.png" style="width:600px;height:400px;">
<caption><center> <u> **Figure 2** </u>: **deep neural network**<br>*LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID*</center></caption>
Let's look at your implementations for forward propagation and backward propagation.
```
def forward_propagation_n(X, Y, parameters):
"""
Implements the forward propagation (and computes the cost) presented in Figure 3.
Arguments:
X -- training set for m examples
Y -- labels for m examples
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape (5, 4)
b1 -- bias vector of shape (5, 1)
W2 -- weight matrix of shape (3, 5)
b2 -- bias vector of shape (3, 1)
W3 -- weight matrix of shape (1, 3)
b3 -- bias vector of shape (1, 1)
Returns:
cost -- the cost function (logistic cost for one example)
"""
# retrieve parameters
m = X.shape[1]
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
# Cost
logprobs = np.multiply(-np.log(A3),Y) + np.multiply(-np.log(1 - A3), 1 - Y)
cost = 1./m * np.sum(logprobs)
cache = (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
return cost, cache
```
Now, run backward propagation.
```
def backward_propagation_n(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input datapoint, of shape (input size, 1)
Y -- true "label"
cache -- cache output from forward_propagation_n()
Returns:
gradients -- A dictionary with the gradients of the cost with respect to each parameter, activation and pre-activation variables.
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1./m * np.dot(dZ3, A2.T)
db3 = 1./m * np.sum(dZ3, axis=1, keepdims = True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1./m * np.dot(dZ2, A1.T)
db2 = 1./m * np.sum(dZ2, axis=1, keepdims = True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1./m * np.dot(dZ1, X.T)
db1 = 1./m * np.sum(dZ1, axis=1, keepdims = True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,
"dA2": dA2, "dZ2": dZ2, "dW2": dW2, "db2": db2,
"dA1": dA1, "dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
```
You obtained some results on the fraud detection test set but you are not 100% sure of your model. Nobody's perfect! Let's implement gradient checking to verify if your gradients are correct.
**How does gradient checking work?**.
As in 1) and 2), you want to compare "gradapprox" to the gradient computed by backpropagation. The formula is still:
$$ \frac{\partial J}{\partial \theta} = \lim_{\varepsilon \to 0} \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon} \tag{1}$$
However, $\theta$ is not a scalar anymore. It is a dictionary called "parameters". We implemented a function "`dictionary_to_vector()`" for you. It converts the "parameters" dictionary into a vector called "values", obtained by reshaping all parameters (W1, b1, W2, b2, W3, b3) into vectors and concatenating them.
The inverse function is "`vector_to_dictionary`" which outputs back the "parameters" dictionary.
<img src="images/dictionary_to_vector.png" style="width:600px;height:400px;">
<caption><center> <u> **Figure 2** </u>: **dictionary_to_vector() and vector_to_dictionary()**<br> You will need these functions in gradient_check_n()</center></caption>
We have also converted the "gradients" dictionary into a vector "grad" using gradients_to_vector(). You don't need to worry about that.
**Exercise**: Implement gradient_check_n().
**Instructions**: Here is pseudo-code that will help you implement the gradient check.
For each i in num_parameters:
- To compute `J_plus[i]`:
1. Set $\theta^{+}$ to `np.copy(parameters_values)`
2. Set $\theta^{+}_i$ to $\theta^{+}_i + \varepsilon$
3. Calculate $J^{+}_i$ using to `forward_propagation_n(x, y, vector_to_dictionary(`$\theta^{+}$ `))`.
- To compute `J_minus[i]`: do the same thing with $\theta^{-}$
- Compute $gradapprox[i] = \frac{J^{+}_i - J^{-}_i}{2 \varepsilon}$
Thus, you get a vector gradapprox, where gradapprox[i] is an approximation of the gradient with respect to `parameter_values[i]`. You can now compare this gradapprox vector to the gradients vector from backpropagation. Just like for the 1D case (Steps 1', 2', 3'), compute:
$$ difference = \frac {\| grad - gradapprox \|_2}{\| grad \|_2 + \| gradapprox \|_2 } \tag{3}$$
```
# GRADED FUNCTION: gradient_check_n
def gradient_check_n(parameters, gradients, X, Y, epsilon = 1e-7):
"""
Checks if backward_propagation_n computes correctly the gradient of the cost output by forward_propagation_n
Arguments:
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
gradients -- output of backward_propagation_n, contains gradients of the cost with respect to the parameters.
x -- input datapoint, of shape (input size, 1)
y -- true "label"
epsilon -- tiny shift to the input to compute approximated gradient with formula(1)
Returns:
difference -- difference (2) between the approximated gradient and the backward propagation gradient
"""
# Set-up variables
parameters_values, _ = dictionary_to_vector(parameters)
grad = gradients_to_vector(gradients)
num_parameters = parameters_values.shape[0]
J_plus = np.zeros((num_parameters, 1))
J_minus = np.zeros((num_parameters, 1))
gradapprox = np.zeros((num_parameters, 1))
# print(parameters_values.shape)
# Compute gradapprox
for i in range(num_parameters):
# Compute J_plus[i]. Inputs: "parameters_values, epsilon". Output = "J_plus[i]".
# "_" is used because the function you have to outputs two parameters but we only care about the first one
### START CODE HERE ### (approx. 3 lines)
thetaplus = np.copy(parameters_values) # Step 1
thetaplus[i][0] += epsilon # Step 2
J_plus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaplus)) # Step 3
### END CODE HERE ###
# Compute J_minus[i]. Inputs: "parameters_values, epsilon". Output = "J_minus[i]".
### START CODE HERE ### (approx. 3 lines)
thetaminus = np.copy(parameters_values) # Step 1
thetaminus[i][0] -= epsilon # Step 2
J_minus[i], _ = forward_propagation_n(X, Y, vector_to_dictionary(thetaminus)) # Step 3
### END CODE HERE ###
# Compute gradapprox[i]
### START CODE HERE ### (approx. 1 line)
gradapprox[i] = (J_plus[i] - J_minus[i]) / (2*epsilon)
### END CODE HERE ###
# Compare gradapprox to backward propagation gradients by computing difference.
### START CODE HERE ### (approx. 1 line)
numerator = np.linalg.norm(grad - gradapprox) # Step 1'
denominator = np.linalg.norm(grad) + np.linalg.norm(gradapprox) # Step 2'
difference = numerator/denominator # Step 3'
### END CODE HERE ###
if difference > 2e-7:
print ("\033[93m" + "There is a mistake in the backward propagation! difference = " + str(difference) + "\033[0m")
else:
print ("\033[92m" + "Your backward propagation works perfectly fine! difference = " + str(difference) + "\033[0m")
return difference
X, Y, parameters = gradient_check_n_test_case()
# print(parameters)
cost, cache = forward_propagation_n(X, Y, parameters)
gradients = backward_propagation_n(X, Y, cache)
difference = gradient_check_n(parameters, gradients, X, Y)
```
**Expected output**:
<table>
<tr>
<td> ** There is a mistake in the backward propagation!** </td>
<td> difference = 0.285093156781 </td>
</tr>
</table>
It seems that there were errors in the `backward_propagation_n` code we gave you! Good that you've implemented the gradient check. Go back to `backward_propagation` and try to find/correct the errors *(Hint: check dW2 and db1)*. Rerun the gradient check when you think you've fixed it. Remember you'll need to re-execute the cell defining `backward_propagation_n()` if you modify the code.
Can you get gradient check to declare your derivative computation correct? Even though this part of the assignment isn't graded, we strongly urge you to try to find the bug and re-run gradient check until you're convinced backprop is now correctly implemented.
**Note**
- Gradient Checking is slow! Approximating the gradient with $\frac{\partial J}{\partial \theta} \approx \frac{J(\theta + \varepsilon) - J(\theta - \varepsilon)}{2 \varepsilon}$ is computationally costly. For this reason, we don't run gradient checking at every iteration during training. Just a few times to check if the gradient is correct.
- Gradient Checking, at least as we've presented it, doesn't work with dropout. You would usually run the gradient check algorithm without dropout to make sure your backprop is correct, then add dropout.
Congrats, you can be confident that your deep learning model for fraud detection is working correctly! You can even use this to convince your CEO. :)
<font color='blue'>
**What you should remember from this notebook**:
- Gradient checking verifies closeness between the gradients from backpropagation and the numerical approximation of the gradient (computed using forward propagation).
- Gradient checking is slow, so we don't run it in every iteration of training. You would usually run it only to make sure your code is correct, then turn it off and use backprop for the actual learning process.
| github_jupyter |
# Basic Pipeline
Our basic pipeline consists of using **Anchor Link** statistics to link an entity mention in the dataset ACY to a Wikipedia page ID in the Kensho-derived Knowledge Graph.
```
# Import necessary packages
# Ensure installation of nltk package in conda environment
import os
import time
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# Add directory above current directory to path so we can import our pre-built package
import sys; sys.path.insert(0, '../..')
from entity_disambiguation.preprocessing import process_input, normalize_text
from nltk.corpus import stopwords
from tqdm import tqdm
try:
stop = stopwords.words('english')
except LookupError:
# Download stopwords if it's your first time
import nltk
nltk.download('stopwords')
```
## 1. Process Aida-Conll-Yago (ACY) dataset into train/test split
We import ACY, provide the ability to split it into train and test (though don't use this yet) and normalize full_mention to lower cases with spaces.
```
# relative path to ACY dataset, should be the same for all users within `entity-disambiguation` repository
acy_path = '../../data/aida-conll-yago-dataset/'
# process_input() takes path location where you've stored AIDA-YAGO2-DATASET.tsv file
# Transforms tsv file into train, test split
# x are words, y are indices
train_x, train_y, test_x, test_y = process_input(acy_path, match="full_mention", train = 1.)
# Display shape of output train data
display((train_x.shape, train_y.shape))
display((test_x.shape, test_y.shape))
# Display x preview
train_x.head(3)
# Display y preview
train_y[0:3]
# Save train x/y into numpy arrays
x_np = train_x.full_mention.values
x_np = np.array([x.lower() if x is not None else x for x in x_np])
y_np = np.array(train_y, dtype = 'int64')
# Preview x_np
x_np[:5], y_np[:5]
```
## Build Kensho Target Dataset
Load `KWNLP` dataset to create our baseline model, i.e. calculate the anchor linkage statistics for Wikipedia pages. Then we apply that to the ACY dataset in the next section.
These baselines so far only consider *exact* matches between a full mention in ACY and an anchor text in KWNLP.
```
# Provide directory path for KWNLP data
# Should be same for all users of `entity-disambiguation` repository
kwnlp_path = '../../data/kwnlp'
# Load article data
article_df = pd.read_csv(os.path.join(kwnlp_path, 'kwnlp-enwiki-20200920-article.csv'))
# Load anchor target counts data
anchor_df = pd.read_csv(os.path.join(kwnlp_path, 'kwnlp-enwiki-20200920-anchor-target-counts.csv'))
# Display article preview
article_df.head()
# Display anchor preview
anchor_df.head()
# Copy anchor_df to new dataframe
at_count_df = anchor_df.copy()
# Normalize anchor_text (lower-case, strip whitespace)
at_count_df["normalized_anchor_text"] = at_count_df["anchor_text"].apply(normalize_text)
# Return all anchor_texts that are non-zero, non-null
at_count_df = at_count_df.loc[at_count_df['normalized_anchor_text'].str.len() > 0, :]
print(len(at_count_df))
at_count_df.head(3)
```
Inner join anchor-target data (mention to linked entity) with Wikipedia page article data. This lets us collate stats like page views with the target (entity) of mentions. Page views serves as another baseline model by selecting the page/entity that is most viewed for that anchor text.
```
# Merge at_count and article stats dataframes
at_count_df = pd.merge(
at_count_df,
article_df,
how="inner",
left_on="target_page_id",
right_on="page_id")
# Rename columns for clarity
at_count_df = at_count_df.rename(columns={
'title': 'target_page_title',
'item_id': 'target_item_id',
'views': 'target_page_views',
'count': 'anchor_target_count',
'page_title': 'target_page_title'})
# Specify column ordering
at_count_df = at_count_df[[
"normalized_anchor_text",
"target_page_id",
"target_item_id",
"target_page_title",
"target_page_views",
"anchor_target_count"]]
# Display preview
at_count_df.head(3)
```
Drop NaNs. These have been encoded as `string` through the text normalisation previously.
```
# Drop NaNs
len_orig = len(at_count_df)
at_count_df = at_count_df.loc[at_count_df['normalized_anchor_text'] != 'nan']
print('Dropped rows:', len_orig - len(at_count_df))
```
This leaves us with our final target baseline. We will now select a page_id for each normalized_anchor_text based on target link count or page views and then try to join full_mention with normalized_anchor_text.
# Develop Two Baseline Models
### i. Anchor Link Count
`pandas` `merge` and `join` can't be used with indices with duplicate values, as it automatically sorts and that can't be undone confidently with duplicates.
```
%%time
# Sort all anchor links by anchor text and then target count
max_anchor_links = at_count_df.sort_values(['normalized_anchor_text', 'anchor_target_count'], ascending = False)
# Keep just most common value (top value after sort)
max_anchor_links.drop_duplicates('normalized_anchor_text', keep = 'first', inplace = True)
# Update index after drops
max_anchor_links.set_index('normalized_anchor_text', inplace = True)
# Show top rows
display(max_anchor_links.head(3))
print('Removed {} mentions with targets with lower anchor counts'.format(len(at_count_df)-len(max_anchor_links)))
assert len(max_anchor_links) == len(set(at_count_df.normalized_anchor_text.values))
# Preview 10 random rows
random_int = np.random.randint(len(max_anchor_links)) # Useful example: 4395177
display(max_anchor_links[random_int:random_int+10][['target_page_id', 'target_page_title', 'anchor_target_count']])
# Print previous random_int in case you want to explore further
print(random_int)
```
The above is our linkage of text to Wikipedia page based on link count.
To create our predictions, we initialize a new DataFrame with our inputs x_np, then for each input (full_mention), search max_anchor_links for matching anchor text string and add the associated page_id and target_page_title.
```
# Create text-to-page-to-title dataframe (as predictions)
preds_anchor = pd.DataFrame({'mention': x_np,
'entity_page_id': None,
'target_page_title': None
})
# tqdm is a progress bar package
# For all anchor texts, update with predictions
for i in tqdm(range(len(preds_anchor))):
try:
preds_anchor.iloc[i, 1:3] = max_anchor_links.loc[preds_anchor.iloc[i, 0], ['target_page_id', 'target_page_title']].values
except KeyError:
# Leave at default None values
continue
# Display entity disambiguation predictions using anchor link statistics
preds_anchor
```
### ii. Page Views
```
# Display dataframe preview
at_count_df
```
Starting with the same dataframe as before, we now sort by page views and take the top result by page views.
```
%%time
# Sort dataframe by anchor text and page views, remove duplicates except top (most popular) and update index
max_page_views = at_count_df.sort_values(['normalized_anchor_text', 'target_page_views'], ascending = False)
max_page_views.drop_duplicates('normalized_anchor_text', keep = 'first', inplace = True)
max_page_views.set_index('normalized_anchor_text', inplace = True)
# Show top rows
display(max_page_views.head(3))
assert len(max_page_views) == len(set(at_count_df.normalized_anchor_text.values))
# Create prediction dataframe with most popular views
preds_page = pd.DataFrame({'mention': x_np,
'entity_page_id': None,
'target_page_title': None
})
# tqdm is progress bar
# For every anchor text, append prediction using page views
for i in tqdm(range(len(preds_page))):
try:
preds_page.iloc[i, 1:3] = max_page_views.loc[preds_page.iloc[i, 0], ['target_page_id', 'target_page_title']].values
except KeyError:
# Leave at default None values
continue
# Display predictions preview
preds_page
```
## Scoring
```
# Accuracy matching full mention with anchor text
print('-- Accuracy on entire ACY dataset using KWNLP --')
print('Anchor Linking: {}%'.format(np.round(100*np.mean(preds_anchor.entity_page_id == y_np), 2)))
print('Page Views: {}%'.format(np.round(100*np.mean(preds_page.entity_page_id == y_np), 2)))
# Accuracy matching single words
print('-- Accuracy on entire ACY dataset using KWNLP --')
print('Anchor Linking: {}%'.format(np.round(100*np.mean(preds_anchor.entity_page_id == y_np), 2)))
print('Page Views: {}%'.format(np.round(100*np.mean(preds_page.entity_page_id == y_np), 2)))
```
If we match just tokens, we see:
Anchor Linking: 33.14%
Page Views: 29.31%
If we match full_mention, we see:
Anchor Linking: 71.92%
Page Views: 61.33%
| github_jupyter |
Copyright 2018 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
https://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
**This tutorial is for educational purposes purposes only and is not intended for use in clinical diagnosis or clinical decision-making or for any other clinical use.**
# Training/Inference on Breast Density Classification Model on Cloud AI Platform
The goal of this tutorial is to train, deploy and run inference on a breast density classification model. Breast density is thought to be a factor for an increase in the risk for breast cancer. This will emphasize using the [Cloud Healthcare API](https://cloud.google.com/healthcare/) in order to store, retreive and transcode medical images (in DICOM format) in a managed and scalable way. This tutorial will focus on using [Cloud AI Platform](https://cloud.google.com/ai-platform/) to scalably train and serve the model.
**Note: This is the Cloud AI Platform version of the AutoML Codelab found [here](./breast_density_auto_ml.ipynb).**
## Requirements
- A Google Cloud project.
- Project has [Cloud Healthcare API](https://cloud.google.com/healthcare/docs/quickstart) enabled.
- Project has [Cloud Machine Learning API ](https://cloud.google.com/ml-engine/docs/tensorflow/getting-started-training-prediction) enabled.
- Project has [Cloud Dataflow API ](https://cloud.google.com/dataflow/docs/quickstarts/quickstart-python) enabled.
- Project has [Cloud Build API](https://cloud.google.com/cloud-build/docs/quickstart-docker) enabled.
- Project has [Kubernetes engine API](https://console.developers.google.com/apis/api/container.googleapis.com/overview?project=) enabled.
- Project has [Cloud Resource Manager API](https://console.cloud.google.com/cloud-resource-manager) enabled.
## Notebook dependencies
We will need to install the hcls_imaging_ml_toolkit package found [here](./toolkit). This toolkit helps make working with DICOM objects and the Cloud Healthcare API easier.
In addition, we will install [dicomweb-client](https://dicomweb-client.readthedocs.io/en/latest/) to help us interact with the DIOCOMWeb API and [pydicom](https://pydicom.github.io/pydicom/dev/index.html) which is used to help up construct DICOM objects.
```
%%bash
pip3 install git+https://github.com/GoogleCloudPlatform/healthcare.git#subdirectory=imaging/ml/toolkit
pip3 install dicomweb-client
pip3 install pydicom
```
## Input Dataset
The dataset that will be used for training is the [TCIA CBIS-DDSM](https://wiki.cancerimagingarchive.net/display/Public/CBIS-DDSM) dataset. This dataset contains ~2500 mammography images in DICOM format. Each image is given a [BI-RADS breast density ](https://breast-cancer.ca/densitbi-rads/) score from 1 to 4. In this tutorial, we will build a binary classifier that distinguishes between breast density "2" (*scattered density*) and "3" (*heterogeneously dense*). These are the two most common and variably assigned scores. In the literature, this is said to be [particularly difficult for radiologists to consistently distinguish](https://aapm.onlinelibrary.wiley.com/doi/pdf/10.1002/mp.12683).
```
project_id = "MY_PROJECT" # @param
location = "us-central1"
dataset_id = "MY_DATASET" # @param
dicom_store_id = "MY_DICOM_STORE" # @param
# Input data used by Cloud ML must be in a bucket with the following format.
cloud_bucket_name = "gs://" + project_id + "-vcm"
%%bash -s {project_id} {location} {cloud_bucket_name}
# Create bucket.
gsutil -q mb -c regional -l $2 $3
# Allow Cloud Healthcare API to write to bucket.
PROJECT_NUMBER=`gcloud projects describe $1 | grep projectNumber | sed 's/[^0-9]//g'`
SERVICE_ACCOUNT="service-${PROJECT_NUMBER}@gcp-sa-healthcare.iam.gserviceaccount.com"
COMPUTE_ENGINE_SERVICE_ACCOUNT="${PROJECT_NUMBER}-compute@developer.gserviceaccount.com"
gsutil -q iam ch serviceAccount:${SERVICE_ACCOUNT}:objectAdmin $3
gsutil -q iam ch serviceAccount:${COMPUTE_ENGINE_SERVICE_ACCOUNT}:objectAdmin $3
gcloud projects add-iam-policy-binding $1 --member=serviceAccount:${SERVICE_ACCOUNT} --role=roles/pubsub.publisher
gcloud projects add-iam-policy-binding $1 --member=serviceAccount:${COMPUTE_ENGINE_SERVICE_ACCOUNT} --role roles/pubsub.admin
# Allow compute service account to create datasets and dicomStores.
gcloud projects add-iam-policy-binding $1 --member=serviceAccount:${COMPUTE_ENGINE_SERVICE_ACCOUNT} --role roles/healthcare.dicomStoreAdmin
gcloud projects add-iam-policy-binding $1 --member=serviceAccount:${COMPUTE_ENGINE_SERVICE_ACCOUNT} --role roles/healthcare.datasetAdmin
import json
import os
import google.auth
from google.auth.transport.requests import AuthorizedSession
from hcls_imaging_ml_toolkit import dicom_path
credentials, project = google.auth.default()
authed_session = AuthorizedSession(credentials)
# Path to Cloud Healthcare API.
HEALTHCARE_API_URL = 'https://healthcare.googleapis.com/v1'
# Create Cloud Healthcare API dataset.
path = os.path.join(HEALTHCARE_API_URL, 'projects', project_id, 'locations', location, 'datasets?dataset_id=' + dataset_id)
headers = {'Content-Type': 'application/json'}
resp = authed_session.post(path, headers=headers)
assert resp.status_code == 200, 'error creating Dataset, code: {0}, response: {1}'.format(resp.status_code, resp.text)
print('Full response:\n{0}'.format(resp.text))
# Create Cloud Healthcare API DICOM store.
path = os.path.join(HEALTHCARE_API_URL, 'projects', project_id, 'locations', location, 'datasets', dataset_id, 'dicomStores?dicom_store_id=' + dicom_store_id)
resp = authed_session.post(path, headers=headers)
assert resp.status_code == 200, 'error creating DICOM store, code: {0}, response: {1}'.format(resp.status_code, resp.text)
print('Full response:\n{0}'.format(resp.text))
dicom_store_path = dicom_path.Path(project_id, location, dataset_id, dicom_store_id)
```
Next, we are going to transfer the DICOM instances to the Cloud Healthcare API.
Note: We are transfering >100GB of data so this will take some time to complete
```
# Store DICOM instances in Cloud Healthcare API.
path = 'https://healthcare.googleapis.com/v1/{}:import'.format(dicom_store_path)
headers = {'Content-Type': 'application/json'}
body = {
'gcsSource': {
'uri': 'gs://gcs-public-data--healthcare-tcia-cbis-ddsm/dicom/**'
}
}
resp = authed_session.post(path, headers=headers, json=body)
assert resp.status_code == 200, 'error creating Dataset, code: {0}, response: {1}'.format(resp.status_code, resp.text)
print('Full response:\n{0}'.format(resp.text))
response = json.loads(resp.text)
operation_name = response['name']
import time
def wait_for_operation_completion(path, timeout, sleep_time=30):
success = False
while time.time() < timeout:
print('Waiting for operation completion...')
resp = authed_session.get(path)
assert resp.status_code == 200, 'error polling for Operation results, code: {0}, response: {1}'.format(resp.status_code, resp.text)
response = json.loads(resp.text)
if 'done' in response:
if response['done'] == True and 'error' not in response:
success = True;
break
time.sleep(sleep_time)
print('Full response:\n{0}'.format(resp.text))
assert success, "operation did not complete successfully in time limit"
print('Success!')
return response
path = os.path.join(HEALTHCARE_API_URL, operation_name)
timeout = time.time() + 40*60 # Wait up to 40 minutes.
_ = wait_for_operation_completion(path, timeout)
```
### Explore the Cloud Healthcare DICOM dataset (optional)
This is an optional section to explore the Cloud Healthcare DICOM dataset. In the following code, we simply just list the studies that we have loaded into the Cloud Healthcare API. You can modify the *num_of_studies_to_print* parameter to print as many studies as desired.
```
num_of_studies_to_print = 2 # @param
path = os.path.join(HEALTHCARE_API_URL, dicom_store_path.dicomweb_path_str, 'studies')
resp = authed_session.get(path)
assert resp.status_code == 200, 'error querying Dataset, code: {0}, response: {1}'.format(resp.status_code, resp.text)
response = json.loads(resp.text)
print(json.dumps(response[:num_of_studies_to_print], indent=2))
```
## Convert DICOM to JPEG
The ML model that we will build requires that the dataset be in JPEG. We will leverage the Cloud Healthcare API to transcode DICOM to JPEG.
First we will create a [Google Cloud Storage](https://cloud.google.com/storage/) bucket to hold the output JPEG files. Next, we will use the ExportDicomData API to transform the DICOMs to JPEGs.
```
jpeg_bucket = cloud_bucket_name + "/images/"
```
Next we will convert the DICOMs to JPEGs using the [ExportDicomData](https://cloud.google.com/sdk/gcloud/reference/beta/healthcare/dicom-stores/export/gcs).
```
%%bash -s {jpeg_bucket} {project_id} {location} {dataset_id} {dicom_store_id}
gcloud beta healthcare --project $2 dicom-stores export gcs $5 --location=$3 --dataset=$4 --mime-type="image/jpeg; transfer-syntax=1.2.840.10008.1.2.4.50" --gcs-uri-prefix=$1
```
We will use the Operation name returned from the previous command to poll the status of ExportDicomData. We will poll for operation completeness, which should take a few minutes. When the operation is complete, the operation's *done* field will be set to true.
Meanwhile, you should be able to observe the JPEG images being added to your Google Cloud Storage bucket.
## Training
We will use [Transfer Learning](https://en.wikipedia.org/wiki/Transfer_learning) to retrain a generically trained trained model to perform breast density classification. Specifically, we will use an [Inception V3](https://github.com/tensorflow/models/tree/master/research/inception) checkpoint as the starting point.
The neural network we will use can roughly be split into two parts: "feature extraction" and "classification". In transfer learning, we take advantage of a pre-trained (checkpoint) model to do the "feature extraction", and add a few layers to perform the "classification" relevant to the specific problem. In this case, we are adding aa [dense](https://www.tensorflow.org/api_docs/python/tf/layers/dense) layer with two neurons to do the classification and a [softmax](https://www.tensorflow.org/api_docs/python/tf/nn/softmax) layer to normalize the classification score. The mammography images will be classified as either "2" (scattered density) or "3" (heterogeneously dense). See below for diagram of the training process:

The "feature extraction" and the "classification" part will be done in the following steps, respectively.
### Preprocess Raw Images using Cloud Dataflow
In this step, we will resize images to 300x300 (required for Inception V3) and will run each image through the checkpoint Inception V3 model to calculate the *bottleneck values*. This is the feature vector for the output of the feature extraction part of the model (the part that is already pre-trained). Since this process is resource intensive, we will utilize [Cloud Dataflow](https://cloud.google.com/dataflow/) in order to do this scalably. We extract the features and calculate the bottleneck values here for performance reasons - so that we don't have to recalculate them during training.
The output of this process will be a collection of [TFRecords](https://www.tensorflow.org/guide/datasets) storing the bottleneck value for each image in the input dataset. This TFRecord format is commonly used to store Tensors in binary format for storage.
Finally, in this step, we will also split the input dataset into *training*, *validation* or *testing*. The percentage of each can be modified using the parameters below.
```
# GCS Bucket to store output TFRecords.
bottleneck_bucket = cloud_bucket_name + "/bottleneck" # @param
# Percentage of dataset to allocate for validation and testing.
validation_percentage = 10 # @param
testing_percentage = 10 # @param
# Number of Dataflow workers. This can be increased to improve throughput.
dataflow_num_workers = 5 # @param
# Staging bucket for training.
staging_bucket = cloud_bucket_name # @param
```
The following command will kick off a Cloud Dataflow pipeline that runs preprocessing. The script that has the relevant code is [preprocess.py](./scripts/trainer/preprocess.py). ***You can check out how the pipeline is progressing [here](https://console.cloud.google.com/dataflow)***.
When the operation is done, we will begin training the classification layers.
```
%%bash -s {project_id} {jpeg_bucket} {bottleneck_bucket} {validation_percentage} {testing_percentage} {dataflow_num_workers} {staging_bucket}
# Install Python library dependencies.
pip install virtualenv
python3 -m virtualenv env
source env/bin/activate
pip install tensorflow==1.15.0 google-apitools apache_beam[gcp]==2.18.0
# Start job in Cloud Dataflow and wait for completion.
python3 -m scripts.preprocess.preprocess \
--project $1 \
--input_path $2 \
--output_path "$3/record" \
--num_workers $6 \
--temp_location "$7/temp" \
--staging_location "$7/staging" \
--validation_percentage $4 \
--testing_percentage $5
```
### Train the Classification Layers of Model using Cloud AI Platform
In this step, we will train the classification layers of the model. This consists of just a [dense](https://www.tensorflow.org/api_docs/python/tf/layers/dense) and [softmax](https://www.tensorflow.org/api_docs/python/tf/nn/softmax) layer. We will use the bottleneck values calculated at the previous step as the input to these layers. We will use Cloud AI Platform to train the model. The output of stage will be a trained model exported to GCS, which can be used for inference.
There are various training parameters below that can be tuned.
```
training_steps = 1000 # @param
learning_rate = 0.01 # @param
# Location of exported model.
exported_model_bucket = cloud_bucket_name + "/models" # @param
# Inference requires the exported model to be versioned (by default we choose version 1).
exported_model_versioned_uri = exported_model_bucket + "/1"
```
We'll invoke Cloud AI Platform with the above parameters. We use a GPU for training to speed up operations. The script that does the training is [model.py](./scripts/trainer/model.py)
```
%%bash -s {location} {bottleneck_bucket} {staging_bucket} {training_steps} {learning_rate} {exported_model_versioned_uri}
# Start training on CAIP.
gcloud ai-platform jobs submit training breast_density \
--python-version 3.7 \
--runtime-version 1.15 \
--scale-tier BASIC_GPU \
--module-name "scripts.trainer.model" \
--package-path scripts \
--staging-bucket $3 \
--region $1 \
-- \
--bottleneck_dir "$2/record" \
--training_steps $4 \
--learning_rate $5 \
--export_model_path $6
```
You can monitor the status of the training job by running the following command. The job can take a few minutes to start-up.
```
!gcloud ai-platform jobs describe breast_density
```
When the job has started, you can observe the logs for the training job by executing the below command (it will poll for new logs every 30 seconds).
As training progresses, the logs will output the accuracy on the training set, validation set, as well as the [cross entropy](http://ml-cheatsheet.readthedocs.io/en/latest/loss_functions.html). You'll generally see that the accuracy goes up, while the cross entropy goes down as the number of training iterations increases.
Finally, when the training is complete, the accuracy of the model on the held-out test set will be output to console. The job can take a few minutes to shut-down.
```
!gcloud ai-platform jobs stream-logs breast_density --polling-interval=30
```
### Deployment and Getting Predictions
Cloud AI Platform (CAIP) can also be used to serve the model for inference. The inference model is composed of the pre-trained Inception V3 checkpoint, along with the classification layers we trained above for breast density. First we set the inference model name/version and select a mammography image to test out.
```
model_name = "breast_density" # @param
deployment_version = "deployment" # @param
# The full name of the model.
full_model_name = "projects/" + project_id + "/models/" + model_name + "/versions/" + deployment_version
!gcloud ai-platform models create $model_name --regions $location
!gcloud ai-platform versions create $deployment_version --model $model_name --origin $exported_model_versioned_uri --runtime-version 1.15 --python-version 3.7
# DICOM Study/Series UID of input mammography image that we'll test.
input_mammo_study_uid = "1.3.6.1.4.1.9590.100.1.2.85935434310203356712688695661986996009" # @param
input_mammo_series_uid = "1.3.6.1.4.1.9590.100.1.2.374115997511889073021386151921807063992" # @param
input_mammo_instance_uid = "1.3.6.1.4.1.9590.100.1.2.289923739312470966435676008311959891294" # @param
```
Let's run inference for the image and observe the results. We should see the returned label as well as the score.
```
from base64 import b64encode, b64decode
import io
from PIL import Image
import tensorflow as tf
_INCEPTION_V3_SIZE = 299
input_file_path = os.path.join(jpeg_bucket, input_mammo_study_uid, input_mammo_series_uid, input_mammo_instance_uid + ".jpg")
with tf.io.gfile.GFile(input_file_path, 'rb') as example_img:
# Resize the image to InceptionV3 input size.
im = Image.open(example_img).resize((_INCEPTION_V3_SIZE,_INCEPTION_V3_SIZE))
imgByteArr = io.BytesIO()
im.save(imgByteArr, format='JPEG')
b64str = b64encode(imgByteArr.getvalue()).decode('utf-8')
with open('input_image.json', 'a') as outfile:
json.dump({'inputs': [{'b64': b64str}]}, outfile)
outfile.write('\n')
predictions = !gcloud ai-platform predict --model $model_name --version $deployment_version --json-instances='input_image.json'
print(predictions)
```
### Getting Explanations
**There are limits and caveats when using the Explainable AI feature on CAIP. Read about them [here](https://cloud.google.com/ai-platform/prediction/docs/ai-explanations/limitations).**
The [Explainable AI](https://cloud.google.com/explainable-ai) feature of CAIP can be used to provide visibility as to why the model returned a prediction for a given input. In this codelab, we are going to use this feature to figure out which pixels in the example mammography image contributed the most to the prediction. This can be useful for debugging model performance and improving the confidence in the model. Read [here](https://cloud.google.com/ai-platform/prediction/docs/ai-explanations/overview) for more details. See below for sample output.

To get started, we will first deploy the model to CAIP that has explainable AI enabled.
```
explainable_version = "explainable_ai" # @param
```
We'll create an Explainable AI configuration file. This will allow us to specify the input and the output tensor to correlate. See [here](https://cloud.google.com/ai-platform/prediction/docs/ai-explanations/preparing-metadata) for more details. Below we'll correlate the input image tensor with the output of the softmax layer. The Explanation AI configuration file is required to be stored in the model directory.
```
import json
import os
import scripts.constants as constants
explainable_metadata = {
"outputs": {
"probability": {
"output_tensor_name": constants.OUTPUT_SOFTMAX_TENSOR_NAME + ":0",
}
},
"inputs": {
"img_bytes": {
"input_tensor_name": constants.INPUT_PIXELS_TENSOR_NAME + ":0",
"input_tensor_type": "numeric",
"modality": "image",
}
},
"framework": "tensorflow"
}
# The configuration file in the CAIP model directory.
with tf.io.gfile.GFile(os.path.join(exported_model_versioned_uri, 'explanation_metadata.json'), 'w') as output_file:
json.dump(explainable_metadata, output_file)
```
Finally, let's deploy the model.
```
!gcloud beta ai-platform versions create $explainable_version \
--model $model_name\
--origin $exported_model_versioned_uri \
--runtime-version 1.15 \
--python-version 3.7 \
--machine-type n1-standard-4 \
--explanation-method integrated-gradients \
--num-integral-steps 25
```
Next, we'll ask for the annotated image that includes the Explainable AI overlay.
```
explanations = !gcloud beta ai-platform explain --model $model_name --version $explainable_version --json-instances='input_image.json'
response = json.loads(explanations.s)
```
Next, lets print the annotated image (with overlay). We can see green highlights for the pixels that give the biggest signal for the highest scoring class.
```
import base64
import io
from PIL import Image
assert len(response['explanations']) == 1
LABELS = ['2', '3']
prediction = response['explanations'][0]
predicted_label = LABELS[prediction['attributions_by_label'][0]['label_index']]
confidence = prediction['attributions_by_label'][0]['example_score']
print('Predicted class: ', predicted_label)
print('Confidence: ', confidence)
b64str = prediction['attributions_by_label'][0]['attributions']['img_bytes']['b64_jpeg']
display(Image.open(io.BytesIO(base64.b64decode(b64str))))
```
## Integration in the clinical workflow
To allow medical imaging ML models to be easily integrated into clinical workflows, an *inference module* can be used. A standalone modality, a PACS system or a DICOM router can push DICOM instances into Cloud Healthcare [DICOM stores](https://cloud.google.com/healthcare/docs/introduction), allowing ML models to be triggered for inference. This inference results can then be structured into various DICOM formats (e.g. DICOM [structured reports](http://dicom.nema.org/MEDICAL/Dicom/2014b/output/chtml/part20/sect_A.3.html)) and stored in the Cloud Healthcare API, which can then be retrieved by the customer.
The inference module is built as a [Docker](https://www.docker.com/) container and deployed using [Kubernetes](https://kubernetes.io/), allowing you to easily scale your deployment. The dataflow for inference can look as follows (see corresponding diagram below):
1. Client application uses [STOW-RS](ftp://dicom.nema.org/medical/Dicom/2013/output/chtml/part18/sect_6.6.html) to push a new DICOM instance to the Cloud Healthcare DICOMWeb API.
2. The insertion of the DICOM instance triggers a [Cloud Pubsub](https://cloud.google.com/pubsub/) message to be published. The *inference module* will pull incoming Pubsub messages and will recieve a message for the previously inserted DICOM instance.
3. The *inference module* will retrieve the instance in JPEG format from the Cloud Healthcare API using [WADO-RS](ftp://dicom.nema.org/medical/Dicom/2013/output/chtml/part18/sect_6.5.html).
4. The *inference module* will send the JPEG bytes to the model hosted on Cloud AI Platform.
5. Cloud AI Platform will return the prediction back to the *inference module*.
6. The *inference module* will package the prediction into a DICOM instance. This can potentially be a DICOM structured report, [presentation state](ftp://dicom.nema.org/MEDICAL/dicom/2014b/output/chtml/part03/sect_A.33.html), or even burnt text on the image. In this codelab, we will focus on just DICOM structured reports, specifically [Comprehensive Structured Reports](http://dicom.nema.org/dicom/2013/output/chtml/part20/sect_A.3.html). The structured report is then stored back in the Cloud Healthcare API using STOW-RS.
7. The client application can query for (or retrieve) the structured report by using [QIDO-RS](http://dicom.nema.org/dicom/2013/output/chtml/part18/sect_6.7.html) or WADO-RS. Pubsub can also be used by the client application to poll for the newly created DICOM structured report instance.

To begin, we will create a new DICOM store that will store our inference source (DICOM mammography instance) and results (DICOM structured report). In order to enable Pubsub notifications to be triggered on inserted instances, we will give the DICOM store a Pubsub channel to publish on.
```
# Pubsub config.
pubsub_topic_id = "MY_PUBSUB_TOPIC_ID" # @param
pubsub_subscription_id = "MY_PUBSUB_SUBSRIPTION_ID" # @param
# DICOM Store for store DICOM used for inference.
inference_dicom_store_id = "MY_INFERENCE_DICOM_STORE" # @param
pubsub_subscription_name = "projects/" + project_id + "/subscriptions/" + pubsub_subscription_id
inference_dicom_store_path = dicom_path.FromPath(dicom_store_path, store_id=inference_dicom_store_id)
%%bash -s {pubsub_topic_id} {pubsub_subscription_id} {project_id} {location} {dataset_id} {inference_dicom_store_id}
# Create Pubsub channel.
gcloud beta pubsub topics create $1
gcloud beta pubsub subscriptions create $2 --topic $1
# Create a Cloud Healthcare DICOM store that published on given Pubsub topic.
TOKEN=`gcloud beta auth application-default print-access-token`
NOTIFICATION_CONFIG="{notification_config: {pubsub_topic: \"projects/$3/topics/$1\"}}"
curl -s -X POST -H "Content-Type: application/json" -H "Authorization: Bearer ${TOKEN}" -d "${NOTIFICATION_CONFIG}" https://healthcare.googleapis.com/v1/projects/$3/locations/$4/datasets/$5/dicomStores?dicom_store_id=$6
# Enable Cloud Healthcare API to publish on given Pubsub topic.
PROJECT_NUMBER=`gcloud projects describe $3 | grep projectNumber | sed 's/[^0-9]//g'`
SERVICE_ACCOUNT="service-${PROJECT_NUMBER}@gcp-sa-healthcare.iam.gserviceaccount.com"
gcloud beta pubsub topics add-iam-policy-binding $1 --member="serviceAccount:${SERVICE_ACCOUNT}" --role="roles/pubsub.publisher"
```
Next, we will building the *inference module* using [Cloud Build API](https://cloud.google.com/cloud-build/docs/api/reference/rest/). This will create a Docker container that will be stored in [Google Container Registry](https://cloud.google.com/container-registry/). The inference module code is found in *[inference.py](./scripts/inference/inference.py)*. The build script used to build the Docker container for this module is *[cloudbuild.yaml](./scripts/inference/cloudbuild.yaml)*. Progress of build may be found on [cloud build dashboard](https://console.cloud.google.com/cloud-build/builds?project=).
```
%%bash -s {project_id}
PROJECT_ID=$1
gcloud builds submit --config scripts/inference/cloudbuild.yaml --timeout 1h scripts/inference
```
Next, we will deploy the *inference module* to Kubernetes.
Then we create a Kubernetes Cluster and a Deployment for the *inference module*.
```
%%bash -s {project_id} {location} {pubsub_subscription_name} {full_model_name} {inference_dicom_store_path}
gcloud container clusters create inference-module --region=$2 --scopes https://www.googleapis.com/auth/cloud-platform --num-nodes=1
PROJECT_ID=$1
SUBSCRIPTION_PATH=$3
MODEL_PATH=$4
INFERENCE_DICOM_STORE_PATH=$5
cat <<EOF | kubectl create -f -
apiVersion: extensions/v1beta1
kind: Deployment
metadata:
name: inference-module
namespace: default
spec:
replicas: 1
template:
metadata:
labels:
app: inference-module
spec:
containers:
- name: inference-module
image: gcr.io/${PROJECT_ID}/inference-module:latest
command:
- "/opt/inference_module/bin/inference_module"
- "--subscription_path=${SUBSCRIPTION_PATH}"
- "--model_path=${MODEL_PATH}"
- "--dicom_store_path=${INFERENCE_DICOM_STORE_PATH}"
- "--prediction_service=CAIP"
EOF
```
Next, we will store a mammography DICOM instance from the TCIA dataset to the DICOM store. This is the image that we will request inference for. Pushing this instance to the DICOM store will result in a Pubsub message, which will trigger the *inference module*.
```
# DICOM Study/Series UID of input mammography image that we'll push for inference.
input_mammo_study_uid = "1.3.6.1.4.1.9590.100.1.2.85935434310203356712688695661986996009"
input_mammo_series_uid = "1.3.6.1.4.1.9590.100.1.2.374115997511889073021386151921807063992"
input_mammo_instance_uid = "1.3.6.1.4.1.9590.100.1.2.289923739312470966435676008311959891294"
from google.cloud import storage
from dicomweb_client.api import DICOMwebClient
from dicomweb_client import session_utils
from pydicom
storage_client = storage.Client()
bucket = storage_client.bucket('gcs-public-data--healthcare-tcia-cbis-ddsm', user_project=project_id)
blob = bucket.blob("dicom/{}/{}/{}.dcm".format(input_mammo_study_uid,input_mammo_series_uid,input_mammo_instance_uid))
blob.download_to_filename('example.dcm')
dataset = pydicom.dcmread('example.dcm')
session = session_utils.create_session_from_gcp_credentials()
study_path = dicom_path.FromPath(inference_dicom_store_path, study_uid=input_mammo_study_uid)
dicomweb_url = os.path.join(HEALTHCARE_API_URL, study_path.dicomweb_path_str)
dcm_client = DICOMwebClient(dicomweb_url, session)
dcm_client.store_instances(datasets=[dataset])
```
You should be able to observe the *inference module*'s logs by running the following command. In the logs, you should observe that the inference module successfully recieved the the Pubsub message and ran inference on the DICOM instance. The logs should also include the inference results. It can take a few minutes to start-up the Kubernetes deployment, so you many have to run this a few times.
```
!kubectl logs -l app=inference-module
```
You can also query the Cloud Healthcare DICOMWeb API (using QIDO-RS) to see that the DICOM structured report has been inserted for the study. The structured report contents can be found under tag **"0040A730"**.
You can optionally also use WADO-RS to recieve the instance (e.g. for viewing).
```
dcm_client.search_for_instances(study_path.study_uid, fields=['all'])
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Automatic differentiation and gradient tape
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/tutorials/eager/automatic_differentiation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
In the previous tutorial we introduced `Tensor`s and operations on them. In this tutorial we will cover [automatic differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation), a key technique for optimizing machine learning models.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
```
## Gradient tapes
TensorFlow provides the [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) API for automatic differentiation - computing the gradient of a computation with respect to its input variables. Tensorflow "records" all operations executed inside the context of a `tf.GradientTape` onto a "tape". Tensorflow then uses that tape and the gradients associated with each recorded operation to compute the gradients of a "recorded" computation using [reverse mode differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation).
For example:
```
x = tf.ones((2, 2))
with tf.GradientTape() as t:
t.watch(x)
y = tf.reduce_sum(x)
z = tf.multiply(y, y)
# Derivative of z with respect to the original input tensor x
dz_dx = t.gradient(z, x)
for i in [0, 1]:
for j in [0, 1]:
assert dz_dx[i][j].numpy() == 8.0
```
You can also request gradients of the output with respect to intermediate values computed during a "recorded" `tf.GradientTape` context.
```
x = tf.ones((2, 2))
with tf.GradientTape() as t:
t.watch(x)
y = tf.reduce_sum(x)
z = tf.multiply(y, y)
# Use the tape to compute the derivative of z with respect to the
# intermediate value y.
dz_dy = t.gradient(z, y)
assert dz_dy.numpy() == 8.0
```
By default, the resources held by a GradientTape are released as soon as GradientTape.gradient() method is called. To compute multiple gradients over the same computation, create a `persistent` gradient tape. This allows multiple calls to the `gradient()` method as resources are released when the tape object is garbage collected. For example:
```
x = tf.constant(3.0)
with tf.GradientTape(persistent=True) as t:
t.watch(x)
y = x * x
z = y * y
dz_dx = t.gradient(z, x) # 108.0 (4*x^3 at x = 3)
dy_dx = t.gradient(y, x) # 6.0
del t # Drop the reference to the tape
```
### Recording control flow
Because tapes record operations as they are executed, Python control flow (using `if`s and `while`s for example) is naturally handled:
```
def f(x, y):
output = 1.0
for i in range(y):
if i > 1 and i < 5:
output = tf.multiply(output, x)
return output
def grad(x, y):
with tf.GradientTape() as t:
t.watch(x)
out = f(x, y)
return t.gradient(out, x)
x = tf.convert_to_tensor(2.0)
assert grad(x, 6).numpy() == 12.0
assert grad(x, 5).numpy() == 12.0
assert grad(x, 4).numpy() == 4.0
```
### Higher-order gradients
Operations inside of the `GradientTape` context manager are recorded for automatic differentiation. If gradients are computed in that context, then the gradient computation is recorded as well. As a result, the exact same API works for higher-order gradients as well. For example:
```
x = tf.Variable(1.0) # Create a Tensorflow variable initialized to 1.0
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
y = x * x * x
# Compute the gradient inside the 't' context manager
# which means the gradient computation is differentiable as well.
dy_dx = t2.gradient(y, x)
d2y_dx2 = t.gradient(dy_dx, x)
assert dy_dx.numpy() == 3.0
assert d2y_dx2.numpy() == 6.0
```
## Next Steps
In this tutorial we covered gradient computation in TensorFlow. With that we have enough of the primitives required to build and train neural networks.
```
```
| github_jupyter |
```
%pylab inline
import io3d
import skelet3d
from pathlib import Path
import copy
import glob
import time
import pandas as pd
# import sed
def vessel_skeleton_extraction(pth1):
datap = io3d.read(pth1, datap=True)
from skimage.filters import threshold_otsu
threshold = threshold_otsu(datap["data3d"].ravel()[::1000])
imthr = datap["data3d"] > threshold
# imshow(imthr[int(imthr.shape[0]/2),:,:])
volume_data = imthr
skelet = skelet3d.skelet3d(volume_data)
return skelet, volume_data, datap["voxelsize_mm"]
def skeleton_analysis(skelet, volume_data, voxelsize_mm):
skan = skelet3d.SkeletonAnalyser(skelet, volume_data=volume_data, voxelsize_mm=voxelsize_mm)
stats = skan.skeleton_analysis()
df = stats_as_dataframe(skan)
return df
def extract_df(df):
dfs = df[["id",
"nodeA_ZYX 0",
"nodeA_ZYX 1",
"nodeA_ZYX 2",
"nodeB_ZYX 0",
"nodeB_ZYX 1",
"nodeB_ZYX 2",
"nodeA_ZYX_mm 0",
"nodeA_ZYX_mm 1",
"nodeA_ZYX_mm 2",
"nodeB_ZYX_mm 0",
"nodeB_ZYX_mm 1",
"nodeB_ZYX_mm 2",
"radius_mm",
"connectedEdgesA 0",
"connectedEdgesA 1",
"connectedEdgesA 2",
"connectedEdgesB 0",
"connectedEdgesB 1",
"connectedEdgesB 2",
]]
dfs.to_csv(pth1 + ".csv")
return dfs
def stats_as_dataframe(self):
import pandas as pd
import exsu.dili
if self.stats is None:
msg = "Run skeleton_analyser before stats_as_dataframe()"
logger.error(msg)
raise RuntimeError(msg)
# import imtools.dili
df = pd.DataFrame()
for stats_key in self.stats:
one_edge = copy.copy(self.stats[stats_key])
k = "orderedPoints_mm_X"
if k in one_edge:
one_edge[k] = str(one_edge[k])
k = "orderedPoints_mm_Y"
if k in one_edge:
one_edge[k] = str(one_edge[k])
k = "orderedPoints_mm_Z"
if k in one_edge:
one_edge[k] = str(one_edge[k])
k = "orderedPoints_mm"
if k in one_edge:
one_edge[k] = str(one_edge[k])
# one_edge[]
one_dct = exsu.dili.flatten_dict_join_keys(one_edge, simplify_iterables=True)
# df_one = pd.DataFrame(one_dct)
df_one = pd.DataFrame([list(one_dct.values())], columns=list(one_dct.keys()))
df = df.append(df_one, ignore_index=True)
return df
```
# Read the data
```
# pth = Path(io3d.datasets.join_path("medical/processed/porcine_liver_ct_raw", get_root=True))
pths = glob.glob("C:/Users/Jirik/Downloads/porcine_liver_ct_raw/*.mhd")
# pth1 = pth / "P01_MakroCT_HEAD_5_0_H31S_0004.mhd"
pth1 = pths[-1]
# pth1 = pths[2]
pths
# pth1 = pth / "P01_MakroCT_HEAD_5_0_H31S_0004.mhd"
# pth1 = "C:/Users/Jirik/Downloads/porcine_liver_ct_raw/P01_MakroCT_HEAD_5_0_H31S_0004.mhd"
# pth1 = r"C:/Users/Jirik/Downloads/porcine_liver_ct_raw/P01_a_MikroCT-nejhrubsi_rozliseni_DICOM_liver-1st-important_Macro_pixel-size53.0585um.mhd"
# pth1 = Path("g:\Můj disk\data\medical\processed\porcine_liver_ct_raw\P01_MakroCT_HEAD_5_0_H31S_0004.mhd")
print(pth1)
print(Path(pth1).exists())
t0 = time.time()
datap = io3d.read(pth1, datap=True)
print(time.time()-t0)
import scipy.stats
data3d = datap["data3d"]
dsc = scipy.stats.describe(data3d.ravel()[::100])
# print(f"mn={np.min(datap['data3d'])}, mx={np.max(datap['data3d'])} ")
dsc
from skimage.filters import threshold_otsu, threshold_multiotsu
# classes = 5 if Path(pth1).name == 'P01_MakroCT_HEAD_5_0_H31S_0004.mhd'else 2
# thresholds = threshold_multiotsu(datap["data3d"].ravel()[::1000], classes=classes)
# threshold = thresholds[-1]
# classes
if Path(pth1).name in ('P01_MakroCT_HEAD_5_0_H31S_0004.mhd', 'P01_MakroCT_po_rozrezani_HEAD_0_6_H20S_0003.mhd'):
threshold = 0
data3d = data3d[:,:400,:] # cut the table
else:
threshold = threshold_otsu(data3d.ravel()[::1000])
imthr = data3d > threshold
imshow(imthr[int(imthr.shape[0]/2),:,:])
```
# Do the skeleton analysis
```
volume_data = imthr
t0 = time.time()
skelet = skelet3d.skelet3d(volume_data)
print(time.time()-t0)
# skan = skelet3d.SkeletonAnalyser(skelet, volume_data=volume_data, voxelsize_mm=datap["voxelsize_mm"])
t0 = time.time()
skan = skelet3d.SkeletonAnalyser(
skelet,
# volume_data=volume_data,
voxelsize_mm=datap["voxelsize_mm"]
)
print(time.time()-t0)
# skan.
skan.sklabel.dtype
# sklabel_nodes = skan.sklabel.copy()
# sklabel_nodes[sklabel_nodes > 0] = 0
nz = np.nonzero(skan.sklabel < 0)
nz
voxelsize_mm = datap["voxelsize_mm"]
dfs = pd.DataFrame({
"node_0_px":nz[0],
"node_1_px":nz[1],
"node_2_px":nz[2],
"node_0_mm":nz[0]*voxelsize_mm[0],
"node_1_mm":nz[1]*voxelsize_mm[1],
"node_2_mm":nz[2]*voxelsize_mm[2],
})
dfs.to_csv(pth1 + ".nodes.csv")
# df["Z_px"]
# io3d.write({"data3d"})
np.savez(pth1 + ".npz", sklabel=skan.sklabel, imthr=imthr)
# dir(skan)#.__connection_analysis
# skan._SkeletonAnalyser__connection_analysis(3)
```
# Time consuming steps
```
t0 = time.time()
stats = skan.skeleton_analysis()
print(time.time()-t0)
df = stats_as_dataframe(skan)
df.keys()
datap["voxelsize_mm"]
dfs = df[["id",
"nodeA_ZYX 0",
"nodeA_ZYX 1",
"nodeA_ZYX 2",
"nodeB_ZYX 0",
"nodeB_ZYX 1",
"nodeB_ZYX 2",
"nodeA_ZYX_mm 0",
"nodeA_ZYX_mm 1",
"nodeA_ZYX_mm 2",
"nodeB_ZYX_mm 0",
"nodeB_ZYX_mm 1",
"nodeB_ZYX_mm 2",
"radius_mm",
"connectedEdgesA 0",
"connectedEdgesA 1",
"connectedEdgesA 2",
"connectedEdgesB 0",
"connectedEdgesB 1",
"connectedEdgesB 2",
]]
dfs
dfs.to_csv(pth1 + ".csv")
df[["id", "lengthEstimation", "radius_mm", "tortuosity", 'phiAc', "phiBa", "phiBb", 'connectedEdgesA 0', 'connectedEdgesA 1',
'connectedEdgesB 0', 'connectedEdgesB 1',
'connectedEdgesA 2',
'connectedEdgesA 3',
'connectedEdgesB 2', 'connectedEdgesB 3']]
```
# Bulk processing
```
import glob
pths = glob.glob("C:/Users/Jirik/Downloads/porcine_liver_ct_raw/*.mhd")
# pth1 = pth / "P01_MakroCT_HEAD_5_0_H31S_0004.mhd"
# pth1
pths
# for pth1 in pths:
# vesse
outputs = vessel_skeleton_extraction(pths[0])
# df = keleton_analysis(*outputs)
# dfs = extract_df(df)
outputs
```
| github_jupyter |
## **K - Nearest Neighbors**
K Nearest Neighbour is a simple algorithm that stores all the available cases and classifies the new data or case based on a similarity measure. It is mostly used to classifies a data point based on how its neighbours are classified.
<img src = "https://miro.medium.com/max/718/1*X1KBJctko0RH6BWBsu-XjA.png">
<img src = "https://res.cloudinary.com/dyd911kmh/image/upload/f_auto,q_auto:best/v1531424125/KNN_final1_ibdm8a.png" >
## **Geting Started with KNN Classifier**
```
#Importing Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn import preprocessing
```
## **DataFraming**
Read .csv data into a Dataframe
```
data = pd.read_csv("/content/drive/My Drive/Datasets/credit_data.csv")
data.head(20)
# A very simple & Logical technique to find total number of defaulters
data.default.sum()
#Finding number of Nan or missing values in all columns
print(data.isna().sum(axis = 0))
# data.isna().sum(axis = 0) # Nan values in every column
# data.isna().sum(axis = 1) # Nan values in every row.
```
## **Features Extraction**
Extracting features and splitting data into test and train.
```
features = data[["income","age","loan"]]
target = data.default
features.head()
target.head()
features.head()
plt.scatter(data.income, data.default)
plt.scatter(data.age, data.default)
plt.scatter(data.loan, data.default)
plt.show()
data.head()
# features.shape
# target.shape
```
## **Preprocessing : Standard Scaling**
```
features = preprocessing.MinMaxScaler().fit_transform(features)
feature_train, feature_test, target_train, target_test = train_test_split(features,target)
```
## **Training the Model**
We are using KNeighborsClassifier model as imported from sklearn.neighbors library and then it's being trained on feature_train and target_train
```
model = KNeighborsClassifier(n_neighbors=32)
fitModel = model.fit(feature_train, target_train)
predictions = fitModel.predict(feature_test)
```
## **Finding Optimal K Value**
We are going to use cross-validation in order to find the optimal k value. This optimal value is not going to have as good accuracy and precision as we have seen for any neighbors close to n but it is going to be much more realistic because we use cross-validation. So we are going to use the cross-validation scores and then we are going to make a simple iteration. Basically we are going to consider k values from 1 up to 100.
```
cross_valid_scores = []
for k in range(1, 100):
knn = KNeighborsClassifier(n_neighbors = k)
scores = cross_val_score(knn,features, target, cv = 10, scoring = 'accuracy')
cross_valid_scores.append(scores.mean())
print("Optimal k with cross-validation: \t",np.argmax(cross_valid_scores))
```
## **Printing an Error Matrix and Accuracy Score**
```
print(confusion_matrix(target_test,predictions))
print(accuracy_score(target_test,predictions))
import seaborn as sns
sns.heatmap(confusion_matrix(target_test,predictions), annot=True, cmap='Blues')
```
| github_jupyter |
# RUL estimation Nasa Randomized dataset
```
import numpy as np
import pandas as pd
import scipy.io
import math
import os
import ntpath
import sys
import logging
import time
import sys
import random
from importlib import reload
import plotly.graph_objects as go
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, regularizers
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras.layers import LSTM, Embedding, RepeatVector, TimeDistributed, Masking, Bidirectional
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LambdaCallback
IS_COLAB = False
IS_TRAINING = True
RESULT_NAME = ""
if IS_COLAB:
from google.colab import drive
drive.mount('/content/drive')
data_path = "/content/drive/My Drive/CEM-Data-Experiment/cem-data-experiment/"
else:
data_path = "../../"
sys.path.append(data_path)
from data_processing.nasa_random_data import NasaRandomizedData
from data_processing.prepare_rul_data import RulHandler
```
### Config logging
```
reload(logging)
logging.basicConfig(format='%(asctime)s [%(levelname)s]: %(message)s', level=logging.DEBUG, datefmt='%Y/%m/%d %H:%M:%S')
```
# Load Data
```
train_names = [
'Battery_Uniform_Distribution_Variable_Charge_Room_Temp_DataSet_2Post/data/Matlab/RW1',
'Battery_Uniform_Distribution_Variable_Charge_Room_Temp_DataSet_2Post/data/Matlab/RW2',
'Battery_Uniform_Distribution_Variable_Charge_Room_Temp_DataSet_2Post/data/Matlab/RW7',
#'Battery_Uniform_Distribution_Discharge_Room_Temp_DataSet_2Post/data/Matlab/RW3',
'Battery_Uniform_Distribution_Discharge_Room_Temp_DataSet_2Post/data/Matlab/RW4',
'Battery_Uniform_Distribution_Discharge_Room_Temp_DataSet_2Post/data/Matlab/RW5',
#'Battery_Uniform_Distribution_Charge_Discharge_DataSet_2Post/data/Matlab/RW9',
#'Battery_Uniform_Distribution_Charge_Discharge_DataSet_2Post/data/Matlab/RW10',
#'Battery_Uniform_Distribution_Charge_Discharge_DataSet_2Post/data/Matlab/RW11',
'RW_Skewed_Low_Room_Temp_DataSet_2Post/data/Matlab/RW13',
'RW_Skewed_Low_Room_Temp_DataSet_2Post/data/Matlab/RW14',
'RW_Skewed_Low_Room_Temp_DataSet_2Post/data/Matlab/RW15',
'RW_Skewed_High_Room_Temp_DataSet_2Post/data/Matlab/RW17',
'RW_Skewed_High_Room_Temp_DataSet_2Post/data/Matlab/RW18',
'RW_Skewed_High_Room_Temp_DataSet_2Post/data/Matlab/RW19',
'RW_Skewed_Low_40C_DataSet_2Post/data/Matlab/RW21',
'RW_Skewed_Low_40C_DataSet_2Post/data/Matlab/RW22',
'RW_Skewed_Low_40C_DataSet_2Post/data/Matlab/RW23',
'RW_Skewed_High_40C_DataSet_2Post/data/Matlab/RW25',
'RW_Skewed_High_40C_DataSet_2Post/data/Matlab/RW26',
'RW_Skewed_High_40C_DataSet_2Post/data/Matlab/RW27',
]
test_names = [
'Battery_Uniform_Distribution_Variable_Charge_Room_Temp_DataSet_2Post/data/Matlab/RW8',
#'Battery_Uniform_Distribution_Discharge_Room_Temp_DataSet_2Post/data/Matlab/RW6',
#'Battery_Uniform_Distribution_Charge_Discharge_DataSet_2Post/data/Matlab/RW12',
'RW_Skewed_Low_Room_Temp_DataSet_2Post/data/Matlab/RW16',
'RW_Skewed_High_Room_Temp_DataSet_2Post/data/Matlab/RW20',
'RW_Skewed_Low_40C_DataSet_2Post/data/Matlab/RW24',
'RW_Skewed_High_40C_DataSet_2Post/data/Matlab/RW28',
]
nasa_data_handler = NasaRandomizedData(data_path)
rul_handler = RulHandler()
```
## Data preparation
```
CAPACITY_THRESHOLDS = None
NOMINAL_CAPACITY = 2.2
N_CYCLE = 500
WARMUP_TRAIN = 15
WARMUP_TEST = 30
(train_x, train_y_soh, test_x, test_y_soh,
train_battery_range, test_battery_range,
time_train, time_test, current_train, current_test) = nasa_data_handler.get_discharge_whole_cycle_future(train_names, test_names)
train_y = rul_handler.prepare_y_future(train_names, train_battery_range, train_y_soh, current_train, time_train, CAPACITY_THRESHOLDS, capacity=NOMINAL_CAPACITY)
del globals()["current_train"]
del globals()["time_train"]
test_y = rul_handler.prepare_y_future(test_names, test_battery_range, test_y_soh, current_test, time_test, CAPACITY_THRESHOLDS, capacity=NOMINAL_CAPACITY)
del globals()["current_test"]
del globals()["time_test"]
train_x, test_x = rul_handler.compress_cycle(train_x, test_x)
x_norm = rul_handler.Normalization()
train_x, test_x = x_norm.normalize(train_x, test_x)
train_x = rul_handler.battery_life_to_time_series(train_x, N_CYCLE, train_battery_range)
test_x = rul_handler.battery_life_to_time_series(test_x, N_CYCLE, test_battery_range)
train_x, train_y, train_battery_range, train_y_soh = rul_handler.delete_initial(train_x, train_y, train_battery_range, train_y_soh, WARMUP_TRAIN)
test_x, test_y, test_battery_range, test_y_soh = rul_handler.delete_initial(test_x, test_y, test_battery_range, test_y_soh, WARMUP_TEST)
train_x, train_y, train_battery_range, train_y_soh = rul_handler.limit_zeros(train_x, train_y, train_battery_range, train_y_soh)
test_x, test_y, test_battery_range, test_y_soh = rul_handler.limit_zeros(test_x, test_y, test_battery_range, test_y_soh)
# first one is SOH, we keep only RUL
train_y = train_y[:,1]
test_y = test_y[:,1]
```
### Y normalization
```
y_norm = rul_handler.Normalization()
train_y, test_y = y_norm.normalize(train_y, test_y)
```
# Model training
```
if IS_TRAINING:
EXPERIMENT = "lstm_rul_nasa_randomized"
experiment_name = time.strftime("%Y-%m-%d-%H-%M-%S") + '_' + EXPERIMENT
print(experiment_name)
# Model definition
opt = tf.keras.optimizers.Adam(lr=0.000003)
model = Sequential()
model.add(Masking(input_shape=(train_x.shape[1], train_x.shape[2])))
model.add(LSTM(128, activation='selu',
return_sequences=True,
kernel_regularizer=regularizers.l2(0.0002)))
model.add(LSTM(64, activation='selu', return_sequences=False,
kernel_regularizer=regularizers.l2(0.0002)))
model.add(Dense(64, activation='selu', kernel_regularizer=regularizers.l2(0.0002)))
model.add(Dense(32, activation='selu', kernel_regularizer=regularizers.l2(0.0002)))
model.add(Dense(1, activation='linear'))
model.summary()
model.compile(optimizer=opt, loss='huber', metrics=['mse', 'mae', 'mape', tf.keras.metrics.RootMeanSquaredError(name='rmse')])
if IS_TRAINING:
history = model.fit(train_x, train_y,
epochs=500,
batch_size=32,
verbose=1,
validation_split=0
)
if IS_TRAINING:
model.save(data_path + 'results/trained_model/%s.h5' % experiment_name)
hist_df = pd.DataFrame(history.history)
hist_csv_file = data_path + 'results/trained_model/%s_history.csv' % experiment_name
with open(hist_csv_file, mode='w') as f:
hist_df.to_csv(f)
history = history.history
if not IS_TRAINING:
history = pd.read_csv(data_path + 'results/trained_model/%s_history.csv' % RESULT_NAME)
model = keras.models.load_model(data_path + 'results/trained_model/%s.h5' % RESULT_NAME)
model.summary()
if not IS_TRAINING:
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
print(history)
```
### Testing
```
results = model.evaluate(test_x, test_y, return_dict = True)
print(results)
max_rmse = 0
for index in range(test_x.shape[0]):
result = model.evaluate(np.array([test_x[index, :, :]]), np.array([test_y[index]]), return_dict = True, verbose=0)
max_rmse = max(max_rmse, result['rmse'])
print("Max rmse: {}".format(max_rmse))
```
# Results Visualization
```
fig = go.Figure()
fig.add_trace(go.Scatter(y=history['loss'],
mode='lines', name='train'))
fig.update_layout(title='Loss trend',
xaxis_title='epoch',
yaxis_title='loss',
width=1400,
height=600)
fig.show()
train_predictions = model.predict(train_x)
train_y = y_norm.denormalize(train_y)
train_predictions = y_norm.denormalize(train_predictions)
a = 0
for b in train_battery_range:
fig = go.Figure()
fig.add_trace(go.Scatter(x=train_y_soh[a:b], y=train_predictions[a:b,0],
mode='lines', name='predicted'))
fig.add_trace(go.Scatter(x=train_y_soh[a:b], y=train_y[a:b],
mode='lines', name='actual'))
fig.update_layout(title='Results on training',
xaxis_title='SoH Capacity',
yaxis_title='Remaining Ah until EOL',
xaxis={'autorange':'reversed'},
width=1400,
height=600)
fig.show()
a = b
a = 0
for b in train_battery_range:
fig = go.Figure()
fig.add_trace(go.Scatter(y=train_predictions[a:b,0],
mode='lines', name='predicted'))
fig.add_trace(go.Scatter(y=train_y[a:b],
mode='lines', name='actual'))
fig.update_layout(title='Results on training',
xaxis_title='Cycle',
yaxis_title='Remaining Ah until EOL',
width=1400,
height=600)
fig.show()
a = b
test_predictions = model.predict(test_x)
test_y = y_norm.denormalize(test_y)
test_predictions = y_norm.denormalize(test_predictions)
a = 0
for b in test_battery_range:
fig = go.Figure()
fig.add_trace(go.Scatter(x=test_y_soh[a:b], y=test_predictions[a:b,0],
mode='lines', name='predicted'))
fig.add_trace(go.Scatter(x = test_y_soh[a:b], y=test_y[a:b],
mode='lines', name='actual'))
fig.update_layout(title='Results on testing',
xaxis_title='SoH Capacity',
yaxis_title='Remaining Ah until EOL',
xaxis={'autorange':'reversed'},
width=1400,
height=600)
fig.show()
a = b
a = 0
for b in test_battery_range:
fig = go.Figure()
fig.add_trace(go.Scatter(y=test_predictions[a:b, 0],
mode='lines', name='predicted'))
fig.add_trace(go.Scatter(y=test_y[a:b],
mode='lines', name='actual'))
fig.update_layout(title='Results on testing',
xaxis_title='Cycle',
yaxis_title='Remaining Ah until EOL',
width=1400,
height=600)
fig.show()
a = b
```
| github_jupyter |
# for GPU-accelerated ELM, install `scikit-cuda` package from Pip
```
import numpy as np
import hpelm
from sklearn.datasets import make_moons, make_regression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
import matplotlib
from matplotlib import pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
matplotlib.rcParams['figure.figsize'] = [10, 8]
```
## Prepare some data
```
X, Y = make_moons(n_samples=2000000, noise=0.3)
Xt, Xs, Yt, Ys = train_test_split(X, Y, test_size=0.3)
subs = 1000 # subsampling for plots
L = 100 # number of non-linear neurons
plt.scatter(Xs[Ys==0, 0][::subs], Xs[Ys==0, 1][::subs], c="r")
plt.scatter(Xs[Ys==1, 0][::subs], Xs[Ys==1, 1][::subs], c="b")
plt.show()
rfc = RandomForestClassifier(n_estimators=8, n_jobs=-1)
%time rfc.fit(Xt, Yt)
Yh = rfc.predict(Xs)
plt.scatter(Xs[Yh==0, 0][::subs], Xs[Yh==0, 1][::subs], c="r")
plt.scatter(Xs[Yh==1, 0][::subs], Xs[Yh==1, 1][::subs], c="b")
plt.show()
```
## Run HPELM
```
elm = hpelm.HPELM(2, 1, precision='double', norm=1)
elm.add_neurons(2, 'lin')
elm.add_neurons(L, 'tanh')
%time elm.train(Xt, Yt)
Yh = elm.predict(Xs)
Yh = np.array(Yh[:,0] > 0.5, dtype=np.int)
plt.scatter(Xs[Yh==0, 0][::subs], Xs[Yh==0, 1][::subs], c="r")
plt.scatter(Xs[Yh==1, 0][::subs], Xs[Yh==1, 1][::subs], c="b")
plt.show()
```
## Run HPELM+GPU
```
import os
elm2 = hpelm.HPELM(2, 1, precision='single', accelerator="GPU", norm=1)
elm2.add_neurons(2, 'lin')
elm2.add_neurons(L, 'tanh')
%time elm2.train(Xt, Yt)
Yh = elm2.predict(Xs)
Yh = np.array(Yh[:,0] > 0.5, dtype=np.int)
plt.scatter(Xs[Yh==0, 0][::subs], Xs[Yh==0, 1][::subs], c="r")
plt.scatter(Xs[Yh==1, 0][::subs], Xs[Yh==1, 1][::subs], c="b")
plt.show()
```
## Dummy task with very large data
```
X_big = np.random.randn(10000000, 200) # 10 million samples
# outputs are sums of first 10 and second 10 input values
Y_big = np.stack((X_big[:, :10].sum(1), X_big[:, 10:20].sum(1))).T
Y_big.shape
elm_cpu = hpelm.HPELM(200, 2)
elm_gpu = hpelm.HPELM(200, 2, precision="single", accelerator="GPU")
elm_cpu.add_neurons(200, 'lin')
elm_cpu.add_neurons(1000, 'tanh')
elm_gpu.add_neurons(200, 'lin')
elm_gpu.add_neurons(1000, 'tanh')
%%time
elm_cpu.train(X_big, Y_big)
%%time
elm_gpu.train(X_big, Y_big)
```
plots checking that output weights correctly correspond
to sum of first 10 inputs and the next 10 inputs
```
B_cpu = elm_cpu.nnet.get_B()
plt.plot(range(1200), B_cpu[:,0])
plt.plot(range(1200), B_cpu[:,1])
plt.xlim([0, 50])
plt.show()
B_gpu = elm_gpu.nnet.get_B() # this corectly downloads weights from GPU back to main memory
plt.plot(range(1200), B_gpu[:,0])
plt.plot(range(1200), B_gpu[:,1])
plt.xlim([0, 50])
plt.show()
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from statsmodels.graphics.tsaplots import plot_pacf
from statsmodels.graphics.tsaplots import plot_acf
from matplotlib.pyplot import figure
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix
```
# Visualisation of Prediction - Artifical Signal
```
artifical_data = pd.read_csv("../../files/classification/MLE/variance_data_new.csv", sep=";")
artifical_data.head()
mean_trainings_data = 0.06844859
variance_trainings_data = 112.93894129
artifical_data["y_true target"] = artifical_data["y_true target"] * np.sqrt(variance_trainings_data)
artifical_data["y_true mu predicted"] = artifical_data["y_true mu predicted"] * np.sqrt(variance_trainings_data)
#artifical_data["y_true sigma predicted"] = artifical_data["y_true sigma predicted"] * np.sqrt(variance_trainings_data)
artifical_data["y_true target"] = artifical_data["y_true target"] + mean_trainings_data
artifical_data["y_true mu predicted"] = artifical_data["y_true mu predicted"] + mean_trainings_data
#artifical_data["y_true sigma predicted"] = artifical_data["y_true sigma predicted"] + mean_trainings_data
artifical_data["constant sigma_1"] = 2
artifical_data["constant sigma_2"] = 5
artifical_data.head()
fig = plt.figure(figsize=(14,6), dpi=200)
ax = fig.add_subplot(111)
time = artifical_data["time"].values
training_data = artifical_data.iloc[:,1].values
# Predicted sigma band
mu_predicted = artifical_data.iloc[:,8].values
sigma_predicted = artifical_data.iloc[:,11].values
lower_bound_predicted = mu_predicted - sigma_predicted
upper_bound_predicted = mu_predicted + sigma_predicted
ax.fill_between(time, lower_bound_predicted, upper_bound_predicted, color="green", alpha=0.3, label="mean RE + 60%")
# True sigma band
mu_true = artifical_data.iloc[:,8].values
sigma_true = artifical_data.iloc[:,10].values
lower_bound_true = mu_true - sigma_true
upper_bound_true = mu_true + sigma_true
ax.fill_between(time, lower_bound_true, upper_bound_true, color="blue", alpha=0.3, label="mean RE + 40%")
"""
# predicted mean
sns.lineplot(
x=time,
y=mu_predicted,
ax=ax,
color="blue",
label="predicted mean",
linewidth=2)
"""
# true mean
sns.lineplot(
x=time,
y=mu_true,
ax=ax,
color="red",
label="predicted mean",
linewidth=2)
# training data
sns.scatterplot(
x=time,
y=training_data,
ax=ax,
color="black",
s=5,
label="trainings data",
linewidth=0.1)
# Set labels etc.
ax.legend(loc='upper center',
bbox_to_anchor=(0.5, -0.15),
fancybox=True,
shadow=True,
ncol=6,
prop={'size': 16})
ax.set_ylabel("")
ax.set_xlim(-2.98,1)
ax.tick_params(labelsize=12)
#ax.set_title("Idealized representation of prediction results", fontsize=18)
plt.tight_layout(pad=1)
plt.savefig("./pictures/prediction_variance_MSE_paper.png")
plt.show()
fig = plt.figure(figsize=(14,6), dpi=200)
ax = fig.add_subplot(111)
time = artifical_data["time"].values
training_data = artifical_data.iloc[:,1].values
# Predicted sigma band
mu_predicted = artifical_data.iloc[:,8].values
sigma_predicted = artifical_data.iloc[:,9].values
lower_bound_predicted = mu_predicted - 2*sigma_predicted
upper_bound_predicted = mu_predicted + 2*sigma_predicted
ax.fill_between(time, lower_bound_predicted, upper_bound_predicted, color="blue", alpha=0.3, label="predicted 2 $\sigma$ uncertainty interval")
"""
# predicted mean
sns.lineplot(
x=time,
y=mu_predicted,
ax=ax,
color="blue",
label="predicted mean",
linewidth=2)
"""
# true mean
sns.lineplot(
x=time,
y=mu_true,
ax=ax,
color="red",
label="predicted mean",
linewidth=2)
# training data
sns.scatterplot(
x=time,
y=training_data,
ax=ax,
color="black",
s=5,
label="trainings data",
linewidth=0.1)
# Set labels etc.
ax.legend(loc='upper center',
bbox_to_anchor=(0.5, -0.15),
fancybox=True,
shadow=True,
ncol=6,
prop={'size': 16})
ax.set_ylabel("")
ax.set_xlim(-2.98,1)
ax.tick_params(labelsize=12)
#ax.set_title("Idealized representation of prediction results", fontsize=18)
plt.tight_layout(pad=1)
plt.savefig("./pictures/prediction_variance_MLE_paper.png")
plt.show()
ax.legend(
fancybox=True,
shadow=True,
ncol=1,
prop={'size': 12})
```
| github_jupyter |
# Image Captioning
**Author:** [A_K_Nain](https://twitter.com/A_K_Nain)<br>
**Date created:** 2021/05/29<br>
**Last modified:** 2021/10/31<br>
**Description:** Implement an image captioning model using a CNN and a Transformer.
## Setup
```
import os
import re
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.applications import efficientnet
from tensorflow.keras.layers import TextVectorization
seed = 111
np.random.seed(seed)
tf.random.set_seed(seed)
```
## Download the dataset
We will be using the Flickr8K dataset for this tutorial. This dataset comprises over
8,000 images, that are each paired with five different captions.
```
!wget -q https://github.com/jbrownlee/Datasets/releases/download/Flickr8k/Flickr8k_Dataset.zip
!wget -q https://github.com/jbrownlee/Datasets/releases/download/Flickr8k/Flickr8k_text.zip
!unzip -qq Flickr8k_Dataset.zip
!unzip -qq Flickr8k_text.zip
!rm Flickr8k_Dataset.zip Flickr8k_text.zip
# Path to the images
IMAGES_PATH = "Flicker8k_Dataset"
# Desired image dimensions
IMAGE_SIZE = (299, 299)
# Vocabulary size
VOCAB_SIZE = 10000
# Fixed length allowed for any sequence
SEQ_LENGTH = 25
# Dimension for the image embeddings and token embeddings
EMBED_DIM = 512
# Per-layer units in the feed-forward network
FF_DIM = 512
# Other training parameters
BATCH_SIZE = 64
EPOCHS = 30
AUTOTUNE = tf.data.AUTOTUNE
```
## Preparing the dataset
```
def load_captions_data(filename):
"""Loads captions (text) data and maps them to corresponding images.
Args:
filename: Path to the text file containing caption data.
Returns:
caption_mapping: Dictionary mapping image names and the corresponding captions
text_data: List containing all the available captions
"""
with open(filename) as caption_file:
caption_data = caption_file.readlines()
caption_mapping = {}
text_data = []
images_to_skip = set()
for line in caption_data:
line = line.rstrip("\n")
# Image name and captions are separated using a tab
img_name, caption = line.split("\t")
# Each image is repeated five times for the five different captions.
# Each image name has a suffix `#(caption_number)`
img_name = img_name.split("#")[0]
img_name = os.path.join(IMAGES_PATH, img_name.strip())
# We will remove caption that are either too short to too long
tokens = caption.strip().split()
if len(tokens) < 5 or len(tokens) > SEQ_LENGTH:
images_to_skip.add(img_name)
continue
if img_name.endswith("jpg") and img_name not in images_to_skip:
# We will add a start and an end token to each caption
caption = "<start> " + caption.strip() + " <end>"
text_data.append(caption)
if img_name in caption_mapping:
caption_mapping[img_name].append(caption)
else:
caption_mapping[img_name] = [caption]
for img_name in images_to_skip:
if img_name in caption_mapping:
del caption_mapping[img_name]
return caption_mapping, text_data
def train_val_split(caption_data, train_size=0.8, shuffle=True):
"""Split the captioning dataset into train and validation sets.
Args:
caption_data (dict): Dictionary containing the mapped caption data
train_size (float): Fraction of all the full dataset to use as training data
shuffle (bool): Whether to shuffle the dataset before splitting
Returns:
Traning and validation datasets as two separated dicts
"""
# 1. Get the list of all image names
all_images = list(caption_data.keys())
# 2. Shuffle if necessary
if shuffle:
np.random.shuffle(all_images)
# 3. Split into training and validation sets
train_size = int(len(caption_data) * train_size)
training_data = {
img_name: caption_data[img_name] for img_name in all_images[:train_size]
}
validation_data = {
img_name: caption_data[img_name] for img_name in all_images[train_size:]
}
# 4. Return the splits
return training_data, validation_data
# Load the dataset
captions_mapping, text_data = load_captions_data("Flickr8k.token.txt")
# Split the dataset into training and validation sets
train_data, valid_data = train_val_split(captions_mapping)
print("Number of training samples: ", len(train_data))
print("Number of validation samples: ", len(valid_data))
```
## Vectorizing the text data
We'll use the `TextVectorization` layer to vectorize the text data,
that is to say, to turn the
original strings into integer sequences where each integer represents the index of
a word in a vocabulary. We will use a custom string standardization scheme
(strip punctuation characters except `<` and `>`) and the default
splitting scheme (split on whitespace).
```
def custom_standardization(input_string):
lowercase = tf.strings.lower(input_string)
return tf.strings.regex_replace(lowercase, "[%s]" % re.escape(strip_chars), "")
strip_chars = "!\"#$%&'()*+,-./:;<=>?@[\]^_`{|}~"
strip_chars = strip_chars.replace("<", "")
strip_chars = strip_chars.replace(">", "")
vectorization = TextVectorization(
max_tokens=VOCAB_SIZE,
output_mode="int",
output_sequence_length=SEQ_LENGTH,
standardize=custom_standardization,
)
vectorization.adapt(text_data)
# Data augmentation for image data
image_augmentation = keras.Sequential(
[
layers.RandomFlip("horizontal"),
layers.RandomRotation(0.2),
layers.RandomContrast(0.3),
]
)
```
## Building a `tf.data.Dataset` pipeline for training
We will generate pairs of images and corresponding captions using a `tf.data.Dataset` object.
The pipeline consists of two steps:
1. Read the image from the disk
2. Tokenize all the five captions corresponding to the image
```
def decode_and_resize(img_path):
img = tf.io.read_file(img_path)
img = tf.image.decode_jpeg(img, channels=3)
img = tf.image.resize(img, IMAGE_SIZE)
img = tf.image.convert_image_dtype(img, tf.float32)
return img
def process_input(img_path, captions):
return decode_and_resize(img_path), vectorization(captions)
def make_dataset(images, captions):
dataset = tf.data.Dataset.from_tensor_slices((images, captions))
dataset = dataset.shuffle(len(images))
dataset = dataset.map(process_input, num_parallel_calls=AUTOTUNE)
dataset = dataset.batch(BATCH_SIZE).prefetch(AUTOTUNE)
return dataset
# Pass the list of images and the list of corresponding captions
train_dataset = make_dataset(list(train_data.keys()), list(train_data.values()))
valid_dataset = make_dataset(list(valid_data.keys()), list(valid_data.values()))
```
## Building the model
Our image captioning architecture consists of three models:
1. A CNN: used to extract the image features
2. A TransformerEncoder: The extracted image features are then passed to a Transformer
based encoder that generates a new representation of the inputs
3. A TransformerDecoder: This model takes the encoder output and the text data
(sequences) as inputs and tries to learn to generate the caption.
```
def get_cnn_model():
base_model = efficientnet.EfficientNetB0(
input_shape=(*IMAGE_SIZE, 3), include_top=False, weights="imagenet",
)
# We freeze our feature extractor
base_model.trainable = False
base_model_out = base_model.output
base_model_out = layers.Reshape((-1, base_model_out.shape[-1]))(base_model_out)
cnn_model = keras.models.Model(base_model.input, base_model_out)
return cnn_model
class TransformerEncoderBlock(layers.Layer):
def __init__(self, embed_dim, dense_dim, num_heads, **kwargs):
super().__init__(**kwargs)
self.embed_dim = embed_dim
self.dense_dim = dense_dim
self.num_heads = num_heads
self.attention_1 = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=embed_dim, dropout=0.0
)
self.layernorm_1 = layers.LayerNormalization()
self.layernorm_2 = layers.LayerNormalization()
self.dense_1 = layers.Dense(embed_dim, activation="relu")
def call(self, inputs, training, mask=None):
inputs = self.layernorm_1(inputs)
inputs = self.dense_1(inputs)
attention_output_1 = self.attention_1(
query=inputs,
value=inputs,
key=inputs,
attention_mask=None,
training=training,
)
out_1 = self.layernorm_2(inputs + attention_output_1)
return out_1
class PositionalEmbedding(layers.Layer):
def __init__(self, sequence_length, vocab_size, embed_dim, **kwargs):
super().__init__(**kwargs)
self.token_embeddings = layers.Embedding(
input_dim=vocab_size, output_dim=embed_dim
)
self.position_embeddings = layers.Embedding(
input_dim=sequence_length, output_dim=embed_dim
)
self.sequence_length = sequence_length
self.vocab_size = vocab_size
self.embed_dim = embed_dim
self.embed_scale = tf.math.sqrt(tf.cast(embed_dim, tf.float32))
def call(self, inputs):
length = tf.shape(inputs)[-1]
positions = tf.range(start=0, limit=length, delta=1)
embedded_tokens = self.token_embeddings(inputs)
embedded_tokens = embedded_tokens * self.embed_scale
embedded_positions = self.position_embeddings(positions)
return embedded_tokens + embedded_positions
def compute_mask(self, inputs, mask=None):
return tf.math.not_equal(inputs, 0)
class TransformerDecoderBlock(layers.Layer):
def __init__(self, embed_dim, ff_dim, num_heads, **kwargs):
super().__init__(**kwargs)
self.embed_dim = embed_dim
self.ff_dim = ff_dim
self.num_heads = num_heads
self.attention_1 = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=embed_dim, dropout=0.1
)
self.attention_2 = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=embed_dim, dropout=0.1
)
self.ffn_layer_1 = layers.Dense(ff_dim, activation="relu")
self.ffn_layer_2 = layers.Dense(embed_dim)
self.layernorm_1 = layers.LayerNormalization()
self.layernorm_2 = layers.LayerNormalization()
self.layernorm_3 = layers.LayerNormalization()
self.embedding = PositionalEmbedding(
embed_dim=EMBED_DIM, sequence_length=SEQ_LENGTH, vocab_size=VOCAB_SIZE
)
self.out = layers.Dense(VOCAB_SIZE, activation="softmax")
self.dropout_1 = layers.Dropout(0.3)
self.dropout_2 = layers.Dropout(0.5)
self.supports_masking = True
def call(self, inputs, encoder_outputs, training, mask=None):
inputs = self.embedding(inputs)
causal_mask = self.get_causal_attention_mask(inputs)
if mask is not None:
padding_mask = tf.cast(mask[:, :, tf.newaxis], dtype=tf.int32)
combined_mask = tf.cast(mask[:, tf.newaxis, :], dtype=tf.int32)
combined_mask = tf.minimum(combined_mask, causal_mask)
attention_output_1 = self.attention_1(
query=inputs,
value=inputs,
key=inputs,
attention_mask=combined_mask,
training=training,
)
out_1 = self.layernorm_1(inputs + attention_output_1)
attention_output_2 = self.attention_2(
query=out_1,
value=encoder_outputs,
key=encoder_outputs,
attention_mask=padding_mask,
training=training,
)
out_2 = self.layernorm_2(out_1 + attention_output_2)
ffn_out = self.ffn_layer_1(out_2)
ffn_out = self.dropout_1(ffn_out, training=training)
ffn_out = self.ffn_layer_2(ffn_out)
ffn_out = self.layernorm_3(ffn_out + out_2, training=training)
ffn_out = self.dropout_2(ffn_out, training=training)
preds = self.out(ffn_out)
return preds
def get_causal_attention_mask(self, inputs):
input_shape = tf.shape(inputs)
batch_size, sequence_length = input_shape[0], input_shape[1]
i = tf.range(sequence_length)[:, tf.newaxis]
j = tf.range(sequence_length)
mask = tf.cast(i >= j, dtype="int32")
mask = tf.reshape(mask, (1, input_shape[1], input_shape[1]))
mult = tf.concat(
[tf.expand_dims(batch_size, -1), tf.constant([1, 1], dtype=tf.int32)],
axis=0,
)
return tf.tile(mask, mult)
class ImageCaptioningModel(keras.Model):
def __init__(
self, cnn_model, encoder, decoder, num_captions_per_image=5, image_aug=None,
):
super().__init__()
self.cnn_model = cnn_model
self.encoder = encoder
self.decoder = decoder
self.loss_tracker = keras.metrics.Mean(name="loss")
self.acc_tracker = keras.metrics.Mean(name="accuracy")
self.num_captions_per_image = num_captions_per_image
self.image_aug = image_aug
def calculate_loss(self, y_true, y_pred, mask):
loss = self.loss(y_true, y_pred)
mask = tf.cast(mask, dtype=loss.dtype)
loss *= mask
return tf.reduce_sum(loss) / tf.reduce_sum(mask)
def calculate_accuracy(self, y_true, y_pred, mask):
accuracy = tf.equal(y_true, tf.argmax(y_pred, axis=2))
accuracy = tf.math.logical_and(mask, accuracy)
accuracy = tf.cast(accuracy, dtype=tf.float32)
mask = tf.cast(mask, dtype=tf.float32)
return tf.reduce_sum(accuracy) / tf.reduce_sum(mask)
def _compute_caption_loss_and_acc(self, img_embed, batch_seq, training=True):
encoder_out = self.encoder(img_embed, training=training)
batch_seq_inp = batch_seq[:, :-1]
batch_seq_true = batch_seq[:, 1:]
mask = tf.math.not_equal(batch_seq_true, 0)
batch_seq_pred = self.decoder(
batch_seq_inp, encoder_out, training=training, mask=mask
)
loss = self.calculate_loss(batch_seq_true, batch_seq_pred, mask)
acc = self.calculate_accuracy(batch_seq_true, batch_seq_pred, mask)
return loss, acc
def train_step(self, batch_data):
batch_img, batch_seq = batch_data
batch_loss = 0
batch_acc = 0
if self.image_aug:
batch_img = self.image_aug(batch_img)
# 1. Get image embeddings
img_embed = self.cnn_model(batch_img)
# 2. Pass each of the five captions one by one to the decoder
# along with the encoder outputs and compute the loss as well as accuracy
# for each caption.
for i in range(self.num_captions_per_image):
with tf.GradientTape() as tape:
loss, acc = self._compute_caption_loss_and_acc(
img_embed, batch_seq[:, i, :], training=True
)
# 3. Update loss and accuracy
batch_loss += loss
batch_acc += acc
# 4. Get the list of all the trainable weights
train_vars = (
self.encoder.trainable_variables + self.decoder.trainable_variables
)
# 5. Get the gradients
grads = tape.gradient(loss, train_vars)
# 6. Update the trainable weights
self.optimizer.apply_gradients(zip(grads, train_vars))
# 7. Update the trackers
batch_acc /= float(self.num_captions_per_image)
self.loss_tracker.update_state(batch_loss)
self.acc_tracker.update_state(batch_acc)
# 8. Return the loss and accuracy values
return {"loss": self.loss_tracker.result(), "acc": self.acc_tracker.result()}
def test_step(self, batch_data):
batch_img, batch_seq = batch_data
batch_loss = 0
batch_acc = 0
# 1. Get image embeddings
img_embed = self.cnn_model(batch_img)
# 2. Pass each of the five captions one by one to the decoder
# along with the encoder outputs and compute the loss as well as accuracy
# for each caption.
for i in range(self.num_captions_per_image):
loss, acc = self._compute_caption_loss_and_acc(
img_embed, batch_seq[:, i, :], training=False
)
# 3. Update batch loss and batch accuracy
batch_loss += loss
batch_acc += acc
batch_acc /= float(self.num_captions_per_image)
# 4. Update the trackers
self.loss_tracker.update_state(batch_loss)
self.acc_tracker.update_state(batch_acc)
# 5. Return the loss and accuracy values
return {"loss": self.loss_tracker.result(), "acc": self.acc_tracker.result()}
@property
def metrics(self):
# We need to list our metrics here so the `reset_states()` can be
# called automatically.
return [self.loss_tracker, self.acc_tracker]
cnn_model = get_cnn_model()
encoder = TransformerEncoderBlock(embed_dim=EMBED_DIM, dense_dim=FF_DIM, num_heads=1)
decoder = TransformerDecoderBlock(embed_dim=EMBED_DIM, ff_dim=FF_DIM, num_heads=2)
caption_model = ImageCaptioningModel(
cnn_model=cnn_model, encoder=encoder, decoder=decoder, image_aug=image_augmentation,
)
```
## Model training
```
# Define the loss function
cross_entropy = keras.losses.SparseCategoricalCrossentropy(
from_logits=False, reduction="none"
)
# EarlyStopping criteria
early_stopping = keras.callbacks.EarlyStopping(patience=3, restore_best_weights=True)
# Learning Rate Scheduler for the optimizer
class LRSchedule(keras.optimizers.schedules.LearningRateSchedule):
def __init__(self, post_warmup_learning_rate, warmup_steps):
super().__init__()
self.post_warmup_learning_rate = post_warmup_learning_rate
self.warmup_steps = warmup_steps
def __call__(self, step):
global_step = tf.cast(step, tf.float32)
warmup_steps = tf.cast(self.warmup_steps, tf.float32)
warmup_progress = global_step / warmup_steps
warmup_learning_rate = self.post_warmup_learning_rate * warmup_progress
return tf.cond(
global_step < warmup_steps,
lambda: warmup_learning_rate,
lambda: self.post_warmup_learning_rate,
)
# Create a learning rate schedule
num_train_steps = len(train_dataset) * EPOCHS
num_warmup_steps = num_train_steps // 15
lr_schedule = LRSchedule(post_warmup_learning_rate=1e-4, warmup_steps=num_warmup_steps)
# Compile the model
caption_model.compile(optimizer=keras.optimizers.Adam(lr_schedule), loss=cross_entropy)
# Fit the model
caption_model.fit(
train_dataset,
epochs=EPOCHS,
validation_data=valid_dataset,
callbacks=[early_stopping],
)
```
## Check sample predictions
```
vocab = vectorization.get_vocabulary()
index_lookup = dict(zip(range(len(vocab)), vocab))
max_decoded_sentence_length = SEQ_LENGTH - 1
valid_images = list(valid_data.keys())
def generate_caption():
# Select a random image from the validation dataset
sample_img = np.random.choice(valid_images)
# Read the image from the disk
sample_img = decode_and_resize(sample_img)
img = sample_img.numpy().clip(0, 255).astype(np.uint8)
plt.imshow(img)
plt.show()
# Pass the image to the CNN
img = tf.expand_dims(sample_img, 0)
img = caption_model.cnn_model(img)
# Pass the image features to the Transformer encoder
encoded_img = caption_model.encoder(img, training=False)
# Generate the caption using the Transformer decoder
decoded_caption = "<start> "
for i in range(max_decoded_sentence_length):
tokenized_caption = vectorization([decoded_caption])[:, :-1]
mask = tf.math.not_equal(tokenized_caption, 0)
predictions = caption_model.decoder(
tokenized_caption, encoded_img, training=False, mask=mask
)
sampled_token_index = np.argmax(predictions[0, i, :])
sampled_token = index_lookup[sampled_token_index]
if sampled_token == " <end>":
break
decoded_caption += " " + sampled_token
decoded_caption = decoded_caption.replace("<start> ", "")
decoded_caption = decoded_caption.replace(" <end>", "").strip()
print("Predicted Caption: ", decoded_caption)
# Check predictions for a few samples
generate_caption()
generate_caption()
generate_caption()
```
## End Notes
We saw that the model starts to generate reasonable captions after a few epochs. To keep
this example easily runnable, we have trained it with a few constraints, like a minimal
number of attention heads. To improve the predictions, you can try changing these training
settings and find a good model for your use case.
| github_jupyter |
# Simple Spark test: read and filter tiny CSV file
# <span style="color:red"> NOTE </span>
Please set the `context_already_defined` flag depending on your context.
If this notebook is started against a pyspark session, there will already be
a defined context called `spark`. Attempts to create a new one will fail.
If this notebook is meant to start a Spark context in local mode,
set `context_already_defined = False`, sine one needs to create a context.
```
from pyspark.sql.types import *
context_already_defined = True
if context_already_defined:
sc = spark.sparkContext
sqlc = SQLContext(sc)
else:
from pyspark import SparkContext, SparkConf, SQLContext
conf = SparkConf().setAppName("msmap-filter").setMaster("local[*]")
sc = SparkContext(conf=conf)
sqlc = SQLContext(sc)
```
## Create tiny CSV file and read it into a pyspak Dataframe
```
tiny_csv="""
A,B
1,4.45
2,4.55
3,7.7
4,8.2"""
with open('test_data.csv', 'w') as tiny_csv_file:
tiny_csv_file.write(tiny_csv)
! cat test_data.csv
test_file = 'test_data.csv'
ms_schema = StructType([StructField("A", ByteType()),
StructField("B", FloatType())])
test_data = sqlc.read.csv(test_file, schema=ms_schema, header="true")
test_data.show(2)
```
## Please inspect the application SparkUI for stage execution trace and SQL
## Stage Details
```
org.apache.spark.sql.Dataset.showString(Dataset.scala:245)
sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
java.lang.reflect.Method.invoke(Method.java:498)
py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
py4j.Gateway.invoke(Gateway.java:280)
py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
py4j.commands.CallCommand.execute(CallCommand.java:79)
py4j.GatewayConnection.run(GatewayConnection.java:214)
java.lang.Thread.run(Thread.java:745)
```
## SQL for stage
WholeStageCodegen: Scan csv number of output rows: 4number of files: 1metadata time (ms): 6
```
== Parsed Logical Plan ==
GlobalLimit 3
+- LocalLimit 3
+- Relation[A#0,B#1] csv
== Analyzed Logical Plan ==
A: tinyint, B: float
GlobalLimit 3
+- LocalLimit 3
+- Relation[A#0,B#1] csv
== Optimized Logical Plan ==
GlobalLimit 3
+- LocalLimit 3
+- Relation[A#0,B#1] csv
== Physical Plan ==
CollectLimit 3
+- *FileScan csv [A#0,B#1] Batched: false, Format: CSV, Location: InMemoryFileIndex[file:<edited>/test_data.csv], PartitionFilters: [], PushedFilters: [], ReadSchema: struct<A:tinyint,B:float>
```
## Simple numerical filter on the inner RDD
```
test_data = test_data.rdd.filter(lambda row: row['A'] % 2 == 0)
test_data.take(4)
```
## Stage Details
```
org.apache.spark.SparkContext.runJob(SparkContext.scala:2062)
org.apache.spark.api.python.PythonRDD$.runJob(PythonRDD.scala:446)
org.apache.spark.api.python.PythonRDD.runJob(PythonRDD.scala)
sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
java.lang.reflect.Method.invoke(Method.java:498)
py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
py4j.Gateway.invoke(Gateway.java:280)
py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
py4j.commands.CallCommand.execute(CallCommand.java:79)
py4j.GatewayConnection.run(GatewayConnection.java:214)
java.lang.Thread.run(Thread.java:745)
```
| github_jupyter |
# Introduction #
So far in this course, we've learned about how neural networks can solve regression problems. Now we're going to apply neural networks to another common machine learning problem: classification. Most everything we've learned up until now still applies. The main difference is in the loss function we use and in what kind of outputs we want the final layer to produce.
# Binary Classification #
Classification into one of two classes is a common machine learning problem. You might want to predict whether or not a customer is likely to make a purchase, whether or not a credit card transaction was fraudulent, whether deep space signals show evidence of a new planet, or a medical test evidence of a disease. These are all **binary classification** problems.
In your raw data, the classes might be represented by strings like `"Yes"` and `"No"`, or `"Dog"` and `"Cat"`. Before using this data we'll assign a **class label**: one class will be `0` and the other will be `1`. Assigning numeric labels puts the data in a form a neural network can use.
# Accuracy and Cross-Entropy #
**Accuracy** is one of the many metrics in use for measuring success on a classification problem. Accuracy is the ratio of correct predictions to total predictions: `accuracy = number_correct / total`. A model that always predicted correctly would have an accuracy score of `1.0`. All else being equal, accuracy is a reasonable metric to use whenever the classes in the dataset occur with about the same frequency.
The problem with accuracy (and most other classification metrics) is that it can't be used as a loss function. SGD needs a loss function that changes smoothly, but accuracy, being a ratio of counts, changes in "jumps". So, we have to choose a substitute to act as the loss function. This substitute is the *cross-entropy* function.
Now, recall that the loss function defines the *objective* of the network during training. With regression, our goal was to minimize the distance between the expected outcome and the predicted outcome. We chose MAE to measure this distance.
For classification, what we want instead is a distance between *probabilities*, and this is what cross-entropy provides. **Cross-entropy** is a sort of measure for the distance from one probability distribution to another.
<figure style="padding: 1em;">
<img src="https://i.imgur.com/DwVV9bR.png" width="400" alt="Graphs of accuracy and cross-entropy.">
<figcaption style="textalign: center; font-style: italic"><center>Cross-entropy penalizes incorrect probability predictions.</center></figcaption>
</figure>
The idea is that we want our network to predict the correct class with probability `1.0`. The further away the predicted probability is from `1.0`, the greater will be the cross-entropy loss.
The technical reasons we use cross-entropy are a bit subtle, but the main thing to take away from this section is just this: use cross-entropy for a classification loss; other metrics you might care about (like accuracy) will tend to improve along with it.
# Making Probabilities with the Sigmoid Function #
The cross-entropy and accuracy functions both require probabilities as inputs, meaning, numbers from 0 to 1. To covert the real-valued outputs produced by a dense layer into probabilities, we attach a new kind of activation function, the **sigmoid activation**.
<figure style="padding: 1em;">
<img src="https://i.imgur.com/FYbRvJo.png" width="400" alt="The sigmoid graph is an 'S' shape with horizontal asymptotes at 0 to the left and 1 to the right. ">
<figcaption style="textalign: center; font-style: italic"><center>The sigmoid function maps real numbers into the interval $[0, 1]$.</center></figcaption>
</figure>
To get the final class prediction, we define a *threshold* probability. Typically this will be 0.5, so that rounding will give us the correct class: below 0.5 means the class with label 0 and 0.5 or above means the class with label 1. A 0.5 threshold is what Keras uses by default with its [accuracy metric](https://www.tensorflow.org/api_docs/python/tf/keras/metrics/BinaryAccuracy).
# Example - Binary Classification #
Now let's try it out!
The [Ionosphere](https://archive.ics.uci.edu/ml/datasets/Ionosphere) dataset contains features obtained from radar signals focused on the ionosphere layer of the Earth's atmosphere. The task is to determine whether the signal shows the presence of some object, or just empty air.
```
#$HIDE_INPUT$
import pandas as pd
from IPython.display import display
ion = pd.read_csv('../input/dl-course-data/ion.csv', index_col=0)
display(ion.head())
df = ion.copy()
df['Class'] = df['Class'].map({'good': 0, 'bad': 1})
df_train = df.sample(frac=0.7, random_state=0)
df_valid = df.drop(df_train.index)
max_ = df_train.max(axis=0)
min_ = df_train.min(axis=0)
df_train = (df_train - min_) / (max_ - min_)
df_valid = (df_valid - min_) / (max_ - min_)
df_train.dropna(axis=1, inplace=True) # drop the empty feature in column 2
df_valid.dropna(axis=1, inplace=True)
X_train = df_train.drop('Class', axis=1)
X_valid = df_valid.drop('Class', axis=1)
y_train = df_train['Class']
y_valid = df_valid['Class']
```
We'll define our model just like we did for the regression tasks, with one exception. In the final layer include a `'sigmoid'` activation so that the model will produce class probabilities.
```
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
layers.Dense(4, activation='relu', input_shape=[33]),
layers.Dense(4, activation='relu'),
layers.Dense(1, activation='sigmoid'),
])
```
Add the cross-entropy loss and accuracy metric to the model with its `compile` method. For two-class problems, be sure to use `'binary'` versions. (Problems with more classes will be slightly different.) The Adam optimizer works great for classification too, so we'll stick with it.
```
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['binary_accuracy'],
)
```
The model in this particular problem can take quite a few epochs to complete training, so we'll include an early stopping callback for convenience.
```
early_stopping = keras.callbacks.EarlyStopping(
patience=10,
min_delta=0.001,
restore_best_weights=True,
)
history = model.fit(
X_train, y_train,
validation_data=(X_valid, y_valid),
batch_size=512,
epochs=1000,
callbacks=[early_stopping],
verbose=0, # hide the output because we have so many epochs
)
```
We'll take a look at the learning curves as always, and also inspect the best values for the loss and accuracy we got on the validation set. (Remember that early stopping will restore the weights to those that got these values.)
```
history_df = pd.DataFrame(history.history)
# Start the plot at epoch 5
history_df.loc[5:, ['loss', 'val_loss']].plot()
history_df.loc[5:, ['binary_accuracy', 'val_binary_accuracy']].plot()
print(("Best Validation Loss: {:0.4f}" +\
"\nBest Validation Accuracy: {:0.4f}")\
.format(history_df['val_loss'].min(),
history_df['val_binary_accuracy'].max()))
```
# Your Turn #
Use a neural network to [**predict cancellations in hotel reservations**](#$NEXT_NOTEBOOK_URL$) with the *Hotel Cancellations* dataset.
| github_jupyter |
```
%matplotlib inline
import numpy as np
import os
import csv
import itertools
from clusim.clustering import Clustering
import clusim.sim as sim
import clusim.clugen as clugen
import clusim.clusimelement as elsim
from sklearn.cluster import AgglomerativeClustering
import matplotlib.pylab as plt
def hex2rgb(value):
value = value.lstrip('#')
lv = len(value)
return tuple(int(value[i:i + lv // 3], 16)/255. for i in range(0, lv, lv // 3))
# define some colors for drawing
try:
import seaborn as sns
sns.set_style('white')
red_color = sns.color_palette()[3]
BiasColorDict = {}
BiasColorDict['jaccard_index'] = sns.color_palette()[9]
BiasColorDict['fowlkes_mallows_index'] = sns.color_palette()[2]
BiasColorDict['adj_rand'] = sns.color_palette()[8]
BiasColorDict['fmeasure'] = sns.color_palette()[6]
BiasColorDict['elmsim'] = hex2rgb('#1F1F1F')
BiasColorDict['nmi'] = sns.color_palette()[1]
BiasColorDict['onmi'] = sns.color_palette()[0]
except ImportError:
red_color = 'r'
BiasColorDict = {}
BiasColorDict['jaccard_index'] = hex2rgb('#6BB5E6')
BiasColorDict['fowlkes_mallows_index'] = hex2rgb('#008000')
BiasColorDict['adj_rand'] = hex2rgb('#F1DE50')
BiasColorDict['fmeasure'] = hex2rgb('#BF82A8')
BiasColorDict['elmsim'] = hex2rgb('#1F1F1F')
BiasColorDict['nmi'] = hex2rgb('#C86629')
BiasColorDict['onmi'] = hex2rgb('#2875AE')
# wrap the Sklearn AgglomerativeClustering
def GetHierCluster(data, membership, K = 4, metric = 'correlation', linkage_method = 'average'):
ac = AgglomerativeClustering(n_clusters=K,
affinity=metric,
linkage=linkage_method)
hierclus = Clustering()
agg_membership = ac.fit_predict(data, membership)
hierclus.from_membership_list(agg_membership)
return hierclus
```
# Example 1
The clustering similarity measures presented here differ in how each evaluates the trade-offs between variation in three primary characteristics of clusterings: the grouping of elements into clusters, the number of clusters, and the size distribution of those clusters. To illustrate these trade-offs, we present three simple examples a,b,c.
```
n_samples = 20 # set to 100 to fully reproduce the results from the paper
n_elements = 2**10
n_clusterfixed = 2**5
def adj_rand(c1, c2):
# set paramter values for adjusted rand index
return sim.adjrand_index(c1, c2, random_model = 'perm')
def elmsim(c1, c2):
# set paramter values for element_centric similarity
return sim.element_sim(c1, c2, alpha = 0.9)
# list of similarity measures to compare
measure_list = ['jaccard_index', 'fmeasure', 'adj_rand', 'fowlkes_mallows_index', 'nmi', 'onmi', 'elmsim']
n_measures = len(measure_list)
```
### Example 1a
In the first example, 1,024 elements are grouped into 32 clusters of equal size and compared against a similar clustering with a fraction of the elements randomly exchanged between the clusters, keeping the same cluster sizes.
```
n_shuffle_points = 40
percent_shuffle_values = np.linspace(0.0, 1.0, n_shuffle_points)
percent_randomize_compare = np.zeros((n_measures, n_shuffle_points, n_samples))
# loop through samples and print status every 10
for isample in range(n_samples):
if isample % 10 == 0: print(isample)
# loop through the shuffle fraction points
for iper in range(n_shuffle_points):
random_cluster = clugen.make_equal_clustering(n_elements, n_clusterfixed)
random_cluster2 = clugen.shuffle_memberships(random_cluster, percent = percent_shuffle_values[iper])
for imeasure in range(n_measures):
if measure_list[imeasure] in ['adj_rand', 'elmsim']:
percent_randomize_compare[imeasure, iper, isample] = eval(measure_list[imeasure]+'(random_cluster, random_cluster2)')
else:
percent_randomize_compare[imeasure, iper, isample] = eval('sim.'+measure_list[imeasure]+'(random_cluster, random_cluster2)')
measure_means = np.mean(percent_randomize_compare, axis = 2)
measure_stds = np.std(percent_randomize_compare, axis = 2)
print("Simulation Finished")
# plot results
fig, ax = plt.subplots(1,1, figsize = (8,8))
for imeasure in range(n_measures):
ax.fill_between(percent_shuffle_values, measure_means[imeasure] - measure_stds[imeasure],
measure_means[imeasure] + measure_stds[imeasure],
color = BiasColorDict[measure_list[imeasure]], alpha = 0.3)
line = ax.plot(percent_shuffle_values, measure_means[imeasure], color = BiasColorDict[measure_list[imeasure]],
label = measure_list[imeasure], alpha = 0.9, lw = 1, dashes = [500,1])
ax.set_ylim([0.0, 1.0])
ax.set_xticks([0.0, 0.5, 1.0])
ax.set_yticks([0.0, 0.5, 1.0])
ax.set_xlabel('Fraction of randomized elements')
ax.set_ylabel('Similarity')
for spine in ['right', 'top']:
ax.spines[spine].set_visible(False)
plt.legend()
plt.show()
```
### Example 1b
In the second example, 1,024 elements are grouped into 32 clusters of equal size and compared against a similar clustering with increasing cluster size skew-ness (measured by the entropy of the cluster size distribution).
```
n_skew_points = 10**2 # increase to 10**4 to incrase the entropy range as in the paper
skew_step_size = 5*10**2
skewed_clusters_compare = np.zeros((n_measures + 1, n_skew_points, n_samples)) # we also need to store the cluster entropy
initial_cluster = clugen.make_equal_clustering(n_elements, n_clusterfixed)
# loop through samples and print status every 10
for isample in range(n_samples):
if isample % 10 == 0: print(isample)
# create a copy of the original clustering with all memberships randomized
random_cluster = clugen.shuffle_memberships(initial_cluster, percent = 1.0)
# loop through the fraction points
for istep in range(n_skew_points):
# record the entropy
skewed_clusters_compare[n_measures, istep, isample] = sim.entropy(np.array(random_cluster.clu_size_seq, dtype = float)/n_elements)
# perform clustering similarity measurements
for imeasure in range(n_measures):
if measure_list[imeasure] in ['adj_rand', 'elmsim']:
skewed_clusters_compare[imeasure, istep, isample] = eval(measure_list[imeasure]+'(initial_cluster, random_cluster)')
else:
skewed_clusters_compare[imeasure, istep, isample] = eval('sim.'+measure_list[imeasure]+'(initial_cluster, random_cluster)')
# now apply the Preferential Attachment Model 'skew_step_size' times
random_cluster = clugen.shuffle_memberships_pa(random_cluster, n_steps=skew_step_size,
constant_num_clusters=True)
print("Simulation Finished")
# once we have finished sampling clusterings, we need to aggregate the comparisons by cluster size entropy
n_bins = 40
def binned_index(value, bins):
return ((value >= bins).argmin().astype(int) - 1)
binned_index = np.vectorize(binned_index, excluded=[1])
entropy_bins = np.linspace(skewed_clusters_compare[-1].min(), skewed_clusters_compare[-1].max(), n_bins + 1)
skewed_clusters_compare[-1] = binned_index(skewed_clusters_compare[-1], entropy_bins)
skew_measure_means = np.zeros((n_measures, n_bins))
skew_measure_stds = np.zeros((n_measures, n_bins))
for ibin in range(n_bins):
skew_measure_means[:,ibin] = np.nanmean(skewed_clusters_compare[:-1][:, skewed_clusters_compare[-1] == ibin], axis = 1)
skew_measure_stds[:,ibin] = np.nanstd(skewed_clusters_compare[:-1][:, skewed_clusters_compare[-1] == ibin], axis = 1)
# plot results
fig, ax = plt.subplots(1,1, figsize = (8,8))
for imeasure in range(n_measures):
ax.fill_between(entropy_bins[:-1], skew_measure_means[imeasure] - skew_measure_stds[imeasure],
skew_measure_means[imeasure] + skew_measure_stds[imeasure],
color = BiasColorDict[measure_list[imeasure]], alpha = 0.3)
line = ax.plot(entropy_bins[:-1], skew_measure_means[imeasure], color = BiasColorDict[measure_list[imeasure]],
label = measure_list[imeasure], alpha = 0.9, lw = 1, dashes = [500,1])
ax.set_ylim([0.0, 0.16])
ax.set_xlim([5, 1.5])
ax.set_xticks([5,3.25, 1.5])
ax.set_yticks([0.0,0.08, 0.16])
ax.set_xlabel('Entropy of B, bits')
ax.set_ylabel('Similarity')
for spine in ['right', 'top']:
ax.spines[spine].set_visible(False)
plt.legend()
plt.show()
```
### Example 1c
In the third example, 1,024 elements are grouped into 8 clusters of equal size and compared against a similar clustering with an increasing number of equal-sized clusters.
```
n_clusterfixed = 2**3
n_cluster_points = 40
number_cluster_values = np.logspace(np.log2(n_clusterfixed), np.log2(n_elements), n_cluster_points, base = 2, dtype = int)
number_cluster_compare = np.zeros((n_measures, n_cluster_points, n_samples))
# loop through samples and print status every 10
for isample in range(n_samples):
if isample % 10 == 0: print(isample)
initial_cluster = clugen.make_equal_clustering(n_elements, n_clusterfixed)
for ipoint in range(n_cluster_points):
random_cluster = clugen.make_equal_clustering(n_elements, number_cluster_values[ipoint])
random_cluster = clugen.shuffle_memberships(random_cluster, percent = 1.0)
# perform clustering similarity measurements
for imeasure in range(n_measures):
if measure_list[imeasure] in ['adj_rand', 'elmsim']:
number_cluster_compare[imeasure, ipoint, isample] = eval(measure_list[imeasure]+'(initial_cluster, random_cluster)')
else:
number_cluster_compare[imeasure, ipoint, isample] = eval('sim.'+measure_list[imeasure]+'(initial_cluster, random_cluster)')
numc_measure_mean = np.mean(number_cluster_compare, axis = 2)
numc_measure_error = np.std(number_cluster_compare, axis = 2)
print("Simulation Finished")
# plot results
fig, ax = plt.subplots(1,1, figsize = (8,8))
ax.set_xscale('log')
for imeasure in range(n_measures):
ax.fill_between(number_cluster_values, numc_measure_mean[imeasure] - numc_measure_error[imeasure],
numc_measure_mean[imeasure] + numc_measure_error[imeasure],
color = BiasColorDict[measure_list[imeasure]], alpha = 0.3)
line = ax.plot(number_cluster_values, numc_measure_mean[imeasure], color = BiasColorDict[measure_list[imeasure]],
label = measure_list[imeasure], alpha = 0.9, lw = 1, dashes = [500,1])
ax.set_ylim([0.0, 0.5])
ax.set_xticks([])
ax.set_yticks([0.0, 0.25, 0.5])
for spine in ['right', 'top']:
ax.spines[spine].set_visible(False)
ax.set_xlabel('Number of clusters in B')
ax.set_ylabel('Similarity')
plt.legend()
plt.show()
```
# Example 2
Evaluating clustering comparisons w.r.t. random models. We illustrate the use of random models by comparing the true classification of cancer types to a clustering derived from Agglomerative Hierarchical Clustering on gene expression data.
```
# Load the Gene Expression Data and ground truth classification
datafile = 'Data/risinger-2003_database.txt'
with open(datafile, 'r') as gefile:
reader = csv.reader(gefile, delimiter="\t")
d = [list(r) for r in reader]
membership = d[1]
membership.pop(0)
clusnames = {c:i for i, c in enumerate(set(membership))}
membership = [clusnames[n] for n in membership]
d = np.array(d[2:])
genetitles = d[:,0]
expressiondata = d[:,1:].astype(np.float)
K = len(clusnames)
ground_truth = Clustering()
ground_truth.from_membership_list(membership)
# find the Hierarchical Clustering
hierclus = GetHierCluster(expressiondata.T, membership, K)
Nsamples = [100, 100**2]
model_list = ['perm', 'num1']
true_value = sim.rand_index(hierclus, ground_truth)
fig, ax = plt.subplots(2,1, sharex = True, figsize = (6, 4))
bins = np.linspace(0.4, 0.7, 41)
titles = ['Permutation Model', 'One-sided Fixed Number of Clusters']
for iax in range(2):
pairwise_comparisons = sim.sample_expected_sim(hierclus, ground_truth, measure = 'rand_index',
random_model = model_list[iax],
n_samples = Nsamples[iax], keep_samples = True)
rand_dist, bins = np.histogram(pairwise_comparisons, bins = bins)
ax[iax].bar(0.5*(bins[:-1] + bins[1:]), rand_dist, width = 0.95*np.abs(bins[:-1] - bins[1:]), alpha = 0.4)
exp_value = sim.expected_rand_index(ground_truth.n_elements, random_model = model_list[iax],
n_clusters1 = hierclus.n_clusters,
n_clusters2 = ground_truth.n_clusters,
clu_size_seq1 = hierclus.clu_size_seq,
clu_size_seq2 = ground_truth.clu_size_seq)
ax[iax].plot([exp_value,exp_value], [0, rand_dist.max()], c = 'k', ls = '--')
ax[iax].plot([true_value,true_value], [0, rand_dist.max()], c = sns.color_palette()[3])
ax[iax].set_title(titles[iax], fontsize = 11)
ax[iax].set_ylabel('# of comparisons', fontsize = 10)
ax[1].set_xlabel('Rand Index', fontsize = 10)
plt.tight_layout()
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/OUCTheoryGroup/colab_demo/blob/master/DSCMR_CVPR2019.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## DSCMR 深度监督跨模态检索
Liangli Zhen, Peng Hu, Xu Wang, Dezhong Peng, Deep Supervised Cross-modal Retrieval. *CVPR 2019*
```
! mkdir pascal
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/test_img.mat
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/test_img_lab.mat
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/test_txt.mat
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/train_img.mat
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/train_img_lab.mat
! wget -P pascal https://raw.githubusercontent.com/penghu-cs/DSCMR/master/data/pascal/train_txt.mat
import torch
from datetime import datetime
import torch.optim as optim
import matplotlib.pyplot as plt
from torch.utils.data.dataset import Dataset
from scipy.io import loadmat, savemat
from torch.utils.data import DataLoader
import numpy as np
import torch.nn as nn
import torch.nn.functional as F
from torchvision import models
import time
import copy
import scipy.spatial
class ImgNN(nn.Module):
"""Network to learn image representations"""
def __init__(self, input_dim=4096, output_dim=1024):
super(ImgNN, self).__init__()
self.denseL1 = nn.Linear(input_dim, output_dim)
def forward(self, x):
out = F.relu(self.denseL1(x))
return out
class TextNN(nn.Module):
"""Network to learn text representations"""
def __init__(self, input_dim=1024, output_dim=1024):
super(TextNN, self).__init__()
self.denseL1 = nn.Linear(input_dim, output_dim)
def forward(self, x):
out = F.relu(self.denseL1(x))
return out
class IDCM_NN(nn.Module):
"""Network to learn text representations"""
def __init__(self, img_input_dim=4096, img_output_dim=2048,
text_input_dim=1024, text_output_dim=2048, minus_one_dim=1024, output_dim=10):
super(IDCM_NN, self).__init__()
self.img_net = ImgNN(img_input_dim, img_output_dim)
self.text_net = TextNN(text_input_dim, text_output_dim)
self.linearLayer = nn.Linear(img_output_dim, minus_one_dim)
self.linearLayer2 = nn.Linear(minus_one_dim, output_dim)
def forward(self, img, text):
view1_feature = self.img_net(img)
view2_feature = self.text_net(text)
view1_feature = self.linearLayer(view1_feature)
view2_feature = self.linearLayer(view2_feature)
view1_predict = self.linearLayer2(view1_feature)
view2_predict = self.linearLayer2(view2_feature)
return view1_feature, view2_feature, view1_predict, view2_predict
class CustomDataSet(Dataset):
def __init__(
self,
images,
texts,
labels):
self.images = images
self.texts = texts
self.labels = labels
def __getitem__(self, index):
img = self.images[index]
text = self.texts[index]
label = self.labels[index]
return img, text, label
def __len__(self):
count = len(self.images)
assert len(
self.images) == len(self.labels)
return count
def ind2vec(ind, N=None):
ind = np.asarray(ind)
if N is None:
N = ind.max() + 1
return np.arange(N) == np.repeat(ind, N, axis=1)
def get_loader(path, batch_size):
img_train = loadmat(path+"train_img.mat")['train_img']
img_test = loadmat(path + "test_img.mat")['test_img']
text_train = loadmat(path+"train_txt.mat")['train_txt']
text_test = loadmat(path + "test_txt.mat")['test_txt']
label_train = loadmat(path+"train_img_lab.mat")['train_img_lab']
label_test = loadmat(path + "test_img_lab.mat")['test_img_lab']
label_train = ind2vec(label_train).astype(int)
label_test = ind2vec(label_test).astype(int)
imgs = {'train': img_train, 'test': img_test}
texts = {'train': text_train, 'test': text_test}
labels = {'train': label_train, 'test': label_test}
dataset = {x: CustomDataSet(images=imgs[x], texts=texts[x], labels=labels[x])
for x in ['train', 'test']}
shuffle = {'train': False, 'test': False}
dataloader = {x: DataLoader(dataset[x], batch_size=batch_size,
shuffle=shuffle[x], num_workers=0) for x in ['train', 'test']}
img_dim = img_train.shape[1]
text_dim = text_train.shape[1]
num_class = label_train.shape[1]
input_data_par = {}
input_data_par['img_test'] = img_test
input_data_par['text_test'] = text_test
input_data_par['label_test'] = label_test
input_data_par['img_train'] = img_train
input_data_par['text_train'] = text_train
input_data_par['label_train'] = label_train
input_data_par['img_dim'] = img_dim
input_data_par['text_dim'] = text_dim
input_data_par['num_class'] = num_class
return dataloader, input_data_par
def train_model(model, data_loaders, optimizer, alpha, beta, device="cpu", num_epochs=500):
since = time.time()
# device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
test_img_acc_history = []
test_txt_acc_history = []
epoch_loss_history =[]
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch+1, num_epochs))
print('-' * 20)
# Each epoch has a training and validation phase
for phase in ['train', 'test']:
if phase == 'train':
# Set model to training mode
model.train()
else:
# Set model to evaluate mode
model.eval()
running_loss = 0.0
running_corrects_img = 0
running_corrects_txt = 0
# Iterate over data.
for imgs, txts, labels in data_loaders[phase]:
# imgs = imgs.to(device)
# txts = txts.to(device)
# labels = labels.to(device)
if torch.sum(imgs != imgs)>1 or torch.sum(txts != txts)>1:
print("Data contains Nan.")
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
# Get model outputs and calculate loss
# Special case for inception because in training it has an auxiliary output. In train
# mode we calculate the loss by summing the final output and the auxiliary output
# but in testing we only consider the final output.
if torch.cuda.is_available():
imgs = imgs.cuda()
txts = txts.cuda()
labels = labels.cuda()
# zero the parameter gradients
optimizer.zero_grad()
# Forward
view1_feature, view2_feature, view1_predict, view2_predict = model(imgs, txts)
loss = calc_loss(view1_feature, view2_feature, view1_predict,
view2_predict, labels, labels, alpha, beta)
img_preds = view1_predict
txt_preds = view2_predict
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item()
running_corrects_img += torch.sum(torch.argmax(img_preds, dim=1) == torch.argmax(labels, dim=1))
running_corrects_txt += torch.sum(torch.argmax(txt_preds, dim=1) == torch.argmax(labels, dim=1))
epoch_loss = running_loss / len(data_loaders[phase].dataset)
# epoch_img_acc = running_corrects_img.double() / len(data_loaders[phase].dataset)
# epoch_txt_acc = running_corrects_txt.double() / len(data_loaders[phase].dataset)
t_imgs, t_txts, t_labels = [], [], []
with torch.no_grad():
for imgs, txts, labels in data_loaders['test']:
if torch.cuda.is_available():
imgs = imgs.cuda()
txts = txts.cuda()
labels = labels.cuda()
t_view1_feature, t_view2_feature, _, _ = model(imgs, txts)
t_imgs.append(t_view1_feature.cpu().numpy())
t_txts.append(t_view2_feature.cpu().numpy())
t_labels.append(labels.cpu().numpy())
t_imgs = np.concatenate(t_imgs)
t_txts = np.concatenate(t_txts)
t_labels = np.concatenate(t_labels).argmax(1)
img2text = fx_calc_map_label(t_imgs, t_txts, t_labels)
txt2img = fx_calc_map_label(t_txts, t_imgs, t_labels)
print('{} Loss: {:.4f} Img2Txt: {:.4f} Txt2Img: {:.4f}'.format(phase, epoch_loss, img2text, txt2img))
# deep copy the model
if phase == 'test' and (img2text + txt2img) / 2. > best_acc:
best_acc = (img2text + txt2img) / 2.
best_model_wts = copy.deepcopy(model.state_dict())
if phase == 'test':
test_img_acc_history.append(img2text)
test_txt_acc_history.append(txt2img)
epoch_loss_history.append(epoch_loss)
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(time_elapsed // 60, time_elapsed % 60))
print('Best average ACC: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
return model, test_img_acc_history, test_txt_acc_history, epoch_loss_history
def calc_loss(view1_feature, view2_feature, view1_predict, view2_predict, labels_1, labels_2, alpha, beta):
term1 = ((view1_predict-labels_1.float())**2).sum(1).sqrt().mean() + ((view2_predict-labels_2.float())**2).sum(1).sqrt().mean()
cos = lambda x, y: x.mm(y.t()) / ((x ** 2).sum(1, keepdim=True).sqrt().mm((y ** 2).sum(1, keepdim=True).sqrt().t())).clamp(min=1e-6) / 2.
theta11 = cos(view1_feature, view1_feature)
theta12 = cos(view1_feature, view2_feature)
theta22 = cos(view2_feature, view2_feature)
Sim11 = calc_label_sim(labels_1, labels_1).float()
Sim12 = calc_label_sim(labels_1, labels_2).float()
Sim22 = calc_label_sim(labels_2, labels_2).float()
term21 = ((1+torch.exp(theta11)).log() - Sim11 * theta11).mean()
term22 = ((1+torch.exp(theta12)).log() - Sim12 * theta12).mean()
term23 = ((1 + torch.exp(theta22)).log() - Sim22 * theta22).mean()
term2 = term21 + term22 + term23
term3 = ((view1_feature - view2_feature)**2).sum(1).sqrt().mean()
im_loss = term1 + alpha * term2 + beta * term3
return im_loss
def calc_label_sim(label_1, label_2):
Sim = label_1.float().mm(label_2.float().t())
return Sim
def fx_calc_map_label(image, text, label, k = 0, dist_method='COS'):
if dist_method == 'L2':
dist = scipy.spatial.distance.cdist(image, text, 'euclidean')
elif dist_method == 'COS':
dist = scipy.spatial.distance.cdist(image, text, 'cosine')
ord = dist.argsort()
numcases = dist.shape[0]
if k == 0:
k = numcases
res = []
for i in range(numcases):
order = ord[i]
p = 0.0
r = 0.0
for j in range(k):
if label[i] == label[order[j]]:
r += 1
p += (r / (j + 1))
if r > 0:
res += [p / r]
else:
res += [0]
return np.mean(res)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# data parameters
DATA_DIR = 'pascal/'
alpha = 1e-3
beta = 1e-1
MAX_EPOCH = 200
batch_size = 100
# batch_size = 512
lr = 1e-4
betas = (0.5, 0.999)
weight_decay = 0
print('...Data loading is beginning...')
data_loader, input_data_par = get_loader(DATA_DIR, batch_size)
print('...Data loading is completed...')
model_ft = IDCM_NN(img_input_dim=input_data_par['img_dim'], text_input_dim=input_data_par['text_dim'], output_dim=input_data_par['num_class']).to(device)
params_to_update = list(model_ft.parameters())
# Observe that all parameters are being optimized
optimizer = optim.Adam(params_to_update, lr=lr, betas=betas)
print('...Training is beginning...')
# Train and evaluate
model_ft, img_acc_hist, txt_acc_hist, loss_hist = train_model(model_ft, data_loader, optimizer, alpha, beta, MAX_EPOCH)
print('...Training is completed...')
print('...Evaluation on testing data...')
view1_feature, view2_feature, view1_predict, view2_predict = model_ft(torch.tensor(input_data_par['img_test']).to(device), torch.tensor(input_data_par['text_test']).to(device))
label = torch.argmax(torch.tensor(input_data_par['label_test']), dim=1)
view1_feature = view1_feature.detach().cpu().numpy()
view2_feature = view2_feature.detach().cpu().numpy()
view1_predict = view1_predict.detach().cpu().numpy()
view2_predict = view2_predict.detach().cpu().numpy()
img_to_txt = fx_calc_map_label(view1_feature, view2_feature, label)
print('...Image to Text MAP = {}'.format(img_to_txt))
txt_to_img = fx_calc_map_label(view2_feature, view1_feature, label)
print('...Text to Image MAP = {}'.format(txt_to_img))
print('...Average MAP = {}'.format(((img_to_txt + txt_to_img) / 2.)))
```
| github_jupyter |
# Gradient Descent
$
\begin{eqnarray}
x_{i+1}&=&x_{t} - \eta \frac{\partial F}{\partial x} \\
x_{t+1} - x_{t}&=& -\eta \frac{\partial F}{\partial x} \\
\Delta x &=& -\eta \frac{\partial F}{\partial x}
\end{eqnarray}
$
```
%matplotlib inline
import numpy as np
import matplotlib.cm as cm
from matplotlib import pyplot as plt
plt.style.use('ggplot')
x=[]
yt=[]
for i in range(100):
x.append( [0.5+np.random.rand(), 0.5+np.random.rand()])
x.append( [-0.5+np.random.rand(), -0.5+np.random.rand()])
yt.append([0 , 1])
yt.append([0 , 1])
for i in range(100):
x.append( [0.5+np.random.rand(), -0.5+np.random.rand()])
x.append( [-0.5+np.random.rand(), 0.5+np.random.rand()])
yt.append([1 , 0])
yt.append([1 , 0])
x=np.array(x)
yt=np.array(yt)
plt.plot( x[yt[:,1]==1,0], x[yt[:,1]==1,1], 'ob')
plt.plot( x[yt[:,0]==1,0], x[yt[:,0]==1,1], 'or')
```
# Single Layer Perceptron
## Error
$
\begin{eqnarray}
E&=&\frac{1}{2} (y_p-y_t)^2\\
\end{eqnarray}
$
## Input and output
$
\begin{eqnarray}
a_0 &\Leftarrow &X\\
y_p& \Leftarrow &a_2\\
\end{eqnarray}
$
## Forward Network
$
\begin{eqnarray}
z_0&=&a_0.w_0+b_0\\
a_1&=&g(z_0)\\
z_1&=&a_1.w_1+b_1\\
a_2&=&g(z_1)\\
\end{eqnarray}
$
# Backpropagation on w0
$
\begin{eqnarray}
\Delta w_0 &=& -\eta \frac{\partial E}{\partial w_0}\\
\frac{\partial E}{\partial w_0} &=& \frac{\partial E}{\partial y_p} \frac{\partial y_p}{\partial a_2} \frac{\partial a_2}{\partial z_1} \frac{\partial z_1}{\partial a_1} \frac{\partial a_1}{\partial z_0} \frac{\partial z_0}{\partial w_0} \\
\frac{\partial E}{\partial y_p} &=& y_p-y_t \\
\frac{\partial y_p}{\partial a_2} &=& 1 \\
\frac{\partial a_2}{\partial z_1} &=& \frac{\partial g(z_1)}{\partial z_1} \\
\frac{\partial z_1}{\partial a_1} &=& w_1 \\
\frac{\partial a_1}{\partial z_0} &=& \frac{\partial g(z_0)}{\partial z_0} \\
\frac{\partial z_0}{\partial w_0} &=& a_0 \\
\end{eqnarray}
$
```
def g(x):
return 1/(1+np.exp(-x))
def grad_g(g):
return (1-g)*g
#random init weight and bias
np.random.seed(1)
N0=4 #first layer 4 nodes
N1=2 #second layer 2 nodes
eta=0.01
a0=np.ones([x.shape[0],x.shape[1]+1]) #[Nx3]
a0[:,:-1]=x
w0 = np.random.random((2,N0)) # 2 inputs x 4 nodes
b0 = np.random.random((1,N0)) # 1 bias x 4 nodes
wb0=np.concatenate((w0, b0), axis=0)
a1=np.ones([x.shape[0],N0+1])
w1 = np.random.random((N0,N1)) # 4 inputs x 2 nodes
b1 = np.random.random((1,N1)) # 1 bias x 2 nodes
wb1=np.concatenate((w1, b1), axis=0)
for i in range(10000):
#forward x=a0, a2=yp
z0=np.dot(a0,wb0)
a1[:,:-1]=g(z0)
#a1=np.concatenate([g(z0),np.ones([a0.shape[0],1])], axis=1)
z1=np.dot(a1,wb1)
a2=g(z1)
#backward
d_a2=yt-a2
d_z1=d_a2*grad_g(a2)
wb1 += eta*np.dot(a1.T,d_z1)
d_a1=np.dot(d_z1,wb1.T)
d_z0=d_a1*grad_g(a1)
wb0 += eta*np.dot(a0.T,d_z0[:,:-1])
if(i % 1000) == 0: # Only print the error every 10000 steps
E=0.5*np.sum(np.square(d_a2))
print("Error: {}".format(E))
my,mx=np.mgrid[slice(-0.6,1.6,0.01),slice(-0.6,1.6,0.01)]
out=np.zeros(mx.shape)
for i in range(mx.shape[0]):
for j in range(mx.shape[1]):
u=[ mx[i,j], my[i,j],1]
#forward
ha1=np.concatenate( [ g(np.dot(u,wb0)), [1]])
ha2=g(np.dot(ha1,wb1))
out[i,j]=ha2[1]-ha2[0]
plt.pcolor(mx,my,out,cmap=cm.RdYlBu)
plt.colorbar()
plt.plot( x[yt[:,1]==1,0], x[yt[:,1]==1,1], 'ob')
plt.plot( x[yt[:,0]==1,0], x[yt[:,0]==1,1], 'or')
wb0.shape
```
| github_jupyter |
# Milky Way CGM X-ray Absorption Spectrum
Figure 8.2 from Chapter 8 of *Interstellar and Intergalactic Medium* by Ryden & Pogge, 2021,
Cambridge University Press.
Plot of detection of OVII and OVIII Lyman-$\alpha$ x-ray absorption lines from the warm-hot and hot
phases of the Milky Way circumgalactic medium (CGM) along the line-of-sight to the x-ray bright quasar
1ES1553+113 obtained using the XMM-Newton X-ray Telescope and the Reflection Grating Spectrometer (RGS)
instrument.
Data from [Das et al. 2019, ApJL, 882, L23](https://ui.adsabs.harvard.edu/abs/2019ApJ...882L..23D),
Figure 2, replotted using data provided by Sanskriti Das to just show the OVII and OVIII absorption lines.
```
%matplotlib inline
import math
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator, LogLocator, NullFormatter
import warnings
warnings.filterwarnings('ignore',category=UserWarning, append=True)
warnings.filterwarnings('ignore',category=RuntimeWarning, append=True)
```
## Standard Plot Format
Setup the standard plotting format and make the plot. Fonts and resolution adopted follow CUP style.
```
figName = 'Fig8_2'
# graphic aspect ratio = width/height
aspect = 2.5
# Text width in inches - don't change, this is defined by the print layout
textWidth = 6.0 # inches
# output format and resolution
figFmt = 'png'
dpi = 600
# Graphic dimensions
plotWidth = dpi*textWidth
plotHeight = plotWidth/aspect
axisFontSize = 10
labelFontSize = 8
lwidth = 0.5
axisPad = 5
wInches = textWidth
hInches = wInches/aspect
# Plot filename
plotFile = f'{figName}.{figFmt}'
# LaTeX is used throughout for markup of symbols, Times-Roman serif font
plt.rc('text', usetex=True)
plt.rc('font', **{'family':'serif','serif':['Times-Roman'],'weight':'bold','size':'16'})
# Font and line weight defaults for axes
matplotlib.rc('axes',linewidth=lwidth)
matplotlib.rcParams.update({'font.size':axisFontSize})
# axis and label padding
plt.rcParams['xtick.major.pad'] = f'{axisPad}'
plt.rcParams['ytick.major.pad'] = f'{axisPad}'
plt.rcParams['axes.labelpad'] = f'{axisPad}'
```
## Convenience function
Define a function to compute a Gaussian emission-line profile given the central wavelength,
full-width at half maximum (FWHM), peak intensity, for an array of wavelengths, lam.
Not pretty, not meant to be.
```
def gauss(lam,cen,fwhm,pk):
sig = fwhm/(2.0*math.sqrt(2.0*math.log(2)))
arg = (lam-cen)*(lam-cen)/(2*sig*sig)
return pk*np.exp(-arg)
```
## Spectral Data
Data are in a 8-column ASCII format, of which we extract 6 columns:
* wavelength - wavelength in Angstroms, range 8-30
* err_wavelength - wavelength error in Angstroms
* flux - flux in Jansky (10$^{-26}$ erg s$^{-1}$ cm$^{-2}$ Hz$^{-1}$)
* err_flux - flux error in Jansky
* model - best fit model
* continuum - continuum model component
We will plot the fluxes normalized relative to the best fit continuum, as is conventional for absorption-line
studies at all wavelengths.
```
# RGS1 data
dataFile = 'Das2019_RGS1.txt'
data = pd.read_csv(dataFile,sep=r'\s+',comment='#')
wave1 = np.array(data['wavelength'])
errWave1 = np.array(data['err_wavelength'])
flux1 = np.array(data['flux'])
errFlux1 = np.array(data['err_flux'])
fluxMod1 = np.array(data['model'])
fluxCont1 = np.array(data['continuum'])
# normalized flux and errors
fluxNorm1 = flux1/fluxCont1
modNorm1 = fluxMod1/fluxCont1
errNorm1 = errFlux1/fluxCont1
# Plotting Limits
xMin1 = 21.4 # Angstroms - OVII 21.2
xMax1 = 21.8
# RGS2 data
dataFile = 'Das2019_RGS2.txt'
data = pd.read_csv(dataFile,sep=r'\s+',comment='#')
wave2 = np.array(data['wavelength'])
errWave2 = np.array(data['err_wavelength'])
flux2 = np.array(data['flux'])
errFlux2 = np.array(data['err_flux'])
fluxMod2 = np.array(data['model'])
fluxCont2 = np.array(data['continuum'])
# normalized flux and errors
fluxNorm2 = flux2/fluxCont2
modNorm2 = fluxMod2/fluxCont2
errNorm2 = errFlux2/fluxCont2
# Plotting Limits
xMin2 = 18.76 # Angstroms - OVIII 18.96
xMax2 = 19.16
# common Y limits
yMin = 0.7 # normalized flux
yMax = 1.2 #
```
## Make the plot
```
fig,ax = plt.subplots()
fig.set_dpi(dpi)
fig.set_size_inches(wInches,hInches,forward=True)
fig.subplots_adjust(wspace=0.2, hspace=0)
# Left panel: OVII, RGS1 only
ax1 = plt.subplot(121)
ax1.set_xlim(xMin1,xMax1)
ax1.set_ylim(yMin,yMax)
ax1.tick_params('both',length=6,width=lwidth,which='major',direction='in',top='on',right='on')
ax1.tick_params('both',length=3,width=lwidth,which='minor',direction='in',top='on',right='on')
ax1.xaxis.set_major_locator(MultipleLocator(0.1))
ax1.xaxis.set_minor_locator(MultipleLocator(0.05))
ax1.yaxis.set_major_locator(MultipleLocator(0.1))
ax1.yaxis.set_minor_locator(MultipleLocator(0.05))
plt.xlabel(r'Wavelength [\AA]', fontsize=axisFontSize)
plt.ylabel(r'Normalized flux',fontsize=axisFontSize)
plt.errorbar(wave1,fluxNorm1,xerr=errWave1,yerr=errNorm1,color='black',fmt='o',ms=1,zorder=10,lw=0.5)
plt.plot(wave1,modNorm1,'-',ds='steps-mid',color='black',lw=0.5)
plt.text(0.5*(xMin1+xMax1),1.125,r'O{\sc vii}',ha='center',va='center',fontsize=axisFontSize)
# Right Panel: OVIII, RGS1 and 2 data
ax2 = plt.subplot(122)
ax2.tick_params('both',length=6,width=lwidth,which='major',direction='in',top='on',right='on')
ax2.tick_params('both',length=3,width=lwidth,which='minor',direction='in',top='on',right='on')
ax2.set_xlim(xMin2,xMax2)
ax2.set_ylim(yMin,yMax)
ax2.xaxis.set_major_locator(MultipleLocator(0.1))
ax2.xaxis.set_minor_locator(MultipleLocator(0.05))
plt.xlabel(r'Wavelength [\AA]', fontsize=axisFontSize)
ax2.yaxis.set_major_locator(MultipleLocator(0.1))
ax2.yaxis.set_minor_locator(MultipleLocator(0.05))
plt.errorbar(wave1,fluxNorm1,xerr=errWave1,yerr=errNorm1,color='black',fmt='o',ms=1,zorder=10,lw=0.5)
plt.errorbar(wave2,fluxNorm2,xerr=errWave2,yerr=errNorm2,color='#666666',fmt='o',ms=1,zorder=10,lw=0.5)
plt.plot(wave1,modNorm1,'-',ds='steps-mid',color='black',lw=0.5)
plt.text(0.5*(xMin2+xMax2),1.125,r'O{\sc viii}',ha='center',va='center',fontsize=axisFontSize)
plt.plot()
plt.savefig(plotFile,bbox_inches='tight',facecolor='white')
```
| github_jupyter |
# Comparing Two Climate Models
In this notebook, I will be comparing two climate reanalysis models:
* NCEP-DOE Reanalysis 2: Surface
* ERA5
I will be looking at the following variables:
* Surface Pressure
* Mean Sea Level Pressure
* Total Column Water
The idea is simple: these two models should have very similar properties. I will be trying to user RBIG in order to assess how similar these models are. I'll be looking at the following IT measures
* Entropy
* Total Correlation
* Mutual Information
If these climate models are that similar, then they should exhibit similar IT measures.
## Data - Climate Models
```
import os, sys
cwd = os.getcwd()
source_path = f"{cwd}/../../../"
sys.path.insert(0, f'{source_path}')
import numpy as np
# Data Loaders
from src.data.climate.amip import DataDownloader
from src.data.climate.amip import DataLoader
from src.features.climate.build_features import get_time_overlap
# ESDC tools
sys.path.insert(0, f'/home/emmanuel/code/py_esdc')
from esdc.standardize import normalize_temporal
from esdc.transform import regrid_data
from esdc.utils import check_time_coords
import pandas as pd
import xarray as xr
from tqdm import tqdm
from sklearn import preprocessing
import seaborn as sns
import matplotlib.pyplot as plt
plt.style.use('ggplot')
%matplotlib inline
%load_ext autoreload
%autoreload 2
amip_data_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/amip/"
era5_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/era5/"
ncep_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/raw/ncep/"
results_path = f"/home/emmanuel/projects/2020_rbig_rs/data/climate/results/"
fig_path = f"/home/emmanuel/projects/2020_rbig_rs/reports/figures/climate/"
```
### Dataset - CMIP5
```
loader = DataLoader()
dataset = 'ipsl_cm5b_lr'
cmip5_data = loader.load_amip_data(dataset)
cmip5_data
```
#### ERA5
```
era5_data = xr.open_mfdataset(f"{era5_path}*.nc", combine="by_coords")
era5_data = era5_data.rename({'msl': 'mslp', 'latitude': 'lat', 'longitude': 'lon'})
# era5_data = era5_data.rename({'latitude': 'lat'})
# era5_data.attrs['model_id'] = 'era5'
# rescale model from 0.25 to 2.5 degrees
# era5_data = era5_data.coarsen(lat=10, lon=10, boundary='pad').mean()
era5_data.attrs['model_id'] = 'era5'
era5_data = era5_data.rename({'mslp': 'psl'})
era5_data
```
## Time Coords Overlap
```
era5_data, cmip5_data = get_time_overlap(era5_data, cmip5_data)
era5_data
cmip5_data
```
## NCEP
```
ncep_data = xr.open_mfdataset(f"{ncep_path}*mon.mean.nc", combine="by_coords")
ncep_data = ncep_data.rename({'mslp': 'psl'})
ncep_data.attrs['model_id'] = 'ncar_ncep_doe_2'
ncep_data
```
#### CMIP5
```
loader = DataLoader()
dataset = 'ipsl_cm5b_lr'
cmip5_data = loader.load_amip_data(dataset)
cmip5_data
ncep_data, cmip5_data = get_time_overlap(ncep_data, cmip5_data)
ncep_data
cmip5_data
```
| github_jupyter |
# Introduction to the enDAQ library
## Introduction
This notebook and accompanying webinar was developed and released by the [enDAQ team](https://endaq.com/). This is the fourth "chapter" of our series on *Python for Mechanical Engineers*:
1. [Get Started with Python](https://colab.research.google.com/drive/1_pcGtgJleapV9tz5WfuRuqfWryjqhPHy#scrollTo=ikUJITDDIp19)
* Blog: [Get Started with Python: Why and How Mechanical Engineers Should Make the Switch](https://blog.endaq.com/get-started-with-python-why-how-mechanical-engineers-should-make-the-switch)
2. [Introduction to Numpy & Pandas for Data Analysis](https://colab.research.google.com/drive/1O-VwAdRoSlcrineAk0Jkd_fcw7mFGHa4#scrollTo=ce97q1ZcBiwj)
3. [Introduction to Plotly for Plotting Data](https://colab.research.google.com/drive/1pag2pKQQW5amWgRykAH8uMAPqHA2yUfU)
4. **Introduction of the enDAQ Library**
* [Watch Recording of This](https://info.endaq.com/simplify-shock-and-vibration-analysis-with-endaq-python-library)
5. [More Custom Examples](https://colab.research.google.com/drive/1cuZa5Yx55qXLhhnBMdzsJ0iiklwVi5Mq)
To sign up for future webinars and watch previous ones, [visit our webinars page](https://endaq.com/pages/shock-vibration-webinars).
## Why Did We Develop This?
When analyzing data you'll need:
- Customize a bit to meet your specific need
- Share the results
- Share the *methodology*
- Reproduce the analysis for future/other data sets
Writing scripts that produce highly interactive and custom plots addresses all of these needs. And that's why we created the open source enDAQ library - to make analysis more convenient, adaptable and reliable!
## Docs
All of our functions are documented at www.docs.endaq.com

The code itself lives on github at: https://github.com/MideTechnology/endaq-python
## Installation
Installing is as easy as `pip install endaq` that is needed once when running locally, but everytime in Google Colab.
```
!pip install -q endaq
exit() #needed in Colab because they pre-load some libraries
import endaq
import plotly.express as px
import plotly.io as pio; pio.renderers.default = "iframe"
import plotly.graph_objects as go
import pandas as pd
import numpy as np
import scipy
```
## Story 1: PSD of Large Time Series
### Accessing Data
This first uses a function `endaq.ide.get_doc()` ([see docs](https://docs.endaq.com/en/latest/endaq/ide.html#endaq.ide.get_doc)) to load in an IDE file. This can accept locations locally, or hosted online.
Please note that Python won't accept backslashes and there are a number of ways around this (libraries!). I typically add an 'r' before the string of the folder like so `r"C:\Users\shanly"+"\\"` which ends up becoming: `'C:\\Users\\shanly\\'`. For more [see Python's docs](https://docs.python.org/3/reference/lexical_analysis.html#string-and-bytes-literals).
```
doc = endaq.ide.get_doc('https://info.endaq.com/hubfs/data/Sys034_1.ide')
```
There are a lot of elements to this object that you can explore with `doc.__dict__` but my favorite is to grab the serial number and part number.
```
print(doc.recorderInfo['PartNumber'])
print(doc.recorderInfo['RecorderSerial'])
```
Once the file is loaded, we are using the `endaq.ide.get_channel_table()` ([see docs](https://docs.endaq.com/en/latest/endaq/ide.html#endaq.ide.get_channel_table)) which will present the contents of the file.
This file is modestly sized at 140 MB.
```
table = endaq.ide.get_channel_table(doc)
table
```
If you need to parse this `table` dataframe to sort it, find channels of interest, etc., you'll need to grab the data like so.
```
table.data
```
Ok, now that we have IDE file, let's get the data out! This is using the `endaq.ide.to_pandas()` ([see docs](https://docs.endaq.com/en/latest/endaq/ide.html#endaq.ide.to_pandas)) to pull out every channel into a dictionary with keys of the channel name.
```
data = {doc.channels[ch].name: endaq.ide.to_pandas(doc.channels[ch], time_mode='datetime') for ch in doc.channels}
data
```
### Dashboard of an IDE File
We can pass this into a plot function to display a dashboard! Here are [the docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.dashboards.rolling_enveloped_dashboard), this accepts a dictionary of dataframes to display a bunch of sub plots for every channel/column.
```
fig = endaq.plot.dashboards.rolling_enveloped_dashboard(
data,
desired_num_points=100,
min_points_to_plot=1,
plot_as_bars=True,
)
fig.show()
```
### New Plotting Theme
We've developed four Plotly themes (although really just two):
1. endaq
2. endaq_light
3. endaq_cloud (Open Sans Font)
4. endaq_cloud_light (Open Sans Font)
The `set_theme()` function creates the above four themes and makes one the default.
This uses a helper function, `define_theme()` which we recommend to those that may want to develop their own theme. [Here are the docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.utilities.define_theme).
```
endaq.plot.utilities.set_theme()
fig.update_layout(
template='endaq'
)
```
Remember these are plotly figures which can be saved as interactive HTML files.
```
fig.write_html('dashboard.html',include_plotlyjs ='cdn')
```
This function allows a lot of customization with how the dashboard is displayed.
```
endaq.plot.dashboards.rolling_enveloped_dashboard(
data,
desired_num_points=100,
min_points_to_plot=1,
plot_as_bars=False,
num_rows=1,
num_cols=None
)
```
This dashboard can be used on any collection of dataframes, not just from an IDE file.
```
csv_data = {
'crash':pd.read_csv('https://info.endaq.com/hubfs/data/Motorcycle-Car-Crash.csv',index_col=0),
'moto':pd.read_csv('https://info.endaq.com/hubfs/data/motorcycle-vibration-moving-frequency.csv',index_col=0),
'instrument':pd.read_csv('https://info.endaq.com/hubfs/data/surgical-instrument.csv',index_col=0),
'rocket':pd.read_csv('https://info.endaq.com/hubfs/data/blushift.csv',index_col=0),
'calibration':pd.read_csv('https://info.endaq.com/hubfs/data/Calibration-Shake.csv',index_col=0),
'baseball':pd.read_csv('https://info.endaq.com/hubfs/data/baseball-throw-acceleration.csv',index_col=0, header=None, names=['X','Y','Z']),
'volleyball':pd.read_csv('https://info.endaq.com/hubfs/data/volleyball-hit-acceleration.csv',index_col=0, header=None, names=['X','Y','Z']),
'football':pd.read_csv('https://info.endaq.com/hubfs/data/football-catch-acceleration.csv',index_col=0, header=None, names=['X','Y','Z']),
}
endaq.plot.dashboards.rolling_enveloped_dashboard(
csv_data,
desired_num_points=100,
min_points_to_plot=1,
plot_as_bars=True,
)
```
We also have a related function to compute some rolling metrics, not just the envelope. Here I will plot the rolling peak and standard deviation (effectively the RMS). [See the docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.dashboards.rolling_metric_dashboard).
```
endaq.plot.dashboards.rolling_metric_dashboard(
csv_data,
desired_num_points=100,
rolling_metrics_to_plot = ('absolute max', 'std')
)
```
No let's go back to that big dataset and look at our dataframes within it.
```
data.keys()
data['100g PE Acceleration']
```
If you want to skip this step we just went through (load all data, present dashboard) and you know which channel is of interest, you can directly load in that data with `endaq.ide.to_pandas()` ([see docs](https://docs.endaq.com/en/latest/endaq/ide.html#endaq.ide.to_pandas)).
This defaults to present the time with datetime objects which is helpful for synchronization. You can pass in `time_mode='seconds'` to just get the time as seconds if preferred.
```
accel = endaq.ide.to_pandas(doc.channels[8])
accel
```
### High Pass Filter
If you noticed, the DC offset on the piezoelectric accelerometer was a bit wonky which should be filtered away. Even if there isn't an egregious DC offset like this example, it is still recommended to apply this filter when doing vibration analysis.
This uses our `endaq.calc.filters.butterworth()` ([see docs](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.filters.butterworth)) to filter intuitively.
```
filtered = endaq.calc.filters.butterworth(accel, low_cutoff=2)
filtered
```
### Plotting Large Time Series
Now here's a function similar to the dashboard above that was based off a dictionary of dataframes with each subchannel/column of data getting its own plot.
Here in `rolling_min_max_envelope()` ([see docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.rolling_min_max_envelope)) though we plot a single dataframe's data on one plot. When using `plot_as_bars` this view will appear identical to loading ALL of the data and plotting it, yet this operation is completed in <10 seconds and will be highly responsive.
We are going to be making a lot of acceleration vs time plots, so I am going to simplify the labeling.
```
accel_time_labels = dict(
xaxis_title_text='',
yaxis_title_text='Acceleration (g)',
legend_title_text=''
)
fig = endaq.plot.plots.rolling_min_max_envelope(
filtered,
desired_num_points=1000,
plot_as_bars=True,
opacity=0.7
)
fig.update_layout(
accel_time_labels,
title_text='Filtered Time Series with 13M Points',
)
```
### Linear PSD
Now we have the data, let's generate a PSD on the whole thing with `psd.welch()`([see docs](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.psd.welch)), and add in the resultant (PSDs are squared, so the resultant is simply the sum).
```
psd = endaq.calc.psd.welch(filtered, bin_width=1)
psd['Resultant'] = psd.sum(axis=1)
psd
```
Remember this is a pandas dataframe, saving to csv is easy with `to_csv()` (or other file type, [see docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html))
We're going to make a lot of PSDs so let's make the labeling easy.
```
psd_labels = dict(
xaxis_title_text='Frequency (Hz)',
yaxis_title_text='Acceleration (g^2/Hz)',
legend_title_text='',
xaxis_type='log',
yaxis_type='log',
)
```
Now let's plot it in Plotly!
```
fig = px.line(psd)
fig.update_layout(
psd_labels,
title_text='Power Spectral Density',
)
```
### Log PSD
Once a linear PSD is computed, we have a function to convert it to octave spacing, [see docs](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.psd.to_octave).
```
oct_psd = endaq.calc.psd.to_octave(psd, fstart=4, octave_bins=3)
oct_psd.head()
```
Using Plotly graph_objects, I'll add these lines to the existing plot.
```
for c in oct_psd.columns:
fig.add_trace(go.Scattergl(
x=oct_psd.index,
y=oct_psd[c],
name=c+' Octave',
line_width=6,
line_dash='dash'
))
fig.show()
```
### Spectrogram
Our spectrogram function ([see docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.octave_spectrogram)) allows for octave spaced frequency bins which drastically reduces the heatmap resolution needed and is arguably a better way to represent the data anyways. This is a spectrogram generated off 13M points and completed in 3 seconds.
```
freqs, bins, Pxx, fig = endaq.plot.octave_spectrogram(filtered[['X (100g)']], window=12, bins_per_octave=6)
fig.show()
```
### Extracting a Subsection of IDE File
With long files there may be subsections we'd like to pull out to save and share. This function does just that, ([see docs](https://docs.endaq.com/en/latest/endaq/ide.html)).
```
endaq.ide.extract_time(doc,
out='extracted.ide',
start=pd.to_datetime('2021-10-22 09:17:15'),
end=pd.to_datetime('2021-10-22 09:17:25'))
```
Within Python and with dataframes, remember we can easily "slice" the data to focus on areas of interest. Here I'll generate a PSD on a 10 second period of a fixed operating state.
```
psd = endaq.calc.psd.welch(filtered['2021-10-22 09:17:15':'2021-10-22 09:17:25'], bin_width=1)
psd['Resultant'] = psd.sum(axis=1)
fig = px.line(psd)
fig.update_layout(
psd_labels,
title_text='Power Spectral Density from 9:17:15 to 9:17:25',
)
```
I knew that to be a particularly interesting time in the file because of the light data this user utilized... clever!
```
fig = px.line(data['Light Sensor'][['Lux']][::4])
fig.update_layout(
showlegend=False,
xaxis_title_text='',
yaxis_title_text="Light (Lux)"
)
```
## Story 2: Multiple Files
We did a quick test with 3 devices on a shaker.

I know I only care about the secondary accelerometer, so I can load that channel (80) directly on these **THREE** files.
```
data = {
'Shaker' : endaq.ide.to_pandas(endaq.ide.get_doc('https://info.endaq.com/hubfs/data/Transfer_Shaker.ide').channels[80]),
'Long Beam' : endaq.ide.to_pandas(endaq.ide.get_doc('https://info.endaq.com/hubfs/data/Transfer_Long_Beam.ide').channels[80]),
'Short Beam' : endaq.ide.to_pandas(endaq.ide.get_doc('https://info.endaq.com/hubfs/data/Transfer_Short_Beam.ide').channels[80])
}
data
```
### Dashboard
With these three files, let's generate that dashboard again and compare them all in one figure (with shared x axes).
```
fig = endaq.plot.dashboards.rolling_enveloped_dashboard(
data,
desired_num_points=500,
min_points_to_plot=1,
plot_as_bars=True,
)
fig.update_xaxes(matches='x')
fig.show()
```
### Single Plot Comparison
We may want to compare them all in one plot, here I'll combine just the Z axis from each file.
```
time_overall = pd.DataFrame()
for device in data.keys():
#Get Y axis and filter
accel = endaq.calc.filters.butterworth(data[device]['Z (40g)'].to_frame(), low_cutoff=2)
#Rename the column as our test/device name
accel.columns = [device]
#Combine Times
time_overall = pd.concat([time_overall,accel])
time_overall
```
There is still 40,000 data points, so I'll use the rolling_envelope plot to simplify the plot.
```
fig = endaq.plot.plots.rolling_min_max_envelope(
time_overall,
desired_num_points=1000,
plot_as_bars=True,
opacity=0.7
)
fig.update_layout(
accel_time_labels,
title_text='Comparison of Acceleration in Time Domain',
)
fig.show()
```
### PSD Comparison
Now let's compute a PSD for all of these, and combine into one dataframe by rounding to a shared frequency bin.
```
psd_overall = pd.DataFrame()
for device in data.keys():
#Get Z axis and filter
accel = endaq.calc.filters.butterworth(data[device]['Z (40g)'].to_frame(), low_cutoff=2)
#Get PSD
psd = endaq.calc.psd.welch(accel, bin_width = 0.5)
#Round to the nearest 0.5 Hz
psd.index = np.round(psd.index*2,0)/2
#Rename the PSD column as our test/device name
psd.columns = [device]
#Combine PSDs
psd_overall = pd.concat([psd_overall,psd], axis=1)
psd_overall
fig = px.line(psd_overall[4:500])
fig.update_layout(
psd_labels,
title_text='PSD Comparison',
)
```
### Octave PSD
```
oct_psd = endaq.calc.psd.to_octave(psd_overall, fstart=4, octave_bins = 0.5)
fig = px.line(oct_psd['Shaker'][:256])
fig.update_layout(
psd_labels,
title_text='PSD Comparison',
template='endaq_light'
)
```
### Transfer Function
I can use the shaker data to compute a transfer function for the other devices on different length beams. This is accomplished by taking the square root of the ratio from the beam PSD to the shaker.
```
transfer = psd_overall[5:240].copy().drop('Shaker',axis=1)
for col in transfer.columns:
transfer[col] = (psd_overall[col]/psd_overall['Shaker']) ** 0.5
transfer
```
Now we can plot it to see the natural frequencies of these two beams.
```
fig = px.line(transfer)
fig.update_layout(
title_text='Transfer Function',
xaxis_title_text='Frequency (Hz)',
yaxis_title_text='Transfer (g/g)',
legend_title_text='',
xaxis_type='log',
yaxis_type='log',
)
```
### Integrate to Displacement
We also have a function to integrate to velocity and displacement which is done here ([see docs](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.integrate.integrals)).
```
displacements = {}
for device in data.keys():
#Get Z axis and filter
accel = data[device]['Z (40g)'].to_frame()
#Double Integrate to Displacement
[df_accel, df_vel] = endaq.calc.integrate.integrals(accel, n=1, highpass_cutoff=2, tukey_percent=0.2)
[df_vel, df_disp] = endaq.calc.integrate.integrals(df_vel, n=1, highpass_cutoff=2, tukey_percent=0.2)
df_disp = df_disp*9.81*39.37 #convert to in
#Rename the column as our test/device name
df_disp.columns = [device]
#Combine Times
displacements[device] = df_disp
displacements
```
### Resample at Slower Rate
Displacement is dominated by lower frequency content, so we can reduce the amount of data by resampling at 200 Hz with another function.
```
displacement = pd.DataFrame()
for device in displacements.keys():
#Resample at 100 Hz
disp_resampled = endaq.calc.utils.resample(displacements[device], sample_rate = 200)
#Rename the index name to help with "melting"
disp_resampled.index.name = 'index'
#Combine Into One DataFrame
displacement = pd.concat([displacement, disp_resampled.reset_index().melt(id_vars='index').dropna()])
displacement
fig = px.line(displacement,
x='index',
y='value',
color='variable')
fig.update_layout(
title_text = 'Z Axis Displacement Resampled at 200 Hz',
yaxis_title_text = 'Displacement (in)',
xaxis_title_text = '',
legend_title_text = '',
)
```
Synchronization is quite impressive if I do say so myself!! Maybe off by about 0.003 seconds?
## Story 3: Shock Event
Now let's analyze some data that was discussed in our [blog on the shock response spectrum](https://blog.endaq.com/shock-analysis-response-spectrum-srs-pseudo-velocity-severity).
```
doc = endaq.ide.get_doc('https://info.endaq.com/hubfs/data/Motorcycle-Car-Crash.ide')
accel = endaq.ide.to_pandas(doc.channels[8])
accel
```
### Plot at Peak Time
The `around_peak()` function ([see docs](https://docs.endaq.com/en/latest/endaq/plot.html#endaq.plot.around_peak)) takes in a dataframe and plots around the peak value across all columns.
```
fig = endaq.plot.plots.around_peak(
accel,
num=1500,
leading_ratio=0.4
)
fig.update_layout(
accel_time_labels,
title_text='Acceleration Around Peak',
)
```
### Low Pass Filter
Now let's apply a bunch of low-pass filters ([see docs](https://docs.endaq.com/en/latest/endaq/calc.html#module-endaq.calc.filters)).
```
accel = accel['Y (500g)'].to_frame() #keep as a dataframe
accel = (accel - accel.median()) * -1 #apply a high pass and make the shock positive acceleration
accel.columns = ['No Low-Pass']
freqs = [1600, 800, 400, 200, 100, 50]
for freq in freqs:
name = 'Filtered at: '+str(freq)+' Hz'
accel[name] = endaq.calc.filters.butterworth(accel['No Low-Pass'].to_frame(), high_cutoff=freq)
accel = accel['2019-07-03 17:05:08.4':'2019-07-03 17:05:08.55'] #isolate the time of interest
accel = accel - accel.iloc[0] #force start to 0 to remove any filtering artifact
accel
fig = px.line(accel)
fig.update_layout(
accel_time_labels,
title_text='Motorcycle Car Crash, Effect of Filtering'
)
```
### Shock Response Spectrum
First we need to define which frequencies we want to calculate and plot the SRS for.
```
freqs = endaq.calc.utils.logfreqs(accel, init_freq=1, bins_per_octave=12)
```
Now we can calculate the shock response spectrum ([see docs](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.shock.shock_spectrum)).
```
srs = endaq.calc.shock.shock_spectrum(accel, freqs=freqs, damp=0.05, mode='srs')
fig = px.line(srs)
fig.update_layout(
title_text='Shock Response Spectrum (SRS)',
xaxis_title_text="Natural Frequency (Hz)",
yaxis_title_text="Peak Acceleration (g)",
legend_title_text='',
xaxis_type="log",
yaxis_type="log",
)
```
### Pseudo Velocity Shock Spectrum
That function also allows us to calculate the PVSS.
```
pvss = endaq.calc.shock.shock_spectrum(accel, freqs=freqs, damp=0.05, mode='pvss')
pvss = pvss*9.81*39.37 #convert to in/s
fig = px.line(pvss)
fig.update_layout(
title_text='Psuedo Velocity Shock Spectrum (PVSS)',
xaxis_title_text="Natural Frequency (Hz)",
yaxis_title_text="Psuedo Velocity (in/s)",
legend_title_text='',
xaxis_type="log",
yaxis_type="log",
)
```
#### Enveloped Half Sine
Once a PVSS is calculated we have a function to find a half-sine pulse that would envelop that PVSS (this can make reproducing the event easy with a drop test). [See the docs for more](https://docs.endaq.com/en/latest/endaq/calc.html#endaq.calc.shock.enveloping_half_sine).
```
t = np.linspace(0,0.1,num=1000) # NOTE: if there aren't enough samples, low-frequency artifacts will appear!
def half_sine_pulse(t, T):
result = np.zeros_like(t)
result[(t > 0) & (t < T)] = np.sin(np.pi*t[(t > 0) & (t < T)] / T)
df_result = pd.DataFrame({'Time':t,
'Pulse':result}).set_index('Time')
return df_result
for c in ['No Low-Pass', 'Filtered at: 200 Hz']:
half_sine_params = endaq.calc.shock.enveloping_half_sine(pvss[c]/9.81/39.37, damp=0.05)
pvss_pulse = endaq.calc.shock.shock_spectrum(half_sine_params[0] * half_sine_pulse(t, T=half_sine_params[1]),
freqs=freqs, damp=0.05, mode='pvss')*9.81*39.37
fig.add_trace(go.Scattergl(
x=pvss_pulse.index,
y=pvss_pulse[pvss_pulse.columns[0]].values,
name='Half Sine of '+c+': '+str(np.round(half_sine_params[0],1))+'g, '+ str(np.round(half_sine_params[1],5)) +'s',
line_width=6,
line_dash='dot'
))
fig.show()
```
#### Impact of Damping
```
pvss_damping = pd.DataFrame()
damps = [0, 0.025, 0.05, 0.10]
for damp in damps:
pvss = endaq.calc.shock.shock_spectrum(accel['No Low-Pass'].to_frame(), freqs=freqs, damp=damp, mode='pvss')
name = 'Damping Ratio = '+str(damp)
pvss_damping[name] = pvss[pvss.columns[0]]*9.81*39.37 #convert to in/s
fig = px.line(pvss_damping)
fig.update_layout(
title_text='Impact of Damping on PVSS',
xaxis_title_text="Natural Frequency (Hz)",
yaxis_title_text="Psuedo Velocity (in/s)",
legend_title_text='',
xaxis_type="log",
yaxis_type="log",
)
```
### Integration to Velocity
```
[df_accel, df_vel] = endaq.calc.integrate.integrals(accel, n=1, highpass_cutoff=None, tukey_percent=0)
df_vel = df_vel-df_vel.iloc[0] #forced the starting velocity to 0
df_vel = df_vel*9.81*39.37 #convert to in/s
fig = px.line(df_vel)
fig.update_layout(
title_text="Integrated Velocity Time History",
xaxis_title_text="",
yaxis_title_text="Velocity (in/s)",
legend_title_text=''
)
fig.show()
```
## Story 4: Moving Frequency
```
df_vibe = pd.read_csv('https://info.endaq.com/hubfs/data/motorcycle-vibration-moving-frequency.csv',index_col=0)
df_vibe = df_vibe - df_vibe.median()
df_vibe
```
### Shaded Plot in Time
```
fig = endaq.plot.plots.rolling_min_max_envelope(
df_vibe,
desired_num_points=1000,
plot_as_bars=True,
opacity=1.0
)
fig.update_layout(
accel_time_labels,
title_text='Engine Reving of Motorcycle',
)
fig.show()
```
### Spectrogram
```
freqs, bins, Pxx, fig = endaq.plot.octave_spectrogram(df_vibe[['Z (40g)']],
window=0.5,
bins_per_octave=12,
max_freq=1000,
freq_start=40)
fig.show()
```
### Plot of Frequency vs Time
```
df_Pxx = pd.DataFrame(Pxx, index= freqs, columns = bins)
df_Pxx = 10 ** (df_Pxx/10)
fig = px.line(df_Pxx[df_Pxx.index<500].idxmax())
fig.update_layout(
title_text="Moving Peak Frequency",
xaxis_title_text="",
yaxis_title_text="Peak Frequency (Hz)",
showlegend=False
)
fig.show()
```
### PSD Animation over Time
Now we can prepare for creating an animation of the PSD over time.
```
df_Pxx.index.name='Frequency (Hz)'
df_Pxx.columns.name = 'Time (s)'
df_Pxx.columns = np.round(df_Pxx.columns,2)
```
Create base figure with animation, this will render fine but we'll want to add the other lines for scaling and reference.
```
fig = px.line(
df_Pxx.reset_index().melt(id_vars='Frequency (Hz)'),
x='Frequency (Hz)',
y='value',
animation_frame='Time (s)',
log_x=True,
log_y=True,
)
```
Add in the max, min, median, and mean lines.
```
def add_line(df_stat,name,dash,color):
fig.add_trace(go.Scattergl(
x=df_stat.index,
y=df_stat.values,
name=name,
line_width=3,
line_dash=dash,
line_color=color
))
#Add max, min, median
for stat,dash,quant in zip(['Max','Min','Median'],
['dash','dash','dot'],
[1.0,0.0,0.5]):
df_stat = df_Pxx.quantile(quant, axis=1)
add_line(df_stat,stat,dash,'#6914F0')
#Add in mean
df_stat = df_Pxx.mean(axis=1)
add_line(df_stat,'Mean','dot','#2DB473')
fig.update_layout(
legend_y=-0.7,
yaxis_title_text='Acceleration (g^2/Hz)'
)
fig.show()
```
### Compare to Overall PSD
```
psd = endaq.calc.psd.welch(df_vibe[['Z (40g)']], bin_width=0.25)
oct_psd = endaq.calc.psd.to_octave(psd,octave_bins=12,fstart=40)
fig = px.line(oct_psd[oct_psd.index<=1000])
fig.update_layout(
psd_labels
)
```
## Story 5: Sound Data
```
mic = endaq.ide.to_pandas(endaq.ide.get_doc('https://info.endaq.com/hubfs/data/sound/gangnam-style.ide').channels[8],time_mode='seconds')
mic
mic = mic*-5.3075 #convert to Pa, this will be natively done with devices starting in the next month or so
fig = endaq.plot.plots.rolling_min_max_envelope(
mic,
desired_num_points=1000,
plot_as_bars=True,
opacity=1.0
)
fig.update_layout(
title_text='Gangnam Style, Then Fan',
yaxis_title_text='Sound Level (Pa)',
xaxis_title_text='',
showlegend=False
)
fig.show()
```
### Save .WAV File
```
import scipy.io.wavfile
mic_normalized = mic.copy()
mic_normalized /= np.max(np.abs(mic_normalized),axis=0)
scipy.io.wavfile.write(
'sound.wav',
int(np.round(1/endaq.calc.utils.sample_spacing(mic))),
mic_normalized.values.astype(np.float32),
)
```
### Play in Notebook
```
import IPython
IPython.display.display(IPython.display.Audio('sound.wav'))
```
### Convert to dB
```
n = int(mic.shape[0]/100)
rolling_pa = mic.rolling(n).std()[::n]
rolling_dB = rolling_pa.apply(endaq.calc.utils.to_dB, reference=endaq.calc.utils.dB_refs["SPL"], raw=True)
fig = px.line(rolling_dB)
fig.update_layout(
title_text='Gangnam Style, Then Fan Sound Level',
yaxis_title_text='Sound Level (dB)',
xaxis_title_text='',
showlegend=False
)
fig.show()
```
### dB vs Frequency
```
df_pascal_psd = endaq.calc.psd.welch(mic, bin_width=1)
df_pascal_octave = endaq.calc.psd.to_octave(df_pascal_psd*df_pascal_psd.index[1],agg="sum",octave_bins=3, fstart=10)
df_audio_psd_dB = df_pascal_octave.apply(endaq.calc.utils.to_dB,
reference=endaq.calc.utils.dB_refs["SPL"]**2,
squared=True,
raw=True)
fig = px.line(df_audio_psd_dB)
fig.update_layout(
title_text='Sound Level vs Frequency',
xaxis_title_text='Frequency (Hz)',
xaxis_type='log',
yaxis_title_text='Sound Level (dB)',
showlegend=False
)
```
### Spectrogram
```
freqs, bins, Pxx, fig = endaq.plot.octave_spectrogram(mic, window=0.5, bins_per_octave=12, freq_start=40, max_freq=5000)
fig.show()
```
## enDAQ Cloud as an Alternative
Our enDAQ cloud ([cloud.endaq.com](https://cloud.endaq.com/)) offers an environment to generate interactive dashboards for free without the need to write Python code.
But what is especially unique is that our cloud also allows paying tiers (starting at $100/month) to customize these dashboards with code to accelerate the analysis cycle and allow deploying customizing dashboard generation to colleagues and customers without ever needing to install anything.
Here is an example that has it's own [unique URL](https://cloud.endaq.com/user/dashboard/4dff0e96-1594-4cef-a364-2476ad1fb99a) for sharing with colleagues (requires log-in).
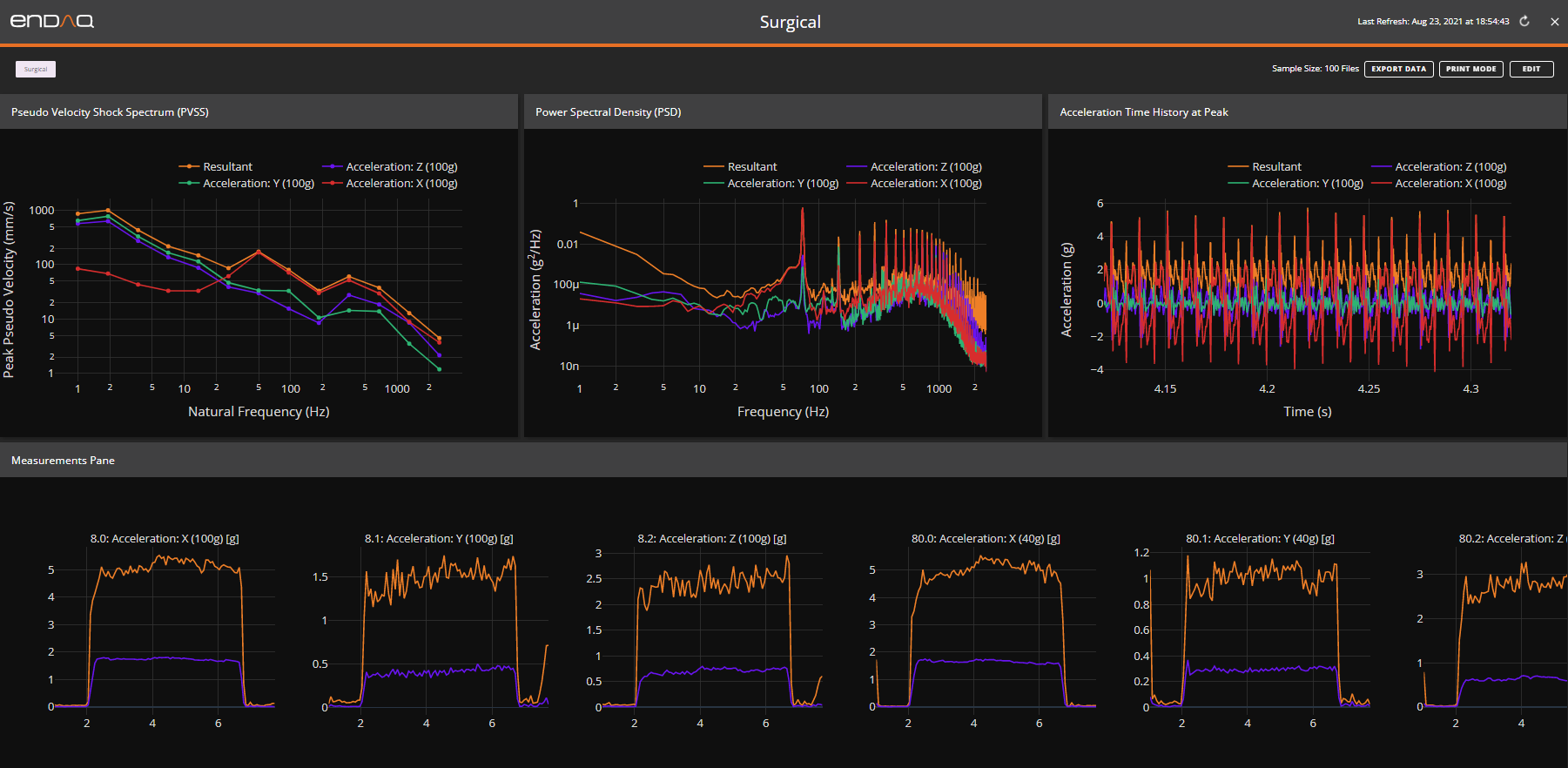
For more on generating these custom reports see our [Help Article](https://support.endaq.com/article/317-dynamic-reports-in-the-cloud).
## What's Coming Next?
More webinars and more functionality!
1. User Requested Examples
2. Release of endaq.batch for Batch Processing
3. Updating enDAQ Cloud to Provide Access to New Python Library
| github_jupyter |
<a href="https://colab.research.google.com/github/raahatg21/Movie-Recommender-System/blob/master/content_recommender.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Content based Recommender System of Movies
Content-based recommenders suggest similar items based on a particular item. This system uses item metadata, such as genre, director, description, actors, etc. for movies, to make these recommendations. The general idea behind these recommender systems is that if a person liked a particular item, he or she will also like an item that is similar to it.
We use the dataset of all movies in IMDB (derived by MovieLens 20M Dataset).
## Plot Description based Recommender
### Workflow
- Select the 'overview' of each movie
- Convert it into TF-IDF vectors and construct TF-IDF matrix
- Calculate the similarity score using cosine similarity
- User inputs a movie title
- We find similarity scores of all movies w.r.t. that title
- Sort them
- Select the 10 most similar movies to input movie
```
import pandas as pd
import numpy as np
import os
# Mounting Google Drive
from google.colab import drive
drive.mount('/content/drive')
BASE_DIR = '/content/drive/My Drive/Recommender System/the-movies-dataset'
metadata_path = os.path.join(BASE_DIR, 'movies_metadata.csv')
credits_path = os.path.join(BASE_DIR, 'credits.csv')
keywords_path = os.path.join(BASE_DIR, 'keywords.csv')
# Load the movies metadata
metadata = pd.read_csv(metadata_path, low_memory = False)
metadata.shape
metadata.head(5)
# Let's look at movie overviews
metadata['overview'].head()
```
Since the entire data is too large, we'll only be using the top 10% popular movies (movies with at least 160 votes)
```
m = metadata['vote_count'].quantile(0.90)
q_metadata = metadata.copy().loc[metadata['vote_count'] >= m]
q_metadata = q_metadata.reset_index()
q_metadata.shape
q_metadata.head()
```
That's better!
```
q_metadata['overview'] = q_metadata['overview'].fillna('')
# Now, processing this text using TF-IDF Vectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf = TfidfVectorizer(stop_words = 'english') # initialising the TF-IDF Vector object
tfidf_matrix = tfidf.fit_transform(q_metadata['overview']) # Constructing the TF-IDF Matrix (no. of movies x every word in vocabulary)
tfidf_matrix.shape
# Computing the cosine similarity
from sklearn.metrics.pairwise import linear_kernel
cosine_sim = linear_kernel(tfidf_matrix, tfidf_matrix) # Constructing the Cosine Similarity Matrix (no. of movies x no. of movies)
cosine_sim.shape
# Construct a reverse map of indices and titles
indices = pd.Series(q_metadata.index, index = q_metadata['title']).drop_duplicates()
# Function that inputs movie titles and outputs top 10 movies similar to it
def get_recommendations(title, cosine_sim = cosine_sim):
idx = indices[title]
sim_scores = list(enumerate(cosine_sim[idx])) # Get the similarity scores of all movies wrt input movie
sim_scores = sorted(sim_scores, key = lambda x : x[1], reverse = True)
sim_scores = sim_scores[1:11]
movie_indices = [i[0] for i in sim_scores]
return q_metadata['title'].iloc[movie_indices]
get_recommendations('The Dark Knight Rises')
get_recommendations('Titanic')
```
While our system has done a decent job of finding movies with similar plot descriptions, the quality of recommendations is not that great. "The Dark Knight Rises" returns all Batman movies while it more likely that the people who liked that movie are more inclined to enjoy other Christopher Nolan movies.
So fix this, we incorporate information about the director, actors and genre of the movie into our model
## Credits, Genres and Keywords based Recommender
### Workflow
- Get the credits and keywords for each movie from separate csv files and merge them into the current dataframe
- Convert them from 'stringified' lists to a from that is more suitable to us
- Clean the data, convert them into lower case, and remove spaces
- Join these features to create a 'metadata soup'
- Convert this soup into vectorised format using Count Vectorizer
- Calculate cosine similarity from the Count matrix
- Use the existing `get_recommendations()` function to get movie recommendations using this cosine similarity
```
credits = pd.read_csv(credits_path)
keywords = pd.read_csv(keywords_path)
# Convert the ids to int and merge the dataframes
keywords['id'] = keywords['id'].astype('int')
credits['id'] = credits['id'].astype('int')
q_metadata['id'] = q_metadata['id'].astype('int')
q_metadata = q_metadata.merge(credits, on='id')
q_metadata = q_metadata.merge(keywords, on='id')
q_metadata.shape
q_metadata.head()
features = ['cast', 'crew', 'keywords', 'genres']
features_d = ['cast', 'keywords', 'genres']
# Parse the stringified features into usable python objects
from ast import literal_eval
for feature in features:
q_metadata[feature] = q_metadata[feature].apply(literal_eval)
q_metadata.head()
# Extracting the director's name
def get_director(x):
for i in x:
if i['job'] == 'Director':
return str(i['name'])
return np.nan
# Extracting top 3 members of any list
def get_list(x):
if isinstance(x, list): # if x is a list
names = [str(i['name']) for i in x]
if len(names) > 3:
names = names[:3]
#names = list(names)
return names
return []
q_metadata['director'] = q_metadata['crew'].apply(get_director)
for f in features_d:
q_metadata[f] = q_metadata[f].apply(get_list)
q_metadata[['title', 'cast', 'director', 'keywords', 'genres']].head()
def clean_data(x):
if isinstance(x, list): # if x is a list
try:
return [str.lower(i.replace(' ', '')) for i in x]
except:
return ''
else:
if isinstance(x, str): # if x is name of director
return str.lower(x.replace(' ', ''))
else:
return ''
for feature in features:
q_metadata[feature] = q_metadata[feature].apply(clean_data)
q_metadata[['title', 'cast', 'director', 'keywords', 'genres']].head()
# Creating the metadata soup
def create_soup(x):
return ' '.join(x['keywords']) + ' ' + ' '.join(x['cast']) + ' ' + x['director'] + ' ' + ' '.join(x['genres'])
q_metadata['soup'] = q_metadata.apply(create_soup, axis = 1)
q_metadata['soup'].head()
# Using Count Vectorizer and creating the count matrix
from sklearn.feature_extraction.text import CountVectorizer
count = CountVectorizer(stop_words = 'english')
count_matrix = count.fit_transform(q_metadata['soup'])
count_matrix.shape
from sklearn.metrics.pairwise import cosine_similarity
cosine_sim2 = cosine_similarity(count_matrix, count_matrix)
cosine_sim2.shape
q_metadata = q_metadata.reset_index()
indices = pd.Series(q_metadata.index, index = q_metadata['title'])
get_recommendations('The Dark Knight Rises', cosine_sim2)
get_recommendations('John Wick', cosine_sim2)
```
| github_jupyter |
# Introduction: What is Logistic Regression?
Logistic regression is a supervised learning classification algorithm used to predict the probability of a target variable. The nature of target or dependent variable is dichotomous, which means there would be only two possible classes.
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
import seaborn as sns
```
## Behind The Mask
The term **logistic** in logistic regression is used because we are applying another function to the weighted sum of input data and parameters of the model and this function is called logit (**sigmoid**) function.
Sigmoid function always outputs values between 0 and 1, mapping values to a range and thus can be used to calculate probabilities of input data belonging to a certain class:
<img src="https://miro.medium.com/max/271/1*Gp5E23P5d2PY5D5kOo8ePw.png" width="200" height="100">
<img src="https://miro.medium.com/max/1280/1*OUOB_YF41M-O4GgZH_F2rw.png" width="400" height="200">
```
def sigmoid(x):
% ====================== YOUR CODE HERE ======================================================
% Instructions: Compute the sigmoid of each value of x (x can be a matrix, vector or scalar).
% ============================================================================================
```
## Performance Measure
Weights (represented by theta in our notation) is a vital part of Logistic Regression and other Machine Learning algorithms and we want to find the best values for them. To start we pick random values and we need a way to measure how well the algorithm performs using those random weights. That measure is computed using the **cost function**.
The cost function is defined as:
<img src="https://miro.medium.com/max/1400/1*2g14OVjyJqio2zXwJxgj2w.png" width="600" height="400">
The above two functions can be compressed into a single function i.e.
<img src="https://miro.medium.com/max/1400/1*_52kKSp8zWgVTNtnE2eYrg.png" width="600" height="400">
```
def compute_cost(X, y, theta):
# Initialize some useful values
m = length(y) % number of training examples
% ====================== YOUR CODE HERE ==========================
% Instructions: Compute the cost of a particular choice of theta.
%
%
% ================================================================
```
## Finding Optimal Parameters
The goal is to minimize the cost by means of increasing or decreasing the weights.
This can be done with a function called **Gradient Descent**. <br>
Gradient descent is just the **derivative of the cost function with respect to its weights**.
Now to minimize our cost function we need to run the gradient descent function on each parameter i.e.
<img src="https://miro.medium.com/max/245/1*1--MUhjPjOL7oYdVo7R6gQ.png">
Where: α = **learning rate** (usually 0.1)
This is implemented as follows : <br>
<img src="https://miro.medium.com/max/1400/1*Ecea3jVIRxK4Mkrh_Nie4w.jpeg" width="500" height="300">
```
def gradient_descent(X, y, theta, learning_rate, iterations):
# Initialize some useful values
m = len(y)
cost_history = np.zeros((iterations,1))
% ====================== YOUR CODE HERE ==============================================
% Instructions: Write a for loop that runs the parameter "iterations" number of times.
In each iteration do the following -
> Update theta according to the formula shown above
> Compute cost for current theta using compute_cost() and enter
it into the array cost_history at that iteration index.
%
% Return (cost_history, theta)
%
%
%
%
% ====================================================================================
```
## Predicting Classes
Now let's write the **prediction function**. <br>
Since we are dealing with probabilities here, if the resulting value is above 0.50, we round it up to 1, meaning the data sample belongs to the class 1. Consequently, if the probability of a data sample belonging to the class 1 is below 0.50, it simply means that it is part of the other class (class 0).<br>
Remember that this is binary classification, so we have only two classes (class 1 and class 0).
```
def predict(X, theta):
# Initialize some useful values
m = size(X, 1); % Number of training examples
% ====================== YOUR CODE HERE =================================
% Instructions: Compute the predictions for X using a threshold at 0.5
(i.e., if sigmoid(theta.T * x) >= 0.5, predict 1)
% =======================================================================
```
## Dataset Generation
After writing the code for the necessary functions, let’s create our very own dataset with **make_classification** function from **sklearn.datasets**. <br>
We will create **500 sample points** with **two classes** and plot the dataset with the help of seaborn library.
```
X, y = make_classification(n_samples=500, n_features=2, n_redundant=0, n_informative=1,
n_clusters_per_class=1, random_state=14)
y = y[:,np.newaxis]
sns.set_style('white')
sns.scatterplot(X[:,0],X[:,1],hue=y.reshape(-1));
```
## Model Testing
Finally, here is the algorithm for implementation, that we built for you using the functions you created above.
Now, let's run it to calculate the parameters of our model.
```
m = len(y)
X = np.hstack((np.ones((m,1)),X))
n = np.size(X,1)
theta = np.zeros((n,1))
iterations = 1500
learning_rate = 0.03
initial_cost = compute_cost(X, y, theta)
print("Initial Cost is: {} \n".format(initial_cost))
(cost_history, params_optimal) = gradient_descent(X, y, params, learning_rate, iterations)
print("Optimal Parameters are: \n", params_optimal, "\n")
```
Now we will plot the cost as a function of number of iterations.
```
plt.figure()
sns.set_style('white')
plt.plot(range(len(cost_history)), cost_history, 'r')
plt.title("Convergence Graph of Cost Function")
plt.xlabel("Number of Iterations")
plt.ylabel("Cost")
plt.show()
```
- Does the cost decrease with every iteration?<br>
- Does it converge to a minimum value?
## Model Accuracy
After running the algorithm and getting the optimal parameters, we want to know how successful our model is at predicting the classes of our data.
```
y_pred = predict(X, params_optimal)
score = float(sum(y_pred == y))/ float(len(y))
print(score)
```
What accuracy score did you achieve?!
## Dataset Plotting with Decision Boundary
Now, for the sake of visualization, let’s plot our dataset along with the **decision boundary** of our model.<br>
We simply calculate the intercept and slope values using the optimal parameters and plot the boundary that classifies the data into two classes.
```
slope = -(params_optimal[1] / params_optimal[2])
intercept = -(params_optimal[0] / params_optimal[2])
sns.set_style('white')
sns.scatterplot(X[:,1],X[:,2],hue=y.reshape(-1));
ax = plt.gca()
ax.autoscale(False)
x_vals = np.array(ax.get_xlim())
y_vals = intercept + (slope * x_vals)
plt.plot(x_vals, y_vals, c="k");
```
Is the classification 100% correct?
If not, how many misclassifications are present?
### Congratulations! You just successfully implemented Logistic regression from scratch!
| github_jupyter |
# Basic core
This module contains all the basic functions we need in other modules of the fastai library (split with [`torch_core`](/torch_core.html#torch_core) that contains the ones requiring pytorch). Its documentation can easily be skipped at a first read, unless you want to know what a given function does.
```
from fastai.gen_doc.nbdoc import *
from fastai.core import *
```
## Global constants
`default_cpus = min(16, num_cpus())` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/core.py#L45">[source]</a></div>
## Check functions
```
show_doc(has_arg)
```
Examples for two [`fastai.core`](/core.html#core) functions. Docstring shown before calling [`has_arg`](/core.html#has_arg) for reference
```
has_arg(download_url,'url')
has_arg(index_row,'x')
has_arg(index_row,'a')
show_doc(ifnone)
param,alt_param = None,5
ifnone(param,alt_param)
param,alt_param = None,[1,2,3]
ifnone(param,alt_param)
show_doc(is1d)
two_d_array = np.arange(12).reshape(6,2)
print( two_d_array )
print( is1d(two_d_array) )
is1d(two_d_array.flatten())
show_doc(is_listy)
```
Check if `x` is a `Collection`. `Tuple` or `List` qualify
```
some_data = [1,2,3]
is_listy(some_data)
some_data = (1,2,3)
is_listy(some_data)
some_data = 1024
print( is_listy(some_data) )
print( is_listy( [some_data] ) )
some_data = dict([('a',1),('b',2),('c',3)])
print( some_data )
print( some_data.keys() )
print( is_listy(some_data) )
print( is_listy(some_data.keys()) )
print( is_listy(list(some_data.keys())) )
show_doc(is_tuple)
```
Check if `x` is a `tuple`.
```
print( is_tuple( [1,2,3] ) )
print( is_tuple( (1,2,3) ) )
```
## Collection related functions
```
show_doc(arange_of)
arange_of([5,6,7])
type(arange_of([5,6,7]))
show_doc(array)
array([1,2,3])
```
Note that after we call the generator, we do not reset. So the [`array`](/core.html#array) call has 5 less entries than it would if we ran from the start of the generator.
```
def data_gen():
i = 100.01
while i<200:
yield i
i += 1.
ex_data_gen = data_gen()
for _ in range(5):
print(next(ex_data_gen))
array(ex_data_gen)
ex_data_gen_int = data_gen()
array(ex_data_gen_int,dtype=int) #Cast output to int array
show_doc(arrays_split)
data_a = np.arange(15)
data_b = np.arange(15)[::-1]
mask_a = (data_a > 10)
print(data_a)
print(data_b)
print(mask_a)
arrays_split(mask_a,data_a)
np.vstack([data_a,data_b]).transpose().shape
arrays_split(mask_a,np.vstack([data_a,data_b]).transpose()) #must match on dimension 0
show_doc(chunks)
```
You can transform a `Collection` into an `Iterable` of 'n' sized chunks by calling [`chunks`](/core.html#chunks):
```
data = [0,1,2,3,4,5,6,7,8,9]
for chunk in chunks(data, 2):
print(chunk)
for chunk in chunks(data, 3):
print(chunk)
show_doc(df_names_to_idx)
ex_df = pd.DataFrame.from_dict({"a":[1,1,1],"b":[2,2,2]})
print(ex_df)
df_names_to_idx('b',ex_df)
show_doc(extract_kwargs)
key_word_args = {"a":2,"some_list":[1,2,3],"param":'mean'}
key_word_args
(extracted_val,remainder) = extract_kwargs(['param'],key_word_args)
print( extracted_val,remainder )
show_doc(idx_dict)
idx_dict(['a','b','c'])
show_doc(index_row)
data = [0,1,2,3,4,5,6,7,8,9]
index_row(data,4)
index_row(pd.Series(data),7)
data_df = pd.DataFrame([data[::-1],data]).transpose()
data_df
index_row(data_df,7)
show_doc(listify)
to_match = np.arange(12)
listify('a',to_match)
listify('a',5)
listify(77.1,3)
listify( (1,2,3) )
listify((1,2,3),('a','b','c'))
show_doc(random_split)
```
Splitting is done here with `random.uniform()` so you may not get the exact split percentage for small data sets
```
data = np.arange(20).reshape(10,2)
data.tolist()
random_split(0.20,data.tolist())
random_split(0.20,pd.DataFrame(data))
show_doc(range_of)
range_of([5,4,3])
range_of(np.arange(10)[::-1])
show_doc(series2cat)
data_df = pd.DataFrame.from_dict({"a":[1,1,1,2,2,2],"b":['f','e','f','g','g','g']})
data_df
data_df['b']
series2cat(data_df,'b')
data_df['b']
series2cat(data_df,'a')
data_df['a']
show_doc(split_kwargs_by_func)
key_word_args = {'url':'http://fast.ai','dest':'./','new_var':[1,2,3],'testvalue':42}
split_kwargs_by_func(key_word_args,download_url)
show_doc(to_int)
to_int(3.1415)
data = [1.2,3.4,7.25]
to_int(data)
show_doc(uniqueify)
uniqueify( pd.Series(data=['a','a','b','b','f','g']) )
```
## Files management and downloads
```
show_doc(download_url)
show_doc(find_classes)
show_doc(join_path)
show_doc(join_paths)
show_doc(loadtxt_str)
show_doc(save_texts)
```
## Multiprocessing
```
show_doc(num_cpus)
show_doc(parallel)
show_doc(partition)
show_doc(partition_by_cores)
```
## Data block API
```
show_doc(ItemBase, title_level=3)
```
All items used in fastai should subclass this. Must have a [`data`](/tabular.data.html#tabular.data) field that will be used when collating in mini-batches.
```
show_doc(ItemBase.apply_tfms)
show_doc(ItemBase.show)
```
The default behavior is to set the string representation of this object as title of `ax`.
```
show_doc(Category, title_level=3)
```
Create a [`Category`](/core.html#Category) with an `obj` of index [`data`](/tabular.data.html#tabular.data) in a certain classes list.
```
show_doc(EmptyLabel, title_level=3)
show_doc(MultiCategory, title_level=3)
```
Create a [`MultiCategory`](/core.html#MultiCategory) with an `obj` that is a collection of labels. [`data`](/tabular.data.html#tabular.data) corresponds to the one-hot encoded labels and `raw` is a list of associated string.
```
show_doc(FloatItem)
```
## Others
```
show_doc(camel2snake)
camel2snake('DeviceDataLoader')
show_doc(even_mults)
show_doc(func_args)
show_doc(noop)
```
Return `x`.
```
show_doc(one_hot)
show_doc(show_some)
show_doc(subplots)
show_doc(text2html_table)
```
## Undocumented Methods - Methods moved below this line will intentionally be hidden
## New Methods - Please document or move to the undocumented section
| github_jupyter |
# Bayesian optimization with `skopt`
(based on scikit-optimize documentation https://scikit-optimize.github.io/notebooks/sklearn-gridsearchcv-replacement.html)
```
# sklearn version fixed to avoid known skopt issue
!pip install scikit-optimize scikit-learn==0.20.3
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
from skopt import BayesSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
```
## Optimising a RandomForest classifier
```
from sklearn.datasets import load_digits
X, y = load_digits(10, True)
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.75,
random_state=0)
```
dimensions for parameters [list, shape=(n_dims,)]: List of search space dimensions. Each search dimension can be defined either as
- a (lower_bound, upper_bound) tuple (for Real or Integer dimensions),
- a (lower_bound, upper_bound, prior) tuple (for Real dimensions),
- as a list of categories (for Categorical dimensions), or
- an instance of a Dimension object (Real, Integer or Categorical).
```
param_dist = {
"max_depth": (3, 10,),
"max_features": (1, 11),
"min_samples_split": <YOUR CODE>, # from 2 to 10
"min_samples_leaf": <YOUR CODE>, # from 1 to 10
"bootstrap": [True, False], # categorical valued parameter
"criterion": <YOUR CODE> # either "gini" or "entropy"
}
clf = RandomForestClassifier(n_estimators=20)
opt = BayesSearchCV(clf, param_dist, n_iter=10, return_train_score=True, cv=3)
opt.fit(X_train, y_train);
print("val. score: %s" % opt.best_score_)
print("test score: %s" % opt.score(X_test, y_test))
# Utility function to report best scores
import pandas as pd
def report(results, n_top=3):
res = pd.DataFrame(results)
res = res.sort_values(by=['mean_test_score'], ascending=False, axis=0)
res.reset_index(inplace = True, drop=True)
# a = res[['mean_test_score', 'std_test_score']]
for candidate in range(0, n_top):
print("Model with rank: {0}".format(candidate))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
res['mean_test_score'][candidate],
res['std_test_score'][candidate]))
print("Parameters: {0}".format(res['params'][candidate]))
print("")
report(opt.cv_results_)
```
## Neural Network
Optimise the neural net from the previous notebook via `BayesSearchCV`
```
import torch
from torch import nn
import torch.nn.functional as F
from skorch import NeuralNetClassifier
torch.manual_seed(0);
from sklearn.datasets import make_classification
X, y = make_classification(1000, 20, n_informative=10, n_classes=2, random_state=0)
X = X.astype(np.float32)
class ClassifierModule(nn.Module):
<CODE OF THE CLASSIFIER FROM NOTEBOOK i-1>
net = NeuralNetClassifier(
ClassifierModule,
max_epochs=20,
lr=0.1,
device='cuda', # comment this to train with CPU
optimizer__momentum=0.9,
verbose=0
)
```
Define a space for parameter sampling in the form of dict, list of dict or list of tuple containing (dict, int). One of these cases:
1. dictionary, where keys are parameter names (strings) and values are skopt.space.Dimension instances (Real, Integer or Categorical) or any other valid value that defines skopt dimension (see skopt.Optimizer docs). Represents search space over parameters of the provided estimator.
2. list of dictionaries: a list of dictionaries, where every dictionary fits the description given in case 1 above. If a list of dictionary objects is given, then the search is performed sequentially for every parameter space with maximum number of evaluations set to self.n_iter.
3. list of (dict, int > 0): an extension of case 2 above, where first element of every tuple is a dictionary representing some search subspace, similarly as in case 2, and second element is a number of iterations that will be spent optimizing over this subspace.
(see [skopt docs](https://scikit-optimize.github.io/#skopt.BayesSearchCV) for details)
```
params = {
'lr': [0.05, 0.1],
'module__num_units': [10, 20, 30], # range from 10 to 30
'module__dropout': [0.1, 0.3], # range from 0.1 to 0.3
'optimizer__nesterov': [False, True],
}
bs = BayesSearchCV(net, params, refit=False, cv=3, scoring='accuracy',
verbose=0, n_jobs=1, n_iter=10, return_train_score=True)
bs.fit(X, y);
report(bs.cv_results_)
```
## Task
add `optimizer__momentum` to the space with [0.5, 1.5] range
```
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
from sklearn.metrics import classification_report, accuracy_score, confusion_matrix
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
import statsmodels.api as sm
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
from sklearn.model_selection import KFold
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
from pylab import rcParams
rcParams['figure.figsize'] = 14, 8
RANDOM_SEED = 42
LABELS = ["Subscribe", "Not_Subscribe"]
from imblearn.under_sampling import NearMiss
from collections import Counter
from imblearn.combine import SMOTETomek
data = pd.read_csv(r"E:\ExcelR Assignment\Assignment 6 - Logistic Regression\bank-full.csv",sep=';')
data.head()
```
### DATA ANALYSIS
```
# Univaraite Analysis
data.describe()
data.isnull().sum()
```
### No NULL Values
```
data.groupby('y').mean()
data['y'].value_counts()
No = len(data[data.y=='no'])
Yes = len(data[data.y=='yes'])
Yes_Percent = (Yes/len(data['y']))*100
No_Percent = (No/len(data['y']))*100
print('Percent of Yes: ' + str(Yes_Percent))
print('Percent of No: ' + str(No_Percent))
count_classes = pd.value_counts(data['y'], sort = True)
count_classes.plot(kind = 'bar', rot=0)
plt.title("Term Deposit Subscription")
plt.xticks(range(2), LABELS)
plt.xlabel("y")
plt.ylabel("Frequency")
```
#### So, this data has many NO compared to YES.
#### Therefore, data is biased to NO and most of the time answer would be no despite it could have been yes.
```
Subscribe = data[data['y']=='yes']
Not_Subscribe = data[data['y']=='no']
print(Subscribe.shape,Not_Subscribe.shape)
```
## Checking behaviour of variables with target variable
#### Age vs Target Variable
```
plt.figure(figsize=(20,8))
sns.countplot(x="age",data=data,hue="y")
```
#### Job - Default
```
plt.figure(figsize=(15,8))
sns.countplot(x="job",data=data,hue="default")
```
#### Job - Personal Loan
```
plt.figure(figsize=(15,8))
sns.countplot(x="job",data=data,hue="loan")
```
#### Job - Housing Loan
```
plt.figure(figsize=(15,8))
sns.countplot(x="job",data=data,hue="housing")
```
### From the above we can say that most of job category are not default,most of them have not taken personal and housing loan.
#### Job - Target Variable
```
plt.figure(figsize=(15,8))
sns.countplot(x="job",data=data,hue="y")
```
### *It is also clear that most of the people with job haven't subscribed to term deposit*
#### Marital - Default
```
sns.countplot(x="marital",data=data,hue="default")
```
#### Marital - Housing Loan
```
sns.countplot(x="marital",data=data,hue="housing")
```
#### Marital - Personal Loan
```
sns.countplot(x="marital",data=data,hue="loan")
```
### From the above we can say that marital class most of them are not default, have housing and personal loan
#### Marital - Target Variable
```
sns.countplot(x="marital",data=data,hue="y")
```
### *It is also clear that mostly all marital class haven't subscribed to term deposit*
#### Education - Default
```
sns.countplot(x="education",data=data,hue="default")
```
#### Education - Housing
```
sns.countplot(x="education",data=data,hue="housing")
```
#### Education - Personal Loan
```
sns.countplot(x="education",data=data,hue="loan")
```
### From the above we can say that education are not default, also primary and secondary class has housing and personal loan
#### Education - Target Variable
```
plt.figure(figsize=(14,8))
sns.countplot(x="education",data=data,hue="y")
```
### *It is also clear that mostly all educated class haven't subscribed to term deposit*
```
sns.pairplot(data=data,hue="y",vars=['age','balance','day','duration'])
```
### Correlation Mapping
```
plt.figure(figsize=(20,10))
sns.heatmap(data=data.corr(), annot=True, cmap='viridis')
```
### Correlation between variables is very less, which could possibly be due to presence of many outliers, so we will apply Transformations
## Checking Outliers in Continuous Variables
```
sns.boxplot(x="y",y="age",data=data)
sns.boxplot(x="y",y="balance",data=data)
sns.boxplot(x="y",y="duration",data=data)
```
### Applying Transformation
```
data['duration'].plot.hist()
from sklearn.preprocessing import PowerTransformer
b = data.drop(['job','marital','education','default','campaign','housing','loan','contact','month','previous','poutcome','y'],axis=1)
b
pt = PowerTransformer(method='yeo-johnson', standardize=True)
skl_yeojohnson = pt.fit(b)
calc_lambdas = skl_yeojohnson.lambdas_
skl_yeojohnson = pt.transform(b)
df_ptdata = pd.DataFrame(data=skl_yeojohnson, columns=['Age','Balance','Day','Duration','Pdays'])
df_ptdata.head()
```
### After Treatment data is tending towards normal distributed
```
import warnings
warnings.filterwarnings("ignore")
plt.figure(figsize=(15,5))
sns.distplot(df_ptdata)
```
### Before Treatment Data was Right Skewed
```
plt.figure(figsize=(15,5))
sns.distplot(b)
```
### By doing Transformation in 5 Columns most of data are now treated
```
DATA = pd.concat([df_ptdata,data],axis=1)
DATA
```
#### Replacing Treated Variables with original data
```
c = DATA.drop(['age','balance','day','duration','pdays'],axis=1)
c
```
### LABEL ENCODING on Certain Ordinal Data
```
label_encoder = preprocessing.LabelEncoder()
data['Job']= label_encoder.fit_transform(data['job'])
data['Marital']= label_encoder.fit_transform(data['marital'])
data['Education']= label_encoder.fit_transform(data['education'])
data['Month']= label_encoder.fit_transform(data['month'])
data['y1']= label_encoder.fit_transform(data['y'])
data
label_data=data.drop(['age','job','marital','education','default','balance','housing','loan','contact','day','duration','campaign','pdays','previous','poutcome','y','month'],axis=1)
label_data
```
### ONE HOT ENCODING on certain non-ordinal variables
```
Dummy_DATA = pd.get_dummies(c,columns=['default','housing','loan','contact','poutcome'])
Dummy_DATA
Dummy_Variables = Dummy_DATA.drop(['Age','Balance','Day','Duration','Pdays','job','marital','education','month','campaign','y','previous'],axis=1)
Dummy_Variables
Nondummy_DATA = Dummy_DATA.drop(['default_no','default_yes','housing_no','housing_yes','loan_no','loan_yes','contact_cellular','contact_telephone','contact_unknown','poutcome_failure','poutcome_other','poutcome_success','poutcome_unknown'],axis=1)
Nondummy_DATA
final_data=pd.concat([Dummy_Variables,Nondummy_DATA,label_data],axis=1)
final_data
final_data.shape
final_cleaned_data=final_data.drop(['job', 'marital', 'education', 'month','y'],axis=1)
final_cleaned_data.shape
final_cleaned_data.columns
```
### Our Model is to be built with 25 Columns where 5 Columns undergo One Hot Encoding and 4 Columns undergo Label Encoding
```
X = final_cleaned_data.iloc[:,:-1]
Y = final_cleaned_data.iloc[:,-1]
X_train,X_test,y_train,y_test = train_test_split(X,Y,test_size = 0.2,random_state = 42)
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
classifier.coef_
classifier.predict_proba(X_test)
y_pred=classifier.predict(X)
final_cleaned_data["y_pred"]=y_pred
final_cleaned_data
y_prob = pd.DataFrame(classifier.predict_proba(X.iloc[:,:]))
new_df = pd.concat([final_cleaned_data,y_prob],axis=1)
new_df
pd.crosstab(Y,y_pred)
print(classification_report(Y,y_pred))
logit_roc_score = roc_auc_score(Y,classifier.predict(X))
logit_roc_score
logit = sm.Logit(Y,X)
logit.fit().summary()
```
### From the analysis it seems that 4 columns have p>0.05, so those variables can be removed inorder to increase ROC Score.
### It also seems that there is Imbalance data that needs to be treated
```
!pip install imblearn
# Implementing Undersampling for Handling Imbalanced
nm = NearMiss()
X_res,y_res=nm.fit_resample(X,Y)
X_res.shape,y_res.shape
print('Original dataset shape {}'.format(Counter(Y)))
print('Resampled dataset shape {}'.format(Counter(y_res)))
X_train,X_test,y_train,y_test = train_test_split(X_res,y_res,test_size = 0.2,random_state = 42)
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
classifier.coef_
classifier.predict_proba(X_test)
y_pred=classifier.predict(X)
final_cleaned_data["y_pred"]=y_pred
final_cleaned_data
y_prob = pd.DataFrame(classifier.predict_proba(X.iloc[:,:]))
new_df = pd.concat([final_cleaned_data,y_prob],axis=1)
new_df
print(classification_report(Y,y_pred))
logit_roc_score = roc_auc_score(Y,classifier.predict(X))
logit_roc_score
logit = sm.Logit(Y,X)
logit.fit().summary()
```
### From Undersampling Technique we loose some part of data, so we should go with Oversampling Technique to get more accuracy
```
# Performing Oversampling method to handle imbalanced data
smk = SMOTETomek(random_state=42)
X_res,y_res = smk.fit_resample(X,Y)
X_res.shape,y_res.shape
print('Original dataset shape {}'.format(Counter(Y)))
print('Resampled dataset shape {}'.format(Counter(y_res)))
X_train,X_test,y_train,y_test = train_test_split(X_res,y_res,test_size = 0.2,random_state = 42)
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
classifier.coef_
classifier.predict_proba(X_test)
y_pred=classifier.predict(X)
final_cleaned_data["y_pred"]=y_pred
final_cleaned_data
print(classification_report(Y,y_pred))
logit_roc_score = roc_auc_score(Y,classifier.predict(X))
logit_roc_score
fpr, tpr, thresholds = roc_curve(Y, classifier.predict_proba (X)[:,1])
auc = roc_auc_score(Y, y_pred)
plt.plot(fpr, tpr, color='navy', label='Logistic Regresiion ROC curve (area = %0.2f)' %logit_roc_score)
plt.plot([0, 1], [0, 1], 'k--')
plt.xlabel('False Positive Rate or [1 - True Negative Rate]')
plt.ylabel('True Positive Rate')
plt.grid()
plt.title("ROC with Over-sampling Method")
plt.legend(loc = "lower right")
```
### From the above analysis it is clear that with Over-Sampling method from SMOTE library Logistic Regression for the following assignment gives good AUC Score which is a measure to check accuracy of model
# Conclusions about Assignment:
> First Check data having Null Values
> Check if Target Variable is Balanced or Imbalanced
> If it is Imblanced, we have to perform Undersampling or Oversmapling whichever gives best result should be considered as Final Model
> For Treating Variables Split Data into 3 Parts:
1. Continuous Variables:
- All continuous Data must be passed through some Transformation so that data is Normalized
2. Categorical Variables with Odinal Type data:
- All Categorical Variables with Ordinal Type should be Label Encoded
3. Categorical Variables with Non-Ordinal Type data:
- All Categorical Variable with Non-Ordinal Type should be One Hot Encoded
> All the categorical variables before modelling must be in continuous type
> To check accuracy of model, do not conclude directly from Accuracy of Classification Report, one must also check AUC Score to give promising result of Accuracy.
| github_jupyter |
# Implementation of the Moho inversion algorithm
This notebook presents a Python class that implements the proposed method. We'll use the [inverse problems framework](http://www.fatiando.org/api/inversion.html) of the library [Fatiando a Terra](http://www.fatiando.org). The class `MohoGravityInvSpherical` is defined in the [`mohoinv.py`](mohoinv.py) file.
## Package imports
```
# Insert plots into the notebook
%matplotlib inline
from __future__ import division, unicode_literals
import numpy as np
import multiprocessing
from IPython.display import Image
import matplotlib.pyplot as plt
import seaborn # Makes the default style of the plots nicer
```
Load the required modules from Fatiando a Terra and show the specific version of the library used.
```
from fatiando.vis import mpl
from fatiando.gravmag import tesseroid
from fatiando import utils, gridder
import fatiando
print("Using Fatiando a Terra version: {}".format(fatiando.__version__))
from mohoinv import TesseroidRelief, MohoGravityInvSpherical, make_mesh
```
Get the number of cores in the computer to run the forward modeling in parallel.
```
ncpu = multiprocessing.cpu_count()
ncpu
```
## Test the class on simple synthetic data
We can test and show how the class works on some simple synthetic data. We'll use the example model from the [tesseroid-relief-example.ipynb](tesseroid-relief-example.ipynb) notebook.
First, make the model of the Moho.
```
# shape is nlat, nlon = the number of points in the grid
shape = (30, 30)
# Make a regular grid inside an area = (s, n, w, e)
area = (20, 60, -40, 40)
lat, lon, h = gridder.regular(area, shape, z=250e3)
# Make a checkerboard relief undulating along the -35km height reference
f = 0.15
reference = -35e3
relief = 10e3*np.sin(1.5*f*lon)*np.cos(f*lat) + reference
# The density contrast is negative if the relief is below the reference
density = 600*np.ones_like(relief)
density[relief < reference] *= -1
model = make_mesh(area, shape, relief, reference)
model.addprop('density', density)
plt.figure(figsize=(9, 4))
plt.title("Synthetic Moho depths")
plt.pcolormesh(model.lons, model.lats, -0.001*model.relief.reshape(model.shape),
cmap='Blues')
plt.colorbar(pad=0.01).set_label('km')
plt.xlim(model.lons.min(), model.lons.max())
plt.ylim(model.lats.min(), model.lats.max())
```
Now, generate some synthetic data by forward modeling.
```
gz = tesseroid.gz(lon, lat, h, model, njobs=ncpu)
plt.figure(figsize=(9, 4))
plt.title('Moho synthetic gravity anomaly')
plt.tricontourf(lon, lat, gz, 30, cmap='RdBu_r')
plt.colorbar(pad=0).set_label('mGal')
```
## Run the inversion
For this test, we'll use a mesh with the same dimensions and the original model.
```
mesh = model.copy(deep=True)
mesh.props['density'] = 600*np.ones(mesh.size)
```
Create the solver object.
```
solver = MohoGravityInvSpherical(lat, lon, h, gz, mesh, njobs=ncpu)
```
Configure the optimization method to Gauss-Newton and set the initial estimate.
```
initial = np.ones(solver.nparams)*(mesh.reference - 30e3)
solver.config('newton', initial=initial, tol=0.2, maxit=10)
```
Run the inversion and time the computation.
```
%time solver.fit()
```
Plot the RMS error (Root Mean Square) per iteration to get an idea of the convergence of the method.
```
rms = np.sqrt(solver.stats_['objective'])/np.sqrt(solver.ndata)
plt.figure()
ax = plt.subplot(111)
ax.set_title('Convergence of {}'.format(solver.stats_['method']))
ax.plot(rms, '.k-')
ax.set_ylabel('RMS (mGal)')
ax.set_xlabel('Iteration')
```
Plot the data misfit and residuals
```
predicted = solver.predicted()
residuals = solver.residuals()
plt.figure(figsize=(9, 4))
levels = mpl.contourf(lon, lat, gz, shape, 12, cmap='RdBu_r')
plt.colorbar(pad=0).set_label('mGal')
mpl.contour(lon, lat, predicted, shape, levels)
plt.title('Observed (color) and predicted (contour) data')
plt.title('Residuals')
plt.hist(residuals, bins=20, normed=True)
plt.xlabel('Residual (mGal)')
print('Mean: {} std: {}'.format(residuals.mean(), residuals.std()))
```
Map the estimated Moho depth.
```
moho = solver.estimate_
plt.figure(figsize=(9, 4))
plt.title("Estimated Moho depth")
plt.pcolormesh(moho.lons, moho.lats, -0.001*moho.relief.reshape(moho.shape),
cmap='Blues')
plt.colorbar(pad=0.01).set_label('km')
plt.xlim(moho.lons.min(), moho.lons.max())
plt.ylim(moho.lats.min(), moho.lats.max())
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
#sys.path.insert(1, '/home/ximo/Documents/GitHub/skforecast')
%config Completer.use_jedi = False
```
## Libraries
```
# Libraries
# ==============================================================================
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from skforecast.ForecasterAutoregMultiOutput import ForecasterAutoregMultiOutput
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_squared_error
```
## Data
```
# Download data
# ==============================================================================
url = ('https://raw.githubusercontent.com/JoaquinAmatRodrigo/skforecast/master/data/h2o.csv')
data = pd.read_csv(url, sep=',', header=0, names=['y', 'datetime'])
# Data preprocessing
# ==============================================================================
data['datetime'] = pd.to_datetime(data['datetime'], format='%Y/%m/%d')
data = data.set_index('datetime')
data = data.asfreq('MS')
data = data['y']
data = data.sort_index()
# Split train-test
# ==============================================================================
steps = 36
data_train = data[:-steps]
data_test = data[-steps:]
# Plot
# ==============================================================================
fig, ax=plt.subplots(figsize=(9, 4))
data_train.plot(ax=ax, label='train')
data_test.plot(ax=ax, label='test')
ax.legend();
```
## Train forecaster
```
# Create and fit forecaster
# ==============================================================================
forecaster = ForecasterAutoregMultiOutput(
regressor = Ridge(),
steps = 36,
lags = 15
)
forecaster.fit(y=data_train)
forecaster
```
## Prediction
```
# Predict
# ==============================================================================
steps = 36
predictions = forecaster.predict(steps=steps)
predictions.head(3)
# Plot predictions
# ==============================================================================
fig, ax=plt.subplots(figsize=(9, 4))
data_train.plot(ax=ax, label='train')
data_test.plot(ax=ax, label='test')
predictions.plot(ax=ax, label='predictions')
ax.legend();
# Prediction error
# ==============================================================================
error_mse = mean_squared_error(
y_true = data_test,
y_pred = predictions
)
print(f"Test error (mse): {error_mse}")
```
## Feature importance
Since `ForecasterAutoregMultiOutput` fits one model per step,it is necessary to specify from which model retrieve its feature importance.
```
print(forecaster.get_coef(step=1).to_markdown(index=False))
```
## Extract training matrix
Two steps are needed. One to create the whole training matrix and a second one to subset the data needed for each model (step).
```
X, y = forecaster.create_train_X_y(data_train)
# X and y to train model for step 1
X_1, y_1 = forecaster.filter_train_X_y_for_step(
step = 1,
X_train = X,
y_train = y,
)
print(X_1.head(4).to_markdown(index=False))
print(y_1.head(4).to_markdown(index=False))
```
| github_jupyter |
**10장 – 케라스를 사용한 인공 신경망 소개**
_이 노트북은 10장에 있는 모든 샘플 코드와 연습문제 해답을 가지고 있습니다._
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/handson-ml2/blob/master/10_neural_nets_with_keras.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩에서 실행하기</a>
</td>
</table>
# 설정
먼저 몇 개의 모듈을 임포트합니다. 맷플롯립 그래프를 인라인으로 출력하도록 만들고 그림을 저장하는 함수를 준비합니다. 또한 파이썬 버전이 3.5 이상인지 확인합니다(파이썬 2.x에서도 동작하지만 곧 지원이 중단되므로 파이썬 3을 사용하는 것이 좋습니다). 사이킷런 버전이 0.20 이상인지와 텐서플로 버전이 2.0 이상인지 확인합니다.
```
# 파이썬 ≥3.5 필수
import sys
assert sys.version_info >= (3, 5)
# 사이킷런 ≥0.20 필수
import sklearn
assert sklearn.__version__ >= "0.20"
# 텐서플로 ≥2.0 필수
import tensorflow as tf
assert tf.__version__ >= "2.0"
# 공통 모듈 임포트
import numpy as np
import os
# 노트북 실행 결과를 동일하게 유지하기 위해
np.random.seed(42)
# 깔끔한 그래프 출력을 위해
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# 그림을 저장할 위치
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "ann"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("그림 저장:", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# 불필요한 경고를 무시합니다 (사이파이 이슈 #5998 참조)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
```
# 퍼셉트론
**노트**: 사이킷런 향후 버전에서 `max_iter`와 `tol` 매개변수의 기본값이 바뀌기 때문에 경고를 피하기 위해 명시적으로 지정합니다.
```
import numpy as np
from sklearn.datasets import load_iris
from sklearn.linear_model import Perceptron
iris = load_iris()
X = iris.data[:, (2, 3)] # 꽃잎 길이, 꽃잎 너비
y = (iris.target == 0).astype(np.int)
per_clf = Perceptron(max_iter=1000, tol=1e-3, random_state=42)
per_clf.fit(X, y)
y_pred = per_clf.predict([[2, 0.5]])
y_pred
a = -per_clf.coef_[0][0] / per_clf.coef_[0][1]
b = -per_clf.intercept_ / per_clf.coef_[0][1]
axes = [0, 5, 0, 2]
x0, x1 = np.meshgrid(
np.linspace(axes[0], axes[1], 500).reshape(-1, 1),
np.linspace(axes[2], axes[3], 200).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_predict = per_clf.predict(X_new)
zz = y_predict.reshape(x0.shape)
plt.figure(figsize=(10, 4))
plt.plot(X[y==0, 0], X[y==0, 1], "bs", label="Not Iris-Setosa")
plt.plot(X[y==1, 0], X[y==1, 1], "yo", label="Iris-Setosa")
plt.plot([axes[0], axes[1]], [a * axes[0] + b, a * axes[1] + b], "k-", linewidth=3)
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#9898ff', '#fafab0'])
plt.contourf(x0, x1, zz, cmap=custom_cmap)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="lower right", fontsize=14)
plt.axis(axes)
save_fig("perceptron_iris_plot")
plt.show()
```
# 활성화 함수
```
def sigmoid(z):
return 1 / (1 + np.exp(-z))
def relu(z):
return np.maximum(0, z)
def derivative(f, z, eps=0.000001):
return (f(z + eps) - f(z - eps))/(2 * eps)
z = np.linspace(-5, 5, 200)
plt.figure(figsize=(11,4))
plt.subplot(121)
plt.plot(z, np.sign(z), "r-", linewidth=1, label="Step")
plt.plot(z, sigmoid(z), "g--", linewidth=2, label="Sigmoid")
plt.plot(z, np.tanh(z), "b-", linewidth=2, label="Tanh")
plt.plot(z, relu(z), "m-.", linewidth=2, label="ReLU")
plt.grid(True)
plt.legend(loc="center right", fontsize=14)
plt.title("Activation functions", fontsize=14)
plt.axis([-5, 5, -1.2, 1.2])
plt.subplot(122)
plt.plot(z, derivative(np.sign, z), "r-", linewidth=1, label="Step")
plt.plot(0, 0, "ro", markersize=5)
plt.plot(0, 0, "rx", markersize=10)
plt.plot(z, derivative(sigmoid, z), "g--", linewidth=2, label="Sigmoid")
plt.plot(z, derivative(np.tanh, z), "b-", linewidth=2, label="Tanh")
plt.plot(z, derivative(relu, z), "m-.", linewidth=2, label="ReLU")
plt.grid(True)
#plt.legend(loc="center right", fontsize=14)
plt.title("Derivatives", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("activation_functions_plot")
plt.show()
def heaviside(z):
return (z >= 0).astype(z.dtype)
def mlp_xor(x1, x2, activation=heaviside):
return activation(-activation(x1 + x2 - 1.5) + activation(x1 + x2 - 0.5) - 0.5)
x1s = np.linspace(-0.2, 1.2, 100)
x2s = np.linspace(-0.2, 1.2, 100)
x1, x2 = np.meshgrid(x1s, x2s)
z1 = mlp_xor(x1, x2, activation=heaviside)
z2 = mlp_xor(x1, x2, activation=sigmoid)
plt.figure(figsize=(10,4))
plt.subplot(121)
plt.contourf(x1, x2, z1)
plt.plot([0, 1], [0, 1], "gs", markersize=20)
plt.plot([0, 1], [1, 0], "y^", markersize=20)
plt.title("Activation function: heaviside", fontsize=14)
plt.grid(True)
plt.subplot(122)
plt.contourf(x1, x2, z2)
plt.plot([0, 1], [0, 1], "gs", markersize=20)
plt.plot([0, 1], [1, 0], "y^", markersize=20)
plt.title("Activation function: sigmoid", fontsize=14)
plt.grid(True)
```
# 이미지 분류기 만들기
먼저 텐서플로와 케라스를 임포트합니다.
```
import tensorflow as tf
from tensorflow import keras
tf.__version__
keras.__version__
```
먼저 MNIST 데이터셋을 로드하겠습니다. 케라스는 `keras.datasets`에 널리 사용하는 데이터셋을 로드하기 위한 함수를 제공합니다. 이 데이터셋은 이미 훈련 세트와 테스트 세트로 나누어져 있습니다. 훈련 세트를 더 나누어 검증 세트를 만드는 것이 좋습니다:
```
fashion_mnist = keras.datasets.fashion_mnist
(X_train_full, y_train_full), (X_test, y_test) = fashion_mnist.load_data()
```
훈련 세트는 60,000개의 흑백 이미지입니다. 각 이미지의 크기는 28x28 픽셀입니다:
```
X_train_full.shape
```
각 픽셀의 강도는 바이트(0~255)로 표현됩니다:
```
X_train_full.dtype
```
전체 훈련 세트를 검증 세트와 (조금 더 작은) 훈련 세트로 나누어 보죠. 또한 픽셀 강도를 255로 나누어 0~1 범위의 실수로 바꾸겠습니다.
```
X_valid, X_train = X_train_full[:5000] / 255., X_train_full[5000:] / 255.
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
X_test = X_test / 255.
```
맷플롯립의 `imshow()` 함수와 `'binary'` 컬러맵을 사용해 이미지를 출력할 수 있습니다:
```
plt.imshow(X_train[0], cmap="binary")
plt.axis('off')
plt.show()
```
레이블은 0에서 9까지 (uint8로 표현된) 클래스 아이디입니다:
```
y_train
```
클래스 이름은 다음과 같습니다:
```
class_names = ["T-shirt/top", "Trouser", "Pullover", "Dress", "Coat",
"Sandal", "Shirt", "Sneaker", "Bag", "Ankle boot"]
```
훈련 세트에 있는 첫 번째 이미지는 코트입니다:
```
class_names[y_train[0]]
```
검증 세트는 5,000개의 이미지를 담고 있고 테스트 세트는 10,000개의 이미지를 가집니다:
```
X_valid.shape
X_test.shape
```
이 데이터셋에 있는 샘플 이미지를 몇 개 출력해 보죠:
```
n_rows = 4
n_cols = 10
plt.figure(figsize=(n_cols * 1.2, n_rows * 1.2))
for row in range(n_rows):
for col in range(n_cols):
index = n_cols * row + col
plt.subplot(n_rows, n_cols, index + 1)
plt.imshow(X_train[index], cmap="binary", interpolation="nearest")
plt.axis('off')
plt.title(class_names[y_train[index]], fontsize=12)
plt.subplots_adjust(wspace=0.2, hspace=0.5)
save_fig('fashion_mnist_plot', tight_layout=False)
plt.show()
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="relu"))
model.add(keras.layers.Dense(100, activation="relu"))
model.add(keras.layers.Dense(10, activation="softmax"))
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
model.layers
model.summary()
keras.utils.plot_model(model, "my_fashion_mnist_model.png", show_shapes=True)
hidden1 = model.layers[1]
hidden1.name
model.get_layer(hidden1.name) is hidden1
weights, biases = hidden1.get_weights()
weights
weights.shape
biases
biases.shape
model.compile(loss="sparse_categorical_crossentropy",
optimizer="sgd",
metrics=["accuracy"])
```
위 코드는 다음과 같습니다:
```python
model.compile(loss=keras.losses.sparse_categorical_crossentropy,
optimizer=keras.optimizers.SGD(),
metrics=[keras.metrics.sparse_categorical_accuracy])
```
```
history = model.fit(X_train, y_train, epochs=30,
validation_data=(X_valid, y_valid))
history.params
print(history.epoch)
history.history.keys()
import pandas as pd
pd.DataFrame(history.history).plot(figsize=(8, 5))
plt.grid(True)
plt.gca().set_ylim(0, 1)
save_fig("keras_learning_curves_plot")
plt.show()
model.evaluate(X_test, y_test)
X_new = X_test[:3]
y_proba = model.predict(X_new)
y_proba.round(2)
y_pred = model.predict_classes(X_new)
y_pred
np.array(class_names)[y_pred]
y_new = y_test[:3]
y_new
plt.figure(figsize=(7.2, 2.4))
for index, image in enumerate(X_new):
plt.subplot(1, 3, index + 1)
plt.imshow(image, cmap="binary", interpolation="nearest")
plt.axis('off')
plt.title(class_names[y_test[index]], fontsize=12)
plt.subplots_adjust(wspace=0.2, hspace=0.5)
save_fig('fashion_mnist_images_plot', tight_layout=False)
plt.show()
```
# 회귀 MLP
캘리포니아 주택 데이터셋을 로드하여 나누고 스케일을 바꾸어 보겠습니다(2장에서 사용한 수정된 버전이 아니라 원본을 사용합니다):
```
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
housing = fetch_california_housing()
X_train_full, X_test, y_train_full, y_test = train_test_split(housing.data, housing.target, random_state=42)
X_train, X_valid, y_train, y_valid = train_test_split(X_train_full, y_train_full, random_state=42)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_valid = scaler.transform(X_valid)
X_test = scaler.transform(X_test)
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=X_train.shape[1:]),
keras.layers.Dense(1)
])
model.compile(loss="mean_squared_error", optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit(X_train, y_train, epochs=20, validation_data=(X_valid, y_valid))
mse_test = model.evaluate(X_test, y_test)
X_new = X_test[:3]
y_pred = model.predict(X_new)
plt.plot(pd.DataFrame(history.history))
plt.grid(True)
plt.gca().set_ylim(0, 1)
plt.show()
y_pred
```
# 함수형 API
모든 신경망 모델이 단순하게 순서대로 나열되지는 않습니다. 어떤 신경망은 매우 복잡한 구조를 가집니다. 여러 개의 입력이 있거나 여러 개의 출력이 있습니다. 예를 들어 와이드 & 딥 신경망([논문](https://ai.google/research/pubs/pub45413) 참조)은 입력의 전체 또는 일부를 출력층에 바로 연결합니다.
```
np.random.seed(42)
tf.random.set_seed(42)
input_ = keras.layers.Input(shape=X_train.shape[1:])
hidden1 = keras.layers.Dense(30, activation="relu")(input_)
hidden2 = keras.layers.Dense(30, activation="relu")(hidden1)
concat = keras.layers.concatenate([input_, hidden2])
output = keras.layers.Dense(1)(concat)
model = keras.models.Model(inputs=[input_], outputs=[output])
model.summary()
model.compile(loss="mean_squared_error", optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit(X_train, y_train, epochs=20,
validation_data=(X_valid, y_valid))
mse_test = model.evaluate(X_test, y_test)
y_pred = model.predict(X_new)
```
와이드나 딥 경로에 다른 입력 특성을 전달하면 어떻게 될까요? (특성 0에서 4까지) 5개의 특성을 와이드 경로에 보내고 (특성 2에서 7까지) 6개의 특성을 딥 경로에 전달하겠습니다. 3개의 특성(특성 2, 3, 4)은 양쪽에 모두 전달됩니다.
```
np.random.seed(42)
tf.random.set_seed(42)
input_A = keras.layers.Input(shape=[5], name="wide_input")
input_B = keras.layers.Input(shape=[6], name="deep_input")
hidden1 = keras.layers.Dense(30, activation="relu")(input_B)
hidden2 = keras.layers.Dense(30, activation="relu")(hidden1)
concat = keras.layers.concatenate([input_A, hidden2])
output = keras.layers.Dense(1, name="output")(concat)
model = keras.models.Model(inputs=[input_A, input_B], outputs=[output])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
X_train_A, X_train_B = X_train[:, :5], X_train[:, 2:]
X_valid_A, X_valid_B = X_valid[:, :5], X_valid[:, 2:]
X_test_A, X_test_B = X_test[:, :5], X_test[:, 2:]
X_new_A, X_new_B = X_test_A[:3], X_test_B[:3]
history = model.fit((X_train_A, X_train_B), y_train, epochs=20,
validation_data=((X_valid_A, X_valid_B), y_valid))
mse_test = model.evaluate((X_test_A, X_test_B), y_test)
y_pred = model.predict((X_new_A, X_new_B))
```
규제를 위한 보조 출력 추가하기:
```
np.random.seed(42)
tf.random.set_seed(42)
input_A = keras.layers.Input(shape=[5], name="wide_input")
input_B = keras.layers.Input(shape=[6], name="deep_input")
hidden1 = keras.layers.Dense(30, activation="relu")(input_B)
hidden2 = keras.layers.Dense(30, activation="relu")(hidden1)
concat = keras.layers.concatenate([input_A, hidden2])
output = keras.layers.Dense(1, name="main_output")(concat)
aux_output = keras.layers.Dense(1, name="aux_output")(hidden2)
model = keras.models.Model(inputs=[input_A, input_B],
outputs=[output, aux_output])
model.compile(loss=["mse", "mse"], loss_weights=[0.9, 0.1], optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit([X_train_A, X_train_B], [y_train, y_train], epochs=20,
validation_data=([X_valid_A, X_valid_B], [y_valid, y_valid]))
total_loss, main_loss, aux_loss = model.evaluate(
[X_test_A, X_test_B], [y_test, y_test])
y_pred_main, y_pred_aux = model.predict([X_new_A, X_new_B])
```
# 서브클래싱 API
```
class WideAndDeepModel(keras.models.Model):
def __init__(self, units=30, activation="relu", **kwargs):
super().__init__(**kwargs)
self.hidden1 = keras.layers.Dense(units, activation=activation)
self.hidden2 = keras.layers.Dense(units, activation=activation)
self.main_output = keras.layers.Dense(1)
self.aux_output = keras.layers.Dense(1)
def call(self, inputs):
input_A, input_B = inputs
hidden1 = self.hidden1(input_B)
hidden2 = self.hidden2(hidden1)
concat = keras.layers.concatenate([input_A, hidden2])
main_output = self.main_output(concat)
aux_output = self.aux_output(hidden2)
return main_output, aux_output
model = WideAndDeepModel(30, activation="relu")
model.compile(loss="mse", loss_weights=[0.9, 0.1], optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit((X_train_A, X_train_B), (y_train, y_train), epochs=10,
validation_data=((X_valid_A, X_valid_B), (y_valid, y_valid)))
total_loss, main_loss, aux_loss = model.evaluate((X_test_A, X_test_B), (y_test, y_test))
y_pred_main, y_pred_aux = model.predict((X_new_A, X_new_B))
model = WideAndDeepModel(30, activation="relu")
```
# 저장과 복원
```
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=[8]),
keras.layers.Dense(30, activation="relu"),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit(X_train, y_train, epochs=10, validation_data=(X_valid, y_valid))
mse_test = model.evaluate(X_test, y_test)
model.save("my_keras_model.h5")
model = keras.models.load_model("my_keras_model.h5")
model.predict(X_new)
model.save_weights("my_keras_weights.ckpt")
model.load_weights("my_keras_weights.ckpt")
```
# 훈련 과정에서 콜백 사용하기
```
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=[8]),
keras.layers.Dense(30, activation="relu"),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
checkpoint_cb = keras.callbacks.ModelCheckpoint("my_keras_model.h5", save_best_only=True)
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid),
callbacks=[checkpoint_cb])
model = keras.models.load_model("my_keras_model.h5") # 최상의 모델로 롤백
mse_test = model.evaluate(X_test, y_test)
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
early_stopping_cb = keras.callbacks.EarlyStopping(patience=10,
restore_best_weights=True)
history = model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[checkpoint_cb, early_stopping_cb])
mse_test = model.evaluate(X_test, y_test)
class PrintValTrainRatioCallback(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
print("\nval/train: {:.2f}".format(logs["val_loss"] / logs["loss"]))
val_train_ratio_cb = PrintValTrainRatioCallback()
history = model.fit(X_train, y_train, epochs=1,
validation_data=(X_valid, y_valid),
callbacks=[val_train_ratio_cb])
```
# 텐서보드
```
root_logdir = os.path.join(os.curdir, "my_logs")
def get_run_logdir():
import time
run_id = time.strftime("run_%Y_%m_%d-%H_%M_%S")
return os.path.join(root_logdir, run_id)
run_logdir = get_run_logdir()
run_logdir
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=[8]),
keras.layers.Dense(30, activation="relu"),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=1e-3))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
history = model.fit(X_train, y_train, epochs=30,
validation_data=(X_valid, y_valid),
callbacks=[checkpoint_cb, tensorboard_cb])
```
텐서보드 서버를 실행하는 한 가지 방법은 터미널에서 직접 실행하는 것입니다. 터미널을 열고 텐서보드가 설치된 가상 환경을 활성화합니다. 그다음 노트북 디렉토리로 이동하여 다음 명령을 입력하세요:
```bash
$ tensorboard --logdir=./my_logs --port=6006
```
그다음 웹 브라우저를 열고 [localhost:6006](http://localhost:6006)에 접속하면 텐서보드를 사용할 수 있습니다. 사용이 끝나면 터미널에서 Ctrl-C를 눌러 텐서보드 서버를 종료하세요.
또는 다음처럼 텐서보드의 주피터 확장을 사용할 수 있습니다(이 명령은 텐서보드가 로컬 컴퓨터에 설치되어 있어야 합니다):
```
%load_ext tensorboard
%tensorboard --logdir=./my_logs --port=6006
run_logdir2 = get_run_logdir()
run_logdir2
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Dense(30, activation="relu", input_shape=[8]),
keras.layers.Dense(30, activation="relu"),
keras.layers.Dense(1)
])
model.compile(loss="mse", optimizer=keras.optimizers.SGD(lr=0.05))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir2)
history = model.fit(X_train, y_train, epochs=30,
validation_data=(X_valid, y_valid),
callbacks=[checkpoint_cb, tensorboard_cb])
```
텐서보드에 실행 결과가 2개 있습니다. 학습 곡선을 비교해 보세요.
사용할 수 있는 로깅 옵션을 확인해 보죠:
```
help(keras.callbacks.TensorBoard.__init__)
```
# 하이퍼파라미터 튜닝
```
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
def build_model(n_hidden=1, n_neurons=30, learning_rate=3e-3, input_shape=[8]):
model = keras.models.Sequential()
model.add(keras.layers.InputLayer(input_shape=input_shape))
for layer in range(n_hidden):
model.add(keras.layers.Dense(n_neurons, activation="relu"))
model.add(keras.layers.Dense(1))
optimizer = keras.optimizers.SGD(lr=learning_rate)
model.compile(loss="mse", optimizer=optimizer)
return model
keras_reg = keras.wrappers.scikit_learn.KerasRegressor(build_model)
keras_reg.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[keras.callbacks.EarlyStopping(patience=10)])
mse_test = keras_reg.score(X_test, y_test)
y_pred = keras_reg.predict(X_new)
np.random.seed(42)
tf.random.set_seed(42)
```
**경고**: 다음 셀은 훈련이 끝날 때 에러가 납니다. 이는 최근 사이킷런의 변화때문에 생긴 [케라스 이슈 #13586](https://github.com/keras-team/keras/issues/13586) 때문입니다. 이 이슈를 해결하기 위한 [풀 리퀘스트 #13598](https://github.com/keras-team/keras/pull/13598)가 있으므로 곧 해결될 것 같습니다.
```
from scipy.stats import reciprocal
from sklearn.model_selection import RandomizedSearchCV
param_distribs = {
"n_hidden": [0, 1, 2, 3],
"n_neurons": np.arange(1, 100),
"learning_rate": reciprocal(3e-4, 3e-2),
}
rnd_search_cv = RandomizedSearchCV(keras_reg, param_distribs, n_iter=10, cv=3, verbose=2)
rnd_search_cv.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[keras.callbacks.EarlyStopping(patience=10)])
rnd_search_cv.best_params_
rnd_search_cv.best_score_
rnd_search_cv.best_estimator_
rnd_search_cv.score(X_test, y_test)
model = rnd_search_cv.best_estimator_.model
model
model.evaluate(X_test, y_test)
```
# 연습문제 해답
## 1. to 9.
부록 A 참조.
## 10.
*문제: 심층 MLP를 MNIST 데이터셋에 훈련해보세요(`keras.datasets.mnist.load_data()` 함수를 사용해 데이터를 적재할 수 있습니다). 98% 이상의 정확도를 얻을 수 있는지 확인해보세요. 이 장에서 소개한 방법을 사용해 최적의 학습률을 찾아보세요(즉 학습률을 지수적으로 증가시키면서 손실을 그래프로 그립니다. 그다음 손실이 다시 증가하는 지점을 찾습니다). 모든 부가 기능을 추가해보세요. 즉, 체크포인트를 저장하고, 조기 종료를 사용하고, 텐서보드를 사용해 학습 곡선을 그려보세요.*
데이터셋을 적재해보죠:
```
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.mnist.load_data()
```
패션 MNIST 데이터셋처럼 MNIST 훈련 세트는 28x28 픽셀의 흑백 이미지 60,000개로 이루어져 있습니다:
```
X_train_full.shape
```
각 픽셀 강도는 바이트(0~255)로 표현됩니다:
```
X_train_full.dtype
```
전체 훈련 세트를 검증 세트와 (더 작은) 훈련 세트로 나누어 보겠습니다. 패션 MNIST처럼 픽셀 강도를 255로 나누어 0-1 범위의 실수로 변환합니다:
```
X_valid, X_train = X_train_full[:5000] / 255., X_train_full[5000:] / 255.
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
X_test = X_test / 255.
```
맷플롯립의 `imshow()` 함수와 `'binary'` 컬러 맵으로 이미지를 출력해 보죠:
```
plt.imshow(X_train[0], cmap="binary")
plt.axis('off')
plt.show()
```
레이블은 (uint8로 표현된) 0에서 9까지 클래스 아이디입니다. 편리하게도 클래스 아이디는 이미지가 나타내는 숫자와 같습니다. 따라서 `class_names` 배열을 만들 필요가 없습니다:
```
y_train
```
검증 세트는 5,000개의 이미지를 담고 있고 테스트 세트는 10,000개의 이미지를 담고 있습니다:
```
X_valid.shape
X_test.shape
```
이 데이터셋에 있는 이미지 샘플 몇 개를 출력해 보죠:
```
n_rows = 4
n_cols = 10
plt.figure(figsize=(n_cols * 1.2, n_rows * 1.2))
for row in range(n_rows):
for col in range(n_cols):
index = n_cols * row + col
plt.subplot(n_rows, n_cols, index + 1)
plt.imshow(X_train[index], cmap="binary", interpolation="nearest")
plt.axis('off')
plt.title(y_train[index], fontsize=12)
plt.subplots_adjust(wspace=0.2, hspace=0.5)
plt.show()
```
간단한 밀집 신경망을 만들고 최적의 학습률을 찾아 보겠습니다. 반복마다 학습률을 증가시키기 위해 콜백을 사용합니다. 이 콜백은 반복마다 학습률과 손실을 기록합니다:
```
K = keras.backend
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_batch_end(self, batch, logs):
self.rates.append(K.get_value(self.model.optimizer.lr))
self.losses.append(logs["loss"])
K.set_value(self.model.optimizer.lr, self.model.optimizer.lr * self.factor)
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
```
작은 학습률 1e-3에서 시작하여 반복마다 0.5%씩 증가합니다:
```
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
expon_lr = ExponentialLearningRate(factor=1.005)
```
모델을 1 에포크만 훈련해 보죠:
```
history = model.fit(X_train, y_train, epochs=1,
validation_data=(X_valid, y_valid),
callbacks=[expon_lr])
```
학습률에 대한 함수로 손실을 그릴 수 있습니다:
```
plt.plot(expon_lr.rates, expon_lr.losses)
plt.gca().set_xscale('log')
plt.hlines(min(expon_lr.losses), min(expon_lr.rates), max(expon_lr.rates))
plt.axis([min(expon_lr.rates), max(expon_lr.rates), 0, expon_lr.losses[0]])
plt.xlabel("Learning rate")
plt.ylabel("Loss")
```
손실이 3e-1에서 갑자기 솟구쳤기 때문에 2e-1을 학습률로 사용하겠습니다:
```
keras.backend.clear_session()
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="relu"),
keras.layers.Dense(100, activation="relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=2e-1),
metrics=["accuracy"])
run_index = 1 # 실행할 때마다 이 값을 늘립니다
run_logdir = os.path.join(os.curdir, "my_mnist_logs", "run_{:03d}".format(run_index))
run_logdir
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
checkpoint_cb = keras.callbacks.ModelCheckpoint("my_mnist_model.h5", save_best_only=True)
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
history = model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=[early_stopping_cb, checkpoint_cb, tensorboard_cb])
model = keras.models.load_model("my_mnist_model.h5") # rollback to best model
model.evaluate(X_test, y_test)
```
98% 정확도를 얻었습니다. 마지막으로 텐서보드를 사용해 학습 곡선을 살펴보겠습니다:
```
%tensorboard --logdir=./my_mnist_logs --port=6006
```
| github_jupyter |
# Method2 SVD+Huffman
## Import Libraries
```
import mne
import numpy as np
from scipy.fft import fft,fftshift
import matplotlib.pyplot as plt
from scipy.signal import butter, lfilter
from scipy.signal import freqz
from scipy import signal
from sklearn.metrics import mean_squared_error
from math import sqrt
import os
import pandas as pd
temp = pd.read_csv('TEMP.csv')
np.savetxt('temp.txt',temp)
temp_array = temp.to_numpy()
temp_array_new = []
for i in temp_array:
temp_array_new.append(list(i)[0])
N = len(temp_array_new)
sampling_freq = 1/4
index = np.linspace(0, round((N-1)*sampling_freq,4), N)
```
## Butterworth Band pass Filter
The butterworth filter were not applied on the TEMP data
because of it's low frequency and short sequence
## Resampling
```
y = temp_array
resampled_signal = signal.resample(y,9025)
#This squared number need to be decided by the users
np.savetxt('processed_temp.txt',resampled_signal)
```
## SVD
```
from scipy import linalg
reshaped_signal = np.reshape(resampled_signal, (95,95))
U, s, Vh = linalg.svd(reshaped_signal)
s[2:] = 0
m = 95
n = 95
sigma = np.zeros((m, n))
for i in range(min(m, n)):
sigma[i, i] = s[i]
reconstructed_signal = np.dot(U, np.dot(sigma, Vh))
deSVD = np.reshape(reconstructed_signal,95*95)
```
## Round the signal and then do Huffman coding
```
round_signal= np.round(deSVD,2)
np.savetxt('deSVD_temp.txt',round_signal,fmt='%.2f')
```
### INSTRUCTION ON HOW TO COMPRESS THE DATA BY HUFFMAN CODING
(I used the package "tcmpr 0.2" and "pyhuff 1.1". These two packages provided the same compression result. So here, we just use "tcmpr 0.2")
1. Open your termial or git bash, enter "pip install tcmpr" to install the "tcmpr 0.2" package
2. Enter the directory which include the file you want to compress OR copy the path of the file you want to compress
3. Enter "tcmpr filename.txt" / "tcmpr filepath" to compress the file
4. Find the compressed file in the same directory of the original file
```
os.system('tcmpr deSVD_temp.txt')
```
You could find a file name "deSVD_temp.txt.huffman" in the current directory
### After this step, you could calculate the compression ratio if you want
## Decode the data
```
os.system('tcmpr -d deSVD_temp.txt.huffman')
decoded_data = np.loadtxt(fname = "deSVD_temp.txt")
decoded_data = decoded_data[..., np.newaxis]
from sklearn.metrics import mean_squared_error
from math import sqrt
from scipy import signal
# original_signal = normalized_signal
# compressed_signal = decoded_data
def PRD_calculation(original_signal, compressed_signal):
PRD = sqrt(sum((original_signal-compressed_signal)**2)/(sum(original_signal**2)))
return PRD
PRD = PRD_calculation(resampled_signal, decoded_data)
print("The PRD is {}%".format(round(PRD*100,3)))
```
CR = 227/21 = 10.8
| github_jupyter |
# DC Resistivity Part 1
**Author:** [Lindsey Heagy](https://github.com/lheagy)
This example examines a model similar to that analyzed in [Kaufman (1990)](https://doi.org/10.1190/1.1442769). Here, we look at the DC electric field, current density, and charge distributions along the well near a source. We show the behaviour in the near and intermediate zones as defined in [Kaufman (1990)](https://doi.org/10.1190/1.1442769).
This notebook was used to produce Figure 5 in Heagy and Oldenburg (2018).
If you encounter problems when running this notebook, please [open an issue](https://github.com/simpeg-research/heagy_2018_emcyl/issues).
## Setup and Software environment
The requirements to run this example are in [requirements.txt](../requirements.txt). Uncomment the following cell if you need to install them.
```
# !pip install -r ../requirements.txt
# core python
import numpy as np
import scipy.sparse as sp
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from scipy.constants import mu_0
import ipywidgets
# finite volume
import discretize
from discretize import utils
# numerical simulation
from SimPEG.EM import TDEM
from SimPEG import Utils, Maps, Versions
# solver
from pymatsolver import Pardiso
# wrappers specific to solving casing problems
import casingSimulations as casingSim
%matplotlib inline
from matplotlib import rcParams
rcParams['font.size'] = 14
```
## Simulation Parameters
Here, we set a simulation directory where we can save the results and set the other main parameters for the example including the casing length and the conductivity of the background
```
simDir = 'DC_Kaufman' # create a simulation directory where results can be saved.
# casing parameters
casing_l = 1000 # 2km long casing
casing_t = 10e-3 # 10mm thick casing
casing_d = 100e-3 # 10cm diameter
sigma_back=1e-2 # 100 Ohm-background
sigma_casing=1e6 # conductivity of the casing
print(
'The casing is {}m long, and the '
'conductivity of the background is {} S/m'.format(
casing_l, sigma_back
)
)
```
We set up 2 models,
- `model` includes the conductive, steel cased well
- `wholespace` does not include a conductive well, it is a wholespace with conductivity 0.01S/m
```
# an object for convienently storing the model parameters
model = casingSim.model.CasingInWholespace(
directory = simDir,
sigma_casing = sigma_casing, # conductivity of the casing (S/m)
sigma_back = sigma_back, # conductivity of the background (S/m)
sigma_inside = sigma_back, # fluid inside the well has same conductivity as the background
casing_d = casing_d-casing_t, # 10c.m is outer casing diameter
casing_l = casing_l,
casing_t = casing_t,
src_a = np.r_[0., 0., -casing_l/2.], # put the A electrode just below the surface
src_b = np.r_[casing_l, 0., -casing_l/2.] # put the return electrode at a distance of 2 x length of well away
)
# Here we print the parameters being used to set up the simulation
model.serialize()
```
copy the model and set the conductivity equal to the background so we have a baseline response to compare to
```
# put the models in a dictionary for convienence
model_names = ['casing']
modelDict = dict(zip(model_names, [model]))
wholespace = model.copy()
wholespace.sigma_casing = wholespace.sigma_back
model_names += ['background']
modelDict['background'] = wholespace
```
## Mesh
Here we set up a 3D cylindrical mesh, discretizing in $x$, $\theta$ and $z$.
To discretize in x, we start by defining the finest region of the mesh, ensuring that we have 4 cells across the thickness of the casing. From there, we expand the cell sizes until we reach the second uniform cell size we want to model at (`csx2`). We then use a constant cell spacing of `csx2` until we have meshed out to the end of the domain in which we want to examine data (`domainx2`). Beyond that, we add padding cells to *"infinity"*
```
# parameters defining the core region of the mesh
# note that the finest re
csx2 = 100. # cell size in the x-direction in the second uniform region of the mesh (where we measure data)
csz = 0.1 # cell size in the z-direction
domainx2 = 100 # go out 500m from the well
# padding parameters
npadx, npadz = 8, 26 # number of padding cells
pfx2 = 1.4 # expansion factor for the padding to infinity in the x-direction
pfz = 1.4
# set up a mesh generator which will build a mesh based on the provided parameters
# and casing geometry
cylMeshGen = casingSim.CasingMeshGenerator(
directory=simDir, # directory where we can save things
modelParameters=model, # casing parameters
npadx=npadx, # number of padding cells in the x-direction
npadz=npadz, # number of padding cells in the z-direction
domain_x=domainx2, # extent of the second uniform region of the mesh
csx1=model.casing_t/4., # use at least 4 cells per across the thickness of the casing
csx2=csx2, # second core cell size
csz=csz, # cell size in the z-direction
pfx2=pfx2, # padding factor to "infinity"
pfz=pfz # padding factor to "infinity" for the z-direction
)
print("the mesh has {} cells".format(cylMeshGen.mesh.nC))
ax = cylMeshGen.mesh.plotGrid()
# uncomment the following to zoom in radially
# ax.set_xlim([0, 0.1])
```
## Put the physical properties on the mesh
Here, we organize models in a dictionary, fro there, we build the physical properties, which describes the conductivity over the entire simulation domain.
```
# Assign physical properties on the mesh
physpropsDict = {
name: casingSim.model.PhysicalProperties(cylMeshGen, mod)
for name, mod in modelDict.items()
}
```
### Plot the models
```
xlim = np.r_[-1, 1] # x-limits in meters
zlim = np.r_[-1.5*model.casing_l, 10.] # z-limits in meters. (z-positive up)
fig, ax = plt.subplots(1, len(model_names), figsize=(6*len(model_names), 5))
if len(model_names) == 1:
ax = [ax]
for a, title in zip(ax, model_names):
pp = physpropsDict[title]
pp.plot_sigma(
ax=a,
pcolorOpts={'norm':LogNorm()} # plot on a log-scale
)
a.set_title('{} \n\n $\sigma$ = {:1.2e}S/m'.format(title, pp.modelParameters.sigma_casing), fontsize=13)
a.set_xlim(xlim)
a.set_ylim(zlim)
plt.tight_layout()
# Plot the source location
symbols = ['rd', 'ws', 'k>', 'mo', 'c*', 'C3s']
fig, ax = plt.subplots(1, 1, figsize=(6, 7))
# cylMeshGen.mesh.plotGrid(ax=ax, slice='theta')
out = physpropsDict['casing'].plot_sigma(
ax=ax,
pcolorOpts={'norm':LogNorm()} # plot on a log-scale
)
cb = out[-1]
cb.set_label('conductivity (S/m)')
ax.set_title('')
# plot the source location
ax.plot(model.src_a[0], model.src_a[2], symbols[0])
ax.plot(model.src_b[0], model.src_b[2], symbols[0])
# set bounds
ax.set_xlim(0.5*np.r_[-1, 1]) #src_b[:, 0].max()])
ax.set_ylim([ -1.25*model.casing_l, 100])
xtext = -0.48
ax.annotate('casing {:1.1e} S/m'.format(model.sigma_casing), xy=(0.055, -50), color='w', fontsize=11)
ax.annotate('halfspace {:1.1e} S/m'.format(model.sigma_back), xy=(xtext, -1.2*model.casing_l), color='w', fontsize=11)
ax.annotate(
'source', xy=(0.055, model.src_a[2]), color='w', fontsize=11,
)
```
## set up a DC simulation
```
simDict = {} # store the simulations in a dictionary
for title in model_names:
simDict[title] = casingSim.run.SimulationDC(
modelParameters=modelDict[title], directory=simDir,
meshGenerator=cylMeshGen,
src_a=modelDict[title].src_a, src_b=modelDict[title].src_b
)
```
## run the DC simulation
```
%%time
fieldsDict = {}
for title in model_names:
print('--- Running {} ---'.format(title))
fieldsDict[title] = simDict[title].run(save=False)
print('\n')
```
## View Fields, Fluxes, and Charges
This is a widget for interrogating the results.
- `max_r`: maximum radial extent of the plot (m)
- `min_depth`: minimum depth (m)
- `max_depth`: maximum depth (m)
- `clim_min`: minimum colorbar limit. If `0`, then the colorbar limits are the plotting defaults
- `clim_max`: maximum colorbar limit. If `0`, then the colorbar limits are the plotting defaults
- `model_key`: model which we are viewing
- `view`: field or physical property that is plotted
- `prim_sec`: `primary` plots the background, `secondary` subtracts the `primary` response from the current value (note that if you select `background` and `secondary` the value will be zero and an error thrown
- `show_mesh`: if checked, the mesh will be plotted on the right hand half of the plot
- `casing_outline`: draws the outline of the casing
```
viewer = casingSim.FieldsViewer(
sim_dict=simDict, fields_dict=fieldsDict, model_keys=model_names, primary_key = "background"
)
viewer.widget_cross_section(
defaults={"min_depth": model.casing_l/2.-2, "max_depth":model.casing_l/2.+2},
fixed={"use_aspect":False}
)
```
## Figure 5 in the paper
```
fig, ax = plt.subplots(2, 2, figsize=(2*5, 2*6))
ax=ax.flatten()
max_r = 0.11
min_depth = model.casing_l/2.-2
max_depth = model.casing_l/2.+2
# model =
xlim = max_r * np.r_[-1, 1]
zlim = np.r_[-max_depth, -min_depth]
def plotme(a, view, clim, **kwargs):
out = viewer.plot_cross_section(
ax=a, model_key='casing', xlim=xlim, zlim=zlim, clim=clim,
view=view, casing_outline=True, **kwargs
)
a.set_xlabel('x (m)')
a.set_ylabel('z (m)')
cb = out[-1]
if view in ['charge', 'charge_density']:
cb.set_ticks(np.linspace(clim[0], clim[1], 5))
cb.formatter.set_powerlimits((0, 0))
cb.update_ticks()
else:
ticks = 10**np.hstack([np.arange(np.log10(clim[0]), np.log10(clim[1])), np.r_[np.log10(clim[1])]])[::2]
cb.set_ticks(ticks)
# out[-1].set_clim(clim)
return out
# total charge density
clim = 1e-6* np.r_[-1, 1]
out = plotme(ax[0], 'charge_density', clim)
out[-1].set_label('total charge density (C/m$^3$)')
ax[0].set_title('(a)')
# secondary charges
clim = 3e-9 * np.r_[-1, 1]
out = plotme(ax[1], 'charge_density', clim, cb_extend="both", prim_sec="secondary")
ax[1].set_title('(b)')
out[-1].set_label('secondary charge density (C/m$^3$)')
# electric field
clim = np.r_[1e-6, 1e5]
out = plotme(ax[2], 'e', clim)
ax[2].set_title('(c)')
out[-1].set_label('electric field (V/m)')
# # current density
clim = np.r_[1e-8, 1e3]
out = plotme(ax[3], 'j', clim)
ax[3].set_title('(d)')
out[-1].set_label('current density (A/m$^2$)')
[a.set_yticks(np.hstack([np.arange(-max_depth, -min_depth),np.r_[-min_depth]])) for a in ax]
plt.tight_layout()
fig.savefig('../figures/kaufman_zones', dpi=350, bbox_inches="tight")
fig.savefig('../arxiv-figures/kaufman_zones', dpi=150, bbox_inches="tight")
Versions()
```
| github_jupyter |
# Introduction to the JupyterLab and Jupyter Notebooks
This is a short introduction to two of the flagship tools created by [the Jupyter Community](https://jupyter.org).
> **Note**: This interface is provided by the [JupyterLite project](https://jupyterlite.readthedocs.io/en/latest/), which embeds an entire JupyterLab interface, with many popular packages for scientific computing, in your browser. There may be some minor differences in behavior between JupyterLite and the JupyterLab you install locally.
## JupyterLab 🧪
**JupyterLab** is a next-generation web-based user interface for Project Jupyter. It enables you to work with documents and activities such as Jupyter notebooks, text editors, terminals, and custom components in a flexible, integrated, and extensible manner. It is the interface that you're looking at right now.
**For an overview of the JupyterLab interface**, see the **JupyterLab Welcome Tour** on this page, by going to `Help -> Welcome Tour` and following the prompts.
> **See Also**: For a more in-depth tour of JupyterLab with a full environment that runs in the cloud, see [the JupyterLab introduction on Binder](https://mybinder.org/v2/gh/jupyterlab/jupyterlab-demo/HEAD?urlpath=lab/tree/demo).
## Jupyter Notebooks 📓
**Jupyter Notebooks** are a community standard for communicating and performing interactive computing. They are a document that blends computations, outputs, explanatory text, mathematics, images, and rich media representations of objects.
JupyterLab is one interface used to create and interact with Jupyter Notebooks.
**For an overview of Jupyter Notebooks**, see the **JupyterLab Welcome Tour** on this page, by going to `Help -> Notebook Tour` and following the prompts.
> **See Also**: For a more in-depth tour of Jupyter Notebooks and the Classic Jupyter Notebook interface, see [the Jupyter Notebook IPython tutorial on Binder](https://mybinder.org/v2/gh/ipython/ipython-in-depth/HEAD?urlpath=tree/binder/Index.ipynb).
## An example: visualizing data in the notebook ✨
Below is an example of a code cell. We'll visualize some simple data using two popular packages in Python. We'll use [NumPy](https://numpy.org/) to create some random data, and [Matplotlib](https://matplotlib.org) to visualize it.
Note how the code and the results of running the code are bundled together.
```
from matplotlib import pyplot as plt
import numpy as np
# Generate 100 random data points along 3 dimensions
x, y, scale = np.random.randn(3, 100)
fig, ax = plt.subplots()
# Map each onto a scatterplot we'll create with Matplotlib
ax.scatter(x=x, y=y, c=scale, s=np.abs(scale)*500)
ax.set(title="Some random data, created with JupyterLab!")
plt.show()
```
## Next steps 🏃
This is just a short introduction to JupyterLab and Jupyter Notebooks. This demonstration contains a lot more that you can play around with. Here are some pointers to help you take the next step. Each of the items below corresponds to a file or folder in the **file browser to the left**.
- [python.ipynb](python.ipynb) is a Jupyter Notebook that shows off some basic Python functionality, including more visualizations, data structures, and scientific computing libraries.
- `pyolite/` is a folder that contains several Jupyter Notebooks that highlight many more things that you can do in JupyterLab / JupyterLite. Explore them for inspiration about what you'd like to do next.
- [p5.ipynb](p5.ipynb) is a Jupyter Notebook that shows off computing with [the p5 platform](https://github.com/processing/p5.js/), which allows you to build visual experiences with Javascript.
- [javascript.ipynb](javascript.ipynb) is a Jupyter Notebook that shows off how you can run Javascript code within an ipynb file.
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
```
Training a Classifier
=====================
About data:
For vision, a package called ``torchvision``,
that has data loaders for common datasets such as
Imagenet, CIFAR10, MNIST, etc.
For this experiment, I use the CIFAR10 dataset.
It has the classes: ‘airplane’, ‘automobile’, ‘bird’, ‘cat’, ‘deer’,
‘dog’, ‘frog’, ‘horse’, ‘ship’, ‘truck’.
The images in CIFAR-10 are of
size 3x32x32, i.e. 3-channel color images of 32x32 pixels in size.
Using ``torchvision``, it’s extremely easy to load CIFAR10.
```
import torch
import torchvision
import torchvision.transforms as transforms
```
The output of torchvision datasets are PILImage images of range [0, 1].
I transform them to Tensors of normalized range [-1, 1].
```
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
```
Lets see some of the training dataset images.
```
import matplotlib.pyplot as plt
import numpy as np
# functions to show an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
print(' '.join('%5s' % classes[labels[j]] for j in range(4)))
```
2. Define a Convolutional Neural Network
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Create a 2 convolution layers (conv) and 3 Fully connected layers (Fc)
```
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5) # 3 input image channel, 6 output channels, 5x5 square convolution kernel
self.pool = nn.MaxPool2d(2, 2) # Max pooling over a (2, 2) window
self.conv2 = nn.Conv2d(6, 16, 5)
#3 Fully connected layers (Fc)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1,16*5*5) #(len(x[0]),len(x))#(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
```
3. Define a Loss function and optimizer
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Classification Cross-Entropy loss and SGD with a Lerarning rate = 0.001.
```
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
```
4. Train the network
^^^^^^^^^^^^^^^^^^^^
I simply have to loop over our training dataset (batches),
and feed the inputs to the network and optimize the weights.
```
for epoch in range(50): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
```
5. Test the network on the test data
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
After 50 passes over the training dataset. I need to check if the network
is learning.
I will check this by predicting the class label that the neural network
outputs, and checking it against the actual labels of the test set.
Okay, first step. Let us display an image from the test set to get familiar.
The outputs are energies (probabilities) for the 10 classes.
The higher the energy for a class, the more the network
thinks that the image is of the particular class.
The results seem pretty good.
Let us look at how the network performs on the whole dataset.
```
correct = 0
total = 0
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(4):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
for i in range(10):
print('Accuracy of %5s : %2d %%' % (
classes[i], 100 * class_correct[i] / class_total[i]))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/perfectpanda-works/machine-learning/blob/master/PyTorchTutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#PyTorch Tutorial
PyTorchの基本的な機能のTensorを利用する
```
from __future__ import print_function
import torch
```
初期化なしでテンソルを作成するemptyメソッド。
2階テンソル(行列)の作成。5行3列
```
x = torch.empty(5, 3)
print(x)
```
ランダムに初期化した値を入れたテンソルの作成。randメソッド。
```
x = torch.rand(5, 3)
print(x)
```
0で初期化したテンソルを作成するzerosメソッド。
dtypeに0の型を指定できる。
```
x = torch.zeros(5, 3, dtype=torch.long)
print(x)
```
自分で初期値を指定してテンソルを作成するtensorメソッド。
1階のテンソル(ベクトル)を作成。
```
x = torch.tensor([5.5, 3])
print(x)
```
2階のテンソル(行列)を作成。
行の間にカンマを忘れがち・・・
```
x = torch.tensor([[5.5, 3],
[6.5, 4]])
print(x)
```
すでに作成したテンソルを再度定義し直す事もできます。
その時、指定していないパラメータは以前のものを引き継ぎます。
new_onesは1で埋めるメソッド。
xを5×3の行列と定義して、1で埋めます。そして、それぞれのデータはdouble型(float64型)とx自身のメソッドで定義し直すことができます。
```
x = x.new_ones(5, 3, dtype=torch.double)
print(x)
```
テンソルをランダムな値で埋めて、データの型をfloat型に変更します。
テンソルのサイズは指定されていないので、xの5×3がそのまま引き継がれます。
```
x = torch.randn_like(x, dtype=torch.float)
print(x)
```
numpyでいう「.shape」は、sizeメソッドになります。
```
print(x.size())
```
#テンソルの計算
慣れないベクトル(テンソル)の計算・・・
##足し算と引き算
###スカラー + ベクトル
```
x = 1
y = torch.tensor([4,5])
print(x)
print(y)
```
それぞれの要素にスカラーが足し合わされる。
```
print(x + y)
```
同じように、addメソッドでも足すことができます。
```
print(torch.add(x, y))
```
「torch.add」では次のように出力を指定してテンソル同士を足し合わせる事もできます。
```
result = torch.empty(1,2)
torch.add(x, y, out=result)
print(result)
```
同様に引き算です。
```
print(x - y)
```
###ベクトル+ベクトル
ベクトル同士の足し算、引き算は、同じサイズ同士である必要がある。
```
x2 = torch.tensor([1,2,3])
y2 = torch.tensor([4,5,6])
print(x2)
print(y2)
print(x2 + y2)
print(x2 - y2)
```
サイズが違う場合エラー
```
x2 = torch.tensor([1,2])
y2 = torch.tensor([4,5,6])
print(x2)
print(y2)
print(x2 + y2)
```
###行列の足し算引き算
スカラー、行列
```
x3 = 1
y3 = torch.tensor([[1,2,3],
[4,5,6]])
print(x3)
print(y3)
print(x3 + y3)
```
ベクトル、行列
行、または列が同じ
```
x4 = torch.tensor([1,2,3])
y4 = torch.tensor([[1,2,3],
[4,5,6]])
print(x4 + y4)
x5 = torch.tensor([[1],
[2]])
y5 = torch.tensor([[1,2,3],
[4,5,6]])
print(x5 + y5)
```
###行列と行列
```
x6 = torch.tensor([[1,2,3],
[3,2,1]])
y6 = torch.tensor([[1,2,3],
[3,2,1]])
print(x6 + y6)
```
上書きして足し合わせる場合、次のような表記方法も可能です。
add_メソッド
```
y6.add_(x6)
print(y6)
```
y6に、x6とy6を足した結果が代入されました。
numpyのスライス表記のようにテンソルの要素を取り出すことができます。
```
print(y6[:,0])
```
y6から、1列目の全ての行を取り出しました。
##テンソルのサイズ変更
```
x = torch.randn(4, 4)
y = x.view(16)
z = x.view(-1, 8) # the size -1 is inferred from other dimensions
print(x.size(), y.size(), z.size())
```
zに指定した-1は、他の要素から推定されるという意味です。例えば、列が8になると、もともとxには16個のデータ(4×4)しかないので、2行になるので、 -1は自動的に2を入れることになります。
次のように、要素数に矛盾が生じる場合、エラーになります。
```
y = x.view(15)
```
1要素のみのテンソルをitemメソッドでPythonの数値として取り出すことができるそうです。
```
x = torch.randn(1)
print(x)
print(x.item())
```
使い所が不明だったので、少し調べたところ、平均を取り出す時に使いそうです。
ちなみに、meanを利用することで、テンソルの平均値が取り出せるそうです。
```
x = torch.tensor([5.0, 4.0, 3.0])
average = x.mean()
print(average)
print(average.item())
```
他にも、たくさんの構文があるので、こちらで確認してね、というチュートリアルになっています・・・
#NumPy←→PyTorch
###PyTorchのtensor型をnumpy配列に変換
```
a = torch.ones(5)
print(a)
b = a.numpy()
print(b)
```
tensorからnumpyに変換したものは、連動しているみたいです。
```
a.add_(1)
print(a)
print(b)
```
###Numpy配列をPyTorchのtensor型に変換
```
import numpy as np
a = np.ones(5)
b = torch.from_numpy(a)
np.add(a, 1, out=a)
print(a)
print(b)
```
```
```
CUDAというNVIDIAのGPU並列処理する仕組みを利用して、CUDAで処理できるようなメソッドが「.to」とのことです。CUDA環境ではないので、実行できませんでしたが、簡単にGPUでテンソルを処理するのか、CPUで処理するのかが記載できます。
```
if torch.cuda.is_available():
device = torch.device("cuda") # a CUDA device object
y = torch.ones_like(x, device=device) # directly create a tensor on GPU
x = x.to(device) # or just use strings ``.to("cuda")``
z = x + y
print(z)
print(z.to("cpu", torch.double)) # ``.to`` can also change dtype together!
```
| github_jupyter |
# NumPy
In this lesson we will learn the basics of numerical analysis using the NumPy package.
<div align="left">
<a href="https://github.com/madewithml/lessons/blob/master/notebooks/01_Foundations/03_NumPy.ipynb" role="button"><img class="notebook-badge-image" src="https://img.shields.io/static/v1?label=&message=View%20On%20GitHub&color=586069&logo=github&labelColor=2f363d"></a>
<a href="https://colab.research.google.com/github/madewithml/lessons/blob/master/notebooks/01_Foundations/03_NumPy.ipynb"><img class="notebook-badge-image" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
</div>
# Set up
```
import numpy as np
# Set seed for reproducibility
np.random.seed(seed=1234)
```
# Basics
Let's take a took at how to create tensors with NumPy.
* **Tensor**: collection of values
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/tensors.png" width="650">
</div>
```
# Scalar
x = np.array(6) # scalar
print ("x: ", x)
# Number of dimensions
print ("x ndim: ", x.ndim)
# Dimensions
print ("x shape:", x.shape)
# Size of elements
print ("x size: ", x.size)
# Data type
print ("x dtype: ", x.dtype)
# Vector
x = np.array([1.3 , 2.2 , 1.7])
print ("x: ", x)
print ("x ndim: ", x.ndim)
print ("x shape:", x.shape)
print ("x size: ", x.size)
print ("x dtype: ", x.dtype) # notice the float datatype
# Matrix
x = np.array([[1,2], [3,4]])
print ("x:\n", x)
print ("x ndim: ", x.ndim)
print ("x shape:", x.shape)
print ("x size: ", x.size)
print ("x dtype: ", x.dtype)
# 3-D Tensor
x = np.array([[[1,2],[3,4]],[[5,6],[7,8]]])
print ("x:\n", x)
print ("x ndim: ", x.ndim)
print ("x shape:", x.shape)
print ("x size: ", x.size)
print ("x dtype: ", x.dtype)
```
NumPy also comes with several functions that allow us to create tensors quickly.
```
# Functions
print ("np.zeros((2,2)):\n", np.zeros((2,2)))
print ("np.ones((2,2)):\n", np.ones((2,2)))
print ("np.eye((2)):\n", np.eye((2))) # identity matrix
print ("np.random.random((2,2)):\n", np.random.random((2,2)))
```
# Indexing
Keep in mind that when indexing the row and column, indices start at 0. And like indexing with lists, we can use negative indices as well (where -1 is the last item).
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/indexing.png" width="300">
</div>
```
# Indexing
x = np.array([1, 2, 3])
print ("x: ", x)
print ("x[0]: ", x[0])
x[0] = 0
print ("x: ", x)
# Slicing
x = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
print (x)
print ("x column 1: ", x[:, 1])
print ("x row 0: ", x[0, :])
print ("x rows 0,1 & cols 1,2: \n", x[0:2, 1:3])
# Integer array indexing
print (x)
rows_to_get = np.array([0, 1, 2])
print ("rows_to_get: ", rows_to_get)
cols_to_get = np.array([0, 2, 1])
print ("cols_to_get: ", cols_to_get)
# Combine sequences above to get values to get
print ("indexed values: ", x[rows_to_get, cols_to_get]) # (0, 0), (1, 2), (2, 1)
# Boolean array indexing
x = np.array([[1, 2], [3, 4], [5, 6]])
print ("x:\n", x)
print ("x > 2:\n", x > 2)
print ("x[x > 2]:\n", x[x > 2])
```
# Arithmetic
```
# Basic math
x = np.array([[1,2], [3,4]], dtype=np.float64)
y = np.array([[1,2], [3,4]], dtype=np.float64)
print ("x + y:\n", np.add(x, y)) # or x + y
print ("x - y:\n", np.subtract(x, y)) # or x - y
print ("x * y:\n", np.multiply(x, y)) # or x * y
```
### Dot product
One of the most common NumPy operations we’ll use in machine learning is matrix multiplication using the dot product. We take the rows of our first matrix (2) and the columns of our second matrix (2) to determine the dot product, giving us an output of `[2 X 2]`. The only requirement is that the inside dimensions match, in this case the first matrix has 3 columns and the second matrix has 3 rows.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/dot.gif" width="450">
</div>
```
# Dot product
a = np.array([[1,2,3], [4,5,6]], dtype=np.float64) # we can specify dtype
b = np.array([[7,8], [9,10], [11, 12]], dtype=np.float64)
c = a.dot(b)
print (f"{a.shape} · {b.shape} = {c.shape}")
print (c)
```
### Axis operations
We can also do operations across a specific axis.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/axis.gif" width="450">
</div>
```
# Sum across a dimension
x = np.array([[1,2],[3,4]])
print (x)
print ("sum all: ", np.sum(x)) # adds all elements
print ("sum axis=0: ", np.sum(x, axis=0)) # sum across rows
print ("sum axis=1: ", np.sum(x, axis=1)) # sum across columns
# Min/max
x = np.array([[1,2,3], [4,5,6]])
print ("min: ", x.min())
print ("max: ", x.max())
print ("min axis=0: ", x.min(axis=0))
print ("min axis=1: ", x.min(axis=1))
```
### Broadcasting
Here, we’re adding a vector with a scalar. Their dimensions aren’t compatible as is but how does NumPy still gives us the right result? This is where broadcasting comes in. The scalar is *broadcast* across the vector so that they have compatible shapes.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/broadcasting.png" width="300">
</div>
```
# Broadcasting
x = np.array([1,2]) # vector
y = np.array(3) # scalar
z = x + y
print ("z:\n", z)
```
# Advanced
### Transposing
We often need to change the dimensions of our tensors for operations like the dot product. If we need to switch two dimensions, we can transpose
the tensor.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/transpose.png" width="400">
</div>
```
# Transposing
x = np.array([[1,2,3], [4,5,6]])
print ("x:\n", x)
print ("x.shape: ", x.shape)
y = np.transpose(x, (1,0)) # flip dimensions at index 0 and 1
print ("y:\n", y)
print ("y.shape: ", y.shape)
```
### Reshaping
Sometimes, we'll need to alter the dimensions of the matrix. Reshaping allows us to transform a tensor into different permissible shapes -- our reshaped tensor has the same amount of values in the tensor. (1X6 = 2X3). We can also use `-1` on a dimension and NumPy will infer the dimension based on our input tensor.
The way reshape works is by looking at each dimension of the new tensor and separating our original tensor into that many units. So here the dimension at index 0 of the new tensor is 2 so we divide our original tensor into 2 units, and each of those has 3 values.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/reshape.png" width="450">
</div>
```
# Reshaping
x = np.array([[1,2,3,4,5,6]])
print (x)
print ("x.shape: ", x.shape)
y = np.reshape(x, (2, 3))
print ("y: \n", y)
print ("y.shape: ", y.shape)
z = np.reshape(x, (2, -1))
print ("z: \n", z)
print ("z.shape: ", z.shape)
```
### Unintended reshaping
Though reshaping is very convenient to manipulate tensors, we must be careful of their pitfalls as well. Let's look at the example below. Suppose we have `x`, which has the shape `[2 X 3 X 4]`.
```
[[[ 1 1 1 1]
[ 2 2 2 2]
[ 3 3 3 3]]
[[10 10 10 10]
[20 20 20 20]
[30 30 30 30]]]
```
We want to reshape x so that it has shape `[3 X 8]` which we'll get by moving the dimension at index 0 to become the dimension at index 1 and then combining the last two dimensions. But when we do this, we want our output
to look like:
✅
```
[[ 1 1 1 1 10 10 10 10]
[ 2 2 2 2 20 20 20 20]
[ 3 3 3 3 30 30 30 30]]
```
and not like:
❌
```
[[ 1 1 1 1 2 2 2 2]
[ 3 3 3 3 10 10 10 10]
[20 20 20 20 30 30 30 30]]
```
even though they both have the same shape `[3X8]`.
```
x = np.array([[[1, 1, 1, 1], [2, 2, 2, 2], [3, 3, 3, 3]],
[[10, 10, 10, 10], [20, 20, 20, 20], [30, 30, 30, 30]]])
print ("x:\n", x)
print ("x.shape: ", x.shape)
```
When we naively do a reshape, we get the right shape but the values are not what we're looking for.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/reshape_wrong.png" width="600">
</div>
```
# Unintended reshaping
z_incorrect = np.reshape(x, (x.shape[1], -1))
print ("z_incorrect:\n", z_incorrect)
print ("z_incorrect.shape: ", z_incorrect.shape)
```
Instead, if we transpose the tensor and then do a reshape, we get our desired tensor. Transpose allows us to put our two vectors that we want to combine together and then we use reshape to join them together.
Always create a dummy example like this when you’re unsure about reshaping. Blindly going by the tensor shape can lead to lots of issues downstream.
<div align="left">
<img src="https://raw.githubusercontent.com/madewithml/images/master/01_Foundations/03_NumPy/reshape_right.png" width="600">
</div>
```
# Intended reshaping
y = np.transpose(x, (1,0,2))
print ("y:\n", y)
print ("y.shape: ", y.shape)
z_correct = np.reshape(y, (y.shape[0], -1))
print ("z_correct:\n", z_correct)
print ("z_correct.shape: ", z_correct.shape)
```
### Adding/removing dimensions
We can also easily add and remove dimensions to our tensors and we'll want to do this to make tensors compatible for certain operations.
```
# Adding dimensions
x = np.array([[1,2,3],[4,5,6]])
print ("x:\n", x)
print ("x.shape: ", x.shape)
y = np.expand_dims(x, 1) # expand dim 1
print ("y: \n", y)
print ("y.shape: ", y.shape) # notice extra set of brackets are added
# Removing dimensions
x = np.array([[[1,2,3]],[[4,5,6]]])
print ("x:\n", x)
print ("x.shape: ", x.shape)
y = np.squeeze(x, 1) # squeeze dim 1
print ("y: \n", y)
print ("y.shape: ", y.shape) # notice extra set of brackets are gone
```
# Additional resources
* **NumPy reference manual**: We don't have to memorize anything here and we will be taking a closer look at NumPy in the later lessons. If you want to learn more checkout the [NumPy reference manual](https://docs.scipy.org/doc/numpy-1.15.1/reference/).
---
Share and discover ML projects at <a href="https://madewithml.com/">Made With ML</a>.
<div align="left">
<a class="ai-header-badge" target="_blank" href="https://github.com/madewithml/lessons"><img src="https://img.shields.io/github/stars/madewithml/lessons.svg?style=social&label=Star"></a>
<a class="ai-header-badge" target="_blank" href="https://www.linkedin.com/company/madewithml"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
<a class="ai-header-badge" target="_blank" href="https://twitter.com/madewithml"><img src="https://img.shields.io/twitter/follow/madewithml.svg?label=Follow&style=social"></a>
</div>
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#### Version Check
Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
```
import plotly
plotly.__version__
```
### Set X and Y Coordinates
To set the `x` and `y` coordinates use `x` and `y` attributes. If `x` coordindate values are ommitted a cheater plot will be created. The plot below has a `y` array specified but requires `a` and `b` parameter values before an axis may be plotted.
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10]
)
data = [trace1]
fig = go.Figure(data = data)
url = py.plot(fig, filename = "carpet/basic")
```
### Add Parameter Values
To save parameter values use the `a` and `b` attributes.
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4, 4, 4.5, 4.5, 4.5, 5, 5, 5, 6, 6, 6],
b = [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3],
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10]
)
data = [trace1]
fig = go.Figure(data = data)
py.iplot(fig, filename = "carpet/add-parameters")
```
### Add A and B axis
Use `aaxis` or `baxis` list to make changes to the axes. For a more detailed list of attributes refer to [R reference](https://plotly.com/r/reference/#carpet-aaxis).
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4, 4, 4.5, 4.5, 4.5, 5, 5, 5, 6, 6, 6],
b = [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3],
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10],
aaxis = dict(
tickprefix = 'a = ',
ticksuffix = 'm',
smoothing = 1,
minorgridcount = 9,
),
baxis = dict(
tickprefix = 'b = ',
ticksuffix = 'pa',
smoothing = 1,
minorgridcount = 9,
)
)
data = [trace1]
fig = go.Figure(data = data)
py.iplot(fig, filename = "carpet/add-axes")
```
### Alternate input format
The data arrays `x`, `y` may either be specified as one-dimensional arrays of data or as arrays of arrays. If one-dimensional, then `x`, `y`, `a`, and `b` should all be the same length. If `x` and `y` are arrays of arrays, then the length of `a` should match the inner dimension and the length of `b` the outer dimension. The plot below represents the same plot as those above.
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4.5, 5, 6],
b = [1, 2, 3],
y = [[2, 3, 5.5, 8],
[3.5, 4.5, 6.5, 8.5],
[4, 5, 7.5, 10]]
)
data = [trace1]
fig = go.Figure(data = data)
py.iplot(fig, filename = "carpet/input-format")
```
### Cheater plot layout
The layout of cheater plots is not unique and depends upon the `cheaterslope` and axis `cheatertype` parameters. If `x` is not specified, each row of the `x` array is constructed based on the the formula `a + cheaterslope * b`, where `a` and `b` are either the value or the integer index of `a` and `b` respectively, depending on the corresponding axis `cheatertype`. Although the layout of the axis below is different than the plots above, it represents the same data as the axes above.
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4.5, 5, 6],
b = [1, 2, 3],
y = [[2, 3, 5.5, 8],
[3.5, 4.5, 6.5, 8.5],
[4, 5, 7.5, 10]],
cheaterslope = -5,
aaxis = dict(cheatertype = 'index'),
baxis = dict(cheatertype = 'value')
)
data = [trace1]
fig = go.Figure(data = data)
py.iplot(fig, filename = "carpet/cheater-layout")
```
### Style A and B axis
```
import plotly.graph_objs as go
import plotly.plotly as py
trace1 = go.Carpet(
a = [4, 4, 4, 4.5, 4.5, 4.5, 5, 5, 5, 6, 6, 6],
b = [1, 2, 3, 1, 2, 3, 1, 2, 3, 1, 2, 3],
y = [2, 3.5, 4, 3, 4.5, 5, 5.5, 6.5, 7.5, 8, 8.5, 10],
aaxis = dict(
tickprefix = 'a = ',
ticksuffix = 'm',
smoothing = 1,
minorgridcount = 9,
minorgridwidth = 0.6,
minorgridcolor = 'white',
gridcolor = 'white',
color = 'white'
),
baxis = dict(
ticksuffix = 'Pa',
smoothing = 1,
minorgridcount = 9,
minorgridwidth = 0.6,
gridcolor = 'white',
minorgridcolor = 'white',
color = 'white'
)
)
data = [trace1]
layout = go.Layout(
plot_bgcolor = 'black',
paper_bgcolor = 'black',
xaxis = dict(
showgrid = False,
showticklabels = False
),
yaxis = dict(
showgrid = False,
showticklabels = False
)
)
fig = go.Figure(data = data, layout = layout)
py.iplot(fig, filename = "carpet/styled")
```
### Add Points and Contours
To add points and lines see [Carpet Scatter Plots](https://plotly.com/python/carpet-scatter) or to add contours see [Carpet Contour Plots](https://plotly.com/python/carpet-contour)
### Reference
See https://plotly.com/python/reference/#carpet for more information and chart attribute options!
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'carpet_plot.ipynb', 'python/carpet-plot/', 'Carpet Plots',
'How to make carpet plots in Python with Plotly.',
title = 'Carpet Plots | Plotly',
has_thumbnail='true', thumbnail='thumbnail/carpet.jpg',
language='python',
# page_type='example_index', // note this is only if you want the tutorial to appear on the main page: plot.ly/python
display_as='scientific', order=26,
ipynb= '~notebook_demo/144')
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.